code stringlengths 2.5k 150k | kind stringclasses 1 value |
|---|---|
### Simple housing version
* State: $[w, n, M, e, \hat{S}, z]$, where $z$ is the stock trading experience, which took value of 0 and 1. And $\hat{S}$ now contains 27 states.
* Action: $[c, b, k, q]$ where $q$ only takes 2 value: $1$ or $\frac{1}{2}$
```
from scipy.interpolate import interpn
from multiprocessing import Pool
from functools import partial
from constant import *
import warnings
warnings.filterwarnings("ignore")
#Define the utility function
def u(c):
return (np.float_power(c, 1-gamma) - 1)/(1 - gamma)
#Define the bequeath function, which is a function of wealth
def uB(tb):
return B*u(tb)
#Calcualte HE
def calHE(x):
# the input x is a numpy array
# w, n, M, e, s, z = x
HE = H*pt - x[:,2]
return HE
#Calculate TB
def calTB(x):
# the input x as a numpy array
# w, n, M, e, s, z = x
TB = x[:,0] + x[:,1] + calHE(x)
return TB
#The reward function
def R(x, a):
'''
Input:
state x: w, n, M, e, s, z
action a: c, b, k, q = a which is a np array
Output:
reward value: the length of return should be equal to the length of a
'''
w, n, M, e, s, z = x
reward = np.zeros(a.shape[0])
# actions with not renting out
nrent_index = (a[:,3]==1)
# actions with renting out
rent_index = (a[:,3]!=1)
# housing consumption not renting out
nrent_Vh = (1+kappa)*H
# housing consumption renting out
rent_Vh = (1-kappa)*(H/2)
# combined consumption with housing consumption
nrent_C = np.float_power(a[nrent_index][:,0], alpha) * np.float_power(nrent_Vh, 1-alpha)
rent_C = np.float_power(a[rent_index][:,0], alpha) * np.float_power(rent_Vh, 1-alpha)
reward[nrent_index] = u(nrent_C)
reward[rent_index] = u(rent_C)
return reward
def transition(x, a, t):
'''
Input: state and action and time, where action is an array
Output: possible future states and corresponding probability
'''
w, n, M, e, s, z = x
s = int(s)
e = int(e)
nX = len(x)
aSize = len(a)
# mortgage payment
m = M/D[T_max-t]
M_next = M*(1+rh) - m
# actions
b = a[:,1]
k = a[:,2]
q = a[:,3]
# transition of z
z_next = np.ones(aSize)
if z == 0:
z_next[k==0] = 0
# we want the output format to be array of all possible future states and corresponding
# probability. x = [w_next, n_next, M_next, e_next, s_next, z_next]
# create the empty numpy array to collect future states and probability
if t >= T_R:
future_states = np.zeros((aSize*nS,nX))
n_next = gn(t, n, x, (r_k+r_b)/2)
future_states[:,0] = np.repeat(b*(1+r_b[s]), nS) + np.repeat(k, nS)*(1+np.tile(r_k, aSize))
future_states[:,1] = np.tile(n_next,aSize)
future_states[:,2] = M_next
future_states[:,3] = 0
future_states[:,4] = np.tile(range(nS),aSize)
future_states[:,5] = np.repeat(z_next,nS)
future_probs = np.tile(Ps[s],aSize)
else:
future_states = np.zeros((2*aSize*nS,nX))
n_next = gn(t, n, x, (r_k+r_b)/2)
future_states[:,0] = np.repeat(b*(1+r_b[s]), 2*nS) + np.repeat(k, 2*nS)*(1+np.tile(r_k, 2*aSize))
future_states[:,1] = np.tile(n_next,2*aSize)
future_states[:,2] = M_next
future_states[:,3] = np.tile(np.repeat([0,1],nS), aSize)
future_states[:,4] = np.tile(range(nS),2*aSize)
future_states[:,5] = np.repeat(z_next,2*nS)
# employed right now:
if e == 1:
future_probs = np.tile(np.append(Ps[s]*Pe[s,e], Ps[s]*(1-Pe[s,e])),aSize)
else:
future_probs = np.tile(np.append(Ps[s]*(1-Pe[s,e]), Ps[s]*Pe[s,e]),aSize)
return future_states, future_probs
# Use to approximate the discrete values in V
class Approxy(object):
def __init__(self, points, Vgrid):
self.V = Vgrid
self.p = points
def predict(self, xx):
pvalues = np.zeros(xx.shape[0])
for e in [0,1]:
for s in range(nS):
for z in [0,1]:
index = (xx[:,3] == e) & (xx[:,4] == s) & (xx[:,5] == z)
pvalues[index]=interpn(self.p, self.V[:,:,:,e,s,z], xx[index][:,:3],
bounds_error = False, fill_value = None)
return pvalues
# used to calculate dot product
def dotProduct(p_next, uBTB, t):
if t >= T_R:
return (p_next*uBTB).reshape((len(p_next)//(nS),(nS))).sum(axis = 1)
else:
return (p_next*uBTB).reshape((len(p_next)//(2*nS),(2*nS))).sum(axis = 1)
# Value function is a function of state and time t < T
def V(x, t, NN):
w, n, M, e, s, z = x
yat = yAT(t,x)
m = M/D[T_max - t]
# If the agent can not pay for the ortgage
if yat + w < m:
return [0, [0,0,0,0,0]]
# The agent can pay for the mortgage
if t == T_max-1:
# The objective functions of terminal state
def obj(actions):
# Not renting out case
# a = [c, b, k, q]
x_next, p_next = transition(x, actions, t)
uBTB = uB(calTB(x_next)) # conditional on being dead in the future
return R(x, actions) + beta * dotProduct(uBTB, p_next, t)
else:
def obj(actions):
# Renting out case
# a = [c, b, k, q]
x_next, p_next = transition(x, actions, t)
V_tilda = NN.predict(x_next) # V_{t+1} conditional on being alive, approximation here
uBTB = uB(calTB(x_next)) # conditional on being dead in the future
return R(x, actions) + beta * (Pa[t] * dotProduct(V_tilda, p_next, t) + (1 - Pa[t]) * dotProduct(uBTB, p_next, t))
def obj_solver(obj):
# Constrain: yat + w - m = c + b + kk
actions = []
budget1 = yat + w - m
for cp in np.linspace(0.001,0.999,11):
c = budget1 * cp
budget2 = budget1 * (1-cp)
#.....................stock participation cost...............
for kp in np.linspace(0,1,11):
# If z == 1 pay for matainance cost Km = 0.5
if z == 1:
# kk is stock allocation
kk = budget2 * kp
if kk > Km:
k = kk - Km
b = budget2 * (1-kp)
else:
k = 0
b = budget2
# If z == 0 and k > 0 payfor participation fee Kc = 5
else:
kk = budget2 * kp
if kk > Kc:
k = kk - Kc
b = budget2 * (1-kp)
else:
k = 0
b = budget2
#..............................................................
# q = 1 not renting in this case
actions.append([c,b,k,1])
# Constrain: yat + w - m + (1-q)*H*pr = c + b + kk
for q in [1,0.5]:
budget1 = yat + w - m + (1-q)*H*pr
for cp in np.linspace(0.001,0.999,11):
c = budget1*cp
budget2 = budget1 * (1-cp)
#.....................stock participation cost...............
for kp in np.linspace(0,1,11):
# If z == 1 pay for matainance cost Km = 0.5
if z == 1:
# kk is stock allocation
kk = budget2 * kp
if kk > Km:
k = kk - Km
b = budget2 * (1-kp)
else:
k = 0
b = budget2
# If z == 0 and k > 0 payfor participation fee Kc = 5
else:
kk = budget2 * kp
if kk > Kc:
k = kk - Kc
b = budget2 * (1-kp)
else:
k = 0
b = budget2
#..............................................................
# i = 0, no housing improvement when renting out
actions.append([c,b,k,q])
actions = np.array(actions)
values = obj(actions)
fun = np.max(values)
ma = actions[np.argmax(values)]
return fun, ma
fun, action = obj_solver(obj)
return np.array([fun, action])
# wealth discretization
ws = np.array([10,25,50,75,100,125,150,175,200,250,500,750,1000,1500,3000])
w_grid_size = len(ws)
# 401k amount discretization
ns = np.array([1, 5, 10, 15, 25, 50, 100, 150, 400, 1000])
n_grid_size = len(ns)
# Mortgage amount
Ms = np.array([0.01*H,0.05*H,0.1*H,0.2*H,0.3*H,0.4*H,0.5*H,0.8*H]) * pt
M_grid_size = len(Ms)
points = (ws,ns,Ms)
# dimentions of the state
dim = (w_grid_size, n_grid_size,M_grid_size,2,nS,2)
dimSize = len(dim)
xgrid = np.array([[w, n, M, e, s, z]
for w in ws
for n in ns
for M in Ms
for e in [0,1]
for s in range(nS)
for z in [0,1]
]).reshape(dim + (dimSize,))
# reshape the state grid into a single line of states to facilitate multiprocessing
xs = xgrid.reshape((np.prod(dim),dimSize))
Vgrid = np.zeros(dim + (T_max,))
cgrid = np.zeros(dim + (T_max,))
bgrid = np.zeros(dim + (T_max,))
kgrid = np.zeros(dim + (T_max,))
qgrid = np.zeros(dim + (T_max,))
print("The size of the grid: ", dim + (T_max,))
%%time
# value iteration part, create multiprocesses 32
pool = Pool()
for t in range(T_max-1,T_max-3, -1):
print(t)
if t == T_max - 1:
f = partial(V, t = t, NN = None)
results = np.array(pool.map(f, xs))
else:
approx = Approxy(points,Vgrid[:,:,:,:,:,:,t+1])
f = partial(V, t = t, NN = approx)
results = np.array(pool.map(f, xs))
Vgrid[:,:,:,:,:,:,t] = results[:,0].reshape(dim)
cgrid[:,:,:,:,:,:,t] = np.array([r[0] for r in results[:,1]]).reshape(dim)
bgrid[:,:,:,:,:,:,t] = np.array([r[1] for r in results[:,1]]).reshape(dim)
kgrid[:,:,:,:,:,:,t] = np.array([r[2] for r in results[:,1]]).reshape(dim)
qgrid[:,:,:,:,:,:,t] = np.array([r[3] for r in results[:,1]]).reshape(dim)
pool.close()
# np.save("Vgrid" + str(H), Vgrid)
# np.save("cgrid" + str(H), cgrid)
# np.save("bgrid" + str(H), bgrid)
# np.save("kgrid" + str(H), kgrid)
# np.save("qgrid" + str(H), qgrid)
```
| github_jupyter |
```
# HIDDEN
from datascience import *
from prob140 import *
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
%matplotlib inline
import math
from scipy import stats
```
### Randomizing a Parameter ###
In an earlier chapter we saw that Poissonizing the number of i.i.d. Bernoulli trials has a remarkable effect on the relation between the number of successes and the number of failures. In other situations too, randomizing the parameter of a standard model can affect supposedly well-understood relations between random variables.
In this section we will study one simple example of how randomizing a parameter affects dependence and independence.
### Tossing a Random Coin ###
Suppose I have three coins. Coin 1 lands heads with chance 0.25, Coin 2 with chance 0.5, and Coin 3 with chance 0.75. I pick a coin at random and toss it twice. Let's define some notation:
- $X$ is the label of the coin that I pick.
- $Y$ is the number of heads in the two tosses.
Then $X$ is uniform on $\{1, 2, 3\}$, and given $X$, the conditional distribution of $Y$ is binomial with $n=2$ and $p$ corresponding to the given coin. Here is the joint distribution table for $X$ and $Y$, along with the marginal of $X$.
```
x = make_array(1, 2, 3)
y = np.arange(3)
def jt(x, y):
if x == 1:
return (1/3)*stats.binom.pmf(y, 2, 0.25)
if x == 2:
return (1/3)*stats.binom.pmf(y, 2, 0.5)
if x == 3:
return (1/3)*stats.binom.pmf(y, 2, 0.75)
dist_tbl = Table().values('X', x, 'Y', y).probability_function(jt)
dist = dist_tbl.toJoint()
dist.marginal('X')
```
And here is the posterior distribution of $X$ given each different value of $Y$:
```
dist.conditional_dist('X', 'Y')
```
As we have seen in earlier examples, when the given number of heads is low, the posterior distribution favors the coin that is biased towards tails. When the given number of heads is high, it favors the coin that is biased towards heads.
### Are the Two Tosses Independent? ###
We have always assumed that tosses of a coin are independent of each other. But within that assumption was another assumption, unspoken: *we knew which coin we were tossing*. That is, the chance of heads $p$ was a fixed number. But now we don't know which coin we are tossing, so we have to be careful.
Let $H_i$ be the event that Toss $i$ lands heads. Then
$$
P(H_1) = \frac{1}{3}\cdot 0.25 ~+~ \frac{1}{3}\cdot 0.5 ~+~ \frac{1}{3}\cdot 0.75 ~=~ 0.5 ~=~ P(H_2)
$$
So each toss is equally likely to be heads or tails. Now let's find $P(H_1H_2)$. If the two tosses are independent, our answer shoud be 0.25.
$$
P(H_1H_2) = \frac{1}{3}\cdot 0.25^2 ~+~ \frac{1}{3}\cdot 0.5^2 ~+~ \frac{1}{3}\cdot 0.75^2 ~=~ 0.2917 ~ \ne P(H_1)P(H_2)
$$
```
(1/3)*(0.25**2 + 0.5**2 + 0.75**2)
```
**The two tosses are not independent.** Because the coin itself is random, knowing the result of Toss 1 tells you something about which coin was picked, and hence affects the probability that Toss 2 lands heads.
$$
P(H_2 \mid H_1) = \frac{P(H_1H_2)}{P(H_1)} = \frac{0.2917}{0.5} = 0.5834 > 0.5 = P(H_2)
$$
Knowing that the first coin landed heads makes it more likely that Coin 3 was picked, and hence increases the conditional chance that that the second toss will be a head.
This example shows that you have to be careful about how data can affect probabilities. To make justifiable conclusions based on your data, keep assumptions in mind when you calculate probabilities, and use the division rule to update probabilities as more data comes in.
| github_jupyter |
```
!pip install -U finance-datareader
import FinanceDataReader as fdr
df_krx = fdr.StockListing('KRX')
import sqlite3
conn = sqlite3.connect('./db.stock')
c = conn.cursor()
c.execute("CREATE TABLE IF NOT EXISTS article (id INTEGER PRIMARY KEY AUTOINCREMENT, date TEXT, time TEXT, title TEXT, press TEXT , stock TEXT, posi_nega TEXT)")
from bs4 import BeautifulSoup
import requests
import pandas as pd
# https://jsikim1.tistory.com/143
from datetime import datetime , timedelta
from dateutil.relativedelta import relativedelta
now = datetime.now()
gap = now - relativedelta(years=1)
# gap = now - timedelta(days=1)
now = str(now)[0:10]
gap = str(gap)[0:10]
dt_index = pd.date_range(start=gap, end=now)
dt_list = dt_index.strftime("%Y%m%d").tolist()
for j in dt_list:
date_cnt_uri = 'https://finance.naver.com/news/news_list.nhn?mode=LSS3D§ion_id=101§ion_id2=258§ion_id3=402&date='+j+'&page=100'
date_cnt_target = date_cnt_uri
date_cnt_req = requests.get(date_cnt_target)
date_cnt_soup = BeautifulSoup(date_cnt_req.content,'html.parser')
date_cnt_page = int(date_cnt_soup.select('td.on > a ')[0].get_text())
uri = 'https://finance.naver.com/news/news_list.nhn?mode=LSS3D§ion_id=101§ion_id2=258§ion_id3=402&date='+j+'&page='
for page in range(1,date_cnt_page+1):
target = uri+str(page)
req = requests.get(target)
soup = BeautifulSoup(req.content,'html.parser')
datas = soup.select('#contentarea_left > ul.realtimeNewsList')
for content in datas:
titles = content.select(' li > dl > dd.articleSubject')
article_date = content.select('li > dl > dd.articleSummary > span.wdate ')
article_press = content.select('li > dl > dd.articleSummary > span.press ')
article_sum = list()
for i in range(0,len(titles)-1):
article_data = list()
data_date = article_date[i].get_text(" ",strip=True)[0:10]
data_time = article_date[i].get_text(" ",strip=True)[11:17]
data_press = article_press[i].get_text(" ",strip=True)
data_title = titles[i].get_text(" ",strip=True)
c.execute("INSERT INTO article( date , time , press , title, stock , posi_nega ) VALUES(?,?,?,?,?,?)",( data_date,data_time,data_press,data_title,'stock','posi_nega'))
conn.commit()
c.close()
conn = sqlite3.connect('/content/db.stock')
c_title = conn.cursor()
c_id = conn.cursor()
c_update = conn.cursor()
c_title.execute("SELECT title FROM article ")
c_id.execute("SELECT * FROM article ")
sql_title = c_title.fetchall()
sql_id = c_id.fetchall()
title_list = [list[0] for list in sql_title ]
id_list = [list[0] for list in sql_id ]
df_krx_list = df_krx['Name'].tolist()
for k in range(0,len(title_list)):
for l in range(0,int(len(title_list[k].split()))):
keyword = title_list[k].split()[l]
if keyword in df_krx_list:
sql_update = 'update article set stock = "' + keyword +'" where id = ' + str(sql_id[k][0])
c_update.execute(sql_update)
conn.commit()
conn = sqlite3.connect('/content/db.stock')
c_select = conn.cursor()
final = c_select.execute("SELECT * FROM article where stock != 'stock' order by date desc , time desc")
df = pd.DataFrame(final)
df.columns= ['id', 'date' , 'time' , 'title', 'press' ,'stock' , 'posi_nega' ]
df
```
| github_jupyter |
# Offline reinforcement learning with Ray AIR
In this example, we'll train a reinforcement learning agent using offline training.
Offline training means that the data from the environment (and the actions performed by the agent) have been stored on disk. In contrast, online training samples experiences live by interacting with the environment.
Let's start with installing our dependencies:
```
!pip install -qU "ray[rllib]" gym
```
Now we can run some imports:
```
import argparse
import gym
import os
import numpy as np
import ray
from ray.air import Checkpoint
from ray.air.config import RunConfig
from ray.train.rl.rl_predictor import RLPredictor
from ray.train.rl.rl_trainer import RLTrainer
from ray.air.result import Result
from ray.rllib.agents.marwil import BCTrainer
from ray.tune.tuner import Tuner
```
We will be training on offline data - this means we have full agent trajectories stored somewhere on disk and want to train on these past experiences.
Usually this data could come from external systems, or a database of historical data. But for this example, we'll generate some offline data ourselves and store it using RLlibs `output_config`.
```
def generate_offline_data(path: str):
print(f"Generating offline data for training at {path}")
trainer = RLTrainer(
algorithm="PPO",
run_config=RunConfig(stop={"timesteps_total": 5000}),
config={
"env": "CartPole-v0",
"output": "dataset",
"output_config": {
"format": "json",
"path": path,
"max_num_samples_per_file": 1,
},
"batch_mode": "complete_episodes",
},
)
trainer.fit()
```
Here we define the training function. It will create an `RLTrainer` using the `PPO` algorithm and kick off training on the `CartPole-v0` environment. It will use the offline data provided in `path` for this.
```
def train_rl_bc_offline(path: str, num_workers: int, use_gpu: bool = False) -> Result:
print("Starting offline training")
dataset = ray.data.read_json(
path, parallelism=num_workers, ray_remote_args={"num_cpus": 1}
)
trainer = RLTrainer(
run_config=RunConfig(stop={"training_iteration": 5}),
scaling_config={
"num_workers": num_workers,
"use_gpu": use_gpu,
},
datasets={"train": dataset},
algorithm=BCTrainer,
config={
"env": "CartPole-v0",
"framework": "tf",
"evaluation_num_workers": 1,
"evaluation_interval": 1,
"evaluation_config": {"input": "sampler"},
},
)
# Todo (krfricke/xwjiang): Enable checkpoint config in RunConfig
# result = trainer.fit()
tuner = Tuner(
trainer,
_tuner_kwargs={"checkpoint_at_end": True},
)
result = tuner.fit()[0]
return result
```
Once we trained our RL policy, we want to evaluate it on a fresh environment. For this, we will also define a utility function:
```
def evaluate_using_checkpoint(checkpoint: Checkpoint, num_episodes) -> list:
predictor = RLPredictor.from_checkpoint(checkpoint)
env = gym.make("CartPole-v0")
rewards = []
for i in range(num_episodes):
obs = env.reset()
reward = 0.0
done = False
while not done:
action = predictor.predict([obs])
obs, r, done, _ = env.step(action[0])
reward += r
rewards.append(reward)
return rewards
```
Let's put it all together. First, we initialize Ray and create the offline data:
```
ray.init(num_cpus=8)
path = "/tmp/out"
generate_offline_data(path)
```
Then, we run training:
```
result = train_rl_bc_offline(path=path, num_workers=2, use_gpu=False)
```
And then, using the obtained checkpoint, we evaluate the policy on a fresh environment:
```
num_eval_episodes = 3
rewards = evaluate_using_checkpoint(result.checkpoint, num_episodes=num_eval_episodes)
print(f"Average reward over {num_eval_episodes} episodes: " f"{np.mean(rewards)}")
```
| github_jupyter |
```
import os
os.chdir('../app')
import matplotlib
print(matplotlib.__version__)
import frontend.stock_analytics as salib
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
from pandas.plotting import register_matplotlib_converters
register_matplotlib_converters()
from datetime import datetime,timedelta
from pprint import pprint
import matplotlib.patches as patches
import time
import numpy as np
import datetime
import copy
import preprocessing.lob.s03_fill_cache as l03
import re
import preprocessing.preglobal as pg
import math
%matplotlib inline
import random
import math
import scipy.optimize
import scipy.optimize
import json
import analysis_lib as al
import scipy.special
import cv2
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
from pymongo import MongoClient, UpdateMany, UpdateOne, InsertOne
import pandas as pd
plt.rcParams['figure.figsize'] = (15, 5)
def binary_search( f, target, cstep=10, stepsize=10, prevturn=True): # mon increasing func
#print(f(cstep), target, cstep, stepsize)
if cstep > 1e5:
return -1
res = target/f(cstep)
if np.abs(res-1) < 1e-4:
return cstep
if res < 1:
stepsize /= 2
prevturn=False
cstep -= stepsize
else:
if prevturn:
stepsize *= 2
else:
stepsize /= 2
cstep += stepsize
return binary_search( f, target, cstep, stepsize,prevturn)
# Simulate using inverse transform
# Theoretische Verteilung
def integral_over_phi_slow(t,deltat, omegak, a, K, phi_0,g):
summand = 0
if len(t) > 0:
for k in range(0,K):
summand += (1-np.exp(-omegak[k]*deltat))*np.sum(a[k]*np.exp(-omegak[k]*(t[-1]-t)))
return deltat*phi_0 + g*summand
def integral_over_phi(t,deltat, omegak, a, K, phi_0,g):
summand = np.sum((1-np.exp(-np.outer(omegak,deltat))).T * np.sum(np.multiply(np.exp(-np.outer(omegak,(t[-1]-t))).T,a), axis=0) ,axis=1) \
if len(t) > 0 else 0
return deltat*phi_0 + g*summand
def probability_for_inter_arrival_time(t, deltat, omegak, a, K, phi_0,g):
x= integral_over_phi(t,deltat, omegak, a, K, phi_0,g)
return 1-np.exp(-x)
def probability_for_inter_arrival_time_slow(t, deltat, omegak, a, K, phi_0,g):
x = np.zeros(len(deltat))
for i in range(0, len(deltat)):
x[i]= integral_over_phi_slow(t,deltat[i], omegak, a, K, phi_0,g)
return 1-np.exp(-x)
g_cache_dict = {}
def simulate_by_itrans(phi_dash, g_params, K, conv1=1e-8, conv2=1e-2, N = 250000, init_array=np.array([]), reseed=True, status_update=True, use_binary_search=True):
# Initialize parameters
g, g_omega, g_beta = g_params
phi_0 = phi_dash * (1-g)
omegak, a = al.generate_series_parameters(g_omega, g_beta, K)
if reseed:
np.random.seed(123)
salib.tic()
i = randii = 0
t = 0.
randpool = np.random.rand(100*N)
# Inverse transform algorithm
init_array = np.array(init_array, dtype='double')
hawkes_array = np.pad(init_array,(0,N-len(init_array)), 'constant', constant_values=0.) #np.zeros(N)
hawkes_array = np.array(hawkes_array, dtype='double')
i = len(init_array)
if i > 0:
t = init_array[-1]
endsize = 20
tau = 0
while i < N:
NN = 10000
u = randpool[randii]
randii+=1
if randii >= len(randpool):
print(i)
if use_binary_search:
f = lambda x: probability_for_inter_arrival_time(hawkes_array[:i],x, omegak, a, K, phi_0, g)
tau = binary_search( f, u,cstep=max(tau,1e-5), stepsize=max(tau,1e-5))
if tau == -1:
return hawkes_array[:i]
else:
notok = 1
while notok>0:
if notok > 10:
NN *= 2
notok = 1
tau_x = np.linspace(0,endsize,NN)
pt = probability_for_inter_arrival_time (hawkes_array[:i],tau_x, omegak, a, K, phi_0, g)
okok = True
if pt[-1]-pt[-2] > conv1:
if status_update:
print('warning, pt does not converge',i,pt[1]-pt[0],pt[-1]-pt[-2])
endsize*=1.1
notok += 1
okok = False
if pt[1]-pt[0] > conv2:
if status_update:
print('warning pt increases to fast',i,pt[1]-pt[0],pt[-1]-pt[-2])
endsize/=1.1
notok +=1
okok = False
if okok:
notok = 0
tt = np.max(np.where(pt < u))
if tt == NN-1:
if status_update:
print('vorzeitig abgebrochen', u, tau_x[tt], pt[tt])
return hawkes_array[:i]
tau = tau_x[tt]
t += tau
hawkes_array[i] = t
i += 1
if status_update and i%(int(N/5))==0:
print(i)
salib.toc()
if status_update:
salib.toc()
return hawkes_array
# SIMULATION USING THINNING
def calc_eff_g(number_of_events, g):
noe_binned_x, noe_binned_y, _ = al.dobins(number_of_events, useinteger=True, N=1000)
noe_binned_y /= noe_binned_y.sum()
assert np.abs(np.sum(noe_binned_y) - 1) < 1e-8
print((noe_binned_x*noe_binned_y).sum(), 'should be', 1/(1-g))
plt.plot(np.log(noe_binned_x),noe_binned_y)
# noe_thin_no_cache_K15
gg = 0.886205
noe_thin_no_cache_K15 = [len(\
simulate_by_thinning(phi_dash=0, g_params=(gg, 0.430042, 0.3),\
K=15, N=1000, reseed=False, status_update=False, caching=False, init_array=np.array([0.]))\
) for i in range(0,10000)]
# noe_thin_cache_K15
gg = 0.886205
noe_thin_cache_K15 = [len(\
simulate_by_thinning(phi_dash=0, g_params=(gg, 0.430042, 0.3),\
K=15, N=1000, reseed=False, status_update=False, caching=True, init_array=np.array([0.]))\
) for i in range(0,10000)]
# noe_itrans_binary_K15
gg = 0.886205
noe_itrans_binary_K15 = [len(\
simulate_by_itrans(phi_dash=0, g_params=(gg, 0.430042, 0.3),\
K=15, N=1000, reseed=False, status_update=False,use_binary_search=True, init_array=np.array([0.]))\
) for i in range(0,10000)]
# noe_itrans_no_binary_K15
gg = 0.886205
noe_itrans_no_binary_K15 = [len(\
simulate_by_itrans(phi_dash=0, g_params=(gg, 0.430042, 0.3),\
K=15, N=1000, reseed=False, status_update=False, use_binary_search=False, init_array=np.array([0.])
, conv1=1e-5, conv2=1e-2
)\
) for i in range(0,10000)]
#noe_thin_cache_K0
gg = 0.886205
noe_thin_cache_K0 = [len(\
simulate_by_thinning(phi_dash=0, g_params=(gg, 2.430042, 0.),\
K=1, N=1000, reseed=False, status_update=False, caching=True, init_array=np.array([0.]))\
) for i in range(0,10000)] # braucht recht lang, weil der cache jedes mal neu aufgebaut wird
# noe_thin_no_cache_K0
gg = 0.886205
noe_thin_no_cache_K0 = [len(\
simulate_by_thinning(phi_dash=0, g_params=(gg, 2.430042, 0.),\
K=1, N=1000, reseed=False, status_update=False, caching=False, init_array=np.array([0.]))\
) for i in range(0,10000)]
# noe_itrans_binary_K0
gg = 0.886205
noe_itrans_binary_K0 = [len(\
simulate_by_itrans(phi_dash=0, g_params=(gg, 2.430042, 0.),\
K=1, N=1000, reseed=False, use_binary_search=True, status_update=False, init_array=np.array([0.]))\
) for i in range(0,10000)]
# noe_itrans_no_binary_K0
gg = 0.886205
noe_itrans_no_binary_K0 = [len(\
simulate_by_itrans(phi_dash=0, g_params=(gg, 2.430042, 0.),\
K=1, N=1000, reseed=False, use_binary_search=False, status_update=False, init_array=np.array([0.]))\
) for i in range(0,10000)]
calc_eff_g(noe_thin_cache_K15,gg)
calc_eff_g(noe_thin_no_cache_K15,gg)
calc_eff_g(noe_thin_cache_K0,gg)
calc_eff_g(noe_thin_no_cache_K0,gg)
calc_eff_g(noe_itrans_binary_K15,gg)
calc_eff_g(noe_itrans_no_binary_K15,gg)
calc_eff_g(noe_itrans_binary_K0,gg)
calc_eff_g(noe_itrans_no_binary_K0,gg)
eff_g_sim = {
'noe_thin_cache_K15':noe_thin_cache_K15,
'noe_thin_no_cache_K15':noe_thin_no_cache_K15,
'noe_thin_cache_K0':noe_thin_cache_K0,
'noe_thin_no_cache_K0':noe_thin_no_cache_K0,
'noe_itrans_binary_K15':noe_itrans_binary_K15,
'noe_itrans_no_binary_K15':noe_itrans_no_binary_K15,
'noe_itrans_binary_K0':noe_itrans_binary_K0,
'noe_itrans_no_binary_K0':noe_itrans_no_binary_K0
}
with open('eff_g_sim.json','w') as f:
json.dump( eff_g_sim, f)
gg
sim_thin_no_cache = simulate_by_thinning(phi_dash=68, g_params=(0.886205, 0.430042, 0.253835), K=15, N=10000, caching=False)
sim_thin_cache = simulate_by_thinning(phi_dash=68, g_params=(0.886205, 0.430042, 0.253835), K=15, N=10000, caching=True)
sim_itrans_binary = simulate_by_itrans(phi_dash=68, g_params=(0.886205, 0.430042, 0.253835), use_binary_search=True, K=15, N=10000, reseed=False)
sim_itrans_nobinary = simulate_by_itrans(phi_dash=68, g_params=(0.886205, 0.430042, 0.253835), use_binary_search=False, K=15, N=10000, reseed=False)
import importlib
importlib.reload(al)
import task_lib as tl
with open('17_simulation.json', 'w') as f:
json.dump([ ('sim_thin_no_cache',sim_thin_no_cache),
('sim_itrans_binary',sim_itrans_binary),
('sim_itrans_nobinary',sim_itrans_nobinary)],f, cls=tl.NumpyEncoder)
al.print_stats([ ('sim_thin_no_cache',sim_thin_no_cache),
('sim_itrans_binary',sim_itrans_binary),
('sim_itrans_nobinary',sim_itrans_nobinary)],
tau = np.logspace(-1,1,20), stepsize_hist=1.)
# Show probability distribution!
tg, tg_omega, tg_beta = (0.786205, 0.430042, 0.253835)
tK = 15
tphi_0 = 0
tomegak, ta = al.generate_series_parameters(tg_omega, tg_beta, K=tK, b=5.)
thawkes_array = np.zeros(10)
thawkes_array[0] = 0
ti = 1
tj = 0
tau_x = np.linspace(0.,100,1000)
pt = probability_for_inter_arrival_time(thawkes_array[tj:ti],tau_x, tomegak, ta, tK, tphi_0, tg)
plt.plot(tau_x,pt,'.')
# TEST IF BOTH ARE THE SAME
tt = np.array([0.01388255])
tdeltat = np.linspace(0,1.2607881726256949,1000)
tomegak = np.array([0.430042, 0.0006565823727274271, 1.0024611832713502e-06, 1.5305443242275112e-09, 2.3368145994246977e-12, 3.567817269741932e-15, 5.447295679084947e-18, 8.31685817180986e-21, 1.2698067798225314e-23, 1.9387240046350152e-26, 2.960017875060756e-29, 4.519315694101945e-32, 6.900030744759242e-35, 1.053487463616628e-37, 1.6084505664563106e-40])
ta = np.array([0.8071834195758446, 0.15563834675047422, 0.030009653805760286, 0.005786358827014711, 0.0011157059222237438, 0.0002151266006998307, 4.1479975508621093e-05, 7.998027034306966e-06, 1.5421522230216854e-06, 2.973525181609743e-07, 5.733449573701989e-08, 1.1055041409263513e-08, 2.1315952811567083e-09, 4.110069129946613e-10, 7.924894750082819e-11])
tK = 15
tphi_0 = 7.738059999999999
tg = 0.886205
assert (np.abs(probability_for_inter_arrival_time_slow(tt, tdeltat, tomegak, ta, tK, tphi_0, tg) - probability_for_inter_arrival_time(tt, tdeltat, tomegak, ta, tK, tphi_0, tg)) < 1e-10).all()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/geantrindade/ConvNet-Performance-Prediction/blob/master/notebooks/dataset_meta_extractor.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# **Imports**
```
!pip install -U pymfe
from pymfe.mfe import MFE
import datetime
import numpy as np
from numpy import savez_compressed
from numpy import load
import torch
from torch.utils.data import DataLoader
from torchvision.datasets import MNIST
from torchvision.transforms import Compose, ToTensor, Normalize
import pandas as pd
import matplotlib.pyplot as plt
```
# **Load and pre-process data**
---
```
def get_data_loaders(train_batch_size, test_batch_size):
mnist = MNIST(download=True, train=True, root=".").data.float()
data_transform = Compose([ToTensor(), Normalize((mnist.mean()/255,), (mnist.std()/255,))])
#data_transform = Compose([ToTensor(), Normalize((0.5,), (0.5,))])
train_loader = DataLoader(MNIST(download=True, root=".", transform=data_transform, train=True),
batch_size=train_batch_size, shuffle=True)
test_loader = DataLoader(MNIST(download=True, root=".", transform=data_transform, train=False),
batch_size=test_batch_size, shuffle=False)
return train_loader, test_loader
train_loader, test_loader = get_data_loaders(60000, 10000)
mnist_batch_train = next(iter(train_loader))
mnist_batch_test = next(iter(test_loader))
whole_mnist_X = torch.cat((mnist_batch_train[0], mnist_batch_test[0]), 0)
whole_mnist_Y = torch.cat((mnist_batch_train[1], mnist_batch_test[1]), 0)
np_mnist_batch_train_X = mnist_batch_train[0].numpy()
np_mnist_batch_train_Y = mnist_batch_train[1].numpy()
np_mnist_batch_test_X = mnist_batch_test[0].numpy()
np_mnist_batch_test_Y = mnist_batch_test[1].numpy()
np_whole_mnist_X = whole_mnist_X.numpy()
np_whole_mnist_Y = whole_mnist_Y.numpy()
#flat pixels
np_mnist_batch_train_X = np_mnist_batch_train_X.reshape((np_mnist_batch_train_X.shape[0], np_mnist_batch_train_X.shape[2] * np_mnist_batch_train_X.shape[3]))
np_mnist_batch_test_X = np_mnist_batch_test_X.reshape((np_mnist_batch_test_X.shape[0], np_mnist_batch_test_X.shape[2] * np_mnist_batch_test_X.shape[3]))
np_whole_mnist_X = np_whole_mnist_X.reshape((np_whole_mnist_X.shape[0], np_whole_mnist_X.shape[2] * np_whole_mnist_X.shape[3]))
```
# **Meta-Feature extraction**
```
def extract_meta_features(pymfe_obj, X_data, Y_data) -> tuple:
pymfe_obj.fit(X_data, Y_data)
begin_t = datetime.datetime.now()
meta_features = pymfe_obj.extract()
end_t = datetime.datetime.now()
print("\n\nmeta-extraction total time: ", end_t - begin_t, "\n")
return meta_features
def print_meta_features(meta_features : tuple):
print("\n".join("{:30} {:30} {:30}".format(x, y, z) for x, y, z in zip(meta_features[0], meta_features[1], meta_features[2])))
print("\nnumber of meta-features: ", len(meta_features[0]))
meta_features_sets = []
```
## Version 1
```
mfe = MFE(features=["attr_ent", "attr_to_inst","can_cor", "class_conc", "class_ent", "cov", "eigenvalues", "eq_num_attr", "freq_class", "gravity", "inst_to_attr",
"iq_range" , "joint_ent", "kurtosis", "mad", "max", "mean", "median", "min", "mut_inf", "nr_attr", "nr_bin", "nr_class", "nr_cor_attr",
"nr_disc", "nr_inst", "nr_norm", "nr_num", "nr_outliers", "ns_ratio", "range", "sd", "skewness", "sparsity", "t_mean", "var", "w_lambda"],
summary=["mean", "sd"],
measure_time="total")
#train
mft_train_v1 = extract_meta_features(mfe, np_mnist_batch_train_X, np_mnist_batch_train_Y)
print_meta_features(mft_train_v1)
#test
mft_test_v1 = extract_meta_features(mfe, np_mnist_batch_test_X, np_mnist_batch_test_Y)
print_meta_features(mft_test_v1)
#whole
mft_whole_v1 = extract_meta_features(mfe, np_whole_mnist_X, np_whole_mnist_Y)
print_meta_features(mft_whole_v1)
meta_features_sets.append((mft_train_v1, mft_test_v1, mft_whole_v1))
```
## Version 2
```
mfe = MFE(features=["attr_ent", "attr_to_inst","can_cor", "class_conc", "class_ent", "cov", "eigenvalues", "eq_num_attr", "freq_class", "gravity", "inst_to_attr",
"iq_range" , "joint_ent", "kurtosis", "mad", "max", "mean", "median", "min", "mut_inf", "nr_attr", "nr_bin", "nr_class", "nr_cor_attr",
"nr_disc", "nr_inst", "nr_norm", "nr_num", "nr_outliers", "ns_ratio", "range", "sd", "skewness", "sparsity", "t_mean", "var", "w_lambda"],
summary=["mean"],
measure_time="total")
#train
mft_train_v2 = extract_meta_features(mfe, np_mnist_batch_train_X, np_mnist_batch_train_Y)
print_meta_features(mft_train_v2)
#test
mft_test_v2 = extract_meta_features(mfe, np_mnist_batch_test_X, np_mnist_batch_test_Y)
print_meta_features(mft_test_v2)
#whole
mft_whole_v2 = extract_meta_features(mfe, np_whole_mnist_X, np_whole_mnist_Y)
print_meta_features(mft_whole_v2)
meta_features_sets.append((mft_train_v2, mft_test_v2, mft_whole_v2))
```
## Version 3
```
mfe = MFE(features=["attr_ent", "attr_to_inst","can_cor", "class_conc", "class_ent", "cov", "eigenvalues", "eq_num_attr", "freq_class", "gravity", "inst_to_attr",
"iq_range" , "joint_ent", "kurtosis", "mad", "max", "mean", "median", "min", "mut_inf", "nr_attr", "nr_bin", "nr_class", "nr_cor_attr",
"nr_disc", "nr_inst", "nr_norm", "nr_num", "nr_outliers", "ns_ratio", "range", "sd", "skewness", "sparsity", "t_mean", "var", "w_lambda"],
summary=["max", "min", "median", "mean", "var", "sd", "kurtosis", "skewness"],
measure_time="total")
#train
mft_train_v3 = extract_meta_features(mfe, np_mnist_batch_train_X, np_mnist_batch_train_Y)
print_meta_features(mft_train_v3)
#test
mft_test_v3 = extract_meta_features(mfe, np_mnist_batch_test_X, np_mnist_batch_test_Y)
print_meta_features(mft_test_v3)
#whole
mft_whole_v3 = extract_meta_features(mfe, np_whole_mnist_X, np_whole_mnist_Y)
print_meta_features(mft_whole_v3)
meta_features_sets.append((mft_train_v3, mft_test_v3, mft_whole_v3))
```
# **DataFrame creation**
```
def print_meta_features_dict(mtf : dict):
print("\nnumber of meta-features: ", len(mtf))
print(mtf)
for idx_v, version in enumerate(meta_features_sets):
for idx_p, partition in enumerate(version):
meta_features_dict = {'dataset.name' : 'mnist'}
for i in range(1, len(partition[0])):
mtf_key, mtf_value = str(partition[0][i]), partition[1][i]
meta_features_dict[mtf_key] = mtf_value
df = pd.DataFrame(data=meta_features_dict, index=[0])
#mnist_metafeatures_train_v1, mnist_metafeatures_test_v1, mnist_metafeatures_whole_v1, cifar_metafeatures_train_v1...
csv_name = meta_features_dict.get('dataset.name') + "_metafeatures_" + ("train" if (idx_p == 0) else "test" if (idx_p == 1) else "whole") + "_v" + str(idx_v + 1)
df.to_csv(csv_name + ".csv", index=False)
print_meta_features_dict(meta_features_dict)
```
# **Debug**
```
print("train batch length: ", len(mnist_batch_train))
print("train batch X shape: ", mnist_batch_train[0].shape)
print("train batch X example: ", mnist_batch_train[0])
print("train batch Y shape: ", mnist_batch_train[1].shape)
print("train batch Y example: ", mnist_batch_train[1])
print("\ntest batch X length: ", len(mnist_batch_test))
print("test batch X shape: ", mnist_batch_test[0].shape)
print("test batch X example: ", mnist_batch_test[0])
print("test batch Y shape: ", mnist_batch_test[1].shape)
print("test batch Y example: ", mnist_batch_test[1])
print("\nwhole_mnist_X length: ", len(whole_mnist_X))
print("whole_mnist_X shape: ", whole_mnist_X[0].shape)
print("whole_mnist_X example: ", whole_mnist_X[0])
print("whole_mnist_Y length: ", len(whole_mnist_Y))
print("whole_mnist_Y shape: ", whole_mnist_Y[0].shape)
print("whole_mnist_Y example: ", whole_mnist_Y[0])
image_index = 0
print(mnist_batch_train[1][image_index]) #label
temp = mnist_batch_train[0].numpy()
print(temp.shape)
temp = temp.reshape((temp.shape[0], temp.shape[2], temp.shape[3]))
print(temp.shape)
plt.imshow(temp[image_index], cmap='Greys')
print(whole_mnist_Y[image_index]) #label
temp = whole_mnist_X[0].numpy()
print(temp.shape)
temp = temp.reshape((temp.shape[0], temp.shape[1], temp.shape[2]))
print(temp.shape)
plt.imshow(temp[image_index], cmap='Greys')
print("train batch length: ", len(np_mnist_batch_train_X))
print("train batch X shape: ", np_mnist_batch_train_X.shape)
print("train batch X example: ", np_mnist_batch_train_X[0])
print("train batch Y shape: ", np_mnist_batch_train_Y.shape)
print("train batch Y example: ", np_mnist_batch_train_Y[0])
print("\ntest batch X length: ", len(np_mnist_batch_test_X))
print("test batch X shape: ", np_mnist_batch_test_X.shape)
print("test batch X example: ", np_mnist_batch_test_X[0])
print("test batch Y shape: ", np_mnist_batch_test_Y.shape)
print("test batch Y example: ", np_mnist_batch_test_Y[0])
print("\nnp_whole_mnist_X length: ", len(np_whole_mnist_X))
print("np_whole_mnist_X shape: ", np_whole_mnist_X.shape)
print("np_whole_mnist_X example: ", np_whole_mnist_X[0])
print("np_whole_mnist_Y length: ", len(np_whole_mnist_Y))
print("np_whole_mnist_Y shape: ", np_whole_mnist_Y.shape)
print("np_whole_mnist_Y example: ", np_whole_mnist_Y[0])
```
| github_jupyter |
#Improving Computer Vision Accuracy using Convolutions
In the previous lessons you saw how to do fashion recognition using a Deep Neural Network (DNN) containing three layers -- the input layer (in the shape of the data), the output layer (in the shape of the desired output) and a hidden layer. You experimented with the impact of different sized of hidden layer, number of training epochs etc on the final accuracy.
For convenience, here's the entire code again. Run it and take a note of the test accuracy that is printed out at the end.
```
import tensorflow as tf
mnist = tf.keras.datasets.fashion_mnist
(training_images, training_labels), (test_images, test_labels) = mnist.load_data()
training_images=training_images / 255.0
test_images=test_images / 255.0
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
model.fit(training_images, training_labels, epochs=5)
test_loss = model.evaluate(test_images, test_labels)
```
Your accuracy is probably about 89% on training and 87% on validation...not bad...But how do you make that even better? One way is to use something called Convolutions. I'm not going to details on Convolutions here, but the ultimate concept is that they narrow down the content of the image to focus on specific, distinct, details.
If you've ever done image processing using a filter (like this: https://en.wikipedia.org/wiki/Kernel_(image_processing)) then convolutions will look very familiar.
In short, you take an array (usually 3x3 or 5x5) and pass it over the image. By changing the underlying pixels based on the formula within that matrix, you can do things like edge detection. So, for example, if you look at the above link, you'll see a 3x3 that is defined for edge detection where the middle cell is 8, and all of its neighbors are -1. In this case, for each pixel, you would multiply its value by 8, then subtract the value of each neighbor. Do this for every pixel, and you'll end up with a new image that has the edges enhanced.
This is perfect for computer vision, because often it's features that can get highlighted like this that distinguish one item for another, and the amount of information needed is then much less...because you'll just train on the highlighted features.
That's the concept of Convolutional Neural Networks. Add some layers to do convolution before you have the dense layers, and then the information going to the dense layers is more focussed, and possibly more accurate.
Run the below code -- this is the same neural network as earlier, but this time with Convolutional layers added first. It will take longer, but look at the impact on the accuracy:
```
import tensorflow as tf
print(tf.__version__)
mnist = tf.keras.datasets.fashion_mnist
(training_images, training_labels), (test_images, test_labels) = mnist.load_data()
training_images=training_images.reshape(60000, 28, 28, 1)
training_images=training_images / 255.0
test_images = test_images.reshape(10000, 28, 28, 1)
test_images=test_images/255.0
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(64, (3,3), activation='relu', input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D(2, 2),
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
model.summary()
model.fit(training_images, training_labels, epochs=5)
test_loss = model.evaluate(test_images, test_labels)
```
It's likely gone up to about 93% on the training data and 91% on the validation data.
That's significant, and a step in the right direction!
Try running it for more epochs -- say about 20, and explore the results! But while the results might seem really good, the validation results may actually go down, due to something called 'overfitting' which will be discussed later.
(In a nutshell, 'overfitting' occurs when the network learns the data from the training set really well, but it's too specialised to only that data, and as a result is less effective at seeing *other* data. For example, if all your life you only saw red shoes, then when you see a red shoe you would be very good at identifying it, but blue suade shoes might confuse you...and you know you should never mess with my blue suede shoes.)
Then, look at the code again, and see, step by step how the Convolutions were built:
Step 1 is to gather the data. You'll notice that there's a bit of a change here in that the training data needed to be reshaped. That's because the first convolution expects a single tensor containing everything, so instead of 60,000 28x28x1 items in a list, we have a single 4D list that is 60,000x28x28x1, and the same for the test images. If you don't do this, you'll get an error when training as the Convolutions do not recognize the shape.
```
import tensorflow as tf
mnist = tf.keras.datasets.fashion_mnist
(training_images, training_labels), (test_images, test_labels) = mnist.load_data()
training_images=training_images.reshape(60000, 28, 28, 1)
training_images=training_images / 255.0
test_images = test_images.reshape(10000, 28, 28, 1)
test_images=test_images/255.0
```
Next is to define your model. Now instead of the input layer at the top, you're going to add a Convolution. The parameters are:
1. The number of convolutions you want to generate. Purely arbitrary, but good to start with something in the order of 32
2. The size of the Convolution, in this case a 3x3 grid
3. The activation function to use -- in this case we'll use relu, which you might recall is the equivalent of returning x when x>0, else returning 0
4. In the first layer, the shape of the input data.
You'll follow the Convolution with a MaxPooling layer which is then designed to compress the image, while maintaining the content of the features that were highlighted by the convlution. By specifying (2,2) for the MaxPooling, the effect is to quarter the size of the image. Without going into too much detail here, the idea is that it creates a 2x2 array of pixels, and picks the biggest one, thus turning 4 pixels into 1. It repeats this across the image, and in so doing halves the number of horizontal, and halves the number of vertical pixels, effectively reducing the image by 25%.
You can call model.summary() to see the size and shape of the network, and you'll notice that after every MaxPooling layer, the image size is reduced in this way.
```
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(32, (3,3), activation='relu', input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D(2, 2),
```
Add another convolution
```
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2)
```
Now flatten the output. After this you'll just have the same DNN structure as the non convolutional version
```
tf.keras.layers.Flatten(),
```
The same 128 dense layers, and 10 output layers as in the pre-convolution example:
```
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(10, activation='softmax')
])
```
Now compile the model, call the fit method to do the training, and evaluate the loss and accuracy from the test set.
```
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
model.fit(training_images, training_labels, epochs=5)
test_loss, test_acc = model.evaluate(test_images, test_labels)
print(test_acc)
```
# Visualizing the Convolutions and Pooling
This code will show us the convolutions graphically. The print (test_labels[;100]) shows us the first 100 labels in the test set, and you can see that the ones at index 0, index 23 and index 28 are all the same value (9). They're all shoes. Let's take a look at the result of running the convolution on each, and you'll begin to see common features between them emerge. Now, when the DNN is training on that data, it's working with a lot less, and it's perhaps finding a commonality between shoes based on this convolution/pooling combination.
```
print(test_labels[:100])
import matplotlib.pyplot as plt
f, axarr = plt.subplots(3,4)
FIRST_IMAGE=0
SECOND_IMAGE=7
THIRD_IMAGE=26
CONVOLUTION_NUMBER = 1
from tensorflow.keras import models
layer_outputs = [layer.output for layer in model.layers]
activation_model = tf.keras.models.Model(inputs = model.input, outputs = layer_outputs)
for x in range(0,4):
f1 = activation_model.predict(test_images[FIRST_IMAGE].reshape(1, 28, 28, 1))[x]
print(f1.shape)
break
axarr[0,x].imshow(f1[0, : , :, CONVOLUTION_NUMBER], cmap='inferno')
axarr[0,x].grid(False)
f2 = activation_model.predict(test_images[SECOND_IMAGE].reshape(1, 28, 28, 1))[x]
axarr[1,x].imshow(f2[0, : , :, CONVOLUTION_NUMBER], cmap='inferno')
axarr[1,x].grid(False)
f3 = activation_model.predict(test_images[THIRD_IMAGE].reshape(1, 28, 28, 1))[x]
axarr[2,x].imshow(f3[0, : , :, CONVOLUTION_NUMBER], cmap='inferno')
axarr[2,x].grid(False)
```
EXERCISES
1. Try editing the convolutions. Change the 32s to either 16 or 64. What impact will this have on accuracy and/or training time.
2. Remove the final Convolution. What impact will this have on accuracy or training time?
3. How about adding more Convolutions? What impact do you think this will have? Experiment with it.
4. Remove all Convolutions but the first. What impact do you think this will have? Experiment with it.
5. In the previous lesson you implemented a callback to check on the loss function and to cancel training once it hit a certain amount. See if you can implement that here!
```
import tensorflow as tf
print(tf.__version__)
mnist = tf.keras.datasets.mnist
(training_images, training_labels), (test_images, test_labels) = mnist.load_data()
training_images=training_images.reshape(60000, 28, 28, 1)
training_images=training_images / 255.0
test_images = test_images.reshape(10000, 28, 28, 1)
test_images=test_images/255.0
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(32, (3,3), activation='relu', input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D(2, 2),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
model.fit(training_images, training_labels, epochs=10)
test_loss, test_acc = model.evaluate(test_images, test_labels)
print(test_acc)
```
| github_jupyter |
# Conservative SDOF - Multiple Scales
- Introduces multiple time scales (Homogenation)
- Treate damped systems easier then L-P
- Built-in stability
Introduce new independent time variables
$$
\begin{gather*}
T_n = \epsilon^n t
\end{gather*}
$$
and
$$
\begin{align*}
\frac{d}{dt} &= \frac{\partial}{\partial T_0}\frac{dT_0}{dt} + \frac{\partial}{\partial T_1}\frac{dT_1}{dt} + \frac{\partial}{\partial T_2}\frac{dT_2}{dt} + \cdots\\
&= \frac{\partial}{\partial T_0} + \epsilon \frac{\partial}{\partial T_1} + \epsilon^2 \frac{\partial}{\partial T_2} + \cdots\\
&= D_0 + \epsilon D_1 + \epsilon^2 D_2 + \cdots
\end{align*}
$$
$$
\begin{align*}
\frac{d^2}{dt^2} &= \left( D_0 + \epsilon D_1 + \epsilon^2 D_2 + \cdots \right)^2
\end{align*}
$$
Introducing the Expansion for $x(t)$
$$
\begin{align*}
x(t) &= x_0(T_0,T_1,T_2,\cdots) + \epsilon x_1(T_0,T_1,T_2,\cdots) + \epsilon^2(T_0,T_1,T_2,\cdots) + \epsilon^3(T_0,T_1,T_2,\cdots) + O(\epsilon^4)
\end{align*}
$$
```
import sympy as sp
from sympy.simplify.fu import TR0, TR7, TR8, TR11
from math import factorial
# Functions for multiple scales
# Function for Time operator
def Dt(f, n, Ts, e=sp.Symbol('epsilon')):
if n==1:
return sp.expand(sum([e**i * sp.diff(f, T_i) for i, T_i in enumerate(Ts)]))
return Dt(Dt(f, 1, Ts, e), n-1, Ts, e)
def collect_epsilon(f, e=sp.Symbol('epsilon')):
N = sp.degree(f, e)
f_temp = f
collected_dict = {}
for i in range(N, 0, -1):
collected_term = f_temp.coeff(e**i)
collected_dict[e**i] = collected_term
delete_terms = sp.expand(e**i * collected_term)
f_temp -= delete_terms
collected_dict[e**0] = f_temp
return collected_dict
N = 3
f = sp.Function('f')
t = sp.Symbol('t', real=True)
# Define the symbolic parameters
epsilon = sp.symbols('epsilon')
T_i = sp.symbols('T_(0:' + str(N) + ')', real=True)
alpha_i = sp.symbols('alpha_(2:' + str(N+1) + ')', real=True)
omega0 = sp.Symbol('omega_0', real=True)
# x0 = sp.Function('x_0')(*T_i)
x1 = sp.Function('x_1')(*T_i)
x2 = sp.Function('x_2')(*T_i)
x3 = sp.Function('x_3')(*T_i)
# Expansion for x(t)
x_e = epsilon*x1 + epsilon**2 * x2 + epsilon**3 * x3
x_e
# Derivatives with time operators
xd = Dt(x_e, 1, T_i, epsilon)
xdd = Dt(x_e, 2, T_i, epsilon)
# EOM
EOM = xdd + sp.expand(omega0**2 * x_e) + sp.expand(sum([alpha_i[i-2] * x_e**i for i in range(2,N+1)]))
EOM
# Ordered Equations by epsilon
epsilon_Eq = collect_epsilon(EOM)
epsilon0_Eq = sp.Eq(epsilon_Eq[epsilon**0], 0)
epsilon0_Eq
epsilon1_Eq = sp.Eq(epsilon_Eq[epsilon**1], 0)
epsilon1_Eq
epsilon2_Eq = sp.Eq(epsilon_Eq[epsilon**2], 0)
epsilon2_Eq
epsilon3_Eq = sp.Eq(epsilon_Eq[epsilon**3], 0)
epsilon3_Eq
# Find the solution for epsilon-1
A = sp.Function('A')(*T_i[1::])
x1_sol = A * sp.exp(sp.I * omega0 * T_i[0]) + sp.conjugate(A) * sp.exp(-sp.I * omega0 * T_i[0])
x1_sol
# Update the epsilon-2 equation
epsilon2_Eq = epsilon2_Eq.subs(x1, x1_sol).doit()
epsilon2_Eq = sp.expand(epsilon2_Eq)
epsilon2_Eq
```
The secular terms will be cancelled out by
$$
\begin{gather*}
D_1 A = 0
\end{gather*}
$$
```
epsilon2_Eq = epsilon2_Eq.subs(sp.diff(A, T_i[1]), 0)
epsilon2_Eq
```
The particular solution of $x_2$ is
$$
\begin{gather*}
x_2 = \frac{\alpha_2 A^2}{3 \omega_0^2} e^{2i\omega_0 T_0} - \frac{\alpha_2 }{\omega^2_0}A\overline{A} + cc
\end{gather*}
$$
```
x2_p = alpha_i[0] * A**2 / 3/omega0**2 * sp.exp(2*sp.I*omega0*T_i[0]) - alpha_i[0]/omega0**2 * A * sp.conjugate(A)
x2_p
epsilon3_Eq = epsilon3_Eq.subs([
(sp.diff(A, T_i[1]), 0), (x1, x1_sol), (x2, x2_p)
]).doit()
epsilon3_Eq = sp.expand(epsilon3_Eq)
epsilon3_Eq = epsilon3_Eq.subs(sp.diff(A, T_i[1], 2), 0)
epsilon3_Eq
```
The to cancel out the secular term we let
$$
\begin{gather*}
2i\omega_0 D_2 A + \dfrac{9\alpha_3 \omega_0^2 - 10\alpha_2^2 }{3\omega_0^2}A^2\overline{A} = 0
\end{gather*}
$$
Question: What if the secular terms arising from $i\omega_0$ and $-i \omega_0$ are handled together - do we get a single real equation?
Substitute the polar $A$
$$
\begin{gather*}
A = \dfrac{1}{2}a e^{i\beta}
\end{gather*}
$$
```
x3_sec = sp.Eq(2*sp.I*omega0*sp.diff(A, T_i[2]) + (9*alpha_i[1]*omega0**2 - 10*alpha_i[0]**2)/3/omega0**2 * A**2 * sp.conjugate(A), 0)
a = sp.Symbol('a', real=True)
beta = sp.Symbol('beta', real=True)
temp = x3_sec.subs(A, a*sp.exp(sp.I * beta)/2)
temp
temp = sp.expand(temp)
temp_im = sp.im(temp.lhs)
temp_im
temp_re = sp.re(temp.lhs)
temp_re
```
Thus separating into real and imaginary parts we obtain
$$
\begin{align*}
\omega_0 D_2 a &=0\\
omega_0 a D_2 \beta + \dfrac{10\alpha_2^2 - 9\alpha_3\omega_0^2}{24\omega_0^2}a^3 &= 0
\end{align*}
$$
$a$ is a constant and
$$
\begin{gather*}
D_2\beta = - \dfrac{10\alpha_2^2 - 9\alpha_3\omega_0^3a}{24\omega_0^2}a^3
\beta = \dfrac{9\alpha_3\omega_0^2 - 10\alpha_2^2 }{24\omega_0^3}a^2 T_2 + \beta_0
\end{gather*}
$$
Here $\beta_0$ is a constant. Now using $T_2 = \epsilon^2 t$ we find that
$$
\begin{gather*}
A = \dfrac{1}{2}a \exp\left[ i\dfrac{9\alpha_3\omega_0^2 - 10\alpha_2^2 }{24\omega_0^3}a^3 \epsilon^2 t + i\beta_0 \right]
\end{gather*}
$$
and substituting in the expressions for $x_1$ and $x_2$ into the equations we have, we obtain the following final results
$$
\begin{gather*}
x = \epsilon a \cos(\omega t + \beta_0) - \dfrac{\epsilon^2 a^2\alpha_2}{2\omega_0^2}\left[ 1 - \dfrac{1}{3}\cos(2\omega t + 2\beta_0) \right] + O(\epsilon^3)
\end{gather*}
$$
where
$$
\begin{gather*}
\omega = \omega_0 \left[ 1 + \dfrac{9\alpha_3 \omega_0^2 - 10\alpha_2^2}{24\omega_0^4}\epsilon^2 a^2 \right] + O(\epsilon^3)
\end{gather*}
$$
| github_jupyter |
**Chapter 2 – End-to-end Machine Learning project**
*Welcome to Machine Learning Housing Corp.! Your task is to predict median house values in Californian districts, given a number of features from these districts.*
*This notebook contains all the sample code and solutions to the exercices in chapter 2.*
# Setup
First, let's make sure this notebook works well in both python 2 and 3, import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures:
```
# To support both python 2 and python 3
from __future__ import division, print_function, unicode_literals
# Common imports
import numpy as np
import numpy.random as rnd
import os
# to make this notebook's output stable across runs
rnd.seed(42)
# To plot pretty figures
%matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['xtick.labelsize'] = 12
plt.rcParams['ytick.labelsize'] = 12
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "end_to_end_project"
def save_fig(fig_id, tight_layout=True):
path = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID, fig_id + ".png")
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format='png', dpi=300)
```
# Get the data
```
DOW = "https://raw.githubusercontent.com/ageron/handson-ml/master/"
import os
import tarfile
from six.moves import urllib
HOUSING_PATH = "datasets/housing/"
HOUSING_URL = DOWNLOAD_ROOT + HOUSING_PATH
def fetch_housing_data(housing_url=HOUSING_URL, housing_path=HOUSING_PATH):
if not os.path.exists(housing_path):
os.makedirs(housing_path)
tgz_path = os.path.join(housing_path, "housing.tgz")
urllib.request.urlretrieve(housing_url, tgz_path)
housing_tgz = tarfile.open(tgz_path)
housing_tgz.extractall(path=housing_path)
housing_tgz.close()
fetch_housing_data()
import pandas as pd
def load_housing_data(housing_path=HOUSING_PATH):
csv_path = os.path.join(housing_path, "housing.csv")
return pd.read_csv(csv_path)
housing = load_housing_data()
housing.head()
housing.info()
housing["ocean_proximity"].value_counts()
print(housing.describe())
%matplotlib inline
import matplotlib.pyplot as plt
housing.hist(bins=50, figsize=(11,8))
save_fig("attribute_histogram_plots")
plt.show()
import numpy as np
import numpy.random as rnd
rnd.seed(42) # to make this notebook's output identical at every run
def split_train_test(data, test_ratio):
shuffled_indices = rnd.permutation(len(data))
test_set_size = int(len(data) * test_ratio)
test_indices = shuffled_indices[:test_set_size]
train_indices = shuffled_indices[test_set_size:]
return data.iloc[train_indices], data.iloc[test_indices]
train_set, test_set = split_train_test(housing, 0.2)
print(len(train_set), len(test_set))
import hashlib
def test_set_check(identifier, test_ratio, hash):
return bytearray(hash(np.int64(identifier)).digest())[-1] < 256 * test_ratio
def split_train_test_by_id(data, test_ratio, id_column, hash=hashlib.md5):
ids = data[id_column]
in_test_set = ids.apply(lambda id_: test_set_check(id_, test_ratio, hash))
return data.loc[~in_test_set], data.loc[in_test_set]
housing_with_id = housing.reset_index() # adds an `index` column
train_set, test_set = split_train_test_by_id(housing_with_id, 0.2, "index")
test_set.head()
from sklearn.model_selection import train_test_split
train_set, test_set = train_test_split(housing, test_size=0.2, random_state=42)
test_set.head()
housing["median_income"].hist()
housing["income_cat"] = np.ceil(housing["median_income"] / 1.5)
housing["income_cat"].where(housing["income_cat"] < 5, 5.0, inplace=True)
housing["income_cat"].value_counts()
from sklearn.model_selection import StratifiedShuffleSplit
split = StratifiedShuffleSplit(n_splits=1, test_size=0.2, random_state=42)
for train_index, test_index in split.split(housing, housing["income_cat"]):
strat_train_set = housing.loc[train_index]
strat_test_set = housing.loc[test_index]
def income_cat_proportions(data):
return data["income_cat"].value_counts() / len(data)
train_set, test_set = train_test_split(housing, test_size=0.2, random_state=42)
compare_props = pd.DataFrame({
"Overall": income_cat_proportions(housing),
"Stratified": income_cat_proportions(strat_test_set),
"Random": income_cat_proportions(test_set),
}).sort_index()
compare_props["Rand. %error"] = 100 * compare_props["Random"] / compare_props["Overall"] - 100
compare_props["Strat. %error"] = 100 * compare_props["Stratified"] / compare_props["Overall"] - 100
compare_props
for set in (strat_train_set, strat_test_set):
set.drop("income_cat", axis=1, inplace=True)
```
# Discover and visualize the data to gain insights
```
housing = strat_train_set.copy()
housing.plot(kind="scatter", x="longitude", y="latitude")
save_fig("bad_visualization_plot")
housing.plot(kind="scatter", x="longitude", y="latitude", alpha=0.1)
save_fig("better_visualization_plot")
housing.plot(kind="scatter", x="longitude", y="latitude",
s=housing['population']/100, label="population",
c="median_house_value", cmap=plt.get_cmap("jet"),
colorbar=True, alpha=0.4, figsize=(10,7),
)
plt.legend()
save_fig("housing_prices_scatterplot")
plt.show()
import matplotlib.image as mpimg
california_img=mpimg.imread(PROJECT_ROOT_DIR + '/images/end_to_end_project/california.png')
ax = housing.plot(kind="scatter", x="longitude", y="latitude", figsize=(10,7),
s=housing['population']/100, label="Population",
c="median_house_value", cmap=plt.get_cmap("jet"),
colorbar=False, alpha=0.4,
)
plt.imshow(california_img, extent=[-124.55, -113.80, 32.45, 42.05], alpha=0.5)
plt.ylabel("Latitude", fontsize=14)
plt.xlabel("Longitude", fontsize=14)
prices = housing["median_house_value"]
tick_values = np.linspace(prices.min(), prices.max(), 11)
cbar = plt.colorbar()
cbar.ax.set_yticklabels(["$%dk"%(round(v/1000)) for v in tick_values], fontsize=14)
cbar.set_label('Median House Value', fontsize=16)
plt.legend(fontsize=16)
save_fig("california_housing_prices_plot")
plt.show()
corr_matrix = housing.corr()
corr_matrix["median_house_value"].sort_values(ascending=False)
housing.plot(kind="scatter", x="median_income", y="median_house_value",
alpha=0.3)
plt.axis([0, 16, 0, 550000])
save_fig("income_vs_house_value_scatterplot")
plt.show()
from pandas.tools.plotting import scatter_matrix
attributes = ["median_house_value", "median_income", "total_rooms", "housing_median_age"]
scatter_matrix(housing[attributes], figsize=(11, 8))
save_fig("scatter_matrix_plot")
plt.show()
housing["rooms_per_household"] = housing["total_rooms"] / housing["population"]
housing["bedrooms_per_room"] = housing["total_bedrooms"] / housing["total_rooms"]
housing["population_per_household"] = housing["population"] / housing["households"]
corr_matrix = housing.corr()
corr_matrix["median_house_value"].sort_values(ascending=False)
housing.plot(kind="scatter", x="rooms_per_household", y="median_house_value",
alpha=0.2)
plt.axis([0, 5, 0, 520000])
plt.show()
housing.describe()
```
# Prepare the data for Machine Learning algorithms
```
housing = strat_train_set.drop("median_house_value", axis=1)
housing_labels = strat_train_set["median_house_value"].copy()
housing_copy = housing.copy().iloc[21:24]
housing_copy
housing_copy.dropna(subset=["total_bedrooms"]) # option 1
housing_copy = housing.copy().iloc[21:24]
housing_copy.drop("total_bedrooms", axis=1) # option 2
housing_copy = housing.copy().iloc[21:24]
median = housing_copy["total_bedrooms"].median()
housing_copy["total_bedrooms"].fillna(median, inplace=True) # option 3
housing_copy
from sklearn.preprocessing import Imputer
imputer = Imputer(strategy='median')
housing_num = housing.drop("ocean_proximity", axis=1)
imputer.fit(housing_num)
X = imputer.transform(housing_num)
housing_tr = pd.DataFrame(X, columns=housing_num.columns)
housing_tr.iloc[21:24]
imputer.statistics_
housing_num.median().values
imputer.strategy
housing_tr = pd.DataFrame(X, columns=housing_num.columns)
housing_tr.head()
from sklearn.preprocessing import LabelEncoder
encoder = LabelEncoder()
housing_cat = housing["ocean_proximity"]
housing_cat_encoded = encoder.fit_transform(housing_cat)
housing_cat_encoded
print(encoder.classes_)
from sklearn.preprocessing import OneHotEncoder
encoder = OneHotEncoder()
housing_cat_1hot = encoder.fit_transform(housing_cat_encoded.reshape(-1,1))
housing_cat_1hot
housing_cat_1hot.toarray()
from sklearn.preprocessing import LabelBinarizer
encoder = LabelBinarizer()
encoder.fit_transform(housing_cat)
from sklearn.base import BaseEstimator, TransformerMixin
rooms_ix, bedrooms_ix, population_ix, household_ix = 3, 4, 5, 6
class CombinedAttributesAdder(BaseEstimator, TransformerMixin):
def __init__(self, add_bedrooms_per_room = True): # no *args or **kargs
self.add_bedrooms_per_room = add_bedrooms_per_room
def fit(self, X, y=None):
return self # nothing else to do
def transform(self, X, y=None):
rooms_per_household = X[:, rooms_ix] / X[:, household_ix]
population_per_household = X[:, population_ix] / X[:, household_ix]
if self.add_bedrooms_per_room:
bedrooms_per_room = X[:, bedrooms_ix] / X[:, rooms_ix]
return np.c_[X, rooms_per_household, population_per_household, bedrooms_per_room]
else:
return np.c_[X, rooms_per_household, population_per_household]
attr_adder = CombinedAttributesAdder(add_bedrooms_per_room=False)
housing_extra_attribs = attr_adder.transform(housing.values)
housing_extra_attribs = pd.DataFrame(housing_extra_attribs, columns=list(housing.columns)+["rooms_per_household", "population_per_household"])
housing_extra_attribs.head()
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
num_pipeline = Pipeline([
('imputer', Imputer(strategy="median")),
('attribs_adder', CombinedAttributesAdder()),
('std_scaler', StandardScaler()),
])
num_pipeline.fit_transform(housing_num)
from sklearn.pipeline import FeatureUnion
class DataFrameSelector(BaseEstimator, TransformerMixin):
def __init__(self, attribute_names):
self.attribute_names = attribute_names
def fit(self, X, y=None):
return self
def transform(self, X):
return X[self.attribute_names].values
num_attribs = list(housing_num)
cat_attribs = ["ocean_proximity"]
num_pipeline = Pipeline([
('selector', DataFrameSelector(num_attribs)),
('imputer', Imputer(strategy="median")),
('attribs_adder', CombinedAttributesAdder()),
('std_scaler', StandardScaler()),
])
cat_pipeline = Pipeline([
('selector', DataFrameSelector(cat_attribs)),
('label_binarizer', LabelBinarizer()),
])
preparation_pipeline = FeatureUnion(transformer_list=[
("num_pipeline", num_pipeline),
("cat_pipeline", cat_pipeline),
])
housing_prepared = preparation_pipeline.fit_transform(housing)
housing_prepared
housing_prepared.shape
```
# Prepare the data for Machine Learning algorithms
```
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression()
lin_reg.fit(housing_prepared, housing_labels)
# let's try the full pipeline on a few training instances
some_data = housing.iloc[:5]
some_labels = housing_labels.iloc[:5]
some_data_prepared = preparation_pipeline.transform(some_data)
print("Predictions:\t", lin_reg.predict(some_data_prepared))
print("Labels:\t\t", list(some_labels))
from sklearn.metrics import mean_squared_error
housing_predictions = lin_reg.predict(housing_prepared)
lin_mse = mean_squared_error(housing_labels, housing_predictions)
lin_rmse = np.sqrt(lin_mse)
lin_rmse
from sklearn.metrics import mean_absolute_error
lin_mae = mean_absolute_error(housing_labels, housing_predictions)
lin_mae
from sklearn.tree import DecisionTreeRegressor
tree_reg = DecisionTreeRegressor()
tree_reg.fit(housing_prepared, housing_labels)
housing_predictions = tree_reg.predict(housing_prepared)
tree_mse = mean_squared_error(housing_labels, housing_predictions)
tree_rmse = np.sqrt(tree_mse)
tree_rmse
```
# Fine-tune your model
```
from sklearn.model_selection import cross_val_score
tree_scores = cross_val_score(tree_reg, housing_prepared, housing_labels,
scoring="neg_mean_squared_error", cv=10)
tree_rmse_scores = np.sqrt(-tree_scores)
def display_scores(scores):
print("Scores:", scores)
print("Mean:", scores.mean())
print("Standard deviation:", scores.std())
display_scores(tree_rmse_scores)
lin_scores = cross_val_score(lin_reg, housing_prepared, housing_labels,
scoring="neg_mean_squared_error", cv=10)
lin_rmse_scores = np.sqrt(-lin_scores)
display_scores(lin_rmse_scores)
from sklearn.ensemble import RandomForestRegressor
forest_reg = RandomForestRegressor()
forest_reg.fit(housing_prepared, housing_labels)
housing_predictions = forest_reg.predict(housing_prepared)
forest_mse = mean_squared_error(housing_labels, housing_predictions)
forest_rmse = np.sqrt(forest_mse)
forest_rmse
from sklearn.model_selection import cross_val_score
forest_scores = cross_val_score(forest_reg, housing_prepared, housing_labels,
scoring="neg_mean_squared_error", cv=10)
forest_rmse_scores = np.sqrt(-forest_scores)
display_scores(forest_rmse_scores)
scores = cross_val_score(lin_reg, housing_prepared, housing_labels, scoring="neg_mean_squared_error", cv=10)
pd.Series(np.sqrt(-scores)).describe()
from sklearn.svm import SVR
svm_reg = SVR(kernel="linear")
svm_reg.fit(housing_prepared, housing_labels)
housing_predictions = svm_reg.predict(housing_prepared)
svm_mse = mean_squared_error(housing_labels, housing_predictions)
svm_rmse = np.sqrt(svm_mse)
svm_rmse
from sklearn.model_selection import GridSearchCV
param_grid = [
{'n_estimators': [3, 10, 30], 'max_features': [2, 4, 6, 8]},
{'bootstrap': [False], 'n_estimators': [3, 10], 'max_features': [2, 3, 4]},
]
forest_reg = RandomForestRegressor()
grid_search = GridSearchCV(forest_reg, param_grid, cv=5, scoring='neg_mean_squared_error')
grid_search.fit(housing_prepared, housing_labels)
grid_search.best_params_
grid_search.best_estimator_
cvres = grid_search.cv_results_
for mean_score, params in zip(cvres["mean_test_score"], cvres["params"]):
print(np.sqrt(-mean_score), params)
pd.DataFrame(grid_search.cv_results_)
from sklearn.model_selection import RandomizedSearchCV
from scipy.stats import randint
param_distribs = {
'n_estimators': randint(low=1, high=200),
'max_features': randint(low=1, high=8),
}
forest_reg = RandomForestRegressor()
rnd_search = RandomizedSearchCV(forest_reg, param_distributions=param_distribs,
n_iter=10, cv=5, scoring='neg_mean_squared_error')
rnd_search.fit(housing_prepared, housing_labels)
cvres = rnd_search.cv_results_
for mean_score, params in zip(cvres["mean_test_score"], cvres["params"]):
print(np.sqrt(-mean_score), params)
feature_importances = grid_search.best_estimator_.feature_importances_
feature_importances
extra_attribs = ["rooms_per_household", "population_per_household", "bedrooms_per_room"]
cat_one_hot_attribs = list(encoder.classes_)
attributes = num_attribs + extra_attribs + cat_one_hot_attribs
sorted(zip(feature_importances, attributes), reverse=True)
final_model = grid_search.best_estimator_
X_test = strat_test_set.drop("median_house_value", axis=1)
y_test = strat_test_set["median_house_value"].copy()
X_test_transformed = preparation_pipeline.transform(X_test)
final_predictions = final_model.predict(X_test_transformed)
final_mse = mean_squared_error(y_test, final_predictions)
final_rmse = np.sqrt(final_mse)
final_rmse
```
# Extra material
## Label Binarizer hack
`LabelBinarizer`'s `fit_transform()` method only accepts one parameter `y` (because it was meant for labels, not predictors), so it does not work in a pipeline where the final estimator is a supervised estimator because in this case its `fit()` method takes two parameters `X` and `y`.
This hack creates a supervision-friendly `LabelBinarizer`.
```
class SupervisionFriendlyLabelBinarizer(LabelBinarizer):
def fit_transform(self, X, y=None):
return super(SupervisionFriendlyLabelBinarizer, self).fit_transform(X)
# Replace the Labelbinarizer with a SupervisionFriendlyLabelBinarizer
cat_pipeline.steps[1] = ("label_binarizer", SupervisionFriendlyLabelBinarizer())
# Now you can create a full pipeline with a supervised predictor at the end.
full_pipeline = Pipeline([
("preparation", preparation_pipeline),
("linear", LinearRegression())
])
full_pipeline.fit(housing, housing_labels)
full_pipeline.predict(some_data)
```
## Model persistence using joblib
```
from sklearn.externals import joblib
joblib.dump(final_model, "my_random_forest_regressor.pkl")
final_model_loaded = joblib.load("my_random_forest_regressor.pkl")
final_model_loaded
```
## Example SciPy distributions for `RandomizedSearchCV`
```
from scipy.stats import geom, expon
geom_distrib=geom(0.5).rvs(10000)
expon_distrib=expon(scale=1).rvs(10000)
plt.hist(geom_distrib, bins=50)
plt.show()
plt.hist(expon_distrib, bins=50)
plt.show()
```
# Exercise solutions
**Coming soon**
| github_jupyter |
# Complex Graphs Metadata Example
## Prerequisites
* A kubernetes cluster with kubectl configured
* curl
* pygmentize
## Setup Seldon Core
Use the setup notebook to [Setup Cluster](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html) to setup Seldon Core with an ingress.
```
!kubectl create namespace seldon
!kubectl config set-context $(kubectl config current-context) --namespace=seldon
```
## Used model
In this example notebook we will use a dummy node that can serve multiple purposes in the graph.
The model will read its metadata from environmental variable (this is done automatically).
Actual logic that happens on each of this endpoint is not subject of this notebook.
We will only concentrate on graph-level metadata that orchestrator constructs from metadata reported by each node.
```
%%writefile models/generic-node/Node.py
import logging
import random
import os
NUMBER_OF_ROUTES = int(os.environ.get("NUMBER_OF_ROUTES", "2"))
class Node:
def predict(self, features, names=[], meta=[]):
logging.info(f"model features: {features}")
logging.info(f"model names: {names}")
logging.info(f"model meta: {meta}")
return features.tolist()
def transform_input(self, features, names=[], meta=[]):
return self.predict(features, names, meta)
def transform_output(self, features, names=[], meta=[]):
return self.predict(features, names, meta)
def aggregate(self, features, names=[], meta=[]):
logging.info(f"model features: {features}")
logging.info(f"model names: {names}")
logging.info(f"model meta: {meta}")
return [x.tolist() for x in features]
def route(self, features, names=[], meta=[]):
logging.info(f"model features: {features}")
logging.info(f"model names: {names}")
logging.info(f"model meta: {meta}")
route = random.randint(0, NUMBER_OF_ROUTES)
logging.info(f"routing to: {route}")
return route
```
### Build image
build image using provided Makefile
```
cd models/generic-node
make build
```
If you are using `kind` you can use `kind_image_install` target to directly
load your image into your local cluster.
## Single Model
In case of single-node graph model-level `inputs` and `outputs`, `x` and `y`, will simply be also the deployment-level `graphinputs` and `graphoutputs`.

```
%%writefile graph-metadata/single.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: graph-metadata-single
spec:
name: test-deployment
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/metadata-generic-node:0.4
name: model
env:
- name: MODEL_METADATA
value: |
---
name: single-node
versions: [ generic-node/v0.4 ]
platform: seldon
inputs:
- messagetype: tensor
schema:
names: [node-input]
shape: [ 1 ]
outputs:
- messagetype: tensor
schema:
names: [node-output]
shape: [ 1 ]
graph:
name: model
type: MODEL
children: []
name: example
replicas: 1
!kubectl apply -f graph-metadata/single.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=graph-metadata-single -o jsonpath='{.items[0].metadata.name}')
```
### Graph Level
Graph level metadata is available at the `api/v1.0/metadata` endpoint of your deployment:
```
import requests
import time
def getWithRetry(url):
for i in range(3):
r = requests.get(url)
if r.status_code == requests.codes.ok:
meta = r.json()
return meta
else:
print("Failed request with status code ",r.status_code)
time.sleep(3)
meta = getWithRetry("http://localhost:8003/seldon/seldon/graph-metadata-single/api/v1.0/metadata")
assert meta == {
"name": "example",
"models": {
"model": {
"name": "single-node",
"platform": "seldon",
"versions": ["generic-node/v0.4"],
"inputs": [
{"messagetype": "tensor", "schema": {"names": ["node-input"], "shape": [1]}}
],
"outputs": [
{"messagetype": "tensor", "schema": {"names": ["node-output"], "shape": [1]}}
],
}
},
"graphinputs": [
{"messagetype": "tensor", "schema": {"names": ["node-input"], "shape": [1]}}
],
"graphoutputs": [
{"messagetype": "tensor", "schema": {"names": ["node-output"], "shape": [1]}}
],
}
meta
```
### Model Level
Compare with `model` metadata available at the `api/v1.0/metadata/model`:
```
import requests
meta = getWithRetry("http://localhost:8003/seldon/seldon/graph-metadata-single/api/v1.0/metadata/model")
assert meta == {
"custom": {},
"name": "single-node",
"platform": "seldon",
"versions": ["generic-node/v0.4"],
"inputs": [{
"messagetype": "tensor", "schema": {"names": ["node-input"], "shape": [1]},
}],
"outputs": [{
"messagetype": "tensor", "schema": {"names": ["node-output"], "shape": [1]},
}],
}
meta
!kubectl delete -f graph-metadata/single.yaml
```
## Two-Level Graph
In two-level graph graph output of the first model is input of the second model, `x2=y1`.
The graph-level input `x` will be first model’s input `x1` and graph-level output `y` will be the last model’s output `y2`.

```
%%writefile graph-metadata/two-levels.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: graph-metadata-two-levels
spec:
name: test-deployment
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/metadata-generic-node:0.4
name: node-one
env:
- name: MODEL_METADATA
value: |
---
name: node-one
versions: [ generic-node/v0.4 ]
platform: seldon
inputs:
- messagetype: tensor
schema:
names: [ a1, a2 ]
shape: [ 2 ]
outputs:
- messagetype: tensor
schema:
names: [ a3 ]
shape: [ 1 ]
- image: seldonio/metadata-generic-node:0.4
name: node-two
env:
- name: MODEL_METADATA
value: |
---
name: node-two
versions: [ generic-node/v0.4 ]
platform: seldon
inputs:
- messagetype: tensor
schema:
names: [ a3 ]
shape: [ 1 ]
outputs:
- messagetype: tensor
schema:
names: [b1, b2]
shape: [ 2 ]
graph:
name: node-one
type: MODEL
children:
- name: node-two
type: MODEL
children: []
name: example
replicas: 1
!kubectl apply -f graph-metadata/two-levels.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=graph-metadata-two-levels -o jsonpath='{.items[0].metadata.name}')
import requests
meta = getWithRetry("http://localhost:8003/seldon/seldon/graph-metadata-two-levels/api/v1.0/metadata")
assert meta == {
"name": "example",
"models": {
"node-one": {
"name": "node-one",
"platform": "seldon",
"versions": ["generic-node/v0.4"],
"inputs": [
{"messagetype": "tensor", "schema": {"names": ["a1", "a2"], "shape": [2]}}
],
"outputs": [
{"messagetype": "tensor", "schema": {"names": ["a3"], "shape": [1]}}
],
},
"node-two": {
"name": "node-two",
"platform": "seldon",
"versions": ["generic-node/v0.4"],
"inputs": [
{"messagetype": "tensor", "schema": {"names": ["a3"], "shape": [1]}}
],
"outputs": [
{"messagetype": "tensor", "schema": {"names": ["b1", "b2"], "shape": [2]}}
],
}
},
"graphinputs": [
{"messagetype": "tensor", "schema": {"names": ["a1", "a2"], "shape": [2]}}
],
"graphoutputs": [
{"messagetype": "tensor", "schema": {"names": ["b1", "b2"], "shape": [2]}}
],
}
meta
!kubectl delete -f graph-metadata/two-levels.yaml
```
## Combiner of two models
In graph with the `combiner` request is first passed to combiner's children and before it gets aggregated by the `combiner` itself.
Input `x` is first passed to both models and their outputs `y1` and `y2` are passed to the combiner.
Combiner's output `y` is the final output of the graph.

```
%%writefile graph-metadata/combiner.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: graph-metadata-combiner
spec:
name: test-deployment
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/metadata-generic-node:0.4
name: node-combiner
env:
- name: MODEL_METADATA
value: |
---
name: node-combiner
versions: [ generic-node/v0.4 ]
platform: seldon
inputs:
- messagetype: tensor
schema:
names: [ c1 ]
shape: [ 1 ]
- messagetype: tensor
schema:
names: [ c2 ]
shape: [ 1 ]
outputs:
- messagetype: tensor
schema:
names: [combiner-output]
shape: [ 1 ]
- image: seldonio/metadata-generic-node:0.4
name: node-one
env:
- name: MODEL_METADATA
value: |
---
name: node-one
versions: [ generic-node/v0.4 ]
platform: seldon
inputs:
- messagetype: tensor
schema:
names: [a, b]
shape: [ 2 ]
outputs:
- messagetype: tensor
schema:
names: [ c1 ]
shape: [ 1 ]
- image: seldonio/metadata-generic-node:0.4
name: node-two
env:
- name: MODEL_METADATA
value: |
---
name: node-two
versions: [ generic-node/v0.4 ]
platform: seldon
inputs:
- messagetype: tensor
schema:
names: [a, b]
shape: [ 2 ]
outputs:
- messagetype: tensor
schema:
names: [ c2 ]
shape: [ 1 ]
graph:
name: node-combiner
type: COMBINER
children:
- name: node-one
type: MODEL
children: []
- name: node-two
type: MODEL
children: []
name: example
replicas: 1
!kubectl apply -f graph-metadata/combiner.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=graph-metadata-combiner -o jsonpath='{.items[0].metadata.name}')
import requests
meta = getWithRetry("http://localhost:8003/seldon/seldon/graph-metadata-combiner/api/v1.0/metadata")
assert meta == {
"name": "example",
"models": {
"node-combiner": {
"name": "node-combiner",
"platform": "seldon",
"versions": ["generic-node/v0.4"],
"inputs": [
{"messagetype": "tensor", "schema": {"names": ["c1"], "shape": [1]}},
{"messagetype": "tensor", "schema": {"names": ["c2"], "shape": [1]}},
],
"outputs": [
{"messagetype": "tensor", "schema": {"names": ["combiner-output"], "shape": [1]}}
],
},
"node-one": {
"name": "node-one",
"platform": "seldon",
"versions": ["generic-node/v0.4"],
"inputs": [
{"messagetype": "tensor", "schema": {"names": ["a", "b"], "shape": [2]}},
],
"outputs": [
{"messagetype": "tensor", "schema": {"names": ["c1"], "shape": [1]}}
],
},
"node-two": {
"name": "node-two",
"platform": "seldon",
"versions": ["generic-node/v0.4"],
"inputs": [
{"messagetype": "tensor", "schema": {"names": ["a", "b"], "shape": [2]}},
],
"outputs": [
{"messagetype": "tensor", "schema": {"names": ["c2"], "shape": [1]}}
],
}
},
"graphinputs": [
{"messagetype": "tensor", "schema": {"names": ["a", "b"], "shape": [2]}},
],
"graphoutputs": [
{"messagetype": "tensor", "schema": {"names": ["combiner-output"], "shape": [1]}}
],
}
meta
!kubectl delete -f graph-metadata/combiner.yaml
```
## Router with two models
In this example request `x` is passed by `router` to one of its children.
Router then returns children output `y1` or `y2` as graph's output `y`.
Here we assume that all children accepts similarly structured input and retun a similarly structured output.

```
%%writefile graph-metadata/router.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: graph-metadata-router
spec:
name: test-deployment
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/metadata-generic-node:0.4
name: node-router
- image: seldonio/metadata-generic-node:0.4
name: node-one
env:
- name: MODEL_METADATA
value: |
---
name: node-one
versions: [ generic-node/v0.4 ]
platform: seldon
inputs:
- messagetype: tensor
schema:
names: [ a, b ]
shape: [ 2 ]
outputs:
- messagetype: tensor
schema:
names: [ node-output ]
shape: [ 1 ]
- image: seldonio/metadata-generic-node:0.4
name: node-two
env:
- name: MODEL_METADATA
value: |
---
name: node-two
versions: [ generic-node/v0.4 ]
platform: seldon
inputs:
- messagetype: tensor
schema:
names: [ a, b ]
shape: [ 2 ]
outputs:
- messagetype: tensor
schema:
names: [ node-output ]
shape: [ 1 ]
graph:
name: node-router
type: ROUTER
children:
- name: node-one
type: MODEL
children: []
- name: node-two
type: MODEL
children: []
name: example
replicas: 1
!kubectl apply -f graph-metadata/router.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=graph-metadata-router -o jsonpath='{.items[0].metadata.name}')
import requests
meta = getWithRetry("http://localhost:8003/seldon/seldon/graph-metadata-router/api/v1.0/metadata")
assert meta == {
"name": "example",
"models": {
'node-router': {
'name': 'seldonio/metadata-generic-node',
'versions': ['0.4'],
'inputs': [],
'outputs': [],
},
"node-one": {
"name": "node-one",
"platform": "seldon",
"versions": ["generic-node/v0.4"],
"inputs": [
{"messagetype": "tensor", "schema": {"names": ["a", "b"], "shape": [2]}}
],
"outputs": [
{"messagetype": "tensor", "schema": {"names": ["node-output"], "shape": [1]}}
],
},
"node-two": {
"name": "node-two",
"platform": "seldon",
"versions": ["generic-node/v0.4"],
"inputs": [
{"messagetype": "tensor", "schema": {"names": ["a", "b"], "shape": [2]}}
],
"outputs": [
{"messagetype": "tensor", "schema": {"names": ["node-output"], "shape": [1]}}
],
}
},
"graphinputs": [
{"messagetype": "tensor", "schema": {"names": ["a", "b"], "shape": [2]}}
],
"graphoutputs": [
{"messagetype": "tensor", "schema": {"names": ["node-output"], "shape": [1]}}
],
}
meta
!kubectl delete -f graph-metadata/router.yaml
```
## Input Transformer
Input transformers work almost exactly the same as chained nodes, see two-level example above.
Following graph is presented in a way that is suppose to make next example (output transfomer) more intuitive.

```
%%writefile graph-metadata/input-transformer.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: graph-metadata-input
spec:
name: test-deployment
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/metadata-generic-node:0.4
name: node-input-transformer
env:
- name: MODEL_METADATA
value: |
---
name: node-input-transformer
versions: [ generic-node/v0.4 ]
platform: seldon
inputs:
- messagetype: tensor
schema:
names: [transformer-input]
shape: [ 1 ]
outputs:
- messagetype: tensor
schema:
names: [transformer-output]
shape: [ 1 ]
- image: seldonio/metadata-generic-node:0.4
name: node
env:
- name: MODEL_METADATA
value: |
---
name: node
versions: [ generic-node/v0.4 ]
platform: seldon
inputs:
- messagetype: tensor
schema:
names: [transformer-output]
shape: [ 1 ]
outputs:
- messagetype: tensor
schema:
names: [node-output]
shape: [ 1 ]
graph:
name: node-input-transformer
type: TRANSFORMER
children:
- name: node
type: MODEL
children: []
name: example
replicas: 1
!kubectl apply -f graph-metadata/input-transformer.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=graph-metadata-input -o jsonpath='{.items[0].metadata.name}')
import requests
meta = getWithRetry("http://localhost:8003/seldon/seldon/graph-metadata-input/api/v1.0/metadata")
assert meta == {
"name": "example",
"models": {
"node-input-transformer": {
"name": "node-input-transformer",
"platform": "seldon",
"versions": ["generic-node/v0.4"],
"inputs": [{
"messagetype": "tensor", "schema": {"names": ["transformer-input"], "shape": [1]},
}],
"outputs": [{
"messagetype": "tensor", "schema": {"names": ["transformer-output"], "shape": [1]},
}],
},
"node": {
"name": "node",
"platform": "seldon",
"versions": ["generic-node/v0.4"],
"inputs": [{
"messagetype": "tensor", "schema": {"names": ["transformer-output"], "shape": [1]},
}],
"outputs": [{
"messagetype": "tensor", "schema": {"names": ["node-output"], "shape": [1]},
}],
}
},
"graphinputs": [{
"messagetype": "tensor", "schema": {"names": ["transformer-input"], "shape": [1]}
}],
"graphoutputs": [{
"messagetype": "tensor", "schema": {"names": ["node-output"], "shape": [1]}
}],
}
meta
!kubectl delete -f graph-metadata/input-transformer.yaml
```
## Output Transformer
Output transformers work almost exactly opposite as chained nodes in the two-level example above.
Input `x` is first passed to the model that is child of the `output-transformer` before it is passed to it.

```
%%writefile graph-metadata/output-transformer.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: graph-metadata-output
spec:
name: test-deployment
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/metadata-generic-node:0.4
name: node-output-transformer
env:
- name: MODEL_METADATA
value: |
---
name: node-output-transformer
versions: [ generic-node/v0.4 ]
platform: seldon
inputs:
- messagetype: tensor
schema:
names: [transformer-input]
shape: [ 1 ]
outputs:
- messagetype: tensor
schema:
names: [transformer-output]
shape: [ 1 ]
- image: seldonio/metadata-generic-node:0.4
name: node
env:
- name: MODEL_METADATA
value: |
---
name: node
versions: [ generic-node/v0.4 ]
platform: seldon
inputs:
- messagetype: tensor
schema:
names: [node-input]
shape: [ 1 ]
outputs:
- messagetype: tensor
schema:
names: [transformer-input]
shape: [ 1 ]
graph:
name: node-output-transformer
type: OUTPUT_TRANSFORMER
children:
- name: node
type: MODEL
children: []
name: example
replicas: 1
!kubectl apply -f graph-metadata/output-transformer.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=graph-metadata-output -o jsonpath='{.items[0].metadata.name}')
import requests
meta = getWithRetry("http://localhost:8003/seldon/seldon/graph-metadata-output/api/v1.0/metadata")
assert meta == {
"name": "example",
"models": {
"node-output-transformer": {
"name": "node-output-transformer",
"platform": "seldon",
"versions": ["generic-node/v0.4"],
"inputs": [{
"messagetype": "tensor", "schema": {"names": ["transformer-input"], "shape": [1]},
}],
"outputs": [{
"messagetype": "tensor", "schema": {"names": ["transformer-output"], "shape": [1]},
}],
},
"node": {
"name": "node",
"platform": "seldon",
"versions": ["generic-node/v0.4"],
"inputs": [{
"messagetype": "tensor", "schema": {"names": ["node-input"], "shape": [1]},
}],
"outputs": [{
"messagetype": "tensor", "schema": {"names": ["transformer-input"], "shape": [1]},
}],
}
},
"graphinputs": [{
"messagetype": "tensor", "schema": {"names": ["node-input"], "shape": [1]}
}],
"graphoutputs": [{
"messagetype": "tensor", "schema": {"names": ["transformer-output"], "shape": [1]}
}],
}
meta
!kubectl delete -f graph-metadata/output-transformer.yaml
```
| github_jupyter |
# INFO
This is my solution for the fourth homework problem.
# **SOLUTION**
# Description
I will use network with:
- input layer with **2 neurons** (two input variables)
- **one** hidden layer with **2 neurons** (I need to split the plane in a nonlinear way, creating a U-shaped plane containing the diagonal points)
- output layer with 1 neuron (result - active or inactive)
Also, as an activation function I will use a sigmoid function - simple, values between (0, 1) and with simple derivative
# CODE
```
import numpy as np
```
Let's define our sigmoid function and its derivative
```
def sigmoid(x, derivative=False):
if derivative:
return x * (1 - x)
else:
return 1 / (1 + np.exp(-x))
```
Now, number of neurons per layer
```
layers_sizes = np.array([2, 2, 1])
```
And layers initialization function
```
def init_layers(sizes):
weights = [np.random.uniform(size=size) for size in zip(sizes[0:-1], sizes[1:])]
biases = [np.random.uniform(size=(size, 1)) for size in sizes[1:]]
return weights, biases
```
Function which execute network (forward propagation).
Takes input layer, following layers weights and biases and activation function. Returns layers outputs.
```
def execute(input, weights, biases, activation_f):
result = [input]
previous_layer = input
for weight, bias in zip(weights, biases):
executed_layer = execute_layer(previous_layer, weight, bias, activation_f)
previous_layer = executed_layer
result.append(executed_layer)
return result
def execute_layer(input_layer, weight, bias, activation_f):
layer_activation = np.dot(input_layer.T, weight).T + bias
return activation_f(layer_activation)
```
And time for the backpropagation function.
Function takes layers outputs, weights, biases and activation function, expected output and learning rate.
```
def backpropagation(layers_outputs,
weights,
biases,
activation_f,
expected_output,
learning_rate):
updated_weights = weights.copy()
updated_biases = biases.copy()
predicted_output = layers_outputs[-1]
output_error = 2 * (expected_output - predicted_output)
output_delta = output_error * activation_f(predicted_output, True)
updated_weights[-1] += layers_outputs[-2].dot(output_delta.T) * learning_rate
updated_biases[-1] += output_delta * learning_rate
next_layer_delta = output_delta
for layer_id in reversed(range(1, len(layers_outputs)-1)):
weight_id = layer_id - 1
error = np.dot(weights[weight_id+1], next_layer_delta)
delta = error * activation_f(layers_outputs[layer_id], True)
updated_weights[weight_id] += layers_outputs[layer_id-1].dot(delta.T) * learning_rate
updated_biases[weight_id] += delta * learning_rate
next_layer_delta = delta
return updated_weights, updated_biases
```
---
Create test set:
```
test_set_X = [np.array([[0], [0]]), np.array([[1], [0]]), np.array([[0], [1]]), np.array([[1], [1]])]
test_set_Y = [np.array([[0]]), np.array([[1]]), np.array([[1]]), np.array([[0]])]
```
And training parameters:
```
learning_rate = 0.07
number_of_iterations = 30000
```
And train out model:
```
weights, biases = init_layers(layers_sizes)
errors = []
for iteration in range(number_of_iterations):
error = 0
for test_x, test_y in zip(test_set_X, test_set_Y):
values = execute(test_x, weights, biases, sigmoid)
predicted_y = values[-1]
error += np.sum((predicted_y - test_y) ** 2) / len(test_y)
new_weights, new_biases = backpropagation(values, weights, biases, sigmoid, test_y, learning_rate)
weights = new_weights
biases = new_biases
print("iteration number {} done! Error: {}".format(iteration, error / len(test_set_X)))
errors.append(error / len(test_set_X))
```
And plot the error over iterations
```
import matplotlib.pyplot as plt
plt.plot(errors)
plt.ylabel('error vs iteration')
plt.show()
```
And print results
```
print("iterations: {}, learning rate: {}".format(number_of_iterations, learning_rate))
for test_x, test_y in zip(test_set_X, test_set_Y):
values = execute(test_x, weights, biases, sigmoid)
predicted_y = values[-1]
print("{} xor {} = {} ({} confidence)".format(test_x[0][0], test_x[1][0], round(predicted_y[0][0]), predicted_y))
```
| github_jupyter |
## Dealing with Missing Data
```{admonition} Quick summary
:class: tip
1. **You should focus more on the "whys" of dealing with missing data rather than mechanics.** (You can look up mechanics later.)
- [These slides](https://github.com/matthewbrems/ODSC-missing-data-may-18/blob/master/Analysis%20with%20Missing%20Data.pdf) on missing data are quite good! [This article](https://www.geeksforgeeks.org/working-with-missing-data-in-pandas/) has examples too.
- On the "whys": With firm level data that investment analysts deal with, the most common approach to missing data is to keep all valid observations (don't drop anything), and for each test you run, use all observations that have no missing values for all the variables in a given test. In the slides above, this is called "Complete-Case Analysis".
- "Complete-Case Analysis" works well, as long as the fact that the variable is missing doesn't indicate a systematic difference between observations that are missing and those that aren't. For example, [I have research that characterizes innovation based on patent text](https://bowen.finance/bfh_data/), which gives researchers a powerful tool to examine the impacts of firm innovation. However, this dataset will lead to missing values for any firm without patents. And firms without patents are systematically differant than firms with patents.
- It is less common corporate finance to impute missing values. However, "deductive imputation" is common when the cost of doing so isn't high, like the height example above.
- Interpolation is done in asset pricing when it's necessary to estimate the pricing of options or other derivatives that aren't actually traded.
1. `df.isnull().sum()` will report missing values by variable.
4. In general, when you can confidently deduce a value (my height this year is the same as last year because I'm a fully grown adult, mostly), go ahead. That is valid data.
2. With new datasets, look out for "missing values" that aren't missing. Some datasets use a certain number to indicate missing data (i.e. -99). Convert these to NaNs with `replace`.
```
### Pandas functions you might use to fill missing values
- `fillna` - any value (strings included) you want, back fill, forward, fill, and more
- `dropna` - might not be explicitly needed, some functions ignore NaNs, but some don't. I tend to drop only as needed for a given estimation, and only temporarily.
- `replace` - some datasets use a certain number to indicate missing data (i.e. -99). Convert these to NaN with this function.
- `interpolate` - e.g. use values in surrounding time periods to fill in gaps
- Deduce. Suppose PPE this year is missing. $PPE_t = PPE_{t-1} + CAPX_t - DP_t $
### Practice
Play around with each of those functions on this dataframe:
```
import pandas as pd
import numpy as np
df = pd.DataFrame({"A":[12, 4, 5, None, 1],
"B":[None, 2, 54, 3, None],
"C":[20, 16, None, 3, 8],
"D":[14, 3, None, None, 6]})
_df1 = df.copy()
_df1['firm'] = 1
_df1['date'] = _df1.index
_df2 = df.copy()
_df2['firm'] = 2
_df2['date'] = _df2.index
df2 = _df1.append(_df2)
```
**Questions for `df`**:
1. Fill all missing values with -1
1. Fill missing values for variable "B" with -1
1. Fill all values with the mean for the variable
1. Fill all values with the median for the variable
1. Fill values by taking the most recent non-missing prior value
**Questions for `df2`**:
- Carry missing values forward without carrying values from firm 1 to firm 2
- Fill missing values with the average for firms on that date
| github_jupyter |
# Homework 2
## Almost Shakespeare
Let's try to generate some Shakespeare poetry using RNNs. The sonnets file is available in the notebook directory.
Text generation can be designed in several steps:
1. Data loading
2. Dictionary generation
3. Data preprocessing
4. Model (neural network) training
5. Text generation (model evaluation)
### Data loading
Shakespeare sonnets are awailable at this [link](http://www.gutenberg.org/ebooks/1041?msg=welcome_stranger). In addition, they are stored in the same directory as this notebook (`sonnetes.txt`).
Simple preprocessing is already done for you in the next cell: all technical info is dropped.
**Alternatively**
You could use file `onegin.txt` with Russian texts or your natve language poetry to be able to assess results quality.
**Note: In case of Onegin text you need to adjust reading procedure yourself!!!** (this file has a bit different format than `sonnets.txt`)
```
!wget -nc https://raw.githubusercontent.com/v-goncharenko/madmo-adv/55d929befa12370fc18109f5333f7cf000ea27ce/homeworks/sonnets.txt
!wget -nc https://raw.githubusercontent.com/v-goncharenko/madmo-adv/55d929befa12370fc18109f5333f7cf000ea27ce/homeworks/onegin.txt
with open("sonnets.txt", "r") as iofile:
text = iofile.readlines()
TEXT_START = 45
TEXT_END = -368
text = text[TEXT_START:TEXT_END]
assert len(text) == 2616
```
In opposite to the in-class practice, this time we want to predict complex text. Let's reduce the complexity of the task and lowercase all the symbols.
Now variable `text` is a list of strings. Join all the strings into one and lowercase it.
```
import string
# Join all the strings into one and lowercase it
# Put result into variable text.
# Your great code here
assert len(text) == 100225, "Are you sure you have concatenated all the strings?"
assert not any([x in set(text) for x in string.ascii_uppercase]), "Uppercase letters are present"
print("OK!")
```
Put all the characters, that you've seen in the text, into variable `tokens`.
```
tokens = sorted(set(text))
```
Create dictionary `token_to_idx = {<char>: <index>}` and dictionary `idx_to_token = {<index>: <char>}`
```
# dict <index>:<char>
# Your great code here
# dict <char>:<index>
# Your great code here
```
*Comment: in this task we have only 38 different tokens, so let's use one-hot encoding.*
### Building the model
Now we want to build and train recurrent neural net which would be able to something similar to Shakespeare's poetry.
Let's use vanilla RNN, similar to the one created during the lesson.
```
# Your code here
```
Plot the loss function (axis X: number of epochs, axis Y: loss function).
```
# Your plot code here
# An example of generated text. There is no function `generate_text` in the code above.
# print(generate_text(length=500, temperature=0.2))
```
### More poetic model
Let's use LSTM instead of vanilla RNN and compare the results.
Plot the loss function of the number of epochs. Does the final loss become better?
```
# Your beautiful code here
```
Generate text using the trained net with different `temperature` parameter: `(0.1, 0.2, 0.5, 1.0, 2.0)`.
Evaluate the results visually, try to interpret them.
```
# Text generation with different temperature values here
```
### Saving and loading models
Save the model to the disk, then load it and generate text.
Follow guides from [this tutorial](https://pytorch.org/tutorials/beginner/saving_loading_models.html).
You need to use `Save/Load state_dict (Recommended)` section aka save state dict.
```
# Saving and loading code here
```
## Additional materials on topic
1. [Andrew Karpathy blog post about RNN.](http://karpathy.github.io/2015/05/21/rnn-effectiveness/)\
There are several examples of genration: Shakespeare texts, Latex formulas, Linux Sourse Code and children names.
2. <a href='https://github.com/karpathy/char-rnn'> Repo with char-rnn code </a>
3. Cool repo with [PyTorch examples](https://github.com/spro/practical-pytorch`)
| github_jupyter |
# The One-Body Problem
## Importing Packages
```
import numpy as np
import matplotlib.pyplot as plt
```
## Solving the problem numerically
### The Dynamics
##### Defining the constants of motion
```
G = 6.673e-11 # Newton's Constant in MKS units
M = 1.99e30 # The mass of the Sun in kilograms
GM = G * M # Defined for ease
AU = 149597870700.0 # Earth-Sun distance
yr = 365.25*24*60*60 # One year in seconds
c = 299792458.0 # Speed of light in MKS units
rs = 1e6*GM/c**2 # Schwarzschild radius of sun
```
##### Defining an dynamics (the acceleration) from Newton's Laws
```
def aVec(rVec):
x = rVec[0] # Extracting x and y
y = rVec[1] # from rVec
r = (x**2 + y**2)**(0.5)
RCube = (x**2 + y**2)**(1.5) # Computing r**3 to simplify calculations
ax = -GM*x/RCube *(1 + 3*rs/r)# Computing the acceleration vector's components
ay = -GM*y/RCube *(1 + 3*rs/r)# from Newton's Laws
return np.array([ax,ay]) # Return a "vector"
```
### The Kinematics
##### Setting the initial conditions
```
t_initial = 0 # Initial time
t_final = 10*yr # Final time
dt = 10.0 # Time-step of 1 hour
N = int((t_final-t_initial)/dt) # Number of time-steps
myr = 88*24*60*60
r0 = 2.0e11#1*AU # Magnitude of distance from the sun
v0 = 0.1e5#2*np.pi*r0/yr # Magnitude of velocity for a circular orbit
r0Vec = r0*np.array([1.0,0.0]) # The initial position vector, along x-axis
v0Vec = 1.0*v0*np.array([0.0,1.0]) # The initial velcoity vector, along y-axis
```
##### Defining the arrays to store data and the leapfrog method
```
rVec = np.zeros((N+1,2),float) # An array of 2D vectors of length N+1 for position
vVec = np.zeros((N+1,2),float) # An array of 2D vectors of length N+1 for velocity
t = np.zeros(N+1,float) # An array of length N+1 for time
rVec[0] = r0Vec # Setting the initial position to r0Vec
vVec[0] = v0Vec + aVec(rVec[0])*dt/2 # Setting the initial velocity using the leapfrog method
### Arrays to test our solution ###
energyArray = np.zeros(N+1, float)
LArray = np.zeros(N+1, float)
energyArray[0] = energyPM(r0Vec,v0Vec)
LArray[0] = LPM(r0Vec,v0Vec)
```
##### Solving the kinematics
```
for i in range(1,N+1):
t[i] = i*dt
rVec[i] = rVec[i-1] + vVec[i-1]*dt
vVec[i] = vVec[i-1] + aVec(rVec[i])*dt
### Calculating our conserved quantities ###
energyArray[i] = energyPM(rVec[i],vVec[i])
LArray[i] = LPM(rVec[i],vVec[i])
```
### Plotting the solution
```
xArray = rVec[:,0]
yArray = rVec[:,1]
#plt.figure(figsize=(20,10))
plt.plot(xArray,yArray)
plt.axis('equal')
plt.xlabel("X(t)")
plt.ylabel("Y(t)")
plt.show()
```
## Testing our solution
### Defining our 'conserved quantity' functions
#### The Energy $E$
The total energy of the object (which is a conserved quantity) depends on the total kinetic energy and the total potential energy. The kinetic energy depends on the velocity `vVec` of the object, and the potential energy $V(r)$ depends on the force law, which in turn depends on the position vector `rVec` of the object.
The total energy is given by $E = T + V$, which, in our case is:
$$ E = \frac{1}{2} m v^2 - \frac{G M m}{r^2}.$$
Clearly, since $m$ (the mass of our object) is unchanging, we can define an 'energy per unit mass' which must also be conserved. $$ E_\text{pm} = \frac{1}{2} v^2 - \frac{G M}{r^2}.$$
Thus, the `energyPM` function depends on `energyPM(rVec,vVec)`. In particular, we also know that they depend on the magnitudes of the `rVec` and `vVec` vectors.
```
def energyPM(rVec, vVec):
# Finding the magnitude of the vectors
r = np.linalg.norm(rVec)
v = np.linalg.norm(vVec)
return 0.5*v**2 - GM/r
```
#### The Angular Momentum $L$
Another conserved quantity in this problem is the angular momentum, which again depends on the position and velocity vectors `rVec` and `vVec`. The angular momentum is defined as $L = \vec{r} \times \vec{p}$. In our case (since the mass is unchanging), this is $$\vec{L} = m \left( \vec{r} \times \vec{v}\right).$$
As before, we can defined an 'angular momentum per unit mass', which must also be conserved. $\vec{L}_\text{pm} = \vec{r} \times \vec{v}$. This is a vector condition. A weaker condition is to say that the magnitude of the angular momentum must also be conserved, i.e. $L_\text{pm} = x v_y - y v_x = \text{constant}$, which is what we will use.
```
def LPM(rVec,vVec):
x = rVec[0]
y = rVec[1]
vx= vVec[0]
vy= vVec[1]
return x*vy - y*vx
```
### Plotting our conserved quantities
```
plt.plot(t,(LArray-LArray[0])/LArray[0])
plt.plot(t,(energyArray-energyArray[0])/energyArray[0])
plt.plot(t,xArray**2 + yArray**2)
```
| github_jupyter |
# Import KBase and cFBA
```
# import kbase
import os
local_cobrakbase_path = 'C:\\Users\\Andrew Freiburger\\Dropbox\\My PC (DESKTOP-M302P50)\\Documents\\UVic Civil Engineering\\Internships\\Agronne\\cobrakbase'
os.environ["HOME"] = local_cobrakbase_path
import cobrakbase
token = 'JOSNYJGASTV5BGELWQTUSATE4TNHZ66U'
kbase = cobrakbase.KBaseAPI(token)
ftp_path = '../../../ModelSEEDDatabase'
# import cFBA
%run ../../modelseedpy/core/mscommunity.py
%run ../../modelseedpy/core/msgapfill.py
%matplotlib inline
```
# 2-member Zahmeeth model
## Unconstained model
### Define and execute the model
```
# import the model
# from modelseedpy.fbapkg import kbasemediapkg
modelInfo_2 = ["CMM_iAH991V2_iML1515.kb",40576]
mediaInfo_2 = ["Btheta_Ecoli_minimal_media",40576]
model = kbase.get_from_ws(modelInfo_2[0],modelInfo_2[1])
media = kbase.get_from_ws(mediaInfo_2[0],mediaInfo_2[1])
# kmp = kbasemediapkg.KBaseMediaPkg(self.model)
# kmp.build_package(media)
# simulate and visualize the model
cfba = MSCommunity(model)
cfba.drain_fluxes(media)
cfba.gapfill(media)
cfba.constrain(media)
solution = cfba.run()
cfba.compute_interactions(solution)
cfba.visualize()
```
## FullThermo-constrained model
### Define and execute the model
```
# import the model
# from modelseedpy.fbapkg import kbasemediapkg
modelInfo_2 = ["CMM_iAH991V2_iML1515.kb",40576]
mediaInfo_2 = ["Btheta_Ecoli_minimal_media",40576]
model = kbase.get_from_ws(modelInfo_2[0],modelInfo_2[1])
media = kbase.get_from_ws(mediaInfo_2[0],mediaInfo_2[1])
# kmp = kbasemediapkg.KBaseMediaPkg(self.model)
# kmp.build_package(media)
# simulate and visualize the model
cfba = MSCommunity(model)
cfba.drain_fluxes(media)
cfba.gapfill(media)
cfba.constrain(media, msdb_path_for_fullthermo = ftp_path, verbose = False)
solution = cfba.run()
cfba.compute_interactions(solution)
cfba.visualize()
```
# 3-member Electrosynth model
## Unconstrained model
### Define and execute the model
```
# import the model
# from modelseedpy.fbapkg import kbasemediapkg
modelInfo_2 = ['electrosynth_comnty.mdl.gf.2021',93204]
mediaInfo_2 = ["CO2_minimal",93204]
model = kbase.get_from_ws(modelInfo_2[0],modelInfo_2[1])
media = kbase.get_from_ws(mediaInfo_2[0],mediaInfo_2[1])
# kmp = kbasemediapkg.KBaseMediaPkg(self.model)
# kmp.build_package(media)
# simulate and visualize the model
cfba = MSCommunity(model)
cfba.drain_fluxes(media)
cfba.gapfill(media)
cfba.constrain(media)
solution = cfba.run()
cfba.compute_interactions(solution)
cfba.visualize()
```
## FullThermo-constrained model
### Define and execute the model
```
# import the model
# from modelseedpy.fbapkg import kbasemediapkg
modelInfo_2 = ['electrosynth_comnty.mdl.gf.2021',93204]
mediaInfo_2 = ["CO2_minimal",93204]
model = kbase.get_from_ws(modelInfo_2[0],modelInfo_2[1])
media = kbase.get_from_ws(mediaInfo_2[0],mediaInfo_2[1])
# kmp = kbasemediapkg.KBaseMediaPkg(self.model)
# kmp.build_package(media)
# simulate and visualize the model
cfba = MSCommunity(model)
cfba.drain_fluxes(media)
cfba.gapfill(media)
cfba.constrain(media, msdb_path_for_fullthermo = ftp_path, verbose = False)
solution = cfba.run()
cfba.compute_interactions(solution)
cfba.visualize()
```
# 2-member Aimee model
## Unconstrained model
### Chitin media
```
# import the model
%run ../../modelseedpy/core/mscommunity.py
# from modelseedpy.fbapkg import kbasemediapkg
modelInfo_2 = ['Cjaponicus_Ecoli_Community',97055]
mediaInfo_2 = ["ChitinM9Media",97055]
model = kbase.get_from_ws(modelInfo_2[0],modelInfo_2[1])
media = kbase.get_from_ws(mediaInfo_2[0],mediaInfo_2[1])
# kmp = kbasemediapkg.KBaseMediaPkg(self.model)
# kmp.build_package(media)
# simulate and visualize the model
cfba = MSCommunity(model)
cfba.drain_fluxes(media)
cfba.gapfill(media)
cfba.constrain(media)
solution = cfba.run()
cfba.compute_interactions(solution)
cfba.visualize()
```
## FullThermo-constrained model
### Define and execute the model
```
# import the model
# from modelseedpy.fbapkg import kbasemediapkg
modelInfo_2 = ['Cjaponicus_Ecoli_Community',97055]
mediaInfo_2 = ["ChitinM9Media",97055]
model = kbase.get_from_ws(modelInfo_2[0],modelInfo_2[1])
media = kbase.get_from_ws(mediaInfo_2[0],mediaInfo_2[1])
# kmp = kbasemediapkg.KBaseMediaPkg(self.model)
# kmp.build_package(media)
# simulate and visualize the model
cfba = MSCommunity(model)
cfba.drain_fluxes(media)
cfba.gapfill(media)
cfba.constrain(media, msdb_path_for_fullthermo = ftp_path, verbose = False)
solution = cfba.run()
cfba.compute_interactions(solution)
cfba.visualize()
```
# 7-member Hotlake model
## Unconstrained model
### Define and execute the model
```
# import the model
%run ../../modelseedpy/core/mscommunity.py
# from modelseedpy.fbapkg import kbasemediapkg
modelInfo_2 = ["Hot_Lake_seven.mdl",93544]
mediaInfo_2 = ["HotLakeMedia",93544]
model = kbase.get_from_ws(modelInfo_2[0],modelInfo_2[1])
media = kbase.get_from_ws(mediaInfo_2[0],mediaInfo_2[1])
# kmp = kbasemediapkg.KBaseMediaPkg(self.model)
# kmp.build_package(media)
# simulate and visualize the model
cfba = MSCommunity(model)
cfba.drain_fluxes(media)
cfba.gapfill(media)
cfba.constrain(media)
solution = cfba.run()
cfba.compute_interactions(solution)
cfba.visualize()
```
## FullThermo-constrained model
### Define and execute the model
```
# import the model
%run ../../modelseedpy/core/mscommunity.py
# from modelseedpy.fbapkg import kbasemediapkg
modelInfo_2 = ["Hot_Lake_seven.mdl",93544]
mediaInfo_2 = ["HotLakeMedia",93544]
model = kbase.get_from_ws(modelInfo_2[0],modelInfo_2[1])
media = kbase.get_from_ws(mediaInfo_2[0],mediaInfo_2[1])
# kmp = kbasemediapkg.KBaseMediaPkg(self.model)
# kmp.build_package(media)
# simulate and visualize the model
cfba = MSCommunity(model)
cfba.drain_fluxes(media)
cfba.gapfill(media)
cfba.constrain(media, msdb_path_for_fullthermo = ftp_path, verbose = False)
solution = cfba.run()
cfba.compute_interactions(solution)
cfba.visualize()
```
| github_jupyter |
<img style="float: left; margin: 30px 15px 15px 15px;" src="https://pngimage.net/wp-content/uploads/2018/06/logo-iteso-png-5.png" width="300" height="500" />
### <font color='navy'> Simulación de procesos financieros.
**Nombres:** Betsy Torres | Eduardo Loza
**Fecha:** 05 de noviembre de 2020.
**Expediente** : 714095 | 713423
**Profesor:** Oscar David Jaramillo Zuluaga.
**Liga GitHub:** https://github.com/BetsyTorres/ProyectoConjunto_TorresBetsy_LozaEduardo/blob/main/Tarea_9.ipynb
# Tarea 9: Clase
# Ejercicio 1
Para ver si la venta de chips de silicio son independientes del punto del ciclo de negocios en que se encuentre la economía del país se han recogido las ventas semanales de una empresa y datos acerca de la economía del país, y se reportan los siguientes resultados:

Realice la prueba de chi-cuadrado para validar la independencia del punto del ciclo de negocios en que se encuentre la economía.
# Ejercicio 2
> Ver en este enlace la forma de probar independencia: https://stattrek.com/chi-square-test/independence.aspx?Tutorial=AP
Del enlace anterior replicar los resultados reportados de `Homogeneidad` y `prueba de bondad y ajuste` en python. De esta manera entender cómo se deben realizar estas dos pruebas adicionales.

# Soluciones Betsy Torres
## Sol 1
```
# Importamos librerías
import numpy as np
from functools import reduce
import time
import matplotlib.pyplot as plt
import scipy.stats as st # Librería estadística
import pandas as pd
from scipy import optimize
data = pd.DataFrame(index=['Pico','Abajo','Subiendo','Bajando','Total'],columns=['Alta','Mediana','Baja','Total'])
data.iloc[0,:-1] = [20,7,3]
data.iloc[1,:-1] = [30,40,30]
data.iloc[2,:-1] = [20,8,2]
data.iloc[3,:-1] = [30,5,5]
data.iloc[4,:-1] = data.sum(axis=0)
data['Total'] = data.sum(axis=1)
data
# Obtengo las ponderaciones
P_pico = data.iloc[0,3]/data.iloc[-1,-1]
P_abajo = data.iloc[1,3]/data.iloc[-1,-1]
P_subiendo = data.iloc[2,3]/data.iloc[-1,-1]
f_o = data.loc[data.index!='Total',data.columns!='Total'].values.flatten()
f_e = np.concatenate([data.iloc[-1,:-1]*(P_pico),data.iloc[-1,:-1]*(P_abajo),data.iloc[-1,:-1]*(P_subiendo),data.iloc[-1,:-1]*(1-(P_pico+P_abajo+P_subiendo))])
f_e,f_o
```
### Método de Chi Cuadrada
```
F_obse = f_o
F_espe = f_e
x2 = st.chisquare(F_obse,F_espe,ddof=3)
print('Valor de chi cuadrado = ',list(x2)[0],',p-value de la prueba=',list(x2)[1])
Ji = st.chi2.ppf(q = 0.9,df=3)
print('Estadístico de Ji = ',Ji)
# st.t.interval
x1 = st.chi2.cdf(list(x2)[0],df=3)
x1
```
**Conclusión:**
>Se rechaza la Hipótesis nula $H_0$ ya que el p-value es muy pequeño, por lo tanto se dice que la venta semanal de chips no depende del ciclo de la economía del país.
## Sol 2
**Problema**
Acme Toy Company imprime tarjetas de béisbol. La compañía afirma que el 30% de las tarjetas son novatos, el 60% veteranos pero no All-Stars, y el 10% son All-Stars veteranos.
Suponga que una muestra aleatoria de 100 cartas tiene 50 novatos, 45 veteranos y 5 All-Stars. ¿Es esto consistente con la afirmación de Acme? Utilice un nivel de significancia de 0.05.
**Hipótesis**
- $H_0$: la proporción de novatos, veteranos y All-Stars es 30%, 60% y 10%, respectivamente.
- $H_a$: al menos una de las proporciones de la hipótesis nula es falsa.
```
alpha = 0.05
acme = pd.DataFrame(index=['Rookies','Veterans not All-Stars','Veteran All-Stars'],columns=['Proporciones', 'Random Sample'])
acme['Proporciones'] = [0.30,0.60,0.10]
acme['Random Sample'] = [50, 45, 5]
acme
df = 2
f_e_acme = np.array(acme['Proporciones']*100, dtype=object)
f_o_acme = np.array(acme['Random Sample'], dtype=object)
f_e_acme, f_o_acme
F_obse_acme = f_o_acme
F_esp_acme = f_e_acme
x2_acme = st.chisquare(F_obse_acme, F_esp_acme, ddof=1.99)
print('Valor de chi cuadrado = ',list(x2_acme)[0],', p-value de la prueba=',list(x2_acme)[1])
Ji_acme = st.chi2.ppf(q = 0.95, df=1.99)
print('Estadístico de Ji = ',Ji_acme)
x1_acme = st.chi2.cdf(list(x2_acme)[0],df=1.99)
x1_acme
```
**Conclusión:**
>Se rechaza la Hipótesis nula $H_0$ ya que el p-value es menor a $\alpha = 0.05 $ por lo tanto al menos una de las proposiciones de la Hiótesis nula $H_0$ es falsa
**Problema**
En un estudio de los hábitos de televisión de los niños, un psicólogo del desarrollo selecciona una muestra aleatoria de 300 alumnos de primer grado: 100 niños y 200 niñas. A cada niño se le pregunta cuál de los siguientes programas de televisión le gusta más: El llanero solitario, Barrio Sésamo o Los Simpson. Los resultados se muestran en la tabla de contingencia a continuación.

¿Las preferencias de los chicos por estos programas de televisión difieren significativamente de las preferencias de las chicas? Utilice un nivel de significancia de 0.05.
**Hipótesis**
- $H_0$: La hipótesis nula establece que la proporción de niños que prefieren el llanero solitario es idéntica a la proporción de niñas. Del mismo modo, para los demás programas. Así,
- $H_o : P chicos como Lone Ranger = P chicas como Lone Ranger$
- $H_o : A los chicos les gusta Barrio Sésamo = A las chicas les gusta Barrio Sésamo$
- $H_o : P chicos como Simpson = P chicas como Simpson$
- $H_a$: al menos uno de los enunciados de hipótesis nula es falso.
```
kids = pd.DataFrame(index=['Boys','Girls','total'],columns=['Lone Ranger','Sesame Street','The Simpsons','total'])
kids.iloc[0,:-1] = [50,30,20]
kids.iloc[1,:-1] = [50,80,70]
kids['total'] = kids.sum(axis=1)
kids.loc['total',:] = kids.sum(axis=0)
kids
P_kids = kids.iloc[0,3]/kids.iloc[-1,-1]
f_o_kids = kids.loc[kids.index!='total',kids.columns!='total'].values.flatten()
f_e_kids= np.concatenate([kids.iloc[-1,:-1]*P_kids,kids.iloc[-1,:-1]*(1-P_kids)])
f_e_kids, f_o_kids
F_obs_k = f_o_kids
F_esp_k = f_e_kids
x2_k = st.chisquare(F_obs_k, F_esp_k, ddof=2)
print('Valor de chi cuadrado = ', list(x2_k)[0],',p-value de la prueba=',list(x2_k)[1])
Ji_k = st.chi2.ppf(q = 0.9, df=2)
print('Estadístico de Ji = ',Ji_k)
# st.t.interval
x1_k = st.chi2.cdf(list(x2_k)[0], df=2)
x1_k
```
**Conclusión**
>Se rechaza la hipótesis nula $H_0$ porque el p-value es menor a 0.05 (alpha), por lo que podemos decir que alguno de los enunciados propuestos en la Hipótesis nula es falso.
# Soluciones Eduardo Loza
## Sol 1
```
#creación de data frame
data = pd.DataFrame(index=['Pico','Abajo','Subiendo', 'Bajando'],columns=['Alta', 'Mediana', 'Baja'])
data['Alta'] = [20,30,20,30]
data['Mediana'] = [7,40,8,5]
data['Baja']=[3,30,2,5]
data['Total'] = data.sum(axis=1)
data.loc['Total',:] = data.sum(axis=0)
data
Pa = data.iloc[4,0]/data.iloc[-1,-1]
Pm = data.iloc[4,1]/data.iloc[-1,-1]
Pb = data.iloc[4,2]/data.iloc[-1,-1]
P = np.array([Pa, Pm, Pb])
f_o = data.loc[data.index!='Total',data.columns!='Total'].values.flatten()
f_e = np.concatenate([data.iloc[0,-1]*P, data.iloc[1,-1]*P, data.iloc[2,-1]*P, data.iloc[3,-1]*P])
f_o, f_e
#### Método chi cuadrado
x2 = st.chisquare(f_o,f_e,ddof=6)
print('Valor de chi cuadrado = ',list(x2)[0],',p-value de la prueba=',list(x2)[1])
Ji = st.chi2.ppf(q = 0.9,df=3)
print('Estadístico de Ji = ',Ji)
x1 = st.chi2.cdf(list(x2)[0],df=6)
x1
```
## Sol 2
## Chi-Square Goodness of Fit Test
### Problem
Acme Toy Company prints baseball cards. The company claims that 30% of the cards are rookies, 60% veterans but not All-Stars, and 10% are veteran All-Stars.
Suppose a random sample of 100 cards has 50 rookies, 45 veterans, and 5 All-Stars. Is this consistent with Acme's claim? Use a 0.05 level of significance.
$H_0 =$ las proporciones de rookies, veterans, y all-starts es $30\%,60\%,10\%$ respectivamente.
$H_a =$ al menos una de las proporciones de la $H_0$ es falsa.
```
alpha = 0.05
cards = pd.DataFrame(index=['Rookies','Veterans not all-stars','Veteran all-satrs'],columns=['Proporciones', 'Random'])
cards['Proporciones'] = [0.30,0.60,0.10]
cards['Random'] = [50, 45, 5]
cards
f_es = np.array(cards['Proporciones']*100)
f_ob = np.array(cards['Random'])
f_ob, f_es
#### Método chi cuadrado
x2 = st.chisquare(f_ob,f_es,ddof=1.99)
print('Valor de chi cuadrado = ',list(x2)[0],',p-value de la prueba=',list(x2)[1])
Ji = st.chi2.ppf(q = 0.9,df=2)
print('Estadístico de Ji = ',Ji)
x1 = st.chi2.cdf(list(x2)[0],df=2)
x1
```
Se rechaza la hipótesis nula $H_0$ porque el p-value es menor a $\alpha=.05$
## Chi-Square Test of Homogeneity
### Problem
In a study of the television viewing habits of children, a developmental psychologist selects a random sample of 300 first graders - 100 boys and 200 girls. Each child is asked which of the following TV programs they like best: The Lone Ranger, Sesame Street, or The Simpsons.
```
tv = pd.DataFrame(index=['Boys','Girls','total'],columns=['Lone Ranger','Sesame Street','The Simpsons','total'])
tv.iloc[0,:-1] = [50,30,20]
tv.iloc[1,:-1] = [50,80,70]
tv['total'] = tv.sum(axis=1)
tv.loc['total',:] = tv.sum(axis=0)
tv
```
$H_0 =$ Proporción de niñas que prefieren Lone Ranger es idéntica a la proporción de niños, de igual manera para los otros programas.
$H_a =$ al menos una de las proporciones de la $H_0$ es falsa.
```
P = tv.iloc[0,3]/tv.iloc[-1,-1]
f_obs = tv.loc[tv.index!='total',tv.columns!='total'].values.flatten()
f_esp = np.concatenate([tv.iloc[-1,:-1]*P,tv.iloc[-1,:-1]*(1-P)])
f_esp, f_obs
x2 = st.chisquare(f_obs, f_esp, ddof=2)
print('Valor de chi cuadrado = ', list(x2)[0],',p-value de la prueba=',list(x2)[1])
Ji = st.chi2.ppf(q = 0.9, df=2)
print('Estadístico de Ji = ',Ji)
x1 = st.chi2.cdf(list(x2)[0], df=2)
x1
```
Se rechaza la hipótesis nula $H_0$ porque el p-value es menor a $\alpha=.05$, lo cual nos dice que no son proporcionales los tipos de programas que ven los niños y las niñas.
| github_jupyter |
<a href="https://colab.research.google.com/github/enakai00/rl_book_solutions/blob/master/Chapter06/SARSA_vs_Q_Learning_vs_MC.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import numpy as np
from numpy import random
from pandas import DataFrame
import copy
class Car:
def __init__(self):
self.path = []
self.actions = [(0, 1), (1, 0), (0, -1), (-1, 0)]
self.episodes = [0]
self.q = {}
self.c ={}
self.restart()
def restart(self):
self.x, self.y = 0, 3
self.path = []
def get_state(self):
return self.x, self.y
def show_path(self):
result = [[' ' for x in range(10)] for y in range(7)]
for c, (x, y, a) in enumerate(self.path):
result[y][x] = str(c)[-1]
result[3][7] = 'G'
return result
def add_episode(self, c=0):
self.episodes.append(self.episodes[-1]+c)
def move(self, action):
self.path.append((self.x, self.y, action))
vx, vy = self.actions[action]
if self.x >= 3 and self.x <= 8:
vy -= 1
if self.x >= 6 and self.x <= 7:
vy -= 1
_x, _y = self.x + vx, self.y + vy
if _x < 0 or _x > 9:
_x = self.x
if _y < 0 or _y > 6:
_y = self.y
self.x, self.y = _x, _y
if (self.x, self.y) == (7, 3): # Finish
return True
return False
def get_action(car, epsilon, default_q=0):
if random.random() < epsilon:
a = random.randint(0, len(car.actions))
else:
a = optimal_action(car, default_q)
return a
def optimal_action(car, default_q=0):
optimal = 0
q_max = 0
initial = True
x, y = car.get_state()
for a in range(len(car.actions)):
sa = "{:02},{:02}:{:02}".format(x, y, a)
if sa not in car.q.keys():
car.q[sa] = default_q
if initial or car.q[sa] > q_max:
q_max = car.q[sa]
optimal = a
initial = False
return optimal
def update_q(car, x, y, a, epsilon, q_learning=False):
sa = "{:02},{:02}:{:02}".format(x, y, a)
if q_learning:
_a = optimal_action(car)
else:
_a = get_action(car, epsilon)
_x, _y = car.get_state()
sa_next = "{:02},{:02}:{:02}".format(_x, _y, _a)
if sa not in car.q.keys():
car.q[sa] = 0
if sa_next not in car.q.keys():
car.q[sa_next] = 0
car.q[sa] += 0.5 * (-1 + car.q[sa_next] - car.q[sa])
if q_learning:
_a = get_action(car, epsilon)
return _a
def trial(car, epsilon = 0.1, q_learning=False):
car.restart()
a = get_action(car, epsilon)
while True:
x, y = car.get_state()
finished = car.move(a)
if finished:
car.add_episode(1)
sa = "{:02},{:02}:{:02}".format(x, y, a)
if sa not in car.q.keys():
car.q[sa] = 0
car.q[sa] += 0.5 * (-1 + 0 - car.q[sa])
break
a = update_q(car, x, y, a, epsilon, q_learning)
car.add_episode(0)
def trial_mc(car, epsilon=0.1):
car.restart()
while True:
x, y = car.get_state()
state = "{:02},{:02}".format(x, y)
a = get_action(car, epsilon, default_q=-10**10)
finished = car.move(a)
if finished:
car.add_episode(1)
g = 0
w = 1
path = copy.copy(car.path)
path.reverse()
for x, y, a in path:
car.x, car.y = x, y
opt_a = optimal_action(car, default_q=-10**10)
sa = "{:02},{:02}:{:02}".format(x, y, a)
g += -1 # Reward = -1 for each step
if sa not in car.c.keys():
car.c[sa] = w
car.q[sa] = g
else:
car.c[sa] += w
car.q[sa] += w*(g-car.q[sa])/car.c[sa]
if opt_a != a:
break
w = w / (1 - epsilon + epsilon/len(car.actions))
break
car.add_episode(0)
car1, car2, car3 = Car(), Car(), Car()
while True:
trial(car1)
if len(car1.episodes) >= 10000:
break
print(car1.episodes[-1])
while True:
trial(car2, q_learning=True)
if len(car2.episodes) >= 10000:
break
print(car2.episodes[-1])
while True:
trial_mc(car3)
if len(car3.episodes) >= 200000:
break
print(car3.episodes[-1])
DataFrame({'SARSA': car1.episodes[:8001],
'Q-Learning': car2.episodes[:8001],
'MC': car3.episodes[:8001]}
).plot()
trial(car1, epsilon=0)
print('SARSA:', len(car1.path))
print ("#" * 12)
for _ in map(lambda lst: ''.join(lst), car1.show_path()):
print('#' + _ + '#')
print ("#" * 12)
print ()
trial(car2, epsilon=0)
print('Q-Learning:', len(car2.path))
print ("#" * 12)
for _ in map(lambda lst: ''.join(lst), car2.show_path()):
print('#' + _ + '#')
print ("#" * 12)
print ()
trial_mc(car3, epsilon=0)
print('MC:', len(car3.path))
print ("#" * 12)
for _ in map(lambda lst: ''.join(lst), car3.show_path()):
print('#' + _ + '#')
print ("#" * 12)
print ()
```
| github_jupyter |
# Regression with Amazon SageMaker XGBoost (Parquet input)
This notebook exhibits the use of a Parquet dataset for use with the SageMaker XGBoost algorithm. The example here is almost the same as [Regression with Amazon SageMaker XGBoost algorithm](xgboost_abalone.ipynb).
This notebook tackles the exact same problem with the same solution, but has been modified for a Parquet input.
The original notebook provides details of dataset and the machine learning use-case.
```
import os
import boto3
import re
from sagemaker import get_execution_role
role = get_execution_role()
region = boto3.Session().region_name
# S3 bucket for saving code and model artifacts.
# Feel free to specify a different bucket here if you wish.
bucket = sagemaker.Session().default_bucket()
prefix = 'sagemaker/DEMO-xgboost-parquet'
bucket_path = 'https://s3-{}.amazonaws.com/{}'.format(region, bucket)
```
We will use [PyArrow](https://arrow.apache.org/docs/python/) library to store the Abalone dataset in the Parquet format.
```
!pip install pyarrow
%%time
import numpy as np
import pandas as pd
import urllib.request
from sklearn.datasets import load_svmlight_file
# Download the dataset and load into a pandas dataframe
FILE_DATA = 'abalone'
urllib.request.urlretrieve("https://archive.ics.uci.edu/ml/machine-learning-databases/abalone/abalone.data", FILE_DATA)
feature_names=['Sex',
'Length',
'Diameter',
'Height',
'Whole weight',
'Shucked weight',
'Viscera weight',
'Shell weight',
'Rings']
data = pd.read_csv('abalone.csv',
header=None,
names=feature_names)
# SageMaker XGBoost has the convention of label in the first column
data = data[feature_names[-1:] + feature_names[:-1]]
# Split the downloaded data into train/test dataframes
train, test = np.split(data.sample(frac=1), [int(.8*len(data))])
# requires PyArrow installed
train.to_parquet('abalone_train.parquet')
test.to_parquet('abalone_test.parquet')
%%time
import sagemaker
sagemaker.Session().upload_data('abalone_train.parquet',
bucket=bucket,
key_prefix=prefix+'/'+'train')
sagemaker.Session().upload_data('abalone_test.parquet',
bucket=bucket,
key_prefix=prefix+'/'+'test')
```
We obtain the new container by specifying the framework version (0.90-1). This version specifies the upstream XGBoost framework version (0.90) and an additional SageMaker version (1). If you have an existing XGBoost workflow based on the previous (0.72) container, this would be the only change necessary to get the same workflow working with the new container.
```
from sagemaker.amazon.amazon_estimator import get_image_uri
container = get_image_uri(region, 'xgboost', '0.90-1')
```
After setting training parameters, we kick off training, and poll for status until training is completed, which in this example, takes between 5 and 6 minutes.
```
%%time
import time
import boto3
from time import gmtime, strftime
job_name = 'xgboost-parquet-example-training-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print("Training job", job_name)
#Ensure that the training and validation data folders generated above are reflected in the "InputDataConfig" parameter below.
create_training_params = {
"AlgorithmSpecification": {
"TrainingImage": container,
"TrainingInputMode": "Pipe"
},
"RoleArn": role,
"OutputDataConfig": {
"S3OutputPath": bucket_path + "/" + prefix + "/single-xgboost"
},
"ResourceConfig": {
"InstanceCount": 1,
"InstanceType": "ml.m5.24xlarge",
"VolumeSizeInGB": 20
},
"TrainingJobName": job_name,
"HyperParameters": {
"max_depth":"5",
"eta":"0.2",
"gamma":"4",
"min_child_weight":"6",
"subsample":"0.7",
"silent":"0",
"objective":"reg:linear",
"num_round":"10"
},
"StoppingCondition": {
"MaxRuntimeInSeconds": 3600
},
"InputDataConfig": [
{
"ChannelName": "train",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": bucket_path + "/" + prefix + "/train",
"S3DataDistributionType": "FullyReplicated"
}
},
"ContentType": "application/x-parquet",
"CompressionType": "None"
},
{
"ChannelName": "validation",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": bucket_path + "/" + prefix + "/test",
"S3DataDistributionType": "FullyReplicated"
}
},
"ContentType": "application/x-parquet",
"CompressionType": "None"
}
]
}
client = boto3.client('sagemaker', region_name=region)
client.create_training_job(**create_training_params)
status = client.describe_training_job(TrainingJobName=job_name)['TrainingJobStatus']
print(status)
while status !='Completed' and status!='Failed':
time.sleep(60)
status = client.describe_training_job(TrainingJobName=job_name)['TrainingJobStatus']
print(status)
%matplotlib inline
from sagemaker.analytics import TrainingJobAnalytics
metric_name = 'validation:rmse'
metrics_dataframe = TrainingJobAnalytics(training_job_name=job_name, metric_names=[metric_name]).dataframe()
plt = metrics_dataframe.plot(kind='line', figsize=(12,5), x='timestamp', y='value', style='b.', legend=False)
plt.set_ylabel(metric_name);
```
| github_jupyter |
# Model Centric Federated Learning - MNIST Example: Create Plan
This notebook is an example of creating a simple model and a training plan
for solving MNIST classification in model-centric (aka cross-device) federated learning fashion.
It consists of the following steps:
* Defining the model
* Defining the Training Plan
* Defining the Averaging Plan & FL configuration
* Hosting everything to PyGrid
* Extra: demonstration of PyGrid API
The process of training a hosted model using existing python FL worker is demonstrated in
the following "[MCFL - Execute Plan](mcfl_execute_plan.ipynb)" notebook.
```
# stdlib
import base64
import json
# third party
import jwt
import requests
import torch as th
from websocket import create_connection
# syft absolute
import syft as sy
from syft import deserialize
from syft import serialize
from syft.core.plan.plan_builder import ROOT_CLIENT
from syft.core.plan.plan_builder import make_plan
from syft.federated.model_centric_fl_client import ModelCentricFLClient
from syft.lib.python.int import Int
from syft.lib.python.list import List
from syft.proto.core.plan.plan_pb2 import Plan as PlanPB
from syft.proto.lib.python.list_pb2 import List as ListPB
th.random.manual_seed(42)
```
## Step 1: Define the model
This model will train on MNIST data, it's very simple yet can demonstrate learning process.
There're 2 linear layers:
* Linear 784x100
* ReLU
* Linear 100x10
```
class MLP(sy.Module):
def __init__(self, torch_ref):
super().__init__(torch_ref=torch_ref)
self.l1 = self.torch_ref.nn.Linear(784, 100)
self.a1 = self.torch_ref.nn.ReLU()
self.l2 = self.torch_ref.nn.Linear(100, 10)
def forward(self, x):
x_reshaped = x.view(-1, 28 * 28)
l1_out = self.a1(self.l1(x_reshaped))
l2_out = self.l2(l1_out)
return l2_out
```
## Step 2: Define Training Plan
```
def set_params(model, params):
for p, p_new in zip(model.parameters(), params):
p.data = p_new.data
def cross_entropy_loss(logits, targets, batch_size):
norm_logits = logits - logits.max()
log_probs = norm_logits - norm_logits.exp().sum(dim=1, keepdim=True).log()
return -(targets * log_probs).sum() / batch_size
def sgd_step(model, lr=0.1):
with ROOT_CLIENT.torch.no_grad():
for p in model.parameters():
p.data = p.data - lr * p.grad
p.grad = th.zeros_like(p.grad.get())
local_model = MLP(th)
@make_plan
def train(
xs=th.rand([64 * 3, 1, 28, 28]),
ys=th.randint(0, 10, [64 * 3, 10]),
params=List(local_model.parameters()),
):
model = local_model.send(ROOT_CLIENT)
set_params(model, params)
for i in range(1):
indices = th.tensor(range(64 * i, 64 * (i + 1)))
x, y = xs.index_select(0, indices), ys.index_select(0, indices)
out = model(x)
loss = cross_entropy_loss(out, y, 64)
loss.backward()
sgd_step(model)
return model.parameters()
```
## Step 3: Define Averaging Plan
Averaging Plan is executed by PyGrid at the end of the cycle,
to average _diffs_ submitted by workers and update the model
and create new checkpoint for the next cycle.
_Diff_ is the difference between client-trained
model params and original model params,
so it has same number of tensors and tensor's shapes
as the model parameters.
We define Plan that processes one diff at a time.
Such Plans require `iterative_plan` flag set to `True`
in `server_config` when hosting FL model to PyGrid.
Plan below will calculate simple mean of each parameter.
```
@make_plan
def avg_plan(
avg=List(local_model.parameters()), item=List(local_model.parameters()), num=Int(0)
):
new_avg = []
for i, param in enumerate(avg):
new_avg.append((avg[i] * num + item[i]) / (num + 1))
return new_avg
```
# Config & keys
```
name = "mnist"
version = "1.0"
client_config = {
"name": name,
"version": version,
"batch_size": 64,
"lr": 0.1,
"max_updates": 1, # custom syft.js option that limits number of training loops per worker
}
server_config = {
"min_workers": 2,
"max_workers": 2,
"pool_selection": "random",
"do_not_reuse_workers_until_cycle": 6,
"cycle_length": 28800, # max cycle length in seconds
"num_cycles": 30, # max number of cycles
"max_diffs": 1, # number of diffs to collect before avg
"minimum_upload_speed": 0,
"minimum_download_speed": 0,
"iterative_plan": True, # tells PyGrid that avg plan is executed per diff
}
def read_file(fname):
with open(fname, "r") as f:
return f.read()
private_key = read_file("example_rsa").strip()
public_key = read_file("example_rsa.pub").strip()
server_config["authentication"] = {
"type": "jwt",
"pub_key": public_key,
}
```
## Step 4: Host in PyGrid
Let's now host everything in PyGrid so that it can be accessed by worker libraries (syft.js, KotlinSyft, SwiftSyft, or even PySyft itself).
# Auth
```
grid_address = "localhost:7000"
grid = ModelCentricFLClient(address=grid_address, secure=False)
grid.connect()
```
# Host
If the process already exists, might you need to clear the db. To do that, set path below correctly and run:
```
# !rm PyGrid/apps/domain/src/nodedatabase.db
response = grid.host_federated_training(
model=local_model,
client_plans={"training_plan": train},
client_protocols={},
server_averaging_plan=avg_plan,
client_config=client_config,
server_config=server_config,
)
response
```
# Authenticate for cycle
```
# Helper function to make WS requests
def sendWsMessage(data):
ws = create_connection("ws://" + grid_address)
ws.send(json.dumps(data))
message = ws.recv()
return json.loads(message)
auth_token = jwt.encode({}, private_key, algorithm="RS256").decode("ascii")
auth_request = {
"type": "model-centric/authenticate",
"data": {
"model_name": name,
"model_version": version,
"auth_token": auth_token,
},
}
auth_response = sendWsMessage(auth_request)
auth_response
```
# Do cycle request
```
cycle_request = {
"type": "model-centric/cycle-request",
"data": {
"worker_id": auth_response["data"]["worker_id"],
"model": name,
"version": version,
"ping": 1,
"download": 10000,
"upload": 10000,
},
}
cycle_response = sendWsMessage(cycle_request)
print("Cycle response:", json.dumps(cycle_response, indent=2).replace("\\n", "\n"))
```
# Download model
```
worker_id = auth_response["data"]["worker_id"]
request_key = cycle_response["data"]["request_key"]
model_id = cycle_response["data"]["model_id"]
training_plan_id = cycle_response["data"]["plans"]["training_plan"]
def get_model(grid_address, worker_id, request_key, model_id):
req = requests.get(
f"http://{grid_address}/model-centric/get-model?worker_id={worker_id}&request_key={request_key}&model_id={model_id}"
)
model_data = req.content
pb = ListPB()
pb.ParseFromString(req.content)
return deserialize(pb)
# Model
model_params_downloaded = get_model(grid_address, worker_id, request_key, model_id)
print("Params shapes:", [p.shape for p in model_params_downloaded])
model_params_downloaded[0]
```
# Download & Execute Plan
```
req = requests.get(
f"http://{grid_address}/model-centric/get-plan?worker_id={worker_id}&request_key={request_key}&plan_id={training_plan_id}&receive_operations_as=list"
)
pb = PlanPB()
pb.ParseFromString(req.content)
plan = deserialize(pb)
xs = th.rand([64 * 3, 1, 28, 28])
ys = th.randint(0, 10, [64 * 3, 10])
(res,) = plan(xs=xs, ys=ys, params=model_params_downloaded)
```
# Report Model diff
```
diff = [orig - new for orig, new in zip(res, local_model.parameters())]
diff_serialized = serialize((List(diff))).SerializeToString()
params = {
"type": "model-centric/report",
"data": {
"worker_id": worker_id,
"request_key": request_key,
"diff": base64.b64encode(diff_serialized).decode("ascii"),
},
}
sendWsMessage(params)
```
# Check new model
```
req_params = {
"name": name,
"version": version,
"checkpoint": "latest",
}
res = requests.get(f"http://{grid_address}/model-centric/retrieve-model", req_params)
params_pb = ListPB()
params_pb.ParseFromString(res.content)
new_model_params = deserialize(params_pb)
new_model_params[0]
# !rm PyGrid/apps/domain/src/nodedatabase.db
```
## Step 5: Train
To train hosted model, you can use existing python FL worker.
See the "[MCFL - Execute Plan](mcfl_execute_plan.ipynb)" notebook that
has example of using Python FL worker.
To understand how to make similar model working for mobile FL workers,
see "[MCFL for Mobile - Create Plan](mcfl_execute_plan_mobile.ipynb)" notebook!
| github_jupyter |
# Malaria Detection
Malaria is a life-threatening disease caused by parasites that are transmitted to people through the bites of infected female Anopheles mosquitoes. It is preventable and curable.
In 2017, there were an estimated 219 million cases of malaria in 90 countries.
Malaria deaths reached 435 000 in 2017.
The WHO African Region carries a disproportionately high share of the global malaria burden. In 2017, the region was home to 92% of malaria cases and 93% of malaria deaths.
Malaria is caused by Plasmodium parasites. The parasites are spread to people through the bites of infected female Anopheles mosquitoes, called *"malaria vectors."* There are 5 parasite species that cause malaria in humans, and 2 of these species – P. falciparum and P. vivax – pose the greatest threat.
**Diagnosis of malaria can be difficult:**
Where malaria is not endemic any more (such as in the United States), health-care providers may not be familiar with the disease. Clinicians seeing a malaria patient may forget to consider malaria among the potential diagnoses and not order the needed diagnostic tests. Laboratorians may lack experience with malaria and fail to detect parasites when examining blood smears under the microscope.
Malaria is an acute febrile illness. In a non-immune individual, symptoms usually appear 10–15 days after the infective mosquito bite. The first symptoms – fever, headache, and chills – may be mild and difficult to recognize as malaria. If not treated within 24 hours, P. falciparum malaria can progress to severe illness, often leading to death.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from fastai import *
from fastai.vision import *
from fastai.callbacks.hooks import *
import os
print(os.listdir("../input/cell-images-for-detecting-malaria/cell_images/cell_images/"))
```
**Dataset**
```
img_dir='../input/cell-images-for-detecting-malaria/cell_images/cell_images/'
path=Path(img_dir)
path
data = ImageDataBunch.from_folder(path, train=".",
valid_pct=0.2,
ds_tfms=get_transforms(flip_vert=True, max_warp=0),
size=224,bs=64,
num_workers=0).normalize(imagenet_stats)
print(f'Classes: \n {data.classes}')
data.show_batch(rows=3, figsize=(7,6))
```
## Model ResNet34
```
learn = cnn_learner(data, models.resnet34, metrics=accuracy, model_dir="/tmp/model/")
learn.lr_find()
learn.recorder.plot()
learn.fit_one_cycle(6,1e-2)
learn.save('stage-2')
learn.recorder.plot_losses()
interp = ClassificationInterpretation.from_learner(learn)
interp.plot_top_losses(9, figsize=(15,11))
```
**Confusion Matrix**
```
interp.plot_confusion_matrix(figsize=(8,8), dpi=60)
interp.most_confused(min_val=2)
pred_data= ImageDataBunch.from_folder(path, train=".",
valid_pct=0.2,
ds_tfms=get_transforms(flip_vert=True, max_warp=0),
size=224,bs=64,
num_workers=0).normalize(imagenet_stats)
predictor=cnn_learner(data, models.resnet34, metrics=accuracy, model_dir="/tmp/model/").load('stage-2')
pred_data.single_from_classes(path, pred_data.classes)
x,y = data.valid_ds[3]
x.show()
data.valid_ds.y[3]
pred_class,pred_idx,outputs = predictor.predict(x)
pred_class
```
## Heatmaps
**The heatmap will help us identify were our model it's looking and it's really useful for decision making**
```
def heatMap(x,y,data, learner, size=(0,224,224,0)):
"""HeatMap"""
# Evaluation mode
m=learner.model.eval()
# Denormalize the image
xb,_ = data.one_item(x)
xb_im = Image(data.denorm(xb)[0])
xb = xb.cuda()
# hook the activations
with hook_output(m[0]) as hook_a:
with hook_output(m[0], grad=True) as hook_g:
preds = m(xb)
preds[0,int(y)].backward()
# Activations
acts=hook_a.stored[0].cpu()
# Avg of the activations
avg_acts=acts.mean(0)
# Show HeatMap
_,ax = plt.subplots()
xb_im.show(ax)
ax.imshow(avg_acts, alpha=0.5, extent=size,
interpolation='bilinear', cmap='magma')
heatMap(x,y,pred_data,learn)
```
***It is very hard to completely eliminate false positives and negatives (in a case like this, it could indicate overfitting, given the relatively small training dataset), but the metric for the suitability of a model for the real world is how the model's sensitivity and specificity compare to that of a group of actual pathologists with domain expertise, when both analyze an identical set of real world data that neither has prior exposure to.***
You might improve the accuracy if you artificially increase the size of the training dataset by changing orientations, mirroring, etc., assuming the orientation of the NIH images of the smears haven't been normalized (I would assume they haven't, but that's a dangerous assumption). I'm also curious if you compared ResNet-34 and -50, as 50 might help your specificity (or not).*
| github_jupyter |
<i>Copyright (c) Microsoft Corporation. All rights reserved.</i>
<i>Licensed under the MIT License.</i>
# Surprise Singular Value Decomposition (SVD)
This notebook serves both as an introduction to the [Surprise](http://surpriselib.com/) library, and also introduces the 'SVD' algorithm which is very similar to ALS presented in the ALS deep dive notebook. This algorithm was heavily used during the Netflix Prize competition by the winning BellKor team.
## 1 Matrix factorization algorithm
The SVD model algorithm is very similar to the ALS algorithm presented in the ALS deep dive notebook. The two differences between the two approaches are:
- SVD additionally models the user and item biases (also called baselines in the litterature) from users and items.
- The optimization technique in ALS is Alternating Least Squares (hence the name), while SVD uses stochastic gradient descent.
### 1.1 The SVD model
In ALS, the ratings are modeled as follows:
$$\hat r_{u,i} = q_{i}^{T}p_{u}$$
SVD introduces two new scalar variables: the user biases $b_u$ and the item biases $b_i$. The user biases are supposed to capture the tendency of some users to rate items higher (or lower) than the average. The same goes for items: some items are usually rated higher than some others. The model is SVD is then as follows:
$$\hat r_{u,i} = \mu + b_u + b_i + q_{i}^{T}p_{u}$$
Where $\mu$ is the global average of all the ratings in the dataset. The regularised optimization problem naturally becomes:
$$ \sum(r_{u,i} - (\mu + b_u + b_i + q_{i}^{T}p_{u}))^2 + \lambda(b_i^2 + b_u^2 + ||q_i||^2 + ||p_u||^2)$$
where $\lambda$ is a the regularization parameter.
### 1.2 Stochastic Gradient Descent
Stochastic Gradient Descent (SGD) is a very common algorithm for optimization where the parameters (here the biases and the factor vectors) are iteratively incremented with the negative gradients w.r.t the optimization function. The algorithm essentially performs the following steps for a given number of iterations:
$$b_u \leftarrow b_u + \gamma (e_{ui} - \lambda b_u)$$
$$b_i \leftarrow b_i + \gamma (e_{ui} - \lambda b_i)$$
$$p_u \leftarrow p_u + \gamma (e_{ui} \cdot q_i - \lambda p_u)$$
$$q_i \leftarrow q_i + \gamma (e_{ui} \cdot p_u - \lambda q_i)$$
where $\gamma$ is the learning rate and $e_{ui} = r_{ui} - \hat r_{u,i} = r_{u,i} - (\mu + b_u + b_i + q_{i}^{T}p_{u})$ is the error made by the model for the pair $(u, i)$.
## 2 Surprise implementation of SVD
SVD is implemented in the [Surprise](https://surprise.readthedocs.io/en/stable/) library as a recommender module.
* Detailed documentations of the SVD module in Surprise can be found [here](https://surprise.readthedocs.io/en/stable/matrix_factorization.html#surprise.prediction_algorithms.matrix_factorization.SVD).
* Source codes of the SVD implementation is available on the Surprise Github repository, which can be found [here](https://github.com/NicolasHug/Surprise/blob/master/surprise/prediction_algorithms/matrix_factorization.pyx).
## 3 Surprise SVD movie recommender
We will use the MovieLens dataset, which is composed of integer ratings from 1 to 5.
Surprise supports dataframes as long as they have three colums reprensenting the user ids, item ids, and the ratings (in this order).
```
import sys
import os
import surprise
import papermill as pm
import scrapbook as sb
import pandas as pd
from recommenders.utils.timer import Timer
from recommenders.datasets import movielens
from recommenders.datasets.python_splitters import python_random_split
from recommenders.evaluation.python_evaluation import (rmse, mae, rsquared, exp_var, map_at_k, ndcg_at_k, precision_at_k,
recall_at_k, get_top_k_items)
from recommenders.models.surprise.surprise_utils import predict, compute_ranking_predictions
print("System version: {}".format(sys.version))
print("Surprise version: {}".format(surprise.__version__))
# Select MovieLens data size: 100k, 1m, 10m, or 20m
MOVIELENS_DATA_SIZE = '100k'
```
### 3.1 Load data
```
data = movielens.load_pandas_df(
size=MOVIELENS_DATA_SIZE,
header=["userID", "itemID", "rating"]
)
data.head()
```
### 3.2 Train the SVD Model
Note that Surprise has a lot of built-in support for [cross-validation](https://surprise.readthedocs.io/en/stable/getting_started.html#use-cross-validation-iterators) or also [grid search](https://surprise.readthedocs.io/en/stable/getting_started.html#tune-algorithm-parameters-with-gridsearchcv) inspired scikit-learn, but we will here use the provided tools instead.
We start by splitting our data into trainset and testset with the `python_random_split` function.
```
train, test = python_random_split(data, 0.75)
```
Surprise needs to build an internal model of the data. We here use the `load_from_df` method to build a `Dataset` object, and then indicate that we want to train on all the samples of this dataset by using the `build_full_trainset` method.
```
# 'reader' is being used to get rating scale (for MovieLens, the scale is [1, 5]).
# 'rating_scale' parameter can be used instead for the later version of surprise lib:
# https://github.com/NicolasHug/Surprise/blob/master/surprise/dataset.py
train_set = surprise.Dataset.load_from_df(train, reader=surprise.Reader('ml-100k')).build_full_trainset()
train_set
```
The [SVD](https://surprise.readthedocs.io/en/stable/matrix_factorization.html#surprise.prediction_algorithms.matrix_factorization.SVD) has a lot of parameters. The most important ones are:
- `n_factors`, which controls the dimension of the latent space (i.e. the size of the vectors $p_u$ and $q_i$). Usually, the quality of the training set predictions grows with as `n_factors` gets higher.
- `n_epochs`, which defines the number of iteration of the SGD procedure.
Note that both parameter also affect the training time.
We will here set `n_factors` to `200` and `n_epochs` to `30`. To train the model, we simply need to call the `fit()` method.
```
svd = surprise.SVD(random_state=0, n_factors=200, n_epochs=30, verbose=True)
with Timer() as train_time:
svd.fit(train_set)
print("Took {} seconds for training.".format(train_time.interval))
```
### 3.3 Prediction
Now that our model is fitted, we can call `predict` to get some predictions. `predict` returns an internal object `Prediction` which can be easily converted back to a dataframe:
```
predictions = predict(svd, test, usercol='userID', itemcol='itemID')
predictions.head()
```
### 3.4 Remove rated movies in the top k recommendations
To compute ranking metrics, we need predictions on all user, item pairs. We remove though the items already watched by the user, since we choose not to recommend them again.
```
with Timer() as test_time:
all_predictions = compute_ranking_predictions(svd, train, usercol='userID', itemcol='itemID', remove_seen=True)
print("Took {} seconds for prediction.".format(test_time.interval))
all_predictions.head()
```
### 3.5 Evaluate how well SVD performs
The SVD algorithm was specifically designed to predict ratings as close as possible to their actual values. In particular, it is designed to have a very low RMSE (Root Mean Squared Error), computed as:
$$\sqrt{\frac{1}{N} \sum(\hat{r_{ui}} - r_{ui})^2}$$
As we can see, the RMSE and MAE (Mean Absolute Error) are pretty low (i.e. good), indicating that on average the error in the predicted ratings is less than 1. The RMSE is of course a bit higher, because high errors are penalized much more.
For comparison with other models, we also display Top-k and ranking metrics (MAP, NDCG, etc.). Note however that the SVD algorithm was designed for achieving high accuracy, not for top-rank predictions.
```
eval_rmse = rmse(test, predictions)
eval_mae = mae(test, predictions)
eval_rsquared = rsquared(test, predictions)
eval_exp_var = exp_var(test, predictions)
k = 10
eval_map = map_at_k(test, all_predictions, col_prediction='prediction', k=k)
eval_ndcg = ndcg_at_k(test, all_predictions, col_prediction='prediction', k=k)
eval_precision = precision_at_k(test, all_predictions, col_prediction='prediction', k=k)
eval_recall = recall_at_k(test, all_predictions, col_prediction='prediction', k=k)
print("RMSE:\t\t%f" % eval_rmse,
"MAE:\t\t%f" % eval_mae,
"rsquared:\t%f" % eval_rsquared,
"exp var:\t%f" % eval_exp_var, sep='\n')
print('----')
print("MAP:\t%f" % eval_map,
"NDCG:\t%f" % eval_ndcg,
"Precision@K:\t%f" % eval_precision,
"Recall@K:\t%f" % eval_recall, sep='\n')
# Record results with papermill for tests
sb.glue("rmse", eval_rmse)
sb.glue("mae", eval_mae)
sb.glue("rsquared", eval_rsquared)
sb.glue("exp_var", eval_exp_var)
sb.glue("map", eval_map)
sb.glue("ndcg", eval_ndcg)
sb.glue("precision", eval_precision)
sb.glue("recall", eval_recall)
sb.glue("train_time", train_time.interval)
sb.glue("test_time", test_time.interval)
```
## References
1. Ruslan Salakhutdinov and Andriy Mnih. Probabilistic matrix factorization. 2008. URL: http://papers.nips.cc/paper/3208-probabilistic-matrix-factorization.pdf
2. Yehuda Koren, Robert Bell, and Chris Volinsky. Matrix factorization techniques for recommender systems. 2009.
3. Francesco Ricci, Lior Rokach, Bracha Shapira, and Paul B. Kantor. Recommender Systems Handbook. 1st edition, 2010.
| github_jupyter |
# Song similarity analyser
```
%matplotlib inline
import matplotlib.pyplot as plt
import librosa.display
import sklearn
```
## Settings
```
import shutil
import os
from pathlib import Path
source_path = Path(os.environ["HOME"] + '/Music')
second_to_split_into = 100000
```
## Functions
```
def split_audio_track(sample, sample_rate, second_to_split_into):
samples_per_split = np.round(sample_rate * second_to_split_into)
array_to_split = np.arange(samples_per_split, sample.shape[0], samples_per_split)
return np.split(sample, array_to_split)
def dont_split_audio_track(sample, sample_rate, samples_per_split):
array_to_split = np.arange(samples_per_split, sample.shape[0], samples_per_split)
return np.split(sample, array_to_split)
import time, sys
from IPython.display import clear_output
def update_progress(progress):
bar_length = 20
if isinstance(progress, int):
progress = float(progress)
if not isinstance(progress, float):
progress = 0
if progress < 0:
progress = 0
if progress >= 1:
progress = 1
block = int(round(bar_length * progress))
clear_output(wait = True)
text = "Progress: [{0}] {1:.1f}%".format( "#" * block + "-" * (bar_length - block), progress * 100)
print(text)
```
# Import Data and Create Features
```
import numpy as np
import pandas as pd
import librosa
import pickle
files = [f for f in source_path.glob( "**/*.*")]
error_formats = []
df = pd.DataFrame(columns=['index', 'source', 'zero_crossing', \
'spectral_centroid_mean', 'spectral_centroid_median', 'spectral_centroid_max', 'spectral_centroid_min', \
'mfccs_mean', 'mfccs_median', 'mfccs_max','mfccs_min'])
for file_index in range(len(files)):
update_progress((file_index+1)/len(files))
file = files[file_index]
try:
sample, sample_rate = librosa.load(file, sr = None)
except Exception as e:
print("Reading of sample %s failed" % file)
error_formats.append(file.suffix)
print(e)
continue
split_audio = dont_split_audio_track(sample, sample_rate, len(sample))
data = {'index':np.arange(0, 1),
"source": file}
# Create DataFrame
sample_df = pd.DataFrame(data)
sample_df.insert(2, "data", split_audio, True)
sample_df["zero_crossing"] = sample_df.apply(lambda x: np.sum(librosa.zero_crossings(x['data']) / len(x['data'])), axis = 1)
sample_df["spectral_centroid"] = sample_df.apply(lambda x: librosa.feature.spectral_centroid(x['data'], sr=sample_rate)[0], axis = 1)
sample_df["spectral_centroid_mean"] = sample_df.apply(lambda x: np.mean(x['spectral_centroid']), axis = 1)
sample_df["spectral_centroid_median"] = sample_df.apply(lambda x: np.median(x['spectral_centroid']), axis = 1)
sample_df["spectral_centroid_max"] = sample_df.apply(lambda x: np.max(x['spectral_centroid']), axis = 1)
sample_df["spectral_centroid_min"] = sample_df.apply(lambda x: np.min(x['spectral_centroid']), axis = 1)
sample_df = sample_df.drop(labels= "spectral_centroid", axis = 1)
sample_df["mfccs"] = sample_df.apply(lambda x: librosa.feature.mfcc(x['data'], sr=sample_rate), axis = 1)
sample_df["mfccs_mean"] = sample_df.apply(lambda x: np.mean(x['mfccs']), axis = 1)
sample_df["mfccs_median"] = sample_df.apply(lambda x: np.median(x['mfccs']), axis = 1)
sample_df["mfccs_max"] = sample_df.apply(lambda x: np.max(x['mfccs']), axis = 1)
sample_df["mfccs_min"] = sample_df.apply(lambda x: np.min(x['mfccs']), axis = 1)
sample_df = sample_df.drop(labels=["mfccs", "data"], axis = 1)
df = df.append(sample_df, ignore_index=True, sort=False)
list(set(error_formats))
df = df.reset_index(drop = True)
df
pickle.dump(df, open("music_backup.p", "wb"))
```
| github_jupyter |
# Moving Square Video Prediction
This is the third toy example from Jason Brownlee's [Long Short Term Memory Networks with Python](https://machinelearningmastery.com/lstms-with-python/). It illustrates using a CNN LSTM, ie, an LSTM with input from CNN. Per section 8.2 of the book:
> The moving square video prediction problem is contrived to demonstrate the CNN LSTM. The
problem involves the generation of a sequence of frames. In each image a line is drawn from left to right or right to left. Each frame shows the extension of the line by one pixel. The task is for the model to classify whether the line moved left or right in the sequence of frames. Technically, the problem is a sequence classification problem framed with a many-to-one prediction model.
```
from __future__ import division, print_function
from sklearn.metrics import accuracy_score, confusion_matrix
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import OneHotEncoder
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
import numpy as np
import matplotlib.pyplot as plt
import os
import shutil
%matplotlib inline
DATA_DIR = "../../data"
MODEL_FILE = os.path.join(DATA_DIR, "torch-08-moving-square-{:d}.model")
TRAINING_SIZE = 5000
VALIDATION_SIZE = 100
TEST_SIZE = 500
SEQUENCE_LENGTH = 50
FRAME_SIZE = 50
BATCH_SIZE = 32
NUM_EPOCHS = 5
LEARNING_RATE = 1e-3
```
## Prepare Data
Our data is going to be batches of sequences of images. Each image will need to be in channel-first format, since Pytorch only supports that format. So our output data will be in the (batch_size, sequence_length, num_channels, height, width) format.
```
def next_frame(frame, x, y, move_right, upd_int):
frame_size = frame.shape[0]
if x is None and y is None:
x = 0 if (move_right == 1) else (frame_size - 1)
y = np.random.randint(0, frame_size, 1)[0]
else:
if y == 0:
y = np.random.randint(y, y + 1, 1)[0]
elif y == frame_size - 1:
y = np.random.randint(y - 1, y, 1)[0]
else:
y = np.random.randint(y - 1, y + 1, 1)[0]
if move_right:
x = x + 1
else:
x = x - 1
new_frame = frame.copy()
new_frame[y, x] = upd_int
return new_frame, x, y
row, col = None, None
frame = np.ones((5, 5))
move_right = 1 if np.random.random() < 0.5 else 0
for i in range(5):
frame, col, row = next_frame(frame, col, row, move_right, 0)
plt.subplot(1, 5, (i+1))
plt.xticks([])
plt.yticks([])
plt.title((col, row, "R" if (move_right==1) else "L"))
plt.imshow(frame, cmap="gray")
plt.tight_layout()
plt.show()
def generate_data(frame_size, sequence_length, num_samples):
assert(frame_size == sequence_length)
xs, ys = [], []
for bid in range(num_samples):
frame_seq = []
row, col = None, None
frame = np.ones((frame_size, frame_size))
move_right = 1 if np.random.random() < 0.5 else 0
for sid in range(sequence_length):
frm, col, row = next_frame(frame, col, row, move_right, 0)
frm = frm.reshape((1, frame_size, frame_size))
frame_seq.append(frm)
xs.append(np.array(frame_seq))
ys.append(move_right)
return np.array(xs), np.array(ys)
X, y = generate_data(FRAME_SIZE, SEQUENCE_LENGTH, 10)
print(X.shape, y.shape)
Xtrain, ytrain = generate_data(FRAME_SIZE, SEQUENCE_LENGTH, TRAINING_SIZE)
Xval, yval = generate_data(FRAME_SIZE, SEQUENCE_LENGTH, VALIDATION_SIZE)
Xtest, ytest = generate_data(FRAME_SIZE, SEQUENCE_LENGTH, TEST_SIZE)
print(Xtrain.shape, ytrain.shape, Xval.shape, yval.shape, Xtest.shape, ytest.shape)
```
## Define Network
We want to build a CNN-LSTM network. Each image in the sequence will be fed to a CNN which will learn to produce a feature vector for the image. The sequence of vectors will be fed into an LSTM and the LSTM will learn to generate a context vector that will be then fed into a FCN that will predict if the square is moving left or right.
<img src="08-network-design.png"/>
```
class CNN(nn.Module):
def __init__(self, input_height, input_width, input_channels,
output_channels,
conv_kernel_size, conv_stride, conv_padding,
pool_size):
super(CNN, self).__init__()
self.conv1 = nn.Conv2d(input_channels, output_channels,
kernel_size=conv_kernel_size,
stride=conv_stride,
padding=conv_padding)
self.relu1 = nn.ReLU()
self.output_height = input_height // pool_size
self.output_width = input_width // pool_size
self.output_channels = output_channels
self.pool_size = pool_size
def forward(self, x):
x = self.conv1(x)
x = self.relu1(x)
x = F.max_pool2d(x, self.pool_size)
x = x.view(x.size(0), self.output_channels * self.output_height * self.output_width)
return x
cnn = CNN(FRAME_SIZE, FRAME_SIZE, 1, 2, 2, 1, 1, 2)
print(cnn)
# size debugging
print("--- size debugging ---")
inp = Variable(torch.randn(BATCH_SIZE, 1, FRAME_SIZE, FRAME_SIZE))
out = cnn(inp)
print(out.size())
class CNNLSTM(nn.Module):
def __init__(self, image_size, input_channels, output_channels,
conv_kernel_size, conv_stride, conv_padding, pool_size,
seq_length, hidden_size, num_layers, output_size):
super(CNNLSTM, self).__init__()
# capture variables
self.num_layers = num_layers
self.seq_length = seq_length
self.image_size = image_size
self.output_channels = output_channels
self.hidden_size = hidden_size
self.lstm_input_size = output_channels * (image_size // pool_size) ** 2
# define network layers
self.cnn = CNN(image_size, image_size, input_channels, output_channels,
conv_kernel_size, conv_stride, conv_padding, pool_size)
self.lstm = nn.LSTM(self.lstm_input_size, hidden_size, num_layers, batch_first=True)
self.fc = nn.Linear(hidden_size, output_size)
self.softmax = nn.Softmax()
def forward(self, x):
if torch.cuda.is_available():
h0 = (Variable(torch.randn(self.num_layers, x.size(0), self.hidden_size).cuda()),
Variable(torch.randn(self.num_layers, x.size(0), self.hidden_size).cuda()))
else:
h0 = (Variable(torch.randn(self.num_layers, x.size(0), self.hidden_size)),
Variable(torch.randn(self.num_layers, x.size(0), self.hidden_size)))
cnn_out = []
for i in range(self.seq_length):
cnn_out.append(self.cnn(x[:, i, :, :, :]))
x = torch.cat(cnn_out, dim=1).view(-1, self.seq_length, self.lstm_input_size)
x, h0 = self.lstm(x, h0)
x = self.fc(x[:, -1, :])
x = self.softmax(x)
return x
model = CNNLSTM(FRAME_SIZE, 1, 2, 2, 1, 1, 2, SEQUENCE_LENGTH, 50, 1, 2)
if torch.cuda.is_available():
model.cuda()
print(model)
# size debugging
print("--- size debugging ---")
inp = Variable(torch.randn(BATCH_SIZE, SEQUENCE_LENGTH, 1, FRAME_SIZE, FRAME_SIZE))
out = model(inp)
print(out.size())
loss_fn = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters(), lr=LEARNING_RATE)
```
## Train Network
Training on GPU is probably preferable for this example, takes a long time on CPU. During some runs, the training and validation accuracies get stuck, possibly because of bad initializations, the fix appears to be to just retry the training until it results in good training and validation accuracies and use the resulting model.
```
def compute_accuracy(pred_var, true_var):
if torch.cuda.is_available():
ypred = pred_var.cpu().data.numpy()
ytrue = true_var.cpu().data.numpy()
else:
ypred = pred_var.data.numpy()
ytrue = true_var.data.numpy()
return accuracy_score(ypred, ytrue)
history = []
for epoch in range(NUM_EPOCHS):
num_batches = Xtrain.shape[0] // BATCH_SIZE
shuffled_indices = np.random.permutation(np.arange(Xtrain.shape[0]))
train_loss, train_acc = 0., 0.
for bid in range(num_batches):
Xbatch_data = Xtrain[shuffled_indices[bid * BATCH_SIZE : (bid+1) * BATCH_SIZE]]
ybatch_data = ytrain[shuffled_indices[bid * BATCH_SIZE : (bid+1) * BATCH_SIZE]]
Xbatch = Variable(torch.from_numpy(Xbatch_data).float())
ybatch = Variable(torch.from_numpy(ybatch_data).long())
if torch.cuda.is_available():
Xbatch = Xbatch.cuda()
ybatch = ybatch.cuda()
# initialize gradients
optimizer.zero_grad()
# forward
Ybatch_ = model(Xbatch)
loss = loss_fn(Ybatch_, ybatch)
# backward
loss.backward()
train_loss += loss.data[0]
_, ybatch_ = Ybatch_.max(1)
train_acc += compute_accuracy(ybatch_, ybatch)
optimizer.step()
# compute training loss and accuracy
train_loss /= num_batches
train_acc /= num_batches
# compute validation loss and accuracy
val_loss, val_acc = 0., 0.
num_val_batches = Xval.shape[0] // BATCH_SIZE
for bid in range(num_val_batches):
# data
Xbatch_data = Xval[bid * BATCH_SIZE : (bid + 1) * BATCH_SIZE]
ybatch_data = yval[bid * BATCH_SIZE : (bid + 1) * BATCH_SIZE]
Xbatch = Variable(torch.from_numpy(Xbatch_data).float())
ybatch = Variable(torch.from_numpy(ybatch_data).long())
if torch.cuda.is_available():
Xbatch = Xbatch.cuda()
ybatch = ybatch.cuda()
Ybatch_ = model(Xbatch)
loss = loss_fn(Ybatch_, ybatch)
val_loss += loss.data[0]
_, ybatch_ = Ybatch_.max(1)
val_acc += compute_accuracy(ybatch_, ybatch)
val_loss /= num_val_batches
val_acc /= num_val_batches
torch.save(model.state_dict(), MODEL_FILE.format(epoch+1))
print("Epoch {:2d}/{:d}: loss={:.3f}, acc={:.3f}, val_loss={:.3f}, val_acc={:.3f}"
.format((epoch+1), NUM_EPOCHS, train_loss, train_acc, val_loss, val_acc))
history.append((train_loss, val_loss, train_acc, val_acc))
losses = [x[0] for x in history]
val_losses = [x[1] for x in history]
accs = [x[2] for x in history]
val_accs = [x[3] for x in history]
plt.subplot(211)
plt.title("Accuracy")
plt.plot(accs, color="r", label="train")
plt.plot(val_accs, color="b", label="valid")
plt.legend(loc="best")
plt.subplot(212)
plt.title("Loss")
plt.plot(losses, color="r", label="train")
plt.plot(val_losses, color="b", label="valid")
plt.legend(loc="best")
plt.tight_layout()
plt.show()
```
## Test/Evaluate Network
```
saved_model = CNNLSTM(FRAME_SIZE, 1, 2, 2, 1, 1, 2, SEQUENCE_LENGTH, 50, 1, 2)
saved_model.load_state_dict(torch.load(MODEL_FILE.format(5)))
if torch.cuda.is_available():
saved_model.cuda()
ylabels, ypreds = [], []
num_test_batches = Xtest.shape[0] // BATCH_SIZE
for bid in range(num_test_batches):
Xbatch_data = Xtest[bid * BATCH_SIZE : (bid + 1) * BATCH_SIZE]
ybatch_data = ytest[bid * BATCH_SIZE : (bid + 1) * BATCH_SIZE]
Xbatch = Variable(torch.from_numpy(Xbatch_data).float())
ybatch = Variable(torch.from_numpy(ybatch_data).long())
if torch.cuda.is_available():
Xbatch = Xbatch.cuda()
ybatch = ybatch.cuda()
Ybatch_ = saved_model(Xbatch)
_, ybatch_ = Ybatch_.max(1)
if torch.cuda.is_available():
ylabels.extend(ybatch.cpu().data.numpy())
ypreds.extend(ybatch_.cpu().data.numpy())
else:
ylabels.extend(ybatch.data.numpy())
ypreds.extend(ybatch_.data.numpy())
print("Test accuracy: {:.3f}".format(accuracy_score(ylabels, ypreds)))
print("Confusion matrix")
print(confusion_matrix(ylabels, ypreds))
for i in range(NUM_EPOCHS):
os.remove(MODEL_FILE.format(i + 1))
```
| github_jupyter |
# Match results prediction - Tutorial
This notebook gives you examples about how to use the **predict.py** script in this respository to predict the result of a Team Match of **Chess.com**.
Other ways to predict the result is running notebooks **MatchResutlEstimation.ipynb** or **MatchVarianceAnalysis.ipynb**. The first one detail the procedure to predict a Match of two teams, the second one is a more complete notebook to predict, where is considered a bias as a variable in the Match.
## Running from Terminal
The default way to execute Match Result Prediction using **predict.py** is using *Terminal*.
1. The first step is clonning this repository into the folder you want:
`$ git clone https://github.com/francof2a/ELO_multiplayer_match.git`
2. You need information about the match over you want to get a prediction:
* **API url**: get the Chess.com url of the match. For example https://api.chess.com/pub/match/995756 is the match **WL2019 R4: Team Argentina vs Team England**.
* **ID**: get the ID (identifier of the match) which is the las part of the url. For the last example, the ID is 995756.
* **chess.com url**: if you only get the match url of chess.com, for the last examplie it would be https://www.chess.com/club/matches/team-argentina/995756, you can get the ID (last field of the url), and build the API url concatenating _https://api.chess.com/pub/match/_ and _ID_, or just use ID.
3. Run the prediction just doing next:
`$ python predict.py -id "995756"`
**Note**: The first time you run a prediction for a match, all the info is downloaded from _chess.com API_ and it will take several minutes depending of connection, so please be patient. A backup of that information is stored in _data_ folder to avoid repeat downloading.
A report like this will be printed in terminal:
```
Match info:
Name: WL2019 R4: Team Argentina vs Team England
Team A: Team Argentina
Team B: Team England
Reading ELOs list
Loading from web ...
Done!
Saving backup file./data/wl2019-r4-team-argentina-vs-team-england_match_stats.xlsx
Simulation of match - Result prediction:
Team A (Team Argentina):
Win chances = 100.00 %
Draw chances = 0.00 %
Lose chances = 0.00 %
Expected final score = 423.88 (±11.16) - 244.12 (±11.16)
Expected effectiveness = 63.45 % - 36.55 %
Calculation of Variance over Team A ELOs
Done!
```
All the plots generated will be stored in **outputs** folder.
## Running from jupyter
The **predict.py** script can be executed from a jupyter notebook (like this) emulating console/terminal command line entry using **!**:
```
!python predict.py -id "995756"
```
All the plots generated will be stored in **outputs** folder.
If you have a **ipython console** or you are running the script in a jupyter notebook, you can show the plots inline adding **-plot** argument, then:
```
%matplotlib inline
%run predict.py -id "995756" -plot
```
Note that a **backup file** is being used instead of downloading data from web again.
## Arguments
Next, the list of arguments supported by **predict.py**, and examples:
* -h : help about arguments.
* -id "< match_id >" : (int) ID assigned to the match by chess.com.
* -url "< API match URL >" : (str) URL of the chess.com API assigned to the match. Don't use if you have already especified ID.
* -N < number of trials > : (int) number of trials to execute during prediction. Default value = 1000.
* -Nb < number of trials > : (int) number of trials to execute during prediction considering bias for a team. Default value = 1000.
* -bias < ELO bias > : (float) ELO bias value (offset) for Team A (first team of the match). Default value = 0. It is necessary specify this value to enable _biased analysis_.
* -plot : enable inline plot in _ipython_ console or jupyter notebook.
* -u : force update match data from the web. It is not necessary use this for the first prediction of a particular match.
Examples:
`$ python predict.py -h"`
`$ python predict.py -id "995756"`
`$ python predict.py -url "https://api.chess.com/pub/match/995756"`
`$ python predict.py -id "995756" -N 500`
`$ python predict.py -id "995756" -bias 21.5`
`$ python predict.py -id "995756" -bias 21.5 -Nb 750`
`$ ipython predict.py -id "995756" -plot`
`$ python predict.py -id "995756" -u`
| github_jupyter |
# Bayesian Optimization for Single-Interface Nanoparticle Discovery
**Notebook last update: 3/26/2021** (clean up)
This notebook contains the entire closed-loop process for SINP discovery with BO through SPBCL synthesis, STEM-EDS characterization, as reported in Wahl et al. *to be submitted* 2021.
```
import pandas as pd
from IPython.display import display
import matplotlib.pyplot as plt
import numpy as np
import os
import itertools
import io
from nanoparticle_project import EmbedCompGPUCB, get_comps, \
get_stoichiometric_formulas, compare_to_seed, load_np_data, update_with_new_data
from matminer.featurizers.composition import ElementProperty
from pymatgen import Composition
path = os.getcwd()
```
We will load our dataset into a pandas Dataframe and prepare for downstream modeling. We will prepare the feature space as described in the manuscript using the composition-derived descriptors of Ward et al.
## Prepare seed data and search space
```
df = pd.read_csv('megalibrary_data.csv')
_elts = ['Au%', 'Ag%', 'Cu%', 'Co%', 'Ni%', 'Pt%', 'Pd%', 'Sn%']
for col in _elts:
df[col] = df[col]/100.0
df = df.sample(frac=1) # shuffling the dataframe
df['target'] = -1*np.abs(df["Interfaces"]-1) # set target to single interface NPs
df = df[~df.duplicated()] # drop duplicates
df['Composition'] = df.apply(get_comps,axis=1)
df['n_elems'] = (df[_elts]>0).sum(axis=1)
ep = ElementProperty.from_preset(preset_name='magpie')
featurized_df = ep.featurize_dataframe(df[ ['Composition','target'] ],'Composition').drop('Composition',axis=1)
```
We should now create our search space *D*. First, we generate the composition grid. Then we featurize it as before using compositional descriptors, to generate our `candidate_feats` to search over using BO. Next, we remove any composition from our search space that is closer to a data point in our experimental seed than 5% on any axis.
```
elements = ['Au%', 'Ag%', 'Cu%', 'Co%', 'Ni%', 'Pd%', 'Sn%'] # We will acquire Pt-free in the following iterations
D = get_stoichiometric_formulas(n_components=7, npoints=11)
candidate_data = pd.DataFrame.from_records(D,columns=elements)
candidate_data['Pt%'] = 0.0
candidate_data[ candidate_data <0.00001 ] = 0.0
candidate_data['Composition'] = candidate_data.apply(get_comps,axis=1)
candidate_feats = ep.featurize_dataframe(candidate_data, 'Composition')
candidate_feats = candidate_feats.drop(elements+['Pt%']+['Composition'],axis=1)
for ind,row in df[_elts].iterrows():
candidate_data = candidate_data[_elts][ np.any(np.abs(row-candidate_data)>=0.05,axis=1) ]
candidate_feats = candidate_feats.loc[candidate_data.index]
candidate_feats.shape
```
# Closed-loop optimization procedure
We track the closed loop iterations below step-by-step, making suggestios and updating seed and candidate space with incoming data in each round. We follow this in unfolded form, so that we can closely inspect inputs and outputs in each round.
This is our initial data and the quaternary search space:
```
seed_df = df
seed_data = featurized_df
quaternaries = candidate_data[ ((candidate_data != 0).sum(axis=1) == 4)]
quaternary_feats = candidate_feats.loc[quaternaries.index]
round_number = 1
```
## Round 1
*Optimization agent's suggestions*:
```
agent = EmbedCompGPUCB(n_query=4)
suggestions = agent.get_hypotheses(candidate_data=quaternary_feats, seed_data=seed_data)
display(quaternaries.loc[ suggestions.index ])
compare_to_seed(quaternaries.loc[ suggestions.index ], seed_df)
```
*Experimental feedback in response to suggestions:*
```
new_raw_data = """
Co% Ni% Cu% Au%
13.886 42.787 21.824 21.502
13.883 43.138 21.701 21.278
13.621 42.33 22.244 21.805
22.188 34.332 9.411 34.069
22.186 33.932 9.799 34.083
21.192 34.426 9.112 35.269
8.453 33.012 6.68 51.855
8.935 34.187 6.161 50.718
8.037 34.035 6.445 51.483
10.357 34.259 6.896 48.487
10.767 35.4 6.482 47.352
10.695 36.379 5.961 46.965
13.172 47.616 9.277 29.935
12.56 49.381 8.816 29.243
12.482 47.937 9.203 30.378
12.804 48.143 8.882 30.172
12.396 48.56 9.302 29.742
"""
seed_df, seed_data, quaternaries, quaternary_feats = update_with_new_data(suggestions, new_raw_data, seed_df, seed_data,
quaternaries, quaternary_feats, round_number=round_number,
elements=elements, measured=0)
round_number+=1
```
## Round 2
*Optimization agent's suggestions*:
```
agent = EmbedCompGPUCB(n_query=4)
suggestions = agent.get_hypotheses(candidate_data=quaternary_feats, seed_data=seed_data)
display(quaternaries.loc[ suggestions.index ])
compare_to_seed(quaternaries.loc[ suggestions.index ], seed_df)
new_raw_data = """
Ni% Cu% Ag% Au% Co%
45.71 7.11 7.81 39.37 0
38.42 6.87 11.52 43.19 0
37.13 6.14 13.34 43.39 0
37.61 6.33 14.41 41.65 0
41.49 6.51 8.42 43.58 0
38.63 6.72 6.61 48.04 0
40.04 5.18 13.91 40.87 0
40.46 4.98 14.55 40.01 0
40.36 6.04 7.95 45.64 0
37.9 5.35 15.51 41.24 0
41.92 5.58 9.75 42.76 0
9.35 14.33 0 33.11 43.21
10.1 15.14 0 31.31 43.45
10.92 14.98 0 30.16 43.95
10.63 14.72 0 31.63 43.01
"""
seed_df, seed_data, quaternaries, quaternary_feats = update_with_new_data(suggestions, new_raw_data, seed_df, seed_data,
quaternaries, quaternary_feats, round_number=round_number,
elements=elements, measured=0)
round_number+=1
```
## Round 3
*Optimization agent's suggestions*:
```
agent = EmbedCompGPUCB(n_query=4)
suggestions = agent.get_hypotheses(candidate_data=quaternary_feats, seed_data=seed_data)
display(quaternaries.loc[ suggestions.index ])
compare_to_seed(quaternaries.loc[ suggestions.index ], seed_df)
new_raw_data = """
Ni% Co% Ag% Au% Cu% Pd%
55.4 29.9 5.8 7.5 1.5 0.0
55.5 29.6 4.5 7.8 2.6 0.0
56.2 29.6 4.2 6.3 3.7 0.0
63.1 30.2 2.8 3.9 0 0.0
63.8 30.2 2.3 3.7 0 0.0
62.9 30.2 1.4 3.5 2.1 0.0
18.8 39.7 0 20.9 20.6 0.0
22.8 40.4 0 28.2 8.5 0.0
20 42 0 19 19 0.0
24.4 24.6 0 35.7 15.3 0.0
22.9 24.5 0 43 9.6 0.0
25.4 26.8 0 28.8 19.1 0.0
25.3 26.1 0 25.3 16.2 0.0
0 55 0 24.6 13.2 7.3
0 55.7 0 24.1 13.5 6.7
0 53.4 0 24.8 14.3 7.4
0 56.4 0 22.7 13.7 7.2
"""
seed_df, seed_data, quaternaries, quaternary_feats = update_with_new_data(suggestions, new_raw_data, seed_df, seed_data,
quaternaries, quaternary_feats, round_number=round_number,
elements=elements, measured=0)
round_number+=1
```
## Exploratory Rounds
### Pentanary SINP discovery
```
pentanaries = candidate_data[ ((candidate_data != 0).sum(axis=1) == 5)]
pentanary_feats = candidate_feats.loc[pentanaries.index]
agent = EmbedCompGPUCB(n_query=10)
suggestions_pentanaries = agent.get_hypotheses(candidate_data=pentanary_feats, seed_data=seed_data)
display(pentanaries.loc[ suggestions_pentanaries.index ])
compare_to_seed(pentanaries.loc[ suggestions_pentanaries.index ], seed_df)
new_raw_data = """
Co% Ni% Cu% Pd% Ag% Au%
32.9 10.6 7.3 12.2 0 37
29.5 9.5 6.4 20.5 0 34.2
33.9 10 7 12.3 0 36.8
32.8 9.9 7 13.6 0 36.7
43.9 18 14.7 9 0 14.4
46.3 18.4 13.5 8.2 0 13.6
44.8 18 14.1 8.8 0 14.2
19.4 39.8 10.4 11 0 19.4
19.5 40.2 10.3 10.8 0 19.3
19.7 40 10.4 10.5 0 19.3
19 40.1 10.2 11 0 19.7
22.9 45.1 7.8 0 3.5 20.6
23.1 44.8 7 0 5.9 19.1
23.5 45 7.3 0 5 19.3
22.8 44 7.2 0 6.6 19.5
8.2 23.5 6.5 0 6.3 55.5
7.6 22.6 6.1 0 9.9 53.8
7.9 24.1 6.2 0 7.8 54
7.8 23.2 6 0 10 53
"""
suggestions_targeted_by_team = [6250,6243,5073,5484,6489]
seed_df, seed_data, pentanaries, pentanary_feats = update_with_new_data(suggestions_pentanaries.loc[suggestions_targeted_by_team],
new_raw_data, seed_df, seed_data,
pentanaries, pentanary_feats,
round_number=round_number,
elements=elements, measured=0)
round_number+=1
```
### Hexanary SINP discovery
```
hexanaries = candidate_data[ ((candidate_data != 0).sum(axis=1) == 6)]
hexanaries_feats = candidate_feats.loc[hexanaries.index]
agent = EmbedCompGPUCB(n_query=10)
suggestions_hexanaries = agent.get_hypotheses(candidate_data=hexanaries_feats, seed_data=seed_data)
display(hexanaries.loc[ suggestions_hexanaries.index ].head(10))
compare_to_seed(hexanaries.loc[ suggestions_hexanaries.index ], seed_df)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/Dogechi/Core_exersices/blob/master/20191211_Cluster_Sampling_Examples.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Cluster Sampling
**K-Means Clustering**
In this session, we are going to see how we can perform clustering in practice. To achieve this, we are going to apply one of the most commonly used mothod called K-Means.
K-means clustering is a very simply algorithm that clusters data into K numbers of clusters. But before we dive into what K-mean is all about, there are few terminologies we need to understand to fully grasp the power of K-means clustering.
Please watch the following short videos which explains a few terminologies we are going to use and an overview of how K-means work.
* [K-mean prerequisites](https://drive.google.com/file/d/1JCrS3sRjnZK6GyNYHNwCS6FjyJYryyIQ/view?usp=sharing)
* [K-mean clustering](https://drive.google.com/file/d/119mx9ftGnLZHpK2MuFu__pYKj8UEMKFW/view?usp=sharing)
#Examples
```
# Let's start by importing the necessary librabries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# Set the styles of all graphs to Seaborn one
sns.set()
# Import the KMeans module so we can perform k-means clustering with sklearn
from sklearn.cluster import KMeans
```
**Load the data**
Here is the dataset that we will be using for this example. [Dataset Download](https://drive.google.com/file/d/1IrGJhPCN0e70xlvK2UvLLsup4Fp9uEdH/view?usp=sharing)
```
# Load the country clusters data
data = pd.read_csv('3.01. Country clusters.csv')
#checking out the data manually
data
```
**Plotting the Data**
```
# Create a scatter plot using the longitude and latitude
# Note that in order to reach a result resembling the world map, we must use the longitude as y, and the latitude as x
plt.scatter(data['Longitude'],data['Latitude'])
# Set limits of the axes, again to resemble the world map
plt.xlim(-180,180)
plt.ylim(-90,90)
plt.show
```
**Selecting the features we want from our Dataset**
```
# Since we want to cluster our dataset depending on location, we are going to extract the longitude and latitude and use them as our input while clustering
# To achieve this we will use panda's method .iloc to slice the columns we want from the dataset.
# The first argument of this method identifies the rows we want to keep
# The second - the columns
# In our case, we want to keep all rows and the 1 and 2 colummn.
# Note: Python column indices start from 0
x = data.iloc[:, 1:3]
# Check if we got the 1 and 2 column
x
```
**Clustering**
```
# Create an object from the KMeans method we imported from sklearn library
# The argument in the Kmean method indicates the number of clusters we are aiming for.
# We will use this object for clustering
kmeans = KMeans(2)
# Clustering itself happens using the fit method like this.Here, we are calling the fit method on the object we created and then using our input as x.
kmeans.fit(x)
```
**Clustering Results**
```
# Now that we have performed clustering, we need to obtain the predicted clusters for each observation by using the fit_predict method from sklearn.
# Create a variable which will contain the predicted clusters for each observation
identified_clusters = kmeans.fit_predict(x)
# Check the result
identified_clusters
# The result is an array containing the predicted clusters. There are 2 clusters indicated by 0 and 1.
# Create a copy of the data
data_with_clusters = data.copy()
# Create a new Series, containing the identified cluster for each observation
data_with_clusters['Cluster'] = identified_clusters
# Check the result
data_with_clusters
# Plot the data using the longitude and the latitude
# c (color) is an argument which could be coded with a variable
# The variable in this case has values 0,1,2, indicating to plt.scatter, that there are three colors (0,1,2)
# All points in cluster 0 will be the same colour, all points in cluster 1 - another one, etc.
# cmap is the color map. Rainbow is a nice one, but you can check others here: https://matplotlib.org/users/colormaps.html
plt.scatter(data_with_clusters['Longitude'],data_with_clusters['Latitude'],c=data_with_clusters['Cluster'],cmap='rainbow')
plt.xlim(-180,180)
plt.ylim(-90,90)
plt.show()
```
#Challenges
**Challenge 1**
```
#Increase the number of clusterds in the above example to see the difference in the scatter plot
```
**Challenge 2**
You are given the following dataset ([Dataset Download](https://drive.google.com/file/d/1y6I2BZnYvFIcgyHpuNHnUAL2v2FIEgwr/view?usp=sharing)) and you are required to group all the countries into 2 clusters.
Try with other numbers of clusters and see if they match your expectations.
Plot the data using the c parameter to separate the data by the clusters we defined.
Note: c stands for color
```
# Your code goes here
```
| github_jupyter |
## NLP Sequence Classification using LSTM Recurrent Neural Network
This exercise reproduce Jason Browlee's post:
https://machinelearningmastery.com/sequence-classification-lstm-recurrent-neural-networks-python-keras/
Sequence classification is a predictive modeling problem where you have some sequence of inputs over space or time and the task is to predict a category for the sequence.
What makes this problem difficult is that the sequences can vary in length, be comprised of a very large vocabulary of input symbols and may require the model to learn the long-term context or dependencies between symbols in the input sequence.
This exercise shows:
1. How to develop an LSTM model for a sequence classification problem.
2. How to reduce overfitting in the LSTM models through the use of dropout.
3. How to combine LSTM models with Convolutional Neural Networks that excel at learning spatial relationships.
### How to modify this code for other dataset
To classify multiple classes of text based data, change dense layer's loss function from *binary_crossentropy* to *categorical_crossentropy*
To output probability of each class, instead of 1 or 0, change dense layer's activation function from *sigmoid* to *softmax*
### Reference
https://machinelearningmastery.com/sequence-classification-lstm-recurrent-neural-networks-python-keras/
```
import numpy
from keras.datasets import imdb
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM
from keras.layers.embeddings import Embedding
from keras.preprocessing import sequence
from keras.utils.vis_utils import plot_model
# fix random seed for reproducibility
numpy.random.seed(7)
output_dir = 'output/lstm/'
```
#### Step 1. Load the IMDB dataset.
This exercise uses IMDB movie review sentiment dataset. Each movie review is a variable sequence of words and the sentiment of each movie review is to be classified. Keras provides access to the IMDB dataset built-in. The imdb.load_data() function allows you to load the dataset in a format that is ready for use in neural network and deep learning models.
Constrain the dataset to the top 5,000 words, also split the dataset into train (50%) and test (50%) sets.
```
# load the dataset but only keep the top n words, zero the rest
top_words = 5000
(X_train, y_train), (X_test, y_test) = imdb.load_data(num_words=top_words)
```
#### Step 2. truncate and pad the input sequences
Truncate and pad the input sequences so that they are all the same length for modeling. The model will learn the zero values carry no information so indeed the sequences are not the same length in terms of content, but same length vectors is required to perform the computation in Keras.
```
# truncate and pad input sequences
max_review_length = 500
X_train = sequence.pad_sequences(X_train, maxlen=max_review_length)
X_test = sequence.pad_sequences(X_test, maxlen=max_review_length)
```
#### Step 3. Define, compile and fit LSTM model.
The first layer is the Embedded layer that uses 32 length vectors to represent each word. The next layer is the LSTM layer with 100 memory units (smart neurons). Finally, because this is a classification problem we use a Dense output layer with a single neuron and a sigmoid activation function to make 0 or 1 predictions for the two classes (good and bad) in the problem.
Because it is a binary classification problem, log loss is used as the loss function (binary_crossentropy in Keras). The efficient ADAM optimization algorithm is used. The model is fit for only 2 epochs because it quickly overfits the problem. A large batch size of 64 reviews is used to space out weight updates.
```
# create the model
embedding_vecor_length = 32
model = Sequential()
model.add(Embedding(top_words, embedding_vecor_length, input_length=max_review_length))
model.add(LSTM(100))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
print(model.summary())
model.fit(X_train, y_train, validation_data=(X_test, y_test), epochs=3, batch_size=64)
```
### Step 4. Estimate the performance of the model on unseen reviews.
```
# Final evaluation of the model
scores = model.evaluate(X_test, y_test, verbose=0)
print("Accuracy: %.2f%%" % (scores[1]*100))
```
### Step 5. Solve overfitting problem using dropout
RNN such as LSTM generally have the problem of overfitting. Dropout can be applied to solve the overfitting problem.
The following is an example of applying dropout between layers using Keras Dropout layer:
model = Sequential()
model.add(Embedding(top_words, embedding_vecor_length, input_length=max_review_length))
model.add(Dropout(0.2))
model.add(LSTM(100))
model.add(Dropout(0.2))
model.add(Dense(1, activation='sigmoid'))
To apply the gate specific dropout on the input and recurrent connections of the memory units of the LSTM, use the following:
model = Sequential()
model.add(Embedding(top_words, embedding_vecor_length, input_length=max_review_length))
model.add(LSTM(100, dropout=0.2, recurrent_dropout=0.2))
model.add(Dense(1, activation='sigmoid'))
```
# LSTM with Dropout for sequence classification in the IMDB dataset
import numpy
from keras.datasets import imdb
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM
from keras.layers import Dropout
from keras.layers.embeddings import Embedding
from keras.preprocessing import sequence
# fix random seed for reproducibility
numpy.random.seed(7)
# load the dataset but only keep the top n words, zero the rest
top_words = 5000
(X_train, y_train), (X_test, y_test) = imdb.load_data(num_words=top_words)
# truncate and pad input sequences
max_review_length = 500
X_train = sequence.pad_sequences(X_train, maxlen=max_review_length)
X_test = sequence.pad_sequences(X_test, maxlen=max_review_length)
# create the model
embedding_vecor_length = 32
model = Sequential()
model.add(Embedding(top_words, embedding_vecor_length, input_length=max_review_length))
model.add(Dropout(0.2))
model.add(LSTM(100, dropout=0.2, recurrent_dropout=0.2))
model.add(Dropout(0.2))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
print(model.summary())
model.fit(X_train, y_train, epochs=3, batch_size=64)
# Final evaluation of the model
scores = model.evaluate(X_test, y_test, verbose=0)
print("Accuracy: %.2f%%" % (scores[1]*100))
```
### Step 6 CNN + LSTM
Convolutional Neural Networks excel at learning the spatial structure in input data.
The IMDB review data has a one-dimensional spatial structure in the sequence of workds in reviews, and CNN should be good at picking out the features. This learned spatial features may then be learned as sequences by an LSTM layer.
We can easily add a one-dimensional CNN and max pooling layers after the Embedding layer which then feed the consolidated features to the LSTM. We can use a smallish set of 32 features with a small filter length of 3. The pooling layer can use the standard length of 2 to halve the feature map size.
For example, we would create the model as follows:
model = Sequential()
model.add(Embedding(top_words, embedding_vecor_length, input_length=max_review_length))
model.add(Conv1D(filters=32, kernel_size=3, padding='same', activation='relu'))
model.add(MaxPooling1D(pool_size=2))
model.add(LSTM(100))
model.add(Dense(1, activation='sigmoid'))
```
# LSTM and CNN for sequence classification in the IMDB dataset
import numpy
from keras.datasets import imdb
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM
from keras.layers.convolutional import Conv1D
from keras.layers.convolutional import MaxPooling1D
from keras.layers.embeddings import Embedding
from keras.preprocessing import sequence
# fix random seed for reproducibility
numpy.random.seed(7)
# load the dataset but only keep the top n words, zero the rest
top_words = 5000
(X_train, y_train), (X_test, y_test) = imdb.load_data(num_words=top_words)
# truncate and pad input sequences
max_review_length = 500
X_train = sequence.pad_sequences(X_train, maxlen=max_review_length)
X_test = sequence.pad_sequences(X_test, maxlen=max_review_length)
# create the model
embedding_vecor_length = 32
model = Sequential()
model.add(Embedding(top_words, embedding_vecor_length, input_length=max_review_length))
model.add(Conv1D(filters=32, kernel_size=3, padding='same', activation='relu'))
model.add(MaxPooling1D(pool_size=2))
model.add(LSTM(100, dropout=0.2, recurrent_dropout=0.2))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
print(model.summary())
model.fit(X_train, y_train, epochs=3, batch_size=64)
# Final evaluation of the model
scores = model.evaluate(X_test, y_test, verbose=0)
print("Accuracy: %.2f%%" % (scores[1]*100))
```
### Step 7. With Dropout layers, and increase epoch
```
# LSTM and CNN for sequence classification in the IMDB dataset
import numpy
from keras.datasets import imdb
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM
from keras.layers.convolutional import Conv1D
from keras.layers.convolutional import MaxPooling1D
from keras.layers.embeddings import Embedding
from keras.preprocessing import sequence
# fix random seed for reproducibility
numpy.random.seed(7)
# load the dataset but only keep the top n words, zero the rest
top_words = 5000
(X_train, y_train), (X_test, y_test) = imdb.load_data(num_words=top_words)
# truncate and pad input sequences
max_review_length = 500
X_train = sequence.pad_sequences(X_train, maxlen=max_review_length)
X_test = sequence.pad_sequences(X_test, maxlen=max_review_length)
# create the model
embedding_vecor_length = 32
model = Sequential()
model.add(Embedding(top_words, embedding_vecor_length, input_length=max_review_length))
model.add(Conv1D(filters=32, kernel_size=3, padding='same', activation='relu'))
model.add(Dropout(0.2))
model.add(MaxPooling1D(pool_size=2))
model.add(LSTM(100, dropout=0.2, recurrent_dropout=0.2))
model.add(Dropout(0.2))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
print(model.summary())
plot_model(model, show_shapes=True, to_file=output_dir+'lstm.png')
model.fit(X_train, y_train, epochs=5, batch_size=64)
model.save(output_dir+'lstm_model.h5')
# Final evaluation of the model
scores = model.evaluate(X_test, y_test, verbose=0)
print("Accuracy: %.2f%%" % (scores[1]*100))
```
<img src='output/lstm/lstm.png'/>
| github_jupyter |
# Challenge 014 - Implement Queues using Stacks
This challenge is taken from LeetCode - 232. Implement Queue using Stacks (https://leetcode.com/problems/implement-queue-using-stacks/).
## Problem
Implement a first in first out (FIFO) queue using only two stacks. The implemented queue should support all the functions of a normal queue (push, peek, pop, and empty).
Implement the MyQueue class:
- void push(int x) Pushes element x to the back of the queue.
- int pop() Removes the element from the front of the queue and returns it.
- int peek() Returns the element at the front of the queue.
- boolean empty() Returns true if the queue is empty, false otherwise.
Notes:
- You must use only standard operations of a stack, which means only push to top, peek/pop from top, size, and is empty operations are valid.
- Depending on your language, the stack may not be supported natively. You may simulate a stack using a list or deque (double-ended queue) as long as you use only a stack's standard operations.
Follow-up: Can you implement the queue such that each operation is amortized O(1) time complexity? In other words, performing n operations will take overall O(n) time even if one of those operations may take longer.
Example 1:
Input
```
["MyQueue", "push", "push", "peek", "pop", "empty"]
[[], [1], [2], [], [], []]
```
Output
```
[null, null, null, 1, 1, false]
```
Explanation
```
MyQueue myQueue = new MyQueue();
myQueue.push(1); // queue is: [1]
myQueue.push(2); // queue is: [1, 2] (leftmost is front of the queue)
myQueue.peek(); // return 1
myQueue.pop(); // return 1, queue is [2]
myQueue.empty(); // return false
```
Constraints:
```
1 <= x <= 9
At most 100 calls will be made to push, pop, peek, and empty.
All the calls to pop and peek are valid.
```
## Solution
```
class MyQueue(object):
def __init__(self):
"""
Initialize your data structure here.
"""
self.array = []
def push(self, x):
"""
Push element x to the back of queue.
:type x: int
:rtype: None
"""
self.array.append(x)
def pop(self):
"""
Removes the element from in front of queue and returns that element.
:rtype: int
"""
if len(self.array) > 0:
temp = self.array[0]
self.array.pop(0)
return temp
return 0
def peek(self):
"""
Get the front element.
:rtype: int
"""
if (len(self.array) > 0):
print(self.array)
return self.array[0]
else:
0
def empty(self):
"""
Returns whether the queue is empty.
:rtype: bool
"""
if len(self.array) == 0:
return True
return False
```
| github_jupyter |
<a href="https://colab.research.google.com/github/mrdbourke/tensorflow-deep-learning/blob/main/09_SkimLit_nlp_milestone_project_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Milestone Project 2: SkimLit 📄🔥
In the previous notebook ([NLP fundamentals in TensorFlow](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/08_introduction_to_nlp_in_tensorflow.ipynb)), we went through some fundamental natural lanuage processing concepts. The main ones being **tokenzation** (turning words into numbers) and **creating embeddings** (creating a numerical representation of words).
In this project, we're going to be putting what we've learned into practice.
More specificially, we're going to be replicating the deep learning model behind the 2017 paper [*PubMed 200k RCT: a Dataset for Sequenctial Sentence Classification in Medical Abstracts*](https://arxiv.org/abs/1710.06071).
When it was released, the paper presented a new dataset called PubMed 200k RCT which consists of ~200,000 labelled Randomized Controlled Trial (RCT) abstracts.
The goal of the dataset was to explore the ability for NLP models to classify sentences which appear in sequential order.
In other words, given the abstract of a RCT, what role does each sentence serve in the abstract?

*Example inputs ([harder to read abstract from PubMed](https://pubmed.ncbi.nlm.nih.gov/28942748/)) and outputs ([easier to read abstract](https://pubmed.ncbi.nlm.nih.gov/32537182/)) of the model we're going to build. The model will take an abstract wall of text and predict the section label each sentence should have.*
### Model Input
For example, can we train an NLP model which takes the following input (note: the following sample has had all numerical symbols replaced with "@"):
> To investigate the efficacy of @ weeks of daily low-dose oral prednisolone in improving pain , mobility , and systemic low-grade inflammation in the short term and whether the effect would be sustained at @ weeks in older adults with moderate to severe knee osteoarthritis ( OA ). A total of @ patients with primary knee OA were randomized @:@ ; @ received @ mg/day of prednisolone and @ received placebo for @ weeks. Outcome measures included pain reduction and improvement in function scores and systemic inflammation markers. Pain was assessed using the visual analog pain scale ( @-@ mm ).
Secondary outcome measures included the Western Ontario and McMaster Universities Osteoarthritis Index scores , patient global assessment ( PGA ) of the severity of knee OA , and @-min walk distance ( @MWD ).,
Serum levels of interleukin @ ( IL-@ ) , IL-@ , tumor necrosis factor ( TNF ) - , and high-sensitivity C-reactive protein ( hsCRP ) were measured.
There was a clinically relevant reduction in the intervention group compared to the placebo group for knee pain , physical function , PGA , and @MWD at @ weeks. The mean difference between treatment arms ( @ % CI ) was @ ( @-@ @ ) , p < @ ; @ ( @-@ @ ) , p < @ ; @ ( @-@ @ ) , p < @ ; and @ ( @-@ @ ) , p < @ , respectively. Further , there was a clinically relevant reduction in the serum levels of IL-@ , IL-@ , TNF - , and hsCRP at @ weeks in the intervention group when compared to the placebo group. These differences remained significant at @ weeks. The Outcome Measures in Rheumatology Clinical Trials-Osteoarthritis Research Society International responder rate was @ % in the intervention group and @ % in the placebo group ( p < @ ). Low-dose oral prednisolone had both a short-term and a longer sustained effect resulting in less knee pain , better physical function , and attenuation of systemic inflammation in older patients with knee OA ( ClinicalTrials.gov identifier NCT@ ).
### Model output
And returns the following output:
```
['###24293578\n',
'OBJECTIVE\tTo investigate the efficacy of @ weeks of daily low-dose oral prednisolone in improving pain , mobility , and systemic low-grade inflammation in the short term and whether the effect would be sustained at @ weeks in older adults with moderate to severe knee osteoarthritis ( OA ) .\n',
'METHODS\tA total of @ patients with primary knee OA were randomized @:@ ; @ received @ mg/day of prednisolone and @ received placebo for @ weeks .\n',
'METHODS\tOutcome measures included pain reduction and improvement in function scores and systemic inflammation markers .\n',
'METHODS\tPain was assessed using the visual analog pain scale ( @-@ mm ) .\n',
'METHODS\tSecondary outcome measures included the Western Ontario and McMaster Universities Osteoarthritis Index scores , patient global assessment ( PGA ) of the severity of knee OA , and @-min walk distance ( @MWD ) .\n',
'METHODS\tSerum levels of interleukin @ ( IL-@ ) , IL-@ , tumor necrosis factor ( TNF ) - , and high-sensitivity C-reactive protein ( hsCRP ) were measured .\n',
'RESULTS\tThere was a clinically relevant reduction in the intervention group compared to the placebo group for knee pain , physical function , PGA , and @MWD at @ weeks .\n',
'RESULTS\tThe mean difference between treatment arms ( @ % CI ) was @ ( @-@ @ ) , p < @ ; @ ( @-@ @ ) , p < @ ; @ ( @-@ @ ) , p < @ ; and @ ( @-@ @ ) , p < @ , respectively .\n',
'RESULTS\tFurther , there was a clinically relevant reduction in the serum levels of IL-@ , IL-@ , TNF - , and hsCRP at @ weeks in the intervention group when compared to the placebo group .\n',
'RESULTS\tThese differences remained significant at @ weeks .\n',
'RESULTS\tThe Outcome Measures in Rheumatology Clinical Trials-Osteoarthritis Research Society International responder rate was @ % in the intervention group and @ % in the placebo group ( p < @ ) .\n',
'CONCLUSIONS\tLow-dose oral prednisolone had both a short-term and a longer sustained effect resulting in less knee pain , better physical function , and attenuation of systemic inflammation in older patients with knee OA ( ClinicalTrials.gov identifier NCT@ ) .\n',
'\n']
```
### Problem in a sentence
The number of RCT papers released is continuing to increase, those without structured abstracts can be hard to read and in turn slow down researchers moving through the literature.
### Solution in a sentence
Create an NLP model to classify abstract sentences into the role they play (e.g. objective, methods, results, etc) to enable researchers to skim through the literature (hence SkimLit 🤓🔥) and dive deeper when necessary.
> 📖 **Resources:** Before going through the code in this notebook, you might want to get a background of what we're going to be doing. To do so, spend an hour (or two) going through the following papers and then return to this notebook:
1. Where our data is coming from: [*PubMed 200k RCT: a Dataset for Sequential Sentence Classification in Medical Abstracts*](https://arxiv.org/abs/1710.06071)
2. Where our model is coming from: [*Neural networks for joint sentence
classification in medical paper abstracts*](https://arxiv.org/pdf/1612.05251.pdf).
## What we're going to cover
Time to take what we've learned in the NLP fundmentals notebook and build our biggest NLP model yet:
* Downloading a text dataset ([PubMed RCT200k from GitHub](https://github.com/Franck-Dernoncourt/pubmed-rct))
* Writing a preprocessing function to prepare our data for modelling
* Setting up a series of modelling experiments
* Making a baseline (TF-IDF classifier)
* Deep models with different combinations of: token embeddings, character embeddings, pretrained embeddings, positional embeddings
* Building our first multimodal model (taking multiple types of data inputs)
* Replicating the model architecture from https://arxiv.org/pdf/1612.05251.pdf
* Find the most wrong predictions
* Making predictions on PubMed abstracts from the wild
## How you should approach this notebook
You can read through the descriptions and the code (it should all run, except for the cells which error on purpose), but there's a better option.
Write all of the code yourself.
Yes. I'm serious. Create a new notebook, and rewrite each line by yourself. Investigate it, see if you can break it, why does it break?
You don't have to write the text descriptions but writing the code yourself is a great way to get hands-on experience.
Don't worry if you make mistakes, we all do. The way to get better and make less mistakes is to write more code.
> 📖 **Resource:** See the full set of course materials on GitHub: https://github.com/mrdbourke/tensorflow-deep-learning
## Confirm access to a GPU
Since we're going to be building deep learning models, let's make sure we have a GPU.
In Google Colab, you can set this up by going to Runtime -> Change runtime type -> Hardware accelerator -> GPU.
If you don't have access to a GPU, the models we're building here will likely take up to 10x longer to run.
```
# Check for GPU
!nvidia-smi -L
```
## Get data
Before we can start building a model, we've got to download the PubMed 200k RCT dataset.
In a phenomenal act of kindness, the authors of the paper have made the data they used for their research availably publically and for free in the form of .txt files [on GitHub](https://github.com/Franck-Dernoncourt/pubmed-rct).
We can copy them to our local directory using `git clone https://github.com/Franck-Dernoncourt/pubmed-rct`.
```
!git clone https://github.com/Franck-Dernoncourt/pubmed-rct.git
!ls pubmed-rct
```
Checking the contents of the downloaded repository, you can see there are four folders.
Each contains a different version of the PubMed 200k RCT dataset.
Looking at the [README file](https://github.com/Franck-Dernoncourt/pubmed-rct) from the GitHub page, we get the following information:
* PubMed 20k is a subset of PubMed 200k. I.e., any abstract present in PubMed 20k is also present in PubMed 200k.
* `PubMed_200k_RCT` is the same as `PubMed_200k_RCT_numbers_replaced_with_at_sign`, except that in the latter all numbers had been replaced by `@`. (same for `PubMed_20k_RCT` vs. `PubMed_20k_RCT_numbers_replaced_with_at_sign`).
* Since Github file size limit is 100 MiB, we had to compress `PubMed_200k_RCT\train.7z` and `PubMed_200k_RCT_numbers_replaced_with_at_sign\train.zip`. To uncompress `train.7z`, you may use 7-Zip on Windows, Keka on Mac OS X, or p7zip on Linux.
To begin with, the dataset we're going to be focused on is `PubMed_20k_RCT_numbers_replaced_with_at_sign`.
Why this one?
Rather than working with the whole 200k dataset, we'll keep our experiments quick by starting with a smaller subset. We could've chosen the dataset with numbers instead of having them replaced with `@` but we didn't.
Let's check the file contents.
```
# Check what files are in the PubMed_20K dataset
!ls pubmed-rct/PubMed_20k_RCT_numbers_replaced_with_at_sign
```
Beautiful, looks like we've got three separate text files:
* `train.txt` - training samples.
* `dev.txt` - dev is short for development set, which is another name for validation set (in our case, we'll be using and referring to this file as our validation set).
* `test.txt` - test samples.
To save ourselves typing out the filepath to our target directory each time, let's turn it into a variable.
```
# Start by using the 20k dataset
data_dir = "pubmed-rct/PubMed_20k_RCT_numbers_replaced_with_at_sign/"
# Check all of the filenames in the target directory
import os
filenames = [data_dir + filename for filename in os.listdir(data_dir)]
filenames
```
## Preprocess data
Okay, now we've downloaded some text data, do you think we're ready to model it?
Wait...
We've downloaded the data but we haven't even looked at it yet.
What's the motto for getting familiar with any new dataset?
I'll give you a clue, the word begins with "v" and we say it three times.
> Vibe, vibe, vibe?
Sort of... we've definitely got to the feel the vibe of our data.
> Values, values, values?
Right again, we want to *see* lots of values but not quite what we're looking for.
> Visualize, visualize, visualize?
Boom! That's it. To get familiar and understand how we have to prepare our data for our deep learning models, we've got to visualize it.
Because our data is in the form of text files, let's write some code to read each of the lines in a target file.
```
# Create function to read the lines of a document
def get_lines(filename):
"""
Reads filename (a text file) and returns the lines of text as a list.
Args:
filename: a string containing the target filepath to read.
Returns:
A list of strings with one string per line from the target filename.
For example:
["this is the first line of filename",
"this is the second line of filename",
"..."]
"""
with open(filename, "r") as f:
return f.readlines()
```
Alright, we've got a little function, `get_lines()` which takes the filepath of a text file, opens it, reads each of the lines and returns them.
Let's try it out on the training data (`train.txt`).
```
train_lines = get_lines(data_dir+"train.txt")
train_lines[:20] # the whole first example of an abstract + a little more of the next one
```
Reading the lines from the training text file results in a list of strings containing different abstract samples, the sentences in a sample along with the role the sentence plays in the abstract.
The role of each sentence is prefixed at the start of each line separated by a tab (`\t`) and each sentence finishes with a new line (`\n`).
Different abstracts are separated by abstract ID's (lines beginning with `###`) and newlines (`\n`).
Knowing this, it looks like we've got a couple of steps to do to get our samples ready to pass as training data to our future machine learning model.
Let's write a function to perform the following steps:
* Take a target file of abstract samples.
* Read the lines in the target file.
* For each line in the target file:
* If the line begins with `###` mark it as an abstract ID and the beginning of a new abstract.
* Keep count of the number of lines in a sample.
* If the line begins with `\n` mark it as the end of an abstract sample.
* Keep count of the total lines in a sample.
* Record the text before the `\t` as the label of the line.
* Record the text after the `\t` as the text of the line.
* Return all of the lines in the target text file as a list of dictionaries containing the key/value pairs:
* `"line_number"` - the position of the line in the abstract (e.g. `3`).
* `"target"` - the role of the line in the abstract (e.g. `OBJECTIVE`).
* `"text"` - the text of the line in the abstract.
* `"total_lines"` - the total lines in an abstract sample (e.g. `14`).
* Abstract ID's and newlines should be omitted from the returned preprocessed data.
Example returned preprocessed sample (a single line from an abstract):
```
[{'line_number': 0,
'target': 'OBJECTIVE',
'text': 'to investigate the efficacy of @ weeks of daily low-dose oral prednisolone in improving pain , mobility , and systemic low-grade inflammation in the short term and whether the effect would be sustained at @ weeks in older adults with moderate to severe knee osteoarthritis ( oa ) .',
'total_lines': 11},
...]
```
```
def preprocess_text_with_line_numbers(filename):
"""Returns a list of dictionaries of abstract line data.
Takes in filename, reads its contents and sorts through each line,
extracting things like the target label, the text of the sentence,
how many sentences are in the current abstract and what sentence number
the target line is.
Args:
filename: a string of the target text file to read and extract line data
from.
Returns:
A list of dictionaries each containing a line from an abstract,
the lines label, the lines position in the abstract and the total number
of lines in the abstract where the line is from. For example:
[{"target": 'CONCLUSION',
"text": The study couldn't have gone better, turns out people are kinder than you think",
"line_number": 8,
"total_lines": 8}]
"""
input_lines = get_lines(filename) # get all lines from filename
abstract_lines = "" # create an empty abstract
abstract_samples = [] # create an empty list of abstracts
# Loop through each line in target file
for line in input_lines:
if line.startswith("###"): # check to see if line is an ID line
abstract_id = line
abstract_lines = "" # reset abstract string
elif line.isspace(): # check to see if line is a new line
abstract_line_split = abstract_lines.splitlines() # split abstract into separate lines
# Iterate through each line in abstract and count them at the same time
for abstract_line_number, abstract_line in enumerate(abstract_line_split):
line_data = {} # create empty dict to store data from line
target_text_split = abstract_line.split("\t") # split target label from text
line_data["target"] = target_text_split[0] # get target label
line_data["text"] = target_text_split[1].lower() # get target text and lower it
line_data["line_number"] = abstract_line_number # what number line does the line appear in the abstract?
line_data["total_lines"] = len(abstract_line_split) - 1 # how many total lines are in the abstract? (start from 0)
abstract_samples.append(line_data) # add line data to abstract samples list
else: # if the above conditions aren't fulfilled, the line contains a labelled sentence
abstract_lines += line
return abstract_samples
```
Beautiful! That's one good looking function. Let's use it to preprocess each of our RCT 20k datasets.
```
# Get data from file and preprocess it
%%time
train_samples = preprocess_text_with_line_numbers(data_dir + "train.txt")
val_samples = preprocess_text_with_line_numbers(data_dir + "dev.txt") # dev is another name for validation set
test_samples = preprocess_text_with_line_numbers(data_dir + "test.txt")
len(train_samples), len(val_samples), len(test_samples)
```
How do our training samples look?
```
# Check the first abstract of our training data
train_samples[:14]
```
Fantastic! Looks like our `preprocess_text_with_line_numbers()` function worked great.
How about we turn our list of dictionaries into pandas DataFrame's so we visualize them better?
```
import pandas as pd
train_df = pd.DataFrame(train_samples)
val_df = pd.DataFrame(val_samples)
test_df = pd.DataFrame(test_samples)
train_df.head(14)
```
Now our data is in DataFrame form, we can perform some data analysis on it.
```
# Distribution of labels in training data
train_df.target.value_counts()
```
Looks like sentences with the `OBJECTIVE` label are the least common.
How about we check the distribution of our abstract lengths?
```
train_df.total_lines.plot.hist();
```
Okay, looks like most of the abstracts are around 7 to 15 sentences in length.
It's good to check these things out to make sure when we do train a model or test it on unseen samples, our results aren't outlandish.
### Get lists of sentences
When we build our deep learning model, one of its main inputs will be a list of strings (the lines of an abstract).
We can get these easily from our DataFrames by calling the `tolist()` method on our `"text"` columns.
```
# Convert abstract text lines into lists
train_sentences = train_df["text"].tolist()
val_sentences = val_df["text"].tolist()
test_sentences = test_df["text"].tolist()
len(train_sentences), len(val_sentences), len(test_sentences)
# View first 10 lines of training sentences
train_sentences[:10]
```
Alright, we've separated our text samples. As you might've guessed, we'll have to write code to convert the text to numbers before we can use it with our machine learning models, we'll get to this soon.
## Make numeric labels (ML models require numeric labels)
We're going to create one hot and label encoded labels.
We could get away with just making label encoded labels, however, TensorFlow's CategoricalCrossentropy loss function likes to have one hot encoded labels (this will enable us to use label smoothing later on).
To numerically encode labels we'll use Scikit-Learn's [`OneHotEncoder`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OneHotEncoder.html) and [`LabelEncoder`](http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.LabelEncoder.html) classes.
```
# One hot encode labels
from sklearn.preprocessing import OneHotEncoder
one_hot_encoder = OneHotEncoder(sparse=False)
train_labels_one_hot = one_hot_encoder.fit_transform(train_df["target"].to_numpy().reshape(-1, 1))
val_labels_one_hot = one_hot_encoder.transform(val_df["target"].to_numpy().reshape(-1, 1))
test_labels_one_hot = one_hot_encoder.transform(test_df["target"].to_numpy().reshape(-1, 1))
# Check what training labels look like
train_labels_one_hot
```
### Label encode labels
```
# Extract labels ("target" columns) and encode them into integers
from sklearn.preprocessing import LabelEncoder
label_encoder = LabelEncoder()
train_labels_encoded = label_encoder.fit_transform(train_df["target"].to_numpy())
val_labels_encoded = label_encoder.transform(val_df["target"].to_numpy())
test_labels_encoded = label_encoder.transform(test_df["target"].to_numpy())
# Check what training labels look like
train_labels_encoded
```
Now we've trained an instance of `LabelEncoder`, we can get the class names and number of classes using the `classes_` attribute.
```
# Get class names and number of classes from LabelEncoder instance
num_classes = len(label_encoder.classes_)
class_names = label_encoder.classes_
num_classes, class_names
```
## Creating a series of model experiments
We've proprocessed our data so now, in true machine learning fashion, it's time to setup a series of modelling experiments.
We'll start by creating a simple baseline model to obtain a score we'll try to beat by building more and more complex models as we move towards replicating the sequence model outlined in [*Neural networks for joint sentence
classification in medical paper abstracts*](https://arxiv.org/pdf/1612.05251.pdf).
For each model, we'll train it on the training data and evaluate it on the validation data.
## Model 0: Getting a baseline
Our first model we'll be a TF-IDF Multinomial Naive Bayes as recommended by [Scikit-Learn's machine learning map](https://scikit-learn.org/stable/tutorial/machine_learning_map/index.html).
To build it, we'll create a Scikit-Learn `Pipeline` which uses the [`TfidfVectorizer`](https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.TfidfVectorizer.html) class to convert our abstract sentences to numbers using the TF-IDF (term frequency-inverse document frequecy) algorithm and then learns to classify our sentences using the [`MultinomialNB`](https://scikit-learn.org/stable/modules/generated/sklearn.naive_bayes.MultinomialNB.html) aglorithm.
```
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.naive_bayes import MultinomialNB
from sklearn.pipeline import Pipeline
# Create a pipeline
model_0 = Pipeline([
("tf-idf", TfidfVectorizer()),
("clf", MultinomialNB())
])
# Fit the pipeline to the training data
model_0.fit(X=train_sentences,
y=train_labels_encoded);
```
Due to the speed of the Multinomial Naive Bayes algorithm, it trains very quickly.
We can evaluate our model's accuracy on the validation dataset using the `score()` method.
```
# Evaluate baseline on validation dataset
model_0.score(X=val_sentences,
y=val_labels_encoded)
```
Nice! Looks like 72.1% accuracy will be the number to beat with our deeper models.
Now let's make some predictions with our baseline model to further evaluate it.
```
# Make predictions
baseline_preds = model_0.predict(val_sentences)
baseline_preds
```
To evaluate our baseline's predictions, we'll import the `calculate_results()` function we created in the [previous notebook](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/08_introduction_to_nlp_in_tensorflow.ipynb) and added it to our [`helper_functions.py` script](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/extras/helper_functions.py) to compare them to the ground truth labels.
More specificially the `calculate_results()` function will help us obtain the following:
* Accuracy
* Precision
* Recall
* F1-score
### Download helper functions script
Let's get our `helper_functions.py` script we've been using to store helper functions we've created in previous notebooks.
```
# Download helper functions script
!wget https://raw.githubusercontent.com/mrdbourke/tensorflow-deep-learning/main/extras/helper_functions.py
```
Now we've got the helper functions script we can import the `caculate_results()` function and see how our baseline model went.
```
# Import calculate_results helper function
from helper_functions import calculate_results
# Calculate baseline results
baseline_results = calculate_results(y_true=val_labels_encoded,
y_pred=baseline_preds)
baseline_results
```
## Preparing our data for deep sequence models
Excellent! We've got a working baseline to try and improve upon.
But before we start building deeper models, we've got to create vectorization and embedding layers.
The vectorization layer will convert our text to numbers and the embedding layer will capture the relationships between those numbers.
To start creating our vectorization and embedding layers, we'll need to import the appropriate libraries (namely TensorFlow and NumPy).
```
import numpy as np
import tensorflow as tf
from tensorflow.keras import layers
```
Since we'll be turning our sentences into numbers, it's a good idea to figure out how many words are in each sentence.
When our model goes through our sentences, it works best when they're all the same length (this is important for creating batches of the same size tensors).
For example, if one sentence is eight words long and another is 29 words long, we want to pad the eight word sentence with zeros so it ends up being the same length as the 29 word sentence.
Let's write some code to find the average length of sentences in the training set.
```
# How long is each sentence on average?
sent_lens = [len(sentence.split()) for sentence in train_sentences]
avg_sent_len = np.mean(sent_lens)
avg_sent_len # return average sentence length (in tokens)
```
How about the distribution of sentence lengths?
```
# What's the distribution look like?
import matplotlib.pyplot as plt
plt.hist(sent_lens, bins=7);
```
Looks like the vast majority of sentences are between 0 and 50 tokens in length.
We can use NumPy's [`percentile`](https://numpy.org/doc/stable/reference/generated/numpy.percentile.html) to find the value which covers 95% of the sentence lengths.
```
# How long of a sentence covers 95% of the lengths?
output_seq_len = int(np.percentile(sent_lens, 95))
output_seq_len
```
Wonderful! It looks like 95% of the sentences in our training set have a length of 55 tokens or less.
When we create our tokenization layer, we'll use this value to turn all of our sentences into the same length. Meaning sentences with a length below 55 get padded with zeros and sentences with a length above 55 get truncated (words after 55 get cut off).
> 🤔 **Question:** Why 95%?
We could use the max sentence length of the sentences in the training set.
```
# Maximum sentence length in the training set
max(sent_lens)
```
However, since hardly any sentences even come close to the max length, it would mean the majority of the data we pass to our model would be zeros (sinces all sentences below the max length would get padded with zeros).
> 🔑 **Note:** The steps we've gone through are good practice when working with a text corpus for a NLP problem. You want to know how long your samples are and what the distribution of them is. See section 4 Data Analysis of the [PubMed 200k RCT paper](https://arxiv.org/pdf/1710.06071.pdf) for further examples.
### Create text vectorizer
Now we've got a little more information about our texts, let's create a way to turn it into numbers.
To do so, we'll use the [`TextVectorization`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/TextVectorization) layer from TensorFlow.
We'll keep all the parameters default except for `max_tokens` (the number of unique words in our dataset) and `output_sequence_length` (our desired output length for each vectorized sentence).
Section 3.2 of the [PubMed 200k RCT paper](https://arxiv.org/pdf/1710.06071.pdf) states the vocabulary size of the PubMed 20k dataset as 68,000. So we'll use that as our `max_tokens` parameter.
```
# How many words are in our vocabulary? (taken from 3.2 in https://arxiv.org/pdf/1710.06071.pdf)
max_tokens = 68000
```
And since discovered a sentence length of 55 covers 95% of the training sentences, we'll use that as our `output_sequence_length` parameter.
```
# Create text vectorizer
from tensorflow.keras.layers.experimental.preprocessing import TextVectorization
text_vectorizer = TextVectorization(max_tokens=max_tokens, # number of words in vocabulary
output_sequence_length=55) # desired output length of vectorized sequences
```
Great! Looks like our `text_vectorizer` is ready, let's adapt it to the training data (let it read the training data and figure out what number should represent what word) and then test it out.
```
# Adapt text vectorizer to training sentences
text_vectorizer.adapt(train_sentences)
# Test out text vectorizer
import random
target_sentence = random.choice(train_sentences)
print(f"Text:\n{target_sentence}")
print(f"\nLength of text: {len(target_sentence.split())}")
print(f"\nVectorized text:\n{text_vectorizer([target_sentence])}")
```
Cool, we've now got a way to turn our sequences into numbers.
> 🛠 **Exercise:** Try running the cell above a dozen or so times. What do you notice about sequences with a length less than 55?
Using the [`get_vocabulary()`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/TextVectorization) method of our `text_vectorizer` we can find out a few different tidbits about our text.
```
# How many words in our training vocabulary?
rct_20k_text_vocab = text_vectorizer.get_vocabulary()
print(f"Number of words in vocabulary: {len(rct_20k_text_vocab)}"),
print(f"Most common words in the vocabulary: {rct_20k_text_vocab[:5]}")
print(f"Least common words in the vocabulary: {rct_20k_text_vocab[-5:]}")
```
And if we wanted to figure out the configuration of our `text_vectorizer` we can use the `get_config()` method.
```
# Get the config of our text vectorizer
text_vectorizer.get_config()
```
### Create custom text embedding
Our `token_vectorization` layer maps the words in our text directly to numbers. However, this doesn't necessarily capture the relationships between those numbers.
To create a richer numerical representation of our text, we can use an **embedding**.
As our model learns (by going through many different examples of abstract sentences and their labels), it'll update its embedding to better represent the relationships between tokens in our corpus.
We can create a trainable embedding layer using TensorFlow's [`Embedding`](https://www.tensorflow.org/tutorials/text/word_embeddings) layer.
Once again, the main parameters we're concerned with here are the inputs and outputs of our `Embedding` layer.
The `input_dim` parameter defines the size of our vocabulary. And the `output_dim` parameter defines the dimension of the embedding output.
Once created, our embedding layer will take the integer outputs of our `text_vectorization` layer as inputs and convert them to feature vectors of size `output_dim`.
Let's see it in action.
```
# Create token embedding layer
token_embed = layers.Embedding(input_dim=len(rct_20k_text_vocab), # length of vocabulary
output_dim=128, # Note: different embedding sizes result in drastically different numbers of parameters to train
# Use masking to handle variable sequence lengths (save space)
mask_zero=True,
name="token_embedding")
# Show example embedding
print(f"Sentence before vectorization:\n{target_sentence}\n")
vectorized_sentence = text_vectorizer([target_sentence])
print(f"Sentence after vectorization (before embedding):\n{vectorized_sentence}\n")
embedded_sentence = token_embed(vectorized_sentence)
print(f"Sentence after embedding:\n{embedded_sentence}\n")
print(f"Embedded sentence shape: {embedded_sentence.shape}")
```
## Create datasets (as fast as possible)
We've gone through all the trouble of preprocessing our datasets to be used with a machine learning model, however, there are still a few steps we can use to make them work faster with our models.
Namely, the `tf.data` API provides methods which enable faster data loading.
> 📖 **Resource:** For best practices on data loading in TensorFlow, check out the following:
* [tf.data: Build TensorFlow input pipelines](https://www.tensorflow.org/guide/data)
* [Better performance with the tf.data API](https://www.tensorflow.org/guide/data_performance)
The main steps we'll want to use with our data is to turn it into a `PrefetchDataset` of batches.
Doing so we'll ensure TensorFlow loads our data onto the GPU as fast as possible, in turn leading to faster training time.
To create a batched `PrefetchDataset` we can use the methods [`batch()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#batch) and [`prefetch()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#prefetch), the parameter [`tf.data.AUTOTUNE`](https://www.tensorflow.org/api_docs/python/tf/data#AUTOTUNE) will also allow TensorFlow to determine the optimal amount of compute to use to prepare datasets.
```
# Turn our data into TensorFlow Datasets
train_dataset = tf.data.Dataset.from_tensor_slices((train_sentences, train_labels_one_hot))
valid_dataset = tf.data.Dataset.from_tensor_slices((val_sentences, val_labels_one_hot))
test_dataset = tf.data.Dataset.from_tensor_slices((test_sentences, test_labels_one_hot))
train_dataset
# Take the TensorSliceDataset's and turn them into prefetched batches
train_dataset = train_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
valid_dataset = valid_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
test_dataset = test_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
train_dataset
```
## Model 1: Conv1D with token embeddings
Alright, we've now got a way to numerically represent our text and labels, time to build a series of deep models to try and improve upon our baseline.
All of our deep models will follow a similar structure:
```
Input (text) -> Tokenize -> Embedding -> Layers -> Output (label probability)
```
The main component we'll be changing throughout is the `Layers` component. Because any modern deep NLP model requires text to be converted into an embedding before meaningful patterns can be discovered within.
The first model we're going to build is a 1-dimensional Convolutional Neural Network.
We're also going to be following the standard machine learning workflow of:
- Build model
- Train model
- Evaluate model (make predictions and compare to ground truth)
```
# Create 1D convolutional model to process sequences
inputs = layers.Input(shape=(1,), dtype=tf.string)
text_vectors = text_vectorizer(inputs) # vectorize text inputs
token_embeddings = token_embed(text_vectors) # create embedding
x = layers.Conv1D(64, kernel_size=5, padding="same", activation="relu")(token_embeddings)
x = layers.GlobalAveragePooling1D()(x) # condense the output of our feature vector
outputs = layers.Dense(num_classes, activation="softmax")(x)
model_1 = tf.keras.Model(inputs, outputs)
# Compile
model_1.compile(loss="categorical_crossentropy", # if your labels are integer form (not one hot) use sparse_categorical_crossentropy
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# Get summary of Conv1D model
model_1.summary()
```
Wonderful! We've got our first deep sequence model built and ready to go.
Checking out the model summary, you'll notice the majority of the trainable parameters are within the embedding layer. If we were to increase the size of the embedding (by increasing the `output_dim` parameter of the `Embedding` layer), the number of trainable parameters would increase dramatically.
It's time to fit our model to the training data but we're going to make a mindful change.
Since our training data contains nearly 200,000 sentences, fitting a deep model may take a while even with a GPU. So to keep our experiments swift, we're going to run them on a subset of the training dataset.
More specifically, we'll only use the first 10% of batches (about 18,000 samples) of the training set to train on and the first 10% of batches from the validation set to validate on.
> 🔑 **Note:** It's a standard practice in machine learning to test your models on smaller subsets of data first to make sure they work before scaling them to larger amounts of data. You should aim to run many smaller experiments rather than only a handful of large experiments. And since your time is limited, one of the best ways to run smaller experiments is to reduce the amount of data you're working with (10% of the full dataset is usually a good amount, as long as it covers a similar distribution).
```
# Fit the model
model_1_history = model_1.fit(train_dataset,
steps_per_epoch=int(0.1 * len(train_dataset)), # only fit on 10% of batches for faster training time
epochs=3,
validation_data=valid_dataset,
validation_steps=int(0.1 * len(valid_dataset))) # only validate on 10% of batches
```
Brilliant! We've got our first trained deep sequence model, and it didn't take too long (and if we didn't prefetch our batched data, it would've taken longer).
Time to make some predictions with our model and then evaluate them.
```
# Evaluate on whole validation dataset (we only validated on 10% of batches during training)
model_1.evaluate(valid_dataset)
# Make predictions (our model outputs prediction probabilities for each class)
model_1_pred_probs = model_1.predict(valid_dataset)
model_1_pred_probs
# Convert pred probs to classes
model_1_preds = tf.argmax(model_1_pred_probs, axis=1)
model_1_preds
# Calculate model_1 results
model_1_results = calculate_results(y_true=val_labels_encoded,
y_pred=model_1_preds)
model_1_results
```
## Model 2: Feature extraction with pretrained token embeddings
Training our own embeddings took a little while to run, slowing our experiments down.
Since we're moving towards replicating the model architecture in [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf), it mentions they used a [pretrained GloVe embedding](https://nlp.stanford.edu/projects/glove/) as a way to initialise their token embeddings.
To emulate this, let's see what results we can get with the [pretrained Universal Sentence Encoder embeddings from TensorFlow Hub](https://tfhub.dev/google/universal-sentence-encoder/4).
> 🔑 **Note:** We could use GloVe embeddings as per the paper but since we're working with TensorFlow, we'll use what's available from TensorFlow Hub (GloVe embeddings aren't). We'll save [using pretrained GloVe embeddings](https://keras.io/examples/nlp/pretrained_word_embeddings/) as an extension.
The model structure will look like:
```
Inputs (string) -> Pretrained embeddings from TensorFlow Hub (Universal Sentence Encoder) -> Layers -> Output (prediction probabilities)
```
You'll notice the lack of tokenization layer we've used in a previous model. This is because the Universal Sentence Encoder (USE) takes care of tokenization for us.
This type of model is called transfer learning, or more specifically, **feature extraction transfer learning**. In other words, taking the patterns a model has learned elsewhere and applying it to our own problem.
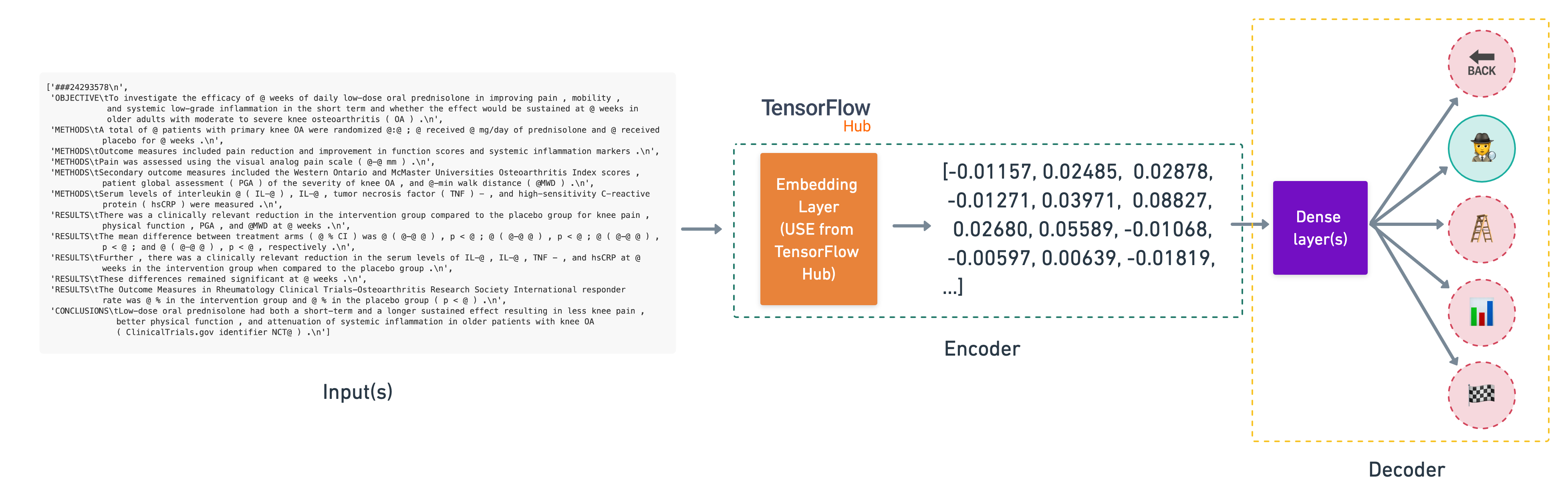
*The feature extractor model we're building using a pretrained embedding from TensorFlow Hub.*
To download the pretrained USE into a layer we can use in our model, we can use the [`hub.KerasLayer`](https://www.tensorflow.org/hub/api_docs/python/hub/KerasLayer) class.
We'll keep the pretrained embeddings frozen (by setting `trainable=False`) and add a trainable couple of layers on the top to tailor the model outputs to our own data.
> 🔑 **Note:** Due to having to download a relatively large model (~916MB), the cell below may take a little while to run.
```
# Download pretrained TensorFlow Hub USE
import tensorflow_hub as hub
tf_hub_embedding_layer = hub.KerasLayer("https://tfhub.dev/google/universal-sentence-encoder/4",
trainable=False,
name="universal_sentence_encoder")
```
Beautiful, now our pretrained USE is downloaded and instantiated as a `hub.KerasLayer` instance, let's test it out on a random sentence.
```
# Test out the embedding on a random sentence
random_training_sentence = random.choice(train_sentences)
print(f"Random training sentence:\n{random_training_sentence}\n")
use_embedded_sentence = tf_hub_embedding_layer([random_training_sentence])
print(f"Sentence after embedding:\n{use_embedded_sentence[0][:30]} (truncated output)...\n")
print(f"Length of sentence embedding:\n{len(use_embedded_sentence[0])}")
```
Nice! As we mentioned before the pretrained USE module from TensorFlow Hub takes care of tokenizing our text for us and outputs a 512 dimensional embedding vector.
Let's put together and compile a model using our `tf_hub_embedding_layer`.
### Building and fitting an NLP feature extraction model from TensorFlow Hub
```
# Define feature extractor model using TF Hub layer
inputs = layers.Input(shape=[], dtype=tf.string)
pretrained_embedding = tf_hub_embedding_layer(inputs) # tokenize text and create embedding
x = layers.Dense(128, activation="relu")(pretrained_embedding) # add a fully connected layer on top of the embedding
# Note: you could add more layers here if you wanted to
outputs = layers.Dense(5, activation="softmax")(x) # create the output layer
model_2 = tf.keras.Model(inputs=inputs,
outputs=outputs)
# Compile the model
model_2.compile(loss="categorical_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# Get a summary of the model
model_2.summary()
```
Checking the summary of our model we can see there's a large number of total parameters, however, the majority of these are non-trainable. This is because we set `training=False` when we instatiated our USE feature extractor layer.
So when we train our model, only the top two output layers will be trained.
```
# Fit feature extractor model for 3 epochs
model_2.fit(train_dataset,
steps_per_epoch=int(0.1 * len(train_dataset)),
epochs=3,
validation_data=valid_dataset,
validation_steps=int(0.1 * len(valid_dataset)))
# Evaluate on whole validation dataset
model_2.evaluate(valid_dataset)
```
Since we aren't training our own custom embedding layer, training is much quicker.
Let's make some predictions and evaluate our feature extraction model.
```
# Make predictions with feature extraction model
model_2_pred_probs = model_2.predict(valid_dataset)
model_2_pred_probs
# Convert the predictions with feature extraction model to classes
model_2_preds = tf.argmax(model_2_pred_probs, axis=1)
model_2_preds
# Calculate results from TF Hub pretrained embeddings results on validation set
model_2_results = calculate_results(y_true=val_labels_encoded,
y_pred=model_2_preds)
model_2_results
```
## Model 3: Conv1D with character embeddings
### Creating a character-level tokenizer
The [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf) paper mentions their model uses a hybrid of token and character embeddings.
We've built models with a custom token embedding and a pretrained token embedding, how about we build one using a character embedding?
The difference between a character and token embedding is that the **character embedding** is created using sequences split into characters (e.g. `hello` -> [`h`, `e`, `l`, `l`, `o`]) where as a **token embedding** is created on sequences split into tokens.
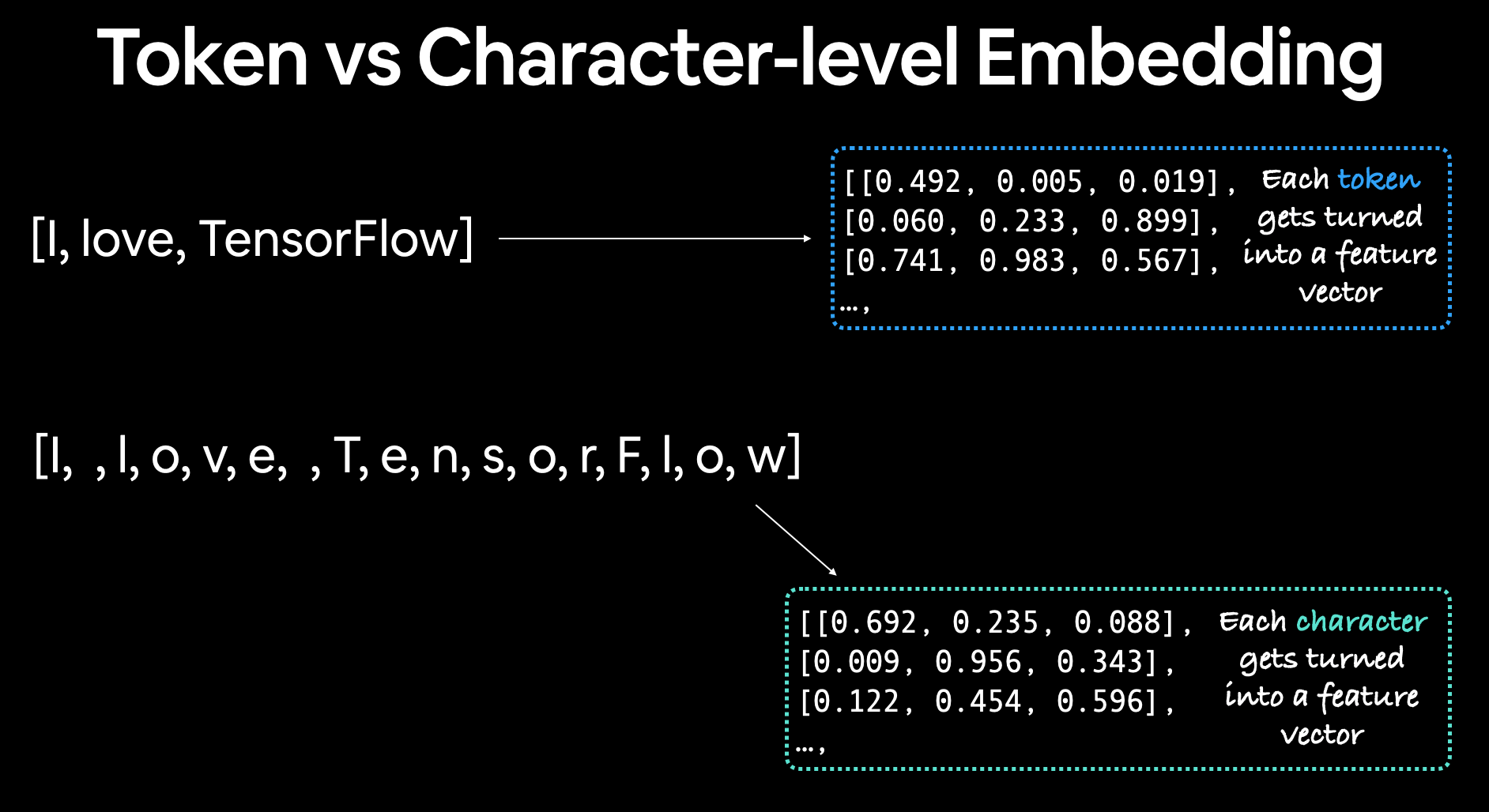

*Token level embeddings split sequences into tokens (words) and embeddings each of them, character embeddings split sequences into characters and creates a feature vector for each.*
We can create a character-level embedding by first vectorizing our sequences (after they've been split into characters) using the [`TextVectorization`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/TextVectorization) class and then passing those vectorized sequences through an [`Embedding`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Embedding) layer.
Before we can vectorize our sequences on a character-level we'll need to split them into characters. Let's write a function to do so.
```
# Make function to split sentences into characters
def split_chars(text):
return " ".join(list(text))
# Test splitting non-character-level sequence into characters
split_chars(random_training_sentence)
```
Great! Looks like our character-splitting function works. Let's create character-level datasets by splitting our sequence datasets into characters.
```
# Split sequence-level data splits into character-level data splits
train_chars = [split_chars(sentence) for sentence in train_sentences]
val_chars = [split_chars(sentence) for sentence in val_sentences]
test_chars = [split_chars(sentence) for sentence in test_sentences]
print(train_chars[0])
```
To figure out how long our vectorized character sequences should be, let's check the distribution of our character sequence lengths.
```
# What's the average character length?
char_lens = [len(sentence) for sentence in train_sentences]
mean_char_len = np.mean(char_lens)
mean_char_len
# Check the distribution of our sequences at character-level
import matplotlib.pyplot as plt
plt.hist(char_lens, bins=7);
```
Okay, looks like most of our sequences are between 0 and 200 characters long.
Let's use NumPy's percentile to figure out what length covers 95% of our sequences.
```
# Find what character length covers 95% of sequences
output_seq_char_len = int(np.percentile(char_lens, 95))
output_seq_char_len
```
Wonderful, now we know the sequence length which covers 95% of sequences, we'll use that in our `TextVectorization` layer as the `output_sequence_length` parameter.
> 🔑 **Note:** You can experiment here to figure out what the optimal `output_sequence_length` should be, perhaps using the mean results in as good results as using the 95% percentile.
We'll set `max_tokens` (the total number of different characters in our sequences) to 28, in other words, 26 letters of the alphabet + space + OOV (out of vocabulary or unknown) tokens.
```
# Get all keyboard characters for char-level embedding
import string
alphabet = string.ascii_lowercase + string.digits + string.punctuation
alphabet
# Create char-level token vectorizer instance
NUM_CHAR_TOKENS = len(alphabet) + 2 # num characters in alphabet + space + OOV token
char_vectorizer = TextVectorization(max_tokens=NUM_CHAR_TOKENS,
output_sequence_length=output_seq_char_len,
standardize="lower_and_strip_punctuation",
name="char_vectorizer")
# Adapt character vectorizer to training characters
char_vectorizer.adapt(train_chars)
```
Nice! Now we've adapted our `char_vectorizer` to our character-level sequences, let's check out some characteristics about it using the [`get_vocabulary()`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/TextVectorization#get_vocabulary) method.
```
# Check character vocabulary characteristics
char_vocab = char_vectorizer.get_vocabulary()
print(f"Number of different characters in character vocab: {len(char_vocab)}")
print(f"5 most common characters: {char_vocab[:5]}")
print(f"5 least common characters: {char_vocab[-5:]}")
```
We can also test it on random sequences of characters to make sure it's working.
```
# Test out character vectorizer
random_train_chars = random.choice(train_chars)
print(f"Charified text:\n{random_train_chars}")
print(f"\nLength of chars: {len(random_train_chars.split())}")
vectorized_chars = char_vectorizer([random_train_chars])
print(f"\nVectorized chars:\n{vectorized_chars}")
print(f"\nLength of vectorized chars: {len(vectorized_chars[0])}")
```
You'll notice sequences with a length shorter than 290 (`output_seq_char_length`) get padded with zeros on the end, this ensures all sequences passed to our model are the same length.
Also, due to the `standardize` parameter of `TextVectorization` being `"lower_and_strip_punctuation"` and the `split` parameter being `"whitespace"` by default, symbols (such as `@`) and spaces are removed.
> 🔑 **Note:** If you didn't want punctuation to be removed (keep the `@`, `%` etc), you can create a custom standardization callable and pass it as the `standardize` parameter. See the [`TextVectorization`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/TextVectorization) class documentation for more.
### Creating a character-level embedding
We've got a way to vectorize our character-level sequences, now's time to create a character-level embedding.
Just like our custom token embedding, we can do so using the [`tensorflow.keras.layers.Embedding`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Embedding) class.
Our character-level embedding layer requires an input dimension and output dimension.
The input dimension (`input_dim`) will be equal to the number of different characters in our `char_vocab` (28). And since we're following the structure of the model in Figure 1 of [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf), the output dimension of the character embedding (`output_dim`) will be 25.
```
# Create char embedding layer
char_embed = layers.Embedding(input_dim=NUM_CHAR_TOKENS, # number of different characters
output_dim=25, # embedding dimension of each character (same as Figure 1 in https://arxiv.org/pdf/1612.05251.pdf)
mask_zero=True,
name="char_embed")
# Test out character embedding layer
print(f"Charified text (before vectorization and embedding):\n{random_train_chars}\n")
char_embed_example = char_embed(char_vectorizer([random_train_chars]))
print(f"Embedded chars (after vectorization and embedding):\n{char_embed_example}\n")
print(f"Character embedding shape: {char_embed_example.shape}")
```
Wonderful! Each of the characters in our sequences gets turned into a 25 dimension embedding.
### Building a Conv1D model to fit on character embeddings
Now we've got a way to turn our character-level sequences into numbers (`char_vectorizer`) as well as numerically represent them as an embedding (`char_embed`) let's test how effective they are at encoding the information in our sequences by creating a character-level sequence model.
The model will have the same structure as our custom token embedding model (`model_1`) except it'll take character-level sequences as input instead of token-level sequences.
```
Input (character-level text) -> Tokenize -> Embedding -> Layers (Conv1D, GlobalMaxPool1D) -> Output (label probability)
```
```
# Make Conv1D on chars only
inputs = layers.Input(shape=(1,), dtype="string")
char_vectors = char_vectorizer(inputs)
char_embeddings = char_embed(char_vectors)
x = layers.Conv1D(64, kernel_size=5, padding="same", activation="relu")(char_embeddings)
x = layers.GlobalMaxPool1D()(x)
outputs = layers.Dense(num_classes, activation="softmax")(x)
model_3 = tf.keras.Model(inputs=inputs,
outputs=outputs,
name="model_3_conv1D_char_embedding")
# Compile model
model_3.compile(loss="categorical_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# Check the summary of conv1d_char_model
model_3.summary()
```
Before fitting our model on the data, we'll create char-level batched `PrefetchedDataset`'s.
```
# Create char datasets
train_char_dataset = tf.data.Dataset.from_tensor_slices((train_chars, train_labels_one_hot)).batch(32).prefetch(tf.data.AUTOTUNE)
val_char_dataset = tf.data.Dataset.from_tensor_slices((val_chars, val_labels_one_hot)).batch(32).prefetch(tf.data.AUTOTUNE)
train_char_dataset
```
Just like our token-level sequence model, to save time with our experiments, we'll fit the character-level model on 10% of batches.
```
# Fit the model on chars only
model_3_history = model_3.fit(train_char_dataset,
steps_per_epoch=int(0.1 * len(train_char_dataset)),
epochs=3,
validation_data=val_char_dataset,
validation_steps=int(0.1 * len(val_char_dataset)))
# Evaluate model_3 on whole validation char dataset
model_3.evaluate(val_char_dataset)
```
Nice! Looks like our character-level model is working, let's make some predictions with it and evaluate them.
```
# Make predictions with character model only
model_3_pred_probs = model_3.predict(val_char_dataset)
model_3_pred_probs
# Convert predictions to classes
model_3_preds = tf.argmax(model_3_pred_probs, axis=1)
model_3_preds
# Calculate Conv1D char only model results
model_3_results = calculate_results(y_true=val_labels_encoded,
y_pred=model_3_preds)
model_3_results
```
## Model 4: Combining pretrained token embeddings + character embeddings (hybrid embedding layer)
Alright, now things are going to get spicy.
In moving closer to build a model similar to the one in Figure 1 of [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf), it's time we tackled the hybrid token embedding layer they speak of.
This hybrid token embedding layer is a combination of token embeddings and character embeddings. In other words, they create a stacked embedding to represent sequences before passing them to the sequence label prediction layer.
So far we've built two models which have used token and character-level embeddings, however, these two models have used each of these embeddings exclusively.
To start replicating (or getting close to replicating) the model in Figure 1, we're going to go through the following steps:
1. Create a token-level model (similar to `model_1`)
2. Create a character-level model (similar to `model_3` with a slight modification to reflect the paper)
3. Combine (using [`layers.Concatenate`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Concatenate)) the outputs of 1 and 2
4. Build a series of output layers on top of 3 similar to Figure 1 and section 4.2 of [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf)
5. Construct a model which takes token and character-level sequences as input and produces sequence label probabilities as output
```
# 1. Setup token inputs/model
token_inputs = layers.Input(shape=[], dtype=tf.string, name="token_input")
token_embeddings = tf_hub_embedding_layer(token_inputs)
token_output = layers.Dense(128, activation="relu")(token_embeddings)
token_model = tf.keras.Model(inputs=token_inputs,
outputs=token_output)
# 2. Setup char inputs/model
char_inputs = layers.Input(shape=(1,), dtype=tf.string, name="char_input")
char_vectors = char_vectorizer(char_inputs)
char_embeddings = char_embed(char_vectors)
char_bi_lstm = layers.Bidirectional(layers.LSTM(25))(char_embeddings) # bi-LSTM shown in Figure 1 of https://arxiv.org/pdf/1612.05251.pdf
char_model = tf.keras.Model(inputs=char_inputs,
outputs=char_bi_lstm)
# 3. Concatenate token and char inputs (create hybrid token embedding)
token_char_concat = layers.Concatenate(name="token_char_hybrid")([token_model.output,
char_model.output])
# 4. Create output layers - addition of dropout discussed in 4.2 of https://arxiv.org/pdf/1612.05251.pdf
combined_dropout = layers.Dropout(0.5)(token_char_concat)
combined_dense = layers.Dense(200, activation="relu")(combined_dropout) # slightly different to Figure 1 due to different shapes of token/char embedding layers
final_dropout = layers.Dropout(0.5)(combined_dense)
output_layer = layers.Dense(num_classes, activation="softmax")(final_dropout)
# 5. Construct model with char and token inputs
model_4 = tf.keras.Model(inputs=[token_model.input, char_model.input],
outputs=output_layer,
name="model_4_token_and_char_embeddings")
```
Woah... There's a lot going on here, let's get a summary and plot our model to visualize what's happening.
```
# Get summary of token and character model
model_4.summary()
# Plot hybrid token and character model
from keras.utils import plot_model
plot_model(model_4)
```
Now that's a good looking model. Let's compile it just as we have the rest of our models.
> 🔑 **Note:** Section 4.2 of [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf) mentions using the SGD (stochastic gradient descent) optimizer, however, to stay consistent with our other models, we're going to use the Adam optimizer. As an exercise, you could try using [`tf.keras.optimizers.SGD`](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers/SGD) instead of [`tf.keras.optimizers.Adam`](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers/Adam) and compare the results.
```
# Compile token char model
model_4.compile(loss="categorical_crossentropy",
optimizer=tf.keras.optimizers.Adam(), # section 4.2 of https://arxiv.org/pdf/1612.05251.pdf mentions using SGD but we'll stick with Adam
metrics=["accuracy"])
```
And again, to keep our experiments fast, we'll fit our token-character-hybrid model on 10% of training and validate on 10% of validation batches. However, the difference with this model is that it requires two inputs, token-level sequences and character-level sequences.
We can do this by create a `tf.data.Dataset` with a tuple as it's first input, for example:
* `((token_data, char_data), (label))`
Let's see it in action.
### Combining token and character data into a `tf.data` dataset
```
# Combine chars and tokens into a dataset
train_char_token_data = tf.data.Dataset.from_tensor_slices((train_sentences, train_chars)) # make data
train_char_token_labels = tf.data.Dataset.from_tensor_slices(train_labels_one_hot) # make labels
train_char_token_dataset = tf.data.Dataset.zip((train_char_token_data, train_char_token_labels)) # combine data and labels
# Prefetch and batch train data
train_char_token_dataset = train_char_token_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
# Repeat same steps validation data
val_char_token_data = tf.data.Dataset.from_tensor_slices((val_sentences, val_chars))
val_char_token_labels = tf.data.Dataset.from_tensor_slices(val_labels_one_hot)
val_char_token_dataset = tf.data.Dataset.zip((val_char_token_data, val_char_token_labels))
val_char_token_dataset = val_char_token_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
# Check out training char and token embedding dataset
train_char_token_dataset, val_char_token_dataset
```
### Fitting a model on token and character-level sequences
```
# Fit the model on tokens and chars
model_4_history = model_4.fit(train_char_token_dataset, # train on dataset of token and characters
steps_per_epoch=int(0.1 * len(train_char_token_dataset)),
epochs=3,
validation_data=val_char_token_dataset,
validation_steps=int(0.1 * len(val_char_token_dataset)))
# Evaluate on the whole validation dataset
model_4.evaluate(val_char_token_dataset)
```
Nice! Our token-character hybrid model has come to life!
To make predictions with it, since it takes multiplie inputs, we can pass the `predict()` method a tuple of token-level sequences and character-level sequences.
We can then evaluate the predictions as we've done before.
```
# Make predictions using the token-character model hybrid
model_4_pred_probs = model_4.predict(val_char_token_dataset)
model_4_pred_probs
# Turn prediction probabilities into prediction classes
model_4_preds = tf.argmax(model_4_pred_probs, axis=1)
model_4_preds
# Get results of token-char-hybrid model
model_4_results = calculate_results(y_true=val_labels_encoded,
y_pred=model_4_preds)
model_4_results
```
## Model 5: Transfer Learning with pretrained token embeddings + character embeddings + positional embeddings
It seems like combining token embeddings and character embeddings gave our model a little performance boost.
But there's one more piece of the puzzle we can add in.
What if we engineered our own features into the model?
Meaning, what if we took our own knowledge about the data and encoded it in a numerical way to give our model more information about our samples?
The process of applying your own knowledge to build features as input to a model is called **feature engineering**.
Can you think of something important about the sequences we're trying to classify?
If you were to look at an abstract, would you expect the sentences to appear in order? Or does it make sense if they were to appear sequentially? For example, sequences labelled `CONCLUSIONS` at the beggining and sequences labelled `OBJECTIVE` at the end?
Abstracts typically come in a sequential order, such as:
* `OBJECTIVE` ...
* `METHODS` ...
* `METHODS` ...
* `METHODS` ...
* `RESULTS` ...
* `CONCLUSIONS` ...
Or
* `BACKGROUND` ...
* `OBJECTIVE` ...
* `METHODS` ...
* `METHODS` ...
* `RESULTS` ...
* `RESULTS` ...
* `CONCLUSIONS` ...
* `CONCLUSIONS` ...
Of course, we can't engineer the sequence labels themselves into the training data (we don't have these at test time), but we can encode the order of a set of sequences in an abstract.
For example,
* `Sentence 1 of 10` ...
* `Sentence 2 of 10` ...
* `Sentence 3 of 10` ...
* `Sentence 4 of 10` ...
* ...
You might've noticed this when we created our `preprocess_text_with_line_numbers()` function. When we read in a text file of abstracts, we counted the number of lines in an abstract as well as the number of each line itself.
Doing this led to the `"line_number"` and `"total_lines"` columns of our DataFrames.
```
# Inspect training dataframe
train_df.head()
```
The `"line_number"` and `"total_lines"` columns are features which didn't necessarily come with the training data but can be passed to our model as a **positional embedding**. In other words, the positional embedding is where the sentence appears in an abstract.
We can use these features because they will be available at test time.

*Since abstracts typically have a sequential order about them (for example, background, objective, methods, results, conclusion), it makes sense to add the line number of where a particular sentence occurs to our model. The beautiful thing is, these features will be available at test time (we can just count the number of sentences in an abstract and the number of each one).*
Meaning, if we were to predict the labels of sequences in an abstract our model had never seen, we could count the number of lines and the track the position of each individual line and pass it to our model.
> 🛠 **Exercise:** Another way of creating our positional embedding feature would be to combine the `"line_number"` and `"total_lines"` columns into one, for example a `"line_position"` column may contain values like `1_of_11`, `2_of_11`, etc. Where `1_of_11` would be the first line in an abstract 11 sentences long. After going through the following steps, you might want to revisit this positional embedding stage and see how a combined column of `"line_position"` goes against two separate columns.
### Create positional embeddings
Okay, enough talk about positional embeddings, let's create them.
Since our `"line_number"` and `"total_line"` columns are already numerical, we could pass them as they are to our model.
But to avoid our model thinking a line with `"line_number"=5` is five times greater than a line with `"line_number"=1`, we'll use one-hot-encoding to encode our `"line_number"` and `"total_lines"` features.
To do this, we can use the [`tf.one_hot`](https://www.tensorflow.org/api_docs/python/tf/one_hot) utility.
`tf.one_hot` returns a one-hot-encoded tensor. It accepts an array (or tensor) as input and the `depth` parameter determines the dimension of the returned tensor.
To figure out what we should set the `depth` parameter to, let's investigate the distribution of the `"line_number"` column.
> 🔑 **Note:** When it comes to one-hot-encoding our features, Scikit-Learn's [`OneHotEncoder`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OneHotEncoder.html) class is another viable option here.
```
# How many different line numbers are there?
train_df["line_number"].value_counts()
# Check the distribution of "line_number" column
train_df.line_number.plot.hist()
```
Looking at the distribution of the `"line_number"` column, it looks like the majority of lines have a position of 15 or less.
Knowing this, let's set the `depth` parameter of `tf.one_hot` to 15.
```
# Use TensorFlow to create one-hot-encoded tensors of our "line_number" column
train_line_numbers_one_hot = tf.one_hot(train_df["line_number"].to_numpy(), depth=15)
val_line_numbers_one_hot = tf.one_hot(val_df["line_number"].to_numpy(), depth=15)
test_line_numbers_one_hot = tf.one_hot(test_df["line_number"].to_numpy(), depth=15)
```
Setting the `depth` parameter of `tf.one_hot` to 15 means any sample with a `"line_number"` value of over 15 gets set to a tensor of all 0's, where as any sample with a `"line_number"` of under 15 gets turned into a tensor of all 0's but with a 1 at the index equal to the `"line_number"` value.
> 🔑 **Note:** We could create a one-hot tensor which has room for all of the potential values of `"line_number"` (`depth=30`), however, this would end up in a tensor of double the size of our current one (`depth=15`) where the vast majority of values are 0. Plus, only ~2,000/180,000 samples have a `"line_number"` value of over 15. So we would not be gaining much information about our data for doubling our feature space. This kind of problem is called the **curse of dimensionality**. However, since this we're working with deep models, it might be worth trying to throw as much information at the model as possible and seeing what happens. I'll leave exploring values of the `depth` parameter as an extension.
```
# Check one-hot encoded "line_number" feature samples
train_line_numbers_one_hot.shape, train_line_numbers_one_hot[:20]
```
We can do the same as we've done for our `"line_number"` column witht he `"total_lines"` column. First, let's find an appropriate value for the `depth` parameter of `tf.one_hot`.
```
# How many different numbers of lines are there?
train_df["total_lines"].value_counts()
# Check the distribution of total lines
train_df.total_lines.plot.hist();
```
Looking at the distribution of our `"total_lines"` column, a value of 20 looks like it covers the majority of samples.
We can confirm this with [`np.percentile()`](https://numpy.org/doc/stable/reference/generated/numpy.percentile.html).
```
# Check the coverage of a "total_lines" value of 20
np.percentile(train_df.total_lines, 98) # a value of 20 covers 98% of samples
```
Beautiful! Plenty of converage. Let's one-hot-encode our `"total_lines"` column just as we did our `"line_number"` column.
```
# Use TensorFlow to create one-hot-encoded tensors of our "total_lines" column
train_total_lines_one_hot = tf.one_hot(train_df["total_lines"].to_numpy(), depth=20)
val_total_lines_one_hot = tf.one_hot(val_df["total_lines"].to_numpy(), depth=20)
test_total_lines_one_hot = tf.one_hot(test_df["total_lines"].to_numpy(), depth=20)
# Check shape and samples of total lines one-hot tensor
train_total_lines_one_hot.shape, train_total_lines_one_hot[:10]
```
### Building a tribrid embedding model
Woohoo! Positional embedding tensors ready.
It's time to build the biggest model we've built yet. One which incorporates token embeddings, character embeddings and our newly crafted positional embeddings.
We'll be venturing into uncovered territory but there will be nothing here you haven't practiced before.
More specifically we're going to go through the following steps:
1. Create a token-level model (similar to `model_1`)
2. Create a character-level model (similar to `model_3` with a slight modification to reflect the paper)
3. Create a `"line_number"` model (takes in one-hot-encoded `"line_number"` tensor and passes it through a non-linear layer)
4. Create a `"total_lines"` model (takes in one-hot-encoded `"total_lines"` tensor and passes it through a non-linear layer)
5. Combine (using [`layers.Concatenate`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Concatenate)) the outputs of 1 and 2 into a token-character-hybrid embedding and pass it series of output to Figure 1 and section 4.2 of [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf)
6. Combine (using [`layers.Concatenate`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Concatenate)) the outputs of 3, 4 and 5 into a token-character-positional tribrid embedding
7. Create an output layer to accept the tribrid embedding and output predicted label probabilities
8. Combine the inputs of 1, 2, 3, 4 and outputs of 7 into a [`tf.keras.Model`](https://www.tensorflow.org/api_docs/python/tf/keras/Model)
Woah! That's alot... but nothing we're not capable of. Let's code it.
```
# 1. Token inputs
token_inputs = layers.Input(shape=[], dtype="string", name="token_inputs")
token_embeddings = tf_hub_embedding_layer(token_inputs)
token_outputs = layers.Dense(128, activation="relu")(token_embeddings)
token_model = tf.keras.Model(inputs=token_inputs,
outputs=token_embeddings)
# 2. Char inputs
char_inputs = layers.Input(shape=(1,), dtype="string", name="char_inputs")
char_vectors = char_vectorizer(char_inputs)
char_embeddings = char_embed(char_vectors)
char_bi_lstm = layers.Bidirectional(layers.LSTM(32))(char_embeddings)
char_model = tf.keras.Model(inputs=char_inputs,
outputs=char_bi_lstm)
# 3. Line numbers inputs
line_number_inputs = layers.Input(shape=(15,), dtype=tf.int32, name="line_number_input")
x = layers.Dense(32, activation="relu")(line_number_inputs)
line_number_model = tf.keras.Model(inputs=line_number_inputs,
outputs=x)
# 4. Total lines inputs
total_lines_inputs = layers.Input(shape=(20,), dtype=tf.int32, name="total_lines_input")
y = layers.Dense(32, activation="relu")(total_lines_inputs)
total_line_model = tf.keras.Model(inputs=total_lines_inputs,
outputs=y)
# 5. Combine token and char embeddings into a hybrid embedding
combined_embeddings = layers.Concatenate(name="token_char_hybrid_embedding")([token_model.output,
char_model.output])
z = layers.Dense(256, activation="relu")(combined_embeddings)
z = layers.Dropout(0.5)(z)
# 6. Combine positional embeddings with combined token and char embeddings into a tribrid embedding
z = layers.Concatenate(name="token_char_positional_embedding")([line_number_model.output,
total_line_model.output,
z])
# 7. Create output layer
output_layer = layers.Dense(5, activation="softmax", name="output_layer")(z)
# 8. Put together model
model_5 = tf.keras.Model(inputs=[line_number_model.input,
total_line_model.input,
token_model.input,
char_model.input],
outputs=output_layer)
```
There's a lot going on here... let's visualize what's happening with a summary by plotting our model.
```
# Get a summary of our token, char and positional embedding model
model_5.summary()
# Plot the token, char, positional embedding model
from tensorflow.keras.utils import plot_model
plot_model(model_5)
```
Visualizing the model makes it much easier to understand.
Essentially what we're doing is trying to encode as much information about our sequences as possible into various embeddings (the inputs to our model) so our model has the best chance to figure out what label belongs to a sequence (the outputs of our model).
You'll notice our model is looking very similar to the model shown in Figure 1 of [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf). However, a few differences still remain:
* We're using pretrained TensorFlow Hub token embeddings instead of GloVe emebddings.
* We're using a Dense layer on top of our token-character hybrid embeddings instead of a bi-LSTM layer.
* Section 3.1.3 of the paper mentions a label sequence optimization layer (which helps to make sure sequence labels come out in a respectable order) but it isn't shown in Figure 1. To makeup for the lack of this layer in our model, we've created the positional embeddings layers.
* Section 4.2 of the paper mentions the token and character embeddings are updated during training, our pretrained TensorFlow Hub embeddings remain frozen.
* The paper uses the [`SGD`](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers/SGD) optimizer, we're going to stick with [`Adam`](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers/Adam).
All of the differences above are potential extensions of this project.
```
# Check which layers of our model are trainable or not
for layer in model_5.layers:
print(layer, layer.trainable)
```
Now our model is constructed, let's compile it.
This time, we're going to introduce a new parameter to our loss function called `label_smoothing`. Label smoothing helps to regularize our model (prevent overfitting) by making sure it doesn't get too focused on applying one particular label to a sample.
For example, instead of having an output prediction of:
* `[0.0, 0.0, 1.0, 0.0, 0.0]` for a sample (the model is very confident the right label is index 2).
It's predictions will get smoothed to be something like:
* `[0.01, 0.01, 0.096, 0.01, 0.01]` giving a small activation to each of the other labels, in turn, hopefully improving generalization.
> 📖 **Resource:** For more on label smoothing, see the great blog post by PyImageSearch, [*Label smoothing with Keras, TensorFlow, and Deep Learning*](https://www.pyimagesearch.com/2019/12/30/label-smoothing-with-keras-tensorflow-and-deep-learning/).
```
# Compile token, char, positional embedding model
model_5.compile(loss=tf.keras.losses.CategoricalCrossentropy(label_smoothing=0.2), # add label smoothing (examples which are really confident get smoothed a little)
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
```
### Create tribrid embedding datasets and fit tribrid model
Model compiled!
Again, to keep our experiments swift, let's fit on 20,000 examples for 3 epochs.
This time our model requires four feature inputs:
1. Train line numbers one-hot tensor (`train_line_numbers_one_hot`)
2. Train total lines one-hot tensor (`train_total_lines_one_hot`)
3. Token-level sequences tensor (`train_sentences`)
4. Char-level sequences tensor (`train_chars`)
We can pass these as tuples to our `tf.data.Dataset.from_tensor_slices()` method to create appropriately shaped and batched `PrefetchedDataset`'s.
```
# Create training and validation datasets (all four kinds of inputs)
train_pos_char_token_data = tf.data.Dataset.from_tensor_slices((train_line_numbers_one_hot, # line numbers
train_total_lines_one_hot, # total lines
train_sentences, # train tokens
train_chars)) # train chars
train_pos_char_token_labels = tf.data.Dataset.from_tensor_slices(train_labels_one_hot) # train labels
train_pos_char_token_dataset = tf.data.Dataset.zip((train_pos_char_token_data, train_pos_char_token_labels)) # combine data and labels
train_pos_char_token_dataset = train_pos_char_token_dataset.batch(32).prefetch(tf.data.AUTOTUNE) # turn into batches and prefetch appropriately
# Validation dataset
val_pos_char_token_data = tf.data.Dataset.from_tensor_slices((val_line_numbers_one_hot,
val_total_lines_one_hot,
val_sentences,
val_chars))
val_pos_char_token_labels = tf.data.Dataset.from_tensor_slices(val_labels_one_hot)
val_pos_char_token_dataset = tf.data.Dataset.zip((val_pos_char_token_data, val_pos_char_token_labels))
val_pos_char_token_dataset = val_pos_char_token_dataset.batch(32).prefetch(tf.data.AUTOTUNE) # turn into batches and prefetch appropriately
# Check input shapes
train_pos_char_token_dataset, val_pos_char_token_dataset
# Fit the token, char and positional embedding model
history_model_5 = model_5.fit(train_pos_char_token_dataset,
steps_per_epoch=int(0.1 * len(train_pos_char_token_dataset)),
epochs=3,
validation_data=val_pos_char_token_dataset,
validation_steps=int(0.1 * len(val_pos_char_token_dataset)))
```
Tribrid model trained! Time to make some predictions with it and evaluate them just as we've done before.
```
# Make predictions with token-char-positional hybrid model
model_5_pred_probs = model_5.predict(val_pos_char_token_dataset, verbose=1)
model_5_pred_probs
# Turn prediction probabilities into prediction classes
model_5_preds = tf.argmax(model_5_pred_probs, axis=1)
model_5_preds
# Calculate results of token-char-positional hybrid model
model_5_results = calculate_results(y_true=val_labels_encoded,
y_pred=model_5_preds)
model_5_results
```
## Compare model results
Far out, we've come a long way. From a baseline model to training a model containing three different kinds of embeddings.
Now it's time to compare each model's performance against each other.
We'll also be able to compare our model's to the [*PubMed 200k RCT:
a Dataset for Sequential Sentence Classification in Medical Abstracts*](https://arxiv.org/pdf/1710.06071.pdf) paper.
Since all of our model results are in dictionaries, let's combine them into a pandas DataFrame to visualize them.
```
# Combine model results into a DataFrame
all_model_results = pd.DataFrame({"baseline": baseline_results,
"custom_token_embed_conv1d": model_1_results,
"pretrained_token_embed": model_2_results,
"custom_char_embed_conv1d": model_3_results,
"hybrid_char_token_embed": model_4_results,
"tribrid_pos_char_token_embed": model_5_results})
all_model_results = all_model_results.transpose()
all_model_results
# Reduce the accuracy to same scale as other metrics
all_model_results["accuracy"] = all_model_results["accuracy"]/100
# Plot and compare all of the model results
all_model_results.plot(kind="bar", figsize=(10, 7)).legend(bbox_to_anchor=(1.0, 1.0));
```
Since the [*PubMed 200k RCT:
a Dataset for Sequential Sentence Classification in Medical Abstracts*](https://arxiv.org/pdf/1710.06071.pdf) paper compares their tested model's F1-scores on the test dataset, let's take at our model's F1-scores.
> 🔑 **Note:** We could've also made these comparisons in TensorBoard using the [`TensorBoard`](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/TensorBoard) callback during training.
```
# Sort model results by f1-score
all_model_results.sort_values("f1", ascending=False)["f1"].plot(kind="bar", figsize=(10, 7));
```
Nice! Based on F1-scores, it looks like our tribrid embedding model performs the best by a fair margin.
Though, in comparison to the results reported in Table 3 of the [*PubMed 200k RCT:
a Dataset for Sequential Sentence Classification in Medical Abstracts*](https://arxiv.org/pdf/1710.06071.pdf) paper, our model's F1-score is still underperforming (the authors model achieves an F1-score of 90.0 on the 20k RCT dataset versus our F1-score of ~82.6).
There are some things to note about this difference:
* Our models (with an exception for the baseline) have been trained on ~18,000 (10% of batches) samples of sequences and labels rather than the full ~180,000 in the 20k RCT dataset.
* This is often the case in machine learning experiments though, make sure training works on a smaller number of samples, then upscale when needed (an extension to this project will be training a model on the full dataset).
* Our model's prediction performance levels have been evaluated on the validation dataset not the test dataset (we'll evaluate our best model on the test dataset shortly).
## Save and load best performing model
Since we've been through a fair few experiments, it's a good idea to save our best performing model so we can reuse it without having to retrain it.
We can save our best performing model by calling the [`save()`](https://www.tensorflow.org/guide/keras/save_and_serialize#the_short_answer_to_saving_loading) method on it.
```
# Save best performing model to SavedModel format (default)
model_5.save("skimlit_tribrid_model") # model will be saved to path specified by string
```
Optional: If you're using Google Colab, you might want to copy your saved model to Google Drive (or [download it](https://colab.research.google.com/notebooks/io.ipynb#scrollTo=hauvGV4hV-Mh)) for more permanent storage (Google Colab files disappear after you disconnect).
```
# Example of copying saved model from Google Colab to Drive (requires Google Drive to be mounted)
# !cp skim_lit_best_model -r /content/drive/MyDrive/tensorflow_course/skim_lit
```
Like all good cooking shows, we've got a pretrained model (exactly the same kind of model we built for `model_5` [saved and stored on Google Storage](https://storage.googleapis.com/ztm_tf_course/skimlit/skimlit_best_model.zip)).
So to make sure we're all using the same model for evaluation, we'll download it and load it in.
And when loading in our model, since it uses a couple of [custom objects](https://www.tensorflow.org/guide/keras/save_and_serialize#custom_objects) (our TensorFlow Hub layer and `TextVectorization` layer), we'll have to load it in by specifying them in the `custom_objects` parameter of [`tf.keras.models.load_model()`](https://www.tensorflow.org/api_docs/python/tf/keras/models/load_model).
```
# Download pretrained model from Google Storage
!wget https://storage.googleapis.com/ztm_tf_course/skimlit/skimlit_tribrid_model.zip
!mkdir skimlit_gs_model
!unzip skimlit_tribrid_model.zip -d skimlit_gs_model
# Import TensorFlow model dependencies (if needed) - https://github.com/tensorflow/tensorflow/issues/38250
import tensorflow_hub as hub
import tensorflow as tf
from tensorflow.keras.layers.experimental.preprocessing import TextVectorization
model_path = "skimlit_gs_model/skimlit_tribrid_model"
# Load downloaded model from Google Storage
loaded_model = tf.keras.models.load_model(model_path,
custom_objects={"TextVectorization": TextVectorization, # required for char vectorization
"KerasLayer": hub.KerasLayer}) # required for token embedding
```
### Make predictions and evalaute them against the truth labels
To make sure our model saved and loaded correctly, let's make predictions with it, evaluate them and then compare them to the prediction results we calculated earlier.
```
# Make predictions with the loaded model on the validation set
loaded_pred_probs = loaded_model.predict(val_pos_char_token_dataset, verbose=1)
loaded_preds = tf.argmax(loaded_pred_probs, axis=1)
loaded_preds[:10]
# Evaluate loaded model's predictions
loaded_model_results = calculate_results(val_labels_encoded,
loaded_preds)
loaded_model_results
```
Now let's compare our loaded model's predictions with the prediction results we obtained before saving our model.
```
# Compare loaded model results with original trained model results (should return no errors)
assert model_5_results == loaded_model_results
```
It's worth noting that loading in a SavedModel unfreezes all layers (makes them all trainable). So if you want to freeze any layers, you'll have to set their trainable attribute to `False`.
```
# Check loaded model summary (note the number of trainable parameters)
loaded_model.summary()
```
## Evaluate model on test dataset
To make our model's performance more comparable with the results reported in Table 3 of the [*PubMed 200k RCT:
a Dataset for Sequential Sentence Classification in Medical Abstracts*](https://arxiv.org/pdf/1710.06071.pdf) paper, let's make predictions on the test dataset and evaluate them.
```
# Create test dataset batch and prefetched
test_pos_char_token_data = tf.data.Dataset.from_tensor_slices((test_line_numbers_one_hot,
test_total_lines_one_hot,
test_sentences,
test_chars))
test_pos_char_token_labels = tf.data.Dataset.from_tensor_slices(test_labels_one_hot)
test_pos_char_token_dataset = tf.data.Dataset.zip((test_pos_char_token_data, test_pos_char_token_labels))
test_pos_char_token_dataset = test_pos_char_token_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
# Check shapes
test_pos_char_token_dataset
# Make predictions on the test dataset
test_pred_probs = loaded_model.predict(test_pos_char_token_dataset,
verbose=1)
test_preds = tf.argmax(test_pred_probs, axis=1)
test_preds[:10]
# Evaluate loaded model test predictions
loaded_model_test_results = calculate_results(y_true=test_labels_encoded,
y_pred=test_preds)
loaded_model_test_results
```
It seems our best model (so far) still has some ways to go to match the performance of the results in the paper (their model gets 90.0 F1-score on the test dataset, where as ours gets ~82.1 F1-score).
However, as we discussed before our model has only been trained on 20,000 out of the total ~180,000 sequences in the RCT 20k dataset. We also haven't fine-tuned our pretrained embeddings (the paper fine-tunes GloVe embeddings). So there's a couple of extensions we could try to improve our results.
## Find most wrong
One of the best ways to investigate where your model is going wrong (or potentially where your data is wrong) is to visualize the "most wrong" predictions.
The most wrong predictions are samples where the model has made a prediction with a high probability but has gotten it wrong (the model's prediction disagreess with the ground truth label).
Looking at the most wrong predictions can give us valuable information on how to improve further models or fix the labels in our data.
Let's write some code to help us visualize the most wrong predictions from the test dataset.
First we'll convert all of our integer-based test predictions into their string-based class names.
```
%%time
# Get list of class names of test predictions
test_pred_classes = [label_encoder.classes_[pred] for pred in test_preds]
test_pred_classes
```
Now we'll enrich our test DataFame with a few values:
* A `"prediction"` (string) column containing our model's prediction for a given sample.
* A `"pred_prob"` (float) column containing the model's maximum prediction probabiliy for a given sample.
* A `"correct"` (bool) column to indicate whether or not the model's prediction matches the sample's target label.
```
# Create prediction-enriched test dataframe
test_df["prediction"] = test_pred_classes # create column with test prediction class names
test_df["pred_prob"] = tf.reduce_max(test_pred_probs, axis=1).numpy() # get the maximum prediction probability
test_df["correct"] = test_df["prediction"] == test_df["target"] # create binary column for whether the prediction is right or not
test_df.head(20)
```
Looking good! Having our data like this, makes it very easy to manipulate and view in different ways.
How about we sort our DataFrame to find the samples with the highest `"pred_prob"` and where the prediction was wrong (`"correct" == False`)?
```
# Find top 100 most wrong samples (note: 100 is an abitrary number, you could go through all of them if you wanted)
top_100_wrong = test_df[test_df["correct"] == False].sort_values("pred_prob", ascending=False)[:100]
top_100_wrong
```
Great (or not so great)! Now we've got a subset of our model's most wrong predictions, let's write some code to visualize them.
```
# Investigate top wrong preds
for row in top_100_wrong[0:10].itertuples(): # adjust indexes to view different samples
_, target, text, line_number, total_lines, prediction, pred_prob, _ = row
print(f"Target: {target}, Pred: {prediction}, Prob: {pred_prob}, Line number: {line_number}, Total lines: {total_lines}\n")
print(f"Text:\n{text}\n")
print("-----\n")
```
What do you notice about the most wrong predictions? Does the model make silly mistakes? Or are some of the labels incorrect/ambiguous (e.g. a line in an abstract could potentially be labelled `OBJECTIVE` or `BACKGROUND` and make sense).
A next step here would be if there are a fair few samples with inconsistent labels, you could go through your training dataset, update the labels and then retrain a model. The process of using a model to help improve/investigate your dataset's labels is often referred to as **active learning**.
## Make example predictions
Okay, we've made some predictions on the test dataset, now's time to really test our model out.
To do so, we're going to get some data from the wild and see how our model performs.
In other words, were going to find an RCT abstract from PubMed, preprocess the text so it works with our model, then pass each sequence in the wild abstract through our model to see what label it predicts.
For an appropriate sample, we'll need to search PubMed for RCT's (randomized controlled trials) without abstracts which have been split up (on exploring PubMed you'll notice many of the abstracts are already preformatted into separate sections, this helps dramatically with readability).
Going through various PubMed studies, I managed to find the following unstructured abstract from [*RCT of a manualized social treatment for high-functioning autism spectrum disorders*](https://pubmed.ncbi.nlm.nih.gov/20232240/):
> This RCT examined the efficacy of a manualized social intervention for children with HFASDs. Participants were randomly assigned to treatment or wait-list conditions. Treatment included instruction and therapeutic activities targeting social skills, face-emotion recognition, interest expansion, and interpretation of non-literal language. A response-cost program was applied to reduce problem behaviors and foster skills acquisition. Significant treatment effects were found for five of seven primary outcome measures (parent ratings and direct child measures). Secondary measures based on staff ratings (treatment group only) corroborated gains reported by parents. High levels of parent, child and staff satisfaction were reported, along with high levels of treatment fidelity. Standardized effect size estimates were primarily in the medium and large ranges and favored the treatment group.
Looking at the large chunk of text can seem quite intimidating. Now imagine you're a medical researcher trying to skim through the literature to find a study relevant to your work.
Sounds like quite the challenge right?
Enter SkimLit 🤓🔥!
Let's see what our best model so far (`model_5`) makes of the above abstract.
But wait...
As you might've guessed the above abstract hasn't been formatted in the same structure as the data our model has been trained on. Therefore, before we can make a prediction on it, we need to preprocess it just as we have our other sequences.
More specifically, for each abstract, we'll need to:
1. Split it into sentences (lines).
2. Split it into characters.
3. Find the number of each line.
4. Find the total number of lines.
Starting with number 1, there are a couple of ways to split our abstracts into actual sentences. A simple one would be to use Python's in-built `split()` string method, splitting the abstract wherever a fullstop appears. However, can you imagine where this might go wrong?
Another more advanced option would be to leverage [spaCy's](https://spacy.io/) (a very powerful NLP library) [`sentencizer`](https://spacy.io/usage/linguistic-features#sbd) class. Which is an easy to use sentence splitter based on spaCy's English language model.
I've prepared some abstracts from PubMed RCT papers to try our model on, we can download them [from GitHub](https://raw.githubusercontent.com/mrdbourke/tensorflow-deep-learning/main/extras/skimlit_example_abstracts.json).
```
# Download and open example abstracts (copy and pasted from PubMed)
!wget https://raw.githubusercontent.com/mrdbourke/tensorflow-deep-learning/main/extras/skimlit_example_abstracts.json
with open("skimlit_example_abstracts.json", "r") as f:
example_abstracts = json.load(f)
example_abstracts
# See what our example abstracts look like
abstracts = pd.DataFrame(example_abstracts)
abstracts
```
Now we've downloaded some example abstracts, let's see how one of them goes with our trained model.
First, we'll need to parse it using spaCy to turn it from a big chunk of text into sentences.
```
# Create sentencizer - Source: https://spacy.io/usage/linguistic-features#sbd
from spacy.lang.en import English
nlp = English() # setup English sentence parser
sentencizer = nlp.create_pipe("sentencizer") # create sentence splitting pipeline object
nlp.add_pipe(sentencizer) # add sentence splitting pipeline object to sentence parser
doc = nlp(example_abstracts[0]["abstract"]) # create "doc" of parsed sequences, change index for a different abstract
abstract_lines = [str(sent) for sent in list(doc.sents)] # return detected sentences from doc in string type (not spaCy token type)
abstract_lines
```
Beautiful! It looks like spaCy has split the sentences in the abstract correctly. However, it should be noted, there may be more complex abstracts which don't get split perfectly into separate sentences (such as the example in [*Baclofen promotes alcohol abstinence in alcohol dependent cirrhotic patients with hepatitis C virus (HCV) infection*](https://pubmed.ncbi.nlm.nih.gov/22244707/)), in this case, more custom splitting techniques would have to be investigated.
Now our abstract has been split into sentences, how about we write some code to count line numbers as well as total lines.
To do so, we can leverage some of the functionality of our `preprocess_text_with_line_numbers()` function.
```
# Get total number of lines
total_lines_in_sample = len(abstract_lines)
# Go through each line in abstract and create a list of dictionaries containing features for each line
sample_lines = []
for i, line in enumerate(abstract_lines):
sample_dict = {}
sample_dict["text"] = str(line)
sample_dict["line_number"] = i
sample_dict["total_lines"] = total_lines_in_sample - 1
sample_lines.append(sample_dict)
sample_lines
```
Now we've got `"line_number"` and `"total_lines"` values, we can one-hot encode them with `tf.one_hot` just like we did with our training dataset (using the same values for the `depth` parameter).
```
# Get all line_number values from sample abstract
test_abstract_line_numbers = [line["line_number"] for line in sample_lines]
# One-hot encode to same depth as training data, so model accepts right input shape
test_abstract_line_numbers_one_hot = tf.one_hot(test_abstract_line_numbers, depth=15)
test_abstract_line_numbers_one_hot
# Get all total_lines values from sample abstract
test_abstract_total_lines = [line["total_lines"] for line in sample_lines]
# One-hot encode to same depth as training data, so model accepts right input shape
test_abstract_total_lines_one_hot = tf.one_hot(test_abstract_total_lines, depth=20)
test_abstract_total_lines_one_hot
```
We can also use our `split_chars()` function to split our abstract lines into characters.
```
# Split abstract lines into characters
abstract_chars = [split_chars(sentence) for sentence in abstract_lines]
abstract_chars
```
Alright, now we've preprocessed our wild RCT abstract into all of the same features our model was trained on, we can pass these features to our model and make sequence label predictions!
```
# Make predictions on sample abstract features
%%time
test_abstract_pred_probs = loaded_model.predict(x=(test_abstract_line_numbers_one_hot,
test_abstract_total_lines_one_hot,
tf.constant(abstract_lines),
tf.constant(abstract_chars)))
test_abstract_pred_probs
# Turn prediction probabilities into prediction classes
test_abstract_preds = tf.argmax(test_abstract_pred_probs, axis=1)
test_abstract_preds
```
Now we've got the predicted sequence label for each line in our sample abstract, let's write some code to visualize each sentence with its predicted label.
```
# Turn prediction class integers into string class names
test_abstract_pred_classes = [label_encoder.classes_[i] for i in test_abstract_preds]
test_abstract_pred_classes
# Visualize abstract lines and predicted sequence labels
for i, line in enumerate(abstract_lines):
print(f"{test_abstract_pred_classes[i]}: {line}")
```
Nice! Isn't that much easier to read? I mean, it looks like our model's predictions could be improved, but how cool is that?
Imagine implementing our model to the backend of the PubMed website to format any unstructured RCT abstract on the site.
Or there could even be a browser extension, called "SkimLit" which would add structure (powered by our model) to any unstructured RCT abtract.
And if showed your medical researcher friend, and they thought the predictions weren't up to standard, there could be a button saying "is this label correct?... if not, what should it be?". That way the dataset, along with our model's future predictions, could be improved over time.
Of course, there are many more ways we could go to improve the model, the usuability, the preprocessing functionality (e.g. functionizing our sample abstract preprocessing pipeline) but I'll leave these for the exercises/extensions.
> 🤔 **Question:** How can we be sure the results of our test example from the wild are truly *wild*? Is there something we should check about the sample we're testing on?
## 🛠 Exercises
1. Train `model_5` on all of the data in the training dataset for as many epochs until it stops improving. Since this might take a while, you might want to use:
* [`tf.keras.callbacks.ModelCheckpoint`](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/ModelCheckpoint) to save the model's best weights only.
* [`tf.keras.callbacks.EarlyStopping`](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/EarlyStopping) to stop the model from training once the validation loss has stopped improving for ~3 epochs.
2. Checkout the [Keras guide on using pretrained GloVe embeddings](https://keras.io/examples/nlp/pretrained_word_embeddings/). Can you get this working with one of our models?
* Hint: You'll want to incorporate it with a custom token [Embedding](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Embedding) layer.
* It's up to you whether or not you fine-tune the GloVe embeddings or leave them frozen.
3. Try replacing the TensorFlow Hub Universal Sentence Encoder pretrained embedding for the [TensorFlow Hub BERT PubMed expert](https://tfhub.dev/google/experts/bert/pubmed/2) (a language model pretrained on PubMed texts) pretrained embedding. Does this effect results?
* Note: Using the BERT PubMed expert pretrained embedding requires an extra preprocessing step for sequences (as detailed in the [TensorFlow Hub guide](https://tfhub.dev/google/experts/bert/pubmed/2)).
* Does the BERT model beat the results mentioned in this paper? https://arxiv.org/pdf/1710.06071.pdf
4. What happens if you were to merge our `line_number` and `total_lines` features for each sequence? For example, created a `X_of_Y` feature instead? Does this effect model performance?
* Another example: `line_number=1` and `total_lines=11` turns into `line_of_X=1_of_11`.
5. Write a function (or series of functions) to take a sample abstract string, preprocess it (in the same way our model has been trained), make a prediction on each sequence in the abstract and return the abstract in the format:
* `PREDICTED_LABEL`: `SEQUENCE`
* `PREDICTED_LABEL`: `SEQUENCE`
* `PREDICTED_LABEL`: `SEQUENCE`
* `PREDICTED_LABEL`: `SEQUENCE`
* ...
* You can find your own unstrcutured RCT abstract from PubMed or try this one from: [*Baclofen promotes alcohol abstinence in alcohol dependent cirrhotic patients with hepatitis C virus (HCV) infection*](https://pubmed.ncbi.nlm.nih.gov/22244707/).
## 📖 Extra-curriculum
* For more on working with text/spaCy, see [spaCy's advanced NLP course](https://course.spacy.io/en/). If you're going to be working on production-level NLP problems, you'll probably end up using spaCy.
* For another look at how to approach a text classification problem like the one we've just gone through, I'd suggest going through [Google's Machine Learning Course for text classification](https://developers.google.com/machine-learning/guides/text-classification).
* Since our dataset has imbalanced classes (as with many real-world datasets), so it might be worth looking into the [TensorFlow guide for different methods to training a model with imbalanced classes](https://www.tensorflow.org/tutorials/structured_data/imbalanced_data).
| github_jupyter |
# Shallow Copy Versus Deep Copy Operations
Here's the issue we are looking at now: when we make a copy of an object that contains other objects, what happens if the object we are copying "contains" other objects. So, if `list_orig` has `inner_list` as one of its members, like...
`list_orig = [1, 2, [3, 5], 4]`
and we make a copy of `list_orig` into `list_copy`...
`list_orig` ---> `list_copy`
does `list_copy` have a **copy** of `inner_list`, or do `list_orig` and `list_copy` share the **same** `inner_list`?
As we will see, the default Python behavior is that `list_orig` and `list_copy` will **share** `inner_list`. That is called a *shallow copy*.
However, Python also permits the programmer to "order up" a *deep copy* so that `inner_list` is copied also.
<img src="https://i.stack.imgur.com/AWKJa.jpg" width="30%">
## Deep copy
Let's first look at a deep copy.
```
# initializing list_a
INNER_LIST_IDX = 2
list_orig = [1, 2, [3, 5], 4]
print ("The original elements before deep copying")
print(list_orig)
print(list_orig[INNER_LIST_IDX][0])
```
We will use deepcopy to deep copy `list_orig` and change an element in the new list.
```
import copy
```
(What's in the `copy` module?)
```
dir(copy)
```
The change is made in list_b:
```
list_copy = copy.deepcopy(list_orig)
# Now change first element of the inner list:
list_copy[INNER_LIST_IDX][0] = 7
print("The new list (list_copy) of elements after deep copying and list modification")
print(list_copy)
```
That change is **not** reflected in original list as we made a deep copy:
```
print ("The original list (list_orig) elements after deep copying")
print(list_orig)
print("The list IDs are:", id(list_orig), id(list_copy))
print("The inner list IDs are:", id(list_orig[INNER_LIST_IDX]),
id(list_copy[INNER_LIST_IDX]))
```
## Shallow copy
Like a "shallow" person, and shallow copy only sees the "surface" of the object it is copying... it doesn't peer further inside.
We'll set up `list_orig` as before:
```
INNER_LIST_IDX = 2
# initializing list_1
list_orig = [1, 2, [3, 5], 4]
# original elements of list
print ("The original elements before shallow copying")
print(list_orig)
```
Using copy to shallow copy adding an element to new list
```
import copy
list_copy = copy.copy(list_orig) # not deepcopy()!
list_copy[INNER_LIST_IDX][0] = 7
```
Let's check the result:
```
print ("The original elements after shallow copying")
print(list_orig)
```
Let's change `inner_list` in `list_orig`:
```
list_orig[INNER_LIST_IDX][0] = "That's different!"
```
And let's see what `list_copy`'s inner list now looks like:
```
print(list_copy)
```
So we can see that `list_orig` and `list_copy` share the same inner list, which is now `["That's different!", 5]`. And their IDs show this:
```
print("The list IDs are:", id(list_orig), id(list_copy))
print("The inner list IDs are:", id(list_orig[INNER_LIST_IDX]),
id(list_copy[INNER_LIST_IDX]))
```
**But**... if we change the outer list element at INNER_LIST_IDX... **that** change is not shared!
```
list_orig[INNER_LIST_IDX] = ["Brand new list!", 16]
print("list_orig:", list_orig)
print("list_copy:", list_copy)
```
### Slicing
We should see which of the above slicing gets us!
```
list_slice = list_orig[:]
print(list_slice)
```
What happens to `list_slice` if we change `list_a`:
```
list_orig[INNER_LIST_IDX][0] = "Did our slice change?"
list_orig[0] = "New value at 0!"
print("Original list:", list_orig)
print("Our slice:", list_slice)
```
So, slicing make a *shallow* copy.
### Assignment
And if we don't even slice, but just assign, even the outer lists will be the same, since we haven't made **any** sort of copy at all... we've just put two labels on the same "box":
```
list_alias = list_orig
another_alias = list_alias
yet_another = list_orig
print(list_alias, end="\n\n")
# change elem 0:
list_orig[0] = "Even the outer elems are the same."
print("List alias has element 0 altered:", list_alias, end="\n\n")
print("List slice does not have element 0 altered:", list_slice, end="\n\n")
# see their IDs:
print("list_orig ID:", id(list_orig), end="\n\n")
print("list_alias ID:", id(list_alias), end="\n\n")
print("another_alias ID:", id(another_alias), end="\n\n")
print("list_slice ID:", id(list_slice), end="\n\n")
```
What does `append()` do in terms of shallow versus deep copy?
```
INNER_LIST_IDX = 3
list_orig = [1, 2, 3, [4, 5]]
list_copy = []
for elem in list_orig:
list_copy.append(elem)
list_orig[INNER_LIST_IDX][0] = "Did the copy's inner list change?"
print("list_copy:", list_copy)
```
## What If We Need Different Copy Behavior for Our Own Class?
**Advanced topic**: Python has *dunder* (double-underscore) methods `__copy__()` and `__deepcopy__()` that we can implement in our own class when we have special copying needs.
## What about Dictionaries?
The above discussion was in terms of lists, but the same considerations apply to dictionaries.
<hr>
*Assignment* just creates an *alias* for a dictionary. *All* changes to the original will be reflected in the alias:
```
original = {"a": 1, "b": 2, "c": {"d": 3, "e": 4}}
dict_alias = original
print("dict_alias:", dict_alias)
original["a"] = "A brand new value!"
print("dict_alias:", dict_alias)
```
<hr>
A *shallow* copy copies the "skin" of the dictionary, but not the "innards":
```
from copy import copy, deepcopy
original = {"a": 1, "b": 2, "c": {"d": 3, "e": 4}}
dict_scopy = copy(original)
print("shallow copy:", dict_scopy)
# change the outer part:
original["a"] = "This won't be in shallow copy!"
print("original:", original)
print("shallow copy:", dict_scopy)
# change the innards:
original["c"]["d"] = "This WILL appear in the shallow copy!"
print("shallow copy:", dict_scopy)
dict_scopy["c"]["e"] = "This WILL appear in the original!"
print("original:", original)
original["c"] = "Hello Monte!"
print("original:", original)
print("shallow copy:", dict_scopy)
```
<hr>
A *deep* copy copies the "innards" of the dictionary as well as the "skin":
```
original = {"a": 1, "b": 2, "c": {"d": 3, "e": 4}}
dict_dcopy = deepcopy(original)
print("deep copy:", dict_dcopy)
original["c"]["d"] = "This WON'T appear in the deep copy!"
print("original:", original)
print("deep copy:", dict_dcopy)
```
| github_jupyter |
## This code aims to generate an animation from the data from the ann
```
import numpy as np
from sklearn.neural_network import MLPRegressor
from sklearn import preprocessing
from sklearn.cross_validation import train_test_split
import pickle #saving to file
import matplotlib.pyplot as plt
import matplotlib.animation as animation
from sklearn.grid_search import GridSearchCV # looking for parameters
import datetime
```
## Preprocessing data
```
#this function reads the file
def read_data(archive, rows, columns):
data = open(archive, 'r')
mylist = data.read().split()
data.close()
myarray = np.array(mylist).reshape(( rows, columns)).astype(float)
return myarray
# getting data and preprocessing
data = read_data('../get_data_example/set.txt',72, 12)
#training_set = [0av_left, 1av_right, 2angles_left,
# 3angles_right, 4aa_left, 5aa_right, 678vxyz_left, 91011vxyz_right];
X = data[:, [4, 5, 6, 7, 8, 9, 10, 11]]
print X.shape
#The angles of the knees[left, right]
angle_knees = np.vstack((data[:,2], data[:, 3])).T
print angle_knees.shape
#print pre_X.shape, data.shape
y = data[:,3] #1
#print y.shape
#getting the time vector for plotting purposes
time_stamp = np.zeros(data.shape[0])
for i in xrange(data.shape[0]):
time_stamp[i] = i*(1.0/60.0)
#print X.shape, time_stamp.shape
X = np.hstack((X, time_stamp.reshape((X.shape[0], 1))))
print X.shape
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=1)
t_test = X_test[:,-1]
t_train = X_train[:, -1]
X_train_std = preprocessing.scale(X_train[:,0:-1])
X_test_std = preprocessing.scale(X_test[:, 0:-1])
print"Test vector parameters", X_test_std.shape
#Here comes the way to sort out the data according to one the elements of it
test_sorted = np.hstack(
(t_test.reshape(X_test_std.shape[0], 1), X_test_std, y_test.reshape(X_test_std.shape[0], 1)))
test_sorted = test_sorted[np.argsort(test_sorted[:,0])] #modified
train_sorted = np.hstack((t_train.reshape(t_train.shape[0], 1), y_train.reshape(y_train.shape[0], 1) ))
train_sorted = train_sorted[np.argsort(train_sorted[:,0])]
#Splitting data for test and for validation
y_validation = y_test[0:int(y_test.shape[0]/2)]
y_test1 = y_test[int(y_test.shape[0]/2):]
X_validation = X_test_std[0:int(y_test.shape[0]/2),:]
X_test_std1 = X_test_std[int(y_test.shape[0]/2):, :]
print y_test.shape, y_validation.shape, y_test1.shape
print X_test_std.shape, X_validation.shape, X_test_std1.shape
```
## Redoing the network for the angles: Animation
```
#Grid search, random state =0: same beginning for all
alpha1 = np.linspace(0.001,0.9, 4).tolist()
momentum1 = np.linspace(0.3,0.9, 4).tolist()
params_dist = {"hidden_layer_sizes":[(10, 15), (15, 20), (15, 10), (15, 7)],
"activation":['tanh','logistic'],"algorithm":['sgd', 'l-bfgs'], "alpha":alpha1,
"learning_rate":['constant'],"max_iter":[500], "random_state":[None],
"verbose": [False], "warm_start":[False], "momentum":momentum1}
grid = GridSearchCV(MLPRegressor(), param_grid=params_dist)
grid.fit(X_validation, y_validation)
print "Best score:", grid.best_score_
print "Best parameter's set found:\n"
print grid.best_params_
new = MLPRegressor(activation= 'logistic', algorithm= 'l-bfgs', alpha= 0.001, batch_size= 'auto', beta_1= 0.9,
beta_2= 0.999, early_stopping= False, epsilon= 1e-08, hidden_layer_sizes= (10, 15),
learning_rate= 'constant', learning_rate_init= 0.001, max_iter= 500, momentum= 0.8,
nesterovs_momentum= True, power_t= 0.5, random_state= None, shuffle= False, tol= 1e-09,
validation_fraction= 0.1, verbose= False,warm_start= False)
new.fit(X_train_std, y_train)
print new.score(X_test_std, y_test)
##Getting the precision(IT MUST BE DONE THE LEAST NUMBER OF TIMES AS POSSIBLE)
print "Last used at:", datetime.datetime.now()
print "Accuracy for the test set", grid.score(X_test_std1, y_test1)
##Plotting it
%matplotlib inline
results = grid.predict(test_sorted[:, 1:-1])
plt.plot(test_sorted[:, 0], results, c='r', label="Predicted") # ( sorted time, results)
plt.plot(train_sorted[:, 0], train_sorted[:,1], c='b', label="Expected" ) #expected
plt.scatter(time_stamp, y, c='k', label="Original")
plt.legend(bbox_to_anchor=(0., 1.02, 1., .102), loc=3, ncol=3, mode="expand", borderaxespad=0.)
plt.xlabel("Time(s)")
plt.ylabel("Angle(rad)")
plt.title("MLP results vs Expected values")
plt.show()
print "Accuracy for the validation set", grid.score(X_validation, y_validation)
##save to file/ load from file
save = "no"
if save == "yes":
f= open("angles.ann", "w")
f.write(pickle.dumps(grid))
f.close
print "Saved to file"
elif save =="load":
f= open("angles.ann", "r")
grid= pickle.loads(f.read())
f.close
print "Loaded from file"
else:
print "Nothing to do"
```
## The animation itself
```
import math
from mpl_toolkits.mplot3d import Axes3D
pox =[]
poy = []
xn = []
yn = []
for i in results:
pox.append(math.cos(i))
poy.append(math.sin(i))
for i in train_sorted[:, 1]:
xn.append(math.cos(i))
yn.append(math.sin(i))
plt.plot(test_sorted[:, 0], pox, '-.', color= 'r', label="x positions")
plt.plot(test_sorted[:, 0], poy, '-.', color= 'b',label="x positions")
plt.legend(bbox_to_anchor=(0., 1.02, 1., .102), loc=1, ncol=2, mode="expand", borderaxespad=0.)
plt.show()
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.plot_wireframe(test_sorted[:, 0], pox, poy)
ax.plot_wireframe(train_sorted[:, 0], xn, yn, color = 'r')
plt.show()
```
| github_jupyter |
```
import tensorflow as tf
import pickle
import numpy as np
def load(data_path):
with open(data_path,'rb') as f:
mnist = pickle.load(f)
return mnist["training_images"], mnist["training_labels"], mnist["test_images"], mnist["test_labels"]
class MnistData:
def __init__(self, filenames, need_shuffle, datatype='training'):
all_data = []
all_labels = []
x_train, y_train, x_test, y_test = load(filenames) #"data/mnist.pkl"
if datatype=='training':
self._data = x_train / 127.5 -1
self._labels = y_train
print(self._data.shape)
print(self._labels.shape)
else:
self._data = x_test / 127.5 -1
self._labels = y_test
print(self._data.shape)
print(self._labels.shape)
self._num_examples = self._data.shape[0]
self._need_shuffle = need_shuffle
self._indicator = 0
if self._need_shuffle:
self._shuffle_data()
def _shuffle_data(self):
# [0,1,2,3,4,5] -> [5,3,2,4,0,1]
p = np.random.permutation(self._num_examples)
self._data = self._data[p]
self._labels = self._labels[p]
def next_batch(self, batch_size):
"""return batch_size examples as a batch."""
end_indicator = self._indicator + batch_size
if end_indicator > self._num_examples:
if self._need_shuffle:
self._shuffle_data()
self._indicator = 0
end_indicator = batch_size
else:
raise Exception("have no more examples")
if end_indicator > self._num_examples:
raise Exception("batch size is larger than all examples")
batch_data = self._data[self._indicator: end_indicator]
batch_labels = self._labels[self._indicator: end_indicator]
self._indicator = end_indicator
return batch_data, batch_labels
train_filenames = "../4_basic_image_recognition/data/mnist.pkl"
train_data = MnistData(train_filenames, True, 'training')
test_data = MnistData(train_filenames, False, 'test')
x = tf.placeholder(tf.float32, [None, 28*28])
y = tf.placeholder(tf.int64, [None])
x_image = tf.reshape(x, [-1, 28, 28, 1])
conv_1 = tf.layers.conv2d(inputs=x_image,
filters=32,
kernel_size=(5, 5),
padding = 'same',
activation=tf.nn.relu,
name='conv1')
pool1 = tf.layers.max_pooling2d(inputs=conv_1,
pool_size=(2, 2),
strides=(2,2),
name='pool1')
conv_2 = tf.layers.conv2d(inputs=pool1,
filters=64,
kernel_size=(5, 5),
padding = 'same',
activation=tf.nn.relu,
name='conv2')
pool2 = tf.layers.max_pooling2d(inputs=conv_2,
pool_size=(2,2),
strides=(2,2),
name='pool2')
# fc layer1
flatten = tf.layers.flatten(pool2, name='flatten')
# fc layer2
y_ = tf.layers.dense(flatten, 10)
loss = tf.losses.sparse_softmax_cross_entropy(labels=y, logits=y_)
predict = tf.argmax(y_, 1)
correct_prediction = tf.equal(predict, y)
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float64))
with tf.name_scope('train_op'):
train_op = tf.train.AdamOptimizer(1e-3).minimize(loss)
init = tf.global_variables_initializer()
batch_size = 20
train_steps = 10000
test_steps = 50
# train 10k: 71.35%
with tf.Session() as sess:
sess.run(init)
for i in range(train_steps):
batch_data, batch_labels = train_data.next_batch(batch_size)
loss_val, acc_val, _ = sess.run([loss, accuracy, train_op], feed_dict={x: batch_data, y: batch_labels})
if (i+1) % 100 == 0:
print('[Train] Step: %d, loss: %4.5f, acc: %4.5f' % (i+1, loss_val, acc_val))
if (i+1) % 1000 == 0:
all_test_acc_val = []
for j in range(test_steps):
test_batch_data, test_batch_labels = test_data.next_batch(batch_size)
test_acc_val = sess.run([accuracy], feed_dict = {x: test_batch_data, y: test_batch_labels})
all_test_acc_val.append(test_acc_val)
test_acc = np.mean(all_test_acc_val)
print('[Test ] Step: %d, acc: %4.5f' % (i+1, test_acc))
```
| github_jupyter |
# MLP example using PySNN
```
import numpy as np
import matplotlib.pyplot as plt
import torch
from torch.utils.data import DataLoader
from torch.utils.tensorboard import SummaryWriter
from torchvision import transforms
from tqdm import tqdm
from pysnn.connection import Linear
from pysnn.neuron import LIFNeuron, Input
from pysnn.learning import MSTDPET
from pysnn.encoding import PoissonEncoder
from pysnn.network import SNNNetwork
from pysnn.datasets import AND, BooleanNoise, Intensity
```
## Parameter defintions
```
# Architecture
n_in = 10
n_hidden = 5
n_out = 1
# Data
duration = 200
intensity = 50
num_workers = 0
batch_size = 1
# Neuronal Dynamics
thresh = 1.0
v_rest = 0
alpha_v = 10
tau_v = 10
alpha_t = 10
tau_t = 10
duration_refrac = 2
dt = 1
delay = 2
i_dynamics = (dt, alpha_t, tau_t, "exponential")
n_dynamics = (thresh, v_rest, alpha_v, alpha_v, dt, duration_refrac, tau_v, tau_t, "exponential")
c_dynamics = (batch_size, dt, delay)
# Learning
epochs = 100
lr = 0.1
w_init = 0.8
a = 0.0
```
## Network definition
The API is mostly the same as for regular PyTorch. The main differences are that layers are composed of a `Neuron` and `Connection` type,
and the layer has to be added to the network by calling the `add_layer` method. Lastly, all objects return both a
spike (or activation potential) object and a trace object.
```
class Network(SNNNetwork):
def __init__(self):
super(Network, self).__init__()
# Input
self.input = Input((batch_size, 1, n_in), *i_dynamics)
# Layer 1
self.mlp1_c = Linear(n_in, n_hidden, *c_dynamics)
self.mlp1_c.reset_weights(distribution="uniform") # initialize uniform between 0 and 1
self.neuron1 = LIFNeuron((batch_size, 1, n_hidden), *n_dynamics)
self.add_layer("fc1", self.mlp1_c, self.neuron1)
# Layer 2
self.mlp2_c = Linear(n_hidden, n_out, *c_dynamics)
self.mlp2_c.reset_weights(distribution="uniform")
self.neuron2 = LIFNeuron((batch_size, 1, n_out), *n_dynamics)
self.add_layer("fc2", self.mlp2_c, self.neuron2)
def forward(self, input):
x, t = self.input(input)
# Layer 1
x, _ = self.mlp1_c(x, t)
x, t = self.neuron1(x)
# Layer out
x, _ = self.mlp2_c(x, t)
x, t = self.neuron2(x)
return x, t
```
## Dataset
Simple Boolean AND dataset, generated to match the input dimensions of the network.
```
data_transform = transforms.Compose(
[
# BooleanNoise(0.2, 0.8),
Intensity(intensity)
]
)
lbl_transform = transforms.Lambda(lambda x: x * intensity)
train_dataset = AND(
data_encoder=PoissonEncoder(duration, dt),
data_transform=data_transform,
lbl_transform=lbl_transform,
repeats=n_in / 2,
)
train_dataloader = DataLoader(
train_dataset, batch_size=batch_size, shuffle=False, num_workers=num_workers
)
# Visualize input samples
_, axes = plt.subplots(1, 4, sharey=True, figsize=(25, 8))
for s in range(len(train_dataset)):
sample = train_dataset[s][0] # Drop label
sample = sample.sum(-1).numpy() # Total spike by summing over time dimension
sample = np.squeeze(sample)
axes[s].bar(range(len(sample)), sample)
axes[s].set_ylabel("Total number of spikes")
axes[s].set_xlabel("Input neuron")
```
## Training
```
device = torch.device("cpu")
net = Network()
# Learning rule definition
layers = net.layer_state_dict()
learning_rule = MSTDPET(layers, 1, 1, lr, np.exp(-1/10))
# Training loop
for _ in tqdm(range(epochs)):
for batch in train_dataloader:
sample, label = batch
# Iterate over input's time dimension
for idx in range(sample.shape[-1]):
input = sample[:, :, :, idx]
spike_net, _ = net(input)
# Determine reward, provide reward of 1 for desired behaviour, 0 otherwise.
# For positive samples (simulating an AND gate) spike as often as possible, for negative samples spike as little as possible.
if spike_net.long().view(-1) == label:
reward = 1
else:
reward = 0
# Perform a single step of the learning rule
learning_rule.step(reward)
# Reset network state (e.g. voltage, trace, spikes)
net.reset_state()
```
## Generate Data for Visualization
```
out_spikes = []
out_voltage = []
out_trace = []
for batch in train_dataloader:
single_out_s = []
single_out_v = []
single_out_t = []
sample, _ = batch
# Iterate over input's time dimension
for idx in range(sample.shape[-1]):
input = sample[:, :, :, idx]
spike_net, trace_net = net(input)
# Single timestep results logging
single_out_s.append(spike_net.clone())
single_out_t.append(trace_net.clone())
single_out_v.append(net.neuron2.v_cell.clone()) # Clone the voltage to make a copy of the value instead of using a pointer to memory
# Store batch results
out_spikes.append(torch.stack(single_out_s, dim=-1).view(-1))
out_voltage.append(torch.stack(single_out_v, dim=-1).view(-1))
out_trace.append(torch.stack(single_out_t, dim=-1).view(-1))
# Reset network state (e.g. voltage, trace, spikes)
net.reset_state()
```
### Visualize output neuron state over time
In the voltage plots the peaks never reach the voltage of 1, this is because the network has already reset the voltage of the spiking neurons during the forward pass. Thus it is not possible to register the exact voltage surpassing the threshold.
```
_, axes = plt.subplots(3, 4, sharey="row", figsize=(25, 12))
# Process every sample separately
for s in range(len(out_spikes)):
ax_col = axes[:, s]
spikes = out_spikes[s]
voltage = out_voltage[s]
trace = out_trace[s]
data_combined = [spikes, trace, voltage]
names = ["Spikes", "Trace", "Voltage"]
# Set column titles
ax_col[0].set_title(f"Sample {s}")
# Plot all states
for ax, data, name in zip(ax_col, data_combined, names):
ax.plot(data, label=name)
ax.legend()
```
| github_jupyter |
<a href="https://csdms.colorado.edu/wiki/ESPIn2020"><img style="float: center; width: 75%" src="../../../media/ESPIn.png"></a>
# Introduction to Landlab: Creating a simple 2D scarp diffusion model
<hr>
<small>For more Landlab tutorials, click here: <a href="https://landlab.readthedocs.io/en/latest/user_guide/tutorials.html">https://landlab.readthedocs.io/en/latest/user_guide/tutorials.html</a></small>
<hr>
This tutorial illustrates how you can use Landlab to construct a simple two-dimensional numerical model on a regular (raster) grid, using a simple forward-time, centered-space numerical scheme. The example is the erosional degradation of an earthquake fault scarp, and which evolves over time in response to the gradual downhill motion of soil. Here we use a simple "geomorphic diffusion" model for landform evolution, in which the downhill flow of soil is assumed to be proportional to the (downhill) gradient of the land surface multiplied by a transport coefficient.
We start by importing the [numpy](https://numpy.org) and [matplotlib](https://matplotlib.org) libraries:
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
## Part 1: 1D version using numpy
This example uses a finite-volume numerical solution to the 2D diffusion equation. The 2D diffusion equation in this case is derived as follows. Continuity of mass states that:
$\frac{\partial z}{\partial t} = -\nabla \cdot \mathbf{q}_s$,
where $z$ is elevation, $t$ is time, the vector $\mathbf{q}_s$ is the volumetric soil transport rate per unit width, and $\nabla$ is the divergence operator (here in two dimensions). (Note that we have omitted a porosity factor here; its effect will be subsumed in the transport coefficient). The sediment flux vector depends on the slope gradient:
$\mathbf{q}_s = -D \nabla z$,
where $D$ is a transport-rate coefficient---sometimes called *hillslope diffusivity*---with dimensions of length squared per time. Combining the two, and assuming $D$ is uniform, we have a classical 2D diffusion equation:
$\frac{\partial z}{\partial t} = -\nabla^2 z$.
In this first example, we will create a our 1D domain in $x$ and $z$, and set a value for $D$.
This means that the equation we solve will be in 1D.
$\frac{d z}{d t} = \frac{d q_s}{dx}$,
where
$q_s = -D \frac{d z}{dx}$
```
dx = 1
x = np.arange(0, 100, dx, dtype=float)
z = np.zeros(x.shape, dtype=float)
D = 0.01
```
Next we must create our fault by uplifting some of the domain. We will increment all elements of `z` in which `x>50`.
```
z[x>50] += 100
```
Finally, we will diffuse our fault for 1,000 years.
We will use a timestep with a [Courant–Friedrichs–Lewy condition](https://en.wikipedia.org/wiki/Courant–Friedrichs–Lewy_condition) of $C_{cfl}=0.2$. This will keep our solution numerically stable.
$C_{cfl} = \frac{\Delta t D}{\Delta x^2} = 0.2$
```
dt = 0.2 * dx * dx / D
total_time = 1e3
nts = int(total_time/dt)
z_orig = z.copy()
for i in range(nts):
qs = -D * np.diff(z)/dx
dzdt = -np.diff(qs)/dx
z[1:-1] += dzdt*dt
plt.plot(x, z_orig, label="Original Profile")
plt.plot(x, z, label="Diffused Profile")
plt.legend()
```
The prior example is pretty simple. If this was all you needed to do, you wouldn't need Landlab.
But what if you wanted...
... to use the same diffusion model in 2D instead of 1D.
... to use an irregular grid (in 1 or 2D).
... wanted to combine the diffusion model with a more complex model.
... have a more complex model you want to use over and over again with different boundary conditions.
These are the sorts of problems that Landlab was designed to solve.
In the next two sections we will introduce some of the core capabilities of Landlab.
In Part 2 we will use the RasterModelGrid, fields, and a numerical utility for calculating flux divergence.
In Part 3 we will use the HexagonalModelGrid.
In Part 4 we will use the LinearDiffuser component.
## Part 2: 2D version using Landlab's Model Grids
The Landlab model grids are data structures that represent the model domain (the variable `x` in our prior example). Here we will use `RasterModelGrid` which creates a grid with regularly spaced square grid elements. The RasterModelGrid knows how the elements are connected and how far apart they are.
Lets start by creating a RasterModelGrid class. First we need to import it.
```
from landlab import RasterModelGrid
```
### (a) Explore the RasterModelGrid
Before we make a RasterModelGrid for our fault example, lets explore the Landlab model grid.
Landlab considers the grid as a "dual" graph. Two sets of points, lines and polygons that represent 2D space.
The first graph considers points called "nodes" that are connected by lines called "links". The area that surrounds each node is called a "cell".
First, the nodes
```
from landlab.plot.graph import plot_graph
grid = RasterModelGrid((4, 5), xy_spacing=(3,4))
plot_graph(grid, at="node")
```
You can see that the nodes are points and they are numbered with unique IDs from lower left to upper right.
Next the links
```
plot_graph(grid, at="link")
```
which are lines that connect the nodes and each have a unique ID number.
And finally, the cells
```
plot_graph(grid, at="cell")
```
which are polygons centered around the nodes.
Landlab is a "dual" graph because it also keeps track of a second set of points, lines, and polygons ("corners", "faces", and "patches"). We will not focus on them further.
### *Exercises for section 2a*
(2a.1) Create an instance of a `RasterModelGrid` with 5 rows and 7 columns, with a spacing between nodes of 10 units. Plot the node layout, and identify the ID number of the center-most node.
```
# (enter your solution to 2a.1 here)
rmg = RasterModelGrid((5, 7), 10.0)
plot_graph(rmg, at='node')
```
(2a.2) Find the ID of the cell that contains this node.
```
# (enter your solution to 2a.2 here)
plot_graph(rmg, at='cell')
```
(2a.3) Find the ID of the horizontal link that connects to the last node on the right in the middle column.
```
# (enter your solution to 2a.3 here)
plot_graph(rmg, at='link')
```
### (b) Use the RasterModelGrid for 2D diffusion
Lets continue by making a new grid that is bigger. We will use this for our next fault diffusion example.
The syntax in the next line says: create a new *RasterModelGrid* object called **mg**, with 25 rows, 40 columns, and a grid spacing of 10 m.
```
mg = RasterModelGrid((25, 40), 10.0)
```
Note the use of object-oriented programming here. `RasterModelGrid` is a class; `mg` is a particular instance of that class, and it contains all the data necessary to fully describe the topology and geometry of this particular grid.
Next we'll add a *data field* to the grid, to represent the elevation values at grid nodes. The "dot" syntax below indicates that we are calling a function (or *method*) that belongs to the *RasterModelGrid* class, and will act on data contained in **mg**. The arguments indicate that we want the data elements attached to grid nodes (rather than links, for example), and that we want to name this data field `topographic__elevation`. The `add_zeros` method returns the newly created NumPy array.
```
z = mg.add_zeros('topographic__elevation', at='node')
```
The above line of code creates space in memory to store 1,000 floating-point values, which will represent the elevation of the land surface at each of our 1,000 grid nodes.
Let's plot the positions of all the grid nodes. The nodes' *(x,y)* positions are stored in the arrays `mg.x_of_node` and `mg.y_of_node`, respectively.
```
plt.plot(mg.x_of_node, mg.y_of_node, '.')
```
If we bothered to count, we'd see that there are indeed 1,000 grid nodes, and a corresponding number of `z` values:
```
len(z)
```
Now for some tectonics. Let's say there's a fault trace that angles roughly east-northeast. We can describe the trace with the equation for a line. One trick here: by using `mg.x_of_node`, in the line of code below, we are calculating a *y* (i.e., north-south) position of the fault trace for each grid node---meaning that this is the *y* coordinate of the trace at the *x* coordinate of a given node.
```
fault_trace_y = 50.0 + 0.25 * mg.x_of_node
```
Here comes the earthquake. For all the nodes north of the fault (i.e., those with a *y* coordinate greater than the corresponding *y* coordinate of the fault trace), we'll add elevation equal to 10 meters plus a centimeter for every meter east along the grid (just to make it interesting):
```
z[mg.y_of_node >
fault_trace_y] += 10.0 + 0.01 * mg.x_of_node[mg.y_of_node > fault_trace_y]
```
(A little bit of Python under the hood: the statement `mg.y_of_node > fault_trace_y` creates a 1000-element long boolean array; placing this within the index brackets will select only those array entries that correspond to `True` in the boolean array)
Let's look at our newly created initial topography using Landlab's *imshow_node_grid* plotting function (which we first need to import).
```
from landlab.plot.imshow import imshow_grid
imshow_grid(mg, 'topographic__elevation')
```
To finish getting set up, we will define two parameters: the transport ("diffusivity") coefficient, `D`, and the time-step size, `dt`. (The latter is set using the Courant condition for a forward-time, centered-space finite-difference solution; you can find the explanation in most textbooks on numerical methods).
```
D = 0.01 # m2/yr transport coefficient
dt = 0.2 * mg.dx * mg.dx / D
dt
```
Boundary conditions: for this example, we'll assume that the east and west sides are closed to flow of sediment, but that the north and south sides are open. (The order of the function arguments is east, north, west, south)
```
mg.set_closed_boundaries_at_grid_edges(True, False, True, False)
```
*A note on boundaries:* with a Landlab raster grid, all the perimeter nodes are boundary nodes. In this example, there are 24 + 24 + 39 + 39 = 126 boundary nodes. The previous line of code set those on the east and west edges to be **closed boundaries**, while those on the north and south are **open boundaries** (the default). All the remaining nodes are known as **core** nodes. In this example, there are 1000 - 126 = 874 core nodes:
```
len(mg.core_nodes)
```
One more thing before we run the time loop: we'll create an array to contain soil flux. In the function call below, the first argument tells Landlab that we want one value for each grid link, while the second argument provides a name for this data *field*:
```
qs = mg.add_zeros('sediment_flux', at='link')
```
And now for some landform evolution. We will loop through 25 iterations, representing 50,000 years. On each pass through the loop, we do the following:
1. Calculate, and store in the array `g`, the gradient between each neighboring pair of nodes. These calculations are done on **links**. The gradient value is a positive number when the gradient is "uphill" in the direction of the link, and negative when the gradient is "downhill" in the direction of the link. On a raster grid, link directions are always in the direction of increasing $x$ ("horizontal" links) or increasing $y$ ("vertical" links).
2. Calculate, and store in the array `qs`, the sediment flux between each adjacent pair of nodes by multiplying their gradient by the transport coefficient. We will only do this for the **active links** (those not connected to a closed boundary, and not connecting two boundary nodes of any type); others will remain as zero.
3. Calculate the resulting net flux at each node (positive=net outflux, negative=net influx). The negative of this array is the rate of change of elevation at each (core) node, so store it in a node array called `dzdt'.
4. Update the elevations for the new time step.
```
for i in range(25):
g = mg.calc_grad_at_link(z)
qs[mg.active_links] = -D * g[mg.active_links]
dzdt = -mg.calc_flux_div_at_node(qs)
z[mg.core_nodes] += dzdt[mg.core_nodes] * dt
```
Let's look at how our fault scarp has evolved.
```
imshow_grid(mg, 'topographic__elevation')
```
Notice that we have just created and run a 2D model of fault-scarp creation and diffusion with fewer than two dozen lines of code. How long would this have taken to write in C or Fortran?
While it was very very easy to write in 1D, writing this in 2D would mean we would have needed to keep track of the adjacency of the different parts of the grid. This is the primary problem that the Landlab grids are meant to solve.
Think about how difficult this would be to hand code if the grid were irregular or hexagonal. In order to conserve mass and implement the differential equation you would need to know how nodes were conected, how long the links were, and how big each cell was.
We do such an example after the next section.
### *Exercises for section 2b*
(2b .1) Create an instance of a `RasterModelGrid` called `mygrid`, with 16 rows and 25 columns, with a spacing between nodes of 5 meters. Use the `plot` function in the `matplotlib` library to make a plot that shows the position of each node marked with a dot (hint: see the plt.plot() example above).
```
# (enter your solution to 2b.1 here)
mygrid = RasterModelGrid((16, 25), xy_spacing=5.0)
plt.plot(mygrid.x_of_node, mygrid.y_of_node, '.')
```
(2b.2) Query the grid variables `number_of_nodes` and `number_of_core_nodes` to find out how many nodes are in your grid, and how many of them are core nodes.
```
# (enter your solution to 2b.2 here)
print(mygrid.number_of_nodes)
print(mygrid.number_of_core_nodes)
```
(2b.3) Add a new field to your grid, called `temperature` and attached to nodes. Have the initial values be all zero.
```
# (enter your solution to 2b.3 here)
temp = mygrid.add_zeros('temperature', at='node')
```
(2b.4) Change the temperature of nodes in the top (north) half of the grid to be 10 degrees C. Use the `imshow_grid` function to display a shaded image of the elevation field.
```
# (enter your solution to 2b.4 here)
temp[mygrid.y_of_node >= 40.0] = 10.0
imshow_grid(mygrid, 'temperature')
```
(2b.5) Use the grid function `set_closed_boundaries_at_grid_edges` to assign closed boundaries to the right and left sides of the grid.
```
# (enter your solution to 2b.5 here)
mygrid.set_closed_boundaries_at_grid_edges(True, False, True, False)
imshow_grid(mygrid, 'temperature', color_for_closed='c')
```
(2b.6) Create a new field of zeros called `heat_flux` and attached to links. Using the `number_of_links` grid variable, verify that your new field array has the correct number of items.
```
# (enter your solution to 2b.6 here)
Q = mygrid.add_zeros('heat_flux', at='link')
print(mygrid.number_of_links)
print(len(Q))
```
(2b.7) Use the `calc_grad_at_link` grid function to calculate the temperature gradients at all the links in the grid. Given the node spacing and the temperatures you assigned to the top versus bottom grid nodes, what do you expect the maximum temperature gradient to be? Print the values in the gradient array to verify that this is indeed the maximum temperature gradient.
```
# (enter your solution to 2b.7 here)
print('Expected max gradient is 2 C/m')
temp_grad = mygrid.calc_grad_at_link(temp)
print(temp_grad)
```
(2b.8) Back to hillslopes: Reset the values in the elevation field of the grid `mg` to zero. Then copy and paste the time loop above (i.e., the block in Section 2b that starts with `for i in range(25):`) below. Modify the last line to add uplift of the hillslope material at a rate `uplift_rate` = 0.0001 m/yr (hint: the amount of uplift in each iteration should be the uplift rate times the time-step duration). Then run the block and plot the resulting topography. Try experimenting with different uplift rates and different values of `D`.
```
# (enter your solution to 2b.8 here)
z[:] = 0.0
uplift_rate = 0.0001
for i in range(25):
g = mg.calc_grad_at_link(z)
qs[mg.active_links] = -D * g[mg.active_links]
dzdt = -mg.calc_flux_div_at_node(qs)
z[mg.core_nodes] += (dzdt[mg.core_nodes] + uplift_rate) * dt
imshow_grid(mg, z)
```
### (c) What's going on under the hood?
This example uses a finite-volume numerical solution to the 2D diffusion equation. The 2D diffusion equation in this case is derived as follows. Continuity of mass states that:
$\frac{\partial z}{\partial t} = -\nabla \cdot \mathbf{q}_s$,
where $z$ is elevation, $t$ is time, the vector $\mathbf{q}_s$ is the volumetric soil transport rate per unit width, and $\nabla$ is the divergence operator (here in two dimensions). (Note that we have omitted a porosity factor here; its effect will be subsumed in the transport coefficient). The sediment flux vector depends on the slope gradient:
$\mathbf{q}_s = -D \nabla z$,
where $D$ is a transport-rate coefficient---sometimes called *hillslope diffusivity*---with dimensions of length squared per time. Combining the two, and assuming $D$ is uniform, we have a classical 2D diffusion equation:
$\frac{\partial z}{\partial t} = -\nabla^2 z$.
For the numerical solution, we discretize $z$ at a series of *nodes* on a grid. The example in this notebook uses a Landlab *RasterModelGrid*, in which every interior node sits inside a cell of width $\Delta x$, but we could alternatively have used any grid type that provides nodes, links, and cells.
The gradient and sediment flux vectors will be calculated at the *links* that connect each pair of adjacent nodes. These links correspond to the mid-points of the cell faces, and the values that we assign to links represent the gradients and fluxes, respectively, along the faces of the cells.
The flux divergence, $\nabla \mathbf{q}_s$, will be calculated by summing, for every cell, the total volume inflows and outflows at each cell face, and dividing the resulting sum by the cell area. Note that for a regular, rectilinear grid, as we use in this example, this finite-volume method is equivalent to a finite-difference method.
To advance the solution in time, we will use a simple explicit, forward-difference method. This solution scheme for a given node $i$ can be written:
$\frac{z_i^{t+1} - z_i^t}{\Delta t} = -\frac{1}{A_i} \sum\limits_{j=1}^{N_i} \delta (l_{ij}) q_s (l_{ij}) \lambda(l_{ij})$.
Here the superscripts refer to time steps, $\Delta t$ is time-step size, $q_s(l_{ij})$ is the sediment flux per width associated with the link that crosses the $j$-th face of the cell at node $i$, $\lambda(l_{ij})$ is the width of the cell face associated with that link ($=\Delta x$ for a regular uniform grid), and $N_i$ is the number of active links that connect to node $i$. The variable $\delta(l_{ij})$ contains either +1 or -1: it is +1 if link $l_{ij}$ is oriented away from the node (in which case positive flux would represent material leaving its cell), or -1 if instead the link "points" into the cell (in which case positive flux means material is entering).
To get the fluxes, we first calculate the *gradient*, $G$, at each link, $k$:
$G(k) = \frac{z(H_k) - z(T_k)}{L_k}$.
Here $H_k$ refers the *head node* associated with link $k$, $T_k$ is the *tail node* associated with link $k$. Each link has a direction: from the tail node to the head node. The length of link $k$ is $L_k$ (equal to $\Delta x$ is a regular uniform grid). What the above equation says is that the gradient in $z$ associated with each link is simply the difference in $z$ value between its two endpoint nodes, divided by the distance between them. The gradient is positive when the value at the head node (the "tip" of the link) is greater than the value at the tail node, and vice versa.
The calculation of gradients in $z$ at the links is accomplished with the `calc_grad_at_link` function. The sediment fluxes are then calculated by multiplying the link gradients by $-D$. Once the fluxes at links have been established, the `calc_flux_div_at_node` function performs the summation of fluxes.
### *Exercises for section 2c*
(2c.1) Make a 3x3 `RasterModelGrid` called `tinygrid`, with a cell spacing of 2 m. Use the `plot_graph` function to display the nodes and their ID numbers.
```
# (enter your solution to 2c.1 here)
tinygrid = RasterModelGrid((3, 3), 2.0)
plot_graph(tinygrid, at='node')
```
(2c.2) Give your `tinygrid` a node field called `height` and set the height of the center-most node to 0.5. Use `imshow_grid` to display the height field.
```
# (enter your solution to 2c.2 here)
ht = tinygrid.add_zeros('height', at='node')
ht[4] = 0.5
imshow_grid(tinygrid, ht)
```
(2c.3) The grid should have 12 links (extra credit: verify this with `plot_graph`). When you compute gradients, which of these links will have non-zero gradients? What will the absolute value(s) of these gradients be? Which (if any) will have positive gradients and which negative? To codify your answers, make a 12-element numpy array that contains your predicted gradient value for each link.
```
# (enter your solution to 2c.3 here)
plot_graph(tinygrid, at='link')
pred_grad = np.array([0, 0, 0, 0.25, 0, 0.25, -0.25, 0, -0.25, 0, 0, 0])
print(pred_grad)
```
(2c.4) Test your prediction by running the `calc_grad_at_link` function on your tiny grid. Print the resulting array and compare it with your predictions.
```
# (enter your solution to 2c.4 here)
grad = tinygrid.calc_grad_at_link(ht)
print(grad)
```
(2c.5) Suppose the flux of soil per unit cell width is defined as -0.01 times the height gradient. What would the flux be at the those links that have non-zero gradients? Test your prediction by creating and printing a new array whose values are equal to -0.01 times the link-gradient values.
```
# (enter your solution to 2c.5 here)
flux = -0.01 * grad
print(flux)
```
(2c.6) Consider the net soil accumulation or loss rate around the center-most node in your tiny grid (which is the only one that has a cell). The *divergence* of soil flux can be represented numerically as the sum of the total volumetric soil flux across each of the cell's four faces. What is the flux across each face? (Hint: multiply by face width) What do they add up to? Test your prediction by running the grid function `calc_flux_div_at_node` (hint: pass your unit flux array as the argument). What are the units of the divergence values returned by the `calc_flux_div_at_node` function?
```
# (enter your solution to 2c.6 here)
print('predicted div is 0 m/yr')
dqsdx = tinygrid.calc_flux_div_at_node(flux)
print(dqsdx)
```
## Part 3: Hexagonal grid
Next we will use an non-raster Landlab grid.
We start by making a random set of points with x values between 0 and 400 and y values of 0 and 250. We then add zeros to our grid at a field called "topographic__elevation" and plot the node locations.
Note that the syntax here is exactly the same as in the RasterModelGrid example (once the grid has been created).
```
from landlab import HexModelGrid
mg = HexModelGrid((25, 40), 10, node_layout="rect")
z = mg.add_zeros('topographic__elevation', at='node')
plt.plot(mg.x_of_node, mg.y_of_node, '.')
```
Next we create our fault trace and uplift the hanging wall.
We can plot just like we did with the RasterModelGrid.
```
fault_trace_y = 50.0 + 0.25 * mg.x_of_node
z[mg.y_of_node >
fault_trace_y] += 10.0 + 0.01 * mg.x_of_node[mg.y_of_node > fault_trace_y]
imshow_grid(mg, "topographic__elevation")
```
And we can use the same code as before to create a diffusion model!
Landlab supports multiple grid types. You can read more about them [here](https://landlab.readthedocs.io/en/latest/reference/grid/index.html).
```
qs = mg.add_zeros('sediment_flux', at='link')
for i in range(25):
g = mg.calc_grad_at_link(z)
qs[mg.active_links] = -D * g[mg.active_links]
dzdt = -mg.calc_flux_div_at_node(qs)
z[mg.core_nodes] += dzdt[mg.core_nodes] * dt
imshow_grid(mg, 'topographic__elevation')
```
### *Exercises for section 3*
(3.1-6) Repeat the exercises from section 2c, but this time using a hexagonal tiny grid called `tinyhex`. Your grid should have 7 nodes: one core node and 6 perimeter nodes. (Hints: use `node_layout = 'hex'`, and make a grid with 3 rows and 2 base-row columns.)
```
# (enter your solution to 3.1 here)
tinyhex = HexModelGrid((3, 2), 2.0)
plot_graph(tinyhex, at='node')
# (enter your solution to 3.2 here)
hexht = tinyhex.add_zeros('height', at='node')
hexht[3] = 0.5
imshow_grid(tinyhex, hexht)
# (enter your solution to 3.3 here)
plot_graph(tinyhex, at='link')
pred_grad = np.array([0, 0, 0.25, 0.25, 0, 0.25, -0.25, 0, -0.25, -0.25, 0, 0])
print(pred_grad)
# (enter your solution to 3.4 here)
hexgrad = tinyhex.calc_grad_at_link(hexht)
print(hexgrad)
# (enter your solution to 3.5 here)
hexflux = -0.01 * hexgrad
print(hexflux)
# (enter your solution to 3.6 here)
print(tinyhex.length_of_face)
print(tinyhex.area_of_cell)
total_outflux = 6 * 0.0025 * tinyhex.length_of_face[0]
divergence = total_outflux / tinyhex.area_of_cell[0]
print(total_outflux)
print(divergence)
```
## Part 4: Landlab Components
Finally we will use a Landlab component, called the LinearDiffuser [link to its documentation](https://landlab.readthedocs.io/en/latest/reference/components/diffusion.html).
Landlab was designed to have many of the utilities like `calc_grad_at_link`, and `calc_flux_divergence_at_node` to help you make your own models. Sometimes, however, you may use such a model over and over and over. Then it is nice to be able to put it in its own python class with a standard interface.
This is what a Landlab Component is.
There is a whole [tutorial on components](../component_tutorial/component_tutorial.ipynb) and a [page on the User Guide](https://landlab.readthedocs.io/en/latest/user_guide/components.html). For now we will just show you what the prior example looks like if we use the LinearDiffuser.
First we import it, set up the grid, and uplift our fault block.
```
from landlab.components import LinearDiffuser
mg = HexModelGrid((25, 40), 10, node_layout="rect")
z = mg.add_zeros('topographic__elevation', at='node')
fault_trace_y = 50.0 + 0.25 * mg.x_of_node
z[mg.y_of_node >
fault_trace_y] += 10.0 + 0.01 * mg.x_of_node[mg.y_of_node > fault_trace_y]
```
Next we instantiate a LinearDiffuser. We have to tell the component what value to use for the diffusivity.
```
ld = LinearDiffuser(mg, linear_diffusivity=D)
```
Finally we run the component forward in time and plot. Like many Landlab components, the LinearDiffuser has a method called "run_one_step" that takes one input, the timestep dt. Calling this method runs the LinearDiffuser forward in time by an increment dt.
```
for i in range(25):
ld.run_one_step(dt)
imshow_grid(mg, 'topographic__elevation')
```
### *Exercises for section 4*
(4.1) Repeat the steps above that instantiate and run a `LinearDiffuser` component, but this time give it a `RasterModelGrid`. Use `imshow_grid` to display the topography below.
```
# (enter your solution to 4.1 here)
rmg = RasterModelGrid((25, 40), 10)
z = rmg.add_zeros('topographic__elevation', at='node')
fault_trace_y = 50.0 + 0.25 * rmg.x_of_node
z[rmg.y_of_node >
fault_trace_y] += 10.0 + 0.01 * rmg.x_of_node[rmg.y_of_node > fault_trace_y]
ld = LinearDiffuser(rmg, linear_diffusivity=D)
for i in range(25):
ld.run_one_step(dt)
imshow_grid(rmg, 'topographic__elevation')
```
(4.2) Using either a raster or hex grid (your choice) with a `topographic__elevation` field that is initially all zeros, write a modified version of the loop that adds uplift to the core nodes each iteration, at a rate of 0.0001 m/yr. Run the model for enough time to accumulate 10 meters of uplift. Plot the terrain to verify that the land surface height never gets higher than 10 m.
```
# (enter your solution to 4.2 here)
rmg = RasterModelGrid((40, 40), 10) # while we're at it, make it a bit bigger
z = rmg.add_zeros('topographic__elevation', at='node')
ld = LinearDiffuser(rmg, linear_diffusivity=D)
for i in range(50):
ld.run_one_step(dt)
z[rmg.core_nodes] += dt * 0.0001
imshow_grid(rmg, 'topographic__elevation')
```
(4.3) Now run the same model long enough that it reaches (or gets very close to) a dynamic equilibrium between uplift and erosion. What shape does the hillslope have?
```
# (enter your solution to 4.3 here)
z[:] = 0.0
uplift_rate = 0.0001
for i in range(4000):
ld.run_one_step(dt)
z[rmg.core_nodes] += dt * uplift_rate
imshow_grid(rmg, 'topographic__elevation')
plt.figure()
plt.plot(rmg.x_of_node, z, '.')
```
(BONUS CHALLENGE QUESTION) Derive an analytical solution for the cross-sectional shape of your steady-state hillslope. Plot this solution next to the actual model's cross-section.
#### *SOLUTION (derivation)*
##### Derivation of the original governing equation
(Note: you could just start with the governing equation and go from there, but we include this here for completeness).
Consider a topographic profile across a hillslope. The horizontal coordinate along the profile is $x$, measured from the left side of the profile (i.e., the base of the hill on the left side, where $x=0$). The horizontal coordinate perpendicular to the profile is $y$. Assume that at any time, the hillslope is perfectly symmetrical in the $y$ direction, and that there is no flow of soil in this direction.
Now consider a vertical column of soil somewhere along the profile. The left side of the column is at position $x$, and the right side is at position $x+\Delta x$, with $\Delta x$ being the width of the column in the $x$ direction. The width of the column in the $y$ direction is $W$. The height of the column, $z$, is also the height of the land surface at that location. Height is measured relative to the height of the base of the slope (in other words, $z(0) = 0$).
The total mass of soil inside the column, and above the slope base, is equal to the volume of soil material times its density times the fraction of space that it fills, which is 1 - porosity. Denoting soil particle density by $\rho$ and porosity by $\phi$, the soil mass in a column of height $h$ is
$m = (1-\phi ) \rho \Delta x W z$.
Conservation of mass dictates that the rate of change of mass equals the rate of mass inflow minus the rate of mass outflow. Assume that mass enters or leaves only by (1) soil creep, and (2) uplift of the hillslope material relative to the elevation of the hillslope base. The rate of the latter, in terms of length per time, will be denoted $U$. The rate of soil creep at a particular position $x$, in terms of bulk volume (including pores) per time per width, will be denoted $q_s(x)$. With this definition in mind, mass conservation dictates that:
$\frac{\partial (1-\phi ) \rho \Delta x W z}{\partial t} = \rho (1-\phi ) \Delta x W U + \rho (1-\phi ) q_s(x) - \rho (1-\phi ) q_s(x+\Delta x)$.
Assume that porosity and density are steady and uniform. Then,
$\frac{\partial z}{\partial t} = U + \frac{q_s(x) - q_s(x+\Delta x)}{\Delta x}$.
Factoring out -1 from the right-most term, and taking the limit as $\Delta x\rightarrow 0$, we get a differential equation that expresses conservation of mass for this situation:
$\frac{\partial z}{\partial t} = U - \frac{\partial q_s}{\partial x}$.
Next, substitute the soil-creep rate law
$q_s = -D \frac{\partial z}{\partial x}$,
to obtain
$\frac{\partial z}{\partial t} = U + D \frac{\partial^2 z}{\partial x^2}$.
##### Steady state
Steady means $dz/dt = 0$. If we go back to the mass conservation law a few steps ago and apply steady state, we find
$\frac{dq_s}{dx} = U$.
If you think of a hillslope that slopes down to the right, you can think of this as indicating that for every step you take to the right, you get another increment of incoming soil via uplift relative to baselevel. (Turns out it works the same way for a slope that angles down the left, but that's less obvious in the above math)
Integrate to get:
$q_s = Ux + C_1$, where $C_1$ is a constant of integration.
To evaluate the integration constant, let's assume the crest of the hill is right in the middle of the profile, at $x=L/2$, with $L$ being the total length of the profile. Net downslope soil flux will be zero at the crest (where the slope is zero), so for this location:
$q_s = 0 = UL/2 + C_1$,
and therefore,
$C_1 = -UL/2$,
and
$q_s = U (x - L/2)$.
Now substitute the creep law for $q_s$ and divide both sides by $-D$:
$\frac{dz}{dx} = \frac{U}{D} (L/2 - x)$.
Integrate:
$z = \frac{U}{D} (Lx/2 - x^2/2) + C_2$.
To evaluate $C_2$, recall that $z(0)=0$ (and also $z(L)=0$), so $C_2=0$. Hence, here's our analytical solution, which describes a parabola:
$\boxed{z = \frac{U}{2D} (Lx - x^2)}$.
```
# (enter your solution to the bonus challenge question here)
L = 390.0 # hillslope length, m
x_analytic = np.arange(0.0, L)
z_analytic = 0.5 * (uplift_rate / D) * (L * x_analytic - x_analytic * x_analytic)
plt.plot(rmg.x_of_node, z, '.')
plt.plot(x_analytic, z_analytic, 'r')
```
Hey, hang on a minute, that's not a very good fit! What's going on?
Turns out our 2D hillslope isn't as tall as the idealized 1D profile because of the boundary conditions: with soil free to flow east and west as well as north and south, the crest ends up lower than it would be if it were perfectly symmetrical in one direction.
So let's try re-running the numerical model, but this time with the north and south boundaries closed so that the hill shape becomes uniform in the $y$ direction:
```
rmg = RasterModelGrid((40, 40), 10)
z = rmg.add_zeros('topographic__elevation', at='node')
rmg.set_closed_boundaries_at_grid_edges(False, True, False, True) # closed on N and S
ld = LinearDiffuser(rmg, linear_diffusivity=D)
for i in range(4000):
ld.run_one_step(dt)
z[rmg.core_nodes] += dt * uplift_rate
imshow_grid(rmg, 'topographic__elevation')
plt.plot(rmg.x_of_node, z, '.')
plt.plot(x_analytic, z_analytic, 'r')
```
That's more like it!
Congratulations on making it to the end of this tutorial!
### Click here for more <a href="https://landlab.readthedocs.io/en/latest/user_guide/tutorials.html">Landlab tutorials</a>
| github_jupyter |
## Padding Characters around Strings
Let us go through how to pad characters to strings using Spark Functions.
```
%%HTML
<iframe width="560" height="315" src="https://www.youtube.com/embed/w85C18tvYNA?rel=0&controls=1&showinfo=0" frameborder="0" allowfullscreen></iframe>
```
* We typically pad characters to build fixed length values or records.
* Fixed length values or records are extensively used in Mainframes based systems.
* Length of each and every field in fixed length records is predetermined and if the value of the field is less than the predetermined length then we pad with a standard character.
* In terms of numeric fields we pad with zero on the leading or left side. For non numeric fields, we pad with some standard character on leading or trailing side.
* We use `lpad` to pad a string with a specific character on leading or left side and `rpad` to pad on trailing or right side.
* Both lpad and rpad, take 3 arguments - column or expression, desired length and the character need to be padded.
Let us start spark context for this Notebook so that we can execute the code provided. You can sign up for our [10 node state of the art cluster/labs](https://labs.itversity.com/plans) to learn Spark SQL using our unique integrated LMS.
```
from pyspark.sql import SparkSession
import getpass
username = getpass.getuser()
spark = SparkSession. \
builder. \
config('spark.ui.port', '0'). \
config("spark.sql.warehouse.dir", f"/user/{username}/warehouse"). \
enableHiveSupport(). \
appName(f'{username} | Python - Processing Column Data'). \
master('yarn'). \
getOrCreate()
```
If you are going to use CLIs, you can use Spark SQL using one of the 3 approaches.
**Using Spark SQL**
```
spark2-sql \
--master yarn \
--conf spark.ui.port=0 \
--conf spark.sql.warehouse.dir=/user/${USER}/warehouse
```
**Using Scala**
```
spark2-shell \
--master yarn \
--conf spark.ui.port=0 \
--conf spark.sql.warehouse.dir=/user/${USER}/warehouse
```
**Using Pyspark**
```
pyspark2 \
--master yarn \
--conf spark.ui.port=0 \
--conf spark.sql.warehouse.dir=/user/${USER}/warehouse
```
### Tasks - Padding Strings
Let us perform simple tasks to understand the syntax of `lpad` or `rpad`.
* Create a Dataframe with single value and single column.
* Apply `lpad` to pad with - to Hello to make it 10 characters.
```
l = [('X',)]
df = spark.createDataFrame(l).toDF("dummy")
from pyspark.sql.functions import lit, lpad
df.select(lpad(lit("Hello"), 10, "-").alias("dummy")).show()
```
* Let’s create the **employees** Dataframe
```
employees = [(1, "Scott", "Tiger", 1000.0,
"united states", "+1 123 456 7890", "123 45 6789"
),
(2, "Henry", "Ford", 1250.0,
"India", "+91 234 567 8901", "456 78 9123"
),
(3, "Nick", "Junior", 750.0,
"united KINGDOM", "+44 111 111 1111", "222 33 4444"
),
(4, "Bill", "Gomes", 1500.0,
"AUSTRALIA", "+61 987 654 3210", "789 12 6118"
)
]
employeesDF = spark.createDataFrame(employees). \
toDF("employee_id", "first_name",
"last_name", "salary",
"nationality", "phone_number",
"ssn"
)
employeesDF.show()
employeesDF.printSchema()
```
* Use **pad** functions to convert each of the field into fixed length and concatenate. Here are the details for each of the fields.
* Length of the employee_id should be 5 characters and should be padded with zero.
* Length of first_name and last_name should be 10 characters and should be padded with - on the right side.
* Length of salary should be 10 characters and should be padded with zero.
* Length of the nationality should be 15 characters and should be padded with - on the right side.
* Length of the phone_number should be 17 characters and should be padded with - on the right side.
* Length of the ssn can be left as is. It is 11 characters.
* Create a new Dataframe **empFixedDF** with column name **employee**. Preview the data by disabling truncate.
```
from pyspark.sql.functions import lpad, rpad, concat
empFixedDF = employeesDF.select(
concat(
lpad("employee_id", 5, "0"),
rpad("first_name", 10, "-"),
rpad("last_name", 10, "-"),
lpad("salary", 10, "0"),
rpad("nationality", 15, "-"),
rpad("phone_number", 17, "-"),
"ssn"
).alias("employee")
)
empFixedDF.show(truncate=False)
```
| github_jupyter |
# NBA Free throw analysis
Now let's see some of these methods in action on real world data.
I'm not a basketball guru by any means, but I thought it would be fun to see whether we can find players that perform differently in free throws when playing at home versus away.
[Basketballvalue.com](http://basketballvalue.com/downloads.php) has
some nice play by play data on season and playoff data between 2007 and 2012, which we will use for this analysis.
It's not perfect, for example it only records player's last names, but it will do for the purpose of demonstration.
## Getting data:
- Download and extract play by play data from 2007 - 2012 data from http://basketballvalue.com/downloads.php
- Concatenate all text files into file called `raw.data`
- Run following to extract free throw data into `free_throws.csv`
```
cat raw.data | ack Free Throw | sed -E 's/[0-9]+([A-Z]{3})([A-Z]{3})[[:space:]][0-9]*[[:space:]].?[0-9]{2}:[0-9]{2}:[0-9]{2}[[:space:]]*\[([A-z]{3}).*\][[:space:]](.*)[[:space:]]Free Throw.*(d|\))/\1,\2,\3,\4,\5/ ; s/(.*)d$/\10/ ; s/(.*)\)$/\11/' > free_throws.csv
```
```
from __future__ import division
import pandas as pd
import numpy as np
import scipy as sp
import scipy.stats
import toyplot as tp
```
## Data munging
Because only last name is included, we analyze "player-team" combinations to avoid duplicates.
This could mean that the same player has multiple rows if he changed teams.
```
df = pd.read_csv('free_throws.csv', names=["away", "home", "team", "player", "score"])
df["at_home"] = df["home"] == df["team"]
df.head()
```
## Overall free throw%
We note that at home the ft% is slightly higher, but there is not much difference
```
df.groupby("at_home").mean()
```
## Aggregating to player level
We use a pivot table to get statistics on every player.
```
sdf = pd.pivot_table(df, index=["player", "team"], columns="at_home", values=["score"],
aggfunc=[len, sum], fill_value=0).reset_index()
sdf.columns = ['player', 'team', 'atm_away', 'atm_home', 'score_away', 'score_home']
sdf['atm_total'] = sdf['atm_away'] + sdf['atm_home']
sdf['score_total'] = sdf['score_away'] + sdf['score_home']
sdf.sample(10)
```
## Individual tests
For each player, we assume each free throw is an independent draw from a Bernoulli distribution with probability $p_{ij}$ of succeeding where $i$ denotes the player and $j=\{a, h\}$ denoting away or home, respectively.
Our null hypotheses are that there is no difference between playing at home and away, versus the alternative that there is a difference.
While you could argue a one-sided test for home advantage is also appropriate, I am sticking with a two-sided test.
$$\begin{aligned}
H_{0, i}&: p_{i, a} = p_{i, h},\\
H_{1, i}&: p_{i, a} \neq p_{i, h}.
\end{aligned}$$
To get test statistics, we conduct a simple two-sample proportions test, where our test statistic is:
$$Z = \frac{\hat p_h - \hat p_a}{\sqrt{\hat p (1-\hat p) (\frac{1}{n_h} + \frac{1}{n_a})}}$$
where
- $n_h$ and $n_a$ are the number of attempts at home and away, respectively
- $X_h$ and $X_a$ are the number of free throws made at home and away
- $\hat p_h = X_h / n_h$ is the MLE for the free throw percentage at home
- likewise, $\hat p_a = X_a / n_a$ for away ft%
- $\hat p = \frac{X_h + X_a}{n_h + n_a}$ is the MLE for overall ft%, used for the pooled variance estimator
Then we know from Stats 101 that $Z \sim N(0, 1)$ under the null hypothesis that there is no difference in free throw percentages.
For a normal approximation to hold, we need $np > 5$ and $n(1-p) > 5$, since $p \approx 0.75$, let's be a little conservative and say we need at least 50 samples for a player to get a good normal approximation.
This leads to data on 936 players, and for each one we compute Z, and the corresponding p-value.
```
data = sdf.query('atm_total > 50').copy()
len(data)
data['p_home'] = data['score_home'] / data['atm_home']
data['p_away'] = data['score_away'] / data['atm_away']
data['p_ovr'] = (data['score_total']) / (data['atm_total'])
# two-sided
data['zval'] = (data['p_home'] - data['p_away']) / np.sqrt(data['p_ovr'] * (1-data['p_ovr']) * (1/data['atm_away'] + 1/data['atm_home']))
data['pval'] = 2*(1-sp.stats.norm.cdf(np.abs(data['zval'])))
# one-sided testing home advantage
# data['zval'] = (data['p_home'] - data['p_away']) / np.sqrt(data['p_ovr'] * (1-data['p_ovr']) * (1/data['atm_away'] + 1/data['atm_home']))
# data['pval'] = (1-sp.stats.norm.cdf(data['zval']))
data.sample(10)
canvas = tp.Canvas(800, 300)
ax1 = canvas.axes(grid=(1, 2, 0), label="Histogram p-values")
hist_p = ax1.bars(np.histogram(data["pval"], bins=50, normed=True), color="steelblue")
hisp_p_density = ax1.plot([0, 1], [1, 1], color="red")
ax2 = canvas.axes(grid=(1, 2, 1), label="Histogram z-values")
hist_z = ax2.bars(np.histogram(data["zval"], bins=50, normed=True), color="orange")
x = np.linspace(-3, 3, 200)
hisp_z_density = ax2.plot(x, sp.stats.norm.pdf(x), color="red")
```
# Global tests
We can test the global null hypothesis, that is, there is no difference in free throw % between playing at home and away for any player using both Fisher's Combination Test and the Bonferroni method.
Which one is preferred in this case? I would expect to see many small difference in effects rather than a few players showing huge effects, so Fisher's Combination Test probably has much better power.
## Fisher's combination test
We expect this test to have good power: if there is a difference between playing at home and away we would expect to see a lot of little effects.
```
T = -2 * np.sum(np.log(data["pval"]))
print 'p-value for Fisher Combination Test: {:.3e}'.format(1 - sp.stats.chi2.cdf(T, 2*len(data)))
```
## Bonferroni's method
The theory would suggest this test has a lot less power, it's unlikely to have a few players where the difference is relatively huge.
```
print '"p-value" Bonferroni: {:.3e}'.format(min(1, data["pval"].min() * len(data)))
```
## Conclusion
Indeed, we find a small p-value for Fisher's Combination Test, while Bonferroni's method does not reject the null hypothesis.
In fact, if we multiply the smallest p-value by the number of hypotheses, we get a number larger than 1, so we aren't even remotely close to any significance.
# Multiple tests
So there definitely seems some evidence that there is a difference in performance.
If you tell a sport's analyst that there is evidence that at least some players that perform differently away versus at home, their first question will be: "So who is?"
Let's see if we can properly answer that question.
## Naive method
Let's first test each null hypothesis ignoring the fact that we are dealing with many hypotheses. Please don't do this at home!
```
alpha = 0.05
data["reject_naive"] = 1*(data["pval"] < alpha)
print 'Number of rejections: {}'.format(data["reject_naive"].sum())
```
If we don't correct for multiple comparisons, there are actually 65 "significant" results (at $\alpha = 0.05$), which corresponds to about 7% of the players.
We expect around 46 rejections by chance, so it's a bit more than expected, but this is a bogus method so no matter what, we should discard the results.
## Bonferroni correction
Let's do it the proper way though, first using Bonferroni correction.
Since this method is basically the same as the Bonferroni global test, we expect no rejections:
```
from statsmodels.sandbox.stats.multicomp import multipletests
data["reject_bc"] = 1*(data["pval"] < alpha / len(data))
print 'Number of rejections: {}'.format(data["reject_bc"].sum())
```
Indeed, no rejections.
## Benjamini-Hochberg
Let's also try the BHq procedure, which has a bit more power than Bonferonni.
```
is_reject, corrected_pvals, _, _ = multipletests(data["pval"], alpha=0.1, method='fdr_bh')
data["reject_fdr"] = 1*is_reject
data["pval_fdr"] = corrected_pvals
print 'Number of rejections: {}'.format(data["reject_fdr"].sum())
```
Even though the BHq procedure has more power, we can't reject any of the individual hypothesis, hence we don't find sufficient evidence for any of the players that free throw performance is affected by location.
# Taking a step back
If we take a step back and take another look at our data, we quickly find that we shouldn't be surprised with our results.
In particular, our tests are clearly underpowered.
That is, the probability of rejecting the null hypothesis when there is a true effect is very small given the effect sizes that are reasonable.
While there are definitely sophisticated approaches to power analysis, we can use a [simple tool](http://statpages.org/proppowr.html) to get a rough estimate.
The free throw% is around 75% percent, and at that level it takes almost 2500 total attempts to detect a difference in ft% of 5% ($\alpha = 0.05$, power = $0.8$), and 5% is a pretty remarkable difference when only looking at home and away difference.
For most players, the observed difference is not even close to 5%, and we have only 11 players in our dataset with more than 2500 free throws.
To have any hope to detect effects for those few players that have plenty of data, the worst thing one can do is throw in a bunch of powerless tests.
It would have been much better to restrict our analysis to players where we have a lot of data.
Don't worry, I've already done that and again we cannot reject a single hypothesis.
So unfortunately it seems we won't be impressing our friends with cool results, more likely we will be the annoying person pointing out the fancy stats during a game don't really mean anything.
There is one cool take-away though: Fisher's combination test did reject the global null hypothesis even though each single test had almost no power, combined they did yield a significant result.
If we aggregate the data across all players first and then conduct a single test of proportions, it turns out we cannot reject that hypothesis.
```
len(data.query("atm_total > 2500"))
reduced_data = data.query("atm_total > 2500").copy()
is_reject2, corrected_pvals2, _, _ = multipletests(reduced_data["pval"], alpha=0.1, method='fdr_bh')
reduced_data["reject_fdr2"] = 1*is_reject2
reduced_data["pval_fdr2"] = corrected_pvals2
print 'Number of rejections: {}'.format(reduced_data["reject_fdr2"].sum())
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import string
import re
from collections import defaultdict
import glob
path = r'/content/drive/My Drive/Instagram_Data'
fileNames = glob.glob(path + "/*.csv")
lst = []
for fileName in fileNames:
print(fileName)
df = pd.read_csv(fileName, index_col=None, header=0)
lst.append(df)
df = pd.concat(lst, axis=0, ignore_index=True)
df.head()
print(len(df))
print(df["text"].iloc[1])
print(df["hashtags"].iloc[1])
print(df["comments"].iloc[1])
```
## Data Processing
### Cleaning Data
```
#https://stackoverflow.com/questions/33404752/removing-emojis-from-a-string-in-python
def deEmojify(text):
regrex_pattern = re.compile(pattern = "["
u"\U0001F600-\U0001F64F" # emoticons
u"\U0001F300-\U0001F5FF" # symbols & pictographs
u"\U0001F680-\U0001F6FF" # transport & map symbols
u"\U0001F1E0-\U0001F1FF" # flags (iOS)
"]+", flags = re.UNICODE)
return regrex_pattern.sub(r'',str(text))
# appending first 25 comments into one big comment
def appendComments(comment):
lst = comment.split("|")
final_comment = ""
for idx,entry in enumerate(lst):
if idx == 25:
break
entry.strip()
final_comment += entry + " "
return final_comment
def cleanComment(comment):
deEmojified_comment = deEmojify(comment)
return appendComments(deEmojified_comment)
df["comments"] = df["comments"].apply(lambda x:cleanComment(x))
#combining comments and actual post
df["text_with_comments"] = df["text"]+ " "+ df["comments"]
# removing punctuations,and converting upper to lower case letters
punctuations = string.punctuation
table = punctuations.maketrans(punctuations+string.ascii_uppercase,
" "*len(punctuations)+string.ascii_lowercase,)
def cleanPosts(sentence):
sentence.strip()
sen = sentence.translate(table)
return sen
def cleanHashTags(sentence):
sentence.strip()
sen = sentence.translate(table)
return "$start " + sen + " end$"
post_df = df["text_with_comments"].apply(lambda w:cleanPosts(str(w)))
hashtag_df = df["hashtags"].apply(lambda w: cleanHashTags(str(w)))
print(hashtag_df.iloc[1])
```
#### Hashtags and Likes
```
# here we only consider hashtags that are appearing atleast 10 times
def count_hashtag_likes(all_posts_hashtags,all_posts_likes):
hashtag_likesCount = defaultdict(int)
hashtagAppearanceCount = defaultdict(int)
hashtag_first10_likesCount = defaultdict(int)
for hashtags,count in zip(all_posts_hashtags,all_posts_likes):
hashtags = str(hashtags).split()
for hashtag in hashtags:
hashtagAppearanceCount[hashtag] += 1
if hashtagAppearanceCount[hashtag] >= 10:
if hashtag not in hashtag_likesCount:
hashtag_likesCount[hashtag] = hashtag_first10_likesCount[hashtag] + count
hashtag_likesCount[hashtag] += count
continue
hashtag_first10_likesCount[hashtag] += count
return hashtag_likesCount
hashtag_likesCount = count_hashtag_likes(hashtag_df.values.tolist(),df["likes"].values.tolist())
print(len(hashtag_likesCount))
# list of hashtags appearing atleast 10 times
frequent_hashtags = hashtag_likesCount.keys()
print(len(frequent_hashtags))
```
#### Splitting Data to Train ,Test and Validation Data
```
from sklearn.model_selection import train_test_split
X, X_val, Y, Y_val = train_test_split(post_df,hashtag_df, test_size=0.10,shuffle = True)
X_train, X_test, Y_train, Y_test = train_test_split(X,Y, test_size=0.10,shuffle = True)
print(X_train.shape,Y_train.shape)
print(X_val.shape,Y_val.shape)
print(X_test.shape,Y_test.shape)
type(X_test)
```
### Creating vector representations
```
class ConstructIndexes():
def __init__(self, data):
self.data = data
self.word2idx = {}
self.idx2word = {}
self.vocab = set()
self.create_index()
def create_index(self):
# we consider words appearing atleas 10 times
wordAppearanceCount = defaultdict(int)
for phrase in self.data:
for word in phrase.split():
wordAppearanceCount[word] += 1
if wordAppearanceCount[word] >= 10:
self.vocab.add(word)
# sort the vocab
self.vocab = sorted(self.vocab)
# add a padding token with index 0
self.word2idx['<pad>'] = 0
# word to index mapping
for index, word in enumerate(self.vocab):
self.word2idx[word] = index + 1 # +1 because of pad token
# index to word mapping
for word, index in self.word2idx.items():
self.idx2word[index] = word
# creating word2idx, idx2word on train data
# we use utils constructed on train to convert test and validation data
post_utils = ConstructIndexes(X_train.values.tolist())
hashtag_utils = ConstructIndexes(Y_train.values.tolist())
print(len(post_utils.word2idx))
print(len(hashtag_utils.word2idx))
# creating numeric vector corresponding to each post+comment
def toNumVec(constructIndexClass,df):
tensor = []
for phrase in df:
phrase_lst = []
for word in phrase.split():
if word in constructIndexClass.word2idx:
phrase_lst.append(constructIndexClass.word2idx[word])
tensor.append(phrase_lst)
return tensor
# Converting X_train,Y_train text sentences into numeric vectors
input_tensor_train = toNumVec(post_utils,X_train)
target_tensor_train = toNumVec(hashtag_utils,Y_train)
print(len(input_tensor_train),len(target_tensor_train))
# Converting X_val,Y_val text sentences into numeric vectors
input_tensor_val = toNumVec(post_utils,X_val)
target_tensor_val = toNumVec(hashtag_utils,Y_val)
print(len(input_tensor_val),len(target_tensor_val))
# Converting X_test,Y_test text sentences into numeric vectors
input_tensor_test = toNumVec(post_utils,X_test)
target_tensor_test = toNumVec(hashtag_utils,Y_test)
print(len(input_tensor_test),len(target_tensor_test))
print(input_tensor_train[1])
print(target_tensor_train[1])
```
#### Padding Numeric Vectors to same size
```
max_length_inp = max([len(t) for t in input_tensor_train])
max_length_tar = max([len(t) for t in target_tensor_train])
print(max_length_inp,max_length_tar)
def pad_sequences(x, max_len):
padded_x = np.zeros((max_len), dtype=np.int64)
if len(x) > max_len:
padded_x = x[:max_len]
else:
padded_x[:len(x)] = x
return padded_x
# padding zeros at the end of train tensors
input_tensor_train = np.array([pad_sequences(x, max_length_inp) for x in input_tensor_train if len(x) > 0])
target_tensor_train = np.array([pad_sequences(x, max_length_tar) for x in target_tensor_train if len(x) > 0])
# padding zeros at the end of validation tensors
input_tensor_val = np.array([pad_sequences(x, max_length_inp) for x in input_tensor_val if len(x) > 0])
target_tensor_val = np.array([pad_sequences(x, max_length_tar) for x in target_tensor_val if len(x) > 0])
# padding zeros at the end of test tensors
input_tensor_test= np.array([pad_sequences(x, max_length_inp) for x in input_tensor_test if len(x) > 0])
target_tensor_test = np.array([pad_sequences(x, max_length_tar) for x in target_tensor_test if len(x) > 0])
```
### Import Pytorch Libraries
```
import torch
import torch.functional as F
import torch.nn as nn
import torch.optim as optim
from torch.nn.utils.rnn import pack_padded_sequence, pad_packed_sequence
import pandas as pd
from sklearn.model_selection import train_test_split
import numpy as np
import unicodedata
import re
import time
print(torch.__version__)
from torch.utils.data import Dataset, DataLoader
```
#### DataLoader for batch inputs
```
# conver the data to tensors and pass to the Dataloader
# to create an batch iterator
class MyData(Dataset):
def __init__(self, X, y):
self.data = X
self.target = y
self.length = [ np.sum(1 - np.equal(x, 0)) for x in X]
def __getitem__(self, index):
x = self.data[index]
y = self.target[index]
x_len = self.length[index]
return x,y,x_len
def __len__(self):
return len(self.data)
```
## Parameters
Let's define the hyperparameters and other things we need for training our NMT model.
```
BUFFER_SIZE = len(input_tensor_train)
BATCH_SIZE = 64
N_BATCH = BUFFER_SIZE//BATCH_SIZE
embedding_dim = 64
units = 128
vocab_inp_size = len(post_utils.word2idx)
vocab_tar_size = len(hashtag_utils.word2idx)
train_dataset = MyData(input_tensor_train, target_tensor_train)
val_dataset = MyData(input_tensor_val, target_tensor_val)
dataset = DataLoader(train_dataset, batch_size = BATCH_SIZE,
drop_last=True,
shuffle=True)
```
### Encoder Model
```
class Encoder(nn.Module):
def __init__(self, vocab_size, embedding_dim, enc_units, batch_sz):
super(Encoder, self).__init__()
self.batch_sz = batch_sz
self.enc_units = enc_units
self.vocab_size = vocab_size
self.embedding_dim = embedding_dim
self.embedding = nn.Embedding(self.vocab_size, self.embedding_dim)
self.gru = nn.GRU(self.embedding_dim, self.enc_units)
def forward(self, x, lens, device):
# x: batch_size, max_length
# x: batch_size, max_length, embedding_dim
x = self.embedding(x)
# x transformed = max_len X batch_size X embedding_dim
# x = x.permute(1,0,2)
x = pack_padded_sequence(x, lens) # unpad
self.hidden = self.initialize_hidden_state(device)
# output: max_length, batch_size, enc_units
# self.hidden: 1, batch_size, enc_units
output, self.hidden = self.gru(x, self.hidden) # gru returns hidden state of all timesteps as well as hidden state at last timestep
# pad the sequence to the max length in the batch
output, _ = pad_packed_sequence(output)
return output, self.hidden
def initialize_hidden_state(self, device):
return torch.zeros((1, self.batch_sz, self.enc_units)).to(device)
### sort batch function to be able to use with pad_packed_sequence
def sort_batch(X, y, lengths):
lengths, indx = lengths.sort(dim=0, descending=True)
X = X[indx]
y = y[indx]
return X.transpose(0,1), y, lengths # transpose (batch x seq) to (seq x batch)
```
### Testing the Encoder
Before proceeding with training, we should always try to test out model behavior such as the size of outputs just to make that things are going as expected. In PyTorch this can be done easily since everything comes in eager execution by default.
```
### Testing Encoder part
# TODO: put whether GPU is available or not
# Device
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
encoder = Encoder(vocab_inp_size, embedding_dim, units, BATCH_SIZE)
encoder.to(device)
# obtain one sample from the data iterator
it = iter(dataset)
x, y, x_len = next(it)
# sort the batch first to be able to use with pac_pack_sequence
xs, ys, lens = sort_batch(x, y, x_len)
enc_output, enc_hidden = encoder(xs.to(device), lens, device)
print(enc_output.size()) # max_length, batch_size, enc_units
```
### Decoder Model
```
class Decoder(nn.Module):
def __init__(self, vocab_size, embedding_dim, dec_units, enc_units, batch_sz):
super(Decoder, self).__init__()
self.batch_sz = batch_sz
self.dec_units = dec_units
self.enc_units = enc_units
self.vocab_size = vocab_size
self.embedding_dim = embedding_dim
self.embedding = nn.Embedding(self.vocab_size, self.embedding_dim)
self.gru = nn.GRU(self.embedding_dim + self.enc_units,
self.dec_units,
batch_first=True)
self.fc = nn.Linear(self.enc_units, self.vocab_size)
# used for attention
self.W1 = nn.Linear(self.enc_units, self.dec_units)
self.W2 = nn.Linear(self.enc_units, self.dec_units)
self.V = nn.Linear(self.enc_units, 1)
def forward(self, x, hidden, enc_output):
# enc_output original: (max_length, batch_size, enc_units)
# enc_output converted == (batch_size, max_length, hidden_size)
enc_output = enc_output.permute(1,0,2)
# hidden shape == (batch_size, hidden size)
# hidden_with_time_axis shape == (batch_size, 1, hidden size)
# we are doing this to perform addition to calculate the score
# hidden shape == (batch_size, hidden size)
# hidden_with_time_axis shape == (batch_size, 1, hidden size)
hidden_with_time_axis = hidden.permute(1, 0, 2)
# score: (batch_size, max_length, hidden_size) # Bahdanaus's
# we get 1 at the last axis because we are applying tanh(FC(EO) + FC(H)) to self.V
# It doesn't matter which FC we pick for each of the inputs
score = torch.tanh(self.W1(enc_output) + self.W2(hidden_with_time_axis))
#score = torch.tanh(self.W2(hidden_with_time_axis) + self.W1(enc_output))
# attention_weights shape == (batch_size, max_length, 1)
# we get 1 at the last axis because we are applying score to self.V
attention_weights = torch.softmax(self.V(score), dim=1)
# context_vector shape after sum == (batch_size, hidden_size)
context_vector = attention_weights * enc_output
context_vector = torch.sum(context_vector, dim=1)
# x shape after passing through embedding == (batch_size, 1, embedding_dim)
# takes case of the right portion of the model above (illustrated in red)
x = self.embedding(x)
# x shape after concatenation == (batch_size, 1, embedding_dim + hidden_size)
#x = tf.concat([tf.expand_dims(context_vector, 1), x], axis=-1)
# ? Looks like attention vector in diagram of source
x = torch.cat((context_vector.unsqueeze(1), x), -1)
# passing the concatenated vector to the GRU
# output: (batch_size, 1, hidden_size)
output, state = self.gru(x)
# output shape == (batch_size * 1, hidden_size)
output = output.view(-1, output.size(2))
# output shape == (batch_size * 1, vocab)
x = self.fc(output)
return x, state, attention_weights
def initialize_hidden_state(self):
return torch.zeros((1, self.batch_sz, self.dec_units))
```
#### Testing the Decoder
```
# Device
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
encoder = Encoder(vocab_inp_size, embedding_dim, units, BATCH_SIZE)
encoder.to(device)
# obtain one sample from the data iterator
it = iter(dataset)
x, y, x_len = next(it)
print("Input: ", x.shape)
print("Output: ", y.shape)
# sort the batch first to be able to use with pac_pack_sequence
xs, ys, lens = sort_batch(x, y, x_len)
enc_output, enc_hidden = encoder(xs.to(device), lens, device)
print("Encoder Output: ", enc_output.shape) # batch_size X max_length X enc_units
print("Encoder Hidden: ", enc_hidden.shape) # batch_size X enc_units (corresponds to the last state)
decoder = Decoder(vocab_tar_size, embedding_dim, units, units, BATCH_SIZE)
decoder = decoder.to(device)
#print(enc_hidden.squeeze(0).shape)
dec_hidden = enc_hidden#.squeeze(0)
dec_input = torch.tensor([[hashtag_utils.word2idx['$start']]] * BATCH_SIZE)
print("Decoder Input: ", dec_input.shape)
print("--------")
for t in range(1, y.size(1)):
# enc_hidden: 1, batch_size, enc_units
# output: max_length, batch_size, enc_units
predictions, dec_hidden, _ = decoder(dec_input.to(device),
dec_hidden.to(device),
enc_output.to(device))
print("Prediction: ", predictions.shape)
print("Decoder Hidden: ", dec_hidden.shape)
#loss += loss_function(y[:, t].to(device), predictions.to(device))
dec_input = y[:, t].unsqueeze(1)
print(dec_input.shape)
break
```
## Training Model
```
def train(epochs,encoder,decoder,device,ytrain_utils,
loss_function,batch_size,optimizer,dataLoader):
for epoch in range(epochs):
start = time.time()
total_loss = 0
encoder.train()
decoder.train()
for (batch, (inp, targ, inp_len)) in enumerate(dataLoader):
loss = 0
xs, ys, lens = sort_batch(inp, targ, inp_len)
enc_output, enc_hidden = encoder(xs.to(device), lens, device)
dec_hidden = enc_hidden
# use teacher forcing - feeding the target as the next input (via dec_input)
dec_input = torch.tensor([[ytrain_utils.word2idx['$start']]] * batch_size)
# run code below for every timestep in the ys batch
for t in range(1, ys.size(1)):
predictions, dec_hidden, _ = decoder(dec_input.to(device),
dec_hidden.to(device),
enc_output.to(device))
loss += loss_function(ys[:, t].to(device), predictions.to(device))
#loss += loss_
dec_input = ys[:, t].unsqueeze(1)
batch_loss = (loss / int(ys.size(1)))
total_loss += batch_loss
optimizer.zero_grad()
loss.backward()
optimizer.step()
'''if batch % 100 == 0:
print('Epoch {} Batch {} Loss {:.4f}'.format(epoch + 1,
batch,
batch_loss.detach().item()))'''
### TODO: Save checkpoint for model
print('Epoch {} Loss {:.4f}'.format(epoch + 1,
total_loss / N_BATCH))
print('Time taken for 1 epoch {} sec\n'.format(time.time() - start))
criterion = nn.CrossEntropyLoss()
def loss_function(real, pred):
""" Only consider non-zero inputs in the loss; mask needed """
#mask = 1 - np.equal(real, 0) # assign 0 to all above 0 and 1 to all 0s
#print(mask)
mask = real.ge(1).type(torch.cuda.FloatTensor)
loss_ = criterion(pred, real) * mask
return torch.mean(loss_)
BATCH_SIZE = 64
## TODO: Combine the encoder and decoder into one class
encoder = Encoder(vocab_inp_size, embedding_dim, units, BATCH_SIZE)
decoder = Decoder(vocab_tar_size, embedding_dim, units, units, BATCH_SIZE)
encoder.to(device)
decoder.to(device)
optimizer = optim.Adam(list(encoder.parameters()) + list(decoder.parameters()),
lr=0.001)
epochs = 50
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
train(epochs,encoder,decoder,device,hashtag_utils,
loss_function,BATCH_SIZE,optimizer,dataset)
```
### Predict hashtags
```
encoder_pred = Encoder(vocab_inp_size, embedding_dim, units, 1)
decoder_pred = Decoder(vocab_tar_size, embedding_dim, units, units,1)
encoder_pred.to(device)
decoder_pred.to(device)
```
#### Loading weights into another model
```
encoder_pred.load_state_dict(encoder.state_dict())
decoder_pred.load_state_dict(decoder.state_dict())
softmax = nn.Softmax(dim=1)
def predict(X_test):
results = []
encoder_pred.eval()
decoder_pred.eval()
for xs in enumerate(X_test):
xs = xs[1]
length = np.sum(1 - np.equal(xs, 0))
xs = torch.tensor(xs).view(-1,1)
length = torch.tensor(length).view(-1,)
enc_output, enc_hidden = encoder_pred(xs.to(device), length, device)
dec_hidden = enc_hidden
dec_input = torch.tensor([[hashtag_utils.word2idx['$start']]])
curr_hashtags = []
# run code below till we generate "end$" tag or 10 hashtags
for t in range(1,10):
predictions, dec_hidden, _ = decoder_pred(dec_input.to(device),
dec_hidden.to(device),
enc_output.to(device))
#print(predictions.size())
top_val,top_idx = softmax(predictions).topk(1,dim = 1)
#print(top_idx.item())
if top_idx == hashtag_utils.word2idx["end$"]:
break
pred_hashtag = hashtag_utils.idx2word[top_idx.item()]
curr_hashtags.append(pred_hashtag)
dec_input = torch.tensor([top_idx]).unsqueeze(1)
results.append(curr_hashtags)
return results
#predict hashtags on test data
predicted_hashtags = predict(input_tensor_test)
for i in range(10):
print(Y_test.iloc[i],predicted_hashtags[i])
```
| github_jupyter |
Temperature animation for Run 08 (sketches for python scripts)
```
#KRM
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
from math import *
import scipy.io
import scipy as spy
#%matplotlib inline
from netCDF4 import Dataset
import pylab as pl
import matplotlib.animation as animation
#'''
#NAME
# Custom Colormaps for Matplotlib
#PURPOSE
# This program shows how to implement make_cmap which is a function that
# generates a colorbar. If you want to look at different color schemes,
# check out https://kuler.adobe.com/create.
#PROGRAMMER(S)
# Chris Slocum
#REVISION HISTORY
# 20130411 -- Initial version created
# 20140313 -- Small changes made and code posted online
# 20140320 -- Added the ability to set the position of each color
#'''
def make_cmap(colors, position=None, bit=False):
#'''
#make_cmap takes a list of tuples which contain RGB values. The RGB
#values may either be in 8-bit [0 to 255] (in which bit must be set to
#rue when called) or arithmetic [0 to 1] (default). make_cmap returns
#a cmap with equally spaced colors.
#Arrange your tuples so that the first color is the lowest value for the
#colorbar and the last is the highest.
#position contains values from 0 to 1 to dictate the location of each color.
#'''
import matplotlib as mpl
import numpy as np
bit_rgb = np.linspace(0,1,256)
if position == None:
position = np.linspace(0,1,len(colors))
else:
if len(position) != len(colors):
sys.exit("position length must be the same as colors")
elif position[0] != 0 or position[-1] != 1:
sys.exit("position must start with 0 and end with 1")
if bit:
for i in range(len(colors)):
colors[i] = (bit_rgb[colors[i][0]],
bit_rgb[colors[i][1]],
bit_rgb[colors[i][2]])
cdict = {'red':[], 'green':[], 'blue':[]}
for pos, color in zip(position, colors):
cdict['red'].append((pos, color[0], color[0]))
cdict['green'].append((pos, color[1], color[1]))
cdict['blue'].append((pos, color[2], color[2]))
cmap = mpl.colors.LinearSegmentedColormap('my_colormap',cdict,256)
return cmap
# Get field from MITgcm netCDF output
#
''' :statefile : string with /path/to/state.0000000000.t001.nc
:fieldname : string with the variable name as written on the netCDF file ('Temp', 'S','Eta', etc.)'''
def getField(statefile, fieldname):
StateOut = Dataset(statefile)
Fld = StateOut.variables[fieldname][:]
shFld = np.shape(Fld)
if len(shFld) == 2:
Fld2 = np.reshape(Fld,(shFld[0],shFld[1])) # reshape to pcolor order
return Fld2
elif len(shFld) == 3:
Fld2 = np.zeros((shFld[0],shFld[1],shFld[2]))
Fld2 = np.reshape(Fld,(shFld[0],shFld[1],shFld[2])) # reshape to pcolor order
return Fld2
elif len(shFld) == 4:
Fld2 = np.zeros((shFld[0],shFld[1],shFld[2],shFld[3]))
Fld2 = np.reshape(Fld,(shFld[0],shFld[1],shFld[2],shFld[3])) # reshape to pcolor order
return Fld2
else:
print (' Check size of field ')
filenameb='/ocean/kramosmu/MITgcm/CanyonUpwelling/180x180x35_BodyForcing_6Tr_LinProfiles/run08/output_0001/state.0000000000.t001.nc'
StateOutb = Dataset(filenameb)
#for dimobj in StateOut.variables.values():
# print dimobj
filename2b='/ocean/kramosmu/MITgcm/CanyonUpwelling/180x180x35_BodyForcing_6Tr_LinProfiles/run08/output_0001/grid.t001.nc'
GridOutb = Dataset(filename2b)
for dimobj in GridOutb.variables.values():
print dimobj
filename3b='/ocean/kramosmu/MITgcm/CanyonUpwelling/180x180x35_BodyForcing_6Tr_LinProfiles/run08/output_0001/ptracers.0000000000.t001.nc'
PtracersOutb = Dataset(filename3b)
#for dimobj in PtracersOut.variables.values():
# print dimobj
# General input
nx = 180
ny = 180
nz = 35
nta = 21 # t dimension size run 04 and 05 (output every 2 hr for 4.5 days)
ntc = 21 # t dimension size run 06 (output every half-day for 4.5 days)
z = StateOutb.variables['Z']
#print(z[:])
Time = StateOutb.variables['T']
print(Time[:])
xc = getField(filename2b, 'XC') # x coords tracer cells
yc = getField(filename2b, 'YC') # y coords tracer cells
zlev = 31 # level 14 corresponds to 162.5 m , near shelf break
timesc = [1,3,5,10,15,20] # These correspond to 1,2,4,6,8,10 days
ugridb = getField(filenameb,'U')
vgridb = getField(filenameb,'V')
print(np.shape(ugridb))
print(np.shape(vgridb))
tempb = getField(filenameb, 'Temp')
temp0b = np.ma.masked_values(tempb, 0)
MASKb = np.ma.getmask(temp0b)
plt.rcParams.update({'font.size':13})
NumLev = 30 # number of levels for contour
#### PLOT ####
def animateTemp(tt):
"""Generate frames for Temperature animation Run10 in 180x180x35_BodyForcing_6Tr_LinProfiles
tt corresponds to the time output
"""
kk=1
plt.cla
### Temperature
ax1 = fig.add_subplot(2, 2, 1)
ax1.set_aspect(0.75)
plt.contourf(xc,yc,temp0b[tt,4,:,:],NumLev,cmap='gist_heat')
plt.ticklabel_format(style='sci', axis='x', scilimits=(0,0))
plt.ticklabel_format(style='sci', axis='y', scilimits=(0,0))
plt.xlabel('m')
plt.ylabel('m')
plt.title(" depth=%1.1f m,%1.1f days " % (z[4],tt/2.))
ax2 = fig.add_subplot(2, 2, 2)
ax2.set_aspect(0.75)
plt.contourf(xc,yc,temp0b[tt,14,:,:],NumLev,cmap='gist_heat')
plt.ticklabel_format(style='sci', axis='x', scilimits=(0,0))
plt.ticklabel_format(style='sci', axis='y', scilimits=(0,0))
plt.xlabel('m')
plt.ylabel('m')
plt.title(" depth=%1.1f m,%1.1f days " % (z[14],tt/2.))
ax3 = fig.add_subplot(2, 2, 3)
ax3.set_aspect(0.75)
plt.contourf(xc,yc,temp0b[tt,24,:,:],NumLev,cmap='gist_heat')
plt.ticklabel_format(style='sci', axis='x', scilimits=(0,0))
plt.ticklabel_format(style='sci', axis='y', scilimits=(0,0))
plt.xlabel('m')
plt.ylabel('m')
plt.title(" depth=%1.1f m,%1.1f days " % (z[24],tt/2.))
ax4 = fig.add_subplot(2, 2, 4)
ax4.set_aspect(0.75)
plt.contourf(xc,yc,temp0b[tt,31,:,:],NumLev,cmap='gist_heat')
plt.ticklabel_format(style='sci', axis='x', scilimits=(0,0))
plt.ticklabel_format(style='sci', axis='y', scilimits=(0,0))
plt.xlabel('m')
plt.ylabel('m')
plt.title(" depth=%1.1f m,%1.1f days " % (z[31],tt/2.))
ax1.set_axis_bgcolor((205/255.0, 201/255.0, 201/255.0))
ax2.set_axis_bgcolor((205/255.0, 201/255.0, 201/255.0))
ax3.set_axis_bgcolor((205/255.0, 201/255.0, 201/255.0))
ax4.set_axis_bgcolor((205/255.0, 201/255.0, 201/255.0))
plt.rcParams.update({'font.size':14})
colorsTemp = [(245.0/255.0,245/255.0,245./255.0), (255/255.0,20/255.0,0)] #(khaki 1246/255.0,143./255.0 ,orangered2)
posTemp = [0, 1]
fig= plt.figure(figsize=(12,9))
#The animation function
anim = animation.FuncAnimation(fig, animateTemp, frames=21)
#cb = plt.colorbar(pad = 0.5)
#cb.set_label(r'$^{\circ}$C',position=(1, 0),rotation=0)
plt.show()
#A line that makes it all work
#mywriter = animation.FFMpegWriter()
#Save in current folder
#anim.save('Run08Temp.mp4',writer=mywriter)
```
| github_jupyter |
```
import os
import shutil
import torch
import torch.nn as nn
import torch.optim
from torch.utils.data import DataLoader
from torch.optim.lr_scheduler import MultiStepLR
import torch.backends.cudnn as cudnn
import numpy as np
from models.grid_proto_fewshot import FewShotSeg
from dataloaders.dev_customized_med import med_fewshot
from dataloaders.GenericSuperDatasetv2 import SuperpixelDataset
from dataloaders.dataset_utils import DATASET_INFO
import dataloaders.augutils as myaug
from util.utils import set_seed, t2n, to01, compose_wt_simple
from util.metric import Metric
from config_ssl_upload import ex
import tqdm
data_name = "CHAOST2_Superpix"
baseset_name = 'CHAOST2'
tr_transforms = myaug.transform_with_label({'aug': myaug.augs['sabs_aug']})
test_labels = DATASET_INFO[baseset_name]['LABEL_GROUP']['pa_all'] - DATASET_INFO[baseset_name]['LABEL_GROUP'][0]
tr_parent = SuperpixelDataset( # base dataset
which_dataset = baseset_name,
base_dir="./data/CHAOST2/chaos_MR_T2_normalized/",
idx_split = 0,
mode='train',
min_fg='1', # dummy entry for superpixel dataset
transforms=tr_transforms,
nsup = 1,
scan_per_load = -1,
exclude_list = [2, 3],
superpix_scale = 'MIDDLE',
fix_length = 1000
)
trainloader = DataLoader(
tr_parent,
batch_size=1,
shuffle=True,
num_workers=0,
pin_memory=True,
drop_last=True
)
for _, sample_batched in enumerate(trainloader):
pass
type(sample_batched)
type(sample_batched['class_ids'])
len(sample_batched['class_ids'])
sample_batched['class_ids'][0]
type(sample_batched['support_images'])
len(sample_batched['support_images'])
type(sample_batched['support_images'][0])
len(sample_batched['support_images'][0])
type(sample_batched['support_images'][0][0])
np.min(sample_batched['support_images'][0][0].numpy()), np.max(sample_batched['support_images'][0][0].numpy())
type(sample_batched['support_mask'])
len(sample_batched['support_mask'])
type(sample_batched['support_mask'][0])
len(sample_batched['support_mask'][0])
type(sample_batched['support_mask'][0][0])
sample_batched['support_mask'][0][0]['fg_mask'].shape
np.min(sample_batched['support_mask'][0][0]['fg_mask'].numpy()), np.max(sample_batched['support_mask'][0][0]['fg_mask'].numpy())
np.min(sample_batched['support_mask'][0][0]['bg_mask'].numpy()), np.max(sample_batched['support_mask'][0][0]['bg_mask'].numpy())
type(sample_batched['query_images'])
len(sample_batched['query_images'])
type(sample_batched['query_images'][0])
sample_batched['query_images'][0].shape
np.min(sample_batched['query_images'][0][0].numpy()), np.max(sample_batched['query_images'][0][0].numpy())
type(sample_batched['query_labels'])
len(sample_batched['query_labels'])
type(sample_batched['query_labels'][0])
sample_batched['query_labels'][0].shape
np.min(sample_batched['query_labels'][0][0].numpy()), np.max(sample_batched['query_labels'][0][0].numpy())
```
| github_jupyter |
# Noise Detection Algorithm
The data presented are measurements of a gaussian beam for varying beam-frequencies and distances. Due to technical difficulties, our measuring device would sometimes crash and provide us with completely noisy data, or data that was only half complete. Our intent was to automize the measuring process, by making the lab-computer automatically evaluate the data. In the case of faulty data, it was supposed to restart the measurement.
# Importing packages
```
# general
import numpy as np
import matplotlib.pyplot as plt
import re # to extract number in name
from pathlib import Path # to extract data from registry
import pandas as pd # data frames
from mpl_toolkits.axes_grid1 import make_axes_locatable #adjust colorbars to axis
from tkinter import Tcl # sort file names
# machine learning
# spli data
from sklearn.model_selection import train_test_split
# process data
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
# classification algorithm
from sklearn.neighbors import KNeighborsClassifier
# pipeline
from sklearn.pipeline import Pipeline
# model evaluation
from sklearn.model_selection import StratifiedKFold
from sklearn.model_selection import cross_val_score
from sklearn.metrics import confusion_matrix
```
# Importing and processing Data
## Importing samples
```
p=Path('.')
# list(path.glob'./*.dat') finds all data ".dat" data in entered directory
paths=list([x for x in p.iterdir() if x.is_dir() and x.name=='Measurements'][0].glob('./*.dat')) #use ** to also include subregistries
#remember to change name if directory name is changed
# generate lists
path_names=list(map(lambda x: x.name, paths)) # get path names
# extract frequencies
pattern=re.compile(r"(\d+)")
freq_list=np.array([pattern.match(x).groups()[0] for x in path_names if x.startswith("Noise")==False]).astype(int)
freq_list=np.sort(freq_list)
# sort paths
sorted_path_names=np.array(Tcl().call('lsort', '-dict', path_names)) # sorted path names
# hack for sorting lists from short too long (this should be the default)
duplicate_freqs=[x for x in np.unique(freq_list) if list(freq_list).count(x)!=1]
for duplis in duplicate_freqs:
duplicate_indices=np.where(freq_list==duplis)
for index in duplicate_indices[0][::-1][:-1]:
backup=np.copy(sorted_path_names[index-1])
sorted_path_names[index-1]=sorted_path_names[index]
sorted_path_names[index]=backup
# Sort Noise-data
for index in np.where(np.array(list(map(lambda x:x.startswith("Noise"),sorted_path_names)))==True)[0][::-1][:-1]:
backup=np.copy(sorted_path_names[index-1])
sorted_path_names[index-1]=sorted_path_names[index]
sorted_path_names[index]=backup
# sort
_,paths=zip(*sorted(zip([list(sorted_path_names).index(path_name) for path_name in path_names], paths)))
path_names=np.copy(sorted_path_names)
del sorted_path_names
#remember to change name if directory name is changed
data_dict={path.name: np.genfromtxt(path,skip_header=1)[:,2] for path in paths}
# add images
size=np.int(np.sqrt(len(data_dict[path_names[0]])))
data_dict["images"]=[data_dict[name].reshape((size,size)) for name in path_names]
len(data_dict["images"])
```
## Assigning Features
```
# targets
# key targets 0=noise, 1=okay data, 2= good data
three_targets={
'70GHz.dat':2,'70GHz-1.dat':2,
'70GHz-2.dat':0,'75GHz.dat':2,
'75GHz-1.dat':2,'80GHz.dat':2,
'80GHz-1.dat':2,'85GHz.dat':2,
'85GHz-1.dat':2,'85GHz-2.dat':0,
'85GHz-3.dat':2,'85GHz-4.dat':0,
'85GHz-5.dat':0,'85GHz-6.dat':0,
'85GHz-7.dat':2,'85GHz-8.dat':1,
'85GHz-9.dat':1,'85GHz-10.dat':2,
'90GHz.dat':2,'90GHz-1.dat':2,
'90GHz-2.dat':2,'90GHz-3.dat':1,
'95GHz.dat':2,'95GHz-1.dat':2,
'95GHz-2.dat':2,'95GHz-3.dat':0,
'95GHz-4.dat':2,'95GHz-5.dat':2,
'95GHz-6.dat':2,'95GHz-7.dat':1,
'95GHz-8.dat':2,'95GHz-9.dat':0,
'95GHz-10.dat':0,'95GHz-11.dat':0,
'100GHz.dat':2,'100GHz-1.dat':2,
'100GHz-2.dat':0,'105GHz.dat':2,
'105GHz-1.dat':2,'105GHz-2.dat':0,
'110GHz.dat':0,'110GHz-1.dat':0,
'110GHz-2.dat':0,'Noise.dat':0,
'Noise-1.dat':0
}
```
## Forming Dictionary out of Data
```
# Create dictionary out of data
y=np.zeros(len(three_targets)).astype(int)
X=np.zeros((len(three_targets),len(data_dict[path_names[0]])))
size=np.int(np.sqrt(len(data_dict[path_names[0]])))
names=len(three_targets)*[""]
# counter
count=0
# format data correctly
for name in path_names:
if name in three_targets:
names[count]=name
y[count]=three_targets[name]
X[count,:]=data_dict[name]
count+=1
else:
print("{} --- not yet labeled".format(name))
beam_data={"data_names":names,"data":X,"target":y,
"target_names":["full noise","half_noise","no noise"],"images":[x.reshape((size,size)) for x in X]}
```
## Plotting all data
```
tes=np.arange(len(path_names))
print (len(tes[15:30]))
del tes
#all data
rows=int(len(path_names) / 3) + (len(path_names) % 3 > 0) # how many rows
# plot
fig = plt.figure(figsize=(18, 5*rows))
print("Yet to be labeled:")
for i,name in enumerate(path_names[30:]):
boolean=False # check if something has to be labeled:
ax=plt.subplot(rows,3,i+1)
ax.set_axis_off() # hide axis
im=ax.imshow(data_dict["images"][list(data_dict.keys()).index(name)],cmap='jet', interpolation='nearest')
if name in beam_data["data_names"]:
ax.set_title("name: {} \n labeled: {}".format(name[:-4],beam_data["target_names"][beam_data["target"][list(beam_data["data_names"]).index(name)]]),fontsize=14)
else:
boolean=True
ax.set_title("name: {}\n not labeled yet".format(name),color="red")
print("'{}'".format(name))
#adjust colorbar to plot
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.1)
cbar=plt.colorbar(im, cax=cax)
cbar.ax.set_ylabel('intensity',fontsize=15)
if boolean==False: print("nothing")
plt.tight_layout()
# for export
#plt.savefig("Images/ml-databatch-3.png")
```
# Feature Engineering
## rescaling
```
def rescale_local_max(X):
return [X[i]/maxi for i,maxi in enumerate(np.amax(X,axis=1))]
```
## rotating for symmetrical prediction (generalization)
In this dataset, the transition to half_noise always occurs from no_noise in the top region to full_noise in the bottom region. To generalize the algorithm for other problems, the half data will now be rotated (Note that this may produce too optimistic evaluations of model predictions for this class).
For even further generalization, all data in 2D representation can be repositioned to have their peak in the center. This way, dispositions of the curve location would not need to be considered within the model, which could improve not only generalization performace, but also general performance in general.
```
# rescale
X_total=np.array(rescale_local_max(beam_data["data"]))
y_total=np.array(beam_data["target"])
# rescale
X_total=rescale_local_max(beam_data["data"])
# take half data
X_half_noise=np.array(X_total)[np.where(beam_data["target"]==1)]
# X_data in total
X_added=np.zeros((X_half_noise.shape[0]*3,X_half_noise.shape[1]))
for i in range(X_half_noise.shape[0]):
X_added[3*i,:]=np.rot90(X_half_noise[i].reshape((size,size))).flatten()
X_added[3*i+1,:]=np.rot90(X_added[3*i,:].reshape((size,size))).flatten()
X_added[3*i+2,:]=np.rot90(X_added[3*i+1,:].reshape((size,size))).flatten()
X_total=np.vstack((X_total,X_added))
y_total=np.concatenate((beam_data["target"],[1]*X_half_noise.shape[0]*3))
# provide example of rotated data
fig=plt.figure(figsize=(10,2.5))
ax=plt.subplot(1,4,1)
ax.imshow(X_half_noise[3,:].reshape((size,size)),cmap="jet")
ax.set_title("original")
ax.set_axis_off() # hide axis
for i in range(3):
ax=plt.subplot(1,4,i+2)
ax.imshow(X_added[9+i,:].reshape((size,size)),cmap="jet")
ax.set_title("rotated by {}°".format(90*(i+1)))
ax.set_axis_off() # hide axis
plt.tight_layout()
plt.savefig("Images/rotated-maps.png")
# split sets
X_train, X_test, y_train, y_test = train_test_split(X_total,y_total, stratify=y_total, random_state=0)
```
# Machine Learning (applying PCA + knn)
```
np.array(X_train).shape
```
In this application we have few samples (40) and many features (441). To reduce the number of features, we apply Prinicipal Component Analysis (PCA), to reduce the dimension in feature space (Here from 221 to 2) and therefore improve the performance of our alogorithm. We use the k-neighbours classifier (knn), as it performs particulary well on small sample sizes.
## PCA and it's data rescaling results
```
# scale to mean=0, std=1
scaler = StandardScaler()
# fit scaling
scaler.fit(X_train)
# apply scaling
scaled_X_train=scaler.transform(X_train)
# n_components=amount of principal components
pca = PCA(n_components=2) # n_components=0.95 alternatively
# fit PCA model to beast cancer data
pca.fit(scaled_X_train)
# transform
pca_X_train = pca.transform(scaled_X_train)
# Create data
# PCA on train data
g0 = pca_X_train[np.where(y_train==0)]
g1 = pca_X_train[np.where(y_train==1)]
g2 = pca_X_train[np.where(y_train==2)]
train_data = (g0, g1, g2)
colors = ("red","orange","blue")
groups = ("full_noise", "half_noise","no_noise")
train_marker=("o")
# PCA on test data
# transform
scaled_X_test=scaler.transform(X_test)
pca_X_test = pca.transform(scaled_X_test)
# # # # #
h0 = pca_X_test[np.where(y_test==0)]
h1 = pca_X_test[np.where(y_test==1)]
h2 = pca_X_test[np.where(y_test==2)]
test_data = (h0, h1, h2)
test_marker=("^")
# Create plot
fig = plt.figure(figsize=(7,7))
ax = fig.add_subplot(1, 1, 1, )
# plot train-transform
for data, color, group in zip(train_data, colors, groups):
x =data[:,0]
y =data[:,1]
ax.scatter(x, y, alpha=0.7, c=color, edgecolors='black', s=50, label="train:"+group, marker=train_marker)
# plot test-transform
for data, color, group in zip(test_data, colors, groups):
x =data[:,0]
y =data[:,1]
ax.scatter(x, y, alpha=0.7, c=color, edgecolors='black', s=100, label="test:"+group, marker=test_marker)
# labels
#plt.title('PCA-transformed plot')
plt.xlabel("prinicipal component Nr.1",fontsize=15)
plt.ylabel("principal component Nr.2",fontsize=15)
plt.legend(loc="best")
plt.tight_layout()
plt.grid(True)
plt.savefig("Images/PCA_map.png", bbox_inches='tight')
plt.show()
# plot components
fig, axes = plt.subplots(1, 2,figsize=(15,6))
for i, (component, ax) in enumerate(zip(pca.components_, axes.ravel())):
im=ax.imshow(component.reshape((size,size)),cmap='jet',interpolation="nearest")
ax.set_title("component Nr.{}".format(i+1),fontsize=30)
ax.set_xlabel("pixel in x-direction",fontsize=25)
ax.set_ylabel("pixel in y-direction",fontsize=25)
fig.colorbar(im, ax=ax, fraction=0.046, pad=0.04)
#fig.suptitle('PCA component weights', fontsize=20)
plt.tight_layout()
plt.savefig("Images/PCA-componenets.png", bbox_inches='tight')
```
## Creating and fitting pipeline
because of small sample-size, no parameter optimization will be conducted
(as splitting the data in another validation set would reduce the already small test set)
```
# create pipeline
pipe = Pipeline([("scaler", StandardScaler()), ("component_analyzer", PCA(n_components=2)),
("classifier", KNeighborsClassifier(n_neighbors=1))])# fitting
pipe.fit(X_train,y_train)
```
## Algorithm performance
### General evaluation via stratified KFold cross validation
stratified makes sure that all classes are represented in each training set. Shuffle makes sure, that the data is shuffled before it is split and is only necessary when the data is sorted.
```
kfold = StratifiedKFold(n_splits=5, shuffle=True, random_state=0)
print("Cross-validation scores:\n{}".format(
cross_val_score(pipe, X_total, y_total, cv=kfold)))
```
### confusion matrix shows, how test samples were classified
```
# cross calidation confusion matrix
splits=list(kfold.split(X_total,y_total))
for i,split in enumerate(splits):
pipe.fit(X_total[split[0]],y_total[split[0]])
print("Split Nr.{}:\n{}\n".format(i,confusion_matrix(pipe.predict(X_total[split[1]]),y_total[split[1]])))
# show incorrectly classified beam map
i=2 # index with non-diagonal confusion matrix
pipe.fit(X_total[splits[i][0]],y_total[splits[i][0]])
false_pred=pipe.predict(X_total[splits[i][1]])[pipe.predict(X_total[splits[i][1]])!=y_total[splits[i][1]]][0]
beam_data["target_names"][false_pred]
# index
index=splits[i][1][np.where(pipe.predict(X_total[splits[i][1]])!=y_total[splits[i][1]])]
# plot
plt.figure(figsize=(5,5.5))
ax=plt.subplot(1,1,1)
ax.set_axis_off() # hide axis
ax.set_title("target= {}\npredicted= {}".format(beam_data["target_names"][y_total[index][0]],
beam_data["target_names"][false_pred]),
fontsize=25)
ax.imshow(X_total[index].reshape((size,size)),
cmap="jet")
#adjust colorbar to plot
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.1)
cbar=plt.colorbar(im, cax=cax)
cbar.ax.set_ylabel('intensity',fontsize=15)
plt.tight_layout()
plt.savefig("Images/false-prediction.png", bbox_inches='tight')
```
## Evaluation of one singular split
### predict test set
```
predictions=pipe.predict(X_test)
print(predictions)
print("confusion matrix:\n{}".format(confusion_matrix(y_test,predictions)))
```
### falsely classified in test data
```
# collect indices
index=[]
predictions=pipe.predict(X_test)
for i in range(len(y_test)):
if y_test[i]!=predictions[i]:
print(y_test[i],predictions[i])
index.append(i)
# show false predictions in test_data
rows=int(len(index) / 3) + (len(index) % 3 > 0) # how many rows
# plot
fig = plt.figure(figsize=(20, 5*rows+1))
for i,ind in enumerate(index):
ax.set_axis_off() # hide axis
ax=plt.subplot(rows,3,i+1)
im=ax.imshow(X_test[ind].reshape((size,size)),cmap='jet', interpolation='nearest')
ax.set_title("pred: {},\n correct: {}".format(beam_data["target_names"][predictions[ind]],
beam_data["target_names"][y_test[ind]]))
#adjust colorbar to plot
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.1)
cbar=plt.colorbar(im, cax=cax)
cbar.ax.set_ylabel('intensity',fontsize=15)
```
### falsely classified in train data (unreasonable here, due to knn:n_neighbours=1)
```
# collect indices
index=[]
predictions=pipe.predict(X_train)
for i in range(len(y_train)):
if y_train[i]!=predictions[i]:
print((y_train[i],predictions[i]))
index.append(i)
# show false predictions in train data
rows=int(len(index) / 3) + (len(index) % 3 > 0) # how many rows
# plot
fig = plt.figure(figsize=(20, 5*rows+1))
for i,ind in enumerate(index):
ax.set_axis_off() # hide axis
ax=plt.subplot(rows,3,i+1)
im=ax.imshow(X_train[ind].reshape((size,size)),cmap='jet', interpolation='nearest')
ax.set_title("pred: {},\n correct: {}".format(beam_data["target_names"][predictions[ind]],
beam_data["target_names"][y_train[ind]]))
#adjust colorbar to plot
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.1)
cbar=plt.colorbar(im, cax=cax)
cbar.ax.set_ylabel('intensity',fontsize=15)
```
# Additional plots for thesis
## 3 categories
```
names=["75GHz.dat","105GHz-2.dat","85GHz-8.dat"]
indici=[beam_data["data_names"].index(name) for name in names]
maps=[beam_data["images"][ind] for ind in indici]
fig=plt.figure(figsize=(11,8))
for i,map_i in enumerate(maps,0):
ax=plt.subplot(1,3,i+1)
ax.set_axis_off() # hide axis
ax.set_title("{}\n labeled:{}".format(names[i],beam_data["target_names"][beam_data["target"][indici][i]]))
#plot
im=ax.imshow(map_i,cmap="jet")
#adjust colorbar to plot
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.1)
cbar=plt.colorbar(im, cax=cax)
cbar.ax.set_ylabel('intensity',fontsize=15)
plt.tight_layout()
plt.savefig("Images/3-categories.png", bbox_inches='tight')
```
## questionable labels
```
names=["85GHz.dat","85GHz-1.dat","95GHz-5.dat"]
indici=[beam_data["data_names"].index(name) for name in names]
maps=[beam_data["images"][ind] for ind in indici]
fig=plt.figure(figsize=(11,8))
for i,map_i in enumerate(maps,0):
ax=plt.subplot(1,3,i+1)
ax.set_axis_off() # hide axis
ax.set_title("{}\n labeled:{}".format(names[i],beam_data["target_names"][beam_data["target"][indici][i]]))
#plot
im=ax.imshow(map_i,cmap="jet")
#adjust colorbar to plot
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.1)
cbar=plt.colorbar(im, cax=cax)
cbar.ax.set_ylabel('intensity',fontsize=15)
plt.tight_layout()
plt.savefig("Images/questionable-labels.png", bbox_inches='tight')
```
| github_jupyter |
# Computer Vision Nanodegree
## Project: Image Captioning
---
In this notebook, you will train your CNN-RNN model.
You are welcome and encouraged to try out many different architectures and hyperparameters when searching for a good model.
This does have the potential to make the project quite messy! Before submitting your project, make sure that you clean up:
- the code you write in this notebook. The notebook should describe how to train a single CNN-RNN architecture, corresponding to your final choice of hyperparameters. You should structure the notebook so that the reviewer can replicate your results by running the code in this notebook.
- the output of the code cell in **Step 2**. The output should show the output obtained when training the model from scratch.
This notebook **will be graded**.
Feel free to use the links below to navigate the notebook:
- [Step 1](#step1): Training Setup
- [Step 2](#step2): Train your Model
- [Step 3](#step3): (Optional) Validate your Model
<a id='step1'></a>
## Step 1: Training Setup
In this step of the notebook, you will customize the training of your CNN-RNN model by specifying hyperparameters and setting other options that are important to the training procedure. The values you set now will be used when training your model in **Step 2** below.
You should only amend blocks of code that are preceded by a `TODO` statement. **Any code blocks that are not preceded by a `TODO` statement should not be modified**.
### Task #1
Begin by setting the following variables:
- `batch_size` - the batch size of each training batch. It is the number of image-caption pairs used to amend the model weights in each training step.
- `vocab_threshold` - the minimum word count threshold. Note that a larger threshold will result in a smaller vocabulary, whereas a smaller threshold will include rarer words and result in a larger vocabulary.
- `vocab_from_file` - a Boolean that decides whether to load the vocabulary from file.
- `embed_size` - the dimensionality of the image and word embeddings.
- `hidden_size` - the number of features in the hidden state of the RNN decoder.
- `num_epochs` - the number of epochs to train the model. We recommend that you set `num_epochs=3`, but feel free to increase or decrease this number as you wish. [This paper](https://arxiv.org/pdf/1502.03044.pdf) trained a captioning model on a single state-of-the-art GPU for 3 days, but you'll soon see that you can get reasonable results in a matter of a few hours! (_But of course, if you want your model to compete with current research, you will have to train for much longer._)
- `save_every` - determines how often to save the model weights. We recommend that you set `save_every=1`, to save the model weights after each epoch. This way, after the `i`th epoch, the encoder and decoder weights will be saved in the `models/` folder as `encoder-i.pkl` and `decoder-i.pkl`, respectively.
- `print_every` - determines how often to print the batch loss to the Jupyter notebook while training. Note that you **will not** observe a monotonic decrease in the loss function while training - this is perfectly fine and completely expected! You are encouraged to keep this at its default value of `100` to avoid clogging the notebook, but feel free to change it.
- `log_file` - the name of the text file containing - for every step - how the loss and perplexity evolved during training.
If you're not sure where to begin to set some of the values above, you can peruse [this paper](https://arxiv.org/pdf/1502.03044.pdf) and [this paper](https://arxiv.org/pdf/1411.4555.pdf) for useful guidance! **To avoid spending too long on this notebook**, you are encouraged to consult these suggested research papers to obtain a strong initial guess for which hyperparameters are likely to work best. Then, train a single model, and proceed to the next notebook (**3_Inference.ipynb**). If you are unhappy with your performance, you can return to this notebook to tweak the hyperparameters (and/or the architecture in **model.py**) and re-train your model.
### Question 1
**Question:** Describe your CNN-RNN architecture in detail. With this architecture in mind, how did you select the values of the variables in Task 1? If you consulted a research paper detailing a successful implementation of an image captioning model, please provide the reference.
**Answer:**
### (Optional) Task #2
Note that we have provided a recommended image transform `transform_train` for pre-processing the training images, but you are welcome (and encouraged!) to modify it as you wish. When modifying this transform, keep in mind that:
- the images in the dataset have varying heights and widths, and
- if using a pre-trained model, you must perform the corresponding appropriate normalization.
### Question 2
**Question:** How did you select the transform in `transform_train`? If you left the transform at its provided value, why do you think that it is a good choice for your CNN architecture?
**Answer:**
### Task #3
Next, you will specify a Python list containing the learnable parameters of the model. For instance, if you decide to make all weights in the decoder trainable, but only want to train the weights in the embedding layer of the encoder, then you should set `params` to something like:
```
params = list(decoder.parameters()) + list(encoder.embed.parameters())
```
### Question 3
**Question:** How did you select the trainable parameters of your architecture? Why do you think this is a good choice?
**Answer:**
### Task #4
Finally, you will select an [optimizer](http://pytorch.org/docs/master/optim.html#torch.optim.Optimizer).
### Question 4
**Question:** How did you select the optimizer used to train your model?
**Answer:**
```
import torch
import torch.nn as nn
from torchvision import transforms
import sys
sys.path.append('/opt/cocoapi/PythonAPI')
from pycocotools.coco import COCO
from data_loader import get_loader
from model import EncoderCNN, DecoderRNN
import math
## TODO #1: Select appropriate values for the Python variables below.
batch_size = ... # batch size
vocab_threshold = ... # minimum word count threshold
vocab_from_file = ... # if True, load existing vocab file
embed_size = ... # dimensionality of image and word embeddings
hidden_size = ... # number of features in hidden state of the RNN decoder
num_epochs = 3 # number of training epochs
save_every = 1 # determines frequency of saving model weights
print_every = 100 # determines window for printing average loss
log_file = 'training_log.txt' # name of file with saved training loss and perplexity
# (Optional) TODO #2: Amend the image transform below.
transform_train = transforms.Compose([
transforms.Resize(256), # smaller edge of image resized to 256
transforms.RandomCrop(224), # get 224x224 crop from random location
transforms.RandomHorizontalFlip(), # horizontally flip image with probability=0.5
transforms.ToTensor(), # convert the PIL Image to a tensor
transforms.Normalize((0.485, 0.456, 0.406), # normalize image for pre-trained model
(0.229, 0.224, 0.225))])
# Build data loader.
data_loader = get_loader(transform=transform_train,
mode='train',
batch_size=batch_size,
vocab_threshold=vocab_threshold,
vocab_from_file=vocab_from_file)
# The size of the vocabulary.
vocab_size = len(data_loader.dataset.vocab)
# Initialize the encoder and decoder.
encoder = EncoderCNN(embed_size)
decoder = DecoderRNN(embed_size, hidden_size, vocab_size)
# Move models to GPU if CUDA is available.
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
encoder.to(device)
decoder.to(device)
# Define the loss function.
criterion = nn.CrossEntropyLoss().cuda() if torch.cuda.is_available() else nn.CrossEntropyLoss()
# TODO #3: Specify the learnable parameters of the model.
params = ...
# TODO #4: Define the optimizer.
optimizer = ...
# Set the total number of training steps per epoch.
total_step = math.ceil(len(data_loader.dataset.caption_lengths) / data_loader.batch_sampler.batch_size)
```
<a id='step2'></a>
## Step 2: Train your Model
Once you have executed the code cell in **Step 1**, the training procedure below should run without issue.
It is completely fine to leave the code cell below as-is without modifications to train your model. However, if you would like to modify the code used to train the model below, you must ensure that your changes are easily parsed by your reviewer. In other words, make sure to provide appropriate comments to describe how your code works!
You may find it useful to load saved weights to resume training. In that case, note the names of the files containing the encoder and decoder weights that you'd like to load (`encoder_file` and `decoder_file`). Then you can load the weights by using the lines below:
```python
# Load pre-trained weights before resuming training.
encoder.load_state_dict(torch.load(os.path.join('./models', encoder_file)))
decoder.load_state_dict(torch.load(os.path.join('./models', decoder_file)))
```
While trying out parameters, make sure to take extensive notes and record the settings that you used in your various training runs. In particular, you don't want to encounter a situation where you've trained a model for several hours but can't remember what settings you used :).
### A Note on Tuning Hyperparameters
To figure out how well your model is doing, you can look at how the training loss and perplexity evolve during training - and for the purposes of this project, you are encouraged to amend the hyperparameters based on this information.
However, this will not tell you if your model is overfitting to the training data, and, unfortunately, overfitting is a problem that is commonly encountered when training image captioning models.
For this project, you need not worry about overfitting. **This project does not have strict requirements regarding the performance of your model**, and you just need to demonstrate that your model has learned **_something_** when you generate captions on the test data. For now, we strongly encourage you to train your model for the suggested 3 epochs without worrying about performance; then, you should immediately transition to the next notebook in the sequence (**3_Inference.ipynb**) to see how your model performs on the test data. If your model needs to be changed, you can come back to this notebook, amend hyperparameters (if necessary), and re-train the model.
That said, if you would like to go above and beyond in this project, you can read about some approaches to minimizing overfitting in section 4.3.1 of [this paper](http://ieeexplore.ieee.org/stamp/stamp.jsp?arnumber=7505636). In the next (optional) step of this notebook, we provide some guidance for assessing the performance on the validation dataset.
```
import torch.utils.data as data
import numpy as np
import os
import requests
import time
# Open the training log file.
f = open(log_file, 'w')
old_time = time.time()
response = requests.request("GET",
"http://metadata.google.internal/computeMetadata/v1/instance/attributes/keep_alive_token",
headers={"Metadata-Flavor":"Google"})
for epoch in range(1, num_epochs+1):
for i_step in range(1, total_step+1):
if time.time() - old_time > 60:
old_time = time.time()
requests.request("POST",
"https://nebula.udacity.com/api/v1/remote/keep-alive",
headers={'Authorization': "STAR " + response.text})
# Randomly sample a caption length, and sample indices with that length.
indices = data_loader.dataset.get_train_indices()
# Create and assign a batch sampler to retrieve a batch with the sampled indices.
new_sampler = data.sampler.SubsetRandomSampler(indices=indices)
data_loader.batch_sampler.sampler = new_sampler
# Obtain the batch.
images, captions = next(iter(data_loader))
# Move batch of images and captions to GPU if CUDA is available.
images = images.to(device)
captions = captions.to(device)
# Zero the gradients.
decoder.zero_grad()
encoder.zero_grad()
# Pass the inputs through the CNN-RNN model.
features = encoder(images)
outputs = decoder(features, captions)
# Calculate the batch loss.
loss = criterion(outputs.view(-1, vocab_size), captions.view(-1))
# Backward pass.
loss.backward()
# Update the parameters in the optimizer.
optimizer.step()
# Get training statistics.
stats = 'Epoch [%d/%d], Step [%d/%d], Loss: %.4f, Perplexity: %5.4f' % (epoch, num_epochs, i_step, total_step, loss.item(), np.exp(loss.item()))
# Print training statistics (on same line).
print('\r' + stats, end="")
sys.stdout.flush()
# Print training statistics to file.
f.write(stats + '\n')
f.flush()
# Print training statistics (on different line).
if i_step % print_every == 0:
print('\r' + stats)
# Save the weights.
if epoch % save_every == 0:
torch.save(decoder.state_dict(), os.path.join('./models', 'decoder-%d.pkl' % epoch))
torch.save(encoder.state_dict(), os.path.join('./models', 'encoder-%d.pkl' % epoch))
# Close the training log file.
f.close()
```
<a id='step3'></a>
## Step 3: (Optional) Validate your Model
To assess potential overfitting, one approach is to assess performance on a validation set. If you decide to do this **optional** task, you are required to first complete all of the steps in the next notebook in the sequence (**3_Inference.ipynb**); as part of that notebook, you will write and test code (specifically, the `sample` method in the `DecoderRNN` class) that uses your RNN decoder to generate captions. That code will prove incredibly useful here.
If you decide to validate your model, please do not edit the data loader in **data_loader.py**. Instead, create a new file named **data_loader_val.py** containing the code for obtaining the data loader for the validation data. You can access:
- the validation images at filepath `'/opt/cocoapi/images/train2014/'`, and
- the validation image caption annotation file at filepath `'/opt/cocoapi/annotations/captions_val2014.json'`.
The suggested approach to validating your model involves creating a json file such as [this one](https://github.com/cocodataset/cocoapi/blob/master/results/captions_val2014_fakecap_results.json) containing your model's predicted captions for the validation images. Then, you can write your own script or use one that you [find online](https://github.com/tylin/coco-caption) to calculate the BLEU score of your model. You can read more about the BLEU score, along with other evaluation metrics (such as TEOR and Cider) in section 4.1 of [this paper](https://arxiv.org/pdf/1411.4555.pdf). For more information about how to use the annotation file, check out the [website](http://cocodataset.org/#download) for the COCO dataset.
```
# (Optional) TODO: Validate your model.
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import math
import time
data = pd.read_csv('mnist.csv')
data.head()
train_data = data.sample(frac=0.8)
test_data = data.drop(train_data.index)
train_labels = train_data['label'].values
train_data = train_data.drop('label', axis=1).values
test_labels = test_data['label'].values
test_data = test_data.drop('label', axis=1).values
num_im = 25
num_cells = math.ceil(math.sqrt(num_im))
plt.figure(figsize=(10, 10))
for i in range(num_im):
pixels = train_data[i]
size = math.ceil(math.sqrt(pixels.size))
pixels = pixels.reshape(size, size)
plt.subplot(num_cells, num_cells, i+1)
plt.title(train_labels[i])
plt.tick_params(axis='both', which='both', bottom=False, left=False, labelbottom=False, labelleft=False)
plt.imshow(pixels)
def sigmoid(x):
return 1 / (1 + np.e ** -x)
x = np.linspace(-10, 10, 50)
y = sigmoid(x)
plt.plot(x, y)
plt.title('Sigmoid Function')
plt.show()
num_samples, num_features = train_data.shape
W = None
train_data = (train_data - train_data.min()) / (train_data.max() - train_data.min())
X = np.c_[np.ones(num_samples), train_data]
num_iters = 5000
lr = 0.001
lambda_ = 0.01
start = time.time()
# train each label using one vs all
for label in range(10):
w = np.random.rand(num_features+1)
y = (train_labels == label).astype(float)
for i in range(num_iters):
diff = sigmoid(X @ w) - y
for wi in range(len(w)):
if wi == 0:
t = diff
reg_term = 0
else:
t = diff @ X[:, wi]
reg_term = lambda_ * X[:, wi]
w[wi] -= lr * np.sum(t - reg_term) / num_samples
if (i + 1) % 1000 == 0:
reg_term = lambda_ * np.sum(w[1:] @ w[1:].T) / num_samples
t = sigmoid(X @ w)
cost = np.sum(-y * np.log(t) - (1-y) * np.log(1-t)) / 2 / num_samples - reg_term
print(f'label={label} iteration={i+1} cost={cost:.5f}')
W = w if W is None else np.vstack((W, w))
print(f'Training Finished | Time Taken = {time.time() - start:.2f}s')
train_pred = np.argmax(X @ W.T, axis=1)
train_acc = np.sum(train_pred == train_labels) / num_samples
print(f'Train Accuracy = {train_acc:.5f}')
test_X = np.c_[np.ones(len(test_data)), test_data]
test_pred = np.argmax(test_X @ W.T, axis=1)
test_acc = np.sum(test_pred == test_labels) / len(test_data)
print(f'Test Accuracy = {test_acc:.5f}')
num_im = 9
num_cells = math.ceil(math.sqrt(num_im))
plt.figure(figsize=(10, 10))
for i in range(num_im):
pixels = W[i][1:]
size = math.ceil(math.sqrt(pixels.size))
pixels = pixels.reshape(size, size)
plt.subplot(num_cells, num_cells, i+1)
plt.title(i)
plt.tick_params(axis='both', which='both', bottom=False, left=False, labelbottom=False, labelleft=False)
plt.imshow(pixels, cmap='Greys')
num_im = 64
num_cells = math.ceil(math.sqrt(num_im))
plt.figure(figsize=(15, 15))
for i in range(num_im):
label = test_labels[i]
pixels = test_data[i]
size = math.ceil(math.sqrt(pixels.size))
pixels = pixels.reshape(size, size)
x = np.concatenate(([1], test_data[i]))
pred = np.argmax(x @ W.T)
plt.subplot(num_cells, num_cells, i+1)
plt.title(pred)
plt.tick_params(axis='both', which='both', bottom=False, left=False, labelbottom=False, labelleft=False)
plt.imshow(pixels, cmap='Greens' if pred == label else 'Reds')
```
| github_jupyter |
```
from toolz import curry
import pandas as pd
import numpy as np
from scipy.special import expit
from linearmodels.panel import PanelOLS
import statsmodels.formula.api as smf
import seaborn as sns
from matplotlib import pyplot as plt
from matplotlib import style
style.use("ggplot")
```
# Difference-in-Diferences: Death and Rebirth
## The Promise of Panel Data
Panel data is when we have multiple units `i` over multiple periods of time `t`. Think about a policy evaluation scenario in the US, where you want to check the effect of cannabis legalization on crime rate. You have crime rate data on multiple states `i` over multiple time perios `t`. You also observe at what point in time each state adopts legislation in the direction of canabis legalization. I hope you can see why this is incredibly powerfull for causal inference. Call canabis legalization the treatment `D` (since `T` is taken; it represents time). We can follow the trend on crime rates for a particular state that eventually gets treated and see if there are any disruptions in the trend at the treatment time. In a way, a state serves as its own control unit, in a sort of before and after comparisson. Furthermore, becase we have multiple states, we can also compare treated states to control states. When we put both comparissons toguether, treated vs control and before and after treatement, we end up with an incredibly powerfull tool to infer counterfactuals and, hence, causal effects.
Panel data methods are often used in govenment policy evaluation, but we can easily make an argument about why it is also incredibly usefull for the (tech) industry. Companies often track user data across multiple periods of time, which results in a rich panel data structure. To expore that idea further, let's consider a hypothetical example of a tech company that traked customers for multiple years. Along those years, it rolled out a new product for some of its customers. More specifically, some customers got acess to the new product in 1985, others in 1994 and others in the year 2000. In causal infererence terms, we can already see that the new product can be seen as a treatment. We call each of those **groups of customers that got treated at the same time a cohort**. In this hypothetical exemple, we want to figure out the impact of the new product on sales. The folowing image shows how sales evolve over time for each of the treated cohorts, plus a never treated group of customers.
```
time = range(1980, 2010)
cohorts = [1985,1994,2000,2011]
units = range(1, 100+1)
np.random.seed(1)
df_hom_effect = pd.DataFrame(dict(
year = np.tile(time, len(units)),
unit = np.repeat(units, len(time)),
cohort = np.repeat(np.random.choice(cohorts, len(units)), len(time)),
unit_fe = np.repeat(np.random.normal(0, 5, size=len(units)), len(time)),
time_fe = np.tile(np.random.normal(size=len(time)), len(units)),
)).assign(
trend = lambda d: (d["year"] - d["year"].min())/8,
post = lambda d: (d["year"] >= d["cohort"]).astype(int),
).assign(
treat = 1,
y0 = lambda d: 10 + d["trend"] + d["unit_fe"] + 0.1*d["time_fe"],
).assign(
treat_post = lambda d: d["treat"]*d["post"],
y1 = lambda d: d["y0"] + 1
).assign(
tau = lambda d: d["y1"] - d["y0"],
sales = lambda d: np.where(d["treat_post"] == 1, d["y1"], d["y0"])
).drop(columns=["unit_fe", "time_fe", "trend", "y0", "y1"])
plt.figure(figsize=(10,4))
[plt.vlines(x=c, ymin=9, ymax=15, color="black", ls="dashed") for c in cohorts[:-1]]
sns.lineplot(
data=(df_hom_effect
.replace({"cohort":{2011:"never-treated"}})
.groupby(["cohort", "year"])["sales"]
.mean()
.reset_index()),
x="year",
y = "sales",
hue="cohort",
);
```
Let's take a momen to appreciate the richness of the data depicted in the above plot. First, we can see that each cohorts have its own baseline level. Thats simply because different customers buy different ammounts. For instance, it looks like customers in the never treated cohort have a higher baseline (of about 11), compared to other cohorts. This means that simply comparing treated cohorts to control cohorts would yield a biased result, since $Y_{0}$ for the neve treated is higher than the $Y_{0}$ for the treated. Fortunatly, we can compare acorss units and time.
Speaking of time, notice how there is an overall upward trend with some wigles (for example, there is a dip in the year 1999). Since latter years have higher $Y_{0}$ than early years, simply comparing the same unit across time would also yield in biased results. Once again, we are fortunate that the pannel data structure allow us to compare not only across time, but also across units.
Another way to see the power of panel data structure in through the lens of linear models and linear regression. Let's say each of our customers `i` has a spend propensity $\gamma$. This is because of indosincasies due to stuff we can't observe, like customer's salary, family sinze and so on. Also, we can say that each year has an sales level $\theta$. Again, maibe because there is a crisis in one year, sales drop. If that is the case, a good way of modeling sales is to say it depends on the customer effect $\gamma$ and the time effect $\theta$, plus some random noise.
$$
Sales_{it} = \gamma_i + \theta_t + e_{it}
$$
To include the treatment in this picture, lets define a variable $D_{it}$ wich is 1 if the unit is treated. In our example, this variable would be always zero for the never treated cohort. It would also be zero for all the other cohorts at the begining, but it would turn into 1 at year 1985 for the cohort treated in 1985 and stay on after that. Same thing for other cohorts, it would turn into 1 at 1994 fot the cohort treated in 1994 and so on. We can include in our model of sales as follows:
$$
Sales_{it} = \tau D_{it} + \gamma_i + \theta_t + e_{it}
$$
Estimating the above model is OLS is what is called the Two-Way Fixed Effects Models (TWFE). Notice that $\tau$ would be the treatment effect, as it tells us how much sales changes once units are treated. Another way of looking at it is to invoke the "holding things constant" propriety of linear regression. If we estimate the above model, we could read the estimate of $\tau$ as how much sales would change if we flip the treatment from 0 to 1 while holding the unit `i` and time `t` fixed. Take a minute to appriciate how bold this is! To say we would hold each unit fixed while seeng how $D$ changes the outcome is to say we are controling for all unit specific characteristic, known and unknown. For example, we would be controling for customers past sales, wich we could measure, but also stuff we have no idea about, like how much the customer like our brand, his salary... The only requirement is that this caracteristic is fixed over the time of the analysis. Moreover, to say we would hold each time period fixed is to say we are controlling for all year specifit characteristic. For instance, since we are holding year fixed, while looking at the effect of $D$, that trend over there would vanish.
To see all this power in action all we have to do is run an OLS model with the treatment indicator $D$ (`treat_post` here), plut dummies for the units and time. In our particular example, I've generated data in such a way that the effect of the treatment (new product) is to increase sales by 1. Notice how TWFE nais in recovering that treatment effect
```
formula = f"""sales ~ treat_post + C(unit) + C(year)"""
mod = smf.ols(formula, data=df_hom_effect)
result = mod.fit()
result.params["treat_post"]
```
Since I've simulated the data above, I know exactly the true individual treatment effect, which is stored in the `tau` column. Since the TWFE recovers the treatment effect on the treated, we can verify that the true ATT matches the one estimated above.
```
df_hom_effect.query("treat_post==1")["tau"].mean()
```
Before anyone comes and say that generating one dummy column for each unit is impossible with big data, let me come foreward and tell you that, yes, that is true. But there is a easy work around. We can use the FWL theorem to partiall that single regression into two. In fact, runing the above model is numerically equivalent to estimating the following model
$$
\tilde{Sales}_{it} = \tau \tilde D_{it} + e_{it}
$$
where
$$
\tilde{Sales}_{it} = Sales_{it} - \underbrace{\frac{1}{T}\sum_{t=0}^T Sales_{it}}_\text{Time Average} - \underbrace{\frac{1}{N}\sum_{i=0}^N Sales_{it}}_\text{Unit Average}
$$
and
$$
\tilde{D}_{it} = D_{it} - \frac{1}{T}\sum_{t=0}^T D_{it} - \frac{1}{N}\sum_{i=0}^N D_{it}
$$
In words now, in case the math is too crowded, we subtract the unit average across time (first term) and the time average across units (second term) from both the treatment indicator and the outcome variable to constrict the residuals. This process is often times called de-meaning, since we subtract the mean from the outcome and treatment. Finally, here is the same exact thing, but in code:
```
@curry
def demean(df, col_to_demean):
return df.assign(**{col_to_demean: (df[col_to_demean]
- df.groupby("unit")[col_to_demean].transform("mean")
- df.groupby("year")[col_to_demean].transform("mean"))})
formula = f"""sales ~ treat_post"""
mod = smf.ols(formula,
data=df_hom_effect
.pipe(demean(col_to_demean="treat_post"))
.pipe(demean(col_to_demean="sales")))
result = mod.fit()
result.summary().tables[1]
```
As we can see, with the alternative implementation, TWFE is also able to perfectly recover the ATT of 1.
## Assuptions
Two
## Death
## Trend in the Effect
```
time = range(1980, 2010)
cohorts = [1985,1994,2000,2011]
units = range(1, 100+1)
np.random.seed(3)
df_trend_effect = pd.DataFrame(dict(
year = np.tile(time, len(units)),
unit = np.repeat(units, len(time)),
cohort = np.repeat(np.random.choice(cohorts, len(units)), len(time)),
unit_fe = np.repeat(np.random.normal(size=len(units)), len(time)),
time_fe = np.tile(np.random.normal(size=len(time)), len(units)),
)).assign(
relative_year = lambda d: d["year"] - d["cohort"],
trend = lambda d: (d["year"] - d["year"].min())/8,
post = lambda d: (d["year"] >= d["cohort"]).astype(int),
).assign(
treat = 1,
y0 = lambda d: 10 + d["unit_fe"] + 0.02*d["time_fe"],
).assign(
y1 = lambda d: d["y0"] + np.minimum(0.2*(np.maximum(0, d["year"] - d["cohort"])), 1)
).assign(
tau = lambda d: d["y1"] - d["y0"],
outcome = lambda d: np.where(d["treat"]*d["post"] == 1, d["y1"], d["y0"])
)
plt.figure(figsize=(10,4))
sns.lineplot(
data=df_trend_effect.groupby(["cohort", "year"])["outcome"].mean().reset_index(),
x="year",
y = "outcome",
hue="cohort",
);
formula = f"""outcome ~ treat:post + C(year) + C(unit)"""
mod = smf.ols(formula, data=df_trend_effect)
result = mod.fit()
result.params["treat:post"]
df_trend_effect.query("treat==1 & post==1")["tau"].mean()
```
### Event Study Desing
```
relative_years = range(-10,10+1)
formula = "outcome~"+"+".join([f'Q({c})' for c in relative_years]) + "+C(unit)+C(year)"
mod = smf.ols(formula,
data=(df_trend_effect.join(pd.get_dummies(df_trend_effect["relative_year"]))))
result = mod.fit()
ax = (df_trend_effect
.query("treat==1")
.query("relative_year>-10")
.query("relative_year<10")
.groupby("relative_year")["tau"].mean().plot())
ax.plot(relative_years, result.params[-len(relative_years):]);
```
## Covariates
## X-Specific Trends
```
time = range(1980, 2000)
cohorts = [1990]
units = range(1, 100+1)
np.random.seed(3)
x = np.random.choice(np.random.normal(size=len(units)//10), size=len(units))
df_cov_trend = pd.DataFrame(dict(
year = np.tile(time, len(units)),
unit = np.repeat(units, len(time)),
cohort = np.repeat(np.random.choice(cohorts, len(units)), len(time)),
unit_fe = np.repeat(np.random.normal(size=len(units)), len(time)),
time_fe = np.tile(np.random.normal(size=len(time)), len(units)),
x = np.repeat(x, len(time)),
)).assign(
trend = lambda d: d["x"]*(d["year"] - d["year"].min())/20,
post = lambda d: (d["year"] >= d["cohort"]).astype(int),
).assign(
treat = np.repeat(np.random.binomial(1, expit(x)), len(time)),
y0 = lambda d: 10 + d["trend"] + 0.5*d["unit_fe"] + 0.01*d["time_fe"],
).assign(
y1 = lambda d: d["y0"] + 1
).assign(
tau = lambda d: d["y1"] - d["y0"],
outcome = lambda d: np.where(d["treat"]*d["post"] == 1, d["y1"], d["y0"])
)
plt.figure(figsize=(10,4))
sns.lineplot(
data=df_cov_trend.groupby(["treat", "year"])["outcome"].mean().reset_index(),
x="year",
y = "outcome",
hue="treat",
);
facet_col = "x"
all_facet_values = sorted(df_cov_trend[facet_col].unique())
g = sns.FacetGrid(df_cov_trend, col=facet_col, sharey=False, sharex=False, col_wrap=4, height=5, aspect=1)
for x, ax in zip(all_facet_values, g.axes):
plot_df = df_cov_trend.query(f"{facet_col}=={x}")
sns.lineplot(
data=plot_df.groupby(["treat", "year"])["outcome"].mean().reset_index(),
x="year",
y = "outcome",
hue="treat",
ax=ax
)
ax.set_title(f"X = {round(x, 2)}")
plt.tight_layout()
formula = f"""outcome ~ treat:post + C(year) + C(unit)"""
mod = smf.ols(formula, data=df_cov_trend)
result = mod.fit()
result.params["treat:post"]
formula = f"""outcome ~ treat:post + x * C(year) + C(unit)"""
mod = smf.ols(formula, data=df_cov_trend)
result = mod.fit()
result.params["treat:post"]
df_cov_trend.query("treat==1 & post==1")["tau"].mean()
```
| github_jupyter |
# Clase 7
El objetivo con esta sesión es entender las nociones básicas y metodología, para realizar una simulación de algún proceso de nuestra vida cotidiana o profesional.
## Etapas para realizar un estudio de simulación
> - *Definición del sistema*: determinar la interacción del sistema con otros sistemas, restricciones, interacción e interrelación de variables de interés y los resultados esperados.
> - *Formulación de un modelo*: Es necesario definir todas las variables que forman parte del sistema y además definir un diagrama de flujo que describa la forma completa del modelo.
> - *Colección de datos*: Definir los datos necesarios para el modelo. Datos pueden provenir de registros contables, órdenes de trabajo, órdenes de compra, opiniones de expertos y si no hay otro remedio por experimentación.
> - *Validación*: En esta etapa es posible detectar deficiencias en la formulación del modelo o en los datos sumunistrados al modelo. Formas de validar un modelo son:
- Opinión de expertos.
- Exactitud con que se predicen datos hitóricos.
- Exactitud de la predicción del futuro.
- La aceptación y confianza en el modelo de la persona que hará uso de los resultados que arroje el experimento de simulación.
> - *Experimentación*: La experimentación con el modelo se realiza después de que éste ha sido validado. La experimentación consiste en generar los datos deseados y en realizar el análisis de sensibilidad de los índices requeridos.
> - *Interpretación*: Se interpretan los resultados que arroja la simulación y en base a esto se toma una decisión.
> - *Documentación*:
- Datos que debe de tener el modelo
- Manual de usurario
# Casos positivos Covid-19 [fuente](https://ourworldindata.org/coronavirus-source-data)
```
# Importación de paquetes
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import os
import scipy.stats as st
# Leer los datos del archivo covid_cases y quedarse únicamente con los datos de méxico
data = pd.read_csv('covid_cases.csv')
data_mx = data[data['location'] == 'Mexico']
data_mx.set_index('date', inplace=True)
data_mx = data_mx.dropna(subset=['total_cases'])
data_mx.head()
# Seleccionar únicamente los casos totales y graficarlos
total_cases = data_mx['total_cases']
total_cases.plot(figsize=[10,5])
# Analizar los datos en el dominio de la frecuencia (histograma)
# total_cases.hist(figsize=[10,5], bins=50)
media = total_cases.mean()
N = 1000
expon = st.expon(loc=0, scale=media).rvs(size=N, random_state=123)
plt.hist(expon, bins=50, density=True);
# Analizar el help de la función st.expon (paŕametros y método rvs)
# Generar 10**4 valores aleatorios de dicha distribución y luego comparar
# el histograma de los datos reales con los datos simulados
N = int(1e4)
expon = st.expon(loc=0, scale=media).rvs(size=N, random_state=123)
fig, ax = plt.subplots(1,2, figsize=[10,5], sharey=True)
# Estos son los datos simulados
ax[0].hist(expon, bins=50, density=True, label='datos simulados')
# Datos reales
total_cases.hist(ax=ax[1], label='datos reales', density=True)
plt.legend()
plt.show()
```
## ¿Cómo usaríamos montecarlo para saber en un futuro cuantas personas es probable que se infecten?
```
escenarios, fecha_futura = 10000, 7
expon = st.expon(loc=0, scale=media).rvs(size=[escenarios, fecha_futura], random_state=123)
expon.mean(axis=0).sum()
```
# Simular dado cargado
Suponga que tenemos un dado cargado que distribuye según un distribución de probabilidad binomial con parámetros `n=5`, `p=0.5`.
```
np.random.seed(344)
# Graficar el histograma de 1000 lanazamientos del dado
d_no_cargado = np.random.randint(1,7,10000)
d_cargado = st.binom(n=5, p=0.1).rvs(size=1000)
y, x = np.histogram(d_cargado, bins=6, density=True)
plt.bar(range(1,7), y)
# d_cargado = st.binom(n=5, p=0.5).rvs(size=1000)
y[-1]
```
# Ejercicio
Si el usuario tira cualquier número entre 1 y 50, el casino gana. Si el usuario tira cualquier número entre 51 y 99, el usuario gana. Si el usuario tira 100, pierde.
```
# Solución
def juego(size=1):
dado = np.random.randint(0,101, size=size)
if 1<= dado <= 50:
return 0
elif 51<= dado <= 99:
return 1
else:
return 0
N = 10000
juegos = [juego() for i in range(N)]
sum(juegos) / N
```
# Tarea 4
# 1
Como ejemplo simple de una simulación de Monte Carlo, considere calcular la probabilidad de una suma particular del lanzamiento de tres dados (cada dado tiene valores del uno al seis). Además cada dado tiene las siguientes carácterísticas: el primer dado no está cargado (distribución uniforme todos son equiprobables); el segundo y tercer dado están cargados basados en una distribución binomial con parámetros (`n=5, p=0.5` y `n=5, p=0.2`). Calcule la probabilidad de que la suma resultante sea 7, 14 o 18.
```
# Solución
```
# 2 Ejercicio de aplicación- Cafetería Central
Premisas para la simulación:
- Negocio de alimentos que vende bebidas y alimentos.
- Negocio dentro del ITESO.
- Negocio en cafetería central.
- Tipo de clientes (hombres y mujeres).
- Rentabilidad del 60%.
## Objetivo
Realizar una simulación estimado el tiempo medio que se tardaran los clientes en ser atendidos entre el horario de 6:30 a 1 pm. Además saber el consumo.
**Analizar supuestos y limitantes**
## Supuestos en simulación
Clasificación de clientes:
- Mujer = 1 $\longrightarrow$ aleatorio < 0.5
- Hombre = 0 $\longrightarrow$ aleatorio $\geq$ 0.5.
Condiciones iniciales:
- Todas las distrubuciones de probabilidad se supondrán uniformes.
- Tiempo de simulación: 6:30 am - 1:30pm $\longrightarrow$ T = 7 horas = 25200 seg.
- Tiempo de llegada hasta ser atendido: Min=5seg, Max=30seg.
- Tiempo que tardan los clientes en ser atendidos:
- Mujer: Min = 1 min= 60seg, Max = 5 min = 300 seg
- Hombre: Min = 40 seg, Max = 2 min= 120 seg
- Consumo según el tipo de cliente:
- Mujer: Min = 30 pesos, Max = 100 pesos
- Hombre: Min = 20 pesos, Max = 80 pesos
Responder las siguientes preguntas basados en los datos del problema:
1. ¿Cuáles fueron los gastos de los hombres y las mujeres en 5 días de trabajo?.
2. ¿Cuál fue el consumo promedio de los hombres y mujeres?
3. ¿Cuál fue el número de personas atendidas por día?
4. ¿Cuál fue el tiempo de atención promedio?
5. ¿Cuánto fue la ganancia promedio de la cafetería en 5 días de trabajo y su respectiva rentabilidad?
```
################## Datos del problema
d = 5
T =25200
T_at_min = 5; T_at_max = 30
T_mujer_min =60; T_mujer_max = 300
T_hombre_min = 40; T_hombre_max = 120
C_mujer_min = 30; C_mujer_max = 100
C_hombre_min = 20; C_hombre_max = 80
```
| github_jupyter |
<a href="https://colab.research.google.com/github/sokrypton/ColabFold/blob/main/beta/AlphaFold2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#AlphaFold2 w/ MMseqs2
Easy to use version of AlphaFold 2 [(Jumper et al. 2021, Nature)](https://www.nature.com/articles/s41586-021-03819-2) a protein structure prediction pipeline, with an API hosted at the Södinglab based on the MMseqs2 server [(Mirdita et al. 2019, Bioinformatics)](https://academic.oup.com/bioinformatics/article/35/16/2856/5280135) for the multiple sequence alignment creation.
**Limitations**
- This notebook does NOT use the AlphaFold2's jackhmmer pipeline for MSA/template generation. It may give better or worse results depending on number of sequences that can be found. Check out the [full AlphaFold2 pipeline](https://github.com/deepmind/alphafold) or Deepmind's official [google-colab notebook](https://colab.research.google.com/github/deepmind/alphafold/blob/main/notebooks/AlphaFold.ipynb).
- For homo-oligomeric setting, amber-relax and templates are currently NOT supported.
- For a typical Google-Colab session, with a `16G-GPU`, the max total length is **1400 residues**. Sometimes a `12G-GPU` is assigned, in which the max length is ~1000 residues.
**WARNING**:
<strong>For detailed instructions, see <a href="#Instructions">bottom</a> of notebook!</strong>
```
#@title Input protein sequence, then hit `Runtime` -> `Run all`
import os
os.environ['TF_FORCE_UNIFIED_MEMORY'] = '1'
os.environ['XLA_PYTHON_CLIENT_MEM_FRACTION'] = '2.0'
import re
import hashlib
from google.colab import files
def add_hash(x,y):
return x+"_"+hashlib.sha1(y.encode()).hexdigest()[:5]
query_sequence = 'PIAQIHILEGRSDEQKETLIREVSEAISRSLDAPLTSVRVIITEMAKGHFGIGGELASK' #@param {type:"string"}
# remove whitespaces
query_sequence = "".join(query_sequence.split())
query_sequence = re.sub(r'[^a-zA-Z]','', query_sequence).upper()
jobname = 'test' #@param {type:"string"}
# remove whitespaces
jobname = "".join(jobname.split())
jobname = re.sub(r'\W+', '', jobname)
jobname = add_hash(jobname, query_sequence)
with open(f"{jobname}.fasta", "w") as text_file:
text_file.write(">1\n%s" % query_sequence)
# number of models to use
#@markdown ---
#@markdown ### Advanced settings
msa_mode = "MMseqs2 (UniRef+Environmental)" #@param ["MMseqs2 (UniRef+Environmental)", "MMseqs2 (UniRef only)","single_sequence","custom"]
num_models = 5 #@param [1,2,3,4,5] {type:"raw"}
use_msa = True if msa_mode.startswith("MMseqs2") else False
use_env = True if msa_mode == "MMseqs2 (UniRef+Environmental)" else False
use_custom_msa = True if msa_mode == "custom" else False
use_amber = False #@param {type:"boolean"}
use_templates = False #@param {type:"boolean"}
use_ptm = True #@param {type:"boolean"}
#@markdown ---
#@markdown ### Experimental options
homooligomer = 1 #@param [1,2,3,4,5,6,7,8] {type:"raw"}
save_to_google_drive = False #@param {type:"boolean"}
#@markdown ---
#@markdown Don't forget to hit `Runtime` -> `Run all` after updating form
if homooligomer > 1:
if use_amber:
print("amber disabled: amber is not currently supported for homooligomers")
use_amber = False
if use_templates:
print("templates disabled: templates are not currently supported for homooligomers")
use_templates = False
# decide which a3m to use
if use_msa:
a3m_file = f"{jobname}.a3m"
elif use_custom_msa:
a3m_file = f"{jobname}.custom.a3m"
if not os.path.isfile(a3m_file):
custom_msa_dict = files.upload()
custom_msa = list(custom_msa_dict.keys())[0]
header = 0
import fileinput
for line in fileinput.FileInput(custom_msa,inplace=1):
if line.startswith(">"):
header = header + 1
if line.startswith("#"):
continue
if line.rstrip() == False:
continue
if line.startswith(">") == False and header == 1:
query_sequence = line.rstrip()
print(line, end='')
os.rename(custom_msa, a3m_file)
print(f"moving {custom_msa} to {a3m_file}")
else:
a3m_file = f"{jobname}.single_sequence.a3m"
with open(a3m_file, "w") as text_file:
text_file.write(">1\n%s" % query_sequence)
if save_to_google_drive == True:
from pydrive.drive import GoogleDrive
from pydrive.auth import GoogleAuth
from google.colab import auth
from oauth2client.client import GoogleCredentials
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
print("You are logged into Google Drive and are good to go!")
#@title Install dependencies
%%bash -s $use_amber $use_msa $use_templates
USE_AMBER=$1
USE_MSA=$2
USE_TEMPLATES=$3
if [ ! -f AF2_READY ]; then
# install dependencies
pip -q install biopython
pip -q install dm-haiku
pip -q install ml-collections
pip -q install py3Dmol
# download model
if [ ! -d "alphafold/" ]; then
git clone https://github.com/deepmind/alphafold.git --quiet
(cd alphafold; git checkout 1e216f93f06aa04aa699562f504db1d02c3b704c --quiet)
mv alphafold alphafold_
mv alphafold_/alphafold .
# remove "END" from PDBs, otherwise biopython complains
sed -i "s/pdb_lines.append('END')//" /content/alphafold/common/protein.py
sed -i "s/pdb_lines.append('ENDMDL')//" /content/alphafold/common/protein.py
fi
# download model params (~1 min)
if [ ! -d "params/" ]; then
wget -qnc https://storage.googleapis.com/alphafold/alphafold_params_2021-07-14.tar
mkdir params
tar -xf alphafold_params_2021-07-14.tar -C params/
rm alphafold_params_2021-07-14.tar
fi
touch AF2_READY
fi
# download libraries for interfacing with MMseqs2 API
if [ ${USE_MSA} == "True" ] || [ ${USE_TEMPLATES} == "True" ]; then
if [ ! -f MMSEQ2_READY ]; then
apt-get -qq -y update 2>&1 1>/dev/null
apt-get -qq -y install jq curl zlib1g gawk 2>&1 1>/dev/null
touch MMSEQ2_READY
fi
fi
# setup conda
if [ ${USE_AMBER} == "True" ] || [ ${USE_TEMPLATES} == "True" ]; then
if [ ! -f CONDA_READY ]; then
wget -qnc https://repo.anaconda.com/miniconda/Miniconda3-latest-Linux-x86_64.sh
bash Miniconda3-latest-Linux-x86_64.sh -bfp /usr/local 2>&1 1>/dev/null
rm Miniconda3-latest-Linux-x86_64.sh
touch CONDA_READY
fi
fi
# setup template search
if [ ${USE_TEMPLATES} == "True" ] && [ ! -f HH_READY ]; then
conda install -y -q -c conda-forge -c bioconda kalign3=3.2.2 hhsuite=3.3.0 python=3.7 2>&1 1>/dev/null
touch HH_READY
fi
# setup openmm for amber refinement
if [ ${USE_AMBER} == "True" ] && [ ! -f AMBER_READY ]; then
conda install -y -q -c conda-forge openmm=7.5.1 python=3.7 pdbfixer 2>&1 1>/dev/null
(cd /usr/local/lib/python3.7/site-packages; patch -s -p0 < /content/alphafold_/docker/openmm.patch)
wget -qnc https://git.scicore.unibas.ch/schwede/openstructure/-/raw/7102c63615b64735c4941278d92b554ec94415f8/modules/mol/alg/src/stereo_chemical_props.txt
mv stereo_chemical_props.txt alphafold/common/
touch AMBER_READY
fi
#@title Import libraries
# setup the model
if "IMPORTED" not in dir():
import numpy as np
import pickle
from string import ascii_uppercase
from alphafold.common import protein
from alphafold.data import pipeline
from alphafold.data import templates
from alphafold.model import data
from alphafold.model import config
from alphafold.model import model
from alphafold.data.tools import hhsearch
from absl import logging
logging.set_verbosity("error")
# plotting libraries
import py3Dmol
import matplotlib.pyplot as plt
IMPORTED = True
if use_amber and "relax" not in dir():
import sys
sys.path.insert(0, '/usr/local/lib/python3.7/site-packages/')
from alphafold.relax import relax
def mk_template(jobname):
template_featurizer = templates.TemplateHitFeaturizer(
mmcif_dir="templates/",
max_template_date="2100-01-01",
max_hits=20,
kalign_binary_path="kalign",
release_dates_path=None,
obsolete_pdbs_path=None)
hhsearch_pdb70_runner = hhsearch.HHSearch(binary_path="hhsearch",databases=[jobname])
a3m_lines = "\n".join(open(f"{jobname}.a3m","r").readlines())
hhsearch_result = hhsearch_pdb70_runner.query(a3m_lines)
hhsearch_hits = pipeline.parsers.parse_hhr(hhsearch_result)
templates_result = template_featurizer.get_templates(query_sequence=query_sequence,
query_pdb_code=None,
query_release_date=None,
hits=hhsearch_hits)
return templates_result.features
def set_bfactor(pdb_filename, bfac, idx_res, chains):
I = open(pdb_filename,"r").readlines()
O = open(pdb_filename,"w")
for line in I:
if line[0:6] == "ATOM ":
seq_id = int(line[22:26].strip()) - 1
seq_id = np.where(idx_res == seq_id)[0][0]
O.write(f"{line[:21]}{chains[seq_id]}{line[22:60]}{bfac[seq_id]:6.2f}{line[66:]}")
O.close()
def predict_structure(prefix, feature_dict, Ls, random_seed=0):
"""Predicts structure using AlphaFold for the given sequence."""
# Minkyung's code
# add big enough number to residue index to indicate chain breaks
idx_res = feature_dict['residue_index']
L_prev = 0
# Ls: number of residues in each chain
for L_i in Ls[:-1]:
idx_res[L_prev+L_i:] += 200
L_prev += L_i
chains = list("".join([ascii_uppercase[n]*L for n,L in enumerate(Ls)]))
feature_dict['residue_index'] = idx_res
plddts = []
paes = []
unrelaxed_pdb_lines = []
relaxed_pdb_lines = []
# Run the models.
if use_templates:
model_names = ["model_1","model_2","model_3","model_4","model_5"][:num_models]
model_start = ["model_1","model_3"]
model_end = ["model_2","model_5"]
else:
model_names = ["model_4","model_1","model_2","model_3","model_5"][:num_models]
model_start = ["model_4"]
model_end = ["model_5"]
for n, model_name in enumerate(model_names):
name = model_name+"_ptm" if use_ptm else model_name
model_config = config.model_config(name)
model_config.data.eval.num_ensemble = 1
if msa_mode == "single_sequence":
model_config.data.common.max_extra_msa = 1
model_config.data.eval.max_msa_clusters = 1
model_params = data.get_model_haiku_params(name, data_dir=".")
if model_name in model_start:
model_runner = model.RunModel(model_config, model_params)
processed_feature_dict = model_runner.process_features(feature_dict,random_seed=0)
else:
# swap params
for k in model_runner.params.keys():
model_runner.params[k] = model_params[k]
print(f"running model_{n+1}")
prediction_result = model_runner.predict(processed_feature_dict)
unrelaxed_protein = protein.from_prediction(processed_feature_dict,prediction_result)
unrelaxed_pdb_lines.append(protein.to_pdb(unrelaxed_protein))
plddts.append(prediction_result['plddt'])
if use_ptm:
paes.append(prediction_result['predicted_aligned_error'])
if use_amber:
# Relax the prediction.
amber_relaxer = relax.AmberRelaxation(max_iterations=0,tolerance=2.39,
stiffness=10.0,exclude_residues=[],
max_outer_iterations=20)
relaxed_pdb_str, _, _ = amber_relaxer.process(prot=unrelaxed_protein)
relaxed_pdb_lines.append(relaxed_pdb_str)
# Delete unused outputs to save memory.
if model_name in model_end:
del model_runner
del processed_feature_dict
del model_params
del prediction_result
# rerank models based on predicted lddt
lddt_rank = np.mean(plddts,-1).argsort()[::-1]
out = {}
print("reranking models based on avg. predicted lDDT")
for n,r in enumerate(lddt_rank):
print(f"model_{n+1} {np.mean(plddts[r])}")
unrelaxed_pdb_path = f'{prefix}_unrelaxed_model_{n+1}.pdb'
with open(unrelaxed_pdb_path, 'w') as f: f.write(unrelaxed_pdb_lines[r])
set_bfactor(unrelaxed_pdb_path, plddts[r], idx_res, chains)
if use_amber:
relaxed_pdb_path = f'{prefix}_relaxed_model_{n+1}.pdb'
with open(relaxed_pdb_path, 'w') as f: f.write(relaxed_pdb_lines[r])
set_bfactor(relaxed_pdb_path, plddts[r], idx_res, chains)
if use_ptm:
out[f"model_{n+1}"] = {"plddt":plddts[r], "pae":paes[r]}
else:
out[f"model_{n+1}"] = {"plddt":plddts[r]}
return out
#@title Call MMseqs2 to get MSA/templates
%%bash -s $use_amber $use_msa $use_templates $jobname $use_env
USE_AMBER=$1
USE_MSA=$2
USE_TEMPLATES=$3
NAME=$4
USE_ENV=$5
if [ ${USE_MSA} == "True" ] || [ ${USE_TEMPLATES} == "True" ]; then
if [ ! -f ${NAME}.mmseqs2.tar.gz ]; then
# query MMseqs2 webserver
echo "submitting job"
MODE=all
if [ ${USE_ENV} == "True" ]; then
MODE=env
fi
ID=$(curl -s -F q=@${NAME}.fasta -F mode=${MODE} https://a3m.mmseqs.com/ticket/msa | jq -r '.id')
STATUS=$(curl -s https://a3m.mmseqs.com/ticket/${ID} | jq -r '.status')
while [ "${STATUS}" == "RUNNING" ] || [ "${STATUS}" == "PENDING" ]; do
STATUS=$(curl -s https://a3m.mmseqs.com/ticket/${ID} | jq -r '.status')
sleep 1
done
if [ "${STATUS}" == "COMPLETE" ]; then
curl -s https://a3m.mmseqs.com/result/download/${ID} > ${NAME}.mmseqs2.tar.gz
tar xzf ${NAME}.mmseqs2.tar.gz
if [ ${USE_ENV} == "True" ]; then
cat uniref.a3m bfd.mgnify30.metaeuk30.smag30.a3m > tmp.a3m
tr -d '\000' < tmp.a3m > ${NAME}.a3m
rm uniref.a3m bfd.mgnify30.metaeuk30.smag30.a3m tmp.a3m
else
tr -d '\000' < uniref.a3m > ${NAME}.a3m
rm uniref.a3m
fi
mv pdb70.m8 ${NAME}.m8
else
echo "MMseqs2 server did not return a valid result."
cp ${NAME}.fasta ${NAME}.a3m
fi
fi
if [ ${USE_MSA} == "True" ]; then
echo "Found $(grep -c ">" ${NAME}.a3m) sequences (after redundacy filtering)"
fi
if [ ${USE_TEMPLATES} == "True" ] && [ ! -f ${NAME}_hhm.ffindex ]; then
echo "getting templates"
if [ -s ${NAME}.m8 ]; then
if [ ! -d templates ]; then
mkdir templates/
fi
printf "pdb\tevalue\n"
head -n 20 ${NAME}.m8 | awk '{print $2"\t"$11}'
TMPL=$(head -n 20 ${NAME}.m8 | awk '{printf $2","}')
curl -s https://a3m-templates.mmseqs.com/template/${TMPL} | tar xzf - -C templates/
mv templates/pdb70_a3m.ffdata ${NAME}_a3m.ffdata
mv templates/pdb70_a3m.ffindex ${NAME}_a3m.ffindex
mv templates/pdb70_hhm.ffdata ${NAME}_hhm.ffdata
mv templates/pdb70_hhm.ffindex ${NAME}_hhm.ffindex
cp ${NAME}_a3m.ffindex ${NAME}_cs219.ffindex
touch ${NAME}_cs219.ffdata
else
echo "no templates found"
fi
fi
fi
#@title Gather input features, predict structure
# parse TEMPLATES
if use_templates and os.path.isfile(f"{jobname}_hhm.ffindex"):
template_features = mk_template(jobname)
else:
use_templates = False
template_features = {}
# parse MSA
a3m_lines = "".join(open(a3m_file,"r").readlines())
msa, deletion_matrix = pipeline.parsers.parse_a3m(a3m_lines)
if homooligomer == 1:
msas = [msa]
deletion_matrices = [deletion_matrix]
else:
# make multiple copies of msa for each copy
# AAA------
# ---AAA---
# ------AAA
#
# note: if you concat the sequences (as below), it does NOT work
# AAAAAAAAA
msas = []
deletion_matrices = []
Ln = len(query_sequence)
for o in range(homooligomer):
L = Ln * o
R = Ln * (homooligomer-(o+1))
msas.append(["-"*L+seq+"-"*R for seq in msa])
deletion_matrices.append([[0]*L+mtx+[0]*R for mtx in deletion_matrix])
# gather features
feature_dict = {
**pipeline.make_sequence_features(sequence=query_sequence*homooligomer,
description="none",
num_res=len(query_sequence)*homooligomer),
**pipeline.make_msa_features(msas=msas,deletion_matrices=deletion_matrices),
**template_features
}
outs = predict_structure(jobname, feature_dict, Ls=[len(query_sequence)]*homooligomer)
#@title Make plots
dpi = 100 #@param {type:"integer"}
# gather MSA info
deduped_full_msa = list(dict.fromkeys(msa))
msa_arr = np.array([list(seq) for seq in deduped_full_msa])
seqid = (np.array(list(query_sequence)) == msa_arr).mean(-1)
seqid_sort = seqid.argsort() #[::-1]
non_gaps = (msa_arr != "-").astype(float)
non_gaps[non_gaps == 0] = np.nan
##################################################################
plt.figure(figsize=(14,4),dpi=dpi)
##################################################################
plt.subplot(1,2,1); plt.title("Sequence coverage")
plt.imshow(non_gaps[seqid_sort]*seqid[seqid_sort,None],
interpolation='nearest', aspect='auto',
cmap="rainbow_r", vmin=0, vmax=1, origin='lower')
plt.plot((msa_arr != "-").sum(0), color='black')
plt.xlim(-0.5,msa_arr.shape[1]-0.5)
plt.ylim(-0.5,msa_arr.shape[0]-0.5)
plt.colorbar(label="Sequence identity to query",)
plt.xlabel("Positions")
plt.ylabel("Sequences")
##################################################################
plt.subplot(1,2,2); plt.title("Predicted lDDT per position")
for model_name,value in outs.items():
plt.plot(value["plddt"],label=model_name)
if homooligomer > 0:
for n in range(homooligomer+1):
x = n*(len(query_sequence)-1)
plt.plot([x,x],[0,100],color="black")
plt.legend()
plt.ylim(0,100)
plt.ylabel("Predicted lDDT")
plt.xlabel("Positions")
plt.savefig(jobname+"_coverage_lDDT.png")
##################################################################
plt.show()
if use_ptm:
print("Predicted Alignment Error")
##################################################################
plt.figure(figsize=(3*num_models,2), dpi=dpi)
for n,(model_name,value) in enumerate(outs.items()):
plt.subplot(1,num_models,n+1)
plt.title(model_name)
plt.imshow(value["pae"],label=model_name,cmap="bwr",vmin=0,vmax=30)
plt.colorbar()
plt.savefig(jobname+"_PAE.png")
plt.show()
##################################################################
#@title Display 3D structure {run: "auto"}
model_num = 1 #@param ["1", "2", "3", "4", "5"] {type:"raw"}
color = "chain" #@param ["chain", "lDDT", "rainbow"]
show_sidechains = False #@param {type:"boolean"}
show_mainchains = False #@param {type:"boolean"}
def plot_plddt_legend():
thresh = ['plDDT:','Very low (<50)','Low (60)','OK (70)','Confident (80)','Very high (>90)']
plt.figure(figsize=(1,0.1),dpi=100)
########################################
for c in ["#FFFFFF","#FF0000","#FFFF00","#00FF00","#00FFFF","#0000FF"]:
plt.bar(0, 0, color=c)
plt.legend(thresh, frameon=False,
loc='center', ncol=6,
handletextpad=1,
columnspacing=1,
markerscale=0.5,)
plt.axis(False)
return plt
def plot_confidence(model_num=1):
model_name = f"model_{model_num}"
"""Plots the legend for plDDT."""
#########################################
if use_ptm:
plt.figure(figsize=(10,3),dpi=100)
plt.subplot(1,2,1)
else:
plt.figure(figsize=(5,3),dpi=100)
plt.title('Predicted lDDT')
plt.plot(outs[model_name]["plddt"])
for n in range(homooligomer+1):
x = n*(len(query_sequence))
plt.plot([x,x],[0,100],color="black")
plt.ylabel('plDDT')
plt.xlabel('position')
#########################################
if use_ptm:
plt.subplot(1,2,2);plt.title('Predicted Aligned Error')
plt.imshow(outs[model_name]["pae"], cmap="bwr",vmin=0,vmax=30)
plt.colorbar()
plt.xlabel('Scored residue')
plt.ylabel('Aligned residue')
#########################################
return plt
def show_pdb(model_num=1, show_sidechains=False, show_mainchains=False, color="lDDT"):
model_name = f"model_{model_num}"
if use_amber:
pdb_filename = f"{jobname}_relaxed_{model_name}.pdb"
else:
pdb_filename = f"{jobname}_unrelaxed_{model_name}.pdb"
view = py3Dmol.view(js='https://3dmol.org/build/3Dmol.js',)
view.addModel(open(pdb_filename,'r').read(),'pdb')
if color == "lDDT":
view.setStyle({'cartoon': {'colorscheme': {'prop':'b','gradient': 'roygb','min':50,'max':90}}})
elif color == "rainbow":
view.setStyle({'cartoon': {'color':'spectrum'}})
elif color == "chain":
for n,chain,color in zip(range(homooligomer),list("ABCDEFGH"),
["lime","cyan","magenta","yellow","salmon","white","blue","orange"]):
view.setStyle({'chain':chain},{'cartoon': {'color':color}})
if show_sidechains:
BB = ['C','O','N']
view.addStyle({'and':[{'resn':["GLY","PRO"],'invert':True},{'atom':BB,'invert':True}]},
{'stick':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
view.addStyle({'and':[{'resn':"GLY"},{'atom':'CA'}]},
{'sphere':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
view.addStyle({'and':[{'resn':"PRO"},{'atom':['C','O'],'invert':True}]},
{'stick':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
if show_mainchains:
BB = ['C','O','N','CA']
view.addStyle({'atom':BB},{'stick':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
view.zoomTo()
return view
if (model_num-1) < num_models:
show_pdb(model_num,show_sidechains, show_mainchains, color).show()
if color == "lDDT": plot_plddt_legend().show()
plot_confidence(model_num).show()
else:
print("this model was not made")
#@title Package and download results
#@markdown If you having issues downloading the result archive, try disabling your adblocker and run this cell again. If that fails click on the little folder icon to the left, navigate to file: `jobname.result.zip`, right-click and select \"Download\" (see [screenshot](https://pbs.twimg.com/media/E6wRW2lWUAEOuoe?format=jpg&name=small)).
with open(f"{jobname}.log", "w") as text_file:
text_file.write(f"num_models={num_models}\n")
text_file.write(f"use_amber={use_amber}\n")
text_file.write(f"use_msa={use_msa}\n")
text_file.write(f"msa_mode={msa_mode}\n")
text_file.write(f"use_templates={use_templates}\n")
text_file.write(f"homooligomer={homooligomer}\n")
text_file.write(f"use_ptm={use_ptm}\n")
citations = {
"Ovchinnikov2021": """@software{Ovchinnikov2021,
author = {Ovchinnikov, Sergey and Steinegger, Martin and Mirdita, Milot},
title = {{ColabFold - Making Protein folding accessible to all via Google Colab}},
year = {2021},
publisher = {Zenodo},
version = {v1.0-alpha},
doi = {10.5281/zenodo.5123297},
url = {https://doi.org/10.5281/zenodo.5123297},
comment = {The AlphaFold notebook}
}""",
"LevyKarin2020": """@article{LevyKarin2020,
author = {{Levy Karin}, Eli and Mirdita, Milot and S{\"{o}}ding, Johannes},
doi = {10.1186/s40168-020-00808-x},
journal = {Microbiome},
number = {1},
title = {{MetaEuk—sensitive, high-throughput gene discovery, and annotation for large-scale eukaryotic metagenomics}},
volume = {8},
year = {2020},
comment = {MetaEuk database}
}""",
"Delmont2020": """@article{Delmont2020,
author = {Delmont, Tom O. and Gaia, Morgan and Hinsinger, Damien D. and Fremont, Paul and Guerra, Antonio Fernandez and Eren, A. Murat and Vanni, Chiara and Kourlaiev, Artem and D'Agata, Leo and Clayssen, Quentin and Villar, Emilie and Labadie, Karine and Cruaud, Corinne and Poulain, Julie and da Silva, Corinne and Wessner, Marc and Noel, Benjamin and Aury, Jean Marc and de Vargas, Colomban and Bowler, Chris and Karsenti, Eric and Pelletier, Eric and Wincker, Patrick and Jaillon, Olivier and Sunagawa, Shinichi and Acinas, Silvia G. and Bork, Peer and Karsenti, Eric and Bowler, Chris and Sardet, Christian and Stemmann, Lars and de Vargas, Colomban and Wincker, Patrick and Lescot, Magali and Babin, Marcel and Gorsky, Gabriel and Grimsley, Nigel and Guidi, Lionel and Hingamp, Pascal and Jaillon, Olivier and Kandels, Stefanie and Iudicone, Daniele and Ogata, Hiroyuki and Pesant, St{\'{e}}phane and Sullivan, Matthew B. and Not, Fabrice and Karp-Boss, Lee and Boss, Emmanuel and Cochrane, Guy and Follows, Michael and Poulton, Nicole and Raes, Jeroen and Sieracki, Mike and Speich, Sabrina},
journal = {bioRxiv},
title = {{Functional repertoire convergence of distantly related eukaryotic plankton lineages revealed by genome-resolved metagenomics}},
year = {2020},
comment = {SMAG database}
}""",
"Mitchell2019": """@article{Mitchell2019,
author = {Mitchell, Alex L and Almeida, Alexandre and Beracochea, Martin and Boland, Miguel and Burgin, Josephine and Cochrane, Guy and Crusoe, Michael R and Kale, Varsha and Potter, Simon C and Richardson, Lorna J and Sakharova, Ekaterina and Scheremetjew, Maxim and Korobeynikov, Anton and Shlemov, Alex and Kunyavskaya, Olga and Lapidus, Alla and Finn, Robert D},
doi = {10.1093/nar/gkz1035},
journal = {Nucleic Acids Res.},
title = {{MGnify: the microbiome analysis resource in 2020}},
year = {2019},
comment = {MGnify database}
}""",
"Eastman2017": """@article{Eastman2017,
author = {Eastman, Peter and Swails, Jason and Chodera, John D. and McGibbon, Robert T. and Zhao, Yutong and Beauchamp, Kyle A. and Wang, Lee-Ping and Simmonett, Andrew C. and Harrigan, Matthew P. and Stern, Chaya D. and Wiewiora, Rafal P. and Brooks, Bernard R. and Pande, Vijay S.},
doi = {10.1371/journal.pcbi.1005659},
journal = {PLOS Comput. Biol.},
number = {7},
title = {{OpenMM 7: Rapid development of high performance algorithms for molecular dynamics}},
volume = {13},
year = {2017},
comment = {Amber relaxation}
}""",
"Jumper2021": """@article{Jumper2021,
author = {Jumper, John and Evans, Richard and Pritzel, Alexander and Green, Tim and Figurnov, Michael and Ronneberger, Olaf and Tunyasuvunakool, Kathryn and Bates, Russ and {\v{Z}}{\'{i}}dek, Augustin and Potapenko, Anna and Bridgland, Alex and Meyer, Clemens and Kohl, Simon A. A. and Ballard, Andrew J. and Cowie, Andrew and Romera-Paredes, Bernardino and Nikolov, Stanislav and Jain, Rishub and Adler, Jonas and Back, Trevor and Petersen, Stig and Reiman, David and Clancy, Ellen and Zielinski, Michal and Steinegger, Martin and Pacholska, Michalina and Berghammer, Tamas and Bodenstein, Sebastian and Silver, David and Vinyals, Oriol and Senior, Andrew W. and Kavukcuoglu, Koray and Kohli, Pushmeet and Hassabis, Demis},
doi = {10.1038/s41586-021-03819-2},
journal = {Nature},
pmid = {34265844},
title = {{Highly accurate protein structure prediction with AlphaFold.}},
year = {2021},
comment = {AlphaFold2 + BFD Database}
}""",
"Mirdita2019": """@article{Mirdita2019,
author = {Mirdita, Milot and Steinegger, Martin and S{\"{o}}ding, Johannes},
doi = {10.1093/bioinformatics/bty1057},
journal = {Bioinformatics},
number = {16},
pages = {2856--2858},
pmid = {30615063},
title = {{MMseqs2 desktop and local web server app for fast, interactive sequence searches}},
volume = {35},
year = {2019},
comment = {MMseqs2 search server}
}""",
"Steinegger2019": """@article{Steinegger2019,
author = {Steinegger, Martin and Meier, Markus and Mirdita, Milot and V{\"{o}}hringer, Harald and Haunsberger, Stephan J. and S{\"{o}}ding, Johannes},
doi = {10.1186/s12859-019-3019-7},
journal = {BMC Bioinform.},
number = {1},
pages = {473},
pmid = {31521110},
title = {{HH-suite3 for fast remote homology detection and deep protein annotation}},
volume = {20},
year = {2019},
comment = {PDB70 database}
}""",
"Mirdita2017": """@article{Mirdita2017,
author = {Mirdita, Milot and von den Driesch, Lars and Galiez, Clovis and Martin, Maria J. and S{\"{o}}ding, Johannes and Steinegger, Martin},
doi = {10.1093/nar/gkw1081},
journal = {Nucleic Acids Res.},
number = {D1},
pages = {D170--D176},
pmid = {27899574},
title = {{Uniclust databases of clustered and deeply annotated protein sequences and alignments}},
volume = {45},
year = {2017},
comment = {Uniclust30/UniRef30 database},
}""",
"Berman2003": """@misc{Berman2003,
author = {Berman, Helen and Henrick, Kim and Nakamura, Haruki},
booktitle = {Nat. Struct. Biol.},
doi = {10.1038/nsb1203-980},
number = {12},
pages = {980},
pmid = {14634627},
title = {{Announcing the worldwide Protein Data Bank}},
volume = {10},
year = {2003},
comment = {templates downloaded from wwPDB server}
}""",
}
to_cite = [ "Jumper2021", "Ovchinnikov2021" ]
if use_msa: to_cite += ["Mirdita2019"]
if use_msa: to_cite += ["Mirdita2017"]
if use_env: to_cite += ["Mitchell2019"]
if use_env: to_cite += ["Delmont2020"]
if use_env: to_cite += ["LevyKarin2020"]
if use_templates: to_cite += ["Steinegger2019"]
if use_templates: to_cite += ["Berman2003"]
if use_amber: to_cite += ["Eastman2017"]
with open(f"{jobname}.bibtex", 'w') as writer:
for i in to_cite:
writer.write(citations[i])
writer.write("\n")
print(f"Found {len(to_cite)} citation{'s' if len(to_cite) > 1 else ''} for tools or databases.")
if use_custom_msa:
print("Don't forget to cite your custom MSA generation method.")
!zip -FSr $jobname".result.zip" $jobname".log" $a3m_file $jobname"_"*"relaxed_model_"*".pdb" $jobname".bibtex" $jobname"_"*".png"
files.download(f"{jobname}.result.zip")
if save_to_google_drive == True and drive != None:
uploaded = drive.CreateFile({'title': f"{jobname}.result.zip"})
uploaded.SetContentFile(f"{jobname}.result.zip")
uploaded.Upload()
print(f"Uploaded {jobname}.result.zip to Google Drive with ID {uploaded.get('id')}")
```
# Instructions <a name="Instructions"></a>
**Quick start**
1. Change the runtime type to GPU at "Runtime" -> "Change runtime type" (improves speed).
2. Paste your protein sequence in the input field below.
3. Press "Runtime" -> "Run all".
4. The pipeline consists of 10 steps. The currently running steps is indicated by a circle with a stop sign next to it.
**Result zip file contents**
1. PDB formatted structures sorted by avg. pIDDT. (relaxed, unrelaxed).
2. Plots of the model quality.
3. Plots of the MSA coverage.
4. Parameter log file.
5. A3M formatted input MSA.
6. BibTeX file with citations for all used tools and databases.
At the end of the job a download modal box will pop up with a `jobname.result.zip` file. Additionally, if the `save_to_google_drive` option was selected, the `jobname.result.zip` will be uploaded to your Google Drive.
**Using a custom MSA as input**
To predict the structure with a custom MSA (A3M formatted): (1) Change the msa_mode: to "custom", (2) Wait for an upload box to appear at the end of the "Input Protein ..." box. Upload your A3M. The first fasta entry of the A3M must be the query sequence without gaps.
To generate good input MSAs the HHblits server can be used here: https://toolkit.tuebingen.mpg.de/tools/hhblits
After submitting your query, click "Query Template MSA" -> "Download Full A3M". Download the a3m file and upload it to the notebook.
**Troubleshooting**
* Try to restart the session "Runtime" -> "Factory reset runtime".
* Check your input sequence.
**Known issues**
* Colab assigns different types of GPUs with varying amount of memory. Some might have not enough memory to predict the structure.
* Your browser can block the pop-up for downloading the result file. You can choose the `save_to_google_drive` option to upload to Google Drive instead or manually download the result file: Click on the little folder icon to the left, navigate to file: `jobname.result.zip`, right-click and select \"Download\" (see [screenshot](https://pbs.twimg.com/media/E6wRW2lWUAEOuoe?format=jpg&name=small)).
**Limitations**
* MSAs: MMseqs2 is very precise and sensitive but might find less hits compared to HHblits/HMMer searched against BFD or Mgnify.
* Computing resources: Our MMseqs2 API can probably handle ~20k requests per day.
* For best results, we recommend using the full pipeline: https://github.com/deepmind/alphafold
**Description of the plots**
* **Number of sequences per position** - We want to see at least 30 sequences per position, for best performance, ideally 100 sequences.
* **Predicted lDDT per position (pLDDT)** - model confidence (out of 100) at each position. Higher the better.
* **Predicted Alignment Error (PAE)** - For homooligomers, this could be a useful metric to assess how confident the model is about the interface. Lower the better.
**Bugs**
- If you encounter any bugs, please report the issue to https://github.com/sokrypton/ColabFold/issues
**Q&A**
- *What is `use_ptm`?* Deepmind finetuned their original 5 trained model params to return PAE and predicted TMscore. We use these models by default to generate the PAE plots. But sometimes using the original models, without finetunning gives different results. Disabling this option maybe useful if you want to reproduce CASP results, or want to get more diversity in the predictions.
**Acknowledgments**
- We would like to thank the AlphaFold team for developing an excellent model and open sourcing the software.
- A colab by Sergey Ovchinnikov ([@sokrypton](https://twitter.com/sokrypton)), Milot Mirdita ([@milot_mirdita](https://twitter.com/milot_mirdita)) and Martin Steinegger ([@thesteinegger](https://twitter.com/thesteinegger)).
- Minkyung Baek ([@minkbaek](https://twitter.com/minkbaek)) and Yoshitaka Moriwaki ([@Ag_smith](https://twitter.com/Ag_smith)) for protein-complex prediction proof-of-concept in AlphaFold2.
- Also, credit to [David Koes](https://github.com/dkoes) for his awesome [py3Dmol](https://3dmol.csb.pitt.edu/) plugin, without whom these notebooks would be quite boring!
- For related notebooks see: [ColabFold](https://github.com/sokrypton/ColabFold)
| github_jupyter |
# Simulating gate noise on the Rigetti Quantum Virtual Machine
© Copyright 2017, Rigetti Computing.
$$
\newcommand{ket}[1]{\left|{#1}\right\rangle}
\newcommand{bra}[1]{\left\langle {#1}\right|}
\newcommand{tr}{\mathrm{Tr}}
$$
## Pure states vs. mixed states
Errors in quantum computing can introduce classical uncertainty in what the underlying state is.
When this happens we sometimes need to consider not only wavefunctions but also probabilistic sums of
wavefunctions when we are uncertain as to which one we have. For example, if we think that an X gate
was accidentally applied to a qubit with a 50-50 chance then we would say that there is a 50% chance
we have the $\ket{0}$ state and a 50% chance that we have a $\ket{1}$ state.
This is called an "impure" or
"mixed"state in that it isn't just a wavefunction (which is pure) but instead a distribution over
wavefunctions. We describe this with something called a density matrix, which is generally an
operator. Pure states have very simple density matrices that we can write as an outer product of a
ket vector $\ket{\psi}$ with its own bra version $\bra{\psi}=\ket{\psi}^\dagger$.
For a pure state the density matrix is simply
$$
\rho_\psi = \ket{\psi}\bra{\psi}.
$$
The expectation value of an operator for a mixed state is given by
$$
\langle X \rangle_\rho = \tr{X \rho}
$$
where $\tr{A}$ is the trace of an operator, which is the sum of its diagonal elements
which is independent of choice of basis.
Pure state density matrices satisfy
$$
\rho \text{ is pure } \Leftrightarrow \rho^2 = \rho
$$
which you can easily verify for $\rho_\psi$ assuming that the state is normalized.
If we want to describe a situation with classical uncertainty between states $\rho_1$ and
$\rho_2$, then we can take their weighted sum
$$
\rho = p \rho_1 + (1-p) \rho_2
$$
where $p\in [0,1]$ gives the classical probability that the state is $\rho_1$.
Note that classical uncertainty in the wavefunction is markedly different from superpositions.
We can represent superpositions using wavefunctions, but use density matrices to describe
distributions over wavefunctions. You can read more about density matrices [here](https://en.wikipedia.org/wiki/Density_matrix).
# Quantum gate errors
## What are they?
For a quantum gate given by its unitary operator $U$, a "quantum gate error" describes the scenario in which the actually induces transformation deviates from $\ket{\psi} \mapsto U\ket{\psi}$.
There are two basic types of quantum gate errors:
1. **coherent errors** are those that preserve the purity of the input state, i.e., instead of the above mapping we carry out a perturbed, but unitary operation $\ket{\psi} \mapsto \tilde{U}\ket{\psi}$, where $\tilde{U} \neq U$.
2. **incoherent errors** are those that do not preserve the purity of the input state,
in this case we must actually represent the evolution in terms of density matrices.
The state $\rho := \ket{\psi}\bra{\psi}$ is then mapped as
$$
\rho \mapsto \sum_{j=1}^n K_j\rho K_j^\dagger,
$$
where the operators $\{K_1, K_2, \dots, K_m\}$ are called Kraus operators and must obey
$\sum_{j=1}^m K_j^\dagger K_j = I$ to conserve the trace of $\rho$.
Maps expressed in the above form are called Kraus maps. It can be shown that every physical map on a finite
dimensional quantum system can be represented as a Kraus map, though this representation is not generally unique.
[You can find more information about quantum operations here](https://en.wikipedia.org/wiki/Quantum_operation#Kraus_operators)
In a way, coherent errors are *in principle* amendable by more precisely calibrated control. Incoherent errors are more tricky.
## Why do incoherent errors happen?
When a quantum system (e.g., the qubits on a quantum processor) is not perfectly isolated from its environment it generally co-evolves with the degrees of freedom it couples to. The implication is that while the total time evolution of system and environment can be assumed to be unitary, restriction to the system state generally is not.
**Let's throw some math at this for clarity:**
Let our total Hilbert space be given by the tensor product of system and environment Hilbert spaces:
$\mathcal{H} = \mathcal{H}_S \otimes \mathcal{H}_E$.
Our system "not being perfectly isolated" must be translated to the statement that the global Hamiltonian contains a contribution that couples the system and environment:
$$
H = H_S \otimes I + I \otimes H_E + V
$$
where $V$ non-trivally acts on both the system and the environment.
Consequently, even if we started in an initial state that factorized over system and environment $\ket{\psi}_{S,0}\otimes \ket{\psi}_{E,0}$
if everything evolves by the Schrödinger equation
$$
\ket{\psi_t} = e^{-i \frac{Ht}{\hbar}} \left(\ket{\psi}_{S,0}\otimes \ket{\psi}_{E,0}\right)
$$
the final state will generally not admit such a factorization.
## A toy model
**In this (somewhat technical) section we show how environment interaction can corrupt an identity gate and derive its Kraus map.**
For simplicity, let us assume that we are in a reference frame in which both the system and environment Hamiltonian's vanish $H_S = 0, H_E = 0$ and where the cross-coupling is small even when multiplied by the duration of the time evolution $\|\frac{tV}{\hbar}\|^2 \sim \epsilon \ll 1$ (any operator norm $\|\cdot\|$ will do here).
Let us further assume that $V = \sqrt{\epsilon} V_S \otimes V_E$ (the more general case is given by a sum of such terms) and that
the initial environment state satisfies $\bra{\psi}_{E,0} V_E\ket{\psi}_{E,0} = 0$. This turns out to be a very reasonable assumption in practice but a more thorough discussion exceeds our scope.
Then the joint system + environment state $\rho = \rho_{S,0} \otimes \rho_{E,0}$ (now written as a density matrix) evolves as
$$
\rho \mapsto \rho' := e^{-i \frac{Vt}{\hbar}} \rho e^{+i \frac{Vt}{\hbar}}
$$
Using the Baker-Campbell-Hausdorff theorem we can expand this to second order in $\epsilon$
$$
\rho' = \rho - \frac{it}{\hbar} [V, \rho] - \frac{t^2}{2\hbar^2} [V, [V, \rho]] + O(\epsilon^{3/2})
$$
We can insert the initially factorizable state $\rho = \rho_{S,0} \otimes \rho_{E,0}$ and trace over the environmental degrees of freedom to obtain
\begin{align}
\rho_S' := \tr_E \rho' & = \rho_{S,0} \underbrace{\tr \rho_{E,0}}_{1} - \frac{i\sqrt{\epsilon} t}{\hbar} \underbrace{\left[ V_S \rho_{S,0} \underbrace{\tr V_E\rho_{E,0}}_{\bra{\psi}_{E,0} V_E\ket{\psi}_{E,0} = 0} - \rho_{S,0}V_S \underbrace{\tr \rho_{E,0}V_E}_{\bra{\psi}_{E,0} V_E\ket{\psi}_{E,0} = 0} \right]}_0 \\
& \qquad - \frac{\epsilon t^2}{2\hbar^2} \left[ V_S^2\rho_{S,0}\tr V_E^2 \rho_{E,0} + \rho_{S,0} V_S^2 \tr \rho_{E,0}V_E^2 - 2 V_S\rho_{S,0}V_S\tr V_E \rho_{E,0}V_E\right] \\
& = \rho_{S,0} - \frac{\gamma}{2} \left[ V_S^2\rho_{S,0} + \rho_{S,0} V_S^2 - 2 V_S\rho_{S,0}V_S\right]
\end{align}
where the coefficient in front of the second part is by our initial assumption very small $\gamma := \frac{\epsilon t^2}{2\hbar^2}\tr V_E^2 \rho_{E,0} \ll 1$.
This evolution happens to be approximately equal to a Kraus map with operators $K_1 := I - \frac{\gamma}{2} V_S^2, K_2:= \sqrt{\gamma} V_S$:
\begin{align}
\rho_S \to \rho_S' &= K_1\rho K_1^\dagger + K_2\rho K_2^\dagger
= \rho - \frac{\gamma}{2}\left[ V_S^2 \rho + \rho V_S^2\right] + \gamma V_S\rho_S V_S + O(\gamma^2)
\end{align}
This agrees to $O(\epsilon^{3/2})$ with the result of our derivation above. This type of derivation can be extended to many other cases with little complication and a very similar argument is used to derive the [Lindblad master equation](https://en.wikipedia.org/wiki/Lindblad_equation).
# Support for noisy gates on the Rigetti QVM
As of today, users of our Forest API can annotate their QUIL programs by certain pragma statements that inform the QVM that a particular gate on specific target qubits should be replaced by an imperfect realization given by a Kraus map.
## But the QVM propagates *pure states*: How does it simulate noisy gates?
It does so by yielding the correct outcomes **in the average over many executions of the QUIL program**:
When the noisy version of a gate should be applied the QVM makes a random choice which Kraus operator is applied to the current state with a probability that ensures that the average over many executions is equivalent to the Kraus map.
In particular, a particular Kraus operator $K_j$ is applied to $\ket{\psi}_S$
$$
\ket{\psi'}_S = \frac{1}{\sqrt{p_j}} K_j \ket{\psi}_S
$$
with probability $p_j:= \bra{\psi}_S K_j^\dagger K_j \ket{\psi}_S$.
In the average over many execution $N \gg 1$ we therefore find that
\begin{align}
\overline{\rho_S'} & = \frac{1}{N} \sum_{n=1}^N \ket{\psi'_n}_S\bra{\psi'_n}_S \\
& = \frac{1}{N} \sum_{n=1}^N p_{j_n}^{-1}K_{j_n}\ket{\psi'}_S \bra{\psi'}_SK_{j_n}^\dagger
\end{align}
where $j_n$ is the chosen Kraus operator label in the $n$-th trial.
This is clearly a Kraus map itself! And we can group identical terms and rewrite it as
\begin{align}
\overline{\rho_S'} & =
\sum_{\ell=1}^n \frac{N_\ell}{N} p_{\ell}^{-1}K_{\ell}\ket{\psi'}_S \bra{\psi'}_SK_{\ell}^\dagger
\end{align}
where $N_{\ell}$ is the number of times that Kraus operator label $\ell$ was selected.
For large enough $N$ we know that $N_{\ell} \approx N p_\ell$ and therefore
\begin{align}
\overline{\rho_S'} \approx \sum_{\ell=1}^n K_{\ell}\ket{\psi'}_S \bra{\psi'}_SK_{\ell}^\dagger
\end{align}
which proves our claim.
**The consequence is that noisy gate simulations must generally be repeated many times to obtain representative results**.
## How do I get started?
1. Come up with a good model for your noise. We will provide some examples below and may add more such
examples to our public repositories over time. Alternatively, you can characterize the gate under
consideration using [Quantum Process Tomography](https://arxiv.org/abs/1202.5344) or
[Gate Set Tomography](http://www.pygsti.info/) and use the resulting process matrices to obtain a
very accurate noise model for a particular QPU.
2. Define your Kraus operators as a list of numpy arrays `kraus_ops = [K1, K2, ..., Km]`.
3. For your QUIL program `p`, call:
```
p.define_noisy_gate("MY_NOISY_GATE", [q1, q2], kraus_ops)
```
where you should replace `MY_NOISY_GATE` with the gate of interest and `q1, q2` the indices of the qubits.
**Scroll down for some examples!**
```
from __future__ import print_function, division
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import binom
import matplotlib.colors as colors
%matplotlib inline
from pyquil.quil import Program, MEASURE
from pyquil.api.qvm import QVMConnection
from pyquil.job_results import wait_for_job
from pyquil.gates import CZ, H, I, X
from scipy.linalg import expm
cxn = QVMConnection()
```
# Example 1: Amplitude damping
Amplitude damping channels are imperfect identity maps with Kraus operators
$$
K_1 = \begin{pmatrix}
1 & 0 \\
0 & \sqrt{1-p}
\end{pmatrix} \\
K_2 = \begin{pmatrix}
0 & \sqrt{p} \\
0 & 0
\end{pmatrix}
$$
where $p$ is the probability that a qubit in the $\ket{1}$ state decays to the $\ket{0}$ state.
```
def damping_channel(damp_prob=.1):
"""
Generate the Kraus operators corresponding to an amplitude damping
noise channel.
:params float damp_prob: The one-step damping probability.
:return: A list [k1, k2] of the Kraus operators that parametrize the map.
:rtype: list
"""
damping_op = np.sqrt(damp_prob) * np.array([[0, 1],
[0, 0]])
residual_kraus = np.diag([1, np.sqrt(1-damp_prob)])
return [residual_kraus, damping_op]
def append_kraus_to_gate(kraus_ops, g):
"""
Follow a gate `g` by a Kraus map described by `kraus_ops`.
:param list kraus_ops: The Kraus operators.
:param numpy.ndarray g: The unitary gate.
:return: A list of transformed Kraus operators.
"""
return [kj.dot(g) for kj in kraus_ops]
def append_damping_to_gate(gate, damp_prob=.1):
"""
Generate the Kraus operators corresponding to a given unitary
single qubit gate followed by an amplitude damping noise channel.
:params np.ndarray|list gate: The 2x2 unitary gate matrix.
:params float damp_prob: The one-step damping probability.
:return: A list [k1, k2] of the Kraus operators that parametrize the map.
:rtype: list
"""
return append_kraus_to_gate(damping_channel(damp_prob), gate)
%%time
# single step damping probability
damping_per_I = 0.02
# number of program executions
trials = 200
results_damping = []
lengths = np.arange(0, 201, 10, dtype=int)
for jj, num_I in enumerate(lengths):
print("{}/{}, ".format(jj, len(lengths)), end="")
p = Program(X(0))
# want increasing number of I-gates
p.inst([I(0) for _ in range(num_I)])
p.inst(MEASURE(0, [0]))
# overload identity I on qc 0
p.define_noisy_gate("I", [0], append_damping_to_gate(np.eye(2), damping_per_I))
cxn.random_seed = int(num_I)
res = cxn.run(p, [0], trials=trials)
results_damping.append([np.mean(res), np.std(res) / np.sqrt(trials)])
results_damping = np.array(results_damping)
dense_lengths = np.arange(0, lengths.max()+1, .2)
survival_probs = (1-damping_per_I)**dense_lengths
logpmf = binom.logpmf(np.arange(trials+1)[np.newaxis, :], trials, survival_probs[:, np.newaxis])/np.log(10)
DARK_TEAL = '#48737F'
FUSCHIA = "#D6619E"
BEIGE = '#EAE8C6'
cm = colors.LinearSegmentedColormap.from_list('anglemap', ["white", FUSCHIA, BEIGE], N=256, gamma=1.5)
plt.figure(figsize=(14, 6))
plt.pcolor(dense_lengths, np.arange(trials+1)/trials, logpmf.T, cmap=cm, vmin=-4, vmax=logpmf.max())
plt.plot(dense_lengths, survival_probs, c=BEIGE, label="Expected mean")
plt.errorbar(lengths, results_damping[:,0], yerr=2*results_damping[:,1], c=DARK_TEAL,
label=r"noisy qvm, errorbars $ = \pm 2\hat{\sigma}$", marker="o")
cb = plt.colorbar()
cb.set_label(r"$\log_{10} \mathrm{Pr}(n_1; n_{\rm trials}, p_{\rm survival}(t))$", size=20)
plt.title("Amplitude damping model of a single qubit", size=20)
plt.xlabel(r"Time $t$ [arb. units]", size=14)
plt.ylabel(r"$n_1/n_{\rm trials}$", size=14)
plt.legend(loc="best", fontsize=18)
plt.xlim(*lengths[[0, -1]])
plt.ylim(0, 1)
```
# Example 2: dephased CZ-gate
Dephasing is usually characterized through a qubit's $T_2$ time.
For a single qubit the dephasing Kraus operators are
$$
K_1(p) = \sqrt{1-p} I_2 \\
K_2(p) = \sqrt{p} \sigma_Z
$$
where $p = 1 - \exp(-T_2/T_{\rm gate})$ is the probability that the qubit is dephased over the time interval of interest, $I_2$ is the $2\times 2$-identity matrix and $\sigma_Z$ is the Pauli-Z operator.
For two qubits, we must construct a Kraus map that has *four* different outcomes:
1. No dephasing
2. Qubit 1 dephases
3. Qubit 2 dephases
4. Both dephase
The Kraus operators for this are given by
\begin{align}
K'_1(p,q) = K_1(p)\otimes K_1(q) \\
K'_2(p,q) = K_2(p)\otimes K_1(q) \\
K'_3(p,q) = K_1(p)\otimes K_2(q) \\
K'_4(p,q) = K_2(p)\otimes K_2(q)
\end{align}
where we assumed a dephasing probability $p$ for the first qubit and $q$ for the second.
Dephasing is a *diagonal* error channel and the CZ gate is also diagonal, therefore we can get the combined map of dephasing and the CZ gate simply by composing $U_{\rm CZ}$ the unitary representation of CZ with each Kraus operator
\begin{align}
K^{\rm CZ}_1(p,q) = K_1(p)\otimes K_1(q)U_{\rm CZ} \\
K^{\rm CZ}_2(p,q) = K_2(p)\otimes K_1(q)U_{\rm CZ} \\
K^{\rm CZ}_3(p,q) = K_1(p)\otimes K_2(q)U_{\rm CZ} \\
K^{\rm CZ}_4(p,q) = K_2(p)\otimes K_2(q)U_{\rm CZ}
\end{align}
**Note that this is not always accurate, because a CZ gate is often achieved through non-diagonal interaction Hamiltonians! However, for sufficiently small dephasing probabilities it should always provide a good starting point.**
```
def dephasing_kraus_map(p=.1):
"""
Generate the Kraus operators corresponding to a dephasing channel.
:params float p: The one-step dephasing probability.
:return: A list [k1, k2] of the Kraus operators that parametrize the map.
:rtype: list
"""
return [np.sqrt(1-p)*np.eye(2), np.sqrt(p)*np.diag([1, -1])]
def tensor_kraus_maps(k1, k2):
"""
Generate the Kraus map corresponding to the composition
of two maps on different qubits.
:param list k1: The Kraus operators for the first qubit.
:param list k2: The Kraus operators for the second qubit.
:return: A list of tensored Kraus operators.
"""
return [np.kron(k1j, k2l) for k1j in k1 for k2l in k2]
%%time
# single step damping probabilities
ps = np.linspace(.001, .5, 200)
# number of program executions
trials = 500
results = []
for jj, p in enumerate(ps):
corrupted_CZ = append_kraus_to_gate(
tensor_kraus_maps(
dephasing_kraus_map(p),
dephasing_kraus_map(p)
),
np.diag([1, 1, 1, -1]))
print("{}/{}, ".format(jj, len(ps)), end="")
# make Bell-state
p = Program(H(0), H(1), CZ(0,1), H(1))
p.inst(MEASURE(0, [0]))
p.inst(MEASURE(1, [1]))
# overload identity I on qc 0
p.define_noisy_gate("CZ", [0, 1], corrupted_CZ)
cxn.random_seed = jj
res = cxn.run(p, [0, 1], trials=trials)
results.append(res)
results = np.array(results)
Z1s = (2*results[:,:,0]-1.)
Z2s = (2*results[:,:,1]-1.)
Z1Z2s = Z1s * Z2s
Z1m = np.mean(Z1s, axis=1)
Z2m = np.mean(Z2s, axis=1)
Z1Z2m = np.mean(Z1Z2s, axis=1)
plt.figure(figsize=(14, 6))
plt.axhline(y=1.0, color=FUSCHIA, alpha=.5, label="Bell state")
plt.plot(ps, Z1Z2m, "x", c=FUSCHIA, label=r"$\overline{Z_1 Z_2}$")
plt.plot(ps, 1-2*ps, "--", c=FUSCHIA, label=r"$\langle Z_1 Z_2\rangle_{\rm theory}$")
plt.plot(ps, Z1m, "o", c=DARK_TEAL, label=r"$\overline{Z}_1$")
plt.plot(ps, 0*ps, "--", c=DARK_TEAL, label=r"$\langle Z_1\rangle_{\rm theory}$")
plt.plot(ps, Z2m, "d", c="k", label=r"$\overline{Z}_2$")
plt.plot(ps, 0*ps, "--", c="k", label=r"$\langle Z_2\rangle_{\rm theory}$")
plt.xlabel(r"Dephasing probability $p$", size=18)
plt.ylabel(r"$Z$-moment", size=18)
plt.title(r"$Z$-moments for a Bell-state prepared with dephased CZ", size=18)
plt.xlim(0, .5)
plt.legend(fontsize=18)
```
| github_jupyter |
Q1: Write a python program to add all the odd numbers from 0 to 20.
```
for i in range (1,21):
if i%2==0:
print(i)
```
Q 2: Write a python program to find the sum of all integers greater than 100 and less than 200.
```
counter = 101
summ = 0
while (counter < 200):
summ=summ+counter
counter = counter+1
# print(counterter)
print(summ)
```
Q3: Write a program to display the sum of square of the first ten even natural numbers
```
# x=1
# total =0
# for i in range(1,50):
# while (i%2==0):
# if(x<12):
# print(i)
# total = total + i*i
# x=x+1
# break
# continue
total=0
counter=0
for i in range(1,50):
if (i%2==0):
if(counter<11):
total = total + (i*i)
counter =counter +1
print(total)
```
Q4: Write a python program to display ascii characters from 65 to 90
```
for i in range(65,91):
print(i,"=",chr(i))
```
Q5: Display ascii characters from 48 to 57.
```
for i in range(48,58):
print(i,"=",chr(i))
```
Q6:Display the following output with the help of Ascii character.
```
for i in range(97,123):
print(i,"=",chr(i))
```
Q7: Write a python program for given a Python list you should be able to display Python list in the following order
L1 = [100, 200, 300, 400, 500]
```
L1 = [100, 200, 300, 400, 500]
L1.sort(reverse=True)
print(L1)
```
Q8: Write a Python program to concatenate following dictionaries to create a new one.
Sample Dictionary :
dic1={1:10, 2:20}
dic2={3:30, 4:40}
dic3={5:50,6:60}
Expected Result : {1: 10, 2: 20, 3: 30, 4: 40, 5: 50, 6: 60}
```
dic1={1:10, 2:20}
dic2={3:30, 4:40}
dic3={5:50,6:60}
dic1.update(dic2)
dic1.update(dic3)
print(dic1)
```
Q9: Write a Python program to add key to a dictionary.
Sample Dictionary : {0: 10, 1: 20}
Expected Result : {0: 10, 1: 20, 2: 30}
```
sample_Dictionary = {0: 10, 1: 20}
sample_Dictionary2 = {2:30}
sample_Dictionary.update(sample_Dictionary2)
print(sample_Dictionary)
sample_Dictionary9 = {0: 10, 1: 20}
sample_Dictionary91 = {2:30}
sample_Dictionary9[2] = 30
print(sample_Dictionary9)
```
Q10: Write a Python program to print out a set containing all the colors from a list which are not present in another list
Test Data :
color_list_1 = set(["White", "Black", "Red"])
color_list_2 = set(["Red", "Green"])
```
color_list_1 = set(["White", "Black", "Red"])
color_list_2 = set(["Red", "Green"])
print(color_list_1.difference(color_list_2))
```
Q11: Given a Python list. Write a python program to turn every item of a list into its square List1 = [1, 2, 3, 4, 5, 6, 7]
Expected output:
[1, 4, 9, 16, 25, 36, 49]
```
List1 = [1, 2, 3, 4, 5, 6, 7]
my_new_list = [i * i for i in List1]
my_new_list
```
Q12: Program to count the number of each vowel in a string.
```
user_str = input('ENTER the string: ')
count_str = 0
vowels=['a','e','i','o','u']
vo_dic={'a': 0,'e':0,'i':0,'o':0,'u':0}
for i in user_str:
if(i in vowels):
count_str = count_str+1
# vo_dic[vowels.append] = 1
print(count_str)
# print(vo_dic)
#Updating a dictionary of vowels
user_str = input('ENTER the string: ')
count_str = 0
vowels=['a','e','i','o','u']
c = {'a': 0,'e':0,'i':0,'o':0,'u':0}
counter = 0
for i in range(len(user_str)):
if user_str[counter] in c:
# print(user_str[counter])
c[user_str[counter]]=c[user_str[counter]]+1
counter = counter + 1
print("Updating a dictionary of vowels",
c)
```
Q13:Write a python program to Access the value of key ‘history’ from the following dictionary-sampleDict = {
"class":{
"student":{
"name":"Mike",
"marks":{
"physics":70,
"history":80
}
}
}
}
```
sampleDict = {
"class":{
"student":{
"name":"Mike",
"marks":{
"physics":70,
"history":80
}
}
}
}
sampleDict['class']['student']['marks']['history']
import string
x = string.punctuation
user_punc = input("Enter string: ")
y=''
rep=user_punc
for i in user_punc:
if i in x:
y = rep.replace(i,'')
rep = y
print(rep)
```
Q15: Write a python program to print the Following:
1
2 1
3 2 1
```
for i in range(1,4):
while(i>0):
print(i, end='')
i=i-1
print()
```
Q16: WAP to print the following asterisk pattern:
```
for i in range(7):
while(i>0):
print('*', end='')
i = i -1
print()
```
Q17: WAP to create a function traiangle to print the following asterisk triangle pattern:
```
for i in range(1,5):
x=0
while(x<i):
print("*", end='')
x = x+1
print('')
for i in range(3,0,-1):
x=0
while(x<i):
print("*", end='')
x = x+1
print('')
```
Q18: Write a python program to print following multiplication table on the screen
```
for a in range(1,11):
print('\t',a,end=' ')
print()
print('\t',("-")*75)
for i in range (1,11):
for j in range(1,11):
# while(i==1):
# print(j,end='_')
# break
# print()
while(j==1):
print(i,end='|')
break
print('\t',i*j, end=' ')
print('','\n')
print()
```
| github_jupyter |
# GC-SAN in PyTorch
## Imports
```
import math
import numpy as np
import torch
from torch import nn
from torch.nn import Parameter
from torch.nn import functional as F
from enum import Enum
```
## Model
```
class SequentialRecommender(AbstractRecommender):
"""
This is a abstract sequential recommender. All the sequential model should implement This class.
"""
def __init__(self, config, dataset):
super(SequentialRecommender, self).__init__()
# load dataset info
self.USER_ID = config['USER_ID_FIELD']
self.ITEM_ID = config['ITEM_ID_FIELD']
self.ITEM_SEQ = self.ITEM_ID + config['LIST_SUFFIX']
self.ITEM_SEQ_LEN = config['ITEM_LIST_LENGTH_FIELD']
self.POS_ITEM_ID = self.ITEM_ID
self.NEG_ITEM_ID = config['NEG_PREFIX'] + self.ITEM_ID
self.max_seq_length = config['MAX_ITEM_LIST_LENGTH']
self.n_items = dataset.num(self.ITEM_ID)
def gather_indexes(self, output, gather_index):
"""Gathers the vectors at the specific positions over a minibatch"""
gather_index = gather_index.view(-1, 1, 1).expand(-1, -1, output.shape[-1])
output_tensor = output.gather(dim=1, index=gather_index)
return output_tensor.squeeze(1)
class MultiHeadAttention(nn.Module):
"""
Multi-head Self-attention layers, a attention score dropout layer is introduced.
Args:
input_tensor (torch.Tensor): the input of the multi-head self-attention layer
attention_mask (torch.Tensor): the attention mask for input tensor
Returns:
hidden_states (torch.Tensor): the output of the multi-head self-attention layer
"""
def __init__(self, n_heads, hidden_size, hidden_dropout_prob, attn_dropout_prob, layer_norm_eps):
super(MultiHeadAttention, self).__init__()
if hidden_size % n_heads != 0:
raise ValueError(
"The hidden size (%d) is not a multiple of the number of attention "
"heads (%d)" % (hidden_size, n_heads)
)
self.num_attention_heads = n_heads
self.attention_head_size = int(hidden_size / n_heads)
self.all_head_size = self.num_attention_heads * self.attention_head_size
self.query = nn.Linear(hidden_size, self.all_head_size)
self.key = nn.Linear(hidden_size, self.all_head_size)
self.value = nn.Linear(hidden_size, self.all_head_size)
self.attn_dropout = nn.Dropout(attn_dropout_prob)
self.dense = nn.Linear(hidden_size, hidden_size)
self.LayerNorm = nn.LayerNorm(hidden_size, eps=layer_norm_eps)
self.out_dropout = nn.Dropout(hidden_dropout_prob)
def transpose_for_scores(self, x):
new_x_shape = x.size()[:-1] + (self.num_attention_heads, self.attention_head_size)
x = x.view(*new_x_shape)
return x.permute(0, 2, 1, 3)
def forward(self, input_tensor, attention_mask):
mixed_query_layer = self.query(input_tensor)
mixed_key_layer = self.key(input_tensor)
mixed_value_layer = self.value(input_tensor)
query_layer = self.transpose_for_scores(mixed_query_layer)
key_layer = self.transpose_for_scores(mixed_key_layer)
value_layer = self.transpose_for_scores(mixed_value_layer)
# Take the dot product between "query" and "key" to get the raw attention scores.
attention_scores = torch.matmul(query_layer, key_layer.transpose(-1, -2))
attention_scores = attention_scores / math.sqrt(self.attention_head_size)
# Apply the attention mask is (precomputed for all layers in BertModel forward() function)
# [batch_size heads seq_len seq_len] scores
# [batch_size 1 1 seq_len]
attention_scores = attention_scores + attention_mask
# Normalize the attention scores to probabilities.
attention_probs = nn.Softmax(dim=-1)(attention_scores)
# This is actually dropping out entire tokens to attend to, which might
# seem a bit unusual, but is taken from the original Transformer paper.
attention_probs = self.attn_dropout(attention_probs)
context_layer = torch.matmul(attention_probs, value_layer)
context_layer = context_layer.permute(0, 2, 1, 3).contiguous()
new_context_layer_shape = context_layer.size()[:-2] + (self.all_head_size,)
context_layer = context_layer.view(*new_context_layer_shape)
hidden_states = self.dense(context_layer)
hidden_states = self.out_dropout(hidden_states)
hidden_states = self.LayerNorm(hidden_states + input_tensor)
return hidden_states
class FeedForward(nn.Module):
"""
Point-wise feed-forward layer is implemented by two dense layers.
Args:
input_tensor (torch.Tensor): the input of the point-wise feed-forward layer
Returns:
hidden_states (torch.Tensor): the output of the point-wise feed-forward layer
"""
def __init__(self, hidden_size, inner_size, hidden_dropout_prob, hidden_act, layer_norm_eps):
super(FeedForward, self).__init__()
self.dense_1 = nn.Linear(hidden_size, inner_size)
self.intermediate_act_fn = self.get_hidden_act(hidden_act)
self.dense_2 = nn.Linear(inner_size, hidden_size)
self.LayerNorm = nn.LayerNorm(hidden_size, eps=layer_norm_eps)
self.dropout = nn.Dropout(hidden_dropout_prob)
def get_hidden_act(self, act):
ACT2FN = {
"gelu": self.gelu,
"relu": fn.relu,
"swish": self.swish,
"tanh": torch.tanh,
"sigmoid": torch.sigmoid,
}
return ACT2FN[act]
def gelu(self, x):
"""Implementation of the gelu activation function.
For information: OpenAI GPT's gelu is slightly different (and gives slightly different results)::
0.5 * x * (1 + torch.tanh(math.sqrt(2 / math.pi) * (x + 0.044715 * torch.pow(x, 3))))
Also see https://arxiv.org/abs/1606.08415
"""
return x * 0.5 * (1.0 + torch.erf(x / math.sqrt(2.0)))
def swish(self, x):
return x * torch.sigmoid(x)
def forward(self, input_tensor):
hidden_states = self.dense_1(input_tensor)
hidden_states = self.intermediate_act_fn(hidden_states)
hidden_states = self.dense_2(hidden_states)
hidden_states = self.dropout(hidden_states)
hidden_states = self.LayerNorm(hidden_states + input_tensor)
return hidden_states
class TransformerLayer(nn.Module):
"""
One transformer layer consists of a multi-head self-attention layer and a point-wise feed-forward layer.
Args:
hidden_states (torch.Tensor): the input of the multi-head self-attention sublayer
attention_mask (torch.Tensor): the attention mask for the multi-head self-attention sublayer
Returns:
feedforward_output (torch.Tensor): The output of the point-wise feed-forward sublayer,
is the output of the transformer layer.
"""
def __init__(
self, n_heads, hidden_size, intermediate_size, hidden_dropout_prob, attn_dropout_prob, hidden_act,
layer_norm_eps
):
super(TransformerLayer, self).__init__()
self.multi_head_attention = MultiHeadAttention(
n_heads, hidden_size, hidden_dropout_prob, attn_dropout_prob, layer_norm_eps
)
self.feed_forward = FeedForward(hidden_size, intermediate_size, hidden_dropout_prob, hidden_act, layer_norm_eps)
def forward(self, hidden_states, attention_mask):
attention_output = self.multi_head_attention(hidden_states, attention_mask)
feedforward_output = self.feed_forward(attention_output)
return feedforward_output
class TransformerEncoder(nn.Module):
r""" One TransformerEncoder consists of several TransformerLayers.
- n_layers(num): num of transformer layers in transformer encoder. Default: 2
- n_heads(num): num of attention heads for multi-head attention layer. Default: 2
- hidden_size(num): the input and output hidden size. Default: 64
- inner_size(num): the dimensionality in feed-forward layer. Default: 256
- hidden_dropout_prob(float): probability of an element to be zeroed. Default: 0.5
- attn_dropout_prob(float): probability of an attention score to be zeroed. Default: 0.5
- hidden_act(str): activation function in feed-forward layer. Default: 'gelu'
candidates: 'gelu', 'relu', 'swish', 'tanh', 'sigmoid'
- layer_norm_eps(float): a value added to the denominator for numerical stability. Default: 1e-12
"""
def __init__(
self,
n_layers=2,
n_heads=2,
hidden_size=64,
inner_size=256,
hidden_dropout_prob=0.5,
attn_dropout_prob=0.5,
hidden_act='gelu',
layer_norm_eps=1e-12
):
super(TransformerEncoder, self).__init__()
layer = TransformerLayer(
n_heads, hidden_size, inner_size, hidden_dropout_prob, attn_dropout_prob, hidden_act, layer_norm_eps
)
self.layer = nn.ModuleList([copy.deepcopy(layer) for _ in range(n_layers)])
def forward(self, hidden_states, attention_mask, output_all_encoded_layers=True):
"""
Args:
hidden_states (torch.Tensor): the input of the TransformerEncoder
attention_mask (torch.Tensor): the attention mask for the input hidden_states
output_all_encoded_layers (Bool): whether output all transformer layers' output
Returns:
all_encoder_layers (list): if output_all_encoded_layers is True, return a list consists of all transformer
layers' output, otherwise return a list only consists of the output of last transformer layer.
"""
all_encoder_layers = []
for layer_module in self.layer:
hidden_states = layer_module(hidden_states, attention_mask)
if output_all_encoded_layers:
all_encoder_layers.append(hidden_states)
if not output_all_encoded_layers:
all_encoder_layers.append(hidden_states)
return all_encoder_layers
class BPRLoss(nn.Module):
""" BPRLoss, based on Bayesian Personalized Ranking
Args:
- gamma(float): Small value to avoid division by zero
Shape:
- Pos_score: (N)
- Neg_score: (N), same shape as the Pos_score
- Output: scalar.
Examples::
>>> loss = BPRLoss()
>>> pos_score = torch.randn(3, requires_grad=True)
>>> neg_score = torch.randn(3, requires_grad=True)
>>> output = loss(pos_score, neg_score)
>>> output.backward()
"""
def __init__(self, gamma=1e-10):
super(BPRLoss, self).__init__()
self.gamma = gamma
def forward(self, pos_score, neg_score):
loss = -torch.log(self.gamma + torch.sigmoid(pos_score - neg_score)).mean()
return loss
class EmbLoss(nn.Module):
""" EmbLoss, regularization on embeddings
"""
def __init__(self, norm=2):
super(EmbLoss, self).__init__()
self.norm = norm
def forward(self, *embeddings):
emb_loss = torch.zeros(1).to(embeddings[-1].device)
for embedding in embeddings:
emb_loss += torch.norm(embedding, p=self.norm)
emb_loss /= embeddings[-1].shape[0]
return emb_loss
class GNN(nn.Module):
r"""Graph neural networks are well-suited for session-based recommendation,
because it can automatically extract features of session graphs with considerations of rich node connections.
"""
def __init__(self, embedding_size, step=1):
super(GNN, self).__init__()
self.step = step
self.embedding_size = embedding_size
self.input_size = embedding_size * 2
self.gate_size = embedding_size * 3
self.w_ih = Parameter(torch.Tensor(self.gate_size, self.input_size))
self.w_hh = Parameter(torch.Tensor(self.gate_size, self.embedding_size))
self.b_ih = Parameter(torch.Tensor(self.gate_size))
self.b_hh = Parameter(torch.Tensor(self.gate_size))
self.linear_edge_in = nn.Linear(self.embedding_size, self.embedding_size, bias=True)
self.linear_edge_out = nn.Linear(self.embedding_size, self.embedding_size, bias=True)
# parameters initialization
self._reset_parameters()
def _reset_parameters(self):
stdv = 1.0 / math.sqrt(self.embedding_size)
for weight in self.parameters():
weight.data.uniform_(-stdv, stdv)
def GNNCell(self, A, hidden):
r"""Obtain latent vectors of nodes via gated graph neural network.
Args:
A (torch.FloatTensor): The connection matrix,shape of [batch_size, max_session_len, 2 * max_session_len]
hidden (torch.FloatTensor): The item node embedding matrix, shape of
[batch_size, max_session_len, embedding_size]
Returns:
torch.FloatTensor: Latent vectors of nodes,shape of [batch_size, max_session_len, embedding_size]
"""
input_in = torch.matmul(A[:, :, :A.size(1)], self.linear_edge_in(hidden))
input_out = torch.matmul(A[:, :, A.size(1):2 * A.size(1)], self.linear_edge_out(hidden))
# [batch_size, max_session_len, embedding_size * 2]
inputs = torch.cat([input_in, input_out], 2)
# gi.size equals to gh.size, shape of [batch_size, max_session_len, embedding_size * 3]
gi = F.linear(inputs, self.w_ih, self.b_ih)
gh = F.linear(hidden, self.w_hh, self.b_hh)
# (batch_size, max_session_len, embedding_size)
i_r, i_i, i_n = gi.chunk(3, 2)
h_r, h_i, h_n = gh.chunk(3, 2)
reset_gate = torch.sigmoid(i_r + h_r)
input_gate = torch.sigmoid(i_i + h_i)
new_gate = torch.tanh(i_n + reset_gate * h_n)
hy = (1 - input_gate) * hidden + input_gate * new_gate
return hy
def forward(self, A, hidden):
for i in range(self.step):
hidden = self.GNNCell(A, hidden)
return hidden
class GCSAN(SequentialRecommender):
r"""GCSAN captures rich local dependencies via graph neural network,
and learns long-range dependencies by applying the self-attention mechanism.
Note:
In the original paper, the attention mechanism in the self-attention layer is a single head,
for the reusability of the project code, we use a unified transformer component.
According to the experimental results, we only applied regularization to embedding.
"""
def __init__(self, config, dataset):
super(GCSAN, self).__init__(config, dataset)
# load parameters info
self.n_layers = config['n_layers']
self.n_heads = config['n_heads']
self.hidden_size = config['hidden_size'] # same as embedding_size
self.inner_size = config['inner_size'] # the dimensionality in feed-forward layer
self.hidden_dropout_prob = config['hidden_dropout_prob']
self.attn_dropout_prob = config['attn_dropout_prob']
self.hidden_act = config['hidden_act']
self.layer_norm_eps = config['layer_norm_eps']
self.step = config['step']
self.device = config['device']
self.weight = config['weight']
self.reg_weight = config['reg_weight']
self.loss_type = config['loss_type']
self.initializer_range = config['initializer_range']
# define layers and loss
self.item_embedding = nn.Embedding(self.n_items, self.hidden_size, padding_idx=0)
self.gnn = GNN(self.hidden_size, self.step)
self.self_attention = TransformerEncoder(
n_layers=self.n_layers,
n_heads=self.n_heads,
hidden_size=self.hidden_size,
inner_size=self.inner_size,
hidden_dropout_prob=self.hidden_dropout_prob,
attn_dropout_prob=self.attn_dropout_prob,
hidden_act=self.hidden_act,
layer_norm_eps=self.layer_norm_eps
)
self.reg_loss = EmbLoss()
if self.loss_type == 'BPR':
self.loss_fct = BPRLoss()
elif self.loss_type == 'CE':
self.loss_fct = nn.CrossEntropyLoss()
else:
raise NotImplementedError("Make sure 'loss_type' in ['BPR', 'CE']!")
# parameters initialization
self.apply(self._init_weights)
def _init_weights(self, module):
""" Initialize the weights """
if isinstance(module, (nn.Linear, nn.Embedding)):
# Slightly different from the TF version which uses truncated_normal for initialization
# cf https://github.com/pytorch/pytorch/pull/5617
module.weight.data.normal_(mean=0.0, std=self.initializer_range)
elif isinstance(module, nn.LayerNorm):
module.bias.data.zero_()
module.weight.data.fill_(1.0)
if isinstance(module, nn.Linear) and module.bias is not None:
module.bias.data.zero_()
def get_attention_mask(self, item_seq):
"""Generate left-to-right uni-directional attention mask for multi-head attention."""
attention_mask = (item_seq > 0).long()
extended_attention_mask = attention_mask.unsqueeze(1).unsqueeze(2) # torch.int64
# mask for left-to-right unidirectional
max_len = attention_mask.size(-1)
attn_shape = (1, max_len, max_len)
subsequent_mask = torch.triu(torch.ones(attn_shape), diagonal=1) # torch.uint8
subsequent_mask = (subsequent_mask == 0).unsqueeze(1)
subsequent_mask = subsequent_mask.long().to(item_seq.device)
extended_attention_mask = extended_attention_mask * subsequent_mask
extended_attention_mask = extended_attention_mask.to(dtype=next(self.parameters()).dtype) # fp16 compatibility
extended_attention_mask = (1.0 - extended_attention_mask) * -10000.0
return extended_attention_mask
def _get_slice(self, item_seq):
items, n_node, A, alias_inputs = [], [], [], []
max_n_node = item_seq.size(1)
item_seq = item_seq.cpu().numpy()
for u_input in item_seq:
node = np.unique(u_input)
items.append(node.tolist() + (max_n_node - len(node)) * [0])
u_A = np.zeros((max_n_node, max_n_node))
for i in np.arange(len(u_input) - 1):
if u_input[i + 1] == 0:
break
u = np.where(node == u_input[i])[0][0]
v = np.where(node == u_input[i + 1])[0][0]
u_A[u][v] = 1
u_sum_in = np.sum(u_A, 0)
u_sum_in[np.where(u_sum_in == 0)] = 1
u_A_in = np.divide(u_A, u_sum_in)
u_sum_out = np.sum(u_A, 1)
u_sum_out[np.where(u_sum_out == 0)] = 1
u_A_out = np.divide(u_A.transpose(), u_sum_out)
u_A = np.concatenate([u_A_in, u_A_out]).transpose()
A.append(u_A)
alias_inputs.append([np.where(node == i)[0][0] for i in u_input])
# The relative coordinates of the item node, shape of [batch_size, max_session_len]
alias_inputs = torch.LongTensor(alias_inputs).to(self.device)
# The connecting matrix, shape of [batch_size, max_session_len, 2 * max_session_len]
A = torch.FloatTensor(A).to(self.device)
# The unique item nodes, shape of [batch_size, max_session_len]
items = torch.LongTensor(items).to(self.device)
return alias_inputs, A, items
def forward(self, item_seq, item_seq_len):
assert 0 <= self.weight <= 1
alias_inputs, A, items = self._get_slice(item_seq)
hidden = self.item_embedding(items)
hidden = self.gnn(A, hidden)
alias_inputs = alias_inputs.view(-1, alias_inputs.size(1), 1).expand(-1, -1, self.hidden_size)
seq_hidden = torch.gather(hidden, dim=1, index=alias_inputs)
# fetch the last hidden state of last timestamp
ht = self.gather_indexes(seq_hidden, item_seq_len - 1)
a = seq_hidden
attention_mask = self.get_attention_mask(item_seq)
outputs = self.self_attention(a, attention_mask, output_all_encoded_layers=True)
output = outputs[-1]
at = self.gather_indexes(output, item_seq_len - 1)
seq_output = self.weight * at + (1 - self.weight) * ht
return seq_output
def calculate_loss(self, interaction):
item_seq = interaction[self.ITEM_SEQ]
item_seq_len = interaction[self.ITEM_SEQ_LEN]
seq_output = self.forward(item_seq, item_seq_len)
pos_items = interaction[self.POS_ITEM_ID]
if self.loss_type == 'BPR':
neg_items = interaction[self.NEG_ITEM_ID]
pos_items_emb = self.item_embedding(pos_items)
neg_items_emb = self.item_embedding(neg_items)
pos_score = torch.sum(seq_output * pos_items_emb, dim=-1) # [B]
neg_score = torch.sum(seq_output * neg_items_emb, dim=-1) # [B]
loss = self.loss_fct(pos_score, neg_score)
else: # self.loss_type = 'CE'
test_item_emb = self.item_embedding.weight
logits = torch.matmul(seq_output, test_item_emb.transpose(0, 1))
loss = self.loss_fct(logits, pos_items)
reg_loss = self.reg_loss(self.item_embedding.weight)
total_loss = loss + self.reg_weight * reg_loss
return total_loss
def predict(self, interaction):
item_seq = interaction[self.ITEM_SEQ]
item_seq_len = interaction[self.ITEM_SEQ_LEN]
test_item = interaction[self.ITEM_ID]
seq_output = self.forward(item_seq, item_seq_len)
test_item_emb = self.item_embedding(test_item)
scores = torch.mul(seq_output, test_item_emb).sum(dim=1) # [B]
return scores
def full_sort_predict(self, interaction):
item_seq = interaction[self.ITEM_SEQ]
item_seq_len = interaction[self.ITEM_SEQ_LEN]
seq_output = self.forward(item_seq, item_seq_len)
test_items_emb = self.item_embedding.weight
scores = torch.matmul(seq_output, test_items_emb.transpose(0, 1)) # [B, n_items]
return scores
```
| github_jupyter |
```
import numpy as np
from tqdm.auto import tqdm
from sklearn.metrics import mean_squared_error
import matplotlib.pyplot as plt
class ANN:
def __init__(self, intermediate_size, activation_fn=np.tanh):
self._parameters = self._init_params([1, intermediate_size, 1])
self._activation_fn = activation_fn
def parameters(self):
return self._parameters
def update_params(self, params):
self._parameters = params
@staticmethod
def _init_params(sizes):
params = {}
for i in range(1, len(sizes)):
params[f"w{i}"] = np.random.normal(scale=0.01, size=(sizes[i], sizes[i - 1]))
params[f"b{i}"] = np.random.normal(scale=0.01, size=(sizes[i], 1))
return params
def forward(self, inputs):
num_layers = len(self._parameters) // 2
activations = {}
for i in range(1, num_layers+1):
if i == 1:
activations[f"z{i}"] = self._parameters[f"w{i}"] @ inputs + self._parameters[f"b{i}"]
activations[f"a{i}"] = self._activation_fn(activations[f"z{i}"])
else:
activations[f"z{i}"] = self._parameters[f"w{i}"] @ activations[f"a{i-1}"] + self._parameters[f"b{i}"]
if i == num_layers:
activations[f"a{i}"] = activations[f"z{i}"]
else:
activations[f"a{i}"] = self._activation_fn(activations[f"z{i}"])
return activations
def backward(self, activations, inputs, targets):
num_layers = len(self._parameters)//2
m = len(targets)
grads = {}
for i in range(num_layers, 0, -1):
if i == num_layers:
dA = 1/m * (activations[f"a{i}"] - targets)
dZ = dA
else:
dA = (self._parameters[f"w{i+1}"].T @ dZ) * (1 - np.tanh(self._parameters[f"w{i+1}"] @ activations[f"a{i}"] + self._parameters[f"b{i}"]) ** 2)
dZ = np.multiply(dA, np.where(activations[f"a{i}"] >= 0, 1, 0))
if i==1:
grads[f"w{i}"] = 1/m * (dZ @ inputs.T)
grads[f"b{i}"] = 1/m * np.sum(dZ, axis=1, keepdims=True)
else:
grads[f"w{i}"] = 1/m * (dZ @ activations[f"a{i-1}"].T)
grads[f"b{i}"] = 1/m * np.sum(dZ, axis=1, keepdims=True)
return grads
def optimizer_step(params, grads, lr):
num_layers = len(params) // 2
new_params = {}
for i in range(1, num_layers+1):
new_params[f"w{i}"] = params[f"w{i}"] - lr * grads[f"w{i}"]
new_params[f"b{i}"] = params[f"b{i}"] - lr * grads[f"b{i}"]
return new_params
def train(X, y, intermediate_size, n_epochs, lr):
model = ANN(intermediate_size)
for _ in range(1, n_epochs + 1):
activations = model.forward(X)
grads = model.backward(activations, X, y.T)
new_params = optimizer_step(model.parameters(), grads, lr)
model.update_params(new_params)
return model
def compute_score(model, X, y):
activations = model.forward(X)
score = np.sqrt(mean_squared_error(y, activations["a2"].T))
return score
ages = [15, 15, 15, 18, 28, 29, 37, 37, 44, 50, 50, 60, 61, 64, 65, 65, 72, 75, 75, 82, 85, 91, 91, 97, 98, 125, 142, 142, 147, 147, 150, 159, 165, 183, 192, 195, 218, 218, 219, 224, 225, 227, 232, 232, 237, 246, 258, 276, 285, 300, 301, 305, 312, 317, 338, 347, 354, 357, 375, 394, 513, 535, 554, 591, 648, 660, 705, 723, 756, 768, 860]
weights = [21.66, 22.75, 22.3, 31.25, 44.79, 40.55, 50.25, 46.88, 52.03, 63.47, 61.13, 81, 73.09, 79.09, 79.51, 65.31, 71.9, 86.1, 94.6, 92.5, 105, 101.7, 102.9, 110, 104.3, 134.9, 130.68, 140.58, 155.3, 152.2, 144.5, 142.15, 139.81, 153.22, 145.72, 161.1, 174.18, 173.03, 173.54, 178.86, 177.68, 173.73, 159.98, 161.29, 187.07, 176.13, 183.4, 186.26, 189.66, 186.09, 186.7, 186.8, 195.1, 216.41, 203.23, 188.38, 189.7, 195.31, 202.63, 224.82, 203.3, 209.7, 233.9, 234.7, 244.3, 231, 242.4, 230.77, 242.57, 232.12, 246.7]
ages, weights = np.array(ages)[None], np.array(weights)
n_epochs = 450
lr = 1e-3
metrics, intermediate_sizes = [], []
for intermediate_size in tqdm(range(10, 1000, 10)):
trained_model = train(ages, weights, intermediate_size, n_epochs, lr)
score = compute_score(trained_model, ages, weights)
metrics.append(score)
intermediate_sizes.append(intermediate_size)
print("Training Loss for various intermediate sizes", metrics)
plt.plot(intermediate_sizes, metrics)
plt.xlabel("Number of neurons in intermediate layer")
plt.ylabel("Training Loss")
plt.show()
```
| github_jupyter |
# Exercise 5 - Variational quantum eigensolver
## Historical background
During the last decade, quantum computers matured quickly and began to realize Feynman's initial dream of a computing system that could simulate the laws of nature in a quantum way. A 2014 paper first authored by Alberto Peruzzo introduced the **Variational Quantum Eigensolver (VQE)**, an algorithm meant for finding the ground state energy (lowest energy) of a molecule, with much shallower circuits than other approaches.[1] And, in 2017, the IBM Quantum team used the VQE algorithm to simulate the ground state energy of the lithium hydride molecule.[2]
VQE's magic comes from outsourcing some of the problem's processing workload to a classical computer. The algorithm starts with a parameterized quantum circuit called an ansatz (a best guess) then finds the optimal parameters for this circuit using a classical optimizer. The VQE's advantage over classical algorithms comes from the fact that a quantum processing unit can represent and store the problem's exact wavefunction, an exponentially hard problem for a classical computer.
This exercise 5 allows you to realize Feynman's dream yourself, setting up a variational quantum eigensolver to determine the ground state and the energy of a molecule. This is interesting because the ground state can be used to calculate various molecular properties, for instance the exact forces on nuclei than can serve to run molecular dynamics simulations to explore what happens in chemical systems with time.[3]
### References
1. Peruzzo, Alberto, et al. "A variational eigenvalue solver on a photonic quantum processor." Nature communications 5.1 (2014): 1-7.
2. Kandala, Abhinav, et al. "Hardware-efficient variational quantum eigensolver for small molecules and quantum magnets." Nature 549.7671 (2017): 242-246.
3. Sokolov, Igor O., et al. "Microcanonical and finite-temperature ab initio molecular dynamics simulations on quantum computers." Physical Review Research 3.1 (2021): 013125.
## Introduction
For the implementation of VQE, you will be able to make choices on how you want to compose your simulation, in particular focusing on the ansatz quantum circuits.
This is motivated by the fact that one of the important tasks when running VQE on noisy quantum computers is to reduce the loss of fidelity (which introduces errors) by finding the most compact quantum circuit capable of representing the ground state.
Practically, this entails to minimizing the number of two-qubit gates (e.g. CNOTs) while not loosing accuracy.
<div class="alert alert-block alert-success">
<b>Goal</b>
Find the shortest ansatz circuits for representing accurately the ground state of given problems. Be creative!
<b>Plan</b>
First you will learn how to compose a VQE simulation for the smallest molecule and then apply what you have learned to a case of a larger one.
**1. Tutorial - VQE for H$_2$:** familiarize yourself with VQE and select the best combination of ansatz/classical optimizer by running statevector simulations.
**2. Final Challenge - VQE for LiH:** perform similar investigation as in the first part but restricting to statevector simulator only. Use the qubit number reduction schemes available in Qiskit and find the optimal circuit for this larger system. Optimize the circuit and use your imagination to find ways to select the best building blocks of parameterized circuits and compose them to construct the most compact ansatz circuit for the ground state, better than the ones already available in Qiskit.
</div>
<div class="alert alert-block alert-danger">
Below is an introduction to the theory behind VQE simulations. You don't have to understand the whole thing before moving on. Don't be scared!
</div>
## Theory
Here below is the general workflow representing how the molecular simulations using VQE are performed on quantum computers.
<img src="resources/workflow.png" width=800 height= 1400/>
The core idea hybrid quantum-classical approach is to outsource to **CPU (classical processing unit)** and **QPU (quantum processing unit)** the parts that they can do best. The CPU takes care of listing the terms that need to be measured to compute the energy and also optimizing the circuit parameters. The QPU implements a quantum circuit representing the quantum state of a system and measures the energy. Some more details are given below:
**CPU** can compute efficiently the energies associated to electron hopping and interactions (one-/two-body integrals by means of a Hartree-Fock calculation) that serve to represent the total energy operator, Hamiltonian. The [Hartree–Fock (HF) method](https://en.wikipedia.org/wiki/Hartree%E2%80%93Fock_method#:~:text=In%20computational%20physics%20and%20chemistry,system%20in%20a%20stationary%20state.) efficiently computes an approximate grounds state wavefunction by assuming that the latter can be represented by a single Slater determinant (e.g. for H$_2$ molecule in STO-3G basis with 4 spin-orbitals and qubits, $|\Psi_{HF} \rangle = |0101 \rangle$ where electrons occupy the lowest energy spin-orbitals). What QPU does later in VQE is finding a quantum state (corresponding circuit and its parameters) that can also represent other states associated missing electronic correlations (i.e. $\sum_i c_i |i\rangle$ states in $|\Psi \rangle = c_{HF}|\Psi_{HF} \rangle + \sum_i c_i |i\rangle $ where $i$ is a bitstring).
After a HF calculation, operators in the Hamiltonian are mapped to measurements on a QPU using fermion-to-qubit transformations (see Hamiltonian section below). One can further analyze the properties of the system to reduce the number of qubits or shorten the ansatz circuit:
- For Z2 symmetries and two-qubit reduction, see [Bravyi *et al*, 2017](https://arxiv.org/abs/1701.08213v1).
- For entanglement forging, see [Eddins *et al.*, 2021](https://arxiv.org/abs/2104.10220v1).
- For the adaptive ansatz see, [Grimsley *et al.*,2018](https://arxiv.org/abs/1812.11173v2), [Rattew *et al.*,2019](https://arxiv.org/abs/1910.09694), [Tang *et al.*,2019](https://arxiv.org/abs/1911.10205). You may use the ideas found in those works to find ways to shorten the quantum circuits.
**QPU** implements quantum circuits (see Ansatzes section below), parameterized by angles $\vec\theta$, that would represent the ground state wavefunction by placing various single qubit rotations and entanglers (e.g. two-qubit gates). The quantum advantage lies in the fact that QPU can efficiently represent and store the exact wavefunction, which becomes intractable on a classical computer for systems that have more than a few atoms. Finally, QPU measures the operators of choice (e.g. ones representing a Hamiltonian).
Below we go slightly more in mathematical details of each component of the VQE algorithm. It might be also helpful if you watch our [video episode about VQE](https://www.youtube.com/watch?v=Z-A6G0WVI9w).
### Hamiltonian
Here we explain how we obtain the operators that we need to measure to obtain the energy of a given system.
These terms are included in the molecular Hamiltonian defined as:
$$
\begin{aligned}
\hat{H} &=\sum_{r s} h_{r s} \hat{a}_{r}^{\dagger} \hat{a}_{s} \\
&+\frac{1}{2} \sum_{p q r s} g_{p q r s} \hat{a}_{p}^{\dagger} \hat{a}_{q}^{\dagger} \hat{a}_{r} \hat{a}_{s}+E_{N N}
\end{aligned}
$$
with
$$
h_{p q}=\int \phi_{p}^{*}(r)\left(-\frac{1}{2} \nabla^{2}-\sum_{I} \frac{Z_{I}}{R_{I}-r}\right) \phi_{q}(r)
$$
$$
g_{p q r s}=\int \frac{\phi_{p}^{*}\left(r_{1}\right) \phi_{q}^{*}\left(r_{2}\right) \phi_{r}\left(r_{2}\right) \phi_{s}\left(r_{1}\right)}{\left|r_{1}-r_{2}\right|}
$$
where the $h_{r s}$ and $g_{p q r s}$ are the one-/two-body integrals (using the Hartree-Fock method) and $E_{N N}$ the nuclear repulsion energy.
The one-body integrals represent the kinetic energy of the electrons and their interaction with nuclei.
The two-body integrals represent the electron-electron interaction.
The $\hat{a}_{r}^{\dagger}, \hat{a}_{r}$ operators represent creation and annihilation of electron in spin-orbital $r$ and require mappings to operators, so that we can measure them on a quantum computer.
Note that VQE minimizes the electronic energy so you have to retrieve and add the nuclear repulsion energy $E_{NN}$ to compute the total energy.
So, for every non-zero matrix element in the $ h_{r s}$ and $g_{p q r s}$ tensors, we can construct corresponding Pauli string (tensor product of Pauli operators) with the following fermion-to-qubit transformation.
For instance, in Jordan-Wigner mapping for an orbital $r = 3$, we obtain the following Pauli string:
$$
\hat a_{3}^{\dagger}= \hat \sigma_z \otimes \hat \sigma_z \otimes\left(\frac{ \hat \sigma_x-i \hat \sigma_y}{2}\right) \otimes 1 \otimes \cdots \otimes 1
$$
where $\hat \sigma_x, \hat \sigma_y, \hat \sigma_z$ are the well-known Pauli operators. The tensor products of $\hat \sigma_z$ operators are placed to enforce the fermionic anti-commutation relations.
A representation of the Jordan-Wigner mapping between the 14 spin-orbitals of a water molecule and some 14 qubits is given below:
<img src="resources/mapping.png" width=600 height= 1200/>
Then, one simply replaces the one-/two-body excitations (e.g. $\hat{a}_{r}^{\dagger} \hat{a}_{s}$, $\hat{a}_{p}^{\dagger} \hat{a}_{q}^{\dagger} \hat{a}_{r} \hat{a}_{s}$) in the Hamiltonian by corresponding Pauli strings (i.e. $\hat{P}_i$, see picture above). The resulting operator set is ready to be measured on the QPU.
For additional details see [Seeley *et al.*, 2012](https://arxiv.org/abs/1208.5986v1).
### Ansatzes
There are mainly 2 types of ansatzes you can use for chemical problems.
- **q-UCC ansatzes** are physically inspired, and roughly map the electron excitations to quantum circuits. The q-UCCSD ansatz (`UCCSD`in Qiskit) possess all possible single and double electron excitations. The paired double q-pUCCD (`PUCCD`) and singlet q-UCCD0 (`SUCCD`) just consider a subset of such excitations (meaning significantly shorter circuits) and have proved to provide good results for dissociation profiles. For instance, q-pUCCD doesn't have single excitations and the double excitations are paired as in the image below.
- **Heuristic ansatzes (`TwoLocal`)** were invented to shorten the circuit depth but still be able to represent the ground state.
As in the figure below, the R gates represent the parametrized single qubit rotations and $U_{CNOT}$ the entanglers (two-qubit gates). The idea is that after repeating certain $D$-times the same block (with independent parameters) one can reach the ground state.
For additional details refer to [Sokolov *et al.* (q-UCC ansatzes)](https://arxiv.org/abs/1911.10864v2) and [Barkoutsos *et al.* (Heuristic ansatzes)](https://arxiv.org/pdf/1805.04340.pdf).
<img src="resources/ansatz.png" width=700 height= 1200/>
### VQE
Given a Hermitian operator $\hat H$ with an unknown minimum eigenvalue $E_{min}$, associated with the eigenstate $|\psi_{min}\rangle$, VQE provides an estimate $E_{\theta}$, bounded by $E_{min}$:
\begin{align*}
E_{min} \le E_{\theta} \equiv \langle \psi(\theta) |\hat H|\psi(\theta) \rangle
\end{align*}
where $|\psi(\theta)\rangle$ is the trial state associated with $E_{\theta}$. By applying a parameterized circuit, represented by $U(\theta)$, to some arbitrary starting state $|\psi\rangle$, the algorithm obtains an estimate $U(\theta)|\psi\rangle \equiv |\psi(\theta)\rangle$ on $|\psi_{min}\rangle$. The estimate is iteratively optimized by a classical optimizer by changing the parameter $\theta$ and minimizing the expectation value of $\langle \psi(\theta) |\hat H|\psi(\theta) \rangle$.
As applications of VQE, there are possibilities in molecular dynamics simulations, see [Sokolov *et al.*, 2021](https://arxiv.org/abs/2008.08144v1), and excited states calculations, see [Ollitrault *et al.*, 2019](https://arxiv.org/abs/1910.12890) to name a few.
<div class="alert alert-block alert-danger">
<b> References for additional details</b>
For the qiskit-nature tutorial that implements this algorithm see [here](https://qiskit.org/documentation/nature/tutorials/01_electronic_structure.html)
but this won't be sufficient and you might want to look on the [first page of github repository](https://github.com/Qiskit/qiskit-nature) and the [test folder](https://github.com/Qiskit/qiskit-nature/tree/main/test) containing tests that are written for each component, they provide the base code for the use of each functionality.
</div>
## Part 1: Tutorial - VQE for H$_2$ molecule
In this part, you will simulate H$_2$ molecule using the STO-3G basis with the PySCF driver and Jordan-Wigner mapping.
We will guide you through the following parts so then you can tackle harder problems.
#### 1. Driver
The interfaces to the classical chemistry codes that are available in Qiskit are called drivers.
We have for example `PSI4Driver`, `PyQuanteDriver`, `PySCFDriver` are available.
By running a driver (Hartree-Fock calculation for a given basis set and molecular geometry), in the cell below, we obtain all the necessary information about our molecule to apply then a quantum algorithm.
```
#from qiskit_nature.drivers import PySCFDriver
#molecule = "H .0 .0 .0; H .0 .0 0.739"
#driver = PySCFDriver(atom=molecule)
#qmolecule = driver.run()
from qiskit_nature.drivers import PySCFDriver
molecule = 'Li 0.0 0.0 0.0; H 0.0 0.0 1.5474'
driver = PySCFDriver(atom=molecule)
qmolecule = driver.run()
```
<div class="alert alert-block alert-danger">
<b> Tutorial questions 1</b>
Look into the attributes of `qmolecule` and answer the questions below.
1. We need to know the basic characteristics of our molecule. What is the total number of electrons in your system?
2. What is the number of molecular orbitals?
3. What is the number of spin-orbitals?
3. How many qubits would you need to simulate this molecule with Jordan-Wigner mapping?
5. What is the value of the nuclear repulsion energy?
You can find the answers at the end of this notebook.
</div>
```
# WRITE YOUR CODE BETWEEN THESE LINES - START
n_el = qmolecule.num_alpha + qmolecule.num_beta
n_mo = qmolecule.num_molecular_orbitals
n_so = 2 * qmolecule.num_molecular_orbitals
n_q = 2* qmolecule.num_molecular_orbitals
e_nn = qmolecule.nuclear_repulsion_energy
# WRITE YOUR CODE BETWEEN THESE LINES - END
```
#### 2. Electronic structure problem
You can then create an `ElectronicStructureProblem` that can produce the list of fermionic operators before mapping them to qubits (Pauli strings).
```
from qiskit_nature.problems.second_quantization.electronic import ElectronicStructureProblem
from qiskit_nature.transformers import FreezeCoreTransformer
#freezeCoreTransformer = FreezeCoreTransformer()
#qmolecule = freezeCoreTransformer.transform(qmolecule)
problem = ElectronicStructureProblem(driver,q_molecule_transformers=[FreezeCoreTransformer(freeze_core=True,remove_orbitals=[3,4])])
#problem = ElectronicStructureProblem(driver)
# Generate the second-quantized operators
second_q_ops = problem.second_q_ops()
# Hamiltonian
main_op = second_q_ops[0]
```
#### 3. QubitConverter
Allows to define the mapping that you will use in the simulation. You can try different mapping but
we will stick to `JordanWignerMapper` as allows a simple correspondence: a qubit represents a spin-orbital in the molecule.
```
from qiskit_nature.mappers.second_quantization import ParityMapper, BravyiKitaevMapper, JordanWignerMapper
from qiskit_nature.converters.second_quantization.qubit_converter import QubitConverter
# Setup the mapper and qubit converter
mapper_type = 'ParityMapper'
if mapper_type == 'ParityMapper':
mapper = ParityMapper()
elif mapper_type == 'JordanWignerMapper':
mapper = JordanWignerMapper()
elif mapper_type == 'BravyiKitaevMapper':
mapper = BravyiKitaevMapper()
converter = QubitConverter(mapper=mapper, two_qubit_reduction=True,z2symmetry_reduction=[1])
#converter = QubitConverter(mapper=mapper, two_qubit_reduction=True)
# The fermionic operators are mapped to qubit operators
num_particles = (problem.molecule_data_transformed.num_alpha,
problem.molecule_data_transformed.num_beta)
qubit_op = converter.convert(main_op, num_particles=num_particles)
```
#### 4. Initial state
As we described in the Theory section, a good initial state in chemistry is the HF state (i.e. $|\Psi_{HF} \rangle = |0101 \rangle$). We can initialize it as follows:
```
from qiskit_nature.circuit.library import HartreeFock
num_particles = (problem.molecule_data_transformed.num_alpha,
problem.molecule_data_transformed.num_beta)
num_spin_orbitals = 2 * problem.molecule_data_transformed.num_molecular_orbitals
init_state = HartreeFock(num_spin_orbitals, num_particles, converter)
print(init_state)
```
#### 5. Ansatz
One of the most important choices is the quantum circuit that you choose to approximate your ground state.
Here is the example of qiskit circuit library that contains many possibilities for making your own circuit.
```
from qiskit.circuit.library import TwoLocal
from qiskit_nature.circuit.library import UCCSD, PUCCD, SUCCD
# Choose the ansatz
ansatz_type = "TwoLocal"
# Parameters for q-UCC antatze
num_particles = (problem.molecule_data_transformed.num_alpha,
problem.molecule_data_transformed.num_beta)
num_spin_orbitals = 2 * problem.molecule_data_transformed.num_molecular_orbitals
# Put arguments for twolocal
if ansatz_type == "TwoLocal":
# Single qubit rotations that are placed on all qubits with independent parameters
rotation_blocks = ['ry', 'rz']
# Entangling gates
entanglement_blocks = 'cx'
# How the qubits are entangled
entanglement = 'full'
# Repetitions of rotation_blocks + entanglement_blocks with independent parameters
repetitions = 3
# Skip the final rotation_blocks layer
skip_final_rotation_layer = True
ansatz = TwoLocal(qubit_op.num_qubits, rotation_blocks, entanglement_blocks, reps=repetitions,
entanglement=entanglement, skip_final_rotation_layer=skip_final_rotation_layer)
# Add the initial state
ansatz.compose(init_state, front=True, inplace=True)
elif ansatz_type == "UCCSD":
ansatz = UCCSD(converter,num_particles,num_spin_orbitals,initial_state = init_state)
elif ansatz_type == "PUCCD":
ansatz = PUCCD(converter,num_particles,num_spin_orbitals,initial_state = init_state)
elif ansatz_type == "SUCCD":
ansatz = SUCCD(converter,num_particles,num_spin_orbitals,initial_state = init_state)
elif ansatz_type == "Custom":
# Example of how to write your own circuit
from qiskit.circuit import Parameter, QuantumCircuit, QuantumRegister
# Define the variational parameter
theta = Parameter('a')
n = qubit_op.num_qubits
# Make an empty quantum circuit
qc = QuantumCircuit(qubit_op.num_qubits)
qubit_label = 0
# Place a Hadamard gate
qc.h(qubit_label)
# Place a CNOT ladder
for i in range(n-1):
qc.cx(i, i+1)
# Visual separator
qc.barrier()
# rz rotations on all qubits
qc.rz(theta, range(n))
ansatz = qc
ansatz.compose(init_state, front=True, inplace=True)
print(ansatz)
```
#### 6. Backend
This is where you specify the simulator or device where you want to run your algorithm.
We will focus on the `statevector_simulator` in this challenge.
```
from qiskit import Aer
backend = Aer.get_backend('statevector_simulator')
```
#### 7. Optimizer
The optimizer guides the evolution of the parameters of the ansatz so it is very important to investigate the energy convergence as it would define the number of measurements that have to be performed on the QPU.
A clever choice might reduce drastically the number of needed energy evaluations.
```
from qiskit.algorithms.optimizers import COBYLA, L_BFGS_B, SPSA, SLSQP
optimizer_type = 'L_BFGS_B'
# You may want to tune the parameters
# of each optimizer, here the defaults are used
if optimizer_type == 'COBYLA':
optimizer = COBYLA(maxiter=500)
elif optimizer_type == 'L_BFGS_B':
optimizer = L_BFGS_B(maxfun=500)
elif optimizer_type == 'SPSA':
optimizer = SPSA(maxiter=500)
elif optimizer_type == 'SLSQP':
optimizer = SLSQP(maxiter=500)
```
#### 8. Exact eigensolver
For learning purposes, we can solve the problem exactly with the exact diagonalization of the Hamiltonian matrix so we know where to aim with VQE.
Of course, the dimensions of this matrix scale exponentially in the number of molecular orbitals so you can try doing this for a large molecule of your choice and see how slow this becomes.
For very large systems you would run out of memory trying to store their wavefunctions.
```
from qiskit_nature.algorithms.ground_state_solvers.minimum_eigensolver_factories import NumPyMinimumEigensolverFactory
from qiskit_nature.algorithms.ground_state_solvers import GroundStateEigensolver
import numpy as np
def exact_diagonalizer(problem, converter):
solver = NumPyMinimumEigensolverFactory()
calc = GroundStateEigensolver(converter, solver)
result = calc.solve(problem)
return result
result_exact = exact_diagonalizer(problem, converter)
exact_energy = np.real(result_exact.eigenenergies[0])
print("Exact electronic energy", exact_energy)
print(result_exact)
# The targeted electronic energy for H2 is -1.85336 Ha
# Check with your VQE result.
```
#### 9. VQE and initial parameters for the ansatz
Now we can import the VQE class and run the algorithm.
```
from qiskit.algorithms import VQE
from IPython.display import display, clear_output
# Print and save the data in lists
def callback(eval_count, parameters, mean, std):
# Overwrites the same line when printing
display("Evaluation: {}, Energy: {}, Std: {}".format(eval_count, mean, std))
clear_output(wait=True)
counts.append(eval_count)
values.append(mean)
params.append(parameters)
deviation.append(std)
counts = []
values = []
params = []
deviation = []
# Set initial parameters of the ansatz
# We choose a fixed small displacement
# So all participants start from similar starting point
try:
initial_point = [0.01] * len(ansatz.ordered_parameters)
except:
initial_point = [0.01] * ansatz.num_parameters
algorithm = VQE(ansatz,
optimizer=optimizer,
quantum_instance=backend,
callback=callback,
initial_point=initial_point)
result = algorithm.compute_minimum_eigenvalue(qubit_op)
print(result)
```
#### 9. Scoring function
We need to judge how good are your VQE simulations, your choice of ansatz/optimizer.
For this, we implemented the following simple scoring function:
$$ score = N_{CNOT}$$
where $N_{CNOT}$ is the number of CNOTs.
But you have to reach the chemical accuracy which is $\delta E_{chem} = 0.004$ Ha $= 4$ mHa, which may be hard to reach depending on the problem.
You have to reach the accuracy we set in a minimal number of CNOTs to win the challenge.
The lower the score the better!
```
# Store results in a dictionary
from qiskit.transpiler import PassManager
from qiskit.transpiler.passes import Unroller
# Unroller transpile your circuit into CNOTs and U gates
pass_ = Unroller(['u', 'cx'])
pm = PassManager(pass_)
ansatz_tp = pm.run(ansatz)
cnots = ansatz_tp.count_ops()['cx']
score = cnots
accuracy_threshold = 4.0 # in mHa
energy = result.optimal_value
if ansatz_type == "TwoLocal":
result_dict = {
'optimizer': optimizer.__class__.__name__,
'mapping': converter.mapper.__class__.__name__,
'ansatz': ansatz.__class__.__name__,
'rotation blocks': rotation_blocks,
'entanglement_blocks': entanglement_blocks,
'entanglement': entanglement,
'repetitions': repetitions,
'skip_final_rotation_layer': skip_final_rotation_layer,
'energy (Ha)': energy,
'error (mHa)': (energy-exact_energy)*1000,
'pass': (energy-exact_energy)*1000 <= accuracy_threshold,
'# of parameters': len(result.optimal_point),
'final parameters': result.optimal_point,
'# of evaluations': result.optimizer_evals,
'optimizer time': result.optimizer_time,
'# of qubits': int(qubit_op.num_qubits),
'# of CNOTs': cnots,
'score': score}
else:
result_dict = {
'optimizer': optimizer.__class__.__name__,
'mapping': converter.mapper.__class__.__name__,
'ansatz': ansatz.__class__.__name__,
'rotation blocks': None,
'entanglement_blocks': None,
'entanglement': None,
'repetitions': None,
'skip_final_rotation_layer': None,
'energy (Ha)': energy,
'error (mHa)': (energy-exact_energy)*1000,
'pass': (energy-exact_energy)*1000 <= accuracy_threshold,
'# of parameters': len(result.optimal_point),
'final parameters': result.optimal_point,
'# of evaluations': result.optimizer_evals,
'optimizer time': result.optimizer_time,
'# of qubits': int(qubit_op.num_qubits),
'# of CNOTs': cnots,
'score': score}
# Plot the results
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1, 1)
ax.set_xlabel('Iterations')
ax.set_ylabel('Energy')
ax.grid()
fig.text(0.7, 0.75, f'Energy: {result.optimal_value:.3f}\nScore: {score:.0f}')
plt.title(f"{result_dict['optimizer']}-{result_dict['mapping']}\n{result_dict['ansatz']}")
ax.plot(counts, values)
ax.axhline(exact_energy, linestyle='--')
fig_title = f"\
{result_dict['optimizer']}-\
{result_dict['mapping']}-\
{result_dict['ansatz']}-\
Energy({result_dict['energy (Ha)']:.3f})-\
Score({result_dict['score']:.0f})\
.png"
fig.savefig(fig_title, dpi=300)
# Display and save the data
import pandas as pd
import os.path
filename = 'results_h2.csv'
if os.path.isfile(filename):
result_df = pd.read_csv(filename)
result_df = result_df.append([result_dict])
else:
result_df = pd.DataFrame.from_dict([result_dict])
result_df.to_csv(filename)
result_df[['optimizer','ansatz', '# of qubits', '# of parameters','rotation blocks', 'entanglement_blocks',
'entanglement', 'repetitions', 'error (mHa)', 'pass', 'score']]
```
<div class="alert alert-block alert-danger">
<b>Tutorial questions 2</b>
Experiment with all the parameters and then:
1. Can you find your best (best score) heuristic ansatz (by modifying parameters of `TwoLocal` ansatz) and optimizer?
2. Can you find your best q-UCC ansatz (choose among `UCCSD, PUCCD or SUCCD` ansatzes) and optimizer?
3. In the cell where we define the ansatz,
can you modify the `Custom` ansatz by placing gates yourself to write a better circuit than your `TwoLocal` circuit?
For each question, give `ansatz` objects.
Remember, you have to reach the chemical accuracy $|E_{exact} - E_{VQE}| \leq 0.004 $ Ha $= 4$ mHa.
</div>
```
# WRITE YOUR CODE BETWEEN THESE LINES - START
# WRITE YOUR CODE BETWEEN THESE LINES - END
```
## Part 2: Final Challenge - VQE for LiH molecule
In this part, you will simulate LiH molecule using the STO-3G basis with the PySCF driver.
</div>
<div class="alert alert-block alert-success">
<b>Goal</b>
Experiment with all the parameters and then find your best ansatz. You can be as creative as you want!
For each question, give `ansatz` objects as for Part 1. Your final score will be based only on Part 2.
</div>
Be aware that the system is larger now. Work out how many qubits you would need for this system by retrieving the number of spin-orbitals.
### Reducing the problem size
You might want to reduce the number of qubits for your simulation:
- you could freeze the core electrons that do not contribute significantly to chemistry and consider only the valence electrons. Qiskit already has this functionality implemented. So inspect the different transformers in `qiskit_nature.transformers` and find the one that performs the freeze core approximation.
- you could use `ParityMapper` with `two_qubit_reduction=True` to eliminate 2 qubits.
- you could reduce the number of qubits by inspecting the symmetries of your Hamiltonian. Find a way to use `Z2Symmetries` in Qiskit.
### Custom ansatz
You might want to explore the ideas proposed in [Grimsley *et al.*,2018](https://arxiv.org/abs/1812.11173v2), [H. L. Tang *et al.*,2019](https://arxiv.org/abs/1911.10205), [Rattew *et al.*,2019](https://arxiv.org/abs/1910.09694), [Tang *et al.*,2019](https://arxiv.org/abs/1911.10205).
You can even get try machine learning algorithms to generate best ansatz circuits.
### Setup the simulation
Let's now run the Hartree-Fock calculation and the rest is up to you!
<div class="alert alert-block alert-danger">
<b>Attention</b>
We give below the `driver`, the `initial_point`, the `initial_state` that should remain as given.
You are free then to explore all other things available in Qiskit.
So you have to start from this initial point (all parameters set to 0.01):
`initial_point = [0.01] * len(ansatz.ordered_parameters)`
or
`initial_point = [0.01] * ansatz.num_parameters`
and your initial state has to be the Hartree-Fock state:
`init_state = HartreeFock(num_spin_orbitals, num_particles, converter)`
For each question, give `ansatz` object.
Remember you have to reach the chemical accuracy $|E_{exact} - E_{VQE}| \leq 0.004 $ Ha $= 4$ mHa.
</div>
```
from qiskit_nature.drivers import PySCFDriver
molecule = 'Li 0.0 0.0 0.0; H 0.0 0.0 1.5474'
driver = PySCFDriver(atom=molecule)
qmolecule = driver.run()
# WRITE YOUR CODE BETWEEN THESE LINES - START
from qiskit_nature.problems.second_quantization.electronic import ElectronicStructureProblem
from qiskit_nature.transformers import FreezeCoreTransformer
from qiskit_nature.mappers.second_quantization import ParityMapper
from qiskit_nature.converters.second_quantization.qubit_converter import QubitConverter
from qiskit_nature.circuit.library import HartreeFock
from qiskit_nature.circuit.library import UCCSD
from qiskit.circuit.library import TwoLocal
from qiskit import Aer
from qiskit.algorithms.optimizers import COBYLA
from qiskit.algorithms.optimizers import SPSA
from qiskit.algorithms.optimizers import L_BFGS_B
from qiskit_nature.algorithms.ground_state_solvers.minimum_eigensolver_factories import NumPyMinimumEigensolverFactory
from qiskit_nature.algorithms.ground_state_solvers import GroundStateEigensolver
import numpy as np
backend = Aer.get_backend('statevector_simulator')
n_el = qmolecule.num_alpha + qmolecule.num_beta
n_mo = qmolecule.num_molecular_orbitals
n_so = 2 * qmolecule.num_molecular_orbitals
n_q = 2* qmolecule.num_molecular_orbitals
e_nn = qmolecule.nuclear_repulsion_energy
############################################################
freezeCoreTransformer = FreezeCoreTransformer()
qmolecule = freezeCoreTransformer.transform(qmolecule)
problem = ElectronicStructureProblem(driver,q_molecule_transformers=[FreezeCoreTransformer(freeze_core=True,remove_orbitals=[3])])
#problem = ElectronicStructureProblem(driver)
# Generate the second-quantized operators
second_q_ops = problem.second_q_ops()
# Hamiltonian
main_op = second_q_ops[0]
############################################################
# Setup the mapper and qubit converter
converter = QubitConverter(mapper=ParityMapper(), two_qubit_reduction=True,z2symmetry_reduction=[1])
#converter = QubitConverter(mapper=mapper, two_qubit_reduction=True)
# The fermionic operators are mapped to qubit operators
num_particles = (problem.molecule_data_transformed.num_alpha,
problem.molecule_data_transformed.num_beta)
qubit_op = converter.convert(main_op, num_particles=num_particles)
###########################################################
num_particles = (problem.molecule_data_transformed.num_alpha,
problem.molecule_data_transformed.num_beta)
num_spin_orbitals = 2 * problem.molecule_data_transformed.num_molecular_orbitals
init_state = HartreeFock(num_spin_orbitals, num_particles, converter)
#print(init_state)
###########################################################
ansatz = TwoLocal(converter,num_particles,num_spin_orbitals,initial_state = init_state)
#print(ansatz)
###########################################################
optimizer = L_BFGS_B(maxfun=500)
###########################################################
def exact_diagonalizer(problem, converter):
solver = NumPyMinimumEigensolverFactory()
calc = GroundStateEigensolver(converter, solver)
result = calc.solve(problem)
return result
result_exact = exact_diagonalizer(problem, converter)
exact_energy = np.real(result_exact.eigenenergies[0])
#print("Exact electronic energy", exact_energy)
#print(result_exact)
########################################################
from qiskit.algorithms import VQE
from IPython.display import display, clear_output
# Print and save the data in lists
def callback(eval_count, parameters, mean, std):
# Overwrites the same line when printing
display("Evaluation: {}, Energy: {}, Std: {}".format(eval_count, mean, std))
clear_output(wait=True)
counts.append(eval_count)
values.append(mean)
params.append(parameters)
deviation.append(std)
counts = []
values = []
params = []
deviation = []
# Set initial parameters of the ansatz
# We choose a fixed small displacement
# So all participants start from similar starting point
try:
initial_point = [0.01] * len(ansatz.ordered_parameters)
except:
initial_point = [0.01] * ansatz.num_parameters
algorithm = VQE(ansatz,
optimizer=optimizer,
quantum_instance=backend,
callback=callback,
initial_point=initial_point)
result = algorithm.compute_minimum_eigenvalue(qubit_op)
print(result)
# WRITE YOUR CODE BETWEEN THESE LINES - END
# Check your answer using following code
from qc_grader import grade_ex5
freeze_core = True # change to True if you freezed core electrons
grade_ex5(ansatz,qubit_op,result,freeze_core)
# Submit your answer. You can re-submit at any time.
from qc_grader import submit_ex5
submit_ex5(ansatz,qubit_op,result,freeze_core)
```
## Answers for Part 1
<div class="alert alert-block alert-danger">
<b>Questions</b>
Look into the attributes of `qmolecule` and answer the questions below.
1. We need to know the basic characteristics of our molecule. What is the total number of electrons in your system?
2. What is the number of molecular orbitals?
3. What is the number of spin-orbitals?
3. How many qubits would you need to simulate this molecule with Jordan-Wigner mapping?
5. What is the value of the nuclear repulsion energy?
</div>
<div class="alert alert-block alert-success">
<b>Answers </b>
1. `n_el = qmolecule.num_alpha + qmolecule.num_beta`
2. `n_mo = qmolecule.num_molecular_orbitals`
3. `n_so = 2 * qmolecule.num_molecular_orbitals`
4. `n_q = 2* qmolecule.num_molecular_orbitals`
5. `e_nn = qmolecule.nuclear_repulsion_energy`
</div>
## Additional information
**Created by:** Igor Sokolov, Junye Huang, Rahul Pratap Singh
**Version:** 1.0.1
| github_jupyter |
## Import Statements
```
import pandas as pd
import numpy as np
```
## Define functions
```
np.random.seed(seed=123456)
def get_initial_center_indices(size=None, k=None):
'''
size: total rows in data ( = highest index + 1)
k = number of clusters
'''
return np.random.choice(a=size, replace=False, size=k)
def calculate_distance(a=None, b=None):
'''
a, b : numpy arrays of same length
'''
return np.sqrt(np.sum((a - b) ** 2))
def get_distance_from_each_center(x=None, centers=None):
'''
x : numpy array for the datapoint
centers : pandas dataframe where each row represents center
This function calculates distance of point x from all centers.
'''
return centers.apply(calculate_distance, axis='columns', b=x)
def get_nearest_clusters_index(x=None, centers=None):
'''
x : numpy array for the datapoint
centers : pandas dataframe where each row represents center
This function calculates distance of point x from all centers.
'''
return np.argmin(get_distance_from_each_center(x, centers).values)
```
## Prepare data
```
# To download data file. Visit : https://raw.githubusercontent.com/mubaris/friendly-fortnight/master/xclara.csv
data = pd.read_csv("~/Downloads/xclara.csv", header=0)
data = data.sample(n=100, random_state=12345)
data.reset_index(inplace=True, drop=True)
```
## Inefficient KMeans!
```
def kmeans_my_1(data=None, k=3, max_iter=200):
import copy
centers = data.loc[np.random.choice(data.shape[0], replace=False, size=k)].reset_index(inplace=False, drop=True)
cur_iter = 0
shouldIterate = True
while shouldIterate:
clusters = data.apply(get_nearest_clusters_index, axis='columns', centers=centers)
old_centers = copy.deepcopy(centers)
# update centers
for i in range(k):
centers.loc[i] = data.loc[clusters == i].mean()
cur_iter += 1
shouldIterate = any(np.sqrt(np.sum((centers - old_centers) ** 2)) > 1) and cur_iter < max_iter
print(centers)
print(cur_iter)
```
## Our solution is accurate, but sklearn is FAST!
~50xfaster!
```
from sklearn.cluster import KMeans
from datetime import datetime
km = KMeans(n_clusters=3)
start = datetime.now()
km.fit(data)
sklearn_time = datetime.now() - start
print("sklearn time : {}".format(sklearn_time))
print(km.cluster_centers_)
start = datetime.now()
kmeans_my_1(data=data, k=3)
our_time = datetime.now() - start
print("solution 1 time : {}".format(our_time))
print("sklearn is faster by {0:.0f}x".format(our_time / sklearn_time))
```
### Let us try to reduce use of dataframe apply function.
```
def kmeans_my_2(data=None, k=3, max_iter=200):
import copy
centers = data.loc[np.random.choice(data.shape[0], replace=False, size=k)].values
clusters = np.array([-1] * data.shape[0])
cur_iter = 0
shouldIterate = True
while shouldIterate:
for nth_row in range(data.shape[0]):
data_row = data.loc[nth_row]
nearest_cluster = -1
min_distance = float("inf")
for cluster_num in range(k):
distance = np.sum((centers[cluster_num] - data_row) ** 2)
if distance < min_distance:
nearest_cluster = cluster_num
min_distance = distance
clusters[nth_row] = nearest_cluster
old_centers = copy.deepcopy(centers)
# update centers
for i in range(k):
centers[i] = data.loc[clusters == i].mean().values
cur_iter += 1
if cur_iter >= max_iter:
break
shouldIterate = False
for i in range(centers.shape[0]):
if np.sum((centers[i] - old_centers[i]) ** 2) > 1:
shouldIterate = True
break
print(centers)
print(cur_iter)
```
## Still sklearn is ~25x faster
```
from sklearn.cluster import KMeans
from datetime import datetime
km = KMeans(n_clusters=3)
start = datetime.now()
km.fit(data)
sklearn_time = datetime.now() - start
print("sklearn time : {}".format(sklearn_time))
print(km.cluster_centers_)
start = datetime.now()
kmeans_my_2(data=data, k=3)
our_time = datetime.now() - start
print("solution 2 time : {}".format(our_time))
print("sklearn is faster by {0:.0f}x".format(our_time / sklearn_time))
```
### Since clusters calculation is bottle neck, let us use numpy there.
```
def kmeans_my_3(data=None, k=3, max_iter=200):
import copy
max_iter = 200
k=3
centers = data.loc[np.random.choice(data.shape[0], replace=False, size=k)].values
cur_iter = 0
shouldIterate = True
clusters = np.array([-1] * data.shape[0])
while shouldIterate:
for nth_row in range(data.shape[0]):
clusters[nth_row] = np.argmin(np.sum((centers - data.loc[nth_row].values) ** 2, axis=1))
old_centers = copy.deepcopy(centers)
# update centers
for i in range(k):
centers[i] = data.loc[clusters == i].mean().values
cur_iter += 1
if cur_iter >= max_iter:
break
shouldIterate = any(np.sum((centers - old_centers) ** 2, axis=1) > 1)
print(centers)
print(cur_iter)
```
### With the use of numpy for norm, we are now ~5x slower than sklearn
```
from sklearn.cluster import KMeans
from datetime import datetime
km = KMeans(n_clusters=3)
start = datetime.now()
km.fit(data)
sklearn_time = datetime.now() - start
print("sklearn time : {}".format(sklearn_time))
print(km.cluster_centers_)
start = datetime.now()
kmeans_my_3(data=data, k=3)
our_time = datetime.now() - start
print("solution 3 time : {}".format(our_time))
print("sklearn is faster by {0:.0f}x".format(our_time / sklearn_time))
```
| github_jupyter |
# Linear Regression Implementation from Scratch
:label:`sec_linear_scratch`
Now that you understand the key ideas behind linear regression,
we can begin to work through a hands-on implementation in code.
In this section, (**we will implement the entire method from scratch,
including the data pipeline, the model,
the loss function, and the minibatch stochastic gradient descent optimizer.**)
While modern deep learning frameworks can automate nearly all of this work,
implementing things from scratch is the only way
to make sure that you really know what you are doing.
Moreover, when it comes time to customize models,
defining our own layers or loss functions,
understanding how things work under the hood will prove handy.
In this section, we will rely only on tensors and auto differentiation.
Afterwards, we will introduce a more concise implementation,
taking advantage of bells and whistles of deep learning frameworks.
```
%matplotlib inline
import random
from mxnet import autograd, np, npx
from d2l import mxnet as d2l
npx.set_np()
```
## Generating the Dataset
To keep things simple, we will [**construct an artificial dataset
according to a linear model with additive noise.**]
Our task will be to recover this model's parameters
using the finite set of examples contained in our dataset.
We will keep the data low-dimensional so we can visualize it easily.
In the following code snippet, we generate a dataset
containing 1000 examples, each consisting of 2 features
sampled from a standard normal distribution.
Thus our synthetic dataset will be a matrix
$\mathbf{X}\in \mathbb{R}^{1000 \times 2}$.
(**The true parameters generating our dataset will be
$\mathbf{w} = [2, -3.4]^\top$ and $b = 4.2$,
and**) our synthetic labels will be assigned according
to the following linear model with the noise term $\epsilon$:
(**$$\mathbf{y}= \mathbf{X} \mathbf{w} + b + \mathbf\epsilon.$$**)
You could think of $\epsilon$ as capturing potential
measurement errors on the features and labels.
We will assume that the standard assumptions hold and thus
that $\epsilon$ obeys a normal distribution with mean of 0.
To make our problem easy, we will set its standard deviation to 0.01.
The following code generates our synthetic dataset.
```
def synthetic_data(w, b, num_examples): #@save
"""Generate y = Xw + b + noise."""
X = np.random.normal(0, 1, (num_examples, len(w)))
y = np.dot(X, w) + b
y += np.random.normal(0, 0.01, y.shape)
return X, y.reshape((-1, 1))
true_w = np.array([2, -3.4])
true_b = 4.2
features, labels = synthetic_data(true_w, true_b, 1000)
```
Note that [**each row in `features` consists of a 2-dimensional data example
and that each row in `labels` consists of a 1-dimensional label value (a scalar).**]
```
print('features:', features[0], '\nlabel:', labels[0])
```
By generating a scatter plot using the second feature `features[:, 1]` and `labels`,
we can clearly observe the linear correlation between the two.
```
d2l.set_figsize()
# The semicolon is for displaying the plot only
d2l.plt.scatter(features[:, (1)].asnumpy(), labels.asnumpy(), 1);
```
## Reading the Dataset
Recall that training models consists of
making multiple passes over the dataset,
grabbing one minibatch of examples at a time,
and using them to update our model.
Since this process is so fundamental
to training machine learning algorithms,
it is worth defining a utility function
to shuffle the dataset and access it in minibatches.
In the following code, we [**define the `data_iter` function**] (~~that~~)
to demonstrate one possible implementation of this functionality.
The function (**takes a batch size, a matrix of features,
and a vector of labels, yielding minibatches of the size `batch_size`.**)
Each minibatch consists of a tuple of features and labels.
```
def data_iter(batch_size, features, labels):
num_examples = len(features)
indices = list(range(num_examples))
# The examples are read at random, in no particular order
random.shuffle(indices)
for i in range(0, num_examples, batch_size):
batch_indices = np.array(indices[i:min(i + batch_size, num_examples)])
yield features[batch_indices], labels[batch_indices]
```
In general, note that we want to use reasonably sized minibatches
to take advantage of the GPU hardware,
which excels at parallelizing operations.
Because each example can be fed through our models in parallel
and the gradient of the loss function for each example can also be taken in parallel,
GPUs allow us to process hundreds of examples in scarcely more time
than it might take to process just a single example.
To build some intuition, let us read and print
the first small batch of data examples.
The shape of the features in each minibatch tells us
both the minibatch size and the number of input features.
Likewise, our minibatch of labels will have a shape given by `batch_size`.
```
batch_size = 10
for X, y in data_iter(batch_size, features, labels):
print(X, '\n', y)
break
```
As we run the iteration, we obtain distinct minibatches
successively until the entire dataset has been exhausted (try this).
While the iteration implemented above is good for didactic purposes,
it is inefficient in ways that might get us in trouble on real problems.
For example, it requires that we load all the data in memory
and that we perform lots of random memory access.
The built-in iterators implemented in a deep learning framework
are considerably more efficient and they can deal
with both data stored in files and data fed via data streams.
## Initializing Model Parameters
[**Before we can begin optimizing our model's parameters**] by minibatch stochastic gradient descent,
(**we need to have some parameters in the first place.**)
In the following code, we initialize weights by sampling
random numbers from a normal distribution with mean 0
and a standard deviation of 0.01, and setting the bias to 0.
```
w = np.random.normal(0, 0.01, (2, 1))
b = np.zeros(1)
w.attach_grad()
b.attach_grad()
```
After initializing our parameters,
our next task is to update them until
they fit our data sufficiently well.
Each update requires taking the gradient
of our loss function with respect to the parameters.
Given this gradient, we can update each parameter
in the direction that may reduce the loss.
Since nobody wants to compute gradients explicitly
(this is tedious and error prone),
we use automatic differentiation,
as introduced in :numref:`sec_autograd`, to compute the gradient.
## Defining the Model
Next, we must [**define our model,
relating its inputs and parameters to its outputs.**]
Recall that to calculate the output of the linear model,
we simply take the matrix-vector dot product
of the input features $\mathbf{X}$ and the model weights $\mathbf{w}$,
and add the offset $b$ to each example.
Note that below $\mathbf{Xw}$ is a vector and $b$ is a scalar.
Recall the broadcasting mechanism as described in :numref:`subsec_broadcasting`.
When we add a vector and a scalar,
the scalar is added to each component of the vector.
```
def linreg(X, w, b): #@save
"""The linear regression model."""
return np.dot(X, w) + b
```
## Defining the Loss Function
Since [**updating our model requires taking
the gradient of our loss function,**]
we ought to (**define the loss function first.**)
Here we will use the squared loss function
as described in :numref:`sec_linear_regression`.
In the implementation, we need to transform the true value `y`
into the predicted value's shape `y_hat`.
The result returned by the following function
will also have the same shape as `y_hat`.
```
def squared_loss(y_hat, y): #@save
"""Squared loss."""
return (y_hat - y.reshape(y_hat.shape))**2 / 2
```
## Defining the Optimization Algorithm
As we discussed in :numref:`sec_linear_regression`,
linear regression has a closed-form solution.
However, this is not a book about linear regression:
it is a book about deep learning.
Since none of the other models that this book introduces
can be solved analytically, we will take this opportunity to introduce your first working example of
minibatch stochastic gradient descent.
[~~Despite linear regression has a closed-form solution, other models in this book don't. Here we introduce minibatch stochastic gradient descent.~~]
At each step, using one minibatch randomly drawn from our dataset,
we will estimate the gradient of the loss with respect to our parameters.
Next, we will update our parameters
in the direction that may reduce the loss.
The following code applies the minibatch stochastic gradient descent update,
given a set of parameters, a learning rate, and a batch size.
The size of the update step is determined by the learning rate `lr`.
Because our loss is calculated as a sum over the minibatch of examples,
we normalize our step size by the batch size (`batch_size`),
so that the magnitude of a typical step size
does not depend heavily on our choice of the batch size.
```
def sgd(params, lr, batch_size): #@save
"""Minibatch stochastic gradient descent."""
for param in params:
param[:] = param - lr * param.grad / batch_size
```
## Training
Now that we have all of the parts in place,
we are ready to [**implement the main training loop.**]
It is crucial that you understand this code
because you will see nearly identical training loops
over and over again throughout your career in deep learning.
In each iteration, we will grab a minibatch of training examples,
and pass them through our model to obtain a set of predictions.
After calculating the loss, we initiate the backwards pass through the network,
storing the gradients with respect to each parameter.
Finally, we will call the optimization algorithm `sgd`
to update the model parameters.
In summary, we will execute the following loop:
* Initialize parameters $(\mathbf{w}, b)$
* Repeat until done
* Compute gradient $\mathbf{g} \leftarrow \partial_{(\mathbf{w},b)} \frac{1}{|\mathcal{B}|} \sum_{i \in \mathcal{B}} l(\mathbf{x}^{(i)}, y^{(i)}, \mathbf{w}, b)$
* Update parameters $(\mathbf{w}, b) \leftarrow (\mathbf{w}, b) - \eta \mathbf{g}$
In each *epoch*,
we will iterate through the entire dataset
(using the `data_iter` function) once
passing through every example in the training dataset
(assuming that the number of examples is divisible by the batch size).
The number of epochs `num_epochs` and the learning rate `lr` are both hyperparameters,
which we set here to 3 and 0.03, respectively.
Unfortunately, setting hyperparameters is tricky
and requires some adjustment by trial and error.
We elide these details for now but revise them
later in
:numref:`chap_optimization`.
```
lr = 0.03
num_epochs = 3
net = linreg
loss = squared_loss
for epoch in range(num_epochs):
for X, y in data_iter(batch_size, features, labels):
with autograd.record():
l = loss(net(X, w, b), y) # Minibatch loss in `X` and `y`
# Because `l` has a shape (`batch_size`, 1) and is not a scalar
# variable, the elements in `l` are added together to obtain a new
# variable, on which gradients with respect to [`w`, `b`] are computed
l.backward()
sgd([w, b], lr, batch_size) # Update parameters using their gradient
train_l = loss(net(features, w, b), labels)
print(f'epoch {epoch + 1}, loss {float(train_l.mean()):f}')
```
In this case, because we synthesized the dataset ourselves,
we know precisely what the true parameters are.
Thus, we can [**evaluate our success in training
by comparing the true parameters
with those that we learned**] through our training loop.
Indeed they turn out to be very close to each other.
```
print(f'error in estimating w: {true_w - w.reshape(true_w.shape)}')
print(f'error in estimating b: {true_b - b}')
```
Note that we should not take it for granted
that we are able to recover the parameters perfectly.
However, in machine learning, we are typically less concerned
with recovering true underlying parameters,
and more concerned with parameters that lead to highly accurate prediction.
Fortunately, even on difficult optimization problems,
stochastic gradient descent can often find remarkably good solutions,
owing partly to the fact that, for deep networks,
there exist many configurations of the parameters
that lead to highly accurate prediction.
## Summary
* We saw how a deep network can be implemented and optimized from scratch, using just tensors and auto differentiation, without any need for defining layers or fancy optimizers.
* This section only scratches the surface of what is possible. In the following sections, we will describe additional models based on the concepts that we have just introduced and learn how to implement them more concisely.
## Exercises
1. What would happen if we were to initialize the weights to zero. Would the algorithm still work?
1. Assume that you are
[Georg Simon Ohm](https://en.wikipedia.org/wiki/Georg_Ohm) trying to come up
with a model between voltage and current. Can you use auto differentiation to learn the parameters of your model?
1. Can you use [Planck's Law](https://en.wikipedia.org/wiki/Planck%27s_law) to determine the temperature of an object using spectral energy density?
1. What are the problems you might encounter if you wanted to compute the second derivatives? How would you fix them?
1. Why is the `reshape` function needed in the `squared_loss` function?
1. Experiment using different learning rates to find out how fast the loss function value drops.
1. If the number of examples cannot be divided by the batch size, what happens to the `data_iter` function's behavior?
[Discussions](https://discuss.d2l.ai/t/42)
| github_jupyter |
# Skip-gram Word2Vec
In this notebook, I'll lead you through using PyTorch to implement the [Word2Vec algorithm](https://en.wikipedia.org/wiki/Word2vec) using the skip-gram architecture. By implementing this, you'll learn about embedding words for use in natural language processing. This will come in handy when dealing with things like machine translation.
## Readings
Here are the resources I used to build this notebook. I suggest reading these either beforehand or while you're working on this material.
* A really good [conceptual overview](http://mccormickml.com/2016/04/19/word2vec-tutorial-the-skip-gram-model/) of Word2Vec from Chris McCormick
* [First Word2Vec paper](https://arxiv.org/pdf/1301.3781.pdf) from Mikolov et al.
* [Neural Information Processing Systems, paper](http://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf) with improvements for Word2Vec also from Mikolov et al.
---
## Word embeddings
When you're dealing with words in text, you end up with tens of thousands of word classes to analyze; one for each word in a vocabulary. Trying to one-hot encode these words is massively inefficient because most values in a one-hot vector will be set to zero. So, the matrix multiplication that happens in between a one-hot input vector and a first, hidden layer will result in mostly zero-valued hidden outputs.
<img src='assets/one_hot_encoding.png' width=50%>
To solve this problem and greatly increase the efficiency of our networks, we use what are called **embeddings**. Embeddings are just a fully connected layer like you've seen before. We call this layer the embedding layer and the weights are embedding weights. We skip the multiplication into the embedding layer by instead directly grabbing the hidden layer values from the weight matrix. We can do this because the multiplication of a one-hot encoded vector with a matrix returns the row of the matrix corresponding the index of the "on" input unit.
<img src='assets/lookup_matrix.png' width=50%>
Instead of doing the matrix multiplication, we use the weight matrix as a lookup table. We encode the words as integers, for example "heart" is encoded as 958, "mind" as 18094. Then to get hidden layer values for "heart", you just take the 958th row of the embedding matrix. This process is called an **embedding lookup** and the number of hidden units is the **embedding dimension**.
<img src='assets/tokenize_lookup.png' width=50%>
There is nothing magical going on here. The embedding lookup table is just a weight matrix. The embedding layer is just a hidden layer. The lookup is just a shortcut for the matrix multiplication. The lookup table is trained just like any weight matrix.
Embeddings aren't only used for words of course. You can use them for any model where you have a massive number of classes. A particular type of model called **Word2Vec** uses the embedding layer to find vector representations of words that contain semantic meaning.
---
## Word2Vec
The Word2Vec algorithm finds much more efficient representations by finding vectors that represent the words. These vectors also contain semantic information about the words.
<img src="assets/context_drink.png" width=40%>
Words that show up in similar **contexts**, such as "coffee", "tea", and "water" will have vectors near each other. Different words will be further away from one another, and relationships can be represented by distance in vector space.
<img src="assets/vector_distance.png" width=40%>
There are two architectures for implementing Word2Vec:
>* CBOW (Continuous Bag-Of-Words) and
* Skip-gram
<img src="assets/word2vec_architectures.png" width=60%>
In this implementation, we'll be using the **skip-gram architecture** because it performs better than CBOW. Here, we pass in a word and try to predict the words surrounding it in the text. In this way, we can train the network to learn representations for words that show up in similar contexts.
---
## Loading Data
Next, we'll ask you to load in data and place it in the `data` directory
1. Load the [text8 dataset](https://s3.amazonaws.com/video.udacity-data.com/topher/2018/October/5bbe6499_text8/text8.zip); a file of cleaned up *Wikipedia article text* from Matt Mahoney.
2. Place that data in the `data` folder in the home directory.
3. Then you can extract it and delete the archive, zip file to save storage space.
After following these steps, you should have one file in your data directory: `data/text8`.
```
# read in the extracted text file
with open('data/text8') as f:
text = f.read()
# print out the first 100 characters
print(text[:100])
```
## Pre-processing
Here I'm fixing up the text to make training easier. This comes from the `utils.py` file. The `preprocess` function does a few things:
>* It converts any punctuation into tokens, so a period is changed to ` <PERIOD> `. In this data set, there aren't any periods, but it will help in other NLP problems.
* It removes all words that show up five or *fewer* times in the dataset. This will greatly reduce issues due to noise in the data and improve the quality of the vector representations.
* It returns a list of words in the text.
This may take a few seconds to run, since our text file is quite large. If you want to write your own functions for this stuff, go for it!
```
import utils
# get list of words
words = utils.preprocess(text)
print(words[:30])
# print some stats about this word data
print("Total words in text: {}".format(len(words)))
print("Unique words: {}".format(len(set(words)))) # `set` removes any duplicate words
```
### Dictionaries
Next, I'm creating two dictionaries to convert words to integers and back again (integers to words). This is again done with a function in the `utils.py` file. `create_lookup_tables` takes in a list of words in a text and returns two dictionaries.
>* The integers are assigned in descending frequency order, so the most frequent word ("the") is given the integer 0 and the next most frequent is 1, and so on.
Once we have our dictionaries, the words are converted to integers and stored in the list `int_words`.
```
vocab_to_int, int_to_vocab = utils.create_lookup_tables(words)
int_words = [vocab_to_int[word] for word in words]
print(int_words[:30])
```
## Subsampling
Words that show up often such as "the", "of", and "for" don't provide much context to the nearby words. If we discard some of them, we can remove some of the noise from our data and in return get faster training and better representations. This process is called subsampling by Mikolov. For each word $w_i$ in the training set, we'll discard it with probability given by
$$ P(w_i) = 1 - \sqrt{\frac{t}{f(w_i)}} $$
where $t$ is a threshold parameter and $f(w_i)$ is the frequency of word $w_i$ in the total dataset.
$$ P(0) = 1 - \sqrt{\frac{1*10^{-5}}{1*10^6/16*10^6}} = 0.98735 $$
I'm going to leave this up to you as an exercise. Check out my solution to see how I did it.
> **Exercise:** Implement subsampling for the words in `int_words`. That is, go through `int_words` and discard each word given the probablility $P(w_i)$ shown above. Note that $P(w_i)$ is the probability that a word is discarded. Assign the subsampled data to `train_words`.
```
from collections import Counter
import random
import numpy as np
threshold = 1e-5
word_counts = Counter(int_words)
print(list(word_counts.items())[0]) # dictionary of int_words, how many times they appear
# discard some frequent words, according to the subsampling equation
# create a new list of words for training
def get_train_words(int_words, word_counts, threshold):
total_words = len(int_words)
drop_prob= {}
for word, count in word_counts.items():
drop_prob[word] = 1 - np.sqrt(threshold/(count/total_words))
train_words = [word for word in int_words if random.random() < (1 - drop_prob[word])]
return train_words
train_words = get_train_words(int_words, word_counts, threshold)
print(train_words[:30])
```
## Making batches
Now that our data is in good shape, we need to get it into the proper form to pass it into our network. With the skip-gram architecture, for each word in the text, we want to define a surrounding _context_ and grab all the words in a window around that word, with size $C$.
From [Mikolov et al.](https://arxiv.org/pdf/1301.3781.pdf):
"Since the more distant words are usually less related to the current word than those close to it, we give less weight to the distant words by sampling less from those words in our training examples... If we choose $C = 5$, for each training word we will select randomly a number $R$ in range $[ 1: C ]$, and then use $R$ words from history and $R$ words from the future of the current word as correct labels."
> **Exercise:** Implement a function `get_target` that receives a list of words, an index, and a window size, then returns a list of words in the window around the index. Make sure to use the algorithm described above, where you chose a random number of words to from the window.
Say, we have an input and we're interested in the idx=2 token, `741`:
```
[5233, 58, 741, 10571, 27349, 0, 15067, 58112, 3580, 58, 10712]
```
For `R=2`, `get_target` should return a list of four values:
```
[5233, 58, 10571, 27349]
```
```
def get_target(words, idx, window_size=5):
''' Get a list of words in a window around an index. '''
# choose random number between 1 and window size
R = np.random.randint(1, window_size+1)
# define start and end indexes
start = idx - R if (idx - R) > 0 else 0
end = idx + R
target_words = words[start:idx] + words[idx+1:end+1]
return list(target_words)
# test your code!
# run this cell multiple times to check for random window selection
int_text = [i for i in range(10)]
print('Input: ', int_text)
idx=5 # word index of interest
target = get_target(int_text, idx=idx, window_size=5)
print('Target: ', target) # you should get some indices around the idx
```
### Generating Batches
Here's a generator function that returns batches of input and target data for our model, using the `get_target` function from above. The idea is that it grabs `batch_size` words from a words list. Then for each of those batches, it gets the target words in a window.
```
def get_batches(words, batch_size, window_size=5):
''' Create a generator of word batches as a tuple (inputs, targets) '''
n_batches = len(words)//batch_size
# only full batches
words = words[:n_batches*batch_size]
for idx in range(0, len(words), batch_size):
x, y = [], []
batch = words[idx:idx+batch_size]
for ii in range(len(batch)):
batch_x = batch[ii]
batch_y = get_target(batch, ii, window_size)
y.extend(batch_y)
x.extend([batch_x]*len(batch_y))
yield x, y
int_text = [i for i in range(20)]
x,y = next(get_batches(int_text, batch_size=4, window_size=5))
print('x\n', x)
print('y\n', y)
```
## Building the graph
Below is an approximate diagram of the general structure of our network.
<img src="assets/skip_gram_arch.png" width=60%>
>* The input words are passed in as batches of input word tokens.
* This will go into a hidden layer of linear units (our embedding layer).
* Then, finally into a softmax output layer.
We'll use the softmax layer to make a prediction about the context words by sampling, as usual.
The idea here is to train the embedding layer weight matrix to find efficient representations for our words. We can discard the softmax layer because we don't really care about making predictions with this network. We just want the embedding matrix so we can use it in _other_ networks we build using this dataset.
---
## Validation
Here, I'm creating a function that will help us observe our model as it learns. We're going to choose a few common words and few uncommon words. Then, we'll print out the closest words to them using the cosine similarity:
<img src="assets/two_vectors.png" width=30%>
$$
\mathrm{similarity} = \cos(\theta) = \frac{\vec{a} \cdot \vec{b}}{|\vec{a}||\vec{b}|}
$$
We can encode the validation words as vectors $\vec{a}$ using the embedding table, then calculate the similarity with each word vector $\vec{b}$ in the embedding table. With the similarities, we can print out the validation words and words in our embedding table semantically similar to those words. It's a nice way to check that our embedding table is grouping together words with similar semantic meanings.
```
def cosine_similarity(embedding, valid_size=16, valid_window=100, device='cpu'):
""" Returns the cosine similarity of validation words with words in the embedding matrix.
Here, embedding should be a PyTorch embedding module.
"""
# Here we're calculating the cosine similarity between some random words and
# our embedding vectors. With the similarities, we can look at what words are
# close to our random words.
# sim = (a . b) / |a||b|
embed_vectors = embedding.weight
# magnitude of embedding vectors, |b|
magnitudes = embed_vectors.pow(2).sum(dim=1).sqrt().unsqueeze(0)
# pick N words from our ranges (0,window) and (1000,1000+window). lower id implies more frequent
valid_examples = np.array(random.sample(range(valid_window), valid_size//2))
valid_examples = np.append(valid_examples,
random.sample(range(1000,1000+valid_window), valid_size//2))
valid_examples = torch.LongTensor(valid_examples).to(device)
valid_vectors = embedding(valid_examples)
similarities = torch.mm(valid_vectors, embed_vectors.t())/magnitudes
return valid_examples, similarities
```
## SkipGram model
Define and train the SkipGram model.
> You'll need to define an [embedding layer](https://pytorch.org/docs/stable/nn.html#embedding) and a final, softmax output layer.
An Embedding layer takes in a number of inputs, importantly:
* **num_embeddings** – the size of the dictionary of embeddings, or how many rows you'll want in the embedding weight matrix
* **embedding_dim** – the size of each embedding vector; the embedding dimension
```
import torch
from torch import nn
import torch.optim as optim
class SkipGram(nn.Module):
def __init__(self, n_vocab, n_embed):
super().__init__()
# complete SkipGram model
self.embed = nn.Embedding(n_vocab, n_embed)
self.output = nn.Linear(n_embed, n_vocab)
self.log_softmax = nn.LogSoftmax(dim=1)
def forward(self, x):
# define the forward behavior
x = self.embed(x)
x = self.output(x)
x = self.log_softmax(x)
return x
```
### Training
Below is our training loop, and I recommend that you train on GPU, if available.
**Note that, because we applied a softmax function to our model output, we are using NLLLoss** as opposed to cross entropy. This is because Softmax in combination with NLLLoss = CrossEntropy loss .
```
# check if GPU is available
device = 'cuda' if torch.cuda.is_available() else 'cpu'
embedding_dim=300 # you can change, if you want
model = SkipGram(len(vocab_to_int), embedding_dim).to(device)
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.parameters(), lr=0.003)
print_every = 500
steps = 0
epochs = 2
# train for some number of epochs
for e in range(epochs):
# get input and target batches
for inputs, targets in get_batches(train_words, 512):
steps += 1
inputs, targets = torch.LongTensor(inputs), torch.LongTensor(targets)
inputs, targets = inputs.to(device), targets.to(device)
log_ps = model(inputs)
loss = criterion(log_ps, targets)
optimizer.zero_grad()
loss.backward()
optimizer.step()
if steps % print_every == 0:
# getting examples and similarities
valid_examples, valid_similarities = cosine_similarity(model.embed, device=device)
_, closest_idxs = valid_similarities.topk(6) # topk highest similarities
valid_examples, closest_idxs = valid_examples.to('cpu'), closest_idxs.to('cpu')
for ii, valid_idx in enumerate(valid_examples):
closest_words = [int_to_vocab[idx.item()] for idx in closest_idxs[ii]][1:]
print(int_to_vocab[valid_idx.item()] + " | " + ', '.join(closest_words))
print("...")
```
## Visualizing the word vectors
Below we'll use T-SNE to visualize how our high-dimensional word vectors cluster together. T-SNE is used to project these vectors into two dimensions while preserving local stucture. Check out [this post from Christopher Olah](http://colah.github.io/posts/2014-10-Visualizing-MNIST/) to learn more about T-SNE and other ways to visualize high-dimensional data.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
# getting embeddings from the embedding layer of our model, by name
embeddings = model.embed.weight.to('cpu').data.numpy()
viz_words = 600
tsne = TSNE()
embed_tsne = tsne.fit_transform(embeddings[:viz_words, :])
fig, ax = plt.subplots(figsize=(16, 16))
for idx in range(viz_words):
plt.scatter(*embed_tsne[idx, :], color='steelblue')
plt.annotate(int_to_vocab[idx], (embed_tsne[idx, 0], embed_tsne[idx, 1]), alpha=0.7)
```
| github_jupyter |
```
# !pip install catboost plotly
```
[Данные](https://yadi.sk/d/CBoVCVIxJ2q2cw)
# Обработка лидарных данных
## Сегментация
### Про лидар

```
from IPython.lib.display import YouTubeVideo
YouTubeVideo('Pa-q5elS_nE')
```
## А что за данные на самом деле
```
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import plotly.offline as py
import plotly.figure_factory as ff
import plotly.graph_objs as go
import tqdm
py.init_notebook_mode(connected=True)
EQUAL_ASPECT_RATIO_LAYOUT = dict(
margin={
'l': 0,
'r': 0,
'b': 0,
't': 0
}, scene=dict(
aspectmode='data'
))
def color(x, cmap='Reds'):
cmap = plt.get_cmap(cmap)
x = (x - np.min(x)) / np.max(x)
return cmap(x)
%matplotlib inline
ds = pd.read_csv('snow/snow.csv')
ds = ds.set_index(['scene_id'])
ds.head()
```
* intensity - ???
* ring - ???
### Кольцо

```
scene = ds.loc[0]
fig = go.Figure(layout=EQUAL_ASPECT_RATIO_LAYOUT)
fig.add_scatter3d(**{
'x': scene.x,
'y': scene.y,
'z': scene.z,
'mode': 'markers',
'marker': {
'size': 1,
'color': color(scene.ring, 'tab20'),
},
'text': scene.ring
})
py.iplot(fig)
```
### Интенсивность

```
fig = go.Figure(layout=EQUAL_ASPECT_RATIO_LAYOUT)
fig.add_scatter3d(**{
'x': scene.x,
'y': scene.y,
'z': scene.z,
'mode': 'markers',
'marker': {
'size': 1,
'color': color(scene.intensity, 'seismic'),
},
'text': scene.intensity
})
py.iplot(fig)
```
# Отфильтруем снег
## Эвристикой
```
def filter_by_intensity(intensity, limit=3):
# TODO
filtered_scene = scene[filter_by_intensity(scene.intensity)]
fig = go.Figure(layout=EQUAL_ASPECT_RATIO_LAYOUT)
fig.add_scatter3d(**{
'x': filtered_scene.x,
'y': filtered_scene.y,
'z': filtered_scene.z,
'mode': 'markers',
'marker': {
'size': 1,
'color': color(filtered_scene.intensity, 'seismic'),
},
'text': filtered_scene.intensity
})
py.iplot(fig)
```
Часть шума остается. При этом есть опасность, что мы вырежем что-то важное.
## Облачные вычисления
```
from sklearn.neighbors import KDTree
class ComputeFeatures(object):
def __init__(self, r=1.0):
self.xyz = None
self.intensity = None
self.ring = None
self.index = None
self.r = r
def _feature_names(self):
# TODO: return list of feature names
return []
def compute_point_features(self, point_id, neighbours):
# TODO: return list of features for given neighbours
# neighbours is list of indices
def get_point_neighbours(self, point_id):
return self.index.query_radius(self.xyz[point_id][np.newaxis, :], r=self.r)[0]
def __call__(self, xyz, intensity, ring):
self.xyz = xyz[:]
self.intensity = intensity[:]
self.ring = ring[:]
self.index = KDTree(self.xyz)
features = []
for point_id in range(len(self.xyz)):
neighbours = self.get_point_neighbours(point_id)
features.append(self.compute_point_features(point_id, neighbours))
return pd.DataFrame(columns=self._feature_names(), data=features)
import os
os.mkdir('features')
features = ComputeFeatures(r=1.0)
for scene_id in tqdm.tqdm(ds.reset_index().scene_id.unique()):
scene = ds.loc[scene_id]
features_df = \
features(scene[['x', 'y', 'z']].values, scene.intensity.values, scene.ring.values)
features_df.to_csv('./features/{}.csv'.format(scene_id))
ds.loc[scene_id].label.values
ds_features = []
for scene in os.listdir('features'):
scene_features = pd.read_csv(os.path.join('features', scene))
scene_id = int(scene.split('.')[0])
scene_features['scene_id'] = scene_id
scene_features['label'] = ds.loc[scene_id].label.values
ds_features.append(scene_features)
ds_features = pd.concat(ds_features)
ds_features = ds_features.drop(['Unnamed: 0'], axis=1)
ds_features.head()
```
# Посмотрим на разметку
```
scene = ds.loc[1]
fig = go.Figure(layout=EQUAL_ASPECT_RATIO_LAYOUT)
fig.add_scatter3d(**{
'x': scene.x,
'y': scene.y,
'z': scene.z,
'mode': 'markers',
'marker': {
'size': 1,
'color': color(scene.label, 'seismic'),
},
'text': scene.label
})
py.iplot(fig)
```
## Поучим что-нибудь
```
train = None # TODO train test split, but how
test = None
val = None
import catboost
def learn(X_train, X_val, y_train, y_val):
clf = catboost.CatBoostClassifier(n_estimators=100)
clf.fit(
X_train, y_train, early_stopping_rounds=10,
use_best_model=True, eval_set=(X_val.values, y_val.values), plot=True, verbose=False)
return clf
X_train = train.drop(["scene_id", "label"], axis=1)
y_train = train.label
X_val = val.drop(["scene_id", "label"], axis=1)
y_val = val.label
cls = learn(X_train, X_val, y_train, y_val)
X_test = test.drop(['scene_id', 'label'], axis=1)
y_test = test.label
from sklearn.metrics import precision_recall_curve, precision_score, recall_score
def test_one(clf, X_test, y_test):
y_test_hat = clf.predict_proba(X_test)
pr, rec, thr = precision_recall_curve(y_test, y_test_hat[:, 1])
ix = np.linspace(1, len(pr)-1, num=2000).astype(int)
return pr[ix], rec[ix], thr[ix - 1]
def heuristic_filter_scoring():
pr = []
rec = []
filter_range = range(1, 10)
for i in filter_range:
y_test_heuristic_hat = np.ones(len(X_test))
y_test_heuristic_hat[filter_by_intensity(test.intensity, i)] = 0
pr.append(precision_score(y_test, y_test_heuristic_hat))
rec.append(recall_score(y_test, y_test_heuristic_hat))
return pr, rec, filter_range
pr_bl, rec_bl, thr_bl = heuristic_filter_scoring()
def plot_pr_rec(*models):
traces = []
for model, clf, X_test, y_test in models:
pr, rec, thr = test_one(clf, X_test, y_test)
pr_rec = go.Scattergl(x = rec, y = pr, mode='lines', text=thr, name=model)
traces.append(pr_rec)
pr_rec_bl = go.Scatter(x = rec_bl, y = pr_bl, mode='lines+markers', text=thr_bl, name='Intensity BL')
layout = go.Layout(
title='Precission-recall',
xaxis=dict(
title='Recall'
),
yaxis=dict(
title='Precission'
))
fig = go.Figure(
data=traces + [pr_rec_bl],
layout=layout)
py.iplot(fig)
models = [('Catboost classifier', cls, X_test, y_test)]
plot_pr_rec(*models)
```
# Повизуализируем
```
y_test_hat = cls.predict_proba(test.drop(['scene_id', 'label'], axis=1))
scene_id = 3
scene = ds.loc[scene_id]
scene_predictions = y_test_hat[test.scene_id == scene_id][:, 1]
fig = go.Figure(layout=EQUAL_ASPECT_RATIO_LAYOUT)
fig.add_scatter3d(**{
'x': scene.x,
'y': scene.y,
'z': scene.z,
'mode': 'markers',
'marker': {
'size': 1,
'color': color(scene_predictions, 'seismic'),
},
'text': scene_predictions
})
py.iplot(fig)
```
| github_jupyter |
## Topics covered in this notebook:
1. What is Linear Regression.
2. Simple 1D Linear Regression.
3. Quality of Fit - R-Squared.
4. Multiple Linear Regression.
5. Probabilistic interpretation of the squared error.
6. L2 Regularization or Ridge Regression.
7. Gradient Descent.
8. L1 Regularization or Lasso.
9. L1 vs. L2 Regularization.
## 1. Linear Regression
Consider a set of questions:
1. How to predict future stock price?
2. How to predict a rating of a movie?
3. How many followers will I get on twitter?
4. How can you predict a price of a house?
What does all these questions have in common?
1. **Outputs**. More specifically continous outputs (price of a house, # of followers etc.). Let's call this **'Y'** or **'y'**
2. Predicting continuous outputs is called **regression**
What do I need to predict outputs?
1. **Features** or Inputs, Let's call this **'X'**.
2. **Training examples**, many X's for which Y's are known.
3. A **model**, a function that represents the relationship between X & Y.
4. A **loss** or **cost** or **objective** function that tells us how our model represents the training examples.
5. **Optimization**, a way we can find the model parameters that minimizes the cost function.
## 2. Simple 1D Linear Regression
So let's re-state our problem in more general terms:
1. We are given a set of points: {(x1,y1),(x2,y2),....(xn,yn)}
2. We plot them in a 2D chart.
3. We find the line of best fit.
```
%%javascript
MathJax.Hub.Config({
TeX: { equationNumbers: { autoNumber: "AMS" } }
});
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
X = np.linspace(0,10,50)
Y = 2*X + np.random.normal(0,1,50)
plt.scatter(X,Y,c = 'black',label = 'Data')
plt.plot(X,2*X,c = 'red',label = 'line of best fit')
plt.xlabel('X')
plt.ylabel('Y')
plt.legend()
plt.show()
```
Our line of best fit is defined as:
\begin{equation}
\large \hat y_i = ax_i+ b
\end{equation}
This is our **model**. <br>
What can we do to make sure $\large y_i$ is closer to $\large \hat y_i$
Can we do this? <br>
\begin{equation}
\large Error = \Sigma (\large y_i - \large \hat y_i)
\end{equation}
No. If the error for a point is -5 and +5 for the other the overall error is zero. However clearly that is not the case.<br>
What we want is this:
1. For any target != predictions, a +ve contribution to the error.
2. Standard way is to square of difference.
3. Called the "sum of the squared errors".
\begin{equation}
\large Error (E) = \Sigma (\large y_i - \large \hat y_i)^2
\end{equation}
Now that we have error we have to minimize it. How do we do that?<br> Take derivative & set it to zero! <br>
Substitute equation (1) in (3):
\begin{equation}
\large E = \Sigma (\large y_i - \large ax_i- b)^2
\end{equation}
<br>
We need to minimize E w.r.t a & b. Hence we need to take partial derivative.<br>
Derivative of above equation w.r.t a & b:
\begin{align}
\large \frac{\partial E}{\partial a} &= \Sigma \,2\, (\large y_i - \large ax_i- b)(-x_i) \\
\large \frac{\partial E}{\partial b} &= \Sigma \,2\, (\large y_i - \large ax_i- b)
\end{align}
<br>
Set that to zero: <br>
\begin{align}
\large a\,\Sigma \large x_i^2 + \large b\,\Sigma \large x_i &= \Sigma \,\large x_i\,\large y_i \\
\large a\,\Sigma \large x_i + \large b\,\large N &= \Sigma \,\large y_i \\
\end{align}
<br>
Solve those 2 equations simultaneously:
\begin{align}
\large a\, &= \frac{N\,\large\Sigma \,\large x_i\,\large y_i - \Sigma \,\large x_i\,\Sigma \,\large y_i}{N\, \large\Sigma \large x_i^2 - (\Sigma \large x_i)^2} \\
\large b\, &= \frac{N\,\large\Sigma \,\large x_i\,\large y_i^2 - \Sigma \,\large x_i\,\Sigma \,\large x_i \large y_i}{N\, \large\Sigma \large x_i^2 - (\Sigma \large x_i)^2} \\
\end{align}
<br>
Simplify it further is diving the above equations by $N^2$ and utilize the defitions of mean:
\begin{align}
\large a\, &= \frac{\overline{\large xy} - \overline{\large x}\,\overline{\large y}}{\overline{\large x^2} - \overline{\large x}^2}\\
\\
\large b\, &= \frac{\overline{\large y}\, \overline{\large x^2} - \overline{\large x}\,\overline{\large xy}}{\overline{\large x^2} - \overline{\large x}^2}\\
\end{align}
where,
\begin{align}
\large \overline{x}\, &= \frac{1}{N} \,\large \Sigma \,\large x_i \\
\large \overline{xy}\, &= \frac{1}{N} \,\large \Sigma \,\large x_i\,y_i \\
\end{align}
## 3. Quality of Fit: R-Sq
\begin{equation}
\large R^2 \, = 1 \, - \frac{SS_{\text{res}}}{SS_{\text{tot}}} \\
\end{equation}
where
\begin{align}
\large SS_{\text{res}}\, &= \large \Sigma (y\, - \hat y_i)^2 \\
\large SS_{\text{tot}}\, &= \large \Sigma (y\, - \overline y_i)^2 \\
\end{align}
<br>
1. Suppose the residual (SS_res) is close to zero. Then the R-Sq will be ~1 -> Perfect correlation.
2. Suppose the R-Sq is zero which means SS_res / SS_tot = 1 -> This means we predicted just the everage of y. Model not good.
3. When is R-Sq < 0? This means SS_res / SS_tot > 1. Model prediction is worse than mean. Again model not good.
## 4. Multiple Linear Regression
The error function still doesn't change from 1D regression, only the expression for the prediction does,
\begin{equation}
\large Error (E) = \Sigma (\large y_i - \large \hat y_i)^2 = \Sigma (\large y_i - \large w^Tx_i)^2
\end{equation}
<br>
We can still take derivative w.r.t to any component of w: j = 1,....D(No. of inputs or feature dimension) <br>
\begin{align}
\large \frac{\partial E}{\partial w_j} &= \Sigma 2(\large y_i - \large w^Tx_i)(- \frac{\partial (w^Tx_i)}{\partial w_j})\\
&= \large \Sigma \,2(\large y_i - \large w^Tx_i)(- x_{ij})
\end{align}
<br>
Set this equation to zero. D equations & D unknowns:
\begin{align}
\large \Sigma \,2(\large y_i - \large w^Tx_i)(- x_{ij}) &= 0\\
\large \Sigma \,w^Tx_ix_{ij} - \large \Sigma y_ix_{ij} &= 0
\end{align}
<br>
Isolate w & represent everything in matrix form using dot product:
<br>
\begin{align}
\large w^T(X^TX)\, = \, y^TX\\
\end{align}
<br>
Take transpose on both sides:
\begin{align}
\large [w^T(X^TX)]^T\, &= \, [y^TX]^T\\
\large (X^TX)w\, &= \, X^T\large y\\
\large w \, &=\, (X^TX)^{-1}\, X^T\large y
\end{align}
<br>
Use numpy linalg.solve to solve the above equation. Elimiates the need to manually taking inverse:<br>
\begin{align}
\large w \, &=\, np.lingalg.solve(X^TX\,, X^T\large y)
\end{align}
## 5. Probabilistic Interpretation of Squared Error:
### Linear Regression is the maximum likelihood solution to the line of best fit.
What is Maximum Likelihood?
1. Assuming Gaussian distribution - If we plot histogram for grades of students we will get a bell curve:<br>
<img src="Images/bell.jpg" alt="Drawing" style="width: 300px;"/>
2. We would do an experiment to collect everyone's grades: {x1, x2,....xn}<br>
3. Intuitively, we know the average grade of the students is:
\begin{align}
\large \mu \, &= \frac{1}{N} \,\large \Sigma \,\large x_i \\
\end{align}
4. Is there a systematic way of getting this answer? Suppose we want to find thr true mean of the Gaussian form which the data arises:<br>
<br>
\begin{align}
find\, \large \mu \,,\, where\,X \, \tilde \, N(\mu , \sigma^2) \\
\end{align}
5. We can write the probability of any single point xi:
<br>
\begin{align}
\large p(x_i) \, &= \frac{1}{\sqrt{2\pi\sigma^2}} \,\large exp^{\frac {1}{2} \, \large \frac {(\large x_i-\ \large mu)^ 2}{\large \sigma^2}}\\
\end{align}
6. Since we know the grades are iid, we can muliply individual probabiities:
\begin{align}
\large p(x_1,x_2,...x_N) \, &= \prod_{i=1}^n\frac{1}{\sqrt{2\pi\sigma^2}} \,\large exp^{\frac {1}{2} \, \large \frac {(\large x_i-\ \large mu)^ 2}{\large \sigma^2}}\\
\end{align}
7. Another way to write this is as 'likelihood' form. Probability of X given the parameter of interest:
1. We want to find mean so that likelihood is maximized. This is called 'maximum likelihood'.
2. We want to finfd the best setting of mean(mu) so that the date we measured is likely to have come from this distribution.
\begin{align}
\large p(X\,|\,\mu) = \large p(x_1,x_2,...x_N) \, &= \prod_{i=1}^n\frac{1}{\sqrt{2\pi\sigma^2}} \,\large exp^{\frac {1}{2} \, \large \frac {(\large x_i-\ \large mu)^ 2}{\large \sigma^2}}\\
\end{align}
8. How to find mu? Take log of the likelihood & set it to zero to make it easy to solve. log() in monotomically increasing function so if A > B -> log(A) > log(B)
\begin{align}
\large l \, &= \large log \prod_{i=1}^n\frac{1}{\sqrt{2\pi\sigma^2}} \,\large exp^{\frac {1}{2} \, \large \frac {(\large x_i-\ \large mu)^ 2}{\large \sigma^2}}\\
\large l \, &= \large [\, \Sigma_{i=1}^n-\frac{1}{2} log(\sqrt{2\pi\sigma^2}) \,- \large {\frac {1}{2} \, \large \frac {(\large x_i-\ \large mu)^ 2}{\large \sigma^2}}\,]\\
\end{align}
Take derivative and set it to zero:
\begin{align}
\large \frac{dl}{d\mu} &= \large \Sigma \, \frac {(x_i - \mu)}{\sigma^2} = 0\\
\large \mu \, &= \frac{1}{N} \,\large \Sigma \,\large x_i \\
\end{align}
<br>
9. If we look at the equation for l and remove some irrelevant terms that are absorbed by zero, we get:
\begin{align}
\large l \, &= \large [\, \Sigma_{i=1}^n-\frac{1}{2} log(\sqrt{2\pi\sigma^2}) \,- \large {\frac {1}{2} \, \large \frac {(\large x_i-\ \large mu)^ 2}{\large \sigma^2}}\,]\\
\large equivalent \,l &= \large - \Sigma \, (x_i - \mu)^2\\
\end{align}
### This means maximizing the log-likelihood is equivalent to minimizing negative of squared errors
Compare this to our error(E):
\begin{align}
\large \,l &= \large - \Sigma \, (x_i - \mu)^2 -> \,maximize\\
\large E & = \large \Sigma\,(\large y_i - \large \hat y_i)^2 -> \, minimize \\
\end{align}
So minimizing E is equivalent to maximizing -E. So when we minize the squared error for linear regression, this is equivalent to maximizing the likelihood. <br>
Equivalent way of writing this:
\begin{align}
\large y \,\tilde \,\, N(w^Tx,\sigma^2)\\
\end{align}
<br>
\begin{align}
\large y & = \, \large w^Tx\, +\, \epsilon,\, \epsilon\, \tilde\, N(0,\sigma^2) \\
\end{align}
<br>
### In other words, linear regression makes the assumption that errors are gaussion and the trend is linear.
## 6. L2 Regularization or Ridge Regression:
Data may have outliers. These outliers are usually measurement errors or data entry errors. In some cases these outliers may actually mean something but in this case lets assume its the former assumption.
#### Outliers pull the line of best fit away from the main trend to minimize the squared error. The idea is to ensure the weights are not overly large weights because that might want to fit to outliers.
Look at the 2 lines below. Which one is the best fit? Red is the standard model which accounts for the outliers whereas black is the L2 regularization model that ignored the effect of outliers.<br>
<img src="Images/L2-Reg.png" alt="Drawing" style="length: 1080px;" style="width: 1080px;"/>
### How does L2 Regularization work?
Modify the cost function (J) such that large weights are penalized.
<br>
\begin{align}
\large J & = \large \Sigma\,(\large y_i - \large \hat y_i)^2 + \lambda \lvert \large w^2\rvert \\
\lvert \large w^2\rvert &= \large w^Tw = \large w1^2 + w2^2 + ....w_D^2
\end{align}
#### Probabilistic Perspective:
1. Plain squared error maximizes likelihood because J = negative log likelihood.
2. Now we are no longer maximizing this, since there are 2 terms. The 2nd is called prior which has the information about the weights irrespective of the data since it is not dependent on x & y.
<img src="Images/L2 Prob.jpg" alt="Drawing" style="width: 400px;"/>
<br>
<br>
3. Looks like Bayes rule.
#### This is called MAP - maximum a posteriori.
<img src="Images/MAP.jpg" alt="Drawing" style="width: 250px;"/>
<br>
<br>
4. Solve for w:
<img src="Images/L2-w.jpg" alt="Drawing" style="width: 300px;"/>
<img src="Images/L2-der.jpg" alt="Drawing" style="width: 300px;"/>
<img src="Images/L2-wFinal.jpg" alt="Drawing" style="width: 200px;"/>
## 7. Gradient Descent:
1. Optimization method.
2. Used extensively in deep learning.
3. Idea: You have a fucntion you want to minimize, J(w) = cost or error. Find optimal inputs to minimize this function.
<img src="Images/GD.png" alt="Drawing" style="width: 500px;"/>
<br>
4. Weight, w = w - learning_rate * dJ/dw.
5. Example: J = w^2:
1. dJ/dw = 2w.
2. Set initial w = 10, learning rate = 0.1.
3. Iteration 1: w = 10 - 0.1 * 20 = 8.
4. Iteration 2: w = 08 - 0.1 * 16 = 6.4.
5. Iteration 3: w = 6.4 - 0.1 * 12.8 = 5.12 and so on..
6. Notice how the weigh 'w' is converging towards the true solution 0. With enough iterations we will coverge.
```
import numpy as np
w = 5
learning_rate = 0.1
for i in range(50):
w -= learning_rate * 2* w
print(w)
print('Notice how w is converging towards zero!')
```
### Gradient Descent for Linear Regression:
Cost function to minimize:
\begin{align}
\large J \, &= \large (Y\, - Xw)^T (Y\, - Xw) \\
\end{align}
<br>
Gradient:
\begin{align}
\large \frac {\partial J}{\partial w} \, &= \large -2X^TY\, +\, 2X^TXw \,=\, 2X^T(\hat Y\, -\, Y) \\
\end{align}
<br>
#### Instead of setting it to 0 & solving for w, we will just take small steps in this direction.
So weight update will be for a certain number of steps: <br>
\begin{align}
\large w \, &= \large w - \eta * X^T(\hat Y\, -\, Y) \\
\end{align}
## 7. L1 Regularization (Lasso):
In general we want no. of features(D) << no. of samples(N). Also, there might be a case where some of the features are just constant or quasi-const, uncorrelated to output etc. In all these cases we could use L1 regularization to eliminate those features from the model.
1. Select a small number of important features that actually predict the trend.
2. Eliminate the noise influence on the output.
3. Similar to L2. It has a penalty term using L1 norm.
<img src="Images/L1-Concept.jpg" alt="Drawing" style="width: 250px;"/>
4. Similar to L2 this also puts a prior on w, so its also a MAP estimation of w. We had a gaussian distribution on the prior for L2. We have Laplace here:
\begin{align}
\large p(w) \, &= \large \frac {\lambda}{2} exp(- \lambda \lvert \large w\rvert) \\
\end{align}
5. Taking derivative of cost function:
\begin{align}
\large \frac {\partial J}{\partial w} \, &= \large -2X^TY\, +\, 2X^TXw \,+\, \lambda sign(w) \\
\end{align}
6. Since this has a sign term we can't used closed form solution to update weights and use gradient descent instead.
## 9. L1 vs L2 Regularization
L2 reduces the effect of outliers on the model by maintaining the weights to a smaller value.
L1 removes unwanted features. Encourages a sparse solution.
Both helps prevent overfitting, by not fitting to noise.
L2 penalty is quadratic: as w -> 0 derivative ->0 <br>
L1 penalty is abs func : as w -> 0 doesn't matter it will fall at a contant rate. When it reaches 0, is stays there forever.
#### Combine L1 & L2 = ElasticNet
## References:
1. An Introduction to Statistical Learning Textbook by Gareth James, Daniela Witten, Trevor Hastie and Robert Tibshirani.
2. University of Michigan EECS 445 - Machine Learning Course (https://github.com/eecs445-f16/umich-eecs445-f16).<br>
3. University of Toronto CSC 411 - Intro. to Machine Learning (http://www.cs.toronto.edu/~urtasun/courses/CSC411_Fall16/CSC411_Fall16.html).<br>
4. Stanford CS109 - Intro. to proabability for computer scientists (https://web.stanford.edu/class/archive/cs/cs109/cs109.1166/). <br>
5. Few online courses on Udemy, Coursera etc.
| github_jupyter |
```
# Importing Libraries
import os
import sys
import numpy as np
import pandas as pd
import glob
# Importing Data
# Load the Drive helper and mount
from google.colab import drive
# This will prompt for authorization.
drive.mount('/content/drive')
# Installing Time Series libraries
pip install pyramid-arima
pip install pmdarima
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.pylab import rcParams
import seaborn as sns
import scipy
%matplotlib inline
import scipy
import statsmodels.api as sm
from statsmodels.tsa.seasonal import seasonal_decompose
import pyramid as pm
from sklearn import metrics
import pmdarima as pm
from pmdarima import auto_arima
import warnings
warnings.filterwarnings('ignore')
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity="all"
# Loading Data
#df = pd.read_csv('content/drive/My Drive/REVA/Interview/inputData.csv")
df = pd.read_csv('/content/drive/My Drive/REVA/Interview/inputData.csv', sep=',')
# Exploratory Data Analysis
df.head()
df.tail()
df.shape
df.describe()
# Missing Values Check
df.info()
# Not found any missing values from below analysis
df["Timestamp"]=pd.to_datetime(df['Timestamp']).dt.strftime('%Y-%m-%d')
pd.pivot_table(data=df, index='Timestamp', values='CPU_Used', aggfunc='mean').reset_index().rename(columns={'CPU_Used': 'average_number_of_CPU_Used'}).round(2)
df_pivot=pd.pivot_table(data=df, index='Timestamp', values='CPU_Used', aggfunc='sum').reset_index().rename(columns={'CPU_Used': 'total_CPU_Used'})
df_pivot.head(12)
df_pivot.plot(figsize=(30,10), linewidth=1, fontsize=20)
plt.xlabel('Unique CPU Used in Days', fontsize=30);
plt.ylabel('CPU_Used', fontsize=30);
month = pd.date_range('20190831', periods = 11, freq = 'M')
month
df_pivot['Timestamp']= month
df_pivot.head()
data=df_pivot.loc[:,('Timestamp','CPU_Used')]
data.head()
data.describe()
data.set_index('Timestamp',inplace=True)
data.head()
plt.figure(figsize = (15,5))
plt.plot(data)
plt.xlabel('Time')
plt.ylabel('Total CPU used in 1000')
plt.title("Number of Total CPU used over time")
plt.show()
decomposition=seasonal_decompose(data,model='multiplicative')
decomposition_ad=seasonal_decompose(data,model='additive')
plt.figure(figsize = (20,10))
trend=decomposition.trend
seasonal=decomposition.seasonal
residual=decomposition.resid
plt.figure(figsize=(15,10))
plt.subplot(221)
plt.plot(data,color="#00b8ff",label="Original")
plt.legend(loc="best")
plt.subplot(222)
plt.plot(trend,'b',label="Trend")
plt.legend(loc="best")
plt.subplot(223)
#plt.plot(data,color="#ff00ff",label="Original")
plt.plot(seasonal,'b',label="Seasonal")
plt.legend(loc="best")
plt.subplot(224)
plt.plot(residual,'y',label="residual")
plt.legend(loc="best")
plt.tight_layout()
plt.show()
plt.figure(figsize = (30,15))
trend_ad=decomposition_ad.trend
seasonal_ad=decomposition_ad.seasonal
residual_ad=decomposition_ad.resid
plt.figure(figsize=(15,10))
plt.subplot(221)
plt.plot(data,color="#00b8ff",label="Original")
plt.legend(loc="best")
plt.subplot(222)
plt.plot(trend_ad,'b',label="Trend")
plt.legend(loc="best")
plt.subplot(223)
#plt.plot(data,color="#ff00ff",label="Original")
plt.plot(seasonal_ad,'b',label="Seasonal")
plt.legend(loc="best")
plt.subplot(224)
plt.plot(residual_ad,'y',label="residual")
plt.legend(loc="best")
plt.tight_layout()
plt.show()
adfTest=pm.arima.ADFTest(alpha=0.05)
adfTest.is_stationary(data)
train, test=data[:20],data[20:]
train.shape
test.shape
plt.figure(figsize=(15,10))
plt.plot(train)
plt.plot(test)
plt.show()
Arima_model=auto_arima(train,start_p=1,start_q=1,max_p=1,max_q=1,start_P=0,start_Q=0,max_P=8,max_Q=8,m=12,seasonal=True,trace=True,d=1,D=1,error_action="warn",suppress_warnings=True,stepwise=True,random_state=20,n_fits=30)
Arima_model.summary()
predictions=pd.DataFrame(Arima_model.predict(n_periods=4),index=test.index)
predictions.columns=["Predicted_CPU_Used"]
plt.figure(figsize = (15,5))
plt.plot(train,label="Training")
plt.plot(test,label="Testing")
plt.plot(predictions,label="Predicted")
plt.legend(loc="upper center")
plt.show()
test["Predicted_Invoiced"]=predictions
test["Error"]=test["total_Invoiced"]-test["Predicted_Invoiced"]
test
metrics.mean_absolute_error(test.total_Invoiced,test.Predicted_Invoiced)
metrics.mean_squared_error(test.total_Invoiced,test.Predicted_Invoiced)
metrics.median_absolute_error(test.total_Invoiced,test.Predicted_Invoiced)
plt.figure(figsize = (20,10))
plt.subplot(121)
plt.plot(test.Error,color="#ff33CC")
plt.title("Error Distribution OverTime")
plt.subplot(122)
scipy.stats.probplot(test.Error,plot=plt)
plt.show()
plt.figure(figsize = (20,10))
pm.autocorr_plot(test.Error)
plt.show()
```
| github_jupyter |
# Multi-Layer Perceptron, MNIST
---
In this notebook, we will train an MLP to classify images from the [MNIST database](http://yann.lecun.com/exdb/mnist/) hand-written digit database.
The process will be broken down into the following steps:
>1. Load and visualize the data
2. Define a neural network
3. Train the model
4. Evaluate the performance of our trained model on a test dataset!
Before we begin, we have to import the necessary libraries for working with data and PyTorch.
```
# import libraries
import torch
import numpy as np
```
---
## Load and Visualize the [Data](http://pytorch.org/docs/stable/torchvision/datasets.html)
Downloading may take a few moments, and you should see your progress as the data is loading. You may also choose to change the `batch_size` if you want to load more data at a time.
This cell will create DataLoaders for each of our datasets.
```
from torchvision import datasets
import torchvision.transforms as transforms
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 20
# convert data to torch.FloatTensor
transform = transforms.ToTensor()
# choose the training and test datasets
train_data = datasets.MNIST(root='data', train=True,
download=True, transform=transform)
test_data = datasets.MNIST(root='data', train=False,
download=True, transform=transform)
# prepare data loaders
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size,
num_workers=num_workers)
```
### Visualize a Batch of Training Data
The first step in a classification task is to take a look at the data, make sure it is loaded in correctly, then make any initial observations about patterns in that data.
```
import matplotlib.pyplot as plt
%matplotlib inline
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy()
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
# print out the correct label for each image
# .item() gets the value contained in a Tensor
ax.set_title(str(labels[idx].item()))
```
### View an Image in More Detail
```
img = np.squeeze(images[1])
fig = plt.figure(figsize = (12,12))
ax = fig.add_subplot(111)
ax.imshow(img, cmap='gray')
width, height = img.shape
thresh = img.max()/2.5
for x in range(width):
for y in range(height):
val = round(img[x][y],2) if img[x][y] !=0 else 0
ax.annotate(str(val), xy=(y,x),
horizontalalignment='center',
verticalalignment='center',
color='white' if img[x][y]<thresh else 'black')
```
---
## Define the Network [Architecture](http://pytorch.org/docs/stable/nn.html)
The architecture will be responsible for seeing as input a 784-dim Tensor of pixel values for each image, and producing a Tensor of length 10 (our number of classes) that indicates the class scores for an input image. This particular example uses two hidden layers and dropout to avoid overfitting.
```
import torch.nn as nn
import torch.nn.functional as F
## TODO: Define the NN architecture
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# linear layer (784 -> 1 hidden node)
self.fc1 = nn.Linear(28 * 28, 1)
def forward(self, x):
# flatten image input
x = x.view(-1, 28 * 28)
# add hidden layer, with relu activation function
x = F.relu(self.fc1(x))
return x
# initialize the NN
model = Net()
print(model)
```
### Specify [Loss Function](http://pytorch.org/docs/stable/nn.html#loss-functions) and [Optimizer](http://pytorch.org/docs/stable/optim.html)
It's recommended that you use cross-entropy loss for classification. If you look at the documentation (linked above), you can see that PyTorch's cross entropy function applies a softmax funtion to the output layer *and* then calculates the log loss.
```
import torch.optim as optim
## TODO: Specify loss and optimization functions
# loss = nn.CrossEntropyLoss()
# specify loss function
criterion = nn.CrossEntropyLoss()
# specify optimizer
# optimizer = optim.Adam([var1, var2], lr=0.0001)
optimizer = optim.SGD(model.parameters(), lr=0.01, momentum=0.9)
```
---
## Train the Network
The steps for training/learning from a batch of data are described in the comments below:
1. Clear the gradients of all optimized variables
2. Forward pass: compute predicted outputs by passing inputs to the model
3. Calculate the loss
4. Backward pass: compute gradient of the loss with respect to model parameters
5. Perform a single optimization step (parameter update)
6. Update average training loss
The following loop trains for 30 epochs; feel free to change this number. For now, we suggest somewhere between 20-50 epochs. As you train, take a look at how the values for the training loss decrease over time. We want it to decrease while also avoiding overfitting the training data.
```
# number of epochs to train the model
n_epochs = 30 # suggest training between 20-50 epochs
model.train() # prep model for training
for epoch in range(n_epochs):
# monitor training loss
train_loss = 0.0
###################
# train the model #
###################
for data, target in train_loader:
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update running training loss
train_loss += loss.item()*data.size(0)
# print training statistics
# calculate average loss over an epoch
train_loss = train_loss/len(train_loader.sampler)
print('Epoch: {} \tTraining Loss: {:.6f}'.format(
epoch+1,
train_loss
))
```
---
## Test the Trained Network
Finally, we test our best model on previously unseen **test data** and evaluate it's performance. Testing on unseen data is a good way to check that our model generalizes well. It may also be useful to be granular in this analysis and take a look at how this model performs on each class as well as looking at its overall loss and accuracy.
#### `model.eval()`
`model.eval(`) will set all the layers in your model to evaluation mode. This affects layers like dropout layers that turn "off" nodes during training with some probability, but should allow every node to be "on" for evaluation!
```
# initialize lists to monitor test loss and accuracy
test_loss = 0.0
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
model.eval() # prep model for *evaluation*
for data, target in test_loader:
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# update test loss
test_loss += loss.item()*data.size(0)
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
correct = np.squeeze(pred.eq(target.data.view_as(pred)))
# calculate test accuracy for each object class
for i in range(len(target)):
label = target.data[i]
class_correct[label] += correct[i].item()
class_total[label] += 1
# calculate and print avg test loss
test_loss = test_loss/len(test_loader.sampler)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(10):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
str(i), 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
```
### Visualize Sample Test Results
This cell displays test images and their labels in this format: `predicted (ground-truth)`. The text will be green for accurately classified examples and red for incorrect predictions.
```
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
# get sample outputs
output = model(images)
# convert output probabilities to predicted class
_, preds = torch.max(output, 1)
# prep images for display
images = images.numpy()
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
ax.set_title("{} ({})".format(str(preds[idx].item()), str(labels[idx].item())),
color=("green" if preds[idx]==labels[idx] else "red"))
```
| github_jupyter |
OK, to begin we need to import some standart Python modules
```
# -*- coding: utf-8 -*-
"""
Created on Fri Feb 12 13:21:45 2016
@author: GrinevskiyAS
"""
from __future__ import division
import numpy as np
from numpy import sin,cos,tan,pi,sqrt
import matplotlib as mpl
import matplotlib.cm as cm
import matplotlib.pyplot as plt
%matplotlib inline
font = {'family': 'Arial', 'weight': 'normal', 'size':14}
mpl.rc('font', **font)
```
First, let us setup the working area.
```
#This would be the size of each grid cell (X is the spatial coordinate, T is two-way time)
xstep=10
tstep=10
#size of the whole grid
xmax = 301
tmax = 201
#that's the arrays of x and t
xarray=np.arange(0, xmax, xstep)
tarray=np.arange(0, tmax, tstep)
#now fimally we created a 2D array img, which is now all zeros, but later we will add some amplitudes there
img=np.zeros((len(xarray), len(tarray)))
```
Let's show our all-zero image
```
plt.imshow(img.T,interpolation='none',cmap=cm.Greys, vmin=-2,vmax=2, extent=[xarray[0]-xstep/2, xarray[-1]+xstep/2, tarray[-1]+tstep/2, tarray[0]-tstep/2])
```
What we are now going to do is create a class named **`Hyperbola`**
Each object of this class is capable of computing traveltimes to a certain subsurface point (diffractor) and plotting this point response on a grid
```
class Hyperbola:
def __init__(self, xarray, tarray, x0, v, t0):
###input parameters define a difractor's position (x0,t0), P-wave velocity of homogeneous subsurface, and x- and t-arrays to compute traveltimes on.
###
self.x=xarray
self.x0=x0
self.t0=t0
self.v=v
#compute traveltimes
self.t=sqrt(t0**2 + (2*(xarray-x0)/v)**2)
#obtain some grid parameters
xstep=xarray[1]-xarray[0]
tbegin=tarray[0]
tend=tarray[-1]
tstep=tarray[1]-tarray[0]
#delete t's and x's for samples where t exceeds maxt
self.x=self.x[ (self.t>=tbegin) & (self.t <= tend) ]
self.t=self.t[ (self.t>=tbegin) & (self.t <= tend) ]
self.imgind=((self.x-xarray[0])/xstep).astype(int)
#compute amplitudes' fading according to geometrical spreading
self.amp = 1/(self.t/self.t0)
self.grid_resample(xarray, tarray)
def grid_resample(self, xarray, tarray):
# that's a function that computes at which 'cells' of image should we place the hyperbola
tend=tarray[-1]
tstep=tarray[1]-tarray[0]
self.xind=((self.x-xarray[0])/xstep).astype(int) #X cells numbers
self.tind=np.round((self.t-tarray[0])/tstep).astype(int) #T cells numbers
self.tind=self.tind[self.tind*tstep<=tarray[-1]] #delete T's exceeding max.T
self.tgrid=tarray[self.tind] # get 'gridded' T-values
self.coord=np.vstack((self.xind,tarray[self.tind]))
def add_to_img(self, img, wavelet):
# puts the hyperbola into the right cells of image with a given wavelet
maxind=np.size(img,1)
wavlen=np.floor(len(wavelet)/2).astype(int)
self.imgind=self.imgind[self.tind < maxind-1]
self.tind=self.tind[self.tind < maxind-1]
ind_begin=self.tind-wavlen
for i,sample in enumerate(wavelet):
img[self.imgind,ind_begin+i]=img[self.imgind,ind_begin+i]+sample
return img
```
For testing purposes, let's create an object named Hyp_test and view its parameters
```
Hyp_test = Hyperbola(xarray, tarray, x0 = 100, t0 = 30, v = 2)
#Create a fugure and add axes to it
fgr_test1 = plt.figure(figsize=(7,5), facecolor='w')
ax_test1 = fgr_test1.add_subplot(111)
#Now plot Hyp_test's parameters: X vs T
ax_test1.plot(Hyp_test.x, Hyp_test.t, 'r', lw = 2)
#and their 'gridded' equivalents
ax_test1.plot(Hyp_test.x, Hyp_test.tgrid, ls='none', marker='o', ms=6, mfc=[0,0.5,1],mec='none')
#Some commands to add gridlines, change the directon of T axis and move x axis to top
ax_test1.set_ylim(tarray[-1],tarray[0])
ax_test1.xaxis.set_ticks_position('top')
ax_test1.grid(True, alpha = 0.1, ls='-',lw=.5)
ax_test1.set_xlabel('X, m')
ax_test1.set_ylabel('T, ms')
ax_test1.xaxis.set_label_position('top')
plt.show()
```
| github_jupyter |
```
%matplotlib inline
import numpy as np
from joblib import Parallel, delayed
from gensim.models.keyedvectors import KeyedVectors
# from numba import jit, autojit
from sklearn.manifold import MDS, TSNE
from tqdm import tqdm
from sklearn.metrics.pairwise import cosine_similarity, cosine_distances, linear_kernel, euclidean_distances
from sklearn.feature_extraction.text import TfidfVectorizer
from glob import glob
import matplotlib.pyplot as plt
import seaborn as sns
import os
import scipy.sparse as sps
# model = KeyedVectors.load_word2vec_format('word2vec-models/lemmas.cbow.s100.w2v.bin', binary=True)
model_sg = KeyedVectors.load_word2vec_format('word2vec-models/lemmas.sg.s100.w2v.bin', binary=True)
model_sg
def n_similarity(s1, s2):
vec1 = np.mean(model[s1.split()], axis=0)
vec2 = np.mean(model[s2.split()], axis=0)
return cosine_similarity([vec1], [vec2])[0][0]
def n_distance(s1, s2):
vec1 = np.mean(model[s1.split()], axis=0)
vec2 = np.mean(model[s2.split()], axis=0)
return cosine_distances([vec1], [vec2])[0][0]
def matrix_row_sim(s1, contexts, row_length):
row = np.empty(row_length)
for j, s2 in enumerate(contexts):
# row[j] = model.n_similarity(s1.split(), s2.split())
row[j] = n_similarity(s1, s2)
return row
def matrix_row_dist(s1, contexts, row_length):
row = np.empty(row_length)
for j, s2 in enumerate(contexts):
row[j] = n_distance(s1, s2)
return row
```
# tf-idf
```
words = [('joogitee', 'sõidutee'),
('õun', 'banaan'),
('õun', 'puder'),
('õun', 'kivi'),
('ämber', 'pang'),
('hea', 'halb'),
('countries', 'cities'),
('Eesti', 'TallinnTartu')]
# words = [('hea', 'halb'),
# ('countries', 'cities'),
# ('Eesti', 'TallinnTartu')]
words
for word1, word2 in words:
print(word1, word2)
for window in [2,3,4]:
for symmetric in [True, False]:
print(window, symmetric)
with open('../datasets/contexts/{}_s_{}_w_{}.txt'.format(word1, symmetric, window)) as f:
contexts1 = f.read().splitlines()
with open('../datasets/contexts/{}_s_{}_w_{}.txt'.format(word2, symmetric, window)) as f:
contexts2 = f.read().splitlines()
contexts = contexts1 + contexts2
# labels = [0]*len(contexts1) + [1]*len(contexts2)
print(len(contexts))
tfidf_vectorizer = TfidfVectorizer()
tfidf = tfidf_vectorizer.fit_transform(contexts)
print('saving')
print()
filename = '../datasets/tfidf-features/{}_{}_w_{}_s_{}.npy'.format(word1, word2, window, symmetric)
print(filename)
# break
# break
# break
np.save(filename, tfidf)
```
# mean-vec
```
for word1, word2 in words:
print(word1, word2)
for window in [2,3,4]:
for symmetric in [True, False]:
print(window, symmetric)
with open('datasets/contexts/{}_s_{}_w_{}.txt'.format(word1, symmetric, window)) as f:
contexts1 = f.read().splitlines()
with open('datasets/contexts/{}_s_{}_w_{}.txt'.format(word2, symmetric, window)) as f:
contexts2 = f.read().splitlines()
contexts_len = min(len(contexts1), len(contexts2))
contexts = contexts1[:contexts_len] + contexts2[:contexts_len]
# labels = [0]*len(contexts1) + [1]*len(contexts2)
print(len(contexts1), len(contexts2), contexts_len, len(contexts))
n = len(contexts)
mean_vectors = np.zeros((n, 100))
for i in range(n):
mean_vectors[i] = np.mean(model_sg[contexts[i].split()], axis=0)
print('saving')
print()
filename = 'datasets/sg/mean-vec/vectors/{}_{}_w_{}_s_{}.npy'.format(word1, word2, window, symmetric)
print(filename)
# break
# break
# break
np.save(filename, mean_vectors)
```
# {angular, euclidean}_distance
```
filename
t = np.load(filename).item()
t.shape
cosine_similarity(features[i], features)
a = glob(os.path.join('../datasets', feature, '*'))[0]
filename
features = np.load(filename)
features = features.item()
feature = 'mean-vec'
feature = 'tfidf-features'
a = glob(os.path.join('../datasets', feature, '*'))[0]
features_to_pairwise(files_5k[0], feature)
files_5k[0]
# doing both angular and euclidean here. change contentds to modify.
def features_to_pairwise(filename):
print(filename)
features = np.load(filename)
# if feature == 'tfidf-features':
# features = features.
# print(features.item().shape)
# features = features.item()
# print(type(features))
n = features.shape[0]
matrix = np.zeros((n,n))
for i in range(n):
row = [features[i]]
# if feature == 'tfidf-features':
# row = row[0]
matrix[i,:] = cosine_similarity(row, features).flatten()
# matrix[i,:] = euclidean_distances(row, features).flatten()
matrix[matrix>1] = 1
matrix = np.arccos(matrix)/np.pi
basename = os.path.basename(filename)
new_path = os.path.join('datasets/sg/mean-vec/angular-distance/', basename)
# new_path = os.path.join('datasets/sg/mean-vec/euclidean-distance/', basename)
print(new_path)
np.save(new_path, matrix)
# LSI COS DISTANCE
def features_to_pairwise_lsi(filename):
print(filename)
features = np.load(filename)
# if feature == 'tfidf-features':
# features = features.
# print(features.item().shape)
# features = features.item()
# print(type(features))
n = features.shape[0]
matrix = np.zeros((n,n))
for i in range(n):
row = [features[i]]
# if feature == 'tfidf-features':
# row = row[0]
matrix[i,:] = cosine_distances(row, features).flatten()
# matrix[i,:] = euclidean_distances(row, features).flatten()
# matrix[matrix>1] = 1
# matrix = np.arccos(matrix)/np.pi
basename = os.path.basename(filename)
new_path = os.path.join('datasets/tfidf/lsi-cos-dist/', basename)
# new_path = os.path.join('datasets/sg/mean-vec/euclidean-distance/', basename)
print(new_path)
np.save(new_path, matrix)
features_to_pairwise(fname)
fname = glob('datasets/tfidf/lsi/*')[0]
all_files = glob(os.path.join('datasets/tfidf/lsi/*'))
len(all_files)/48
# all_files = glob(os.path.join('../datasets', feature, '*'))
# files_5k = [filename for filename in all_files if ('hea' in filename or 'countries' in filename or 'Eesti' in filename)]
Parallel(n_jobs=25)(delayed(features_to_pairwise_lsi)(filename) for filename in all_files)
basename = os.path.basename(glob('../datasets/mean-vec/*')[0])
new_path = os.path.join('../datasets/angular-distance/mean-vec/', basename)
np.arccos(1.1)
matrix[matrix>1] = 1
matrix[matrix>1] = 1
angular_sim = 1-np.arccos(matrix)/np.pi
```
# WMD
# Old code
```
for window in [2,3,4]:
for symmetric in [True, False]:
print(metric.__name__, window, symmetric)
apple_contexts = open('../datasets/apple_contexts_s_{}_w_{}.txt'.format(symmetric, window)).read().splitlines()
rock_contexts = open('../datasets/rock_contexts_s_{}_w_{}.txt'.format(symmetric, window)).read().splitlines()
pear_contexts = open('../datasets/pear_contexts_s_{}_w_{}.txt'.format(symmetric, window)).read().splitlines()
contexts = apple_contexts + rock_contexts + pear_contexts
labels = [0]*len(apple_contexts) + [1]*len(rock_contexts) + [2]*len(pear_contexts)
n = len(contexts)
matrix = np.empty((n,n))
matrix[:] = np.NAN
print('constructing matrix')
tfidf_vectorizer = TfidfVectorizer()
tfidf = tfidf_vectorizer.fit_transform(contexts)
for i in tqdm(range(n)):
matrix[i,:] = metric(tfidf[i], tfidf).flatten()
print('saving')
filename = '../datasets/apple-rock-pear/tfidf_{}_w_{}_s_{}.npy'.format(metric.__name__, window, symmetric)
np.save(filename, matrix)
for metric in [cosine_similarity, cosine_distances]:
for window in [2,3,4]:
for symmetric in [True, False]:
print(metric.__name__, window, symmetric)
apple_contexts = open('../datasets/apple_contexts_s_{}_w_{}.txt'.format(symmetric, window)).read().splitlines()
rock_contexts = open('../datasets/rock_contexts_s_{}_w_{}.txt'.format(symmetric, window)).read().splitlines()
pear_contexts = open('../datasets/pear_contexts_s_{}_w_{}.txt'.format(symmetric, window)).read().splitlines()
contexts = apple_contexts + rock_contexts + pear_contexts
labels = [0]*len(apple_contexts) + [1]*len(rock_contexts) + [2]*len(pear_contexts)
n = len(contexts)
matrix = np.zeros((n,n))
print('constructing matrix')
tfidf_vectorizer = TfidfVectorizer()
tfidf = tfidf_vectorizer.fit_transform(contexts)
for i in tqdm(range(n)):
matrix[i,:] = metric(tfidf[i], tfidf).flatten()
print('saving')
filename = '../datasets/apple-rock-pear/tfidf_{}_w_{}_s_{}.npy'.format(metric.__name__, window, symmetric)
np.save(filename, matrix)
```
| github_jupyter |
## cuDF perf tests
### Loading financial time-series (per-minute ETFs) data from CSV files into a cuDF and running the queries
```
data_path = '/workspace/data/datasets/unianalytica/group/analytics-perf-tests/symbols/'
import sys
import os
import csv
import pandas as pd
import numpy as np
import cudf
from pymapd import connect
import pyarrow as pa
import pandas as pd
from datetime import datetime
import pytz
import time
```
### 1.Load up all files to one cuDF DataFrame
#### Reading the CSV files into a Pandas DF (takes about 2 minutes - 63 files, 3.5 GB CSV format total size)
```
symbol_dfs_list = []
records_count = 0
symbols_files = sorted(os.listdir(data_path))
for ix in range(len(symbols_files)):
current_symbol_df = pd.read_csv(data_path + symbols_files[ix], parse_dates=[2], infer_datetime_format=True,
names=['symbol_record_id', 'symbol', 'datetime', 'open', 'high', 'low', 'close', 'volume', 'split_factor', 'earnings', 'dividends'])
records_count = records_count + len(current_symbol_df)
symbol_dfs_list.append(current_symbol_df)
print('Finished reading; now concatenating the DFs...')
symbols_pandas_df = pd.concat(symbol_dfs_list)
symbols_pandas_df.index = np.arange(records_count)
del(symbol_dfs_list)
print('Built a Pandas DF of {} records.'.format(records_count))
symbols_pandas_df.head()
```
#### Building a cuDF from Pandas DF:
Replacing the `symbol` column here with `symbol_id`, as cuDF still cannot handle strings.
```
symbols_list = sorted(pd.unique(symbols_pandas_df.symbol))
print(symbols_list)
keys = symbols_list
values = list(range(1, len(symbols_list)+1))
dictionary = dict(zip(keys, values))
symbols_pandas_df.insert(0, 'symbol_id', np.array([dictionary[x] for x in symbols_pandas_df.symbol.values]))
symbols_pandas_df_cudf = symbols_pandas_df.drop('symbol', axis=1)
symbols_pandas_df_cudf.head()
symbols_pandas_df_cudf.dtypes
```
### Now, building the cuDF from Pandas DF:
```
symbols_gdf = cudf.DataFrame.from_pandas(symbols_pandas_df_cudf)
del(symbols_pandas_df_cudf)
print(symbols_gdf)
```
### 2.Perf Tests
#### 2.1 Descriptive statistics
```
%%timeit -n1 -r3
print('Trading volume stats: mean of {}, variance of {}'.format(symbols_gdf['volume'].mean(), symbols_gdf['volume'].var()))
```
#### 2.2 Sorting
```
%%timeit -n1 -r3
print(symbols_gdf[['symbol_id', 'datetime', 'volume']].sort_values(by='volume', ascending=False).head(1))
```
#### 2.3 Mixed analytics (math ops + sorting) [finding the top per-minute return]:
```
%%timeit -n1 -r3
symbols_gdf['return'] = 100*(symbols_gdf['close']-symbols_gdf['open'])/symbols_gdf['open']
print(symbols_gdf[['symbol_id', 'datetime', 'return']].sort_values(by='return', ascending=False).head(1))
```
## License
Copyright (c) 2019, PatternedScience Inc.
This code was originally run on the [UniAnalytica](https://www.unianalytica.com) platform, is published by PatternedScience Inc. on [GitHub](https://github.com/patternedscience/GPU-Analytics-Perf-Tests) and is licensed under the terms of Apache License 2.0; a copy of the license is available in the GitHub repository.
| github_jupyter |
# Tutorial 3: SQL data source
## Preparing
### Step 1. Install LightAutoML
Uncomment if doesn't clone repository by git. (ex.: colab, kaggle version)
```
#! pip install -U lightautoml
```
### Step 2. Import necessary libraries
```
# Standard python libraries
import os
import time
import requests
# Installed libraries
import numpy as np
import pandas as pd
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import train_test_split
import torch
# Imports from our package
import gensim
from lightautoml.automl.presets.tabular_presets import TabularAutoML, TabularUtilizedAutoML
from lightautoml.dataset.roles import DatetimeRole
from lightautoml.tasks import Task
```
### Step 3. Parameters
```
N_THREADS = 8 # threads cnt for lgbm and linear models
N_FOLDS = 5 # folds cnt for AutoML
RANDOM_STATE = 42 # fixed random state for various reasons
TEST_SIZE = 0.2 # Test size for metric check
TIMEOUT = 300 # Time in seconds for automl run
TARGET_NAME = 'TARGET' # Target column name
```
### Step 4. Fix torch number of threads and numpy seed
```
np.random.seed(RANDOM_STATE)
torch.set_num_threads(N_THREADS)
```
### Step 5. Example data load
Load a dataset from the repository if doesn't clone repository by git.
```
DATASET_DIR = '../data/'
DATASET_NAME = 'sampled_app_train.csv'
DATASET_FULLNAME = os.path.join(DATASET_DIR, DATASET_NAME)
DATASET_URL = 'https://raw.githubusercontent.com/sberbank-ai-lab/LightAutoML/master/examples/data/sampled_app_train.csv'
%%time
if not os.path.exists(DATASET_FULLNAME):
os.makedirs(DATASET_DIR, exist_ok=True)
dataset = requests.get(DATASET_URL).text
with open(DATASET_FULLNAME, 'w') as output:
output.write(dataset)
%%time
data = pd.read_csv(DATASET_FULLNAME)
data.head()
```
### Step 6. (Optional) Some user feature preparation
Cell below shows some user feature preparations to create task more difficult (this block can be omitted if you don't want to change the initial data):
```
%%time
data['BIRTH_DATE'] = (np.datetime64('2018-01-01') + data['DAYS_BIRTH'].astype(np.dtype('timedelta64[D]'))).astype(str)
data['EMP_DATE'] = (np.datetime64('2018-01-01') + np.clip(data['DAYS_EMPLOYED'], None, 0).astype(np.dtype('timedelta64[D]'))
).astype(str)
data['constant'] = 1
data['allnan'] = np.nan
data['report_dt'] = np.datetime64('2018-01-01')
data.drop(['DAYS_BIRTH', 'DAYS_EMPLOYED'], axis=1, inplace=True)
```
### Step 7. (Optional) Data splitting for train-test
Block below can be omitted if you are going to train model only or you have specific train and test files:
```
%%time
train_data, test_data = train_test_split(data,
test_size=TEST_SIZE,
stratify=data[TARGET_NAME],
random_state=RANDOM_STATE)
print('Data splitted. Parts sizes: train_data = {}, test_data = {}'
.format(train_data.shape, test_data.shape))
train_data.head()
```
### Step 8. (Optional) Reading data from SqlDataSource
#### Preparing datasets as SQLite data bases
```
import sqlite3 as sql
for _fname in ('train.db', 'test.db'):
if os.path.exists(_fname):
os.remove(_fname)
train_db = sql.connect('train.db')
train_data.to_sql('data', train_db)
test_db = sql.connect('test.db')
test_data.to_sql('data', test_db)
```
#### Using dataset wrapper for a connection
```
from lightautoml.reader.tabular_batch_generator import SqlDataSource
# train_data is replaced with a wrapper for an SQLAlchemy connection
# Wrapper requires SQLAlchemy connection string and query to obtain data from
train_data = SqlDataSource('sqlite:///train.db', 'select * from data', index='index')
test_data = SqlDataSource('sqlite:///test.db', 'select * from data', index='index')
```
## AutoML preset usage
### Step 1. Create Task
```
%%time
task = Task('binary', )
```
### Step 2. Setup columns roles
Roles setup here set target column and base date, which is used to calculate date differences:
```
%%time
roles = {'target': TARGET_NAME,
DatetimeRole(base_date=True, seasonality=(), base_feats=False): 'report_dt',
}
```
### Step 3. Create AutoML from preset
To create AutoML model here we use `TabularAutoML` preset, which looks like:

All params we set above can be send inside preset to change its configuration:
```
%%time
automl = TabularAutoML(task = task,
timeout = TIMEOUT,
general_params = {'nested_cv': False, 'use_algos': [['linear_l2', 'lgb', 'lgb_tuned']]},
reader_params = {'cv': N_FOLDS, 'random_state': RANDOM_STATE},
tuning_params = {'max_tuning_iter': 20, 'max_tuning_time': 30},
lgb_params = {'default_params': {'num_threads': N_THREADS}})
oof_pred = automl.fit_predict(train_data, roles = roles)
print('oof_pred:\n{}\nShape = {}'.format(oof_pred, oof_pred.shape))
```
### Step 4. Predict to test data and check scores
```
%%time
test_pred = automl.predict(test_data)
print('Prediction for test data:\n{}\nShape = {}'
.format(test_pred, test_pred.shape))
print('Check scores...')
print('OOF score: {}'.format(roc_auc_score(train_data.data[TARGET_NAME].values, oof_pred.data[:, 0])))
print('TEST score: {}'.format(roc_auc_score(test_data.data[TARGET_NAME].values, test_pred.data[:, 0])))
```
### Step 5. Create AutoML with time utilization
Below we are going to create specific AutoML preset for TIMEOUT utilization (try to spend it as much as possible):
```
%%time
automl = TabularUtilizedAutoML(task = task,
timeout = TIMEOUT,
general_params = {'nested_cv': False, 'use_algos': [['linear_l2', 'lgb', 'lgb_tuned']]},
reader_params = {'cv': N_FOLDS, 'random_state': RANDOM_STATE},
tuning_params = {'max_tuning_iter': 20, 'max_tuning_time': 30},
lgb_params = {'default_params': {'num_threads': N_THREADS}})
oof_pred = automl.fit_predict(train_data, roles = roles)
print('oof_pred:\n{}\nShape = {}'.format(oof_pred, oof_pred.shape))
```
### Step 6. Predict to test data and check scores for utilized automl
```
%%time
test_pred = automl.predict(test_data)
print('Prediction for test data:\n{}\nShape = {}'
.format(test_pred, test_pred.shape))
print('Check scores...')
print('OOF score: {}'.format(roc_auc_score(train_data.data[TARGET_NAME].values, oof_pred.data[:, 0])))
print('TEST score: {}'.format(roc_auc_score(test_data.data[TARGET_NAME].values, test_pred.data[:, 0])))
```
| github_jupyter |
# Chapter3 ニューラルネットワークの基本
## 3. 糖尿病の予後予測【サンプルコード】
```
# 必要なパッケージのインストール
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.datasets import load_diabetes
from sklearn.model_selection import train_test_split
import torch
from torch.utils.data import TensorDataset, DataLoader
from torch import nn
import torch.nn.functional as F
from torch import optim
```
## 3.1. 糖尿病(Diabetes)データセット
```
# データセットのロード
diabetes = load_diabetes()
# データセットの説明
print(diabetes.DESCR)
# データフレームに変換
df = pd.DataFrame(diabetes.data, columns=diabetes.feature_names)
# 1年後の進行度の追加
df['target'] = diabetes.target
print(df.head())
# 基本統計量の確認
print(df.describe())
# データセットの可視化
sns.pairplot(df, x_vars=diabetes.feature_names, y_vars='target')
plt.show()
```
## 3.2. 前準備
```
# データセットの読み込み
diabetes = load_diabetes()
data = diabetes.data # 特徴量
label = diabetes.target.reshape(-1, 1) # 一年後の糖尿病の進行度
# データセットのサイズの確認
print("data size: {}".format(data.shape))
print("label size: {}".format(label.shape))
```
## 3.3. 訓練データとテストデータの用意
```
# 学習データとテストデータを分割
train_data, test_data, train_label, test_label = train_test_split(
data, label, test_size=0.2)
# 学習データとテストデータのサイズの確認
print("train_data size: {}".format(len(train_data)))
print("test_data size: {}".format(len(test_data)))
print("train_label size: {}".format(len(train_label)))
print("test_label size: {}".format(len(test_label)))
# ndarrayをPyTorchのTensorに変換
train_x = torch.Tensor(train_data)
test_x = torch.Tensor(test_data)
train_y = torch.Tensor(train_label) # torch.float32のデータ型に
test_y = torch.Tensor(test_label) # torch.float32のデータ型に
# 特徴量とラベルを結合したデータセットを作成
train_dataset = TensorDataset(train_x, train_y)
test_dataset = TensorDataset(test_x, test_y)
# ミニバッチサイズを指定したデータローダーを作成
train_batch = DataLoader(
dataset=train_dataset, # データセットの指定
batch_size=20, # バッチサイズの指定
shuffle=True, # シャッフルするかどうかの指定
num_workers=2) # コアの数
test_batch = DataLoader(
dataset=test_dataset,
batch_size=20,
shuffle=False,
num_workers=2)
# ミニバッチデータセットの確認
for data, label in train_batch:
print("batch data size: {}".format(data.size())) # バッチの入力データサイズ
print("batch label size: {}".format(label.size())) # バッチのラベルサイズ
break
```
## 3.4. ニューラルネットワークの定義
```
# ニューラルネットワークの定義
class Net(nn.Module):
def __init__(self, D_in, H, D_out):
super(Net, self).__init__()
self.linear1 = nn.Linear(D_in, H)
self.linear2 = nn.Linear(H, H) # 追加
self.linear3 = nn.Linear(H, D_out)
self.dropout = nn.Dropout(p=0.5)
def forward(self, x):
x = F.relu(self.linear1(x))
x = F.relu(self.linear2(x)) # 追加
x = F.relu(self.linear2(x)) # 追加
x = self.dropout(x) # 追加
x = self.linear3(x)
return x
# ハイパーパラメータの定義
D_in = 10 # 入力次元: 10
H = 200 # 隠れ層次元: 200
D_out = 1 # 出力次元: 1
epoch = 100 # 学習回数: 100
# ネットワークのロード
# CPUとGPUどちらを使うかを指定
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
net = Net(D_in, H, D_out).to(device)
# デバイスの確認
print("Device: {}".format(device))
```
## 3.5. 損失関数と最適化関数の定義
```
# 損失関数の定義
criterion = nn.MSELoss() # 今回の損失関数(平均二乗誤差: MSE)
criterion2 = nn.L1Loss() # 参考用(平均絶対誤差: MAE)
# 最適化関数の定義
optimizer = optim.Adam(net.parameters())
```
## 3.6. 学習
```
# 損失を保存するリストを作成
train_loss_list = [] # 学習損失(MSE)
test_loss_list = [] # 評価損失(MSE)
train_mae_list = [] # 学習MAE
test_mae_list = [] # 評価MAE
# 学習(エポック)の実行
for i in range(epoch):
# エポックの進行状況を表示
print('---------------------------------------------')
print("Epoch: {}/{}".format(i+1, epoch))
# 損失の初期化
train_loss = 0 # 学習損失(MSE)
test_loss = 0 # 評価損失(MSE)
train_mae = 0 # 学習MAE
test_mae = 0 # 評価MAE
# ---------学習パート--------- #
# ニューラルネットワークを学習モードに設定
net.train()
# ミニバッチごとにデータをロードし学習
for data, label in train_batch:
# GPUにTensorを転送
data = data.to(device)
label = label.to(device)
# 勾配を初期化
optimizer.zero_grad()
# データを入力して予測値を計算(順伝播)
y_pred = net(data)
# 損失(誤差)を計算
loss = criterion(y_pred, label) # MSE
mae = criterion2(y_pred, label) # MAE
# 勾配の計算(逆伝搬)
loss.backward()
# パラメータ(重み)の更新
optimizer.step()
# ミニバッチごとの損失を蓄積
train_loss += loss.item() # MSE
train_mae += mae.item() # MAE
# ミニバッチの平均の損失を計算
batch_train_loss = train_loss / len(train_batch)
batch_train_mae = train_mae / len(train_batch)
# ---------学習パートはここまで--------- #
# ---------評価パート--------- #
# ニューラルネットワークを評価モードに設定
net.eval()
# 評価時の計算で自動微分機能をオフにする
with torch.no_grad():
for data, label in test_batch:
# GPUにTensorを転送
data = data.to(device)
label = label.to(device)
# データを入力して予測値を計算(順伝播)
y_pred = net(data)
# 損失(誤差)を計算
loss = criterion(y_pred, label) # MSE
mae = criterion2(y_pred, label) # MAE
# ミニバッチごとの損失を蓄積
test_loss += loss.item() # MSE
test_mae += mae.item() # MAE
# ミニバッチの平均の損失を計算
batch_test_loss = test_loss / len(test_batch)
batch_test_mae = test_mae / len(test_batch)
# ---------評価パートはここまで--------- #
# エポックごとに損失を表示
print("Train_Loss: {:.4f} Train_MAE: {:.4f}".format(
batch_train_loss, batch_train_mae))
print("Test_Loss: {:.4f} Test_MAE: {:.4f}".format(
batch_test_loss, batch_test_mae))
# 損失をリスト化して保存
train_loss_list.append(batch_train_loss)
test_loss_list.append(batch_test_loss)
train_mae_list.append(batch_train_mae)
test_mae_list.append(batch_test_mae)
```
## 3.7. 結果の可視化
```
# 損失(MSE)
plt.figure()
plt.title('Train and Test Loss')
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.plot(range(1, epoch+1), train_loss_list, color='blue',
linestyle='-', label='Train_Loss')
plt.plot(range(1, epoch+1), test_loss_list, color='red',
linestyle='--', label='Test_Loss')
plt.legend() # 凡例
# MAE
plt.figure()
plt.title('Train and Test MAE')
plt.xlabel('Epoch')
plt.ylabel('MAE')
plt.plot(range(1, epoch+1), train_mae_list, color='blue',
linestyle='-', label='Train_MAE')
plt.plot(range(1, epoch+1), test_mae_list, color='red',
linestyle='--', label='Test_MAE')
plt.legend() # 凡例
# 表示
plt.show()
```
| github_jupyter |
```
import pandas as pd
from sklearn import model_selection
from sklearn.ensemble import AdaBoostRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import GridSearchCV
from sklearn import metrics
import time
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.pylab import rcParams
import threading
import lightgbm as lgb
import catboost as cb
import xgboost as xgb
%matplotlib inline
import imblearn
train_df = pd.read_csv('train_small.csv', header=None)
test_df = pd.read_csv('test_small.csv')
irregulars = [14, 103, 128, 136, 152, 155, 160, 166, 177, 179, 182, 198]
train_df = train_df.iloc[:,[0, 1]+irregulars]
test_df = test_df.iloc[:, np.array([1] + irregulars)-1]
train_df
print(train_df.mean())
print(test_df.mean())
n_train = 60000
train_targets = train_df.iloc[:n_train,1]
train_data = train_df.iloc[:n_train,2:]
test_targets = train_df.iloc[n_train:,1]
test_data = train_df.iloc[n_train:,2:]
def auc2(m, train, test):
return (metrics.trainPredstrain_targets,m.predict(train)),
metrics.roc_auc_score(test_targets,m.predict(test)))
# Organise the data into numpy arrays so that it is handled correctly by imblearn
nump_train_data = train_data.to_numpy()
nump_train_data = nump_train_data.astype(float)
nump_train_targets = train_targets.to_numpy()
nump_train_targets = nump_train_targets.astype(int)
nump_test_data = test_data.to_numpy()
nump_test_data = nump_test_data.astype(float)
nump_test_targets = test_targets.to_numpy()
nump_test_targets = nump_test_targets.astype(int)
rus = imblearn.under_sampling.RandomUnderSampler(sampling_strategy='auto',return_indices=True)
X_rus, y_rus, id_rus = rus.fit_sample(nump_train_data, nump_train_targets)
lg = lgb.LGBMRegressor(silent=False)
param_dist = {"max_depth": [1,2, 3],
"learning_rate" : [0.1,0.5,1],
# "num_leaves": [10,15,20],
"num_leaves": [2,3],
"n_estimators": [300]
}
grid_search = GridSearchCV(lg, n_jobs=-1, param_grid=param_dist, cv = 3, scoring="roc_auc", verbose=5)
grid_search.fit(train_data,train_targets)
grid_search.best_estimator_
lg
auc2(grid_search.best_estimator_, train_data, test_data)
```
### Best predictors
500 rows 150 estimators (12s) : depth 3; rate 0.5; 10 leaves;
1000 rows 200 estimators (19s) : depth 2; rate 0.1; 10 leaves; 0.7361517358288425 auc (1 with training set)
10000 rows 300 estimators (1min1s) : depth 1; rate 0.5; 10 leaves; 0.8508816764850232 auc (0.941760012507842 with training set)
Does number of leaves impact ? NO default 31
60000 rows; 300 estiators; depth 1; rate 0.5; 10 leaves : (0.8986034603281355, 0.8795559277631042) (train then test)
60000 rows; 1000 estiators; depth 1; rate 0.5; 10 leaves : (0.9183773853908943, 0.8924150185745882) (train then test)
#### Shows that overfitting is not good ? Or just harder to have high auc with bigger amount of data.
60000 rows; 10000 estiators; depth 1; rate 0.5; 10 leaves : (0.9433394334525005, 0.8876156885481759) (train then test)
#### Now it seems clear
```
lggood = lgb.LGBMRegressor(silent=False, max_depth=1, learning_rate=0.5, n_estimators=1000)
start = time.time()
lggood.fit(train_data,train_targets)
end = time.time()
print(end-start)
auc2(lggood, train_data, test_data)
print(lggood)
```
rate 0.5 1000 estim : (0.9183773853908943, 0.8924150185745882) in 6.60472559928894s
rate 0.3 1000 estim : (0.9158483146682737, 0.8915313835384053) in 5.39690375328064s
0.3 800 5.174034118652344
(0.91017955230423, 0.8890010522021087)
0.5 1500 8.940380811691284
(0.9225535053779779, 0.892192660353454)
#### Overfitting :
rate 0.5 1000 estim depth 2 : (0.9736699095251268, 0.8562574888874572) 9.129353523254395s
rate 0.5 300 estim depth 2 : (0.9332266802884277, 0.8697272863922353) kinda 3.4s
rate 1 200 est depth 5 : (0.9997954560831425, 0.7009104769267109) in 5.8162806034088135
rate 1 200 est depth 5 : (1.0, 0.6860658377890871) in 26.31138586997986s
#### Mmmmmh
rate 0.1 200 est depth 5 : (0.9999726974419348, 0.8615919388434367) in 23.673893928527832s
0.03 1000 est : 25.953459978103638s (0.9924414078232998, 0.8748606369043784)
0.01 : 25.04612970352173 (0.9542057987408105, 0.8484477871545448)
0.01 3000 est : (0.9925341998563096, 0.8746814404432133) 78.72248482704163s (meh)
#### To few estimators in this case (?):
rate 0.1 300 estim depth 2 : (0.8918921360973638, 0.8568826902016364) 3.3066024780273438s
rate 0.1 1000 estim depth 2 : (0.9356584753554953, 0.8837796602890337) 8.71259069442749s
```
lgrus = lgb.LGBMRegressor(silent=False, max_depth=1, learning_rate=0.1, n_estimators=3000)
start = time.time()
lgrus.fit(X_rus,y_rus)
end = time.time()
print(end-start)
auc2(lgrus, train_data, test_data)
```
#### Balancing data
scale_pos_weight useless
with RUS:
rate 0.5 1000 estim : 1.704514980316162s : (0.9156733819245538, 0.8793312074556034)
rate 0.1 3000 estim : (0.9144309306569556, 0.888129979170693) 4.4809534549713135s
rate 0.1 4000 estim : (0.9172283755513937, 0.887266314501063)
rate 0.08 5000 estim : 7.250913619995117s (0.9161398215981195, 0.8879848182267172)
```
class Bag_pred:
def __init__(self, predictors=[], weights=None, scatter=True):
self.predictors = predictors
if weights==None:
self.weights = list(np.ones(len(predictors)))
else:
self.weights = list(weights)
self.scatter = scatter
def predict(self, Y):
predictions = [pred.predict(Y) for pred in self.predictors]
weights = np.array(self.weights)/sum(self.weights)
total = weights[0]*np.array(self.predictors[0].predict(Y))
total.shape = len(total)
for pred, weight in zip(predictions, weights[1:]):
prediction = weight*np.array(pred)
prediction.shape = len(prediction)
total += prediction
return total
def predict1(self, Y):
def Lambda(predictor,Y,predicts,i):
predicts[i] = predictor.predict(Y)
threads = []
predictions = 5*[0]
for i, pred in enumerate(self.predictors):
threads.append(threading.Thread(target=Lambda, args=(pred,Y,predictions,i)))
threads[i].start()
for i, _ in enumerate(self.predictors):
threads[i].join()
weights = np.array(self.weights)/sum(self.weights)
total = weights[0]*np.array(predictions[0])
total.shape = len(total)
for pred, weight in zip(predictions, weights[1:]):
prediction = weight*np.array(pred)
prediction.shape = len(prediction)
total += prediction
return total
def fit(self, X, y):
if(self.scatter):
factor = int(X/len(self.predictors))
for i, pred in in self.predictors:
low, high = factor*i, factor*(i+1)
yi = train_df.iloc[low:high,1]
Xi = train_df.iloc[low:high,2:]
pred.fit(Xi, yi)
else:
for pred in self.predictors:
pred.fit(X,y)
def append(self, pred, weight=1):
self.predictors.append(pred)
self.weights.append(weight)
# Let's BAG
Bag = Bag_pred()
start = time.time()
for i in range(6):
low, high = 10000*i, 10000*(i+1)
train_targetsi = train_df.iloc[low:high,1]
train_datai = train_df.iloc[low:high,2:]
def auci(m, train, test):
return (metrics.roc_auc_score(train_targetsi,m.predict(train)),
metrics.roc_auc_score(test_targets,m.predict(test)))
lgover = lgb.LGBMRegressor(silent=False, max_depth=5, learning_rate=0.035, n_estimators=200)
#lgover.fit(train_datai, train_targetsi)
#print(i, auci(lgover, train_datai, test_data))
Bag.append(lgover)
Bag.fit(train_data, train_targets)
end = time.time()
print(end-start)
start = time.time()
print(auc2(Bag, train_data, test_data))
end = time.time()
print(end-start)
# No reinit of the bag ....
6 depth 5 r 0.035 n 200 (0.9345731839134271, 0.8709206768451114) = with n=400
6 depth 3 r 0.035 n 200 (0.9265064602771006, 0.8644267645858833)
6 depth 3 r 0.035 n 400 (0.9235606891946174, 0.8654291481457622) in 13s
6 depth 5 r 0.035 n 100 ~0.86
6 depth 5 r 0.035 n 50 (0.9311799920441334, 0.8681047477935966) in 6 + 43
6 depth 5 r 0.035 n 50
cbr = cb.CatBoostRegressor(silent=True, depth=1, learning_rate=0.3, iterations=1000)
# l2 leaf reg ??
start = time.time()
cbr.fit(train_data,train_targets)
end = time.time()
print(end-start)
auc2(cbr, train_data, test_data)
```
rate 0.5 : (0.917555524639378, 0.8912588846657648) in 16.263858318328857s
rate 0.3 : (0.9127614850298509, 0.8919345487341365) in 14.216562747955322s
```
xgbr = xgb.XGBRegressor(silent=False, max_depth=1, learning_rate=0.5, n_estimators=300, objective='binary:logistic')
# min_child_weight ??
start = time.time()
xgbr.fit(train_data,train_targets)
end = time.time()
print(end-start)
auc2(xgbr, train_data, test_data)
xgbr
```
300 estimators : (0.9068527447700975, 0.8810668040971461) in 76.34813570976257
500 estimators : (0.9178513085660571, 0.8891569499022955) in 137.13004994392395
XGBoost is not very Parallel with stamps
```
lgr = lgb.LGBMRegressor(silent=False)
param_dist = {"max_depth": [1],
"learning_rate" : np.arange(0.2,0.8,0.1),
# "num_leaves": [10,15,20],
#"num_leaves": [2,3],
"n_estimators": [1000]
}
grid_search = GridSearchCV(lgr, n_jobs=-1, param_grid=param_dist, cv = 3, scoring="roc_auc", verbose=5)
start = time.time()
grid_search.fit(train_data,train_targets)
end = time.time()
print(end-start)
print(grid_search.best_estimator_.learning_rate)
print(auc2(grid_search.best_estimator_, train_data, test_data))
```
Learning rate does not seem to impact much
```
lgr = lgb.LGBMRegressor(silent=False)
param_dist = {"max_depth": [1],
"learning_rate" : [0.5],
# "num_leaves": [10,15,20],
#"num_leaves": [2,3],
"n_estimators": [800, 1000, 1500, 2000, 3000]
}
grid_search = GridSearchCV(lgr, n_jobs=-1, param_grid=param_dist, cv = 3, scoring="roc_auc", verbose=5)
start = time.time()
grid_search.fit(train_data,train_targets)
end = time.time()
print(end-start)
print(grid_search.best_estimator_.n_estimators)
print(auc2(grid_search.best_estimator_, train_data, test_data))
```
neither a precise number estims ?
```
from mlxtend.classifier import StackingCVClassifier
meta = DecisionTreeRegressor()
sclf = StackingCVClassifier(classifiers=[lggood, xgbr, cbr, lgrus],
meta_classifier=meta,
random_state=785)
sclf.fit(train_data,train_targets)
sclf.meta_classifier.fit(train_data,train_targets)
sclf.meta_classifier.tree_.
auc2(sclf.meta_classifier, train_data, test_data)
stack = [lggood, xgbr, cbr, lgrus, Bag]
mat = []
for model in stack:
mat.append(model.predict(train_data))
meta_df = pd.DataFrame(np.transpose(np.array(mat)))
from sklearn.linear_model import LogisticRegression
meta = LogisticRegression()
meta.fit(meta_df, train_targets)
stack = [lggood, xgbr, cbr, lgrus, Bag]
mat = []
for model in stack:
mat.append(model.predict(test_data))
meta_df_test = pd.DataFrame(np.transpose(np.array(mat)))
print(meta_df)
auc2(meta, meta_df, meta_df_test)
estimator = meta
n_nodes = estimator.tree_.node_count
children_left = estimator.tree_.children_left
children_right = estimator.tree_.children_right
feature = estimator.tree_.feature
threshold = estimator.tree_.threshold
# The tree structure can be traversed to compute various properties such
# as the depth of each node and whether or not it is a leaf.
node_depth = np.zeros(shape=n_nodes, dtype=np.int64)
is_leaves = np.zeros(shape=n_nodes, dtype=bool)
stack = [(0, -1)] # seed is the root node id and its parent depth
while len(stack) > 0:
node_id, parent_depth = stack.pop()
node_depth[node_id] = parent_depth + 1
# If we have a test node
if (children_left[node_id] != children_right[node_id]):
stack.append((children_left[node_id], parent_depth + 1))
stack.append((children_right[node_id], parent_depth + 1))
else:
is_leaves[node_id] = True
print("The binary tree structure has %s nodes and has "
"the following tree structure:"
% n_nodes)
for i in range(n_nodes):
if is_leaves[i]:
print("%snode=%s leaf node." % (node_depth[i] * "\t", i))
else:
print("%snode=%s test node: go to node %s if X[:, %s] <= %s else to "
"node %s."
% (node_depth[i] * "\t",
i,
children_left[i],
feature[i],
threshold[i],
children_right[i],
))
print()
# First let's retrieve the decision path of each sample. The decision_path
# method allows to retrieve the node indicator functions. A non zero element of
# indicator matrix at the position (i, j) indicates that the sample i goes
# through the node j.
node_indicator = estimator.decision_path(X_test)
# Similarly, we can also have the leaves ids reached by each sample.
leave_id = estimator.apply(X_test)
# Now, it's possible to get the tests that were used to predict a sample or
# a group of samples. First, let's make it for the sample.
sample_id = 0
node_index = node_indicator.indices[node_indicator.indptr[sample_id]:
node_indicator.indptr[sample_id + 1]]
print('Rules used to predict sample %s: ' % sample_id)
for node_id in node_index:
if leave_id[sample_id] == node_id:
continue
if (X_test[sample_id, feature[node_id]] <= threshold[node_id]):
threshold_sign = "<="
else:
threshold_sign = ">"
print("decision id node %s : (X_test[%s, %s] (= %s) %s %s)"
% (node_id,
sample_id,
feature[node_id],
X_test[sample_id, feature[node_id]],
threshold_sign,
threshold[node_id]))
# For a group of samples, we have the following common node.
sample_ids = [0, 1]
common_nodes = (node_indicator.toarray()[sample_ids].sum(axis=0) ==
len(sample_ids))
common_node_id = np.arange(n_nodes)[common_nodes]
print("\nThe following samples %s share the node %s in the tree"
% (sample_ids, common_node_id))
print("It is %s %% of all nodes." % (100 * len(common_node_id) / n_nodes,))
start = time.time()
scores = model_selection.cross_val_score(lggood, train_data, train_targets, cv=7, scoring='roc_auc', verbose=Tue)
end = time.time()
print(end-start)
print(scores)
print(scores.mean())
```
| github_jupyter |
```
from IPython.display import HTML
HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide();
} else {
$('div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
The raw code for this Jupyter notebook is by default hidden for easier reading.
To toggle on/off the raw code, click <a href="javascript:code_toggle()">here</a>.
<style>
.output_png {
display: table-cell;
text-align: center;
vertical-align: middle;
}
</style>
''')
```

### <h1><center>Module 2: Terminology of Digital Signal Processing</center></h1>
[Digital Signal Processing](https://en.wikipedia.org/wiki/Digital_signal_processing) (or DSP) is one of the *most powerful technologies* that will shape science and engineering in the twenty-first century. Revolutionary changes have already been made in a broad range of fields: communications, medical imaging, radar & sonar, high fidelity music reproduction, and oil prospecting, to name just a few.
Each of these areas has developed a deep DSP technology, with its own algorithms, mathematics, and specialized techniques. This combination of breath and depth makes it *impossible* for any one individual to master all of the DSP technology that has been developed. DSP education involves two tasks:
* learning general concepts that apply to the field as a whole; and
* learning specialized techniques for your particular area of interest.
The purpose of this module is to provide you with some of the key terminology that we will be covering in this DSP course: **signals**; **continuous, discrete** and **digital**; **systems**; and **processing**.
## Signal
A [signal](https://en.wikipedia.org/wiki/Signal) is anything that conveys **information** and is a description of how one (or a set of) **parameter(s)** relates to another parameter(s) (e.g., amplitude or voltage as a function of time). Examples of signals are everywhere, including:
* Seismic or radar pulse
* Speech
* DNA sequence
* Stock price
* Image
* Video
A signal can have single or multiple independent variables. For example, you'll be familiar with the following examples:
* 1D - Speech: $s=s(t)$
* 2D - Image: $I=I(x,y)$, Topography map: $elev=elev(lat,long)$
* 3D - 3D Seismic/GPR shot gather: $S = S(t,r_x,r_y)$
* 4D - EM/Seismic wavefield: $W=W(t,x,y,z)$
* 5D - 3D Seismic data set: $D=D(t,r_x,r_y,s_x,s_y)$
## Continuous, Discrete and Digital
There are three types of signals that are functions of *time*:
1. **Continuous-time** (analog) - $x(t)$: defined on a continuous range of time *t*, amplitude can be any value.
2. **Discrete-time** - $x(nT)$: defined only at discrete instants of time: $t=...-T,0,T,2T...$ where the amplitude can be any value.
3. **Digital** (quantized) - $x_Q[nT]$: both time and amplitude are discrete. Signals only defined at $t=...,-T,0,T,2T,...$ and amplitude is confined to a finite set of numbers.
<img src="Fig/1-SignalTypes.png" width="700">
**Figure 1. Illustrating the differences between continuous-time, discrete-time and digital signals.**
In DSP we deal with $x_Q[nT]$ because this corresponds with computer-based processing (which is quantized by definition - e.g. 16-bit vs 32-bit system). In this course we will assume that **discrete-time signal** is equivalent to a **digital signal** (equivalent to saying that the quantizer has infinite resolution). Thus, we will commonly write continuous and discrete (and quantized) signal as $x(t)$ and $x[nT]$, where parentheses and square brackets will denote continuity and discreteness, respectively.
## Systems
A **system** is a mathematical model or abstraction of a physical process that relates **input** to **output**. Any system that processes [digital](https://en.wikipedia.org/wiki/Digital_signal) signals is called a **digital system** or **digital signal processor**.
Examples include:
* Amplifier
* input: ${\rm cos}\,\omega t$
* output: $10\,{\rm cos}\,\omega t$
* Delay
* input: $f[nT]$
* output: $g[(n+p)T]$ where integer $p>0$
* Feature extraction
* Input: ${\rm cos}\,\omega_1 t + {\rm cos}\,\omega_2 t$
* Output: [$\omega_1,\omega_2$]
* Cellphone communication
* Input: Voice
* Output: CDMA signal
## Processing
**Processing** performs a particular function by passing a [signal](https://en.wikipedia.org/wiki/Signal)
through a **system**. Examples include:
* [Analog](https://en.wikipedia.org/wiki/Analog_signal) processing of **analog** signal
<img src="Fig/1-ASP.png" width="250">
**Figure 2. Illustrating the analog processing of an analog signal.**
* Digital processing of analog signal
<img src="Fig/1-A2D2A.png" width="700">
**Figure 3. Illustrating the steps required to digitially process an analog signal.**
## Signals vs. Underlying Processes
In most cases we aim to model geophysical phenomena using **deterministic equations** (e.g., acoustic wave equation, Maxwell's equations). Given an known input, these equations will generate a predictable output (for a noise-less system). However, distinguishing between the acquired **signal** and the **underlying stochastic process** is often very important.
For example, imagine creating a 1000 point signal by flipping a coin 1000 times. If the coin flip is heads, the corresponding sample is made a value of one. On tails, the sample is set to zero. The process that created this signal has a mean of exactly 0.5, determined by the relative probability of each possible outcome: 50% heads, 50% tails. However, it is unlikely that the actual 1000 point signal will have a mean of exactly 0.5. Random chance will make the number of ones and zeros slightly different each time the signal is generated. The probabilities of the underlying process are constant, but the statistics of the acquired signal change each time the experiment is repeated. This random irregularity found in actual data is called by such names as: **statistical variation**, **statistical fluctuation**, and **statistical noise**.
Finally, we live in a world where even though we understand many deterministic equations and systems of equations, the data we record and use are contaminated with both **coherent and incoherent noise**. Much of the signal processing work that you will do is trying to limit these sources of noise in order to enhance the signal. This is one of the main reasons why it is good to study digital signal processing!
## DSP versus ASP
It is worth mentioning that one can also perform various [Analog Signal Processing](https://en.wikipedia.org/wiki/Analog_signal_processing) (ASP) tasks. ASP is any type of signal processing conducted on continuous analog signals by some analog means. Examples include "bass", "treble" and "volume" controls on stereos, and "tint" controls on TVs.
There are a number of key advanages of DSP over ASP:
* Allows development with use of computers (e.g., with Python, Matlab)
* *Robust tool kits and modularity* - Leverage significant number of complex tools in Python/Matlab without having to redesign physical hardware every time.
* Allows *flexibility* in reconfiguring the DSP operators by changing the program (not the hardware!)
* *Reliable*: processing of 0's and 1's is almost immune to noise, and data are easily stored without deterioration
* *Security* through encryption/scrambling
* *Simplicity* - most operators are additions and subtractions (can be scalar-scalar, vector-scalar, vector-vector, matrix-vector)
However, there are also a number of advantages of ASP over DSP:
* Excellent for throughput signals
* Preclude the need for data storage and interface with computer CPU
| github_jupyter |
# Tutorial 0: Getting Started
Welcome to TenSEAL's first tutorial of a serie aiming at introducing homomorphic encryption and the capabilities of the library.
TenSEAL is a library for doing homomorphic encryption operations on tensors. It's built on top of [Microsoft SEAL](https://github.com/Microsoft/SEAL), a C++ library implementing the BFV and CKKS homomorphic encryption schemes. TenSEAL provides ease of use through a Python API, while preserving efficiency by implementing most of its operations using C++, so TenSEAL is a C++ library that have a Python interface.
Let's now start the tutorial with a brief review of what homomorphic encryption is, but keep in mind that you don't need to be a crypto expert to use the library.
Authors:
- Ayoub Benaissa - Twitter: [@y0uben11](https://twitter.com/y0uben11)
## Homomorphic Encryption
__Definition__ : Homomorphic encription (HE) is an encryption technique that allows computations to be made on ciphertexts and generates results that when decrypted, corresponds to the result of the same computations made on plaintexts.
<img src="assets/he-black-box.png" alt="he-black-box" width="600"/>
This means that an HE scheme lets you encrypt two numbers *X* and *Y*, add their encrypted versions so that it gets decrypted to *X + Y*, the addition could have been a multiplication as well. If we translate this to python, it may look something like this:
```python
x = 7
y = 3
x_encrypted = HE.encrypt(x)
y_encrypted = HE.encrypt(y)
z_encrypted = x_encrypted + y_encrypted
# z should now be x + y = 10
z = HE.decrypt(z_encrypted)
```
Many details are hidden in this python script, things like key generation doesn't appear, and that `+` operation over encrypted numbers isn't the usual `+` over integers, but a special evaluation algorithm that can evaluate addition over encrypted numbers. TenSEAL supports addition, substraction and multiplication of encrypted vectors of either integers (using BFV) or real numbers (using CKKS).
Next we will look at the most important object of the library, the TenSEALContext.
## TenSEALContext
The TenSEALContext is a special object that holds different encryption keys and parameters for you, so that you only need to lift a single object to make your encrypted computation instead of managing all they keys and the HE details. Basically, you will want to create a single TenSEALContext before doing your encrypted computation. Let's see how to create one !
```
import tenseal as ts
context = ts.context(ts.SCHEME_TYPE.BFV, poly_modulus_degree=4096, plain_modulus=1032193)
context
```
That's it ! We need to specify the HE scheme (BFV here) that we want to use, as well as its parameters. Don't worry about the parameters now, you will learn more about them in upcoming tutorials.
An important thing to note is that the TenSEALContext now is holding the secret-key and you can decrypt without the need to provide it, however, you can choose to manage it as a separate object and will need to pass it to functions that requires the secret-key. Let's see how this translates into python
```
public_context = ts.context(ts.SCHEME_TYPE.BFV, poly_modulus_degree=4096, plain_modulus=1032193)
print("Is the context private?", ("Yes" if public_context.is_private() else "No"))
print("Is the context public?", ("Yes" if public_context.is_public() else "No"))
sk = public_context.secret_key()
# the context will drop the secret-key at this point
public_context.make_context_public()
print("Secret-key dropped")
print("Is the context private?", ("Yes" if public_context.is_private() else "No"))
print("Is the context public?", ("Yes" if public_context.is_public() else "No"))
```
You can now try to fetch the secret-key from the `public_context` and see that it raises an error. We will now continue using our first created TenSEALContext `context` which is holding the secret-key.
## Encryption and Evaluation
The next step after holding our TenSEALContext is to start doing some encrypted computation. First, we create an encrypted vector of integer.
```
plain_vector = [60, 66, 73, 81, 90]
encrypted_vector = ts.bfv_vector(context, plain_vector)
print("We just encrypted our plaintext vector of size:", encrypted_vector.size())
encrypted_vector
```
Here we encrypted a vector of integers into a BFVVector, a vector type that uses the BFV scheme, now we can do both addition, substraction and multiplication in an element-wise fashion with other encrypted or plain vectors.
```
add_result = encrypted_vector + [1, 2, 3, 4, 5]
print(add_result.decrypt())
sub_result = encrypted_vector - [1, 2, 3, 4, 5]
print(sub_result.decrypt())
mul_result = encrypted_vector * [1, 2, 3, 4, 5]
print(mul_result.decrypt())
encrypted_add = add_result + sub_result
print(encrypted_add.decrypt())
encrypted_sub = encrypted_add - encrypted_vector
print(encrypted_sub.decrypt())
encrypted_mul = encrypted_add * encrypted_sub
print(encrypted_mul.decrypt())
```
We just made both ciphertext to plaintext (c2p) and ciphertext to ciphertext (c2c) evaluations (add, sub and mul). An important thing to note is that you should never encrypts your plaintext values to evaluate them with ciphertexts if there is no reason to do so, that's because c2p evaluations are more efficient than c2c. Look at the below script to see how a c2p multiplication is faster than a c2c one.
```
from time import time
t_start = time()
_ = encrypted_add * encrypted_mul
t_end = time()
print("c2c multiply time: {} ms".format((t_end - t_start) * 1000))
t_start = time()
_ = encrypted_add * [1, 2, 3, 4, 5]
t_end = time()
print("c2p multiply time: {} ms".format((t_end - t_start) * 1000))
```
## More about TenSEALContext
TenSEALContext is holding more attributes than what we have seen so far, so it's worth mentionning some of the interesting ones. The coolest attributes (at least for me) are the ones for setting automatic relinearization, rescaling (for CKKS only) and modulus switching, these features are enabled by defaut as you can see below
```
print("Automatic relinearization is:", ("on" if context.auto_relin else "off"))
print("Automatic rescaling is:", ("on" if context.auto_rescale else "off"))
print("Automatic modulus switching is:", ("on" if context.auto_mod_switch else "off"))
```
Experienced users can choose to disable one or many of these features and manage when and how to do these operations.
TenSEALContext can also holds a `global_scale` (only used when using CKKS) so that it's used as a default scale value when the user doesn't provide one. As most often users will define a single value to be used as scale during their HE computation, defining it globally can be helpful instead of passing it to every function call.
```
# this should throw an error as the global_scale isn't defined yet
try:
print("global_scale:", context.global_scale)
except ValueError:
print("The global_scale isn't defined yet")
# you can define it to 2 ** 20 for instance
context.global_scale = 2 ** 20
print("global_scale:", context.global_scale)
```
# Congratulations!!! - Time to Join the Community!
Congratulations on completing this notebook tutorial! If you enjoyed this and would like to join the movement toward privacy preserving, decentralized ownership of AI and the AI supply chain (data), you can do so in the following ways!
### Star TenSEAL on GitHub
The easiest way to help our community is just by starring the Repos! This helps raise awareness of the cool tools we're building.
- [Star TenSEAL](https://github.com/OpenMined/TenSEAL)
### Join our Slack!
The best way to keep up to date on the latest advancements is to join our community! You can do so by filling out the form at [http://slack.openmined.org](http://slack.openmined.org)
### Donate
If you don't have time to contribute to our codebase, but would still like to lend support, you can also become a Backer on our Open Collective. All donations go toward our web hosting and other community expenses such as hackathons and meetups!
[OpenMined's Open Collective Page](https://opencollective.com/openmined)
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
!wget --no-check-certificate \
https://storage.googleapis.com/laurencemoroney-blog.appspot.com/bbc-text.csv \
-O /tmp/bbc-text.csv
import csv
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
#Stopwords list from https://github.com/Yoast/YoastSEO.js/blob/develop/src/config/stopwords.js
# Convert it to a Python list and paste it here
stopwords = [ "a", "about", "above", "after", "again", "against", "all", "am", "an", "and", "any", "are", "as", "at", "be", "because", "been", "before", "being", "below", "between", "both", "but", "by", "could", "did", "do", "does", "doing", "down", "during", "each", "few", "for", "from", "further", "had", "has", "have", "having", "he", "he'd", "he'll", "he's", "her", "here", "here's", "hers", "herself", "him", "himself", "his", "how", "how's", "i", "i'd", "i'll", "i'm", "i've", "if", "in", "into", "is", "it", "it's", "its", "itself", "let's", "me", "more", "most", "my", "myself", "nor", "of", "on", "once", "only", "or", "other", "ought", "our", "ours", "ourselves", "out", "over", "own", "same", "she", "she'd", "she'll", "she's", "should", "so", "some", "such", "than", "that", "that's", "the", "their", "theirs", "them", "themselves", "then", "there", "there's", "these", "they", "they'd", "they'll", "they're", "they've", "this", "those", "through", "to", "too", "under", "until", "up", "very", "was", "we", "we'd", "we'll", "we're", "we've", "were", "what", "what's", "when", "when's", "where", "where's", "which", "while", "who", "who's", "whom", "why", "why's", "with", "would", "you", "you'd", "you'll", "you're", "you've", "your", "yours", "yourself", "yourselves" ]
sentences = []
labels = []
with open("/tmp/bbc-text.csv", 'r') as csvfile:
fr = csv.reader(csvfile, delimiter=',')
next(fr)
for row in fr:
labels.append(row[0])
sentence = row[1]
for word in stopwords:
token = " "+word+" "
sentence = sentence.replace(token, " ")
sentence = sentence.replace(" ", " ")
sentences.append(sentence)
print(len(sentences))
print(sentences[0])
#Expected output
# 2225
# tv future hands viewers home theatre systems plasma high-definition tvs digital video recorders moving living room way people watch tv will radically different five years time. according expert panel gathered annual consumer electronics show las vegas discuss new technologies will impact one favourite pastimes. us leading trend programmes content will delivered viewers via home networks cable satellite telecoms companies broadband service providers front rooms portable devices. one talked-about technologies ces digital personal video recorders (dvr pvr). set-top boxes like us s tivo uk s sky+ system allow people record store play pause forward wind tv programmes want. essentially technology allows much personalised tv. also built-in high-definition tv sets big business japan us slower take off europe lack high-definition programming. not can people forward wind adverts can also forget abiding network channel schedules putting together a-la-carte entertainment. us networks cable satellite companies worried means terms advertising revenues well brand identity viewer loyalty channels. although us leads technology moment also concern raised europe particularly growing uptake services like sky+. happens today will see nine months years time uk adam hume bbc broadcast s futurologist told bbc news website. likes bbc no issues lost advertising revenue yet. pressing issue moment commercial uk broadcasters brand loyalty important everyone. will talking content brands rather network brands said tim hanlon brand communications firm starcom mediavest. reality broadband connections anybody can producer content. added: challenge now hard promote programme much choice. means said stacey jolna senior vice president tv guide tv group way people find content want watch simplified tv viewers. means networks us terms channels take leaf google s book search engine future instead scheduler help people find want watch. kind channel model might work younger ipod generation used taking control gadgets play them. might not suit everyone panel recognised. older generations comfortable familiar schedules channel brands know getting. perhaps not want much choice put hands mr hanlon suggested. end kids just diapers pushing buttons already - everything possible available said mr hanlon. ultimately consumer will tell market want. 50 000 new gadgets technologies showcased ces many enhancing tv-watching experience. high-definition tv sets everywhere many new models lcd (liquid crystal display) tvs launched dvr capability built instead external boxes. one example launched show humax s 26-inch lcd tv 80-hour tivo dvr dvd recorder. one us s biggest satellite tv companies directtv even launched branded dvr show 100-hours recording capability instant replay search function. set can pause rewind tv 90 hours. microsoft chief bill gates announced pre-show keynote speech partnership tivo called tivotogo means people can play recorded programmes windows pcs mobile devices. reflect increasing trend freeing multimedia people can watch want want.
tokenizer = Tokenizer(oov_token="<OOV>")
tokenizer.fit_on_texts(sentences)
word_index = tokenizer.word_index
print(len(word_index))
# Expected output
# 29714
sequences = tokenizer.texts_to_sequences(sentences)
padded = pad_sequences(sequences, padding='post')
print(padded[0])
print(padded.shape)
# Expected output
# [ 96 176 1158 ... 0 0 0]
# (2225, 2442)
label_tokenizer = Tokenizer()
label_tokenizer.fit_on_texts(labels)
label_word_index = label_tokenizer.word_index
label_seq = label_tokenizer.texts_to_sequences(labels)
print(label_seq)
print(label_word_index)
# Expected Output
# [[4], [2], [1], [1], [5], [3], [3], [1], [1], [5], [5], [2], [2], [3], [1], [2], [3], [1], [2], [4], [4], [4], [1], [1], [4], [1], [5], [4], [3], [5], [3], [4], [5], [5], [2], [3], [4], [5], [3], [2], [3], [1], [2], [1], [4], [5], [3], [3], [3], [2], [1], [3], [2], [2], [1], [3], [2], [1], [1], [2], [2], [1], [2], [1], [2], [4], [2], [5], [4], [2], [3], [2], [3], [1], [2], [4], [2], [1], [1], [2], [2], [1], [3], [2], [5], [3], [3], [2], [5], [2], [1], [1], [3], [1], [3], [1], [2], [1], [2], [5], [5], [1], [2], [3], [3], [4], [1], [5], [1], [4], [2], [5], [1], [5], [1], [5], [5], [3], [1], [1], [5], [3], [2], [4], [2], [2], [4], [1], [3], [1], [4], [5], [1], [2], [2], [4], [5], [4], [1], [2], [2], [2], [4], [1], [4], [2], [1], [5], [1], [4], [1], [4], [3], [2], [4], [5], [1], [2], [3], [2], [5], [3], [3], [5], [3], [2], [5], [3], [3], [5], [3], [1], [2], [3], [3], [2], [5], [1], [2], [2], [1], [4], [1], [4], [4], [1], [2], [1], [3], [5], [3], [2], [3], [2], [4], [3], [5], [3], [4], [2], [1], [2], [1], [4], [5], [2], [3], [3], [5], [1], [5], [3], [1], [5], [1], [1], [5], [1], [3], [3], [5], [4], [1], [3], [2], [5], [4], [1], [4], [1], [5], [3], [1], [5], [4], [2], [4], [2], [2], [4], [2], [1], [2], [1], [2], [1], [5], [2], [2], [5], [1], [1], [3], [4], [3], [3], [3], [4], [1], [4], [3], [2], [4], [5], [4], [1], [1], [2], [2], [3], [2], [4], [1], [5], [1], [3], [4], [5], [2], [1], [5], [1], [4], [3], [4], [2], [2], [3], [3], [1], [2], [4], [5], [3], [4], [2], [5], [1], [5], [1], [5], [3], [2], [1], [2], [1], [1], [5], [1], [3], [3], [2], [5], [4], [2], [1], [2], [5], [2], [2], [2], [3], [2], [3], [5], [5], [2], [1], [2], [3], [2], [4], [5], [2], [1], [1], [5], [2], [2], [3], [4], [5], [4], [3], [2], [1], [3], [2], [5], [4], [5], [4], [3], [1], [5], [2], [3], [2], [2], [3], [1], [4], [2], [2], [5], [5], [4], [1], [2], [5], [4], [4], [5], [5], [5], [3], [1], [3], [4], [2], [5], [3], [2], [5], [3], [3], [1], [1], [2], [3], [5], [2], [1], [2], [2], [1], [2], [3], [3], [3], [1], [4], [4], [2], [4], [1], [5], [2], [3], [2], [5], [2], [3], [5], [3], [2], [4], [2], [1], [1], [2], [1], [1], [5], [1], [1], [1], [4], [2], [2], [2], [3], [1], [1], [2], [4], [2], [3], [1], [3], [4], [2], [1], [5], [2], [3], [4], [2], [1], [2], [3], [2], [2], [1], [5], [4], [3], [4], [2], [1], [2], [5], [4], [4], [2], [1], [1], [5], [3], [3], [3], [1], [3], [4], [4], [5], [3], [4], [5], [2], [1], [1], [4], [2], [1], [1], [3], [1], [1], [2], [1], [5], [4], [3], [1], [3], [4], [2], [2], [2], [4], [2], [2], [1], [1], [1], [1], [2], [4], [5], [1], [1], [4], [2], [4], [5], [3], [1], [2], [3], [2], [4], [4], [3], [4], [2], [1], [2], [5], [1], [3], [5], [1], [1], [3], [4], [5], [4], [1], [3], [2], [5], [3], [2], [5], [1], [1], [4], [3], [5], [3], [5], [3], [4], [3], [5], [1], [2], [1], [5], [1], [5], [4], [2], [1], [3], [5], [3], [5], [5], [5], [3], [5], [4], [3], [4], [4], [1], [1], [4], [4], [1], [5], [5], [1], [4], [5], [1], [1], [4], [2], [3], [4], [2], [1], [5], [1], [5], [3], [4], [5], [5], [2], [5], [5], [1], [4], [4], [3], [1], [4], [1], [3], [3], [5], [4], [2], [4], [4], [4], [2], [3], [3], [1], [4], [2], [2], [5], [5], [1], [4], [2], [4], [5], [1], [4], [3], [4], [3], [2], [3], [3], [2], [1], [4], [1], [4], [3], [5], [4], [1], [5], [4], [1], [3], [5], [1], [4], [1], [1], [3], [5], [2], [3], [5], [2], [2], [4], [2], [5], [4], [1], [4], [3], [4], [3], [2], [3], [5], [1], [2], [2], [2], [5], [1], [2], [5], [5], [1], [5], [3], [3], [3], [1], [1], [1], [4], [3], [1], [3], [3], [4], [3], [1], [2], [5], [1], [2], [2], [4], [2], [5], [5], [5], [2], [5], [5], [3], [4], [2], [1], [4], [1], [1], [3], [2], [1], [4], [2], [1], [4], [1], [1], [5], [1], [2], [1], [2], [4], [3], [4], [2], [1], [1], [2], [2], [2], [2], [3], [1], [2], [4], [2], [1], [3], [2], [4], [2], [1], [2], [3], [5], [1], [2], [3], [2], [5], [2], [2], [2], [1], [3], [5], [1], [3], [1], [3], [3], [2], [2], [1], [4], [5], [1], [5], [2], [2], [2], [4], [1], [4], [3], [4], [4], [4], [1], [4], [4], [5], [5], [4], [1], [5], [4], [1], [1], [2], [5], [4], [2], [1], [2], [3], [2], [5], [4], [2], [3], [2], [4], [1], [2], [5], [2], [3], [1], [5], [3], [1], [2], [1], [3], [3], [1], [5], [5], [2], [2], [1], [4], [4], [1], [5], [4], [4], [2], [1], [5], [4], [1], [1], [2], [5], [2], [2], [2], [5], [1], [5], [4], [4], [4], [3], [4], [4], [5], [5], [1], [1], [3], [2], [5], [1], [3], [5], [4], [3], [4], [4], [2], [5], [3], [4], [3], [3], [1], [3], [3], [5], [4], [1], [3], [1], [5], [3], [2], [2], [3], [1], [1], [1], [5], [4], [4], [2], [5], [1], [3], [4], [3], [5], [4], [4], [2], [2], [1], [2], [2], [4], [3], [5], [2], [2], [2], [2], [2], [4], [1], [3], [4], [4], [2], [2], [5], [3], [5], [1], [4], [1], [5], [1], [4], [1], [2], [1], [3], [3], [5], [2], [1], [3], [3], [1], [5], [3], [2], [4], [1], [2], [2], [2], [5], [5], [4], [4], [2], [2], [5], [1], [2], [5], [4], [4], [2], [2], [1], [1], [1], [3], [3], [1], [3], [1], [2], [5], [1], [4], [5], [1], [1], [2], [2], [4], [4], [1], [5], [1], [5], [1], [5], [3], [5], [5], [4], [5], [2], [2], [3], [1], [3], [4], [2], [3], [1], [3], [1], [5], [1], [3], [1], [1], [4], [5], [1], [3], [1], [1], [2], [4], [5], [3], [4], [5], [3], [5], [3], [5], [5], [4], [5], [3], [5], [5], [4], [4], [1], [1], [5], [5], [4], [5], [3], [4], [5], [2], [4], [1], [2], [5], [5], [4], [5], [4], [2], [5], [1], [5], [2], [1], [2], [1], [3], [4], [5], [3], [2], [5], [5], [3], [2], [5], [1], [3], [1], [2], [2], [2], [2], [2], [5], [4], [1], [5], [5], [2], [1], [4], [4], [5], [1], [2], [3], [2], [3], [2], [2], [5], [3], [2], [2], [4], [3], [1], [4], [5], [3], [2], [2], [1], [5], [3], [4], [2], [2], [3], [2], [1], [5], [1], [5], [4], [3], [2], [2], [4], [2], [2], [1], [2], [4], [5], [3], [2], [3], [2], [1], [4], [2], [3], [5], [4], [2], [5], [1], [3], [3], [1], [3], [2], [4], [5], [1], [1], [4], [2], [1], [5], [4], [1], [3], [1], [2], [2], [2], [3], [5], [1], [3], [4], [2], [2], [4], [5], [5], [4], [4], [1], [1], [5], [4], [5], [1], [3], [4], [2], [1], [5], [2], [2], [5], [1], [2], [1], [4], [3], [3], [4], [5], [3], [5], [2], [2], [3], [1], [4], [1], [1], [1], [3], [2], [1], [2], [4], [1], [2], [2], [1], [3], [4], [1], [2], [4], [1], [1], [2], [2], [2], [2], [3], [5], [4], [2], [2], [1], [2], [5], [2], [5], [1], [3], [2], [2], [4], [5], [2], [2], [2], [3], [2], [3], [4], [5], [3], [5], [1], [4], [3], [2], [4], [1], [2], [2], [5], [4], [2], [2], [1], [1], [5], [1], [3], [1], [2], [1], [2], [3], [3], [2], [3], [4], [5], [1], [2], [5], [1], [3], [3], [4], [5], [2], [3], [3], [1], [4], [2], [1], [5], [1], [5], [1], [2], [1], [3], [5], [4], [2], [1], [3], [4], [1], [5], [2], [1], [5], [1], [4], [1], [4], [3], [1], [2], [5], [4], [4], [3], [4], [5], [4], [1], [2], [4], [2], [5], [1], [4], [3], [3], [3], [3], [5], [5], [5], [2], [3], [3], [1], [1], [4], [1], [3], [2], [2], [4], [1], [4], [2], [4], [3], [3], [1], [2], [3], [1], [2], [4], [2], [2], [5], [5], [1], [2], [4], [4], [3], [2], [3], [1], [5], [5], [3], [3], [2], [2], [4], [4], [1], [1], [3], [4], [1], [4], [2], [1], [2], [3], [1], [5], [2], [4], [3], [5], [4], [2], [1], [5], [4], [4], [5], [3], [4], [5], [1], [5], [1], [1], [1], [3], [4], [1], [2], [1], [1], [2], [4], [1], [2], [5], [3], [4], [1], [3], [4], [5], [3], [1], [3], [4], [2], [5], [1], [3], [2], [4], [4], [4], [3], [2], [1], [3], [5], [4], [5], [1], [4], [2], [3], [5], [4], [3], [1], [1], [2], [5], [2], [2], [3], [2], [2], [3], [4], [5], [3], [5], [5], [2], [3], [1], [3], [5], [1], [5], [3], [5], [5], [5], [2], [1], [3], [1], [5], [4], [4], [2], [3], [5], [2], [1], [2], [3], [3], [2], [1], [4], [4], [4], [2], [3], [3], [2], [1], [1], [5], [2], [1], [1], [3], [3], [3], [5], [3], [2], [4], [2], [3], [5], [5], [2], [1], [3], [5], [1], [5], [3], [3], [2], [3], [1], [5], [5], [4], [4], [4], [4], [3], [4], [2], [4], [1], [1], [5], [2], [4], [5], [2], [4], [1], [4], [5], [5], [3], [3], [1], [2], [2], [4], [5], [1], [3], [2], [4], [5], [3], [1], [5], [3], [3], [4], [1], [3], [2], [3], [5], [4], [1], [3], [5], [5], [2], [1], [4], [4], [1], [5], [4], [3], [4], [1], [3], [3], [1], [5], [1], [3], [1], [4], [5], [1], [5], [2], [2], [5], [5], [5], [4], [1], [2], [2], [3], [3], [2], [3], [5], [1], [1], [4], [3], [1], [2], [1], [2], [4], [1], [1], [2], [5], [1], [1], [4], [1], [2], [3], [2], [5], [4], [5], [3], [2], [5], [3], [5], [3], [3], [2], [1], [1], [1], [4], [4], [1], [3], [5], [4], [1], [5], [2], [5], [3], [2], [1], [4], [2], [1], [3], [2], [5], [5], [5], [3], [5], [3], [5], [1], [5], [1], [3], [3], [2], [3], [4], [1], [4], [1], [2], [3], [4], [5], [5], [3], [5], [3], [1], [1], [3], [2], [4], [1], [3], [3], [5], [1], [3], [3], [2], [4], [4], [2], [4], [1], [1], [2], [3], [2], [4], [1], [4], [3], [5], [1], [2], [1], [5], [4], [4], [1], [3], [1], [2], [1], [2], [1], [1], [5], [5], [2], [4], [4], [2], [4], [2], [2], [1], [1], [3], [1], [4], [1], [4], [1], [1], [2], [2], [4], [1], [2], [4], [4], [3], [1], [2], [5], [5], [4], [3], [1], [1], [4], [2], [4], [5], [5], [3], [3], [2], [5], [1], [5], [5], [2], [1], [3], [4], [2], [1], [5], [4], [3], [3], [1], [1], [2], [2], [2], [2], [2], [5], [2], [3], [3], [4], [4], [5], [3], [5], [2], [3], [1], [1], [2], [4], [2], [4], [1], [2], [2], [3], [1], [1], [3], [3], [5], [5], [3], [2], [3], [3], [2], [4], [3], [3], [3], [3], [3], [5], [5], [4], [3], [1], [3], [1], [4], [1], [1], [1], [5], [4], [5], [4], [1], [4], [1], [1], [5], [5], [2], [5], [5], [3], [2], [1], [4], [4], [3], [2], [1], [2], [5], [1], [3], [5], [1], [1], [2], [3], [4], [4], [2], [2], [1], [3], [5], [1], [1], [3], [5], [4], [1], [5], [2], [3], [1], [3], [4], [5], [1], [3], [2], [5], [3], [5], [3], [1], [3], [2], [2], [3], [2], [4], [1], [2], [5], [2], [1], [1], [5], [4], [3], [4], [3], [3], [1], [1], [1], [2], [4], [5], [2], [1], [2], [1], [2], [4], [2], [2], [2], [2], [1], [1], [1], [2], [2], [5], [2], [2], [2], [1], [1], [1], [4], [2], [1], [1], [1], [2], [5], [4], [4], [4], [3], [2], [2], [4], [2], [4], [1], [1], [3], [3], [3], [1], [1], [3], [3], [4], [2], [1], [1], [1], [1], [2], [1], [2], [2], [2], [2], [1], [3], [1], [4], [4], [1], [4], [2], [5], [2], [1], [2], [4], [4], [3], [5], [2], [5], [2], [4], [3], [5], [3], [5], [5], [4], [2], [4], [4], [2], [3], [1], [5], [2], [3], [5], [2], [4], [1], [4], [3], [1], [3], [2], [3], [3], [2], [2], [2], [4], [3], [2], [3], [2], [5], [3], [1], [3], [3], [1], [5], [4], [4], [2], [4], [1], [2], [2], [3], [1], [4], [4], [4], [1], [5], [1], [3], [2], [3], [3], [5], [4], [2], [4], [1], [5], [5], [1], [2], [5], [4], [4], [1], [5], [2], [3], [3], [3], [4], [4], [2], [3], [2], [3], [3], [5], [1], [4], [2], [4], [5], [4], [4], [1], [3], [1], [1], [3], [5], [5], [2], [3], [3], [1], [2], [2], [4], [2], [4], [4], [1], [2], [3], [1], [2], [2], [1], [4], [1], [4], [5], [1], [1], [5], [2], [4], [1], [1], [3], [4], [2], [3], [1], [1], [3], [5], [4], [4], [4], [2], [1], [5], [5], [4], [2], [3], [4], [1], [1], [4], [4], [3], [2], [1], [5], [5], [1], [5], [4], [4], [2], [2], [2], [1], [1], [4], [1], [2], [4], [2], [2], [1], [2], [3], [2], [2], [4], [2], [4], [3], [4], [5], [3], [4], [5], [1], [3], [5], [2], [4], [2], [4], [5], [4], [1], [2], [2], [3], [5], [3], [1]]
# {'sport': 1, 'business': 2, 'politics': 3, 'tech': 4, 'entertainment': 5}
```
| github_jupyter |
# Adadelta
:label:`sec_adadelta`
Adadelta is yet another variant of AdaGrad (:numref:`sec_adagrad`). The main difference lies in the fact that it decreases the amount by which the learning rate is adaptive to coordinates. Moreover, traditionally it referred to as not having a learning rate since it uses the amount of change itself as calibration for future change. The algorithm was proposed in :cite:`Zeiler.2012`. It is fairly straightforward, given the discussion of previous algorithms so far.
## The Algorithm
In a nutshell, Adadelta uses two state variables, $\mathbf{s}_t$ to store a leaky average of the second moment of the gradient and $\Delta\mathbf{x}_t$ to store a leaky average of the second moment of the change of parameters in the model itself. Note that we use the original notation and naming of the authors for compatibility with other publications and implementations (there is no other real reason why one should use different Greek variables to indicate a parameter serving the same purpose in momentum, Adagrad, RMSProp, and Adadelta).
Here are the technical details of Adadelta. Given the parameter du jour is $\rho$, we obtain the following leaky updates similarly to :numref:`sec_rmsprop`:
$$\begin{aligned}
\mathbf{s}_t & = \rho \mathbf{s}_{t-1} + (1 - \rho) \mathbf{g}_t^2.
\end{aligned}$$
The difference to :numref:`sec_rmsprop` is that we perform updates with the rescaled gradient $\mathbf{g}_t'$, i.e.,
$$\begin{aligned}
\mathbf{x}_t & = \mathbf{x}_{t-1} - \mathbf{g}_t'. \\
\end{aligned}$$
So what is the rescaled gradient $\mathbf{g}_t'$? We can calculate it as follows:
$$\begin{aligned}
\mathbf{g}_t' & = \frac{\sqrt{\Delta\mathbf{x}_{t-1} + \epsilon}}{\sqrt{{\mathbf{s}_t + \epsilon}}} \odot \mathbf{g}_t, \\
\end{aligned}$$
where $\Delta \mathbf{x}_{t-1}$ is the leaky average of the squared rescaled gradients $\mathbf{g}_t'$. We initialize $\Delta \mathbf{x}_{0}$ to be $0$ and update it at each step with $\mathbf{g}_t'$, i.e.,
$$\begin{aligned}
\Delta \mathbf{x}_t & = \rho \Delta\mathbf{x}_{t-1} + (1 - \rho) {\mathbf{g}_t'}^2,
\end{aligned}$$
and $\epsilon$ (a small value such as $10^{-5}$) is added to maintain numerical stability.
## Implementation
Adadelta needs to maintain two state variables for each variable, $\mathbf{s}_t$ and $\Delta\mathbf{x}_t$. This yields the following implementation.
```
%matplotlib inline
from d2l import tensorflow as d2l
import tensorflow as tf
def init_adadelta_states(feature_dim):
s_w = tf.Variable(tf.zeros((feature_dim, 1)))
s_b = tf.Variable(tf.zeros(1))
delta_w = tf.Variable(tf.zeros((feature_dim, 1)))
delta_b = tf.Variable(tf.zeros(1))
return ((s_w, delta_w), (s_b, delta_b))
def adadelta(params, grads, states, hyperparams):
rho, eps = hyperparams['rho'], 1e-5
for p, (s, delta), grad in zip(params, states, grads):
s[:].assign(rho * s + (1 - rho) * tf.math.square(grad))
g = (tf.math.sqrt(delta + eps) / tf.math.sqrt(s + eps)) * grad
p[:].assign(p - g)
delta[:].assign(rho * delta + (1 - rho) * g * g)
```
Choosing $\rho = 0.9$ amounts to a half-life time of 10 for each parameter update. This tends to work quite well. We get the following behavior.
```
data_iter, feature_dim = d2l.get_data_ch11(batch_size=10)
d2l.train_ch11(adadelta, init_adadelta_states(feature_dim),
{'rho': 0.9}, data_iter, feature_dim);
```
For a concise implementation we simply use the `adadelta` algorithm from the `Trainer` class. This yields the following one-liner for a much more compact invocation.
```
# adadelta is not converging at default learning rate
# but it's converging at lr = 5.0
trainer = tf.keras.optimizers.Adadelta
d2l.train_concise_ch11(trainer, {'learning_rate':5.0, 'rho': 0.9}, data_iter)
```
## Summary
* Adadelta has no learning rate parameter. Instead, it uses the rate of change in the parameters itself to adapt the learning rate.
* Adadelta requires two state variables to store the second moments of gradient and the change in parameters.
* Adadelta uses leaky averages to keep a running estimate of the appropriate statistics.
## Exercises
1. Adjust the value of $\rho$. What happens?
1. Show how to implement the algorithm without the use of $\mathbf{g}_t'$. Why might this be a good idea?
1. Is Adadelta really learning rate free? Could you find optimization problems that break Adadelta?
1. Compare Adadelta to Adagrad and RMS prop to discuss their convergence behavior.
[Discussions](https://discuss.d2l.ai/t/1077)
| github_jupyter |
```
import math
import numpy as np
from scipy.integrate import odeint
import matplotlib.pyplot as plt
m = 1 # kg
k_air=0.00324844 # kg/m
F_M = 12.2 # N/kg
g = 9.8 # m/s^2
epsilon = 0.4
miu = 0.6
r_wheel = 0.75 # m
theta = 10
A=F_M-(m*g*miu)/(r_wheel)-m*g*(math.sin(math.pi*theta/180.00))
# function that returns dy/dt
def velocity(v,t):
dvdt = ( F_M-k_air*(v**2)-(m*g*miu)/(r_wheel)-m*g*(math.sin(math.pi*theta/180.00)) )/m
return dvdt
# initial condition
v0 = 0
# time points
t = np.linspace(0,70)
# solve ODE
v = odeint(velocity,v0,t)
analytical = [0 for i in range(len(t))]
for i in range(len(t)):
analytical[i] = (math.sqrt(A)*np.tanh( (t[i]*math.sqrt(A*k_air))/m ))/(math.sqrt(k_air))
l = [23.69322586 for i in range (len(t))]
fig = plt.figure(figsize=(5,5), dpi=80)
# plot results
plt.plot(t,v)
plt.plot(t,l,':')
plt.plot(t,analytical)
plt.xlabel('time')
plt.ylabel('velocity')
plt.legend(['y=v(t)','maximum possible velocity with human force\n 23.69 m/s','analytical solution'])
plt.show()
import math
import numpy as np
from scipy.integrate import odeint
import matplotlib.pyplot as plt
import scipy.integrate as integrate
from scipy.integrate import quad
LEFT = 0
RIGHT = 40
DOWN = -500
UP = 3000
sigma = 41.5
E_total = 2403.5
INTERVAL = 0.1
t=np.arange(LEFT,RIGHT,INTERVAL)
def integrand(t):
return (math.sqrt(A)*np.tanh( (t*math.sqrt(A*k_air))/m ))/(math.sqrt(k_air))
def with_bonds(a,b):
inte=quad(integrand,a,b)
return inte[0]
integral_result = [0 for i in range(len(t))]
for i in range(len(t)):
integral_result[i] = with_bonds(0,t[i])
energy = [0 for i in range(len(t))]
for i in range(len(t)):
energy[i] = sigma*t[i] - epsilon*F_M*integral_result[i]
energy_final = [energy[i]+E_total for i in range(len(t))]
# for the figure itself
fig = plt.figure(figsize=(5,5), dpi=80)
def find_first_zero(theta):
for i in range(len(t)-1):
if(energy_final[i]*energy_final[i+1]<=0):
return round(i*INTERVAL,2)
else:
continue
fstzero = find_first_zero(theta)
plt.plot(t,energy_final)
plt.plot(fstzero,0,'ro')
plt.annotate('('+str(fstzero)+',0)',xy = (fstzero,0),xytext = (fstzero+INTERVAL,100))
plt.xlabel('time')
plt.ylabel('energy')
plt.legend(['the remaining energy','transition point'])
plt.xticks(np.arange(LEFT+5,RIGHT,5))
plt.yticks(np.arange(DOWN,UP,400))
ax = plt.gca()
ax.spines['right'].set_color('none')
ax.spines['top'].set_color('none')
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
ax.spines['bottom'].set_position(('data', 0))
ax.spines['left'].set_position(('data', 0))
plt.show()
print(energy_final[195])
print(find_first_zero(theta))
import math
import numpy as np
from scipy.integrate import odeint
import matplotlib.pyplot as plt
m = 1 # kg
k_air=0.00324844 # kg/m
F_M = 12.2 # N/kg
g = 9.8 # m/s^2
miu = 0.6
r_wheel = 0.75 # m
theta = 4
A=F_M-(m*g*miu)/(r_wheel)-m*g*(math.sin(math.pi*theta/180.00))
# function that returns dy/dt
def velocity(v,t):
dvdt = ( F_M-k_air*(v**2)-(m*g*miu)/(r_wheel)-m*g*(math.sin(math.pi*theta/180.00)) )/m
return dvdt
# initial condition
v0 = 0
# time points
t = np.linspace(0,50)
# solve ODE
v = odeint(velocity,v0,t)
analytical = [0 for i in range(len(t))]
for i in range(len(t)):
analytical[i] = (math.sqrt(A)*np.tanh( (t[i]*math.sqrt(A*k_air))/m ))/(math.sqrt(k_air))
l = [analytical[len(t)-1] for i in range (len(t))]
fig = plt.figure(figsize=(5,5), dpi=80)
print(with_bonds(0,19.4))
# plot results
plt.plot(t,v)
plt.plot(t,l,':')
plt.plot(t,analytical)
plt.xlabel('time')
plt.ylabel('velocity')
plt.legend(['y=v(t)','maximum possible velocity with human force\n'+str(round(analytical[len(t)-1],2))+'m/s','analytical solution'])
plt.show()
import math
import numpy as np
from scipy.integrate import odeint
import matplotlib.pyplot as plt
m = 1 # kg
k_air=0.00324844 # kg/m
F_M = 12.2 # N/kg
g = 9.8 # m/s^2
miu = 0.6
r_wheel = 0.75 # m
# theta = 15
INF = 999999
def get_A(theta):
return (F_M-(m*g*miu)/(r_wheel)-m*g*(math.sin(math.pi*theta/180.00)))
t = np.linspace(0,50)
theta=[0,4,8]
upper=[0,0,0]
A=[0,0,0]
analytical=[[0 for i in range(len(t))] for j in range(3)]
for i in range(3):
A[i]=get_A(theta[i])
for j in range(len(t)):
analytical[i][j] = (math.sqrt(A[i])*np.tanh( (t[j]*math.sqrt(A[i]*k_air))/m ))/(math.sqrt(k_air))
upper[i] = (math.sqrt(A[i])*np.tanh( (INF*math.sqrt(A[i]*k_air))/m ))/(math.sqrt(k_air))
newupper=[[0 for j in range(len(t))] for i in range(3)]
for i in range(3):
newupper[i]=[upper[i] for j in range(len(t))]
colorset=['purple','darkblue','blue']
fig = plt.figure(figsize=(5,5), dpi=80)
for i in range(3):
plt.plot(t,analytical[i],c=colorset[i])
plt.plot(t,newupper[i],':',c=colorset[i])
plt.xlabel('time')
plt.ylabel('velocity')
plt.legend(['theta=0 degree','v_max='+str(round(upper[0],2))+'m/s',
'theta=4 degrees','v_max='+str(round(upper[1],2))+'m/s',
'theta=8 degrees','v_max='+str(round(upper[2],2))+'m/s'])
plt.show()
import numpy as np
from scipy.integrate import odeint
from scipy.interpolate import UnivariateSpline
import matplotlib.pyplot as plt
def bicycle(v,t,m,k,A,E):
( m*dfdt*f* + (1/2)*k*(f**3) )/( m*(ddfdtdt)*f + (dfdt**2) + (3/2)*k*(f**2) ) = A*x+E
import matplotlib.pyplot as plt
from scipy.interpolate import UnivariateSpline
import numpy as np
x = np.array([ 120. , 121.5, 122. , 122.5, 123. , 123.5, 124. , 124.5,
125. , 125.5, 126. , 126.5, 127. , 127.5, 128. , 128.5,
129. , 129.5, 130. , 130.5, 131. , 131.5, 132. , 132.5,
133. , 133.5, 134. , 134.5, 135. , 135.5, 136. , 136.5,
137. , 137.5, 138. , 138.5, 139. , 139.5, 140. , 140.5,
141. , 141.5, 142. , 142.5, 143. , 143.5, 144. , 144.5,
145. , 145.5, 146. , 146.5, 147. ])
y = np.array([ 1.25750000e+01, 1.10750000e+01, 1.05750000e+01,
1.00750000e+01, 9.57500000e+00, 9.07500000e+00,
8.57500000e+00, 8.07500000e+00, 7.57500000e+00,
7.07500000e+00, 6.57500000e+00, 6.07500000e+00,
5.57500000e+00, 5.07500000e+00, 4.57500000e+00,
4.07500000e+00, 3.57500000e+00, 3.07500000e+00,
2.60500000e+00, 2.14500000e+00, 1.71000000e+00,
1.30500000e+00, 9.55000000e-01, 6.65000000e-01,
4.35000000e-01, 2.70000000e-01, 1.55000000e-01,
9.00000000e-02, 5.00000000e-02, 2.50000000e-02,
1.50000000e-02, 1.00000000e-02, 1.00000000e-02,
1.00000000e-02, 1.00000000e-02, 1.00000000e-02,
1.00000000e-02, 1.00000000e-02, 5.00000000e-03,
5.00000000e-03, 5.00000000e-03, 5.00000000e-03,
5.00000000e-03, 5.00000000e-03, 5.00000000e-03,
5.00000000e-03, 5.00000000e-03, 5.00000000e-03,
5.00000000e-03, 5.00000000e-03, 5.00000000e-03,
5.00000000e-03, 5.00000000e-03])
y_spl = UnivariateSpline(x,y,s=0,k=4)
plt.semilogy(x,y,'ro',label = 'data')
x_range = np.linspace(x[0],x[-1],1000)
plt.semilogy(x_range,y_spl(x_range))
y_spl_2d = y_spl.derivative(n=2)
plt.plot(x_range,y_spl_2d(x_range))
from PIL import Image
import numpy as np
im=Image.open(r"H:\stOOrz-Mathematical-Modelling-Group\MCM-ICM_2022\Images\uci-2.png")
maxns=[0 for i in range(10000)]
for x in range(im.size[0]):
tot=0
# print(x)
for y in range(im.size[1]):
pix=im.getpixel((x,y))
if(pix[3]==255):
tot+=1
else:
maxns[x]=max(maxns[x],tot)
tot=0
pre=pix
maxns[x]=max(maxns[x],tot)
# print(maxns[sz-50:sz])
import matplotlib.pyplot as plt
datax = datay = [0 for i in range(im.size[0])]
datax=range(im.size[0])
for i in range(im.size[0]):
datay[i]=maxns[datax[i]]
sz = im.size[0]
new_data = [0 for i in range(sz)]
stretch = sz/43.3
newdatax = [(i-11)/stretch for i in range(sz)]
newdatay = [datay[i]*52.5/83 for i in range(sz)]
fig=plt.figure(figsize=(10,2),dpi=80)
plt.plot(newdatax[12:sz-9],newdatay[12:sz-9])
plt.ylim(0,70)
plt.xlim(0,43.3)
plt.xlabel('distance from the start(km)')
plt.ylabel('elevation(m)')
plt.show()
1586/43.3
95
math.atan(1000/18000)/math.pi*180
from PIL import Image
im=Image.open(r"H:\stOOrz-Mathematical-Modelling-Group\MCM-ICM_2022\Images\tokyo-course-adjusted2.png")
maxns=[0 for i in range(10000)]
for x in range(im.size[0]):
tot=0
# print(x)
for y in range(im.size[1]):
pix=im.getpixel((x,y))
if pix!=(0, 0, 0, 255):
# print(pix)
tot=tot+1
maxns[x]=tot
import matplotlib.pyplot as plt
datax = datay = [0 for i in range(im.size[0])]
datax=range(im.size[0])
for i in range(im.size[0]):
datay[i]=maxns[datax[i]]
sz = im.size[0]
new_data = [0 for i in range(sz)]
stretch = sz/234
newdatax = [(i-49)/stretch for i in range(sz)]
newdatay = [datay[i]*1400/200 for i in range(sz)]
fig=plt.figure(figsize=(10,2),dpi=800)
plt.plot(newdatax,newdatay)
plt.ylim(0,1500)
plt.xlim(0,234)
plt.xlabel('distance from the start(km)')
plt.ylabel('elevation(m)')
plt.show()
len('0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, ')
# the relationship between
import math
import numpy as np
from scipy.integrate import odeint
import matplotlib.pyplot as plt
m = 1 # kg
k_air=0.00324844 # kg/m
F_M = 12.2 # N/kg
g = 9.8 # m/s^2
miu = 0.6
r_wheel = 0.75 # m
theta = 4
def max_first_phase_distance(theta):
A=F_M-(m*g*miu)/(r_wheel)-m*g*(math.sin(math.pi*theta/180.00))
# function that returns dy/dt
def velocity(v,t):
dvdt = ( F_M-k_air*(v**2)-(m*g*miu)/(r_wheel)-m*g*(math.sin(math.pi*theta/180.00)) )/m
return dvdt
# initial condition
v0 = 0
# time points
t = np.linspace(0,50)
# solve ODE
v = odeint(velocity,v0,t)
analytical = [0 for i in range(len(t))]
for i in range(len(t)):
analytical[i] = (math.sqrt(A)*np.tanh( (t[i]*math.sqrt(A*k_air))/m ))/(math.sqrt(k_air))
l = [analytical[len(t)-1] for i in range (len(t))]
return with_bonds(0,find_first_zero(theta))
print(max_first_phase_distance(4))
dtx = np.arange(-8,8,0.01)
# dty = [max_first_phase_distance(dtx[i]) for i in range(len(dtx))]
# plt.plot(dtx,dty)
# plt.show()
# for i in range(len(dtx)):
# print(max_first_phase_distance(dtx[i]))
```
| github_jupyter |
<a href="https://githubtocolab.com/giswqs/geemap/blob/master/examples/notebooks/65_vector_styling.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab"/></a>
Uncomment the following line to install [geemap](https://geemap.org) if needed.
```
# !pip install geemap
```
**Styling Earth Engine vector data**
```
import ee
import geemap
# geemap.update_package()
```
## Use the default style
```
Map = geemap.Map()
states = ee.FeatureCollection("TIGER/2018/States")
Map.addLayer(states, {}, "US States")
Map
```
## Use Image.paint()
```
Map = geemap.Map()
states = ee.FeatureCollection("TIGER/2018/States")
image = ee.Image().paint(states, 0, 3)
Map.addLayer(image, {'palette': 'red'}, "US States")
Map
```
## Use FeatureCollection.style()
```
Map = geemap.Map()
states = ee.FeatureCollection("TIGER/2018/States")
style = {'color': '0000ffff', 'width': 2, 'lineType': 'solid', 'fillColor': '00000080'}
Map.addLayer(states.style(**style), {}, "US States")
Map
```
## Use add_styled_vector()
```
Map = geemap.Map()
states = ee.FeatureCollection("TIGER/2018/States")
vis_params = {
'color': '000000',
'colorOpacity': 1,
'pointSize': 3,
'pointShape': 'circle',
'width': 2,
'lineType': 'solid',
'fillColorOpacity': 0.66,
}
palette = ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']
Map.add_styled_vector(
states, column="NAME", palette=palette, layer_name="Styled vector", **vis_params
)
Map
import geemap.colormaps as cm
Map = geemap.Map()
states = ee.FeatureCollection("TIGER/2018/States")
vis_params = {
'color': '000000',
'colorOpacity': 1,
'pointSize': 3,
'pointShape': 'circle',
'width': 2,
'lineType': 'solid',
'fillColorOpacity': 0.66,
}
palette = list(cm.palettes.gist_earth.n12)
Map.add_styled_vector(
states, column="NAME", palette=palette, layer_name="Styled vector", **vis_params
)
Map
Map = geemap.Map()
states = ee.FeatureCollection("TIGER/2018/States").filter(
ee.Filter.inList('NAME', ['California', 'Nevada', 'Utah', 'Arizona'])
)
palette = {
'California': 'ff0000',
'Nevada': '00ff00',
'Utah': '0000ff',
'Arizona': 'ffff00',
}
vis_params = {
'color': '000000',
'colorOpacity': 1,
'width': 2,
'lineType': 'solid',
'fillColorOpacity': 0.66,
}
Map.add_styled_vector(
states, column="NAME", palette=palette, layer_name="Styled vector", **vis_params
)
Map
```
## Use interactive GUI
```
Map = geemap.Map()
states = ee.FeatureCollection("TIGER/2018/States")
Map.addLayer(states, {}, "US States")
Map
```
| github_jupyter |
<a href="https://colab.research.google.com/github/hoops92/DS-Unit-2-Kaggle-Challenge/blob/master/module4-classification-metrics/Scott_LS_DS10_224_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
Lambda School Data Science
*Unit 2, Sprint 2, Module 4*
---
# Classification Metrics
## Assignment
- [ ] If you haven't yet, [review requirements for your portfolio project](https://lambdaschool.github.io/ds/unit2), then submit your dataset.
- [ ] Plot a confusion matrix for your Tanzania Waterpumps model.
- [ ] Continue to participate in our Kaggle challenge. Every student should have made at least one submission that scores at least 70% accuracy (well above the majority class baseline).
- [ ] Submit your final predictions to our Kaggle competition. Optionally, go to **My Submissions**, and _"you may select up to 1 submission to be used to count towards your final leaderboard score."_
- [ ] Commit your notebook to your fork of the GitHub repo.
- [ ] Read [Maximizing Scarce Maintenance Resources with Data: Applying predictive modeling, precision at k, and clustering to optimize impact](https://towardsdatascience.com/maximizing-scarce-maintenance-resources-with-data-8f3491133050), by Lambda DS3 student Michael Brady. His blog post extends the Tanzania Waterpumps scenario, far beyond what's in the lecture notebook.
## Stretch Goals
### Reading
- [Attacking discrimination with smarter machine learning](https://research.google.com/bigpicture/attacking-discrimination-in-ml/), by Google Research, with interactive visualizations. _"A threshold classifier essentially makes a yes/no decision, putting things in one category or another. We look at how these classifiers work, ways they can potentially be unfair, and how you might turn an unfair classifier into a fairer one. As an illustrative example, we focus on loan granting scenarios where a bank may grant or deny a loan based on a single, automatically computed number such as a credit score."_
- [Notebook about how to calculate expected value from a confusion matrix by treating it as a cost-benefit matrix](https://github.com/podopie/DAT18NYC/blob/master/classes/13-expected_value_cost_benefit_analysis.ipynb)
- [Simple guide to confusion matrix terminology](https://www.dataschool.io/simple-guide-to-confusion-matrix-terminology/) by Kevin Markham, with video
- [Visualizing Machine Learning Thresholds to Make Better Business Decisions](https://blog.insightdatascience.com/visualizing-machine-learning-thresholds-to-make-better-business-decisions-4ab07f823415)
### Doing
- [ ] Share visualizations in our Slack channel!
- [ ] RandomizedSearchCV / GridSearchCV, for model selection. (See module 3 assignment notebook)
- [ ] More Categorical Encoding. (See module 2 assignment notebook)
- [ ] Stacking Ensemble. (See below)
### Stacking Ensemble
Here's some code you can use to "stack" multiple submissions, which is another form of ensembling:
```python
import pandas as pd
# Filenames of your submissions you want to ensemble
files = ['submission-01.csv', 'submission-02.csv', 'submission-03.csv']
target = 'status_group'
submissions = (pd.read_csv(file)[[target]] for file in files)
ensemble = pd.concat(submissions, axis='columns')
majority_vote = ensemble.mode(axis='columns')[0]
sample_submission = pd.read_csv('sample_submission.csv')
submission = sample_submission.copy()
submission[target] = majority_vote
submission.to_csv('my-ultimate-ensemble-submission.csv', index=False)
```
```
%%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Kaggle-Challenge/master/data/'
!pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
import pandas as pd
# Merge train_features.csv & train_labels.csv
train = pd.merge(pd.read_csv(DATA_PATH+'waterpumps/train_features.csv'),
pd.read_csv(DATA_PATH+'waterpumps/train_labels.csv'))
# Read test_features.csv & sample_submission.csv
test = pd.read_csv(DATA_PATH+'waterpumps/test_features.csv')
sample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')
```
## Define a function to wrangle train, validate and test sets in the same way. Clean outliers and engineer features
```
%matplotlib inline
import category_encoders as ce
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from sklearn.impute import SimpleImputer
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
from sklearn.ensemble import RandomForestClassifier
def wrangle(X):
"""Wrangle train, validate, and test sets in the same way"""
# Prevent SettingWithCopyWarning
X = X.copy()
# About 3% of the time, latitude has small values near zero,
# outside Tanzania, so we'll treat these values like zero.
X['latitude'] = X['latitude'].replace(-2e-08, 0)
# When columns have zeros and shouldn't, they are like null values.
# So we will replace the zeros with nulls, and impute missing values later.
cols_with_zeros = ['longitude', 'latitude', 'construction_year',
'gps_height','population','amount_tsh']
for col in cols_with_zeros:
X[col] = X[col].replace(0, np.nan)
X[col+'_MISSING'] = X[col].isnull()
# Approximate distance from 'Null Island'
X['distance'] = ((X['latitude']+10.99846435)**2 + (X['longitude']-19.6071219)**2)**.5
# Convert to datetime and create year_ month_ & day_recorded
X['date_recorded'] = pd.to_datetime(X['date_recorded'], infer_datetime_format=True)
X['year_recorded'] = X['date_recorded'].dt.year
X['month_recorded'] = X['date_recorded'].dt.month
X['day_recorded'] = X['date_recorded'].dt.day
X = X.drop(columns='date_recorded')
# Engineer feature: how many years from construction_year to date_recorded
X['years'] = X['year_recorded'] - X['construction_year']
X['years_MISSING'] = X['years'].isnull()
# region_code & district_code are numeric columns, but should be categorical features,
# so convert it from a number to a string
X['region_code'] = X['region_code'].astype(str)
X['district_code'] = X['district_code'].astype(str)
# quantity & quantity_group are duplicates, so drop one
X = X.drop(columns='quantity_group')
# source, source_class & source_type are almost identical.
# source has higher level of detail.
X = X.drop(columns=['source_class','source_type'])
# recorded_by has single value, so drop.
X = X.drop(columns='recorded_by')
X = X.drop(columns='id')
# water_quality & quality_group are almost identical.
# water_quality has higher level of detail.
X = X.drop(columns='quality_group')
# waterpoint_type & waterpoint_type_group are almost identical.
# waterpoint_type has higher level of detail.
X = X.drop(columns='waterpoint_type_group')
# payment & payment_type are duplicates, so drop one
X = X.drop(columns='payment_type')
# extraction_type, extraction_type_class & extraction_type_group are almost identical.
# extraction_type has higher level of detail.
X = X.drop(columns=['extraction_type_class','extraction_type_group'])
# installer & funder are almost identical.
# funder has higher level of detail.
X = X.drop(columns='installer')
# management & management_group are almost identical.
# management has higher level of detail.
X = X.drop(columns='management_group')
# region_code & region are almost identical.
# region_code has higher level of detail.
X = X.drop(columns='region')
# return the wrangled dataframe
return X
# Merge train_features.csv & train_labels.csv
train = pd.merge(pd.read_csv(DATA_PATH+'waterpumps/train_features.csv'),
pd.read_csv(DATA_PATH+'waterpumps/train_labels.csv'))
# Read test_features.csv & sample_submission.csv
test = pd.read_csv(DATA_PATH+'waterpumps/test_features.csv')
sample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')
# Split train into train & val. Make val the same size as test.
target = 'status_group'
train, val = train_test_split(train, train_size=.8, test_size=.2,
stratify=train[target], random_state=42)
# Wrangle train, validate, and test sets in the same way
train = wrangle(train)
val = wrangle(val)
test = wrangle(test)
# Arrange data into X features matrix and y target vector
X_train = train.drop(columns=target)
y_train = train[target]
X_val = val.drop(columns=target)
y_val = val[target]
X_test = test
# Make pipeline!
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='mean'),
RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)
)
# Fit on train, score on val
pipeline.fit(X_train, y_train)
y_pred = pipeline.predict(X_val)
print('Validation Accuracy', accuracy_score(y_val, y_pred))
```
## Plot a confusion matrix for your Tanzania Waterpumps model
```
from sklearn.metrics import confusion_matrix
confusion_matrix(y_val, y_pred)
# We need to get labels
from sklearn.utils.multiclass import unique_labels
unique_labels(y_val)
import seaborn as sns
def plot_confusion_matrix(y_true, y_pred):
labels = unique_labels(y_true)
columns = [f'Predicted {label}' for label in labels]
index = [f'Actual {label}' for label in labels]
table = pd.DataFrame(confusion_matrix(y_true, y_pred),
columns=columns, index=index)
return sns.heatmap(table, annot=True, fmt='d', cmap='viridis')
plot_confusion_matrix(y_val, y_pred);
```
| github_jupyter |
<a href="https://colab.research.google.com/github/mscouse/TBS_investment_management/blob/main/PM_labs_part_4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
[](https://colab.research.google.com/drive/1F1J2rObxMwR11cnRzm5m_b1cLwW1U-nL?usp=sharing)
# <strong> Investment Management 1</strong>
---
#<strong> Part 4: Data sources & data collection in Python.</strong>
In the course repository on GitHub, you will find several introductory Colab notebooks covering the following topics:
**Part 1: Introduction to Python and Google Colab notebooks.**
**Part 2: Getting started with Colab notebooks & basic features.**
**Part 3: Data visualisation libraries.**
**Part 4: Data sources & data collection in Python (CURRENT NOTEBOOK).**
**Part 5: Basic financial calculations in python.**
The notebooks have been designed to help you get started with Python and Google Colab. See the **“1_labs_introduction”** folder for more information. Each notebook contains all necessary libraries and references to the required subsets of data.
# <strong>Data sources and data collection</strong>
To perform data analysis, the first step is to load a file containing the pertinent data – such as a CSV or Excel file - into Colab. There are several ways to do so. You can import your own data into Colab notebooks from Google Drive, GitHub and many other sources. Some of these are discussed below.
To find out more about importing data, and how Colab can be used for data analysis, see the <a href="https://github.com/mscouse/TBS_investment_management/blob/main/Python_workspace.pdf">Python Workspace</a> document in the course GitHub repository or a more <a href="https://neptune.ai/blog/google-colab-dealing-with-files">comprehensive guide</a> prepared by Siddhant Sadangi of Reuters.
##1. Uploading files from your local drive
It is easy to upload your locally stored data files. To upload the data from your local drive, type in the following code in a new “Code” cell in Colab (as demonstrated below):
```
from google.colab import files
files.upload()
```
Once executed, the code will prompt you to select a file containing your data. Click on **“Choose Files”** then select and upload the file. Wait for the file to be 100% uploaded. You should see the name of the file in the code cell once it is uploaded.
On the left side of Colab interface, there is a **"Files/ Folder"** tab. You can find the uploaded file in that directory.
If you want to read the uploaded data into a Pandas dataframe (named `df` in this example), use the following code in a new code cell. The **'filename.csv'** should match the name of the uploaded file, including the `.csv` extension:
```
import pandas as pd
df = pd.read_csv('filename.csv')
```
```
from google.colab import files
files.upload()
```
##2. Upload files from GitHub (via its RAW URL)
You can either clone an entire GitHub repository to your Colab environment or access individual files from their raw link. We use the latter method throughout the course/assignments.
**Clone a GitHub repository**
You can clone a GitHub repository into your Colab environment in the same way as you would on your local machine, using `!git clone` followed by the clone URL of the repository:
```
# use the correct URL
!git clone https://github.com/repository_name.git
```
Once the repository is cloned, refresh the file-explorer to browse through its contents. Then you can simply read the files as you would in your local machine (see above).
**Load GitHub files using raw links**
There is no neeed to clone the repository to Colab if you need to work with only a few files from that repository. You can load individual files directly from GitHub using thier raw links, as follows:
1. click on the file in the repository;
2. click on `View Raw`;
3. copy the URL of the raw file,
4. use this URL as the location of your file (see sample code below)
```
import pandas as pd
# step 1: store the link to your dataset as a string titled "url"
url="https://raw.githubusercontent.com/mscouse/TBS_investment_management/main/1_labs_introduction/stock_prices_1.csv"
# step 2: Load the dataset into pandas. The dataset is stored as a pandas dataframe "df".
df = pd.read_csv(url)
```
Try doing it yourself using the code cells below.
```
# import any required libraries
import pandas as pd
# store the URL link to your GitHub dataset as a string titled "url"
url = 'https://raw.githubusercontent.com/mscouse/TBS_investment_management/main/1_labs_introduction/stock_prices_1.csv'
# load the dataset into Pandas. The dataset will be stored as a Pandas Dataframe "df".
# Note that the file we deal with in this example contains dates in the first column.
# Therefore, we parse the dates using "parse_dates" and set the date column to be
# the index of the dataframe (using the "index_col" parameter)
df = pd.read_csv(url, parse_dates=['date'], index_col=['date'])
df.head()
```
##3. Accessing financial data
There are several open source Python library designed to help researchers access financial data. One example is `yfinance` (formerly known as `fix-yahoo-finance`). It is a popular library, developed as a means to access the financial data available on Yahoo Finance.
Other widely used libraries are `pandas_datareader`, `yahoo_fin`, `ffn`, `PyNance`, and `alpha vantage`.
In this section we focus on the former library, `yfinance`. As this library is not pre-installed in Google Colab by default, we will first execute the following code to install it:
```
!pip install yfinance
```
The `!pip install <package>` command looks for the latest version of the package and installs it. This only needs to be done once per session.
```
# install the yfinance library
!pip install yfinance
```
As you may know, **Yahoo Finance** offers historical market data on stocks, bonds, cryptocurrencies, and currencies. It also aggregates companies' fundamental data.
We will be using several modules and functions included with the `yfinance` library to download historical market data from Yahoo Finance. For more information on the library, see <a href="https://pypi.org/project/yfinance/">here</a>.
**Company information**
The first `module` of the `yfinance` library we consider is `Ticker`. By using the `Ticker` function we pass the stock symbol for which we need to download the data. It allows us to access ticker-specific data, such as stock info, corporate actions, company financials, etc. In the example below we are working with Apple - its ticker is”AAPL”. The first step is to call the `Ticker` function to initialize the stock we work with.
```
# import required libraries (note that yfinance needs to be imported in addition to being installed)
import yfinance as yf
# assign ticker to Python variable
aapl = yf.Ticker("AAPL")
# get stock info
aapl.info
```
**Downloading stock data**
To download the historical stock data, we need to use the `history` function. As arguments, we can pass **start** and **end** dates to set a specific time period. Otherwise, we can set the period to **max** which will return all the stock data available on Yahoo for the chosen ticker.
Available paramaters for the `history()` method are:
* period: data period to download (either use `period` parameter or use `start` and `end`). Valid periods are: 1d, 5d, 1mo, 3mo, 6mo, 1y, 2y, 5y, 10y, ytd, max;
* interval: data interval (intraday data cannot extend past 60 days). Valid intervals are: 1m, 2m, 5m, 15m, 30m, 60m, 90m, 1h, 1d, 5d, 1wk, 1mo, 3mo;
* start: if not using `period` - download start date string (YYYY-MM-DD) or datetime;
* end: if not using `period` - download end date string (YYYY-MM-DD) or datetime;
```
# get historical market data
hist = aapl.history(period="max")
hist.head()
```
**Displaying corporate actions and analysts recommendations**
To display information about the dividends and stock splits, or the analysts recommendations use the `actions` and `recommendations` functions.
```
# show company corporate actions, such as dividends and stock splits
aapl.actions
# show analysts recommendations
aapl.recommendations
```
**Data for multiple stocks**
To download data for multiple tickers, we need to use the `download()` method, as follows:
```
# Version 1
import yfinance as yf
stock_data = yf.download("AAPL MSFT BRK-A", start="2015-01-01", end="2021-01-20")
```
Alternatively, we can rewrite the code above as:
```
# Version 2
import yfinance as yf
tickers = "AAPL MSFT BRK-A"
date_1 = "2015-01-01"
date_2 = "2021-01-20"
stock_data = yf.download(tickers, start=date_1, end=date_2)
```
To access the closing adjusted price data for the tickers in the `stock_data` dataframe the code above creates, you should use: `stock_data['Adj Close']`. To access the closing adjusted price data for 'AAPL' only, use: `stock_data['Adj Close']['AAPL']`.
```
# Version 1
# import required libraries
import yfinance as yf
# fetch data for multiple tickers
stock_data = yf.download("AAPL MSFT BRK-A", start="2015-01-01", end="2021-01-20")
# display the last 5 rows of the dataframe; we choose to display the "Adj Close" column only
stock_data["Adj Close"].tail()
# Version 2
import yfinance as yf
# assign required values to variables
tickers = "AAPL MSFT BRK-A"
date_1 = "2015-01-01"
date_2 = "2021-01-20"
# fetch data for multiple tickers
stock_data = yf.download(tickers, start=date_1, end=date_2)
# display the last 5 rows of the dataframe; we choose to display the "Adj Close" column only
stock_data["Adj Close"].tail()
# display the last 5 rows of AAPL adjusted close prices
stock_data['Adj Close']['AAPL'].tail()
```
However, if you want to group stock data by ticker, use the following code:
```
# Version 3
import yfinance as yf
tickers = "AAPL MSFT BRK-A"
date_1 = "2015-01-01"
date_2 = "2021-01-20"
stock_data = yf.download(tickers, start=date_1, end=date_2, group_by="ticker")
```
To access the closing adjusted price data for 'AAPL' only, use: `stock_data['AAPL']['Adj Close']`.
```
# Version 3
import yfinance as yf
# assign required values to variables
tickers = "AAPL MSFT BRK-A"
date_1 = "2015-01-01"
date_2 = "2021-01-20"
# fetch data for multiple tickers
stock_data = yf.download(tickers, start=date_1, end=date_2, group_by="ticker")
# display the last 5 rows of "Adj Close" prices for AAPL only
stock_data["AAPL"]["Adj Close"].tail()
```
| github_jupyter |
# Crisis Sentiment Analysis
This activity is a mini-project where students will create a data visualization dashboard, they have to analyze sentiment and tone about the news related to the financial crisis of 2008 that where published along the last month. Students will retrieve the news articles from the News API; by default, the developer account gives access to news articles up to a month old.
In this activity, students will use their new sentiment analysis skills, in combination to some of the skills they already master such as: Pandas, Pyviz, Plotly Express and PyViz Panel.
This Jupyter notebook is a sandbox where students will conduct the sentiment analysis tasks and charts creation before assembling the dashboard.
```
# Initial imports
import os
from path import Path
import pandas as pd
import numpy as np
import hvplot.pandas
import nltk
from wordcloud import WordCloud
from nltk.sentiment.vader import SentimentIntensityAnalyzer
from newsapi import NewsApiClient
from ibm_watson import ToneAnalyzerV3
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
import plotly.express as px
import matplotlib.pyplot as plt
import matplotlib as mpl
import panel as pn
plt.style.use("seaborn-whitegrid")
pn.extension("plotly")
```
## Instructions
### Fetching the Latests News Metions About the Crisis of 2008
Using the News API, get all the news in English about the financial crisis of 2008 using the keywords `"financial AND crisis AND 2008"` in the `q` parameter. Define a `page_size=100` to have at least 100 news articles to analyze.
```
# Retrieve the News API key
news_api = os.getenv("news_api")
# Create the newsapi client
# Fetch the news articles about the financial crisis on 2008 in English
# Show the total number of news
```
### Creating a VADER Sentiment Scoring Function
Use the VADER sentiment scoring function from `NLTK` to score the sentiment polarity of the 100 news you fetched. Just for convenience, start downloading the `vader_lexicon` in order to initialize the VADER sentiment analyzer.
```
# Download/Update the VADER Lexicon
nltk.download("vader_lexicon")
# Initialize the VADER sentiment analyzer
```
In order to score the VADER sentiment, create a function named `get_sentiment_scores(text, date, source, url)` that will receive four parameters.
* `text` is the text whose sentiment will be scored.
* `date` the date the news article was published using the format `YYYY-MM-DD`.
* `source` is the name of the news article's source.
* `url` is the URL that points to the article.
The `get_sentiment_score()` function should return a Python dictionary with the scoring results. This dictionary is going to be used in the next section to create a DataFrame; the structure of the dictionary is the following:
* `date` the date passed as parameter to the function.
* `text` the text passed a parameter to the function.
* `source` the source passed as parameter to the function.
* `url` the URL passed as parameter to the function.
* `compound` the compound score from the VADER sentiment analyzer.
* `pos` the positive score from the VADER sentiment analyzer.
* `neu` the neutral score from the VADER sentiment analyzer.
* `neg` the negative score from the VADER sentiment analyzer.
* `normalized` the normalized scored based on the `compound` results. Its value should be `1` for positive sentiment, `-1` for negative sentiment, and `0` for neutral sentiment.
This is an example of the function's return value:
```python
{'date': '2019-06-24',
'text': '\nMore than a decade since the global economic meltdown of 2008
devastated lives across the world, no one who caused the crisis has
been held responsible.\n\n"The 2008 financial crisis displayed what
the world now identifies as financial contagion," says Philip J Baker,
the former managing partner of a US-based \nhedge fund that collapsed
during the financial crisis.\n\nDespite this, "zero Wall Street chief
executives have been to prison, even though there is today absolutely
no doubt that Wall Street executives and politicians \nwere complicit
in creating the crisis," he says. \n\nBaker was among the few
relatively smaller players imprisoned for the part they played.\n\n
In July 2009, he was arrested in Germany and extradited to the
United States where he faced federal court on charges of fraud and
financial crimes.\n\nHe pled guilty and was sentenced to 20 years
in prison for costing some 900 investors about $294mn worldwide.
He served eight years in jail and is now on \nparole and advocates
against financial crime.\n',
'source': 'aljazeera',
'url': 'https://www.aljazeera.com/programmes/specialseries/2019/06/men-stole-world-2008-financial-crisis-190611124411311.html',
'compound': -0.9911,
'pos': 0.048,
'neu': 0.699,
'neg': 0.254,
'normalized': -1}
```
```
# Define a function to get the sentiment scores
def get_sentiment_scores(text, date, source, url):
sentiment_scores = {}
return sentiment_scores
```
### Creating the News Articles' Sentiments DataFrame
In this section you have to create a DataFrame that is going to be used to plot the sentiment analysis results. Using a `for-loop`, iterate across all the news articles you fetched to create the DataFrame structure; define an empty list to append the sentiment scoring results for each news article and create the DataFrame using the list as data source.
Once you create the DataFrame do the following:
* Sort the DataFrame rows by the `date` column.
* Define the `date` column as the DataFrame index.
* Save the DataFrame as a CSV file in order to use it on the sentiment analysis dashboard creation.
```
# Empty list to store the DataFrame structure
sentiments_data = []
# Loop through all the news articles
for article in crisis_news_en["articles"]:
try:
# Get sentiment scoring using the get_sentiment_score() function
except AttributeError:
pass
# Create a DataFrame with the news articles' data and their sentiment scoring results
# Sort the DataFrame rows by date
# Define the date column as the DataFrame's index
# Save the news articles DataFrame with VADER Sentiment scoring as a CSV file
```
### Creating the Average Sentiment Chart
Use `hvPlot` to create a two lines chart that compares the average `compound` and `normalized` sentiment scores along the last month.
```
# Define the average sentiment DataFrame
# Create the two lines chart
```
### Creating the Sentiment Distribution Chart
Based on the `normalized` sentiment score, create a bar chart using `hvPlot` that shows the number of negative, neutral and positive news articles. This chart represents the overall sentiment distribution.
```
# Define the sentiment distribution DataFrame
# Create the sentiment distribution bar chart
```
### Getting the Top 10 Positive and Negative News Articles
In this section you have to create two DataFrames, one with the top 10 positive news according to the `compound` score, and other with the top 10 negative news. Refer to the [`hvplot.table()` documentation](https://hvplot.pyviz.org/user_guide/Plotting.html#Tables) to create two tables presenting the following columns of these news articles:
* Date
* Source
* Text
* URL
```
# Getting Top 10 positive news articles
# Create a table with hvplot
# Getting Top 10 negative news articles
# Create a table with hvplot
```
### Creating the Sentiment Distribution by News Article's Source
In this section, use `hvPlot` to create a bar chart that presents the distribution of negative, neutral and positive news according to the `normalized` score; the results should be grouped by `source`.
```
# Create the sentiment distribution by news articles' source DataFrame
# Create the sentiment distribution by news articles' source bar chart
```
### Creating the Word Clouds
In this section you will create two word clouds, one using the bag-of-words method and other using TF-IDF.
#### Bag-of-Words' Word Cloud
Use the `CountVectorizer` module from `sklearn` to create a word cloud with the top 20 words with the highest counting. Save the DataFrame with the top 20 words as a CSV file named `top_words_data.csv` for future use on the dashboard creation.
```
# Creating the CountVectorizer instance defining the stopwords in English to be ignored
# Getting the tokenization and occurrence counting
# Retrieve unique words list
# Get the last 100 word (just as a sample)
# Getting the bag of words as DataFrame
# Sorting words by 'Word_Count' in descending order
# Get top 20 words with the highest counting
# Save the top words DataFrame
# Create a string list of terms to generate the bag-of-words word cloud
# Create the bag-of-words word cloud
```
#### TF-IDF Wordcloud
Use the `TfidfVectorizer` module from `sklearn` to create a word cloud with the top 20 words with the highest frequency. Save the DataFrame with the top 20 words as a CSV file named `top_wors_tfidf_data.csv` for future use on the dashboard creation.
```
# Getting the TF-IDF
# Retrieve words list from corpous
# Get the last 100 word (just as a sample)
# Creating a DataFrame Representation of the TF-IDF results
# Sorting words by 'Frequency' in descending order
# Get 20 top words
# Save the top words TF-IDF DataFrame
# Create a string list of terms to generate the tf-idf word cloud
# Create the tf-idf word cloud
```
## Challenge: Radar Chart with Tone Analysis
In this challenge section, you have to use Plotly Express and IBM Watson Tone Analyzer to create a radar chart presenting the tone of all the news articles that you retrieved.
Refer to the [polar coordinates chart demo](https://plot.ly/python/plotly-express/#polar-coordinates) and the [Plotly Express reference documentation](https://www.plotly.express/plotly_express/#plotly_express.scatter_polar) to learn more about how to create this chart.
```
# Get the Tone Analyzer API Key and URL
tone_api = os.getenv("tone_api")
tone_url = os.getenv("tone_url")
# Initialize Tone Analyser Client
```
In order to create the radar chart, you need to score the tone of each article and retrieve the `document_tone`. Create a function named `get_tone(text,url)` that will receive two parameters and will get the tone score for a particular article.
* `text` the content of the article.
* `url` the URL pointing to the article.
The `get_tone()` function will use the `tone()` method from the `ToneAnalyzerV3` module to score the article's tone. Remember that for each document (or text), the `tone()` method of IBM Watson Tone Analyzer [scores one or more overall document tones](https://cloud.ibm.com/apidocs/tone-analyzer#analyze-general-tone-get), you can also get and empty result if no tone were scored; this function should return a dictionary with the first document tone's score with the following structure:
* `score` refers to the first `tone` from the `document_tone`.
* `tone_id` refers to the `tone_id` from the first `tone`.
* `tone_name` refers to the `tone_name` from the first `tone`.
* `text` the text passed as parameter.
* `url` the URL passed as parameter.
This is an example of the function's return value:
```python
{'score': 0.616581,
'tone_id': 'sadness',
'tone_name': 'Sadness',
'text': '\nMore than a decade since the global economic meltdown of 2008
devastated lives across the world, no one who caused the crisis has
been held responsible.\n\n"The 2008 financial crisis displayed what
the world now identifies as financial contagion," says Philip J Baker,
the former managing partner of a US-based \nhedge fund that collapsed
during the financial crisis.\n\nDespite this, "zero Wall Street chief
executives have been to prison, even though there is today absolutely
no doubt that Wall Street executives and politicians \nwere complicit
in creating the crisis," he says. \n\nBaker was among the few
relatively smaller players imprisoned for the part they played.\n\n
In July 2009, he was arrested in Germany and extradited to the
United States where he faced federal court on charges of fraud and
financial crimes.\n\nHe pled guilty and was sentenced to 20 years
in prison for costing some 900 investors about $294mn worldwide.
He served eight years in jail and is now on \nparole and advocates
against financial crime.\n',
'url': 'https://www.aljazeera.com/programmes/specialseries/2019/06/men-stole-world-2008-financial-crisis-190611124411311.html'}
```
```
# Create a function to analyze the text's tone with the 'tone()' method of IBM Watson Tone Analyzer.
def get_tone(text, url):
try:
except:
pass
```
Create a DataFrame with the tone scoring from all the news articles. Use an empty list to create a the DataFrame's structure and a `for-loop` to iterate across all the news to score their tone using the `get_tone()` function.
```
# Create an empty list to create the DataFrame's structure
# Iterate across all the news articles to score their tone.
print(f"Analyzing tone from {crisis_news_df.shape[0]} articles...")
for index, row in crisis_news_df.iterrows():
try:
print("*", end="")
# Get news article's tone
except:
pass
print("\nDone :-)")
# Create the DateFrame containing the news articles and their tone scoring results.
```
Save the DataFrame as a CSV file named `tone_data.csv` for further use on the dashboard creation.
```
```
Create a radar chart using the `scatter_polar()` method from Plotly Express as follows:
* Use the `score` column for the `r` and `color` parameters.
* Use the `tone_name` column for the `theta` parameter.
* Use the `url` column for the `hover_data` parameter.
* Define a `title` for the chart.
```
# Create the radar chart
```
| github_jupyter |
# Project Setup
* This section describes how to create and configure a project
* This is the same as creating a new project in the editor and going through all of the steps.
* When a user creates a project with client.create_project() the project is not ready for labeling.
* An ontology must be set
* datasets must be attached
```
!pip install labelbox
from labelbox import Client, Project, LabelingFrontend
from labelbox.schema.ontology import Tool, OntologyBuilder
from getpass import getpass
import os
# If you don't want to give google access to drive you can skip this cell
# and manually set `API_KEY` below.
COLAB = "google.colab" in str(get_ipython())
if COLAB:
!pip install colab-env -qU
from colab_env import envvar_handler
envvar_handler.envload()
API_KEY = os.environ.get("LABELBOX_API_KEY")
if not os.environ.get("LABELBOX_API_KEY"):
API_KEY = getpass("Please enter your labelbox api key")
if COLAB:
envvar_handler.add_env("LABELBOX_API_KEY", API_KEY)
# Set this to a project that is already set up
PROJECT_ID = "ckm4xyfncfgja0760vpfdxoro"
# Only update this if you have an on-prem deployment
ENDPOINT = "https://api.labelbox.com/graphql"
client = Client(api_key=API_KEY, endpoint=ENDPOINT)
```
### Identify project, dataset, and ontology
* Pick the project to setup
* Dataset(s) to attach to that project
* Configure the ontology for the project
```
# Use bounding boxes to label cats
ontology_builder = OntologyBuilder(
tools=[Tool(name="cat", tool=Tool.Type.BBOX)])
project = client.create_project(name="my_new_project")
dataset = client.create_dataset(name="my_new_dataset")
# Add data_rows since this is a new dataset (see basics/data_rows.ipynb for more information on this)
test_img_url = "https://upload.wikimedia.org/wikipedia/commons/thumb/0/08/Kitano_Street_Kobe01s5s4110.jpg/2560px-Kitano_Street_Kobe01s5s4110.jpg"
dataset.create_data_row(row_data=test_img_url)
# Unless you are using a custom editor you should always use the following editor:
editor = next(
client.get_labeling_frontends(where=LabelingFrontend.name == "Editor"))
# Note that you can use any dataset or ontology even if they already exist.
existing_project = client.get_project(PROJECT_ID)
# We are not using this, but it is possible to copy the ontology to the new project
ontology = existing_project.ontology()
```
### Setup and attach dataset
* Setting up a project will add an ontology and will enable labeling to begin
* Attaching dataset(s) will add all data_rows belonging to the dataset to the queue.
```
project.setup(editor, ontology_builder.asdict())
# Could also do if ontology is a normalized ontology
# project.setup(editor, ontology.normalized)
# Run this for each dataset we want to attach
project.datasets.connect(dataset)
# project.datasets.connect(another_dataset)
```
### Review
```
# Note setup_complete will be None if it fails.
print(project.setup_complete)
print(project.ontology)
print([ds.name for ds in project.datasets()])
print(f"https://app.labelbox.com/projects/{project.uid}")
```
| github_jupyter |
# Sequential MNIST results from the paper by Rui Costa et al.:<br/>"Cortical microcircuits as gated-recurrent neural networks"
## Implementation done in the scope of the nurture.ai NIPS 2017 paper implementation challenge
- nurture.ai challenge: https://nurture.ai/nips-challenge
- Paper: http://papers.nips.cc/paper/6631-cortical-microcircuits-as-gated-recurrent-neural-networks
- Credits:<br/>
Training logic based on the r2rt LSTM tutorial (https://r2rt.com/recurrent-neural-networks-in-tensorflow-ii.html).<br/>
Model definition based on KnHuq implementation (https://github.com/KnHuq/Dynamic-Tensorflow-Tutorial/blob/master/LSTM/LSTM.py
).
## This notebook compare the results of models with 2 layers (not done in the paper)
### Loading Librairies and Models
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
import numpy as np
import sys
#import LSTM and subLSMT cell models with 2 layers
sys.path.append('../models/')
from LSTMTwoLayers import *
from subLSTMTwoLayers import *
from parameters import *
sys.path.append('../src/common/')
import helper as hp
```
### Loading MNIST dataset
```
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
```
### Training Function
```
def train_network(g, batch_size=50, n_epoch=10, verbose=False, save=False, patience=25, min_delta=0.01):
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
# parameters for early stopping
patience_cnt = 0
max_test_accuracy = 0.0
#Iterations to do trainning
for epoch in range(n_epoch):
X, Y = mnist.train.next_batch(batch_size)
X = X.reshape(batch_size, 1, g['input_size'])
sess.run(g['train_step'],feed_dict={g['rnn']._inputs:X, g['y']:Y})
if epoch % 1000 == 0:
Loss=str(sess.run(g['cross_entropy'],feed_dict={g['rnn']._inputs:X, g['y']:Y}))
Train_accuracy=str(sess.run(g['accuracy'],feed_dict={g['rnn']._inputs:X, g['y']:Y}))
X_test = mnist.test.images.reshape(10000,1,g['input_size'])
Test_accuracy=str(sess.run(g['accuracy'],feed_dict={g['rnn']._inputs:X_test, g['y']:mnist.test.labels}))
if verbose:
print("\rIteration: %s Loss: %s Train Accuracy: %s Test Accuracy: %s"%(epoch,Loss,Train_accuracy,Test_accuracy))
# early stopping
if float(Test_accuracy) > max_test_accuracy:
max_test_accuracy = float(Test_accuracy)
patience_cnt = 0
else:
patience_cnt += 1
if patience_cnt > patience:
print("early stopping at epoch: ", epoch)
break
if isinstance(save, str):
g['saver'].save(sess, save)
return max_test_accuracy
```
### Building Graph Model Function
```
def build_graph(cell_type = None, load_parameters = False):
# define initial parameters
input_size = 784
output_size = 10
optimizer = 'Adam'
momentum = False
learning_rate = 0.001
hidden_units = 10
if load_parameters:
#load parameters from file
if cell_type == 'LSTM':
parameters = LSTM_parameters()
elif cell_type == 'sub_LSTM':
parameters = SubLSTM_parameters()
elif cell_type == 'fix_sub_LSTM':
parameters = Fix_subLSTM_parameters()
else:
print("No cell_type selected! Use LSTM cell")
parameters = LSTM_parameters()
input_size = parameters.mnist['input_size']
output_size = parameters.mnist['output_size']
optimizer = parameters.mnist['optimizer']
momentum = parameters.mnist['momentum']
learning_rate = parameters.mnist['learning_rate']
hidden_units = parameters.mnist['hidden_units']
# reset graph
if 'sess' in globals() and sess:
sess.close()
tf.reset_default_graph()
# Initializing rnn object
if cell_type == 'LSTM':
rnn = LSTM_cell(input_size, hidden_units, output_size)
elif cell_type == 'sub_LSTM':
rnn = subLSTM_cell(input_size, hidden_units, output_size)
elif cell_type == 'fix_sub_LSTM':
print("TODO!")
else:
rnn = LSTM_cell(input_size, hidden_units, output_size)
#input label placeholder
y = tf.placeholder(tf.float32, [None, output_size])
# Getting all outputs from rnn
outputs = rnn.get_outputs()
# Getting final output through indexing after reversing
last_output = outputs[-1]
# As rnn model output the final layer through Relu activation softmax is
# used for final output
output = tf.nn.softmax(last_output)
# Computing the Cross Entropy loss
cross_entropy = -tf.reduce_sum(y * tf.log(output))
# setting optimizer
if optimizer == 'Adam':
# Trainning with Adam Optimizer
train_step = tf.train.AdamOptimizer(learning_rate).minimize(cross_entropy)
elif optimizer == 'RMSProp':
# Trainning with RMSProp Optimizer
train_step = tf.train.RMSPropOptimizer(learning_rate).minimize(cross_entropy)
else:
#if nothing is define use Adam optimizer
train_step = tf.train.AdamOptimizer(learning_rate).minimize(cross_entropy)
# Calculation of correct prediction and accuracy
correct_prediction = tf.equal(tf.argmax(y, 1), tf.argmax(output, 1))
accuracy = (tf.reduce_mean(tf.cast(correct_prediction, tf.float32))) * 100
return dict(
rnn = rnn,
y = y,
input_size = input_size,
output = output,
cross_entropy = cross_entropy,
train_step = train_step,
preds = correct_prediction,
accuracy = accuracy,
saver = tf.train.Saver()
)
```
### Simulation Parameters
```
n_simulation = 5
batch_size = 500
n_epoch = 150000
```
### LSTM training
```
%%time
lstm_accuracies = []
print('Traning begins for: ', n_simulation, ' simulation(s)')
for n in range(n_simulation):
print('simulation ', n, ' running')
g = build_graph(cell_type='LSTM', load_parameters=True)
test_accuracy = train_network(g, batch_size, n_epoch, verbose=False)
lstm_accuracies.append(test_accuracy)
lstm_mean_accuracy = np.mean(lstm_accuracies)
lstm_std_accuracy = np.std(lstm_accuracies)
lstm_best_accuracy = np.amax(lstm_accuracies)
print("The mean test accuracy of the simulation is:", lstm_mean_accuracy)
print("the standard deviation is:", lstm_std_accuracy)
print("The best test accuracy obtained was:", lstm_best_accuracy)
```
### SubLSTM training
```
%%time
sub_lstm_accuracies = []
print('Traning begins for: ', n_simulation, ' simulation(s)')
for n in range(n_simulation):
print('simulation ', n, ' running')
g = build_graph(cell_type='sub_LSTM', load_parameters=True)
test_accuracy = train_network(g, batch_size, n_epoch, verbose = False)
sub_lstm_accuracies.append(test_accuracy)
sub_lstm_mean_accuracy = np.mean(sub_lstm_accuracies)
sub_lstm_std_accuracy = np.std(sub_lstm_accuracies)
sub_lstm_best_accuracy = np.amax(sub_lstm_accuracies)
print("The mean test accuracy of the simulation is:", sub_lstm_mean_accuracy)
print("the standard deviation is:", sub_lstm_std_accuracy)
print("The best test accuracy obtained was:", sub_lstm_best_accuracy)
```
### Plot test mean accuracies and std
```
objects = ('LSTM', 'SubLSTM')
mean_accuracies = [lstm_mean_accuracy, sub_lstm_mean_accuracy]
std_accuracies = [lstm_std_accuracy, sub_lstm_std_accuracy]
accuracies = [lstm_accuracies, sub_lstm_accuracies]
hp.bar_plot(objects, mean_accuracies, std_accuracies, accuracies)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/michalspiegel/freecodecamp_projects/blob/main/fcc_sms_text_classification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
*Note: You are currently reading this using Google Colaboratory which is a cloud-hosted version of Jupyter Notebook. This is a document containing both text cells for documentation and runnable code cells. If you are unfamiliar with Jupyter Notebook, watch this 3-minute introduction before starting this challenge: https://www.youtube.com/watch?v=inN8seMm7UI*
---
In this challenge, you need to create a machine learning model that will classify SMS messages as either "ham" or "spam". A "ham" message is a normal message sent by a friend. A "spam" message is an advertisement or a message sent by a company.
You should create a function called `predict_message` that takes a message string as an argument and returns a list. The first element in the list should be a number between zero and one that indicates the likeliness of "ham" (0) or "spam" (1). The second element in the list should be the word "ham" or "spam", depending on which is most likely.
For this challenge, you will use the [SMS Spam Collection dataset](http://www.dt.fee.unicamp.br/~tiago/smsspamcollection/). The dataset has already been grouped into train data and test data.
The first two cells import the libraries and data. The final cell tests your model and function. Add your code in between these cells.
```
# import libraries
try:
# %tensorflow_version only exists in Colab.
!pip install tf-nightly
except Exception:
pass
import tensorflow as tf
import pandas as pd
from tensorflow import keras
!pip install tensorflow-datasets
import tensorflow_datasets as tfds
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
# get data files
!wget https://cdn.freecodecamp.org/project-data/sms/train-data.tsv
!wget https://cdn.freecodecamp.org/project-data/sms/valid-data.tsv
train_file_path = "train-data.tsv"
test_file_path = "valid-data.tsv"
# To DataFrame
df_train = pd.read_csv(train_file_path, names=['label', 'message'], sep='\t', lineterminator='\n')
df_test = pd.read_csv(test_file_path, names=['label', 'message'], sep='\t', lineterminator='\n')
# Pop off labels
train_labels = df_train.pop('label')
test_labels = df_test.pop('label')
MAXLEN = 250
# Number of unique words
uniq_words = set()
df = df_train['message']#.append(df_test['message'])
df = df.str.lower().str.split().apply(uniq_words.update)
VOCAB_SIZE = len(uniq_words)
BUFFER_SIZE = 10000
BATCH_SIZE = 64
# Encoding binary labels
train_labels = np.where(train_labels.str.contains("spam"), 1, 0)
test_labels = np.where(test_labels.str.contains("spam"), 1, 0)
# Convert into tf Dataset
train_dataset = tf.data.Dataset.from_tensor_slices((df_train.values, train_labels))
test_dataset = tf.data.Dataset.from_tensor_slices((df_test.values, test_labels))
train_dataset = train_dataset.shuffle(BUFFER_SIZE).batch(BATCH_SIZE).prefetch(tf.data.AUTOTUNE)
test_dataset = test_dataset.batch(BATCH_SIZE).prefetch(tf.data.AUTOTUNE)
# Encoder - text to vector (text to int matrix)
encoder = tf.keras.layers.experimental.preprocessing.TextVectorization(
max_tokens=VOCAB_SIZE)
encoder.adapt(train_dataset.map(lambda text, label: text))
model = tf.keras.Sequential([
encoder,
keras.layers.Embedding(
input_dim=len(encoder.get_vocabulary()),
output_dim=64,
# Use masking to handle the variable sequence lengths
mask_zero=True),
keras.layers.Bidirectional(keras.layers.LSTM(64)),
keras.layers.Dense(64, activation='relu'),
keras.layers.Dense(1, activation='sigmoid')
])
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
history = model.fit(train_dataset, epochs=10,
validation_data=test_dataset,
validation_steps=30)
# function to predict messages based on model
# (should return list containing prediction and label, ex. [0.008318834938108921, 'ham'])
def predict_message(pred_text):
prediction = model.predict(np.array([pred_text]))[0][0]
label = "ham"
if round(prediction) == 1:
label = "spam"
return [prediction, label]
pred_text = "how are you doing today?"
prediction = predict_message(pred_text)
print(prediction)
# Run this cell to test your function and model. Do not modify contents.
def test_predictions():
test_messages = ["how are you doing today",
"sale today! to stop texts call 98912460324",
"i dont want to go. can we try it a different day? available sat",
"our new mobile video service is live. just install on your phone to start watching.",
"you have won £1000 cash! call to claim your prize.",
"i'll bring it tomorrow. don't forget the milk.",
"wow, is your arm alright. that happened to me one time too"
]
test_answers = ["ham", "spam", "ham", "spam", "spam", "ham", "ham"]
passed = True
for msg, ans in zip(test_messages, test_answers):
prediction = predict_message(msg)
if prediction[1] != ans:
passed = False
if passed:
print("You passed the challenge. Great job!")
else:
print("You haven't passed yet. Keep trying.")
test_predictions()
```
| github_jupyter |
# Section 2.2: Naive Bayes
In contrast to *k*-means clustering, Naive Bayes is a supervised machine-learning (ML) algorithm. It provides good speed and good accuracy and is often used in aspects of natural-language processing such text classification or, in our case in this section, spam detection.
Spam emails are more than just a nuisance. As recently as 2008, spam constituted an apocalyptic 97.8 percent of all email traffic according to a [2009 Microsoft security report](http://download.microsoft.com/download/4/3/8/438BE24D-4D58-4D9A-900A-A1FC58220813/Microsoft_Security_Intelligence_Report _volume8_July-Dec2009_English.pdf). That tide has thankfully turned and, as of May 2019, spam makes up only about [85 percent of email traffic](https://www.talosintelligence.com/reputation_center/email_rep) — thanks, in no small part, to Naive Bayes spam filters.
Naive Bayes is a convenient algorithm for spam detection because it does not require encoding complex rules. All it needs is training examples, of which there are plenty when it comes to email spam. Naive Bayes does all this through the use of [conditional probability](https://en.wikipedia.org/wiki/Conditional_probability).
> **Learning objective:** By the end of this section, you should have a basic understanding of how naive Bayes works and some of the reasons for its popularity.
## Conditional probability
Ordinary probability deals with the likelihood of isolated events occurring. For example, rolling a 6 on a fair six-sided die will occur, on average, on one out of six rolls. Mathematicians express this probability as $P({\rm die}=6)=\frac{1}{6}$.
Conditional probability concerns itself with the contingencies of interconnected events: what is the probability of event $A$ happening if event $B$ occurs. Mathematicians denote this as $P(A|B)$, or "the probability of $A$ given $B$."
In order to compute the probability of conditional events, we use the following equation:
$P(A \mid B)=\cfrac{P(A \cap B)}{P(B)}$
This equation is nice, but it assumes that we know the joint probability $P(A\cap B)$, which we often don't. Instead, we often need to know something about $A$ but all we can directly observe is $B$. For instance, when we want to infer whether an email is spam only by knowing the words it contains. For this, we need Bayes' law.
## Bayes' law
Bayes' law takes its name from the eighteenth-century English statistician and philosopher Thomas Bayes, who described the probability of an event based solely on prior knowledge of conditions that might be related to that event thus:
$P(A \mid B)=\cfrac{P(B \mid A)P(A)}{P(B)}$
In words, Bayes' Law says that if I know the prior probabilities $P(A)$ and $P(B)$, in addition to the likelihood (even just an assumed likelihood) $P(B \mid A)$, I can compute the posterior probability $P(A \mid B)$. Let's apply this to spam.
<img align="center" style="padding-right:10px;" src="Images/spam.png" border="5">
In order to use Bayesian probability on spam email messages like this one, consider it (and all other emails, spam or ham) to be bags of words. We don't care about word order or even word meaning. We just want to count the frequency of certain words in spam messages versus the frequency of those same words in valid email messages.
Let's say that, after having counted the words in hundreds of emails that we have received, we determine the probability of the word "debt" appearing in any kind of email message (spam or ham) to be 0.157, with the probability of "debt" appearing in spam messages being 0.309. Further more, let's say that we assume that there is a 50 percent chance that any given email message we receive is spam (for this example, we don't know either way what type of email it might be, so it's a coin flip). Mathematically, we could thus say:
- Probability that a given message is spam: $P({\rm S})=0.5$
- Probability that “debt” appears in a given message: $P({\rm debt})=0.157$
- Probability that “debt” appears in a spam message: $P({\rm debt} \mid {\rm S})=0.309$
Plugging this in to Bayes' law, we get the following probability that an email message containing the word "debt" is spam:
$P({\rm S} \mid {\rm debt})=\cfrac{P({\rm debt} \mid {\rm S})P({\rm S})}{P({\rm debt})}=\cfrac{(0.309)(0.5)}{0.157}=\cfrac{0.1545}{0.157}=0.984$
Thus if an email contains the word "debt," we calculate that it is 98.4 percent likely to be spam.
## What makes it naive?
Our above calculation is great for looking at individual words, but emails contain several words that can give us clues to an email's relative likelihood of being spam or ham. For example, say we wanted to determine whether an email is spam given that it contains the words "debt" and "bills." We can begin by reasoning that the probability that an email containing "debt" and "bills" is spam is, if not equal, at least proportional to the probability of "debt" and "bills" appearing in known spam messages times the probability of any given message being spam:
$P({\rm S} \mid {\rm debt, bills}) \propto P({\rm debt, bills} \mid {\rm S})P({\rm S})$
(**Mathematical note:** The symbol ∝ represents proportionality rather than equality.)
Now if we assume that the occurrence of the words "debt" and "bills" are independent events, we can extend this proportionality:
$P({\rm S} \mid {\rm debt, bills}) \propto P({\rm debt} \mid {\rm S})P({\rm bills} \mid {\rm S})P({\rm S})$
We should state here that this assumption of independence is generally not true. Just look at the example spam message above. The probability that "bills" will appears in a spam message containing "debt" is probably quite high. However, assuming that the probabilities of words occurring in our email messages are independent is useful and works surprising well. This assumption of independence is the naive part of the Baysian probabilities that we will use in this section; expressed mathematically, the working assumption that will underpin the ML in this section is that for any collection of $n$ words:
$P({\rm S}\mid {\rm word_1}, {\rm word_2},\ldots, {\rm word}_n)=P({\rm S})P({\rm word_1}\mid {\rm S})P({\rm word_2}\mid {\rm S})\cdots P({\rm word}_n\mid {\rm S})$
> **Key takeaway:** We cannot emphasize enough that this chain rule expressed in the equation above—that the probability of a message being spam based on the words in it is equal to the product of the likelihoods of those individual words appearing in messages known to be spam is ***not*** true. But it gets good results and, in the world of data science, fast and good enough always trump mathematical fidelity.
## Import the dataset
In this section, we'll use the [SMS Spam Collection dataset](https://archive.ics.uci.edu/ml/datasets/SMS+Spam+Collection). It contains 5,574 messages collected for SMS spam research and tagged as "spam" or "ham." The dataset files contain one message per line with each line being composed of the tag and the raw text of the SMS message. For example:
| Class | Message |
|:------|:------------------------------|
| ham | What you doing?how are you? |
| ham | Ok lar... Joking wif u oni... |
Let’s now import pandas and load the dataset. (Note that the path name is case sensitive.)
```
import pandas as pd
df = pd.read_csv('Data/SMSSpamCollection', sep='\t', names=['Class', 'Message'])
```
> **Question**
>
> What do the `sep` and `names` parameters do in the code cell above? (**Hint:** If you are unsure, you can refer to the built-in Help documentation using `pd.read_csv?` in the code cell below.)
Let's take an initial look at what's in the dataset.
```
df.head()
```
Note that several entries in the `Message` column are truncated. We can use the `set_option()` function to set pandas to display the maximum width of each entry.
```
pd.set_option('display.max_colwidth', -1)
df.head()
```
> **Question**
>
> What do you think the purpose of the `-1` parameter passed to `pd.set_option()` is in the code cell above?
Alternatively, we can dig into individual messages.
```
df['Message'][13]
```
## Explore the data
Now that we have an idea of some of the individual entries in the dataset, let's get a better sense of the dataset as a whole.
```
df.info()
```
> **Exercise**
>
> Now run the `describe()` method on `df`. Does it provide much useful information about this dataset? If not, why not?
> **Possible exercise solution**
```
df.describe()
```
We can also visualize the dataset to graphically see the mix of spam to ham. (Note that we need to include the `%matplotlib inline` magic command in order to actually see the bar chart here in the notebook.)
```
%matplotlib inline
df.groupby('Class').count().plot(kind='bar')
```
> **Key takeaway:** Notice that here an in previous sections we have stuck together several methods to run on a `DataFrame`. This kind of additive method-stacking is part of what makes Python and pandas such a power combination for the rough-and-ready data exploration that is a crucial part of data science.
## Explore the data using word clouds
Because our data is largely not numeric, you might have noticed that some of our go-to data exploration tools (such as bar charts and the `describe()` method) have been of limited use in exploring this data. Instead, word clouds can be a powerful way of getting a quick glance at what's represented in text data as a whole.
```
!pip install wordcloud
```
We will have to supply a number of parameters to the `WordCloud()` function and to matplotlib in order to render the word clouds, so we will save ourselves some redundant work by writing a short function to handle it. Parameters for `WordCloud()` will include the stop words we want to ignore and font size for the words in the cloud. For matplotlib, these parameters will include instructions for rendering the word cloud.
```
from wordcloud import WordCloud, STOPWORDS
import matplotlib.pyplot as plt
def get_wordcloud(text_data,title):
wordcloud = WordCloud(background_color='black',
stopwords=set(STOPWORDS),
max_font_size=40,
relative_scaling=1.0,
random_state=1
).generate(str(text_data))
fig = plt.figure(1, figsize=(12, 12))
plt.axis('off')
plt.title(title)
plt.imshow(wordcloud)
plt.show()
```
Now it is time to plot the word clouds.
```
spam_msg = df.loc[df['Class']=='spam']['Message']
get_wordcloud(spam_msg,'Spam Cloud')
ham_msg = df.loc[df['Class']=='ham']['Message']
get_wordcloud(ham_msg,'Ham Cloud')
```
Looking at the two word clouds, it is immediately apparent that the frequency of the most common words is different between our spam and our ham messages, which will form the primary basis of our spam detection.
## Explore the data numerically
Just because the data does not naturally lend itself to numerical analysis "out of the box" does not mean that we can't do so. We can also analyze the average length of spam and ham messages to see if there are differences. For this, we need to create a new column.
```
df['Length_of_msg'] = df['Message'].apply(len)
df.head()
```
> **Question**
>
> What does the `apply()` method do in the code cell above? (**Hint:** If you are unsure, you can refer to [this page](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.apply.html).)
Now that we have the length of each message, we can visualize those message lengths using a histogram.
```
df.groupby('Class')['Length_of_msg'].plot(kind='hist', bins=50)
```
The orange histogram is the spam messages. Because there are so many more ham messages than spam, let's break these out separately to see the details more clearly.
```
df.hist(bins=50,by='Class', column='Length_of_msg')
```
Spam messages skew much longer than ham messages.
> **Question**
>
> Why does it appear in the details histograms that there is almost no overlap between the lengths of ham and spam text messages? What do the differences in scale tell us (and what could they inadvertently obscure)?
Let's look at the differences in length of the two classes of message numerically.
```
df.groupby('Class').mean()
```
These numbers accord with what we saw in the histograms.
Now, let's get to the actual modeling and spam detection.
## Prepare the data for modeling
One of the great strengths of naive Bayes analysis is that we don't have to go too deep into text processing in order to develop robust spam detection. However, the text is raw and it does require a certain amount of cleaning. To do this, we will use one of the most commonly used text-analytics libraries in Python, the Natural Language Toolkit (NLTK). However, before we can import it, we will need to first install it.
```
!pip install nltk
```
We can now import NLTK, in addition to the native Python string library to help with our text manipulation. We will also download the latest list of stop words (such as 'the', 'is', and 'are') for NLTK.
```
import string
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
```
Part of our data preparation will be *vectorizing* the text data. Recall that earlier in the section when we first introduced naive Bayes analysis, we stated that we wanted to treat our messages as "bags of words" rather than as English-language messages. Vectorization is the process by which we convert our collection of text messages to a matrix of word counts.
Part of the vectorization process will be for us to remove punctuation from the messages and exclude stop words from our analysis. We will write a function to perform those tasks here, because we will want to access those actions later on.
```
def txt_preprocess(text):
#Remove punctuation
temp = [w for w in text if w not in string.punctuation]
temp = ''.join(temp)
#Exclude stopwords
processedtext = [w for w in temp.split() if w.lower() not in stopwords.words('english')]
return processedtext
```
Scikit-learn provides a count-vectorizer function. We will now import it and then use the `txt_preprocess()` function we just wrote as a custom analyzer for it.
```
from sklearn.feature_extraction.text import CountVectorizer
X = df['Message']
y = df['Class']
CountVect = CountVectorizer(analyzer=txt_preprocess).fit(X)
```
> **Technical note:** The convention of using an upper-case `X` to represent the independent variables (the predictors) and a lower-case `y` to represent the dependent variable (the response) comes from statistics and is commonly used by data scientists.
In order to see how the vectorizer transformed the words, let's check it against a common English word like "go."
```
print(CountVect.vocabulary_.get('go'))
```
So "go" appears 6,864 times in our dataset.
Now, before we transform the entire dataset and train the model, we have the final preparatory step of splitting our data into training and test data to perform.
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=50)
```
Finally, we will transform our training messages into a [document-term matrix](https://en.wikipedia.org/wiki/Document-term_matrix). "Document" might sound a little grandiose in this case as it refers to individual text messages, but it is a term of art for text analysis.
```
X_train_data = CountVect.transform(X_train)
```
This can be a tricky concept, so let's look at the training-text matrix directly.
```
print(X_train_data)
X_train_data.shape
```
`X_train_data` is now a 3900x11425 matrix, where each of the 3,900 rows represents a text ("document") from the training dataset and each column is a specific word (11,425 of them in this case).
> **Key takeaway:** Putting our bag of words into a document-term matrix like this is a standard tool of natural-language processing and text analysis, and it is used in contexts beyond naive Bayes analysis in which word-frequency is important, such as [term frequency–inverse document frequency (TF-IDF)](https://en.wikipedia.org/wiki/Tf%E2%80%93idf).
## Train the model
Now it is time to train our naive Bayes model. For our model, we will use the multinomial naive Bayes classifier. "Multinomial" in this case derives from our assumption that, for our bag of $n$ words, $P({\rm S}\mid {\rm word_1}, {\rm word_2},\ldots, {\rm word}_n)=P({\rm S})P({\rm word_1}\mid {\rm S})P({\rm word_2}\mid {\rm S})\cdots P({\rm word}_n\mid {\rm S})$ and that we don't assume that our word likelihoods follow a normal distribution.
```
from sklearn.naive_bayes import MultinomialNB
naivebayes_model = MultinomialNB()
naivebayes_model.fit(X_train_data,y_train)
```
Our model is now fitted. However, before we run our predictions on all of our test data, let's see what our model says about some artificial data in order to get a better sense of what our model will do with all of the messages in our test dat. From the word clouds we constructed earlier, we can see that "call" and "free" are both prominent words among our spam messages, so let's create our own spam message and see how our model classifies it.
```
pred = naivebayes_model.predict(CountVect.transform(['Call for a free offer!']))
pred
```
As we expected, our model correctly classified this message as spam.
> **Exercise**
>
> Review the ham word cloud above, construct a ham message, and then run it against the model to see how it is classified.
> **Possible exercise solution**
```
pred2 = naivebayes_model.predict(CountVect.transform(['Let me know what time we should go.']))
pred2
```
Now let's run our test data through the model. First, we need to transform it to a document-term matrix.
```
X_test_data = CountVect.transform(X_test)
X_test_data.shape
```
> **Exercise**
>
> Run the predictions for the test data.
> **Exercise solution**
```
predictions = naivebayes_model.predict(X_test_data)
predictions
```
Now it's time to evaluate our model's performance.
```
from sklearn.metrics import classification_report, confusion_matrix
print(classification_report(predictions, y_test))
```
> **Exercise**
>
> Overall, our model is good for spam detection, but our recall score (the proportion of actual positives that were identified correctly) is surprisingly low. Why might this be? What implications does it have for spam detection? (**Hint:** Use the scikit-learn `confusion_matrix()` function to better understand the specific performance of the model. For help interpreting the confusion matrix, see [this page](https://en.wikipedia.org/wiki/Confusion_matrix).)
> **Possible exercise solution**
```
print(confusion_matrix(y_test, predictions))
```
> **Takeaway**
>
> The performance of our naive Bayes model helps underscore the algorithm's popularity, particularly for spam detection. Even untuned, we got good performance, performance that would only continue to improve in production as users submitted more examples of spam messages.
## Further exploration
Beyond detecting spam, we can use ML to explore the SMS data more deeply. To do so, we can use sophisticated, cloud-based cognitive tools such as Microsoft Azure Cognitive Services.
### Azure Cognitive Services
The advantage of using cloud-based services is that they provide cutting-edge models that you can access without having to train the models. This can help accelerate both your exploration and your use of ML.
Azure provides Cognitive Services APIs that can be consumed using Python to conduct image recognition, speech recognition, and text recognition, just to name a few. For the purposes of this subsection, we're going to look at using the Azure Text Analytics API.
First, we’ll start by obtaining a Cognitive Services API key. Note that you can get a free key for seven days (after which you'll be required to pay for continued access to the API).
To learn more about pricing for Cognitive Services, see https://azure.microsoft.com/en-us/pricing/details/cognitive-services/
Browse to **Try Azure Cognitive Services** at https://azure.microsoft.com/en-us/try/cognitive-services/
1. Click **Language APIs**.
2. By **Text Analytics**, click **Get API key**.
3. In the **Try Cognitive Services for free** window, under **7-day trial**, click **Get stared**.
4. In the **Microsoft Cognitive Services Terms** window, accept the terms of the free trial and click **Next**.
5. In the **Sign-in to Continue** window, select your preferred means of signing in to your Azure account.
Once you have your API keys in hand, you're ready to start. Substitute the API key that you get for the seven-day trial below where it reads `ACCOUNT_KEY`.
```
# subscription_key = 'ACCOUNT_KEY'
subscription_key = '8efb79ce8fd84c95bd1aa2f9d68ae734'
assert subscription_key
# If using a Free Trial account, this URL does not need to be updated.
# If using a paid account, verify that it matches the region where the
# Text Analytics Service was setup.
text_analytics_base_url = "https://westcentralus.api.cognitive.microsoft.com/text/analytics/v2.1/"
```
We will also need to import the NumPy and requests modules.
```
import numpy as np
import requests
```
The Azure Text Analytics API has a hard limit of 1,000 calls at a time, so we will need to split our 5,572 SMS messages into at least six chunks to run them through Azure.
```
chunks = np.array_split(df, 6)
for chunk in chunks:
print(len(chunk))
```
Two of the things that cognitives services like those provided by Azure offer are language identification and sentiment analysis. Both are relevant for our dataset, so we will prepare our data for both by submitting them as JavaScript Object Notation (JSON) documents. We'll prepare the data for language identification first.
```
# Prepare the header for the JSON document including your subscription key
headers = {"Ocp-Apim-Subscription-Key": subscription_key}
# Supply the URL for the language-identification API.
language_api_url = text_analytics_base_url + "languages"
# Iterate over the chunked DataFrame.
for i in range(len(chunks)):
# Reset the indexes within the chunks to avoid problems later on.
chunks[i] = chunks[i].reset_index()
# Split up the message from the DataFrame and put them in JSON format.
documents = {'documents': []}
for j in range(len(chunks[i]['Message'])):
documents['documents'].append({'id': str(j), 'text': chunks[i]['Message'][j]})
# Call the API and capture the responses.
response = requests.post(language_api_url, headers=headers, json=documents)
languages = response.json()
# Put the identified languages in a list.
lang_list = []
for document in languages['documents']:
lang_list.append(document['detectedLanguages'][0]['name'])
# Put the list of identified languages in a new column of the chunked DataFrame.
chunks[i]['Language'] = np.array(lang_list)
```
Now we need perform similar preparation of the data for sentiment analysis.
```
# Supply the URL for the sentiment-analysis API.
sentiment_api_url = text_analytics_base_url + "sentiment"
# Iterate over the chunked DataFrame.
for i in range(len(chunks)):
# We have alread reset the chunk-indexes, so we don't need to do again.
# Split up the messages from the DataFrame and put them in JSON format.
documents = {'documents': []}
for j in range(len(chunks[i]['Message'])):
documents['documents'].append({'id': str(j), 'text': chunks[i]['Message'][j]})
# Call the API and capture the responses.
response = requests.post(sentiment_api_url, headers=headers, json=documents)
sentiments = response.json()
# Put the identified sentiments in a list.
sent_list = []
for document in sentiments['documents']:
sent_list.append(document['score'])
# Put the list of identified sentiments in a new column of the chunked DataFrame.
chunks[i]['Sentiment'] = np.array(sent_list)
```
We now need to re-assembled our chunked DataFrame.
```
azure_df = pd.DataFrame(columns=['Index', 'Class', 'Message', 'Language', 'Sentiment'])
for i in range(len(chunks)):
azure_df = pd.concat([azure_df, chunks[i]])
if i == 0:
azure_df['index'] = chunks[i].index
azure_df.set_index('index', inplace=True)
azure_df.drop(['Index'], axis=1, inplace=True)
azure_df.head()
```
We can also look at the tail of the `DataFrame` to check that our indexing worked as expected.
```
azure_df.tail()
```
Let's now see if all of the SMS messages were in English (and, if not, how many messages of which languages we are looking at).
```
azure_df.groupby('Language')['Message'].count().plot(kind='bar')
```
So the overwhelming majority of the messages are in English, though we have several additional languages in our dataset. Let's look at the actual numbers.
> **Exercise**
>
> Now use the `groupby` method to display actual counts of the languages detected in the dataset rather than a bar chart of them.
> **Exercise solution**
```
azure_df.groupby('Language')['Message'].count()
```
We have a surprising array of languages, perhaps, but the non-English messages are really just outliers and should have no real impact on the spam detection.
Now let's look at the sentiment analysis for our messages.
```
azure_df.groupby('Class')['Sentiment'].plot(kind='hist', bins=50)
```
It is perhaps not too surprising that the sentiments represented in the dataset should be bifurcated: SMS is a medium that captures extremes better than nuanced middle ground. That said, the number of dead-center messages is interesting. The proportion of spam messages right in the middle is also interesting. Let's break the two classes (ham and spam) into separate histograms to get a better look.
> **Exercise**
>
> Break out the single histogram above into two histograms (one for each class of message). (**Hint:** Refer back to the code we used to do this earlier in the section.)
> **Exercise solution**
```
azure_df.hist(bins=50,by='Class', column='Sentiment')
```
The number of spam messages in our dataset is about a tenth of the amount of ham, yet the number of spam messages with exactly neutral sentiment is about half that of the ham, indicating that spam messages, on average, tend to be more neutral than legitimate messages. We can also notice that non-neutral spam messages tend to have more positive than negative sentiment, which makes intuitive sense.
> **Takeaway**
>
> Beyond providing additional insight into our data, sophisticated language-identification and sentiment-analysis algorithms provided by cloud-based services like Azure can provide additional details that could potentially help improve spam detection. For example, how patterns of sentiments in spam differ from those in legitimate messages.
| github_jupyter |
# Computing FSAs
**(C) 2017-2019 by [Damir Cavar](http://damir.cavar.me/)**
**Version:** 1.0, September 2019
**License:** [Creative Commons Attribution-ShareAlike 4.0 International License](https://creativecommons.org/licenses/by-sa/4.0/) ([CA BY-SA 4.0](https://creativecommons.org/licenses/by-sa/4.0/))
## Introduction
Consider the following automaton:
<img src="NDFSAMatrixOp.png" caption="Non-deterministic Finite State Automaton" style="width: 200px;"/>
We can represent it in terms of transition tables. We will use the Python numpy module for that.
```
from numpy import array
```
The transitions are coded in terms of state to state transitions. The columns and rows represent the states 0, 1, and 2. The following transition matrix shows all transitions that are associated with the label "a", that is from 0 to 0, from 0 to 1, and from 1 to 0.
```
a = array([
[1, 1, 0],
[1, 0, 0],
[0, 0, 0]
])
```
The following transition matrix shows that for the transitions associated with "b".
```
b = array([
[0, 1, 0],
[0, 1, 0],
[0, 0, 0]
])
```
The following transition matrix shows this for the transitions associated with "c".
```
c = array([
[0, 0, 0],
[0, 0, 1],
[0, 0, 0]
])
```
We can define the start state using an init vector. This init vector indicates that the start state should be 0.
```
init = array([
[1, 0, 0]
])
```
The set of final states can be encoded as a column vector that in this case defines state 3 as the only final state.
```
final = array([
[0],
[0],
[1]
])
```
If we want to compute the possibility for a sequence like "aa" to be accepted by this automaton, we could compute the dot product of the init-vector and the a matrices, with the dot product of the final state.
```
init.dot(a).dot(c).dot(final)
```
The 0 indicates that there is no path from the initial state to the final state based on a sequence "aa".
Let us verify this for a sequence "bc", for which we know that there is such a path:
```
init.dot(b).dot(c).dot(final)
```
Just to verify once more, let us consider the sequence "aabc":
```
init.dot(a).dot(a).dot(b).dot(c).dot(final)
```
There are obviously three paths in our Non-deterministic Finite State Automaton that generate the sequence "aabc".
## Wrapping the Process into a Function
We could define the FSA above as a 5-tuple $(\Sigma, Q, i, F, E)$, with:
$\Sigma = \{a, b, c\}$, the set of symbols.
$Q = \{ 0, 1, 2 \}$, the set of states.
$i \in Q$, with $i = 0$, the initial state.
$F \subseteq Q$, with $F = \{ 2 \}$, the set of final states.
$E \subseteq Q \times (\Sigma \cup \epsilon) \times Q$, the set of transitions.
$E$ is the subset of tuples determined by the cartesian product of the set of states, the set of symbols including the empty set, and the set of states. This tuple defines a transition from one state to another state with a specific symbol.
$E$ could also be defined in terms of a function $\delta(\sigma, q)$, with $\sigma$ an input symbol and $q$ the current state. $\delta(\sigma, q)$ returns the new state of the transition, or a failure. The possible transitions for any given symbol from any state can be defined in a transition table:
| | a | b | c |
| :---: | :---: | :---: | :---: |
| **0** | 0, 1 | 1 | - |
| **1** | 0 | 1 | 2 |
| **2:** | - | - | - |
We can define the automaton in Python:
```
S = set( ['a', 'b', 'c'] )
Q = set( [0, 1, 2] )
i = 0
F = set( [ 2 ] )
td = { (0, 'a'): [0, 1],
(1, 'a'): [0],
(0, 'b'): [1],
(1, 'b'): [1],
(1, 'c'): [2]
}
def df(state, symbol):
print(state, symbol)
return td.get(tuple( [state, symbol] ), [])
def accept(sequence):
agenda = []
state = i
count = len(sequence)
agenda.append((state, 0))
while agenda:
print(agenda)
if not agenda:
break
state, pos = agenda.pop()
states = df(state, sequence[pos])
if not states:
print("No transition")
return False
state = states[0]
if pos == count - 1:
print("Reached end")
if F.intersection(set(states)):
return True
break
for s in states[1:]:
agenda.append( (s, pos+1) )
if state in F:
print("Not final state")
return True
return False
accept("aac")
alphabetMatrices = {}
alphabetMatrices["a"] = array([
[1, 1, 0],
[1, 0, 0],
[0, 0, 0]
])
alphabetMatrices["b"] = array([
[0, 1, 0],
[0, 1, 0],
[0, 0, 0]
])
alphabetMatrices["c"] = array([
[0, 0, 0],
[0, 0, 1],
[0, 0, 0]
])
alphabetMatrices["default"] = array([
[0, 0, 0],
[0, 0, 0],
[0, 0, 0]
])
def paths(seq):
res = init
for x in seq:
res = res.dot( alphabetMatrices.get(x, alphabetMatrices["default"]) )
return res.dot(array([
[0],
[0],
[1]
]))[0][0]
paths("aabc")
```
**(C) 2016-2019 by [Damir Cavar](http://damir.cavar.me/) <<dcavar@iu.edu>>**
| github_jupyter |
```
%matplotlib inline
%load_ext autoreload
%autoreload 2
from constants_and_util import *
import matplotlib.pyplot as plt
import pandas as pd
import random
import numpy as np
from copy import deepcopy
from scipy.signal import argrelextrema
import statsmodels.api as sm
from scipy.special import expit
from scipy.stats import scoreatpercentile
import pickle
import os
from collections import Counter
import dataprocessor
import compare_to_seasonal_cycles
assert not USE_SIMULATED_DATA
import sys
import cPickle
assert sys.version[0] == '2'
import seaborn as sns
import generate_results_for_paper
generate_results_for_paper.make_figure_to_illustrate_data_for_one_user()
generate_results_for_paper.make_maps_of_countries_with_clue_data()
results = compare_to_seasonal_cycles.load_all_results()
# Sentence about relative change in happy/sad curve.
happy_sad_curve = compare_to_seasonal_cycles.convert_regression_format_to_simple_mean_format(
results['emotion*happy_versus_emotion*sad']['by_very_active_northern_hemisphere_loggers'][True]['linear_regression'],
'linear_regression')
cycle_amplitude = compare_to_seasonal_cycles.get_cycle_amplitude(happy_sad_curve,
cycle='date_relative_to_period',
metric_to_use='max_minus_min',
hourly_period_to_exclude=None)
overall_sad_frac = 1 - results['emotion*happy_versus_emotion*sad']['by_very_active_northern_hemisphere_loggers'][True]['overall_positive_frac']
print("Happy/sad frac: %2.3f; period cycle amplitude: %2.3f; relative change: %2.3f" % (overall_sad_frac,
cycle_amplitude,
cycle_amplitude/overall_sad_frac))
generate_results_for_paper.make_cycle_amplitudes_bar_plot_for_figure_1(results)
generate_results_for_paper.make_happiness_by_date_date_trump_effects_plot_for_figure_1(results)
generate_results_for_paper.make_happiness_by_date_date_trump_effects_plot_for_figure_1(results,
plot_red_line=False)
compare_to_seasonal_cycles.make_four_cycle_plots(results,
['by_very_active_northern_hemisphere_loggers'],
['emotion*happy_versus_emotion*sad'],
ylimits_by_pair={'emotion*happy_versus_emotion*sad':4},
figname='figures_for_paper/four_cycle_plot.png',
suptitle=False,
include_amplitudes_in_title=False,
different_colors_for_each_cycle=True)
```
# Figure 2
this has already been filtered for countries with MIN_USERS_FOR_SUBGROUP and MIN_OBS_FOR_SUBGROUP
```
generate_results_for_paper.make_maps_for_figure_2(results)
```
# Figure 3: age effects.
this has already been filtered for ages with MIN_USERS_FOR_SUBGROUP and MIN_OBS_FOR_SUBGROUP
```
opposite_pairs_to_plot = ['emotion*happy_versus_emotion*sad',
'continuous_features*heart_rate_versus_continuous_features*null',
'continuous_features*bbt_versus_continuous_features*null',
'continuous_features*weight_versus_continuous_features*null']
generate_results_for_paper.make_age_trend_plot(results,
opposite_pairs_to_plot=opposite_pairs_to_plot,
specifications_to_plot=['age'],
figname='figures_for_paper/main_fig4.pdf',
plot_curves_for_two_age_groups=True,
n_subplot_rows=2,
n_subplot_columns=4,
figsize=[14, 8],
subplot_kwargs={'wspace':.3,
'hspace':.65,
'right':.95,
'left':.15,
'top':.92,
'bottom':.1},
plot_yerr=True)
generate_results_for_paper.make_age_trend_plot(results,
opposite_pairs_to_plot=ORDERED_SYMPTOM_NAMES,
specifications_to_plot=['age',
'country+age',
'country+age+behavior',
'country+age+behavior+app usage'],
figname='figures_for_paper/age_trend_robustness.png',
plot_curves_for_two_age_groups=False,
n_subplot_rows=5,
n_subplot_columns=3,
figsize=[12, 15],
subplot_kwargs={'wspace':.7,
'hspace':.95,
'right':.72,
'left':.12,
'top':.95,
'bottom':.1},
age_ticks_only_at_bottom=False,
label_kwargs={'fontsize':11},
linewidth=1,
plot_legend=True,
include_ylabel=False)
```
| github_jupyter |
# PyCity Schools Analysis & Observations
• Overall, all schools performed better in reading (100%) rather than Math (92%) since the average reading score is higher than the average math score at every school in the city. • The charter schools performed better at math and reading than the district schools • Performance did not correlate to amount spent per student, but to the size of the school and type. The smaller schools, which were all charter schools, performed better. • District schools performed far low across the board in math and reading scores than charter schools and the lowest scores were also observed among schools that spend the most per student, and the largest schools) • Higher Per Student Budget not necessary lead to a higher grade, for example Cabrera High School, who has the highest passing rate spent less than 600 dollar budget per student, and the bottom 5 school all have budget per student higher than 600 dollar. In addition, schools spent lower than 615 dollar per student has significant (15%) higher scores than over 615 dollar. The highest spending category 645-675 got the lowest score. • School type matters. All Top 5 school are Charter school, and 5 Bottom schools are District Schools. Overall variance between these two on passing rate are 23%, Charter schools are higher. Charter school performed especially well in math score. • Larger size schools doesn’t show good performance in score, compared to median size and small size school. However once the school size goes below 2000 students, the change are not significant.
### Note
* Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
```
# Dependencies and Setup
import pandas as pd
# File to Load (Remember to Change These)
school_data_to_load = "Resources/schools_complete.csv"
student_data_to_load = "Resources/students_complete.csv"
# Read School and Student Data File and store into Pandas DataFrames
school_data = pd.read_csv(school_data_to_load)
student_data = pd.read_csv(student_data_to_load)
# Combine the data into a single dataset.
school_data_complete = pd.merge(student_data, school_data, how="left", on=["school_name", "school_name"])
school_data_complete.head()
```
## District Summary
* Calculate the total number of schools
* Calculate the total number of students
* Calculate the total budget
* Calculate the average math score
* Calculate the average reading score
* Calculate the percentage of students with a passing math score (70 or greater)
* Calculate the percentage of students with a passing reading score (70 or greater)
* Calculate the percentage of students who passed math **and** reading (% Overall Passing)
* Create a dataframe to hold the above results
* Optional: give the displayed data cleaner formatting
```
#Calculate the total number of schools
num_of_schools = school_data['school_name'].count()
#print(num_of_schools )
#Calculate the total number of students
num_of_students = student_data['Student ID'].count()
#print(num_of_students)
#Calculate the total budget
total_budget = school_data['budget'].sum()
#print(total_budget)
#Calculate the average math score
avg_math_score = school_data_complete['math_score'].mean()
#print(avg_math_score)
#Calculate the average reading score
avg_reading_score = school_data_complete['reading_score'].mean()
#print(avg_reading_score)
#Calculate the percentage of students with a passing math score (70 or greater)
pass_math = school_data_complete[(school_data_complete['math_score'] >= 70)].count() ['student_name']
#print(pass_math)
math_percent = (pass_math / float(num_of_students))*100
#print(math_percent)
#Calculate the percentage of students with a passing reading score (70 or greater)
pass_reading = school_data_complete[(school_data_complete['reading_score'] >= 70)].count() ['student_name']
#print(pass_reading)
reading_percent = (pass_reading / float(num_of_students))*100
#print(reading_percent)
#Calculate the percentage of students who passed math **and** reading (% Overall Passing)
pass_math_reading = school_data_complete[(school_data_complete['math_score'] >= 70) & (school_data_complete['reading_score'] >= 70) ].count() ['student_name']
#print(pass_math_reading)
math_reading_percent = (pass_math_reading / float(num_of_students))*100
#print(math_reading_percent)
```
The brackets are unnecessary when referencing variables - this is in the dictionary for district_summary.
#ValueError: If using all scalar values, you must pass an index
```
#Create a dataframe to hold the above results
#Optional: give the displayed data cleaner formatting
# district_summary = pd.DataFrame ({'total_schools': [num_of_schools],'total_students': [num_of_students],
# 'total_budget': [total_budget], 'avg_math_score': [avg_math_score],
# 'avg_reading_score': [avg_reading_score],'percentage_pass_math': [math_percent],
# 'percentage_pass_reading': [reading_percent], 'overall pass percent': [math_reading_percent]
# })
district_summary = pd.DataFrame({'total_schools': num_of_schools,'total_students': num_of_students,
'total_budget': total_budget, 'avg_math_score': avg_math_score,
'avg_reading_score': avg_reading_score,'percentage_pass_math': math_percent,
'percentage_pass_reading': reading_percent, 'overall pass percent': math_reading_percent}
,index =[0] )
district_summary['total_students'] = district_summary['total_students'].map("{:,}".format)
district_summary['total_budget'] = district_summary['total_budget'].map("${:,.2f}".format)
district_summary
```
## School Summary
* Create an overview table that summarizes key metrics about each school, including:
* School Name
* School Type
* Total Students
* Total School Budget
* Per Student Budget
* Average Math Score
* Average Reading Score
* % Passing Math
* % Passing Reading
* % Overall Passing (The percentage of students that passed math **and** reading.)
* Create a dataframe to hold the above results
Why do you set the school type for school_type to the index?
school_type = school_data['type'] - when i tried like this i got all NAN values in the rows from 0-14. So i thought i need to set the index.
```
#School Summary - School name
school_summary = school_data_complete.groupby("school_name")
#print(school_summary["school_name"].unique())
#school Type
school_type = school_data.set_index(["school_name"])['type']
#school_type = school_data['type']
#print(school_type)
#Total number of students per school
total_students = school_data_complete.groupby(["school_name"]).count()['Student ID']
#print(total_students)
#Total School Budget
total_school_budget = school_data_complete.groupby(["school_name"]).mean()['budget']
#print(total_school_budget)
#Per Student Budget
per_student_budget = total_school_budget/total_students
#print(per_student_budget)
#Average Math score and Passing Percecntage
avg_math_score_per_student = school_summary['math_score'].mean()
#print(avg_math_score_per_student)
passing_math = school_data_complete[(school_data_complete['math_score'] >= 70)]
#print(passing_math)
percent_passing_math = (passing_math.groupby(["school_name"]).count()['Student ID'] / total_students)*100
#print(percent_passing_math)
#Average Reading score and Passing Percentage
avg_reading_score_per_student = school_summary['reading_score'].mean()
#print(avg_reading_score_per_student)
passing_reading = school_data_complete[(school_data_complete['reading_score'] >= 70)]
#print(passing_reading)
percent_passing_reading = (passing_reading.groupby(["school_name"]).count()['Student ID'] / total_students)*100
#print(percent_passing_reading)
#Overall Passing Percentage
overall_passing = school_data_complete[(school_data_complete['math_score'] >= 70) & (school_data_complete['reading_score'] >= 70)]
#print(overall_passing)
overall_passing_percent = (overall_passing.groupby(["school_name"]).count()['Student ID'] / total_students)*100
#print(overall_passing_percent)
schools_summary = pd.DataFrame ({'School Type': school_type,'Total students': total_students,
'Total School Budget': total_school_budget,
'Per Student Budget': per_student_budget,
'Average Math Score': avg_math_score_per_student,
'Average Reading Score': avg_reading_score_per_student,
'% Passing Math': percent_passing_math,
'% Passing Reading': percent_passing_reading,
'% Overall Passing': overall_passing_percent
})
schools_summary['Total School Budget'] = schools_summary['Total School Budget'].map("${:,.2f}".format)
schools_summary['Per Student Budget'] = schools_summary['Per Student Budget'].map("${:.2f}".format)
schools_summary
```
## Top Performing Schools (By % Overall Passing)
* Sort and display the top five performing schools by % overall passing.
```
top_performing = schools_summary.sort_values("% Overall Passing", ascending = False)
top_performing.head()
```
## Bottom Performing Schools (By % Overall Passing)
* Sort and display the five worst-performing schools by % overall passing.
```
bottom_performing = schools_summary.sort_values("% Overall Passing")
bottom_performing.head()
```
## Math Scores by Grade
* Create a table that lists the average Reading Score for students of each grade level (9th, 10th, 11th, 12th) at each school.
* Create a pandas series for each grade. Hint: use a conditional statement.
* Group each series by school
* Combine the series into a dataframe
* Optional: give the displayed data cleaner formatting
```
ninth_grade_math = student_data.loc[student_data['grade'] == '9th'].groupby('school_name')["math_score"].mean()
tenth_grade_math = student_data.loc[student_data['grade'] == '10th'].groupby('school_name')["math_score"].mean()
eleventh_grade_math = student_data.loc[student_data['grade'] == '11th'].groupby('school_name')["math_score"].mean()
twelvth_grade_math = student_data.loc[student_data['grade'] == '12th'].groupby('school_name')["math_score"].mean()
math_scores_grade = pd.DataFrame({
"9th": ninth_grade_math,
"10th": tenth_grade_math,
"11th": eleventh_grade_math,
"12th": twelvth_grade_math
})
math_scores_grade.head(15)
```
## Reading Score by Grade
* Perform the same operations as above for reading scores
```
ninth_grade_reading = student_data.loc[student_data['grade'] == '9th'].groupby('school_name')["reading_score"].mean()
tenth_grade_reading = student_data.loc[student_data['grade'] == '10th'].groupby('school_name')["reading_score"].mean()
eleventh_grade_reading = student_data.loc[student_data['grade'] == '11th'].groupby('school_name')["reading_score"].mean()
twelvth_grade_reading = student_data.loc[student_data['grade'] == '12th'].groupby('school_name')["reading_score"].mean()
reading_scores_grade = pd.DataFrame({
"9th": ninth_grade_reading,
"10th": tenth_grade_reading,
"11th": eleventh_grade_reading,
"12th": twelvth_grade_reading
})
reading_scores_grade.head(15)
```
## Scores by School Spending
* Create a table that breaks down school performances based on average Spending Ranges (Per Student). Use 4 reasonable bins to group school spending. Include in the table each of the following:
* Average Math Score
* Average Reading Score
* % Passing Math
* % Passing Reading
* Overall Passing Rate (Average of the above two)
```
bins = [0,585,630,645,675]
group_names = ["< $585","$585 - $629","$630 - $644","$645 - $675"]
school_data_complete['Spending Ranges (Per Student)'] = pd.cut(school_data_complete['budget']/school_data_complete['size'], bins, labels = group_names)
score_by_budget = school_data_complete.groupby('Spending Ranges (Per Student)')
avg_math = score_by_budget['math_score'].mean()
avg_read = score_by_budget['reading_score'].mean()
pass_math = school_data_complete[school_data_complete['math_score'] >= 70].groupby('Spending Ranges (Per Student)')['Student ID'].count()/score_by_budget['Student ID'].count() * 100
pass_read = school_data_complete[school_data_complete['reading_score'] >= 70].groupby('Spending Ranges (Per Student)')['Student ID'].count()/score_by_budget['Student ID'].count() * 100
overall = school_data_complete[(school_data_complete['math_score'] >= 70) & (school_data_complete['reading_score'] >= 70)].groupby('Spending Ranges (Per Student)')['Student ID'].count()/score_by_budget['Student ID'].count() * 100
scores_by_budget = pd.DataFrame({
"Average Math Score": avg_math,
"Average Reading Score": avg_read,
"% Passing Math": pass_math,
"% Passing Reading": pass_read,
"% Overall Passing": overall
})
scores_by_budget['Average Math Score'] = scores_by_budget['Average Math Score'].map("{:,.2f}".format)
scores_by_budget['Average Reading Score'] = scores_by_budget['Average Reading Score'].map("{:,.2f}".format)
scores_by_budget['% Passing Math'] = scores_by_budget['% Passing Math'].map("{:,.2f}".format)
scores_by_budget['% Passing Reading'] = scores_by_budget['% Passing Reading'].map("{:,.2f}".format)
scores_by_budget['% Overall Passing'] = scores_by_budget['% Overall Passing'].map("{:,.2f}".format)
scores_by_budget
```
## Scores by School Size
* Perform the same operations as above, based on school size.
```
bins = [0, 1000, 2000, 5000]
group_names = ["Small(<1000)", "Medium (1000 - 2000)" , "Large (2000 - 5000)"]
school_data_complete['School Size'] = pd.cut(school_data_complete['size'], bins, labels = group_names)
score_by_size = school_data_complete.groupby('School Size')
avg_math = score_by_size['math_score'].mean()
avg_read = score_by_size['reading_score'].mean()
pass_math = school_data_complete[school_data_complete['math_score'] >= 70].groupby('School Size')['Student ID'].count()/score_by_size['Student ID'].count() * 100
pass_read = school_data_complete[school_data_complete['reading_score'] >= 70].groupby('School Size')['Student ID'].count()/score_by_size['Student ID'].count() * 100
overall = school_data_complete[(school_data_complete['math_score'] >= 70) & (school_data_complete['reading_score'] >= 70)].groupby('School Size')['Student ID'].count()/score_by_size['Student ID'].count() * 100
scores_by_size = pd.DataFrame({
"Average Math Score": avg_math,
"Average Reading Score": avg_read,
"% Passing Math": pass_math,
"% Passing Reading": pass_read,
"% Overall Passing ": overall
})
scores_by_size
```
## Scores by School Type
* Perform the same operations as above, based on school type
```
score_by_type = school_data_complete.groupby('type')
avg_math = score_by_type['math_score'].mean()
avg_read = score_by_type['reading_score'].mean()
pass_math = school_data_complete[school_data_complete['math_score'] >= 70].groupby('type')['Student ID'].count()/score_by_type['Student ID'].count() * 100
pass_read = school_data_complete[school_data_complete['reading_score'] >= 70].groupby('type')['Student ID'].count()/score_by_type['Student ID'].count() * 100
overall = school_data_complete[(school_data_complete['math_score'] >= 70) & (school_data_complete['reading_score'] >= 70)].groupby('type')['Student ID'].count()/score_by_type['Student ID'].count() * 100
scores_by_type = pd.DataFrame({
"Average Math Score": avg_math,
"Average Reading Score": avg_read,
"% Passing Math": pass_math,
"% Passing Reading": pass_read,
"% Overall Passing": overall})
scores_by_type.index.names = ['School Type']
scores_by_type
```
| github_jupyter |
```
# coding: utf-8
# In[1]:
#trying to rewrite so faster using numpy historgram
#plotting and data analysis for global cold wakes
#from netCDF4 import Dataset # http://code.google.com/p/netcdf4-python/
import os
import time
import datetime as dt
import xarray as xr
from datetime import datetime
import pandas
import matplotlib.pyplot as plt
import numpy as np
import math
import geopy.distance
from math import sin, pi
from scipy import interpolate
from scipy import stats
#functions for running storm data
import sys
####################you will need to change some paths here!#####################
#list of input directories
dir_storm_info='f:/data/tc_wakes/database/info/'
dir_out='f:/data/tc_wakes/database/sst/'
#################################################################################
#start to look at data and make some pdfs
date_1858 = dt.datetime(1858,11,17,0,0,0) # start date is 11/17/1958
map_lats=np.arange(-90,90,.25)
map_lons=np.arange(-180,180,.25)
imap_lats = map_lats.size
imap_lons = map_lons.size
#for iyr_storm in range(2002,2017):
for iyr_storm in range(2002,2018):
#init arrays
init_data=0
map_sum,map_cnt,map_max = np.zeros([imap_lats,imap_lons]),np.zeros([imap_lats,imap_lons]),np.zeros([imap_lats,imap_lons])
map_sum_recov,map_cnt_recov = np.zeros([imap_lats,imap_lons]),np.zeros([imap_lats,imap_lons])
# for inum_storm in range(0,100):
for inum_storm in range(16,17):
filename = dir_out + str(iyr_storm) + '/' + str(inum_storm).zfill(3) + '_interpolated_track.nc'
exists = os.path.isfile(filename)
if not exists:
continue
print(filename)
ds_storm_info=xr.open_dataset(filename)
ds_storm_info = ds_storm_info.sel(j2=0)
ds_storm_info.close()
filename = dir_out + str(iyr_storm) + '/' + str(inum_storm).zfill(3) + '_combined_data.nc'
ds_all = xr.open_dataset(filename)
ds_all['spd']=np.sqrt(ds_all.uwnd**2+ds_all.vwnd**2)
ds_all.close()
filename = dir_out + str(iyr_storm) + '/' + str(inum_storm).zfill(3) + '_MLD_data_v2.nc'
ds_all2 = xr.open_dataset(filename)
ds_all2.close()
if abs(ds_all.lon[-1]-ds_all.lon[0])>180:
ds_all.coords['lon'] = np.mod(ds_all['lon'], 360)
ds_storm_info['lon'] = np.mod(ds_storm_info['lon'], 360)
max_lat = ds_storm_info.lat.max()
#remove all data outsice 100km/800km or cold wake >0 or <-10
if max_lat<0:
cond = ((((ds_all.dist_from_storm_km<100) & (ds_all.side_of_storm<=0)) |
((ds_all.dist_from_storm_km<800) & (ds_all.side_of_storm>0)))
& (ds_all.coldwake_max<=-.1) & (ds_all.coldwake_max>=-10))
else:
cond = ((((ds_all.dist_from_storm_km<800) & (ds_all.side_of_storm<0)) |
((ds_all.dist_from_storm_km<100) & (ds_all.side_of_storm>=0)))
& (ds_all.coldwake_max<=-.1) & (ds_all.coldwake_max>=-10))
subset = ds_all.where(cond)
subset2 = ds_all2.where(cond)
#create coldwake anomaly with nan for all values before wmo storm time
subset['sst_anomaly']=subset.analysed_sst-subset.sst_prestorm
# for i in range(0,xdim):
# for j in range (0,ydim):
# if np.isnan(subset.closest_storm_index[j,i]):
# continue
# iend = subset.closest_storm_index[j,i].data.astype(int)
# subset.sst_anomaly[:iend,j,i]=np.nan
#create array with day.frac since closest storm passage
tdif_dy = (subset.time-subset.closest_storm_time_np64)/np.timedelta64(1, 'D')
# tdif_dy = tdif_dy.where(tdif_dy>=0,np.nan)
subset['tdif_dy']=tdif_dy
xdim,ydim,tdim = ds_all.lon.shape[0],ds_all.lat.shape[0],ds_all.time.shape[0]
pdim=xdim*ydim
pdim3=tdim*xdim*ydim
print(xdim*ydim)
# data = subset.coldwake_max
# cbin1 = np.arange(-10, 0, 0.1) #cold wake bins
# bins=cbin1
# hist1,mids = np.histogram(data,bins)[0],0.5*(bins[1:]+bins[:-1])
# sum1 = np.cumsum(mids*hist1)
cbin1 = np.arange(-10, 0, 0.1) #cold wake bins
bins=cbin1
x= np.reshape(subset.coldwake_max.data,(pdim))
v = np.reshape(subset.coldwake_max.data,(pdim))
hist1,mids1=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum1=stats.binned_statistic(x,v, 'sum', bins)[0]
cbin2 = np.arange(0,8) #day to max, plot histogram of when cold wake max happens
bins=cbin2
x= np.reshape(subset.coldwake_hrtomaxcold.data/24,(pdim))
v = np.reshape(subset.coldwake_max.data,(pdim))
hist2,mids2=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum2=stats.binned_statistic(x,v, 'sum', bins)[0]
cbin3 = np.arange(0,50) #dy to recovery
bins=cbin3
x= np.reshape(subset.coldwake_dytorecovery.data,(pdim))
v = np.reshape(subset.coldwake_max.data,(pdim))
hist3,mids3=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum3=stats.binned_statistic(x,v, 'sum', bins)[0]
cbin4 = np.arange(0,400,10) #max cold wake as function of MLD at start of storm
bins=cbin4
x= np.reshape(subset.dbss_obml[0,:,:].data,(pdim))
v = np.reshape(subset.coldwake_max.data,(pdim))
hist4,mids4=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum4=stats.binned_statistic(x,v, 'sum', bins)[0]
cbin4a = np.arange(0,400,10) #max cold wake as function of MLD at start of storm
bins=cbin4a
x= np.reshape(subset2.mxldepth[1,:,:].data,(pdim))
v = np.reshape(subset.coldwake_max.data,(pdim))
hist4a,mids4a=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum4a=stats.binned_statistic(x,v, 'sum', bins)[0]
cbin5 = np.arange(0,200,5) #max cold wake as function of wmo max storm wind speed
bins=cbin5
x= np.reshape(subset.wmo_storm_wind.data,(pdim))
v = np.reshape(subset.coldwake_max.data,(pdim))
hist5,mids5=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum5=stats.binned_statistic(x,v, 'sum', bins)[0]
cbin6 = np.arange(0,200,5) #max cold wake as function of wmo max storm translation speed
bins=cbin6
x= np.reshape(subset.wmo_storm_speed_kmhr.data,(pdim))
v = np.reshape(subset.coldwake_max.data,(pdim))
hist6,mids6=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum6=stats.binned_statistic(x,v, 'sum', bins)[0]
cbin7 = np.arange(-10,50,1) #cold wake recovery as function of time
bins = cbin7
x= np.reshape(subset.tdif_dy.data,(pdim3))
v = np.reshape(subset.sst_anomaly.data,(pdim3))
hist7,mids7=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum7=stats.binned_statistic(x,v, 'sum', bins)[0]
x1= np.reshape(subset.wmo_storm_speed_kmhr.data,(pdim))
x3= np.reshape(subset.wmo_storm_wind.data,(pdim))
b1= cbin6 #np.arange(0,200,1)
b3= cbin5 #np.arange(0,200,1)
dbins=np.vstack((b1,b3)).T
v = np.reshape(subset.coldwake_max.data,(pdim))
hist8=stats.binned_statistic_2d(x1,x3,v,'count', bins=dbins.T)[0]
sum8=stats.binned_statistic_2d(x1,x3,v, 'sum', bins=dbins.T)[0]
x1= np.reshape(subset.wmo_storm_speed_kmhr.data,(pdim))
x2= np.reshape(subset.dbss_obml[0,:,:].data,(pdim))
x3= np.reshape(subset.wmo_storm_wind.data,(pdim))
x=np.vstack((x1,x2,x3))
b1= cbin6 #np.arange(0,200,1)
b2= cbin4 #np.arange(0,600,3)
b3= cbin5 #np.arange(0,200,1)
dbins=np.vstack((b1,b2,b3)).T
v = np.reshape(subset.coldwake_max.data,(pdim))
hist9=stats.binned_statistic_dd(x.T,v,'count', bins=dbins.T)[0]
sum9=stats.binned_statistic_dd(x.T,v, 'sum', bins=dbins.T)[0]
if init_data == 0:
sv_sum1,sv_cnt1,sv_bin1 = sum1,hist1,cbin1
sv_sum2,sv_cnt2,sv_bin2 = sum2,hist2,cbin2
sv_sum3,sv_cnt3,sv_bin3 = sum3,hist3,cbin3
sv_sum4,sv_cnt4,sv_bin4 = sum4,hist4,cbin4
sv_sum4a,sv_cnt4a,sv_bin4a = sum4a,hist4a,cbin4a
sv_sum5,sv_cnt5,sv_bin5 = sum5,hist5,cbin5
sv_sum6,sv_cnt6,sv_bin6 = sum6,hist6,cbin6
sv_sum7,sv_cnt7,sv_bin7 = sum7,hist7,cbin7
sv_sum8,sv_cnt8 = sum8,hist8
sv_sum9,sv_cnt9 = sum9,hist9
init_data=1
else:
sv_sum1+= sum1
sv_cnt1+= hist1
sv_sum2+= sum2
sv_cnt2+= hist2
sv_sum3+= sum3
sv_cnt3+= hist3
sv_sum4+= sum4
sv_cnt4+= hist4
sv_sum4a+= sum4a
sv_cnt4a+= hist4a
sv_sum5+= sum5
sv_cnt5+= hist5
sv_sum6+= sum6
sv_cnt6+= hist6
sv_sum7+= sum7
sv_cnt7+= hist7
sv_sum8+= sum8
sv_cnt8+= hist8
sv_sum9+= sum9
sv_cnt9+= hist9
#put on global map
tem = subset.coldwake_max.interp(lat=map_lats,lon=map_lons)
tem=tem.fillna(0)
temc=(tem/tem).fillna(0)
map_sum+=tem
map_cnt+=temc
map_max=np.where(tem.data < map_max, tem,map_max) #where tem<max put tem value in otherwise leave max
tem = subset.coldwake_dytorecovery.interp(lat=map_lats,lon=map_lons)
tem=tem.fillna(0)
temc=(tem/tem).fillna(0)
map_sum_recov+=tem
map_cnt_recov+=temc
m1=xr.DataArray(map_sum, coords={'lat': map_lats, 'lon':map_lons}, dims=('lat', 'lon'))
m2=xr.DataArray(map_cnt, coords={'lat': map_lats, 'lon':map_lons}, dims=('lat', 'lon'))
m3=xr.DataArray(map_max, coords={'lat': map_lats, 'lon':map_lons}, dims=('lat', 'lon'))
m4=xr.DataArray(map_sum_recov, coords={'lat': map_lats, 'lon':map_lons}, dims=('lat', 'lon'))
m5=xr.DataArray(map_cnt_recov, coords={'lat': map_lats, 'lon':map_lons}, dims=('lat', 'lon'))
ds=xr.Dataset(data_vars={'sum1': (('coldw'),sv_sum1),
'cnt1': (('coldw'),sv_cnt1),
'sum2': (('dymax'),sv_sum2),
'cnt2': (('dymax'),sv_cnt2),
'sum3': (('dyrec'),sv_sum3),
'cnt3': (('dyrec'),sv_cnt3),
'sum4': (('mld'),sv_sum4),
'cnt4': (('mld'),sv_cnt4),
'sum4a': (('mld2'),sv_sum4a),
'cnt4a': (('mld2'),sv_cnt4a),
'sum5': (('wnd'),sv_sum5),
'cnt5': (('wnd'),sv_cnt5),
'sum6': (('tspd'),sv_sum6),
'cnt6': (('tspd'),sv_cnt6),
'sum7': (('dtime'),sv_sum7),
'cnt7': (('dtime'),sv_cnt7),
'sum8': (('tspd','wnd'),sv_sum8),
'cnt8': (('tspd','wnd'),sv_cnt8),
'sum9': (('tspd','mld','wnd'),sv_sum9),
'cnt9': (('tspd','mld','wnd'),sv_cnt9),
'map_sum': (('lat','lon'),m1),
'map_cnt': (('lat','lon'),m2),
'map_max': (('lat','lon'),m3),
'map_sum_recov': (('lat','lon'),m4),
'map_cnt_recov': (('lat','lon'),m5)
},
coords={'coldw':cbin1[0:-1],
'dymax':cbin2[0:-1],
'dyrec':cbin3[0:-1],
'mld':cbin4[0:-1],
'wnd':cbin5[0:-1],
'tspd':cbin6[0:-1],
'dtime':cbin7[0:-1],
'lat':map_lats,'lon':map_lons})
# filename='f:/data/tc_wakes/database/results/hist_sum_'+str(iyr_storm)+'.nc'
# ds.to_netcdf(filename)
ds.map_sum.plot()
(ds_all2.dbss_obml[4,:,:]-ds_all2.dbss_obml[10,:,:]).plot()
(ds_all2.mxldepth[4,:,:]-ds_all2.mxldepth[10,:,:]).plot()
# coding: utf-8
# In[1]:
#trying to rewrite so faster using numpy historgram
#plotting and data analysis for global cold wakes
#from netCDF4 import Dataset # http://code.google.com/p/netcdf4-python/
import os
import time
import datetime as dt
import xarray as xr
from datetime import datetime
import pandas
import matplotlib.pyplot as plt
import numpy as np
import math
import geopy.distance
from math import sin, pi
from scipy import interpolate
from scipy import stats
#functions for running storm data
import sys
####################you will need to change some paths here!#####################
#list of input directories
dir_storm_info='f:/data/tc_wakes/database/info/'
dir_out='f:/data/tc_wakes/database/sst/'
#################################################################################
#start to look at data and make some pdfs
date_1858 = dt.datetime(1858,11,17,0,0,0) # start date is 11/17/1958
map_lats=np.arange(-90,90,.25)
map_lons=np.arange(-180,180,.25)
imap_lats = map_lats.size
imap_lons = map_lons.size
#for iyr_storm in range(2002,2017):
for iyr_storm in range(2002,2003): #2002,2018):
#init arrays
init_data=0
map_sum,map_cnt,map_max = np.zeros([imap_lats,imap_lons]),np.zeros([imap_lats,imap_lons]),np.zeros([imap_lats,imap_lons])
map_sum_recov,map_cnt_recov = np.zeros([imap_lats,imap_lons]),np.zeros([imap_lats,imap_lons])
# for inum_storm in range(0,100):
for inum_storm in range(16,20):
filename = dir_out + str(iyr_storm) + '/' + str(inum_storm).zfill(3) + '_interpolated_track.nc'
exists = os.path.isfile(filename)
if not exists:
continue
print(filename)
ds_storm_info=xr.open_dataset(filename)
ds_storm_info = ds_storm_info.sel(j2=0)
ds_storm_info.close()
filename = dir_out + str(iyr_storm) + '/' + str(inum_storm).zfill(3) + '_combined_data.nc'
ds_all = xr.open_dataset(filename)
ds_all['spd']=np.sqrt(ds_all.uwnd**2+ds_all.vwnd**2)
ds_all.close()
filename = dir_out + str(iyr_storm) + '/' + str(inum_storm).zfill(3) + '_MLD_data_v2.nc'
ds_all2 = xr.open_dataset(filename)
ds_all2.close()
if abs(ds_all.lon[-1]-ds_all.lon[0])>180:
ds_all.coords['lon'] = np.mod(ds_all['lon'], 360)
ds_storm_info['lon'] = np.mod(ds_storm_info['lon'], 360)
max_lat = ds_storm_info.lat.max()
#remove all data outsice 100km/800km or cold wake >0 or <-10
if max_lat<0:
cond = ((((ds_all.dist_from_storm_km<100) & (ds_all.side_of_storm<=0)) |
((ds_all.dist_from_storm_km<800) & (ds_all.side_of_storm>0)))
& (ds_all.coldwake_max<=-.1) & (ds_all.coldwake_max>=-10))
else:
cond = ((((ds_all.dist_from_storm_km<800) & (ds_all.side_of_storm<0)) |
((ds_all.dist_from_storm_km<100) & (ds_all.side_of_storm>=0)))
& (ds_all.coldwake_max<=-.1) & (ds_all.coldwake_max>=-10))
subset = ds_all.where(cond)
subset2 = ds_all2.where(cond)
#create coldwake anomaly with nan for all values before wmo storm time
subset['sst_anomaly']=subset.analysed_sst-subset.sst_prestorm
# for i in range(0,xdim):
# for j in range (0,ydim):
# if np.isnan(subset.closest_storm_index[j,i]):
# continue
# iend = subset.closest_storm_index[j,i].data.astype(int)
# subset.sst_anomaly[:iend,j,i]=np.nan
#create array with day.frac since closest storm passage
tdif_dy = (subset.time-subset.closest_storm_time_np64)/np.timedelta64(1, 'D')
# tdif_dy = tdif_dy.where(tdif_dy>=0,np.nan)
subset['tdif_dy']=tdif_dy
xdim,ydim,tdim = ds_all.lon.shape[0],ds_all.lat.shape[0],ds_all.time.shape[0]
#only keep sst_anomaly from 5 days before storm to cold wake recovery
ds_all['sst_anomaly']=ds_all.analysed_sst-ds_all.analysed_sst_clim-ds_all.sst_prestorm_clim
for i in range(ydim):
for j in range(xdim):
if np.isnan(ds_all.coldwake_max[i,j]):
continue
istart = int(ds_all.closest_storm_index[i,j].data-5)
if istart<0:
istart=0
iend = int(ds_all.closest_storm_index[i,j].data+ds_all.coldwake_dytorecovery[i,j].data)
if iend>tdim:
iend=tdim
ds_all.sst_anomaly[:istart,i,j]=np.nan
ds_all.sst_anomaly[iend:,i,j]=np.nan
cond = ((((ds_all.dist_from_storm_km<800) & (ds_all.side_of_storm<0)) |
((ds_all.dist_from_storm_km<100) & (ds_all.side_of_storm>=0)))
& (ds_all.coldwake_max<=-.1) & (ds_all.coldwake_max>=-10))
subset_cold = ds_all.where(cond)
#end only keep sst_anomaly code
pdim=xdim*ydim
pdim3=tdim*xdim*ydim
print(xdim*ydim)
# data = subset.coldwake_max
# cbin1 = np.arange(-10, 0, 0.1) #cold wake bins
# bins=cbin1
# hist1,mids = np.histogram(data,bins)[0],0.5*(bins[1:]+bins[:-1])
# sum1 = np.cumsum(mids*hist1)
cbin1 = np.arange(-10, 0, 0.1) #cold wake bins
bins=cbin1
x= np.reshape(subset.coldwake_max.data,(pdim))
v = np.reshape(subset.coldwake_max.data,(pdim))
hist1,mids1=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum1=stats.binned_statistic(x,v, 'sum', bins)[0]
cbin2 = np.arange(0,8) #day to max, plot histogram of when cold wake max happens
bins=cbin2
x= np.reshape(subset.coldwake_hrtomaxcold.data/24,(pdim))
v = np.reshape(subset.coldwake_max.data,(pdim))
hist2,mids2=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum2=stats.binned_statistic(x,v, 'sum', bins)[0]
cbin3 = np.arange(0,50) #dy to recovery
bins=cbin3
x= np.reshape(subset.coldwake_dytorecovery.data,(pdim))
v = np.reshape(subset.coldwake_max.data,(pdim))
hist3,mids3=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum3=stats.binned_statistic(x,v, 'sum', bins)[0]
cbin4 = np.arange(0,400,10) #max cold wake as function of MLD at start of storm
bins=cbin4
x= np.reshape(subset.dbss_obml[0,:,:].data,(pdim))
v = np.reshape(subset.coldwake_max.data,(pdim))
hist4,mids4=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum4=stats.binned_statistic(x,v, 'sum', bins)[0]
cbin4a = np.arange(0,400,10) #max cold wake as function of MLD at start of storm
bins=cbin4a
x= np.reshape(subset2.mxldepth[1,:,:].data,(pdim))
v = np.reshape(subset.coldwake_max.data,(pdim))
hist4a,mids4a=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum4a=stats.binned_statistic(x,v, 'sum', bins)[0]
cbin5 = np.arange(0,200,5) #max cold wake as function of wmo max storm wind speed
bins=cbin5
x= np.reshape(subset.wmo_storm_wind.data,(pdim))
v = np.reshape(subset.coldwake_max.data,(pdim))
hist5,mids5=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum5=stats.binned_statistic(x,v, 'sum', bins)[0]
cbin6 = np.arange(0,200,5) #max cold wake as function of wmo max storm translation speed
bins=cbin6
x= np.reshape(subset.wmo_storm_speed_kmhr.data,(pdim))
v = np.reshape(subset.coldwake_max.data,(pdim))
hist6,mids6=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum6=stats.binned_statistic(x,v, 'sum', bins)[0]
cbin7 = np.arange(-10,50,1) #cold wake recovery as function of time
bins = cbin7
x= np.reshape(tdif_dy.data,(pdim3))
v = np.reshape(subset_cold.sst_anomaly.data,(pdim3))
x = x[~np.isnan(v)]
v = v[~np.isnan(v)]
hist7,mids7=stats.binned_statistic(x,v,'count', bins)[0],0.5*(bins[1:]+bins[:-1])
sum7=stats.binned_statistic(x,v, 'sum', bins)[0]
plt.plot(bins[0:-1],sum7/hist7)
plt.plot(x,v)
#ds_all.sst_prestorm_clim.plot()
#((ds_all.analysed_sst[15,:,:]-ds_all.analysed_sst_clim[15,:,:])-ds_all.sst_prestorm_clim ).plot()
#((ds_all.analysed_sst[15,:,:]-ds_all.sst_prestorm_clim) ).plot()
#((ds_all.analysed_sst[30,:,:]-ds_all.analysed_sst_clim[30,:,:])).plot()
#plt.plot(ds_all.analysed_sst[40,40,50]-ds_all.analysed_sst_clim[40,40,50])-ds_all.sst_prestorm_clim[40,50])
#ds_all.sst_anomaly[15,:,:].plot()
ds_all.coldwake_maxindex.plot()
plt.pcolormesh(tdif_dy[20,:,:])
plt.plot(tdif_dy[:,60,60],subset.sst_anomaly[:,60,60])
ds_all.coldwake_dytorecovery.plot()
cond = ((((ds_all.dist_from_storm_km<800) & (ds_all.side_of_storm<0)) |
((ds_all.dist_from_storm_km<100) & (ds_all.side_of_storm>=0)))
& (ds_all.coldwake_max<=-.1) & (ds_all.coldwake_max>=-10))
subset_cold = ds_all.where(cond)
# tdif_dy = (subset.time-subset.closest_storm_time_np64)/np.timedelta64(1, 'D')
subset_cold.sst_anomaly[45,:,:].plot()
tdif_dy[45,:,:].plot()
j
subset_cold
```
| github_jupyter |
# Collaboration and Competition
---
In this notebook, you will learn how to use the Unity ML-Agents environment for the third project of the [Deep Reinforcement Learning Nanodegree](https://www.udacity.com/course/deep-reinforcement-learning-nanodegree--nd893) program.
### 1. Start the Environment
We begin by importing the necessary packages. If the code cell below returns an error, please revisit the project instructions to double-check that you have installed [Unity ML-Agents](https://github.com/Unity-Technologies/ml-agents/blob/master/docs/Installation.md) and [NumPy](http://www.numpy.org/).
```
from unityagents import UnityEnvironment
import gym
import random
import torch
import numpy as np
from collections import deque
import matplotlib.pyplot as plt
%matplotlib inline
from ddpg_agent import Agent
```
Next, we will start the environment! **_Before running the code cell below_**, change the `file_name` parameter to match the location of the Unity environment that you downloaded.
- **Mac**: `"path/to/Tennis.app"`
- **Windows** (x86): `"path/to/Tennis_Windows_x86/Tennis.exe"`
- **Windows** (x86_64): `"path/to/Tennis_Windows_x86_64/Tennis.exe"`
- **Linux** (x86): `"path/to/Tennis_Linux/Tennis.x86"`
- **Linux** (x86_64): `"path/to/Tennis_Linux/Tennis.x86_64"`
- **Linux** (x86, headless): `"path/to/Tennis_Linux_NoVis/Tennis.x86"`
- **Linux** (x86_64, headless): `"path/to/Tennis_Linux_NoVis/Tennis.x86_64"`
For instance, if you are using a Mac, then you downloaded `Tennis.app`. If this file is in the same folder as the notebook, then the line below should appear as follows:
```
env = UnityEnvironment(file_name="Tennis.app")
```
```
env = UnityEnvironment(file_name="./Tennis_Windows_x86_64/Tennis.exe")
```
Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.
```
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
```
### 2. Examine the State and Action Spaces
In this environment, two agents control rackets to bounce a ball over a net. If an agent hits the ball over the net, it receives a reward of +0.1. If an agent lets a ball hit the ground or hits the ball out of bounds, it receives a reward of -0.01. Thus, the goal of each agent is to keep the ball in play.
The observation space consists of 8 variables corresponding to the position and velocity of the ball and racket. Two continuous actions are available, corresponding to movement toward (or away from) the net, and jumping.
Run the code cell below to print some information about the environment.
```
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents
num_agents = len(env_info.agents)
print('Number of agents:', num_agents)
# size of each action
action_size = brain.vector_action_space_size
print('Size of each action:', action_size)
# examine the state space
states = env_info.vector_observations
state_size = states.shape[1]
print('There are {} agents. Each observes a state with length: {}'.format(states.shape[0], state_size))
print('The state for the first agent looks like:', states[0])
```
### 3. Take Random Actions in the Environment
In the next code cell, you will learn how to use the Python API to control the agents and receive feedback from the environment.
Once this cell is executed, you will watch the agents' performance, if they select actions at random with each time step. A window should pop up that allows you to observe the agents.
Of course, as part of the project, you'll have to change the code so that the agents are able to use their experiences to gradually choose better actions when interacting with the environment!
```
# for i in range(1, 6): # play game for 5 episodes
# env_info = env.reset(train_mode=False)[brain_name] # reset the environment
# states = env_info.vector_observations # get the current state (for each agent)
# scores = np.zeros(num_agents) # initialize the score (for each agent)
# while True:
# actions = np.random.randn(num_agents, action_size) # select an action (for each agent)
# actions = np.clip(actions, -1, 1) # all actions between -1 and 1
# env_info = env.step(actions)[brain_name] # send all actions to tne environment
# next_states = env_info.vector_observations # get next state (for each agent)
# rewards = env_info.rewards # get reward (for each agent)
# dones = env_info.local_done # see if episode finished
# scores += env_info.rewards # update the score (for each agent)
# states = next_states # roll over states to next time step
# if np.any(dones): # exit loop if episode finished
# break
# print('Score (max over agents) from episode {}: {}'.format(i, np.max(scores)))
# instantiate the agent
agent = Agent(num_agents=num_agents, state_size=state_size, action_size=action_size, random_seed=0)
def ddpg(n_episodes=10000, max_t=1000):
scores_deque = deque(maxlen=100)
scores = []
for i_episode in range(1, n_episodes+1):
env_info = env.reset(train_mode=True)[brain_name] # reset the environment
states = env_info.vector_observations # get the current state (for each agent)
score = np.zeros(num_agents) # initialize the score (for each agent)
for t in range(max_t):
actions = agent.act(states)
env_info = env.step(actions)[brain_name] # send all actions to tne environment
next_states = env_info.vector_observations # get next state (for each agent)
rewards = env_info.rewards # get reward (for each agent)
dones = env_info.local_done # see if episode finished
agent.step(states, actions, rewards, next_states, dones)
score += rewards # update the score (for each agent)
states = next_states # roll over states to next time step
if np.any(dones):
break
scores_deque.append(np.mean(score))
scores.append(np.mean(score))
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_deque)), end="")
if i_episode % 100 == 0:
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_deque)))
if np.mean(scores_deque)>=0.5:
print('\nEnvironment solved in {:d} episodes!\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_deque)))
torch.save(agent.actor_local.state_dict(), 'checkpoint_actor.pth')
torch.save(agent.critic_local.state_dict(), 'checkpoint_critic.pth')
break
return scores
scores = ddpg()
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(1, len(scores)+1), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
```
When finished, you can close the environment.
```
env.close()
```
### 4. It's Your Turn!
Now it's your turn to train your own agent to solve the environment! When training the environment, set `train_mode=True`, so that the line for resetting the environment looks like the following:
```python
env_info = env.reset(train_mode=True)[brain_name]
```
| github_jupyter |
# smFRET Analysis
This notebook is for simple analysis of smFRET data, starting with an hdf5 file and ending with a FRET efficiency histogram that can be fitted with a gaussians. Burst data can be exported as a .csv for analysis elsewhere.
You can analysis uncorrected data if you are simply looking for relative changes in the conformational ensemble, or accurate FRET correction parameters can be supplied if you want FRET efficiencies that can be converted to distances.
# Import packages
```
from fretbursts import *
sns = init_notebook()
import lmfit
import phconvert
import os
import matplotlib.pyplot as plt
import matplotlib as mpl
from fretbursts.burstlib_ext import burst_search_and_gate
```
# Name and Load in data
First name the data file and check it exists, it will look for the file starting from wherever this notebook is saved.
```
filename = "definitiveset/1cx.hdf5"
if os.path.isfile(filename):
print("File found")
else:
print("File not found, check file name is correct")
```
Load in the file and set correction factors. If you aren't using accurate correction factors, set d.leakage and d.dir_ex to 0 and d.gamma to 1
You may get warnings that some parameters are not defined in the file, this is fine as they will be defined in this workbook anyway
```
d = loader.photon_hdf5(filename)
for i in range(0, len(d.ph_times_t)): #sorting code
indices = d.ph_times_t[i].argsort()
d.ph_times_t[i], d.det_t[i] = d.ph_times_t[i][indices], d.det_t[i][indices]
d.leakage = 0.081 #alpha
d.dir_ex = 0.076 #delta
d.gamma = 0.856
d.beta = 0.848
#d.leakage = 0.
#d.dir_ex = 0.
#d.gamma = 1.
```
# Check alternation cycle is correct
We need to check that the ALEX parameters defined in the HDF5 file are appropriate for the laser cycle used in the experiment. If this is correct, the following histogram should look correct. It is a combined plot of every photon that arrives over the supplied alternation periods.
```
bpl.plot_alternation_hist(d)
```
IF THE ABOVE HISTOGRAM LOOKS CORRECT: then run loader.alex_apply_period, which rewrites the time stamps into groups based on their excitation period. If you want to change the alternation period after this you will have to reload the data into FRET bursts.
IF THE ABOVE HISTOGRAM LOOKS WRONG: then the supplied alternation parameters do not match up to the alternation of the lasers in the data. This could be because the lasers were actually on a different alternation, or because the data set doesn't start at zero so is frame shifted etc.
In this case, you can un-hash the code below and alter the parameters manually.
```
#d.add(det_donor_accept = (0, 1),
# alex_period = 10000,
# offset = 0,
# D_ON = (0, 4500),
# A_ON = (5000, 9500))
loader.alex_apply_period(d)
time = d.time_max
print('Total data time = %s'%time)
```
The following will plot a time trace of the first second of your experiment.
```
dplot(d, timetrace, binwidth=1e-3, tmin=0, tmax=15, figsize=(8,5))
plt.xlim(0,1);
plt.ylim(-45,45);
plt.ylabel(" Photons/ms in APD0 Photons/ms in APD1");
```
# Background Estimation
Background estimation works by plotting log of photons by the delay between them, assuming a poisson distribution of photon arrivals and fitting a line. The plot will contain single molecule bursts however, so a threshold (in microseconds) has to be defined where the fit begins.
The variable "time_s" defines the size of the windows in which background is recalculated. Lower values will make the experiment more accurately sensisitive to fluctuations in background however higher values will give more photons with which to calculate a more precise average background. If the fit fails you may need to increase this value.
```
threshold = 1500
d.calc_bg(bg.exp_fit, time_s=300, tail_min_us=(threshold),)
dplot(d, hist_bg, show_fit=True)
```
This code will plot the calculated background in each window and acts as a good reporter of whether anything major has happened to the solution over the time course of the experiment
```
dplot(d, timetrace_bg);
```
# Burst Searching and Selecting
"d.burst_search()" can be used to do an all photon burst search (APBS), however "burst_search_and_gate(d)" will apply a DCBS / dual channel burst search (Nir 2006), this effectively does independent searches in the DD+DA channel and the AA channel, and then returns the intersecton of these bursts, ensuring that any FRET information is only included whilst an acceptor is still active in the detection volume.
The two numbers given to "F=" are the signal to background threshold in the DD+DA and AA channels respectively. If your background is particularly high in one but not the other you may want to change these independently.
```
bursts=burst_search_and_gate(d, F=(20, 20), m=10, mute=True)
```
The following will plot a graph of burst number vs sizes which can inform your selection thresholding.
```
sizes = bursts.burst_sizes_ich(add_naa=True)
plt.hist(sizes, cumulative=-1, bins = 30, range = (0, 100), histtype="stepfilled", density=False)
plt.xlabel('Burst size (n photons)')
plt.ylabel("N bursts with > n")
```
We can now set thresholds on how many photons we want in each burst, this can be done on all channels together, or just one channel. Thresholding DD+DA will reduce the width in E, thresholding in AA will ensure there are no donor only bursts.
```
bursts = bursts.select_bursts(select_bursts.size, add_naa=True, th1=20,) #all channels
bursts = bursts.select_bursts(select_bursts.size, th1=50) #DD + DA
bursts = bursts.select_bursts(select_bursts.naa , th1=10) #AA
```
# Histograms
Now we can start plotting and fitting the data
```
g=alex_jointplot(bursts)
```
This will fit a gaussian to the E values.
If you set pdf=True then the data will be displayed as a probability density function, pdf=False will give it as bursts instead
```
model = mfit.factory_gaussian()
model.set_param_hint('center', value=1.1, vary=True)
bursts.E_fitter.fit_histogram(model=model, verbose=False, pdf=False)
dplot(bursts, hist_fret, pdf=False, show_model=True, show_fit_value=True, fit_from='center');
bursts.E_fitter.params
```
This will export the data to a .csv file, type the save location between the ""'s. This csv file can be opened in excel or origin and contains information about each burst, most importantly E and S but also things like burst length and width.
```
csvfile = "1cx.csv"
burstmatrix = bext.burst_data(bursts)
burstmatrix.to_csv(csvfile)
```
| github_jupyter |
# Training Neural Networks
The network we built in the previous part isn't so smart, it doesn't know anything about our handwritten digits. Neural networks with non-linear activations work like universal function approximators. There is some function that maps your input to the output. For example, images of handwritten digits to class probabilities. The power of neural networks is that we can train them to approximate this function, and basically any function given enough data and compute time.
<img src="assets/function_approx.png" width=500px>
At first the network is naive, it doesn't know the function mapping the inputs to the outputs. We train the network by showing it examples of real data, then adjusting the network parameters such that it approximates this function.
To find these parameters, we need to know how poorly the network is predicting the real outputs. For this we calculate a **loss function** (also called the cost), a measure of our prediction error. For example, the mean squared loss is often used in regression and binary classification problems
$$
\large \ell = \frac{1}{2n}\sum_i^n{\left(y_i - \hat{y}_i\right)^2}
$$
where $n$ is the number of training examples, $y_i$ are the true labels, and $\hat{y}_i$ are the predicted labels.
By minimizing this loss with respect to the network parameters, we can find configurations where the loss is at a minimum and the network is able to predict the correct labels with high accuracy. We find this minimum using a process called **gradient descent**. The gradient is the slope of the loss function and points in the direction of fastest change. To get to the minimum in the least amount of time, we then want to follow the gradient (downwards). You can think of this like descending a mountain by following the steepest slope to the base.
<img src='assets/gradient_descent.png' width=350px>
## Backpropagation
For single layer networks, gradient descent is straightforward to implement. However, it's more complicated for deeper, multilayer neural networks like the one we've built. Complicated enough that it took about 30 years before researchers figured out how to train multilayer networks.
Training multilayer networks is done through **backpropagation** which is really just an application of the chain rule from calculus. It's easiest to understand if we convert a two layer network into a graph representation.
<img src='assets/backprop_diagram.png' width=550px>
In the forward pass through the network, our data and operations go from bottom to top here. We pass the input $x$ through a linear transformation $L_1$ with weights $W_1$ and biases $b_1$. The output then goes through the sigmoid operation $S$ and another linear transformation $L_2$. Finally we calculate the loss $\ell$. We use the loss as a measure of how bad the network's predictions are. The goal then is to adjust the weights and biases to minimize the loss.
To train the weights with gradient descent, we propagate the gradient of the loss backwards through the network. Each operation has some gradient between the inputs and outputs. As we send the gradients backwards, we multiply the incoming gradient with the gradient for the operation. Mathematically, this is really just calculating the gradient of the loss with respect to the weights using the chain rule.
$$
\large \frac{\partial \ell}{\partial W_1} = \frac{\partial L_1}{\partial W_1} \frac{\partial S}{\partial L_1} \frac{\partial L_2}{\partial S} \frac{\partial \ell}{\partial L_2}
$$
**Note:** I'm glossing over a few details here that require some knowledge of vector calculus, but they aren't necessary to understand what's going on.
We update our weights using this gradient with some learning rate $\alpha$.
$$
\large W^\prime_1 = W_1 - \alpha \frac{\partial \ell}{\partial W_1}
$$
The learning rate $\alpha$ is set such that the weight update steps are small enough that the iterative method settles in a minimum.
## Losses in PyTorch
Let's start by seeing how we calculate the loss with PyTorch. Through the `nn` module, PyTorch provides losses such as the cross-entropy loss (`nn.CrossEntropyLoss`). You'll usually see the loss assigned to `criterion`. As noted in the last part, with a classification problem such as MNIST, we're using the softmax function to predict class probabilities. With a softmax output, you want to use cross-entropy as the loss. To actually calculate the loss, you first define the criterion then pass in the output of your network and the correct labels.
Something really important to note here. Looking at [the documentation for `nn.CrossEntropyLoss`](https://pytorch.org/docs/stable/nn.html#torch.nn.CrossEntropyLoss),
> This criterion combines `nn.LogSoftmax()` and `nn.NLLLoss()` in one single class.
>
> The input is expected to contain scores for each class.
This means we need to pass in the raw output of our network into the loss, not the output of the softmax function. This raw output is usually called the *logits* or *scores*. We use the logits because softmax gives you probabilities which will often be very close to zero or one but floating-point numbers can't accurately represent values near zero or one ([read more here](https://docs.python.org/3/tutorial/floatingpoint.html)). It's usually best to avoid doing calculations with probabilities, typically we use log-probabilities.
```
import torch
from torch import nn
import torch.nn.functional as F
from torchvision import datasets, transforms
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,)),
])
# Download and load the training data
trainset = datasets.MNIST('~/.pytorch/MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
```
### Note
If you haven't seen `nn.Sequential` yet, please finish the end of the Part 2 notebook.
```
# Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10))
# Define the loss
criterion = nn.CrossEntropyLoss()
# Get our data
images, labels = next(iter(trainloader))
# Flatten images
images = images.view(images.shape[0], -1)
# Forward pass, get our logits
logits = model(images)
# Calculate the loss with the logits and the labels
loss = criterion(logits, labels)
print(loss)
```
In my experience it's more convenient to build the model with a log-softmax output using `nn.LogSoftmax` or `F.log_softmax` ([documentation](https://pytorch.org/docs/stable/nn.html#torch.nn.LogSoftmax)). Then you can get the actual probabilities by taking the exponential `torch.exp(output)`. With a log-softmax output, you want to use the negative log likelihood loss, `nn.NLLLoss` ([documentation](https://pytorch.org/docs/stable/nn.html#torch.nn.NLLLoss)).
>**Exercise:** Build a model that returns the log-softmax as the output and calculate the loss using the negative log likelihood loss. Note that for `nn.LogSoftmax` and `F.log_softmax` you'll need to set the `dim` keyword argument appropriately. `dim=0` calculates softmax across the rows, so each column sums to 1, while `dim=1` calculates across the columns so each row sums to 1. Think about what you want the output to be and choose `dim` appropriately.
```
# TODO: Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
# TODO: Define the loss
criterion = nn.NLLLoss()
### Run this to check your work
# Get our data
images, labels = next(iter(trainloader))
# Flatten images
images = images.view(images.shape[0], -1)
# Forward pass, get our logits
logits = model(images)
# Calculate the loss with the logits and the labels
loss = criterion(logits, labels)
print(loss)
```
## Autograd
Now that we know how to calculate a loss, how do we use it to perform backpropagation? Torch provides a module, `autograd`, for automatically calculating the gradients of tensors. We can use it to calculate the gradients of all our parameters with respect to the loss. Autograd works by keeping track of operations performed on tensors, then going backwards through those operations, calculating gradients along the way. To make sure PyTorch keeps track of operations on a tensor and calculates the gradients, you need to set `requires_grad = True` on a tensor. You can do this at creation with the `requires_grad` keyword, or at any time with `x.requires_grad_(True)`.
You can turn off gradients for a block of code with the `torch.no_grad()` content:
```python
x = torch.zeros(1, requires_grad=True)
>>> with torch.no_grad():
... y = x * 2
>>> y.requires_grad
False
```
Also, you can turn on or off gradients altogether with `torch.set_grad_enabled(True|False)`.
The gradients are computed with respect to some variable `z` with `z.backward()`. This does a backward pass through the operations that created `z`.
```
x = torch.randn(2,2, requires_grad=True)
print(x)
y = x**2
print(y)
```
Below we can see the operation that created `y`, a power operation `PowBackward0`.
```
## grad_fn shows the function that generated this variable
print(y.grad_fn)
```
The autograd module keeps track of these operations and knows how to calculate the gradient for each one. In this way, it's able to calculate the gradients for a chain of operations, with respect to any one tensor. Let's reduce the tensor `y` to a scalar value, the mean.
```
z = y.mean()
print(z)
```
You can check the gradients for `x` and `y` but they are empty currently.
```
print(x.grad)
```
To calculate the gradients, you need to run the `.backward` method on a Variable, `z` for example. This will calculate the gradient for `z` with respect to `x`
$$
\frac{\partial z}{\partial x} = \frac{\partial}{\partial x}\left[\frac{1}{n}\sum_i^n x_i^2\right] = \frac{x}{2}
$$
```
z.backward()
print(x.grad)
print(x/2)
```
These gradients calculations are particularly useful for neural networks. For training we need the gradients of the cost with respect to the weights. With PyTorch, we run data forward through the network to calculate the loss, then, go backwards to calculate the gradients with respect to the loss. Once we have the gradients we can make a gradient descent step.
## Loss and Autograd together
When we create a network with PyTorch, all of the parameters are initialized with `requires_grad = True`. This means that when we calculate the loss and call `loss.backward()`, the gradients for the parameters are calculated. These gradients are used to update the weights with gradient descent. Below you can see an example of calculating the gradients using a backwards pass.
```
# Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
images, labels = next(iter(trainloader))
images = images.view(images.shape[0], -1)
logits = model(images)
loss = criterion(logits, labels)
print('Before backward pass: \n', model[0].weight.grad)
loss.backward()
print('After backward pass: \n', model[0].weight.grad)
```
## Training the network!
There's one last piece we need to start training, an optimizer that we'll use to update the weights with the gradients. We get these from PyTorch's [`optim` package](https://pytorch.org/docs/stable/optim.html). For example we can use stochastic gradient descent with `optim.SGD`. You can see how to define an optimizer below.
```
from torch import optim
# Optimizers require the parameters to optimize and a learning rate
optimizer = optim.SGD(model.parameters(), lr=0.01)
```
Now we know how to use all the individual parts so it's time to see how they work together. Let's consider just one learning step before looping through all the data. The general process with PyTorch:
* Make a forward pass through the network
* Use the network output to calculate the loss
* Perform a backward pass through the network with `loss.backward()` to calculate the gradients
* Take a step with the optimizer to update the weights
Below I'll go through one training step and print out the weights and gradients so you can see how it changes. Note that I have a line of code `optimizer.zero_grad()`. When you do multiple backwards passes with the same parameters, the gradients are accumulated. This means that you need to zero the gradients on each training pass or you'll retain gradients from previous training batches.
```
print('Initial weights - ', model[0].weight)
images, labels = next(iter(trainloader))
images.resize_(64, 784)
# Clear the gradients, do this because gradients are accumulated
optimizer.zero_grad()
# Forward pass, then backward pass, then update weights
output = model(images)
loss = criterion(output, labels)
loss.backward()
print('Gradient -', model[0].weight.grad)
# Take an update step and few the new weights
optimizer.step()
print('Updated weights - ', model[0].weight)
```
### Training for real
Now we'll put this algorithm into a loop so we can go through all the images. Some nomenclature, one pass through the entire dataset is called an *epoch*. So here we're going to loop through `trainloader` to get our training batches. For each batch, we'll doing a training pass where we calculate the loss, do a backwards pass, and update the weights.
>**Exercise:** Implement the training pass for our network. If you implemented it correctly, you should see the training loss drop with each epoch.
```
## Your solution here
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
optimizer = optim.SGD(model.parameters(), lr=0.003)
epochs = 5
for e in range(epochs):
running_loss = 0
for images, labels in trainloader:
# Flatten MNIST images into a 784 long vector
images = images.view(images.shape[0], -1)
# TODO: Training pass
optimizer.zero_grad()
output = model(images)
loss = criterion(output, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
else:
print(f"Training loss: {running_loss/len(trainloader)}")
```
With the network trained, we can check out it's predictions.
```
%matplotlib inline
import helper
images, labels = next(iter(trainloader))
img = images[0].view(1, 784)
# Turn off gradients to speed up this part
with torch.no_grad():
logps = model(img)
# Output of the network are log-probabilities, need to take exponential for probabilities
ps = torch.exp(logps)
helper.view_classify(img.view(1, 28, 28), ps)
```
Now our network is brilliant. It can accurately predict the digits in our images. Next up you'll write the code for training a neural network on a more complex dataset.
| github_jupyter |
# Verzweigung
#### Marcel Lüthi, Departement Mathematik und Informatik, Universität Basel
### If-Anweisung

### Anweisungsblöcke
Anweisungsblöcke sind geklammerte Folgen von Anweisungen:
```
{
Anweisung1;
Anweisung2;
...
Anweisung3;
}
```
``then`` und ``else``-Zweig in der ``if``-Anweisung entsprechen jeweils Anweisungsblöcken.
### Randbemerkung: Einrückungen
> Anweisungen in Anweisungsblöcken sollten eingerückt werden
<div style="display: block; margin-top:0.0cm">
<div style="display: inline-block">
Gut
<pre><code class="language-java" data-trim>
if (n != 0) {
n = n / 2;
n--;
} else {
n = n * 2;
n++;
}
</code></pre></div>
<div style="display: inline-block">
Schlecht
<pre><code class="language-java" data-trim>
if (n != 0) {
n = n / 2;
n--;
} else {
n = n * 2;
n++;
}
</code></pre></div>
</div>
* Einrückungen nicht wichtig für Java - aber für den Leser
### Vergleichsoperatoren
Vergleich zweier Werte liefert wahr (``true``) oder falsch (``false``)
| | Bedeutung | Beispiel |
|------|-----------|----------|
| == | gleich | x == 3 |
| != | ungleich | x != y |
| > | grösser | 4 > 3 |
| < | kleiner | x + 1 < 0 |
| >= | grösser oder gleich | x >= y|
| <= | kleiner oder gleich | x <= y|
Wird z.B. in If-Anweisung verwendet
#### Miniübung
* Vervollständigen Sie das Programm, so dass jeweils nur die zutreffende Aussage für die Zahl z ausgegeben wird.
```
int z = 5;
System.out.println("z ist eine gerade, positive Zahl");
System.out.println("z ist eine gerade, negative Zahl");
System.out.println("z ist eine ungerade, positive Zahl");
System.out.println("z ist eine ungerade, negative Zahl");
```
### Zusammengesetzte Vergleiche
Und (&&) und Oder (||) Verknüpfung
| x | y | x ``&&`` y| x | | y |
|---|---|--------|----------------|
| true | true | true | true |
| true | false | false | true |
| false | true | false | true |
| false | false | false | false |
! Nicht-Verknüpfung
| x | !x |
|---|---|
| true | false |
| false | true |
Beispiel:
```java
if (x >= 0 && x <= 10 || x >= 100 && x <= 110) {
x = y;
}
```
### Datentyp boolean
Datentyp wie ``int``, aber mit nur zwei Werten ``true`` und ``false``.
Beispiel:
```
int x = 1;
boolean p = false;
boolean q = x > 0;
p = p || q && x < 10
```
#### Beachte
* Boolesche Werte können mit &&, || und ! verknüpft werden.
* Jeder Vergleich liefert einen Wert vom Typ boolean.
* Boolesche Werte können in boolean- Variablen abgespeichert werden ("flags").
* Namen für boolean- Variablen sollten mit Adjektiv beginnen: equal, full.
# Übungen
### Übung 1: Maximum dreier Zahlen
* Schreiben Sie ein Programm, welches das Maximum dreier Zahlen berechnet.
* Schreiben Sie das Programm jeweils mit einfachen als auch mit zusammengesetzten Bedingungen

### Übung 2: Parametrisieren Sie das gezeichnete Haus
In längeren Programmen kommt es häufig vor, dass sich eine komplexe Anweisungsfolge nur in kleinen Teilen unterscheidet. Dies ist in folgendem Programm illustriert, wo wir mal wieder die Turtle Grafik verwenden.
* Führen Sie boolsche Variablen ```hasWindow``` und ``hasChimney`` ein, welche es erlauben ein Haus wahlweise mit Kamin, mit Fenster oder mit beidem zu zeichnen.
* Führen Sie Variablen ein, um die Zeichnung einfacher parametrisieren zu können.
```
// Laden der Turtle Bibliothek
// Diese Kommandos funktionieren nur in Jupyter-notebooks und entsprechen nicht gültigem Java.
%mavenRepo bintray https://dl.bintray.com/egp/maven
%maven ch.unibas.informatik:jturtle:0.5
import ch.unibas.informatik.jturtle.Turtle;
Turtle t = new Turtle();
// head
t.home();
t.penDown();
t.backward(50);
t.forward(50);
t.turnRight(45);
t.forward(50);
t.turnRight(90);
t.forward(20);
t.turnLeft(135);
t.forward(10);
t.turnRight(90);
t.forward(10);
t.turnRight(90);
t.forward(20);
t.turnLeft(45);
t.forward(15);
t.turnRight(45);
t.forward(50);
t.turnRight(90);
t.forward(70);
t.toImage();
```
#### Mini Übung:
* Fügen Sie eine Verzweigung ein, die ein Fenster zeichnet, wenn eine Variable ```hasWindow``` auf true gesetzt ist.
* Führen Sie Variablen ein, um die Zeichnung zu parametrisieren (Höhe/Breite des Hauses, etc.)
| github_jupyter |
```
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from sklearn.datasets import load_breast_cancer
from sklearn.preprocessing import StandardScaler
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (8,10)
class CancelOut(keras.layers.Layer):
'''
CancelOut layer, keras implementation.
'''
def __init__(self, activation='sigmoid', cancelout_loss=True, lambda_1=0.002, lambda_2=0.001):
super(CancelOut, self).__init__()
self.lambda_1 = lambda_1
self.lambda_2 = lambda_2
self.cancelout_loss = cancelout_loss
if activation == 'sigmoid': self.activation = tf.sigmoid
if activation == 'softmax': self.activation = tf.nn.softmax
def build(self, input_shape):
self.w = self.add_weight(
shape=(input_shape[-1],),
initializer=tf.keras.initializers.Constant(1),
trainable=True,
)
def call(self, inputs):
if self.cancelout_loss:
self.add_loss( self.lambda_1 * tf.norm(self.w, ord=1) + self.lambda_2 * tf.norm(self.w, ord=2))
return tf.math.multiply(inputs, self.activation(self.w))
def get_config(self):
return {"activation": self.activation}
def plot_importance(importances):
indices = np.argsort(importances)
plt.title('Feature Importances')
plt.barh(range(len(indices)), importances[indices], color='b', align='center')
plt.yticks(range(len(indices)), [features[i] for i in indices])
plt.xlabel('Relative Importance')
plt.show()
scaler = StandardScaler()
X = scaler.fit_transform(load_breast_cancer()['data'])
y = load_breast_cancer()['target']
features = load_breast_cancer()['feature_names']
```
### Sigmoid + Loss
```
inputs = keras.Input(shape=(X.shape[1],))
x = CancelOut(activation='sigmoid')(inputs)
x = layers.Dense(32, activation="relu")(x)
#x = CancelOut()(x)
x = layers.Dense(32, activation="relu")(x)
#x = CancelOut()(x)
outputs = layers.Dense(1, activation='sigmoid')(x)
model = keras.Model(inputs=inputs, outputs=outputs)
model.compile(loss='binary_crossentropy',
optimizer='adam')
model.summary()
model.fit(X, y, epochs=20, batch_size=8)
cancelout_feature_importance_sigmoid = model.get_weights()[0]
cancelout_feature_importance_sigmoid
```
### Softmax + No Loss
```
inputs = keras.Input(shape=(X.shape[1],))
x = CancelOut(activation='softmax', cancelout_loss=False)(inputs)
x = layers.Dense(32, activation="relu")(x)
#x = CancelOut()(x)
x = layers.Dense(32, activation="relu")(x)
#x = CancelOut()(x)
outputs = layers.Dense(1, activation='sigmoid')(x)
model = keras.Model(inputs=inputs, outputs=outputs)
model.compile(loss='binary_crossentropy',
optimizer='adam')
model.summary()
model.fit(X, y, epochs=20, batch_size=8)
cancelout_feature_importance_softmax = model.get_weights()[0]
cancelout_feature_importance_softmax
```
### RandomForest Feature Importance
```
from sklearn.ensemble import RandomForestClassifier
rnd_clf = RandomForestClassifier(n_estimators=500, n_jobs=-1, random_state=42).fit(X,y)
rf_importances = rnd_clf.feature_importances_
```
### Plots
```
print('Sigmoid + Loss')
plot_importance(cancelout_feature_importance_sigmoid)
print('Softmax + No Loss')
plot_importance(cancelout_feature_importance_softmax)
print("Random Forest")
plot_importance(rf_importances)
```
| github_jupyter |
# Collapse trees to shared clades for comparison
The script prunes and collapses two or more trees to a shared set of clades. Note that the goals are "**clades**" rather than "**taxa**". A shared clade is defined as a clade present in all trees, with the same set of descendants. The resulting trees can then be used for back-to-back comparison (e.g., using [Dendroscope](http://dendroscope.org/)'s tanglegram function).
Finding a shared set of clades is not trivial. In computer science it is a special form of the [covering problem](https://en.wikipedia.org/wiki/Covering_problems). While exploring the optimal solution is beyond the scope of this study, I designed and implemented a working heuristic to achieve the goal.
```
import scipy as sp
import matplotlib.pyplot as plt
from skbio import TreeNode
```
Number of clades to retain.
```
clades_to_retain = 50
```
Minimum number of descendants for a retained clade.
```
clade_min_size = 50
trees = [
TreeNode.read('astral.cons.nid.e5p68057.nwk'),
TreeNode.read('concat.cons.nid.b50.nwk')
]
node2taxa = []
for tree in trees:
n2t = {}
for node in tree.postorder():
if node.is_tip():
n2t[node.name] = [node.name]
else:
n2t[node.name] = []
for child in node.children:
n2t[node.name].extend(n2t[child.name])
node2taxa.append(n2t)
for i in range(len(node2taxa)):
node2taxa[i] = {k: v for k, v in node2taxa[i].items() if len(v) >= clade_min_size}
for i in range(len(node2taxa)):
for node in node2taxa[i]:
node2taxa[i][node] = set(node2taxa[i][node])
matches = {}
for node1, taxa1 in node2taxa[0].items():
for node2, taxa2 in node2taxa[1].items():
if taxa1 == taxa2:
matches[','.join((node1, node2))] = taxa1
print('Matching pairs: %d.' % len(matches))
sp.special.comb(len(matches), clades_to_retain)
```
Here is my heuristic for the covering problem. It starts at a given size of clade: *k*, and progressively recruits clades from left and right, until the desired number of clades: *n*, is reached.
```
universe = set().union(*[v for k, v in matches.items()])
total_size = len(universe)
print('Taxa under matching pairs: %d.' % total_size)
mean_clade_size = round(len(universe) / clades_to_retain)
mean_clade_size
def select_clades(start_size):
res = []
for match, taxa in sorted(matches.items(), key=lambda x: abs(start_size - len(x[1]))):
is_unique = True
for m in res:
if not taxa.isdisjoint(matches[m]):
is_unique = False
break
if is_unique is True:
res.append(match)
if len(res) == clades_to_retain:
break
return res
```
Test multiple starting sizes.
```
x, y, z = [], [], []
selections = []
start_size = mean_clade_size
while True:
selected_matches = select_clades(start_size)
if len(selected_matches) < clades_to_retain:
break
selections.append(selected_matches)
covered = set().union(*[matches[x] for x in selected_matches])
cv = sp.stats.variation([len(matches[x]) for x in selected_matches])
x.append(start_size)
y.append(len(covered) / total_size)
z.append(cv)
print('Starting at %d, covering %d taxa, CV = %.3f' % (start_size, len(covered), cv))
start_size += 1
```
Plot them out.
```
plt.plot(x, y, 'r')
plt.plot(x, z, 'b')
plt.xlabel('start size')
plt.text(400, 0.55, '% taxa covered', color='r', ha='right')
plt.text(400, 0.52, 'coefficient of variation', color='b', ha='right');
```
The final starting size *k* is manually determined based on the plot.
```
start_size = 348
selected_matches = select_clades(start_size)
for m in selected_matches:
print('%s: %d taxa.' % (m, len(matches[m])))
```
Write selected clades.
```
with open('selected_matches.txt', 'w') as f:
for m in selected_matches:
f.write('%s\n' % m.replace(',', '\t'))
```
Export collapsed trees. Each selected clade becomes a tip, with its height equaling the median of its descendants.
```
def get_clade_height(node):
return np.median([x.accumulate_to_ancestor(node) for x in node.tips()])
def collapse_clades(tree, clades):
tcopy = tree.copy()
clades = set(clades)
nodes_to_remove = []
for node in tcopy.non_tips():
if node.name in clades:
node.length += get_clade_height(node)
nodes_to_remove.extend(node.children)
tcopy.remove_deleted(lambda x: x in nodes_to_remove)
tcopy = tcopy.shear(clades)
tcopy.prune()
return tcopy
for i, tree in enumerate(trees):
clades = [x.split(',')[i] for x in selected_matches]
tcopy = collapse_clades(tree, clades)
tcopy.write('tree%d.nwk' % (i + 1))
for tip in tcopy.tips():
tip.name = 'X%d' % (clades.index(tip.name) + 1)
tcopy.write('tree%d.trans.nwk' % (i + 1))
```
| github_jupyter |
# Deep Learning and Transfer Learning with pre-trained models
This notebook uses a pretrained model to build a classifier (CNN)
```
# import required libs
import os
import keras
import numpy as np
from keras import backend as K
from keras import applications
from keras.datasets import cifar10
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
import matplotlib.pyplot as plt
params = {'legend.fontsize': 'x-large',
'figure.figsize': (15, 5),
'axes.labelsize': 'x-large',
'axes.titlesize':'x-large',
'xtick.labelsize':'x-large',
'ytick.labelsize':'x-large'}
plt.rcParams.update(params)
%matplotlib inline
```
## Load VGG
```
vgg_model = applications.VGG19(include_top=False, weights='imagenet')
vgg_model.summary()
```
Set Parameters
```
batch_size = 128
num_classes = 10
epochs = 50
bottleneck_path = r'F:\work\kaggle\cifar10_cnn\bottleneck_features_train_vgg19.npy'
```
## Get CIFAR10 Dataset
```
# the data, shuffled and split between train and test sets
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
y_train.shape
```
## Pretrained Model for Feature Extraction
```
if not os.path.exists(bottleneck_path):
bottleneck_features_train = vgg_model.predict(x_train,verbose=1)
np.save(open(bottleneck_path, 'wb'),
bottleneck_features_train)
else:
bottleneck_features_train = np.load(open(bottleneck_path,'rb'))
bottleneck_features_train[0].shape
bottleneck_features_test = vgg_model.predict(x_test,verbose=1)
```
## Custom Classifier
```
clf_model = Sequential()
clf_model.add(Flatten(input_shape=bottleneck_features_train.shape[1:]))
clf_model.add(Dense(512, activation='relu'))
clf_model.add(Dropout(0.5))
clf_model.add(Dense(256, activation='relu'))
clf_model.add(Dropout(0.5))
clf_model.add(Dense(num_classes, activation='softmax'))
```
## Visualize the network architecture
```
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
SVG(model_to_dot(clf_model, show_shapes=True,
show_layer_names=True, rankdir='TB').create(prog='dot', format='svg'))
```
## Compile the model
```
clf_model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
```
## Train the classifier
```
clf_model.fit(bottleneck_features_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1)
```
## Predict and test model performance
```
score = clf_model.evaluate(bottleneck_features_test, y_test, verbose=1)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
```
### Assign label to a test image
```
def predict_label(img_idx,show_proba=True):
plt.imshow(x_test[img_idx],aspect='auto')
plt.title("Image to be Labeled")
plt.show()
print("Actual Class:{}".format(np.nonzero(y_test[img_idx])[0][0]))
test_image =np.expand_dims(x_test[img_idx], axis=0)
bf = vgg_model.predict(test_image,verbose=0)
pred_label = clf_model.predict_classes(bf,batch_size=1,verbose=0)
print("Predicted Class:{}".format(pred_label[0]))
if show_proba:
print("Predicted Probabilities")
print(clf_model.predict_proba(bf))
img_idx = 3999 # sample indices : 999,1999 and 3999
for img_idx in [999,1999,3999]:
predict_label(img_idx)
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Feature columns visualization
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github.com/tensorflow/examples/blob/master/community/en/hashing_trick.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/examples/tree/master/community/en/hashing_trick.ipynb">
<img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
View source on GitHub</a>
</td>
</table>
This example demonstrates the use `tf.feature_column.crossed_column` on some simulated Atlanta housing price data.
This spatial data is used primarily so the results can be easily visualized.
These functions are designed primarily for categorical data, not to build interpolation tables.
If you actually want to build smart interpolation tables in TensorFlow you may want to consider [TensorFlow Lattice](https://research.googleblog.com/2017/10/tensorflow-lattice-flexibility.html).
## Setup
```
import os
import subprocess
import tempfile
import tensorflow as tf
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rcParams['figure.figsize'] = 12, 6
mpl.rcParams['image.cmap'] = 'viridis'
```
## Build Synthetic Data
```
# Define the grid
min_latitude = 33.641336
max_latitude = 33.887157
delta_latitude = max_latitude-min_latitude
min_longitude = -84.558798
max_longitude = -84.287259
delta_longitude = max_longitude-min_longitude
resolution = 100
# Use RandomState so the behavior is repeatable.
R = np.random.RandomState(1)
# The price data will be a sum of Gaussians, at random locations.
n_centers = 20
centers = R.rand(n_centers, 2) # shape: (centers, dimensions)
# Each Gaussian has a maximum price contribution, at the center.
# Price_
price_delta = 0.5+2*R.rand(n_centers)
# Each Gaussian also has a standard-deviation and variance.
std = 0.2*R.rand(n_centers) # shape: (centers)
var = std**2
def price(latitude, longitude):
# Convert latitude, longitude to x,y in [0,1]
x = (longitude - min_longitude)/delta_longitude
y = (latitude - min_latitude)/delta_latitude
# Cache the shape, and flatten the inputs.
shape = x.shape
assert y.shape == x.shape
x = x.flatten()
y = y.flatten()
# Convert x, y examples into an array with shape (examples, dimensions)
xy = np.array([x,y]).T
# Calculate the square distance from each example to each center.
components2 = (xy[:,None,:] - centers[None,:,:])**2 # shape: (examples, centers, dimensions)
r2 = components2.sum(axis=2) # shape: (examples, centers)
# Calculate the z**2 for each example from each center.
z2 = r2/var[None,:]
price = (np.exp(-z2)*price_delta).sum(1) # shape: (examples,)
# Restore the original shape.
return price.reshape(shape)
# Build the grid. We want `resolution` cells between `min` and `max` on each dimension
# so we need `resolution+1` evenly spaced edges. The centers are at the average of the
# upper and lower edge.
latitude_edges = np.linspace(min_latitude, max_latitude, resolution+1)
latitude_centers = (latitude_edges[:-1] + latitude_edges[1:])/2
longitude_edges = np.linspace(min_longitude, max_longitude, resolution+1)
longitude_centers = (longitude_edges[:-1] + longitude_edges[1:])/2
latitude_grid, longitude_grid = np.meshgrid(
latitude_centers,
longitude_centers)
# Evaluate the price at each center-point
actual_price_grid = price(latitude_grid, longitude_grid)
price_min = actual_price_grid.min()
price_max = actual_price_grid.max()
price_mean = actual_price_grid.mean()
price_mean
def show_price(price):
plt.imshow(
price,
# The color axis goes from `price_min` to `price_max`.
vmin=price_min, vmax=price_max,
# Put the image at the correct latitude and longitude.
extent=(min_longitude, max_longitude, min_latitude, max_latitude),
# Make the image square.
aspect = 1.0*delta_longitude/delta_latitude)
show_price(actual_price_grid)
```
## Build Datasets
```
# For test data we will use the grid centers.
test_features = {'latitude':latitude_grid.flatten(), 'longitude':longitude_grid.flatten()}
test_ds = tf.data.Dataset.from_tensor_slices((test_features,
actual_price_grid.flatten()))
test_ds = test_ds.cache().batch(512).prefetch(1)
# For training data we will use a set of random points.
train_latitude = min_latitude + np.random.rand(50000)*delta_latitude
train_longitude = min_longitude + np.random.rand(50000)*delta_longitude
train_price = price(train_latitude, train_longitude)
train_features = {'latitude':train_latitude, 'longitude':train_longitude}
train_ds = tf.data.Dataset.from_tensor_slices((train_features, train_price))
train_ds = train_ds.cache().shuffle(100000).batch(512).prefetch(1)
```
## Generate a plot from an Estimator
```
ag = actual_price_grid.reshape(resolution, resolution)
ag.shape
def plot_model(model, ds = test_ds):
# Create two plot axes
actual, predicted = plt.subplot(1,2,1), plt.subplot(1,2,2)
# Plot the actual price.
plt.sca(actual)
plt.pcolor(actual_price_grid.reshape(resolution, resolution))
# Generate predictions over the grid from the estimator.
pred = model.predict(ds)
# Convert them to a numpy array.
pred = np.fromiter((item for item in pred), np.float32)
# Plot the predictions on the secodn axis.
plt.sca(predicted)
plt.pcolor(pred.reshape(resolution, resolution))
```
## A linear regressor
```
# Use `normalizer_fn` so that the model only sees values in [0, 1]
norm_latitude = lambda latitude:(latitude-min_latitude)/delta_latitude - 0.5
norm_longitude = lambda longitude:(longitude-min_longitude)/delta_longitude - 0.5
linear_fc = [tf.feature_column.numeric_column('latitude', normalizer_fn = norm_latitude),
tf.feature_column.numeric_column('longitude', normalizer_fn = norm_longitude)]
# Build and train the model
model = tf.keras.Sequential([
tf.keras.layers.DenseFeatures(feature_columns=linear_fc),
tf.keras.layers.Dense(1),
])
model.compile(optimizer=tf.keras.optimizers.Adam(), loss=tf.keras.losses.MeanAbsoluteError())
model.fit(train_ds, epochs=200, validation_data=test_ds)
plot_model(model)
```
## A DNN regressor
Important: Pure categorical data doesn't the spatial relationships that make this example possible. Embeddings are a way your model can learn spatial relationships.
```
# Build and train the model
model = tf.keras.Sequential([
tf.keras.layers.DenseFeatures(feature_columns=linear_fc),
tf.keras.layers.Dense(100, activation='elu'),
tf.keras.layers.Dense(100, activation='elu'),
tf.keras.layers.Dense(1),
])
model.compile(optimizer=tf.keras.optimizers.Adam(), loss=tf.keras.losses.MeanAbsoluteError())
model.fit(train_ds, epochs=200, validation_data=test_ds)
plot_model(model)
```
# Linear model with buckets
```
# Bucketize the latitude and longitude usig the `edges`
latitude_bucket_fc = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column('latitude'),
list(latitude_edges))
longitude_bucket_fc = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column('longitude'),
list(longitude_edges))
seperable_fc = [
latitude_bucket_fc,
longitude_bucket_fc]
# Build and train the model
model = tf.keras.Sequential([
tf.keras.layers.DenseFeatures(feature_columns=seperable_fc),
tf.keras.layers.Dense(1),
])
model.compile(optimizer=tf.keras.optimizers.Adam(), loss=tf.keras.losses.MeanAbsoluteError())
model.fit(train_ds, epochs=200, validation_data=test_ds)
plot_model(model)
```
# Using `crossed_column` on its own.
```
# Cross the bucketized columns, using 5000 hash bins (for an average weight sharing of 2).
crossed_lat_lon_fc = tf.feature_column.crossed_column(
[latitude_bucket_fc, longitude_bucket_fc], 2000)
crossed_lat_lon_fc = tf.feature_column.indicator_column(crossed_lat_lon_fc)
crossed_fc = [crossed_lat_lon_fc]
# Build and train the model
model = tf.keras.Sequential([
tf.keras.layers.DenseFeatures(feature_columns=crossed_fc),
tf.keras.layers.Dense(1),
])
model.compile(optimizer=tf.keras.optimizers.Adam(), loss=tf.keras.losses.MeanAbsoluteError())
model.fit(train_ds, epochs=200, validation_data=test_ds)
plot_model(model)
```
# Using raw categories with `crossed_column`
The model generalizes better if it also has access to the raw categories, outside of the cross.
```
# Build and train the model
model = tf.keras.Sequential([
tf.keras.layers.DenseFeatures(feature_columns=crossed_fc+seperable_fc),
tf.keras.layers.Dense(1, kernel_regularizer=tf.keras.regularizers.l2(0.0001)),
])
model.compile(optimizer=tf.keras.optimizers.Adam(),
loss=tf.keras.losses.MeanAbsoluteError(),
metrics=[tf.keras.losses.MeanAbsoluteError()])
model.fit(train_ds, epochs=200, validation_data=test_ds)
plot_model(model)
```
| github_jupyter |
# Example 1b: Spin-Bath model (Underdamped Case)
### Introduction
The HEOM method solves the dynamics and steady state of a system and its environment, the latter of which is encoded in a set of auxiliary density matrices.
In this example we show the evolution of a single two-level system in contact with a single Bosonic environment. The properties of the system are encoded in Hamiltonian, and a coupling operator which describes how it is coupled to the environment.
The Bosonic environment is implicitly assumed to obey a particular Hamiltonian (see paper), the parameters of which are encoded in the spectral density, and subsequently the free-bath correlation functions.
In the example below we show how to model the underdamped Brownian motion Spectral Density.
### Drude-Lorentz (overdamped) spectral density
Note that in the above, and the following, we set $\hbar = k_\mathrm{B} = 1$.
### Brownian motion (underdamped) spectral density
The underdamped spectral density is:
$$J_U = \frac{\alpha^2 \Gamma \omega}{(\omega_c^2 - \omega^2)^2 + \Gamma^2 \omega^2)}.$$
Here $\alpha$ scales the coupling strength, $\Gamma$ is the cut-off frequency, and $\omega_c$ defines a resonance frequency. With the HEOM we must use an exponential decomposition:
The Matsubara decomposition of this spectral density is, in real and imaginary parts:
\begin{equation*}
c_k^R = \begin{cases}
\alpha^2 \coth(\beta( \Omega + i\Gamma/2)/2)/4\Omega & k = 0\\
\alpha^2 \coth(\beta( \Omega - i\Gamma/2)/2)/4\Omega & k = 0\\
-2\alpha^2\Gamma/\beta \frac{\epsilon_k }{((\Omega + i\Gamma/2)^2 + \epsilon_k^2)(\Omega - i\Gamma/2)^2 + \epsilon_k^2)} & k \geq 1\\
\end{cases}
\end{equation*}
\begin{equation*}
\nu_k^R = \begin{cases}
-i\Omega + \Gamma/2, i\Omega +\Gamma/2, & k = 0\\
{2 \pi k} / {\beta } & k \geq 1\\
\end{cases}
\end{equation*}
\begin{equation*}
c_k^I = \begin{cases}
i\alpha^2 /4\Omega & k = 0\\
-i\alpha^2 /4\Omega & k = 0\\
\end{cases}
\end{equation*}
\begin{equation*}
\nu_k^I = \begin{cases}
i\Omega + \Gamma/2, -i\Omega + \Gamma/2, & k = 0\\
\end{cases}
\end{equation*}
Note that in the above, and the following, we set $\hbar = k_\mathrm{B} = 1$.
```
%pylab inline
from qutip import *
%load_ext autoreload
%autoreload 2
from bofin.heom import BosonicHEOMSolver
def cot(x):
return 1./np.tan(x)
def coth(x):
"""
Calculates the coth function.
Parameters
----------
x: np.ndarray
Any numpy array or list like input.
Returns
-------
cothx: ndarray
The coth function applied to the input.
"""
return 1/np.tanh(x)
# Defining the system Hamiltonian
eps = .5 # Energy of the 2-level system.
Del = 1.0 # Tunnelling term
Hsys = 0.5 * eps * sigmaz() + 0.5 * Del* sigmax()
# Initial state of the system.
rho0 = basis(2,0) * basis(2,0).dag()
# System-bath coupling (Drude-Lorentz spectral density)
Q = sigmaz() # coupling operator
#solver time steps
nsteps = 1000
tlist = np.linspace(0, 50, nsteps)
#correlation function plotting time steps
tlist_corr = np.linspace(0, 20, 1000)
#Bath properties:
gamma = .1 # cut off frequency
lam = .5 # coupling strenght
w0 = 1 #resonance frequency
T = 1
beta = 1./T
#HEOM parameters
NC = 10 # cut off parameter for the bath
#Spectral Density
wlist = np.linspace(0, 5, 1000)
pref = 1.
J = [lam**2 * gamma * w / ((w0**2-w**2)**2 + (gamma**2)*(w**2)) for w in wlist]
# Plot the results
fig, axes = plt.subplots(1, 1, sharex=True, figsize=(8,8))
axes.plot(wlist, J, 'r', linewidth=2)
axes.set_xlabel(r'$\omega$', fontsize=28)
axes.set_ylabel(r'J', fontsize=28)
#first of all lets look athe correlation functions themselves
Nk = 3 # number of exponentials
Om = np.sqrt(w0**2 - (gamma/2)**2)
Gamma = gamma/2.
#mats
def Mk(t,k):
ek = 2*pi*k/beta
return (-2*lam**2*gamma/beta)*ek*exp(-ek*abs(t))/(((Om+1.0j*Gamma)**2+ek**2)*((Om-1.0j*Gamma)**2+ek**2))
def c(t):
Cr = coth(beta*(Om+1.0j*Gamma)/2)*exp(1.0j*Om*t)+coth(beta*(Om-1.0j*Gamma)/2)*exp(-1.0j*Om*t)
#Cr = coth(beta*(Om+1.0j*Gamma)/2)*exp(1.0j*Om*t)+conjugate(coth(beta*(Om+1.0j*Gamma)/2)*exp(1.0j*Om*t))
Ci = exp(-1.0j*Om*t)-exp(1.0j*Om*t)
return (lam**2/(4*Om))*exp(-Gamma*abs(t))*(Cr+Ci) + sum([Mk(t,k) for k in range(1,Nk+1)])
plt.figure(figsize=(8,8))
plt.plot(tlist_corr ,[real(c(t)) for t in tlist_corr ], '-', color="black", label="Re[C(t)]")
plt.plot(tlist_corr ,[imag(c(t)) for t in tlist_corr ], '-', color="red", label="Im[C(t)]")
plt.legend()
plt.show()
#The Matsubara terms modify the real part
Nk = 3# number of exponentials
Om = np.sqrt(w0**2 - (gamma/2)**2)
Gamma = gamma/2.
#mats
def Mk(t,k):
ek = 2*pi*k/beta
return (-2*lam**2*gamma/beta)*ek*exp(-ek*abs(t))/(((Om+1.0j*Gamma)**2+ek**2)*((Om-1.0j*Gamma)**2+ek**2))
plt.figure(figsize=(8,8))
plt.plot(tlist_corr ,[sum([real(Mk(t,k)) for k in range(1,4)]) for t in tlist_corr ], '-', color="black", label="Re[M(t)] Nk=3")
plt.plot(tlist_corr ,[sum([real(Mk(t,k)) for k in range(1,6)]) for t in tlist_corr ], '--', color="red", label="Re[M(t)] Nk=5")
plt.legend()
plt.show()
#Lets collate the parameters for the HEOM
ckAR = [(lam**2/(4*Om))*coth(beta*(Om+1.0j*Gamma)/2),(lam**2/(4*Om))*coth(beta*(Om-1.0j*Gamma)/2)]
ckAR.extend([(-2*lam**2*gamma/beta)*( 2*pi*k/beta)/(((Om+1.0j*Gamma)**2+ (2*pi*k/beta)**2)*((Om-1.0j*Gamma)**2+( 2*pi*k/beta)**2))+0.j for k in range(1,Nk+1)])
vkAR = [-1.0j*Om+Gamma,1.0j*Om+Gamma]
vkAR.extend([2 * np.pi * k * T + 0.j for k in range(1,Nk+1)])
factor=1./4.
ckAI =[-factor*lam**2*1.0j/(Om),factor*lam**2*1.0j/(Om)]
vkAI = [-(-1.0j*(Om) - Gamma),-(1.0j*(Om) - Gamma)]
NC=14
NR = len(ckAR)
NI = len(ckAI)
Q2 = [Q for kk in range(NR+NI)]
print(Q2)
options = Options(nsteps=15000, store_states=True, rtol=1e-14, atol=1e-14)
HEOM = BosonicHEOMSolver(Hsys, Q2, ckAR, ckAI, vkAR, vkAI, NC, options=options)
result = HEOM.run(rho0, tlist)
# Define some operators with which we will measure the system
# Define some operators with which we will measure the system
# 1,1 element of density matrix - corresonding to groundstate
P11p=basis(2,0) * basis(2,0).dag()
P22p=basis(2,1) * basis(2,1).dag()
# 1,2 element of density matrix - corresonding to coherence
P12p=basis(2,0) * basis(2,1).dag()
# Calculate expectation values in the bases
P11 = expect(result.states, P11p)
P22 = expect(result.states, P22p)
P12= expect(result.states, P12p)
#DL = " 2*pi* 2.0 * {lam} / (pi * {gamma} * {beta}) if (w==0) else 2*pi*(2.0*{lam}*{gamma} *w /(pi*(w**2+{gamma}**2))) * ((1/(exp((w) * {beta})-1))+1)".format(gamma=gamma, beta = beta, lam = lam)
UD = " 2* {lam}**2 * {gamma} / ( {w0}**4 * {beta}) if (w==0) else 2* ({lam}**2 * {gamma} * w /(({w0}**2 - w**2)**2 + {gamma}**2 * w**2)) * ((1/(exp((w) * {beta})-1))+1)".format(gamma = gamma, beta = beta, lam = lam, w0 = w0)
optionsODE = Options(nsteps=15000, store_states=True,rtol=1e-12,atol=1e-12)
outputBR = brmesolve(Hsys, rho0, tlist, a_ops=[[sigmaz(),UD]], options = optionsODE)
# Calculate expectation values in the bases
P11BR = expect(outputBR.states, P11p)
P22BR = expect(outputBR.states, P22p)
P12BR = expect(outputBR.states, P12p)
#Prho0BR = expect(outputBR.states,rho0)
#This Thermal state of a reaction coordinate should, at high temperatures and not to broad baths, tell us the steady-state
dot_energy, dot_state = Hsys.eigenstates()
deltaE = dot_energy[1] - dot_energy[0]
gamma2 = gamma
wa = w0 # reaction coordinate frequency
g = lam/sqrt(2*wa)
#nb = (1 / (np.exp(wa/w_th) - 1))
NRC = 10
Hsys_exp = tensor(qeye(NRC), Hsys)
Q_exp = tensor(qeye(NRC), Q)
a = tensor(destroy(NRC), qeye(2))
H0 = wa * a.dag() * a + Hsys_exp
# interaction
H1 = (g * (a.dag() + a) * Q_exp)
H = H0 + H1
#print(H.eigenstates())
energies, states = H.eigenstates()
rhoss = 0*states[0]*states[0].dag()
for kk, energ in enumerate(energies):
rhoss += (states[kk]*states[kk].dag()*exp(-beta*energies[kk]))
rhoss = rhoss/rhoss.norm()
P12RC = tensor(qeye(NRC), basis(2,0) * basis(2,1).dag())
P12RC = expect(rhoss,P12RC)
P11RC = tensor(qeye(NRC), basis(2,0) * basis(2,0).dag())
P11RC = expect(rhoss,P11RC)
matplotlib.rcParams['figure.figsize'] = (7, 5)
matplotlib.rcParams['axes.titlesize'] = 25
matplotlib.rcParams['axes.labelsize'] = 30
matplotlib.rcParams['xtick.labelsize'] = 28
matplotlib.rcParams['ytick.labelsize'] = 28
matplotlib.rcParams['legend.fontsize'] = 28
matplotlib.rcParams['axes.grid'] = False
matplotlib.rcParams['savefig.bbox'] = 'tight'
matplotlib.rcParams['lines.markersize'] = 5
matplotlib.rcParams['font.family'] = 'STIXgeneral'
matplotlib.rcParams['mathtext.fontset'] = 'stix'
matplotlib.rcParams["font.serif"] = "STIX"
matplotlib.rcParams['text.usetex'] = False
# Plot the results
fig, axes = plt.subplots(1, 1, sharex=True, figsize=(12,7))
plt.yticks([P11RC,0.6,1.0],[0.38,0.6,1])
axes.plot(tlist, np.real(P11BR), 'y--', linewidth=3, label="Bloch-Redfield")
axes.plot(tlist, np.real(P11), 'b', linewidth=3, label="Matsubara $N_k=3$")
axes.plot(tlist, [P11RC for t in tlist], color='black', linestyle="-.",linewidth=2, label="Thermal state")
axes.locator_params(axis='y', nbins=6)
axes.locator_params(axis='x', nbins=6)
axes.set_ylabel(r'$\rho_{11}$',fontsize=30)
axes.set_xlabel(r'$t \Delta$',fontsize=30)
axes.locator_params(axis='y', nbins=4)
axes.locator_params(axis='x', nbins=4)
axes.legend(loc=0)
fig.savefig("figures/fig3.pdf")
from qutip.ipynbtools import version_table
version_table()
```
| github_jupyter |
## 表格图像数据生成
```
import numpy as np
import pandas as pd
import cv2
def get_neighbor_units_direction(r_i, c_i, units_comb_directions):
# Get the direction indexes of the 4 neighbor units (left, right, up and down)
# input: r_i, the row index of the given unit
# input: c_i, the column index of the given unit
# input: units_comb_directions, the direction status matrix of the units
# return: l_direction, u_direction, r_direction, d_direction,
# the direction indexes of the neighbor units
# left
mask1 = units_comb_directions['r_i'] == r_i
mask2 = units_comb_directions['c_i'] == c_i - 1
mask = mask1 & mask2
if np.sum(mask) == 1:
l_direction = units_comb_directions.loc[mask, 'direction'].values[0]
else:
l_direction = None
# right
mask1 = units_comb_directions['r_i'] == r_i
mask2 = units_comb_directions['c_i'] == c_i + 1
mask = mask1 & mask2
if np.sum(mask) == 1:
r_direction = units_comb_directions.loc[mask, 'direction'].values[0]
else:
r_direction = None
# up
mask1 = units_comb_directions['r_i'] == r_i - 1
mask2 = units_comb_directions['c_i'] == c_i
mask = mask1 & mask2
if np.sum(mask) == 1:
u_direction = units_comb_directions.loc[mask, 'direction'].values[0]
else:
u_direction = None
# down
mask1 = units_comb_directions['r_i'] == r_i + 1
mask2 = units_comb_directions['c_i'] == c_i
mask = mask1 & mask2
if np.sum(mask) == 1:
d_direction = units_comb_directions.loc[mask, 'direction'].values[0]
else:
d_direction = None
return l_direction, u_direction, r_direction, d_direction
def check_comb_direction_OK(r_i, c_i, d, units_comb_directions):
# Check if the selected combining direction of the unit (denoted by r_i and c_i) is applicable.
# input: r_i, the row index of the given unit
# input: c_i, the column index of the given unit
# input: d, the target direction of (r_i, c_i) indexed unit
# input: units_comb_directions, the direction status matrix of the units
# return: 0, can not be combined, 1, OK to be combined
is_exist = check_comb_target_exist(r_i, c_i, d)
if is_exist == False:
return 0
# the combining direction for the given unit
# should not be vertical to the up unit nor the left unit
l_d, u_d, r_d, d_d = get_neighbor_units_direction(r_i, c_i, units_comb_directions)
if l_d is not None and l_d != 0 and np.dot(comb_v[d], comb_v[l_d]) == 0:
return 0
if u_d is not None and u_d != 0 and np.dot(comb_v[d], comb_v[u_d]) == 0:
return 0
if r_d is not None and r_d != 0 and np.dot(comb_v[d], comb_v[r_d]) == 0:
return 0
if d_d is not None and d_d != 0 and np.dot(comb_v[d], comb_v[d_d]) == 0:
return 0
return 1
def check_comb_target_exist(r_i, c_i, d):
# Check if the selected combining direction is beyond the arange of the pre-defined table rows and columns or not.
# input: r_i, the row index of the given unit
# input: c_i, the column index of the given unit
# input: d, the target direction of (r_i, c_i) indexed unit
# return: False can not be existed, True the combined target is existed
t_c_i, t_r_i, t_d = get_combined_unit(r_i, c_i, d)
if t_c_i not in np.arange(col_units):
return False
if t_r_i not in np.arange(row_units):
return False
return True
def get_combined_unit(r_i, c_i, d):
# Get the (r_i, c_i) index of the target unit, and the corresponding direction index
# if the combined is conducted.
# input: r_i, the row index of the given unit
# input: c_i, the column index of the given unit
# input: d, the target direction of (r_i, c_i) indexed unit
# return: target_unit[0] is the c_i of the target
# target_unit[1] is the r_i of the target
# t_d is the assigned direction of the target, actually is the reverse direction of the given d
target_unit = np.array([c_i, r_i]) + np.array(comb_v[d])
# t_d is the direction of the target unit of the combing
# the direction should be the reverse direction of the give d
if d == 1:
t_d = 3
elif d == 2:
t_d = 4
elif d == 3:
t_d = 1
elif d == 4:
t_d = 2
# [x , y]
# x is col index
# y is row index
return target_unit[0], target_unit[1], t_d
def generate_table(combined_records, units_pos, units_comb_directions):
# Generate a tabel by randomly combing neighbor units.
# input: units_pos, dataframe to record the r_i, c_i, l_x, l_y, r_x, r_y for each unit
# input: units_comb_directions, dataframe to record the r_i, c_i, direction for each unit
# combined_records = []
for r_i in np.arange(row_units):
unit_h = unit_h_list[r_i]
for c_i in np.arange(col_units):
unit_w = unit_w_list[c_i]
## for the constaint width and height of the unit
# x_center = int((1 + c_i * 2) * (unit_w / 2) + padding / 2)
# y_center = int((1 + r_i * 2) * (unit_h / 2) + padding / 2)
# l_x = x_center - int(unit_w / 2)
# l_y = y_center - int(unit_h / 2)
# r_x = x_center + int(unit_w / 2)
# r_y = y_center + int(unit_h / 2)
## for the randomly set widht and height of the unit
l_x = int(np.sum(unit_w_list[0:c_i]) + padding_l)
l_y = int(np.sum(unit_h_list[0:r_i]) + padding_u)
r_x = int(np.sum(unit_w_list[0:c_i]) + unit_w_list[c_i] + padding_l)
r_y = int(np.sum(unit_h_list[0:r_i]) + unit_h_list[r_i] + padding_u)
# print(l_x, l_y, r_x, r_y)
# l_x = x_center - int(unit_w / 2)
# l_y = y_center - int(unit_h / 2)
# r_x = x_center + int(unit_w / 2)
# r_y = y_center + int(unit_h / 2)
units_pos.iloc[c_i + r_i * col_units, :] = [r_i, c_i, l_x, l_y, r_x, r_y]
# !!! Attention wrong
# units_comb_directions.iloc[c_i + r_i * col_units, :] = [r_i, c_i, 0]
# select the comining direction
# 0 none, 1 right, 2 up, 3 left, 4 down
# randomly decide to combine or not
is_comb = np.random.randint(0,2)
if is_comb:
# randomly select the combining direction
direction = np.random.randint(1,5)
# check if the combining direction is OK or NOT
comb_OK = check_comb_direction_OK(r_i, c_i, direction, units_comb_directions)
# print(r_i, c_i, direction, comb_OK)
if comb_OK == 1:
# record the combined diection for the current unit
units_comb_directions.iloc[c_i + r_i * col_units, :] = [r_i, c_i, direction]
target_c_i, target_r_i, target_d = get_combined_unit(r_i, c_i, direction)
# print("=============")
# ld, ud, rd, dd = get_neighbor_units_direction(r_i, c_i)
# print(ld, ud, rd, dd)
# print(r_i, c_i, direction, '==>', target_r_i, target_c_i, target_d)
# record the combined diection for the combined unit, actually is the reverse direction of the current one
units_comb_directions.iloc[target_c_i + target_r_i * col_units, :] = [target_r_i, target_c_i, target_d]
# print(units_comb_directions)
# record the combined units index
u_a = "{0}-{1}".format(r_i, c_i)
u_b = "{0}-{1}".format(target_r_i, target_c_i)
update_combined_records(combined_records, u_a, u_b)
def update_combined_records(combined_records, u_a, u_b):
# update if u_a, u_b combining reocrd is existed in combined_records
for i in np.arange(len(combined_records)):
one = combined_records[i]
if u_a in one and u_b in one:
return combined_records
elif u_a in one:
combined_records[i].append(u_b)
return combined_records
elif u_b in one:
combined_records[i].append(u_a)
return combined_records
combined_records.append([u_a, u_b])
return combined_records
def check_if_combined(r_i, c_i, combined_records):
# Check if the given r_i, c_i unit is combined or not
u_a = "{0}-{1}".format(r_i, c_i)
for one in combined_records:
if u_a in one:
return True
return False
def get_unit_position(r_i, c_i, units_pos):
mask1 = units_pos['r_i'] == r_i
mask2 = units_pos['c_i'] == c_i
mask = mask1 & mask2
if np.sum(mask) == 1:
one_row = units_pos.loc[mask,:]
l_x, l_y, r_x, r_y = one_row['l_x'].values[0], one_row['l_y'].values[0], one_row['r_x'].values[0], one_row['r_y'].values[0]
return l_x, l_y, r_x, r_y
return None, None, None, None
def get_combined_position(one_record, units_pos):
x_s = []
y_s = []
for one in one_record:
r_i = int(one.split('-')[0])
c_i = int(one.split('-')[1])
l_x, l_y, r_x, r_y = get_unit_position(r_i, c_i,units_pos)
x_s.append(l_x)
y_s.append(l_y)
x_s.append(r_x)
y_s.append(r_y)
return np.min(x_s), np.max(y_s), np.max(x_s), np.min(y_s)
def check_txt_area_size_OK(txt_string, s_x, s_y, lx, ly, rx, ry, textSize=15):
fontStyle = ImageFont.truetype(
"Font/simsun.ttc", textSize, encoding="utf-8")
lines = txt_string.split('\n')
widths = []
height = 0
for one in lines:
(width_i, height_i), (offset_x, offset_y) = fontStyle.font.getsize(one.strip())
widths.append(width_i)
height = height + height_i
width = np.max(widths)
if (s_x - np.min([lx, rx]) + width) > (np.abs(lx - rx) + 2):
return False
elif (s_y - np.min([ly, ry]) + height) > (np.abs(ly - ry) + 2):
return False
return True
def assign_text(image, txts, lx, ly, rx, ry):
# get the text area location: s_x: left, s_y: top
# s_x = np.random.choice(np.arange(np.min([lx, rx]), np.max([lx, rx])), 1)
# s_y = np.random.choice(np.arange(np.min([ly, ry]), np.max([ly, ry])), 1)
s_x = np.min([lx, rx]) + np.random.choice([2, int(np.abs(lx - rx) / 3)])
s_y = np.min([ly, ry]) + np.random.choice([2, int(np.abs(ly - ry)/ 3)])
max_lines = int((np.abs(ly - ry) - (s_y - np.min([ly, ry]))) / 13)
try_i = 0
while(1):
txt_lines = np.random.choice([1, np.ceil(max_lines * 2 / 3)])
txt_strs = []
for i in np.arange(txt_lines):
txt_i = np.random.randint(0, len(txts))
txt_strs.append(txts[txt_i])
txt_string = "\n".join(txt_strs)
flag = check_txt_area_size_OK(txt_string, s_x, s_y, lx, ly, rx, ry)
try_i = try_i + 1
if flag == True or try_i > 10:
break
# print(txt_strs, txt_lines)
# assign txt_string, and get the text area size: w, h
image, (w, h) = cv2ImgAddText(image, txt_string, s_x, s_y, (0,0,0))
return image, txt_string, s_x, s_y, w, h
def generate_save_talbes(table_i, img_path, label_path):
# unit_h_max = np.max(unit_h_list)
# unit_w_max = np.max(unit_w_list)
# table_i = i
line_w = line_w_list[table_i]
img_h = np.sum(unit_h_list) + padding_u + padding_d
img_w = np.sum(unit_w_list) + padding_l + padding_r
img_data = np.zeros((img_h, img_w), np.uint8)
img_data.fill(255)
image = cv2.cvtColor(img_data, cv2.COLOR_GRAY2BGR)
# r_i, index of row
# c_i index of column
# l_x, l_y, left down point x, y
# r_x, r_y, right up point x, y
units_pos = pd.DataFrame(np.zeros((row_units * col_units, 6)),
columns=['r_i', 'c_i', 'l_x', 'l_y', 'r_x', 'r_y'], dtype=int)
com_dir_data = np.zeros((row_units * col_units, 3))
com_dir_data.fill(-1)
units_comb_directions = pd.DataFrame(com_dir_data,
columns=['r_i', 'c_i', 'direction'], dtype=int)
combined_records = []
# opencv table first row is at the bottom
# opencv table first col is at the left
generate_table(combined_records, units_pos, units_comb_directions)
unit_pos_final = []
for r_i in np.arange(row_units):
for c_i in np.arange(col_units):
if not check_if_combined(r_i, c_i, combined_records):
l_x, l_y, r_x, r_y = get_unit_position(r_i, c_i, units_pos)
cv2.rectangle(image, (l_x,l_y), (r_x, r_y), (0, 0, 0), line_w)
# txt_i = np.random.randint(0, len(words))
# cv2.putText(image, words[txt_i], (l_x,l_y),
# fontFace=cv2.FONT_HERSHEY_SIMPLEX, fontScale=1, color=(0, 0, 0))
# image = cv2ImgAddText(image, words[txt_i], l_x + txt_padding, l_y + txt_padding, (0,0,0))
if np.random.rand() < 0.5:
words = words_ZH
else:
words = words_EN
image, txt_string, s_x, s_y, w, h = assign_text(image, words, l_x, l_y, r_x, r_y)
# cv2.rectangle(image, (s_x,s_y), (s_x + w, s_y + h), (0, 255, 0), 1)
# top left -> bottom right
# unit_pos_final.append([l_x, l_y, r_x, r_y, words[txt_i]])
unit_txt = txt_string.replace('\n', '--')
unit_pos_final.append([l_x, l_y, r_x, r_y, line_w, s_x, s_y, w, h, unit_txt])
for one_record in combined_records:
l_x, l_y, r_x, r_y = get_combined_position(one_record, units_pos)
cv2.rectangle(image, (l_x,l_y), (r_x, r_y), (0, 0, 0), line_w)
# txt_i = np.random.randint(0, len(words))
# cv2.putText(image, words[txt_i], (l_x,l_y),
# fontFace=cv2.FONT_HERSHEY_SIMPLEX, fontScale=1, color=(0, 0, 0))
# image = cv2ImgAddText(image, words[txt_i], l_x + txt_padding, r_y + txt_padding, (0, 0, 0))
if np.random.rand() < 0.5:
words = words_ZH
else:
words = words_EN
image, txt_string, s_x, s_y, w, h = assign_text(image, words, l_x, l_y, r_x, r_y)
# cv2.rectangle(image, (s_x,s_y), (s_x + w, s_y + h), (0, 0, 255), 1)
# top left -> bottom right
# unit_pos_final.append([l_x, r_y, r_x, l_y, words[txt_i]])
unit_txt = txt_string.replace('\n', '--')
unit_pos_final.append([l_x, l_y, r_x, r_y, line_w, s_x, s_y, w, h, unit_txt])
title_words = len(unit_w_list) #np.random.randint(int(len(unit_w_list)/2), )
title_words_ids = np.random.randint(0, len(words), title_words)
title_strs = [words[i] for i in title_words_ids]
title_string = "".join(title_strs)
image, (w, h) = cv2ImgAddText(image, title_string, padding_l*4, int(padding_u / 2), (0,0,0))
# label the title text area
unit_pos_final.append([-1, -1, -1, -1, 0, padding_l*4, int(padding_u / 2), w, h, title_string])
unit_pos_final_df = pd.DataFrame(unit_pos_final)
unit_pos_final_df.to_csv(label_path + ".label")
cv2.imwrite(img_path, image)
# cv2.imshow('Table',image)
# cv2.waitKey(0)
from PIL import Image, ImageDraw, ImageFont
def cv2ImgAddText(img, text, left, top, textColor=(0, 0, 0), textSize=15):
if (isinstance(img, np.ndarray)): # 判断是否OpenCV图片类型
img = Image.fromarray(cv2.cvtColor(img, cv2.COLOR_BGR2RGB))
# 创建一个可以在给定图像上绘图的对象
draw = ImageDraw.Draw(img)
# 字体的格式
fontStyle = ImageFont.truetype(
"Font/simsun.ttc", textSize, encoding="utf-8")
# 绘制文本
draw.text((left, top), text, textColor, font=fontStyle)
lines = text.split('\n')
widths = []
height = 0
for one in lines:
(width_i, height_i), (offset_x, offset_y) = fontStyle.font.getsize(one.strip())
widths.append(width_i)
height = 15 * len(widths)
# 转换回OpenCV格式
# width, height 打印文本的区域宽、高
return cv2.cvtColor(np.asarray(img), cv2.COLOR_RGB2BGR), (np.max(widths), height)
# row_units = 5
# col_units = 10
with open("./Text_ZH/CollectedWords.txt", 'r', encoding='utf-8') as txt_f:
words = txt_f.readlines()
words_ZH = [one.strip() for one in words]
with open("./Text_EN/CollectedWords_EN.txt", 'r', encoding='utf-8') as txt_f:
words = txt_f.readlines()
words_EN = [one.strip() for one in words]
table_nums = 20000
line_w_list = np.random.randint(1, 5, table_nums)
padding_u = 50
padding_l = 20
padding_d = 20
padding_r = 20
txt_padding = 5
# 0 none, 1 right, 2 up, 3 left, 4 down
# combine dirction vectors
# x-axis: right
# y-axis: up
comb_v = {1: [1, 0],
2: [0, -1],
3: [-1, 0],
4: [0, 1]}
dataset = "./dataset04/"
img_folder = dataset + "imgs/"
label_folder = dataset + "labels/"
for i in np.arange(table_nums):
filename = "{0}.png".format(i).rjust(10, '0')
img_path = img_folder + filename
label_path = label_folder + filename
row_units = np.random.randint(3, 10)
col_units = np.random.randint(3, 10)
unit_w_list = np.random.randint(50, 150, col_units)
unit_h_list = np.random.randint(20, 100, row_units)
generate_save_talbes(i, img_path, label_path)
print("Saving table {0} to image".format(i))
```
| github_jupyter |
# Project: Linear Regression
Reggie is a mad scientist who has been hired by the local fast food joint to build their newest ball pit in the play area. As such, he is working on researching the bounciness of different balls so as to optimize the pit. He is running an experiment to bounce different sizes of bouncy balls, and then fitting lines to the data points he records. He has heard of linear regression, but needs your help to implement a version of linear regression in Python.
_Linear Regression_ is when you have a group of points on a graph, and you find a line that approximately resembles that group of points. A good Linear Regression algorithm minimizes the _error_, or the distance from each point to the line. A line with the least error is the line that fits the data the best. We call this a line of _best fit_.
We will use loops, lists, and arithmetic to create a function that will find a line of best fit when given a set of data.
## Part 1: Calculating Error
The line we will end up with will have a formula that looks like:
```
y = m*x + b
```
`m` is the slope of the line and `b` is the intercept, where the line crosses the y-axis.
Create a function called `get_y()` that takes in `m`, `b`, and `x` and returns what the `y` value would be for that `x` on that line!
```
def get_y(m, b, x):
y = m*x + b
return y
get_y(1, 0, 7) == 7
get_y(5, 10, 3) == 25
```
Reggie wants to try a bunch of different `m` values and `b` values and see which line produces the least error. To calculate error between a point and a line, he wants a function called `calculate_error()`, which will take in `m`, `b`, and an [x, y] point called `point` and return the distance between the line and the point.
To find the distance:
1. Get the x-value from the point and store it in a variable called `x_point`
2. Get the y-value from the point and store it in a variable called `y_point`
3. Use `get_y()` to get the y-value that `x_point` would be on the line
4. Find the difference between the y from `get_y` and `y_point`
5. Return the absolute value of the distance (you can use the built-in function `abs()` to do this)
The distance represents the error between the line `y = m*x + b` and the `point` given.
```
def calculate_error(m, b, point):
x_point, y_point = point
y = m*x_point + b
distance = abs(y - y_point)
return distance
```
Let's test this function!
```
#this is a line that looks like y = x, so (3, 3) should lie on it. thus, error should be 0:
print(calculate_error(1, 0, (3, 3)))
#the point (3, 4) should be 1 unit away from the line y = x:
print(calculate_error(1, 0, (3, 4)))
#the point (3, 3) should be 1 unit away from the line y = x - 1:
print(calculate_error(1, -1, (3, 3)))
#the point (3, 3) should be 5 units away from the line y = -x + 1:
print(calculate_error(-1, 1, (3, 3)))
```
Great! Reggie's datasets will be sets of points. For example, he ran an experiment comparing the width of bouncy balls to how high they bounce:
```
datapoints = [(1, 2), (2, 0), (3, 4), (4, 4), (5, 3)]
```
The first datapoint, `(1, 2)`, means that his 1cm bouncy ball bounced 2 meters. The 4cm bouncy ball bounced 4 meters.
As we try to fit a line to this data, we will need a function called `calculate_all_error`, which takes `m` and `b` that describe a line, and `points`, a set of data like the example above.
`calculate_all_error` should iterate through each `point` in `points` and calculate the error from that point to the line (using `calculate_error`). It should keep a running total of the error, and then return that total after the loop.
```
def calculate_all_error(m, b, points):
total_error = 0
for point in datapoints:
point_error = calculate_error(m, b, point)
total_error += point_error
return total_error
```
Let's test this function!
```
#every point in this dataset lies upon y=x, so the total error should be zero:
datapoints = [(1, 1), (3, 3), (5, 5), (-1, -1)]
print(calculate_all_error(1, 0, datapoints))
#every point in this dataset is 1 unit away from y = x + 1, so the total error should be 4:
datapoints = [(1, 1), (3, 3), (5, 5), (-1, -1)]
print(calculate_all_error(1, 1, datapoints))
#every point in this dataset is 1 unit away from y = x - 1, so the total error should be 4:
datapoints = [(1, 1), (3, 3), (5, 5), (-1, -1)]
print(calculate_all_error(1, -1, datapoints))
#the points in this dataset are 1, 5, 9, and 3 units away from y = -x + 1, respectively, so total error should be
# 1 + 5 + 9 + 3 = 18
datapoints = [(1, 1), (3, 3), (5, 5), (-1, -1)]
print(calculate_all_error(-1, 1, datapoints))
```
Great! It looks like we now have a function that can take in a line and Reggie's data and return how much error that line produces when we try to fit it to the data.
Our next step is to find the `m` and `b` that minimizes this error, and thus fits the data best!
## Part 2: Try a bunch of slopes and intercepts!
The way Reggie wants to find a line of best fit is by trial and error. He wants to try a bunch of different slopes (`m` values) and a bunch of different intercepts (`b` values) and see which one produces the smallest error value for his dataset.
Using a list comprehension, let's create a list of possible `m` values to try. Make the list `possible_ms` that goes from -10 to 10 inclusive, in increments of 0.1.
Hint (to view this hint, either double-click this cell or highlight the following white space): <font color="white">you can go through the values in range(-100, 100) and multiply each one by 0.1</font>
```
possible_ms = [m * 0.1 for m in range(-100, 101)]
```
Now, let's make a list of `possible_bs` to check that would be the values from -20 to 20 inclusive, in steps of 0.1:
```
possible_bs = [b * 0.1 for b in range(-200, 201)]
```
We are going to find the smallest error. First, we will make every possible `y = m*x + b` line by pairing all of the possible `m`s with all of the possible `b`s. Then, we will see which `y = m*x + b` line produces the smallest total error with the set of data stored in `datapoint`.
First, create the variables that we will be optimizing:
* `smallest_error` — this should start at infinity (`float("inf")`) so that any error we get at first will be smaller than our value of `smallest_error`
* `best_m` — we can start this at `0`
* `best_b` — we can start this at `0`
We want to:
* Iterate through each element `m` in `possible_ms`
* For every `m` value, take every `b` value in `possible_bs`
* If the value returned from `calculate_all_error` on this `m` value, this `b` value, and `datapoints` is less than our current `smallest_error`,
* Set `best_m` and `best_b` to be these values, and set `smallest_error` to this error.
By the end of these nested loops, the `smallest_error` should hold the smallest error we have found, and `best_m` and `best_b` should be the values that produced that smallest error value.
Print out `best_m`, `best_b` and `smallest_error` after the loops.
```
datapoints = [(1, 2), (2, 0), (3, 4), (4, 4), (5, 3)]
smallest_error = float("inf")
best_m = 0
best_b = 0
for m in possible_ms:
for b in possible_bs:
error = calculate_all_error(m, b, datapoints)
if error < smallest_error:
best_m = m
best_b = b
smallest_error = error
print(best_m, best_b, smallest_error)
```
## Part 3: What does our model predict?
Now we have seen that for this set of observations on the bouncy balls, the line that fits the data best has an `m` of 0.3 and a `b` of 1.7:
```
y = 0.3x + 1.7
```
This line produced a total error of 5.
Using this `m` and this `b`, what does your line predict the bounce height of a ball with a width of 6 to be?
In other words, what is the output of `get_y()` when we call it with:
* m = 0.3
* b = 1.7
* x = 6
```
get_y(0.3, 1.7, 6)
```
Our model predicts that the 6cm ball will bounce 3.5m.
Now, Reggie can use this model to predict the bounce of all kinds of sizes of balls he may choose to include in the ball pit!
| github_jupyter |
# Dataprep
### Objective
Crawls through raw_data directory and converts diffusion and flair into a data array
### Prerequisites
All diffusion and FLAIR should be registrated and put in a NIFTI file format.
### Data organisation
- All b0 diffusion should be named "patientid_hX_DWIb0.nii.gz" where "hX" corresponds to time delay and can be "h0" or "h1" (to stratify on delay)
- All b1000 diffusion should be named "patientid_hX_DWIb1000.nii.gz" where "hX" corresponds to time delay and can be "h0" or "h1" (to stratify on delay)
- All corresponding FLAIR sequences should be named: "patientid_hX_qX_FLAIR.nii.gz" where "qX" corresponds to quality and can be "q0" or "q1" or "q2" (to stratify on quality)
- Optionally, you can add a weighted mask "patientid_hX_MASK.nii.gz" with values between 0 (background), 1 (brain mask) and 2 (stroke region) that will be used for training weight. If you don't provide it, a crude stroke segmentation with ADC < 600 will be used as a weighting map.
## Load modules
```
import os, glob, h5py
import numpy as np
from skimage.morphology import dilation, opening
from modules.niftitools import twoniftis2array, flairnifti2array, masknifti2array
```
## Crawl through files
```
dwifiles_precheck = glob.glob(os.path.join("raw_data", "*_DWIb0.nii.gz"))
patnames, timepoints, qualities, b0files, b1000files, flairfiles, maskfiles = [], [], [], [], [], [], []
num_patients = 0
for dwifile in dwifiles_precheck:
name, timepoint, _ = os.path.basename(dwifile).split("_")
timepoint = int(timepoint.replace("h",""))
matchesb1000 = glob.glob(os.path.join("raw_data", name+"_h"+str(timepoint)+"_DWIb1000.nii.gz"))
matchesFlair = glob.glob(os.path.join("raw_data", name+"_h"+str(timepoint)+"_q*_FLAIR.nii.gz"))
if len(matchesFlair) and len(matchesb1000):
_, _, quality, _ = os.path.basename(matchesFlair[0]).split("_")
patnames.append(name)
timepoints.append(timepoint)
qualities.append(int(quality.replace("q","")))
b0files.append(dwifile)
b1000files.append(matchesb1000[0])
flairfiles.append(matchesFlair[0])
matchesMask = glob.glob(os.path.join("raw_data", name+"_h"+str(timepoint)+"_MASK.nii.gz"))
if len(matchesMask):
maskfiles.append(matchesMask[0])
else:
maskfiles.append(None)
num_patients += 1
```
## Create data arrays
```
z_slices = 25
outputdir = "data"
with h5py.File(os.path.join(outputdir,"metadata.hdf5"), "w") as metadata:
metadata.create_dataset("patientnames", data=np.array(patnames, dtype="S"))
metadata.create_dataset("shape_x", data=(num_patients,256,256,z_slices,3))
metadata.create_dataset("shape_y", data=(num_patients,256,256,z_slices,1))
metadata.create_dataset("shape_mask", data=(num_patients,256,256,z_slices,1))
metadata.create_dataset("shape_meta", data=(num_patients,2))
fx = np.memmap(os.path.join(outputdir,"data_x.dat"), dtype="float32", mode="w+",
shape=(num_patients,256,256,z_slices,3))
fy = np.memmap(os.path.join(outputdir,"data_y.dat"), dtype="float32", mode="w+",
shape=(num_patients,256,256,z_slices,1))
fmask = np.memmap(os.path.join(outputdir,"data_mask.dat"), dtype="uint8", mode="w+",
shape=(num_patients,256,256,z_slices,1))
fmeta = np.memmap(os.path.join(outputdir,"data_meta.dat"), dtype="float32", mode="w+",
shape=(num_patients,2))
if num_patients > 0:
print("Imported following patients:", end=" ")
for i in range(num_patients):
if i>0:
print(", ",end="")
fmeta[i,0] = qualities[i]
fmeta[i,1] = timepoints[i]
Xdata, mask, _ = twoniftis2array(b0files[i], b1000files[i],z_slices)
Xdata = Xdata.transpose(1,2,3,0)
fx[i] = Xdata
if maskfiles[i] is not None:
fmask[i] = masknifti2array(maskfiles[i],z_slices)[...,np.newaxis]
else:
crudemask = dilation(dilation(dilation(opening(np.logical_and(mask, Xdata[...,2]<600)))))
crudemask = crudemask.astype("uint8") + mask.astype("uint8")
fmask[i] = crudemask[...,np.newaxis]
fy[i] = flairnifti2array(flairfiles[i],mask,z_slices)[...,np.newaxis]
print(name, end="")
del fx, fy, fmask, fmeta
```
| github_jupyter |
# Process Mining Disease Trajectory
```
import pandas as pd
# Change the 'source' variable with the appropriate file location in your computer
source = "M:\PM_DTM\ToyData_PM_DTM.csv"
# Load the event log into a data frame
eventlog = pd.read_csv(source)
# See the event data
eventlog
# Removing the duplicate events and put them into a new data frame
# Dataframe 'filtered_eventlog' will be created
## Select and then uncomment the necessary statement below
removeduplicate = True
#removeduplicate = False
if removeduplicate == True:
filtered_eventlog = eventlog.drop_duplicates(subset= {'Subject_id', 'Diagnosis'}, keep='first')
else:
filtered_eventlog = eventlog.copy()
filtered_eventlog
# Optional
# Save the filtered event log into a .csv file
#filtered_eventlog.to_csv("M:\PM_STM\Filtered_eventlog.csv", index=False, encoding='utf-8')
```
# Graph Isomorphism Checker
```
import networkx as nx
from networkx.algorithms import isomorphism
import matplotlib.pyplot as plt
# Create the graph data from the trajectory model adapted from A. B. Jensen et al. (2014)
# (see the Figure 1 from the paper)
g_Jensen = nx.DiGraph()
g_Jensen.add_edges_from([('I20', 'I25'),
('I20', 'I48'),
('I20', 'I50'),
('I20', 'K29'),
('I21', 'I25'),
('I25', 'J18'),
('I25', 'N30'),
('I25', 'M10'),
('I48', 'M10'),
('I50', 'M10'),
('I50', 'J15'),
('K29', 'I50'),
('J15', 'M10'),])
# Visualise the trajectory model adapted from A. B. Jensen et al. (2014)
pos = nx.spring_layout(g_Jensen)
nx.draw_networkx(g_Jensen, pos)
plt.show()
# Create the graph data from the trajectory model created by process mining tool (Disco)
# (see the Figure 3 from the paper)
g_promin = nx.DiGraph()
g_promin.add_edges_from([('I20', 'K29'),
('I20', 'I48'),
('I20', 'I50'),
('I20', 'I25'),
('I25', 'J18'),
('I25', 'N30'),
('I25', 'M10'),
('I21', 'I25'),
('I50', 'J15'),
('I50', 'M10'),
('K29', 'I50'),
('I48', 'M10'),
('J15', 'M10'),])
# Visualise the trajectory model created by process mining tool (Disco)
pos = nx.spring_layout(g_promin)
nx.draw_networkx(g_promin, pos)
plt.show()
# Check whether the graphs are isomorphic
nx.is_isomorphic(g_Jensen, g_promin)
```
| github_jupyter |
## Enap - Especialização em Ciência de Dados aplicada a Políticas Públicas
# D20 - Monitoria em Ciência da Computação
## Atividade Avaliativa 01
Nessa atividade, você poderá avaliar seu progresso nos conceitos e habilidades trabalhados no primeiro ciclo da disciplina, isto é, o conteúdo de todas as aulas ministradas em 2021.
Para iniciar sua avaliação, execute a célula abaixo, que permitirá o registro de suas respostas no Dashboard de notas. Depois, responda às questões escrevendo uma função Python que atenda ao comando de cada questão. Note que cada questão é composta por três células: a primeira célula para sua resposta, a segunda célula para testes e a terceira para validação e registro da resposta.
```
#@title Célula de inicialização. Por favor, execute.
import sys, os, subprocess
from datetime import datetime
import requests
import time
from requests.utils import quote
url = 'https://docs.google.com/forms/d/e/1FAIpQLSeASSC8-w8FmfodZ4lBnuSEAvYuE4vatIBowLIREG1f-2pIpA/formResponse?usp=pp_url&entry.1986154915=mbacd&entry.513694412=2021&entry.1914621244=MonitoriaCienciaComputacao'
def format_values(values):
return {
"entry.1269959472": values['student_id'],
"entry.1799867692": str(values['exercise_number']).replace(".", "_"),
"entry.886231469": values['exercise_score'],
"entry.1491599254": values['exercise_extra'],
"entry.1342537331": values['id']
}
def send_attendance(url, data):
count = 0
while count < 3:
count += 1
try:
r = requests.post(url, data=data)
break
except:
print("Error Occured!")
time.sleep(2)
os.system('pip install virtualenv')
def validate(func, inputs, outfunc, outputs, exercise_number, exercise_extra):
answers_status = True
outputs = [True for x in inputs] if outputs == None else outputs
for k, v in zip(inputs, outputs):
ans = func(*k)
result = None
try:
result = outfunc(ans, k) == v
if not result:
answers_status = False
print(f"Resposta incorreta. {func.__name__}({k}) deveria ser {v}, mas retornou {ans}")
except ValueError:
pass
if not result.all():
answers_status = False
print(f"Resposta incorreta. {func.__name__}({k}) deveria ser {v}, mas retornou {ans}")
if answers_status:
student_email=!gcloud config get-value account
if not student_email or 'unset' in student_email[0]:
!gcloud auth login
student_email=!gcloud config get-value account
exercise_score = True
values = {"exercise_number": exercise_number, "student_id": student_email[0],
"exercise_points": 1, "exercise_score": exercise_score,
"exercise_extra": exercise_extra,
"id": f"{student_email[0]}_{exercise_number}"}
final_data = format_values(values)
send_attendance(f"{url}&emailAddress={quote(str(student_email[0]))}", final_data)
print("Parabéns")
```
### Questão 01
Crie uma função que retorne o string `"pelo menos um verdadeiro"` quando o resultado da operação lógica de **conjunção** entre dois parâmetros for verdadeira ($x \vee y = True$) ou o string vazio `""`, quando for falsa ($x \vee y = False$):
```
def conjuncao(x, y):
# Escreva sua resposta aqui
return "pelo menos um verdadeiro" if (x or y) else ""
# Utilize esta célula para testar sua resposta
# Execute esta célula para realizar a validação e registro da sua resposta
entradas = [[False, False], [False, True], [True, False], [True, True]]
saidas = ["", "pelo menos um verdadeiro", "pelo menos um verdadeiro", "pelo menos um verdadeiro"]
validate(conjuncao, entradas, lambda x, y: x if y[0] or y[1] else "", saidas, "a1.1", False)
```
### Questão 02
Crie uma função que retorne o resultado da operação lógica de **implicação** entre dois parâmetros ($x \rightarrow y$):
```
def implicacao(x, y):
# Escreva sua resposta aqui
return (not x) or y
# Utilize esta célula para testar sua resposta
# Execute esta célula para realizar a validação e registro da sua resposta
entradas = [[False, False], [False, True], [True, False], [True, True]]
saidas = [True, True, False, True]
validate(implicacao, entradas, lambda x, y: x, saidas, "a1.2", False)
```
### Questão 03
Crie uma função que retorne o resultado da operação lógica de *exclusão* entre dois parâmetros ($x \veebar y$)
```
def exclusao(x, y):
# Escreva sua resposta aqui
return (not x) != (not y)
# Utilize esta célula para testar sua resposta
# Execute esta célula para realizar a validação e registro da sua resposta
entradas = [[False, False], [False, True], [True, False], [True, True]]
saidas = [False, True, True, False]
validate(exclusao, entradas, lambda x, y: x, saidas, "a1.3", False)
```
### Questão 04
Crie uma função que crie um arquivo com o nome indicado no parâmetro `nome`:
* Dica: você pode usar os métodos `os.system('cmd')` ou `subprocess.getoutput('cmd')` para executar comandos de terminal
```
def criar_arquivo(nome):
# Escreva sua resposta aqui
os.system(f'touch {nome}')
# Utilize esta célula para testar sua resposta
# Execute esta célula para realizar a validação e registro da sua resposta
entradas = [['file1'], ['file2'], ['file3']]
validate(criar_arquivo, entradas, lambda x, y: os.system(f'cat {y[0]}') == 0, None, "a1.4", False)
```
### Questão 05
Crie uma função que crie um diretório com o nome indicado no parâmetro `nome`:
```
def criar_diretorio(nome):
# Escreva sua resposta aqui
os.system(f'mkdir {nome}')
# Utilize esta célula para testar sua resposta
# Execute esta célula para realizar a validação e registro da sua resposta
entradas = [['dir1'], ['dir2'], ['dir3']]
validate(criar_diretorio, entradas, lambda x, y: os.system(f'ls -las {y[0]}') == 0, None, "a1.5", False)
```
### Questão 06
Crie uma função que crie um arquivo `nomeArquivo` dentro do diretório `nomeDiretorio`. A função deve criar o diretório, caso esse ainda não exista.
* Dica 1: você pode usar o comando `os.chdir(path)` para mudar de diretório
* Dica 2: você também pode executar uma composição de comandos de terminal utilizando a expressão `&&`, como em `os.system('cmd1 && cmd2')`:
```
def criar_arquivo_no_diretorio(nomeArquivo, nomeDiretorio):
# Escreva sua resposta aqui
os.system(f'mkdir -p {nomeDiretorio} && touch {nomeDiretorio}/{nomeArquivo}')
# Utilize esta célula para testar sua resposta
# Execute esta célula para realizar a validação e registro da sua resposta
entradas = [['fileA1', 'dirA'], ['fileA2', 'dirA'], ['fileB', 'dirB']]
validate(criar_arquivo_no_diretorio, entradas, lambda x, y: os.system(f'cat {y[1]}/{y[0]}')==0, None, "a1.6", False)
```
### Questão 07
Crie uma função que crie um diretório, dentro do diretório recém-criado, crie um ambiente virtual Python
* Dica: utilize o comando de terminal `virtualenv`, conforme apresentado em aula
```
def criar_projeto_python(nomeProjeto):
# Escreva sua resposta aqui
os.system(f'mkdir -p {nomeProjeto} && cd {nomeProjeto} && virtualenv venv')
# Utilize esta célula para testar sua resposta
# Execute esta célula para realizar a validação e registro da sua resposta
entradas = [['proj1'], ['proj2']]
validate(criar_projeto_python, entradas, lambda x, y: bool(len(subprocess.getoutput(f'find {y} -name site-packages'))), None, "a1.7", False)
```
### Questão 08
Crie uma função que crie um diretório, dentro do diretório recém-criado, inicie um repositório Git e configure um nome e e-mail para o repositório:
* Dica: use os comandos `git init` e `git config`, conforme apresentados em aula
```
def criar_repo_git(nomeRepositorio):
# Escreva sua resposta aqui
os.system(f'mkdir -p {nomeRepositorio} && cd {nomeRepositorio} && git init && git config user.name "John Doe" && git config user.email "johndoe@example.com"')
# Utilize esta célula para testar sua resposta
# Execute esta célula para realizar a validação e registro da sua resposta
entradas = [['repo1'], ['repo2'], ['repo3']]
validate(criar_repo_git, entradas, lambda x, y: bool(len(subprocess.getoutput(f'find {y[0]} -name HEAD'))), None, "a1.8", False)
```
### Questão 09
Crie uma função que crie um arquivo `nomeArquivo` dentro do repositório `nomeRepositorio` e faça o commit do arquivo naquele repositório:
* Observação: o repositório já foi criado no exercício anterior
* Dica 1: não se esqueça da opção `-m "comment"` para adicionar a descrição do commit.
* Dica 2: você também pode informar o autor do commit com a opção `--author="Bruce Wayne <bruce@wayne.com>"`
```
def create_add_commit(nomeArquivo, nomeRepositorio):
# Escreva sua resposta aqui
os.system(f'cd {nomeRepositorio} && touch {nomeArquivo} && git add -A && git commit -am "Coisa"')
# Utilize esta célula para testar sua resposta
# Execute esta célula para realizar a validação e registro da sua resposta
entradas = [['file1', 'repo1'], ['file2', 'repo2'], ['file3', 'repo3']]
validate(create_add_commit, entradas, lambda x, y: y[0] == subprocess.getoutput(f'cd {y[1]} && git ls-files {y[0]}'), None, "a1.9", False)
```
### Questão 10
Crie uma função que crie o branch `nomeBranch` dentro do repositório `nomeRepositorio`:
* Observação: o repositório já foi criado na Questão 08
```
def create_branch(nomeBranch, nomeRepositorio):
# Escreva sua resposta aqui
os.system(f'cd {nomeRepositorio} && git branch {nomeBranch}')
# Utilize esta célula para testar sua resposta
# Execute esta célula para realizar a validação e registro da sua resposta
entradas = [['b1', 'repo1'], ['b2', 'repo2'], ['b3', 'repo3']]
validate(create_branch, entradas, lambda x, y: y[0] == subprocess.getoutput(f'cd {y[1]} && git branch --list {y[0]}').strip(), None, "a1.10", False)
```
| github_jupyter |
# DAY0 - Looking for Dataset + Problem
```
# needed to make web requests
import requests
#store the data we get as a dataframe
import pandas as pd
#convert the response as a structured json
import json
#mathematical operations on lists
import numpy as np
#parse the datetimes we get from NOAA
from datetime import datetime
%matplotlib inline
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from datetime import timedelta
from sklearn.metrics import accuracy_score
#add the access token you got from NOAA
Token = 'xKIlBHakeOEdyBfhPkKcDKyLzjofRpNY'
#MIAMI INTERNATIONAL AIRPORT, FL US station
station_id = 'GHCND:USW00012839'
# https://www.ncdc.noaa.gov/cdo-web/datatools/findstation
#initialize lists to store data
dates_temp = []
dates_prcp = []
temps = []
prcp = []
#for each year from 2015-2019 ...
for year in range(2015, 2020):
year = str(year)
print('working on year '+year)
#make the api call
r = requests.get('https://www.ncdc.noaa.gov/cdo-web/api/v2/data?datasetid=GHCND&datatypeid=TAVG&limit=1000&stationid=GHCND:USW00023129&startdate='+year+'-01-01&enddate='+year+'-12-31', headers={'token':Token})
#load the api response as a json
d = json.loads(r.text)
#get all items in the response which are average temperature readings
avg_temps = [item for item in d['results'] if item['datatype']=='TAVG']
#get the date field from all average temperature readings
dates_temp += [item['date'] for item in avg_temps]
#get the actual average temperature from all average temperature readings
temps += [item['value'] for item in avg_temps]
#initialize dataframe
df_temp = pd.DataFrame()
#populate date and average temperature fields (cast string date to datetime and convert temperature from tenths of Celsius to Fahrenheit)
df_temp['date'] = [datetime.strptime(d, "%Y-%m-%dT%H:%M:%S") for d in dates_temp]
df_temp['avgTemp'] = [float(v)/10.0*1.8 + 32 for v in temps]
df_temp['date'].head()
df_temp['avgTemp'].head()
# ...not taking this approach...
```
# new approach ->
# DAY1 - Brainstorming & Data Preparation
Idea generation & planning
Data gathering & cleaning
Data storage
# Let's start with the Solar Dataset
```
import pandas as pd
solar = pd.read_csv('/Users/gracemartinez/Downloads/solar.csv')
solar.head()
solar.shape
solar.columns
# replacing space between words with underscore
solar.columns = solar.columns.str.replace(' ','_')
solar.columns
```
```
# verifying types of data
solar.dtypes
# need to know what each column means/represents to know if they're a correct type
'''
- Need to change:
Calendar_Date object -> (-)Y/M/D, ex. -1997 May 22
Eclipse_Time object -> Date
- will drop for python, have again for tableau:
Latitude object, separate into 2 columns: decimal# & Letter
Longitude object, separate into 2 columns: decimal# & Letter
- i dont think i need them: need to see the correlation with the Y
Path_Width_(km) object 1/3 of null values.
Central_Duration object 1/3 of null values.
# how many of each type in the data
solar['Eclipse_Type'].value_counts()
# we will data clean by putting all common categories together
# only 4 types of lunar eclipse: P, A, T, H
def eclipseClean(x):
if 'P' in x:
return('P')
if 'A' in x:
return('A')
if 'T' in x:
return('T')
if 'H' in x:
return('H')
# map function; recount the types in new category types
solar['Eclipse_Type'] = list(map(eclipseClean,solar['Eclipse_Type']))
solar['Eclipse_Type'].value_counts()
len(solar['Eclipse_Type'].value_counts())
# so now we have the 4 main categories of 'Eclipse Type',
# but we can drop 'H' column because it's "Hybrid or Annular/Total Eclipse."
# and we're only dealing with Partial, Annular, or Total Eclipses.
# we're removing the hybrid category type 'H'
# because the data will be compromised/disparity. might give unnecessary outliers
solar = solar.drop(solar[solar.Eclipse_Type == 'H'].index)
# recount the new 3 categories
solar['Eclipse_Type'].value_counts()
# now let's look at the Latitude & Longitude columns
solar.Latitude.head()
# Using regex to separate Latitude & Longitude columns
import re
solar['Latitude_Number'] = solar['Latitude'].str.replace('([A-Z]+)', '')
solar['Latitude_Letter'] = solar['Latitude'].str.extract('([A-Z]+)')
solar.head()
# Same with Logitude
solar['Longitude_Number'] = solar['Longitude'].str.replace('([A-Z]+)', '')
solar['Longitude_Letter'] = solar['Longitude'].str.extract('([A-Z]+)')
solar.head()
# Dropping original Latitude & Longitude columns
solar.drop(columns =["Latitude", "Longitude"], inplace = True)
solar.head()
solar.isnull().sum()
# how much correlation do the columns with Null have
solar.isnull().sum() / solar.shape[0]
# Need to drop 2 columns with high missing null values
solar = solar.drop(["Path_Width_(km)", "Central_Duration"], axis=1)
# Also drop 'Catalog_Number' column since it is just like the index, hence unnecessary
solar = solar.drop(["Catalog_Number"], axis=1)
solar.head()
len(solar.columns)
# make new column with no negative symbol
def c0(x):
if '-' in x:
x = x.replace('-','')
return x
solar['Calendar_Date_Clean'] = list(map(c0, solar['Calendar_Date']))
solar.head()
# Look for months only
import re
re.findall('[A-z]+' , solar['Calendar_Date_Clean'][0])
# too much time trying to convert to correct datetime format,
# used simple regex to remove negative symbol and extracted month
def c1(x):
if '-' in x:
x = x.replace('-','')
return((re.findall('[A-z]+', x))[0])
solar['Calendar_Date_Month'] = list(map(c1, solar['Calendar_Date']))
solar['Calendar_Date_Month']
# Look for years only
def c2(x):
if '-' in x:
x = x.replace('-','')
temp = re.findall('\d\d\d\d', x)
if len(temp)>0:
return temp[0]
else:
return temp
solar['Calendar_Date_Year'] = list(map(c2, solar['Calendar_Date']))
solar.head()
# drop original column
solar = solar.drop(["Calendar_Date"], axis=1)
solar.head()
# Use only first 800 rows, encompassing dates beginning from the 17th century. (1601-1999)
# Gregorian calendar starts after 1582
solar[0:800]
# Saving for future Tableau usage
solar.to_csv('Solar_tableau.csv')
SolarCategoricals = solar.select_dtypes(object)
SolarCategoricals
# Need to convert Latitude_Number & Longitude_Number to float64 type
solar["Latitude_Number"] = pd.to_numeric(solar["Latitude_Number"])
solar.head()
solar["Longitude_Number"] = pd.to_numeric(solar["Longitude_Number"])
solar.head()
solar.dtypes
SolarCategoricals = solar.select_dtypes(object)
SolarCategoricals.head()
len(SolarCategoricals.columns)
# Total columns
len(solar.dtypes)
solar = solar.drop(['Latitude_Number', 'Longitude_Number', 'Latitude_Letter', 'Longitude_Letter', 'Eclipse_Time'], axis=1)
SolarNumericals = solar._get_numeric_data()
SolarNumericals
# use corr function, will untilize number of numerical columns
S_corr_matrix = SolarNumericals.corr()
S_corr_matrix.head()
# set fig size to have better readibility of heatmap
fig, ax = plt.subplots(figsize=(8,8))
S_heatmap = sns.heatmap(S_corr_matrix, annot =True, ax=ax)
S_heatmap
# Due to high correlation, we are dropping 'Saros_Number' & 'Lunation_Number'
solar = solar.drop(['Saros_Number', 'Lunation_Number'], axis=1)
# update
SolarNumericals = solar._get_numeric_data()
SolarNumericals
S_corr_matrix = SolarNumericals.corr()
S_corr_matrix.head()
fig, ax = plt.subplots(figsize=(8,8))
S_heatmap = sns.heatmap(S_corr_matrix, annot =True, ax=ax)
S_heatmap
plt.hist(solar["Eclipse_Type"], bins = len(solar["Eclipse_Type"].unique()))
plt.xticks(rotation='vertical')
# normalize numerical values
import pandas as pd
from sklearn import preprocessing
x = SolarNumericals.values #returns a numpy array
min_max_scaler = preprocessing.MinMaxScaler()
x_scaled = min_max_scaler.fit_transform(x)
df = pd.DataFrame(x_scaled)
df.head()
solar.head()
# Now let's play with the categories!
solar.dtypes
SolarCategoricals = solar.select_dtypes(object)
SolarCategoricals.head()
from sklearn import preprocessing
solar.head()
solar.columns
# there are 2 categorical & numericals
'''
The Target[Y] is finding the Eclipse_Type
The Features[X] are what will determine the best outcome for Y
need to determine which are the best features to use to get the prediction
while having a high measurement of acuracy.
We will do a Train/Test Split in order to verify.
'''
```
```
solar
numericals = solar._get_numeric_data()
numericals = pd.DataFrame(numericals)
numericals
# Normalize x values
from sklearn.preprocessing import Normalizer
transformer = Normalizer().fit(numericals)
normalized_x = transformer.transform(numericals)
pd.DataFrame(normalized_x)
categoricals = solar.select_dtypes('object')
categoricals = categoricals['Eclipse_Type']
y = categoricals
# importing the necessary libraries
import pandas as pd
from sklearn import linear_model
from sklearn.model_selection import train_test_split
from matplotlib import pyplot as plt
# defining the target variable (dependent variable) as y
y = solar.Eclipse_Type
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
y = le.fit_transform(y)
y
# creating training and testing variables
# test_size = the percentage of the data for testing. It’s usually around 80/20 or 70/30. In this case 80/20
X_train, X_test, y_train, y_test = train_test_split(normalized_x, y, test_size=0.2)
print(X_train.shape, y_train.shape)
print(X_test.shape, y_test.shape)
```
# LINEAR REGRESSION
```
'''
# fitting the model on the training data
lm = linear_model.LinearRegression()
model = lm.fit(X_train, y_train)
predictions = lm.predict(X_test)
# ...nvm i have to use logistic regression for this CLASSIFICATION PROBLEM *eye roll, sweats*
'''
# show first five predicted values
predictions[0:5]
# plotting the model - The line / model
plt.scatter(y_test, predictions)
plt.xlabel(“True_Values”)
plt.ylabel(“Predictions”)
```
# LOGISTIC REGRESSION
```
from sklearn.datasets import load_iris
from sklearn.linear_model import LogisticRegression
# Training the Logistic Regression Model:
# Split data into 'X' features and 'y' target label sets
X = normalized_x
y = le.fit_transform(y)
# Import module to split dataset
from sklearn.model_selection import train_test_split
# Split data set into training and test sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2,) # random_state= _no._ simply sets a seed to the random generator, so that your train-test splits are always deterministic. If you don't set a seed, it is different each time.
# Import module for fitting
from sklearn.linear_model import LogisticRegression
# Create instance (i.e. object) of LogisticRegression
logmodel = LogisticRegression()
# Fit the model using the training data
# X_train -> parameter supplies the data features
# y_train -> parameter supplies the target labels
logmodel.fit(X_train, y_train)
"""
NOW,
Evaluate the Model by reviewing the classification report or confusion matrix.
By reviewing these tables, we are able to evaluate the model.
Below we are able to identify that the model has a precision of 51.4% accuracy.
To improve this we could gather more data, conduct further feature engineering and more to continue to adjust.
"""
pd.DataFrame(y_test)
from sklearn.metrics import classification_report, accuracy_score
predictions = logmodel.predict(pd.DataFrame(X_test))
print(classification_report(y_test, predictions))
print(accuracy_score(y_test, predictions))
# has a 53% model accuracy
# varies everytime i run it.
```
# RANDOM FOREST ALGORITHM
```
# Dividing data into training and testing sets:
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
# Feature Scaling
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# training our random forests to solve this classification problem
from sklearn.ensemble import RandomForestClassifier
randomForestClassification = RandomForestClassifier(n_estimators=100,random_state=259)
randomForestClassification.fit(X_train, y_train)
y_pred = randomForestClassification.predict(X_test)
pd.Series(y_pred).value_counts()
# Evaluating the Algorithm -
"""
For classification problems the metrics used to evaluate an algorithm are
accuracy, confusion matrix, precision recall, and F1 values. (also known as balanced F-score or F-measure. The F1 score can be interpreted as a weighted average of the precision and recall, where an F1 score reaches its best value at 1 and worst score at 0.)
Executing the following script to find these values:
"""
from sklearn.metrics import classification_report, confusion_matrix, accuracy_score
# print(confusion_matrix(y_test,y_pred))
print(classification_report(y_test,y_pred))
print(accuracy_score(y_test, y_pred))
# The accuracy achieved for by our random forest classifier with 100 trees is 85%.
# varies everytime i run it.
randomForestClassification.feature_importances_
```
# SVM ALGORITHM - Support Vector Machine / Classification
```
# X = normalized_x
# y = le.fit_transform(y)
# Fitting a Support Vector Machine
# import support vector classifier
from sklearn.svm import SVC # "Support Vector Classifier"
clf = SVC(kernel='linear')
# fitting x samples and y classes
clf.fit(X, y)
print(classification_report(y_test, predictions))
print(accuracy_score(y_test, predictions))
# The accuracy achieved by our svm classifier is 37%.
# varies everytime i run it.
```
# NOW LUNAR DATASET
```
lunar = pd.read_csv('/Users/gracemartinez/Downloads/lunar.csv')
lunar.head()
lunar.shape
lunar.columns
lunar.columns = lunar.columns.str.replace(' ','_')
lunar.columns
# number of columns of data
len(lunar.columns)
# different types of each column
lunar.dtypes
# need to know what each column mean/represents to know if they're a correct type
# amount of different individual types of each lunar type
lunar['Eclipse_Type'].value_counts()
# lenght of
len(lunar['Eclipse_Type'].value_counts())
# only 3 types of lunar eclipse: N, P, T
def eclipsetypeClean(x):
if 'N' in x:
return('N')
if 'P' in x:
return('P')
if 'T' in x:
return('T')
# map function to clean up the amount of different types
lunar['Eclipse_Type'] = list(map(eclipsetypeClean,lunar['Eclipse_Type']))
# recounting to make sure the different type of lunar types are accounted for
lunar['Eclipse_Type'].value_counts()
# amount of different types of lunar eclipses
len(lunar['Eclipse_Type'].value_counts())
# so there would be 3 different categories of 'Eclipse Type'
lunar.isnull().sum()
# there are no missing values
# not necessary for estimation of problem in python
lunar = lunar.drop(["Latitude", "Longitude"], axis=1)
lunar.head()
len(lunar.columns)
LunarCategoricals = lunar.select_dtypes(object)
LunarCategoricals
len(LunarCategoricals.columns)
LunarNumericals = lunar._get_numeric_data()
LunarNumericals
len(LunarNumericals.columns)
L_corr_matrix = LunarNumericals.corr()
# set fig size to have better readibility of heatmap
fig, ax = plt.subplots(figsize=(8,8))
L_heatmap = sns.heatmap(L_corr_matrix, annot =True, ax=ax)
L_heatmap
plt.hist(lunar["Eclipse_Type"], bins = len(lunar["Eclipse_Type"].unique()))
plt.xticks(rotation='vertical')
lunar.columns = lunar.columns.str.replace(' ','_')
lunar.columns
```
| github_jupyter |
# Mask R-CNN
This notebook shows how to train a Mask R-CNN object detection and segementation model on a custom coco-style data set.
```
import os
import sys
import random
import math
import re
import time
import numpy as np
import cv2
import matplotlib
import matplotlib.pyplot as plt
sys.path.insert(0, '../libraries')
from mrcnn.config import Config
import mrcnn.utils as utils
import mrcnn.model as modellib
import mrcnn.visualize as visualize
from mrcnn.model import log
import mcoco.coco as coco
import mextra.utils as extra_utils
%matplotlib inline
%config IPCompleter.greedy=True
HOME_DIR = '/home/keras'
DATA_DIR = os.path.join(HOME_DIR, "data/shapes")
WEIGHTS_DIR = os.path.join(HOME_DIR, "data/weights")
MODEL_DIR = os.path.join(DATA_DIR, "logs")
# Local path to trained weights file
COCO_MODEL_PATH = os.path.join(WEIGHTS_DIR, "mask_rcnn_coco.h5")
# Download COCO trained weights from Releases if needed
if not os.path.exists(COCO_MODEL_PATH):
utils.download_trained_weights(COCO_MODEL_PATH)
def get_ax(rows=1, cols=1, size=8):
"""Return a Matplotlib Axes array to be used in
all visualizations in the notebook. Provide a
central point to control graph sizes.
Change the default size attribute to control the size
of rendered images
"""
_, ax = plt.subplots(rows, cols, figsize=(size*cols, size*rows))
return ax
```
# Dataset
Organize the dataset using the following structure:
```
DATA_DIR
│
└───annotations
│ │ instances_<subset><year>.json
│
└───<subset><year>
│ image021.jpeg
│ image022.jpeg
```
```
dataset_train = coco.CocoDataset()
dataset_train.load_coco(DATA_DIR, subset="shapes_train", year="2018")
dataset_train.prepare()
dataset_validate = coco.CocoDataset()
dataset_validate.load_coco(DATA_DIR, subset="shapes_validate", year="2018")
dataset_validate.prepare()
dataset_test = coco.CocoDataset()
dataset_test.load_coco(DATA_DIR, subset="shapes_test", year="2018")
dataset_test.prepare()
# Load and display random samples
image_ids = np.random.choice(dataset_train.image_ids, 4)
for image_id in image_ids:
image = dataset_train.load_image(image_id)
mask, class_ids = dataset_train.load_mask(image_id)
visualize.display_top_masks(image, mask, class_ids, dataset_train.class_names)
```
# Configuration
```
image_size = 64
rpn_anchor_template = (1, 2, 4, 8, 16) # anchor sizes in pixels
rpn_anchor_scales = tuple(i * (image_size // 16) for i in rpn_anchor_template)
class ShapesConfig(Config):
"""Configuration for training on the shapes dataset.
"""
NAME = "shapes"
# Train on 1 GPU and 2 images per GPU. Put multiple images on each
# GPU if the images are small. Batch size is 2 (GPUs * images/GPU).
GPU_COUNT = 1
IMAGES_PER_GPU = 1
# Number of classes (including background)
NUM_CLASSES = 1 + 3 # background + 3 shapes (triangles, circles, and squares)
# Use smaller images for faster training.
IMAGE_MAX_DIM = image_size
IMAGE_MIN_DIM = image_size
# Use smaller anchors because our image and objects are small
RPN_ANCHOR_SCALES = rpn_anchor_scales
# Aim to allow ROI sampling to pick 33% positive ROIs.
TRAIN_ROIS_PER_IMAGE = 32
STEPS_PER_EPOCH = 400
VALIDATION_STEPS = STEPS_PER_EPOCH / 20
config = ShapesConfig()
config.display()
```
# Model
```
model = modellib.MaskRCNN(mode="training", config=config, model_dir=MODEL_DIR)
inititalize_weights_with = "coco" # imagenet, coco, or last
if inititalize_weights_with == "imagenet":
model.load_weights(model.get_imagenet_weights(), by_name=True)
elif inititalize_weights_with == "coco":
model.load_weights(COCO_MODEL_PATH, by_name=True,
exclude=["mrcnn_class_logits", "mrcnn_bbox_fc",
"mrcnn_bbox", "mrcnn_mask"])
elif inititalize_weights_with == "last":
# Load the last model you trained and continue training
model.load_weights(model.find_last()[1], by_name=True)
```
# Training
Training in two stages
## Heads
Only the heads. Here we're freezing all the backbone layers and training only the randomly initialized layers (i.e. the ones that we didn't use pre-trained weights from MS COCO). To train only the head layers, pass layers='heads' to the train() function.
```
model.train(dataset_train, dataset_validate,
learning_rate=config.LEARNING_RATE,
epochs=2,
layers='heads')
```
## Fine-tuning
Fine-tune all layers. Pass layers="all to train all layers.
```
model.train(dataset_train, dataset_validate,
learning_rate=config.LEARNING_RATE / 10,
epochs=3, # starts from the previous epoch, so only 1 additional is trained
layers="all")
```
# Detection
```
class InferenceConfig(ShapesConfig):
GPU_COUNT = 1
IMAGES_PER_GPU = 1
inference_config = InferenceConfig()
# Recreate the model in inference mode
model = modellib.MaskRCNN(mode="inference",
config=inference_config,
model_dir=MODEL_DIR)
# Get path to saved weights
# Either set a specific path or find last trained weights
# model_path = os.path.join(ROOT_DIR, ".h5 file name here")
print(model.find_last()[1])
model_path = model.find_last()[1]
# Load trained weights (fill in path to trained weights here)
assert model_path != "", "Provide path to trained weights"
print("Loading weights from ", model_path)
model.load_weights(model_path, by_name=True)
```
### Test on a random image from the test set
First, show the ground truth of the image, then show detection results.
```
image_id = random.choice(dataset_test.image_ids)
original_image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset_test, inference_config,
image_id, use_mini_mask=False)
log("original_image", original_image)
log("image_meta", image_meta)
log("gt_class_id", gt_class_id)
log("gt_bbox", gt_bbox)
log("gt_mask", gt_mask)
visualize.display_instances(original_image, gt_bbox, gt_mask, gt_class_id,
dataset_train.class_names, figsize=(8, 8))
results = model.detect([original_image], verbose=1)
r = results[0]
visualize.display_instances(original_image, r['rois'], r['masks'], r['class_ids'],
dataset_test.class_names, r['scores'], ax=get_ax())
```
# Evaluation
Use the test dataset to evaluate the precision of the model on each class.
```
predictions =\
extra_utils.compute_multiple_per_class_precision(model, inference_config, dataset_test,
number_of_images=250, iou_threshold=0.5)
complete_predictions = []
for shape in predictions:
complete_predictions += predictions[shape]
print("{} ({}): {}".format(shape, len(predictions[shape]), np.mean(predictions[shape])))
print("--------")
print("average: {}".format(np.mean(complete_predictions)))
print(model.find_last()[1])
```
## Convert result to COCO
Converting the result back to a COCO-style format for further processing
```
import json
import pylab
import matplotlib.pyplot as plt
from tempfile import NamedTemporaryFile
from pycocotools.coco import COCO
coco_dict = extra_utils.result_to_coco(results[0], dataset_test.class_names,
np.shape(original_image)[0:2], tolerance=0)
with NamedTemporaryFile('w') as jsonfile:
json.dump(coco_dict, jsonfile)
jsonfile.flush()
coco_data = COCO(jsonfile.name)
category_ids = coco_data.getCatIds(catNms=['square', 'circle', 'triangle'])
image_data = coco_data.loadImgs(1)[0]
image = original_image
plt.imshow(image); plt.axis('off')
pylab.rcParams['figure.figsize'] = (8.0, 10.0)
annotation_ids = coco_data.getAnnIds(imgIds=image_data['id'], catIds=category_ids, iscrowd=None)
annotations = coco_data.loadAnns(annotation_ids)
coco_data.showAnns(annotations)
```
| github_jupyter |
```
import sys
sys.path.append('./../')
%load_ext autoreload
%autoreload 2
from ontology import get_ontology
ontology = get_ontology('../data/doid.obo')
name2doid = {term.name: term.id for term in ontology.get_terms()}
doid2name = {term.id: term.name for term in ontology.get_terms()}
import numpy as np
import re
```
# Wiki links from obo descriptions
```
import wiki
lst = wiki.get_links_from_ontology(ontology)
print r'example:{:}'.format(repr(lst[10]))
```
### urllib2 to read page in html
```
page = wiki.get_html(lst[101])
page[:1000]
```
# Fuzzy logic
```
import fuzzywuzzy.process as fuzzy_process
from fuzzywuzzy import fuzz
string = "ventricular arrhythmia"
names = np.sort(name2doid.keys())
print fuzzy_process.extractOne(string, names, scorer=fuzz.token_set_ratio)
string = "Complete remission of hairy cell leukemia variant (HCL-v) complicated by red cell aplasia post treatment with rituximab."
print fuzzy_process.extractOne(string, names, scorer=fuzz.partial_ratio)
```
# Wikipedia search engine: headers
```
query = "ventricular arrhythmia"
top = wiki.get_top_headers(query)
top
for header in top:
results = fuzzy_process.extractOne(header, names, scorer=fuzz.token_set_ratio)
print results
page = wikipedia.WikipediaPage(title='Cell_proliferation')
page.summary
```
[name for name in names if len(re.split(' ', name)) > 3]
### pub-med
```
import pubmed
query = 'hcl-v'
titles = pubmed.get(query)
titles_len = [len(title) for title in titles]
for i, string in enumerate(titles):
print("%d) %s" % (i+1, string))
print fuzzy_process.extractOne(string, names, scorer=fuzz.partial_ratio)
print
```
def find_synonym(s_ref, s):
last = s_ref.find('(' + s + ')')
if last == -1:
return None
n_upper = len(''.join([c for c in s if c.isupper()]))
first = [(i,c) for i, c in enumerate(s_ref[:last]) if c.isupper()][-n_upper][0]
return s_ref[first:last-1]
print find_synonym('Wolff-Parkinson-White syndrome (WPW) and athletes: Darwin at play?',
'WPW')
### synonyms
```
import utils
print utils.find_synonym('Wolff-Parkinson-White syndrome (WPW) and athletes: Darwin at play?', 'WPW')
print utils.find_synonym('Complete remission of hairy cell leukemia variant (HCL-v)...', 'hcl-v')
```
### Assymetric distance
```
s_ref = 'artery disease'
s = 'nonartery'
print utils.assym_dist(s, s_ref)
```
### Length statistics
```
print 'Mean term name length:', np.mean([len(term.name) for term in ontology.get_terms()])
print 'Mean article title length:', np.mean(titles_len)
```
### Unique words
```
words = [re.split(' |-', term.name) for term in ontology.get_terms()]
words = np.unique([l for sublist in words for l in sublist if len(l) > 0])
words = [w for w in words if len(w) >= 4]
words[:10]
```
# Threading
```
from threading import Thread
from time import sleep
from ontology import get_ontology
query_results = None
def fn_get_q(query):
global query_results
query_results = fuzzy_process.extractOne(query, names, scorer=fuzz.ratio)
return True
wiki_results = None
def fn_get_wiki(query):
global wiki_results
header = wiki.get_top_headers(query, 1)[0]
wiki_results = fuzzy_process.extractOne(header, names, scorer=fuzz.ratio)
#sleep(0.1)
return True
pubmed_results = None
def fn_get_pubmed(query):
global pubmed_results
string = pubmed.get(query, topK=1)
if string is not None:
string = string[0]
print string
pubmed_results = fuzzy_process.extractOne(string, names, scorer=fuzz.partial_ratio)
return True
else:
return False
'''main'''
## from bot
query = 'valve disease'
def find_answer(query):
query = query.lower()
# load ontology
ontology = get_ontology('../data/doid.obo')
name2doid = {term.name: term.id for term in ontology.get_terms()}
doid2name = {term.id: term.name for term in ontology.get_terms()}
## exact match
if query in name2doid.keys():
doid = name2doid[query]
else:
# exact match -- no
th_get_q = Thread(target = fn_get_q, args = (query,))
th_get_wiki = Thread(target = fn_get_wiki, args = (query,))
th_get_pubmed = Thread(target = fn_get_pubmed, args = (query,))
th_get_q.start()
th_get_wiki.start()
th_get_pubmed.start()
## search engine query --> vertices, p=100(NLP??); synonyms
## new thread for synonyms???
## synonyms NLP
## new thread for NLP
## tree search on vertices (returned + synonyms)
## sleep ?
th_get_q.join()
print query_results
th_get_wiki.join()
print wiki_results
th_get_pubmed.join()
print pubmed_results
## final answer
## draw graph
doid = None
graph = None
return doid, graph
```
| github_jupyter |
<a href="https://colab.research.google.com/github/mirianfsilva/The-Heat-Diffusion-Equation/blob/master/FiniteDiff_test.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
### Implementation of schemes for the Heat Equation:
- Forward Time, Centered Space;
- Backward Time, Centered Space;
- Crank-Nicolson.
\begin{equation}
\partial_{t}u = \partial^2_{x}u , \quad 0 < x < 1, \quad t > 0 \\
\end{equation}
\begin{equation}
\partial_{x}u(0,t) = 0, \quad \partial_x{u}(1,t) = 0\\
\end{equation}
\begin{equation}
u(x, 0) = cos(\pi x)
\end{equation}
### Exact Solution:
\begin{equation}
u(x,t) = e^{-\pi^2t}cos(\pi x)
\end{equation}
```
#Numerical Differential Equations - Federal University of Minas Gerais
""" Utils """
import math, sys
import numpy as np
import sympy as sp
from scipy import sparse
from sympy import fourier_series, pi
from scipy.fftpack import *
from scipy.sparse import diags
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from matplotlib import cm
from os import path
count = 0
#Heat Diffusion in one dimensional wire within the Explicit Method
"""
λ = 2, λ = 1/2 e λ = 1/6
M = 4, M = 8, M = 10, M = 12 e M = 14
"""
#Heat function exact solution
def Solution(x, t):
return np.exp((-np.pi**2)*t)*np.cos(np.pi*x)
# ---- Surface plot ----
def surfaceplot(U, Uexact, tspan, xspan, M):
N = M**2
#meshgrid : Return coordinate matrices from coordinate vectors
X, T = np.meshgrid(tspan, xspan)
fig = plt.figure(figsize=plt.figaspect(0.3))
#fig2 = plt.figure(figsize=plt.figaspect(0.5))
#fig3 = plt.figure(figsize=plt.figaspect(0.5))
# ---- Exact Solution ----
ax = fig.add_subplot(1, 4, 1,projection='3d')
surf = ax.plot_surface(X, T, Uexact, linewidth=0, cmap=cm.jet, antialiased=True)
ax.set_title('Exact Solution')
ax.set_xlabel('Time')
ax.set_ylabel('Space')
ax.set_zlabel('U')
# ---- Method Aproximation Solution ----
ax1 = fig.add_subplot(1, 4, 2,projection='3d')
surf = ax1.plot_surface(X, T, U, linewidth=0, cmap=cm.jet, antialiased=True)
ax1.set_title('Approximation')
ax1.set_xlabel('Time')
ax1.set_ylabel('Space')
ax1.set_zlabel('U')
plt.tight_layout()
ax.view_init(30,230)
ax1.view_init(30,230)
fig.savefig(path.join("plot_METHOD{0}.png".format(count)),dpi=600)
plt.draw()
'''
Exact Solution for 1D reaction-diffusion equation:
u_t = k * u_xx
with Neumann boundary conditions
at x=0: u_x(0,t) = 0
at x=L: u_x(L,t) = 0
with L = 1 and initial conditions:
u(x,0) = np.cos(np.pi*x)
'''
def ExactSolution(M, T = 0.5, L = 1):
N = (M**2) #GRID POINTS on time interval
xspan = np.linspace(0, L, M)
tspan = np.linspace(0, T, N)
Uexact = np.zeros((M, N))
for i in range(0, M):
for j in range(0, N):
Uexact[i][j] = Solution(xspan[i], tspan[j])
return (Uexact, tspan, xspan)
'''
Forward method to solve 1D reaction-diffusion equation:
u_t = k * u_xx
with Neumann boundary conditions
at x=0: u_x(0,t) = 0 = sin(2*np.pi)
at x=L: u_x(L,t) = 0 = sin(2*np.pi)
with L = 1 and initial conditions:
u(x,0) = (1.0/2.0)+ np.cos(2.0*np.pi*x) - (1.0/2.0)*np.cos(3*np.pi*x)
u_x(x,t) = (-4.0*(np.pi**2))np.exp(-4.0*(np.pi**2)*t)*np.cos(2.0*np.pi*x) +
(9.0/2.0)*(np.pi**2)*np.exp(-9.0*(np.pi**2)*t)*np.cos(3*np.pi*x))
'''
def ForwardEuler(M, lambd, T = 0.5, L = 1, k = 1):
#Parameters needed to solve the equation within the explicit method
#M = GRID POINTS on space interval
N = (M**2) #GRID POINTS on time interval
# ---- Length of the wire in x direction ----
x0, xL = 0, L
# ----- Spatial discretization step -----
dx = (xL - x0)/(M-1)
# ---- Final time ----
t0,tF = 0, T
# ----- Time step -----
dt = (tF - t0)/(N-1)
#lambd = dt*k/dx**2
# ----- Creates grids -----
xspan = np.linspace(x0, xL, M)
tspan = np.linspace(t0, tF, N)
# ----- Initializes matrix solution U -----
U = np.zeros((M, N))
# ----- Initial condition -----
U[:,0] = np.cos(np.pi*xspan)
# ----- Neumann boundary conditions -----
"""
To implement these boundary conditions, we again use “false points”, x_0 and x_N+1 which are external points.
We use a difference to approximate ∂u/∂x (xL,t) and set it equal to the desired boundary condition:
"""
f = np.arange(1, N+1)
f = (-3*U[0,:] + 4*U[1,:] - U[2,:])/2*dx
U[0,:] = (4*U[1,:] - U[2,:])/3
g = np.arange(1, N+1)
g = (-3*U[-1,:] + 4*U[-2,:] - U[-3,:])/2*dx
U[-1,:] = (4*U[-2,:] - U[-3,:])/3
# ----- ftcs -----
for k in range(0, N-1):
for i in range(1, M-1):
U[i, k+1] = lambd*U[i-1, k] + (1-2*lambd)*U[i,k] + lambd*U[i+1,k]
return (U, tspan, xspan)
U, tspan, xspan = ForwardEuler(M = 14, lambd = 1.0/6.0)
Uexact, x, t = ExactSolution(M = 14)
surfaceplot(U, Uexact, tspan, xspan, M = 14)
'''
Backward method to solve 1D reaction-diffusion equation:
u_t = k * u_xx
with Neumann boundary conditions
at x=0: u_x(0,t) = 0 = sin(2*np.pi)
at x=L: u_x(L,t) = 0 = sin(2*np.pi)
with L = 1 and initial conditions:
u(x,0) = (1.0/2.0)+ np.cos(2.0*np.pi*x) - (1.0/2.0)*np.cos(3*np.pi*x)
u_x(x,t) = (-4.0*(np.pi**2))np.exp(-4.0*(np.pi**2)*t)*np.cos(2.0*np.pi*x) +
(9.0/2.0)*(np.pi**2)*np.exp(-9.0*(np.pi**2)*t)*np.cos(3*np.pi*x))
'''
def BackwardEuler(M, lambd, T = 0.5, L = 1, k = 1):
#Parameters needed to solve the equation within the explicit method
# M = GRID POINTS on space interval
N = (M**2) #GRID POINTS on time interval
# ---- Length of the wire in x direction ----
x0, xL = 0, L
# ----- Spatial discretization step -----
dx = (xL - x0)/(M-1)
# ---- Final time ----
t0, tF = 0, T
# ----- Time step -----
dt = (tF - t0)/(N-1)
# k = 1.0 Diffusion coefficient
#lambd = dt*k/dx**2
a = 1 + 2*lambd
xspan = np.linspace(x0, xL, M)
tspan = np.linspace(t0, tF, N)
main_diag = (1 + 2*lambd)*np.ones((1,M))
off_diag = -lambd*np.ones((1, M-1))
a = main_diag.shape[1]
diagonals = [main_diag, off_diag, off_diag]
#Sparse Matrix diagonals
A = sparse.diags(diagonals, [0,-1,1], shape=(a,a)).toarray()
A[0,1] = -2*lambd
A[M-1,M-2] = -2*lambd
# --- Initializes matrix U -----
U = np.zeros((M, N))
# --- Initial condition -----
U[:,0] = np.cos(np.pi*xspan)
# ---- Neumann boundary conditions -----
f = np.arange(1, N+1) #LeftBC
#(-3*U[i,j] + 4*U[i-1,j] - U[i-2,j])/2*dx = 0
f = U[0,:] = (4*U[1,:] - U[2,:])/3
g = np.arange(1, N+1) #RightBC
#(-3*U[N,j] + 4*U[N-1,j] - U[N-2,j])/2*dx = 0
g = U[-1,:] = (4*U[-2,:] - U[-3,:])/3
for i in range(1, N):
c = np.zeros((M-2,1)).ravel()
b1 = np.asarray([2*lambd*dx*f[i], 2*lambd*dx*g[i]])
b1 = np.insert(b1, 1, c)
b2 = np.array(U[0:M, i-1])
b = b1 + b2 # Right hand side
U[0:M, i] = np.linalg.solve(A,b) # Solve x=A\b
return (U, tspan, xspan)
U, tspan, xspan = BackwardEuler(M = 14, lambd = 1.0/6.0)
Uexact, x, t = ExactSolution(M = 14)
surfaceplot(U, Uexact, tspan, xspan, M = 14)
'''
Crank-Nicolson method to solve 1D reaction-diffusion equation:
u_t = D * u_xx
with Neumann boundary conditions
at x=0: u_x = sin(2*pi)
at x=L: u_x = sin(2*pi)
with L=1 and initial condition:
u(x,0) = u(x,0) = (1.0/2.0)+ np.cos(2.0*np.pi*x) - (1.0/2.0)*np.cos(3*np.pi*x)
'''
def CrankNicolson(M, lambd, T = 0.5, L = 1, k = 1):
#Parameters needed to solve the equation within the explicit method
# M = GRID POINTS on space interval
N = (M**2) #GRID POINTS on time interval
# ---- Length of the wire in x direction ----
x0, xL = 0, L
# ----- Spatial discretization step -----
dx = (xL - x0)/(M-1)
# ---- Final time ----
t0, tF = 0, T
# ----- Time step -----
dt = (tF - t0)/(N-1)
#lambd = dt*k/(2.0*dx**2)
a0 = 1 + 2*lambd
c0 = 1 - 2*lambd
xspan = np.linspace(x0, xL, M)
tspan = np.linspace(t0, tF, N)
maindiag_a0 = a0*np.ones((1,M))
offdiag_a0 = (-lambd)*np.ones((1, M-1))
maindiag_c0 = c0*np.ones((1,M))
offdiag_c0 = lambd*np.ones((1, M-1))
#Left-hand side tri-diagonal matrix
a = maindiag_a0.shape[1]
diagonalsA = [maindiag_a0, offdiag_a0, offdiag_a0]
A = sparse.diags(diagonalsA, [0,-1,1], shape=(a,a)).toarray()
A[0,1] = (-2)*lambd
A[M-1,M-2] = (-2)*lambd
#Right-hand side tri-diagonal matrix
c = maindiag_c0.shape[1]
diagonalsC = [maindiag_c0, offdiag_c0, offdiag_c0]
Arhs = sparse.diags(diagonalsC, [0,-1,1], shape=(c,c)).toarray()
Arhs[0,1] = 2*lambd
Arhs[M-1,M-2] = 2*lambd
# ----- Initializes matrix U -----
U = np.zeros((M, N))
#----- Initial condition -----
U[:,0] = np.cos(np.pi*xspan)
#----- Neumann boundary conditions -----
#Add one line above and one line below using finit differences
f = np.arange(1, N+1) #LeftBC
#(-3*U[i,j] + 4*U[i-1,j] - U[i-2,j])/2*dx = 0
f = U[0,:] = (4*U[1,:] - U[2,:])/3
g = np.arange(1, N+1) #RightBC
#(-3*U[N,j] + 4*U[N-1,j] - U[N-2,j])/2*dx = 0
g = U[-1,:] = (4*U[-2,:] - U[-3,:])/3
for k in range(1, N):
ins = np.zeros((M-2,1)).ravel()
b1 = np.asarray([4*lambd*dx*f[k], 4*lambd*dx*g[k]])
b1 = np.insert(b1, 1, ins)
b2 = np.matmul(Arhs, np.array(U[0:M, k-1]))
b = b1 + b2 # Right hand side
U[0:M, k] = np.linalg.solve(A,b) # Solve x=A\b
return (U, tspan, xspan)
U, tspan, xspan = CrankNicolson(M = 14, lambd = 1.0/6.0)
Uexact, x, t = ExactSolution(M = 14)
surfaceplot(U, Uexact, tspan, xspan, M = 14)
```
| github_jupyter |
# Overview
The tool serves to let you create task files from CSVs and zip files that you upload through the browser
```
import ipywidgets as ipw
import pandas as pd
import json, io, os, tempfile
import fileupload as fu
from IPython.display import display, FileLink
def upload_as_file_widget(callback=None):
"""Create an upload files button that creates a temporary file and calls a function with the path.
"""
_upload_widget = fu.FileUploadWidget()
def _virtual_file(change):
file_ext = os.path.splitext(change['owner'].filename)[-1]
print('Uploaded `{}`'.format(change['owner'].filename))
if callback is not None:
with tempfile.NamedTemporaryFile(suffix=file_ext) as f:
f.write(change['owner'].data)
callback(f.name)
_upload_widget.observe(_virtual_file, names='data')
return _upload_widget
def make_task(in_df,
image_path='Image Index',
output_labels='Finding Labels',
base_image_directory = 'sample_data'):
return {
'google_forms': {'form_url': 'https://docs.google.com/forms/d/e/1FAIpQLSfBmvqCVeDA7IZP2_mw_HZ0OTgDk2a0JN4VlY5KScECWC-_yw/viewform',
'sheet_url': 'https://docs.google.com/spreadsheets/d/1T02tRhe3IUUHYsMchc7hmH8nVI3uR0GffdX1PNxKIZA/edit?usp=sharing'
},
'dataset': {
'image_path': image_path, # column name
'output_labels': output_labels, # column name
'dataframe': in_df.to_dict(),
'base_image_directory': base_image_directory # path
}
}
def save_task(annotation_task, out_path='task.json'):
with open(out_path, 'w') as f:
json.dump(annotation_task, f)
return out_path
```
## Instructions
Load a CSV file and select the columns for the image path, labels and the name of the directory where the images are located
```
def _load_csv_app(in_path):
"""
A callback to create an app from an uploaded CSV file
>>> _load_csv_app('sample_data/dataset_overview.csv')
"""
ds_df = pd.read_csv(in_path)
table_viewer = ipw.HTML(value=ds_df.sample(3).T.style.render(), layout = ipw.Layout(width="45%"))
image_path_widget = ipw.Dropdown(
options=ds_df.columns,
value=ds_df.columns[0],
description='Image Path Column:',
disabled=False
)
output_labels_widget = ipw.Dropdown(
options=ds_df.columns,
value=ds_df.columns[0],
description='Label Column:',
disabled=False
)
all_dir_list = [p for p, _, _ in os.walk('.') if os.path.isdir(p) and not any([k.startswith('.') and len(k)>1 for k in p.split('/')])]
base_image_directory_widget = ipw.Select(
options=all_dir_list,
value=None,
rows=5,
description='Local Image Folder:',
disabled=False
)
def _create_task(btn):
c_task = make_task(ds_df,
image_path = image_path_widget.value,
output_labels = output_labels_widget.value,
base_image_directory = base_image_directory_widget.value
)
display(FileLink(save_task(c_task)))
create_but = ipw.Button(description='Create Task')
create_but.on_click(_create_task)
controls = ipw.VBox([image_path_widget, output_labels_widget,
base_image_directory_widget, create_but])
out_widget = ipw.HBox([controls,
table_viewer])
display(out_widget)
return out_widget
upload_as_file_widget(_load_csv_app)
```
| github_jupyter |
# Fully-Connected Neural Nets
In the previous homework you implemented a fully-connected two-layer neural network on CIFAR-10. The implementation was simple but not very modular since the loss and gradient were computed in a single monolithic function. This is manageable for a simple two-layer network, but would become impractical as we move to bigger models. Ideally we want to build networks using a more modular design so that we can implement different layer types in isolation and then snap them together into models with different architectures.
In this exercise we will implement fully-connected networks using a more modular approach. For each layer we will implement a `forward` and a `backward` function. The `forward` function will receive inputs, weights, and other parameters and will return both an output and a `cache` object storing data needed for the backward pass, like this:
```python
def layer_forward(x, w):
""" Receive inputs x and weights w """
# Do some computations ...
z = np.matmul(x, w) # ... some intermediate value
# Do some more computations ...
out = np.max(z, 0) # the output
cache = (x, w, z, out) # Values we need to compute gradients
return out, cache
```
The backward pass will receive upstream derivatives and the `cache` object, and will return gradients with respect to the inputs and weights, like this:
```python
def layer_backward(dout, cache):
"""
Receive derivative of loss with respect to outputs and cache,
and compute derivative with respect to inputs.
"""
# Unpack cache values
x, w, z, out = cache
# Use values in cache to compute derivatives
dx = dout * w # Derivative of loss with respect to x
dw = dout * x # Derivative of loss with respect to w
return dx, dw
```
After implementing a bunch of layers this way, we will be able to easily combine them to build classifiers with different architectures.
In addition to implementing fully-connected networks of arbitrary depth, we will also explore different update rules for optimization, and introduce Dropout as a regularizer and Batch Normalization as a tool to more efficiently optimize deep networks.
```
# As usual, a bit of setup
import time
import numpy as np
import matplotlib.pyplot as plt
from cs231n.classifiers.fc_net import affine_norm_relu_forward, affine_norm_relu_backward
from cs231n.data_utils import get_CIFAR10_data
from cs231n.gradient_check import eval_numerical_gradient, eval_numerical_gradient_array
from cs231n.solver import Solver
from cs231n.layers import *
%matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# for auto-reloading external modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
%load_ext autoreload
%autoreload 2
def rel_error(x, y):
""" returns relative error """
return np.max(np.abs(x - y) / (np.maximum(1e-8, np.abs(x) + np.abs(y))))
# Load the (preprocessed) CIFAR10 data.
# unicode --> pickle.load(f, encoding='latin1')
DIR_CS231n = 'd:/lecture/CS231/assignment2/'
data = get_CIFAR10_data(DIR_CS231n)
for k, v in data.items():
print ('%s: ' % k, v.shape)
```
# Affine layer: foward
Open the file `cs231n/layers.py` and implement the `affine_forward` function.
Once you are done you can test your implementaion by running the following:
```
# Test the affine_forward function
num_inputs = 2
input_shape = (4, 5, 6)
output_dim = 3
input_size = num_inputs * np.prod(input_shape)
weight_size = output_dim * np.prod(input_shape)
x = np.linspace(-0.1, 0.5, num=input_size).reshape(num_inputs, *input_shape)
w = np.linspace(-0.2, 0.3, num=weight_size).reshape(np.prod(input_shape), output_dim)
b = np.linspace(-0.3, 0.1, num=output_dim)
out, _ = affine_forward(x, w, b)
correct_out = np.array([[ 1.49834967, 1.70660132, 1.91485297],
[ 3.25553199, 3.5141327, 3.77273342]])
# Compare your output with ours. The error should be around 1e-9.
print ('Testing affine_forward function:')
print ('difference: ', rel_error(out, correct_out))
```
# Affine layer: backward
Now implement the `affine_backward` function and test your implementation using numeric gradient checking.
```
# Test the affine_backward function
x = np.random.randn(10, 2, 3)
w = np.random.randn(6, 5)
b = np.random.randn(5)
dout = np.random.randn(10, 5)
dx_num = eval_numerical_gradient_array(lambda x: affine_forward(x, w, b)[0], x, dout)
dw_num = eval_numerical_gradient_array(lambda w: affine_forward(x, w, b)[0], w, dout)
db_num = eval_numerical_gradient_array(lambda b: affine_forward(x, w, b)[0], b, dout)
_, cache = affine_forward(x, w, b)
dx, dw, db = affine_backward(dout, cache)
# The error should be around 1e-10
print ('Testing affine_backward function:')
print ('dx error: ', rel_error(dx_num, dx))
print ('dw error: ', rel_error(dw_num, dw))
print ('db error: ', rel_error(db_num, db))
```
# ReLU layer: forward
Implement the forward pass for the ReLU activation function in the `relu_forward` function and test your implementation using the following:
```
# Test the relu_forward function
x = np.linspace(-0.5, 0.5, num=12).reshape(3, 4)
out, _ = relu_forward(x)
correct_out = np.array([[ 0., 0., 0., 0., ],
[ 0., 0., 0.04545455, 0.13636364,],
[ 0.22727273, 0.31818182, 0.40909091, 0.5, ]])
# Compare your output with ours. The error should be around 1e-8
print ('Testing relu_forward function:')
print ('difference: ', rel_error(out, correct_out))
```
# ReLU layer: backward
Now implement the backward pass for the ReLU activation function in the `relu_backward` function and test your implementation using numeric gradient checking:
```
x = np.random.randn(10, 10)
dout = np.random.randn(*x.shape)
dx_num = eval_numerical_gradient_array(lambda x: relu_forward(x)[0], x, dout)
_, cache = relu_forward(x)
dx = relu_backward(dout, cache)
# The error should be around 1e-12
print ('Testing relu_backward function:')
print ('dx error: ', rel_error(dx_num, dx))
```
# "Sandwich" layers
There are some common patterns of layers that are frequently used in neural nets. For example, affine layers are frequently followed by a ReLU nonlinearity. To make these common patterns easy, we define several convenience layers in the file `cs231n/layer_utils.py`.
For now take a look at the `affine_relu_forward` and `affine_relu_backward` functions, and run the following to numerically gradient check the backward pass:
```
from cs231n.layer_utils import affine_relu_forward, affine_relu_backward
x = np.random.randn(2, 3, 4)
w = np.random.randn(12, 10)
b = np.random.randn(10)
dout = np.random.randn(2, 10)
out, cache = affine_relu_forward(x, w, b)
dx, dw, db = affine_relu_backward(dout, cache)
dx_num = eval_numerical_gradient_array(lambda x: affine_relu_forward(x, w, b)[0], x, dout)
dw_num = eval_numerical_gradient_array(lambda w: affine_relu_forward(x, w, b)[0], w, dout)
db_num = eval_numerical_gradient_array(lambda b: affine_relu_forward(x, w, b)[0], b, dout)
print ('Testing affine_relu_forward:')
print ('dx error: ', rel_error(dx_num, dx))
print ('dw error: ', rel_error(dw_num, dw))
print ('db error: ', rel_error(db_num, db))
```
# Loss layers: Softmax and SVM
You implemented these loss functions in the last assignment, so we'll give them to you for free here. You should still make sure you understand how they work by looking at the implementations in `cs231n/layers.py`.
You can make sure that the implementations are correct by running the following:
```
num_classes, num_inputs = 10, 50
x = 0.001 * np.random.randn(num_inputs, num_classes)
y = np.random.randint(num_classes, size=num_inputs)
dx_num = eval_numerical_gradient(lambda x: svm_loss(x, y)[0], x, verbose=False)
loss, dx = svm_loss(x, y)
# Test svm_loss function. Loss should be around 9 and dx error should be 1e-9
print ('Testing svm_loss:')
print ('loss: ', loss)
print ('dx error: ', rel_error(dx_num, dx))
dx_num = eval_numerical_gradient(lambda x: softmax_loss(x, y)[0], x, verbose=False)
loss, dx = softmax_loss(x, y)
# Test softmax_loss function. Loss should be 2.3 and dx error should be 1e-8
print ('\nTesting softmax_loss:')
print ('loss: ', loss)
print ('dx error: ', rel_error(dx_num, dx))
```
# Two-layer network
In the previous assignment you implemented a two-layer neural network in a single monolithic class. Now that you have implemented modular versions of the necessary layers, you will reimplement the two layer network using these modular implementations.
Open the file `cs231n/classifiers/fc_net.py` and complete the implementation of the `TwoLayerNet` class. This class will serve as a model for the other networks you will implement in this assignment, so read through it to make sure you understand the API. You can run the cell below to test your implementation.
```
from cs231n.classifiers.fc_net import *
N, D, H, C = 3, 5, 50, 7
X = np.random.randn(N, D)
y = np.random.randint(C, size=N)
std = 1e-2
model = TwoLayerNet(input_dim=D, hidden_dim=H, num_classes=C, weight_scale=std)
print ('Testing initialization ... ')
W1_std = abs(model.params['W1'].std() - std)
b1 = model.params['b1']
W2_std = abs(model.params['W2'].std() - std)
b2 = model.params['b2']
assert W1_std < std / 10, 'First layer weights do not seem right'
assert np.all(b1 == 0), 'First layer biases do not seem right'
assert W2_std < std / 10, 'Second layer weights do not seem right'
assert np.all(b2 == 0), 'Second layer biases do not seem right'
print ('Testing test-time forward pass ... ')
model.params['W1'] = np.linspace(-0.7, 0.3, num=D*H).reshape(D, H)
model.params['b1'] = np.linspace(-0.1, 0.9, num=H)
model.params['W2'] = np.linspace(-0.3, 0.4, num=H*C).reshape(H, C)
model.params['b2'] = np.linspace(-0.9, 0.1, num=C)
X = np.linspace(-5.5, 4.5, num=N*D).reshape(D, N).T
scores = model.loss(X)
correct_scores = np.asarray(
[[11.53165108, 12.2917344, 13.05181771, 13.81190102, 14.57198434, 15.33206765, 16.09215096],
[12.05769098, 12.74614105, 13.43459113, 14.1230412, 14.81149128, 15.49994135, 16.18839143],
[12.58373087, 13.20054771, 13.81736455, 14.43418138, 15.05099822, 15.66781506, 16.2846319 ]])
scores_diff = np.abs(scores - correct_scores).sum()
assert scores_diff < 1e-6, 'Problem with test-time forward pass'
print ('Testing training loss (no regularization)')
y = np.asarray([0, 5, 1])
loss, grads = model.loss(X, y)
correct_loss = 3.4702243556
assert abs(loss - correct_loss) < 1e-10, 'Problem with training-time loss'
model.reg = 1.0
loss, grads = model.loss(X, y)
correct_loss = 26.5948426952
assert abs(loss - correct_loss) < 1e-10, 'Problem with regularization loss'
for reg in [0.0, 0.7]:
print ('Running numeric gradient check with reg = ', reg)
model.reg = reg
loss, grads = model.loss(X, y)
for name in sorted(grads):
f = lambda _: model.loss(X, y)[0]
grad_num = eval_numerical_gradient(f, model.params[name], verbose=False)
print ('%s relative error: %.2e' % (name, rel_error(grad_num, grads[name])))
```
# Solver
In the previous assignment, the logic for training models was coupled to the models themselves. Following a more modular design, for this assignment we have split the logic for training models into a separate class.
Open the file `cs231n/solver.py` and read through it to familiarize yourself with the API. After doing so, use a `Solver` instance to train a `TwoLayerNet` that achieves at least `50%` accuracy on the validation set.
```
model = TwoLayerNet(hidden_dim = 200,reg = 0.5)
solver = Solver(model, data,
update_rule='sgd',
optim_config={
'learning_rate': 1e-3,
},
lr_decay=0.95,
num_epochs=2, batch_size=250,
print_every=100)
solver.train()
# Run this cell to visualize training loss and train / val accuracy
plt.subplot(2, 1, 1)
plt.title('Training loss')
plt.plot(solver.loss_history, 'o')
plt.xlabel('Iteration')
plt.subplot(2, 1, 2)
plt.title('Accuracy')
plt.plot(solver.train_acc_history, '-o', label='train')
plt.plot(solver.val_acc_history, '-o', label='val')
plt.plot([0.5] * len(solver.val_acc_history), 'k--')
plt.xlabel('Epoch')
plt.legend(loc='lower right')
plt.gcf().set_size_inches(15, 12)
plt.show()
```
# Multilayer network
Next you will implement a fully-connected network with an arbitrary number of hidden layers.
Read through the `FullyConnectedNet` class in the file `cs231n/classifiers/fc_net.py`.
Implement the initialization, the forward pass, and the backward pass. For the moment don't worry about implementing dropout or batch normalization; we will add those features soon.
## Initial loss and gradient check
As a sanity check, run the following to check the initial loss and to gradient check the network both with and without regularization. Do the initial losses seem reasonable?
For gradient checking, you should expect to see errors around 1e-6 or less.
```
N, D, H1, H2,H3, C = 2, 15, 20, 30,40, 10
X = np.random.randn(N, D)
y = np.random.randint(C, size=(N,))
for reg in [0, 3.14]:
print ('Running check with reg = ', reg)
model = FullyConnectedNet([H1, H2,H3], input_dim=D, num_classes=C,
reg=reg, weight_scale=5e-2, dtype=np.float64)
loss, grads = model.loss(X, y)
print ('Initial loss: ', loss)
for name in sorted(grads):
f = lambda _: model.loss(X, y)[0]
grad_num = eval_numerical_gradient(f, model.params[name], verbose=False, h=1e-5)
print ('%s relative error: %.2e' % (name, rel_error(grad_num, grads[name])))
```
As another sanity check, make sure you can overfit a small dataset of 50 images. First we will try a three-layer network with 100 units in each hidden layer. You will need to tweak the learning rate and initialization scale, but you should be able to overfit and achieve 100% training accuracy within 20 epochs.
```
# TODO: Use a three-layer Net to overfit 50 training examples.
num_train = 50
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],}
weight_scale = 1e-2
learning_rate = 1e-2
model = FullyConnectedNet([100, 100],
weight_scale=weight_scale, dtype=np.float64)
solver = Solver(model, small_data,
print_every=10, num_epochs=20, batch_size=25,
update_rule='sgd',
optim_config={
'learning_rate': learning_rate,})
solver.train()
plt.plot(solver.loss_history, 'o')
plt.title('Training loss history')
plt.xlabel('Iteration')
plt.ylabel('Training loss')
plt.show()
```
Now try to use a five-layer network with 100 units on each layer to overfit 50 training examples. Again you will have to adjust the learning rate and weight initialization, but you should be able to achieve 100% training accuracy within 20 epochs.
```
def run_model(weight_scale,learning_rate):
model = FullyConnectedNet([100, 100,100,100],
weight_scale=weight_scale, dtype=np.float64)
solver = Solver(model, small_data,
print_every=10, num_epochs=20, batch_size=25,
update_rule='sgd',
optim_config={
'learning_rate': learning_rate,
}
)
solver.train()
return solver.train_acc_history
np.random.uniform(-4,4)
# TODO: Use a five-layer Net to overfit 50 training examples.
num_train = 50
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],
}
not_reach = True
while not_reach:
weight_scale = 10**(np.random.uniform(-6,-1))
learning_rate = 10**(np.random.uniform(-4,-1))
train_acc_hist = run_model(weight_scale,learning_rate)
if max(train_acc_hist) == 1.0:
not_reach = False
lr = learning_rate
ws = weight_scale
print ('Has worked with %f and %f'%(lr,ws))
plt.plot(solver.loss_history, 'o')
plt.title('Training loss history')
plt.xlabel('Iteration')
plt.ylabel('Training loss')
plt.show()
num_train = 50
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],
}
run_model(ws,lr)
print ('Has worked with %f and %f'%(lr,ws))
plt.plot(solver.loss_history, 'o')
plt.title('Training loss history')
plt.xlabel('Iteration')
plt.ylabel('Training loss')
plt.show()
```
# Inline question:
Did you notice anything about the comparative difficulty of training the three-layer net vs training the five layer net?
# Update rules
So far we have used vanilla stochastic gradient descent (SGD) as our update rule. More sophisticated update rules can make it easier to train deep networks. We will implement a few of the most commonly used update rules and compare them to vanilla SGD.
# SGD+Momentum
Stochastic gradient descent with momentum is a widely used update rule that tends to make deep networks converge faster than vanilla stochstic gradient descent.
Open the file `cs231n/optim.py` and read the documentation at the top of the file to make sure you understand the API. Implement the SGD+momentum update rule in the function `sgd_momentum` and run the following to check your implementation. You should see errors less than 1e-8.
```
from cs231n.optim import sgd_momentum
N, D = 4, 5
w = np.linspace(-0.4, 0.6, num=N*D).reshape(N, D)
dw = np.linspace(-0.6, 0.4, num=N*D).reshape(N, D)
v = np.linspace(0.6, 0.9, num=N*D).reshape(N, D)
config = {'learning_rate': 1e-3, 'velocity': v}
next_w, _ = sgd_momentum(w, dw, config=config)
expected_next_w = np.asarray([
[ 0.1406, 0.20738947, 0.27417895, 0.34096842, 0.40775789],
[ 0.47454737, 0.54133684, 0.60812632, 0.67491579, 0.74170526],
[ 0.80849474, 0.87528421, 0.94207368, 1.00886316, 1.07565263],
[ 1.14244211, 1.20923158, 1.27602105, 1.34281053, 1.4096 ]])
expected_velocity = np.asarray([
[ 0.5406, 0.55475789, 0.56891579, 0.58307368, 0.59723158],
[ 0.61138947, 0.62554737, 0.63970526, 0.65386316, 0.66802105],
[ 0.68217895, 0.69633684, 0.71049474, 0.72465263, 0.73881053],
[ 0.75296842, 0.76712632, 0.78128421, 0.79544211, 0.8096 ]])
print( 'next_w error: ', rel_error(next_w, expected_next_w))
print ('velocity error: ', rel_error(expected_velocity, config['velocity']))
```
Once you have done so, run the following to train a six-layer network with both SGD and SGD+momentum. You should see the SGD+momentum update rule converge faster.
```
num_train = 4000
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],
}
solvers = {}
for update_rule in ['sgd', 'sgd_momentum']:
print ('running with ', update_rule)
model = FullyConnectedNet([100, 100, 100, 100, 100], weight_scale=5e-2)
solver = Solver(model, small_data,
num_epochs=5, batch_size=100,
update_rule=update_rule,
optim_config={
'learning_rate': 1e-2,
},
verbose=True)
solvers[update_rule] = solver
solver.train()
print()
plt.subplot(3, 1, 1)
plt.title('Training loss')
plt.xlabel('Iteration')
plt.subplot(3, 1, 2)
plt.title('Training accuracy')
plt.xlabel('Epoch')
plt.subplot(3, 1, 3)
plt.title('Validation accuracy')
plt.xlabel('Epoch')
for update_rule, solver in solvers.items():
plt.subplot(3, 1, 1)
plt.plot(solver.loss_history, 'o', label=update_rule)
plt.subplot(3, 1, 2)
plt.plot(solver.train_acc_history, '-o', label=update_rule)
plt.subplot(3, 1, 3)
plt.plot(solver.val_acc_history, '-o', label=update_rule)
for i in [1, 2, 3]:
plt.subplot(3, 1, i)
plt.legend(loc='upper center', ncol=4)
plt.gcf().set_size_inches(15, 15)
plt.show()
```
# RMSProp and Adam
RMSProp [1] and Adam [2] are update rules that set per-parameter learning rates by using a running average of the second moments of gradients.
In the file `cs231n/optim.py`, implement the RMSProp update rule in the `rmsprop` function and implement the Adam update rule in the `adam` function, and check your implementations using the tests below.
[1] Tijmen Tieleman and Geoffrey Hinton. "Lecture 6.5-rmsprop: Divide the gradient by a running average of its recent magnitude." COURSERA: Neural Networks for Machine Learning 4 (2012).
[2] Diederik Kingma and Jimmy Ba, "Adam: A Method for Stochastic Optimization", ICLR 2015.
```
# Test RMSProp implementation; you should see errors less than 1e-7
from cs231n.optim import rmsprop
N, D = 4, 5
w = np.linspace(-0.4, 0.6, num=N*D).reshape(N, D)
dw = np.linspace(-0.6, 0.4, num=N*D).reshape(N, D)
cache = np.linspace(0.6, 0.9, num=N*D).reshape(N, D)
config = {'learning_rate': 1e-2, 'cache': cache}
next_w, _ = rmsprop(w, dw, config=config)
expected_next_w = np.asarray([
[-0.39223849, -0.34037513, -0.28849239, -0.23659121, -0.18467247],
[-0.132737, -0.08078555, -0.02881884, 0.02316247, 0.07515774],
[ 0.12716641, 0.17918792, 0.23122175, 0.28326742, 0.33532447],
[ 0.38739248, 0.43947102, 0.49155973, 0.54365823, 0.59576619]])
expected_cache = np.asarray([
[ 0.5976, 0.6126277, 0.6277108, 0.64284931, 0.65804321],
[ 0.67329252, 0.68859723, 0.70395734, 0.71937285, 0.73484377],
[ 0.75037008, 0.7659518, 0.78158892, 0.79728144, 0.81302936],
[ 0.82883269, 0.84469141, 0.86060554, 0.87657507, 0.8926 ]])
print ('next_w error: ', rel_error(expected_next_w, next_w))
print ('cache error: ', rel_error(expected_cache, config['cache']))
# Test Adam implementation; you should see errors around 1e-7 or less
from cs231n.optim import adam
N, D = 4, 5
w = np.linspace(-0.4, 0.6, num=N*D).reshape(N, D)
dw = np.linspace(-0.6, 0.4, num=N*D).reshape(N, D)
m = np.linspace(0.6, 0.9, num=N*D).reshape(N, D)
v = np.linspace(0.7, 0.5, num=N*D).reshape(N, D)
config = {'learning_rate': 1e-2, 'm': m, 'v': v, 't': 5}
next_w, _ = adam(w, dw, config=config)
expected_next_w = np.asarray([
[-0.40094747, -0.34836187, -0.29577703, -0.24319299, -0.19060977],
[-0.1380274, -0.08544591, -0.03286534, 0.01971428, 0.0722929],
[ 0.1248705, 0.17744702, 0.23002243, 0.28259667, 0.33516969],
[ 0.38774145, 0.44031188, 0.49288093, 0.54544852, 0.59801459]])
expected_v = np.asarray([
[ 0.69966, 0.68908382, 0.67851319, 0.66794809, 0.65738853,],
[ 0.64683452, 0.63628604, 0.6257431, 0.61520571, 0.60467385,],
[ 0.59414753, 0.58362676, 0.57311152, 0.56260183, 0.55209767,],
[ 0.54159906, 0.53110598, 0.52061845, 0.51013645, 0.49966, ]])
expected_m = np.asarray([
[ 0.48, 0.49947368, 0.51894737, 0.53842105, 0.55789474],
[ 0.57736842, 0.59684211, 0.61631579, 0.63578947, 0.65526316],
[ 0.67473684, 0.69421053, 0.71368421, 0.73315789, 0.75263158],
[ 0.77210526, 0.79157895, 0.81105263, 0.83052632, 0.85 ]])
print ('next_w error: ', rel_error(expected_next_w, next_w))
print ('v error: ', rel_error(expected_v, config['v']))
print('m error: ', rel_error(expected_m, config['m']))
```
Once you have debugged your RMSProp and Adam implementations, run the following to train a pair of deep networks using these new update rules:
```
learning_rates = {'rmsprop': 1e-4, 'adam': 1e-3}
for update_rule in ['adam', 'rmsprop']:
print ('running with ', update_rule)
model = FullyConnectedNet([100, 100, 100, 100, 100], weight_scale=5e-2)
solver = Solver(model, small_data,
num_epochs=10, batch_size=100,
update_rule=update_rule,
optim_config={
'learning_rate': learning_rates[update_rule]
},
verbose=True)
solvers[update_rule] = solver
solver.train()
print()
plt.subplot(3, 1, 1)
plt.title('Training loss')
plt.xlabel('Iteration')
plt.subplot(3, 1, 2)
plt.title('Training accuracy')
plt.xlabel('Epoch')
plt.subplot(3, 1, 3)
plt.title('Validation accuracy')
plt.xlabel('Epoch')
for update_rule, solver in solvers.items():
plt.subplot(3, 1, 1)
plt.plot(solver.loss_history, 'o', label=update_rule)
plt.subplot(3, 1, 2)
plt.plot(solver.train_acc_history, '-o', label=update_rule)
plt.subplot(3, 1, 3)
plt.plot(solver.val_acc_history, '-o', label=update_rule)
for i in [1, 2, 3]:
plt.subplot(3, 1, i)
plt.legend(loc='upper center', ncol=4)
plt.gcf().set_size_inches(15, 15)
plt.show()
```
# Train a good model!
Train the best fully-connected model that you can on CIFAR-10, storing your best model in the `best_model` variable. We require you to get at least 50% accuracy on the validation set using a fully-connected net.
If you are careful it should be possible to get accuracies above 55%, but we don't require it for this part and won't assign extra credit for doing so. Later in the assignment we will ask you to train the best convolutional network that you can on CIFAR-10, and we would prefer that you spend your effort working on convolutional nets rather than fully-connected nets.
You might find it useful to complete the `BatchNormalization.ipynb` and `Dropout.ipynb` notebooks before completing this part, since those techniques can help you train powerful models.
```
X_train = data['X_train']
y_train = data['y_train']
X_val = data['X_val']
y_val = data['y_val']
X_test = data['X_test']
y_test = data['y_test']
best_model = None
################################################################################
# TODO: Train the best FullyConnectedNet that you can on CIFAR-10. You might #
# batch normalization and dropout useful. Store your best model in the #
# best_model variable. #
################################################################################
best_model = FullyConnectedNet([100, 100, 100, 100], weight_scale=1e-1, reg=0.01, use_batchnorm=True)
solver = Solver(best_model, small_data, num_epochs=1000, batch_size=256, update_rule='adam',
optim_config={'learning_rate': 1e-3}, verbose=True)
solver.train()
################################################################################
# END OF YOUR CODE #
################################################################################
```
# Test you model
Run your best model on the validation and test sets. You should achieve above 50% accuracy on the validation set.
```
y_test_pred = np.argmax(best_model.loss(data['X_test']), axis=1)
y_val_pred = np.argmax(best_model.loss(data['X_val']), axis=1)
print ('Validation set accuracy: ', (y_val_pred == y_val).mean())
print ('Test set accuracy: ', (y_test_pred == y_test).mean())
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.