code stringlengths 2.5k 150k | kind stringclasses 1 value |
|---|---|
# Домашняя работа №4
```
pip install scikit-uplift
pip install causalml
pip install catboost
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import itertools
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import cross_val_score, train_test_split
from sklearn.metrics import precision_recall_curve, roc_curve, roc_auc_score, confusion_matrix, log_loss
from sklearn.pipeline import Pipeline, make_pipeline, FeatureUnion
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.preprocessing import StandardScaler
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier
from sklift.metrics import uplift_at_k
from sklift.viz import plot_uplift_preds
from sklift.models import SoloModel, TwoModels, ClassTransformation
from IPython.display import Image
from causalml.inference.tree import UpliftTreeClassifier, UpliftRandomForestClassifier
from causalml.inference.tree import uplift_tree_string, uplift_tree_plot
from catboost import CatBoostClassifier
from scipy.sparse import hstack
from google.colab import drive
drive.mount('/gdrive')
df = pd.read_csv('/content/data.csv')
df.head()
df.channel.value_counts()
df.zip_code.value_counts()
df.recency.value_counts()
df.offer.value_counts()
df.rename(columns={df.columns[-1]: 'target'}, inplace=True)
df.rename(columns={df.columns[-2]: 'treatment'}, inplace=True)
df.head()
df.treatment
df.loc[df['treatment'] == 'Buy One Get One', 'treatment'] = 1
df.loc[df['treatment'] == 'Discount', 'treatment'] = 1
df.loc[df['treatment'] == 'No Offer', 'treatment'] = 0
df.head()
X_train_df, X_test_df, y_train_df, y_test_df = train_test_split(df.drop(['target'], 1),
df.target, random_state=15, test_size=0.3,)
X_train_t = X_train_df.drop('treatment', 1)
indices_train = X_train_df.index
indices_test = X_test_df.index
indices_learn, indices_valid = train_test_split(X_train_df.index, test_size=0.3, random_state=123)
X_train_df.shape
X_train_df
X_train = X_train_t.loc[indices_learn, :]
y_train = y_train_df.loc[indices_learn]
treat_train = X_train_df.loc[indices_learn, 'treatment']
X_val = X_train_t.loc[indices_valid, :]
y_val = y_train_df.loc[indices_valid]
treat_val = X_train_df.loc[indices_valid, 'treatment']
X_train_full = X_train_t.loc[indices_train, :]
y_train_full = y_train_df.loc[:]
treat_train_full = X_train_df.loc[:, 'treatment']
X_test = X_test_df.loc[indices_test, :]
cat_features = ['zip_code', 'channel', 'recency']
models_results = {
'approach': [],
'uplift@20%': [],
'uplift@10%': []
}
cat_features = ['zip_code', 'channel', 'recency']
sm = SoloModel(CatBoostClassifier(iterations=20, thread_count=2, random_state=15, silent=True))
sm = sm.fit(X_train, y_train, treat_train, estimator_fit_params={'cat_features': cat_features})
uplift_sm = sm.predict(X_val)
sm_score = uplift_at_k(y_true=y_val, uplift=uplift_sm, treatment=treat_val, strategy='by_group', k=0.2)
models_results['approach'].append('SoloModel')
models_results['uplift@20%'].append(sm_score)
ct = ClassTransformation(CatBoostClassifier(iterations=20, thread_count=2, random_state=15, silent=True))
ct = ct.fit(X_train, y_train, treat_train, estimator_fit_params={'cat_features': cat_features})
uplift_ct = ct.predict(X_val)
ct_score = uplift_at_k(y_true=y_val, uplift=uplift_ct, treatment=treat_val, strategy='by_group', k=0.2)
models_results['approach'].append('ClassTransformation')
models_results['uplift@20%'].append(ct_score)
tm = TwoModels(
estimator_trmnt=CatBoostClassifier(iterations=20, thread_count=2, random_state=15, silent=True),
estimator_ctrl=CatBoostClassifier(iterations=20, thread_count=2, random_state=15, silent=True),
method='vanilla'
)
tm = tm.fit(
X_train, y_train, treat_train,
estimator_trmnt_fit_params={'cat_features': cat_features},
estimator_ctrl_fit_params={'cat_features': cat_features}
)
uplift_tm = tm.predict(X_val)
tm_score = uplift_at_k(y_true=y_val, uplift=uplift_tm, treatment=treat_val, strategy='by_group', k=0.2)
models_results['approach'].append('TwoModels')
models_results['uplift@20%'].append(tm_score)
sm = SoloModel(CatBoostClassifier(iterations=20, thread_count=2, random_state=15, silent=True))
sm = sm.fit(X_train, y_train, treat_train, estimator_fit_params={'cat_features': cat_features})
uplift_sm = sm.predict(X_val)
sm_score = uplift_at_k(y_true=y_val, uplift=uplift_sm, treatment=treat_val, strategy='by_group', k=0.1)
models_results['uplift@10%'].append(sm_score)
ct = ClassTransformation(CatBoostClassifier(iterations=20, thread_count=2, random_state=15, silent=True))
ct = ct.fit(X_train, y_train, treat_train, estimator_fit_params={'cat_features': cat_features})
uplift_ct = ct.predict(X_val)
ct_score = uplift_at_k(y_true=y_val, uplift=uplift_ct, treatment=treat_val, strategy='by_group', k=0.1)
models_results['uplift@10%'].append(ct_score)
tm = TwoModels(
estimator_trmnt=CatBoostClassifier(iterations=20, thread_count=2, random_state=15, silent=True),
estimator_ctrl=CatBoostClassifier(iterations=20, thread_count=2, random_state=15, silent=True),
method='vanilla'
)
tm = tm.fit(
X_train, y_train, treat_train,
estimator_trmnt_fit_params={'cat_features': cat_features},
estimator_ctrl_fit_params={'cat_features': cat_features}
)
uplift_tm = tm.predict(X_val)
tm_score = uplift_at_k(y_true=y_val, uplift=uplift_tm, treatment=treat_val, strategy='by_group', k=0.1)
models_results['uplift@10%'].append(tm_score)
models_results
table = pd.DataFrame(models_results)
table
zip_dummie = pd.get_dummies(X_train.zip_code, prefix='zip')
X_train = X_train.join(zip_dummie).drop('zip_code', 1)
channel_dummie = pd.get_dummies(X_train.channel, prefix='channel')
X_train.join(channel_dummie).drop('channel', 1)
X_train_tree = X_train.copy()
features = [col for col in X_train_tree]
X_train_tree.shape
uplift_model = UpliftTreeClassifier(max_depth=4, min_samples_leaf=200, min_samples_treatment=50,
n_reg=100, evaluationFunction='KL', control_name='control')
uplift_model.fit(X_train_tree.values,
treatment=treat_train.map({1: 'treatment1', 0: 'control'}).values,
y=y_train)
graph = uplift_tree_plot(uplift_model.fitted_uplift_tree, features)
Image(graph.create_png())
uplift_model = UpliftTreeClassifier(max_depth=3, min_samples_leaf=200, min_samples_treatment=50,
n_reg=100, evaluationFunction='KL', control_name='control')
uplift_model.fit(X_train_tree.values,
treatment=treat_train.map({1: 'treatment1', 0: 'control'}).values,
y=y_train)
graph = uplift_tree_plot(uplift_model.fitted_uplift_tree, features)
Image(graph.create_png())
```
Самый левый лист - 2570 человек, которые использовали скидку и у которых zip-code Rural. Им нельзя звонить!
У всех остальных p_value = 0, это означает, что они положительно реагируют на взаимодействия с ними.
```
```
| github_jupyter |
# Working with Python: functions and modules
## Session 4: Using third party libraries
- [Matplotlib](#Matplotlib)
- [Exercise 4.1](#Exercise-4.1)
- [BioPython](#BioPython)
- [Working with sequences](#Working-with-sequences)
- [Connecting with biological databases](#Connecting-with-biological-databases)
- [Exercise 4.2](#Exercise-4.2)
## Matplotlib
[matplotlib](http://matplotlib.org/) is probably the single most used Python package for graphics. It provides both a very quick way to visualize data from Python and publication-quality figures in many formats.
matplotlib.pyplot is a collection of command style functions that make matplotlib work like MATLAB. Each pyplot function makes some change to a figure: e.g., creates a figure, creates a plotting area in a figure, plots some lines in a plotting area, decorates the plot with labels, etc.
Let's start with a very simple plot.
```
import matplotlib.pyplot as mpyplot
mpyplot.plot([1,2,3,4])
mpyplot.ylabel('some numbers')
mpyplot.show()
```
`plot()` is a versatile command, and will take an arbitrary number of arguments. For example, to plot x versus y, you can issue the command:
```
mpyplot.plot([1,2,3,4], [1,4,9,16])
```
For every x, y pair of arguments, there is an **optional third argument** which is the format string that indicates the color and line type of the plot. The letters and symbols of the format string are from MATLAB, and you concatenate a color string with a line style string. The default format string is `b-`, which is a solid blue line. For example, to plot the above with red circles, you would chose `ro`.
```
import matplotlib.pyplot as mpyplot
mpyplot.plot([1,2,3,4], [1,4,9,16], 'ro')
mpyplot.axis([0, 6, 0, 20])
mpyplot.show()
```
`matplotlib` has a few methods in the **`pyplot` module** that make creating common types of plots faster and more convenient because they automatically create a Figure and an Axes object. The most widely used are:
- [mpyplot.bar](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.bar) – creates a bar chart.
- [mpyplot.boxplot](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.boxplot) – makes a box and whisker plot.
- [mpyplot.hist](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.hist) – makes a histogram.
- [mpyplot.plot](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.plot) – creates a line plot.
- [mpyplot.scatter](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.scatter) – makes a scatter plot.
Calling any of these methods will automatically setup `Figure` and `Axes` objects, and draw the plot. Each of these methods has different parameters that can be passed in to modify the resulting plot.
The [Pyplot tutorial](http://matplotlib.org/users/pyplot_tutorial.html) is where these simple examples above are coming from. More could be learn from it if you wish during your own time.
Let's now try to plot the GC content along the chain we have calculated during the previous session, while solving the Exercises 3.3 and 3.4.
```
seq = 'ATGGTGCATCTGACTCCTGAGGAGAAGTCTGCCGTTACTGCCCTGTGGGGCAAGGTG'
gc = [40.0, 60.0, 80.0, 60.0, 40.0, 60.0, 40.0, 40.0, 40.0, 60.0,
40.0, 60.0, 60.0, 60.0, 60.0, 60.0, 60.0, 60.0, 60.0, 60.0,
60.0, 40.0, 40.0, 40.0, 40.0, 40.0, 60.0, 60.0, 80.0, 80.0,
80.0, 60.0, 40.0, 40.0, 20.0, 40.0, 60.0, 80.0, 80.0, 80.0,
80.0, 60.0, 60.0, 60.0, 80.0, 80.0, 100.0, 80.0, 60.0, 60.0,
60.0, 40.0, 60.0]
window_ids = range(len(gc))
import matplotlib.pyplot as mpyplot
mpyplot.plot(window_ids, gc, '--' )
mpyplot.xlabel('5 bases window id along the sequence')
mpyplot.ylabel('%GC')
mpyplot.title('GC plot for sequence\n' + seq)
mpyplot.show()
```
## Exercise 4.1
Re-use the GapMinder dataset to plot, in Jupyter using Matplotlib, from the world data the life expectancy against GDP per capita for 1957 and 2007 using a scatter plot, add title to your graph as well as a legend.
## BioPython
The goal of Biopython is to make it as easy as possible to use Python for bioinformatics by creating high-quality, reusable modules and classes. Biopython features include parsers for various Bioinformatics file formats (BLAST, Clustalw, FASTA, Genbank,...), access to online services (NCBI, Expasy,...), interfaces to common and not-so-common programs (Clustalw, DSSP, MSMS...), a standard sequence class, various clustering modules, a KD tree data structure etc. and documentation as well as a tutorial: http://biopython.org/DIST/docs/tutorial/Tutorial.html.
## Working with sequences
We can create a sequence by defining a `Seq` object with strings. `Bio.Seq()` takes as input a string and converts in into a Seq object. We can print the sequences, individual residues, lengths and use other functions to get summary statistics.
```
# Creating sequence
from Bio.Seq import Seq
my_seq = Seq("AGTACACTGGT")
print(my_seq)
print(my_seq[10])
print(my_seq[1:5])
print(len(my_seq))
print(my_seq.count("A"))
```
We can use functions from `Bio.SeqUtils` to get idea about a sequence
```
# Calculate the molecular weight
from Bio.SeqUtils import GC, molecular_weight
print(GC(my_seq))
print(molecular_weight(my_seq))
```
One letter code protein sequences can be converted into three letter codes using `seq3` utility
```
from Bio.SeqUtils import seq3
print(seq3(my_seq))
```
Alphabets defines how the strings are going to be treated as sequence object. `Bio.Alphabet` module defines the available alphabets for Biopython. `Bio.Alphabet.IUPAC` provides basic definition for DNA, RNA and proteins.
```
from Bio.Alphabet import IUPAC
my_dna = Seq("AGTACATGACTGGTTTAG", IUPAC.unambiguous_dna)
print(my_dna)
print(my_dna.alphabet)
my_dna.complement()
my_dna.reverse_complement()
my_dna.translate()
```
### Parsing sequence file format: FASTA files
Sequence files can be parsed and read the same way we read other files.
```
with open( "data/glpa.fa" ) as f:
print(f.read())
```
Biopython provides specific functions to allow parsing/reading sequence files.
```
# Reading FASTA files
from Bio import SeqIO
with open("data/glpa.fa") as f:
for protein in SeqIO.parse(f, 'fasta'):
print(protein.id)
print(protein.seq)
```
Sequence objects can be written into files using file handles with the function `SeqIO.write()`. We need to provide the name of the output sequence file and the sequence file format.
```
# Writing FASTA files
from Bio import SeqIO
from Bio.SeqRecord import SeqRecord
from Bio.Seq import Seq
from Bio.Alphabet import IUPAC
sequence = 'MYGKIIFVLLLSEIVSISASSTTGVAMHTSTSSSVTKSYISSQTNDTHKRDTYAATPRAHEVSEISVRTVYPPEEETGERVQLAHHFSEPEITLIIFG'
seq = Seq(sequence, IUPAC.protein)
protein = [SeqRecord(seq, id="THEID", description='a description'),]
with open( "biopython.fa", "w") as f:
SeqIO.write(protein, f, 'fasta')
with open( "biopython.fa" ) as f:
print(f.read())
```
## Connecting with biological databases
Sequences can be searched and downloaded from public databases.
```
# Read FASTA file from NCBI GenBank
from Bio import Entrez
Entrez.email = 'A.N.Other@example.com' # Always tell NCBI who you are
handle = Entrez.efetch(db="nucleotide", id="71066805", rettype="gb")
seq_record = SeqIO.read(handle, "gb")
handle.close()
print(seq_record.id, 'with', len(seq_record.features), 'features')
print(seq_record.seq)
print(seq_record.format("fasta"))
# Read SWISSPROT record
from Bio import ExPASy
handle = ExPASy.get_sprot_raw('HBB_HUMAN')
prot_record = SeqIO.read(handle, "swiss")
handle.close()
print(prot_record.description)
print(prot_record.seq)
```
## Exercise 4.2
- Retrieve a FASTA file named `data/sample.fa` using BioPython and answer the following questions:
- How many sequences are in the file?
- What are the IDs and the lengths of the longest and the shortest sequences?
- Select sequences longer than 500bp. What is the average length of these sequences?
- Calculate and print the percentage of GC in each of the sequences.
- Write the newly created sequences into a FASTA file named `long_sequences.fa`
## Congratulation! You reached the end!
| github_jupyter |
# 02_hsn_v1_lean-voc2012
```
import time
import skimage.io as imgio
import pandas as pd
import numpy.matlib
from adp_cues import ADPCues
from utilities import *
from dataset import Dataset
MODEL_CNN_ROOT = '../database/models_cnn'
MODEL_WSSS_ROOT = '../database/models_wsss'
dataset = 'VOC2012'
model_type = 'VGG16'
batch_size = 16
sess_id = dataset + '_' + model_type
if model_type in ['VGG16', 'VGG16bg']:
size = 321
else:
size = 224
should_saveimg = False
is_verbose = True
if model_type in ['VGG16', 'VGG16bg']:
img_size = 321
else:
img_size = 224
sess_id = dataset + '_' + model_type
model_dir = os.path.join(MODEL_CNN_ROOT, sess_id)
if is_verbose:
print('Predict: dataset=' + dataset + ', model=' + model_type)
database_dir = os.path.join(os.path.dirname(os.getcwd()), 'database')
if dataset == 'VOC2012':
devkit_dir = os.path.join(database_dir, 'VOCdevkit', 'VOC2012')
fgbg_modes = ['fg', 'bg']
OVERLAY_R = 0.75
elif 'DeepGlobe' in dataset:
devkit_dir = os.path.join(database_dir, 'DGdevkit')
fgbg_modes = ['fg']
OVERLAY_R = 0.25
img_dir = os.path.join(devkit_dir, 'JPEGImages')
gt_dir = os.path.join(devkit_dir, 'SegmentationClassAug')
out_dir = os.path.join('./out', sess_id)
if not os.path.exists(out_dir):
os.makedirs(out_dir)
eval_dir = os.path.join('./eval', sess_id)
if not os.path.exists(eval_dir):
os.makedirs(eval_dir)
```
## Load network and data
```
# Load network and thresholds
mdl = {}
thresholds = {}
alpha = {}
final_layer = {}
for fgbg_mode in fgbg_modes:
mdl[fgbg_mode] = build_model(model_dir, sess_id)
thresholds[fgbg_mode] = load_thresholds(model_dir, sess_id)
thresholds[fgbg_mode] = np.maximum(np.minimum(thresholds[fgbg_mode], 0), 1 / 3)
alpha[fgbg_mode], final_layer[fgbg_mode] = get_grad_cam_weights(mdl[fgbg_mode],
np.zeros((1, img_size, img_size, 3)))
# Load data and classes
ds = Dataset(data_type=dataset, size=img_size, batch_size=batch_size)
class_names, seg_class_names = load_classes(dataset)
colours = get_colours(dataset)
if 'DeepGlobe' in dataset:
colours = colours[:-1]
gen_curr = ds.set_gens[ds.sets[ds.is_evals.index(True)]]
```
## Generate segmentations for single batch
```
# Process images in batches
intersects = np.zeros((len(colours)))
unions = np.zeros((len(colours)))
confusion_matrix = np.zeros((len(colours), len(colours)))
gt_count = np.zeros((len(colours)))
n_batches = 1
for iter_batch in range(n_batches):
batch_start_time = time.time()
if is_verbose:
print('\tBatch #%d of %d' % (iter_batch + 1, n_batches))
start_idx = iter_batch * batch_size
end_idx = min(start_idx + batch_size - 1, len(gen_curr.filenames) - 1)
cur_batch_sz = end_idx - start_idx + 1
# Image reading
start_time = time.time()
img_batch_norm, img_batch = read_batch(gen_curr.directory, gen_curr.filenames[start_idx:end_idx + 1],
cur_batch_sz, (img_size, img_size), dataset)
if is_verbose:
print('\t\tImage read time: %0.5f seconds (%0.5f seconds / image)' % (time.time() - start_time,
(time.time() - start_time) / cur_batch_sz))
# Generate patch confidence scores
start_time = time.time()
predicted_scores = {}
is_pass_threshold = {}
for fgbg_mode in fgbg_modes:
predicted_scores[fgbg_mode] = mdl[fgbg_mode].predict(img_batch_norm)
is_pass_threshold[fgbg_mode] = np.greater_equal(predicted_scores[fgbg_mode], thresholds[fgbg_mode])
if is_verbose:
print('\t\tGenerating patch confidence scores time: %0.5f seconds (%0.5f seconds / image)' %
(time.time() - start_time, (time.time() - start_time) / cur_batch_sz))
# Generate Grad-CAM
start_time = time.time()
H = {}
for fgbg_mode in fgbg_modes:
H[fgbg_mode] = grad_cam(mdl[fgbg_mode], alpha[fgbg_mode], img_batch_norm, is_pass_threshold[fgbg_mode],
final_layer[fgbg_mode], predicted_scores[fgbg_mode], orig_sz=[img_size, img_size],
should_upsample=True)
H[fgbg_mode] = np.transpose(H[fgbg_mode], (0, 3, 1, 2))
if is_verbose:
print('\t\tGenerating Grad-CAM time: %0.5f seconds (%0.5f seconds / image)' % (time.time() - start_time,
(time.time() - start_time) / cur_batch_sz))
# Modify fg Grad-CAM with bg activation
start_time = time.time()
if dataset == 'VOC2012':
Y_gradcam = np.zeros((cur_batch_sz, len(seg_class_names), img_size, img_size))
mode = 'mult'
if mode == 'mult':
X_bg = np.sum(H['bg'], axis=1)
Y_gradcam[:, 0] = 0.15 * scipy.special.expit(np.max(X_bg) - X_bg)
Y_gradcam[:, 1:] = H['fg']
elif 'DeepGlobe' in dataset:
Y_gradcam = H['fg'][:, :-1, :, :]
if is_verbose:
print('\t\tFg/Bg modifications time: %0.5f seconds (%0.5f seconds / image)' % (time.time() - start_time,
(time.time() - start_time) / cur_batch_sz))
# FC-CRF
start_time = time.time()
if dataset == 'VOC2012':
dcrf_config = np.array([3 / 4, 3, 80 / 4, 13, 10, 10]) # test (since 2448 / 500 = 4.896 ~= 4)
elif 'DeepGlobe' in dataset:
dcrf_config = np.array([3, 3, 80, 13, 10, 10]) # test
Y_crf = dcrf_process(Y_gradcam, img_batch, dcrf_config)
if is_verbose:
print('\t\tCRF time: %0.5f seconds (%0.5f seconds / image)' % (time.time() - start_time,
(time.time() - start_time) / cur_batch_sz))
elapsed_time = time.time() - batch_start_time
if is_verbose:
print('\t\tElapsed time: %0.5f seconds (%0.5f seconds / image)' % (elapsed_time, elapsed_time / cur_batch_sz))
if dataset == 'VOC2012':
for iter_file, filename in enumerate(gen_curr.filenames[start_idx:end_idx + 1]):
# Load GT segmentation
gt_filepath = os.path.join(gt_dir, filename.replace('.jpg', '.png'))
gt_idx = cv2.cvtColor(cv2.imread(gt_filepath), cv2.COLOR_BGR2RGB)[:, :, 0]
# Load predicted segmentation
pred_idx = cv2.resize(np.uint8(Y_crf[iter_file]), (gt_idx.shape[1], gt_idx.shape[0]),
interpolation=cv2.INTER_NEAREST)
pred_segmask = np.zeros((gt_idx.shape[0], gt_idx.shape[1], 3))
# Evaluate predicted segmentation
for k in range(len(colours)):
intersects[k] += np.sum((gt_idx == k) & (pred_idx == k))
unions[k] += np.sum((gt_idx == k) | (pred_idx == k))
confusion_matrix[k, :] += np.bincount(pred_idx[gt_idx == k], minlength=len(colours))
pred_segmask += np.expand_dims(pred_idx == k, axis=2) * \
np.expand_dims(np.expand_dims(colours[k], axis=0), axis=0)
gt_count[k] += np.sum(gt_idx == k)
# Save outputted segmentation to file
if should_saveimg:
orig_filepath = os.path.join(img_dir, filename)
orig_img = cv2.cvtColor(cv2.imread(orig_filepath), cv2.COLOR_BGR2RGB)
imgio.imsave(os.path.join(out_dir, filename.replace('.jpg', '') + '.png'), pred_segmask / 256.0)
imgio.imsave(os.path.join(out_dir, filename.replace('.jpg', '') + '_overlay.png'),
(1 - OVERLAY_R) * orig_img / 256.0 +
OVERLAY_R * pred_segmask / 256.0)
elif 'DeepGlobe' in dataset:
for iter_file, filename in enumerate(gen_curr.filenames[start_idx:end_idx + 1]):
# Load GT segmentation
gt_filepath = os.path.join(gt_dir, filename.replace('.jpg', '.png'))
gt_curr = cv2.cvtColor(cv2.imread(gt_filepath), cv2.COLOR_BGR2RGB)
gt_r = gt_curr[:, :, 0]
gt_g = gt_curr[:, :, 1]
gt_b = gt_curr[:, :, 2]
# Load predicted segmentation
pred_idx = cv2.resize(np.uint8(Y_crf[iter_file]), (gt_curr.shape[1], gt_curr.shape[0]),
interpolation=cv2.INTER_NEAREST)
pred_segmask = np.zeros((gt_curr.shape[0], gt_curr.shape[1], 3))
# Evaluate predicted segmentation
for k, gt_colour in enumerate(colours):
gt_mask = (gt_r == gt_colour[0]) & (gt_g == gt_colour[1]) & (gt_b == gt_colour[2])
pred_mask = pred_idx == k
intersects[k] += np.sum(gt_mask & pred_mask)
unions[k] += np.sum(gt_mask | pred_mask)
confusion_matrix[k, :] += np.bincount(pred_idx[gt_mask], minlength=len(colours))
pred_segmask += np.expand_dims(pred_mask, axis=2) * \
np.expand_dims(np.expand_dims(colours[k], axis=0), axis=0)
gt_count[k] += np.sum(gt_mask)
# Save outputted segmentation to file
if should_saveimg:
orig_filepath = os.path.join(img_dir, filename)
orig_img = cv2.cvtColor(cv2.imread(orig_filepath), cv2.COLOR_BGR2RGB)
orig_img = cv2.resize(orig_img, (orig_img.shape[0] // 4, orig_img.shape[1] // 4))
pred_segmask = cv2.resize(pred_segmask, (pred_segmask.shape[0] // 4, pred_segmask.shape[1] // 4),
interpolation=cv2.INTER_NEAREST)
imgio.imsave(os.path.join(out_dir, filename.replace('.jpg', '') + '.png'), pred_segmask / 256.0)
imgio.imsave(os.path.join(out_dir, filename.replace('.jpg', '') + '_overlay.png'),
(1 - OVERLAY_R) * orig_img / 256.0 + OVERLAY_R * pred_segmask / 256.0)
```
## Show sample segmentations
```
img_filepath = os.path.join(gen_curr.directory, gen_curr.filenames[0])
I = cv2.cvtColor(cv2.imread(img_filepath), cv2.COLOR_BGR2RGB)
gt_filepath = os.path.join(gt_dir, gen_curr.filenames[0].replace('.jpg', '.png'))
gt_idx = np.expand_dims(cv2.cvtColor(cv2.imread(gt_filepath), cv2.COLOR_BGR2RGB)[:, :, 0], axis=0)
G = maxconf_class_as_colour(gt_idx, colours, gt_idx.shape[1:3])
plt.figure
plt.subplot(121)
plt.imshow(I.astype('uint8'))
plt.title('Original image')
plt.subplot(122)
plt.imshow(G[0].astype('uint8'))
plt.title('Ground truth\n segmentation')
# Load predicted segmentation
pred_idx = cv2.resize(np.uint8(Y_crf[0]), (gt_idx.shape[2], gt_idx.shape[1]), interpolation=cv2.INTER_NEAREST)
Y = np.zeros((gt_idx.shape[1], gt_idx.shape[2], 3))
# Evaluate predicted segmentation
for k in range(len(colours)):
Y += np.expand_dims(pred_idx == k, axis=2) * np.expand_dims(np.expand_dims(colours[k], axis=0), axis=0)
# Obtain overlay
Y_overlay = (1 - OVERLAY_R) * I.astype('uint8') + OVERLAY_R * Y
plt.figure
plt.subplot(121)
plt.imshow(Y.astype('uint8'))
plt.title('Predicted\n Segmentation')
plt.subplot(122)
plt.imshow(Y_overlay.astype('uint8'))
plt.title('Overlaid\n Segmentation')
```
| github_jupyter |
# Auxiliary Lines in Planar Geometry
## Preface
Proving a proposition in planar geometry is like an outdoor exploration -- to find a path from the starting point (the problem) to the destination (the conclusion). Yet the path can be a broad highway, or a meandering trail, or -- you may even find yourself in front of a river.
The auxiliary lines are the bridges to get you across. Such lines are indispensible in many problems, or can drastically simplify the proof in others. Just like there is no universal rule on where or how to build the bridge for all kind of terrains, the auxiliary lines have to be designed for each individual problem. Difficult as it can be for beginners, the process of analyzing the problem and finding the solution is rigorous, creative, fascinating and extremely rewarding. This booklet is intended to give you a helping hand.
## Basic problems
Let's look at a simple example. In $\triangle ABC$, $AD$ is a median. $DE$ extends $AD$ and $DE = AD$ (Figure 1). Show that $BE \| AC$ and $BE = AC$.

**Figure 1**
It's obvious that to prove $BE \| AC$, we can start from proving $\angle EBC = \angle ACB$, or $\angle BEA = \angle CAE$. To prove $BE = AC$, we can try to prove $\triangle BED \cong \triangle CAD$. Note that as the corresponding angles are equal in congruent triangles, $\angle EBC = \angle ACB$ or $\angle BEA = \angle CAE$ will already be implied when the congruency is established, which is all we need. The proof can be written as follows:
$\because$ $AD$ is a median of $\triangle ABC$
$\therefore BD = DC$
Also $\because$ $\angle ADC = \angle BDE$ and $DE = AD$
$\therefore \triangle BED \cong \triangle CAD$ (SAS)
$\therefore BE = AC, \angle EBD = \angle ACD$
$\therefore BE \| AC$.
Given the drawing, we came to the solution fairly smoothly in this example. But in many cases, the drawing itself may not be enough, and auxiliary lines are required. Even in this example, if we connect $CE$, we get $\text{▱}ABEC$, from which we can easily prove $BE = AC$ and $BE \| AC$.
Let's see some more examples.
### Medians

**Figure 2**

**Figure 3**

**Figure 4**

**Figure 5**

**Figure 6**
### Midpoints
### Angle bisects
### Trapezoids, squares and triangles

**Figure 20**

**Figure 21**

**Figure 22**

**Figure 23**

**Figure 24**

**Figure 25**

**Figure 26**

**Figure 27**

**Figure 28**

**Figure 29**
### Special points in triangles
### Double angles
### Right triangles
### Proportional segments

**Figure 47**

**Figure 48**

**Figure 49**

**Figure 50**

**Figure 51**

**Figure 52**

**Figure 53**

**Figure 54**

**Figure 55**

**Figure 56**

**Figure 57**
### Concyclic points

**Figure 66**

**Figure 66**

**Figure 66**
   
**Figure 69**
### Tangent and intersecting circles
#### Exercises
1. Two circles intersect at $A$ and $B$. $AC$ and $AD$ are the diameters of two circles respectively. Show that $C, B$, and $D$ are collinear.
2. Two circles intersect at $A$ and $B$. $AD$ and $BF$ are the chords of the two circles, and intersect with the other circle at $C$ and $E$ respectively. Show that $CF\|DE$.
3. Two circles are tangent at $P$. Chord $AB$ of the first circle is tangent to the second circle at $C$. The extension of $AP$ intersects the second circle at $D$. Show that $\angle BPC=\angle CPD$.
4. Given semicircle $O$ with $AB$ as a diameter, $C$ is a point on the semicircle, and $CD\bot AB$ at $D$. $\odot P$ is tangent to $\odot O$ externally at $E$, and to line $CD$ at $F$. $A$ and $E$ are on the same side of $CD$. Show that $A, E, F$ are collinear.
## Problem Set 1
1. $AD$ is a median of $\triangle ABC$, and $AE$ is a median of $\triangle ABD$. $BA=BD$. Show that $AC=2AE$.

**Problem 1**
2. Prove the perimeter of a triangle is greater than the sum of the three medians.
3. Given triangle $ABC$, $P$ is a point on the exterior bisector of angle $A$. Show that $PB+PC > AB+AC$.

**Problem 3**
4. For right triangle $ABC$ with $AB$ as the hypotenuse, the perpendicular bisector $ME$ of $AB$ intersects the angle bisector of $C$ at $E$. Show that $MC=ME$.

**Problem 4**
5. For isosceles triangle $ABC$ with $AB=AC$, $CX$ is the altitude on $AB$. $XP\bot BC$ at $P$. Show that $AB^2=PA^2+PX^2$.

**Problem 5**
6. Show that the diagonal of a rectangle is longer than any line segment between opposite sides.
7. For square $ABCD$, $E$ is the midpoint of $CD$. $BF\bot AE$ at $F$. Show that $CF=CB$.

**Problem 7**
8. In isosceles triangle $ABC$ with $AB=AC$, the circle with $AB$ as a diameter intersects $AC$ and $BC$ at $E$ and $D$ respectively. Make $DF\bot AC$ at $F$. Show that $DF^2=EF\cdot FA$.

**Problem 8**
9. Show that for a triangle, the reflection points of the orthocenter along three sides are on the circumcircle.
10. As shown in the figure, $AB$ is a diameter of $\odot O$, and $AT$ is a tangent line of $\odot O$. $P$ is on the extension of $BM$ such that $PT\bot AT$ and $PT=PM$. Show that $PB=AB$.

**Problem 10**
11. As shown in the figure, $AB$ is a diameter of $\odot O$, and $P$ is a point on the circle. $Q$ is the midpoint of arc $\widearc{BP}$ and tangent $QH$ intersects $AP$ at $H$. Show that $QH\bot AP$.

**Problem 11**
12. As shown in the figure, two circles are tangent internally at $P$. A secant intersects the two circles at $A, B, C, D$. Show that $\angle APB=\angle CPD$.

**Problem 12**
13. As shown in the figure, two circles are tangent externally at $P$. A secant intersects the two circles at $A, B, C, D$. Show that $\angle APD+\angle BPC=180\degree$.

**Problem 13**
14. Two circles intersect at $A,B$. A line through $A$ intersects the two circles at $C$ and $D$. The tangent lines at $C$ and $D$ intersect at $P$. Show that $B, C, P, D$ are concyclic.

**Problem 14**
15. In $\triangle ABC$, $\angle C=90\degree$, and $CD$ is an altitude. The circle with $CD$ as a diameter intersects $AC$ and $BC$ at $E$ and $F$ respectively. Show that $\frac{BF}{AE}=\frac{BC^3}{AC^3}$.

**Problem 15**
16. In $\triangle ABC$, $\angle B=3\angle C$. $AD$ is the angle bisector of $\angle A$. $BD\bot AD$. Show that $BD=\frac{1}{2}(AC-AB)$.

**Problem 16**
17. In right triangle $ABC$, $\angle A=90\degree$, and $AD$ is the altitude on $BC$. $BF$ is the angle bisector of $\angle B$, and $AD$ and $BF$ intersect at $E$. $EG\|BC$. Show that $CG=AF$.

**Problem 17**
18. As shown in the figure, $D$ and $E$ are the midpoints of $AB$ and $AC$ respectively. $AB>AC$. $F$ is a point between $B$ and $D$ such that $DF=AE$. $AH$ is the angle bisector of $\angle BAC$. $FH\bot AH$, and $FH$ intersects $BC$ at $M$. Show that $BM=MC$.

**Problem 18**
19. In trapezoid $ABCD$, $AD\|BC$ and $AD+BC=AB$. $F$ is the midpoint of $CD$. Show that the angle bisectors of $\angle A$ and $\angle B$ intersect at $F$.

**Problem 19**
20. In $\triangle ABC$, $AC=BC$, and $\angle B=2\angle C$. Show that $AC^2=AB^2+AC\cdot AB$.
21.

**Problem 21**
22.

**Problem 22**
23.

**Problem 23**
24.

**Problem 24**
25.

**Problem 25**
26.

**Problem 26**
27.
28.

**Problem 28**
29.

**Problem 29**
30.
31.

**Problem 31**
32.

**Problem 32**
33.

**Problem 33**
34.

**Problem 34**
35.

**Problem 35**
36.

**Problem 36**
37.

**Problem 37**
38.

**Problem 38**
39.

**Problem 39**
## Advanced problems
### Problem Set 2

**Problem 2**

**Problem 3**

**Problem 4**

**Problem 5**

**Problem 6**

**Problem 7**

**Problem 8**

**Problem 9**

**Problem 10**

**Problem 11**

**Problem 12**

**Problem 13**

**Problem 14**

**Problem 15**

**Problem 19**

**Problem 20**
## Hints and answer keys
| github_jupyter |
***
## Vamos con el laboratorio 3
Crear una calculadora sencilla sin definir funciones - parte 1
***
### lab 3 - Crear una calculadora **sencilla** sin definir funciones - parte 1
Cree una calculadora sencilla que acepte un primer número, una operación y un segundo número.
Este es un resultado de ejemplo, en el que un usuario ha escrito *4*, *** y *5* en los cuadros de entrada:
```output
Simple calculator!
First number? 4
Operation? *
Second number? 5
product of 4 * 5 equals 20
```
El programa debe aceptar un símbolo, como el símbolo de asterisco (*), para realizar una multiplicación y generar un producto. Asegúrese de implementar lógica para estos resultados:
- Suma (+)
- Resta (-)
- Multiplicación (*)
- Cociente (/)
- Exponente (//)
- Módulo (%)
Si el usuario no escribe un valor numérico, muestre este mensaje:
```output
Please input a number.
```
Si el usuario escribe una operación que no se reconoce, muestre este mensaje:
```output
Operation not recognized.
```
Tanto si tiene dificultades y necesita echar un vistazo a la solución como si finaliza el ejercicio correctamente, continúe para ver una solución a este desafío.
#### PSEUDOCODIGO
Declaramos dos valores de entrada
```
a (input)
b (input)
```
Declarar el tipo de operación a realizar
```
operacion (input)
```
Creamos una variable con una lista de operadores
```
operadores = ["+", "-", "*", "/", "//", "%" ]
```
Declaramos la condición de evaluación
SI el operador no se encuentra en la lista de Operadores
``
print (Operación no reconocida)
```
ELSE
SI el operacion es "+"
ELSE RESULTADO a+b
SI EL operacion es "-"
ELSE RESULTADO a-b
... así para el resto de operadores
print(resultado)
```
```
print("¡Hola! Bienvenido a la calculadora.")
print("A continuación escribe los datos que requiere el programa para hacer tus cálculos:")
numero_1 = int(input("Escribe el primer número: "))
numero_2 = int(input("Escribe el segundo número: "))
operador = input("Escribe el operador número: ")
if operador == "+":
resultado_suma = numero_1 + numero_2
print("El resultado de tu suma es: ",resultado_suma)
elif operador == "-":
resultado_resta = numero_1 - numero_2
print("El resultado de tu resta es: ",resultado_resta)
elif operador == "*":
resultado_multi = numero_1 * numero_2
print("El resultado de tu multiplicación es: ",resultado_multi)
elif operador == "/":
resultado_divi = numero_1 / numero_2
print("El resultado de tu división es: ",resultado_divi)
elif operador == "//":
resultado_div = numero_1 // numero_2
print("El resultado de tu división entera es: ",resultado_divi)
elif operador == "%":
resultado_resto = numero_1 % numero_2
print("El resultado de tu resto es: ",resultado_resto)
else:
print("El operador no es válido")
```
| github_jupyter |
```
#importing nevesary libraries
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
print("Done")
df_cust = pd.read_csv("Customer List.csv")
df_cust.head()
df_cust.tail()
df_cust.info()
df_cust.isnull().sum()
from sklearn.preprocessing import LabelEncoder
lb = LabelEncoder()
df_cust['Gender'] = lb.fit_transform(df_cust['Gender'])
df_cust
X = df_cust.drop(columns='Purchased')
y = df_cust['Purchased']
from sklearn.model_selection import train_test_split
```
# Feature Scaling
```
from sklearn.preprocessing import MinMaxScaler
min_max = MinMaxScaler()
X = min_max.fit_transform(X)
print(X)
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.25, random_state =42)
X_train
#using knn
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier()
knn.fit(X_train, y_train)
knn_pred = knn.predict(X_test)
knn_pred
from sklearn.metrics import accuracy_score
acc_knn = accuracy_score(y_test, knn_pred)
acc_knn
knn_prob = knn.predict_proba(X_test)
knn_prob = pd.DataFrame(knn_prob, columns=['yes', 'no'])
knn_prob
sns.pairplot(knn_prob)
plt.show()
```
# GridSearch for hyperparameter tuning
```
from sklearn.model_selection import GridSearchCV
knn_param = [{'n_neighbors': [3,5,7,9],
'weights': ['uniform', 'distance'],
'algorithm': ['auto', 'brute', 'kd_tree'],
'leaf_size': [5,15,20]}]
gs_knn = GridSearchCV(knn,
param_grid=knn_param,
scoring= 'recall',
cv=5)
gs_knn.fit(X_train, y_train)
#
gs_knn.best_params_
gs_knn.score(X_test, y_test)
from sklearn.svm import SVC
svm_ = SVC(kernel='linear')
svm_.fit(X_train, y_train)
svm_pred = svm_.predict(X_test)
svm_pred
svm_acc = accuracy_score(y_test, svm_pred)
svm_acc
from imblearn.over_sampling import SMOTE
sm = SMOTE(random_state=42)
X_res, y_res = sm.fit_resample(X,y)
X_res = min_max.fit_transform(X_res)
X_train1, X_test1, y_train1, y_test1 = train_test_split(X_res,y_res, test_size = 0.25, random_state =42)
y_res.value_counts()
knn.fit(X_train1, y_train1)
knn_pred2 = knn.predict(X_test1)
knn_pred2
knn_acc2 = accuracy_score(y_test1, knn_pred2)
knn_acc2
svm_.fit(X_train1, y_train1)
svm_pred2 = svm_.predict(X_test1)
svm_pred2
svm_acc2 = accuracy_score(y_test1, svm_pred2)
svm_acc2
from sklearn.metrics import f1_score, confusion_matrix, recall_score, precision_score, precision_recall_fscore_support
#modelling using XGB classifier with thread
import xgboost as xg
xg_ = xg.XGBClassifier(nthreads = -1)
xg_.fit(X_train, y_train)
pred_xgb = xg_.predict(X_test)
pred_xgb
# Check the accuracy of the model on train and test dataset.
score_xgb = accuracy_score(y_test, pred_xgb)
score_xgb
#prec = precision_score(y_test, knn_pred2)
#prec
xg_.fit(X_train1, y_train1)
pred_xgb2 = xg_.predict(X_test1)
pred_xgb2
# Check the accuracy of the model on train and test dataset.
score_xgb2 = accuracy_score(y_test1, pred_xgb2)
score_xgb2
cm_bal= confusion_matrix(y_test1, knn_pred2)
sns.heatmap(cm_bal)
plt.show()
from sklearn.metrics import classification_report
cr_bal = classification_report(y_test1, knn_pred2)
print(cr_bal)
cr_ubal = classification_report(y_test, knn_pred)
print(cr_ubal)
plt.figure(figsize=(5,4))
sns.countplot(svm_pred2)
plt.show()
from sklearn.metrics import roc_auc_score
roc_acc = roc_auc_score(y_test1, knn_pred2)
roc_acc
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y_test1, knn_pred2)
fpr
thresholds
tpr
def plot_roc_curve(fpr, tpr, label=None):
plt.plot(fpr, tpr, linewidth=2, label=label)
plt.plot([0, 1], [0, 1], 'k--') # dashed diagonal
[...] # Add axis labels and grid
plot_roc_curve(fpr, tpr)
plt.show()
```
| github_jupyter |
# Optimize, print and plot
You will learn how to work with numerical data (**numpy**) and solve simple numerical optimization problems (**scipy.optimize**) and report the results both in text (**print**) and in figures (**matplotlib**).
**Links:**:
- **print**: [examples](https://www.python-course.eu/python3_formatted_output.php) (very detailed)
- **numpy**: [detailed tutorial](https://www.python-course.eu/numpy.php)
- **matplotlib**: [examples](https://matplotlib.org/tutorials/introductory/sample_plots.html#sphx-glr-tutorials-introductory-sample-plots-py), [documentation](https://matplotlib.org/users/index.html), [styles](https://matplotlib.org/3.1.0/gallery/style_sheets/style_sheets_reference.html)
- **scipy-optimize**: [documentation](https://docs.scipy.org/doc/scipy/reference/optimize.html)
# The consumer problem
Consider the following 2-good consumer problem with
* utility function $u(x_1,x_2):\mathbb{R}^2_{+}\rightarrow\mathbb{R}$,
* exogenous income $I$, and
* price-vector $(p_1,p_2)$,
given by
$$
\begin{aligned}
V(p_{1},p_{2},I) & = \max_{x_{1},x_{2}}u(x_{1},x_{2})\\
\text{s.t.}\\
p_{1}x_{1}+p_{2}x_{2} & \leq I,\,\,\,p_{1},p_{2},I>0\\
x_{1},x_{2} & \geq 0
\end{aligned}
$$
**Specific example:** Let the utility function be Cobb-Douglas,
$$
u(x_1,x_2) = x_1^{\alpha}x_2^{1-\alpha}
$$
We then know the solution is given by
$$
\begin{aligned}
x_1^{\ast} &= \alpha \frac{I}{p_1} \\
x_2^{\ast} &= (1-\alpha) \frac{I}{p_2}
\end{aligned}
$$
which implies that $\alpha$ is the budget share of the first good and $1-\alpha$ is the budget share of the second good.
# Numerical python (numpy)
```
import numpy as np # import the numpy module
```
A **numpy array** is like a list, but with two important differences:
1. Elements must be of **one homogenous type**
2. A **slice returns a view** rather than extract content
## Basics
Numpy arrays can be **created from lists** and can be **multi-dimensional**:
```
A = np.array([0, 1, 2, 3, 4, 5, 6, 7, 8, 9]) # one dimension
B = np.array([[3.4, 8.7, 9.9],
[1.1, -7.8, -0.7],
[4.1, 12.3, 4.8]]) # two dimensions
print(type(A),type(B)) # type
print(A.dtype,B.dtype) # data type
print(A.ndim,B.ndim) # dimensions
print(A.shape,B.shape) # shape (1d: (columns,), 2d: (row,columns))
print(A.size,B.size) # size
```
**Slicing** a numpy array returns a **view**:
```
A = np.array([0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
B = A.copy() # a copy of A
C = A[2:6] # a view into A
C[0] = 0
C[1] = 0
print(A) # changed
print(B) # not changed
```
Numpy array can also be created using numpy functions:
```
print(np.ones((2,3)))
print(np.zeros((4,2)))
print(np.linspace(0,1,6)) # linear spacing
```
**Tip 1:** Try pressing <kbd>Shift</kbd>+<kbd>Tab</kbd> inside a function.<br>
**Tip 2:** Try to write `?np.linspace` in a cell
```
?np.linspace
```
## Math
Standard **mathematical operations** can be applied:
```
A = np.array([[1,0],[0,1]])
B = np.array([[2,2],[2,2]])
print(A+B)
print(A-B)
print(A*B) # element-by-element product
print(A/B) # element-by-element division
print(A@B) # matrix product
```
If arrays does not fit together **broadcasting** is applied. Here is an example with multiplication:
```
A = np.array([ [10, 20, 30], [40, 50, 60] ]) # shape = (2,3)
B = np.array([1, 2, 3]) # shape = (3,) = (1,3)
C = np.array([[1],[2]]) # shape = (2,1)
print(A)
print(A*B) # every row is multiplied by B
print(A*C) # every column is multiplied by C
```
**General rule:** Numpy arrays can be added/substracted/multiplied/divided if they in all dimensions have the same size or one of them has a size of one. If the numpy arrays differ in number of dimensions, this only has to be true for the (inner) dimensions they share.
**More on broadcasting:** [Documentation](https://docs.scipy.org/doc/numpy-1.13.0/user/basics.broadcasting.html).
A lot of **mathematical procedures** can easily be performed on numpy arrays.
```
A = np.array([3.1, 2.3, 9.1, -2.5, 12.1])
print(np.min(A)) # find minimum
print(np.argmin(A)) # find index for minimum
print(np.mean(A)) # calculate mean
print(np.sort(A)) # sort (ascending)
```
**Note:** Sometimes a method can be used instead of a function, e.g. ``A.mean()``. Personally, I typically stick to functions because that always works.
## Indexing
**Multi-dimensional** indexing is done as:
```
X = np.array([ [11, 12, 13], [21, 22, 23] ])
print(X)
print(X[0,0]) # first row, first column
print(X[0,1]) # first row, second column
print(X[1,2]) # second row, third column
X[0] # first row
```
Indexes can be **logical**. Logical 'and' is `&` and logical 'or' is `|`.
```
A = np.array([1,2,3,4,1,2,3,4])
B = np.array([3,3,3,3,2,2,2,2])
I = (A < 3) & (B == 3) # note & instead of 'and'
print(type(I),I.dtype)
print(I)
print(A[I])
I = (A < 3) | (B == 3) # note | instead of 'or'
print(A[I])
```
## List of good things to know
**Attributes and methods** to know:
- size / ndim / shape
- ravel / reshape / sort
- copy
**Functions** to know:
- array / empty / zeros / ones / linspace
- mean / median / std / var / sum / percentile
- min/max, argmin/argmax / fmin / fmax / sort / clip
- meshgrid / hstack / vstack / concatenate / tile / insert
- allclose / isnan / isinf / isfinite / any / all
**Concepts** to know:
- view vs. copy
- broadcasting
- logical indexing
**Question:** Consider the following code:
```
A = np.array([1,2,3,4,5])
B = A[3:]
B[:] = 0
```
What is `np.sum(A)` equal to?
- **A:** 15
- **B:** 10
- **C:** 6
- **D:** 0
- **E:** Don't know
## Extra: Memory
Memory is structured in **rows**:
```
A = np.array([[3.1,4.2],[5.7,9.3]])
B = A.ravel() # one-dimensional view of A
print(A.shape,A[0,:])
print(B.shape,B)
```
# Utility function
Define the utility function:
```
def u_func(x1,x2,alpha=0.50):
return x1**alpha*x2**(1-alpha)
# x1,x2 are positional arguments
# alpha is a keyword argument with default value 0.50
```
## Print to screen
Print a **single evaluation** of the utility function.
```
x1 = 1
x2 = 3
u = u_func(x1,x2)
# f'text' is called a "formatted string"
# {x1:.3f} prints variable x1 as floating point number with 3 decimals
print(f'x1 = {x1:.3f}, x2 = {x2:.3f} -> u = {u:.3f}')
```
Print **multiple evaluations** of the utility function.
```
x1_list = [2,4,6,8,10,12]
x2 = 3
for x1 in x1_list: # loop through each element in x1_list
u = u_func(x1,x2,alpha=0.25)
print(f'x1 = {x1:.3f}, x2 = {x2:.3f} -> u = {u:.3f}')
```
And a little nicer...
```
for i,x1 in enumerate(x1_list): # i is a counter
u = u_func(x1,x2,alpha=0.25)
print(f'{i:2d}: x1 = {x1:<6.3f} x2 = {x2:<6.3f} -> u = {u:<6.3f}')
# {i:2d}: integer a width of 2 (right-aligned)
# {x1:<6.3f}: float width of 6 and 3 decimals (<, left-aligned)
```
**Task**: Write a loop printing the results shown in the answer below.
```
# write your code here
```
**Answer:**
```
for i,x1 in enumerate(x1_list): # i is a counter
u = u_func(x1,x2,alpha=0.25)
print(f'{i:2d}: u({x1:.2f},{x2:.2f}) = {u:.4f}')
```
**More formatting options?** See these [examples](https://www.python-course.eu/python3_formatted_output.php).
## Print to file
Open a text-file and write lines in it:
```
with open('somefile.txt', 'w') as the_file: # 'w' is for 'write'
for i, x1 in enumerate(x1_list):
u = u_func(x1,x2,alpha=0.25)
text = f'{i+10:2d}: x1 = {x1:<6.3f} x2 = {x2:<6.3f} -> u = {u:<6.3f}'
the_file.write(text + '\n') # \n gives a lineshift
# note: the with clause ensures that the file is properly closed afterwards
```
Open a text-file and read the lines in it and then print them:
```
with open('somefile.txt', 'r') as the_file: # 'r' is for 'read'
lines = the_file.readlines()
for line in lines:
print(line,end='') # end='' removes the extra lineshift print creates
```
> **Note:** You could also write tables in LaTeX format and the import them in your LaTeX document.
## Calculate the utility function on a grid
**Calculate the utility function** on a 2-dimensional grid with $N$ elements in each dimension:
```
# a. settings
N = 100 # number of elements
x_max = 10 # maximum value
# b. allocate numpy arrays
shape_tuple = (N,N)
x1_values = np.empty(shape_tuple) # allocate 2d numpy array with shape=(N,N)
x2_values = np.empty(shape_tuple)
u_values = np.empty(shape_tuple)
# c. fill numpy arrays
for i in range(N): # 0,1,...,N-1
for j in range(N): # 0,1,...,N-1
x1_values[i,j] = (i/(N-1))*x_max # in [0,x_max]
x2_values[i,j] = (j/(N-1))*x_max # in [0,x_max]
u_values[i,j] = u_func(x1_values[i,j],x2_values[i,j],alpha=0.25)
```
**Alternatively:** Use internal numpy functions:
```
x_vec = np.linspace(0,x_max,N)
x1_values_alt,x2_values_alt = np.meshgrid(x_vec,x_vec,indexing='ij')
u_values_alt = u_func(x1_values_alt,x2_values_alt,alpha=0.25)
```
Test whether the results are the same:
```
# a. maximum absolute difference
max_abs_diff = np.max(np.abs(u_values-u_values_alt))
print(max_abs_diff) # very close to zero
# b. test if all values are "close"
print(np.allclose(u_values,u_values_alt))
```
**Note:** The results are not exactly the same due to floating point arithmetics.
## Plot the utility function
Import modules and state that the figures should be inlined:
```
%matplotlib inline
import matplotlib.pyplot as plt # baseline modul
from mpl_toolkits.mplot3d import Axes3D # for 3d figures
plt.style.use('seaborn-whitegrid') # whitegrid nice with 3d
```
Construct the actual plot:
```
fig = plt.figure() # create the figure
ax = fig.add_subplot(1,1,1,projection='3d') # create a 3d axis in the figure
ax.plot_surface(x1_values,x2_values,u_values); # create surface plot in the axis
# note: fig.add_subplot(a,b,c) creates the c'th subplot in a grid of a times b plots
```
Make the figure **zoomable** and **panable** using a widget:
```
%matplotlib widget
fig = plt.figure() # create the figure
ax = fig.add_subplot(1,1,1,projection='3d') # create a 3d axis in the figure
ax.plot_surface(x1_values,x2_values,u_values); # create surface plot in the axis
```
Turn back to normal inlining:
```
%matplotlib inline
```
**Extensions**: Use a colormap, make it pretier, and save to disc.
```
from matplotlib import cm # for colormaps
# a. actual plot
fig = plt.figure()
ax = fig.add_subplot(1,1,1,projection='3d')
ax.plot_surface(x1_values,x2_values,u_values,cmap=cm.jet)
# b. add labels
ax.set_xlabel('$x_1$')
ax.set_ylabel('$x_2$')
ax.set_zlabel('$u$')
# c. invert xaxis
ax.invert_xaxis()
# d. save
fig.tight_layout()
fig.savefig('someplot.pdf') # or e.g. .png
```
**More formatting options?** See these [examples](https://matplotlib.org/tutorials/introductory/sample_plots.html#sphx-glr-tutorials-introductory-sample-plots-py).
**Task**: Construct the following plot:

**Answer:**
```
# write your code here
# a. actual plot
fig = plt.figure()
ax = fig.add_subplot(1,1,1,projection='3d')
ax.plot_wireframe(x1_values,x2_values,u_values,edgecolor='black')
# b. add labels
ax.set_xlabel('$x_1$')
ax.set_ylabel('$x_2$')
ax.set_zlabel('$u$')
# c. invert xaxis
ax.invert_xaxis()
# e. save
fig.tight_layout()
fig.savefig('someplot_wireframe.png')
fig.savefig('someplot_wireframe.pdf')
```
## Summary
We have talked about:
1. Print (to screen and file)
2. Figures (matplotlib)
**Other plotting libraries:** [seaborn](https://seaborn.pydata.org/) and [bokeh](https://bokeh.pydata.org/en/latest/).
# Algorithm 1: Simple loops
Remember the problem we wanted to solve:
$$
\begin{aligned}
V(p_{1},p_{2},I) & = \max_{x_{1},x_{2}}u(x_{1},x_{2})\\
& \text{s.t.}\\
p_{1}x_{1}+p_{2}x_{2} & \leq I,\,\,\,p_{1},p_{2},I>0\\
x_{1},x_{2} & \geq 0
\end{aligned}
$$
**Idea:** Loop through a grid of $N_1 \times N_2$ possible solutions. This is the same as solving:
$$
\begin{aligned}
V(p_{1},p_{2},I) & = \max_{x_{1}\in X_1,x_{2} \in X_2} x_1^{\alpha}x_2^{1-\alpha}\\
& \text{s.t.}\\
X_1 & = \left\{0,\frac{1}{N_1-1}\frac{I}{p_1},\frac{2}{N_1-1}\frac{I}{p_1},\dots,\frac{I}{p_1}\right\} \\
X_2 & = \left\{0,\frac{1}{N_2-1}\frac{I}{p_2},\frac{2}{N_2-1}\frac{ I}{p_2},\dots,\frac{ I}{p_2}\right\} \\
p_{1}x_{1}+p_{2}x_{2} & \leq I\\
\end{aligned}
$$
Function doing just this:
```
def find_best_choice(alpha,I,p1,p2,N1,N2,do_print=True):
# a. allocate numpy arrays
shape_tuple = (N1,N2)
x1_values = np.empty(shape_tuple)
x2_values = np.empty(shape_tuple)
u_values = np.empty(shape_tuple)
# b. start from guess of x1=x2=0
x1_best = 0
x2_best = 0
u_best = u_func(0,0,alpha=alpha)
# c. loop through all possibilities
for i in range(N1):
for j in range(N2):
# i. x1 and x2 (chained assignment)
x1_values[i,j] = x1 = (i/(N1-1))*I/p1
x2_values[i,j] = x2 = (j/(N2-1))*I/p2
# ii. utility
if p1*x1+p2*x2 <= I: # u(x1,x2) if expenditures <= income
u_values[i,j] = u_func(x1,x2,alpha=alpha)
else: # u(0,0) if expenditures > income
u_values[i,j] = u_func(0,0,alpha=alpha)
# iii. check if best sofar
if u_values[i,j] > u_best:
x1_best = x1_values[i,j]
x2_best = x2_values[i,j]
u_best = u_values[i,j]
# d. print
if do_print:
print_solution(x1_best,x2_best,u_best,I,p1,p2)
return x1_best,x2_best,u_best,x1_values,x2_values,u_values
# function for printing the solution
def print_solution(x1,x2,u,I,p1,p2):
print(f'x1 = {x1:.8f}')
print(f'x2 = {x2:.8f}')
print(f'u = {u:.8f}')
print(f'I-p1*x1-p2*x2 = {I-p1*x1-p2*x2:.8f}')
```
Call the function:
```
sol = find_best_choice(alpha=0.25,I=10,p1=1,p2=2,N1=500,N2=400)
```
Plot the solution:
```
%matplotlib widget
# a. unpack solution
x1_best,x2_best,u_best,x1_values,x2_values,u_values = sol
# b. setup figure
fig = plt.figure(dpi=100,num='')
ax = fig.add_subplot(1,1,1,projection='3d')
# c. plot 3d surface of utility values for different choices
ax.plot_surface(x1_values,x2_values,u_values,cmap=cm.jet)
ax.invert_xaxis()
# d. plot optimal choice
ax.scatter(x1_best,x2_best,u_best,s=50,color='black');
%matplotlib inline
```
**Task**: Can you find a better solution with higher utility and lower left-over income, $I-p_1 x_1-p_2 x_2$?
```
# write your code here
# sol = find_best_choice()
```
**Answer:**
```
sol = find_best_choice(alpha=0.25,I=10,p1=1,p2=2,N1=1000,N2=1000)
```
# Algorithm 2: Use monotonicity
**Idea:** Loop through a grid of $N$ possible solutions for $x_1$ and assume the remainder is spent on $x_2$. This is the same as solving:
$$
\begin{aligned}
V(p_{1},p_{2},I) & = \max_{x_{1}\in X_1} x_1^{\alpha}x_2^{1-\alpha}\\
\text{s.t.}\\
X_1 & = \left\{0,\frac{1}{N-1}\frac{}{p_1},\frac{2}{N-1}\frac{I}{p_1},\dots,\frac{I}{p_1}\right\} \\
x_{2} & = \frac{I-p_{1}x_{1}}{p_2}\\
\end{aligned}
$$
Function doing just this:
```
def find_best_choice_monotone(alpha,I,p1,p2,N,do_print=True):
# a. allocate numpy arrays
shape_tuple = (N)
x1_values = np.empty(shape_tuple)
x2_values = np.empty(shape_tuple)
u_values = np.empty(shape_tuple)
# b. start from guess of x1=x2=0
x1_best = 0
x2_best = 0
u_best = u_func(0,0,alpha)
# c. loop through all possibilities
for i in range(N):
# i. x1
x1_values[i] = x1 = i/(N-1)*I/p1
# ii. implied x2
x2_values[i] = x2 = (I-p1*x1)/p2
# iii. utility
u_values[i] = u_func(x1,x2,alpha)
if u_values[i] >= u_best:
x1_best = x1_values[i]
x2_best = x2_values[i]
u_best = u_values[i]
# d. print
if do_print:
print_solution(x1_best,x2_best,u_best,I,p1,p2)
return x1_best,x2_best,u_best,x1_values,x2_values,u_values
sol_monotone = find_best_choice_monotone(alpha=0.25,I=10,p1=1,p2=2,N=1000)
```
Plot the solution:
```
plt.style.use("seaborn")
# a. create the figure
fig = plt.figure(figsize=(10,4))# figsize is in inches...
# b. unpack solution
x1_best,x2_best,u_best,x1_values,x2_values,u_values = sol_monotone
# c. left plot
ax_left = fig.add_subplot(1,2,1)
ax_left.plot(x1_values,u_values)
ax_left.scatter(x1_best,u_best)
ax_left.set_title('value of choice, $u(x_1,x_2)$')
ax_left.set_xlabel('$x_1$')
ax_left.set_ylabel('$u(x_1,(I-p_1 x_1)/p_2)$')
ax_left.grid(True)
# c. right plot
ax_right = fig.add_subplot(1,2,2)
ax_right.plot(x1_values,x2_values)
ax_right.scatter(x1_best,x2_best)
ax_right.set_title('implied $x_2$')
ax_right.set_xlabel('$x_1$')
ax_right.set_ylabel('$x_2$')
ax_right.grid(True)
```
# Algorithm 3: Call a solver
```
from scipy import optimize
```
Choose paramters:
```
alpha = 0.25 # preference parameter
I = 10 # income
p1 = 1 # price 1
p2 = 2 # price 2
```
**Case 1**: Scalar solver using monotonicity.
```
# a. objective funciton (to minimize)
def value_of_choice(x1,alpha,I,p1,p2):
x2 = (I-p1*x1)/p2
return -u_func(x1,x2,alpha)
# b. call solver
sol_case1 = optimize.minimize_scalar(
value_of_choice,method='bounded',
bounds=(0,I/p1),args=(alpha,I,p1,p2))
# c. unpack solution
x1 = sol_case1.x
x2 = (I-p1*x1)/p2
u = u_func(x1,x2,alpha)
print_solution(x1,x2,u,I,p1,p2)
```
**Case 2**: Multi-dimensional constrained solver.
```
# a. objective function (to minimize)
def value_of_choice(x,alpha,I,p1,p2):
# note: x is a vector
x1 = x[0]
x2 = x[1]
return -u_func(x1,x2,alpha)
# b. constraints (violated if negative) and bounds
constraints = ({'type': 'ineq', 'fun': lambda x: I-p1*x[0]-p2*x[1]})
bounds = ((0,I/p1),(0,I/p2))
# c. call solver
initial_guess = [I/p1/2,I/p2/2]
sol_case2 = optimize.minimize(
value_of_choice,initial_guess,args=(alpha,I,p1,p2),
method='SLSQP',bounds=bounds,constraints=constraints)
# d. unpack solution
x1 = sol_case2.x[0]
x2 = sol_case2.x[1]
u = u_func(x1,x2,alpha)
print_solution(x1,x2,u,I,p1,p2)
```
**Case 3**: Multi-dimensional unconstrained solver with constrains implemented via penalties.
```
# a. objective function (to minimize)
def value_of_choice(x,alpha,I,p1,p2):
# i. unpack
x1 = x[0]
x2 = x[1]
# ii. penalty
penalty = 0
E = p1*x1+p2*x2 # total expenses
if E > I: # expenses > income -> not allowed
fac = I/E
penalty += 1000*(E-I) # calculate penalty
x1 *= fac # force E = I
x2 *= fac # force E = I
return -u_func(x1,x2,alpha)
# b. call solver
initial_guess = [I/p1/2,I/p2/2]
sol_case3 = optimize.minimize(
value_of_choice,initial_guess,method='Nelder-Mead',
args=(alpha,I,p1,p2))
# c. unpack solution
x1 = sol_case3.x[0]
x2 = sol_case3.x[1]
u = u_func(x1,x2,alpha)
print_solution(x1,x2,u,I,p1,p2)
```
**Task:** Find the error in the code in the previous cell.
```
# write your code here
```
**Answer:**
```
# a. objective function (to minimize)
def value_of_choice(x,alpha,I,p1,p2):
# i. unpack
x1 = x[0]
x2 = x[1]
# ii. penalty
penalty = 0
E = p1*x1+p2*x2 # total expenses
if E > I: # expenses > income -> not allowed
fac = I/E
penalty += 1000*(E-I) # calculate penalty
x1 *= fac # force E = I
x2 *= fac # force E = I
return -u_func(x1,x2,alpha) + penalty # the error
# b. call solver
initial_guess = [I/p1/2,I/p2/2]
sol_case3 = optimize.minimize(
value_of_choice,initial_guess,method='Nelder-Mead',
args=(alpha,I,p1,p2))
# c. unpack solution
x1 = sol_case3.x[0]
x2 = sol_case3.x[1]
u = u_func(x1,x2,alpha)
print_solution(x1,x2,u,I,p1,p2)
```
# Indifference curves
Remember that the indifference curve through the point $(y_1,y_2)$ is given by
$$
\big\{(x_1,x_2) \in \mathbb{R}^2_+ \,|\, u(x_1,x_2) = u(y_1,y_2)\big\}
$$
To find the indifference curve, we can fix a grid for $x_2$, and then find the corresponding $x_1$ which solves $u(x_1,x_2) = u(y_1,y_2)$ for each value of $x_2$.
```
def objective(x1,x2,alpha,u):
return u_func(x1,x2,alpha)-u
# = 0 then on indifference curve with utility = u
def find_indifference_curve(y1,y2,alpha,N,x2_max):
# a. utiltty in (y1,y2)
u_y1y2 = u_func(y1,y2,alpha)
# b. allocate numpy arrays
x1_vec = np.empty(N)
x2_vec = np.linspace(1e-8,x2_max,N)
# c. loop through x2
for i,x2 in enumerate(x2_vec):
x1_guess = 0 # initial guess
sol = optimize.root(objective, x1_guess, args=(x2,alpha,u_y1y2))
# optimize.root -> solve objective = 0 starting from x1 = x1_guess
x1_vec[i] = sol.x[0]
return x1_vec,x2_vec
```
Find and plot an inddifference curve:
```
# a. find indifference curve through (4,4) for x2 in [0,10]
x2_max = 10
x1_vec,x2_vec = find_indifference_curve(y1=4,y2=4,alpha=0.25,N=100,x2_max=x2_max)
# b. plot inddifference curve
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(1,1,1)
ax.plot(x1_vec,x2_vec)
ax.set_xlabel('$x_1$')
ax.set_ylabel('$x_2$')
ax.set_xlim([0,x2_max])
ax.set_ylim([0,x2_max])
ax.grid(True)
```
**Task:** Find the indifference curve through $x_1 = 15$ and $x_2 = 3$ with $\alpha = 0.5$.
```
# write your code here
x2_max = 20
x1_vec,x2_vec = find_indifference_curve(y1=15,y2=3,alpha=0.5,N=100,x2_max=x2_max)
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(1,1,1)
ax.plot(x1_vec,x2_vec)
ax.set_xlabel('$x_1$')
ax.set_ylabel('$x_2$')
ax.set_xlim([0,x2_max])
ax.set_ylim([0,x2_max])
ax.grid(True)
```
# A classy solution
> **Note:** This section is advanced due to the use of a module with a class. It is, however, a good example of how to structure code for solving and illustrating a model.
**Load module** I have written (consumer_module.py in the same folder as this notebook).
```
from consumer_module import consumer
```
## Jeppe
Give birth to a consumer called **jeppe**:
```
jeppe = consumer() # create an instance of the consumer class called jeppe
print(jeppe)
```
Solve **jeppe**'s problem.
```
jeppe.solve()
print(jeppe)
```
## Mette
Create a new consumer, called Mette, and solve her problem.
```
mette = consumer(alpha=0.25)
mette.solve()
mette.find_indifference_curves()
print(mette)
```
Make an illustration of Mette's problem and it's solution:
```
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(1,1,1)
mette.plot_indifference_curves(ax)
mette.plot_budgetset(ax)
mette.plot_solution(ax)
mette.plot_details(ax)
```
# Summary
**This lecture:** We have talked about:
1. Numpy (view vs. copy, indexing, broadcasting, functions, methods)
2. Print (to screen and file)
3. Figures (matplotlib)
4. Optimization (using loops or scipy.optimize)
5. Advanced: Consumer class
Most economic models contain optimizing agents solving a constrained optimization problem. The tools applied in this lecture is not specific to the consumer problem in anyway.
**Your work:** Before solving Problem Set 1 read through this notebook and play around with the code. To solve the problem set, you only need to modify the code used here slightly.
**Next lecture:** Random numbers and simulation.
| github_jupyter |
# 3.3 MNIST Handwritten Digits
```
import os
import numpy as np
from matplotlib import pyplot as plt
from keras.datasets import mnist
from keras.layers import *
from keras.models import Sequential, Model
from keras.optimizers import Adam, SGD
from keras.utils import to_categorical
from keras.losses import categorical_crossentropy
from keras import backend as K
from keras.backend.tensorflow_backend import set_session
import tensorflow as tf
os.environ['CUDA_VISIBLE_DEVICES'] = '1'
# GPU memory usage
config = tf.ConfigProto()
config.gpu_options.per_process_gpu_memory_fraction = 0.07
set_session(tf.Session(config=config))
plt.style.use('seaborn-darkgrid')
```
## Define Class
```
class PPAN:
def __init__(self,
img_size=(28, 28),
channels=1,
noise_dim=20,
learning_rates=[2e-4, 2e-4, 1e-3],
utility_weight=2,
valid_weight=1):
'''
Build PPAN model
Args:
img_size: tuple, input image size, default=(28, 28)
channels: int, input image channels, default=1
noise_dim: int, noise input of mechanism network, default=20
learning_rates: list, list of learning rates of adversary, discriminator and mechanism, default=[1e-4, 1e-4, 1e-3]
utility_weight: float, weight of the utility loss, default=2
valid_weight: float, weight of the discriminator loss, default=1
Returns:
PPAN object
'''
# Get variables
self.img_size = img_size
self.channels = channels
self.noise_dim = noise_dim
self.utility_weight = utility_weight
self.valid_weight = valid_weight
# Get networks
self.mechanism = self.build_mechanism()
self.adversary = self.build_adversary()
self.discriminator = self.build_discriminator()
# Setup optimizers
a_optimizer = SGD(learning_rates[0])
d_optimizer = SGD(learning_rates[1])
m_optimizer = Adam(learning_rates[2], 0.5)
# Compile discriminators
self.adversary.compile(
loss=self.negative_mutual_info_privacy_loss,
optimizer=a_optimizer,
weighted_metrics=["accuracy"])
self.discriminator.compile(
loss="binary_crossentropy",
optimizer=d_optimizer,
weighted_metrics=["accuracy"])
# Combined two discriminators with the mechanism(generator)
noised_img = Input(
shape=(self.img_size[0] * self.img_size[1] + self.noise_dim,))
gen_img = self.mechanism(noised_img)
# Freeze discriminators
self.adversary.trainable = False
self.discriminator.trainable = False
# Get predictions from discriminators
pred_label = self.adversary(gen_img)
pred_valid = self.discriminator(gen_img)
self.ppan = Model(
inputs=noised_img, outputs=[pred_label, gen_img, pred_valid])
# Compile mechanism
self.ppan.compile(
loss=[
self.mutual_info_privacy_loss,
"binary_crossentropy",
"binary_crossentropy"
],
loss_weights=[1.0, self.utility_weight, self.valid_weight],
optimizer=m_optimizer)
def mutual_info_privacy_loss(self, y_true, y_pred):
conditional_entropy = -K.mean(
K.sum(K.log(y_pred + 1e-9) * y_true, axis=1)) # K.sum(axis=1) turn the vector into scalar
return conditional_entropy
def negative_mutual_info_privacy_loss(self, y_true, y_pred):
conditional_entropy = -K.mean(
K.sum(K.log(y_pred + 1e-9) * y_true, axis=1)) # K.sum(axis=1) turn the vector into scalar
return -conditional_entropy
def build_mechanism(self):
model = Sequential(name="Mechanism")
model.add(
Dense(
1000,
activation="tanh",
input_dim=self.img_size[0] * self.img_size[1] * self.channels +
self.noise_dim,
name="fc1"))
model.add(Dense(1000, activation="tanh", name="fc2"))
model.add(Dense(self.img_size[0] * \
self.img_size[1], activation="sigmoid", name="output"))
print("\n=== Mechanism Summary ===")
model.summary()
return model
def build_adversary(self):
model = Sequential(name="Adversary")
model.add(
Dense(
1000,
activation="tanh",
input_dim=self.img_size[0] * self.img_size[1] * self.channels,
name="fc1"))
model.add(Dense(1000, activation="tanh", name="fc2"))
model.add(Dense(10, activation="softmax", name="output"))
print("\n=== Adversary Summary ===")
model.summary()
return model
def build_discriminator(self):
model = Sequential(name="Discriminator")
model.add(
Dense(
500,
activation="tanh",
input_dim=self.img_size[0] * self.img_size[1] * self.channels,
name="fc1"))
model.add(Dense(1, activation="sigmoid", name="output"))
print("\n=== Discriminator Summary ===")
model.summary()
return model
def train(self,
x_train,
y_train,
x_test,
y_test,
epochs,
batch_size,
save_path,
print_interval=10,
save_interval=50):
'''
Train model
Args:
x_train: ndarray, training images
y_train: ndarray, training labels
x_test: ndarray, testing images
y_test: ndarray, testing labels
epochs: int, training epochs
batch_size: int, training batch size
save_path: string, saving path for generated images during training process
print_interval: int, print accuracy & loss every print_interval epochs, default=10
save_interval: int, save generated images every save_interval, default=50
Returns:
None
'''
# Flatten images data
x_train = x_train.reshape(
(x_train.shape[0],
self.img_size[0] * self.img_size[1] * self.channels))
x_train = x_test.reshape(
(x_test.shape[0],
self.img_size[0] * self.img_size[1] * self.channels))
# Set labels for discriminator
valid = np.ones(shape=(batch_size, 1))
fake = np.zeros(shape=(batch_size, 1))
# Store training loss and accuracy in an array
# privacy/realistic/utility loss (a/d/m networks)
self.losses = np.zeros(shape=(epochs, 3))
# adversary/discriminator accuracy
self.accuracies = np.zeros(shape=(epochs, 2))
# Training
for e in range(epochs):
# -------------------------
# Train Discriminators
# -------------------------
# Random sample batch_size images from data
idx = np.random.randint(
low=0, high=x_train.shape[0], size=batch_size)
original_imgs, labels = x_train[idx], y_train[idx]
# Sample noise and generate a batch of new images
noise = np.random.uniform(
low=-1, high=1, size=(batch_size, self.noise_dim))
gen_imgs = self.mechanism.predict(np.hstack((original_imgs, noise)))
# Train adversary
self.losses[e, 0], self.accuracies[
e, 0] = self.adversary.train_on_batch(gen_imgs, labels)
# Train discriminator
d_loss_valid = self.discriminator.train_on_batch(
original_imgs, valid)
d_loss_fake = self.discriminator.train_on_batch(gen_imgs, fake)
self.losses[e, 1], self.accuracies[e, 1] = 0.5 * \
np.add(d_loss_valid, d_loss_fake)
# -------------------------
# Train Generator
# -------------------------
# Sample batch_size images and noise
idx = np.random.randint(
low=0, high=x_train.shape[0], size=batch_size)
original_imgs, labels = x_train[idx], y_train[idx]
noise = np.random.uniform(
low=-1, high=1, size=(batch_size, self.noise_dim))
# Train mechanism
m_loss = self.ppan.train_on_batch(
np.hstack((original_imgs, noise)),
[labels, original_imgs, valid])
self.losses[e, 2] = m_loss[0] + m_loss[
1] * self.utility_weight + m_loss[2] * self.valid_weight
# Print loss at the beginning/end and every print_interval
# if (e+1 % print_interval == 0) or (e == 0) or (e+1 == epochs):
print(
"%dth epoch\tA loss: %.3f, acc: %.2f%%\tD loss: %.3f, acc: %.2f%%\tM privacy: %.3f, utility: %.3f, realistic: %.3f, weighted: %.3f"
% (e + 1, self.losses[e, 0], self.accuracies[e, 0] * 100,
self.losses[e, 1], self.accuracies[e, 1] * 100, m_loss[0],
m_loss[1], m_loss[2], self.losses[e, 2]))
# Generate images and save at the beginning/end and every save_interval
# if e+1 % save_interval == 0 or e == 0 or e+1 == self.epochs:
# self.generate(self.noise, "%s%d.png" % (save_path, e))
# def gen_noise(self):
# def generate(self)
```
## Load Data
```
(x_train, y_train), (x_test, y_test) = mnist.load_data()
```
## Data Preprocessing
### Normalize
```
x_train = x_train / 255.0
x_test = x_test / 255.0
```
### One-Hot Encoded
```
Y_train = to_categorical(y_train, num_classes=10)
Y_test = to_categorical(y_test, num_classes=10)
```
## Main
### Build Model
```
ppan = PPAN(utility_weight=2, valid_weight=1)
```
### Train Model
```
%%time
ppan.train(x_train, Y_train, x_test, Y_test, 1000, 32, save_path="generated/")
plt.style.use('seaborn-dark') # have to call this function twice to make it work
plt.figure(figsize=(12, 4))
plt.plot(ppan.losses[:,0], label="Adversary")
plt.plot(ppan.losses[:,2], label="Mechanism")
plt.legend(loc='best')
plt.title("Learning Curve")
plt.xlabel("Epochs")
plt.show()
plt.style.use('seaborn-dark') # have to call this function twice to make it work
plt.figure(figsize=(12, 4))
plt.plot(ppan.losses[:,1], label="Discriminator")
plt.legend(loc='best')
plt.title("Learning Curve")
plt.xlabel("Epochs")
plt.show()
```
## Testing
```
test_sample = [1,3,5,7,2,0,13,15,17,19]
for i in test_sample:
test_img = x_train[i].reshape((1, 784))
test_noise = np.random.uniform(low=-1, high=1, size=(1, 20))
test_result = ppan.mechanism.predict(np.hstack((test_img, test_noise))).reshape((28, 28))
plt.imshow(test_img.reshape((28, 28)), cmap="gray")
plt.show()
plt.imshow(test_result, cmap="gray")
plt.show()
print(y_train[i])
```
| github_jupyter |
```
import string
from tensorflow.keras.utils import plot_model
from os import listdir
from tensorflow.keras.layers import Dropout
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.applications.vgg16 import preprocess_input
from tensorflow.keras.layers import Embedding
from tensorflow.keras.preprocessing.image import img_to_array
from tensorflow.keras.applications.vgg16 import VGG16
from tensorflow.keras.layers import LSTM
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.layers import Input
from tensorflow.keras.models import Model, load_model
from tensorflow.keras.layers import Add
from pickle import load,dump
from tensorflow.keras.callbacks import ModelCheckpoint
from tensorflow.keras.preprocessing.image import load_img
from numpy import array, argmax
from nltk.translate.bleu_score import corpus_bleu
from tensorflow.keras.layers import Dense
from tensorflow.keras.preprocessing.text import Tokenizer
# from google.colab import drive
# drive.mount('/content/drive')
def feature_extraction(dir):
model=VGG16()
model=Model(inputs=model.Input,outputs=model.layers[-2].output)
print(model.summary())
feat=dict()
for nm in listdir(dir):
f_name=dir + '/' + nm
pict=load_img(f_name, target_size=(224, 224))
pict=img_to_array(pict)
pict=pict.reshape((1,pict.shape[0],pict.shape[1],pict.shape[2]))
pict=preprocess_input(pict)
featr=model.predict(pict,verbose=0)
pict_id=nm.split('.')[0]
feat[pict_id]=featr
print('>%s'%nm)
return feat
'''
This code section is used to generate features. Generated features are already saved into features.pkl
dir = '/content/drive/MyDrive/Colab_Notebooks/archive/Images'
feat = feature_extraction(dir)
print('Extracted Features: %d'%len(feat))
'''
#dump(feat, open('/content/drive/MyDrive/Colab_Notebooks/archive/features.pkl', 'wb'))
def document_loading(f_name):
file=open(f_name,'r')
txt=file.read()
file.close()
return txt
def decs_load(doc):
map_desc=dict()
for ln in doc.split('\n'):
tkn=ln.split()
if len(ln)<2:
continue
pict_id,pict_desc=tkn[0],tkn[1:]
pict_id=pict_id.split('.')[0]
pict_desc = ' '.join(pict_desc)
if pict_id not in map_desc:
map_desc[pict_id]=list()
map_desc[pict_id].append(pict_desc)
return map_desc
def desc_cleaning(descs):
tb=str.maketrans('', '', string.punctuation)
for k, desc_list in descs.items():
for i in range(len(desc_list)):
desc=desc_list[i]
desc=desc.split()
desc=[wd.lower() for wd in desc]
desc=[w.translate(tb) for w in desc]
desc=[wd for wd in desc if len(wd)>1]
desc=[wd for wd in desc if wd.isalpha()]
desc_list[i] = ' '.join(desc)
def vocab_conv(descs):
desc_all=set()
for k in descs.keys():
[desc_all.update(d.split()) for d in descs[k]]
return desc_all
def desc_saving(descs,f_name):
lns=list()
for k,desc_list in descs.items():
for desc in desc_list:
lns.append(k+' '+desc)
dta='\n'.join(lns)
file=open(f_name,'w')
file.write(dta)
file.close()
f_name='C:/Users/Kunj/Downloads/recommender/Flickr8k.token.txt'
doc = document_loading(f_name)
descs=decs_load(doc)
print('Loaded: %d '%len(descs))
desc_cleaning(descs)
vblry=vocab_conv(descs)
print('Vocabulary Size: %d'%len(vblry))
desc_saving(descs,'descriptions.txt')
def dset_load(f_name):
doc = document_loading(f_name)
dset = list()
for ln in doc.split('\n'):
if len(ln)<1:
continue
idf=ln.split('.')[0]
dset.append(idf)
return set(dset)
def cleaned_desc_load(f_name,dset):
doc=document_loading(f_name)
descs=dict()
for ln in doc.split('\n'):
tkn=ln.split()
pict_id,pict_desc=tkn[0],tkn[1:]
if pict_id in dset:
if pict_id not in descs:
descs[pict_id]=list()
desc='startingsequence ' + ' '.join(pict_desc) + ' endingsequence'
descs[pict_id].append(desc)
return descs
def img_features_load(f_name,dset):
allfeatr=load(open(f_name,'rb'))
ftrs={k: allfeatr[k] for k in dset}
return ftrs
def into_lns(descs):
desc_all=list()
for k in descs.keys():
[desc_all.append(d) for d in descs[k]]
return desc_all
def generate_Tokenizer(descs):
lns=into_lns(descs)
tokenizer=Tokenizer()
tokenizer.fit_on_texts(lns)
return tokenizer
def most_lngth(descs):
lns=into_lns(descs)
max_lngth=max(len(d.split()) for d in lns)
return max_lngth
def build_seq(tokenizer,most_lngth,desc_list,photo,vocb_size):
aa1,aa2,y=list(),list(),list()
for desc in desc_list:
seq=tokenizer.texts_to_sequences([desc])[0]
for i in range(1, len(seq)):
input_sequence,output_sequence=seq[:i],seq[i]
input_sequence=pad_sequences([input_sequence],maxlen=most_lngth)[0]
output_sequence=to_categorical([output_sequence],num_classes=vocb_size)[0]
aa1.append(photo)
aa2.append(input_sequence)
y.append(output_sequence)
return array(aa1),array(aa2),array(y)
def defn_modl(vocb_size,most_lngth):
inp1=Input(shape=(4096,))
fe1=Dropout(0.5)(inp1)
fe2=Dense(256,activation='relu')(fe1)
inp2=Input(shape=(most_lngth,))
ese1=Embedding(vocb_size,256,mask_zero=True)(inp2)
ese2=Dropout(0.5)(ese1)
ese3=LSTM(256)(ese2)
dec1=Add()([fe2,ese3])
dec2=Dense(256,activation='relu')(dec1)
oupts=Dense(vocb_size,activation='softmax')(dec2)
model=Model(inputs=[inp1, inp2],outputs=oupts)
model.compile(loss='categorical_crossentropy',optimizer='adam')
model.summary()
plot_model(model,to_file='model.png',show_shapes=True)
return model
def data_creator(descs,photos,tokenizer,most_lngth,vocb_size):
while 1:
for k, desc_list in descs.items():
photo=photos[k][0]
in_img,input_sequence,out_word=build_seq(tokenizer,most_lngth,desc_list,photo,vocb_size)
yield [in_img,input_sequence],out_word
f_name='C:/Users/Kunj/Downloads/recommender/Flickr_8k.trainImages.txt'
train=dset_load(f_name)
print('Dataset: %d'%len(train))
train_desc=cleaned_desc_load('descriptions.txt',train)
print('Descriptions: train=%d'%len(train_desc))
train_feat=img_features_load('C:/Users/Kunj/Downloads/recommender/features.pkl',train)
print('Photos: train=%d'%len(train_feat))
tokenizer=generate_Tokenizer(train_desc)
vocb_size=len(tokenizer.word_index)+1
print('Size of Vocabulary=%d'%vocb_size)
most_lngth=most_lngth(train_desc)
print('Description Length: %d'%most_lngth)
model=defn_modl(vocb_size,most_lngth)
# epochs=20
# stps=len(train_desc)
# for i in range(epochs):
# generator=data_creator(train_desc,train_feat,tokenizer,most_lngth,vocb_size)
# model.fit_generator(generator,epochs=1,steps_per_epoch=stps,verbose=1)
# model.save("model_'+str(i)+'.h5")
def word_for_id(itgr,tokenizer):
tkzz=tokenizer
for word,index in tkzz.word_index.items():
if index==itgr:
return word
return None
def generate_desc(model,tkzz,photo,maxim_lgth):
in_text='startingsequence'
for i in range(maxim_lgth):
sequence=tkzz.texts_to_sequences([in_text])[0]
sequence=pad_sequences([sequence],maxlen=maxim_lgth)
yhat=model.predict([photo,sequence],verbose=0)
yhat=argmax(yhat)
word=word_for_id(yhat,tkzz)
if word is None:
break
in_text+=' '+word
if word=='endingsequence':
break
return in_text
def model_evaluation(model,descriptions,photos,tokenizer,maxim_lgth):
actual, predicted=list(),list()
for key,desc_list in descriptions.items():
yhat=generate_desc(model,tokenizer,photos[key],maxim_lgth)
references=[d.split() for d in desc_list]
actual.append(references)
predicted.append(yhat.split())
print('BLEU-1:%f'%corpus_bleu(actual,predicted,weights=(1.0,0,0,0)))
print('BLEU-2:%f'%corpus_bleu(actual,predicted,weights=(0.5,0.5,0,0)))
print('BLEU-3:%f'%corpus_bleu(actual,predicted,weights=(0.3,0.3,0.3,0)))
print('BLEU-4:%f'%corpus_bleu(actual,predicted,weights=(0.25,0.25,0.25,0.25)))
name_of_file='C:/Users/Kunj/Downloads/recommender/Flickr_8k.testImages.txt'
test=dset_load(name_of_file)
print('Dataset: %d'%len(test))
test_descriptions=cleaned_desc_load('C:/Users/Kunj/Downloads/recommender/descriptions.txt',test)
print('Descriptions: test=%d'%len(test_descriptions))
test_features=img_features_load('C:/Users/Kunj/Downloads/recommender/features.pkl',test)
print('Photos: test=%d'%len(test_features))
name_of_file='C:/Users/Kunj/Downloads/recommender/model/model_19.h5'
model=load_model(name_of_file)
print("evaluation start")
model_evaluation(model,test_descriptions,test_features,tokenizer,most_lngth)
name_of_file='C:/Users/Kunj/Downloads/recommender/Flickr_8k.trainImages.txt'
train=dset_load(name_of_file)
print('Dataset: %d'%len(train))
train_descriptions=cleaned_desc_load('C:/Users/Kunj/Downloads/recommender/descriptions.txt',train)
print('Descriptions: train=%d'%len(train_descriptions))
tokenizer=generate_Tokenizer(train_descriptions)
dump(tokenizer,open('C:/Users/Kunj/Downloads/recommender/tokenizer.pkl','wb'))
tokenizer=load(open('C:/Users/Kunj/Downloads/recommender/tokenizer.pkl','rb'))
maxim_lgth=34
model=load_model('C:/Users/Kunj/Downloads/recommender/model/model_19.h5')
def extract_features(name_of_file):
model=VGG16()
model=Model(inputs=model.inputs,outputs=model.layers[-2].output)
image=load_img(name_of_file,target_size=(224,224))
image=img_to_array(image)
image=image.reshape((1,image.shape[0],image.shape[1],image.shape[2]))
image=preprocess_input(image)
feature=model.predict(image,verbose=0)
return feature
photo=extract_features('C:/Users/Kunj/Downloads/recommender/example.jpg')
padding_words=['startingsequence','endingsequence']
description=generate_desc(model,tokenizer,photo,maxim_lgth)
print(description)
```
| github_jupyter |
# Jupyter lab on Sunbird using port forwarding
An excellent manual for Classic Jupyter: https://github.com/McWilliamsCenter/slurm_jupyter
I've tailored this manual for Swansea Sunbird for both Jupyter and Jupyter lab.
What is Swansea Sunbird ? https://portal.supercomputing.wales/index.php/about-sunbird/
# Installing Jupyter lab
* Create a conda environment or load an existing one as `source activate ml`
* By default a package named `six` is missing.
```sh
(pytorch) [s.1915438@sl2 ~]$ jupyter notebook --generate-config
Traceback (most recent call last):
File "/apps/languages/anaconda3/bin/jupyter-notebook", line 7, in <module>
from notebook.notebookapp import main
File "/apps/languages/anaconda3/lib/python3.6/site-packages/notebook/__init__.py", line 25, in <module>
from .nbextensions import install_nbextension
File "/apps/languages/anaconda3/lib/python3.6/site-packages/notebook/nbextensions.py", line 31, in <module>
from .config_manager import BaseJSONConfigManager
File "/apps/languages/anaconda3/lib/python3.6/site-packages/notebook/config_manager.py", line 14, in <module>
from six import PY3
ModuleNotFoundError: No module named 'six'
```
So, it is better to install new Jupyter lab or Jupyter as follows:
* Jupyter lab: `pip install jupyterlab`
* Jupyter: `pip install notebook`
In the root i.e. `/home/s.1915438` run this command to generate the config file and store password for secure port forwarding. Apperently this command does not work.
```sh
(ml) [s.1915438@sl2 ~]$ pwd
/home/s.1915438
(ml) [s.1915438@sl2 ~]$ jupyter-lab --generate-config
(ml) [s.1915438@sl2 ~]$ jupyter-lab password
Enter password:
Verify password:
Traceback (most recent call last):
File "/home/s.1915438/.conda/envs/ml/bin/jupyter-lab", line 8, in <module>
sys.exit(main())
File "/home/s.1915438/.conda/envs/ml/lib/python3.9/site-packages/jupyter_server/extension/application.py", line 602, in launch_instance
serverapp.start()
File "/home/s.1915438/.conda/envs/ml/lib/python3.9/site-packages/jupyter_server/serverapp.py", line 2760, in start
self.start_app()
File "/home/s.1915438/.conda/envs/ml/lib/python3.9/site-packages/jupyter_server/serverapp.py", line 2658, in start_app
super(ServerApp, self).start()
File "/home/s.1915438/.conda/envs/ml/lib/python3.9/site-packages/jupyter_core/application.py", line 253, in start
self.subapp.start()
File "/home/s.1915438/.conda/envs/ml/lib/python3.9/site-packages/jupyter_server/serverapp.py", line 492, in start
set_password(config_file=self.config_file)
File "/home/s.1915438/.conda/envs/ml/lib/python3.9/site-packages/jupyter_server/auth/security.py", line 172, in set_password
hashed_password = passwd(password)
File "/home/s.1915438/.conda/envs/ml/lib/python3.9/site-packages/jupyter_server/auth/security.py", line 63, in passwd
import argon2
File "/home/s.1915438/.conda/envs/ml/lib/python3.9/site-packages/argon2/__init__.py", line 7, in <module>
from . import exceptions, low_level, profiles
File "/home/s.1915438/.conda/envs/ml/lib/python3.9/site-packages/argon2/low_level.py", line 15, in <module>
from _argon2_cffi_bindings import ffi, lib
File "/home/s.1915438/.conda/envs/ml/lib/python3.9/site-packages/_argon2_cffi_bindings/__init__.py", line 3, in <module>
from ._ffi import ffi, lib
ImportError: libffi.so.7: cannot open shared object file: No such file or directory
```
# Tinkering configuration file
So, we will go without the password. But there is a Jupyter server token for security also the public IP is also hidden in ssh-tunnel.
**Update**: When I was using ssh through VS Code's extention I was able to run this command `jupyter-lab password` and was able to setup the password. If you have setup the password, Jupyter server will not generate a token.
Now head on to `/lustrehome/home/s.1915438/.jupyter` and open the file, un-comment and change the following lines.
* Jupyter lab:
* `c.LabApp.open_browser = False`
* `c.ServerApp.port = 8888`
* Jupyter:
* `c.NotebookApp.open_browser = False`
* `c.NotebookApp.port = 8888` # (You can set this to any four-digit integer)
Obviously, we don't want to the Notebook to open in the browser as there is no browser on Sunbird. Lol.
Also, we want to set a specific port on server (Sunbird) for port forwarding.
# Port forwarding
This command is used to run commands on the server (Sunbird) while sshing.
`ssh -L <port>:localhost:<port> -t <user>@<server> "jupyter notebook"`
where `<port>` is the port you set earlier, `<user>` is your cluster user id, and `<server>` is the address of the login server. The `-L` flag tells ssh to tunnel the `localhost:<port>` of the remote server to that of your local machine. The `-t` flag opens up the connection as an interactive session, allowing you to pass `SIGINT` (Ctrl-C) to end the jupyter notebook before killing the ssh connection. To open your jupyter notebook within a specific conda environment (e.g. `<env>`), replace the command in quotations with `source activate <env>; jupyter notebook`.
# Running Jupyter lab on login node
Here is out turn to become creative with the command.
```sh
(base) hell@Dell-Precision-T1600:~/Desktop/repos/Sunbird/Jupyter_lab_port_forwarding$ ssh -L 8888:localhost:8888 -t s.1915438@sunbird.swansea.ac.uk "module load anaconda/3;source activate ml;jupyter-lab"
```
Here, `8888` on left, in green colour is my server (Sunbird) port and `8888` on the right is my local port and `-L` tunnel both the ports.
`-t` opens an interactive session in which we can any command that falls inside those inverted commas `""`.
Now, we can run our Jupyter lab as follows:
* Load Anaconda: `module load anaconda/3` the module name might change time to time. Check the module name using `module avail`.
* Activate the conda environment where you installed the Jupyter Lab. In my case it was `ml` conda environment. So, I would type `source activate ml`.
* `source activate base` and then `conda activate ml` is equivalent to above command.
* Now run `jupyter-lab` as you were using `anaconda prompt` in Windows.
All these things can be passed to ssh as `"module load anaconda/3;source activate ml;jupyter-lab"`.
As you run ssh command you will see Jupyter lab is starting.
```sh
(base) hell@Dell-Precision-T1600:~/Desktop/repos/Sunbird/Jupyter_lab_port_forwarding$ ssh -L 8888:localhost:8888 -t s.1915438@sunbird.swansea.ac.uk "module load anaconda/3;source activate ml;jupyter-lab"
[I 2022-03-19 22:39:59.593 ServerApp] jupyterlab | extension was successfully linked.
[I 2022-03-19 22:39:59.602 ServerApp] nbclassic | extension was successfully linked.
[I 2022-03-19 22:40:00.377 ServerApp] notebook_shim | extension was successfully linked.
[I 2022-03-19 22:40:00.432 ServerApp] notebook_shim | extension was successfully loaded.
[I 2022-03-19 22:40:00.434 LabApp] JupyterLab extension loaded from /home/s.1915438/.conda/envs/ml/lib/python3.9/site-packages/jupyterlab
[I 2022-03-19 22:40:00.434 LabApp] JupyterLab application directory is /lustrehome/home/s.1915438/.conda/envs/ml/share/jupyter/lab
[I 2022-03-19 22:40:00.438 ServerApp] jupyterlab | extension was successfully loaded.
[I 2022-03-19 22:40:00.452 ServerApp] nbclassic | extension was successfully loaded.
[I 2022-03-19 22:40:00.452 ServerApp] Serving notebooks from local directory: /lustrehome/home/s.1915438
[I 2022-03-19 22:40:00.453 ServerApp] Jupyter Server 1.15.6 is running at:
[I 2022-03-19 22:40:00.453 ServerApp] http://localhost:8888/lab?token=d9d5dd555ef63d682ec2b68232f493a4818db4bb71f1f6a1
[I 2022-03-19 22:40:00.453 ServerApp] or http://127.0.0.1:8888/lab?token=d9d5dd555ef63d682ec2b68232f493a4818db4bb71f1f6a1
[I 2022-03-19 22:40:00.453 ServerApp] Use Control-C to stop this server and shut down all kernels (twice to skip confirmation).
[C 2022-03-19 22:40:00.459 ServerApp]
To access the server, open this file in a browser:
file:///lustrehome/home/s.1915438/.local/share/jupyter/runtime/jpserver-89059-open.html
Or copy and paste one of these URLs:
http://localhost:8888/lab?token=d9d5dd555ef63d682ec2b68232f493a4818db4bb71f1f6a1
or http://127.0.0.1:8888/lab?token=d9d5dd555ef63d682ec2b68232f493a4818db4bb71f1f6a1
[W 2022-03-19 22:40:30.098 LabApp] Could not determine jupyterlab build status without nodejs
[I 2022-03-19 22:42:43.725 ServerApp] New terminal with automatic name: 1
TermSocket.open: 1
TermSocket.open: Opened 1
[I 2022-03-19 22:43:22.523 ServerApp] Writing notebook-signing key to /lustrehome/home/s.1915438/.local/share/jupyter/notebook_secret
[W 2022-03-19 22:43:22.526 ServerApp] Notebook ipynb_try/2D heat conduction.ipynb is not trusted
[I 2022-03-19 22:43:24.066 ServerApp] Kernel started: 45842dc9-891f-4ce4-94d1-12e6aa528f09
```
Copy and paste the one the link the `localhost` one or the `127.0.0.1:8888` one in your favourite browser. Bingo. It is working.
Remember, if you managed to setup a password, then you won't get the token. Simply open `http://localhost:<local_por>/` or `http://localhost:8888/`.
A screenshot:

To close the session. Go to File -> Shut Down.
```sh
[I 2022-03-19 23:33:38.361 ServerApp] Terminal 1 closed
Websocket closed
[I 2022-03-19 23:33:39.321 ServerApp] Shutting down on /api/shutdown request.
[I 2022-03-19 23:33:39.322 ServerApp] Shutting down 3 extensions
[I 2022-03-19 23:33:39.322 ServerApp] Shutting down 0 kernels
[I 2022-03-19 23:33:39.322 ServerApp] Shutting down 0 terminals
[I 2022-03-19 23:47:36.316 ServerApp] Shutting down 0 terminals
Connection to sunbird.swansea.ac.uk closed.
```
The Sunbird has `htop` preinstalled so you can see how much memory are you using. Also, you can double check if the Jupyter Lab server is still running or not.
# Wrapping ssh command in Bash file
Instead of typing this `ssh -L 8888:localhost:8888 -t s.1915438@sunbird.swansea.ac.uk "module load anaconda/3;source activate ml;jupyter-lab"` every single time, we can write a bash script.
Just create a new text file using `gedit` or `nano` and paste the above command.
Now where do you create this bash file. On Sunbird? Obviously not. If you have this question in your mind `rm -rf /` your PC.
| github_jupyter |
```
# Jupyter notebook for analyzing DOGAMI data, see Scott Tse's emergence-response notebook at
# https://github.com/hackoregon/emergency-response/blob/analytics/notebooks/census_eda_geo.ipynb
# Import modules included in jupyter container, plus modules from "kitchen-sink" script
import geopandas as gpd
import matplotlib.pyplot as plt
import numpy as np
import os
import gdal
from osgeo import osr, ogr
import pandas as pd
import psycopg2
from pyproj import Geod
import seaborn as sns
from shapely.geometry import Polygon
import sys
# Import modules NOT included in "kitchen-sink"
from dotenv import load_dotenv, find_dotenv # install python-dotenv
import geoplot as gplt # requires cython
from rasterstats import zonal_stats
%matplotlib inline
```
F.D. Pearce, 04/16/18
Notebook for computing statistics on raster pixel values contained within a geometry (shape) file
```
# Define ALL parameters in dictionary (convert to json config file!)
params = {
'raster': {
'root': './CSZ_M9p0_',
'names': ['pgv_site', 'PGD_landslide_dry', 'PGD_landslide_wet', 'PGD_liquefaction_wet'],
'ext': '.tif'
},
'geometry': {
# Try reading geometry 'from_point', if not defined, then try reading 'from_file',
# if not defined, then finally try reading 'from_postgis'
#'from_point': {
# lon_lat list in decimal degrees, xy_offset is half the width ( or height)
# of a rectangle centered on lon_lat
# xy_units MUST be m, ToDo: implement handling of different units
# 'lon_lat': [-122.6263038077892, 45.4585072924327],
# 'xy_offset': 300,
# 'xy_units': 'm'
#},
'from_file': {
'name':'./Data/DisasterNeighborhoods_FIXED_final/RLIS_ST_clips_pdx_jurisa_FIXED.dbf',
'geom_col': 'geometry'
},
'from_postgis': {
# You MUST have a .env file specifying parameters for postgis db in order to
# read data in from postgis! See sample_env.txt at
# https://github.com/hackoregon/disaster-resilience/blob/analytics/notebooks
'query': {
'table_name': 'neighborhood_units',
'select_cols': 'nuid',
#'select_cols': 'jurisdiction',
'geometry_col': 'wkb_geometry',
'epsg_code': 4326
}
}
},
'zonal_stats': {
'layer': 1,
'stats': ['count', 'min', 'max', 'mean', 'std']
},
'stats_classification': {
'stats_to_class': ['min', 'max', 'mean'],
'pgv_site': {
'levels': [-9999, 0.1, 1.1, 3.4, 8.1, 16, 31, 60, 116, 9999],
'level_labels': ['Not felt (I)', 'Weak (II-III)', 'Light (IV)',
'Moderate (V)', 'Strong (VI)', 'Very Strong (VII)',
'Severe (VIII)', 'Violent (IX)', 'Extreme (X)'],
'class_name': 'Modified Mercalli Intensity',
'class_tag': 'MMI'
},
'PGD_landslide_dry': {
'levels': [-9999, 0, 10, 30, 100, 9999],
'level_labels': ['None', 'Low', 'Moderate', 'High', 'Very High'],
'class_name': 'Landslide Intensity (Dry)',
'class_tag': 'DI'
},
'PGD_landslide_wet': {
'levels': [-9999, 0, 10, 30, 100, 9999],
'level_labels': ['None', 'Low', 'Moderate', 'High', 'Very High'],
'class_name': 'Landslide Intensity (Wet)',
'class_tag': 'DI'
},
'PGD_liquefaction_wet': {
'levels': [-9999, 0, 10, 30, 100, 9999],
'level_labels': ['None', 'Low', 'Moderate', 'High', 'Very High'],
'class_name': 'Liquefaction Intensity (Wet)',
'class_tag': 'DI'
},
'PGD_total_wet': {
'levels': [-9999, 0, 10, 30, 100, 9999],
'level_labels': ['None', 'Low', 'Moderate', 'High', 'Very High'],
'class_name': 'Total Deformation Intensity (Wet)',
'class_tag': 'DI'
}
},
'write_csv': {
'name': "./DisasterNeighborhoodsFinal_DogamiRaster_stats.csv"
#'name': "./DOGAMI_neighborhoodunits_raster_stats_v3.csv"
}
}
# Functions for obtaining geopandas dataframe containing geometry column from either
# 1) a local file (e.g. .dba file) or 2) interacting with PostGres database
def pgconnect():
"""Establish connection to PostGres database using the parameters specified in .env file.
First, walk root diretory to find and load .env file w/ PostGres variables defining database,
user, host, password, and port variables.
Then, return connection to database from psycopg2.connect
"""
try:
load_dotenv(find_dotenv())
conn = psycopg2.connect(database=os.environ.get("PG_DATABASE"), user=os.environ.get("PG_USER"),
password = os.environ.get("PG_PASSWORD"),
host=os.environ.get("PG_HOST"), port=os.environ.get("PG_PORT"))
print("Opened database successfully\n")
return conn
except psycopg2.Error as e:
print("Unable to connect to the database\n")
print(e)
print(e.pgcode)
print(e.pgerror)
#print(traceback.format_exc())
return None
def get_query_string(table_name, select_cols, geometry_col, epsg_code):
"""Build query string from parameter inputs defining table name, all the columns
to select, select_cols, the column that defines the geometry, geometry_col, and
the epsg code that defines the ellipsoid.
"""
query_string = 'SELECT ' + select_cols + ', ' + \
'ST_TRANSFORM({}, {}) AS geometry'.format(geometry_col, epsg_code)
return query_string + ' FROM {}'.format(table_name)
def get_geometry_from_postgis(postgis_params):
'''
This function takes a dictionary containing parameters for building a SQL query,
as defined in get_query_string, then connects to a postgis db, selects the
data specified in the query, and finally returns a geodataframe with a single
column named geometry that contains shape data.
'''
query_string = get_query_string(**postgis_params['query'])
conn = pgconnect()
#cur = conn.cursor()
print("SQL QUERY = "+query_string+'\r\n')
try:
geo_df = gpd.GeoDataFrame.from_postgis(
query_string,
conn,
geom_col='geometry',
crs={'init': u'epsg:{}'.format(postgis_params['query']['epsg_code'])},
coerce_float=False
)
return geo_df
except Exception as e:
print(e)
finally:
conn.close()
def get_geometry_from_file(name, geom_col='geometry'):
"""Import geometry from a file using geopandas.read_file
Returns only the geometry column!
"""
gdf = gpd.read_file(name)
#print(type(gdf))
return gdf
def calc_square_lonlat(lon_lat, xy_offset):
"""Calculate the longitude and latitude corresponding to the upper-right and lower-left
corners of a square box centered on lat_lon, with a width of 2*xy_offset.
xy_offset MUST be in meters. Uses WGS84 (epsg=4326) ellipsoid.
"""
az = [45, 225]
lon = 2*[lon_lat[0]]
lat = 2*[lon_lat[1]]
mag = 2*[np.sqrt(2)*xy_offset]
g = Geod(ellps='WGS84')
rl_lon, tb_lat, _ = g.fwd(lon, lat, az, mag)
return (rl_lon, tb_lat)
def calc_square_polygon(lon_lat, xy_offset):
"""Calculate polygon defining square box centered on lon, lat (decimal degrees)
with a width of 2*xy_offset (meters).
"""
rl_lon, tb_lat = calc_square_lonlat(lon_lat, xy_offset)
poly = [Polygon((
(rl_lon[0], tb_lat[0]),
(rl_lon[0], tb_lat[1]),
(rl_lon[1], tb_lat[1]),
(rl_lon[1], tb_lat[0])
))]
return poly
def get_geometry_from_point(lon_lat, xy_offset, xy_units="m"):
"""Returns a geodataframe containing a single geometry column that
defines a square box centered on a point, specified as a lat, lon pair,
The input parameter xy_offset defines the box half-width
1) Calculate the top-right corner and bottom-left corner of square box
centered on lon_lat, using the WGS84 ellipsoid.
2) Use the lon, lat of each corner to build a rectangular
polygon using shapely Polygon.
3) Convert polygon to pandas geodataframe, set coordinate reference to
epsg 4326 (equivalent to WGS84).
"""
if xy_units == "m":
poly = calc_square_polygon(lon_lat, xy_offset)
# Build geodataframe with one row, column
gdf = gpd.GeoDataFrame(poly, columns=['geometry'], geometry='geometry')
gdf.crs = {'init' :'epsg:4326'}
return gdf
else:
print("Error: input xy_offset MUST be in meters!!!")
def get_geodf_geometry(**kwargs):
"""Import geometry either from point and size, a file, or from postgis db
"""
if 'from_point' in kwargs:
return get_geometry_from_point(**kwargs['from_point'])
elif 'from_file' in kwargs:
return get_geometry_from_file(**kwargs['from_file'])
elif 'from_postgis' in kwargs:
return get_geometry_from_postgis(kwargs['from_postgis'])
# Functions for manipulating geoshapes and raster files
def get_gdfcrs_epsg(gdf):
"""Return integer EPSG code corresponding to Coordinate Reference
used in input geodataframe, gdf. Attribute gdf.crs must contain
a dict with key = 'init' that contains a string starting with 'epsg',
followed by a colon, followed by an integer as a string.
"""
try:
dfepsg = gdf.crs['init'].split(':')
if dfepsg[0] == 'epsg':
return int(dfepsg[1])
except:
print('Error: geodataframe crs = {}, unrecognized EPSG integer'.format(gdf.crs['init']))
def get_raster_nodatavalue(rasterfn):
raster = gdal.Open(rasterfn)
band = raster.GetRasterBand(1)
return band.GetNoDataValue()
def get_raster_info_crs(raster_file, print_info=True):
"""Print information about raster file, and return its
spatial reference system using gdal.
"""
if print_info:
try:
print(gdal.Info(raster_file))
except:
print("Error reading info from raster file = {}".format(raster_file))
try:
raster = gdal.Open(raster_file)
except:
print("Error opening raster file = {}".format(raster_file))
else:
raster_crs = osr.SpatialReference()
raster_crs.ImportFromWkt(raster.GetProjection())
return raster_crs.ExportToProj4()
def transform_gdf_to_crsout(gdf, geom_col, crs_out):
"""Transform list of georeferenced polygon geometries from geopandas
dataframe geometry column, geom_col, to the desired output Spatial Reference, srs_out.
"""
geom_out = gdf[geom_col].copy()
return geom_out.to_crs(crs_out)
def transform_polygons_from_srsinp_to_srsout(geom_col, srs_out):
"""Transform list of georeferenced polygon geometries from geopandas
dataframe geometry column, geom_col, to the desired output Spatial Reference, srs_out.
The input geometry column, geom_col, MUST have a valid epsg code defining its Spatial
Reference (SRS)."""
# Define input spatial reference using epsg code from gdf
srs_inp = ogr.osr.SpatialReference()
srs_inp.ImportFromEPSG(get_gdfcrs_epsg(geom_col))
poly_out = []
# Define list of polygons in transformed spatial reference
for g in geom_col:
# If MultiPolygon, then assume it contains only one Polygon
if g.type == 'MultiPolygon':
poly = ogr.CreateGeometryFromWkt(g.geoms[0].wkt)
elif g.type == 'Polygon':
# Need to test this
poly = ogr.CreateGeometryFromWkt(g.wkt)
else:
print("Error: geometry = {}, MUST be Polygon or MultiPolygon".format(g.type))
poly.AssignSpatialReference(srs_inp)
# Transform point co-ordinates so that they are in same projection as raster
poly.Transform(osr.CoordinateTransformation(srs_inp, srs_out))
poly_out.append(poly.ExportToWkt())
return poly_out
# Functions for computing raster statistics
def get_raster_stats_df(geom_ras, df_index, raster_file, raster_name, **kwargs):
"""Compute raster statistics for input geometry and raster file.
Return results in dataframe
"""
kwargs.update({'nodata_value': get_raster_nodatavalue(raster_file)})
#print(kwargs)
geomstats = zonal_stats(geom_ras, raster_file, **kwargs)
df_gs = pd.DataFrame(geomstats, index=df_index)
df_gs.rename(columns={co: raster_name+'_'+co for co in df_gs.columns}, inplace=True)
return df_gs
# Functions for classifying raster statistics
def get_stats_classification(gdf, **kwargs):
"""Classify raster statistics using specified parameters in kwargs"""
raster_names = [rn for rn in kwargs.keys() if rn != "stats_to_class"]
for rn in raster_names:
stats_to_class = [rn+'_'+sc for sc in kwargs['stats_to_class']]
levels = kwargs[rn]['levels']
labels = kwargs[rn]['level_labels']
ctag = '_' + kwargs[rn]['class_tag']
for s2c in stats_to_class:
try:
gdf[s2c+ctag] = pd.cut(gdf[s2c], levels, right=True, labels=labels)
except KeyError:
print("Key Error exception occurred for raster stat key = {}".format(s2c))
except Exception as e:
print("A non-key error exception occurred: {}".format(e))
return gdf
#help(gpd.read_file)
# Step 1) Select geometry column either from Postgis db (implemented), or
# from shapefile (not yet implemented). In eithe case, make sure geometry
# has a valid epsg Spatial reference assigned to it, such as 4326 (lon/lat)
# For a Postgis-derived geometry, this is done on the db-side using ST_TRANSFORM
# Note a copy of the original geopandas dataframe is made to preserve the dataframe
# obtained from postgis for debugging :-)
gdf = get_geodf_geometry(**params['geometry'])
gdf_merge = gdf.copy()
gdf_merge.info()
#gdf_merge = gdf.iloc[0:5].copy()
#gdf.info()
#gdf_merge.info()
#print(gdf['geometry'][0].type)
#print(gdf['geometry'].crs)
# Steps 2 through 5 are repeated for each raster file
# the results for each raster are appended to gdf_merge
for raster_name in params['raster']['names']:
# Step 2) Print info about tif file (optional) and get its spatial reference info
raster_file = params['raster']['root'] + raster_name + params['raster']['ext']
print("Computing statistics for raster file = {}".format(raster_file))
crs_raster = get_raster_info_crs(raster_file, print_info=False)
# Step 3) Generate a list of polygons transformed from the srs used in the
# input geodataframe, gdf, to the srs used in the raster file, crs_raster
geom_ras = transform_gdf_to_crsout(gdf_merge, 'geometry', crs_raster)
#geom_ras = transform_polygons_from_srsinp_to_srsout(gdf_merge['geometry'], srs_raster)
# Step 4) Use rasterstats to compute analytics on pixel values within specified geometry,
# MUST be polygon or multipolygon and transformed to srs_raster!
# Add stats from pixel values into geodataframe that defines geometry
df_gs = get_raster_stats_df(geom_ras, gdf_merge.index, raster_file, raster_name,
**params['zonal_stats']
)
# Step 5) Aggregate the statistics from each raster into a final merged geodataframe
gdf_merge = gdf_merge.join(df_gs)
# Compute total deformation for wet conditions by adding the mean landslide deformation
# to the mean liquefaction deformation. This should NOT be interpreted as the total
# deformation for a particular point, but instead represents the overall risk of permanent
# deformation to a given neighborhood. Only do this for the mean as is adding together any
# of the other statistics doesn't seem justifiable to me.
gdf_merge['PGD_total_wet_mean'] = gdf_merge['PGD_landslide_wet_mean'] + gdf_merge['PGD_liquefaction_wet_mean']
# Step 6) Classify a subset of the geometry statistics, converting the calculated
# stat in pixel values to a label describing the stats intensity bin
if 'stats_classification' in params:
gdf_merge_class = get_stats_classification(gdf_merge, **params['stats_classification'])
gdf_merge_class.info()
#print(gdf_merge_class[['PGD_landslide_dry_max', 'PGD_landslide_dry_max_DI']])
# Remove columns that aren't required
gdf_merge_class = gdf_merge_class.drop(columns=['fid_1', 'area', 'sqmile', 'Area_Recal', 'Perim', 'Isop'])
gdf_merge_class.info()
# Write results to csv file
if 'write_csv' in params:
gdf_merge_class.to_csv(params['write_csv']['name'])
```
### Testing
```
gdf_merge_class.info()
gdf_merge_class_nonull = gdf_merge_class[~gdf_merge_class.isnull().any(axis=1)].copy()
gdf_merge_class_nonull['PGD_liquefaction_wet_max_DI'] = pd.Categorical(
gdf_merge_class_nonull['PGD_liquefaction_wet_max_DI'],
categories=params['stats_classification']['PGD_liquefaction_wet']['level_labels'],
dtype="category", ordered=True
)
gdf_merge_class_nonull['PGD_liquefaction_wet_max_DI']
#If geometry was build from point, then check the length of square to make
# sure it matches the input length, xy_offset
if 'from_point' in params['geometry']:
poly_coords = list(gdf_merge_class['geometry'][0].exterior.coords)
print(poly_coords)
g = Geod(ellps='WGS84')
az12,az21,dist = g.inv(poly_coords[0][0], poly_coords[0][1], poly_coords[1][0], poly_coords[1][1])
print("The returned azimuth, {}, should point N-S (i.e. 0 or 180)".format(az12))
print("The returned distance, {}, should be equal to 2*xy_offset, {}, within roundoff."
.format(dist, 2*params['geometry']['from_point']['xy_offset']
))
```
### Plotting
```
# Plot column value for each geometry contained in gdf
# there is a bug in geopandas plot function when column value
# is categorical and ordered! Misorders labels and/or omits labels!
col2plot = 'PGD_total_wet_mean'
#col2plot = 'pgv_site_mean'
#help(gdf_merge_class.plot)
categorical = False
savefig = False
if categorical:
k = None
scheme = None
else:
k = 7
scheme = 'Equal_interval'
# Plotting
fig, ax = plt.subplots(1, figsize=(8, 8))
basemap = gdf_merge_class.plot(column=col2plot, cmap='Purples', ax=ax, k=k,
categorical=categorical, scheme=scheme, legend=True, linewidth=1, edgecolor='k'
)
#basemap = gplt.choropleth(gdf_merge_class, hue=col2plot, projection=gplt.crs.PlateCarree(),
# cmap='Purples', categorical=categorical, legend=True, linewidth=1, edgecolor='k',
# legend_labels=['None', 'Low', 'Moderate', 'High', 'Very High']
#)
ax.set_aspect('equal')
ax.set_title(col2plot, fontsize=16)
plt.tight_layout()
if savefig:
plt.savefig(col2plot+"final.png", bbox_inches='tight', pad_inches=0.1)
gdf_merge_class['PGD_landslide_dry_max_DI']
#help(gdf_merge_class.plot)
#help(gplt.choropleth)
# Plot a single class label within a given categorical column
# This is for categorical data ONLY!
#cmap = plt.cm.Purples(0.5)
#print(cmap)
colbase = 'PGD_total_wet'
col2plot = colbase + '_mean_DI'
#colbase = 'pgv_site'
#col2plot = colbase + '_mean_MMI'
categorical = True
# Plotting
num_labels = len(params['stats_classification'][colbase]['level_labels'])
fig, ax = plt.subplots(1, figsize=(8, 8))
legend_patches = []
import matplotlib.patches as mpatches
for ind, lab in enumerate(params['stats_classification'][colbase]['level_labels']):
#print(ind/num_labels)
gdf_catplot = gdf_merge_class.loc[gdf_merge_class[col2plot]==lab, ['geometry', col2plot]]
#print(gdf_catplot)
if not gdf_catplot.empty:
ph = gdf_catplot.plot(color=plt.cm.Purples(ind/(num_labels-1)), ax=ax,
categorical=categorical, linewidth=1, edgecolor='k', legend=True
)
legend_patches.append(mpatches.Patch(color=plt.cm.Purples(ind/(num_labels-1)), label=lab))
ax.set_aspect('equal')
ax.set_title(col2plot, fontsize=16)
ax.legend(handles=legend_patches)
#plt.legend(handles=[red_patch])
#ax.legend(labels=params['stats_classification'][colbase]['level_labels'])
plt.tight_layout()
plt.savefig(col2plot+"final.png", bbox_inches='tight', pad_inches=0.1)
gdf_catplot = gdf_merge_class.loc[gdf_merge_class[col2plot]=="None", ['geometry', col2plot]]
print(gdf_catplot)
gdf_catplot = gdf_merge_class.loc[gdf_merge_class[col2plot]=="Moderate", ['geometry', col2plot]]
print(not gdf_catplot.empty)
```
| github_jupyter |
# Transfer Learning with TensorFlow Hub for TFLite
## Set up library versions for TF2
```
# !pip uninstall tensorflow --yes
!pip install -U --pre -q tensorflow-gpu==2.0.0-beta1
# !pip install -U --pre -q tf-nightly-gpu-2.0-preview==2.0.0.dev20190715
# Last tested version: 2.0.0-dev20190704
# !pip install -U --pre -q tf-estimator-nightly==1.14.0.dev2019071001
# !pip uninstall tensorflow-hub --yes
# !pip install -U --pre -q tf-hub-nightly==0.6.0.dev201907150002
# Last tested version: Hub version: 0.6.0.dev201907160002
from __future__ import absolute_import, division, print_function
import os
import matplotlib.pylab as plt
import numpy as np
import tensorflow as tf
import tensorflow_hub as hub
print("Version: ", tf.__version__)
print("Eager mode: ", tf.executing_eagerly())
print("Hub version: ", hub.__version__)
print("GPU is", "available" if tf.test.is_gpu_available() else "NOT AVAILABLE")
```
## Select the Hub/TF2 module to use
Hub modules for TF 1.x won't work here, please use one of the selections provided.
```
module_selection = ("mobilenet_v2", 224, 1280) #@param ["(\"mobilenet_v2\", 224, 1280)", "(\"inception_v3\", 299, 2048)"] {type:"raw", allow-input: true}
handle_base, pixels, FV_SIZE = module_selection
MODULE_HANDLE ="https://tfhub.dev/google/tf2-preview/{}/feature_vector/4".format(handle_base)
IMAGE_SIZE = (pixels, pixels)
print("Using {} with input size {} and output dimension {}".format(
MODULE_HANDLE, IMAGE_SIZE, FV_SIZE))
```
## Data preprocessing
Use [TensorFlow Datasets](http://tensorflow.org/datasets) to load the cats and dogs dataset.
This `tfds` package is the easiest way to load pre-defined data. If you have your own data, and are interested in importing using it with TensorFlow see [loading image data](../load_data/images.ipynb)
```
import tensorflow_datasets as tfds
tfds.disable_progress_bar()
```
The `tfds.load` method downloads and caches the data, and returns a `tf.data.Dataset` object. These objects provide powerful, efficient methods for manipulating data and piping it into your model.
Since `"cats_vs_dog"` doesn't define standard splits, use the subsplit feature to divide it into (train, validation, test) with 80%, 10%, 10% of the data respectively.
```
splits = tfds.Split.ALL.subsplit(weighted=(80, 10, 10))
splits, info = tfds.load('cats_vs_dogs', with_info=True, as_supervised=True, split = splits)
(train_examples, validation_examples, test_examples) = splits
num_examples = info.splits['train'].num_examples
num_classes = info.features['label'].num_classes
```
### Format the Data
Use the `tf.image` module to format the images for the task.
Resize the images to a fixes input size, and rescale the input channels
```
def format_image(image, label):
image = tf.image.resize(image, IMAGE_SIZE) / 255.0
return image, label
```
Now shuffle and batch the data
```
BATCH_SIZE = 32 #@param {type:"integer"}
train_batches = train_examples.shuffle(num_examples // 4).map(format_image).batch(BATCH_SIZE).prefetch(1)
validation_batches = validation_examples.map(format_image).batch(BATCH_SIZE).prefetch(1)
test_batches = test_examples.map(format_image).batch(1)
```
Inspect a batch
```
for image_batch, label_batch in train_batches.take(1):
pass
image_batch.shape
```
## Defining the model
All it takes is to put a linear classifier on top of the `feature_extractor_layer` with the Hub module.
For speed, we start out with a non-trainable `feature_extractor_layer`, but you can also enable fine-tuning for greater accuracy.
```
do_fine_tuning = False #@param {type:"boolean"}
```
Load TFHub Module
```
feature_extractor = hub.KerasLayer(MODULE_HANDLE,
input_shape=IMAGE_SIZE + (3,),
output_shape=[FV_SIZE],
trainable=do_fine_tuning)
print("Building model with", MODULE_HANDLE)
model = tf.keras.Sequential([
feature_extractor,
tf.keras.layers.Dense(num_classes, activation='softmax')
])
model.summary()
#@title (Optional) Unfreeze some layers
NUM_LAYERS = 10 #@param {type:"slider", min:1, max:50, step:1}
if do_fine_tuning:
feature_extractor.trainable = True
for layer in model.layers[-NUM_LAYERS:]:
layer.trainable = True
else:
feature_extractor.trainable = False
```
## Training the model
```
if do_fine_tuning:
model.compile(
optimizer=tf.keras.optimizers.SGD(lr=0.002, momentum=0.9),
loss=tf.keras.losses.SparseCategoricalCrossentropy(),
metrics=['accuracy'])
else:
model.compile(
optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
EPOCHS = 5
hist = model.fit(train_batches,
epochs=EPOCHS,
validation_data=validation_batches)
```
## Export the model
```
CATS_VS_DOGS_SAVED_MODEL = "exp_saved_model"
```
Export the SavedModel
```
tf.saved_model.save(model, CATS_VS_DOGS_SAVED_MODEL)
%%bash -s $CATS_VS_DOGS_SAVED_MODEL
saved_model_cli show --dir $1 --tag_set serve --signature_def serving_default
loaded = tf.saved_model.load(CATS_VS_DOGS_SAVED_MODEL)
print(list(loaded.signatures.keys()))
infer = loaded.signatures["serving_default"]
print(infer.structured_input_signature)
print(infer.structured_outputs)
```
## Convert using TFLite's Converter
```
```
Load the TFLiteConverter with the SavedModel
```
converter = tf.lite.TFLiteConverter.from_saved_model(CATS_VS_DOGS_SAVED_MODEL)
```
### Post-training quantization
The simplest form of post-training quantization quantizes weights from floating point to 8-bits of precision. This technique is enabled as an option in the TensorFlow Lite converter. At inference, weights are converted from 8-bits of precision to floating point and computed using floating-point kernels. This conversion is done once and cached to reduce latency.
To further improve latency, hybrid operators dynamically quantize activations to 8-bits and perform computations with 8-bit weights and activations. This optimization provides latencies close to fully fixed-point inference. However, the outputs are still stored using floating point, so that the speedup with hybrid ops is less than a full fixed-point computation.
```
converter.optimizations = [tf.lite.Optimize.DEFAULT]
```
### Post-training integer quantization
We can get further latency improvements, reductions in peak memory usage, and access to integer only hardware accelerators by making sure all model math is quantized. To do this, we need to measure the dynamic range of activations and inputs with a representative data set. You can simply create an input data generator and provide it to our converter.
```
def representative_data_gen():
for input_value, _ in test_batches.take(100):
yield [input_value]
converter.representative_dataset = representative_data_gen
```
The resulting model will be fully quantized but still take float input and output for convenience.
Ops that do not have quantized implementations will automatically be left in floating point. This allows conversion to occur smoothly but may restrict deployment to accelerators that support float.
### Full integer quantization
To require the converter to only output integer operations, one can specify:
```
converter.target_spec.supported_ops = [tf.lite.OpsSet.TFLITE_BUILTINS_INT8]
```
### Finally convert the model
```
tflite_model = converter.convert()
tflite_model_file = 'converted_model.tflite'
with open(tflite_model_file, "wb") as f:
f.write(tflite_model)
```
##Test the TFLite model using the Python Interpreter
```
# Load TFLite model and allocate tensors.
interpreter = tf.lite.Interpreter(model_path=tflite_model_file)
interpreter.allocate_tensors()
input_index = interpreter.get_input_details()[0]["index"]
output_index = interpreter.get_output_details()[0]["index"]
from tqdm import tqdm
# Gather results for the randomly sampled test images
predictions = []
test_labels, test_imgs = [], []
for img, label in tqdm(test_batches.take(10)):
interpreter.set_tensor(input_index, img)
interpreter.invoke()
predictions.append(interpreter.get_tensor(output_index))
test_labels.append(label.numpy()[0])
test_imgs.append(img)
#@title Utility functions for plotting
# Utilities for plotting
class_names = ['cat', 'dog']
def plot_image(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array[i], true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
img = np.squeeze(img)
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'green'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
```
NOTE: Colab runs on server CPUs. At the time of writing this, TensorFlow Lite doesn't have super optimized server CPU kernels. For this reason post-training full-integer quantized models may be slower here than the other kinds of optimized models. But for mobile CPUs, considerable speedup can be observed.
```
#@title Visualize the outputs { run: "auto" }
index = 1 #@param {type:"slider", min:0, max:9, step:1}
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(index, predictions, test_labels, test_imgs)
plt.show()
```
Download the model
```
from google.colab import files
files.download('converted_model.tflite')
labels = ['cat', 'dog']
with open('labels.txt', 'w') as f:
f.write('\n'.join(labels))
files.download('labels.txt')
```
# Prepare the test images for download (Optional)
This part involves downloading additional test images for the Mobile Apps only in case you need to try out more samples
```
!mkdir -p test_images
from PIL import Image
for index, (image, label) in enumerate(test_batches.take(50)):
image = tf.cast(image * 255.0, tf.uint8)
image = tf.squeeze(image).numpy()
pil_image = Image.fromarray(image)
pil_image.save('test_images/{}_{}.jpg'.format(class_names[label[0]], index))
!ls test_images
!zip -qq cats_vs_dogs_test_images.zip -r test_images/
files.download('cats_vs_dogs_test_images.zip')
```
| github_jupyter |
# HW #2, Introduction to Big Data Analytics, Fall 2019
## 2. Dimensionality reduction
### Exercise 11.1.7
```
import numpy as np
from numpy.linalg import norm, matrix_rank
def principal_eigenpair(M, epsilon=1e-6):
'''
Find principal eigenvector and eigenvalue using power iteration method.
'''
M = np.array(M)
nCol, nRow = M.shape
x = np.ones((nRow, 1))
x_prev = np.full((nRow, 1), fill_value=np.inf)
while norm(x - x_prev) > epsilon:
p = np.matmul(M, x)
x_prev, x = x, p/norm(p)
l = np.matmul(np.matmul(x.T, M), x).flatten()
return x, l
def subtract_eigenpair(M, e, l):
return M - l*np.matmul(e, e.T)
def find_all_eigenpairs(M):
M_ = []
e_ = []
l_ = []
for _ in range(matrix_rank(M)):
e, l = principal_eigenpair(M)
M_.append(M)
e_.append(e)
l_.append(l)
M = subtract_eigenpair(M, e, l)
return M_, e_, l_
''' problem (a) and (b) '''
M = np.array([[1,1,1], [1,2,3], [1,3,6]])
e, l = principal_eigenpair(M)
print e, l
''' problem (c) '''
print subtract_eigenpair(M, e, l)
''' problem (d) and (e) '''
print find_all_eigenpairs(M)
```
### Exercise 11.3.1
```
import numpy as np
np.set_printoptions(precision=3, suppress=True)
M = np.array([[1, 2, 3],
[3, 4, 5],
[5, 4, 3],
[0, 2, 4],
[1, 3, 5]])
''' problem (a) '''
mtm = np.matmul(M.T, M)
mmt = np.matmul(M, M.T)
print mtm
print mmt
''' problem (b) '''
# eigenpairs of MtM
mtm_eigvals, mtm_eigvecs = np.linalg.eig(mtm)
print mtm_eigvals
print mtm_eigvecs
# eigenpairs of MMt
mmt_eigvals, mmt_eigvecs = np.linalg.eig(mmt)
mmt_eigvals = np.real(mmt_eigvals)
mmt_eigvecs = np.real(mmt_eigvecs)
print mmt_eigvals
print mmt_eigvecs
''' problem (c) '''
# construct V
V = mtm_eigvecs[:, [0,1]]
# construct U
U = mmt_eigvecs[:, [0,2]]
U[:, 0] = -U[:, 0]
# construct sigma
d = np.sqrt(mtm_eigvals[[0,1]])
sigma = np.diagflat(d)
print U
print sigma
print V
'''problem (d)'''
V_ = V[:, 0:1]
U_ = U[:, 0:1]
sigma_ = sigma[0:1, 0:1]
M_ = np.matmul(np.matmul(U_, sigma_), V_.T)
print M_
'''problem (e)'''
retained_energy = (sigma[0,0]**2) / (sigma[0,0]**2 + sigma[1,1]**2)
print retained_energy
```
### Exercise 11.4.2 (CUR decomposition)
```
import numpy as np
from numpy.linalg import eig, norm
M = np.array([[1,1,1,0,0],
[3,3,3,0,0],
[4,4,4,0,0],
[5,5,5,0,0],
[0,0,0,4,4],
[0,0,0,5,5],
[0,0,0,2,2]])
def CUR_verify(M, i, j, r=2):
p = (np.sum(M**2, axis=1)/norm(M)**2).flatten() # distribution of i
q = (np.sum(M**2, axis=0)/norm(M)**2).flatten() # distribution of j
C = M[:, j]
R = M[i, :]
C = C / np.sqrt(r*q[j][np.newaxis, ...])
R = R / np.sqrt(r*p[i][..., np.newaxis])
W = M[i, :][:, j]
X, s, Yh = np.linalg.svd(W)
S = np.diagflat(s)
Sp = np.linalg.pinv(S)
print Sp
U = np.matmul(np.matmul(Yh.T, np.matmul(Sp, Sp)), X.T)
return C, U, R
'''problem (a)'''
i, j = [1,2], [0,1]
# construct C and R
p = (np.sum(M**2, axis=1)/norm(M)**2).flatten() # distribution of i
q = (np.sum(M**2, axis=0)/norm(M)**2).flatten() # distribution of j
C = M[:, j] / np.sqrt(r*q[j][np.newaxis, ...])
R = M[i, :] / np.sqrt(r*p[i][..., np.newaxis])
# construct U
W = M[i, :][:, j]
# SVD of W
d, Y = np.linalg.eig(np.matmul(W.T, W))
_, X = np.linalg.eig(np.matmul(W, W.T))
X = -X[:, [1,0]]
sigma = np.diagflat(np.sqrt(d))
# compute Moore-Penrose pseudoinverse of sigma
d[0] = 1/d[0]
sigma_p = np.diagflat(np.sqrt(d))
U = np.matmul(np.matmul(Y,np.matmul(sigma_p, sigma_p)), X.T)
print C
print U
print R
'''problem (b)'''
i, j = [3,4], [1,2]
# construct C and R
p = (np.sum(M**2, axis=1)/norm(M)**2).flatten() # distribution of i
q = (np.sum(M**2, axis=0)/norm(M)**2).flatten() # distribution of j
C = M[:, j] / np.sqrt(r*q[j][np.newaxis, ...])
R = M[i, :] / np.sqrt(r*p[i][..., np.newaxis])
# construct U
W = M[i, :][:, j]
# SVD of W
d, Y = np.linalg.eig(np.matmul(W.T, W))
_, X = np.linalg.eig(np.matmul(W, W.T))
sigma = np.diagflat(np.sqrt(d))
# compute Moore-Penrose pseudoinverse of sigma
d[0] = 1/d[0]
sigma_p = np.diagflat(np.sqrt(d))
U = np.matmul(np.matmul(Y,np.matmul(sigma_p, sigma_p)), X.T)
print C
print U
print R
'''problem (c)'''
i, j = [0,6], [0,4]
# construct C and R
p = (np.sum(M**2, axis=1)/norm(M)**2).flatten() # distribution of i
q = (np.sum(M**2, axis=0)/norm(M)**2).flatten() # distribution of j
C = M[:, j] / np.sqrt(r*q[j][np.newaxis, ...])
R = M[i, :] / np.sqrt(r*p[i][..., np.newaxis])
# construct U
W = M[i, :][:, j]
# SVD of W
d, Y = np.linalg.eig(np.matmul(W.T, W))
_, X = np.linalg.eig(np.matmul(W, W.T))
sigma = np.diagflat(np.sqrt(d))
# compute Moore-Penrose pseudoinverse of sigma
d = 1/d
sigma_p = np.diagflat(np.sqrt(d))
U = np.matmul(np.matmul(Y,np.matmul(sigma_p, sigma_p)), X.T)
print C
print U
print R
```
| github_jupyter |
<a id="title_ID"></a>
# JWST Pipeline Validation Notebook: calwebb_detector1, firstframe unit tests
<span style="color:red"> **Instruments Affected**</span>: MIRI
### Table of Contents
<div style="text-align: left">
<br> [Introduction](#intro)
<br> [JWST Unit Tests](#unit)
<br> [Defining Terms](#terms)
<br> [Test Description](#description)
<br> [Data Description](#data_descr)
<br> [Imports](#imports)
<br> [Convenience Functions](#functions)
<br> [Perform Tests](#testing)
<br> [About This Notebook](#about)
<br>
</div>
<a id="intro"></a>
# Introduction
This is the validation notebook that displays the unit tests for the Firstframe step in calwebb_detector1. This notebook runs and displays the unit tests that are performed as a part of the normal software continuous integration process. For more information on the pipeline visit the links below.
* Pipeline description: https://jwst-pipeline.readthedocs.io/en/latest/jwst/firstframe/index.html
* Pipeline code: https://github.com/spacetelescope/jwst/tree/master/jwst/
[Top of Page](#title_ID)
<a id="unit"></a>
# JWST Unit Tests
JWST unit tests are located in the "tests" folder for each pipeline step within the [GitHub repository](https://github.com/spacetelescope/jwst/tree/master/jwst/), e.g., ```jwst/firstframe/tests```.
* Unit test README: https://github.com/spacetelescope/jwst#unit-tests
[Top of Page](#title_ID)
<a id="terms"></a>
# Defining Terms
These are terms or acronymns used in this notebook that may not be known a general audience.
* JWST: James Webb Space Telescope
* NIRCam: Near-Infrared Camera
[Top of Page](#title_ID)
<a id="description"></a>
# Test Description
Unit testing is a software testing method by which individual units of source code are tested to determine whether they are working sufficiently well. Unit tests do not require a separate data file; the test creates the necessary test data and parameters as a part of the test code.
[Top of Page](#title_ID)
<a id="data_descr"></a>
# Data Description
Data used for unit tests is created on the fly within the test itself, and is typically an array in the expected format of JWST data with added metadata needed to run through the pipeline.
[Top of Page](#title_ID)
<a id="imports"></a>
# Imports
* tempfile for creating temporary output products
* pytest for unit test functions
* jwst for the JWST Pipeline
* IPython.display for display pytest reports
[Top of Page](#title_ID)
```
import tempfile
import pytest
import jwst
from IPython.display import IFrame
```
<a id="functions"></a>
# Convenience Functions
Here we define any convenience functions to help with running the unit tests.
[Top of Page](#title_ID)
```
def display_report(fname):
'''Convenience function to display pytest report.'''
return IFrame(src=fname, width=700, height=600)
```
<a id="testing"></a>
# Perform Tests
Below we run the unit tests for the Firstframe step.
[Top of Page](#title_ID)
```
with tempfile.TemporaryDirectory() as tmpdir:
!pytest jwst/firstframe -v --ignore=jwst/associations --ignore=jwst/datamodels --ignore=jwst/stpipe --ignore=jwst/regtest --html=tmpdir/unit_report.html --self-contained-html
report = display_report('tmpdir/unit_report.html')
report
```
<a id="about"></a>
## About This Notebook
**Author:** Alicia Canipe, Staff Scientist, NIRCam
<br>**Updated On:** 01/07/2021
[Top of Page](#title_ID)
<img style="float: right;" src="./stsci_pri_combo_mark_horizonal_white_bkgd.png" alt="stsci_pri_combo_mark_horizonal_white_bkgd" width="200px"/>
| github_jupyter |
#タンパク質折りたたみ問題
量子アニーリングを用いた創薬関連のタンパク質折りたたみ問題がハーバード大学の先生によって2012年に発表されていました。そのタンパク質折りたたみ問題の論文を元に実際にwildqatで解いてみたいと思います。
##参考にする論文
natureに掲載されているこちらの論文をベースにします。
Finding low-energy conformations of lattice protein models by quantum annealing
Alejandro Perdomo-Ortiz, Neil Dickson, Marshall Drew-Brook, Geordie Rose & Alán Aspuru-Guzik
Scientific Reports volume 2, Article number: 571 (2012)
https://www.nature.com/articles/srep00571
##問題の概要とHPモデル、MJモデル
タンパク質を単純にモデル化をしてそれをイジングモデルモデルに落とし込むという試みです。
まずは、HPモデルというモデルを使用しています。
HPモデル
• アミノ酸をH(疎水性、非極性アミノ酸)とP(親水性、極性アミノ酸)のいずれかに分ける。
• Hは、水を嫌い、互いに引き付けあう
参考:HPモデルhttp://www.iba.t.u-tokyo.ac.jp/iba/AI/HP.pdf
Mijazawa-Jernigan (MJ) model
今回は単純化されたMJモデルを利用します。
##MJモデルのQUBOへの適用
用意された塩基列を特定の方向に回転させる操作をイジングモデルに対応させています。
<img src="https://github.com/Blueqat/Wildqat/blob/master/examples_ja/img/024_5.png?raw=1">
引用:https://www.nature.com/articles/srep00571
今回使用する塩基列は、PSVKMAの配列で、
下記のように特定の塩基列が隣接すると安定状態になり、エネルギーがへります。このエネルギーの安定化を使っってコスト関数を最小化させることを狙います。
<img src="https://github.com/Blueqat/Wildqat/blob/master/examples_ja/img/024_0.png?raw=1">
また、今回塩基列を全て一度に処理するのは難しいのでいくつかのパターンに分けます。
<img src="https://github.com/Blueqat/Wildqat/blob/master/examples_ja/img/024_1.png?raw=1">
引用:https://www.nature.com/articles/srep00571
上記のようにすでにいくつかの折りたたまれたパターンから出発して安定状態を求めます。数が多くないので書き出すことができ、もっとも低いエネルギー安定状態は下記のようになります。それぞれのパターンに対して安定状態に到達できる形状が異なるようなので、どれか1つのschemeを取り上げて一番エネルギーの低い状態を立式から導き出したいと思います。
<img src="https://github.com/Blueqat/Wildqat/blob/master/examples_ja/img/024_2.png?raw=1">
引用:https://www.nature.com/articles/srep00571
##コスト関数
今回のコスト関数は、
$$E_p = E_{onsite} + E_{px} + E_{ext}$$
となります。1項目はタンパク質の塩基列が重ならないという条件、2項目は塩基列同士の近接の相互作用のエネルギー関数、3項目は外部からの影響です。今回3項目は使用しないということなので、
$$E_p = E_{onsite} + E_{pw}$$
となります。
##モデル
今回、論文中にも触れられている実験3をやって見たいと思います。塩基列は、
<img src="https://github.com/Blueqat/Wildqat/blob/master/examples_ja/img/024_3.png?raw=1">
この順番ですが、今回はある程度折りたたまれた状態で始まります。
<img src="https://github.com/Blueqat/Wildqat/blob/master/examples_ja/img/024_4.png?raw=1">
この場合、PSKVまでは決まっていて、Mは下か左が決まっていますので、回転方向はPから順番に書いてみると、
$$010010q_10q_2q_3$$
となり、3量子ビットの式を解けばよいことになります。$01$は右、$00$は下、$10$は左、Mは下か左しかないので、$00$か$10$なので、$0$は決まっていて、残りの$q_1$から$q_3$をイジング問題で求めます。
コスト関数は、
$$E = -1-4q_3+9q_1q_3+9q_2q_3-16q_1q_2q_3$$
##3体問題の2体問題への分解
ここで、QUBOではそのままでは解けない3体問題がでてきます。この際には数学変換で2体問題へと分解をします。詳しくは他のチュートリアルを参照ください。
新しい量子ビット$q_4$を導入して、
$$q_4=q_1q_2$$
これにペナルティ項を導入するとコスト関数は、
$$E = -1-4q_3+9q_1q_3+9q_2q_3-16q_3q_4+\delta(3q_4+q_1q_2-2q_1q_4-2q_2q_4)$$
こちらをWildqatに入れて計算をして見ます。
##Wildqatへ実装
ここで上記のQUBOを実装します。
デルタの値を10として見てとくと、
```
!pip install blueqat
import blueqat.wq as wq
a = wq.Opt()
d = 10;
a.qubo = [[0,d,9,-2*d],[0,0,9,-2*d],[0,0,-4,-16],[0,0,0,3*d]]
a.sa()
```
答えは$0010$となりました、QUBOで表現されたタンパク質の折りたたみは、
$0100100010$
という回転を表現することになり、下記のようなものがもっとも安定なものとなります。
<img src="https://github.com/Blueqat/Wildqat/blob/master/examples_ja/img/024_7.png?raw=1">
| github_jupyter |
Najprej skonstruiramo random slovar. Kateri nam vrne za vsako povezavo svoj uncertainty set.
```
#ta sedaj c-je poda kot seznam seznamov
def seznam_cen(st_opazanj, st_pov_grafa, min_cena, max_cena):
sur_podatki = []
for i in range(0, st_opazanj):
mer_povezav = []
for j in range(0, st_pov_grafa):
mer_povezav.append(random.randint(min_cena, max_cena))
sur_podatki.append(mer_povezav)
return(sur_podatki)
utezi = seznam_cen(5, 10, 1, 5)
#print(utezi[2])
#print(utezi)
#print(len(utezi))
#print([sum(x) for x in zip(*utezi)])
#povp_pov = [i * 1/len(utezi) for i in [sum(x) for x in zip(*utezi)]]
#povp_pov
def mnoz(vektor1, vektor2):
matrika = []
for i in range(0,len(vektor1)):
vrstica =[]
for j in range(0, len(vektor2)):
vrstica.append(vektor1[i] * vektor2[j])
matrika.append(vrstica)
return(matrika)
#np.transpose(x)
#numpy.subtract()
#a = np.matrix([[1, 2], [3, 4]])
#b = np.matrix([[2, 2], [2, 2]])
#
#>>> a+b
#matrix([[3, 4],
# [5, 6]])
#np.dot(A,v)
#naračuna kovariančno matriko, ce mu podamo vsa opažanja
def kov_matrika(vse_utezi):
avr_pov = [i * 1/len(vse_utezi) for i in [sum(x) for x in zip(*vse_utezi)]]
zac_mat = numpy.matrix(mnoz(numpy.subtract(vse_utezi[0],avr_pov),numpy.subtract(vse_utezi[0],avr_pov)))
for i in range(1, len(vse_utezi)):
zac_mat += numpy.matrix(mnoz(numpy.subtract(vse_utezi[i],avr_pov),numpy.subtract(vse_utezi[i],avr_pov)))
kon_mat = 1/len(vse_utezi)*zac_mat
return(kon_mat)
#kov_matrika(utezi)
testna_kov_mat = kov_matrika([[1,2,3],[1,2,1]])
print(testna_kov_mat)
numpy.diag(testna_kov_mat)
#mnoz([1,2,3],[1,2,3])
#numpy.dot([1,2,3],numpy.transpose([1,2,3])) #to je skalarni produkt
def graf(st_opazanj, st_vozlisc, st_pov_grafa, min_cena, max_cena):
utezi2 = seznam_cen(st_opazanj, st_pov_grafa, min_cena, max_cena)
graf_seznam = vsa_vozlisca(st_vozlisc, st_pov_grafa)
nov = [] #najprej spremeniva v list
for i in range(0,len(graf_seznam)):
nov.append(list(graf_seznam[i]))
konec = [] #dodava še ceno, v obliki ki jo sprjema digraph
seznam_gra = []
for j in range(0,len(utezi2)):
zadnji = []
for i in range(0,len(nov)):
vmesni = []
vmesni.append(nov[i][0])
vmesni.append(nov[i][1])
vmesni.append(utezi2[j][i])
zadnji.append(tuple(vmesni))
konec.append(zadnji)
gr = DiGraph(zadnji)
seznam_gra.append(gr)
#seznam_gra[0].show(edge_labels = True)
return(seznam_gra, konec)
((graf(5, 100, 20, 1, 5))[0][0]).show(edge_labels = True)
#ggrafi = gen_grafov(cene)
#ggrafi[0].show(edge_labels = True)
#graf_seznam = graphs.RandomGNM(5, 10).edges(labels=False)
#print(g)
#gr = DiGraph(g)
#gr.show(edge_labels=False)
#
#nov = []
#for i in range(0,len(graf_seznam)):
# nov.append(list(graf_seznam[i]))
#print(nov)
#
#konec = []
#seznam_gra = []
#for j in range(0,len(utezi2)):
# zadnji = []
# for i in range(0,len(nov)):
# vmesni = []
# vmesni.append(nov[i][0])
# vmesni.append(nov[i][1])
# vmesni.append(utezi2[j][i])
# zadnji.append(tuple(vmesni))
# konec.append(zadnji)
# gr = DiGraph(zadnji)
# seznam_gra.append(gr)
# gr.show(edge_labels=False)
#print(konec)
#print(seznam_gra)
#Kako seznam spremenis v set, ki bi ga nato lahko podal v DiGraph?
sezam = [1,2,3]
s = set(sezam)
msezam = set(sezam)
print(msezam)
import random
import sage.graphs.graph_plot
from sage.graphs.base.boost_graph import *
import numpy
def gen_vh_pod(st_opazanj, st_pov_grafa):
surovi_podatki = {}
for i in range(0, st_opazanj):
mer_povezav = []
for j in range(0, st_pov_grafa):
mer_povezav.append(random.randint(1,5))
surovi_podatki[i] = mer_povezav
return(surovi_podatki)
#c=gen_vh_pod(3,10)
#print(c)
#M = Matrix([[0, c[0][0], c[0][1], c[0][2], 0],[0, 0, c[0][3], 0, 0],[0, c[0][4], 0, 0, c[0][5]],[c[0][6], 0, c[0][7], 0, c[0][8]],[0, c[0][9], 0, 0, 0]])
#M
cene = gen_vh_pod(10, 10)
print(cene)
def gen_grafov(c):
g_matrike={}
for i in range(0,len(c)):
g_matrike[i] = [[0, c[i][0], c[i][1], c[i][2], 0],[0, 0, c[i][3], 0, 0],[0, c[i][4], 0, 0, c[i][5]],[c[i][6], 0, c[i][7], 0, c[i][8]],[0, c[i][9], 0, 0, 0]]
print(g_matrike)
grafi = {}
for j in range(0, len(g_matrike)):
grafi[j] = DiGraph(Matrix(g_matrike[j]), sparse = True, weighted = True)
#return(g_matrike)
return(grafi)
#ZGLEDI ZA GENERIRANJE GRAFOV
#ggrafi = gen_grafov(cene)
#ggrafi[0].show(edge_labels = True)
#ggrafi[1].show(edge_labels = True)
#ggrafi[5].show(edge_labels = True)
#ggrafi[8].show(edge_labels = True)
#ggraf[8].all_paths(zacetno, koncno, use_multiedges=True, report_edges=False, labels=False)
#print(ggrafi[8].edges())
#ggrafi[5].show(edge_labels = True)
#m = gen_grafov(cene)
#DiGraph(Matrix(m[0]), sparse = True, weighted = True).show(edge_labels = True)
#DiGraph(Matrix(m[1]), sparse = True, weighted = True).show(edge_labels = True)
#DiGraph(Matrix(m[2]), sparse = True, weighted = True).show(edge_labels = True)
#c[0][0] #na tak nacin klicemo znotraj slovarjev
#type(c) vrne dictionary
cene.values()
lists_of_lists = [[1, 2, 3], [4, 5, 6]]
[sum(x) for x in zip(*lists_of_lists)]
# -> [5, 7, 9]
#JAN PROBAVA KAJ SPREMENITI
#nase_st_vozlisc = Grafek.order()
#start = random.randint(1, nase_st_vozlisc)
#end = random.randint(1, nase_st_vozlisc)
#poti = Grafek.all_paths(start, end, use_multiedges=True, report_edges=False, labels=False)
#
#st_vozlisc_grafa = graf.order()
#zacetno = 3
#koncno = 4
#vse_poti = graf.all_paths(zacetno, koncno, use_multiedges=True, report_edges=False, labels=False)
#definicija za delanje vseh vektorjev ki so v seznamu:-to je to kar v resnici rabimo
def vektorcki_x(graf, zacetno, koncno):
vse_poti = graf.all_paths(zacetno, koncno, use_multiedges=True, report_edges=False, labels=False)
vse_povezave = graf.edges()
par = []
for i in range(0,len(vse_povezave)):
par.append(vse_povezave[i][0:2])
seznam_vektorjev = []
for i in range(0, len(vse_poti)):
pot = (vse_poti[i])
prvi_oklepaj1 = list(zip(pot, pot[1:] + pot[:1]))
prvi_oklepaj = prvi_oklepaj1[:-1]
x=[]
for i in range(0, len(par)):
if par[i] in prvi_oklepaj:
x.append(1)
else:
x.append(0)
seznam_vektorjev.append(x)
return(seznam_vektorjev)
#naključna vglišča
def nakljucno_vozlisce(st_vozlisc1):
start = random.randint(0, st_vozlisc1)
end = random.randint(0,st_vozlisc1)
if start == end:
return nakljucno_vozlisce(st_vozlisc1)
else:
return[start, end]
#nakljucno_vozlisce(ggrafi[8])
def vsa_vozlisca(koliko, st_pov1):
vozlisca = []
for i in range(0, st_pov1):
x = nakljucno_vozlisce(koliko)
while x in vozlisca:
x = nakljucno_vozlisce(koliko)
vozlisca.append(x)
return vozlisca
vsa_vozlisca(50,1000)
```
Skalarni produkt
```
cene = gen_vh_pod(10, 10)
def povprecje(cene1):
seznam_cen = list(cene1.values())
povprecne_cene = []
for i in range(0, len(seznam_cen)):
povprecne_cene.append(numpy.average(seznam_cen[i]))
return(povprecne_cene)
print(povprecje(cene))
def vektor_skalarni_produkt(cene1, start, end, graf):
cene_poti = {}
for i in range(0,len(cene1)):
x = vektorcki_x(graf[0], start, end)
vmesni = []
for j in range(0,len(x)):
vmesni.append(numpy.dot(cene[i], x[j]))
cene_poti[i] = vmesni
return
vektor_skalarni_produkt(cene, 2, 4, ggrafi)
```
The average objective value
```
#M3 = Matrix([[0,1,-1],[-1,0,-1/2],[1,1/2,0]])
#M4 = Matrix([[0,1.2,-3],[-1,0,-0.35],[1,0.4,0]])
#G3 = DiGraph(M3,sparse=True,weighted=True)
#G4 = DiGraph(M4, sparse= True, weighted=True)
##g = graphs.RandomGNM(15, 20) # 15 vertices and 20 edges
##show(g)
##g.incidence_matrix()
#M3
#G3.show(edge_labels=True)
#M4
#G4.show(edge_labels=True)
seznam1 = gen_vh_pod(3,10)
print(seznam1)
#G_seznam = [(1,2,seznam1[0][0]),(2,1,seznam1[0][1]),(1,4,seznam1[0][2]),(4,1,seznam1[0][3]),(1,5,seznam1[0][4]),(5,1,seznam1[0][5]),(2,3,seznam1[0][6]),
# (3,4,seznam1[0][7]),(3,5,seznam1[0][8]),(5,3,seznam1[0][9])]
#Grafek = DiGraph(G_seznam, weighted=True)
#Grafek.show(edge_labels=True)
G_seznam = [{1,2,seznam1[0][0]},{2,1,seznam1[0][1]},{1,4,seznam1[0][2]}]
Grafek = DiGraph(G_seznam, weighted=True)
Grafek.show(edge_labels=True)
st_vozlisc = Grafek.order()
#Grafek.incidence_matrix() #izpiše incindenčno matriko grafa
#Grafek.weighted_adjacency_matrix()
#def risanje_grafa(seznam,cene_povezav):
# seznam
# for i in range(0, length(cene_povezav)):
# for i in range(0, length(list(cene_povezav)[i])):
# seznam
#DELUJOČ
seznam1 = gen_vh_pod(5,10)
print(seznam1)
graf_seznam = []
for j in range(0, len(seznam1)):
a = [(1,2,seznam1[j][0]),(2,1,seznam1[j][1]),(1,4,seznam1[j][2]),(4,1,seznam1[j][3]),(1,5,seznam1[j][4]),(5,1,seznam1[j][5]),(2,3,seznam1[j][6]), (3,4,seznam1[j][7]),(3,5,seznam1[j][8]),(5,3,seznam1[j][9])]
graf_seznam.append(a)
Grafek = DiGraph(a, weighted=True)
Grafek.show(edge_labels=True)
st_vozlisc = Grafek.order()
print(graf_seznam)
print(st_vozlisc)
#G_seznam = [(1,2,seznam1[0][0]),(2,1,seznam1[0][1]),(1,4,seznam1[0][2]),(4,1,seznam1[0][3]),(1,5,seznam1[0][4]),(5,1,seznam1[0][5]),(2,3,seznam1[0][6]),
# (3,4,seznam1[0][7]),(3,5,seznam1[0][8]),(5,3,seznam1[0][9])]
#Grafek = DiGraph(G_seznam, weighted=True)
#Grafek.show(edge_labels=True)
#def generiranje_uncertainty_sets(st_poti):
# nakljucna_cela = {}
# for j in range(1, st_poti+1):
# nakljucna = []
# for i in range(0,4): #koliko števil je v množici
# n = random.randint(1,30)
# nakljucna.append(n)
# nakljucna_cela[j] = nakljucna
# return(nakljucna_cela)
#
#generiranje_uncertainty_sets(10)
#
```
Narišimo najprej za majhne grafe. Graf z 7 vozlišči.
```
#seznam = [(1,3,3),(1,5,3),(5,1,4),(4,2,4),(5,2,5),(3,4,6),(3,5,7),(5,4,8),(2,6,9),(4,6,13),(6,7,13)]
#d=DiGraph([[1..7],seznam], weighted=True)
#d.show(edge_labels=True)
#
#short = shortest_paths(d, start = 1, weight_function = None, algorithm=None)
#print(short)
#d.all_paths(start=1, end=7, use_multiedges=True, report_edges=False, labels=False)
```
Naredimo funkcijo, ki nam išče najkrajšo pot v grafu. (Tega za zdaj ne rabimo.)
```
def BFS_SP(graph, start, goal):
explored = []
queue = [[start]]
if start == goal:
print("Same Node")
return
while queue:
path = queue.pop(0)
node = path[-1]
if node not in explored:
neighbours = graph[node]
for neighbour in neighbours:
new_path = list(path)
new_path.append(neighbour)
queue.append(new_path)
if neighbour == goal:
print("Shortest path = ", *new_path)
return(new_path)
explored.append(node)
print("So sorry, but a connecting path doesn't exist.")
return(new_path)
new = BFS_SP(d,1,7)
print(new)
#nase_st_vozlisc = Grafek.order()
#
#def kon_zac(graf):
# par_voz =[]
# st_voz = graf.order()
# start = random.randint(0,st_voz)
# end = random.randint(0,st_voz)
# if end == start:
# kon_zac(graf)
# else:
# start_end(stevilo)
#
#print(start_end(nase_st_vozlisc))
```
Sestavimo vektor x
```
#to je za prvi oklepaj v vseh poteh
#pot = (vse_poti[0])
#print(pot)
#prvi_oklepaj = list(zip(pot, pot[1:] + pot[:1]))
#print(prvi_oklepaj[:-1])
#to nam vzame samo prve dve številke v seznamu od grafa
#par = []
#for i in range(0,len(G_seznam)):
# par.append(G_seznam[i][0:2])
#print(par)
#
#vektor x za prvo pot
#x=[]
#for i in range(0, len(par)):
# if par[i] in prvi_oklepaj:
# x.append(1)
# else:
# x.append(0)
#print(x)
#nase_st_vozlisc = Grafek.order()
#start = random.randint(1, nase_st_vozlisc)
#end = random.randint(1, nase_st_vozlisc)
start = 3
end = 4
poti = Grafek.all_paths(start, end, use_multiedges=True, report_edges=False, labels=False)
#definicija za delanje vseh vektorjev ki so v seznamu:-to je to kar v resnici rabimo
def vektorji_x(vse_poti):
par = []
for i in range(0,len(G_seznam)):
par.append(G_seznam[i][0:2])
print(par)
seznam_vektorjev = []
for i in range(0, len(vse_poti)):
pot = (vse_poti[i])
prvi_oklepaj1 = list(zip(pot, pot[1:] + pot[:1]))
prvi_oklepaj = prvi_oklepaj1[:-1]
x=[]
for i in range(0, len(par)):
if par[i] in prvi_oklepaj:
x.append(1)
else:
x.append(0)
seznam_vektorjev.append(x)
return(seznam_vektorjev)
vektorji_x(poti)
```
tega naprej načeloma ne rabiva (da je gor cela množica):
```
#a = generiranje_uncertainty_sets(10).values()
#print(a)
#print(list(a)[0])
#
#u=DiGraph([[1..7],[(1,3,list(a)[0]),(1,5,list(a)[1]),(4,2,list(a)[2]),(5,2,list(a)[3]),(3,4,list(a)[4]),(3,5,list(a)[5]),(5,4,list(a)[6]),(2,6,list(a)[7]),(4,6,list(a)[8]),(6,7,list(a)[9])]])
#u.show(edge_labels=True)
```
Skalarni produkt
```
a = [0,1,2]
b=[3,4,5]
print(numpy.dot(a,b))
```
| github_jupyter |
```
# the code is for 6-class sub-independent test, no data saved to HD
from __future__ import division, print_function, absolute_import
import tensorflow as tf
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import mne
import sklearn.cross_validation as cross_validation
from mne.io import RawArray
from mne.epochs import concatenate_epochs
from mne import create_info, find_events, Epochs
from mne.channels import read_custom_montage
from sklearn.pipeline import make_pipeline
from sklearn.linear_model import LogisticRegression
from sklearn.cross_validation import cross_val_score, LeaveOneLabelOut
from glob import glob
from mpl_toolkits.axes_grid1 import make_axes_locatable
from scipy.signal import welch
from mne import pick_types
from tensorflow.examples.tutorials.mnist import input_data
import os.path
import scipy.io as sio
##########################
# create MNE readable file
##########################
def creat_mne_raw_object(fname):
"""Create a mne raw instance from csv file"""
# Read EEG file
data = pd.read_csv(fname)
# get chanel names
ch_names = list(data.columns[1:])
# read EEG standard montage from mne
montage = 'standard_1005'
# events file
ev_fname = fname.replace('_data','_events')
# read event file
events = pd.read_csv(ev_fname)
events_names = events.columns[1:]
events_data = np.array(events[events_names]).T
# concatenate event file and data
data = np.concatenate((1e-6*np.array(data[ch_names]).T,events_data))
# define channel type, the first is EEG, the last 6 are stimulations
ch_type = ['eeg']*len(ch_names) + ['stim']*6
# create and populate MNE info structure
ch_names.extend(events_names)
info = create_info(ch_names,sfreq=500.0,ch_types=ch_type,montage = montage)
# info.set_montage(montage)
#info['filename'] = fname
# create raw object
raw = RawArray(data,info,verbose=False)
return raw
####################################################################
# Data preprocessing, band_pass, event related 1.5s + 1.5s featuring
####################################################################
def data_gen(subjects):
X_train=[]
y_train=[]
n = 0
for i in range(len(subjects)):
n+=1
subject = subjects[i]
epochs_tot = []
epochs_rest_tot = []
fnames = glob('../EEG/train/subj%d_series*_data.csv' % (subject))
y = []
for i,fname in enumerate(fnames):
# read data
raw = creat_mne_raw_object(fname)
#raw.plot(block=True)
# pick eeg signal
picks = pick_types(raw.info,eeg=True)
raw.filter(7,35, picks=picks, method='iir', n_jobs=-1, verbose=False)
# Filter data for alpha frequency and beta band
##########
# Events #
##########
###############id = 'Replace'#################
# get event posision corresponding to Replace
events = find_events(raw,stim_channel='Replace', verbose=False)
# epochs signal for 1.5 second before the movement
epochs = Epochs(raw, events, {'after' : 1}, 0.5, 2, proj=False, picks=picks, baseline=None,
preload=True, verbose=False)
epochs_tot.append(epochs)
# epochs signal for 1.5 second after the movement, this correspond to the rest period.
epochs_rest = Epochs(raw, events, {'during' : 1}, -2, -0.5, proj=False, picks=picks, baseline=None,
preload=True, verbose=False)
# Workaround to be able to concatenate epochs
epochs_rest.shift_time(2.5, relative = True)
epochs_rest_tot.append(epochs_rest)
# adding "Replace" lable "5"
y.extend([5]*len(epochs_rest))
#################### id = 'HandStart'####################
# get event posision corresponding to HandStart
events = find_events(raw,stim_channel='HandStart', verbose=False)
# epochs signal for 1.5 second before the movement
epochs = Epochs(raw, events, {'during' : 1}, 0.5, 2, proj=False, picks=picks, baseline=None,
preload=True, verbose=False)
epochs_tot.append(epochs)
# epochs signal for 1.5 second after the movement, this correspond to the rest period.
epochs_rest = Epochs(raw, events, {'before' : 1}, -2, -0.5, proj=False, picks=picks, baseline=None,
preload=True, verbose=False)
# Workaround to be able to concatenate epochs
epochs_rest.shift_time(2.5, relative = True)
epochs_rest_tot.append(epochs_rest)
# adding lable "1" of "HandStart"
y.extend([1]*len(epochs_rest))
#################### id = 'FirstDigitTouch'####################
# get event posision corresponding to FirstDigitTouch
events = find_events(raw,stim_channel='FirstDigitTouch', verbose=False)
epochs = Epochs(raw, events, {'during' : 1}, 0.5, 2, proj=False, picks=picks, baseline=None,
preload=True, verbose=False)
epochs_tot.append(epochs)
epochs_rest = Epochs(raw, events, {'before' : 1}, -2, -0.5, proj=False, picks=picks, baseline=None,
preload=True, verbose=False)
epochs_rest.shift_time(2.5, relative = True)
epochs_rest_tot.append(epochs_rest)
# adding lable "2" of "FirstDigitTouch"
y.extend([2]*len(epochs_rest))
#################### id = 'BothStartLoadPhase'####################
# get event posision corresponding to BothStartLoadPh - truncated to 15 characters
events = find_events(raw,stim_channel='BothStartLoadPh', verbose=False)
epochs = Epochs(raw, events, {'during' : 1}, 0.5, 2, proj=False, picks=picks, baseline=None,
preload=True, verbose=False)
epochs_tot.append(epochs)
epochs_rest = Epochs(raw, events, {'before' : 1}, -2, -0.5, proj=False, picks=picks, baseline=None,
preload=True, verbose=False)
epochs_rest.shift_time(2.5, relative = True)
epochs_rest_tot.append(epochs_rest)
# adding lable "3" of "FirstDigitTouch"
y.extend([3]*len(epochs_rest))
#################### id = 'Liftoff'####################
# get event posision corresponding to Liftoff
events = find_events(raw,stim_channel='LiftOff', verbose=False)
epochs = Epochs(raw, events, {'during' : 1}, 0.5, 2, proj=False, picks=picks, baseline=None,
preload=True, verbose=False)
epochs_tot.append(epochs)
epochs_rest = Epochs(raw, events, {'before' : 1}, -2, -0.5, proj=False, picks=picks, baseline=None,
preload=True, verbose=False)
epochs_rest.shift_time(2.5, relative = True)
epochs_rest_tot.append(epochs_rest)
# adding lable "4" of "FirstDigitTouch"
y.extend([4]*len(epochs_rest))
#################### id = 'BothReleased'####################
# get event posision corresponding to BothReleased
events = find_events(raw,stim_channel='BothReleased', verbose=False)
epochs = Epochs(raw, events, {'during' : 1}, 0.5, 2, proj=False, picks=picks, baseline=None,
preload=True, verbose=False)
epochs_tot.append(epochs)
epochs_rest = Epochs(raw, events, {'before' : 1}, -2, -0.5, proj=False, picks=picks, baseline=None,
preload=True, verbose=False)
epochs_rest.shift_time(2.5, relative = True)
epochs_rest_tot.append(epochs_rest)
# adding lable "6" of "BothReleased"
y.extend([6]*len(epochs_rest))
epochs_during = concatenate_epochs(epochs_tot)
epochs_rests = concatenate_epochs(epochs_rest_tot)
#get data
X_during = epochs_during.get_data()
X_rests = epochs_rests.get_data()
X = np.concatenate((X_during,X_rests),axis=1)
y = np.array(y)
#y = np.array(y)
print ("subject",subject,X.shape)
if n == 1:
X_train = X
y_train = y
else:
X_train = np.append(X_train,X,axis =0)
y_train = np.append(y_train,y,axis =0)
print ("data stack shape:",X_train.shape,y_train.shape)
# generate feature dataset for next process
X_train = np.array(X_train)
y_train = np.array(y_train)
print ('return data shape: ',X_train.shape,y_train.shape)
return (X_train, y_train)
for j in range(10,13):
train_subject = [k for k in range(1,13) if k != j]
test_subject = [j]
train_X,train_y = data_gen(train_subject)
test_X,test_y = data_gen(test_subject)
idx = list(range(len(train_y)))
np.random.shuffle(idx)
train_X = train_X[idx]
train_y = train_y[idx]
# sio.savemat('/home/yaoxiaojian/Desktop/kaggle/EEG/TACR_indenpendent_6-class/T-set_sub'+str(j)+'.mat', {"train_x": train_X, "train_y": train_y, "test_x": test_X, "test_y": test_y})
import tensorflow as tf
from cnn_class import cnn
import sklearn.cross_validation as cross_validation
from sklearn.cross_validation import cross_val_score
import time
from sklearn.metrics import classification_report, roc_auc_score, auc, roc_curve, f1_score
from RnnAttention.attention import attention
from scipy import interp
import tensorflow as tf
# train_X preprocess
X_inputs = np.transpose(train_X, [1, 0, 2])
X_inputs=X_inputs.reshape((X_inputs.shape[0],(X_inputs.shape[1]*X_inputs.shape[2])))
X_inputs = np.transpose(X_inputs, [1, 0])
X_inputs = X_inputs*100000
print ('X_inputs',X_inputs.shape)
# test_X preprocess
X_inputs1 = np.transpose(test_X, [1, 0, 2])
X_inputs1 = X_inputs1.reshape((X_inputs1.shape[0],(X_inputs1.shape[1]*X_inputs1.shape[2])))
X_inputs1 = np.transpose(X_inputs1, [1, 0])
X_inputs1 = X_inputs1*100000
print ('X_inputs1',X_inputs1.shape)
# X_inputs and Y_targets are np.array, while X,Y are tf.tensor class
learning_rate = 0.01
training_epochs = 5
batch_size = 751
display_step = 1
n_input = 64
X = tf.placeholder("float", [None, n_input])
n_hidden_1 = 88
n_hidden_2 = 44
n_hidden_3 = 22
weights = {
'encoder_h1': tf.Variable(tf.truncated_normal([n_input, n_hidden_1], )),
'encoder_h2': tf.Variable(tf.truncated_normal([n_hidden_1, n_hidden_2], )),
'encoder_h3': tf.Variable(tf.truncated_normal([n_hidden_2, n_hidden_3], )),
'decoder_h1': tf.Variable(tf.truncated_normal([n_hidden_3, n_hidden_2], )),
'decoder_h2': tf.Variable(tf.truncated_normal([n_hidden_2, n_hidden_1], )),
'decoder_h3': tf.Variable(tf.truncated_normal([n_hidden_1, n_input], )),
}
biases = {
'encoder_b1': tf.Variable(tf.random_normal([n_hidden_1])),
'encoder_b2': tf.Variable(tf.random_normal([n_hidden_2])),
'encoder_b3': tf.Variable(tf.random_normal([n_hidden_3])),
'decoder_b1': tf.Variable(tf.random_normal([n_hidden_2])),
'decoder_b2': tf.Variable(tf.random_normal([n_hidden_1])),
'decoder_b3': tf.Variable(tf.random_normal([n_input])),
}
def encoder(x):
layer_1 = tf.nn.sigmoid(tf.add(tf.matmul(x, weights['encoder_h1']),
biases['encoder_b1']))
layer_2 = tf.nn.sigmoid(tf.add(tf.matmul(layer_1, weights['encoder_h2']),
biases['encoder_b2']))
layer_3 = tf.add(tf.matmul(layer_2, weights['encoder_h3']),
biases['encoder_b3'])
return layer_3
def decoder(x):
layer_1 = tf.nn.sigmoid(tf.add(tf.matmul(x, weights['decoder_h1']),
biases['decoder_b1']))
layer_2 = tf.nn.sigmoid(tf.add(tf.matmul(layer_1, weights['decoder_h2']),
biases['decoder_b2']))
layer_3 = tf.nn.sigmoid(tf.add(tf.matmul(layer_2, weights['decoder_h3']),
biases['decoder_b3']))
return layer_3
# iteration
encoder_op = encoder(X)
decoder_op = decoder(encoder_op)
y_pred = decoder_op
y_true = X
cost = tf.reduce_mean(tf.pow(y_true - y_pred, 2)) # cost func. MSE
optimizer = tf.train.AdamOptimizer(learning_rate).minimize(cost)
encoder_result = []
with tf.Session() as sess:
init = tf.global_variables_initializer()
sess.run(init)
total_batch = int(X_inputs.shape[0] / batch_size)
for epoch in range(training_epochs):
for i in range(total_batch):
offset = (i * batch_size)
batch_xs = X_inputs[offset:(offset + batch_size), :]
#batch_ys = Y_targets[offset:(offset + batch_size), :]
print ('.',end = '')
_, c = sess.run([optimizer, cost], feed_dict={X: batch_xs})
if epoch % display_step == 0:
print()
print("Epoch:", '%04d' % (epoch + 1), "cost=", "{:.9f}".format(c))
print("Optimization Finished!")
# build encoder_result array as feature vector
train_X=sess.run(encoder_op, feed_dict={X: X_inputs})
train_X=train_X.reshape(int(train_X.shape[0]/751),751,22)
train_X=np.transpose(train_X,[0,2,1])
test_X=sess.run(encoder_op, feed_dict={X: X_inputs1})
test_X=test_X.reshape(int(test_X.shape[0]/751),751,22)
test_X=np.transpose(test_X,[0,2,1])
print (train_X.shape, test_X.shape, train_y.shape,test_y.shape)
display (test_X)
sess.close()
test_y = test_y.ravel()
train_y = train_y.ravel()
display (test_y)
train_y = np.asarray(pd.get_dummies(train_y), dtype = np.int8)
test_y = np.asarray(pd.get_dummies(test_y), dtype = np.int8)
#############
# Set window
#############
window_size = 100
step = 50
n_channel = 22
def windows(data, size, step):
start = 0
while ((start+size) < data.shape[0]):
yield int(start), int(start + size)
start += step
def segment_signal_without_transition(data, window_size, step):
segments = []
for (start, end) in windows(data, window_size, step):
if(len(data[start:end]) == window_size):
segments = segments + [data[start:end]]
return np.array(segments)
def segment_dataset(X, window_size, step):
win_x = []
for i in range(X.shape[0]):
win_x = win_x + [segment_signal_without_transition(X[i], window_size, step)]
win_x = np.array(win_x)
return win_x
train_raw_x = np.transpose(train_X, [0, 2, 1])
test_raw_x = np.transpose(test_X, [0, 2, 1])
train_win_x = segment_dataset(train_raw_x, window_size, step)
print("train_win_x shape: ", train_win_x.shape)
test_win_x = segment_dataset(test_raw_x, window_size, step)
print("test_win_x shape: ", test_win_x.shape)
# [trial, window, channel, time_length]
train_win_x = np.transpose(train_win_x, [0, 1, 3, 2])
print("train_win_x shape: ", train_win_x.shape)
test_win_x = np.transpose(test_win_x, [0, 1, 3, 2])
print("test_win_x shape: ", test_win_x.shape)
# [trial, window, channel, time_length, 1]
train_x = np.expand_dims(train_win_x, axis = 4)
test_x = np.expand_dims(test_win_x, axis = 4)
num_timestep = train_x.shape[1]
###########################################################################
# set model parameters
###########################################################################
# kernel parameter
kernel_height_1st = 22
kernel_width_1st = 45
kernel_stride = 1
conv_channel_num = 40
# pooling parameter
pooling_height_1st = 1
pooling_width_1st = 56
pooling_stride_1st = 10
# full connected parameter
attention_size = 512
n_hidden_state = 64
###########################################################################
# set dataset parameters
###########################################################################
# input channel
input_channel_num = 1
# input height
input_height = train_x.shape[2]
# input width
input_width = train_x.shape[3]
# prediction class
num_labels = 6
###########################################################################
# set training parameters
###########################################################################
# set learning rate
learning_rate = 1e-4
# set maximum traing epochs
training_epochs = 120
# set batch size
batch_size = 10
# set dropout probability
dropout_prob = 0.5
# set train batch number per epoch
batch_num_per_epoch = train_x.shape[0]//batch_size
# instance cnn class
padding = 'VALID'
cnn_2d = cnn(padding=padding)
# input placeholder
X = tf.placeholder(tf.float32, shape=[None, input_height, input_width, input_channel_num], name = 'X')
Y = tf.placeholder(tf.float32, shape=[None, num_labels], name = 'Y')
train_phase = tf.placeholder(tf.bool, name = 'train_phase')
keep_prob = tf.placeholder(tf.float32, name='keep_prob')
# first CNN layer
conv_1 = cnn_2d.apply_conv2d(X, kernel_height_1st, kernel_width_1st, input_channel_num, conv_channel_num, kernel_stride, train_phase)
print("conv 1 shape: ", conv_1.get_shape().as_list())
pool_1 = cnn_2d.apply_max_pooling(conv_1, pooling_height_1st, pooling_width_1st, pooling_stride_1st)
print("pool 1 shape: ", pool_1.get_shape().as_list())
pool1_shape = pool_1.get_shape().as_list()
pool1_flat = tf.reshape(pool_1, [-1, pool1_shape[1]*pool1_shape[2]*pool1_shape[3]])
fc_drop = tf.nn.dropout(pool1_flat, keep_prob)
lstm_in = tf.reshape(fc_drop, [-1, num_timestep, pool1_shape[1]*pool1_shape[2]*pool1_shape[3]])
################
# pipline design
#################
########################## RNN ########################
cells = []
for _ in range(2):
cell = tf.contrib.rnn.BasicLSTMCell(n_hidden_state, forget_bias=1.0, state_is_tuple=True)
cell = tf.contrib.rnn.DropoutWrapper(cell, output_keep_prob=keep_prob)
cells.append(cell)
lstm_cell = tf.contrib.rnn.MultiRNNCell(cells)
init_state = lstm_cell.zero_state(batch_size, dtype=tf.float32)
# output ==> [batch, step, n_hidden_state]
rnn_op, states = tf.nn.dynamic_rnn(lstm_cell, lstm_in, initial_state=init_state, time_major=False)
########################## attention ########################
with tf.name_scope('Attention_layer'):
attention_op, alphas = attention(rnn_op, attention_size, time_major = False, return_alphas=True)
attention_drop = tf.nn.dropout(attention_op, keep_prob)
########################## readout ########################
y_ = cnn_2d.apply_readout(attention_drop, rnn_op.shape[2].value, num_labels)
# probability prediction
y_prob = tf.nn.softmax(y_, name = "y_prob")
# class prediction
y_pred = tf.argmax(y_prob, 1, name = "y_pred")
########################## loss and optimizer ########################
# cross entropy cost function
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=y_, labels=Y), name = 'loss')
update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
with tf.control_dependencies(update_ops):
# set training SGD optimizer
optimizer = tf.train.AdamOptimizer(learning_rate).minimize(cost)
# get correctly predicted object
correct_prediction = tf.equal(tf.argmax(tf.nn.softmax(y_), 1), tf.argmax(Y, 1))
########################## define accuracy ########################
# calculate prediction accuracy
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32), name = 'accuracy')
#############
# train test
#############
def multiclass_roc_auc_score(y_true, y_score):
assert y_true.shape == y_score.shape
fpr = dict()
tpr = dict()
roc_auc = dict()
n_classes = y_true.shape[1]
# compute ROC curve and ROC area for each class
for i in range(n_classes):
fpr[i], tpr[i], _ = roc_curve(y_true[:, i], y_score[:, i])
roc_auc[i] = auc(fpr[i], tpr[i])
# compute micro-average ROC curve and ROC area
fpr["micro"], tpr["micro"], _ = roc_curve(y_true.ravel(), y_score.ravel())
roc_auc["micro"] = auc(fpr["micro"], tpr["micro"])
# compute macro-average ROC curve and ROC area
# First aggregate all false probtive rates
all_fpr = np.unique(np.concatenate([fpr[i] for i in range(n_classes)]))
# Then interpolate all ROC curves at this points
mean_tpr = np.zeros_like(all_fpr)
for i in range(n_classes):
mean_tpr += interp(all_fpr, fpr[i], tpr[i])
# Finally average it and compute AUC
mean_tpr /= n_classes
fpr["macro"] = all_fpr
tpr["macro"] = mean_tpr
roc_auc["macro"] = auc(fpr["macro"], tpr["macro"])
return roc_auc
# run with gpu memory growth
config = tf.ConfigProto()
# config.gpu_options.allow_growth = True
train_acc = []
test_acc = []
best_test_acc = []
train_loss = []
with tf.Session(config=config) as session:
session.run(tf.global_variables_initializer())
best_acc = 0
for epoch in range(training_epochs):
pred_test = np.array([])
true_test = []
prob_test = []
########################## training process ########################
for b in range(batch_num_per_epoch):
offset = (b * batch_size) % (train_y.shape[0] - batch_size)
batch_x = train_x[offset:(offset + batch_size), :, :, :, :]
batch_x = batch_x.reshape([len(batch_x)*num_timestep, n_channel, window_size, 1])
batch_y = train_y[offset:(offset + batch_size), :]
_, c = session.run([optimizer, cost], feed_dict={X: batch_x, Y: batch_y, keep_prob: 1-dropout_prob, train_phase: True})
# calculate train and test accuracy after each training epoch
if(epoch%1 == 0):
train_accuracy = np.zeros(shape=[0], dtype=float)
test_accuracy = np.zeros(shape=[0], dtype=float)
train_l = np.zeros(shape=[0], dtype=float)
test_l = np.zeros(shape=[0], dtype=float)
# calculate train accuracy after each training epoch
for i in range(batch_num_per_epoch):
########################## prepare training data ########################
offset = (i * batch_size) % (train_y.shape[0] - batch_size)
train_batch_x = train_x[offset:(offset + batch_size), :, :, :]
train_batch_x = train_batch_x.reshape([len(train_batch_x)*num_timestep, n_channel, window_size, 1])
train_batch_y = train_y[offset:(offset + batch_size), :]
########################## calculate training results ########################
train_a, train_c = session.run([accuracy, cost], feed_dict={X: train_batch_x, Y: train_batch_y, keep_prob: 1.0, train_phase: False})
train_l = np.append(train_l, train_c)
train_accuracy = np.append(train_accuracy, train_a)
print("("+time.asctime(time.localtime(time.time()))+") Epoch: ", epoch+1, " Training Cost: ", np.mean(train_l), "Training Accuracy: ", np.mean(train_accuracy))
train_acc = train_acc + [np.mean(train_accuracy)]
train_loss = train_loss + [np.mean(train_l)]
# calculate test accuracy after each training epoch
for j in range(batch_num_per_epoch):
########################## prepare test data ########################
offset = (j * batch_size) % (test_y.shape[0] - batch_size)
test_batch_x = test_x[offset:(offset + batch_size), :, :, :]
test_batch_x = test_batch_x.reshape([len(test_batch_x)*num_timestep, n_channel, window_size, 1])
test_batch_y = test_y[offset:(offset + batch_size), :]
########################## calculate test results ########################
test_a, test_c, prob_v, pred_v = session.run([accuracy, cost, y_prob, y_pred], feed_dict={X: test_batch_x, Y: test_batch_y, keep_prob: 1.0, train_phase: False})
test_accuracy = np.append(test_accuracy, test_a)
test_l = np.append(test_l, test_c)
pred_test = np.append(pred_test, pred_v)
true_test.append(test_batch_y)
prob_test.append(prob_v)
if np.mean(test_accuracy) > best_acc :
best_acc = np.mean(test_accuracy)
true_test = np.array(true_test).reshape([-1, num_labels])
prob_test = np.array(prob_test).reshape([-1, num_labels])
auc_roc_test = multiclass_roc_auc_score(y_true=true_test, y_score=prob_test)
f1 = f1_score (y_true=np.argmax(true_test, axis = 1), y_pred=pred_test, average = 'macro')
print("("+time.asctime(time.localtime(time.time()))+") Epoch: ", epoch+1, "Test Cost: ", np.mean(test_l),
"Test Accuracy: ", np.mean(test_accuracy),
"Test f1: ", f1,
"Test AUC: ", auc_roc_test['macro'], "\n")
input()
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/geemap/tree/master/examples/notebooks/geemap_and_earthengine.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/geemap/blob/master/examples/notebooks/geemap_and_earthengine.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/geemap/blob/master/examples/notebooks/geemap_and_earthengine.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as emap
except:
import geemap as emap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
```
Map = emap.Map(center=(40, -100), zoom=4)
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
image = ee.Image('USGS/SRTMGL1_003')
# Set visualization parameters.
vis_params = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
# Print the elevation of Mount Everest.
xy = ee.Geometry.Point([86.9250, 27.9881])
elev = image.sample(xy, 30).first().get('elevation').getInfo()
print('Mount Everest elevation (m):', elev)
# Add Earth Engine layers to Map
Map.addLayer(image, vis_params, 'SRTM DEM', True, 0.5)
Map.addLayer(xy, {'color': 'red'}, 'Mount Everest')
```
## Change map positions
For example, center the map on an Earth Engine object:
```
Map.centerObject(ee_object=xy, zoom=13)
```
Set the map center using coordinates (longitude, latitude)
```
Map.setCenter(lon=-100, lat=40, zoom=4)
```
## Extract information from Earth Engine data based on user inputs
```
import ee
import geemap
from ipyleaflet import *
from ipywidgets import Label
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
Map = geemap.Map(center=(40, -100), zoom=4)
Map.default_style = {'cursor': 'crosshair'}
# Add Earth Engine dataset
image = ee.Image('USGS/SRTMGL1_003')
# Set visualization parameters.
vis_params = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
# Add Earth Eninge layers to Map
Map.addLayer(image, vis_params, 'STRM DEM', True, 0.5)
latlon_label = Label()
elev_label = Label()
display(latlon_label)
display(elev_label)
coordinates = []
markers = []
marker_cluster = MarkerCluster(name="Marker Cluster")
Map.add_layer(marker_cluster)
def handle_interaction(**kwargs):
latlon = kwargs.get('coordinates')
if kwargs.get('type') == 'mousemove':
latlon_label.value = "Coordinates: {}".format(str(latlon))
elif kwargs.get('type') == 'click':
coordinates.append(latlon)
# Map.add_layer(Marker(location=latlon))
markers.append(Marker(location=latlon))
marker_cluster.markers = markers
xy = ee.Geometry.Point(latlon[::-1])
elev = image.sample(xy, 30).first().get('elevation').getInfo()
elev_label.value = "Elevation of {}: {} m".format(latlon, elev)
Map.on_interaction(handle_interaction)
Map
import ee
import geemap
from ipyleaflet import *
from bqplot import pyplot as plt
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
Map = geemap.Map(center=(40, -100), zoom=4)
Map.default_style = {'cursor': 'crosshair'}
# Compute the trend of nighttime lights from DMSP.
# Add a band containing image date as years since 1990.
def createTimeBand(img):
year = img.date().difference(ee.Date('1991-01-01'), 'year')
return ee.Image(year).float().addBands(img)
NTL = ee.ImageCollection('NOAA/DMSP-OLS/NIGHTTIME_LIGHTS') \
.select('stable_lights')
# Fit a linear trend to the nighttime lights collection.
collection = NTL.map(createTimeBand)
fit = collection.reduce(ee.Reducer.linearFit())
image = NTL.toBands()
figure = plt.figure(1, title='Nighttime Light Trend', layout={'max_height': '250px', 'max_width': '400px'})
count = collection.size().getInfo()
start_year = 1992
end_year = 2013
x = range(1, count+1)
coordinates = []
markers = []
marker_cluster = MarkerCluster(name="Marker Cluster")
Map.add_layer(marker_cluster)
def handle_interaction(**kwargs):
latlon = kwargs.get('coordinates')
if kwargs.get('type') == 'click':
coordinates.append(latlon)
markers.append(Marker(location=latlon))
marker_cluster.markers = markers
xy = ee.Geometry.Point(latlon[::-1])
y = image.sample(xy, 500).first().toDictionary().values().getInfo()
plt.clear()
plt.plot(x, y)
# plt.xticks(range(start_year, end_year, 5))
Map.on_interaction(handle_interaction)
# Display a single image
Map.addLayer(ee.Image(collection.select('stable_lights').first()), {'min': 0, 'max': 63}, 'First image')
# Display trend in red/blue, brightness in green.
Map.setCenter(30, 45, 4)
Map.addLayer(fit,
{'min': 0, 'max': [0.18, 20, -0.18], 'bands': ['scale', 'offset', 'scale']},
'stable lights trend')
fig_control = WidgetControl(widget=figure, position='bottomright')
Map.add_control(fig_control)
Map
```
| github_jupyter |
<center>
<img src="https://gitlab.com/ibm/skills-network/courses/placeholder101/-/raw/master/labs/module%201/images/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
</center>
# **Space X Falcon 9 First Stage Landing Prediction**
## Web scraping Falcon 9 and Falcon Heavy Launches Records from Wikipedia
Estimated time needed: **40** minutes
In this lab, you will be performing web scraping to collect Falcon 9 historical launch records from a Wikipedia page titled `List of Falcon 9 and Falcon Heavy launches`
[https://en.wikipedia.org/wiki/List_of_Falcon\_9\_and_Falcon_Heavy_launches](https://en.wikipedia.org/wiki/List_of_Falcon\_9\_and_Falcon_Heavy_launches?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01)

Falcon 9 first stage will land successfully

Several examples of an unsuccessful landing are shown here:

More specifically, the launch records are stored in a HTML table shown below:

## Objectives
Web scrap Falcon 9 launch records with `BeautifulSoup`:
* Extract a Falcon 9 launch records HTML table from Wikipedia
* Parse the table and convert it into a Pandas data frame
First let's import required packages for this lab
```
!pip3 install beautifulsoup4
!pip3 install requests
import sys
import requests
from bs4 import BeautifulSoup
import re
import unicodedata
import pandas as pd
```
and we will provide some helper functions for you to process web scraped HTML table
```
def date_time(table_cells):
"""
This function returns the data and time from the HTML table cell
Input: the element of a table data cell extracts extra row
"""
return [data_time.strip() for data_time in list(table_cells.strings)][0:2]
def booster_version(table_cells):
"""
This function returns the booster version from the HTML table cell
Input: the element of a table data cell extracts extra row
"""
out=''.join([booster_version for i,booster_version in enumerate( table_cells.strings) if i%2==0][0:-1])
return out
def landing_status(table_cells):
"""
This function returns the landing status from the HTML table cell
Input: the element of a table data cell extracts extra row
"""
out=[i for i in table_cells.strings][0]
return out
def get_mass(table_cells):
mass=unicodedata.normalize("NFKD", table_cells.text).strip()
if mass:
mass.find("kg")
new_mass=mass[0:mass.find("kg")+2]
else:
new_mass=0
return new_mass
def extract_column_from_header(row):
"""
This function returns the landing status from the HTML table cell
Input: the element of a table data cell extracts extra row
"""
if (row.br):
row.br.extract()
if row.a:
row.a.extract()
if row.sup:
row.sup.extract()
colunm_name = ' '.join(row.contents)
# Filter the digit and empty names
if not(colunm_name.strip().isdigit()):
colunm_name = colunm_name.strip()
return colunm_name
```
To keep the lab tasks consistent, you will be asked to scrape the data from a snapshot of the `List of Falcon 9 and Falcon Heavy launches` Wikipage updated on
`9th June 2021`
```
static_url = "https://en.wikipedia.org/w/index.php?title=List_of_Falcon_9_and_Falcon_Heavy_launches&oldid=1027686922"
```
Next, request the HTML page from the above URL and get a `response` object
### TASK 1: Request the Falcon9 Launch Wiki page from its URL
First, let's perform an HTTP GET method to request the Falcon9 Launch HTML page, as an HTTP response.
```
# use requests.get() method with the provided static_url
# assign the response to a object
import requests
response = requests.get(static_url)
type(response)
```
Create a `BeautifulSoup` object from the HTML `response`
```
# Use BeautifulSoup() to create a BeautifulSoup object from a response text content
soup = BeautifulSoup(response.content,'html.parser')
```
Print the page title to verify if the `BeautifulSoup` object was created properly
```
# Use soup.title attribute
soup.title
```
### TASK 2: Extract all column/variable names from the HTML table header
Next, we want to collect all relevant column names from the HTML table header
Let's try to find all tables on the wiki page first. If you need to refresh your memory about `BeautifulSoup`, please check the external reference link towards the end of this lab
```
# Use the find_all function in the BeautifulSoup object, with element type `table`
# Assign the result to a list called `html_tables`
html_tables = soup.find_all('table')
```
Starting from the third table is our target table contains the actual launch records.
```
# Let's print the third table and check its content
first_launch_table = html_tables[2]
```
You should able to see the columns names embedded in the table header elements `<th>` as follows:
```
<tr>
<th scope="col">Flight No.
</th>
<th scope="col">Date and<br/>time (<a href="/wiki/Coordinated_Universal_Time" title="Coordinated Universal Time">UTC</a>)
</th>
<th scope="col"><a href="/wiki/List_of_Falcon_9_first-stage_boosters" title="List of Falcon 9 first-stage boosters">Version,<br/>Booster</a> <sup class="reference" id="cite_ref-booster_11-0"><a href="#cite_note-booster-11">[b]</a></sup>
</th>
<th scope="col">Launch site
</th>
<th scope="col">Payload<sup class="reference" id="cite_ref-Dragon_12-0"><a href="#cite_note-Dragon-12">[c]</a></sup>
</th>
<th scope="col">Payload mass
</th>
<th scope="col">Orbit
</th>
<th scope="col">Customer
</th>
<th scope="col">Launch<br/>outcome
</th>
<th scope="col"><a href="/wiki/Falcon_9_first-stage_landing_tests" title="Falcon 9 first-stage landing tests">Booster<br/>landing</a>
</th></tr>
```
Next, we just need to iterate through the `<th>` elements and apply the provided `extract_column_from_header()` to extract column name one by one
```
column_names = []
# Apply find_all() function with `th` element on first_launch_table
# Iterate each th element and apply the provided extract_column_from_header() to get a column name
# Append the Non-empty column name (`if name is not None and len(name) > 0`) into a list called column_names
html_th = first_launch_table.find_all('th')
for th in html_th:
if(th != None and len(th)>0):
column_names.append(extract_column_from_header(th))
```
Check the extracted column names
```
print(column_names)
```
## TASK 3: Create a data frame by parsing the launch HTML tables
We will create an empty dictionary with keys from the extracted column names in the previous task. Later, this dictionary will be converted into a Pandas dataframe
```
launch_dict= dict.fromkeys(column_names)
# Remove an irrelvant column
del launch_dict['Date and time ( )']
# Let's initial the launch_dict with each value to be an empty list
launch_dict['Flight No.'] = []
launch_dict['Launch site'] = []
launch_dict['Payload'] = []
launch_dict['Payload mass'] = []
launch_dict['Orbit'] = []
launch_dict['Customer'] = []
launch_dict['Launch outcome'] = []
# Added some new columns
launch_dict['Version Booster']=[]
launch_dict['Booster landing']=[]
launch_dict['Date']=[]
launch_dict['Time']=[]
```
Next, we just need to fill up the `launch_dict` with launch records extracted from table rows.
Usually, HTML tables in Wiki pages are likely to contain unexpected annotations and other types of noises, such as reference links `B0004.1[8]`, missing values `N/A [e]`, inconsistent formatting, etc.
To simplify the parsing process, we have provided an incomplete code snippet below to help you to fill up the `launch_dict`. Please complete the following code snippet with TODOs or you can choose to write your own logic to parse all launch tables:
```
extracted_row = 0
#Extract each table
for table_number,table in enumerate(soup.find_all('table',"wikitable plainrowheaders collapsible")):
# get table row
for rows in table.find_all("tr"):
#check to see if first table heading is as number corresponding to launch a number
if rows.th:
if rows.th.string:
flight_number=rows.th.string.strip()
flag=flight_number.isdigit()
else:
flag=False
#get table element
row = rows.find_all('td')
#if it is number save cells in a dictonary
if flag:
extracted_row += 1
# Flight Number value
# TODO: Append the flight_number into launch_dict with key `Flight No.`
launch_dict['Flight No.'].append(flight_number)
datatimelist=date_time(row[0])
# Date value
# TODO: Append the date into launch_dict with key `Date`
date = datatimelist[0].strip(',')
launch_dict['Date'].append(date)
# Time value
# TODO: Append the time into launch_dict with key `Time`
time = datatimelist[1]
launch_dict['Time'].append(time)
# Booster version
# TODO: Append the bv into launch_dict with key `Version Booster`
bv=booster_version(row[1])
if not(bv):
bv = row[1].string
launch_dict['Version Booster'].append(bv)
# Launch Site
# TODO: Append the bv into launch_dict with key `Launch Site`
launch_site = row[2].string
launch_dict['Launch site'].append(launch_site)
# Payload
# TODO: Append the payload into launch_dict with key `Payload`
payload = row[3].string
launch_dict['Payload'].append(payload)
# Payload Mass
# TODO: Append the payload_mass into launch_dict with key `Payload mass`
payload_mass = get_mass(row[4])
launch_dict['Payload mass'].append(payload_mass)
# Orbit
# TODO: Append the orbit into launch_dict with key `Orbit`
orbit = row[5].a.string
launch_dict['Orbit'].append(orbit)
#print(orbit)
# Customer
# TODO: Append the customer into launch_dict with key `Customer`
customer = row[6].string
launch_dict['Customer'].append(customer)
# Launch outcome
# TODO: Append the launch_outcome into launch_dict with key `Launch outcome`
launch_outcome = list(row[7].strings)[0]
launch_dict['Launch outcome'].append(launch_outcome)
# Booster landing
# TODO: Append the launch_outcome into launch_dict with key `Booster landing`
booster_landing = landing_status(row[8])
launch_dict['Booster landing'].append(booster_landing)
```
After you have fill in the parsed launch record values into `launch_dict`, you can create a dataframe from it.
```
df=pd.DataFrame(launch_dict)
```
We can now export it to a <b>CSV</b> for the next section, but to make the answers consistent and in case you have difficulties finishing this lab.
Following labs will be using a provided dataset to make each lab independent.
<code>df.to_csv('spacex_web_scraped.csv', index=False)</code>
## Authors
<a href="https://www.linkedin.com/in/yan-luo-96288783/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01">Yan Luo</a>
<a href="https://www.linkedin.com/in/nayefaboutayoun/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01">Nayef Abou Tayoun</a>
## Change Log
| Date (YYYY-MM-DD) | Version | Changed By | Change Description |
| ----------------- | ------- | ---------- | --------------------------- |
| 2021-06-09 | 1.0 | Yan Luo | Tasks updates |
| 2020-11-10 | 1.0 | Nayef | Created the initial version |
Copyright © 2021 IBM Corporation. All rights reserved.
| github_jupyter |
# Homework 05
### Exercise 1 - Terminology
Describe the following terms with your own words:
***boolean array:*** An array consisting of boolean values, that indicate if a condidtion is True or False regarding every element in an array
***shape:*** how many elements a dismension contains
***axis:*** represents the dimension
```
Answer the following questions:
***Which ways are there to select one or more elements from a Numpy array?*** fancy indexing, slicing, numpy functions
***What is the difference between Numpy and Scipy?*** numpy is the base library and scipy is a package building on top of numpy
```
### Exercise 2 - Download data from entsoe-e for Lecture 6
For lecture 6, we need to download data from the Entso-e [transparency platform](https://transparency.entsoe.eu/): Entso-e provides (almost) real-time data on European electricity systems. We will download hourly load data (i.e. electricity demand) for all systems in Europe. First, you need to get a user account at Entsoe-e [here](https://transparency.entsoe.eu/usrm/user/createPublicUser).
We are going to use the S-FTP server of Entso-e. To use S-FTP in Python, you have to install the package pysftp. You can do so here in the notebook by executing the following command (please be aware that this may take some time):
```
!conda install -c conda-forge pysftp --yes
```
Now we are ready to download the data. In principle, you simply have to fill out your account information (by setting ```USER``` and ```PWD```), decide where to put the data locally by assigning a path to a ```DOWNLOAD_DIR``` and run the 4 cells below. If the download directory does not exist, it will be created. The download will take some time, so you may want to run the script overnight.
If the download fails at some point, you can restart it by simply executing the cell again. Files which are already downloaded will not be downloaded again. ***Hint:*** I had problems downloading to a directoy which was on a google drive - so if you run into an error message, which says ```OSError: size mismatch in get!``` you may want to choose a directory which is not on a google drive or possibly a dropbox. Also, this error may occur if your disk is full.
```
import os
import pysftp
# if you want, you can modify this too, per default it will create a folder
# in the parant folder of the homework repository:
DOWNLOAD_DIR = 'F:\\scientific_computing'
CATEGORIES = [
'ActualTotalLoad'
]
# To avoid storing the user credentials in the public Github repository,
# these commands will ask you to enter them interactively:
from getpass import getpass
user = getpass('Username for ENTSO-E API:')
pwd = getpass('Password for ENTSO-E API:')
def download_entsoe_data(user, pwd, category, output_dir, server_uri='sftp-transparency.entsoe.eu'):
"""Download a dataset from ENTSO-E's transparency data sftp server.
Contact ENTSO-E to receive login credentials:
https://transparency.entsoe.eu/usrm/user/createPublicUser
:param user: user name required for connecting with sftp server
:param pwd: password required for connecting with sftp server
:param category: ENTSO-E data category to be downloaded
:param output_dir: directory where downloaded data is saved to, a separate
subdirectory is created for each category.
:param server_uri: URI of ENTSO-E transparency server (default last updated on 2020-05-01)
"""
abspath = os.path.abspath(output_dir)
# check if local_dir exists and create if it doesn't
if not os.path.exists(abspath):
os.mkdir(abspath)
print (f'Successfully created the directory {abspath} and using it for download')
else:
print (f'{abspath} exists and will be used for download')
print("\nCopy this path for other notebooks, e.g. the next lecture or homework:\n"
f"DOWNLOAD_DIR = '{abspath}'\n")
cnopts = pysftp.CnOpts()
cnopts.hostkeys = None
# connect to entsoe server via sFTP
entsoe_dir = f'/TP_export/{category}'
with pysftp.Connection(server_uri, username=user, password=pwd, cnopts=cnopts) as sftp:
sftp.chdir(entsoe_dir)
files_entsoe = sftp.listdir()
to_download = list(files_entsoe)
print(f'In total, {len(to_download)} files are going to be downloaded')
# download files not on disk
for file in to_download:
print(f'Downloading file {file}...')
dest_file = os.path.join(abspath, file)
if not os.path.exists(dest_file):
temp_file = os.path.join(abspath, f'{file}.partial')
sftp.get(f'{entsoe_dir}/{file}', temp_file)
os.rename(temp_file, dest_file)
print(f'{file} downloaded successfully.')
else:
print(f'{file} already present locally, skipping download.')
sftp.close()
print("All downloads completed")
# download data...
for category in CATEGORIES:
download_entsoe_data(user, pwd, category, DOWNLOAD_DIR)
```
**Privacy note:** If you don't want to publish the path to your repository on Github (it may contain your Windows user name for example), clear the output of the cell above before saving the Notebook! (In the menu via Cell -> Current outputs -> Clear.)
### Exercise 3 - Create a diagonal matrix
Create a matrix `m` with shape `(4, 4)` by using `np.zeros()` and set the 4 diagonal elements to `1` by using indexing using `np.arange()`. Do not use more two assign statements in total for this exercise!
Bonus: Find multiple ways to avoid calling `np.arange()` twice and analyze which is the best regarding readability, performance and memory usage!
Note: Normally you would use `np.diag()` to do this. You can also have a look into the code using `np.diag??`, but it's probably easier to write your own implementation (which might be less generic and slower, but way simpler).
```
import numpy as np
import math
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
import urllib
import os.path
from pathlib import Path
```
### Versuch ohne np.arange
```
x = np.zeros((4, 4))
x[0, 0] = 1
x[1, 1] = 1
x[2, 2] = 1
x[3, 3] = 1
x
```
### Versuch mit np.arange
```
z = np.zeros ((4, 4))
z[np.arange(4), np.arange(4)]= 1
z
```
### Exercise 4 - Invasion
Create a canvas using `np.zeros()` of shape `(8, 11)`. Then set the following elements to one using fancy slicing techniques:
- Rows 4 and 5 completely.
- In row 3 all elements except the first one.
- In row 2 all elements except the first two ones.
- The two elements defined by: `row_idcs, column_idcs = [0, 1], [2, 3]`
- In row 6 the elements in column 0 and 2.
- In row 7 all elements except the first three and the last three.
And then afterwards the following elements to zero:
- The three elements defined by: `row_idcs, column_idcs = [3, 5, 7], [3, 1, 5]`
As a last step, set assign the content of the first five columns to the last five columns in reversed order. This can be done by using a `step=-1` and starting with 4, i.e. the first five columns in reversed order are indexed by `canvas[:, 4::-1]`.
Then plot the canvas using `plt.imshow()` with the parameter `cmap='gray'`!
**Hint:** it helps a lot to have all commands in one cell (including the `imshow()` command) and execute the cell often, to check the result.
**Note:** When ever the instruction says "first element" it is something like `x[0]`, because it refers to the first one in the array. If it is column 1 or row 1 it is `x[1]`, because it refers then to the index of the column/row.
**Note:** It is `canvas[row_index, column_index]`, so if you are thinking in x/y coordinates, it is `canvas[y, x]` and the y axis goes downwards.
```
canvas = np.zeros((8,11))
canvas[4:6] = 1
canvas[3,1:] = 1
canvas[2,2:] = 1
canvas[[0,1],[2,3]] = 1
canvas[[6], [0,2]] = 1
canvas[7, 3:-3] = 1
canvas[[3,5,7],[3,1,5]] = 0
x = canvas[:, 4::-1]
canvas[:,6:] = x
print(x)
print(canvas)
plt.imshow(canvas, cmap='gray')
```
### Exercise 5 - Draw a circle
Draw a full circle: first define a resolution e.g. $N=50$. Then define coordinates $x$ and $y$ using `np.linspace()` and pass the resolution as parameter `num=N`. Use `np.meshgrid()` to define a grid `xx` and `yy`. Define a canvas of shape `(N, N)` using `np.zeros()`. Then use the circle formula $x^2 + y^2 < r^2$ to define all circle points on the grid (use $r=2$). Then use the boolean 2D expression to set the inside of the circle to 1. Finally plot the canvas using `imshow()`.
## First and Unfinished Try:
```
N = 50
x = np.linspace(0, 250, num=N)
y = np.linspace(0, 250, num=N)
xx, yy = np.meshgrid(x, y)
#grid = np.array([xx, yy])[:, np.newaxis, :, :]
#grid.shape
a = np.zeros((N,N))
r = 2
is_circle = xx**2 + yy**2 < r**2
#plt.imshow(is_circle)
#plt.imshow(grid)
```
## Second try after feedback:
```
N = 50
x = np.linspace(-5, 5, num=N)
y = np.linspace(-5, 5, num=N)
xx, yy = np.meshgrid(x, y)
a = np.zeros((N,N))
#print(a)
r = 2
is_circle = xx**2 + yy**2 < r**2
#plt.imshow(is_circle)
#print(is_circle)
new_a = 1 * is_circle
#print(new_a)
plt.imshow(new_a)
```
### Exercise 6 - Frequency of shades of gray
Convert the picture `numpy-meme.png` to gray scale and plot a histogram!
**Instructions:** Load the image by using `plt.imread()`. This will return a three dimensional array (width, height and colors) with values between zero and one. Using the formula `gray = red * 0.2125 + green * 0.7154 + blue * 0.0721`, convert the picture to shades of gray. Look at the shape of the image and pick the right axis by looking at the length of the array in this axis! You can first calculate a weighted version of the array by multiplying with a vector of length 3 (and the three weights) and then sum along the right axis. Check the shape of the gray image afterwards and plot it using `plt.imshow()` with the parameter `cmap='gray'`. It should be only two dimensional now. Use `image_gray.flatten()` to get all pixels as one-dimensional vector and pass this to the function `plt.hist()` with the parameter `bins=50` to get 50 bins with different gray values.
```
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
image1=plt.imread("numpy-meme.png")
image1.shape
red = image1[:,:,0]
green = image1[:,:,1]
blue = image1[:,:,2]
gray = red * 0.2125 + green * 0.7154 + blue * 0.0721
plt.imshow(gray, cmap='gray')
image_gray=gray.flatten()
plt.hist(image_gray, bins=50)
matplotlib.rc('figure', figsize=(15, 10))
```
### Exercise 7 - Count colors (optional)
Calculate the number of colors used in the picture `numpy-meme.png` and the percentage of the color space (3 x 8bit, i.e. 256 values per color) used!
**Instructions:** Load the image by using `plt.imread()`. This will return a three dimensional array (width, height and colors) with values between zero and one. Multiplying the array with 255 will restore the original 8bit values (integer values between 0 and 255). After multiplying by 255 use `image = image.astype(int)` to convert the image to integer type. Plot the `image` using `plt.imshow()` to see the image and guess the result. Check the shape of the array. One of the axes is of length three - this is the color axis (red, green and blue). We want to map all colors to unique integers. This can be done by defining `colors = red + green * 256 + blue * 256**2`. This is a unique mapping between the triples `(red, green, blue)` and the integers `color` similar to decimal digits (three values between 0 and 9 e.g. `(3, 5, 1)` can be mapped to a three digit number `3 + 5 * 10 + 1 * 100 = 153`). Then use `np.unique()` to get an array with unique colors (in the mapped form as in `color`). This can be used to determine the number of unique colors in the image. This value can also be used to calculate the percentage of the color space used.
<small>Image source: https://me.me/i/1-import-numpy-1-import-numpy-as-np-there-is-e4a6fb9cf75b413dbb3154794fd3d603</small>
```
plt.imread('numpy-meme.png').shape
```
Inspired by [this exercise](https://github.com/rougier/numpy-100/blob/master/100_Numpy_exercises_with_solutions.md#66-considering-a-wh3-image-of-dtypeubyte-compute-the-number-of-unique-colors-) (MIT licensed, [DOI](https://zenodo.org/badge/latestdoi/10173/rougier/numpy-100))
| github_jupyter |
Copyright (c) 2021, salesforce.com, inc.\
All rights reserved.\
SPDX-License-Identifier: BSD-3-Clause\
For full license text, see the LICENSE file in the repo root or https://opensource.org/licenses/BSD-3-Clause
### Colab
Try this notebook on [Colab](http://colab.research.google.com/github/salesforce/warp-drive/blob/master/tutorials/tutorial-5-training_with_warp_drive.ipynb)!
## ⚠️ PLEASE NOTE:
This notebook runs on a GPU runtime.\
If running on Colab, choose Runtime > Change runtime type from the menu, then select 'GPU' in the dropdown.
# Introduction
In this tutorial, we describe how to
- Use the WarpDrive framework to perform end-to-end training of multi-agent reinforcement learning (RL) agents.
- Visualize the behavior using the trained policies.
In case you haven't familiarized yourself with WarpDrive, please see the other tutorials we have prepared for you
- [WarpDrive basics](https://www.github.com/salesforce/warp-drive/blob/master/tutorials/tutorial-1-warp_drive_basics.ipynb)
- [WarpDrive sampler](https://www.github.com/salesforce/warp-drive/blob/master/tutorials/tutorial-2-warp_drive_sampler.ipynb)
- [WarpDrive reset and log controller](https://www.github.com/salesforce/warp-drive/blob/master/tutorials/tutorial-3-warp_drive_reset_and_log.ipynb)
Please also see our [tutorial](https://www.github.com/salesforce/warp-drive/blob/master/tutorials/tutorial-4-create_custom_environments.ipynb) on creating your own RL environment in CUDA C. Once you have your own environment in CUDA C, this tutorial explains how to integrate it with the WarpDrive framework to perform training.
## Dependencies
You can install the warpdrive package using
- the pip package manager OR
- by cloning the warp_drive package and installing the requirements (we shall use this when running on Colab).
```
import sys
IN_COLAB = 'google.colab' in sys.modules
if IN_COLAB:
! git clone https://github.com/salesforce/warp-drive.git
% cd warp-drive
! pip install -e .
% cd tutorials
else:
! pip install rl_warp_drive
from warp_drive.env_wrapper import EnvWrapper
from warp_drive.training.models.fully_connected import FullyConnected
from example_envs.tag_continuous.tag_continuous import TagContinuous
from utils.generate_rollout_animation import generate_env_rollout_animation
from gym.spaces import Discrete, MultiDiscrete
from IPython.display import HTML
import json
import numpy as np
import torch
```
## Training the tag-continuous environment with WarpDrive
For your convenience, there are end-to-end RL training scripts at `warp_drive/training/example_training_scripts.py`. Currently, it supports training both the discrete and the continuous versions of Tag.
In order to run the training for these environments, we first need to configure the *run config*: the set of environment, training, and model parameters.
The run configs for each of the environments are listed in `warp_drive/training/run_configs`, and a sample set of good configs for the **tag-continuous** environment is shown below.
In this tutorial, we'll use $5$ taggers and $100$ runners in a $20 \times 20$ square grid. The taggers and runners have the same skill level, i.e., the runners can move just as fast as the taggers.
```yaml
# YAML configuration for the tag continuous environment
name: "tag_continuous"
# Environment settings
env:
num_taggers: 5
num_runners: 100
grid_length: 20
episode_length: 500
max_acceleration: 0.1
min_acceleration: -0.1
max_turn: 2.35 # 3*pi/4 radians
min_turn: -2.35 # -3*pi/4 radians
num_acceleration_levels: 20
num_turn_levels: 20
skill_level_runner: 1
skill_level_tagger: 1
seed: 274880
use_full_observation: False
runner_exits_game_after_tagged: True
num_other_agents_observed: 10
tag_reward_for_tagger: 10.0
tag_penalty_for_runner: -10.0
step_penalty_for_tagger: -0.00
step_reward_for_runner: 0.00
edge_hit_penalty: -0.0
end_of_game_reward_for_runner: 1.0
tagging_distance: 0.02
# Trainer settings
trainer:
num_envs: 1 # Number of environment replicas
num_episodes: 1000000000 # Number of episodes to run the training for
train_batch_size: 100 # total batch size used for training per iteration (across all the environments)
algorithm: "A2C" # trainer algorithm
vf_loss_coeff: 1 # loss coefficient for the value function loss
entropy_coeff: 0.05 # coefficient for the entropy component of the loss
clip_grad_norm: True # fla indicating whether to clip the gradient norm or not
max_grad_norm: 0.5 # when clip_grad_norm is True, the clip level
normalize_advantage: False # flag indicating whether to normalize advantage or not
normalize_return: False # flag indicating whether to normalize return or not
# Policy network settings
policy: # list all the policies below
runner:
to_train: True
name: "fully_connected"
gamma: 0.98 # discount rate gamms
lr: 0.005 # learning rate
model:
fc_dims: [256, 256] # dimension(s) of the fully connected layers as a list
model_ckpt_filepath: ""
tagger:
to_train: True
name: "fully_connected"
gamma: 0.98
lr: 0.002
model:
fc_dims: [256, 256]
model_ckpt_filepath: ""
# Checkpoint saving setting
saving:
print_metrics_freq: 100 # How often (in iterations) to print the metrics
save_model_params_freq: 5000 # How often (in iterations) to save the model parameters
basedir: "/tmp" # base folder used for saving
tag: "800runners_5taggers_bs100"
```
Next, we also need to specify a mapping from the policy to agent indices trained using that policy. This needs to be set in `warp_drive/training/example_training_script.py`. As such, we have the tagger and runner policies, and we map those to the corresponding agents, as in
```python
policy_tag_to_agent_id_map = {
"tagger": list(envObj.env.taggers),
"runner": list(envObj.env.runners),
}
```
Note that if you wish to use just a single policy across all the agents, or many other policies, you will need to update the run configuration as well as the policy_to_agent_id_mappping.
For example, for using a shared policy across all agents (say `shared_policy`), for example, you can just use the run configuration as
```python
"policy": {
"shared_policy": {
"to_train": True,
"name": "fully_connected",
"gamma": 0.98,
"lr": 0.002,
"model": {
"num_fc": 2,
"fc_dim": 256,
"model_ckpt_filepath": "",
},
},
},
```
and also set all the agent ids to use this shared policy
```python
policy_tag_to_agent_id_map = {
"shared_policy": np.arange(envObj.env.num_agents),
}
```
**Note: make sure the `policy` keys and the `policy_tag_to_agent_id_map` keys are identical.**
Once the run configuration and the policy to agent id mapping are set, you can invoke training by using
```shell
python warp_drive/training/example_training_script.py --env <ENV-NAME>
```
where `<ENV-NAME>` can be `tag_gridworld` or `tag_continuous` (or any new env that you build). And that's it!
The training script performs the following in order
1. Creates the pertinent environment object (with the `use_cuda` flag set to True).
2. Creates and pushes observtion, action, reward and done placeholder data arrays to the device.
3. Creates the trainer object using the environment object, the run configuration, and policy to agent id mapping.
4. Invokes trainer.train()
## Visualizing the trainer policies
In the run config, there's a `save_model_params_freq` parameter that can be set to frequently keep saving model checkpoints. With the model checkpoints, we can initialize the neural network weights and generate a full episode rollout.
We can find an example run config and the trained tagger and runner policy model weights (after about 20M steps) in the `assets/tag_continuous_training/` folder.
```
# Load the run config.
with open("assets/tag_continuous_training/run_config.json") as f:
run_config = json.load(f)
# Create the environment object.
env_wrapper = EnvWrapper(TagContinuous(**run_config['env']))
```
The taggers (runners) use a shared tagger (runner) policy model. The `policy_tag_to_agent_id_map` describes this mapping.
```
# Define the policy tag to agent id mapping.
policy_tag_to_agent_id_map = {
"tagger": list(env_wrapper.env.taggers),
"runner": list(env_wrapper.env.runners),
}
# Step through the environment.
# The environment(s) store and update the rollout data internally in env.global_state.
def generate_rollout_inplace(env_wrapper, run_config, load_model_weights=False):
assert env_wrapper is not None
assert run_config is not None
obs = env_wrapper.reset_all_envs()
action_space = env_wrapper.env.action_space[0]
# Instantiate the policy models.
policy_models = {}
for policy in policy_tag_to_agent_id_map:
policy_config = run_config["policy"][policy]
if policy_config["name"] == "fully_connected":
policy_models[policy] = FullyConnected(
env=env_wrapper,
model_config=policy_config["model"],
policy=policy,
policy_tag_to_agent_id_map=policy_tag_to_agent_id_map,
)
else:
raise NotImplementedError
if load_model_weights:
print(f"Loading saved weights into the policy models...")
for policy in policy_models:
state_dict_filepath = f"assets/tag_continuous_training/{policy}_after_training.state_dict"
policy_models[policy].load_state_dict(torch.load(state_dict_filepath))
print(f"Loaded ckpt {state_dict_filepath} for {policy} policy model.")
for t in range(env_wrapper.env.episode_length):
stacked_obs = np.stack(obs.values()).astype(np.float32)
# Create dict to collect the actions for all agents.
if isinstance(action_space, Discrete):
actions = {agent_id: 0 for agent_id in range(env_wrapper.env.num_agents)}
elif isinstance(action_space, MultiDiscrete):
actions = {agent_id: [0, 0] for agent_id in range(env_wrapper.env.num_agents)}
else:
raise NotImplementedError
# Sample actions for all agents.
for policy in policy_models:
agent_ids = policy_tag_to_agent_id_map[policy]
probabilities, vals = policy_models[policy](
obs=torch.from_numpy(stacked_obs[agent_ids])
)
if isinstance(action_space, Discrete):
for idx, probs in enumerate(probabilities):
sampled_actions = torch.multinomial(probs, num_samples=1)
for sample_action_idx, action in enumerate(sampled_actions):
actions[agent_ids[sample_action_idx]] = action.numpy()[0]
elif isinstance(action_space, MultiDiscrete):
for idx, probs in enumerate(probabilities):
sampled_actions = torch.multinomial(probs, num_samples=1)
for sample_action_idx, action in enumerate(sampled_actions):
actions[agent_ids[sample_action_idx]][idx] = action.numpy()[0]
else:
raise NotImplementedError
# Execute actions in the environment.
obs, rew, done, info = env_wrapper.step(actions)
if done["__all__"]:
break
```
## Visualize the environment before training
```
generate_rollout_inplace(env_wrapper, run_config)
# Visualize the env at t=0
anim = generate_env_rollout_animation(env_wrapper.env, i_start=1, fps=50, fig_width=6, fig_height=6)
# Now, visualize the entire episode roll-out
HTML(anim.to_html5_video())
```
In the visualization above, the large purple dots represent the taggers, while the smaller blue dots represent the runners. Before training, the runners and taggers move around randomly, and that only results in some runners getting tagged, just by chance.
## Visualize the environment after training (for about 20M steps)
```
generate_rollout_inplace(env_wrapper, run_config, load_model_weights=True)
# Visualize the env at t=0
anim = generate_env_rollout_animation(env_wrapper.env, i_start=1, fps=50, fig_width=6, fig_height=6)
# Now, visualize the entire episode roll-out
HTML(anim.to_html5_video())
```
After training, the runners learn to run away from the taggers, and the taggers learn to chase them; there are some instances where we see that taggers also team up to chase and tag the runners. Overall, about 80% of the runners are caught now.
# Learn More and Explore our Tutorials!
You've now seen the entire end-to-end multi-agent RL pipeline!
For your reference, all our tutorials are here:
- [A simple end-to-end RL training example](https://www.github.com/salesforce/warp-drive/blob/master/tutorials/simple-end-to-end-example.ipynb)
- [WarpDrive basics](https://www.github.com/salesforce/warp-drive/blob/master/tutorials/tutorial-1-warp_drive_basics.ipynb)
- [WarpDrive sampler](https://www.github.com/salesforce/warp-drive/blob/master/tutorials/tutorial-2-warp_drive_sampler.ipynb)
- [WarpDrive reset and log](https://www.github.com/salesforce/warp-drive/blob/master/tutorials/tutorial-3-warp_drive_reset_and_log.ipynb)
- [Creating custom environments](https://www.github.com/salesforce/warp-drive/blob/master/tutorials/tutorial-4-create_custom_environments.ipynb)
- [Training with WarpDrive](https://www.github.com/salesforce/warp-drive/blob/master/tutorials/tutorial-5-training_with_warp_drive.ipynb)
| github_jupyter |
#### Copyright 2017 Google LLC.
```
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Sparsity and L1 Regularization
**Learning Objectives:**
* Calculate the size of a model
* Apply L1 regularization to reduce the size of a model by increasing sparsity
One way to reduce complexity is to use a regularization function that encourages weights to be exactly zero. For linear models such as regression, a zero weight is equivalent to not using the corresponding feature at all. In addition to avoiding overfitting, the resulting model will be more efficient.
L1 regularization is a good way to increase sparsity.
## Setup
Run the cells below to load the data and create feature definitions.
```
from __future__ import print_function
import math
from IPython import display #for displaying multiple tables using the same code block
from matplotlib import cm
from matplotlib import gridspec
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
from sklearn import metrics
import tensorflow as tf
from tensorflow.python.data import Dataset
tf.logging.set_verbosity(tf.logging.ERROR)
pd.options.display.max_rows = 10
pd.options.display.float_format = '{:.1f}'.format
california_housing_dataframe = pd.read_csv("https://dl.google.com/mlcc/mledu-datasets/california_housing_train.csv", sep=",")
#reindex the indices for random draws
california_housing_dataframe = california_housing_dataframe.reindex(
np.random.permutation(california_housing_dataframe.index))
def preprocess_features(california_housing_dataframe):
"""Prepares input features from California housing data set.
Args:
california_housing_dataframe: A Pandas DataFrame expected to contain data
from the California housing data set.
Returns:
A DataFrame that contains the features to be used for the model, including
synthetic features.
"""
selected_features = california_housing_dataframe[
["latitude",
"longitude",
"housing_median_age",
"total_rooms",
"total_bedrooms",
"population",
"households",
"median_income"]]
processed_features = selected_features.copy() #copy() is a pandas data frame method
# Create a synthetic feature.
processed_features["rooms_per_person"] = (
california_housing_dataframe["total_rooms"] /
california_housing_dataframe["population"])
return processed_features
def preprocess_targets(california_housing_dataframe):
"""Prepares target features (i.e., labels) from California housing data set.
Args:
california_housing_dataframe: A Pandas DataFrame expected to contain data
from the California housing data set.
Returns:
A DataFrame that contains the target feature.
"""
output_targets = pd.DataFrame()
# Create a boolean categorical feature representing whether the
# median_house_value is above a set threshold.
output_targets["median_house_value_is_high"] = (
california_housing_dataframe["median_house_value"] > 265000).astype(float)
return output_targets
#for clarity
(california_housing_dataframe["median_house_value"] > 265000).astype(float)
# Choose the first 12000 (out of 17000) examples for training.
training_examples = preprocess_features(california_housing_dataframe.head(12000))
training_targets = preprocess_targets(california_housing_dataframe.head(12000))
# Choose the last 5000 (out of 17000) examples for validation.
validation_examples = preprocess_features(california_housing_dataframe.tail(5000))
validation_targets = preprocess_targets(california_housing_dataframe.tail(5000))
# Double-check that we've done the right thing.
print("Training examples summary:")
display.display(training_examples.describe())
print("Validation examples summary:")
display.display(validation_examples.describe())
print("Training targets summary:")
display.display(training_targets.describe())
print("Validation targets summary:")
display.display(validation_targets.describe())
def my_input_fn(features, targets, batch_size=1, shuffle=True, num_epochs=None):
"""Trains a linear regression model.
Args:
features: pandas DataFrame of features
targets: pandas DataFrame of targets
batch_size: Size of batches to be passed to the model
shuffle: True or False. Whether to shuffle the data.
num_epochs: Number of epochs for which data should be repeated. None = repeat indefinitely
Returns:
Tuple of (features, labels) for next data batch
"""
# Convert pandas data into a dict of np arrays.
features = {key:np.array(value) for key,value in dict(features).items()}
# Construct a dataset, and configure batching/repeating.
ds = Dataset.from_tensor_slices((features,targets)) # warning: 2GB limit
ds = ds.batch(batch_size).repeat(num_epochs)
# Shuffle the data, if specified.
if shuffle:
ds = ds.shuffle(10000)
# Return the next batch of data.
features, labels = ds.make_one_shot_iterator().get_next()
return features, labels
{key:np.array(value) for key,value in dict(training_examples).items()}
def get_quantile_based_buckets(feature_values, num_buckets):
quantiles = feature_values.quantile(
[(i+1.)/(num_buckets + 1.) for i in range(num_buckets)])
return [quantiles[q] for q in quantiles.keys()]
def construct_feature_columns():
"""Construct the TensorFlow Feature Columns.
Returns:
A set of feature columns
"""
bucketized_households = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("households"),
boundaries=get_quantile_based_buckets(training_examples["households"], 10))
bucketized_longitude = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("longitude"),
boundaries=get_quantile_based_buckets(training_examples["longitude"], 50))
bucketized_latitude = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("latitude"),
boundaries=get_quantile_based_buckets(training_examples["latitude"], 50))
bucketized_housing_median_age = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("housing_median_age"),
boundaries=get_quantile_based_buckets(
training_examples["housing_median_age"], 10))
bucketized_total_rooms = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("total_rooms"),
boundaries=get_quantile_based_buckets(training_examples["total_rooms"], 10))
bucketized_total_bedrooms = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("total_bedrooms"),
boundaries=get_quantile_based_buckets(training_examples["total_bedrooms"], 10))
bucketized_population = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("population"),
boundaries=get_quantile_based_buckets(training_examples["population"], 10))
bucketized_median_income = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("median_income"),
boundaries=get_quantile_based_buckets(training_examples["median_income"], 10))
bucketized_rooms_per_person = tf.feature_column.bucketized_column(
tf.feature_column.numeric_column("rooms_per_person"),
boundaries=get_quantile_based_buckets(
training_examples["rooms_per_person"], 10))
long_x_lat = tf.feature_column.crossed_column(
set([bucketized_longitude, bucketized_latitude]), hash_bucket_size=1000)
feature_columns = set([
long_x_lat,
bucketized_longitude,
bucketized_latitude,
bucketized_housing_median_age,
bucketized_total_rooms,
bucketized_total_bedrooms,
bucketized_population,
bucketized_households,
bucketized_median_income,
bucketized_rooms_per_person])
return feature_columns
```
## Calculate the Model Size
To calculate the model size, we simply count the number of parameters that are non-zero. We provide a helper function below to do that. The function uses intimate knowledge of the Estimators API - don't worry about understanding how it works.
```
def model_size(estimator):
variables = estimator.get_variable_names()
size = 0
for variable in variables:
if not any(x in variable
for x in ['global_step',
'centered_bias_weight',
'bias_weight',
'Ftrl']
):
size += np.count_nonzero(estimator.get_variable_value(variable))
return size
```
## Reduce the Model Size
Your team needs to build a highly accurate Logistic Regression model on the *SmartRing*, a ring that is so smart it can sense the demographics of a city block ('median_income', 'avg_rooms', 'households', ..., etc.) and tell you whether the given city block is high cost city block or not.
Since the SmartRing is small, the engineering team has determined that it can only handle a model that has **no more than 600 parameters**. On the other hand, the product management team has determined that the model is not launchable unless the **LogLoss is less than 0.35** on the holdout test set.
Can you use your secret weapon—L1 regularization—to tune the model to satisfy both the size and accuracy constraints?
### Task 1: Find a good regularization coefficient.
**Find an L1 regularization strength parameter which satisfies both constraints — model size is less than 600 and log-loss is less than 0.35 on validation set.**
The following code will help you get started. There are many ways to apply regularization to your model. Here, we chose to do it using `FtrlOptimizer`, which is designed to give better results with L1 regularization than standard gradient descent.
Again, the model will train on the entire data set, so expect it to run slower than normal.
```
def train_linear_classifier_model(
learning_rate,
regularization_strength,
steps,
batch_size,
feature_columns,
training_examples,
training_targets,
validation_examples,
validation_targets):
"""Trains a linear regression model.
In addition to training, this function also prints training progress information,
as well as a plot of the training and validation loss over time.
Args:
learning_rate: A `float`, the learning rate.
regularization_strength: A `float` that indicates the strength of the L1
regularization. A value of `0.0` means no regularization.
steps: A non-zero `int`, the total number of training steps. A training step
consists of a forward and backward pass using a single batch.
feature_columns: A `set` specifying the input feature columns to use.
training_examples: A `DataFrame` containing one or more columns from
`california_housing_dataframe` to use as input features for training.
training_targets: A `DataFrame` containing exactly one column from
`california_housing_dataframe` to use as target for training.
validation_examples: A `DataFrame` containing one or more columns from
`california_housing_dataframe` to use as input features for validation.
validation_targets: A `DataFrame` containing exactly one column from
`california_housing_dataframe` to use as target for validation.
Returns:
A `LinearClassifier` object trained on the training data.
"""
periods = 7
steps_per_period = steps / periods
# Create a linear classifier object.
my_optimizer = tf.train.FtrlOptimizer(learning_rate=learning_rate, l1_regularization_strength=regularization_strength)
my_optimizer = tf.contrib.estimator.clip_gradients_by_norm(my_optimizer, 5.0)
linear_classifier = tf.estimator.LinearClassifier(
feature_columns=feature_columns,
optimizer=my_optimizer
)
# Create input functions.
training_input_fn = lambda: my_input_fn(training_examples,
training_targets["median_house_value_is_high"],
batch_size=batch_size)
predict_training_input_fn = lambda: my_input_fn(training_examples,
training_targets["median_house_value_is_high"],
num_epochs=1,
shuffle=False) #num_epochs=1 ensures data repeats for only 1 cycle
predict_validation_input_fn = lambda: my_input_fn(validation_examples,
validation_targets["median_house_value_is_high"],
num_epochs=1,
shuffle=False) #num_epochs=1 ensures data repeats for only 1 cycle
# Train the model, but do so inside a loop so that we can periodically assess
# loss metrics.
print("Training model...")
print("LogLoss (on validation data):")
training_log_losses = []
validation_log_losses = []
for period in range (0, periods):
# Train the model, starting from the prior state.
linear_classifier.train(
input_fn=training_input_fn,
steps=steps_per_period
)
# Take a break and compute predictions.
training_probabilities = linear_classifier.predict(input_fn=predict_training_input_fn)
training_probabilities = np.array([item['probabilities'] for item in training_probabilities])
validation_probabilities = linear_classifier.predict(input_fn=predict_validation_input_fn)
validation_probabilities = np.array([item['probabilities'] for item in validation_probabilities])
# Compute training and validation loss.
training_log_loss = metrics.log_loss(training_targets, training_probabilities)
validation_log_loss = metrics.log_loss(validation_targets, validation_probabilities)
# Occasionally print the current loss.
print(" period %02d : %0.2f" % (period, validation_log_loss))
# Add the loss metrics from this period to our list.
training_log_losses.append(training_log_loss)
validation_log_losses.append(validation_log_loss)
print("Model training finished.")
# Output a graph of loss metrics over periods.
plt.ylabel("LogLoss")
plt.xlabel("Periods")
plt.title("LogLoss vs. Periods")
plt.tight_layout()
plt.plot(training_log_losses, label="training")
plt.plot(validation_log_losses, label="validation")
plt.legend()
return linear_classifier
linear_classifier = train_linear_classifier_model(
learning_rate=0.1,
# TWEAK THE REGULARIZATION VALUE BELOW
regularization_strength=0.1,
steps=300,
batch_size=100,
feature_columns=construct_feature_columns(),
training_examples=training_examples,
training_targets=training_targets,
validation_examples=validation_examples,
validation_targets=validation_targets)
print("Model size:", model_size(linear_classifier))
linear_classifier = train_linear_classifier_model(
learning_rate=0.1,
# TWEAK THE REGULARIZATION VALUE BELOW
regularization_strength=0.6,
steps=300,
batch_size=100,
feature_columns=construct_feature_columns(),
training_examples=training_examples,
training_targets=training_targets,
validation_examples=validation_examples,
validation_targets=validation_targets)
print("Model size:", model_size(linear_classifier))
variables= linear_classifier.get_variable_names()
size= []
for variable in variables:
if not any(x in variable
for x in ['global_step',
'centered_bias_weight',
'bias_weight',
'Ftrl']
):
size.append(linear_classifier.get_variable_value(variable))
tot= 0
for i in xrange(len(size)):
tot+= len(size[i])
tot
```
| github_jupyter |
```
import tensorflow as tf
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import pickle
import keras
import random
from keras.models import Sequential
from keras.layers.core import Dense, Activation, Dropout
from keras.preprocessing.sequence import pad_sequences
from keras.layers import Dense, Input, LSTM, Embedding, Dropout, Activation, CuDNNGRU, Conv1D, CuDNNLSTM, Flatten
from keras.layers import Lambda, AveragePooling1D, MaxPooling1D, Bidirectional, GlobalMaxPool1D, Concatenate, GlobalAveragePooling1D, GlobalMaxPooling1D,concatenate
from keras.layers import SpatialDropout1D
from keras.models import Sequential
from keras.callbacks import Callback
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from keras.layers import Input, Add, Dense, Activation, ZeroPadding1D, BatchNormalization, Flatten, Conv1D, AveragePooling1D, MaxPooling1D, GlobalMaxPooling1D
from keras.initializers import glorot_uniform
from keras.models import Model, load_model
from keras.optimizers import SGD,Adam
import os
import warnings
from keras import backend as K
warnings.filterwarnings('ignore')
os.environ['PYTHONHASHSEED'] = '0'
np.random.seed(42)
random.seed(12345)
session_conf = tf.ConfigProto(intra_op_parallelism_threads=1, inter_op_parallelism_threads=1)
tf.set_random_seed(1234)
sess = tf.Session(graph=tf.get_default_graph(), config=session_conf)
K.set_session(sess)
from keras.regularizers import l2
def W_init(shape,name=None):
"""Initialize weights as in paper"""
values = np.random.normal(loc=0,scale=1e-2,size=shape)
return K.variable(values,name=name)
#//TODO: figure out how to initialize layer biases in keras.
def b_init(shape,name=None):
"""Initialize bias as in paper"""
values=np.random.normal(loc=0.5,scale=1e-2,size=shape)
return K.variable(values,name=name)
input_shape = (1001, 1)
left_input = Input(input_shape)
right_input = Input(input_shape)
#build convnet to use in each siamese 'leg'
convnet = Sequential()
convnet.add(Conv1D(64,10,activation='relu',input_shape=input_shape,
kernel_initializer=W_init,kernel_regularizer=l2(2e-4)))
convnet.add(MaxPooling1D())
convnet.add(Conv1D(128,7,activation='relu',
kernel_regularizer=l2(2e-4),kernel_initializer=W_init,bias_initializer=b_init))
convnet.add(MaxPooling1D())
convnet.add(Conv1D(128,4,activation='relu',kernel_initializer=W_init,kernel_regularizer=l2(2e-4),bias_initializer=b_init))
convnet.add(MaxPooling1D())
convnet.add(Conv1D(256,4,activation='relu',kernel_initializer=W_init,kernel_regularizer=l2(2e-4),bias_initializer=b_init))
convnet.add(Flatten())
convnet.add(Dense(4096,activation="sigmoid",kernel_regularizer=l2(1e-3),kernel_initializer=W_init,bias_initializer=b_init))
#call the convnet Sequential model on each of the input tensors so params will be shared
encoded_l = convnet(left_input)
encoded_r = convnet(right_input)
#layer to merge two encoded inputs with the l1 distance between them
L1_layer = Lambda(lambda tensors:K.abs(tensors[0] - tensors[1]))
#call this layer on list of two input tensors.
L1_distance = L1_layer([encoded_l, encoded_r])
prediction = Dense(1,activation='sigmoid',bias_initializer=b_init)(L1_distance)
siamese_net = Model(inputs=[left_input,right_input],outputs=prediction)
optimizer = Adam(0.00006)
#//TODO: get layerwise learning rates and momentum annealing scheme described in paperworking
siamese_net.compile(loss="binary_crossentropy",optimizer=optimizer)
siamese_net.count_params()
file_to_read = open('new_data.pickle', 'rb')
#通过pickle的load函数读取data1.pkl中的对象,并赋值给data2
tmp = pickle.load(file_to_read)
y_train = pd.get_dummies(Data.iloc[all_index,:]['label'])
cnn_model = Sequential()
cnn_model.add(Conv1D(6, 5, strides = 1, activation = 'relu', input_shape = (1001, 1)))
cnn_model.add(MaxPooling1D(2, strides = 2))
cnn_model.add(Dropout(0.2))
cnn_model.add(Conv1D(16, 5, strides = 1, activation = 'relu'))
cnn_model.add(MaxPooling1D(2, strides = 2))
cnn_model.add(Dropout(0.2))
cnn_model.add(Conv1D(32, 5, strides = 1, activation = 'relu'))
cnn_model.add(MaxPooling1D(2, strides = 2))
cnn_model.add(Dropout(0.2))
cnn_model.add(Flatten())
cnn_model.add(Dense(120, activation = 'relu'))
cnn_model.add(Dense(84, activation = 'relu'))
cnn_model.add(Dense(186, activation = 'softmax', activity_regularizer = keras.regularizers.l2(0.1)))
cnn_model.compile(loss='categorical_crossentropy', optimizer = 'adam', metrics = ['accuracy'])
cnn_model.fit(X_train.reshape(118880, 1001, 1), y_train.values, batch_size = 64, epochs = 100, verbose = 1)
```
## 搭建SiameseNet模型
基本上是加深版本的Lenet 5
```
input_shape = (2251, 1)
left_input = Input(input_shape)
right_input = Input(input_shape)
#build convnet to use in each siamese 'leg'
# Construct CNN layers
# cnn_model = Sequential()
# cnn_model.add(Conv1D(64, 3, strides = 1, activation = 'relu', input_shape = (2251, 1)))
# cnn_model.add(Conv1D(64, 3, strides = 1, activation = 'relu'))
# cnn_model.add(MaxPooling1D(2, strides = 2))
# cnn_model.add(Dropout(0.2))
# cnn_model.add(Conv1D(128, 3, strides = 1, activation = 'relu'))
# cnn_model.add(Conv1D(128, 3, strides = 1, activation = 'relu'))
# cnn_model.add(MaxPooling1D(2, strides = 2))
# cnn_model.add(Dropout(0.2))
# cnn_model.add(Conv1D(256, 3, strides = 1, activation = 'relu'))
# # cnn_model.add(Conv1D(256, 3, strides = 1, activation = 'relu'))
# cnn_model.add(MaxPooling1D(2, strides = 2))
# cnn_model.add(Dropout(0.2))
# cnn_model.add(Conv1D(512, 3, strides = 1, activation = 'relu'))
# #cnn_model.add(Conv1D(512, 3, strides = 1, activation = 'relu'))
# cnn_model.add(MaxPooling1D(2, strides = 2))
# cnn_model.add(Dropout(0.2))
# cnn_model.add(Flatten())
# cnn_model.add(Dense(120, activation = 'relu'))
# cnn_model.add(Dense(84, activation = 'relu'))
# Lenet 5
cnn_model = Sequential()
cnn_model.add(Conv1D(6, 5, strides = 1, activation = 'relu', input_shape = (2251, 1)))
cnn_model.add(MaxPooling1D(2, strides = 2))
cnn_model.add(Dropout(0.2))
cnn_model.add(Conv1D(16, 5, strides = 1, activation = 'relu'))
cnn_model.add(Conv1D(16, 5, strides = 1, activation = 'relu'))
cnn_model.add(MaxPooling1D(2, strides = 2))
cnn_model.add(Dropout(0.2))
cnn_model.add(Conv1D(32, 5, strides = 1, activation = 'relu'))
cnn_model.add(Conv1D(32, 5, strides = 1, activation = 'relu'))
cnn_model.add(MaxPooling1D(2, strides = 2))
cnn_model.add(Dropout(0.2))
cnn_model.add(Conv1D(64, 5, strides = 1, activation = 'relu'))
cnn_model.add(Conv1D(64, 5, strides = 1, activation = 'relu'))
cnn_model.add(MaxPooling1D(2, strides = 2))
cnn_model.add(Dropout(0.2))
cnn_model.add(Flatten())
cnn_model.add(Dense(120, activation = 'relu'))
cnn_model.add(Dense(84, activation = 'relu'))
cnn_model.add(Dense(186, activation = 'softmax', activity_regularizer = keras.regularizers.l2(0.1)))
#call the convnet Sequential model on each of the input tensors so params will be shared
encoded_l = cnn_model(left_input)
encoded_r = cnn_model(right_input)
#layer to merge two encoded inputs with the l1 distance between them
L1_layer = Lambda(lambda tensors:K.abs(tensors[0] - tensors[1]))
#call this layer on list of two input tensors.
L1_distance = L1_layer([encoded_l, encoded_r])
#prediction = Dense(1,activation='sigmoid',bias_initializer=b_init)(L1_distance)
prediction = Dense(1,activation='sigmoid')(L1_distance)
siamese_net = Model(inputs=[left_input,right_input],outputs=prediction)
optimizer = Adam(0.00016)
#//TODO: get layerwise learning rates and momentum annealing scheme described in paperworking
siamese_net.compile(loss="binary_crossentropy",optimizer=optimizer)
siamese_net.count_params()
cnn_model.summary()
siamese_net.summary()
# file_to_read = open('train_pairs.pickle', 'rb')
# train_pairs = pickle.load(file_to_read)
# file_to_read = open('train_targets.pickle', 'rb')
# train_targets = pickle.load(file_to_read)
file_to_read = open('test_pairs.pickle', 'rb')
test_pairs = pickle.load(file_to_read)
file_to_read = open('test_targets.pickle', 'rb')
test_targets = pickle.load(file_to_read)
# new train data
#打开一个名为data1.pkl的文件,打开方式为二进制读取(参数‘rb’)
file_to_read = open('new_data.pickle', 'rb')
#通过pickle的load函数读取data1.pkl中的对象,并赋值给data2
tmp = pickle.load(file_to_read)
# train_data = tmp
t = np.array(tmp)
t.shape
train_targets, train_pairs = [],[]
pair_1, pair_2 = [], []
merge_label_pair = []
# label 1 pairs
for n in range(186):
locfix = n*13
for i in range(13):
loc = n*13+i
for j in range(i+1, 13):
merge_label_pair.append(train_data[loc][:-1])
merge_label_pair.append(train_data[locfix+j][:-1])
merge_label_pair.append(1)
# merge_label_pair = np.array(merge_label_pair)
# print(merge_label_pair.shape)
# print(merge_label_pair[2])
# merge_label_pair = np.array(merge_label_pair).reshape(186*78,3)
# print(merge_label_pair.shape)
# label 0 pairs
for n in range(186):
locfix = n*13
index = np.arange(2418)
delete_index = list(range(n*13,(n+1)*13))
rand_index = np.random.choice(np.delete(index,delete_index,0),6)
for i in range(13):
loc = n*13+i
for j in range(6):
merge_label_pair.append(train_data[loc][:-1])
merge_label_pair.append(train_data[rand_index[j]][:-1])
merge_label_pair.append(0)
merge_label_pair = np.array(merge_label_pair)
print(merge_label_pair.shape)
merge_label_pair = merge_label_pair.reshape(186*78*2,3)
print(merge_label_pair.shape)
# print(merge_label_pair[1][1])
# a = np.array(merge_label_pair[:][1])
# print(a.shape)
np.random.shuffle(merge_label_pair)
#split
pair_1, pair_2 = [], []
for i in range(29016):
pair_1.append(merge_label_pair[i][0])
pair_2.append(merge_label_pair[i][1])
train_pairs = [pair_1, pair_2]
# print(train_pairs)
train_pairs = np.array(train_pairs)
print(train_pairs.shape)
train_pairs = train_pairs.reshape(2,186*78*2,2251,1)
print(train_pairs.shape)
train_pairs = train_pairs.tolist()
train_targets = merge_label_pair[:, -1]
# print(train_targets[1])
print(train_targets.shape)
train_targets = train_targets.tolist()
# print(train_targets)
file_to_read = open('train_low_None_.pickle', 'rb')
tmp1 = pickle.load(file_to_read)
train_data = tmp1.values
train_targets, train_pairs = [],[]
pair_1, pair_2 = [], []
index = np.arange(800)
# print(index)
rand_index = np.random.choice(index,185)
# print(rand_index)
for i in range(185):
for j in range(185):
pair_1.append(train_data[rand_index[i]][:-1])
pair_2.append(train_data[rand_index[j]][:-1])
train_targets.append(1)
pair_1.append(train_data[rand_index[i]][:-1])
pair_2.append(train_data[rand_index[j]+800*(j+1)][:-1])
train_targets.append(0)
#print(pairs,targets)
train_pairs = [pair_1, pair_2]
train_pairs = np.array(train_pairs)
print(train_pairs.shape)
train_pairs = train_pairs.reshape(2,185*185*2,1001,1)
print(train_pairs)
print(train_pairs.shape)
# print(train_pairs[:][:10][:10][:])
# train_pairs = train_pairs.tolist()
output = open('train_pairs.pickle', 'wb')
pickle.dump(train_pairs,output)
output.close()
output = open('train_targets.pickle', 'wb')
pickle.dump(train_targets,output)
output.close()
print(train_pairs[0][0][-1][:])
print(train_pairs[1][0][-1][:])
print(train_targets[1])
siamese_net.fit(train_pairs, train_targets, batch_size = 64, epochs = 100, verbose = 1)
#cnn_model.fit(X_train.reshape(118880, 1001, 1), y_train.values, batch_size = 64, epochs = 100, verbose = 1)
```
# 创立训练数据
```
import numpy as np
import pickle
#trainfile="/home/suliangbu/work/wanghong/train_low_None_.pickle"
file_to_read = open('train_low_None_.pickle', 'rb')
tmp = pickle.load(file_to_read)
data = tmp.values
featur = np.zeros((186, 800, 1001))
for i in range(186):
featur[i] = data[i * 800:(i+1) * 800, :1001]
def get_batch(batch_size,featur):
"""Create batch of n pairs, half same class, half different class"""
n_examples = 800
n_classes = 186
categories = np.random.choice(n_classes,size=(batch_size,),replace=False)
pairs = [np.zeros((batch_size, 1001, 1)) for i in range(2)]
targets=np.zeros((batch_size))
targets[batch_size//2:] = 1
for i in range(batch_size):
category = categories[i]
idx_1 = np.random.randint(0, n_examples)
pairs[0][i] = featur[category, idx_1].reshape(1001,1)
idx_2 = np.random.randint(0, n_examples)
if i >= batch_size // 2:
category_2 = category
else:
category_2 = (category + np.random.randint(1,n_classes)) % n_classes
pairs[1][i] = featur[category_2,idx_2].reshape(1001,1)
#pairs = np.array(pairs)
#print(pairs.shape)
return pairs, targets
file_to_read = open('T_origin.pickle', 'rb')
tmp = pickle.load(file_to_read)
test_data = tmp.values
print(test_data[:,-1])
print(test_data.shape)
test_data = np.load('mix_testdata_and_label.npy')
test_targets, test_pairs = [], []
pair_1,pair_2 = [], []
for i in range(14):
for j in range(i + 1, 14):
if test_data[i][-1] == test_data[j][-1]:
pair_1.append(test_data[i][:-1])
pair_2.append(test_data[j][:-1])
#test_pairs.append([test_data[i][:-1], test_data[j][:-1]])
test_targets.append(1)
else:
#test_pairs.append([test_data[i][:-1], test_data[j][:-1]])
pair_1.append(test_data[i][:-1])
pair_2.append(test_data[j][:-1])
test_targets.append(0)
test_pairs = [pair_1,pair_2]
test_pairs = np.array(test_pairs).reshape(2,91,2251,1)
test_pairs = test_pairs.tolist()
output = open('test_pairs.pickle', 'wb')
pickle.dump(test_pairs,output)
output.close()
output1 = open('test_targets.pickle', 'wb')
pickle.dump(test_targets,output1)
output1.close()
```
# 创立测试数据
```
file_to_read = open('test__None_.pickle', 'rb')
tmp = pickle.load(file_to_read)
print(tmp)
test_data = tmp.values
test_data
test_targets, test_pairs = [], []
pair_1,pair_2 = [], []
for i in range(14):
for j in range(i + 1, 14):
if test_data[i][-1] == test_data[j][-1]:
pair_1.append(test_data[i][:-1])
pair_2.append(test_data[j][:-1])
#test_pairs.append([test_data[i][:-1], test_data[j][:-1]])
test_targets.append(1)
else:
#test_pairs.append([test_data[i][:-1], test_data[j][:-1]])
pair_1.append(test_data[i][:-1])
pair_2.append(test_data[j][:-1])
test_targets.append(0)
test_pairs = [pair_1,pair_2]
# test_pairs = np.array(test_pairs)
# print(test_pairs.shape)
test_pairs = np.array(test_pairs).reshape(2,91,1002,1)
test_pairs = test_pairs.tolist()
#test_targets = np.array(test_targets)
#test_pairs.shape
#print(pairs,targets)
print(test_pairs[0][3][-1][:])
print(test_pairs[1][3][-1][:])
print(test_targets[3])
```
# 训练
```
batch_size = 32
loss_every = 500
for i in range(1,50000):
(inputs,targets)=get_batch(batch_size,featur)
loss=siamese_net.train_on_batch(inputs,targets)
#print(loss)
if i % loss_every == 0:
print("iteration {}, training loss: {:.2f},".format(i,loss))
probs = siamese_net.predict(test_pairs)
n_correct = 0
print(probs)
for i in range(len(probs)):
if probs[i] <= 0.5:
probs[i] = 0
else:
probs[i] = 1
print(probs)
for i in range(len(probs)):
if probs[i] == test_targets[i]:
n_correct += 1
percent_correct = (100.0*n_correct / 91)
print(percent_correct)
# if np.argmax(probs) == np.argmax(test_targets):
# n_correct+=1
# percent_correct = (100.0*n_correct / 91)
#print(percent_correct)
#print(len(probs))
print(percent_correct)
# print(probs)
for i in range(91):
if(probs[i]==1):
print(i)
for i in range(14):
print(test_data[i][-1])
for i in range(91):
if(test_targets[i]==1):
print(i)
# import h5py
# model = siamese_net
# model.save('model_12_18.h5')
# file_to_read = open('new_test_pairs.pickle', 'rb')
# test_pairs = pickle.load(file_to_read)
# file_to_read = open('new_test_targets.pickle', 'rb')
# test_targets = pickle.load(file_to_read)
from keras.models import load_model
siamese_net= load_model('model_12_18.h5')
print(test_pairs)
test_pairs = test_pairs.tolist()
print(test_pairs)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib
from matplotlib import pylab as plt
from scipy.stats import linregress
df = pd.read_csv('../data/News_pro.csv')
# max(df['Facebook_pro'].values)
df_preprocessed = df.drop(columns=['IDLink','Topic','Facebook','GooglePlus','LinkedIn'])
corrmat = df_preprocessed.corr()
all_cols = corrmat.sort_values('BestPlat',ascending=False)['BestPlat'].index
cols = all_cols # negatively correlated features
cm = corrmat.loc[cols,cols]
plt.figure(figsize=(10,10))
plt.matshow(cm,vmin=-1,vmax=1,cmap='seismic',fignum=0)
plt.colorbar(label='corr. coeff.')
plt.xticks(np.arange(cm.shape[0]),list(cols),rotation=90)
plt.yticks(np.arange(cm.shape[0]),list(cols))
plt.tight_layout()
# plt.savefig('figures/corr_coeff_dummies.png',dpi=300)
plt.show()
label = 'BestPlat'
topic = ['x0_economy','x0_microsoft','x0_obama','x0_palestine']
# i = 1
for x in topic:
count_matrix = df.groupby([x, label]).size().unstack()
count_matrix_norm = count_matrix.div(count_matrix.sum(axis=1),axis=0)
count_matrix_norm.plot(kind='bar', stacked=True, figsize=(10,5))
# plt.subplot(2,2,i)
plt.xlabel(x,fontsize=18)
plt.ylabel('Best Platform to post news',fontsize=16)
plt.title("The best platform to post news of a specific topic",fontsize=16)
plt.legend(['Not post','Facebook','GooglePlus','LinkedIn','Indifferent','Others'],loc='best')
plt.show()
# i += 1
labels = ['Not post','Facebook','GooglePlus','LinkedIn','Indifferent','Others']
share = [0.061616, 0.576164, 0.024636, 0.129828, 0.159536, 0.048220]
explode = [0.1, 0, 0, 0, 0, 0.1]
plt.figure(figsize=(10,8))
plt.axes(aspect=1)
plt.pie(share, explode = explode,
labels = labels, autopct = '%3.1f%%',
startangle = 180, textprops={'fontsize': 14})
plt.title('the percentage of BestPlat',fontsize = 16)
plt.show()
labels = ['','Facebook','GooglePlus','LinkedIn','Indifferent','']
share = [0,0.647257, 0.027675, 0.145847, 0.179221,0]
explode = [0, 0, 0, 0, 0, 0]
plt.figure(figsize=(10,8))
plt.axes(aspect=1)
plt.pie(share, explode = explode,
labels = labels, autopct = '%3.1f%%',
startangle = 180, textprops={'fontsize': 14})
plt.title('the percentage of BestPlat',fontsize = 16)
plt.show()
label = 'Topic'
df[['Facebook_pro',label]].boxplot(by=label,figsize=(10,5))
plt.xlabel(label)
plt.ylabel('Facebook')
plt.show()
label = 'Topic'
df[['GooglePlus_pro',label]].boxplot(by=label,figsize=(10,5))
plt.xlabel(label)
plt.ylabel('GooglePlus')
plt.show()
label = 'Topic'
df[['LinkedIn_pro',label]].boxplot(by=label,figsize=(10,5))
plt.xlabel(label)
plt.ylabel('LinkedIn')
plt.show()
h = []
x = df['Facebook_pro']
y = df['x0_economy']
n = 0
m = 0
for i in range(len(y)):
if y[i] == 1:
h.append(x[i])
n += 1
if 0 < x[i] < 0.0003:
m += 1
plt.hist(h,range=(0,0.0003), alpha=0.7, rwidth=0.85)
plt.xlabel('Popularity on Facebook (preprocessed)')
plt.ylabel('frequency')
plt.title("Histogram for popularity on Facebook")
str(m/n*100)
plt.scatter(x = df['PublishDate'], y = df['Facebook_pro'], alpha=0.5)
plt.title('publish date - popularity on Facebook')
plt.xlabel('PublishDate (unix time)')
plt.ylabel('popularity on Facebook')
plt.xlim(0.95, 1.0)
plt.show()
plt.figure(figsize=(8,6))
X = df['PublishDate']
y = df['Facebook_pro']
slope, intercept, r_value, p_value, std_err = linregress(X,y)
plt.plot(X, slope*X+intercept)
a = p_value
print(r_value)
X = df['PublishDate']
y = df['GooglePlus_pro']
slope, intercept, r_value, p_value, std_err = linregress(X,y)
plt.plot(X, slope*X+intercept)
b = p_value
print(r_value)
X = df['PublishDate']
y = df['LinkedIn_pro']
slope, intercept, r_value, p_value, std_err = linregress(X,y)
plt.plot(X, slope*X+intercept)
c = p_value
print(r_value)
plt.legend(['Facebook','GooglePlus','LinkedIn'],loc="best" )
plt.xlabel('Publish time (Unix time)')
plt.ylabel('Popularity')
plt.title("Simple linear regression between publish time and popularity")
plt.legend(['Facebook: p-value = '+str(a),'GooglePlus: p-value = '+str(b),'LinkedIn: p-value = '+str(c)],loc='best')
0.048220
```
| github_jupyter |
# Functionally align representations across neural networks
<a href="https://colab.research.google.com/github/qihongl/demo-nnalign/blob/master/demo-nnalign.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
<br>
<br>
This is a tutorial based on <a href="http://arxiv.org/abs/1811.11684">Lu et al. 2018</a>.
Here's the <a href="https://github.com/qihongl/nnsrm-neurips18">repo</a> for that project,
which contains the code for the experiments/simulations documented in the paper.
<br>
This notebook is hosted <a href="https://github.com/qihongl/demo-nnalign">here</a>.
Here's a one sentence summary of the result:
- different neural networks with the same learning experience acquire representations of the same "shape"
The goal of this script is to demonstrate the point above, by...
- training two networks on the same task
- then show that i) they have the same "shape"; ii) they are mis-aligned
- align their representations
```
# get brainiak
!pip install pip==9.0.1
!pip install git+https://github.com/brainiak/brainiak
!pip install keract
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from brainiak.funcalign.srm import SRM
from sklearn.preprocessing import StandardScaler
from sklearn.datasets import make_blobs, make_classification
from keract import get_activations
from keras.models import Sequential
from keras.layers import Dense
sns.set(style='white', context='talk', palette='colorblind')
np.random.seed(0)
```
## 0. Make some simulated data
First of all, we need generate a learning problem to train some neural networks. In this notebook, we will use a "noisy XOR task". The figure below shows the training set. The test set is independently generated.
```
def make_xor_data(n_samples=200):
center_locs = np.array([[1,1],[-1,-1],[1,-1],[-1,1]])
cluster_std = .4
n_features = 2
n_classes = 2
# gen pts
coords, cluster_ids = make_blobs(
n_features=n_features,
n_samples=n_samples,
shuffle=False,
cluster_std=cluster_std,
centers=center_locs
)
points_per_class = n_samples // n_classes
class_labels = np.repeat([0,1], points_per_class)
return coords, class_labels
# generate some XOR data
n_examples = 200
x_train, y_train = make_xor_data(n_examples)
x_test, y_test = make_xor_data(n_examples)
# plot the data
cur_palette = sns.color_palette(n_colors=2)
f, ax = plt.subplots(1,1, figsize=(6,5))
for i, y_val in enumerate(np.unique(y_train)):
ax.scatter(
x_train[y_val == y_train,0],x_train[y_val == y_train,1],
color=cur_palette[i],
)
ax.set_xlabel('x1')
ax.set_ylabel('x2')
ax.set_title('the training data')
sns.despine()
```
## 1. Train some neural networks on a common training set
```
# help funcs
hidden_layer_name = 'hidden'
def get_net(n_hidden):
"""define a simple neural network with some hidden units"""
model = Sequential()
model.add(Dense(n_hidden, input_dim=2, activation='tanh',name=hidden_layer_name))
model.add(Dense(1, activation='sigmoid',name='output'))
# Compile model
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
return model
def get_hidden_act(model, data):
"""get neural network activity"""
acts = get_activations(model, data)
for layer_name in acts.keys():
if hidden_layer_name in layer_name:
return acts[layer_name]
# model params
n_hidden = 50
# training params
batch_size = 64
shuffle=True
n_nets = 2
n_epochs = 500
# train several models
models = []
records = []
for i in range(n_nets):
model_i = get_net(n_hidden)
record_i = model_i.fit(
x_train, y_train, epochs=n_epochs,
validation_data = (x_test,y_test),
batch_size=batch_size, shuffle=shuffle, verbose=0
)
models.append(model_i)
records.append(record_i)
```
Here's the classification accuracy for both the training set and the test set, just to confirm that these network learned the task reasonably.
```
# plot the learning curves
f, axes = plt.subplots(1,n_nets, figsize=(13,4),sharey=True)
for i,ax in enumerate(axes):
ax.plot(records[i].history['accuracy'],label='train')
ax.plot(records[i].history['val_accuracy'],label='test')
ax.set_title(f'Learning curve, network {i+1}')
ax.set_xlabel('Epochs')
ax.set_ylabel('Accuracy')
ax.legend(frameon=False, bbox_to_anchor=(.7, .6), loc=2, borderaxespad=0)
sns.despine()
f.tight_layout()
```
## 2. Some observations
#### Observation 1: "representational similarity matrix (RSM)" are similar across networks
The following figure shows within-subject RSM, which compares the similarity between ...
- the evoked response of stimulus i in one network
- the evoked response of stimulus j in **the same** network
The result shows that the two networks learn basically the same "representational similarity structure". That is, the relation between stimulus i and stimulus j is similar across the two networks. We call this the shared representational structure across these two networks.
```
# test the network and get its hidden layer activity
hidden_act_mats = [get_hidden_act(model, x_test) for model in models]
within_subject_RSMs = [np.corrcoef(h_i) for h_i in hidden_act_mats]
f,axes = plt.subplots(1, 2, figsize=(10,5))
for i in range(n_nets):
axes[i].imshow(within_subject_RSMs[i], cmap='viridis')
axes[i].set_xlabel('stimuli id')
axes[i].set_ylabel('stimuli id')
axes[i].set_title(f'Within network RSM, network {i+1}')
f.tight_layout()
```
#### Observation 2: however, the same stimulus evokes different response patterns across the two networks
The following figure shows the intersubject RSM in their native spaces, which compares ...
- the evoked hidden activity pattern of stimulus i from one network
- the evoked hidden activity pattern of stimulus j from **another** network
You can see that the following "intersubject" RSM doesn't reflect the representational similarity structure that's shared across the two networks.
```
inter_subject_RSM_native = np.corrcoef(
hidden_act_mats[0],hidden_act_mats[1]
)[:n_examples,n_examples:]
f, ax = plt.subplots(1, 1, figsize=(6,5))
ax.imshow(inter_subject_RSM_native, cmap='viridis')
ax.set_xlabel('stimuli id')
ax.set_ylabel('stimuli id')
ax.set_title('intersubject RSM, native spaces')
f.tight_layout()
```
**Question** So what's similar across these two networks? How do these two networks encode the same similarity structure with different patterns of neural activity?
**One answer:** We observed that the main reason is that they are misaligned. That is to say, their representational structures really have the same "shape", but oriented differently. Empirically, we show that a rigid-body-transformation (i.e. an orthogonal matrix) is usually enough to align them.
In the two subject case, we can solve the optimal orthogonal transformation by solving the procrustes problem. However, in the more general case of aligning n networks, we can use the shared response model.
## 3. Align representations across networks with the shared response model (SRM)
The following code blocks provide a minimal example of the SRM alignment pipeline.
*In practice, the number of components for SRM can be tuned like how you tune PCA (e.g. measure variance explained).
```
# step 1
# get neural network activity matrices ...
h_act_train = [get_hidden_act(model, x_train).T for model in models]
h_act_test = [get_hidden_act(model, x_test).T for model in models]
# step 2: normalize the data
for i in range(n_nets):
sscalar = StandardScaler()
sscalar.fit(h_act_train[i])
h_act_train[i] = sscalar.transform(h_act_train[i])
h_act_test[i] = sscalar.transform(h_act_test[i])
# step 3: fit SRM
n_components = n_hidden
srm = SRM(features=n_components)
# train SRM on the training set
h_act_train_shared = srm.fit_transform(h_act_train)
# use the trained SRM to transform the test set activity
h_act_test_shared = srm.transform(h_act_test)
```
Once aligned, the intersubject RSM is become similar to the within-subject RSM.
```
# compute inter-subject RSM in for the transformed activities (in the shared space)
inter_subject_RSM_shared = np.corrcoef(
h_act_test_shared[0].T,h_act_test_shared[1].T
)[:n_examples,n_examples:]
f, ax = plt.subplots(1, 1, figsize=(6,5))
ax.imshow(inter_subject_RSM_shared, cmap='viridis')
ax.set_xlabel('stimuli id')
ax.set_ylabel('stimuli id')
ax.set_title('intersubject RSM, shared space')
f.tight_layout()
```
We can put the three matrices ...
- intersubject RSM in the shared space
- averaged within-subject RSM
- intersubject RSM in the native space
... side by side. This is similar to Fig 2B in the paper.
Again the point is that, **once aligned in some common space, the intersubject RSM is similar to the within-subject RSM**. In other words, although both stimulus i and stimulus j evoke different responses across these two networks, the relation between stimulus i and stimulus j is represented similarity (across these two networks).
```
# put the 3 matrices side by side
titles = [
'intersubject RSM\n shared space, aligned',
'within-subject RSM\n native spaces',
'intersubject RSM\n native spaces, misaligned'
]
mats = [
inter_subject_RSM_shared,
np.mean(within_subject_RSMs,axis=0),
inter_subject_RSM_native
]
f, axes = plt.subplots(1, 3, figsize=(13,5))
for i,ax in enumerate(axes):
ax.imshow(mats[i], cmap='viridis')
ax.set_title(titles[i])
ax.set_xlabel('stimuli id')
ax.set_ylabel('stimuli id')
f.tight_layout()
```
### Thank you very much to your time!
| github_jupyter |
```
# Ensure the scenepic library will auto reload
%load_ext autoreload
# Imports
import json
import math
import os
import numpy as np
import scenepic as sp
%autoreload
# Seed random number generator for consistency
np.random.seed(0)
ASSET_DIR = os.path.join("..", "ci", "assets")
def asset_path(filename):
return os.path.join(ASSET_DIR, filename)
```
# ScenePic Python Tutorials
These tutorials provide practical examples that highlight most of the functionality supported by ScenePic. While by no means exhaustive, they should give you a solid start towards building useful and insightful 3D visualizations of your own. If there is something you feel is missing from this tutorial, or if there is something you would like to contribute, please contact the maintainers via GitHub Issues.
```
# Tutorial 1 - Scene and Canvas basics
# Create a Scene, the top level container in ScenePic
scene = sp.Scene()
# A Scene can contain many Canvases
# For correct operation, you should create these using scene1.create_canvas() (rather than constructing directly using sp.Canvas(...))
canvas_1 = scene.create_canvas_3d(width = 300, height = 300)
canvas_2 = scene.create_canvas_3d(width = 100, height = 300)
# ScenePic has told Jupyter how to display scene objects
scene
# Tutorial 2 - Meshes and Frames
# Create a scene
scene = sp.Scene()
# A Mesh is a vertex/triangle/line buffer with convenience methods
# Meshes "belong to" the Scene, so should be created using create_mesh()
# Meshes can be re-used across multiple frames/canvases
my_first_mesh = scene.create_mesh(shared_color = sp.Color(1.0, 0.0, 1.0)) # If shared_color is not provided, you can use per-vertex coloring
my_first_mesh.add_cube(transform = sp.Transforms.Scale(0.1)) # Adds a unit cube centered at the origin
my_first_mesh.add_cube(transform = np.dot(sp.Transforms.Translate([-1.0, 1.0, -1.0]), sp.Transforms.Scale(0.5)))
my_first_mesh.add_sphere(transform = sp.Transforms.Translate([1.0, 1.0, 1.0]))
# A Canvas is a 3D rendering panel
canvas = scene.create_canvas_3d(width = 300, height = 300)
# Create an animation with multiple Frames
# A Frame references a set of Meshes
# Frames are created from the Canvas not the Scene
for i in range(10):
frame = canvas.create_frame()
frame.add_mesh(my_first_mesh, transform = sp.Transforms.Translate([i / 10.0, 0.0, 0.0])) # An arbitrary rigid transform can optionally be specified.
mesh2 = scene.create_mesh(shared_color = sp.Color(1.0,0.0,0.0),camera_space=True)
mesh2.add_cube(transform = np.dot(sp.Transforms.Translate([0.0, 0.0, -5.0]), sp.Transforms.Scale(0.5)))
frame.add_mesh(mesh2)
label = scene.create_label(text = "Hi", color = sp.Colors.White, size_in_pixels = 80, offset_distance = 0.6, camera_space = True)
frame.add_label(label = label, position = [0.0, 0.0, -5.0])
# Display the Scene in Jupyter
scene
# Tutorial 3 - Point clouds 1
# Create a scene
scene = sp.Scene()
# Create a mesh that we'll turn in to a point-cloud using enable_instancing()
mesh = scene.create_mesh(shared_color = sp.Color(0,1,0))
mesh.add_cube() # Unit diameter cube that will act as primitive
mesh.apply_transform(sp.Transforms.Scale(0.01)) # Scale the primitive
mesh.enable_instancing(positions = 2 * np.random.rand(10000, 3) - 1) # Cause the mesh to be replicated across many instances with the provided translations. You can optionally also provide per-instance colors and quaternion rotations.
# Create Canvas and Frame, and add Mesh to Frame
canvas = scene.create_canvas_3d(width = 300, height = 300, shading=sp.Shading(bg_color=sp.Colors.White))
frame = canvas.create_frame()
frame.add_mesh(mesh)
scene
# Tutorial 4 - Points clouds 2
# Note that the point cloud primitive can be arbitrarily complex.
# The primitive geometry will only be stored once for efficiency.
# Some parameters
disc_thickness = 0.2
normal_length = 1.5
point_size = 0.1
# A helper Mesh which we won't actually use for rendering - just to find the points and normals on a sphere to be used in mesh2 below
# NB this is created using the sp.Mesh() constructor directly so it doesn't get added automatically to the Scene
sphere_mesh = sp.Mesh()
sphere_mesh.add_sphere(transform = sp.Transforms.Scale(2.0), color = sp.Color(1.0, 0.0, 0.0))
N = sphere_mesh.count_vertices()
points = sphere_mesh.vertex_buffer['pos']
normals = sphere_mesh.vertex_buffer['norm']
# Convert the normals into quaternion rotations
rotations = np.zeros((N, 4))
for i in range(0, N):
rotations[i, :] = sp.Transforms.QuaternionToRotateXAxisToAlignWithAxis(normals[i, :])
# Generate some random colors
colors = np.random.rand(N,3)
# Create a scene
scene = sp.Scene()
# Create a mesh that we'll turn in to a point-cloud using enable_instancing()
mesh = scene.create_mesh(shared_color = sp.Color(0,1,0), double_sided = True) # shared_color will be overridden in a moment
# Add the primitive to the Mesh - a disc and a thickline showing the normal
mesh.add_disc(segment_count = 20, transform = sp.Transforms.Scale([disc_thickness, 1.0, 1.0]))
mesh.add_thickline(start_point = np.array([disc_thickness * 0.5, 0.0, 0.0]), end_point = np.array([normal_length, 0.0, 0.0]), start_thickness = 0.2, end_thickness = 0.1)
mesh.apply_transform(sp.Transforms.Scale(point_size))
# Now turn the mesh into a point-cloud
mesh.enable_instancing(positions = points, rotations = rotations, colors = colors) # Both rotations and colors are optional
# Create Canvas and Frame, and add Mesh to Frame
canvas = scene.create_canvas_3d(width = 300, height = 300)
frame = canvas.create_frame()
frame.add_mesh(mesh)
scene
# Tutorial 5 - Misc Meshes
# Scene is the top level container in ScenePic
scene = sp.Scene()
# Ok - let's start by creating some Mesh objects
# Mesh 1 - contains a cube and a sphere
# Mesh objects can contain arbitrary triangle mesh and line geometry
# Meshes can belong to "layers" which can be controlled by the user interactively
mesh1 = scene.create_mesh(layer_id = "Sphere+") # No shared_color provided, so per-vertex coloring enabled
mesh1.add_cylinder(color = sp.Color(1.0, 0.0, 0.0), transform = sp.Transforms.Translate([-2.0, 0.0, -2.0]))
mesh1.add_uv_sphere(color = sp.Color(0.0, 0.0, 1.0), transform = np.dot(sp.Transforms.Translate([-1.0, 1.0, 0.0]), sp.Transforms.Scale(1.8)), fill_triangles = False, add_wireframe = True)
mesh1.add_icosphere(color = sp.Color(0.0, 1.0, 1.0), transform = np.dot(sp.Transforms.Translate([2.0, 1.0, 0.0]), sp.Transforms.Scale(1.8)), fill_triangles = False, add_wireframe = True, steps = 2)
# Mesh 2 - coordinate axes
mesh2 = scene.create_mesh(layer_id = "Coords")
mesh2.add_coordinate_axes(transform = sp.Transforms.Translate([0.0, 0.0, 0.0]))
# Mesh 3 - example of Loop Subdivision on a cube
cube_verts = np.array([[-0.5, -0.5, -0.5], [+0.5, -0.5, -0.5], [-0.5, +0.5, -0.5], [+0.5, +0.5, -0.5], [-0.5, -0.5, +0.5], [+0.5, -0.5, +0.5], [-0.5, +0.5, +0.5], [+0.5, +0.5, +0.5]])
cube_tris = np.array([[0, 2, 3], [0, 3, 1], [1, 3, 7], [1, 7, 5], [4, 5, 7], [4, 7, 6], [4, 6, 2], [4, 2, 0], [2, 6, 7], [2, 7, 3], [4, 0, 1], [4, 1, 5]])
cube_verts_a, cube_tris_a = sp.LoopSubdivStencil(cube_tris, 2, False).apply(cube_verts) # Two steps of subdivision, no projection to limit surface. Stencils could be reused for efficiency for other meshes with same triangle topology.
cube_verts_b, cube_tris_b = sp.LoopSubdivStencil(cube_tris, 2, True).apply(cube_verts) # Two steps of subdivision, projection to limit surface. Stencils could be reused for efficiency for other meshes with same triangle topology.
mesh3 = scene.create_mesh(shared_color = sp.Color(1.0, 0.8, 0.8))
mesh3.add_mesh_without_normals(cube_verts, cube_tris, transform = sp.Transforms.Translate([-1.0, 0.0, 0.0])) # Add non-subdivided cube
mesh3.add_mesh_without_normals(cube_verts_a, cube_tris_a)
mesh3.add_mesh_without_normals(cube_verts_b, cube_tris_b, transform = sp.Transforms.Translate([+1.0, 0.0, 0.0]))
# Mesh 4 - line example
mesh4 = scene.create_mesh()
Nsegs = 7000
positions = np.cumsum(np.random.rand(Nsegs, 3) * 0.2, axis = 0)
colored_points = np.concatenate((positions, np.random.rand(Nsegs, 3)), axis = 1)
mesh4.add_lines(colored_points[0:-1, :], colored_points[1:, :])
mesh4.add_camera_frustum(color = sp.Color(1.0,1.0,0.0))
# Let's create two Canvases this time
canvas1 = scene.create_canvas_3d(width = 300, height = 300)
canvas2 = scene.create_canvas_3d(width = 300, height = 300)
# We can link their keyboard/mouse/etc. input events to keep the views in sync
scene.link_canvas_events(canvas1, canvas2)
# And we can specify that certain named "mesh collections" should have user-controlled visibility and opacity
# Meshs without mesh_collection set, or without specified visibilities will always be visible and opaque
canvas1.set_layer_settings({"Coords" : { "opacity" : 0 }, "Sphere+" : { "opacity" : 1 }})
# A Frame contains an array of meshes
frame11 = canvas1.create_frame(meshes = [mesh1, mesh2]) # Note that Frames are created from the Canvas not the Scene
frame21 = canvas2.create_frame(meshes = [mesh2, mesh3])
frame22 = canvas2.create_frame(meshes = [mesh4, mesh1])
# ScenePic has told Jupyter how to display scene objects
scene
# Tutorial 6 - Images and Textures
# Scene is the top level container in ScenePic
scene = sp.Scene()
# Create and populate an Image object
image1 = scene.create_image(image_id = "PolarBear")
image1.load(asset_path("PolarBear.png")) # This will preserve the image data in compressed PNG format
# Create a texture map
texture = scene.create_image(image_id = "texture")
texture.load(asset_path("uv.png")) # we can use this image to skin meshes
# Example of a mesh that is defined in camera space not world space
# This will not move as the virtual camera is moved with the mouse
cam_space_mesh = scene.create_mesh(shared_color = sp.Color(1.0, 0.0, 0.0), camera_space = True)
cam_space_mesh.add_sphere(transform = np.dot(sp.Transforms.Translate([10, -10, -20.0]), sp.Transforms.Scale(1.0)))
# Some textured primitives
sphere = scene.create_mesh(texture_id=texture.image_id, nn_texture = False)
sphere.add_icosphere(steps=4)
cube = scene.create_mesh(texture_id=texture.image_id)
transform = sp.Transforms.translate([-1, 0, 0]) @ sp.Transforms.scale(0.5)
cube.add_cube(transform=transform)
# Show images in 3D canvas
canvas = scene.create_canvas_3d(shading=sp.Shading(bg_color=sp.Colors.White))
mesh1 = scene.create_mesh(texture_id = "PolarBear")
mesh1.add_image() # Adds image in canonical position
# Add an animation that rigidly transforms each image
n_frames = 20
for i in range(n_frames):
angle = 2 * math.pi * i / n_frames
c, s = math.cos(angle), math.sin(angle)
# Create a focus point that allows you to "lock" the camera's translation and optionally orientation by pressing the "l" key
axis = np.array([1.0, 0.0, 1.0])
axis /= np.linalg.norm(axis)
focus_point = sp.FocusPoint([c,s,0], orientation_axis_angle = axis * angle)
mesh = scene.create_mesh()
mesh.add_coordinate_axes(transform = np.dot(sp.Transforms.Translate(focus_point.position), sp.Transforms.RotationMatrixFromAxisAngle(axis, angle)))
im_size = 15
im_data = np.random.rand(im_size, im_size, 4)
im_data[:,:,3] = 0.5 + 0.5 * im_data[:,:,3]
imageB = scene.create_image()
imageB.from_numpy(im_data) # Converts data to PNG format
meshB = scene.create_mesh(texture_id = imageB, is_billboard = True, use_texture_alpha=True)
meshB.add_image(transform = np.dot(sp.Transforms.Scale(2.0), sp.Transforms.Translate([0,0,-1])))
frame = canvas.create_frame(focus_point = focus_point)
frame.add_mesh(mesh1, transform = sp.Transforms.Translate([c,s,0]))
frame.add_mesh(meshB, transform = np.dot(sp.Transforms.Scale(i * 1.0 / n_frames), sp.Transforms.Translate([-c,-s,0])))
frame.add_mesh(cam_space_mesh)
frame.add_mesh(sphere, transform=sp.Transforms.rotation_about_y(np.pi * 2 * i / n_frames))
frame.add_mesh(cube, transform=sp.Transforms.rotation_about_y(-np.pi * 2 * i / n_frames))
frame.add_mesh(mesh)
# Show Scene
scene
# Tutorial 7 - 2D canvases
# Scene is the top level container in ScenePic
scene = sp.Scene()
# Load an image
image1 = scene.create_image(image_id = "PolarBear")
image1.load(asset_path("PolarBear.png")) # This will preserve the image data in compressed PNG format
# Create and populate an Image object
image2 = scene.create_image(image_id = "Random")
image2.from_numpy(np.random.rand(20, 30, 3) * 128 / 255.0) # Converts data to PNG format
# Create a 2D canvas demonstrating different image positioning options
canvas1 = scene.create_canvas_2d(width = 400, height = 300, background_color = sp.Colors.White)
canvas1.create_frame().add_image(image1, "fit")
canvas1.create_frame().add_image(image1, "fill")
canvas1.create_frame().add_image(image1, "stretch")
canvas1.create_frame().add_image(image1, "manual", x = 50, y= 50, scale = 0.3)
# You can composite images and primitives too
canvas2 = scene.create_canvas_2d(width = 300, height = 300)
f = canvas2.create_frame()
f.add_image(image2, "fit")
f.add_image(image1, "manual", x = 30, y= 30, scale = 0.2)
f.add_circle(200, 200, 40, fill_color = sp.Colors.Black, line_width = 10, line_color = sp.Colors.Blue)
f.add_rectangle(200, 100, 50, 25, fill_color = sp.Colors.Green, line_width = 0)
f.add_text("Hello World", 30, 100, sp.Colors.White, 100, "segoe ui light")
scene.framerate = 2
scene
# Tutorial 8 - a mix of transparent and opaque objects, with labels
np.random.seed(55)
scene = sp.Scene()
canvas = scene.create_canvas_3d(width = 700, height = 700)
frame = canvas.create_frame()
# Create a mesh that we'll turn in to a point-cloud using enable_instancing()
layer_settings = { "Labels" : { "opacity" : 1.0 }}
N = 20
for i in range(N):
# Sample object
geotype = np.random.randint(2)
color = np.random.rand(3)
size = 0.3 * np.random.rand() + 0.2
position = 3.0 * np.random.rand(3) - 1.5
opacity = 1.0 if np.random.randint(2) == 0 else np.random.uniform(0.45, 0.55)
# Generate geometry
layer_id = "Layer" + str(i)
mesh = scene.create_mesh(shared_color = color, layer_id = layer_id)
layer_settings[layer_id] = { "opacity" : opacity }
if geotype == 0:
mesh.add_cube()
elif geotype == 1:
mesh.add_sphere()
mesh.apply_transform(sp.Transforms.Scale(size)) # Scale the primitive
mesh.apply_transform(sp.Transforms.Translate(position))
frame.add_mesh(mesh)
# Add label
text = "{0:0.2f} {1:0.2f} {2:0.2f} {3:0.2f}".format(color[0], color[1], color[2], opacity)
horizontal_align = ["left", "center", "right"][np.random.randint(3)]
vertical_align = ["top", "middle", "bottom"][np.random.randint(3)]
if geotype == 0:
if horizontal_align != "center" and vertical_align != "middle":
offset_distance = size * 0.7
else:
offset_distance = size * 0.9
else:
if horizontal_align != "center" and vertical_align != "middle":
offset_distance = size * 0.5 * 0.8
else:
offset_distance = size * 0.6
label = scene.create_label(text = text, color = color, layer_id = "Labels", font_family = "consolas", size_in_pixels = 80 * size, offset_distance = offset_distance, vertical_align = vertical_align, horizontal_align = horizontal_align)
frame.add_label(label = label, position = position)
canvas.set_layer_settings(layer_settings)
scene
# Tutorial 9 - mesh animation
# let's create our mesh to get started
scene = sp.Scene()
canvas = scene.create_canvas_3d(width=700, height=700)
# Load a mesh to animate
jelly_mesh = sp.load_obj(asset_path("jelly.obj"))
texture = scene.create_image("texture")
texture.load(asset_path("jelly.png"))
# create a base mesh for the animation. The animation
# will only change the vertex positions, so this mesh
# is used to set everything else, e.g. textures.
base_mesh = scene.create_mesh("jelly_base")
base_mesh.texture_id = texture.image_id
base_mesh.use_texture_alpha = True
base_mesh.add_mesh(jelly_mesh)
def random_linspace(min_val, max_val, num_samples):
vals = np.linspace(min_val, max_val, num_samples)
np.random.shuffle(vals)
return vals
# this base mesh will be instanced, so we can animate each
# instance individual using rigid transforms, in this case
# just translation.
marbles = scene.create_mesh("marbles_base")
num_marbles = 10
marbles.add_sphere(sp.Colors.White, transform=sp.Transforms.Scale(0.2))
marble_positions = np.zeros((num_marbles, 3), np.float32)
marble_positions[:, 0] = random_linspace(-0.6, 0.6, num_marbles)
marble_positions[:, 2] = random_linspace(-1, 0.7, num_marbles)
marble_offsets = np.random.uniform(0, 2*np.pi, size=num_marbles).astype(np.float32)
marble_colors_start = np.random.uniform(0, 1, size=(num_marbles, 3)).astype(np.float32)
marble_colors_end = np.random.uniform(0, 1, size=(num_marbles, 3)).astype(np.float32)
marbles.enable_instancing(marble_positions, colors=marble_colors_start)
for i in range(60):
# animate the wave mesh by updating the vertex positions
positions = jelly_mesh.positions.copy()
delta_x = (positions[:, 0] + 0.0838 * i) * 10
delta_z = (positions[:, 2] + 0.0419 * i) * 10
positions[:, 1] = positions[:, 1] + 0.1 * (np.cos(delta_x) + np.sin(delta_z))
# we create a mesh update with the new posiitons. We can use this mesh update
# just like a new mesh, because it essentially is one, as ScenePic will create
# a new mesh from the old one using these new positions.
jelly_update = scene.update_mesh_positions("jelly_base", positions)
frame = canvas.create_frame(meshes=[jelly_update])
# this is a simpler form of animation in which we will change the position
# and colors of the marbles
marble_y = np.sin(0.105 * i + marble_offsets)
positions = np.stack([marble_positions[:, 0], marble_y, marble_positions[:, 2]], -1)
alpha = ((np.sin(marble_y) + 1) * 0.5).reshape(-1, 1)
beta = 1 - alpha
colors = alpha * marble_colors_start + beta * marble_colors_end
marbles_update = scene.update_instanced_mesh("marbles_base", positions, colors=colors)
frame.add_mesh(marbles_update)
scene.quantize_updates()
scene
# Tutorial 10 - Instanced Animation
# In this tutorial we will explore how we can use mesh updates on
# instanced meshes as well. We will begin by creating a simple primitive
# and use instancing to create a cloud of stylized butterflies. We will
# then using mesh updates on the instances to make the butterflies
# fly.
scene = sp.Scene()
butterflies = scene.create_mesh("butterflies", double_sided=True)
# the primitive will be a single wing, and we'll use instancing to create
# all the butterflies
butterflies.add_quad(sp.Colors.Blue, [0, 0, 0], [0.1, 0, 0.04], [0.08, 0, -0.06], [0.015, 0, -0.03])
rotate_back = sp.Transforms.quaternion_from_axis_angle([1, 0, 0], -np.pi / 6)
num_butterflies = 100
num_anim_frames = 20
# this will make them flap their wings independently
start_frames = np.random.randint(0, num_anim_frames, num_butterflies)
rot_angles = np.random.uniform(-1, 1, num_butterflies)
rotations = np.zeros((num_butterflies * 2, 4), np.float32)
positions = np.random.uniform(-1, 1, (num_butterflies * 2, 3))
colors = np.random.random((num_butterflies * 2, 3))
for b, angle in enumerate(rot_angles):
rot = sp.Transforms.quaternion_from_axis_angle([0, 1, 0], angle)
rotations[2 * b] = rotations[2 * b + 1] = rot
# we will use the second position per butterfly as a destination
dx = np.sin(angle) * 0.1
dy = positions[2 * b + 1, 1] - positions[2 * b, 1]
dy = np.sign(angle) * min(abs(angle), 0.1)
dz = np.cos(angle) * 0.1
positions[2 * b + 1] = positions[2 * b] + [dx, dy, dz]
butterflies.enable_instancing(positions, rotations, colors)
canvas = scene.create_canvas_3d("main", 700, 700)
canvas.shading = sp.Shading(sp.Colors.White)
start = -np.pi / 6
end = np.pi / 2
delta = (end - start) / (num_anim_frames // 2 - 1)
# let's construct the animation frame by frame
animation = []
for i in range(num_anim_frames):
frame_positions = np.zeros_like(positions)
frame_rotations = np.zeros_like(rotations)
frame_colors = np.zeros_like(colors)
for b, start_frame in enumerate(start_frames):
frame = (i + start_frame) % num_anim_frames
if frame < num_anim_frames // 2:
angle = start + delta * frame
else:
angle = end + delta * (frame - num_anim_frames // 2)
right = sp.Transforms.quaternion_from_axis_angle([0, 0, 1], angle)
right = sp.Transforms.quaternion_multiply(rotate_back, right)
right = sp.Transforms.quaternion_multiply(rotations[2 * b], right)
left = sp.Transforms.quaternion_from_axis_angle([0, 0, 1], np.pi - angle)
left = sp.Transforms.quaternion_multiply(rotate_back, left)
left = sp.Transforms.quaternion_multiply(rotations[2 * b + 1], left)
frame_rotations[2 * b] = right
frame_rotations[2 * b + 1] = left
progress = np.sin((frame * 2 * np.pi) / num_anim_frames)
progress = (progress + 1) * 0.5
# we move the butterfly along its path
pos = (1 - progress) * positions[2 * b] + progress * positions[2 * b + 1]
pos[1] -= np.sin(angle) * 0.02
frame_positions[2 * b : 2 * b + 2, :] = pos
# finally we alter the color
color = (1 - progress) * colors[2 * b] + progress * colors[2 * b + 1]
frame_colors[2 * b : 2 * b + 2, :] = color
# now we create the update. Here we update position, rotation,
# and color, but you can update them separately as well by passing
# the `*None()` versions of the buffers to this function.
update = scene.update_instanced_mesh("butterflies", frame_positions, frame_rotations, frame_colors)
animation.append(update)
# now we create the encapsulating animation which will move the camera
# around the butterflies. The inner animation will loop as the camera moves.
num_frames = 300
cameras = sp.Camera.orbit(num_frames, 3, 2)
for i, camera in enumerate(cameras):
frame = canvas.create_frame()
frame.add_mesh(animation[i % num_anim_frames])
frame.camera = camera
scene
# Tutorial 11 - camera movement
# in this tutorial we will show how to create per-frame camera movement.
# while the user can always choose to override this behavior, having a
# camera track specified can be helpful for demonstrating particular
# items in 3D. We will also show off the flexible GLCamera class.
scene = sp.Scene()
spin_canvas = scene.create_canvas_3d("spin")
spiral_canvas = scene.create_canvas_3d("spiral")
# let's create some items in the scene so we have a frame of reference
polar_bear = scene.create_image(image_id="polar_bear")
polar_bear.load(asset_path("PolarBear.png"))
uv_texture = scene.create_image(image_id = "texture")
uv_texture.load(asset_path("uv.png"))
cube = scene.create_mesh("cube", texture_id=polar_bear.image_id)
cube.add_cube()
sphere = scene.create_mesh("sphere", texture_id=uv_texture.image_id)
sphere.add_icosphere(steps=4, transform=sp.Transforms.translate([0, 1, 0]))
num_frames = 60
for i in range(num_frames):
angle = i*np.pi*2/num_frames
# for the first camera we will spin in place on the Z axis
rotation = sp.Transforms.rotation_about_z(angle)
spin_camera = sp.Camera(center=[0, 0, 4], rotation=rotation, fov_y_degrees=30.0)
# for the second camera, we will spin the camera in a spiral around the scene
# we can do this using the look-at initialization, which provides a straightforward
# "look at" interface for camera placement.
camera_center = [4*np.cos(angle), i*4/num_frames - 2, 4*np.sin(angle)]
spiral_camera = sp.Camera(camera_center, look_at=[0, 0.5, 0])
# we can add frustums directly using the ScenePic camera objects
frustums = scene.create_mesh()
frustums.add_camera_frustum(spin_camera, sp.Colors.Red)
frustums.add_camera_frustum(spiral_camera, sp.Colors.Green)
spin_frame = spin_canvas.create_frame()
spin_frame.camera = spin_camera # each frame can have its own camera object
spin_frame.add_meshes([cube, sphere, frustums])
spiral_frame = spiral_canvas.create_frame()
spiral_frame.camera = spiral_camera
spiral_frame.add_meshes([cube, sphere, frustums])
scene.link_canvas_events(spin_canvas, spiral_canvas)
scene
# Tutorial 12 - audio tracks
# in this tutorial we'll show how to attach audio tracks to canvases. ScenePic
# supports any audio file format supported by the browser.
def _set_audio(scene, canvas, path):
audio = scene.create_audio()
audio.load(path)
canvas.media_id = audio.audio_id
scene = sp.Scene()
names = ["red", "green", "blue"]
colors = [sp.Colors.Red, sp.Colors.Green, sp.Colors.Blue]
frequencies = [0, 1, 0.5]
graph = scene.create_graph("graph", width=900, height=150)
for name, color, frequency in zip(names, colors, frequencies):
mesh = scene.create_mesh()
mesh.add_cube(color)
canvas = scene.create_canvas_3d(name, width=300, height=300)
_set_audio(scene, canvas, asset_path(name + ".ogg"))
values = []
for j in range(60):
frame = canvas.create_frame()
scale = math.sin(j * 2 * math.pi * frequency / 30)
frame.add_mesh(mesh, sp.Transforms.scale((scale + 1) / 2 + 0.5))
values.append(scale)
graph.add_sparkline(name, values, color)
graph.media_id = canvas.media_id
names.append("graph")
scene.grid("600px", "1fr auto", "1fr 1fr 1fr")
scene.place("graph", "2", "1 / span 3")
scene.link_canvas_events(*names)
scene
# Tutorial 13 - video
# It is also possible to attach videos to ScenePic scenes. Once attached, you can draw the
# frames of those videos to canvases in the same way as images, and can draw the same
# video to multiple frames. Once a media file (video or audio) has been attached to a
# canvas, that file will be used to drive playback. In practical terms, this means that
# ScenePic will display frames such that they line up with the timestamps of the video
# working on the assumption that ScenePic frames are displayed at the framerate of the video.
def _angle_to_pos(angle, radius):
return np.cos(angle) * radius + 200, np.sin(angle) * radius + 200
scene = sp.Scene()
video = scene.create_video()
video.load(asset_path("circles.mp4"))
tracking = scene.create_canvas_2d("tracking", background_color=sp.Colors.White)
tracking.media_id = video.video_id
multi = scene.create_canvas_2d("multi", background_color=sp.Colors.White)
multi.media_id = video.video_id
angles = np.linspace(0, 2 * np.pi, 360, endpoint=False)
for angle in angles:
# if a 2D canvas has an associated video
# then a frame of that video can be added
# via the add_video method.
frame = tracking.create_frame()
frame.add_video(layer_id="video")
red_pos = _angle_to_pos(angle, 160)
frame.add_rectangle(red_pos[0] - 11, red_pos[1] - 11, 22, 22, [255, 0, 0], 2, layer_id="rect")
frame.add_circle(red_pos[0], red_pos[1], 10, fill_color=[255, 0, 0], layer_id="dot")
green_pos = _angle_to_pos(-2*angle, 80)
frame.add_rectangle(green_pos[0] - 11, green_pos[1] - 11, 22, 22, [0, 255, 0], 2, layer_id="rect")
frame.add_circle(green_pos[0], green_pos[1], 10, fill_color=[0, 255, 0], layer_id="dot")
blue_pos = _angle_to_pos(4*angle, 40)
frame.add_rectangle(blue_pos[0] - 11, blue_pos[1] - 11, 22, 22, [0, 0, 255], 2, layer_id="rect")
frame.add_circle(blue_pos[0], blue_pos[1], 10, fill_color=[0, 0, 255], layer_id="dot")
frame = multi.create_frame()
frame.add_video("manual", red_pos[0] - 40, red_pos[1] - 40, 0.2, layer_id="red")
frame.add_video("manual", green_pos[0] - 25, green_pos[1] - 25, 0.125, layer_id="green")
frame.add_video("manual", 160, 160, 0.2, layer_id="blue")
tracking.set_layer_settings({
"rect": {"render_order": 0},
"video": {"render_order": 1},
"dot": {"render_order": 2}
})
scene.link_canvas_events("tracking", "multi")
scene
# Tutorial 14 - Multiview Visualization
# One common and useful scenario for ScenePic is to visualize the result of multiview 3D reconstruction.
# In this tutorial we'll show how to load some geometry, assocaited camera calibration
# information, and images to create a visualization depicting the results.
def _load_camera(camera_info):
# this function loads an "OpenCV"-style camera representation
# and converts it to a GL style for use in ScenePic
location = np.array(camera_info["location"], np.float32)
euler_angles = np.array(camera_info["rotation"], np.float32)
rotation = sp.Transforms.euler_angles_to_matrix(euler_angles, "XYZ")
translation = sp.Transforms.translate(location)
extrinsics = translation @ rotation
world_to_camera = sp.Transforms.gl_world_to_camera(extrinsics)
aspect_ratio = camera_info["width"] / camera_info["height"]
projection = sp.Transforms.gl_projection(camera_info["fov"], aspect_ratio, 0.01, 100)
return sp.Camera(world_to_camera, projection)
def _load_cameras():
with open(asset_path("cameras.json")) as file:
cameras = json.load(file)
return [_load_camera(cameras[key])
for key in cameras]
scene = sp.Scene()
# load the fitted cameras
cameras = _load_cameras()
# this textured cube will stand in for a reconstructed mesh
texture = scene.create_image("texture")
texture.load(asset_path("PolarBear.png"))
cube = scene.create_mesh("cube")
cube.texture_id = texture.image_id
cube.add_cube(transform=sp.Transforms.scale(2))
# construct all of the frustums
# and camera images
frustums = scene.create_mesh("frustums", layer_id="frustums")
colors = [sp.Colors.Red, sp.Colors.Green, sp.Colors.Blue]
paths = [asset_path(name) for name in ["render0.png", "render1.png", "render2.png"]]
camera_images = []
images = []
for i, (color, path, camera) in enumerate(zip(colors, paths, cameras)):
image = scene.create_image(path)
image.load(path)
frustums.add_camera_frustum(camera, color)
image_mesh = scene.create_mesh("image{}".format(i),
layer_id="images",
shared_color=sp.Colors.Gray,
double_sided=True,
texture_id=image.image_id)
image_mesh.add_camera_image(camera)
images.append(image)
camera_images.append(image_mesh)
# create one canvas for each camera to show the scene from
# that camera's viewpoint
width = 640
for i, camera in enumerate(cameras):
height = width / camera.aspect_ratio
canvas = scene.create_canvas_3d("hand{}".format(i), width, height, camera=camera)
frame = canvas.create_frame()
frame.add_mesh(cube)
frame.add_mesh(frustums)
frame.camera = camera
for cam_mesh in camera_images:
frame.add_mesh(cam_mesh)
scene
# Tutorial 15 - Frame Layer Settings
# It is possible to use the per-frame layer settings to automatically
# change various layer properties, for example to fade meshes in and
# out of view. The user can still override this manually using the
# controls, of course, but this feature can help guide the user through
# more complex animations.
scene = sp.Scene()
# In this tutorial we will fade out one mesh (the cube) and fade
# another in (the sphere).
cube = scene.create_mesh(layer_id="cube")
cube.add_cube(sp.Colors.Green)
sphere = scene.create_mesh(layer_id="sphere")
sphere.add_sphere(sp.Colors.Red)
canvas = scene.create_canvas_3d()
for i in range(60):
sphere_opacity = i / 59
cube_opacity = 1 - sphere_opacity
frame = canvas.create_frame()
frame.add_mesh(cube)
frame.add_mesh(sphere)
# the interface here is the same as with how layer settings
# usually works at the canvas level.
frame.set_layer_settings({
"cube": {"opacity": cube_opacity},
"sphere": {"opacity": sphere_opacity}
})
scene
```
| github_jupyter |
```
############## PLEASE RUN THIS CELL FIRST! ###################
# import everything and define a test runner function
from importlib import reload
from helper import run
import helper, op, script, tx
```
### Exercise 1
#### Make [this test](/edit/session5/tx.py) pass: `tx.py:TxTest:test_verify_p2pkh`
```
# Exercise 1
reload(tx)
run(tx.TxTest('test_verify_p2pkh'))
# Transaction Construction Example
from ecc import PrivateKey
from helper import decode_base58, SIGHASH_ALL
from script import p2pkh_script, Script
from tx import Tx, TxIn, TxOut
# Step 1
tx_ins = []
prev_tx = bytes.fromhex('95ae966a54493321316799d5c4ba7f48faadd653b7eb725ea2c5ea7d38f65cff')
prev_index = 0
tx_ins.append(TxIn(prev_tx, prev_index))
# Step 2
tx_outs = []
h160 = decode_base58('mzx5YhAH9kNHtcN481u6WkjeHjYtVeKVh2')
tx_outs.append(TxOut(
amount=int(0.37*100000000),
script_pubkey=p2pkh_script(h160),
))
h160 = decode_base58('mnrVtF8DWjMu839VW3rBfgYaAfKk8983Xf')
tx_outs.append(TxOut(
amount=int(0.1*100000000),
script_pubkey=p2pkh_script(h160),
))
tx_obj = Tx(1, tx_ins, tx_outs, 0, network="signet")
# Step 3
z = tx_obj.sig_hash(0)
pk = PrivateKey(secret=8675309)
der = pk.sign(z).der()
sig = der + SIGHASH_ALL.to_bytes(1, 'big')
sec = pk.point.sec()
tx_obj.tx_ins[0].script_sig = Script([sig, sec])
print(tx_obj.serialize().hex())
```
### Exercise 2
#### Make [this test](/edit/session5/tx.py) pass: `tx.py:TxTest:test_sign_input`
```
# Exercise 2
reload(tx)
run(tx.TxTest('test_sign_input'))
```
### Exercise 3
You have been sent 100,000 Sats on the Signet network. Send 40,000 Sats to this address: `mqYz6JpuKukHzPg94y4XNDdPCEJrNkLQcv` and send the rest back to yourself.
#### Send your transaction here: https://mempool.space/signet/tx/push
```
# Exercise 3
from tx import Tx, TxIn, TxOut
from helper import decode_base58, hash256, little_endian_to_int
from script import p2pkh_script
prev_tx = bytes.fromhex('d487dc425d0103aee99c46f8ab58a01ee99a19d9ce9f4db07e196597c4e73a29')
prev_index = 0
target_address = 'mqYz6JpuKukHzPg94y4XNDdPCEJrNkLQcv'
target_amount = 40000
fee = 300
passphrase = b'Jimmy Song'
secret = little_endian_to_int(hash256(passphrase))
private_key = PrivateKey(secret=secret)
change_address = private_key.point.address(network="signet")
# initialize inputs
tx_ins = []
# create a new tx input with prev_tx, prev_index
tx_ins.append(TxIn(prev_tx, prev_index))
# initialize outputs
tx_outs = []
# decode the hash160 from the target address
target_h160 = decode_base58(target_address)
# convert hash160 to p2pkh script
target_script_pubkey = p2pkh_script(target_h160)
# create a new tx output for target with amount and script_pubkey
tx_outs.append(TxOut(target_amount, target_script_pubkey))
# decode the hash160 from the change address
change_h160 = decode_base58(change_address)
# convert hash160 to p2pkh script
change_script_pubkey = p2pkh_script(change_h160)
# get the value for the transaction input (remember network="signet")
prev_amount = tx_ins[0].value(network="signet")
# calculate change_amount based on previous amount, target_amount & fee
change_amount = prev_amount - target_amount - fee
# create a new tx output for change with amount and script_pubkey
tx_outs.append(TxOut(change_amount, change_script_pubkey))
# create the transaction (name it tx_obj and set network="signet")
tx_obj = Tx(1, tx_ins, tx_outs, 0, network="signet")
# now sign the 0th input with the private_key using sign_input
tx_obj.sign_input(0, private_key)
# SANITY CHECK: change address corresponds to private key
if private_key.point.address(network="signet") != change_address:
raise RuntimeError('Private Key does not correspond to Change Address, check priv_key and change_address')
# SANITY CHECK: output's script_pubkey is the same one as your address
if tx_ins[0].script_pubkey(network="signet").commands[2] != decode_base58(change_address):
raise RuntimeError('Output is not something you can spend with this private key. Check that the prev_tx and prev_index are correct')
# SANITY CHECK: fee is reasonable
if tx_obj.fee() > 100000 or tx_obj.fee() <= 0:
raise RuntimeError(f'Check that the change amount is reasonable. Fee is {tx_obj.fee()}')
# serialize and hex()
print(tx_obj.serialize().hex())
```
### Exercise 4
#### Bonus Question. Only attempt if you've finished Exercise 3 and have time to try it.
Get some signet coins from a faucet and spend both outputs (one from your change address and one from the signet faucet) to
`mqYz6JpuKukHzPg94y4XNDdPCEJrNkLQcv`
#### You can get some free signet coins at: https://signet.bc-2.jp/
```
# Exercise 4
# Bonus
from tx import Tx, TxIn, TxOut
from helper import decode_base58, hash256, little_endian_to_int
from script import p2pkh_script
prev_tx_1 = bytes.fromhex('c2cfd11f33297c36c61f8911529b05fc14eab09e610461768f9eab6cc2576acf')
prev_index_1 = 1
prev_tx_2 = bytes.fromhex('368941a52f2eb3a87b84b85396db0d2203a0eb32c95559d4249a8f02afa5e239')
prev_index_2 = 0
target_address = 'mqYz6JpuKukHzPg94y4XNDdPCEJrNkLQcv'
fee = 400
passphrase = b'Jimmy Song'
secret = little_endian_to_int(hash256(passphrase))
private_key = PrivateKey(secret=secret)
# initialize inputs
tx_ins = []
# create the first tx input with prev_tx_1, prev_index_1
tx_ins.append(TxIn(prev_tx_1, prev_index_1))
# create the second tx input with prev_tx_2, prev_index_2
tx_ins.append(TxIn(prev_tx_2, prev_index_2))
# initialize outputs
tx_outs = []
# decode the hash160 from the target address
h160 = decode_base58(target_address)
# convert hash160 to p2pkh script
script_pubkey = p2pkh_script(h160)
# calculate target amount by adding the input values and subtracting the fee
target_satoshis = tx_ins[0].value(network="signet") + tx_ins[1].value(network="signet") - fee
# create a single tx output for target with amount and script_pubkey
tx_outs.append(TxOut(target_satoshis, script_pubkey))
# create the transaction
tx_obj = Tx(1, tx_ins, tx_outs, 0, network="signet")
# sign both inputs with the private key using sign_input
tx_obj.sign_input(0, private_key)
tx_obj.sign_input(1, private_key)
# SANITY CHECK: output's script_pubkey is the same one as your address
if tx_ins[0].script_pubkey(network="signet").commands[2] != decode_base58(private_key.point.address(network="signet")):
raise RuntimeError('Output is not something you can spend with this private key. Check that the prev_tx and prev_index are correct')
# SANITY CHECK: fee is reasonable
if tx_obj.fee() > 100000 or tx_obj.fee() <= 0:
raise RuntimeError('Check that the change amount is reasonable. Fee is {tx_obj.fee()}')
# serialize and hex()
print(tx_obj.serialize().hex())
# op_checkmultisig
def op_checkmultisig(stack, z):
if len(stack) < 1:
return False
n = decode_num(stack.pop())
if len(stack) < n + 1:
return False
sec_pubkeys = []
for _ in range(n):
sec_pubkeys.append(stack.pop())
m = decode_num(stack.pop())
if len(stack) < m + 1:
return False
der_signatures = []
for _ in range(m):
# signature is assumed to be using SIGHASH_ALL
der_signatures.append(stack.pop()[:-1])
# OP_CHECKMULTISIG bug
stack.pop()
try:
raise NotImplementedError
except (ValueError, SyntaxError):
return False
return True
```
### Exercise 5
#### Make [this test](/edit/session5/op.py) pass: `op.py:OpTest:test_op_checkmultisig`
```
# Exercise 5
reload(op)
reload(script)
run(op.OpTest('test_op_checkmultisig'))
```
### Exercise 6
Find the hash160 of the RedeemScript
```
5221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152ae
```
```
# Exercise 6
from helper import hash160
hex_redeem_script = '5221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152ae'
# bytes.fromhex script
redeem_script = bytes.fromhex(hex_redeem_script)
# hash160 result
h160 = hash160(redeem_script)
# hex() to display
print(h160.hex())
# P2SH address construction example
from helper import encode_base58_checksum
print(encode_base58_checksum(b'\x05'+bytes.fromhex('74d691da1574e6b3c192ecfb52cc8984ee7b6c56')))
```
### Exercise 7
#### Make [this test](/edit/session5/helper.py) pass: `helper.py:HelperTest:test_p2pkh_address`
```
# Exercise 7
reload(helper)
run(helper.HelperTest('test_p2pkh_address'))
```
### Exercise 8
#### Make [this test](/edit/session5/helper.py) pass: `helper.py:HelperTest:test_p2sh_address`
```
# Exercise 8
reload(helper)
run(helper.HelperTest('test_p2sh_address'))
```
### Exercise 9
#### Make [this test](/edit/session5/script.py) pass: `script.py:ScriptTest:test_address`
```
# Exercise 9
reload(script)
run(script.ScriptTest('test_address'))
# z for p2sh example
from helper import hash256
h256 = hash256(bytes.fromhex('0100000001868278ed6ddfb6c1ed3ad5f8181eb0c7a385aa0836f01d5e4789e6bd304d87221a000000475221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152aeffffffff04d3b11400000000001976a914904a49878c0adfc3aa05de7afad2cc15f483a56a88ac7f400900000000001976a914418327e3f3dda4cf5b9089325a4b95abdfa0334088ac722c0c00000000001976a914ba35042cfe9fc66fd35ac2224eebdafd1028ad2788acdc4ace020000000017a91474d691da1574e6b3c192ecfb52cc8984ee7b6c56870000000001000000'))
z = int.from_bytes(h256, 'big')
print(hex(z))
# p2sh verification example
from ecc import S256Point, Signature
from helper import hash256
h256 = hash256(bytes.fromhex('0100000001868278ed6ddfb6c1ed3ad5f8181eb0c7a385aa0836f01d5e4789e6bd304d87221a000000475221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152aeffffffff04d3b11400000000001976a914904a49878c0adfc3aa05de7afad2cc15f483a56a88ac7f400900000000001976a914418327e3f3dda4cf5b9089325a4b95abdfa0334088ac722c0c00000000001976a914ba35042cfe9fc66fd35ac2224eebdafd1028ad2788acdc4ace020000000017a91474d691da1574e6b3c192ecfb52cc8984ee7b6c56870000000001000000'))
z = int.from_bytes(h256, 'big')
point = S256Point.parse(bytes.fromhex('022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb70'))
sig = Signature.parse(bytes.fromhex('3045022100dc92655fe37036f47756db8102e0d7d5e28b3beb83a8fef4f5dc0559bddfb94e02205a36d4e4e6c7fcd16658c50783e00c341609977aed3ad00937bf4ee942a89937'))
print(point.verify(z, sig))
```
### Exercise 10
Validate the second signature of the first input
```
0100000001868278ed6ddfb6c1ed3ad5f8181eb0c7a385aa0836f01d5e4789e6bd304d87221a000000db00483045022100dc92655fe37036f47756db8102e0d7d5e28b3beb83a8fef4f5dc0559bddfb94e02205a36d4e4e6c7fcd16658c50783e00c341609977aed3ad00937bf4ee942a8993701483045022100da6bee3c93766232079a01639d07fa869598749729ae323eab8eef53577d611b02207bef15429dcadce2121ea07f233115c6f09034c0be68db99980b9a6c5e75402201475221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152aeffffffff04d3b11400000000001976a914904a49878c0adfc3aa05de7afad2cc15f483a56a88ac7f400900000000001976a914418327e3f3dda4cf5b9089325a4b95abdfa0334088ac722c0c00000000001976a914ba35042cfe9fc66fd35ac2224eebdafd1028ad2788acdc4ace020000000017a91474d691da1574e6b3c192ecfb52cc8984ee7b6c568700000000
```
The sec pubkey of the second signature is:
```
03b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb71
```
The der signature of the second signature is:
```
3045022100da6bee3c93766232079a01639d07fa869598749729ae323eab8eef53577d611b02207bef15429dcadce2121ea07f233115c6f09034c0be68db99980b9a6c5e75402201475221022
```
The redeemScript is:
```
475221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152ae
```
```
# Exercise 10
from io import BytesIO
from ecc import S256Point, Signature
from helper import int_to_little_endian, SIGHASH_ALL
from script import Script
from tx import Tx
hex_sec = '03b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb71'
hex_der = '3045022100da6bee3c93766232079a01639d07fa869598749729ae323eab8eef53577d611b02207bef15429dcadce2121ea07f233115c6f09034c0be68db99980b9a6c5e754022'
hex_redeem_script = '475221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152ae'
sec = bytes.fromhex(hex_sec)
der = bytes.fromhex(hex_der)
redeem_script_stream = BytesIO(bytes.fromhex(hex_redeem_script))
redeem_script = Script.parse(redeem_script_stream)
hex_tx = '0100000001868278ed6ddfb6c1ed3ad5f8181eb0c7a385aa0836f01d5e4789e6bd304d87221a000000db00483045022100dc92655fe37036f47756db8102e0d7d5e28b3beb83a8fef4f5dc0559bddfb94e02205a36d4e4e6c7fcd16658c50783e00c341609977aed3ad00937bf4ee942a8993701483045022100da6bee3c93766232079a01639d07fa869598749729ae323eab8eef53577d611b02207bef15429dcadce2121ea07f233115c6f09034c0be68db99980b9a6c5e75402201475221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152aeffffffff04d3b11400000000001976a914904a49878c0adfc3aa05de7afad2cc15f483a56a88ac7f400900000000001976a914418327e3f3dda4cf5b9089325a4b95abdfa0334088ac722c0c00000000001976a914ba35042cfe9fc66fd35ac2224eebdafd1028ad2788acdc4ace020000000017a91474d691da1574e6b3c192ecfb52cc8984ee7b6c568700000000'
stream = BytesIO(bytes.fromhex(hex_tx))
# parse the S256Point and Signature
point = S256Point.parse(sec)
sig = Signature.parse(der)
# parse the Tx
t = Tx.parse(stream)
# change the first input's ScriptSig to RedeemScript
t.tx_ins[0].script_sig = redeem_script
# get the serialization
ser = t.serialize()
# add the sighash (4 bytes, little-endian of SIGHASH_ALL)
ser += int_to_little_endian(SIGHASH_ALL, 4)
# hash256 the result
h256 = hash256(ser)
# your z is the hash256 as a big-endian number: use int.from_bytes(x, 'big')
z = int.from_bytes(h256, 'big')
# now verify the signature using point.verify
print(point.verify(z, sig))
```
| github_jupyter |
# Under the Hood
*Modeling and Simulation in Python*
Copyright 2021 Allen Downey
License: [Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International](https://creativecommons.org/licenses/by-nc-sa/4.0/)
```
# download modsim.py if necessary
from os.path import basename, exists
def download(url):
filename = basename(url)
if not exists(filename):
from urllib.request import urlretrieve
local, _ = urlretrieve(url, filename)
print('Downloaded ' + local)
download('https://raw.githubusercontent.com/AllenDowney/' +
'ModSimPy/master/modsim.py')
# import functions from modsim
from modsim import *
```
In this chapter we "open the hood," looking more closely at how some of
the tools we have used---`run_solve_ivp`, `root_scalar`, and `maximize_scalar`---work.
Most of the time you don't need to know, which is why I left this chapter for last.
But you might be curious. And if nothing else, I have found that I can remember how to use these tools more easily because I know something about how they work.
## How run_solve_ivp Works
`run_solve_ivp` is a function in the ModSimPy library that checks for common errors in the parameters and then calls `solve_ip`, which is the function in the SciPy library that does the actual work.
By default, `solve_ivp` uses the Dormand-Prince method, which is a kind of Runge-Kutta method. You can read about it at
<https://en.wikipedia.org/wiki/Dormand-Prince_method>, but I'll give you a sense of
it here.
The key idea behind all Runge-Kutta methods is to evaluate the slope function several times at each time step and use a weighted average of the computed slopes to estimate the value at the next time step.
Different methods evaluate the slope function in different places and compute the average with different weights.
So let's see if we can figure out how `solve_ivp` works.
As an example, we'll solve the following differential equation:
$$\frac{dy}{dt}(t) = y \sin t$$
Here's the slope function we'll use:
```
import numpy as np
def slope_func(t, state, system):
y, = state
dydt = y * np.sin(t)
return dydt
```
I'll create a `State` object with the initial state and a `System` object with the end time.
```
init = State(y=1)
system = System(init=init, t_end=3)
```
Now we can call `run_solve_ivp`.
```
results, details = run_solve_ivp(system, slope_func)
details
```
One of the variables in `details` is `nfev`, which stands for "number of function evaluations", that is, the number of times `solve_ivp` called the slope function.
This example took 50 evaluations.
Keep that in mind.
Here are the first few time steps in `results`:
```
results.head()
```
And here is the number of time steps.
```
len(results)
```
`results` contains 101 points that are equally spaced in time.
Now you might wonder, if `solve_ivp` ran the slope function 50 times, how did we get 101 time steps?
To answer that question, we need to know more how the solver works.
There are actually three steps:
1. For each time step, `solve_ivp` evaluates the slope function seven times, with different values of `t` and `y`.
2. Using the results, it computes the best estimate for the value `y` at the next time step.
3. After computing all of the time steps, it uses interpolation to compute equally spaced points that connect the estimates from the previous step.
So we can see what's happening, I will run `run_solve_ivp` with the keyword argument `dense_output=False`, which skips the interpolation step and returns time steps that are not equally spaced (that is, not "dense").
While we're at it, I'll modify the slope function so that every time it runs, it adds the values of `t`, `y`, and `dydt` to a list called `evals`.
```
def slope_func(t, state, system):
y, = state
dydt = y * np.sin(t)
evals.append((t, y, dydt))
return dydt
```
Now, before we call `run_solve_ivp` again, I'll initialize `evals` with an empty list.
```
evals = []
results2, details = run_solve_ivp(system, slope_func, dense_output=False)
```
Here are the results:
```
results2
```
It turns out there are only eight time steps, and the first five of them only cover 0.11 seconds.
The time steps are not equal because the Dormand-Prince method is *adaptive*.
At each time step, it actually computes two estimates of the next
value. By comparing them, it can estimate the magnitude of the error,
which it uses to adjust the time step. If the error is too big, it uses
a smaller time step; if the error is small enough, it uses a bigger time
step. By adjusting the time step in this way, it minimizes the number
of times it calls the slope function to achieve a given level of
accuracy.
Because we saved the values of `y` and `t`, we can plot the locations where the slope function was evaluated.
I'll need to use a couple of features we have not seen before, if you don't mind.
First we'll unpack the values from `evals` using `np.transpose`.
Then we can use trigonometry to convert the slope, `dydt`, to components called `u` and `v`.
```
t, y, slope = np.transpose(evals)
theta = np.arctan(slope)
u = np.cos(theta)
v = np.sin(theta)
```
Using these values, we can generate a *quiver plot* that shows an arrow for each time the slope function ran.
The location of the each arrow represents the values of `t` and `y`; the orientation of the arrow shows the slope that was computed.
```
import matplotlib.pyplot as plt
plt.quiver(t, y, u, v, pivot='middle',
color='C1', alpha=0.4, label='evaluation points')
results2['y'].plot(style='o', color='C0', label='solution points')
results['y'].plot(lw=1, label='interpolation')
decorate(xlabel='Time (t)',
ylabel='Quantity (y)')
```
In this figure, the arrows show where the slope function was executed;
the dots show the best estimate of `y` for each time step; and the line shows the interpolation that connects the estimates.
Notice that many of the arrows do not fall on the line; `solve_ivp` evaluated the slope function at these locations in order to compute the solution, but as it turned out, they are not part of the solution.
This is good to know when you are writing a slope function; you should not assume that the time and state you get as input variables are correct.
## How root_scalar Works
`root_scalar` in the ModSim library is a wrapper for a function in the SciPy library with the same name.
Like `run_solve_ivp`, it checks for common errors and changes some of the parameters in a way that makes the SciPy function easier to use (I hope).
According to the documentation, `root_scalar` uses "a combination of bisection, secant, and inverse quadratic interpolation methods." (See
<https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.root_scalar.html>)
To understand what that means, suppose we're trying to find a root of a
function of one variable, $f(x)$, and assume we have evaluated the
function at two places, $x_1$ and $x_2$, and found that the results have
opposite signs. Specifically, assume $f(x_1) > 0$ and $f(x_2) < 0$, as
shown in the following diagram:

If $f$ is a continuous function, there must be at least one root in this
interval. In this case we would say that $x_1$ and $x_2$ *bracket* a
root.
If this were all you knew about $f$, where would you go looking for a
root? If you said "halfway between $x_1$ and $x_2$," congratulations!
`You just invented a numerical method called *bisection*!
If you said, "I would connect the dots with a straight line and compute
the zero of the line," congratulations! You just invented the *secant
method*!
And if you said, "I would evaluate $f$ at a third point, find the
parabola that passes through all three points, and compute the zeros of
the parabola," congratulations, you just invented *inverse quadratic
interpolation*!
That's most of how `root_scalar` works. The details of how these methods are
combined are interesting, but beyond the scope of this book. You can
read more at <https://en.wikipedia.org/wiki/Brents_method>.
## How maximize_scalar Works
`maximize_scalar` in the ModSim library is a wrapper for a function in the SciPy library called `minimize_scalar`.
You can read about it at <https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.minimize_scalar.html>.
By default, it uses Brent's method, which is related to the method I described in the previous section for root-finding.
Brent's method for finding a maximum or minimum is based on a simpler algorithm:
the *golden-section search*, which I will explain.
Suppose we're trying to find the minimum of a function of a single variable, $f(x)$.
As a starting place, assume that we have evaluated the function at three
places, $x_1$, $x_2$, and $x_3$, and found that $x_2$ yields the lowest
value. The following diagram shows this initial state.

We will assume that $f(x)$ is continuous and *unimodal* in this range,
which means that there is exactly one minimum between $x_1$ and $x_3$.
The next step is to choose a fourth point, $x_4$, and evaluate $f(x_4)$.
There are two possible outcomes, depending on whether $f(x_4)$ is
greater than $f(x_2)$ or not.
The following figure shows the two possible states.

If $f(x_4)$ is less than $f(x_2)$ (shown on the left), the minimum must
be between $x_2$ and $x_3$, so we would discard $x_1$ and proceed with
the new bracket $(x_2, x_4, x_3)$.
If $f(x_4)$ is greater than $f(x_2)$ (shown on the right), the local
minimum must be between $x_1$ and $x_4$, so we would discard $x_3$ and
proceed with the new bracket $(x_1, x_2, x_4)$.
Either way, the range gets smaller and our estimate of the optimal value
of $x$ gets better.
This method works for almost any value of $x_4$, but some choices are
better than others. You might be tempted to bisect the interval between
$x_2$ and $x_3$, but that turns out not to be optimal. You can
read about a better option at <https://greenteapress.com/matlab/golden>.
## Chapter Review
The information in this chapter is not strictly necessary; you can use
these methods without knowing much about how they work. But there are
two reasons you might want to know.
One reason is pure curiosity. If you use these methods, and especially
if you come to rely on them, you might find it unsatisfying to treat
them as "black boxes." At the risk of mixing metaphors, I hope you
enjoyed opening the hood.
The other reason is that these methods are not infallible; sometimes
things go wrong. If you know how they work, at least in a general sense,
you might find it easier to debug them.
With that, you have reached the end of the book, so congratulations! I
hope you enjoyed it and learned a lot. I think the tools in this book
are useful, and the ways of thinking are important, not just in
engineering and science, but in practically every field of inquiry.
Models are the tools we use to understand the world: if you build good
models, you are more likely to get things right. Good luck!
| github_jupyter |
# What you will learn from this notebook
This notebook is supposed to demonstrate a simplified version of an actual analysis you might want to run. In the real world steps would be probably the same but the dataset itself would be much, much noisier (meaning it would take some effort to put it into the required shape) and much bigger (I mean, nowadays in the industry we are dealing with more than ~30 samples!).
```
# general packages
import pandas as pd
import numpy as np
# specialized stats packages
from lifelines import KaplanMeierFitter
# plotting
import matplotlib.pyplot as plt
import seaborn as sns
# preferences
%matplotlib inline
import warnings
warnings.filterwarnings('ignore')
```
# Data
I will use one of default datasets from lifetimes library. I don't know much about it and would prefer to avoid jumping to conclusions so I will pretend this data comes actually from a survey among 26 vampires collected a 100 years ago.
In that survey scientists collected information about how many years ago the vampire became un-dead (in other words was bitten by another vampire and turned into one), how old they were at the time of their transformation, whether they identified themselves as binary or non-binary and whether they have experienced depression symptoms yet.
```
# data
from lifelines.datasets import load_psychiatric_patients
df = load_psychiatric_patients()
df.head()
```
Alright, so we have vampires at different age when they tranformed (`Age` column), they reported how many years have passed since transformation (`T` column), whether they have experienced depression symptoms (`C` column) and what gender they identify with (`sex` column, I'm gonna assume `1` is binary and `2` is non-binary because why not).
# Plotting lifetimes and very basic data exploration
There aren't many variables to work with and I will first show you how to plot lifetimes (assuming *now* is at 25, change `current_time` to see how the plot changes):
```
current_time = 25
observed_lifetimes = df['T'].values
observed = observed_lifetimes < current_time
# I'm using slightly modified function from lifetimes library. See the end of this notebook for details.
# If you are running this notebook yourself first execute the cell with function definition at the bottom
# of this notebook
plot_lifetimes(observed_lifetimes, event_observed=observed, block=True)
```
Next I will see whether experiencing depression symptoms is somehow related to age at which the transformation into a vampire took place:
```
sns.catplot(x="C", y="Age", kind="boxen",
data=df.sort_values("C"));
plt.xlabel('Vampire experienced depression or not', size=18)
plt.ylabel('Age', size=18)
plt.title('Vampire survival as a function of age', size=18);
```
Looks like it does! Appears that vampires who have experienced depressive symptoms were on average older when they were bitten and consequently turned into vampires. This is very interesting! Let's look at Kaplan-Meier curves, and hazard curves to check whether gender has anything to do with depressive symptoms.
# Kaplan-Meier curve
```
kmf = KaplanMeierFitter()
T = df["T"] # time since vampire transformation
C = df["C"] # whether they experienced depression symptoms
kmf.fit(T,C);
kmf.survival_function_
kmf.median_
kmf.plot(figsize=[10,6])
```
## Kaplan-Meier curve plotted separately for vampires who define themselves as binary and non-binary
```
# plot both genders on the same plot
plt.figure(figsize=[10,6])
groups = df['sex']
ix = (groups == 1)
kmf.fit(T[~ix], C[~ix], label='binary vampires')
ax = kmf.plot(figsize=[10,10]);
kmf.fit(T[ix], C[ix], label='non-binary vampires')
kmf.plot(ax=ax);
```
Our sample size is small so error bars are relatively large. It looks like in the early years after vampire tranformation more binary (blue line) than non-binary (orange line) vampires experienced depressive symptoms. Maybe non-binary vampires were in a honeymoon stage with vampirism? However, the error bars are pretty much overlapping starting at 20 years past transformation so likely the differences are not statistically significant. But let's look at the hazard rate first.
# Hazard rate using Nelson-Aalen estimator
```
from lifelines import NelsonAalenFitter
naf = NelsonAalenFitter()
naf.fit(T,event_observed=C);
naf.plot(figsize=[10,6]);
naf.fit(T[~ix], C[~ix], label='binary vampires')
ax = naf.plot(figsize=[10,10])
naf.fit(T[ix], C[ix], label='non-binary vampires')
naf.plot(ax=ax);
```
Okay, so it looks like hazard rate increases with time for both groups which we could already deduce from survival curves. Interestingly, it seems that the hazard rate for non-binary vampires increases rapidly around 35 years compared to previous period (I'm ignoring error bars for the moment).
# Statistical analysis of differences
Is there a difference between hazard rate for binary and non-binary vampires? Let's run a log rank test. It will look at random combinations of samples from the two distributions and calculate how many times one had a higher value than the other.
A very important point to remember is that this analysis will not tell us anything about the hazard rates themselves but rather whether one is different from the other - so it signals only relative differences.
```
from lifelines.statistics import logrank_test
results = logrank_test(T[ix], T[~ix], event_observed_A=C[ix], event_observed_B=C[~ix])
results.print_summary()
```
Looks like indeed there are no significant differences between binary and non-binary vampires but for the sake of exercise let's see how to get from the test statistic to difference in hazard rate:
$$ log{\lambda} = Z \sqrt{ \frac{4}{D} } $$
```
Z = results.test_statistic
D = C.count()
log_lambda = Z * np.sqrt (D / 4)
log_lambda
```
Okay, so if the test was significant we could conclude that the hazard rate for binary versus non-binary vampires is roughly 4 times higher which means they are more likely to suffer from depressive symptoms
## What factors influence vampire's survival? Cox Proportional Hazards Model
Alright, and lets say now we want to look at how age and gender identity shape vampire's future. We want to train the model on one set of samples and then use it to predict relative hazard increases (it's always relative to other vampires, never absolute hazard!) during vampire's lifetime.
```
from lifelines import CoxPHFitter
cph = CoxPHFitter()
cph.fit(df, duration_col='T', event_col='C', show_progress=True)
cph.print_summary()
```
It looks like age is significantly related to the occurence of depressive symptoms, just like our EDA indicated at the beginning. If we had some new data we could use the beta values calculated in by fitting method in the previous step to predict relative changes in hazard rates of new vampires (using `cph.predict_cumulative_hazard(new_df)`. This is a semi-parametric model which means that it assumes the same constant rate of change during lifetime for all vampires. There are also models which take into account time covariates but they are beyond the scope of this short notebook. Thanks for reading and good luck with your own explorations!
## Helper function
```
# the function below is a modified version of plotting function from the lifetimes library. All credit should go to
# them and all faults are mine.
def plot_lifetimes(lifetimes, event_observed=None, birthtimes=None,
order=False, block=True):
"""
Parameters:
lifetimes: an (n,) numpy array of lifetimes.
event_observed: an (n,) numpy array of booleans: True if event observed, else False.
birthtimes: an (n,) numpy array offsetting the births away from t=0.
Creates a lifetime plot, see
examples:
"""
from matplotlib import pyplot as plt
N = lifetimes.shape[0]
if N > 100:
print("warning: you may want to subsample to less than 100 individuals.")
if event_observed is None:
event_observed = np.ones(N, dtype=bool)
if birthtimes is None:
birthtimes = np.zeros(N)
if order:
"""order by length of lifetimes; probably not very informative."""
ix = np.argsort(lifetimes, 0)
lifetimes = lifetimes[ix, 0]
event_observed = event_observed[ix, 0]
birthtimes = birthtimes[ix]
fig, ax = plt.subplots(figsize=[15,5], frameon=False)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
for i in range(N):
c = "#663366" if event_observed[i] else "green"
l = 'burned by the sun rays or an angry mob' if event_observed[i] else "alive"
plt.hlines(N - 1 - i, birthtimes[i], birthtimes[i] + lifetimes[i], color=c, lw=3, label=l if (i == 0) or (i==40) else "")
m = "|" if not event_observed[i] else 'o'
plt.scatter((birthtimes[i]) + lifetimes[i], N - 1 - i, color=c, s=30, marker=m)
plt.legend(fontsize=16)
plt.xlabel("Number of years since becoming a vampire", size=18)
plt.ylabel("Individual vampires", size=20)
plt.vlines(current_time, 0, N, lw=2, linestyles='--', alpha=0.5)
plt.xticks(fontsize=18)
plt.ylim(-0.5, N)
return
```
| github_jupyter |
<br><br><br><br><br>
# Awkward datasets
<br><br><br><br><br>
<br><br><br><br><br>
It's not uncommon for data to be non-rectangular. Jagged ("ragged") arrays, cross-references, trees, and graphs are frequently encountered, but difficult to cast as Numpy arrays or Pandas DataFrames.
<br>
**Let's start with NASA's exoplanet database:** each star can have an arbitrary number of planets (jagged array).
<br><br><br><br><br>
```
import pandas
# NASA provides this dataset as a CSV file, which suggests a rectangular table: one row per planet.
exoplanets = pandas.read_csv("data/nasa-exoplanets.csv")
exoplanets
# Quite a few planets in this table have the same star ("host") name.
numplanets = exoplanets.groupby("pl_hostname").size()
numplanets[numplanets > 1]
# Use Pandas's MultiIndex to represent a sparse, 2D index (stars × planets without missing values).
exoplanets.index = pandas.MultiIndex.from_arrays([exoplanets["pl_hostname"], exoplanets["pl_letter"]])
exoplanets.index.names = ["star", "planet"]
exoplanets
# Simplify the table to show 5 star attributes and 5 planet attributes. Star attributes are repeated.
df = exoplanets[["ra", "dec", "st_dist", "st_mass", "st_rad", "pl_orbsmax", "pl_orbeccen", "pl_orbper", "pl_bmassj", "pl_radj"]]
df.columns = pandas.MultiIndex.from_arrays([["star"] * 5 + ["planet"] * 5,
["right asc. (deg)", "declination (deg)", "distance (pc)", "mass (solar)", "radius (solar)", "orbit (AU)", "eccen.", "period (days)", "mass (Jupiter)", "radius (Jupiter)"]])
df
# DataFrame.unstack moves the sparse planet index into a dense set of columns.
# Every column (reduced to 2: orbit and mass) is duplicated 8 times because one star has 8 planets.
df[[("planet", "orbit (AU)"), ("planet", "mass (Jupiter)")]].unstack("planet")
# We can also select a cross-section (xs) of the index by planet letter to focus on one at a time.
df.xs("b", level="planet") # try "c", "d", "e", "f", "g", "h", "i"
```
<br><br><br><br><br>
### Alternative: stars and planets as nested objects
<br><br><br><br><br>
```
# Despite the nice tools Pandas provides, it's easier to think of stars and planets as objects.
stardicts = []
for (starname, planetname), row in df.iterrows():
if len(stardicts) == 0 or stardicts[-1]["name"] != starname:
stardicts.append({"name": starname,
"ra": row["star", "right asc. (deg)"],
"dec": row["star", "declination (deg)"],
"dist": row["star", "distance (pc)"],
"mass": row["star", "mass (solar)"],
"radius": row["star", "radius (solar)"],
"planets": []})
stardicts[-1]["planets"].append({"name": planetname,
"orbit": row["planet", "orbit (AU)"],
"eccen": row["planet", "eccen."],
"period": row["planet", "period (days)"],
"mass": row["planet", "mass (Jupiter)"],
"radius": row["planet", "radius (Jupiter)"]})
stardicts[:30]
# But this destroys Numpy's array-at-a-time performance and (in some cases) convenience.
# Here's a way to get both (disclosure: I'm the author).
import awkward
stars = awkward.fromiter(stardicts)
stars
# The data are logically a collection of nested lists and dicts...
stars[:30].tolist()
# ...but they have been entirely converted into arrays.
for starattr in "name", "ra", "dec", "dist", "mass", "radius":
print("{:15s} =".format("stars[{!r:}]".format(starattr)), stars[starattr])
print()
for planetattr in "name", "orbit", "eccen", "period", "mass", "radius":
print("{:26s} =".format("stars['planets'][{!r:}]".format(planetattr)), stars["planets"][planetattr])
# The object structure is a façade, built on Numpy arrays.
planet_masses = stars["planets"]["mass"]
# It appears to be a list of lists;
print("\nplanet_masses =", planet_masses)
# but it is a JaggedArray class instance;
print("\ntype(planet_masses) =", type(planet_masses))
# whose numerical data are in a content array;
print("\nplanet_masses.content =", planet_masses.content)
# and divisions between stars are encoded in an offsets array.
print("\nplanet_masses.offsets =", planet_masses.offsets)
# Pandas's unstack becomes...
stars["planets"][["orbit", "mass"]].pad(8).tolist()
# ...which can be used to produce regular Numpy arrays.
maxplanets = stars["planets"].counts.max()
stars["planets"]["mass"].pad(maxplanets).fillna(float("nan")).regular()
# Pandas's cross-section becomes...
stars["planets"][:, 0].tolist()
# ...though the first dimension must be selected for >= n subelements to ask for the nth subelement.
print("stars['planets'].counts =", stars["planets"].counts)
atleast3 = (stars["planets"].counts >= 3)
print("atleast3 =", atleast3)
stars["planets"][atleast3, 2].tolist()
# Motivated by particle physics analyses, which have particularly complex events.
import uproot
# Open a simplified file (for tutorials).
lhc_data = uproot.open("data/HZZ.root")["events"]
# Read columns of data for particle energies.
particle_energies = lhc_data.arrays(["*_E"], namedecode="utf-8")
# There's a different number of particles for each particle type in each event.
for name, array in particle_energies.items():
print("\nparticle_energies['{}'] = {}".format(name, array))
```
<br><br>
### Overview of Awkward Arrays
Awkward Array (`import awkward`) has been designed to resemble a generalization of Numpy to
* jagged arrays
* non-rectangular tables
* nullable types
* heterogeneous lists
* cross-references and cyclic references
* non-contiguous arrays
* virtual data and objects
<br><br>
```
# Generate simple data or convert from JSON using fromiter.
a = awkward.fromiter([[1.1, 2.2, 3.3], [], [4.4, 5.5]])
# Columnar structure is built into the resulting object.
print("\na =", a)
print("\ntype(a) =", type(a))
print("\na.content =", a.content)
print("\na.offsets =", a.offsets)
# Numpy ufuncs pass through the structure for array-at-a-time calculations.
# (Uses the same __array_ufunc__ trick as CuPy and Dask...)
import numpy
a = awkward.fromiter([[1.1, 2.2, 3.3], [], [4.4, 5.5]])
print(numpy.sqrt(a))
# Array-at-a-time calculations are only possible if all arguments have the same structure.
a = awkward.fromiter([[1.1, 2.2, 3.3], [], [4.4, 5.5]])
b = awkward.fromiter([[100, 200, 300], [], [400, 500]])
print("a + b =", a + b)
# In Numpy, scalars can be "broadcasted" to be used in calculations with arrays.
# Generalizing this, Numpy arrays can be "broadcasted" to fit jagged arrays.
a = awkward.fromiter([[1.1, 2.2, 3.3], [], [4.4, 5.5]])
b = numpy.array([100, 200, 300])
print("a + b =", a + b)
# Slicing works like Numpy.
a = awkward.fromiter([[1.1, 2.2, 3.3, 4.4], [5.5, 6.6], [7.7, 8.8, 9.9]])
# Take the first two outer lists.
print("\na[:2] =", a[:2])
# Take the first two of each inner list.
print("\na[:, :2] =", a[:, :2])
# Masking works like Numpy, but with new capabilities for jagged masks.
a = awkward.fromiter([[ 1.1, 2.2, 3.3, 4.4], [ 5.5, 6.6], [ 7.7, 8.8, 9.9]])
mask = awkward.fromiter([True, False, True])
jaggedmask = awkward.fromiter([[True, False, False, True], [False, True], [False, False, False]])
# Filter outer lists.
print("\na[mask] =", a[mask])
# Filter inner lists.
print("\na[jaggedmask] =", a[jaggedmask])
# Integer indexing works like Numpy, but with new capabilities for jagged indexes.
a = awkward.fromiter([[1.1, 2.2, 3.3, 4.4], [5.5, 6.6], [7.7, 8.8, 9.9]])
index = awkward.fromiter([2, 1, 1, 0])
jaggedindex = awkward.fromiter([[3, 0, 0, 1, 2], [], [-1]])
# Apply an integer function to outer lists.
print("\na[index] =", a[index])
# Apply an integer function to inner lists.
print("\na[jaggedindex] =", a[jaggedindex])
# In Numpy, "reducers" turn arrays into scalars.
# Generalizing this, jagged arrays can be "reduced" to Numpy arrays.
a = awkward.fromiter([[1.1, 2.2, 3.3], [], [4.4, 5.5]])
print("\na.sum() =", a.sum())
print("\na.max() =", a.max())
# Like Numpy, argmax and argmin produce integer indexes appropriate for application to arrays.
a = awkward.fromiter([[1.1, 2.2, 3.3], [], [4.4, 5.5]])
b = awkward.fromiter([[100, 200, 300], [], [400, 500]])
indexes = a.argmax()
print("\nindexes =", indexes)
print("\nb[indexes] =", b[indexes])
# Since we often deal with different numbers of objects in the same event, we need ways to
# match them for comparison.
a = awkward.fromiter([[1.1, 2.2, 3.3], [], [4.4, 5.5]])
b = awkward.fromiter([[10, 20], [30], [40]])
print("\na.cross(b) =", a.cross(b))
print("\na.cross(b).i0 (lefts) =", a.cross(b).i0)
print("\na.cross(b).i1 (rights) =", a.cross(b).i1)
```
<br><br><br><br><br>
### Application to a realistic problem
Based on a typical case in particle physics, but general enough for all sciences.
<br><br><br><br><br>
```
# Suppose we have a variable number of real objects in each event.
import collections
T = collections.namedtuple("T", ["x", "y"])
truth = []
for i in range(10):
truth.append([])
for j in range(numpy.random.poisson(2)):
truth[-1].append(T(*numpy.random.randint(0, 100, 2)/100))
truth
# When we try to reconstruct these objects from the signals they produce,
# the measurements have error, some unlucky objects are lost, and some spurious noise is added.
M = collections.namedtuple("M", ["x", "y"])
error = lambda: numpy.random.normal(0, 0.001)
unlucky = lambda: numpy.random.uniform(0, 1) < 0.2
observed = []
for event in truth:
observed.append([M(x + error(), y + error()) for x, y in event if not unlucky()])
for j in range(numpy.random.poisson(0.25)):
observed[-1].append(M(*numpy.random.normal(0.5, 0.25, 2)))
observed
# So the simulated data look like this:
data = awkward.Table(truth=awkward.fromiter(truth), observed=awkward.fromiter(observed))
data.tolist()
# The measured objects were reconstructed from raw signals in our simulation by a complex process.
# We want to match real and measured to learn what the simulation is telling us about measurement
# errors, missing fraction, and spurious fraction.
pairs = data["truth"].cross(data["observed"], nested=True) # pairs for all combinations
distances = numpy.sqrt((pairs.i0["x"] - pairs.i1["x"])**2 + # compute distance for all
(pairs.i0["y"] - pairs.i1["y"])**2)
best = distances.argmin() # pick smallest distance
print("\nbest =", best)
good_enough = (distances[best] < 0.005) # exclude if the distance is too large
print("\ngood_enough =", good_enough)
good_pairs = pairs[best][good_enough].flatten(axis=1) # select best and good enough; reduce
print("\ngood_pairs[0] =", good_pairs[0])
```
#### **Explode:** create deeper structures by combining the ones we have
<center><img src="img/explode.png" width="25%"></center>
#### **Flat:** compute something in a vectorized way
<center><img src="img/flat.png" width="25%"></center>
#### **Reduce:** use the new values to eliminate structure (max, sum, mean...)
<center><img src="img/reduce.png" width="25%"></center>
```
# Other awkward types: nullable, heterogeneous lists, nested records...
a = awkward.fromiter([[1.1, 2.2, None, 3.3, None],
[4.4, [5.5]],
[{"x": 6, "y": {"z": 7}}, None, {"x": 8, "y": {"z": 9}}]
])
# Array type as a function signature
print(a.type)
print()
# Vectorized operations all the way down
(a + 100).tolist()
# Cross-references
data = awkward.fromiter([
{"tracks": [{"phi": 1.0}, {"phi": 2.0}],
"hits": [{"id": 100, "pos": 3.7}, {"id": 50, "pos": 2.1}, {"id": 75, "pos": 2.5}]},
{"tracks": [{"phi": 1.5}],
"hits": [{"id": 100, "pos": 1.4}, {"id": 50, "pos": 0.7}, {"id": 75, "pos": 3.0}]}])
data["tracks"]["hits-on-track"] = \
awkward.JaggedArray.fromcounts([2, 1],
awkward.JaggedArray.fromcounts([2, 2, 1, 1],
awkward.IndexedArray([0, 1, 1, 2, 3, 5],
data["hits"].content)))
data.tolist()
# Cyclic references
tree = awkward.fromiter([
{"value": 1.23, "left": 1, "right": 2}, # node 0
{"value": 3.21, "left": 3, "right": 4}, # node 1
{"value": 9.99, "left": 5, "right": 6}, # node 2
{"value": 3.14, "left": 7, "right": None}, # node 3
{"value": 2.71, "left": None, "right": 8}, # node 4
{"value": 5.55, "left": None, "right": None}, # node 5
{"value": 8.00, "left": None, "right": None}, # node 6
{"value": 9.00, "left": None, "right": None}, # node 7
{"value": 0.00, "left": None, "right": None}, # node 8
])
left = tree.contents["left"].content
right = tree.contents["right"].content
left[(left < 0) | (left > 8)] = 0 # satisfy overzealous validity checks
right[(right < 0) | (right > 8)] = 0
tree.contents["left"].content = awkward.IndexedArray(left, tree)
tree.contents["right"].content = awkward.IndexedArray(right, tree)
tree[0].tolist()
```
| Array type | Purpose | Members | Usage |
|:-----------|:--------|:--------|:------|
| JaggedArray | variable-sized data structures | starts, stops, content | ubiquitous |
| Table | struct-like objects in columns | contents _(dict)_ | ubiquitous |
| ObjectArray | arbitrary Python types on demand | generator, content | common |
| Methods | mix-in methods and properties on any array type | _(none)_ | common |
| MaskedArray | allow nullable values (`None`) | mask _(bytes)_, content | occasional |
| BitMaskedArray | same, but with a bit-mask | mask _(bits)_, content | from Arrow |
| IndexedMaskedArray | same, but with dense content | mask-index _(integers)_ content | rare |
| IndexedArray | lazy integer indexing: "pointers" | index, content | rare |
| SparseArray | huge array defined at a few indexes | index, content, default | rare |
| UnionArray | heterogeneous types or data sources | tags, index, contents _(list)_ | rare |
| StringArray | special case: jagged array of characters | starts, stops, content, string methods | common |
| ChunkedArray | discontiguous array presented as a whole | counts, chunks _(lists)_ | from Parquet |
| AppendableArray | chunked allocation for efficient appending | counts, chunks _(lists)_ | rare |
| VirtualArray | array generated from a function when needed | generator, possible cached array | from Parquet |
| github_jupyter |
# VacationPy
----
#### Note
* Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
```
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import gmaps
import os
# Import API key
from api_keys import g_key
```
### Store Part I results into DataFrame
* Load the csv exported in Part I to a DataFrame
```
#load cvs file and exclude na
weather = pd.read_csv("../output_data/cities.csv")
weather_df=weather.dropna()
weather_df
```
### Humidity Heatmap
* Configure gmaps.
* Use the Lat and Lng as locations and Humidity as the weight.
* Add Heatmap layer to map.
```
#gmaps access
gmaps.configure(g_key)
#Lat and Lng as locations and Humidity as the weight
locations = weather_df[["Lat", "Lng"]].astype(float)
humidity = weather_df["Humidity"].astype(float)
fig = gmaps.figure()
heat_layer = gmaps.heatmap_layer(locations, weights=humidity,
dissipating=False, max_intensity=100,
point_radius = 2)
fig.add_layer(heat_layer)
plt.savefig("vacationheatmap.png")
fig
```
### Create new DataFrame fitting weather criteria
* Narrow down the cities to fit weather conditions.
* Drop any rows will null values.
```
# A max temperature lower than 80 degrees but higher than 70.
max_temp = weather_df.loc[(weather_df["Max Temp"]>70) & (weather_df["Max Temp"]<80),:]
#Wind speed less than 10 mph.
wind_speed = max_temp.loc[max_temp["Wind Speed"]<10,:]
# Zero cloudiness.
final_condition = wind_speed.loc[wind_speed["Cloudiness"]==0,:]
# Drop any rows that don't contain all three conditions. You want to be sure the weather is ideal.
final_condition.dropna(inplace=True)
final_condition
```
### Hotel Map
* Store into variable named `hotel_df`.
* Add a "Hotel Name" column to the DataFrame.
* Set parameters to search for hotels with 5000 meters.
* Hit the Google Places API for each city's coordinates.
* Store the first Hotel result into the DataFrame.
* Plot markers on top of the heatmap.
```
# Store filtered data to hotel_df and add "Hotel Name"
final_hotel_df = final_condition
final_hotel_df['Hotel Name'] = " "
final_hotel_df.head()
#Set parameters to search for hotels with 5000 meters
url = "https://maps.googleapis.com/maps/api/place/nearbysearch/json"
params = {
"radius": "5000",
"type": "lodging",
"key": g_key,
}
response = requests.get(url, params=params).json()
#Google Places API for each city's coordinates.
for index, row in hotel_df.iterrows():
# get locations of the city
latitude=row["Lat"]
longitude=row["Lng"]
location=f"{latitude},{longitude}"
city_name = row["City"]
params['location'] = location
response = requests.get(url, params=params).json()
# extract results
results = response['results']
try:
hotel = results[0]["name"]
hotel_df.loc[index, 'Hotel Name'] = hotel
except (KeyError, IndexError):
print("No Data....")
#Store the first Hotel result into the DataFrame
hotel_df
#Plot markers on top of the heatmap
# NOTE: Do not change any of the code in this cell
# Using the template add the hotel marks to the heatmap
info_box_template = """
<dl>
<dt>Name</dt><dd>{Hotel Name}</dd>
<dt>City</dt><dd>{City}</dd>
<dt>Country</dt><dd>{Country}</dd>
</dl>
"""
# Store the DataFrame Row
# NOTE: be sure to update with your DataFrame name
hotel_info = [info_box_template.format(**row) for index, row in hotel_df.iterrows()]
locations = hotel_df[["Lat", "Lng"]]
```
| github_jupyter |
```
import warnings
import numpy as np
import os
import keras
from keras.models import Model
from keras import layers
from keras.layers import Activation
from keras.layers import Dense
from keras.layers import Input
from keras.layers import BatchNormalization
from keras.layers import Conv3D
from keras.layers import MaxPooling3D
from keras.layers import AveragePooling3D
from keras.layers import Dropout
from keras.layers import Reshape
from keras.layers import Lambda
from keras.layers import GlobalAveragePooling3D,Flatten
from keras.models import Sequential
from keras.engine.topology import get_source_inputs
from keras.utils import layer_utils
from keras.utils.data_utils import get_file
from keras import backend as K
import tensorflow as tf
import matplotlib.pyplot as plt
# 그림 그리기
def plot_training_loss(H):
plt.style.use("ggplot")
plt.figure()
plt.plot(H.history["loss"], label="train_loss")
plt.plot(H.history["val_loss"], label="val_loss")
plt.title("Training Loss")
plt.xlabel("Epoch #")
plt.ylabel("Loss")
plt.legend(loc="lower left")
# 그림 그리기
def plot_training_acc(H):
plt.style.use("ggplot")
plt.figure()
plt.plot(H.history["accuracy"], label="train_acc")
plt.plot(H.history["val_accuracy"], label="val_acc")
plt.title("Training acc")
plt.xlabel("Epoch #")
plt.ylabel("acc")
plt.legend(loc="lower left")
```
# Pre trained I3D
```
WEIGHTS_NAME = ['rgb_kinetics_only', 'flow_kinetics_only', 'rgb_imagenet_and_kinetics', 'flow_imagenet_and_kinetics']
# path to pretrained models with top (classification layer)
WEIGHTS_PATH = {
'rgb_kinetics_only' : 'https://github.com/dlpbc/keras-kinetics-i3d/releases/download/v0.2/rgb_inception_i3d_kinetics_only_tf_dim_ordering_tf_kernels.h5',
'flow_kinetics_only' : 'https://github.com/dlpbc/keras-kinetics-i3d/releases/download/v0.2/flow_inception_i3d_kinetics_only_tf_dim_ordering_tf_kernels.h5',
'rgb_imagenet_and_kinetics' : 'https://github.com/dlpbc/keras-kinetics-i3d/releases/download/v0.2/rgb_inception_i3d_imagenet_and_kinetics_tf_dim_ordering_tf_kernels.h5',
'flow_imagenet_and_kinetics' : 'https://github.com/dlpbc/keras-kinetics-i3d/releases/download/v0.2/flow_inception_i3d_imagenet_and_kinetics_tf_dim_ordering_tf_kernels.h5'
}
# path to pretrained models with no top (no classification layer)
WEIGHTS_PATH_NO_TOP = {
'rgb_kinetics_only' : 'https://github.com/dlpbc/keras-kinetics-i3d/releases/download/v0.2/rgb_inception_i3d_kinetics_only_tf_dim_ordering_tf_kernels_no_top.h5',
'flow_kinetics_only' : 'https://github.com/dlpbc/keras-kinetics-i3d/releases/download/v0.2/flow_inception_i3d_kinetics_only_tf_dim_ordering_tf_kernels_no_top.h5',
'rgb_imagenet_and_kinetics' : 'https://github.com/dlpbc/keras-kinetics-i3d/releases/download/v0.2/rgb_inception_i3d_imagenet_and_kinetics_tf_dim_ordering_tf_kernels_no_top.h5',
'flow_imagenet_and_kinetics' : 'https://github.com/dlpbc/keras-kinetics-i3d/releases/download/v0.2/flow_inception_i3d_imagenet_and_kinetics_tf_dim_ordering_tf_kernels_no_top.h5'
}
def _obtain_input_shape(input_shape,
default_frame_size,
min_frame_size,
default_num_frames,
min_num_frames,
data_format,
require_flatten,
weights=None):
if weights != 'kinetics_only' and weights != 'imagenet_and_kinetics' and input_shape and len(input_shape) == 4:
if data_format == 'channels_first':
if input_shape[0] not in {1, 3}:
warnings.warn(
'This model usually expects 1 or 3 input channels. '
'However, it was passed an input_shape with ' +
str(input_shape[0]) + ' input channels.')
default_shape = (input_shape[0], default_num_frames, default_frame_size, default_frame_size)
else:
if input_shape[-1] not in {1, 3}:
warnings.warn(
'This model usually expects 1 or 3 input channels. '
'However, it was passed an input_shape with ' +
str(input_shape[-1]) + ' input channels.')
default_shape = (default_num_frames, default_frame_size, default_frame_size, input_shape[-1])
else:
if data_format == 'channels_first':
default_shape = (3, default_num_frames, default_frame_size, default_frame_size)
else:
default_shape = (default_num_frames, default_frame_size, default_frame_size, 3)
if (weights == 'kinetics_only' or weights == 'imagenet_and_kinetics') and require_flatten:
if input_shape is not None:
if input_shape != default_shape:
raise ValueError('When setting`include_top=True` '
'and loading `imagenet` weights, '
'`input_shape` should be ' +
str(default_shape) + '.')
return default_shape
if input_shape:
if data_format == 'channels_first':
if input_shape is not None:
if len(input_shape) != 4:
raise ValueError(
'`input_shape` must be a tuple of four integers.')
if input_shape[0] != 3 and (weights == 'kinetics_only' or weights == 'imagenet_and_kinetics'):
raise ValueError('The input must have 3 channels; got '
'`input_shape=' + str(input_shape) + '`')
if input_shape[1] is not None and input_shape[1] < min_num_frames:
raise ValueError('Input number of frames must be at least ' +
str(min_num_frames) + '; got '
'`input_shape=' + str(input_shape) + '`')
if ((input_shape[2] is not None and input_shape[2] < min_frame_size) or
(input_shape[3] is not None and input_shape[3] < min_frame_size)):
raise ValueError('Input size must be at least ' +
str(min_frame_size) + 'x' + str(min_frame_size) + '; got '
'`input_shape=' + str(input_shape) + '`')
else:
if input_shape is not None:
if len(input_shape) != 4:
raise ValueError(
'`input_shape` must be a tuple of four integers.')
if input_shape[-1] != 3 and (weights == 'kinetics_only' or weights == 'imagenet_and_kinetics'):
raise ValueError('The input must have 3 channels; got '
'`input_shape=' + str(input_shape) + '`')
if input_shape[0] is not None and input_shape[0] < min_num_frames:
raise ValueError('Input number of frames must be at least ' +
str(min_num_frames) + '; got '
'`input_shape=' + str(input_shape) + '`')
if ((input_shape[1] is not None and input_shape[1] < min_frame_size) or
(input_shape[2] is not None and input_shape[2] < min_frame_size)):
raise ValueError('Input size must be at least ' +
str(min_frame_size) + 'x' + str(min_frame_size) + '; got '
'`input_shape=' + str(input_shape) + '`')
else:
if require_flatten:
input_shape = default_shape
else:
if data_format == 'channels_first':
input_shape = (3, None, None, None)
else:
input_shape = (None, None, None, 3)
if require_flatten:
if None in input_shape:
raise ValueError('If `include_top` is True, '
'you should specify a static `input_shape`. '
'Got `input_shape=' + str(input_shape) + '`')
return input_shape
def conv3d_bn(x,
filters,
num_frames,
num_row,
num_col,
padding='same',
strides=(1, 1, 1),
use_bias = False,
use_activation_fn = True,
use_bn = True,
name=None):
if name is not None:
bn_name = name + '_bn'
conv_name = name + '_conv'
else:
bn_name = None
conv_name = None
x = Conv3D(
filters, (num_frames, num_row, num_col),
strides=strides,
padding=padding,
use_bias=use_bias,
name=conv_name)(x)
if use_bn:
if K.image_data_format() == 'channels_first':
bn_axis = 1
else:
bn_axis = 4
x = BatchNormalization(axis=bn_axis, scale=False, name=bn_name)(x)
if use_activation_fn:
x = Activation('relu', name=name)(x)
return x
def Inception_Inflated3d(include_top=True,
weights=None,
input_tensor=None,
input_shape=None,
dropout_prob=0.0,
endpoint_logit=True,
classes=400):
if not (weights in WEIGHTS_NAME or weights is None or os.path.exists(weights)):
raise ValueError('The `weights` argument should be either '
'`None` (random initialization) or %s' %
str(WEIGHTS_NAME) + ' '
'or a valid path to a file containing `weights` values')
if weights in WEIGHTS_NAME and include_top and classes != 400:
raise ValueError('If using `weights` as one of these %s, with `include_top`'
' as true, `classes` should be 400' % str(WEIGHTS_NAME))
# Determine proper input shape
input_shape = _obtain_input_shape(
input_shape,
default_frame_size=224,
min_frame_size=80,
default_num_frames=32,
min_num_frames=32,
data_format=K.image_data_format(),
require_flatten=include_top,
weights=weights)
if input_tensor is None:
img_input = Input(shape=input_shape)
else:
if not K.is_keras_tensor(input_tensor):
img_input = Input(tensor=input_tensor, shape=input_shape)
else:
img_input = input_tensor
if K.image_data_format() == 'channels_first':
channel_axis = 1
else:
channel_axis = 4
# Downsampling via convolution (spatial and temporal)
x = conv3d_bn(img_input, 64, 7, 7, 7, strides=(2, 2, 2), padding='same', name='Conv3d_1a_7x7')
# Downsampling (spatial only)
x = MaxPooling3D((1, 3, 3), strides=(1, 2, 2), padding='same', name='MaxPool2d_2a_3x3')(x)
x = conv3d_bn(x, 64, 1, 1, 1, strides=(1, 1, 1), padding='same', name='Conv3d_2b_1x1')
x = conv3d_bn(x, 192, 3, 3, 3, strides=(1, 1, 1), padding='same', name='Conv3d_2c_3x3')
# Downsampling (spatial only)
x = MaxPooling3D((1, 3, 3), strides=(1, 2, 2), padding='same', name='MaxPool2d_3a_3x3')(x)
# Mixed 3b
branch_0 = conv3d_bn(x, 64, 1, 1, 1, padding='same', name='Conv3d_3b_0a_1x1')
branch_1 = conv3d_bn(x, 96, 1, 1, 1, padding='same', name='Conv3d_3b_1a_1x1')
branch_1 = conv3d_bn(branch_1, 128, 3, 3, 3, padding='same', name='Conv3d_3b_1b_3x3')
branch_2 = conv3d_bn(x, 16, 1, 1, 1, padding='same', name='Conv3d_3b_2a_1x1')
branch_2 = conv3d_bn(branch_2, 32, 3, 3, 3, padding='same', name='Conv3d_3b_2b_3x3')
branch_3 = MaxPooling3D((3, 3, 3), strides=(1, 1, 1), padding='same', name='MaxPool2d_3b_3a_3x3')(x)
branch_3 = conv3d_bn(branch_3, 32, 1, 1, 1, padding='same', name='Conv3d_3b_3b_1x1')
x = layers.concatenate(
[branch_0, branch_1, branch_2, branch_3],
axis=channel_axis,
name='Mixed_3b')
# Mixed 3c
branch_0 = conv3d_bn(x, 128, 1, 1, 1, padding='same', name='Conv3d_3c_0a_1x1')
branch_1 = conv3d_bn(x, 128, 1, 1, 1, padding='same', name='Conv3d_3c_1a_1x1')
branch_1 = conv3d_bn(branch_1, 192, 3, 3, 3, padding='same', name='Conv3d_3c_1b_3x3')
branch_2 = conv3d_bn(x, 32, 1, 1, 1, padding='same', name='Conv3d_3c_2a_1x1')
branch_2 = conv3d_bn(branch_2, 96, 3, 3, 3, padding='same', name='Conv3d_3c_2b_3x3')
branch_3 = MaxPooling3D((3, 3, 3), strides=(1, 1, 1), padding='same', name='MaxPool2d_3c_3a_3x3')(x)
branch_3 = conv3d_bn(branch_3, 64, 1, 1, 1, padding='same', name='Conv3d_3c_3b_1x1')
x = layers.concatenate(
[branch_0, branch_1, branch_2, branch_3],
axis=channel_axis,
name='Mixed_3c')
# Downsampling (spatial and temporal)
x = MaxPooling3D((3, 3, 3), strides=(2, 2, 2), padding='same', name='MaxPool2d_4a_3x3')(x)
# Mixed 4b
branch_0 = conv3d_bn(x, 192, 1, 1, 1, padding='same', name='Conv3d_4b_0a_1x1')
branch_1 = conv3d_bn(x, 96, 1, 1, 1, padding='same', name='Conv3d_4b_1a_1x1')
branch_1 = conv3d_bn(branch_1, 208, 3, 3, 3, padding='same', name='Conv3d_4b_1b_3x3')
branch_2 = conv3d_bn(x, 16, 1, 1, 1, padding='same', name='Conv3d_4b_2a_1x1')
branch_2 = conv3d_bn(branch_2, 48, 3, 3, 3, padding='same', name='Conv3d_4b_2b_3x3')
branch_3 = MaxPooling3D((3, 3, 3), strides=(1, 1, 1), padding='same', name='MaxPool2d_4b_3a_3x3')(x)
branch_3 = conv3d_bn(branch_3, 64, 1, 1, 1, padding='same', name='Conv3d_4b_3b_1x1')
x = layers.concatenate(
[branch_0, branch_1, branch_2, branch_3],
axis=channel_axis,
name='Mixed_4b')
# Mixed 4c
branch_0 = conv3d_bn(x, 160, 1, 1, 1, padding='same', name='Conv3d_4c_0a_1x1')
branch_1 = conv3d_bn(x, 112, 1, 1, 1, padding='same', name='Conv3d_4c_1a_1x1')
branch_1 = conv3d_bn(branch_1, 224, 3, 3, 3, padding='same', name='Conv3d_4c_1b_3x3')
branch_2 = conv3d_bn(x, 24, 1, 1, 1, padding='same', name='Conv3d_4c_2a_1x1')
branch_2 = conv3d_bn(branch_2, 64, 3, 3, 3, padding='same', name='Conv3d_4c_2b_3x3')
branch_3 = MaxPooling3D((3, 3, 3), strides=(1, 1, 1), padding='same', name='MaxPool2d_4c_3a_3x3')(x)
branch_3 = conv3d_bn(branch_3, 64, 1, 1, 1, padding='same', name='Conv3d_4c_3b_1x1')
x = layers.concatenate(
[branch_0, branch_1, branch_2, branch_3],
axis=channel_axis,
name='Mixed_4c')
# Mixed 4d
branch_0 = conv3d_bn(x, 128, 1, 1, 1, padding='same', name='Conv3d_4d_0a_1x1')
branch_1 = conv3d_bn(x, 128, 1, 1, 1, padding='same', name='Conv3d_4d_1a_1x1')
branch_1 = conv3d_bn(branch_1, 256, 3, 3, 3, padding='same', name='Conv3d_4d_1b_3x3')
branch_2 = conv3d_bn(x, 24, 1, 1, 1, padding='same', name='Conv3d_4d_2a_1x1')
branch_2 = conv3d_bn(branch_2, 64, 3, 3, 3, padding='same', name='Conv3d_4d_2b_3x3')
branch_3 = MaxPooling3D((3, 3, 3), strides=(1, 1, 1), padding='same', name='MaxPool2d_4d_3a_3x3')(x)
branch_3 = conv3d_bn(branch_3, 64, 1, 1, 1, padding='same', name='Conv3d_4d_3b_1x1')
x = layers.concatenate(
[branch_0, branch_1, branch_2, branch_3],
axis=channel_axis,
name='Mixed_4d')
# Mixed 4e
branch_0 = conv3d_bn(x, 112, 1, 1, 1, padding='same', name='Conv3d_4e_0a_1x1')
branch_1 = conv3d_bn(x, 144, 1, 1, 1, padding='same', name='Conv3d_4e_1a_1x1')
branch_1 = conv3d_bn(branch_1, 288, 3, 3, 3, padding='same', name='Conv3d_4e_1b_3x3')
branch_2 = conv3d_bn(x, 32, 1, 1, 1, padding='same', name='Conv3d_4e_2a_1x1')
branch_2 = conv3d_bn(branch_2, 64, 3, 3, 3, padding='same', name='Conv3d_4e_2b_3x3')
branch_3 = MaxPooling3D((3, 3, 3), strides=(1, 1, 1), padding='same', name='MaxPool2d_4e_3a_3x3')(x)
branch_3 = conv3d_bn(branch_3, 64, 1, 1, 1, padding='same', name='Conv3d_4e_3b_1x1')
x = layers.concatenate(
[branch_0, branch_1, branch_2, branch_3],
axis=channel_axis,
name='Mixed_4e')
# Mixed 4f
branch_0 = conv3d_bn(x, 256, 1, 1, 1, padding='same', name='Conv3d_4f_0a_1x1')
branch_1 = conv3d_bn(x, 160, 1, 1, 1, padding='same', name='Conv3d_4f_1a_1x1')
branch_1 = conv3d_bn(branch_1, 320, 3, 3, 3, padding='same', name='Conv3d_4f_1b_3x3')
branch_2 = conv3d_bn(x, 32, 1, 1, 1, padding='same', name='Conv3d_4f_2a_1x1')
branch_2 = conv3d_bn(branch_2, 128, 3, 3, 3, padding='same', name='Conv3d_4f_2b_3x3')
branch_3 = MaxPooling3D((3, 3, 3), strides=(1, 1, 1), padding='same', name='MaxPool2d_4f_3a_3x3')(x)
branch_3 = conv3d_bn(branch_3, 128, 1, 1, 1, padding='same', name='Conv3d_4f_3b_1x1')
x = layers.concatenate(
[branch_0, branch_1, branch_2, branch_3],
axis=channel_axis,
name='Mixed_4f')
# Downsampling (spatial and temporal)
x = MaxPooling3D((2, 2, 2), strides=(2, 2, 2), padding='same', name='MaxPool2d_5a_2x2')(x)
# Mixed 5b
branch_0 = conv3d_bn(x, 256, 1, 1, 1, padding='same', name='Conv3d_5b_0a_1x1')
branch_1 = conv3d_bn(x, 160, 1, 1, 1, padding='same', name='Conv3d_5b_1a_1x1')
branch_1 = conv3d_bn(branch_1, 320, 3, 3, 3, padding='same', name='Conv3d_5b_1b_3x3')
branch_2 = conv3d_bn(x, 32, 1, 1, 1, padding='same', name='Conv3d_5b_2a_1x1')
branch_2 = conv3d_bn(branch_2, 128, 3, 3, 3, padding='same', name='Conv3d_5b_2b_3x3')
branch_3 = MaxPooling3D((3, 3, 3), strides=(1, 1, 1), padding='same', name='MaxPool2d_5b_3a_3x3')(x)
branch_3 = conv3d_bn(branch_3, 128, 1, 1, 1, padding='same', name='Conv3d_5b_3b_1x1')
x = layers.concatenate(
[branch_0, branch_1, branch_2, branch_3],
axis=channel_axis,
name='Mixed_5b')
# Mixed 5c
branch_0 = conv3d_bn(x, 384, 1, 1, 1, padding='same', name='Conv3d_5c_0a_1x1')
branch_1 = conv3d_bn(x, 192, 1, 1, 1, padding='same', name='Conv3d_5c_1a_1x1')
branch_1 = conv3d_bn(branch_1, 384, 3, 3, 3, padding='same', name='Conv3d_5c_1b_3x3')
branch_2 = conv3d_bn(x, 48, 1, 1, 1, padding='same', name='Conv3d_5c_2a_1x1')
branch_2 = conv3d_bn(branch_2, 128, 3, 3, 3, padding='same', name='Conv3d_5c_2b_3x3')
branch_3 = MaxPooling3D((3, 3, 3), strides=(1, 1, 1), padding='same', name='MaxPool2d_5c_3a_3x3')(x)
branch_3 = conv3d_bn(branch_3, 128, 1, 1, 1, padding='same', name='Conv3d_5c_3b_1x1')
x = layers.concatenate(
[branch_0, branch_1, branch_2, branch_3],
axis=channel_axis,
name='Mixed_5c')
if include_top:
# Classification block
x = AveragePooling3D((2, 7, 7), strides=(1, 1, 1), padding='valid', name='global_avg_pool')(x)
x = Dropout(dropout_prob)(x)
x = conv3d_bn(x, classes, 1, 1, 1, padding='same',
use_bias=True, use_activation_fn=False, use_bn=False, name='Conv3d_6a_1x1')
num_frames_remaining = int(x.shape[1])
x = Reshape((num_frames_remaining, classes))(x)
# logits (raw scores for each class)
x = Lambda(lambda x: K.mean(x, axis=1, keepdims=False),
output_shape=lambda s: (s[0], s[2]))(x)
if not endpoint_logit:
x = Activation('softmax', name='prediction')(x)
else:
h = int(x.shape[2])
w = int(x.shape[3])
x = AveragePooling3D((2, h, w), strides=(1, 1, 1), padding='valid', name='global_avg_pool')(x)
inputs = img_input
# create model
model = Model(inputs, x, name='i3d_inception')
# load weights
if weights in WEIGHTS_NAME:
if weights == WEIGHTS_NAME[0]: # rgb_kinetics_only
if include_top:
weights_url = WEIGHTS_PATH['rgb_kinetics_only']
model_name = 'i3d_inception_rgb_kinetics_only.h5'
else:
weights_url = WEIGHTS_PATH_NO_TOP['rgb_kinetics_only']
model_name = 'i3d_inception_rgb_kinetics_only_no_top.h5'
elif weights == WEIGHTS_NAME[1]: # flow_kinetics_only
if include_top:
weights_url = WEIGHTS_PATH['flow_kinetics_only']
model_name = 'i3d_inception_flow_kinetics_only.h5'
else:
weights_url = WEIGHTS_PATH_NO_TOP['flow_kinetics_only']
model_name = 'i3d_inception_flow_kinetics_only_no_top.h5'
elif weights == WEIGHTS_NAME[2]: # rgb_imagenet_and_kinetics
if include_top:
weights_url = WEIGHTS_PATH['rgb_imagenet_and_kinetics']
model_name = 'i3d_inception_rgb_imagenet_and_kinetics.h5'
else:
weights_url = WEIGHTS_PATH_NO_TOP['rgb_imagenet_and_kinetics']
model_name = 'i3d_inception_rgb_imagenet_and_kinetics_no_top.h5'
elif weights == WEIGHTS_NAME[3]: # flow_imagenet_and_kinetics
if include_top:
weights_url = WEIGHTS_PATH['flow_imagenet_and_kinetics']
model_name = 'i3d_inception_flow_imagenet_and_kinetics.h5'
else:
weights_url = WEIGHTS_PATH_NO_TOP['flow_imagenet_and_kinetics']
model_name = 'i3d_inception_flow_imagenet_and_kinetics_no_top.h5'
downloaded_weights_path = get_file(model_name, weights_url, cache_subdir='models')
model.load_weights(downloaded_weights_path)
if K.backend() == 'theano':
layer_utils.convert_all_kernels_in_model(model)
if K.image_data_format() == 'channels_first' and K.backend() == 'tensorflow':
warnings.warn('You are using the TensorFlow backend, yet you '
'are using the Theano '
'image data format convention '
'(`image_data_format="channels_first"`). '
'For best performance, set '
'`image_data_format="channels_last"` in '
'your keras config '
'at ~/.keras/keras.json.')
elif weights is not None:
model.load_weights(weights)
return model
model=Inception_Inflated3d(include_top=False,
weights='rgb_imagenet_and_kinetics',
input_tensor=None,
input_shape=(32,224,224,3),
dropout_prob=0.3,
endpoint_logit=True,
classes=6)
model.summary()
trasfer_model = Sequential()
trasfer_model.add(model)
trasfer_model.add(Flatten())
trasfer_model.add(Dropout(0.3))
trasfer_model.add(Dense(6,activation = 'softmax'))
trasfer_model.summary()
import numpy as np
saved_npz_train = np.load('video_dataset_train.npz')
saved_npz_test = np.load('video_dataset_test.npz')
X_train=saved_npz_train['X']
y_train = saved_npz_train['Y']
X_test=saved_npz_test['X']
y_test = saved_npz_test['Y']
saved_npz_train.close()
saved_npz_test.close()
X_train=X_train.astype('float16')
X_train=X_train/255.0
X_test=X_test.astype('float16')
X_test=X_test/255.0
opt = keras.optimizers.Adam(learning_rate = 0.00001)
trasfer_model.compile(loss='categorical_crossentropy', optimizer=opt,metrics=['accuracy'])
history = trasfer_model.fit(
X_train, y_train,
batch_size=8,
epochs=30,
validation_data=(X_test,y_test)
)
trasfer_model.save('tran_video_model.h5')
plot_training_loss(history)
plot_training_acc(history)
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc" style="margin-top: 1em;"><ul class="toc-item"></ul></div>
# Saving TF Models with SavedModel for TF Serving <a class="tocSkip">
```
import math
import os
import numpy as np
np.random.seed(123)
print("NumPy:{}".format(np.__version__))
import tensorflow as tf
tf.set_random_seed(123)
print("TensorFlow:{}".format(tf.__version__))
DATASETSLIB_HOME = os.path.expanduser('~/dl-ts/datasetslib')
import sys
if not DATASETSLIB_HOME in sys.path:
sys.path.append(DATASETSLIB_HOME)
%reload_ext autoreload
%autoreload 2
import datasetslib
from datasetslib import util as dsu
datasetslib.datasets_root = os.path.join(os.path.expanduser('~'),'datasets')
models_root = os.path.join(os.path.expanduser('~'),'models')
```
# Serving Model in TensorFlow
# Saving model with SavedModel
```
# Restart kernel to run the flag setting again
#tf.flags.DEFINE_integer('model_version', 1, 'version number of the model.')
model_name = 'mnist'
model_version = '1'
model_dir = os.path.join(models_root,model_name,model_version)
# get the MNIST Data
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets(os.path.join(datasetslib.datasets_root,'mnist'), one_hot=True)
x_train = mnist.train.images
x_test = mnist.test.images
y_train = mnist.train.labels
y_test = mnist.test.labels
# parameters
pixel_size = 28
num_outputs = 10 # 0-9 digits
num_inputs = 784 # total pixels
def mlp(x, num_inputs, num_outputs,num_layers,num_neurons):
w=[]
b=[]
for i in range(num_layers):
# weights
w.append(tf.Variable(tf.random_normal( \
[num_inputs if i==0 else num_neurons[i-1], \
num_neurons[i]]), \
name="w_{0:04d}".format(i) \
) \
)
# biases
b.append(tf.Variable(tf.random_normal( \
[num_neurons[i]]), \
name="b_{0:04d}".format(i) \
) \
)
w.append(tf.Variable(tf.random_normal(
[num_neurons[num_layers-1] if num_layers > 0 else num_inputs,
num_outputs]),name="w_out"))
b.append(tf.Variable(tf.random_normal([num_outputs]),name="b_out"))
# x is input layer
layer = x
# add hidden layers
for i in range(num_layers):
layer = tf.nn.relu(tf.matmul(layer, w[i]) + b[i])
# add output layer
layer = tf.matmul(layer, w[num_layers]) + b[num_layers]
model = layer
probs = tf.nn.softmax(model)
return model,probs
tf.reset_default_graph()
# input images
serialized_tf_example = tf.placeholder(tf.string, name='tf_example')
feature_configs = {'x': tf.FixedLenFeature(shape=[784], dtype=tf.float32),}
tf_example = tf.parse_example(serialized_tf_example, feature_configs)
x_p = tf.identity(tf_example['x'], name='x_p') # use tf.identity() to assign name
# target output
y_p = tf.placeholder(dtype=tf.float32, name="y_p", shape=[None, num_outputs])
num_layers = 2
num_neurons = []
for i in range(num_layers):
num_neurons.append(256)
learning_rate = 0.01
n_epochs = 50
batch_size = 100
n_batches = mnist.train.num_examples//batch_size
model,probs = mlp(x=x_p,
num_inputs=num_inputs,
num_outputs=num_outputs,
num_layers=num_layers,
num_neurons=num_neurons)
# loss function
#loss = tf.reduce_mean(-tf.reduce_sum(y * tf.log(model), axis=1))
loss = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=model, labels=y_p))
# optimizer function
optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate)
train_op = optimizer.minimize(loss)
predictions_check = tf.equal(tf.argmax(probs,1), tf.argmax(y_p,1))
accuracy_function = tf.reduce_mean(tf.cast(predictions_check, tf.float32))
values, indices = tf.nn.top_k(probs, 10)
table = tf.contrib.lookup.index_to_string_table_from_tensor(
tf.constant([str(i) for i in range(10)]))
prediction_classes = table.lookup(tf.to_int64(indices))
with tf.Session() as tfs:
tfs.run(tf.global_variables_initializer())
for epoch in range(n_epochs):
epoch_loss = 0.0
for batch in range(n_batches):
x_batch, y_batch = mnist.train.next_batch(batch_size)
_,batch_loss = tfs.run([train_op,loss], feed_dict={x_p: x_batch, y_p: y_batch})
epoch_loss += batch_loss
average_loss = epoch_loss / n_batches
print("epoch: {0:04d} loss = {1:0.6f}".format(epoch,average_loss))
accuracy_score = tfs.run(accuracy_function, feed_dict={x_p: x_test, y_p: y_test })
print("accuracy={0:.8f}".format(accuracy_score))
# save the model
# definitions for saving the models
builder = tf.saved_model.builder.SavedModelBuilder(model_dir)
# build signature_def_map
classification_inputs = tf.saved_model.utils.build_tensor_info(
serialized_tf_example)
classification_outputs_classes = tf.saved_model.utils.build_tensor_info(
prediction_classes)
classification_outputs_scores = tf.saved_model.utils.build_tensor_info(values)
classification_signature = (
tf.saved_model.signature_def_utils.build_signature_def(
inputs={
tf.saved_model.signature_constants.CLASSIFY_INPUTS:
classification_inputs
},
outputs={
tf.saved_model.signature_constants.CLASSIFY_OUTPUT_CLASSES:
classification_outputs_classes,
tf.saved_model.signature_constants.CLASSIFY_OUTPUT_SCORES:
classification_outputs_scores
},
method_name=tf.saved_model.signature_constants.CLASSIFY_METHOD_NAME))
tensor_info_x = tf.saved_model.utils.build_tensor_info(x_p)
tensor_info_y = tf.saved_model.utils.build_tensor_info(probs)
prediction_signature = (
tf.saved_model.signature_def_utils.build_signature_def(
inputs={'inputs': tensor_info_x},
outputs={'outputs': tensor_info_y},
method_name=tf.saved_model.signature_constants.PREDICT_METHOD_NAME))
legacy_init_op = tf.group(tf.tables_initializer(), name='legacy_init_op')
builder.add_meta_graph_and_variables(
tfs, [tf.saved_model.tag_constants.SERVING],
signature_def_map={
'predict_images':
prediction_signature,
tf.saved_model.signature_constants.DEFAULT_SERVING_SIGNATURE_DEF_KEY:
classification_signature,
},
legacy_init_op=legacy_init_op)
builder.save()
print('Run following command:')
print('tensorflow_model_server --model_name=mnist --model_base_path={}'
.format(os.path.join(models_root,model_name)))
```
| github_jupyter |
```
'''
Cleanly written notebook of python commands for producing blog figures
'''
# Initial library imports
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# Import ALL data
## We will merge 'character_stats' and 'superheroes_power_matrix'
csv_files = !ls -1 'Marvel_Data'/Marvel-Superheroes/*.csv
csv_files
# Example csv
pd.read_csv('Marvel_Data/Marvel-Superheroes/marvel_dc_characters.csv',encoding = "ISO-8859-1").head()
# Put all dataframes into a datafram dictionary: df_dict2
## 'First appearance' feature above in marvel_dc_characters needs special encoding, see above
df_dict2 = {}
for csv_f in csv_files:
k = csv_f.split('/')[-1]
k = k.split('.')[0] #to get name with the csv extenstion
#print(k)
if[k=="marvel_dc_characters"]:
df_dict2[k] = pd.read_csv(csv_f,encoding = "ISO-8859-1")
else:
df_dict2[k] = pd.read_csv(csv_f)
df_dict2.keys()
# Shapes
print(df_dict2['characters'].shape, df_dict2['charactersToComics'].shape, df_dict2['characters_stats'].shape,"\n")
print(df_dict2['comics'].shape, df_dict2['hero-network'].shape, df_dict2['marvel_characters_info'].shape,"\n")
print(df_dict2['marvel_dc_characters'].shape, df_dict2['superheroes_power_matrix'].shape)
# Superpower list for each character. 168 total powers, we will not use so many for our model...
df_dict2['superheroes_power_matrix'].head()
# character_stats sample data
## notice: not all stats are included, we will count how many like this later...
df_dict2['characters_stats'].sample(10)
# Watch out for FRANKLIN STORM, the only superhero with Power=0 and ACTUAL STATS
df_dict2['characters_stats'][(df_dict2['characters_stats']['Power']==0) &
(df_dict2['characters_stats']['Intelligence']>1)]
# Merge data
super_data = df_dict2['characters_stats']
super_data = super_data.merge(df_dict2['superheroes_power_matrix'], on='Name')
super_data
# Our total observations = 519
super_data.shape
# 104 observations with NO STATS will need to impute
super_data[super_data['Power'] ==0].shape
# We've now cut 39,648 total superhero characters in Marvel/DC-Comics Universe to only
# 519 characters with stats AND superpowers :(
# This list is mostly among the MOST popular characters in the superhero universe
# Before we get started, let's see the publication years of our TOTAL comicbook dataset
# We see our comics span 1939-2019. An 80 year span!
#-----Unused-----------
## Creating a filter to find the comics with least 8 characters in
comicsWith7orMoreCharacterFilter=df_dict2['charactersToComics'].groupby(by='comicID').count()['characterID']>7
## Applying the filter and find comics with 7+ characters in it
comicsWith7orMoreCharacterData=df_dict2['charactersToComics'].groupby(by='comicID').count()[comicsWith7orMoreCharacterFilter]
#-----Unused-----------
#Finding year of the comics
## First, creating a function to extract the year from comics title column
def extractYear(txt):
startChr=txt.find('(')+1
endChr=txt.find(')')
try:
result=int(txt[startChr:endChr])
return result
except:
pass
# Creating a copy of comics data frame
comicsWyear=df_dict2['comics'].copy()
# Applying extractYear function and creating a new column that contains year of the comics
comicsWyear['year']=df_dict2['comics']['title'].apply(extractYear)
# Creating a bar chart
comicsWyear.groupby(by='year').count()['comicID'].plot(kind='bar',figsize=(18,12));
plt.title('Comics Printed per Year')
del comicsWyear
# Work in progress... need to show popularity of our tiny subsample as valid representation
# of 'good' vs 'bad' divide.
# Improve model: add popularity feature (comic book appearances)
# add 'year-character-created'
# Before we move on...
## Three alginment types will be reduced to two
## for more accurate labeling of data and to go easier on our model
# BEFORE
super_data['Alignment'].value_counts()
#AFTER
super_data['Alignment'] = super_data['Alignment'].apply(lambda x: 'bad' if x=="neutral" or x=="bad"
else 'good')
super_data['Alignment'].value_counts()
# Getting a feel for our stats data...
## Let's first isolate for only data with data ... cutting ~20% data that we will return shortly
super_data_w_stats = super_data[super_data['Power'] != 0]
super_data_w_stats.shape
# Total power stat of our subsample of Heroes w Stats shows:
## 'bad' superheroes have higher stats
import seaborn as sns
#print(df_dict2['characters_stats'].groupby(by='Alignment').agg(['count','mean','sum'])['Total'])
print('Total power of \'bad\' guys are greater than \'good\' guys')
print(super_data_w_stats.groupby(by='Alignment').agg(['count','mean','sum'])['Total'])
sns.boxplot(x='Alignment',
y='Total',
# data=df_dict2['characters_stats']);
data=super_data_w_stats);
# Looking at the boxplot another way...
# Our stat distributions look mostly gaussian.
# 'bad' types are slightly left-skewed: No weak bad guys, superstrong are restricted to bad
#Filter Good and Bad Guys
filterGood=super_data_w_stats['Alignment'].isin(['good'])
filterBad =super_data_w_stats['Alignment'].isin(['bad'])
#Creating a figure for 2 plots
f,(ax1,ax2)=plt.subplots(2,1,figsize=(7,7), sharex=True, sharey=True);
#Plot the data and show the distribution
sns.distplot(super_data_w_stats[filterGood]['Total'],ax=ax1);
ax1.set_ylabel("Good");
sns.distplot(super_data_w_stats[filterBad]['Total'],ax=ax2);
ax2.set_ylabel("Bad");
# Individual stats show 'good' vs 'bad' discrepancy mostly exists in Intelligence, Strength, and Durability
stat_cols = ['Intelligence_x', 'Strength', 'Speed', 'Durability_x', 'Combat', 'Power']
super_data_w_stats [ super_data_w_stats['Alignment'] == 'bad'][stat_cols].mean()
super_data_w_stats [ super_data_w_stats['Alignment'] == 'good'][stat_cols].mean()
# A sample scatter plot to see rough feel for correlation of stats
super_data_w_stats.plot(kind='scatter', x='Strength',y='Intelligence_x',alpha=0.5,color='blue')
super_data_w_stats.xlabel1=('Strength')
super_data_w_stats.ylabel1=('Intelligence')
plt.title('Intelligence vs Strength');
# Now, let's switch to getting a feel of our superpowers
## See our 168 types of superpowers
superhero_powers = df_dict2['superheroes_power_matrix']
superpower_columns = list(superhero_powers.columns)
superpower_columns.remove('Name')
superpower_list = list(superhero_powers.columns)
superpower_list.sort()
superpower_list
# We'll want to cut some of our superpower features to limit our model for now
## We'll show the powers with less than 20 characters than hold them
## Preferable to keep and manage smartly for model
columns_to_remove=[]
for column in superpower_columns:
if (int(superhero_powers[column].sum()) < 30):
print(column, superhero_powers[column].sum())
columns_to_remove.append(column)
print("\nWe will cut", len(columns_to_remove), "superpower features.")
# Work in progress ... try to show superpowers correlated to other superpowers
# AND superpowers correlated to good vs bad
# Now let's remind our superhero data
super_data
# Wrangle our data to impute stats for no-stat-characters and cut some superpowers
def wrangle(X):
X = X.copy()
# IMPUTATION
stat_cols = ['Intelligence_x', 'Strength', 'Speed', 'Durability_x', 'Combat', 'Power']
for col in stat_cols:
impute_val = X[X['Power']!=0][col].mean()
def impute_stat(row):
if row['Power']==0:
return impute_val
else:
return row[col]
X[col] = X.apply(impute_stat, axis=1)
if col==stat_cols[0]:
X['Total']=0
X['Total'] = X['Total'] + X[col]
# CUT SUPERPOWERS
## Remove superpower columns with num superheroes listed below cutoff
## Should probably fix double-features 'Durability' and 'Intelligence' stats vs powers
cutoff = 30
superpower_columns = list(df_dict2['superheroes_power_matrix'].columns)
superpower_columns.remove('Name')
columns_to_remove=[]
for col in superpower_columns:
if (int(df_dict2['superheroes_power_matrix'][col].sum()) < cutoff):
#print(col, superhero_powers[col].sum())
columns_to_remove.append(col)
X = X.drop(list(columns_to_remove),1)
return X
super_data_wrangled = wrangle(super_data)
super_data_wrangled
super_data_wrangled.shape
super_data_wrangled.columns
super_data_wrangled['Alignment'].value_counts()
# MODEL TARGET = 'Alignment'
target = 'Alignment'
train = super_data_wrangled
X_train = train.drop(columns=target)
X_train = X_train.drop('Name',axis=1)
y_train = super_data_wrangled[target]
y_train.value_counts()
# BASELINE ACCURACY
majority_class = y_train.mode()[0]
y_pred = [majority_class]*len(y_train)
from sklearn.metrics import accuracy_score
accuracy_score(y_train,y_pred)
# Study model quality with train and val data. ( 80% train | 20% val )
from sklearn.model_selection import train_test_split
X_train, X_val, y_train, y_val = train_test_split(X_train, y_train, test_size=0.2,
stratify=train[target], random_state=42)
X_train.shape, X_val.shape, y_train.shape, y_val.shape
# Check alignment proportion pt1
y_train.value_counts(normalize=True)
# Check alignment proportion pt2
y_val.value_counts(normalize=True)
# Initally, RANDOM FOREST MODEL
# By the way, if only 'stats' columns are used, accuracy score = .70
# if only 'superpower' columns are used, accuracy score = .64
# Therefore, inclusion of both features crucial for beating baseline model
from sklearn.ensemble import RandomForestClassifier
model = RandomForestClassifier(n_estimators=1000, random_state=42, n_jobs=-1)
model.fit(X_train, y_train)
y_pred = model.predict(X_val)
accuracy_score(y_val,y_pred)
%matplotlib inline
import matplotlib.pyplot as plt
plt.figure(figsize=(10,10))
importances = pd.Series(model.feature_importances_, X_train.columns)
importances.sort_values().plot.barh(color='grey');
# Next, Improved model: XGBoost
from xgboost import XGBClassifier
eval_set = [
(X_train, y_train),
(X_val, y_val)
]
#model = XGBClassifier(n_estimators=2000, n_jobs=-1)
model = XGBClassifier(objective = 'multi:softmax', booster = 'gbtree', nrounds = 'min.error.idx',
num_class = 3, maximize = False, eval_metric = 'merror', eta = .1,
max_depth = 20, colsample_bytree = .4, n_jobs = -1, random_state=42,
)
model.fit(X_train,y_train,
eval_set=eval_set,
early_stopping_rounds=20
)
# XGBOOST SCORE
y_pred = model.predict(X_val)
accuracy_score(y_val,y_pred)
# Feature importance drastically changes upon hypertuning optimization is welcomed
importances = pd.Series(model.feature_importances_, X_train.columns)
# Plot feature importances
n = len(X_train.columns)
plt.figure(figsize=(10,n/2))
plt.title(f'Top {n} features')
importances.sort_values()[-n:].plot.barh(color='grey');
# Example of some strong predictors. Normally, good:bad ratio is about 2.4 : 1
## Strong predictor 1 - good:bad ratio almost 1 highly doesn't match blind ratio...
train[train['Molecular Manipulation']==True]['Alignment'].value_counts()
## Strong predictor 2 - high frequency superpower &
## ratio of ~1.75 doesn't match blind good:ratio
train[train['Super Strength']==True]['Alignment'].value_counts()
# Weak predictor
train[train['Cold Resistance']==True]['Alignment'].value_counts()
# PERMUTATION IMPORTANCE
## (note: high errors)
import eli5
from eli5.sklearn import PermutationImportance
permuter = PermutationImportance(model, scoring='accuracy',
cv='prefit', n_iter=5, random_state=42)
permuter.fit(X_val, y_val)
feature_names = X_val.columns.tolist()
eli5.show_weights(permuter, top=None, feature_names=feature_names)
# Partial Dependece Plots (PDP)
# PDP Intelligence
## Moderate intelligence suggests "good" alignment but high intelligence suggest "bad"
from pdpbox.pdp import pdp_isolate, pdp_plot
feature = 'Intelligence_x'
isolated = pdp_isolate(
model=model,
dataset=X_val,
model_features=X_val.columns,
feature=feature
)
pdp_plot(isolated, feature_name=feature);
# PDP Total stat
## Total stat indicates super-duper statatiscally strong characters typically 'bad'
from pdpbox.pdp import pdp_isolate, pdp_plot
feature = 'Total'
isolated = pdp_isolate(
model=model,
dataset=X_val,
model_features=X_val.columns,
feature=feature
)
pdp_plot(isolated, feature_name=feature);
# PDP Super Strength power
## Superpower PDP's all show a small, high variance effect for prediction
from pdpbox.pdp import pdp_isolate, pdp_plot
feature = 'Super Strength'
isolated = pdp_isolate(
model=model,
dataset=X_val,
model_features=X_val.columns,
feature=feature
)
pdp_plot(isolated, feature_name=feature);
# PDP Super Strength power
## Superpower PDP's all show high variance in effect
from pdpbox.pdp import pdp_isolate, pdp_plot
feature = 'Time Travel'
isolated = pdp_isolate(
model=model,
dataset=X_val,
model_features=X_val.columns,
feature=feature
)
pdp_plot(isolated, feature_name=feature);
#-------------------------DONE-----------------------------------------------#
# cross-validation ought to be performed for our small/median dataset
# Plotly-dash is work in progess... When user inputs stats/superpowers,
# tell user their character is most like _______ (ex. Thanos i.e. similarity score _____ -> bad)
# Restrict this data to MARVEL data NOT DC-Comics
# Nemesis Predictor Model
# SHAPLEY PLOTS (still in development)
## Pick out some characters to briefly look at: Thanos, Spider-Man
## Preliminary test results show reverse of model expectation
df_dict2['characters_stats'][df_dict2['characters_stats']['Name']=='Thanos']
data_for_prediction = X_train[101:102]
data_for_prediction
y_pred = model.predict(X_train[101:102])
y_pred
import shap
shap.initjs()
explainer = shap.TreeExplainer(model)
shap_values = explainer.shap_values(data_for_prediction)
shap_values
shap.force_plot(explainer.expected_value[0], shap_values[0], data_for_prediction)
df_dict2['characters_stats'][df_dict2['characters_stats']['Name']=='Spider-Man']
data_for_prediction = X_train[171:172]
data_for_prediction
import shap
shap.initjs()
explainer = shap.TreeExplainer(model)
shap_values = explainer.shap_values(data_for_prediction)
shap_values
shap.force_plot(explainer.expected_value[0], shap_values[0], data_for_prediction)
```
| github_jupyter |
# Face Recognition with SphereFace
Paper: https://arxiv.org/abs/1704.08063
Repo: https://github.com/wy1iu/sphereface
```
import cv2
import numpy as np
import pandas as pd
from tqdm import tqdm
import matplotlib.pyplot as plt
#We are going to use deepface to detect and align faces
#Repo: https://github.com/serengil/deepface
#!pip install deepface
from deepface.commons import functions
```
### Pre-trained model
```
#Structure: https://github.com/wy1iu/sphereface/blob/master/train/code/sphereface_deploy.prototxt
#Weights: https://drive.google.com/open?id=0B_geeR2lTMegb2F6dmlmOXhWaVk
model = cv2.dnn.readNetFromCaffe("sphereface_deploy.prototxt", "sphereface_model.caffemodel")
#SphereFace input shape. You can verify this in the prototxt.
input_shape = (112, 96)
```
### Common functions
```
#Similarity metrics tutorial: https://sefiks.com/2018/08/13/cosine-similarity-in-machine-learning/
def findCosineDistance(source_representation, test_representation):
a = np.matmul(np.transpose(source_representation), test_representation)
b = np.sum(np.multiply(source_representation, source_representation))
c = np.sum(np.multiply(test_representation, test_representation))
return 1 - (a / (np.sqrt(b) * np.sqrt(c)))
def findEuclideanDistance(source_representation, test_representation):
euclidean_distance = source_representation - test_representation
euclidean_distance = np.sum(np.multiply(euclidean_distance, euclidean_distance))
euclidean_distance = np.sqrt(euclidean_distance)
return euclidean_distance
```
### Data set
```
# Master.csv: https://github.com/serengil/deepface/blob/master/tests/dataset/master.csv
# Images: https://github.com/serengil/deepface/tree/master/tests/dataset
df = pd.read_csv("../deepface/tests/dataset/master.csv")
df.head()
euclidean_distances = []; cosine_distances = []
for index, instance in tqdm(df.iterrows(), total = df.shape[0]):
img1_path = instance["file_x"]
img2_path = instance["file_y"]
target_label = instance["Decision"]
#----------------------------------
#detect and align
img1 = functions.preprocess_face("../deepface/tests/dataset/%s" % (img1_path), target_size=input_shape)[0]
img2 = functions.preprocess_face("../deepface/tests/dataset/%s" % (img2_path), target_size=input_shape)[0]
#----------------------------------
#reshape images to expected shapes
img1_blob = cv2.dnn.blobFromImage(img1)
img2_blob = cv2.dnn.blobFromImage(img2)
if img1_blob.shape != (1, 3, 96, 112):
raise ValueError("img shape must be (1, 3, 96, 112) but it has a ", img1_blob.shape," shape")
#----------------------------------
#representation
model.setInput(img1_blob)
img1_representation = model.forward()[0]
model.setInput(img2_blob)
img2_representation = model.forward()[0]
#----------------------------------
euclidean_distance = findEuclideanDistance(img1_representation, img2_representation)
euclidean_distances.append(euclidean_distance)
cosine_distance = findCosineDistance(img1_representation, img2_representation)
cosine_distances.append(cosine_distance)
df['euclidean'] = euclidean_distances
df['cosine'] = cosine_distances
df.head()
```
### Visualize distributions
```
df[df.Decision == "Yes"]['euclidean'].plot(kind='kde', title = 'euclidean', label = 'Yes', legend = True)
df[df.Decision == "No"]['euclidean'].plot(kind='kde', title = 'euclidean', label = 'No', legend = True)
plt.show()
df[df.Decision == "Yes"]['cosine'].plot(kind='kde', title = 'cosine', label = 'Yes', legend = True)
df[df.Decision == "No"]['cosine'].plot(kind='kde', title = 'cosine', label = 'No', legend = True)
plt.show()
```
### Find the best threshold
```
#Repo: https://github.com/serengil/chefboost
#!pip install chefboost
from chefboost import Chefboost as chef
config = {'algorithm': 'C4.5'}
df[['euclidean', 'Decision']].head()
euclidean_tree = chef.fit(df[['euclidean', 'Decision']].copy(), config)
cosine_tree = chef.fit(df[['cosine', 'Decision']].copy(), config)
#stored in outputs/rules
euclidean_threshold = 17.212238311767578 #euclidean
cosine_threshold = 0.4668717384338379 #cosine
```
### Predictions
```
df['prediction_by_euclidean'] = 'No'
df['prediction_by_cosine'] = 'No'
df.loc[df[df['euclidean'] <= euclidean_threshold].index, 'prediction_by_euclidean'] = 'Yes'
df.loc[df[df['cosine'] <= cosine_threshold].index, 'prediction_by_cosine'] = 'Yes'
df.sample(5)
euclidean_positives = 0; cosine_positives = 0
for index, instance in df.iterrows():
target = instance['Decision']
prediction_by_euclidean = instance['prediction_by_euclidean']
prediction_by_cosine = instance['prediction_by_cosine']
if target == prediction_by_euclidean:
euclidean_positives = euclidean_positives + 1
if target == prediction_by_cosine:
cosine_positives = cosine_positives + 1
print("Accuracy (euclidean): ",round(100 * euclidean_positives/df.shape[0], 2))
print("Accuracy (cosine): ",round(100 * cosine_positives/df.shape[0], 2))
```
### Production
```
def verifyFaces(img1_path, img2_path):
print("Verify ",img1_path," and ",img2_path)
#------------------------------------
#detect and align
img1 = functions.preprocess_face(img1_path, target_size=input_shape)[0]
img2 = functions.preprocess_face(img2_path, target_size=input_shape)[0]
img1_blob = cv2.dnn.blobFromImage(img1)
img2_blob = cv2.dnn.blobFromImage(img2)
#------------------------------------
#representation
model.setInput(img1_blob)
img1_representation = model.forward()[0]
model.setInput(img2_blob)
img2_representation = model.forward()[0]
#------------------------------------
#verify
euclidean_distance = findEuclideanDistance(img1_representation, img2_representation)
print("Found euclidean distance is ",euclidean_distance," whereas required threshold is ",euclidean_threshold)
fig = plt.figure()
ax1 = fig.add_subplot(1,2,1)
plt.imshow(img1[:,:,::-1])
plt.axis('off')
ax2 = fig.add_subplot(1,2,2)
plt.imshow(img2[:,:,::-1])
plt.axis('off')
plt.show()
if euclidean_distance <= euclidean_threshold:
print("they are same person")
else:
print("they are not same person")
```
### True positive examples
```
verifyFaces("../deepface/tests/dataset/img1.jpg", "../deepface/tests/dataset/img2.jpg")
verifyFaces("../deepface/tests/dataset/img54.jpg", "../deepface/tests/dataset/img3.jpg")
verifyFaces("../deepface/tests/dataset/img42.jpg", "../deepface/tests/dataset/img45.jpg")
verifyFaces("../deepface/tests/dataset/img9.jpg", "../deepface/tests/dataset/img49.jpg")
```
### True negative examples
```
verifyFaces("../deepface/tests/dataset/img1.jpg", "../deepface/tests/dataset/img3.jpg")
verifyFaces("../deepface/tests/dataset/img1.jpg", "../deepface/tests/dataset/img45.jpg")
verifyFaces("../deepface/tests/dataset/img1.jpg", "../deepface/tests/dataset/img49.jpg")
```
| github_jupyter |
# Supervised learning (without calculus!)
This tutorial wants to show the (possibly) most naive way to train the most naive neural network on the [MNIST database](https://en.wikipedia.org/wiki/MNIST_database). For our model we only need to known matrix multiplication, for loops. Importantly, no knowledge of calculus (gradient descent) is required and coding syntax is explained in detail. The reader expect to:
- present key ideas in machine learning in simple setting,
- be impressed by the power of simple matrix multiplication,
- build intuition for issues such as overfitting and
- have plenty of extensions of the model to experiment.
(This tutorial was originally made as a concrete application of the linear algebra course at Otago University.)
## Dataset and packages
We first import some basic packages along with the training/testing and test dataset and we reduce the size of the training dataset. (Note that tensorflow is only used to import the data.)
```
# We import the packages that we will use
import matplotlib.pyplot as plt
import tensorflow as tf
import numpy as np
from numpy.linalg import norm as Euclid_norm
from time import time
#We import the dataset
mnist = tf.keras.datasets.mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
size_train_small = 200
x_train, y_train = x_train[:size_train_small], y_train[:size_train_small]
```
## Data visualisation
Our aim is to "train" a "machine" capable of recognising the ten digits $\{0,1,2,3,4,5,6,7,8,9\}$ when hand-written. It is natural to show the machine a bunch of these digits and then to train it to choose correctly by rewarding the machine when correct, and penalising it otherwise (much like humans do). Once trained we send the machine into the unkown territory of the testing data set, and see how well it performs. Lukily, images are matrices (tabular arrays) where each position represent a pixel and the value of this entry is the color (and machines love matrices). Let's see this.
```
fig, ax = plt.subplots(1,5,figsize=(10,4))
fig.suptitle('.jpg files as 28x28 matrices with the respective labels', fontsize=15)
for j in range(5):
ax[j].imshow(x_train[j:j+1][0])
ax[j].set_title(f'Label: {y_train[j]}');
plt.show();
print("To see the first image in matrix notation type x_train[0] in a cell and run it. ");
```
## The neural network
So an image is a $28\times 28$ matrix, or equivalenty a $28^2\times 1$ vector/matrix, obtained by flattening it (stack all rows in one line). For each image/vector the machine needs to return a number between 0 and 9 (according to what the image is understood to be). What is the easiest (reasonable) way to obtain ten evaluations out of a $28^2\times 1$ vector $x$. Of course it is matrix multiplication by a $10\times 28^2$ matrix `W`. The matrix `W` is called the weights matrix (it weights `x` to to measure its value out of 10 different options) and our model follows the simple decision ruled
**PREDICTION**: `x` is the digit equal to the index of the biggest entry in `Wx`.
For example if `Wx` $ = [0,0,0,1,0,3,-\pi, 7,100]^T$, then the machine tells us that `x` is the picture of a 9. That simple.
**TRAINING** = finding a good `W`. What's "good"? It's up to us to decide. For example, an option is to tell the machine that $W x$ needs to have a very high third entry if the label of $x$ is 2, this is the same as asking that
- $\|W x-y \|_2$ is small,
- $y = [0,0,C,0,0,0,0,0,0,0]$,
- $C>0$ is very big and
where $\|$ `x` $ \|_2=(x_0^2+...+x^2_9)^{\frac12}$ is the 10-$d$ Euclidean norm. So we require $\|W x-y \|_2$ to be samll for all images in the traning set by requiring that the loss function
$$
L(W)= \frac1n\sum_{i=1}^n\|Wx_i - y_i\|_2
$$
is small, where $\{(x_i,y_i)\}_{i=1}^n$ are the pairs of images with their vector label and $n$ is `size_train_small`. Mathematically, we want a `W` that solves the optimisation problem $\arg\min_W L(W)$. SoHow do we train this model? With a
**Trivialized gradient descent alghoritym**:
Initialize at the entry `(i, j) = (0, 0)`, then
1. Take the current `W` and create a copy `W_temp` that equals `W` apart from `W_temp[i,j] = W[i,j] + 1`,
2. if `L(W_temp) < L(W)` make `W_temp` the new `W`.
3. Fix a new entry `(i,j)` to move horizontally in the $10\times 28^2$ matrix `W` (skipping lines once at the last column and staring back from `(0,0)` once all entries have been visited)
4. start back at 1.
## Model construction
We first flatten all images and construct labeld vectors in `y_label`.
```
#Flatten images in training set
x_train = x_train.reshape(x_train.shape[0],28*28)
x_test = x_test.reshape(x_test.shape[0],28**2)
#Create 10x1 vectors with a 1-entry corresponding the respective
#image label
y_label = np.zeros((y_train.shape[0], 10))
for i in range(y_train.shape[0]):
y_label[i,y_train[i]] = 1
```
The following is the class cotaining our model. It performs three main tasks:
I. `nl = NL()`: the model is created and named `nl`, and the size of each image input vector is set to 28*28, the output vector size to 10, the weight matrix `W` is initialised to a $10\times28*28$ matrix where each entry is a 0, the number of times we run through all the entries of `W` in our gradiant descent (in `self.steps`).
II. `nl.fit(x_train,y_label)`: this is the trivialized gradient descent, check the description of the steps!
III. `nl.predict(x_test):` the decision making process. The `nl` computes the matrix product `W@x` with the current `W` for each image `x` in `x_test` and returns its decisions, i.e. the index of the biggest entry in `W@x`.
The remaining two functions simply compute the average correct predictions of `nl` and allow to return the current weights matrix `W`.
```
class NL:
def __init__(self):
self.size_in = 28*28 #Lenght of a flattened image
self.size_out = 10 #Ten possible choices for the possible digit in the image
self.weights = np.zeros((self.size_out, self.size_in)) #Initialize W as a zero matrix
self.steps = 10 #How many times the descent algorythm is performed on all entries of W
def fit(self,x_train,y_label):
"""Train the neural network by performing the trivialized
gradient descent."""
t_start = time() #Time at start of training
steps = self.steps*(self.size_in*self.size_out) #Number of iteration of the descent algo
W = self.weights
h, C = 2., 10**5. #Set the learning step h and the big constant C to scale y_label
y_label = C*y_label
#Algorythm starts here:
for i in range(steps): #Steps 3 and 4 happen here
#Step 1: make a copy of W
W_temp = W.copy()
#Change the current entry of W_temp by h
j = i % (self.size_in*self.size_out -1)
W_temp.reshape(np.prod(W_temp.shape))[j] = W_temp.reshape(np.prod(W_temp.shape))[j] + h
#Step 2: Create a matrix contaning all vectors Wx - y
Wx_minus_y = np.ones(x_train.shape[0]*10).reshape(x_train.shape[0],10)
for m in range(x_train.shape[0]):
Wx_minus_y[m] = W@x_train[m]- y_label[m]
#and create a matrix contaning all vectors W_temp x - y
W_temp_x_minus_y = np.ones(x_train.shape[0]*10).reshape(x_train.shape[0],10)
for m in range(x_train.shape[0]):
W_temp_x_minus_y[m] = W_temp@x_train[m] - y_label[m]
#and then compute the loss under the current and temporary weight matrices
Loss_W = Euclid_norm(Wx_minus_y,axis=0).mean()
Loss_W_temp = Euclid_norm(W_temp_x_minus_y,axis=0).mean()
#and decide whether we descend with W_temp
if Loss_W_temp < Loss_W:
W = W_temp
t_end = time() #Time at end of training
self.weights = W # The training has finished and the last W is set to be the model weight matrix
print("Training time:", np.round(t_end-t_start,2),"s")
def predict(self,x_test):
"""Predict the digit in each image x in x_test by returning
the position where W@x is the highest.
"""
y_pred = np.ones(x_test.shape[0])
for i in range(x_test.shape[0]):
y_pred[i] = np.where(self.weights@x_test[i]
== np.max(self.weights@x_test[i]))[0][0]
return y_pred
def return_weights(self):
return self.weights
def accuracy(self,y_test,y_pred):
return len(np.where((y_pred - y_test) == 0.)[0])/len(y_test)
```
## Training and predictions
So initialise/instantiate the model and check its predictions before training, which are of course goiong to be pretty bad and expected to be around 10% accuracy (why?).
```
nl = NL()
#Compute prediction accuracy without training
y_pred = nl.predict(x_test)
print("Probability of correct prediction before training:\n", nl.accuracy(y_pred,y_test))
```
Finally, we are ready to train (or fit) the model. So lets feed to `nl` the training images with the respective labels and let do its homework.
```
nl.fit(x_train,y_label)
```
Now that the model is trained, let's check its predictions.
```
y_pred_in_sample = nl.predict(x_train)
print("Probability of correct prediction after training\n\n- on traning set:",
nl.accuracy(y_pred_in_sample,y_train))
y_pred_out_of_sample = nl.predict(x_test)
print("\n- on test set:",
nl.accuracy(y_pred_out_of_sample,y_test))
```
## Discussion and exercises
So we trained the machine to recognise digits with more than 60% accuracy with rather trivial mathematics!!! This is very exciting because 60% is so much higher than 10%, meaning that the machine did learn a good deal about digits during its first five minutes of training.
However, note that on the training set the machine, or better, the weight matrix performs much better, close to 90%. This is a classic case of overfitting, mainly due to the small sized training set that we provided. So we leave the reader with some questions to think about:
- How can we interpret the rows of `W`?
- How can you fix the overfitting?
- Can you speed up the calculations to allow for bigger size traning sets (would dropouts make sense here)?
- Can you improve the model by allowing $h$ and $-h$ learning steps?
## Tensorflow equivalent
We now train our model using TensorFlow on the same dataset and then on the full dataset. We first reimport the data and create a small and a large training set. Then instantiate the first neural network, display its description and its performance before traning. (Notice that the tf model performs the flattening for us.)
```
#reimport data and create a big and small dataset
(x_train_big, y_train_big), (x_test, y_test) = mnist.load_data()
size_train_big = 20000
x_train_big, y_train_big = x_train_big[:size_train_big], y_train_big[:size_train_big]
x_train_small, y_train_small = x_train[:size_train_small], y_train[:size_train_small]
#Model instantiation and summary
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(10,use_bias=False)
])
print("This is the model we instantiated:\n")
print(model.summary())
#Choose a loss function
loss_fn = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
model.compile(optimizer='adam',
loss=loss_fn,
metrics=['accuracy'])
#Check accuracy before training
print('\n\nUntrained accuracy:',
np.round(model.evaluate(x_test, y_test, verbose=0)[1],4))
#Train the model
print(f"\n\nNow we fit/train the model on the small data set",
f"with {size_train_small} images.\n")
model.fit(x_train, y_train, epochs=5)
#Check accuracy in/out-of-sample
print("\n\nProbability of correct prediction after training\n\n- on traning set:",
np.round(model.evaluate(x_train, y_train, verbose=0)[1],4))
print("\n- on test set:",
np.round(model.evaluate(x_test, y_test, verbose=0)[1],4),"\n");
```
So the TensorFlow training, though blindingly fast and using gradient descent, yields a model performs similarly to our naively trained model! In particular it seems to be also suffering from overfitting. Lets try now to train the same model on a bigger set.
```
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(10,use_bias=False)
])
model.compile(optimizer='adam',
loss=loss_fn,
metrics=['accuracy'])
print(f"\n\nWe fit/train the model on the bit data set",
f"with {size_train_big} images.\n\n")
model.fit(x_train_big, y_train_big, epochs=5)
print("\n\nProbability of correct prediction after training\n\n- on traning set:",
np.round(model.evaluate(x_train_big, y_train_big, verbose=0)[1],4))
print("\n- on test set:",
np.round(model.evaluate(x_test, y_test, verbose=0)[1],4));
```
Great news! The in-sample and out-of-sample accuracy are high and very close, so we can conclude that the model is not overfitting the traning set.
Have fun!
---
Author: Lorenzo Toniazzi
Github: [ltoniazzi](https://github.com/ltoniazzi)
### Future additions
- Speed up this code with JAX.
| github_jupyter |
# Example 12.1: Clapeyron Equation
*John F. Maddox, Ph.D., P.E.<br>
University of Kentucky - Paducah Campus<br>
ME 321: Engineering Thermodynamics II<br>*
## Problem Statement
Determine the enthalpy of vaporization for water at 100 kPa using:
* (a) the Clapeyron equation
* (b) the Clapeyron-Clausius equation
* (c) the steam tables
## Solution
__[Video Explanation](https://uky.yuja.com/V/Video?v=3074201&node=10465100&a=2106948292&autoplay=1)__
### Python Initialization
We'll start by importing the libraries we will use for our analysis and initializing dictionaries to hold the properties we will be usings.
```
from kilojoule.templates.default import *
fluid = realfluid.Properties('Water')
p[1] = Quantity(100,'kPa')
x[1] = 0
states.fix(1,fluid)
p[2] = Quantity(125,'kPa')
x[2] = 0
states.fix(2,fluid)
p[3] = Quantity(75,'kPa')
x[3] = 0
states.fix(3,fluid)
p[4] = p[1]
x[4] = 1
states.fix(4,fluid)
states.display();
```
### (a) Clapeyron equation
The Clapeyron equation relates the enthalpy of vaporization to easily measureable properties, $T$, $p$, and $v$
$$ \left( \frac{\partial p}{\partial T} \right)_{sat} = \frac{h_{fg}}{T_{sat}v_{fg}}$$
This can be solved for the enthalpy
$$ h_{fg} = Tv_{fg}\left(\frac{\partial P}{\partial T}\right)_{sat} $$
The application of this equation depends on knowing the partial derivative of the pressure with respect to the temperature during phase change. This can be visualized as the slope of the saturation curve on a $p$-$T$ diagram at the pressure of interest
```
%%showcalc
v_fg = v[4]-v[1]
pT = fluid.property_diagram(x="T",y="p",saturation=False)
pT.ax.set_ylim(bottom=50,top=150)
# pT.ax.set_xlim(left=80,right=120)
# Label phases
pT.plot_iso_line("x",0,y_range=[50,150]*units('kPa'),label='saturated',ycoor=55,color='gray')
pT.text(0.1,.9,'liquid',axcoor=True)
pT.text(0.9,.1,'vapor',axcoor=True)
# Plot states
pT.plot_state(states[1],label_loc='north west',gridlines=True)
pT.plot_state(states[2],label_loc='north west')
pT.plot_state(states[3],label_loc='north west')
```
#### Forward Difference
```
%%showcalc
dPdT_forward = (p[2]-p[1])/(T[2]-T[1])
h_fg_forward = T[1].to('K')*v_fg*dPdT_forward
h_fg_forward.ito('kJ/kg')
x = Quantity(pT.xlim,'degC')
y = dPdT_forward*(x-T[1]) + p[1]
pT.plot(x,y,color='green',linewidth=.5,label='forward')
pT.fig.legend()
pT.show()
```
#### Backward Difference
```
%%showcalc
dPdT_backward = (p[1]-p[3])/(T[1]-T[3])
h_fg_backward = T[1].to('K')*v_fg*dPdT_backward
h_fg_backward.ito('kJ/kg')
y = dPdT_backward*(x-T[1]) + p[1]
pT.ax.plot(x,y,color='blue',linewidth=.5,label='backward')
pT.fig.legend()
pT.show()
```
#### Central Difference
```
%%showcalc
dPdT_central = (p[2]-p[3])/(T[2]-T[3])
h_fg_central = T[1].to('K')*v_fg*dPdT_central
h_fg_central.ito('kJ/kg')
y = dPdT_central*(x-T[1]) + p[1]
pT.ax.plot(x,y,color='red',linewidth=.5,label='central')
pT.fig.legend()
pT.show()
```
### (b) Clapeyron-Clausius equation
$$\ln\left(\frac{p_B}{p_A}\right) = \frac{h_{fg}}{R}\left(\frac{1}{T_A}-\frac{1}{T_B}\right)$$
```
%%showcalc
from math import log
R = fluid.R
h_fg = (R*log(p[3]/p[2]))/(1/T[2].to('K')-1/T[3].to('K'))
h_fg.ito('kJ/kg')
```
### (c) Steam Tables
```
%%showcalc
h_fg = h[4]-h[1]
```
| github_jupyter |
Old Guestbook IP Extraction
===
This script processes the json guestbook in the old (2016) dataset to a CSV file containing the IP metadata.
```
%reload_ext autoreload
%autoreload 2
%matplotlib inline
import os
import re
import pandas as pd
import numpy as np
from collections import Counter
import sqlite3
from nltk import word_tokenize
from html.parser import HTMLParser
from tqdm import tqdm
import random
import pickle
import json
from datetime import datetime
from pprint import pprint
import matplotlib.pyplot as plt
import matplotlib.dates as md
import matplotlib
import pylab as pl
from IPython.core.display import display, HTML
from pathlib import Path
git_root_dir = !git rev-parse --show-toplevel
git_root_dir = Path(git_root_dir[0].strip())
git_root_dir
import sys
caringbridge_core_path = "/home/srivbane/levon003/repos/caringbridge_core"
sys.path.append(caringbridge_core_path)
import cbcore.data.paths as paths
import cbcore.data.dates as dates
raw_data_dir = paths.raw_data_2016_filepath
guestbook_filepath = os.path.join(raw_data_dir, "guestbook_scrubbed.json")
working_dir = "/home/srivbane/shared/caringbridge/data/projects/sna-social-support/geo_data"
os.makedirs(working_dir, exist_ok=True)
assert os.path.exists(working_dir)
```
### Load and convert journal file
```
output_filepath = os.path.join(working_dir, "gb_ip_raw.csv")
bad_ips = []
with open(output_filepath, 'w') as outfile:
with open(guestbook_filepath, encoding='utf-8') as infile:
for line in tqdm(infile, total=82980359):
gb = json.loads(line)
if "ip" not in gb:
continue
ip = gb['ip']
if not re.match(r"[0-9]+\.[0-9]+\.[0-9]+\.[0-9]+$", ip):
bad_ips.append(ip)
continue
created_at = int(gb["createdAt"]["$date"])
updated_at = int(gb["updatedAt"]["$date"]) if "updatedAt" in gb else ""
outfile.write(f"{int(gb['userId'])},{ip},{int(gb['siteId'])},{created_at},{updated_at}\n")
len(bad_ips)
Counter(bad_ips).most_common()[:10]
!wc -l {output_filepath}
```
### Add geo information to data
This is now in a separate PBS script for easier execution as an independent, long-running job.
```
raw_ip_filepath = os.path.join(working_dir, "gb_ip_raw.csv")
geo_info_added_filepath = os.path.join(working_dir, "gb_geo_data.csv")
# read the geoip2 database
import geoip2
import geoip2.database
city_database_filepath = "/home/srivbane/shared/caringbridge/data/derived/geolite2/GeoLite2-City_20190813/GeoLite2-City.mmdb"
reader = geoip2.database.Reader(city_database_filepath)
with open(raw_ip_filepath, 'r') as infile:
with open(geo_info_added_filepath, 'w') as outfile:
for line in tqdm(infile, total=76810342):
tokens = line.strip().split(",")
if len(tokens) != 5:
raise ValueError(f"Too many values to unpack: {line}")
user_id, ip, site_id, created_at, updated_at = tokens
try:
g = reader.city(ip)
except geoip2.errors.AddressNotFoundError:
outfile.write(f"{user_id},{site_id},{created_at},NOT_FOUND,,,,,,\n")
continue
country = g.country.iso_code
subdiv_count = len(g.subdivisions)
state = g.subdivisions.most_specific.iso_code
city = g.city.name
lat = g.location.latitude
long = g.location.longitude
acc_radius = g.location.accuracy_radius
outfile.write(f"{user_id},{site_id},{created_at},{country},{subdiv_count},{state},{city},{lat},{long},{acc_radius}\n")
```
### Read csv file
```
header = ['user_id','site_id','created_at','country','subdiv_count','state','city','lat','long','acc_radius']
df = pd.read_csv(geo_info_added_filepath, header=None, names=header)
len(df)
df.head()
Counter(df.country).most_common()[:15]
Counter(df.state).most_common()[:15]
Counter(df.city).most_common()[:15]
Counter(df.acc_radius).most_common()[:15]
Counter(df.subdiv_count).most_common()[:3]
# multiple subdivisions is a non-US thing
Counter(df[df.subdiv_count == 2].country).most_common()
plt.hist([int(ar) for ar in df.acc_radius if ar is not None and ar != 'None' and not pd.isnull(ar)], log=True)
plt.title("Accuracy radius for the classified points")
plt.show()
```
### Visualization by US state and location
```
us_df = df[(df.subdiv_count == 1)&(df.country=='US')]
len(us_df), len(us_df) / len(df)
us_df.head()
import geopandas as gpd
import geopandas.datasets
import shapely
#from quilt.data.ResidentMario import geoplot_data
#world = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
#world = gpd.read_file(geoplot_data.contiguous_usa())
world = gpd.read_file(geopandas.datasets.get_path('naturalearth_lowres'))
fig, ax = plt.subplots(1,1, figsize=(12,12))
ax = world.plot(ax=ax, color="seagreen")
marker_size_map = {'1000': 9,
'500': 8,
'200': 7,
'100': 6,
'50': 5,
'20': 4,
'10': 3,
'5': 2,
'1': 1
}
marker_color_map = {'1000': plt.cm.Greys(0.7, alpha=0.01),
'500': plt.cm.Greys(0.7, alpha=0.05),
'200': plt.cm.Greys(0.8, alpha=0.2),
'100': plt.cm.Greys(0.8, alpha=0.3),
'50': plt.cm.Greys(0.9, alpha=0.4),
'20': plt.cm.Greys(0.9, alpha=0.4),
'10': plt.cm.Greys(1, alpha=0.5),
'5': plt.cm.Greys(1, alpha=0.5),
'1': plt.cm.Greys(1, alpha=0.5)
}
geometry = [shapely.geometry.Point(xy) for xy in zip(us_df.long.astype("float"), us_df.lat.astype("float"))]
crs = {'init': 'epsg:4326'}
gdf = gpd.GeoDataFrame(us_df, crs=crs, geometry=geometry)
gdf = gdf.to_crs(world.crs) # just to ensure the coordinate system is the same
markersizes = [marker_size_map[ar] for ar in us_df.acc_radius]
colors = [marker_color_map[ar] for ar in us_df.acc_radius]
ax = gdf.plot(ax=ax, color=colors, markersize=markersizes)
# try to apply different colors to the response categories in order to sanity check the demo_country variable
#ax = gdf[gdf["demo_country"] == "USA"].plot(ax=ax, color='blue', markersize=4)
#ax = gdf[gdf["demo_country"] == "Canada"].plot(ax=ax, color='red', markersize=4)
#ax = gdf[gdf["demo_country"] == "Other"].plot(ax=ax, color='white', markersize=2)
# some hacky estimates of North America coords
ax.set_xlim((-141,-62))
ax.set_ylim((14,72))
_ = ax.set_title("US Guestbook Locations (From IP)")
```
| github_jupyter |
# Combining Thompson Sampling Results
```
import pinot
ds = pinot.data.moonshot()
actual_best = max([d[1].item() for d in ds])
import pandas as pd
best_human = pd.read_csv('best_Human.csv', index_col=0)
pro_human = pd.read_csv('pro_Human.csv', index_col=0)
retro_human = pd.read_csv('retro_Human.csv', index_col=0)
for df in [best_human, pro_human, retro_human]:
df['Type'] = 'Human'
best_ei = pd.read_csv('best_ExpectedImprovement.csv', index_col=0)
pro_ei = pd.read_csv('pro_ExpectedImprovement.csv', index_col=0)
retro_ei = pd.read_csv('retro_ExpectedImprovement.csv', index_col=0)
for df in [best_ei, pro_ei, retro_ei]:
df['Type'] = 'ExpectedImprovement'
best = pd.concat([best_human, best_ei])
pro = pd.concat([pro_human, pro_ei])
retro = pd.concat([retro_human, retro_ei])
def larger_font(ylabel):
plt.xticks(size=20)
plt.xlabel('Round', size=20)
plt.yticks(size=20)
plt.ylabel(ylabel, size=20)
import matplotlib.pyplot as plt
import seaborn as sns
sns.catplot(x='Round', y='Value',
hue='Type',
data=retro,
kind='violin',
height=10,
aspect=2,
# split=True
palette='tab10'
)
larger_font('Thompson Estimates of $y_{max}$')
plt.axhline(actual_best, color='black')
import seaborn as sns
fig, ax = plt.subplots(figsize=(20, 10))
# plt.axhline(actual_best, color='black')
sns.lineplot(x='Round', y='Value', hue='Type', data=best, ax=ax)
larger_font('$y_{max}^i$')
import torch
import numpy as np
import pandas as pd
import seaborn as sns
improvement_list = []
for type_ in ['Human', 'ExpectedImprovement']:
pro_subset = pro[pro['Type'] == type_]
best_subset = best[best['Type'] == type_]
for trial in pro_subset.Trial.unique():
for round_ in pro_subset.Round.unique():
round_values = pro_subset[np.logical_and(pro_subset['Round'] == round_, pro_subset['Trial'] == trial)]['Value']
round_best = best[np.logical_and(best['Round'] == round_, best['Trial'] == trial)]['Value'].iloc[0]
improvement_list.append({'Acquisition Function': 'ExpectedImprovement',
'Trial': trial,
'Round': round_,
'Type': type_,
'ProbabilityImprovement': (round_values > round_best).mean(),
'ExpectedImprovement': (np.maximum(round_values - round_best, 0)).mean()})
improvement_df = pd.DataFrame(improvement_list)
import matplotlib.pyplot as plt
import pandas as pd
fig, ax = plt.subplots(figsize=(10, 10))
sns.swarmplot(x='Round', y='ProbabilityImprovement', hue='Type', data=improvement_df, ax=ax)
larger_font('$P$(Thompson Estimate > $y_{max}^i$)')
fig, ax = plt.subplots(figsize=(10, 10))
sns.swarmplot(x='Round', y='ExpectedImprovement', hue='Type', data=improvement_df, ax=ax)
plt.ylabel('Thompson Estimates of $y_{max}$')
larger_font('$E$($\max$(Thompson Estimate - $y_{max}^i$, 0)')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/sazio/NMAs/blob/main/Data_Loader.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Exploratory Data Analysis of Stringer Dataset
@authors: Simone Azeglio, Chetan Dhulipalla , Khalid Saifullah
Part of the code here has been taken from [Neuromatch Academy's Computational Neuroscience Course](https://compneuro.neuromatch.io/projects/neurons/README.html), and specifically from [this notebook](https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/projects/neurons/load_stringer_spontaneous.ipynb)
# to do list
1. custom normalization: dividing by mean value per neuron
1a. downsampling: convolve then downsample by 5
2. training validation split: withhold last 20 percent of time series for testing
3. RNN for each layer: a way to capture the dynamics inside each layer instead of capturing extra dynamics from inter-layer interactions. it will be OK to compare the different RNNs. maintain same neuron count in each layer to reduce potential bias
4. layer weight regularization: L2
5. early stopping , dropout?
```
#%%capture
#!pip install wandb --upgrade --quiet
import wandb
wandb.login()
```
## Loading of Stringer spontaneous data
```
#@title Data retrieval
import os, requests
fname = "stringer_spontaneous.npy"
url = "https://osf.io/dpqaj/download"
if not os.path.isfile(fname):
try:
r = requests.get(url)
except requests.ConnectionError:
print("!!! Failed to download data !!!")
else:
if r.status_code != requests.codes.ok:
print("!!! Failed to download data !!!")
else:
with open(fname, "wb") as fid:
fid.write(r.content)
#@title Import matplotlib and set defaults
from matplotlib import rcParams
from matplotlib import pyplot as plt
rcParams['figure.figsize'] = [20, 4]
rcParams['font.size'] =15
rcParams['axes.spines.top'] = False
rcParams['axes.spines.right'] = False
rcParams['figure.autolayout'] = True
```
## Exploratory Data Analysis (EDA)
```
#@title Data loading
import numpy as np
dat = np.load('stringer_spontaneous.npy', allow_pickle=True).item()
print(dat.keys())
# functions
def moving_avg(array, factor = 5):
"""Reducing the number of compontents by averaging of N = factor
subsequent elements of array"""
zeros_ = np.zeros((array.shape[0], 2))
array = np.hstack((array, zeros_))
array = np.reshape(array, (array.shape[0], int(array.shape[1]/factor), factor))
array = np.mean(array, axis = 2)
return array
```
## Extracting Data for RNN (or LFADS)
The first problem to address is that for each layer we don't have the exact same number of neurons. We'd like to have a single RNN encoding all the different layers activities, to make it easier we can take the number of neurons ($N_{neurons} = 1131$ of the least represented class (layer) and level out each remaining class.
```
# Extract labels from z - coordinate
from sklearn import preprocessing
x, y, z = dat['xyz']
le = preprocessing.LabelEncoder()
labels = le.fit_transform(z)
### least represented class (layer with less neurons)
n_samples = np.histogram(labels, bins=9)[0][-1]
### Data for LFADS / RNN
import pandas as pd
dataSet = pd.DataFrame(dat["sresp"])
dataSet["label"] = labels
# it can be done in one loop ...
data_ = []
for i in range(0, 9):
data_.append(dataSet[dataSet["label"] == i].sample(n = n_samples).iloc[:,:-1].values)
dataRNN = np.zeros((n_samples*9, dataSet.shape[1]-1))
for i in range(0,9):
# dataRNN[n_samples*i:n_samples*(i+1), :] = data_[i]
## normalized by layer
dataRNN[n_samples*i:n_samples*(i+1), :] = data_[i]/np.mean(np.asarray(data_)[i,:,:], axis = 0)
## shuffling for training purposes
#np.random.shuffle(dataRNN)
#unshuffled = np.array(data_)
#@title Convolutions code
# convolution moving average
# kernel_length = 50
# averaging_kernel = np.ones(kernel_length) / kernel_length
# dataRNN.shape
# avgd_dataRNN = list()
# for neuron in dataRNN:
# avgd_dataRNN.append(np.convolve(neuron, averaging_kernel))
# avg_dataRNN = np.array(avgd_dataRNN)
# print(avg_dataRNN.shape)
# @title Z Score Code
# from scipy.stats import zscore
# neuron = 500
# scaled_all = zscore(avg_dataRNN)
# scaled_per_neuron = zscore(avg_dataRNN[neuron, :])
# scaled_per_layer = list()
# for layer in unshuffled:
# scaled_per_layer.append(zscore(layer))
# scaled_per_layer = np.array(scaled_per_layer)
# plt.plot(avg_dataRNN[neuron, :])
# plt.plot(avg_dataRNN[2500, :])
# plt.figure()
# plt.plot(dataRNN[neuron, :])
# plt.figure()
# plt.plot(scaled_all[neuron, :])
# plt.plot(scaled_per_neuron)
# plt.figure()
# plt.plot(scaled_per_layer[0,neuron,:])
# custom normalization
normed_dataRNN = list()
for neuron in dataRNN:
normed_dataRNN.append(neuron)# / neuron.mean())
normed_dataRNN = np.array(normed_dataRNN)
# downsampling and averaging
#avgd_normed_dataRNN = dataRNN#
avgd_normed_dataRNN = dataRNN #moving_avg(dataRNN, factor=2)
plt.plot(avgd_normed_dataRNN[0,:])
```
issue: does the individual scaling by layer introduce bias that may artificially increase performance of the network?
## Data Loader
```
import torch
import torch.nn as nn
import torch.nn.functional as F
torch.cuda.empty_cache()
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# set the seed
np.random.seed(42)
# number of neurons
NN = dataRNN.shape[0]
# swapping the axes to maintain consistency with seq2seq notebook in the following code - the network takes all the neurons at a time step as input, not just one neuron
# avgd_normed_dataRNN = np.swapaxes(avgd_normed_dataRNN, 0, 1)
avgd_normed_dataRNN.shape
frac = 4/5
#x1 = torch.from_numpy(dataRNN[:,:int(frac*dataRNN.shape[1])]).to(device).float().unsqueeze(0)
#x2 = torch.from_numpy(dataRNN[:,int(frac*dataRNN.shape[1]):]).to(device).float().unsqueeze(0)
#x1 = torch.from_numpy(avgd_normed_dataRNN[:1131,:]).to(device).float().unsqueeze(2)
#x2 = torch.from_numpy(avgd_normed_dataRNN[:1131,:]).to(device).float().unsqueeze(2)
n_neurs = 1131
# let's use n_neurs/10 latent components
ncomp = int(n_neurs/10)
x1_train = torch.from_numpy(avgd_normed_dataRNN[:n_neurs,:int(frac*avgd_normed_dataRNN.shape[1])]).to(device).float().unsqueeze(2)
x2_train = torch.from_numpy(avgd_normed_dataRNN[:n_neurs,:int(frac*avgd_normed_dataRNN.shape[1])]).to(device).float().unsqueeze(2)
x1_valid = torch.from_numpy(avgd_normed_dataRNN[:n_neurs,int(frac*avgd_normed_dataRNN.shape[1]):]).to(device).float().unsqueeze(2)
x2_valid = torch.from_numpy(avgd_normed_dataRNN[:n_neurs,int(frac*avgd_normed_dataRNN.shape[1]):]).to(device).float().unsqueeze(2)
NN1 = x1_train.shape[0]
NN2 = x2_train.shape[0]
class Net(nn.Module):
def __init__(self, ncomp, NN1, NN2, num_layers = 1, n_comp = 50, dropout= 0, bidi=True):
super(Net, self).__init__()
# play with some of the options in the RNN!
self.rnn = nn.LSTM(NN1, ncomp, num_layers = num_layers, dropout = dropout,
bidirectional = bidi)
"""
self.rnn = nn.RNN(NN1, ncomp, num_layers = 1, dropout = 0,
bidirectional = bidi, nonlinearity = 'tanh')
self.rnn = nn.GRU(NN1, ncomp, num_layers = 1, dropout = 0,
bidirectional = bidi)
"""
self.mlp = nn.Sequential(
nn.Linear(ncomp, ncomp*2),
nn.Mish(),
nn.Linear(ncomp*2, ncomp*2),
nn.Mish(),
nn.Dropout(0.25),
nn.Linear(ncomp*2, ncomp),
nn.Mish())
self.fc = nn.Linear(ncomp, NN2)
def forward(self, x):
x = x.permute(1, 2, 0)
#print(x.shape)
# h_0 = torch.zeros(2, x.size()[1], self.ncomp).to(device)
y, h_n = self.rnn(x)
#print(y.shape)
#print(h_n.shape)
if self.rnn.bidirectional:
# if the rnn is bidirectional, it concatenates the activations from the forward and backward pass
# we want to add them instead, so as to enforce the latents to match between the forward and backward pass
q = (y[:, :, :ncomp] + y[:, :, ncomp:])/2
else:
q = y
q = self.mlp(q)
# the softplus function is just like a relu but it's smoothed out so we can't predict 0
# if we predict 0 and there was a spike, that's an instant Inf in the Poisson log-likelihood which leads to failure
#z = F.softplus(self.fc(q), 10)
#print(q.shape)
z = self.fc(q).permute(2, 0, 1)
# print(z.shape)
return z, q
sweep_config = {
'method': 'random'
}
metric = {
'name': 'loss',
'goal': 'minimize'
}
sweep_config['metric'] = metric
parameters_dict = {
'optimizer': {
'values': ['adam']
},
'num_layers': {
'values': [1]
},
'n_comp': {
'values': [100, 200, 300, 400, 500]#[50, 75, 100, 125, 150]
},
'dropout': {
'values': [0.1, 0.2, 0.3, 0.5]
},
'weight_decay': {
'values': [0., 1e-5, 5e-5, 1e-6]
},
}
sweep_config['parameters'] = parameters_dict
parameters_dict.update({
'epochs': {
'value': 4000}
})
import math
parameters_dict.update({
'learning_rate': {
# a flat distribution between 0 and 0.1
'distribution': 'log_uniform',
'min': -9.9,
'max': -5.3
},
})
import pprint
pprint.pprint(sweep_config)
sweep_id = wandb.sweep(sweep_config, project="NMAs-Full-Signals")
# you can keep re-running this cell if you think the cost might decrease further
cost = nn.MSELoss()
# rnn_loss = 0.2372, lstm_loss = 0.2340, gru_lstm = 0.2370
```
## Training
```
from tqdm import tqdm
def build_optimizer(network, optimizer, learning_rate, weight_decay):
optimizer = torch.optim.Adam(network.parameters(),
lr=learning_rate, weight_decay=weight_decay)
return optimizer
def train(config=None):
# Initialize a new wandb run
with wandb.init(config=config):
# If called by wandb.agent, as below,
# this config will be set by Sweep Controller
config = wandb.config
# loader = build_dataset(config.batch_size)
# Net(ncomp, NN1, NN2, bidi = True).to(device)
network = Net(ncomp, NN1, NN2, config.num_layers, config.dropout).to(device)
optimizer = build_optimizer(network, config.optimizer, config.learning_rate, config.weight_decay)
for epoch in range(config.epochs):
# avg_loss = train_epoch(network, loader, optimizer)
network.train()
# the networkwork outputs the single-neuron prediction and the latents
z, y = network(x1_train)
# our cost
loss = cost(z, x2_train)
# train the networkwork as usual
loss.backward()
optimizer.step()
optimizer.zero_grad()
with torch.no_grad():
network.eval()
valid_loss = cost(network(x1_valid)[0], x2_valid)
if epoch % 50 == 0:
with torch.no_grad():
network.eval()
valid_loss = cost(network(x1_valid)[0], x2_valid)
print(f' iteration {epoch}, train cost {loss.item():.4f}, valid cost {valid_loss.item():.4f}')
wandb.log({"train_loss": loss.item(), 'valid_loss': valid_loss.item(), "epoch": epoch})
wandb.agent(sweep_id, train, count= 50)
```
| github_jupyter |
```
#export
from fastai2.torch_basics import *
from fastai2.data.all import *
from fastai2.text.core import *
from nbdev.showdoc import *
#default_exp text.data
#default_cls_lvl 3
```
# Text data
> Functions and transforms to help gather text data in a `DataSource`
## Numericalizing
```
#export
def make_vocab(count, min_freq=3, max_vocab=60000):
"Create a vocab of `max_vocab` size from `Counter` `count` with items present more than `min_freq`"
vocab = [o for o,c in count.most_common(max_vocab) if c >= min_freq]
for o in reversed(defaults.text_spec_tok): #Make sure all special tokens are in the vocab
if o in vocab: vocab.remove(o)
vocab.insert(0, o)
vocab = vocab[:max_vocab]
return vocab + [f'xxfake' for i in range(0, 8-len(vocab)%8)]
count = Counter(['a', 'a', 'a', 'a', 'b', 'b', 'c', 'c', 'd'])
test_eq(set([x for x in make_vocab(count) if not x.startswith('xxfake')]),
set(defaults.text_spec_tok + 'a'.split()))
test_eq(len(make_vocab(count))%8, 0)
test_eq(set([x for x in make_vocab(count, min_freq=1) if not x.startswith('xxfake')]),
set(defaults.text_spec_tok + 'a b c d'.split()))
test_eq(set([x for x in make_vocab(count,max_vocab=12, min_freq=1) if not x.startswith('xxfake')]),
set(defaults.text_spec_tok + 'a b c'.split()))
#export
class TensorText(TensorBase): pass
class LMTensorText(TensorText): pass
# export
class Numericalize(Transform):
"Reversible transform of tokenized texts to numericalized ids"
def __init__(self, vocab=None, min_freq=3, max_vocab=60000, sep=' '):
self.vocab,self.min_freq,self.max_vocab,self.sep = vocab,min_freq,max_vocab,sep
self.o2i = None if vocab is None else defaultdict(int, {v:k for k,v in enumerate(vocab)})
def setups(self, dsrc):
if dsrc is None: return
if self.vocab is None:
count = dsrc.counter if hasattr(dsrc, 'counter') else Counter(p for o in dsrc for p in o)
self.vocab = make_vocab(count, min_freq=self.min_freq, max_vocab=self.max_vocab)
self.o2i = defaultdict(int, {v:k for k,v in enumerate(self.vocab) if v != 'xxfake'})
def encodes(self, o): return TensorText(tensor([self.o2i [o_] for o_ in o]))
def decodes(self, o): return TitledStr(self.sep.join([self.vocab[o_] for o_ in o if self.vocab[o_] != PAD]))
num = Numericalize(min_freq=1, sep=' ')
num.setup(L('This is an example of text'.split(), 'this is another text'.split()))
test_eq(set([x for x in num.vocab if not x.startswith('xxfake')]),
set(defaults.text_spec_tok + 'This is an example of text this another'.split()))
test_eq(len(num.vocab)%8, 0)
start = 'This is an example of text'
t = num(start.split())
test_eq(t, tensor([11, 9, 12, 13, 14, 10]))
test_eq(num.decode(t), start)
num = Numericalize(min_freq=2, sep=' ')
num.setup(L('This is an example of text'.split(), 'this is another text'.split()))
test_eq(set([x for x in num.vocab if not x.startswith('xxfake')]),
set(defaults.text_spec_tok + 'is text'.split()))
test_eq(len(num.vocab)%8, 0)
t = num(start.split())
test_eq(t, tensor([0, 9, 0, 0, 0, 10]))
test_eq(num.decode(t), f'{UNK} is {UNK} {UNK} {UNK} text')
#hide
df = pd.DataFrame({'texts': ['This is an example of text', 'this is another text']})
tl = TfmdList(df, [attrgetter('text'), Tokenizer.from_df('texts'), Numericalize(min_freq=2, sep=' ')])
test_eq(tl, [tensor([2, 8, 9, 10, 0, 0, 0, 11]), tensor([2, 9, 10, 0, 11])])
```
## LM_DataLoader -
```
#export
def _maybe_first(o): return o[0] if isinstance(o, tuple) else o
#export
#TODO: add backward
@delegates()
class LMDataLoader(TfmdDL):
def __init__(self, dataset, lens=None, cache=2, bs=64, seq_len=72, num_workers=0, **kwargs):
self.items = ReindexCollection(dataset, cache=cache, tfm=_maybe_first)
self.seq_len = seq_len
if lens is None: lens = [len(o) for o in self.items]
self.lens = ReindexCollection(lens, idxs=self.items.idxs)
# The "-1" is to allow for final label, we throw away the end that's less than bs
corpus = round_multiple(sum(lens)-1, bs, round_down=True)
self.bl = corpus//bs #bl stands for batch length
self.n_batches = self.bl//(seq_len) + int(self.bl%seq_len!=0)
self.last_len = self.bl - (self.n_batches-1)*seq_len
self.make_chunks()
super().__init__(dataset=dataset, bs=bs, num_workers=num_workers, **kwargs)
self.n = self.n_batches*bs
def make_chunks(self): self.chunks = Chunks(self.items, self.lens)
def shuffle_fn(self,idxs):
self.items.shuffle()
self.make_chunks()
return idxs
def create_item(self, seq):
if seq>=self.n: raise IndexError
sl = self.last_len if seq//self.bs==self.n_batches-1 else self.seq_len
st = (seq%self.bs)*self.bl + (seq//self.bs)*self.seq_len
txt = self.chunks[st : st+sl+1]
return LMTensorText(txt[:-1]),txt[1:]
#hide
bs,sl = 4,3
ints = L([0,1,2,3,4],[5,6,7,8,9,10],[11,12,13,14,15,16,17,18],[19,20],[21,22]).map(tensor)
dl = LMDataLoader(ints, bs=bs, seq_len=sl)
list(dl)
test_eq(list(dl),
[[tensor([[0, 1, 2], [5, 6, 7], [10, 11, 12], [15, 16, 17]]),
tensor([[1, 2, 3], [6, 7, 8], [11, 12, 13], [16, 17, 18]])],
[tensor([[3, 4], [8, 9], [13, 14], [18, 19]]),
tensor([[4, 5], [9, 10], [14, 15], [19, 20]])]])
bs,sl = 4,3
ints = L([0,1,2,3,4],[5,6,7,8,9,10],[11,12,13,14,15,16,17,18],[19,20],[21,22,23],[24]).map(tensor)
dl = LMDataLoader(ints, bs=bs, seq_len=sl)
test_eq(list(dl),
[[tensor([[0, 1, 2], [6, 7, 8], [12, 13, 14], [18, 19, 20]]),
tensor([[1, 2, 3], [7, 8, 9], [13, 14, 15], [19, 20, 21]])],
[tensor([[3, 4, 5], [ 9, 10, 11], [15, 16, 17], [21, 22, 23]]),
tensor([[4, 5, 6], [10, 11, 12], [16, 17, 18], [22, 23, 24]])]])
#hide
#Check lens work
dl = LMDataLoader(ints, lens=ints.map(len), bs=bs, seq_len=sl)
test_eq(list(dl),
[[tensor([[0, 1, 2], [6, 7, 8], [12, 13, 14], [18, 19, 20]]),
tensor([[1, 2, 3], [7, 8, 9], [13, 14, 15], [19, 20, 21]])],
[tensor([[3, 4, 5], [ 9, 10, 11], [15, 16, 17], [21, 22, 23]]),
tensor([[4, 5, 6], [10, 11, 12], [16, 17, 18], [22, 23, 24]])]])
dl = LMDataLoader(ints, bs=bs, seq_len=sl, shuffle=True)
for x,y in dl: test_eq(x[:,1:], y[:,:-1])
((x0,y0), (x1,y1)) = tuple(dl)
#Second batch begins where first batch ended
test_eq(y0[:,-1], x1[:,0])
test_eq(type(x0), LMTensorText)
```
### Showing
```
#export
@patch
def truncate(self:TitledStr, n):
words = self.split(' ')[:n]
return TitledStr(' '.join(words))
#export
@typedispatch
def show_batch(x: TensorText, y, samples, ctxs=None, max_n=10, trunc_at=150, **kwargs):
if ctxs is None: ctxs = get_empty_df(min(len(samples), max_n))
samples = L((s[0].truncate(trunc_at),*s[1:]) for s in samples)
ctxs = show_batch[object](x, y, samples, max_n=max_n, ctxs=ctxs, **kwargs)
display_df(pd.DataFrame(ctxs))
return ctxs
#export
@typedispatch
def show_batch(x: LMTensorText, y, samples, ctxs=None, max_n=10, **kwargs):
return show_batch[TensorText](x, None, samples, ctxs=ctxs, max_n=max_n, **kwargs)
```
## Integration example
```
path = untar_data(URLs.IMDB_SAMPLE)
df = pd.read_csv(path/'texts.csv')
df.head(2)
splits = ColSplitter()(df)
tfms = [attrgetter('text'), Tokenizer.from_df('text'), Numericalize()]
dsrc = DataSource(df, [tfms], splits=splits, dl_type=LMDataLoader)
dbunch = dsrc.databunch(bs=16, seq_len=72)
dbunch.show_batch(max_n=6)
b = dbunch.one_batch()
test_eq(type(x), LMTensorText)
test_eq(len(dbunch.valid_ds[0][0]), dbunch.valid_dl.lens[0])
```
## Classification
```
#export
def pad_input(samples, pad_idx=1, pad_fields=0, pad_first=False, backwards=False):
"Function that collect samples and adds padding. Flips token order if needed"
pad_fields = L(pad_fields)
max_len_l = pad_fields.map(lambda f: max([len(s[f]) for s in samples]))
if backwards: pad_first = not pad_first
def _f(field_idx, x):
if field_idx not in pad_fields: return x
idx = pad_fields.items.index(field_idx) #TODO: remove items if L.index is fixed
sl = slice(-len(x), sys.maxsize) if pad_first else slice(0, len(x))
pad = x.new_zeros(max_len_l[idx]-x.shape[0])+pad_idx
x1 = torch.cat([pad, x] if pad_first else [x, pad])
if backwards: x1 = x1.flip(0)
return retain_type(x1, x)
return [tuple(map(lambda idxx: _f(*idxx), enumerate(s))) for s in samples]
test_eq(pad_input([(tensor([1,2,3]),1), (tensor([4,5]), 2), (tensor([6]), 3)], pad_idx=0),
[(tensor([1,2,3]),1), (tensor([4,5,0]),2), (tensor([6,0,0]), 3)])
test_eq(pad_input([(tensor([1,2,3]), (tensor([6]))), (tensor([4,5]), tensor([4,5])), (tensor([6]), (tensor([1,2,3])))], pad_idx=0, pad_fields=1),
[(tensor([1,2,3]),(tensor([6,0,0]))), (tensor([4,5]),tensor([4,5,0])), ((tensor([6]),tensor([1, 2, 3])))])
test_eq(pad_input([(tensor([1,2,3]),1), (tensor([4,5]), 2), (tensor([6]), 3)], pad_idx=0, pad_first=True),
[(tensor([1,2,3]),1), (tensor([0,4,5]),2), (tensor([0,0,6]), 3)])
test_eq(pad_input([(tensor([1,2,3]),1), (tensor([4,5]), 2), (tensor([6]), 3)], pad_idx=0, backwards=True),
[(tensor([3,2,1]),1), (tensor([5,4,0]),2), (tensor([6,0,0]), 3)])
x = test_eq(pad_input([(tensor([1,2,3]),1), (tensor([4,5]), 2), (tensor([6]), 3)], pad_idx=0, backwards=True),
[(tensor([3,2,1]),1), (tensor([5,4,0]),2), (tensor([6,0,0]), 3)])
#hide
#Check retain type
x = [(TensorText([1,2,3]),1), (TensorText([4,5]), 2), (TensorText([6]), 3)]
y = pad_input(x, pad_idx=0)
for s in y: test_eq(type(s[0]), TensorText)
#export
def _default_sort(x): return len(x[0])
@delegates(TfmdDL)
class SortedDL(TfmdDL):
def __init__(self, dataset, sort_func=None, res=None, **kwargs):
super().__init__(dataset, **kwargs)
self.sort_func = _default_sort if sort_func is None else sort_func
self.res = [self.sort_func(self.do_item(i)) for i in range_of(self.dataset)] if res is None else res
self.idx_max = np.argmax(self.res)
def get_idxs(self):
idxs = super().get_idxs()
if self.shuffle: return idxs
return sorted(idxs, key=lambda i: self.res[i], reverse=True)
def shuffle_fn(self,idxs):
idxs = np.random.permutation(len(self.dataset))
idx_max = np.where(idxs==self.idx_max)[0][0]
idxs[0],idxs[idx_max] = idxs[idx_max],idxs[0]
sz = self.bs*50
chunks = [idxs[i:i+sz] for i in range(0, len(idxs), sz)]
chunks = [sorted(s, key=lambda i: self.res[i], reverse=True) for s in chunks]
sort_idx = np.concatenate(chunks)
sz = self.bs
batches = [sort_idx[i:i+sz] for i in range(0, len(sort_idx), sz)]
sort_idx = np.concatenate(np.random.permutation(batches[1:-1])) if len(batches) > 2 else np.array([],dtype=np.int)
sort_idx = np.concatenate((batches[0], sort_idx) if len(batches)==1 else (batches[0], sort_idx, batches[-1]))
return iter(sort_idx)
ds = [(tensor([1,2]),1), (tensor([3,4,5,6]),2), (tensor([7]),3), (tensor([8,9,10]),4)]
dl = SortedDL(ds, bs=2, before_batch=partial(pad_input, pad_idx=0))
test_eq(list(dl), [(tensor([[ 3, 4, 5, 6], [ 8, 9, 10, 0]]), tensor([2, 4])),
(tensor([[1, 2], [7, 0]]), tensor([1, 3]))])
ds = [(tensor(range(random.randint(1,10))),i) for i in range(101)]
dl = SortedDL(ds, bs=2, create_batch=partial(pad_input, pad_idx=-1), shuffle=True, num_workers=0)
batches = list(dl)
max_len = len(batches[0][0])
for b in batches:
assert(len(b[0])) <= max_len
test_ne(b[0][-1], -1)
splits = RandomSplitter()(range_of(df))
dsrc = DataSource(df, splits=splits, tfms=[tfms, [attrgetter("label"), Categorize()]], dl_type=SortedDL)
dbch = dsrc.databunch(before_batch=pad_input)
dbch.show_batch(max_n=2)
```
## TransformBlock for text
```
#export
class TextBlock(TransformBlock):
def __init__(self, tok_tfm, vocab=None, is_lm=False):
return super().__init__(type_tfms=[tok_tfm, Numericalize(vocab)],
dl_type=LMDataLoader if is_lm else SortedDL,
dbunch_kwargs={} if is_lm else {'before_batch': pad_input})
@classmethod
@delegates(Tokenizer.from_df, keep=True)
def from_df(cls, text_cols, vocab=None, is_lm=False, **kwargs):
return cls(Tokenizer.from_df(text_cols, **kwargs), vocab=vocab, is_lm=is_lm)
@classmethod
@delegates(Tokenizer.from_folder, keep=True)
def from_folder(cls, path, vocab=None, is_lm=False, **kwargs):
return cls(Tokenizer.from_folder(path, **kwargs), vocab=vocab, is_lm=is_lm)
```
## TextDataBunch -
```
#export
class TextDataBunch(DataBunch):
@classmethod
@delegates(DataBunch.from_dblock)
def from_folder(cls, path, train='train', valid='valid', valid_pct=None, seed=None, vocab=None, text_vocab=None, is_lm=False,
tok_tfm=None, **kwargs):
"Create from imagenet style dataset in `path` with `train`,`valid`,`test` subfolders (or provide `valid_pct`)."
splitter = GrandparentSplitter(train_name=train, valid_name=valid) if valid_pct is None else RandomSplitter(valid_pct, seed=seed)
blocks = [TextBlock.from_folder(path, text_vocab, is_lm) if tok_tfm is None else TextBlock(tok_tfm, text_vocab, is_lm)]
if not is_lm: blocks.append(CategoryBlock(vocab=vocab))
get_items = partial(get_text_files, folders=[train,valid]) if valid_pct is None else get_text_files
dblock = DataBlock(blocks=blocks,
get_items=get_items,
splitter=splitter,
get_y=None if is_lm else parent_label)
return cls.from_dblock(dblock, path, path=path, **kwargs)
@classmethod
@delegates(DataBunch.from_dblock)
def from_df(cls, df, path='.', valid_pct=0.2, seed=None, text_col=0, label_col=1, label_delim=None, y_block=None,
text_vocab=None, is_lm=False, valid_col=None, tok_tfm=None, **kwargs):
blocks = [TextBlock.from_df(text_col, text_vocab, is_lm) if tok_tfm is None else TextBlock(tok_tfm, text_vocab, is_lm)]
if y_block is None and not is_lm:
blocks.append(MultiCategoryBlock if is_listy(label_col) and len(label_col) > 1 else CategoryBlock)
if y_block is not None and not is_lm: blocks += (y_block if is_listy(y_block) else [y_block])
splitter = RandomSplitter(valid_pct, seed=seed) if valid_col is None else ColSplitter(valid_col)
dblock = DataBlock(blocks=blocks,
get_x=ColReader(text_col),
get_y=None if is_lm else ColReader(label_col, label_delim=label_delim),
splitter=splitter)
return cls.from_dblock(dblock, df, path=path, **kwargs)
@classmethod
def from_csv(cls, path, csv_fname='labels.csv', header='infer', delimiter=None, **kwargs):
df = pd.read_csv(Path(path)/csv_fname, header=header, delimiter=delimiter)
return cls.from_df(df, path=path, **kwargs)
TextDataBunch.from_csv = delegates(to=TextDataBunch.from_df)(TextDataBunch.from_csv)
```
## Export -
```
#hide
from nbdev.export import notebook2script
notebook2script()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/JoanYego/Supervised-and-Unsupervised-Learning-in-R/blob/main/Ad_clicks.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Defining the Research Problem
## Specifying the Research Question
The goal of this analysis is to predict individuals who are most likely to click on a cryptography course advertisement. This project uses R for analysis.
## Defining the Metric for Success
The project will be considered a success when we are able to clean and analyse past data to segment blog users and predict individuals who should be targeted for an advertisement.
## Understanding the Context
A Kenyan entrepreneur has created an online cryptography course and would want to advertise it on her blog. She currently targets audiences originating from various countries. In the past, she ran ads to advertise a related course on the same blog and collected data in the process. She has employed my services as a Data Science Consultant to help her identify which individuals are most likely to click on her ads.
## Recording the Exprimental Design
Below are the steps that will be followed in this analysis in order to respond to the research question satisfactorily:
>* Read the Data
>* Check the Data
>* Data Cleaning
>* Univariate Analysis
>* Bivariate Analysis
>* Implementing the Solution (Modeling)
>* Conclusion and Recommendation
## Data Relevance
The data used for the project was collected from a prior advertisement for a similar course on the same platform. The dataset contains 10 attributes and 1,000 records. These attributes contain descriptive information of past blog users. Some of the attributes include country, age and gender of the user among others.
# Importing Relevant Libraries
```
# Installing data.table package
install.packages("data.table", dependencies=TRUE)
```
# Reading the Data
```
# Reading the data into R from the csv file
library(data.table)
ad <- read.csv('advertising.csv')
head(ad)
```
# Checking the Data
```
# Checking the top 6 records
head(ad)
# Checking the bottom 6 records
tail(ad)
# Checking the total number of records
nrow(ad)
# Checking the total number of columns
ncol(ad)
# Checking all column names
names(ad)
# Checking the data types of each column
str(ad)
# Checking the number of unique values in each column
lengths(lapply(ad, unique))
```
- There are 969 unique cities and 237 unique countries in the dataset.
- There are only 43 unique ages in the dataset.
```
# Checking the summary of the data
summary(ad)
```
# Data Cleaning
Missing Data
```
# Checking the existence of missing values
colSums(is.na(ad))
```
- No missing values in any columns of the dataframe
Outliers
```
# creating a variable with only numeric columns
library(tidyverse)
my_data <- ad %>% select(1,2,3,4,7,10)
# Previewing outliers for numeric columns using boxplots
boxplot(my_data)
```
- We see that 'area income' is the only attribute with outliers. We shall investigate each column individually for further analysis.
```
# Boxplot for daily time spent variable
boxplot(ad$Daily.Time.Spent.on.Site)
# Boxplot for age variable
boxplot(ad$Age)
# Boxplot for daily internet usage variable
boxplot(ad$Daily.Internet.Usage)
# Boxplot for area income variable
boxplot(ad$Area.Income)
```
- From the above graphs, no other columns have outliers except the 'area income' attribute.
```
# Displaying all outliers in the income column
boxplot.stats(ad$Area.Income)$out
# Checking the countries associated with outlier incomes
ad$Country[ad$Area.Income %in% c(17709.98, 18819.34, 15598.29, 15879.1, 14548.06, 13996.5, 14775.5, 18368.57)]
```
- We observe that the really low 'outlier' income numbers are associated with developing countries. This is consistent with observations in the real world therefore we will keep the ouliers.
Anomalies
```
# Checking for duplicate data
duplicated_rows <- ad[duplicated(ad),]
duplicated_rows
```
- No duplicate records in the dataset
# Exploratory Data Analysis
## Univariate Analysis
- In this section, we will investigate each variable individually. The steps here include calculating and interpreting measures of central tendency (mode, median, mean) as well as computing and explaining the range, the interquartile range, the standard deviation, variance, skewness, and kurtosis
```
# Calculating the mean for all numeric columns
lapply(my_data,FUN=mean)
```
- Average age of the blog users is 36 while the average income is 55,000.
```
# Calculating the median for all numeric columns
lapply(my_data,FUN=median)
# Calculating the mode for all numeric columns
getmode <- function(v) {
uniqv <- unique(v)
uniqv[which.max(tabulate(match(v, uniqv)))]
}
lapply(my_data,FUN=getmode)
```
- Most occuring age is 31 and the median age is 35.
- Most occuring gender is female.
```
# Calculating the minimum value for all numeric columns
lapply(my_data,FUN=min)
# Calculating the maximum value for all numeric columns
lapply(my_data,FUN=max)
```
- Lowest income is ~14,000 while the highest is ~79,500
- The youngest age is 19 and the oldest blog user's age is 61
```
# Checking the range for all numeric columns
lapply(my_data,FUN=range)
# Calculating the quantiles for all numeric columns
lapply(my_data,FUN=quantile)
# Calculating the variance for all numeric columns
lapply(my_data,FUN=var)
# Calculating the standard deviation for all numeric columns
lapply(my_data,FUN=sd)
# Plotting a histogram for age variable
hist(ad$Age)
```
- The frequency distribution above depicts a relatively normal distribution for the age attribute. Most individuals' age is centered around the mean.
```
# Plotting a histogram for area income variable
hist(ad$Area.Income)
```
- Income distribution is skewed to the left.
```
# Plotting a histogram for daily time variable
hist(ad$Daily.Time.Spent.on.Site)
# Plotting a histogram for daily internet variable
hist(ad$Daily.Internet.Usage)
# Plotting a histogram for gender variable
hist(ad$Male)
```
- The number of males and females is fairly balanced.
```
# Plotting a histogram for clicked ad variable
hist(ad$Clicked.on.Ad)
```
- The target variable for this analysis has equal observations for both classes.
```
# Checking actual number of male vs females
table(ad$Male)
# Confirming distribution of classes
table(ad$Clicked.on.Ad)
# Bar plot of the age variable
age <- ad$Age
age_freq <- table(age)
barplot(age_freq)
# Checking distribution of each country
table(ad$Country)
```
## Bivariate Analysis
In this section, we investigate the relationship of different variables by creating relevant visualizations such asscatter plots, correlation matrix and Pearson correlation coefficient.
```
# Checking the correlation coefficients for numeric variables
install.packages("ggcorrplot")
library(ggcorrplot)
corr = round(cor(select_if(my_data, is.numeric)), 2)
ggcorrplot(corr, hc.order = T, ggtheme = ggplot2::theme_gray,
colors = c("#6D9EC1", "white", "#E46726"), lab = T)
```
- There's a relatively strong negative correlation between daily internet usage, area income, daily time spent on site vs clicked on ad.
```
# Scatter plot to compare age vs income
plot(ad$Age, ad$Area.Income, xlab="Age", ylab="Income")
```
- Most high income individuals are between the ages of 30 to 50.
```
# Scatter plot to compare income vs Clicked on ad
plot(ad$Clicked.on.Ad, ad$Area.Income, xlab="Clicked on Ad", ylab="Income")
```
- Most low income individuals clicked on the ad.
```
# Scatter plot to compare age vs daily time spent
plot(ad$Age, ad$Daily.Time.Spent.on.Site, xlab="Age", ylab="Time Spent")
# Scatter plot to compare clicked on ad vs time spent
plot(ad$Clicked.on.Ad, ad$Daily.Time.Spent.on.Site, xlab="Clicked on Ad", ylab="Time Spent")
```
- Most users who spent the least amount of time on the blog clicked on the ad.
```
# Scatter plot to compare clicked on ad vs internet usage
plot(ad$Clicked.on.Ad, ad$Daily.Internet.Usage, xlab="Clicked on Ad", ylab="Internet Usage")
# Scatter plot to compare age vs Clicked on ad
plot(ad$Clicked.on.Ad, ad$`Age`, xlab="Clicked on Ad", ylab="Age")
```
# Implementing the Solution
## Decision Trees
```
# Importing relevant libraries
install.packages("rpart")
install.packages("rpart.plot")
install.packages("mlbench")
install.packages("caret")
library(rpart)
library(rpart.plot)
library(mlbench)
library(caret)
# Defining features and target variables
ad <- ad %>% select(1,2,3,4,7,8,10)
head(ad)
# Converting the country variable to numeric data type
ad$Country <- as.integer(as.factor(ad$Country))
# Normalizing relevant features
normalize <- function(x){
return ((x-min(x)) / (max(x)-min(x)))}
ad$Daily.Time.Spent.on.Site <- normalize(ad$Daily.Time.Spent.on.Site )
ad$Age <- normalize(ad$Age)
ad$Area.Income <- normalize(ad$Area.Income)
ad$Daily.Internet.Usage <- normalize(ad$Daily.Internet.Usage)
ad$Country <- normalize(ad$Country)
# Confirming the dimensions of the dataset
dim(ad)
# Creating the test and train sets. We can do a 800/200 split.
data_train <- ad[1:800, ]
data_test <- ad[801:1000,]
# Confirming the dimensions of the train and test sets
dim(data_train)
dim(data_test)
# Fitting the decision tree model
install.packages('tree')
library(tree)
model <- rpart(Clicked.on.Ad~., data = data_train, method = 'class')
rpart.plot(model)
# Visualizing the model
rpart.plot(model)
# Making a prediction on the test data
predict_unseen <-predict(fit, data_test, type = 'class')
table_mat <- table(data_test$Clicked.on.Ad, predict_unseen)
table_mat
```
- The model accurately predicted 85 users who did not click on the ad and 104 users who clicked on the ad. The total number of incorrect predictions is 11. This a fairly good prediction model.
```
# Calculating the accuracy score of the model
accuracy_score <- sum(diag(table_mat)) / sum(table_mat)
accuracy_score
```
- The model has an accuracy of 94.5%. This high prediction value is quite acceptable since we do not want the model to overfit.
Hyperparameter Tuning to Optimize the Model
```
# Adjusting the maximum depth as well as minimum sample of a node
accuracy_tune <- function(fit) {
predict_unseen <- predict(fit, data_test, type = 'class')
table_mat <- table(data_test$Clicked.on.Ad, predict_unseen)
accuracy_Test <- sum(diag(table_mat)) / sum(table_mat)
accuracy_Test
}
control <- rpart.control(minsplit = 4,
minbucket = round(5 / 3),
maxdepth = 3,
cp = 0)
tune_fit <- rpart(Clicked.on.Ad~., data = data_train, method = 'class', control = control)
accuracy_tune(tune_fit)
```
- Performing hyperparameter tuning improves the model performance slightly by 0.5%
# Conclusion and Recommendations
From the above analysis, below are some of the conclusions and recommendations that were obtained from data exploration (EDA):
- There is no need to have a time threshold before one can view the course advertisement on the blog. This is because most users who spent a relatively short time on the blog clicked on a previous similar ad.
- Users who are relatively older (above 50) are most likely to click on the ad. The advertisement can therefore target that age group more compared to other age groups.
- Low income areas should still be included in the target group of the ad since users in that income bracket are most likely to view the advertisement. Areas that experience low internet usage should also be included in the target audience.
In terms of predicting blog users who are likely to click on a course advertisement, Desicion Tree Classifier provides a descent model with a high prediction accuracy. However, other machine learning algorithms such as SVM and Random Forests can be investigated to further challenge the solution provided in this analysis.
| github_jupyter |
# 12-1 datetime 오브젝트
datetime 라이브러리는 날짜와 시간을 처리하는 등의 다양한 기능을 제공하는 파이썬 라이브러리입니다. datetime 라이브러리에는 날짜를 처리하는 date 오브젝트, 시간을 처리하는 time 오브젝트, 날짜와 시간을 모두 처리하는 datetime 오브젝트가 포함되어 있습니다. 앞으로 3개의 오브젝트를 명확히 구분하기 위해 영문을 그대로 살려 date, time, datetime 오브젝트라고 부르겠습니다.
### datetime 오브젝트 사용하기
#### 1.
datetime 오브젝트를 사용하기 위해 datetime 라이브러리를 불러옵니다.
```
from datetime import datetime
```
#### 2.
now, today 메서드를 사용하면 다음과 같이 현재 시간을 출력할 수 있습니다.
```
now1=datetime.now()
print(now1)
now2=datetime.today()
print(now2)
```
#### 3.
다음은 datetime 오브젝트를 생성할 때 시간을 직접 입력하여 인자로 전달한 것입니다. 각 변수를 출력하여 확인해 보면 입력한 시간을 바탕으로 datetime 오브젝트가 생성된 것을 알 수 있습니다.
```
t1=datetime.now()
t2=datetime(1970,1,1)
t3=datetime(1970,12,12,13,24,34)
print(t1)
print(t2)
print(t3)
```
#### 4.
datetime 오브젝트를 사용하는 이유 중 하나는 시간 계산을 할 수 있다는 점입니다. 다음은 두 datetime 오브젝트의 차이를 구한 것입니다.
```
diff1=t1-t2
print(diff1)
print(type(diff1))
diff2=t2-t1
print(diff2)
print(type(diff2))
```
## datetime 오브텍트로 변환하기 - to_datetime 메서드
경우에 따라서는 시게열 데이터를 문자열로 저장해야 할 때도 있습니다. 하지만 문자열은 시간 계산을 할 수 없기 때문에 datetime 오브젝트로 변환해 주어야 합니다. 이번에는 to_datetime 메서드를 사용하여 문자열을 datetime 오브젝트로 변환하는 방법에 대해 알아보겠습니다.
### 문자열 datetime 오브젝트로 변환하기
#### 1.
먼저 ebola 데이터 집합을 불러옵니다.
```
import pandas as pd
import os
ebola=pd.read_csv('data/country_timeseries.csv')
```
#### 2.
ebola 데이터프레임을 보면 문자열로 지정된 Date 열이 있는 것을 알 수 있습니다.
```
print(ebola.info())
```
#### 3.
to_datetime 메서드를 사용하면 Date 열의 자료형을 datetime 오브젝트로 변환할 수 있습니다. 다음과 같이 to_datetime 메서드를 사용하여 Date 열의 자료형을 datetime 오르젝트로 변환한 다음 ebola 데이터프레임에 새로운 열로 추가합니다.
```
ebola['date_dt']=pd.to_datetime(ebola['Date'])
print(ebola.info())
```
#### 4.
to_datetime 메서드를 좀 더 자세히 알아볼까요? 시간 형식 지정자와 기호를 적절히 조합하여 format 인자에 전달하면 그 형식에 맞게 정리된 datetime 오브젝트를 얻을 수 있습니다. 다음 실습을 참고하여 format 인자의 사용법을 꼭 익혀두세요.
```
test_df1=pd.DataFrame({'order_day':['01/01/15','02/01/15','03/01/15']})
test_df1['date_dt1']=pd.to_datetime(test_df1['order_day'],format='%d/%m/%y')
test_df1['date_dt2']=pd.to_datetime(test_df1['order_day'],format='%m/%d/%y')
test_df1['date_dt3']=pd.to_datetime(test_df1['order_day'],format='%y/%m/%d')
print(test_df1)
test_df2=pd.DataFrame({'order_day':['01-01-15','02-01-15','03-01-15']})
test_df2['date_dt']=pd.to_datetime(test_df2['order_day'],format='%d-%m-%y')
print(test_df2)
```
## 시간 형식 지정자
다음은 시간 형식 지정자를 정리한 표입니다. 이 장의 실습에서 종종 사용하므로 한 번 읽고 넘어가기 바랍니다.
### 시간 형식 지정자
- 시간 형식 지정자 : 의미 : 결과
- %a : 요일 출력 : Sun,Mon,...Set
- %A : 요일 출력(긴 이름) : Sunday,Monday, ...,Saturday
- %w : 요일 출력(숫자,0부터 일요일) : 0,1,....,6
- %d : 날짜 출력(2자리로 표시) : 01,02,....,31
- %b : 월 출력 : Jan,Feb,....Dec
- %B : 월 출력(긴 이름) : January,February,...December
- %m : 월 출력(숫자) : 01,02,...,12
- %y : 년 출력(2자리로 표시) : 00,01,...,99
- %Y : 년 출력(4자리로 표시) : 0001,0002,...,2013,2014,...,9999
- %H : 시간 출력(24시간) : 01,02,...,23
- %I : 시간 출력(12시간) : 01,02,...,12
- %p : AM 또는 PM 출력 : AM,PM
- %M : 분 출력(2자리로 표시) : 00,01,...,59
- %S : 초 출력(2자리로 표시) : 00,01,...,59
- %f : 마이크로초 출력 : 000000,000001,...,999999
- %z : UTC 차이 출력(+HHMM이나 -HHMM형태) : (None),+0000,-0400,+1030
- %Z : 기준 지역 이름 출력 : (None),UTC,EST,CST
- %j : 올해의 지난 일 수 출력(1일,2일,...) : 001,002,...,366
- %U : 올해의 지난 주 수 출력(1주,2주,...) : 00,01,...,53
- %c : 날짜와 시간 출력 : Tue Aug 16 21:30:00 1988
- %x : 날짜 출력 : 08/16/88(None);08/16/1988
- %X : 시간 출력 : 21:30:00
- %G : 년 출력(ISO 8601 형식) : 0001,0002,...,2013,2014,...,9999
- %u : 요일 출력(ISO 8601 형식) : 1,2,...,7
- %V : 올해의 지난 주 수 출력(ISO 8601 형식) : 01,02,...,53
### 시계열 데이터를 구분해서 추출하고 싶어요
now 메서드로 얻은 현재 시간의 시계열 데이터는 아주 정밀한 단위까지 시간을 표현합니다. 하지만 원하는 시계열 데이터의 시간 요소가 연도, 월, 일뿐이라면 now 메서드로 얻은 시계열 데이터를 잘라내야 합니다. 다음은 strftime 메서드와 시간 형식 지정자를 이용하여 시게열 데이터를 잘라낸 것입니다.
```
now=datetime.now()
print(now)
nowDate=now.strftime('%Y-%m-%d')
print(nowDate)
nowTime=now.strftime('%H:%M:%S')
print(nowTime)
nowDatetime=now.strftime('%Y-%m-%d %H:%M:%S')
print(nowDatetime)
```
## datetime 오브젝트로 변환하기 - read_csv 메서드
앞에서는 to_datetime 메서드를 사용하여 문자열로 저장되어 있는 Date 열을 datetime오브젝트로 변환했습니다. 하지만 datetime 오브젝트로 변환하려는 열을 지정하여 데이터 집합을 불러오는 것이 더 간단합니다. 다음 실습을 통해 알아보겠습니다.
### datetime 오브젝트로 변환하려는 열을 지정하여 데이터 집합 불러오기
#### 1.
다음은 read_csv 메서드의 parse_dates 인자에 datetime 오브젝트로 변환하고자 하는 열의 이름을 전달하여 데이터 집합을 불러온 것입니다. 결과를 보면 Date 열이 문자열이 아니라 datetime 오브젝트라는 것을 확인할 수 있습니다.
```
ebola1 =pd.read_csv('data/country_timeseries.csv',parse_dates=['Date'])
print(ebola1.info())
```
## datetime 오브젝트에서 날짜 정보 추출하기
datetime 오브젝트에 년,월,일과 같은 날짜 정보를 따로 저장하고 있는 속성이 이미 준비되어 있습니다. 다음 실습을 통해 datetime 오브젝트에서 날짜 정보를 하나씩 추출해 보겠습니다
### datetime 오브젝트에서 날짜 정보 추출하기
#### 1.
다음은 문자열로 저장된 날짜를 시리즈에 담아 datetime 오브젝트로 변환한 것입니다.
```
date_series=pd.Series(['2018-05-16','2018-05-17','2018-05-18'])
d1=pd.to_datetime(date_series)
print(d1)
```
#### 2.
datetime 오브젝트의 year,month,day 속성을 이용하면 년,월,일 정보를 바로 추출할 수 있습니다.
```
print(d1[0].year)
print(d1[0].month)
print(d1[0].day)
```
## dt 접근자 사용하기
문자열을 처리하려면 str 접근자를 사용한 다음 문자열 속성이나 메서드를 사용해야 했습니다. datetime 오브젝트도 마찬가지로 dt 접근자를 사용하면 datetime 속성이나 메서드를 사용하여 시계열 데이터를 처리할 수 있습니다.
### dt 접근자로 시계열 데이터 정리하기
#### 1.
먼저 ebola 데이터 집합을 불러온 다음 Date 열을 datetime 오브젝트로 변환하여 새로운 열로 추가합니다.
```
ebola=pd.read_csv('data/country_timeseries.csv')
ebola['date_dt']=pd.to_datetime(ebola['Date'])
```
#### 2.
다음은 dt 접근자를 사용하지 않고 인덱스가 3인 데이터의 년,월,일 데이터를 추출한 것입니다.
```
print(ebola[['Date', 'date_dt']].head())
print(ebola['date_dt'][3].year)
print(ebola['date_dt'][3].month)
print(ebola['date_dt'][3].day)
```
#### 3.
과정 2와 같은 방법은 date_dt 열의 특정 데이터를 인덱스로 접근해야 하기 때문에 불편합니다. 다음은 dt 접근자로 date_dt 열에 한 번에 접근한 다음 year 속성을 이용하여 연도값을 추출한 것입니다. 추출한 연도값은 ebola 데이터프레임의 새로운 열로 추가했습니다.
```
ebola['year']=ebola['date_dt'].dt.year
print(ebola[['Date','date_dt','year']].head())
```
#### 4.
다음은 과정 3을 응용하여 월,일 데이터를 한 번에 추출해서 새로운 열로 추가한 것입니다.
```
ebola['month'],ebola['day']=(ebola['date_dt'].dt.month,ebola['date_dt'].dt.day)
print(ebola[['Date','date_dt','year','month','day']].head())
```
#### 5.
다음은 ebola 데이터프레임에 새로 추가한 date_dt, year, month,day 열의 자료형을 출력한 것입니다. date_dt열은 datetime 오브젝트이고 나머지는 정수형이라는 것을 알 수 있습니다.
```
print(ebola.info())
```
# 12-2 사례별 시계열 데이터 계산하기
### 에볼라 최초 발병일 계산하기
#### 1.
ebola 데이터프레임의 마지막 행과 열을 5개씩만 살펴보겠습니다. ebola 데이터프레임은 데이터가 시간 역순으로 정렬되어 있습니다. 즉, 시간 순으로 데이터를 살펴보려면 데이터프레임의 마지막부터 살펴봐야 합니다.
```
print(ebola.iloc[-5:,:5])
```
#### 2.
121행에서 볼 수 있듯이 에볼라가 발생하기 시작한 날은 2014년 03월 22일입니다. 다음은 min. 메서드를 사용하여 에볼라의 최초 발병일을 찾은 것입니다.
```
print(ebola['date_dt'].min())
print(type(ebola['date_dt'].min()))
```
#### 3.
에볼라의 최초 발병일을 알아냈으니 Date 열에서 에볼라의 최초 발병일을 빼면 에볼라의 진행 정도를 알 수 있습니다.
```
ebola['outbreak_d']=ebola['date_dt']-ebola['date_dt'].min()
print(ebola[['Date','Day','outbreak_d']].head())
```
### 파산한 은행의 개수 계산하기
이번에는 파산한 은행 데이터를 불러와 분기별로 파산한 은행이 얼마나 되는지 계산해 보겠습니다. 그리고 이번에는 그래프로도 시각화해 보겠습니다.
#### 1.
다음은 파산한 은행 데이터 집합을 불러온 것입니다. banks 데이터프레임의 앞부분을 살펴보면 Closing Date, Updated Date 열의 데이터 자료형이 시계열 데이터라는 것을 알 수 있습니다.
```
banks=pd.read_csv('data/banklist.csv')
print(banks.head())
```
#### 2.
Closing date,Update Date 열의 데이터 자료형은 문자열입니다. 다음은 read_csv 메서드의 parse_dates 속성을 이용하여 문자열로 저장된 두 열을 datetime 오브젝트로 변환하여 불러온 것입니다.
```
banks_no_dates=pd.read_csv('data/banklist.csv')
print(banks_no_dates.info())
banks=pd.read_csv('data/banklist.csv',parse_dates=[5,6])
print(banks.info())
```
#### 3.
dt 접근자와 quater 속성을 이용하면 은행이 파산한 분기를 알 수 있습니다. 다음은 dt 접근자와 year,quarter 속성을 이용하여 은행이 파산한 연도, 분기를 새로운 열로 추가한 것입니다.
```
banks['closing_quarter'],banks['closing_year']=(banks['Closing Date'].dt.quarter, banks['Closing Date'].dt.year)
print(banks.head())
```
#### 4.
이제 연도별로 파산한 은행이 얼마나 되는지를 알아볼까요? grouby 메서드를 사용하면 연도별로 파산한 은행의 개수를 구할 수 있습니다.
```
closing_year=banks.groupby(['closing_year']).size()
print(closing_year)
```
#### 5.
각 연도별, 분기별로 파산한 은행의 개수도 알아보겠습니다. 다음은 banks 데이터프레임을 연도별로 그룹화한 다음 다시 분기별로 그룹화하여 출력한 것입니다.
```
closing_year_q=banks.groupby(['closing_year','closing_quarter']).size()
print(closing_year_q)
```
#### 6.
다음은 과정 5에서 얻은 값으로 그래프를 그린 것입니다.
```
import matplotlib.pyplot as plt
fig, ax= plt.subplots()
ax=closing_year.plot()
plt.show()
fig, ax=plt.subplots()
ax=closing_year_q.plot()
plt.show()
```
### 테슬라 주식 데이터로 시간 계산하기
이번에는 pandas=datareader 라이브러리를 이용하여 주식 데이터를 불러오겠습니다. 이라이브러리는 지금까지 설치한 적이 없는 라이브러리입니다. 다음을 아나콘다 프롬프트에 입력하여 pandas-datareader 라이브러리를 설치하세요.
```
pip install pandas-datareader
```
#### 1.
다음은 get_data_quanal 메서드에 TSLA라는 문자열을 전달하여 테슬라의 주식 데이터를 내려받은 다음 to_csv 메서드를 사용하여 data 폴더 안에 'tesla_stock_quandl.csv'라는 이름으로 저장한 것입니다.
```
pd.core.common.is_list_like=pd.api.types.is_list_like
import pandas_datareader as pdr
tesla=pdr.get_data_quandl('TSLA', api_key = 'errmVW9g1S9WR_xmBHon')
tesla.to_csv('data/tesla_stock_quandl.csv')
```
#### 2.
tesla 데이터프레임의 Date 열은 문자열로 저장되어 있습니다. 즉,datetime 오브젝트로 자료형을 변환해야 시간 계산을 할 수 있습니다.
```
print(tesla.head())
```
#### 3.
Date 열을 Datetime 형으로 변환하려면 read_csv 메서드로 데이터 집합을 불러올 때 parse_dates 인자에 Date 열을 전달하면 됩니다.
```
tesla=pd.read_csv('data/tesla_stock_quandl.csv',parse_dates=[0])
print(tesla.info())
```
#### 4.
Date 열의 자료형이 datetime 오브젝트로 변환되었습니다. 이제dt 접근자를 사용할 수 있습니다. 다음은 불린 추출로 2010년 6월의 데이터만 추출한 것입니다.
```
print(tesla.loc[(tesla.Date.dt.year ==2010) & (tesla.Date.dt.month ==6)])
```
## detetime 오브젝트와 인덱스 - DatetimeIndex
지금까지의 실습은 대부분 데이터프레임의 행 번호를 인덱스로 사용했지만 datetime 오브젝트를 데이터프레임의 인덱스로 설정하면 원하는 시간의 데이터를 바로 추출할 수 있어 편리합니다. 이번에는 datetime 오브젝트를 인덱스로 지정하는 방법에 대해 알아보겠습니다.
### datetime 오브젝트를 인덱스로 설정해 데이터 추출하기
#### 1.
계속해서 테슬라 주식 데이터를 사용하여 실습을 진행하겠습니다. 다음은 Date 열을 tesla 데이터프레임의 인덱스로 지정한 겁입니다.
```
tesla.index=tesla['Date']
print(tesla.index)
```
#### 2.
datetime 오브젝트를 인덱스로 지정하면 다음과 같은 방법으로 원하는 시간의 데이터를 바로 추출할 수 있습니다. 다음은 2015년의 데이터를 추출한 것입니다.
```
print(tesla['2015'].iloc[:5,:5])
```
#### 3.
다음은 2010년 6월의 데이터를 추출한 것입니다.
```
print(tesla['2010-06'].iloc[:,:5])
```
## 시간 간격과 인덱스 - TimedeltaIndex
예를 들어 주식 데이터에서 최소 5일간 수집된 데이터만 살펴보고 싶다면 어떻게 해야 할까요? 이런 경우에는 시간 간격을 인덱스로 지정하여 데이터를 추출하면 됩니다. 이번에는 datetime 오브젝트를 인덱스로 지정하는 것이 아니라 시간 간격을 인덱스로 지정하여 진행하겠습니다.
### 시간 간격을 인덱스로 지정해 데이터 추출하기
#### 1.
Date 열에서 Date 열의 최솟값을 빼면 데이터를 수집한 이후에 시간이 얼마나 흘렀는지 알 수 있습니다. Date 열에서 Date 열의 최솟값을 뺀 다음 ref_date열로 추가한 것입니다.
```
tesla['ref_date']=tesla['Date']-tesla['Date'].min()
print(tesla.head())
```
#### 2.
다음과 같이 ref_date 열을 인덱스로 지정했습니다. 이제 시간 간격을 이용하여 데이터를 추출할 수 있습니다.
```
tesla.index=tesla['ref_date']
print(tesla.iloc[:5,:5])
```
#### 3.
다음은 데이터를 수집한 이후 최초 5일의 데이터를 추출한 것입니다.
```
print(tesla['5 days':].iloc[:5, :5])
```
## 시간 범위와 인덱스
### 시간범위 생성해 인덱스로 지정하기
#### 1.
테슬라 주식 데이터는 특정 일에 누락된 데이터가 없습니다. 그래서 이번에는 에볼라 데이터 집합을 사용하겠습니다. 가장 앞쪽의 데이터를 살펴보면 2015년 01월 01일의 데이터가 누락된 것을 알 수 있습니다.
```
ebola=pd.read_csv('data/country_timeseries.csv',parse_dates=[0])
print(ebola.iloc[:5,:5])
```
#### 2.
뒤쪽의 데이터도 마찬가지입니다. 2014년 03월 23일의 데이터가 누락되었습니다.
```
print(ebola.iloc[-5:,:5])
```
#### 3.
다음은 date_range 메서드를 사용하여 2014년 12월 31일 부터 2015년 01월 05일 사요의 시간 인덱스를 생성한 것입니다.
```
head_range=pd.date_range(start='2014-12-31', end='2015-01-05')
print(head_range)
```
#### 4.
다음은 원본 데이터를 손상시키는 것을 방지하기 위해 ebola 데이터프레임의 앞쪽 5개의 데이터를 추출하여 새로운 데이터프레임을 만든 것입니다. 이때 Date 열을 인덱스로 먼저 지정하지 않으면 오류가 발생합니다. 반드시 Date 열을 인덱스로 지정한 다음 과정 3에서 생성한 시간 범위를 인덱스로 지정해야 합니다.
```
ebola_5=ebola.head()
ebola_5.index=ebola_5['Date']
ebola_5.reindex(head_range)
print(ebola_5.iloc[:5,:5])
```
### 시간 범위의 주기 설정하기
시간 범위를 인덱스로 지정하면 DatetimeIndex 자료형이 만들어집니다. 그리고 DatetimeIndex에는 freq 속성이 포함되어 있죠 freq 속성값을 지정하면 시간 간격을 조절하여 DatetimeIndex 를 만들수 있습니다. 아래에 freq 속성값으로 사용할 수 있는 시간 주기를 표로 정리했습니다.
#### freq 속성값으로 사용할 수 있는 시간 주기
- 시간 주기 : 설명
- B : 평일만 포함
- C : 사용자가 정의한 평일만 포함
- D : 달력 일자 단위
- W : 주간 단위
- M : 월 마지막 날만 포함
- SM : 15일과 월 마지막 날만 포함
- BM : M 주기의 값이 휴일이면 제외하고 평일만 포함
- CBM : BM에 사용자 정의 평일을 적용
- MS : 월 시작일만 포함
- SMS : 월 시작일과 15일만 포함
- BMS : MS 주기의 값이 휴일이면 제외하고 평일만 포함
- CBMS : BMS에 사용자 정의 평일을 적용
- Q : 3,6,9,12월 분기 마지막 날만 포함
- BQ : 3,6,9,12월 분기 마지막 날이 휴일이면 제외하고 평일만 포함
- QS : 3,6,9,12월 분기 시작일만 포함
- BQS : 3,6,9,12월 분기 시작일이 휴일이면 제외하고 평일만 포함
- A : 년의 마지막 날이 휴일이면 제외하고 평일만 포함
- BA : 년의 마지막 날이 휴일이면 제외하고 평일만 포함
- AS :년의 시작일만 포함
- BAS : 년의 시작일이 휴일이면 제외하고 평일만 포함
- BH : 평일을 시간 단위로 포함(09:00~16:00)
- H : 시간 단위로 포함(00:00~00:00)
- T : 분 단위 포함
- S : 초 단위 포함
- L : 밀리초 단위 포함
- U : 마이크로초 단위 포함
- N : 나노초 단위 포함<br>
다음은 date_range 메서드의 freq 인잣값을 B로 설정하여 평일만 포함시킨 DatetimeIndex를 만든 것입니다.
```
print(pd.date_range('2017-01-01','2017-01-07',freq='B'))
```
## 시간 범위 수정하고 데이터 밀어내기 - shift 메서드
만약 나라별로 에볼라의 확산 속도를 비교하려면 발생하기 시작한 날짜를 옮기는 것이 좋습니다. 왜 그럴까요? 일단 ebola 데이터프레임으로 그래프를 그려보고 에볼라의 확산 속도를 비교하는 데 어떤 문제가 있는지 그리고 해결 방법은 무엇인지 알아보겠습니다.
### 에볼라의 확산 속도 비교하기
#### 1.
다음은 ebola 데이터프레임의 Date 열을 인덱스로 지정한 다음 x축을 Date 열로, y축을 사망자 수로 지정하여 그린 그래프입니다.
```
import matplotlib.pyplot as plt
ebola.index=ebola['Date']
fig, ax=plt.subplots()
ax=ebola.iloc[0:,1:].plot(ax=ax)
ax.legend(fontsize=7, loc=2, borderaxespad=0.)
plt.show()
```
#### 2.
그런데 과정 1의 그래프는 각 나라의 에볼라 발병일이 달라 그래프가 그려지기 시작한 지점도 다릅니다. 달리기 속도를 비교하려면 같은 출발선에서 출발하여 시간을 측정해야겠죠? 에볼라의 확산 속도도 같은 방법으로 측정해야 합니다. 즉, 각나라의 발병일을 가장 처음 에볼라가 발병한 Guinea와 동일한 위치로 옮겨야 나라별 에볼라의 확산 속도를 제대로 비교할 수 있습니다.
```
ebola_sub=ebola[['Day', 'Cases_Guinea', 'Cases_Liberia']]
print(ebola_sub.tail(10))
```
#### 3. 그래프를 그리기 위한 데이터프레임 준비하기
다음은 Date 열의 자료형을 datetime 오브젝트로 변환하여 ebola 데이터프레임을 다시 생성한 것입니다. 그런데 중간에 아에 날짜가 없는 데이터도 있습니다. 이데이터도 포함시켜야 확산 속도를 제대로 비교할 수 있습니다.
```
ebola=pd.read_csv('data/country_timeseries.csv', parse_dates=['Date'])
print(ebola.head().iloc[:,:5])
print(ebola.tail().iloc[:,:5])
```
#### 4.
다음은 Date 열을 인덱스로 지정한 다음 ebola 데이터프레임의 Date 열의 최댓값과 최솟값으로 시간 범위를 생성하여 new_idx에 저장한 것입니다. 이렇게 하면 날짜가 아예 없었던 데이터의 인덱스를 생성할 수 있습니다.
```
ebola.index=ebola['Date']
new_idx=pd.date_range(ebola.index.min(),ebola.index.max())
```
#### 5.
그런데 new_idx를 살펴보면 ebola 데이터 집합에 있는 시간 순서와 반대로 생성되어 있습니다. 다음은 시간 순서를 맞추기 위해 reversed 메서드를 사용하여 인덱스를 반대로 뒤집은 것입니다.
```
print(new_idx)
```
#### 6.
다음은 reindex 메서드를 사용하여 새로 생성한 인덱스를 새로운 인덱스로 지정한 것입니다. 그러면 2015년 01월 01일 데이터와 같은 ebola 데이터프레임에 아예 없었던 날짜가 추가됩니다. 이제 그래프를 그리기 위한 데이터프레임이 준비되었습니다.
```
ebloa=ebola.reindex(new_idx)
print(ebola.head().iloc[:,:5])
print(ebola.tail().iloc[:,:5])
```
#### 7. 각 나라의 에볼라 발병일 옮기기
다음은 last_valid_index, first_valid_index 메서드를 사용하여 각 나라의 에볼라 발병일을 구한 것입니다. 각각의 메서드는 유효한 값이 있는 첫 번째와 마지막 인덱스를 반환합니다. 다음을 입력하고 결과를 확인해 보세요.
```
last_valid=ebola.apply(pd.Series.last_valid_index)
print(last_valid)
first_valid=ebola.apply(pd.Series.first_valid_index)
print(first_valid)
```
#### 8.
각 나라의 에볼라 발병일을 동일한 출발선으로 옮기려면 에볼라가 가장 처음 발병한 날에서 각 나라의 에볼라 발병일을 뺀 만큼만 옯기면 됩니다.
```
earliest_date=ebola.index.min()
print(earliest_date)
shift_values = last_valid - earliest_date
print(shift_values)
```
#### 9.
이제 각 나라의 에볼라 발병일을 옮기면 됩니다. 다음은 shift 메서드를 사용하여 모든 열의 값을 shift_values 값만큼 옯긴 것입니다. shift 메서드는 인잣값만큼 데이터를 밀어내는 메서드입니다.
```
ebola_dict={}
for idx, col in enumerate(ebola):
d= shift_values[idx].days
shifted=ebola[col].shift(d)
ebola_dict[col]=shifted
```
#### 10.
ebola_dict에는 시간을 다시 설정한 데이터가 딕셔너리 형태로 저장되어 있습니다. 다음은 DataFarme 메서드를 사용하여 ebola_dict의 값을 데이터프레임으로 변환한 것입니다.
```
ebola_shift=pd.DataFrame(ebola_dict)
```
#### 11.
이제 에볼라의 최초 발병일을 기준으로 모든 열의 데이터가 옮겨졌습니다.
```
print(ebola_shift.tail())
```
#### 12.
마지막으로 인덱스를 Day 열로 지정하고 그래프에 필요 없는 Date, Day 열을 삭제하면 그래프를 그리기 위한 데이터프레임이 완성됩니다.
```
ebola_shift.index=ebola_shift['Day']
ebola_shift=ebola_shift.drop(['Date', 'Day'], axis=1)
print(ebola_shift.tail())
```
#### 13.
다음은 지금까지 만든 데이터프레임으로 다시 그린 그래프입니다.
```
fig, ax=plt.subplots()
ax=ebola_shift.iloc[:,:].plot(ax=ax)
ax.legend(fontsize=7, loc=2, borderaxespad=0.)
plt.show()
```
## 마무리하며
판다스 라이브러리는 시간을 다룰 수 있는 다양한 기능을 제공합니다. 이 장에서는 시계열 데이터와 깊은 연관성이 있는 에볼라 데이터 및 주식 데이터를 주로 다루었습니다. 우리 주변의 상당수의 데이터는 시간과 깊은 연관성이 있는 경우가 많습니다. 시계열 데이터를 능숙하게 다루는 것은 데이터 분석가의 기본 소양이므로 이 장의 내용을 반드시 익혀두기 바랍니다.
출처 : "do it 데이터분석을 위한 판다스 입문"
| github_jupyter |
# Simplified solid lubricant wear model
<a href="../handbook/reliability_prediction/structural_models_equations.html?highlight=fluid lubricant wear#solid-lubricant-wear" class="back-forward-button">Go to handbook</a>
## Model description
Solid lubricant wear modelling is described on the example of a ball bearing. The modelling is applicable to other cases of solid lubricant wear, however, the number of revolution has to be substituted with another measure of sliding distance.
For solid lubricant reservoir wear (e.g., cage of a ball bearing), the limit state function for the adhesive wear model is formulated for single time interval (simplification) as follows:
$$
g\left( V_{\text{lim}}, K_H, \alpha, \Theta \right) = {V_{{\text{lim}}}} - \Theta {{K_{H}} {\alpha} r}
$$
Where $\alpha$ denotes the average work of ball/cage interaction forces per revolution and $r$ is the number of revolutions in the considered time interval. The parameter $\alpha$ will typically be estimated from tests and thus depends on the wear rate $K_H$, which is used to estimate $\alpha$ from the test results. A summary of these variables and their meaning is given in {numref}`solid_lubricant_model_inputs_table`.
```{list-table} Input variables for reliability analysis
:header-rows: 1
:widths: 15 45 20 20
:name: solid_lubricant_model_inputs_table
* - Name
- Description
- Unit
- Type
* - $V_{\text{lim}}$/V_lim
- Limiting value (worn volume),
- $m^3/10^6$
- uncertain
* - $K_H$/K_H
- Specific wear rate
- $Pa^{-1}=m^2/N/10^{12}$
- uncertain
* - $\alpha$/alpha
- Ball-cage interaction
- $N/m$
- uncertain
* - $r$/r
- Nominal number of revolutions
- $-$
- deterministic
* - $\Theta$/Theta
- Model uncertainty
- $-$
- uncertain
```
## Interactive reliability prediction
This page offers an interactive reliability prediction that lets the user specify the properties of all variables listed in {numref}`solid_lubricant_model_inputs_table`. The value of **deterministic variables** can be selected with a slider. **Uncertain variables** are characterized by:
- _Distribution_ denoted by "Dist" and can be choosen from a set of parametric probability distributions;
- _Mean_ value denoted by "E" and can be selected with a slider;
- _Coefficient of variation_ denoted by "C.o.V." and can be selected with a slider.
The variable $r_h$ denotes the average number of revolutions per hour and is only required for plotting the probability of failure as a function of time.
```{note}
To run the interactive reliability prediction on this page, click the {fa}`rocket` --> {guilabel}`Live Code` button on the top of the page. Wait a few seconds until the Kernel has loaded and run the cell below with {guilabel}`Run`.
```
```{admonition} Under construction
:class: todo
The numerical values and bounds for the input variables are not finalized yet. Therefore, computed failure probabilities might not be representative of the considered components.
```
```
from nrpmint.booktools import solid_lubricant_wear
# start the web user-interface
solid_lubricant_wear.web_ui()
```
| github_jupyter |
# Download ECG data
This notebook downloads ECG data from the [MIT-BIH Arrhythmia Database Directory](https://archive.physionet.org/physiobank/database/html/mitdbdir/mitdbdir.htm)
Copyright 2020 Dr. Klaus G. Paul
Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
```
from IPython.display import display
from ipywidgets import IntProgress
import numpy as np
import os
import pandas as pd
import wfdb
import requests
import zipfile, io
```
Download the zip archive and extract all the files
```
r = requests.get("https://storage.googleapis.com/mitdb-1.0.0.physionet.org/mit-bih-arrhythmia-database-1.0.0.zip")
z = zipfile.ZipFile(io.BytesIO(r.content))
if not os.path.exists("./mit-data"):
os.mkdir("./mit-data")
z.extractall("./mit-data")
```
This is not very generic, but the example does not need a concise dataset.
```
r = set()
for f in os.listdir("./mit-data/mit-bih-arrhythmia-database-1.0.0/"):
s = f.split(".")[0][:3]
if s.isdigit():
r.add(s)
wIP = IntProgress(min=0,max=len(r))
display(wIP)
allAbnormalities = []
allData = []
for rec in r:
record = wfdb.rdrecord('./mit-data/mit-bih-arrhythmia-database-1.0.0/{}'.format(rec))
dfHB = pd.DataFrame(record.p_signal)
dfHB.rename(columns={0:record.sig_name[0],1:record.sig_name[1]},inplace=True)
dfHB["record"] = rec # this is the reference between time series and markup data
dfHB["sample"] = dfHB.index
# this is known
freq = 360
period = '{}N'.format(int(1e9 / freq))
index = pd.date_range(pd.to_datetime("2020-01-01 12:00"), periods=len(dfHB), freq=period)
dfHB["Timestamp"] = index
# need to reduce the amount of data
dfHB = dfHB[dfHB.Timestamp < pd.to_datetime("2020-01-01 12:02:30")]
dfHB.index = dfHB["Timestamp"]
# else bokeh may complain about identical names
del dfHB.index.name
dfHB.to_parquet("../data/{}.parquet".format(rec), use_deprecated_int96_timestamps=True)
allData.append(dfHB)
ann = wfdb.rdann('./mit-data/mit-bih-arrhythmia-database-1.0.0/{}'.format(rec),
extension='atr',return_label_elements=['symbol', 'label_store', 'description'])
ann.create_label_map()
dfAnn = pd.DataFrame({"annotation":ann.description,"sample":ann.sample,"symbol":ann.symbol})
dfAnn = dfAnn[dfAnn["sample"] <= len(dfHB)]
dfAnn = pd.merge(dfAnn,dfHB,on="sample")
dfAnn["record"] = rec
# else bokeh may complain about identical names
del dfAnn.index.name
# uncomment this if you think you need the individual files
#dfAnn.to_csv("../data/ann.{}.csv".format(rec))
#dfAnn[dfAnn.symbol != "N"].to_csv("../data/ann.abnormalities.{}.csv".format(rec))
allAbnormalities.append(dfAnn[dfAnn.symbol != "N"][["Timestamp","annotation","symbol","record"]])
wIP.value += 1
pd.DataFrame().append(allAbnormalities,sort=False).to_parquet("../data/abnormalities.parquet", use_deprecated_int96_timestamps=True)
#pd.DataFrame().append(allData,sort=False).to_parquet("../data/ecg.parquet", use_deprecated_int96_timestamps=True)
```
| github_jupyter |
```
# https://github.com/timestocome
# Lovecraft Corpus
# https://github.com/vilmibm/lovecraftcorpus
# use tfidf to cluster Lovecraft stories by similarity to each other
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# silence is golden
import warnings
warnings.filterwarnings("ignore")
warnings.filterwarnings(action="ignore",category=DeprecationWarning)
warnings.filterwarnings(action="ignore",category=FutureWarning)
# list all files under the input directory
import os
fNames = []
for dirname, _, filenames in os.walk('lovecraftcorpus'):
for filename in filenames:
fNames.append(os.path.join(dirname, filename))
print(fNames)
print(len(fNames))
# read in all files, split into sentences, do a bit of cleanup to reduce vocabulary size
from nltk.tokenize import sent_tokenize
import functools
import re
stories = []
for f in fNames:
fp = open(f)
story = fp.read()
story = story.lower()
story = re.sub('-', ' ', story)
story = re.sub(" \'", ' ', story)
story = re.sub('\"', ' ', story)
story = re.sub('\d', '9', story)
stories.append(sent_tokenize(story))
# flatten stories into sentences
sentences = functools.reduce(lambda x, y: x+y, stories)
n_sentences = len(sentences)
n_stories = len(stories)
# assign a story number as a target for each sentence
targets = []
for i in range(len(stories)):
n_sent = len(stories[i])
t = [i] * n_sent
targets.extend( t )
print('targets %d data %d' %(len(targets), len(sentences)))
# store sentences and targets in a df
train = pd.DataFrame(targets)
train.columns = ['target']
train['sentences'] = sentences
print(train.tail())
# tokenize, vectorize story sentences by story
from sklearn.feature_extraction.text import TfidfVectorizer
from nltk.tokenize.casual import casual_tokenize
tfidf_model = TfidfVectorizer(tokenizer=casual_tokenize)
tfidf_model.fit(raw_documents=train['sentences'])
x = []
y = []
for i in range(n_stories):
docs = train[train['target'] == i]['sentences']
t = tfidf_model.transform(docs)
x.append(t.mean(axis=0).sum())
y.append(t.mean(axis=1).sum())
# split out story names from file list for plot
import re
story_names = []
for i in range(len(fNames)):
stry = re.search('\/(.*).txt', fNames[i])
story_names.append(' ' + stry.group(1))
print(story_names)
# plot stories tfidf mean
import matplotlib.pyplot as plt
font = {'family' : 'sans-serif',
'weight' : 'normal',
'size' : 13}
plt.rc('font', **font)
fig, ax = plt.subplots(figsize=(16,16))
ax.scatter(x, y)
# label points
n = np.arange(0, n_stories)
for i, txt in enumerate(story_names):
#for i, txt in enumerate(n):
ax.annotate(txt, (x[i], y[i]))
plt.title('Similarity of Lovecraft Stories')
plt.show()
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import scipy.stats as st
import probability_kernels as pk
```
#### Note to users
This Jupyter Notebook is for creating the figures in the paper. It also demonstrates how percentile transition matrices can be calculatd using the python file `probability_kernels`.
```
save = True
```
### Figure of the Peason data
```
# Load the data frame (female) -> dff
dff = pd.read_csv('data/pearson-lee-mother-daughter.csv')
# x values (mothers)
xf = dff.Parent.to_numpy()
# y values (daughters)
yf = dff.Child.to_numpy()
# Load the data frame (male) -> dfm
dfm = pd.read_csv('data/pearson-lee-father-son.csv')
# x values (fathers)
xm = dfm.Parent.to_numpy()
# y values (sons)
ym = dfm.Child.to_numpy()
%%time
# Create an empty list of size three, that will store
matrices_p = [None] * 3
matrices_p[0] = pk.get_matrix_data(xf, yf)
matrices_p[1] = pk.get_matrix_data(xm, ym)
matrices_p[2] = pk.get_matrix(r=0.54, rs=0.96, num_iters=1_000_000, trim_score=6)
# Pearson male is exactly the same: pk.get_matrix(r=0.51, rs=0.89)
fig, axes = plt.subplots(3, 1, figsize=(13*1*0.95*0.75, 8*3/0.95*0.75))
titles_p = ['Pearson data, Mother-Daughter', 'Pearson data, Father-Son',
'Pearson data, simulation of estimated parameters']
for i in range(3):
pk.plot_ax(ax=axes.ravel()[i], matrix=matrices_p[i], i=0,
title=titles_p[i], title_loc='center', child=True)
plt.tight_layout()
legend = ['Descendant in\nTop Quintile', 'Fourth Quintile',
'Third Quintile', 'Second Quintile', 'Bottom Quintile']
fig.legend(legend, bbox_to_anchor=(1.27, 0.9805), fontsize=15)
if save:
plt.savefig('latex/figures/quintile-pearson.png', dpi=300)
plt.show()
```
### Figure for multigenerational mobility, standard parameters
```
r = 0.5
rs = pk.stable_rs(r)
num_steps = 6
matrices = [None] * num_steps
print('r_s =', round(rs, 5))
%%time
for i in range(num_steps):
matrices[i] = pk.get_matrix(r=r, rs=rs, n=i+1, num_iters=1_000_000, trim_score=6)
fig, axes = plt.subplots(3, 2, figsize=(13*2*0.95*0.75, 8*3/0.95*0.75))
for i in range(num_steps):
pk.plot_ax(ax=axes.ravel()[i], matrix=matrices[i], i=i, j=i,
title="$n = {}$".format(str(i+1)), title_loc='center', x_label=True, child=False)
plt.tight_layout()
if save:
plt.savefig('latex/figures/quintile-r=0.5-stable.png', dpi=300)
plt.show()
```
### Figure for the mobility measure
```
mv = np.array([12, 6, 3, 2, 1.4, 1])
m = mv.size
rv, rsv = pk.get_rv_rsv(mv)
matrices_m = [None] * m
%%time
for i in range(m):
matrices_m[i] = pk.get_matrix(r=rv[i], rs=rsv[i], n=1, num_iters=1_000_000, trim_score=6)
```
There are `num_iters` number of iterations over the simulated integral for each probability calculation. Therefore, $5\times 5 \times$ `num_iters` total for one quintile transition matrix. Here we make six matrices in 23 seconds. Therefore, about 6.5 million computations per second - due to vectorization.
```
fig, axes = plt.subplots(3, 2, figsize=(13*2*0.95*0.75, 8*3/0.95*0.75))
for i in range(m):
pk.plot_ax(ax=axes.ravel()[i], matrix=matrices_m[i], i=0, j=i,
title=pk.report_mobility(mv, rv, rsv, i), title_loc='center',
x_label=False, child=True)
plt.tight_layout()
if save:
plt.savefig('latex/figures/quintile-mobility.png', dpi=300)
plt.show()
```
### Figure for the Chetty data

```
chetty = np.array(
[[0.337, 0.242, 0.178, 0.134, 0.109],
[0.28, 0.242, 0.198, 0.16, 0.119],
[0.184, 0.217, 0.221, 0.209, 0.17],
[0.123, 0.176, 0.22, 0.244, 0.236],
[0.075, 0.123, 0.183, 0.254, 0.365]])
pk.plot_matrix(chetty, child=True, legend=False)
plt.tight_layout()
if save:
plt.savefig('latex/figures/quintile-chetty.png', dpi=300)
r_chetty = 0.31
pk.plot_matrix(
pk.get_matrix(r=r_chetty, rs=pk.stable_rs(r_chetty),
n=1, num_iters=100_000, trim_score=6))
pk.stable_rs(r_chetty) / r_chetty
```
### Reference
```
r_ref = 0.5
ref = pk.get_matrix(r=r_ref, rs=pk.stable_rs(r_ref), n=3, num_iters=1_000_000, trim_score=6)
fig, axes = plt.subplots(1, 1, figsize=(13*1*0.95*0.75, 8*1/0.95*0.75))
pk.plot_ax(axes, matrix=ref, i=2, j=2, x_label=True, child=False)
plt.tight_layout()
if save:
plt.savefig("latex/figures/quintile_reference.png", dpi=300)
```
#### Test symmetry (proof in paper)
```
def get_sigma(r, rs, n):
return np.sqrt((r**2+rs**2)**n)
def joint(v1, v2, r, rs, n):
return st.norm.pdf(v2,
scale=pk.get_sigma_tilda(1, r, rs, n),
loc=pk.get_mu_tilda(v1, r, n)) * st.norm.pdf(v1)
def check_vs(va, vb, r, rs, n):
va_vb = joint(va, vb, r, rs, n)
vb_va = joint(vb, va, r, rs, n)
return va_vb, vb_va
# Stable population variance
r_c = 0.3
check_vs(va=0.3, vb=0.7, r=r_c, rs=pk.stable_rs(r_c), n=3)
# (Not) stable population variance
check_vs(va=0.3, vb=0.7, r=r_c, rs=0.7, n=3)
pa = 0.17
pb = 0.64
def per_to_v1(p1):
return st.norm.ppf(p1)
def per_to_v2(p2, r, rs, n):
return st.norm.ppf(p2, scale=get_sigma(r, rs, n))
def check_ps(pa, pb, r, rs, n):
va_vb = joint(per_to_v1(pa), per_to_v2(pb, r, rs, n), r, rs, n)
vb_va = joint(per_to_v1(pb), per_to_v2(pa, r, rs, n), r, rs, n)
return va_vb, vb_va
# (Not) stable population variance, but index by percentile
check_ps(pa=0.17, pb=0.64, r=r_c, rs=0.7, n=3)
```
### Pearson summary stats
```
rawm = pk.get_matrix_data(xm, ym, return_raw=True)
rawf = pk.get_matrix_data(xf, yf, return_raw=True)
raws = np.ravel((rawm + rawf) / 2)
np.quantile(raws, (0.25, 0.5, 0.75))
min(np.min(rawm), np.min(rawf))
max(np.max(rawm), np.max(rawf))
np.mean(raws)
```
### Top two quintiles
```
# Stature
100-(25+25+43+25)/2
# Income
100-(25+24+36+24)/2
```
### Archive
```
# r2v = np.arange(0.05, 0.6, 0.1)
# rv = np.sqrt(r2v)
# rsv = pk.stable_rs(rv)
# mv = rsv / rv
# for r in np.arange(0.2, 0.9, 0.1):
# plot_matrix(get_matrix(r=r, rs=stable_rs(r)))
# plt.title(str(round(r, 2)) + ', ' + str(round(stable_rs(r), 2)) + ', ' + str(round(stable_rs(r) / r, 2)))
# plt.show()
```
| github_jupyter |
# Custom Layers
One factor behind deep learning's success
is the availability of a wide range of layers
that can be composed in creative ways
to design architectures suitable
for a wide variety of tasks.
For instance, researchers have invented layers
specifically for handling images, text,
looping over sequential data,
and
performing dynamic programming.
Sooner or later, you will encounter or invent
a layer that does not exist yet in the deep learning framework.
In these cases, you must build a custom layer.
In this section, we show you how.
## Layers without Parameters
To start, we construct a custom layer
that does not have any parameters of its own.
This should look familiar if you recall our
introduction to block in :numref:`sec_model_construction`.
The following `CenteredLayer` class simply
subtracts the mean from its input.
To build it, we simply need to inherit
from the base layer class and implement the forward propagation function.
```
import torch
from torch import nn
from torch.nn import functional as F
class CenteredLayer(nn.Module):
def __init__(self):
super().__init__()
def forward(self, X):
return X - X.mean()
```
Let us verify that our layer works as intended by feeding some data through it.
```
layer = CenteredLayer()
layer(torch.FloatTensor([1, 2, 3, 4, 5]))
```
We can now incorporate our layer as a component
in constructing more complex models.
```
net = nn.Sequential(nn.Linear(8, 128), CenteredLayer())
```
As an extra sanity check, we can send random data
through the network and check that the mean is in fact 0.
Because we are dealing with floating point numbers,
we may still see a very small nonzero number
due to quantization.
```
Y = net(torch.rand(4, 8))
Y.mean()
```
## Layers with Parameters
Now that we know how to define simple layers,
let us move on to defining layers with parameters
that can be adjusted through training.
We can use built-in functions to create parameters, which
provide some basic housekeeping functionality.
In particular, they govern access, initialization,
sharing, saving, and loading model parameters.
This way, among other benefits, we will not need to write
custom serialization routines for every custom layer.
Now let us implement our own version of the fully-connected layer.
Recall that this layer requires two parameters,
one to represent the weight and the other for the bias.
In this implementation, we bake in the ReLU activation as a default.
This layer requires to input arguments: `in_units` and `units`, which
denote the number of inputs and outputs, respectively.
```
class MyLinear(nn.Module):
def __init__(self, in_units, units):
super().__init__()
self.weight = nn.Parameter(torch.randn(in_units, units))
self.bias = nn.Parameter(torch.randn(units,))
def forward(self, X):
linear = torch.matmul(X, self.weight.data) + self.bias.data
return F.relu(linear)
```
Next, we instantiate the `MyDense` class
and access its model parameters.
```
dense = MyLinear(5, 3)
dense.weight
```
We can directly carry out forward propagation calculations using custom layers.
```
dense(torch.rand(2, 5))
```
We can also construct models using custom layers.
Once we have that we can use it just like the built-in fully-connected layer.
```
net = nn.Sequential(MyLinear(64, 8), MyLinear(8, 1))
net(torch.rand(2, 64))
```
## Summary
* We can design custom layers via the basic layer class. This allows us to define flexible new layers that behave differently from any existing layers in the library.
* Once defined, custom layers can be invoked in arbitrary contexts and architectures.
* Layers can have local parameters, which can be created through built-in functions.
## Exercises
1. Design a layer that takes an input and computes a tensor reduction,
i.e., it returns $y_k = \sum_{i, j} W_{ijk} x_i x_j$.
1. Design a layer that returns the leading half of the Fourier coefficients of the data.
[Discussions](https://discuss.d2l.ai/t/59)
| github_jupyter |
## This notebook will help you train a raw Point-Cloud GAN.
(Assumes latent_3d_points is in the PYTHONPATH and that a trained AE model exists)
```
from tqdm import trange
import sys
sys.path.append('/latent_3d')
import os
os.environ["CUDA_VISIBLE_DEVICES"]="2"
import numpy as np
import os.path as osp
import matplotlib.pylab as plt
from latent_3d_points.src.autoencoder import Configuration as Conf
from latent_3d_points.src.neural_net import MODEL_SAVER_ID
from latent_3d_points.src.in_out import snc_category_to_synth_id, create_dir, PointCloudDataSet, \
load_all_point_clouds_under_folder
from latent_3d_points.src.general_utils import plot_3d_point_cloud
from latent_3d_points.src.tf_utils import reset_tf_graph
from latent_3d_points.src.vanilla_gan import Vanilla_GAN
from latent_3d_points.src.w_gan_gp import W_GAN_GP
from latent_3d_points.src.generators_discriminators import point_cloud_generator,\
mlp_discriminator, leaky_relu
%load_ext autoreload
%autoreload 2
%matplotlib inline
# Use to save Neural-Net check-points etc.
top_out_dir = '../data/'
# Top-dir of where point-clouds are stored.
top_in_dir = '../data/shape_net_core_uniform_samples_2048/'
experiment_name = 'raw_gan_with_w_gan_loss'
n_pc_points = 2048 # Number of points per model.
class_name = raw_input('Give me the class name (e.g. "chair"): ').lower()
# Load point-clouds.
syn_id = snc_category_to_synth_id()[class_name]
class_dir = osp.join(top_in_dir , syn_id)
all_pc_data = load_all_point_clouds_under_folder(class_dir, n_threads=8, file_ending='.ply', verbose=True)
print 'Shape of DATA =', all_pc_data.point_clouds.shape
```
Set GAN parameters.
```
use_wgan = True # Wasserstein with gradient penalty, or not?
n_epochs = 10 # Epochs to train.
plot_train_curve = True
save_gan_model = True
saver_step = np.hstack([np.array([1, 5, 10]), np.arange(50, n_epochs + 1, 50)])
# If true, every 'saver_step' epochs we produce & save synthetic pointclouds.
save_synthetic_samples = True
# How many synthetic samples to produce at each save step.
n_syn_samples = all_pc_data.num_examples
# Optimization parameters
init_lr = 0.0001
batch_size = 50
noise_params = {'mu':0, 'sigma': 0.2}
noise_dim = 128
beta = 0.5 # ADAM's momentum.
n_out = [n_pc_points, 3] # Dimensionality of generated samples.
discriminator = mlp_discriminator
generator = point_cloud_generator
if save_synthetic_samples:
synthetic_data_out_dir = osp.join(top_out_dir, 'OUT/synthetic_samples/', experiment_name)
create_dir(synthetic_data_out_dir)
if save_gan_model:
train_dir = osp.join(top_out_dir, 'OUT/raw_gan', experiment_name)
create_dir(train_dir)
reset_tf_graph()
if use_wgan:
lam = 10
disc_kwargs = {'b_norm': False}
gan = W_GAN_GP(experiment_name, init_lr, lam, n_out, noise_dim,
discriminator, generator,
disc_kwargs=disc_kwargs, beta=beta)
else:
leak = 0.2
disc_kwargs = {'non_linearity': leaky_relu(leak), 'b_norm': False}
gan = Vanilla_GAN(experiment_name, init_lr, n_out, noise_dim,
discriminator, generator, beta=beta, disc_kwargs=disc_kwargs)
accum_syn_data = []
train_stats = []
# Train the GAN.
for _ in trange(n_epochs):
loss, duration = gan._single_epoch_train(all_pc_data, batch_size, noise_params)
epoch = int(gan.sess.run(gan.increment_epoch))
print epoch, loss
if save_gan_model and epoch in saver_step:
checkpoint_path = osp.join(train_dir, MODEL_SAVER_ID)
gan.saver.save(gan.sess, checkpoint_path, global_step=gan.epoch)
if save_synthetic_samples and epoch in saver_step:
syn_data = gan.generate(n_syn_samples, noise_params)
np.savez(osp.join(synthetic_data_out_dir, 'epoch_' + str(epoch)), syn_data)
for k in range(3): # plot three (synthetic) random examples.
plot_3d_point_cloud(syn_data[k][:, 0], syn_data[k][:, 1], syn_data[k][:, 2],
in_u_sphere=True)
train_stats.append((epoch, ) + loss)
if plot_train_curve:
x = range(len(train_stats))
d_loss = [t[1] for t in train_stats]
g_loss = [t[2] for t in train_stats]
plt.plot(x, d_loss, '--')
plt.plot(x, g_loss)
plt.title('GAN training. (%s)' %(class_name))
plt.legend(['Discriminator', 'Generator'], loc=0)
plt.tick_params(axis='x', which='both', bottom='off', top='off')
plt.tick_params(axis='y', which='both', left='off', right='off')
plt.xlabel('Epochs.')
plt.ylabel('Loss.')
```
| github_jupyter |
## Load the data
## Modelling (BaseLine)
## Modelling (Testing)
## Modelling (Final)
```
import os
import cv2
import torch
import numpy as np
BATCH_SIZE = 32
from tqdm import tqdm
def load_data(img_size=112):
data = []
index = -1
labels = {}
for directory in os.listdir('./data/')[:37]:
index += 1
labels[f'./data/{directory}/'] = [index,-1]
for label in tqdm(labels):
for file in os.listdir(label):
filepath = label + file
img = cv2.imread(filepath)
img = cv2.resize(img,(img_size,img_size))
# img = img / 255.0
data.append([
np.array(img),
labels[label][0]
])
labels[label][1] += 1
for _ in range(12):
np.random.shuffle(data)
print(len(data))
# np.save('./data.npy',data)
return data
import torch
def other_loading_data_proccess(data):
print(len(data))
X = []
y = []
print('going through the data..')
for d in data:
X.append(d[0])
y.append(d[1])
print('splitting the data')
VAL_SPLIT = 0.25
VAL_SPLIT = len(X)*VAL_SPLIT
VAL_SPLIT = int(VAL_SPLIT)
X_train = X[:-VAL_SPLIT]
y_train = y[:-VAL_SPLIT]
X_test = X[-VAL_SPLIT:]
y_test = y[-VAL_SPLIT:]
print('turning data to tensors')
X_train = torch.from_numpy(np.array(X_train))
print(len(X_train))
y_train = torch.from_numpy(np.array(y_train))
print(len(y_train))
X_test = torch.from_numpy(np.array(X_test))
print(len(X_test))
y_test = torch.from_numpy(np.array(y_test))
print(len(y_test))
return [X_train,X_test,y_train,y_test]
REBUILD_DATA = True
if REBUILD_DATA:
data = load_data()
np.random.shuffle(data)
X_train,X_test,y_train,y_test = other_loading_data_proccess(data)
import torch
import torch.nn as nn
from torchvision import transforms
import matplotlib.pyplot as plt
transform_train = transforms.Compose([
transforms.RandomCrop(112, padding=2),
transforms.RandomHorizontalFlip(),
transforms.CenterCrop(112),
transforms.RandomHorizontalFlip(),
transforms.ColorJitter(0.25,0.25),
transforms.ToTensor(),
transforms.Normalize((0.4914, 0.4822, 0.4465), (0.2023, 0.1994, 0.2010)),
])
from PIL import Image
X_train_new = []
import random
for index in range(len(X_train)):
testing = X_train[index]
testing = np.array(testing)
testing = Image.fromarray(testing)
X_train_new.append(np.array(transform_train(testing)))
X_train_new = np.array(X_train_new)
X_train_new = X_train_new.astype(int)
X_train = torch.from_numpy(X_train_new)
X_train.shape
```
## Modelling BaseLine
```
import torch
import torch.nn as nn
from torchvision import models
device = torch.device('cuda')
model = models.resnet18(pretrained=True).to(device)
model = model.to(device)
num_ftrs = model.fc.in_features
model.fc = nn.Linear(num_ftrs, 37)
optimizer = torch.optim.SGD(model.parameters(),lr=0.1)
criterion = nn.CrossEntropyLoss()
EPOCHS = 12
BATCH_SIZE = 32
from tqdm import tqdm
loss_logs = []
for _ in tqdm(range(EPOCHS)):
for i in range(0,len(X_train),BATCH_SIZE):
X_batch = X_train[i:i+BATCH_SIZE].view(-1,3,112,112).to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model = model.to(device)
preds = model(X_batch.float().to(device))
preds = preds.to(device)
loss = criterion(preds,y_batch)
optimizer.zero_grad()
loss.backward()
optimizer.step()
loss_logs.append(loss.item())
plt.plot(loss_logs)
y_batch.shape
preds.shape
preds_new = []
for pred in preds:
preds_new.append(int(torch.argmax(torch.round(pred))))
def get_loss(criterion,y,model,X):
model.to('cpu')
preds = model(X.view(-1,1,112,112).to('cpu').float())
preds.to('cpu')
loss = criterion(preds,torch.tensor(y,dtype=torch.long).to('cpu'))
loss.backward()
return loss.item()
def test(net,X,y):
device = 'cpu'
net.to(device)
correct = 0
total = 0
net.eval()
with torch.no_grad():
for i in range(len(X)):
real_class = torch.argmax(y[i]).to(device)
net_out = net(X[i].view(-1,1,112,112).to(device).float())
net_out = net_out[0]
predictied_class = torch.argmax(net_out)
if predictied_class == real_class:
correct += 1
total += 1
net.train()
net.to('cuda')
return round(correct/total,3)
PROJECT_NAME = 'fruits-37'
import wandb
optimizers = [torch.optim.Adam,torch.optim.AdamW,torch.optim.Adamax,torch.optim.SGD]
for optimizer in optimizers:
model = models.resnet18(pretrained=True).to(device)
model = model.to(device)
num_ftrs = model.fc.in_features
model.fc = nn.Linear(num_ftrs, 37)
optimizer = optimizer(model.parameters(),lr=0.1)
criterion = nn.CrossEntropyLoss()
wandb.init(project=PROJECT_NAME,name=f'optimizer-{optimizer}')
for _ in tqdm(range(EPOCHS)):
for i in range(0,len(X_train),BATCH_SIZE):
X_batch = X_train[i:i+BATCH_SIZE].view(-1,3,112,112).to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model = model.to(device)
preds = model(X_batch.float().to(device))
preds = preds.to(device)
loss = criterion(preds,y_batch)
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item(),'accuracy':test(model,X_train,y_train),'val_accuracy':test(model,X_test.view(-1,3,112,112),y_test),'val_loss':get_loss(criterion,y_test,model,X_test)})
```
| github_jupyter |
### Analyse node statistics for benchmark results
In this notebook we analyse the node statistics, such as e.g. average degree, for correctly and
misclassified nodes, given the benchmark results of any community detection method.
First, we import the necessary packages.
```
%reload_ext autoreload
%autoreload 2
import os
import matplotlib.pyplot as plt
import numpy as np
from clusim.clustering import Clustering
from src.utils.cluster_analysis import get_cluster_properties, get_node_properties
from src.utils.plotting import plot_histogram, init_plot_style
from src.wrappers.igraph import read_graph
%matplotlib
init_plot_style()
color_dict = {'infomap': 'tab:blue', 'synwalk': 'tab:orange', 'walktrap': 'tab:green', 'louvain': 'tab:red',
'graphtool': 'tab:purple'}
```
First, we specify the network to be analyzed, load the network and glance at its basic properties.
```
# select network
network = 'pennsylvania-roads'
# assemble paths
graph_file = '../data/empirical/clean/' + network + '.txt'
results_dir = '../results/empirical/' + network + '/'
os.makedirs(results_dir, exist_ok=True)
# output directory for storing generated figures
fig_dir = '../figures/'
os.makedirs(fig_dir, exist_ok=True)
# load network
graph = read_graph(graph_file)
node_degrees = graph.degree()
avg_degree = np.mean(node_degrees)
print(f'Network size is {len(graph.vs)} nodes, {len(graph.es)} edges')
print (f'Min/Max/Average degrees are {np.min(node_degrees)}, {np.max(node_degrees)}, {avg_degree}.')
```
Here we compute single-number characteristics of the detected clusters.
```
# methods = ['infomap', 'synwalk', 'walktrap']
methods = ['synwalk', 'louvain', 'graphtool']
colors = [color_dict[m] for m in methods]
graph = read_graph(graph_file)
for method in methods:
clu = Clustering().load(results_dir + 'clustering_' + method + '.json')
trivial_clu_sizes = [len(cluster) for cluster in clu.to_cluster_list() if len(cluster) < 3]
num_trivial = len(trivial_clu_sizes)
num_non_trivial = clu.n_clusters - num_trivial
print ('\nCluster statistics for ' + method + ': ')
print (f'Number of detected clusters: {clu.n_clusters}')
# print (f'Number of trivial clusters: {clu.n_clusters - num_non_trivial}')
print (f'Number of non-trivial clusters: {num_non_trivial}')
print (f'Fraction of non-trivial clusters: {num_non_trivial/clu.n_clusters}')
print (f'Fraction of nodes in non-trivial clusters: {1.0 - sum(trivial_clu_sizes)/clu.n_elements}')
print (f'Modularity: {graph.modularity(clu.to_membership_list())}')
```
Here we plot the degree occurances of the network.
```
# plot parameters
bin_size = 1 # integer bin size for aggregating degrees
save_figure = False # if True, we save the figure as .pdf in ´fig_dir´
plt.close('all')
graph = read_graph(graph_file)
node_degrees = graph.degree()
avg_degree = np.mean(node_degrees)
# compute degree pmf
min_deg = np.min(node_degrees)
max_deg = np.max(node_degrees)
bin_edges = np.array(range(min_deg - 1, max_deg+1, bin_size)) + 0.5
bin_centers = bin_edges[:-1] + 0.5
occurances,_ = np.histogram(node_degrees, bins=bin_edges, density=True)
# plot the degree distribution
fig, ax = plt.subplots(figsize=(12,9))
ax.plot(bin_centers, occurances, 'x', label=f'Node Degrees')
ax.plot([avg_degree, avg_degree], [0, np.max(occurances)], color='crimson',
label=fr'Average Degree, $\bar{{k}} = {avg_degree:.2f}$')
ax.set_ylabel(r'Probability Mass, $p(k_\alpha)$')
ax.set_xlabel(r'Node Degree, $k_\alpha$')
ax.loglog()
ax.legend(loc='upper right')
plt.tight_layout()
# save figure as .pdf
if save_figure:
fig_path = fig_dir + 'degrees_' + network + '.pdf'
plt.savefig(fig_path, dpi=600, format='pdf')
plt.close()
```
The next cell plots the histogram of cluster sizes.
```
feature = 'size'
n_bins = 25
xmax = 1e3
plt.close('all')
save_figure = True # if True, we save the figure as .pdf in ´fig_dir´
# compute cluster properties
data = []
for method in methods:
clu = Clustering().load(results_dir + 'clustering_' + method + '.json')
data.append(get_cluster_properties(graph, clu, feature=feature))
# plot histogram
_, ax = plt.subplots(figsize=(12,9))
plot_histogram(ax, data, methods, n_bins, normalization = 'pmf', log_scale=True, xmax=xmax, colors=colors)
ax.set_xlabel(r'Cluster sizes, $|\mathcal{Y}_i|$')
ax.set_ylabel(r'Bin Probability Mass, $p(|\mathcal{Y}_i|)$')
ax.legend(loc='best')
plt.tight_layout()
# save figure as .pdf
if save_figure:
fig_path = fig_dir + feature + '_' + network + '.pdf'
plt.savefig(fig_path, dpi=600, format='pdf')
plt.close()
```
The next cell plots the histogram of cluster densities.
```
feature = 'density'
xmin=1e-2
n_bins = 25
plt.close('all')
save_figure = True # if True, we save the figure as .pdf in ´fig_dir´
# compute cluster properties
data = []
for method in methods:
clu = Clustering().load(results_dir + 'clustering_' + method + '.json')
data.append(get_cluster_properties(graph, clu, feature=feature))
# plot histogram
_, ax = plt.subplots(figsize=(12,9))
plot_histogram(ax, data, methods, n_bins, normalization = 'pmf', log_scale=True, xmin=xmin, colors=colors)
ax.set_xlabel(r'Cluster Density, $\rho(\mathcal{Y}_i)$')
ax.set_ylabel(r'Bin Probability Mass, $p(\rho(\mathcal{Y}_i))$')
ax.legend(loc='best')
plt.tight_layout()
# save figure as .pdf
if save_figure:
fig_path = fig_dir + feature + '_' + network + '.pdf'
plt.savefig(fig_path, dpi=600, format='pdf')
plt.close()
```
The next cell plots the histogram of clustering coefficients.
```
feature = 'clustering_coefficient'
n_bins = 25
xmin = 1e-2
plt.close('all')
save_figure = True # if True, we save the figure as .pdf in ´fig_dir´
# compute cluster properties
data = []
for method in methods:
clu = Clustering().load(results_dir + 'clustering_' + method + '.json')
data.append(get_cluster_properties(graph, clu, feature=feature))
# plot histogram
_, ax = plt.subplots(figsize=(12,9))
plot_histogram(ax, data, methods, n_bins, normalization = 'pmf', log_scale=True, xmin=xmin, colors=colors)
ax.set_xlabel(r'Clustering coefficient, $c(\mathcal{Y}_i)$')
ax.set_ylabel(r'Bin Probability Mass, $p(c(\mathcal{Y}_i))$')
ax.legend(loc='best')
plt.tight_layout()
# save figure as .pdf
if save_figure:
fig_path = fig_dir + feature + '_' + network + '.pdf'
plt.savefig(fig_path, dpi=600, format='pdf')
plt.close()
```
The next cell plots the histogram of cluster conductances.
```
feature = 'conductance'
n_bins = 25
plt.close('all')
save_figure = True # if True, we save the figure as .pdf in ´fig_dir´
# compute cluster properties
data = []
for method in methods:
clu = Clustering().load(results_dir + 'clustering_' + method + '.json')
data.append(get_cluster_properties(graph, clu, feature=feature))
# plot histogram
_, ax = plt.subplots(figsize=(12,9))
plot_histogram(ax, data, methods, n_bins, normalization = 'pmf', log_scale=False, colors=colors)
ax.set_xlabel(r'Conductance, $\kappa(\mathcal{Y}_i)$')
ax.set_ylabel(r'Bin Probability Mass, $p(\kappa(\mathcal{Y}_i))$')
ax.legend(loc='best')
plt.tight_layout()
# save figure as .pdf
if save_figure:
fig_path = fig_dir + feature + '_' + network + '.pdf'
plt.savefig(fig_path, dpi=600, format='pdf')
plt.close()
```
The next cell plots the histogram of cluster cut ratios.
```
feature = 'cut_ratio'
xmin = None
n_bins = 25
plt.close('all')
save_figure = True # if True, we save the figure as .pdf in ´fig_dir´
# compute cluster properties
data = []
for method in methods:
clu = Clustering().load(results_dir + 'clustering_' + method + '.json')
data.append(get_cluster_properties(graph, clu, feature=feature))
# plot histogram
_, ax = plt.subplots(figsize=(12,9))
plot_histogram(ax, data, methods, n_bins, normalization = 'pmf', log_scale=True, xmin=xmin, colors=colors)
ax.set_xlabel(r'Cut Ratio, $\xi(\mathcal{Y}_i)$')
ax.set_ylabel(r'Bin Probability Mass, $p(\xi(\mathcal{Y}_i))$')
ax.legend(loc='best')
plt.tight_layout()
# save figure as .pdf
if save_figure:
fig_path = fig_dir + feature + '_' + network + '.pdf'
plt.savefig(fig_path, dpi=600, format='pdf')
plt.close()
```
The next cell plots the histogram of node mixing parameters.
```
feature = 'mixing_parameter'
xmin = 1e-2
n_bins = 15
plt.close('all')
save_figure = True # if True, we save the figure as .pdf in ´fig_dir´
# compute cluster properties
data = []
for method in methods:
clu = Clustering().load(results_dir + 'clustering_' + method + '.json')
data.append(get_node_properties(graph, clu, feature=feature))
# plot histogram
_, ax = plt.subplots(figsize=(12,9))
plot_histogram(ax, data, methods, n_bins, normalization = 'pmf', log_scale=True, xmin=xmin, colors=colors)
ax.set_xlabel(r'Mixing parameter, $\mu_\alpha$')
ax.set_ylabel(r'Bin Probability Mass, $p(\mu_\alpha)$')
ax.legend(loc='best')
plt.tight_layout()
# save figure as .pdf
if save_figure:
fig_path = fig_dir + feature + '_' + network + '.pdf'
plt.savefig(fig_path, dpi=600, format='pdf')
plt.close()
```
The next cell plots the histogram of normalized local degrees.
```
feature = 'nld'
n_bins = 25
plt.close('all')
save_figure = True # if True, we save the figure as .pdf in ´fig_dir´
# compute cluster properties
data = []
for method in methods:
clu = Clustering().load(results_dir + 'clustering_' + method + '.json')
data.append(get_node_properties(graph, clu, feature=feature))
# plot histogram
_, ax = plt.subplots(figsize=(12,9))
plot_histogram(ax, data, methods, n_bins, normalization = 'pmf', log_scale=True, colors=colors)
ax.set_xlabel(r'Normalized local degree, $\hat{k}_\alpha$')
ax.set_ylabel(r'Probability Mass, $p(\hat{k}_\alpha)$')
ax.legend(loc='best')
plt.tight_layout()
# save figure as .pdf
if save_figure:
fig_path = fig_dir + feature + '_' + network + '.pdf'
plt.savefig(fig_path, dpi=600, format='pdf')
plt.close()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/derek-shing/DS-Unit-2-Sprint-3-Advanced-Regression/blob/master/LS_DS2_234_Ridge_Regression.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Lambda School Data Science - Ridge Regression
Regularize your way to a better tomorrow.
# Lecture
Data science depends on math, and math is generally focused on situations where:
1. a solution exists,
2. the solution is unique,
3. the solution's behavior changes continuously with the initial conditions.
These are known as [well-posed problems](https://en.wikipedia.org/wiki/Well-posed_problem), and are the sorts of assumptions so core in traditional techniques that it is easy to forget about them. But they do matter, as there can be exceptions:
1. no solution - e.g. no $x$ such that $Ax = b$
2. multiple solutions - e.g. several $x_1, x_2, ...$ such that $Ax = b$
3. "chaotic" systems - situations where small changes in initial conditions interact and reverberate in essentially unpredictable ways - for instance, the difficulty in longterm predictions of weather (N.B. not the same thing as longterm predictions of *climate*) - you can think of this as models that fail to generalize well, because they overfit on the training data (the initial conditions)
Problems suffering from the above are called ill-posed problems. Relating to linear algebra and systems of equations, the only truly well-posed problems are those with a single unique solution.

Think for a moment - what would the above plot look like if there was no solution? If there were multiple solutions? And how would that generalize to higher dimensions?
# Well-Posed problems in Linear Algebra
A lot of what you covered with linear regression was about getting matrices into the right shape for them to be solvable in this sense. But some matrices just won't submit to this, and other problems may technically "fit" linear regression but still be violating the above assumptions in subtle ways.
[Overfitting](https://en.wikipedia.org/wiki/Overfitting) is in some ways a special case of this - an overfit model uses more features/parameters than is "justified" by the data (essentially by the *dimensionality* of the data, as measured by $n$ the number of observations). As the number of features approaches the number of observations, linear regression still "works", but it starts giving fairly perverse results. In particular, it results in a model that fails to *generalize* - and so the core goal of prediction and explanatory power is undermined.
How is this related to well and ill-posed problems? It's not clearly a no solution or multiple solution case, but it does fall in the third category - overfitting results in fitting to the "noise" in the data, which means the particulars of one random sample or another (different initial conditions) will result in dramatically different models.
## Two Equations with Two Unknowns (well-posed)
\begin{align}
x-y = -1
\end{align}
\begin{align}
3x+y = 9
\end{align}
\begin{align}
\begin{bmatrix}
1 & -1 \\
3 & 1
\end{bmatrix}
\begin{bmatrix}
x \\
y
\end{bmatrix}
=
\begin{bmatrix}
-1 \\
9
\end{bmatrix}
\end{align}
```
import numpy as np
A = np.array([[1, -1], [3, 1]])
b = [[-1],[9]]
solution = np.linalg.solve(A, b)
print(solution)
x = solution[0][0]
y = solution[1][0]
print('\n')
print("x:", x)
print("y:", y)
```
## Two Equations with Three Unknowns (not well-posed)
\begin{align}
x-y+z = -1
\end{align}
\begin{align}
3x+y-2z = 9
\end{align}
\begin{align}
\begin{bmatrix}
1 & -1 & 1 \\
3 & 1 & -2
\end{bmatrix}
\begin{bmatrix}
x \\
y \\
z
\end{bmatrix}
=
\begin{bmatrix}
-1 \\
9
\end{bmatrix}
\end{align}
```
import numpy as np
A = np.array([[1, -1, 1], [3, 1, -2]])
b = [[-1],[9]]
solution = np.linalg.solve(A, b)
print(solution)
```
You can reduce these formulas, but there is no single solution, there are infinitely many solutions where the solution to at least one of these variables must be a function of the other variables.
Example:
<https://www.youtube.com/watch?v=tGPSEXVYw_o>
# Generalization in Machine Learning
The goal of machine learning is to end up with a model that can predict well on new data that it has never seen before. This is sometimes called "out of sample accuracy". This is what we are simulating when we do a train-test-split. We allow or model to fit to the training dataset and then we test its ability to generalize to new data by evaluating its accuracy on a test dataset. We want models that will be usable on new data indefinitely that way we can train them once and then reap the rewards of accurate predictions for a long time to come.
## Underfitting
An underfit model will not perform well on the test data and will also not generalize to new data. Because of this, we can usually detect it easily (it just performs poorly in all situations). Because it's easy to identify we either remedy it quickly or move onto new methods.
```
X = np.array([1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20])
y = np.array([10,9,8,7,6,5,4,3,2,1,1,2,3,4,5,6,7,8,9,10])
import matplotlib.pyplot as plt
plt.scatter(X,y)
plt.show()
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=42)
plt.scatter(X_train, y_train)
plt.show()
plt.scatter(X_test, y_test)
plt.show()
from sklearn.linear_model import LinearRegression
X_train = X_train.reshape(-1, 1)
model = LinearRegression().fit(X_train, y_train)
model.score(X_train, y_train)
beta_0 = model.intercept_
beta_1 = model.coef_[0]
print("Slope Coefficient: ", beta_1)
print("\nIntercept Value: ", beta_0)
plt.scatter(X_train, y_train)
y_hat = [beta_1*x + beta_0 for x in X]
plt.plot(x, y_hat)
plt.show()
plt.scatter(X_test, y_test)
y_hat = [beta_1*x + beta_0 for x in X]
plt.plot(x, y_hat)
plt.show()
```
## Overfitting
Lets explore the problem of overfitting (and possible remedy - Ridge Regression) in the context of some housing data.
```
import pandas as pd
from sklearn.datasets import load_boston
from sklearn.preprocessing import scale
# Load and Scale the Data
boston = load_boston()
boston.data = scale(boston.data) # Very helpful for regularization!
# Put it in a dataframe
df = pd.DataFrame(boston.data, columns=boston.feature_names)
df['Price'] = boston.target
df.head()
```
## preprocessing.scale(x) does the same thing as preprocessing.StandardScaler()
The difference is that `.scale(x)` is a function (lowercase naming convention)
`StandardScaler()` is a class (uppercase naming convention) with some extra functionality, they will both scale our data equally well.
```
?scale
from sklearn.preprocessing import StandardScaler
# Load and scale the data
boston = load_boston()
scaler = StandardScaler()
boston.data = scaler.fit_transform(boston.data)
# Put it in a dataframe
df = pd.DataFrame(boston.data, columns=boston.feature_names)
df['Price'] = boston.target
df.head()
df.shape
```
## OLS Baseline Model
```
# Let's try good old least squares!
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
X = df.drop('Price', axis='columns')
y = df.Price
lin_reg = LinearRegression().fit(X, y)
mean_squared_error(y, lin_reg.predict(X))
```
That seems like a pretty good score, but...

Chances are this doesn't generalize very well. You can verify this by splitting the data to properly test model validity.
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=43)
lin_reg_split = LinearRegression().fit(X_train, y_train)
print(mean_squared_error(y, lin_reg_split.predict(X)))
print(mean_squared_error(y_test, lin_reg_split.predict(X_test)))
```
Oops! 💥 - You have overfitting if you are fitting well to training data, but not generalizing well to test data.
### What can we do?
- Use fewer features - sure, but it can be a lot of work to figure out *which* features, and (in cases like this) there may not be any good reason to really favor some features over another.
- Get more data! This is actually a pretty good approach in tech, since apps generate lots of data all the time (and we made this situation by artificially constraining our data). But for case studies, existing data, etc. it won't work.
- **Regularize!**
## Regularization just means "add bias"
OK, there's a bit more to it than that. But that's the core intuition - the problem is the model working "too well", so fix it by making it harder for the model!
It may sound strange - a technique that is purposefully "worse" - but in certain situations, it can really get results.
What's bias? In the context of statistics and machine learning, bias is when a predictive model fails to identify relationships between features and the output. In a word, bias is *underfitting*.
We want to add bias to the model because of the [bias-variance tradeoff](https://en.wikipedia.org/wiki/Bias%E2%80%93variance_tradeoff) - variance is the sensitivity of a model to the random noise in its training data (i.e. *overfitting*), and bias and variance are naturally (inversely) related. Increasing one will always decrease the other, with regards to the overall generalization error (predictive accuracy on unseen data).
Visually, the result looks like this:

The blue line is overfit, using more dimensions than are needed to explain the data and so much of the movement is based on noise and won't generalize well. The green line still fits the data, but is less susceptible to the noise - depending on how exactly we parameterize "noise" we may throw out actual correlation, but if we balance it right we keep that signal and greatly improve generalizability.
### Look carefully at the above plot and think of ways you can quantify the difference between the blue and green lines...
```
# Now with regularization via ridge regression
from sklearn.linear_model import Ridge
ridge_reg = Ridge().fit(X, y)
mean_squared_error(y, ridge_reg.predict(X))
# The score is a bit worse than OLS - but that's expected (we're adding bias)
# Let's try split
ridge_reg_split = Ridge(alpha=0).fit(X_train, y_train)
mean_squared_error(y_test, ridge_reg_split.predict(X_test))
# A little better (to same test split w/OLS) - can we improve it further?
# We just went with defaults, but as always there's plenty of parameters
help(Ridge)
```
How to tune alpha? For now, let's loop and try values.
(For longterm/stretch/next week, check out [cross-validation](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.RidgeCV.html#sklearn.linear_model.RidgeCV).)
```
alphas = []
mses = []
for alpha in range(0, 200, 1):
ridge_reg_split = Ridge(alpha=alpha).fit(X_train, y_train)
mse = mean_squared_error(y_test, ridge_reg_split.predict(X_test))
print(alpha, mse)
alphas.append(alpha)
mses.append(mse)
from matplotlib.pyplot import scatter
scatter(alphas, mses);
```
## What's the intuition? What are we doing?
The `alpha` parameter corresponds to the weight being given to the extra penalty being calculated by [Tikhonov regularization](https://en.wikipedia.org/wiki/Tikhonov_regularization) (this parameter is sometimes referred to as $\lambda$ in the context of ridge regression).
Normal linear regression (OLS) minimizes the **sum of square error of the residuals**.
Ridge regression minimizes the **sum of square error of the residuals** *AND* **the squared slope of the fit model, times the alpha parameter**.
This is why the MSE for the first model in the for loop (`alpha=0`) is the same as the MSE for linear regression - it's the same model!
As `alpha` is increased, we give more and more penalty to a steep slope. In two or three dimensions this is fairly easy to visualize - beyond, think of it as penalizing coefficient size. Each coefficient represents the slope of an individual dimension (feature) of the model, so ridge regression is just squaring and summing those.
So while `alpha=0` reduces to OLS, as `alpha` approaches infinity eventually the penalty gets so extreme that the model will always output every coefficient as 0 (any non-zero coefficient resulting in a penalty that outweighs whatever improvement in the residuals), and just fit a flat model with intercept at the mean of the dependent variable.
Of course, what we want is somewhere in-between these extremes. Intuitively, what we want to do is apply an appropriate "cost" or penalty to the model for fitting parameters, much like adjusted $R^2$ takes into account the cost of adding complexity to a model. What exactly is an appropriate penalty will vary, so you'll have to put on your model comparison hat and give it a go!
PS - scaling the data helps, as that way this cost is consistent and can be added uniformly across features, and it is simpler to search for the `alpha` parameter.
### Bonus - magic! ✨
Ridge regression doesn't just reduce overfitting and help with the third aspect of well-posed problems (poor generalizability). It can also fix the first two (no unique solution)!
```
df_tiny = df.sample(10, random_state=27)
print(df_tiny.shape)
X = df_tiny.drop('Price', axis='columns')
y = df_tiny.Price
lin_reg = LinearRegression().fit(X, y)
lin_reg.score(X, y) # Perfect multi-collinearity!
# NOTE - True OLS would 💥 here
# scikit protects us from actual error, but still gives a poor model
ridge_reg = Ridge().fit(X, y)
ridge_reg.score(X, y) # More plausible (not "perfect")
# Using our earlier test split
mean_squared_error(y_test, lin_reg.predict(X_test))
# Ridge generalizes *way* better (and we've not even tuned alpha)
mean_squared_error(y_test, ridge_reg.predict(X_test))
# e.g. (x1^2 + x2^2 + ...) * alpha is the extra penalty from Ridge
from sklearn.linear_model import RidgeCV
ridgecv = RidgeCV(alphas=[1e-3, 1e-2, 1e-1, 1, 10, 100, 1000]).fit(X, y)
ridgecv.score(X, y)
mean_squared_error(y_test, ridgecv.predict(X_test))
```
## And a bit more math
The regularization used by Ridge Regression is also known as **$L^2$ regularization**, due to the squaring of the slopes being summed. This corresponds to [$L^2$ space](https://en.wikipedia.org/wiki/Square-integrable_function), a metric space of square-integrable functions that generally measure what we intuitively think of as "distance" (at least, on a plane) - what is referred to as Euclidean distance.
The other famous norm is $L^1$, also known as [taxicab geometry](https://en.wikipedia.org/wiki/Taxicab_geometry), because it follows the "grid" to measure distance like a car driving around city blocks (rather than going directly like $L^2$). When referred to as a distance this is called "Manhattan distance", and can be used for regularization (see [LASSO](https://en.wikipedia.org/wiki/Lasso_(statistics%29), which [uses the $L^1$ norm](https://www.quora.com/What-is-the-difference-between-L1-and-L2-regularization-How-does-it-solve-the-problem-of-overfitting-Which-regularizer-to-use-and-when)).
All this comes down to - regularization means increasing model bias by "watering down" coefficients with a penalty typically based on some sort of distance metric, and thus reducing variance (overfitting the model to the noise in the data). It gives us another lever to try and another tool for our toolchest!
## Putting it all together - one last example
The official scikit-learn documentation has many excellent examples - [this one](https://scikit-learn.org/stable/auto_examples/linear_model/plot_ols_ridge_variance.html#sphx-glr-auto-examples-linear-model-plot-ols-ridge-variance-py) illustrates how ridge regression effectively reduces the variance, again by increasing the bias, penalizing coefficients to reduce the effectiveness of features (but also the impact of noise).
```
Due to the few points in each dimension and the straight line that linear regression uses to follow these points as well as it can, noise on the observations will cause great variance as shown in the first plot. Every line’s slope can vary quite a bit for each prediction due to the noise induced in the observations.
Ridge regression is basically minimizing a penalised version of the least-squared function. The penalising shrinks the value of the regression coefficients. Despite the few data points in each dimension, the slope of the prediction is much more stable and the variance in the line itself is greatly reduced, in comparison to that of the standard linear regression
```
```
# Code source: Gaël Varoquaux
# Modified for documentation by Jaques Grobler
# License: BSD 3 clause
import numpy as np
import matplotlib.pyplot as plt
from sklearn import linear_model
X_train = np.c_[.5, 1].T
y_train = [.5, 1]
X_test = np.c_[0, 2].T
np.random.seed(0)
classifiers = dict(ols=linear_model.LinearRegression(),
ridge=linear_model.Ridge(alpha=.1))
for name, clf in classifiers.items():
fig, ax = plt.subplots(figsize=(4, 3))
for _ in range(6):
this_X = .1 * np.random.normal(size=(2, 1)) + X_train
clf.fit(this_X, y_train)
ax.plot(X_test, clf.predict(X_test), color='gray')
ax.scatter(this_X, y_train, s=3, c='gray', marker='o', zorder=10)
clf.fit(X_train, y_train)
ax.plot(X_test, clf.predict(X_test), linewidth=2, color='blue')
ax.scatter(X_train, y_train, s=30, c='red', marker='+', zorder=10)
ax.set_title(name)
ax.set_xlim(0, 2)
ax.set_ylim((0, 1.6))
ax.set_xlabel('X')
ax.set_ylabel('y')
fig.tight_layout()
plt.show()
```
Between the first and the second graph, we have decreased the slope (penalized or watered-down coefficients), but we have less variance between our lines)
# Assignment
Following is data describing characteristics of blog posts, with a target feature of how many comments will be posted in the following 24 hours.
https://archive.ics.uci.edu/ml/datasets/BlogFeedback
Investigate - you can try both linear and ridge. You can also sample to smaller data size and see if that makes ridge more important. Don't forget to scale!
Focus on the training data, but if you want to load and compare to any of the test data files you can also do that.
Note - Ridge may not be that fundamentally superior in this case. That's OK! It's still good to practice both, and see if you can find parameters or sample sizes where ridge does generalize and perform better.
When you've fit models to your satisfaction, answer the following question:
```
Did you find cases where Ridge performed better? If so, describe (alpha parameter, sample size, any other relevant info/processing). If not, what do you think that tells you about the data?
```
You can create whatever plots, tables, or other results support your argument. In this case, your target audience is a fellow data scientist, *not* a layperson, so feel free to dig in!
```
# TODO - write some code!
from google.colab import drive
drive.mount('/content/gdrive')
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import RidgeCV
from sklearn.linear_model import Ridge
from sklearn.preprocessing import scale
from sklearn.metrics import mean_squared_error
import pandas as pd
df = pd.read_csv('/content/gdrive/My Drive/blogData_train.csv', header=None)
df.head()
df.isnull().sum()
df = scale(df)
df
df = pd.DataFrame(df)
df
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=45)
y = df[280]
X = df.drop(280, axis='columns')
X.head()
lin_reg_split = LinearRegression().fit(X_train, y_train)
print(mean_squared_error(y, lin_reg_split.predict(X)))
print(mean_squared_error(y_test, lin_reg_split.predict(X_test)))
ridge = Ridge(300).fit(X_train, y_train)
print(mean_squared_error(y_train, ridge.predict(X_train)))
print(mean_squared_error(y_test, ridge.predict(X_test)))
ridge_cv = RidgeCV([.001,.01,1,10,100,10000]).fit(X_train, y_train)
print(mean_squared_error(y_train, ridge_cv.predict(X_train)))
print(mean_squared_error(y_test, ridge_cv.predict(X_test)))
```
# Resources and stretch goals
Resources:
- https://www.quora.com/What-is-regularization-in-machine-learning
- https://blogs.sas.com/content/subconsciousmusings/2017/07/06/how-to-use-regularization-to-prevent-model-overfitting/
- https://machinelearningmastery.com/introduction-to-regularization-to-reduce-overfitting-and-improve-generalization-error/
- https://towardsdatascience.com/ridge-and-lasso-regression-a-complete-guide-with-python-scikit-learn-e20e34bcbf0b
- https://stats.stackexchange.com/questions/111017/question-about-standardizing-in-ridge-regression#111022
Stretch goals:
- Revisit past data you've fit OLS models to, and see if there's an `alpha` such that ridge regression results in a model with lower MSE on a train/test split
- Yes, Ridge can be applied to classification! Check out [sklearn.linear_model.RidgeClassifier](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.RidgeClassifier.html#sklearn.linear_model.RidgeClassifier), and try it on a problem you previous approached with a different classifier (note - scikit LogisticRegression also automatically penalizes based on the $L^2$ norm, so the difference won't be as dramatic)
- Implement your own function to calculate the full cost that ridge regression is optimizing (the sum of squared residuals + `alpha` times the sum of squared coefficients) - this alone won't fit a model, but you can use it to verify cost of trained models and that the coefficients from the equivalent OLS (without regularization) may have a higher cost
| github_jupyter |
# Data Visualization in Python
## Introduction
In this module, you will learn to quickly and flexibly make a wide series of visualizations for exploratory data analysis and communicating to your audience. This module contains a practical introduction to data visualization in Python and covers important rules that any data visualizer should follow.
## Learning Objectives
* Learn critical rules about data visualization (using the correct graph types, correctly labeling all visual encodings, properly sourcing data).
* Become familiar with a core base of data visualization tools in Python - specifically matplotlib and seaborn.
* Start to develop the ability to conceptualize what visualizations are going to best reveal various types of patterns in your data.
* Learn more about Illinois administrative data with exploratory analyses.
### Table of contents
+ [Setup - Load Python Packages](#Setup---Load-Python-Packages)
+ [Setup - Loading Data](#Setup---Loading-Data)
+ [Our First Chart in matplotlib](#Our-First-Chart-in-matplotlib)
+ [A note on data sourcing](#A-Note-on-Data-Sourcing)
+ [Layering in matplotlib](#Layering-in-Matplotlib)
+ [Our first chart in Seaborn](#Our-First-Chart-in-seaborn)
+ [Choosing a Data Visualization package](#Choosing-a-Data-Visualization-Package)
## Setup - Load Python Packages
```
import pandas as pd
import matplotlib as mplib
import matplotlib.pyplot as plt # visualization package
import seaborn as sns
# database connections
from sqlalchemy import create_engine # to get data from database
from sqlalchemy import __version__ as sql_version
from sqlalchemy import inspect
# so images get plotted in the notebook
%matplotlib inline
```
## Setup - Loading Data
```
# set up sqlalchemy engine
engine = create_engine('postgresql://10.10.2.10/appliedda')
# See all available schemas:
pd.read_sql("SELECT schema_name FROM information_schema.schemata LIMIT 10;", engine)
# We can look at the tables within each schema:
# pd.read_sql("SELECT * FROM pg_tables WHERE schemaname = 'ides'",engine)
pd.read_sql("SELECT * FROM pg_tables WHERE schemaname = 'ada_class3'",engine)
# We can look at column names within tables:
pd.read_sql("SELECT * FROM information_schema.columns WHERE table_schema = 'ada_class3' AND table_name = 'il_wage_hh_recipient'",engine)
select_string = "SELECT ssn, year, SUM(wage) total_wages FROM ada_class3.il_wage_hh_recipient"
select_string += " WHERE year in (2005, 2010, 2015) AND quarter = 1"
select_string += " GROUP BY ssn, year"
print(select_string)
person_wages = pd.read_sql(select_string, engine)
print("Number of rows returned: " + str(len(person_wages)))
```
## Our First Chart in matplotlib
- [Back to top](#Introduction)
Below, we make our first chart in matplotlib. We'll come back to the choice of this particular library in a second, but for now just appreciate that the visualization is creating sensible scales, tick marks, and gridlines on its own.
```
## Wages often have a very strong right skew:
max_wage = person_wages["total_wages"].max()
print("Maximum wage = " + str(max_wage))
## But most people earn under 15,000 in a quarter:
(person_wages["total_wages"] < 15000).value_counts()
## So let's just look at the wages under $15,000
person_wages_lim = person_wages[person_wages["total_wages"] < 15000]
# Make a simple histogram:
plt.hist(person_wages_lim.total_wages)
plt.show()
## We can change options within the hist function (e.g. number of bins, color, transparency:
plt.hist(person_wages_lim.total_wages, bins=20, facecolor="purple", alpha=0.5)
## And we can affect the plot options too:
plt.xlabel('Quarterly Income')
plt.ylabel('Number of Recipients')
plt.title('Most Recipients Earn Under $6,000 per Quarter')
## And add Data sourcing:
### xy are measured in percent of axes length, from bottom left of graph:
plt.annotate('Source: IL IDES & IDHS', xy=(0.7,-0.2), xycoords="axes fraction")
## We use plt.show() to display the graph once we are done setting options:
plt.show()
```
### A Note on Data Sourcing
Data sourcing is a critical aspect of any data visualization. Although here we are simply referencing the agencies that created the data, it is ideal to provide as direct of a path as possible for the viewer to find the data the graph is based on. When this is not possible (e.g. the data is sequestered), directing the viewer to documentation or methodology for the data is a good alternative. Regardless, providing clear sourcing for the underlying data is an **absolutely requirement** of any respectable visualization, and further builds trusts and enables reproducibility.
### Layering in Matplotlib
This functionality - where we can make consecutive changes to the same plot - also allows us to layer on multiple plots. By default, the first graph you create will be at the bottom, with ensuing graphs on top.
Below, we see the 2005 histogram, in blue, is beneath the 2015 histogram, in orange. You might also notice that the distribution of income for welfare recipients has shifted upward over that ten year period.
```
plt.hist(person_wages_lim[person_wages_lim["year"] == 2005].total_wages, facecolor="blue", alpha=0.5)
plt.hist(person_wages_lim[person_wages_lim["year"] == 2015].total_wages, facecolor="orange", alpha=0.5)
plt.annotate('Source: IL IDES & IDHS', xy=(0.7,-0.2), xycoords="axes fraction")
plt.show()
```
### Our First Chart in seaborn
Below, we quickly use pandas to create an aggregation of our wages data - the average wages by year. Then we pass the data to the barplot function in the `seaborn` function, which recall we imported as `sns` for short.
```
## Calculate average wages by year:
avg_annual_wages = person_wages.groupby('year')['total_wages'].mean().reset_index()
avg_annual_wages.columns = ['year','average_wages']
print(type(avg_annual_wages))
print("***********")
print(avg_annual_wages)
## Barplot function
# Note we can reference column names (in quotes) in the specified data:
sns.barplot(x='year', y='average_wages', data=avg_annual_wages)
plt.show()
```
You might notice that if you don't include plt.show(), Jupyter will still produce a chart. However this is not the case in other environments. So we will continue using plt.show() to more formally ask for Python to display the chart we have constructed, after adding all layers and setting all options.
```
## Seaborn has a great series of charts for showing distributions across a categorical variable:
sns.factorplot(x='year', y='total_wages', hue='year', data=person_wages_lim, kind='box')
plt.show()
## Other options for the 'kind' argument include 'bar' and 'violin'
```
Already you might notice some differences between matplotlib and seaborn - at the very least seaborn allows us to more easily reference column names within a pandas dataframe, whereas matplotlib clearly has a plethora of options.
## Choosing a Data Visualization Package
- [Back to top](#Introduction)
There are many excellent data visualiation modules available in Python, but for the tutorial we will stick to the tried and true combination of `matplotlib` and `seaborn`. You can read more about different options for data visualization in Python in the [More Resources](#More-Resources:) section at the bottom of this notebook.
`matplotlib` is very expressive, meaning it has functionality that can easily account for fine-tuned graph creation and adjustment. However, this also means that `matplotlib` is somewhat more complex to code.
`seaborn` is a higher-level visualization module, which means it is much less expressive and flexible than matplotlib, but far more concise and easier to code.
It may seem like we need to choose between these two approaches, but this is not the case! Since `seaborn` is itself written in `matplotlib` (you will sometimes see `seaborn` be called a `matplotlib` 'wrapper'), we can use `seaborn` for making graphs quickly and then `matplotlib` for specific adjustments. When you see `plt` referenced in the code below, we are using `matplotlib`'s pyplot submodule.
`seaborn` also improves on `matplotlib` in important ways, such as the ability to more easily visualize regression model results, creating small multiples, enabling better color palettes, and improve default aesthetics. From [`seaborn`'s documentation](https://seaborn.pydata.org/introduction.html):
> If matplotlib 'tries to make easy things easy and hard things possible', seaborn tries to make a well-defined set of hard things easy too.
```
## Seaborn offers a powerful tool called FacetGrid for making small multiples of matplotlib graphs:
### Create an empty set of grids:
facet_histograms = sns.FacetGrid(person_wages_lim, col='year', hue='year')
## "map' a histogram to each grid:
facet_histograms = facet_histograms.map(plt.hist, 'total_wages')
## Data Sourcing:
plt.annotate('Source: IL IDES & IDHS', xy=(0.6,-0.35), xycoords="axes fraction")
plt.show()
## Alternatively, you can create and save several charts:
for i in set(person_wages_lim["year"]):
tmp = person_wages_lim[person_wages_lim["year"] == i]
plt.hist(tmp["total_wages"])
plt.xlabel('Total Wages')
plt.ylabel('Number of Recipients')
plt.title(str(i))
plt.annotate('Source: IL IDES & IDHS', xy=(0.7,-0.2), xycoords="axes fraction")
filename = "graph_" + str(i) + ".pdf"
plt.savefig(filename)
plt.show()
```
### Seaborn and matplotlib
Below, we use seaborn for setting an overall aesthetic style and then faceting (created small multiples). We then use matplotlib to set very specific adjustments - things like adding the title, adjusting the locations of the plots, and sizing th graph space. This is a pretty protoyptical use of the power of these two libraries together.
More on [Seaborn's set_style function](https://seaborn.pydata.org/generated/seaborn.set_style.html).
More on [matplotlib's figure (fig) API](https://matplotlib.org/api/figure_api.html).
```
# Seaborn's set_style function allows us to set many aesthetic parameters.
sns.set_style("whitegrid")
facet_histograms = sns.FacetGrid(person_wages_lim, col='year', hue='year')
facet_histograms.map(plt.hist, 'total_wages')
## We can still change options with matplotlib, using facet_histograms.fig
facet_histograms.fig.subplots_adjust(top=0.85)
facet_histograms.fig.suptitle("Recipients Income Gains Jumped in 2015", fontsize=14)
facet_histograms.fig.set_size_inches(10,5)
## Add a legend for hue (color):
facet_histograms = facet_histograms.add_legend()
## Data Sourcing:
plt.annotate('Source: IL DOC & IDES', xy=(0.6,-0.35), xycoords="axes fraction")
plt.show()
```
## Visual Encodings
We often start with charts that use 2-dimensional position (like a scatterplot) or that use height (like histograms and bar charts). This is because these visual encodings - the visible mark that represents the data - are particularly perceptually strong. This means that when humans view these visual encodings, they are more accurate in estimating the underlying numbers than encodings like size (think circle size in a bubble chart) or angle (e.g. pie chart).
For more information on visual encodings and data visualization theory, see:
* [Designing Data Visualizations, Chapter 4](http://www.safaribooksonline.com/library/view/designing-data-visualizations/9781449314774/ch04.html) by Julie Steele and Noah Iliinsky
* Now You See It - book by Stephen Few
```
select_string = "SELECT year, avg(total_wages) avg_wages, count(*) num_recipients"
select_string += " FROM (SELECT ssn, year, SUM(wage) total_wages FROM ada_class3.il_wage_hh_recipient"
select_string += " WHERE quarter = 1"
select_string += " GROUP BY ssn, year) tmp"
select_string += " GROUP BY year"
select_string += " ORDER BY year"
print(select_string)
yearly_avg_wages = pd.read_sql(select_string, engine)
yearly_avg_wages = yearly_avg_wages.sort_values('year')
yearly_avg_wages.head()
## We can pass a single value to a the tsplot function to get a simple line chart:
sns.tsplot(data=yearly_avg_wages['avg_wages'], color="#179809")
## Data Sourcing:
plt.annotate('Source: IL IDES & IHDS', xy=(0.8,-0.20), xycoords="axes fraction")
plt.show()
```
### Using Hex Codes for Color
- [Back to top](#Introduction)
In the graph above, you can see I set the color of the graph with pund sign `#` followed by a series of six numbers. This is a hexcode - which is short for hexadecimal code. A hexadecimal code lets you specify one of over 16 million colors using combinations of red, green, and blue. It first has two digits for red, then two digits for green, and lastly two digits for blue: `#RRGGBB`
Further, these codes allow for you to specify sixteen integers (thus hexadecimal) for each digits, in this order:
(0,1,2,3,4,5,6,7,8,9,A,B,C,D,E,F)
Over time, it gets easier to read these codes. For instance, above, I used the hex code "#179809". Understanding how hex codes work, I can see that there is a relatively low number for red (17) and fairly high number for green (98) and another low number for blue (09). Thus it shouldn't be too surprising that a green color resulted in the graph.
Tools like [Adobe Color](https://color.adobe.com) and this [Hex Calculator](https://www.w3schoosl.com/colors/colors_hexadecimal.asp) can help you get used to this system.
Most modern browsers also support eight digit hex codes, in which the first two enable transparency, which is often called 'alpha' in data visualization: `#AARRGGBB`
```
## We can add the time argument to set the x-axis correctly. And let's change the color, since we can:
sns.tsplot(data=yearly_avg_wages['avg_wages'], time=yearly_avg_wages['year'], color="#B088CD")
# Color Note: B088CD
## The highest values are red 'B0' and blue 'CD', so we can expect a mix of those
## Further this is high in all three colors, so it'll be light, not dark
## Data Sourcing:
plt.annotate('Source: IL IDES & IDHS', xy=(0.8,-0.20), xycoords="axes fraction")
plt.show()
```
### Saving Charts As a Variable
Although as you can see above, we can immediately print our plots on a page, it is generally better to save them as variable. We can then alter the charts over several lines before finally displaying them with the `show()` function, which comes from the `matplotlib` `pyplot` module we loaded earlier.
```
## Save the line chart as 'graph'
graph = sns.tsplot(data=yearly_avg_wages['avg_wages'], time=yearly_avg_wages['year'])
## To add data labels, we loop over each row and use graph.text()
for i, row, in yearly_avg_wages.iterrows():
graph.text(row["year"] + 0.05, row["avg_wages"] - 50, int(row["year"]))
## Now change x-axis and y-axis labels:
graph.set(xlabel="Year", ylabel="Average Annual Wages")
graph.set(title="Average Wages Over Time")
plt.annotate('Source: IL IDES & IDHS', xy=(0.8,-0.20), xycoords="axes fraction")
## Then display the plot:
plt.show()
# We can also look at a scatterplot of the number of people and averages wages in each year:
scatter = sns.lmplot(x='num_recipients', y='avg_wages', data=yearly_avg_wages, fit_reg=False)
scatter.set(xlabel="Number of Recipients", ylabel="Average Annual Wages", title="Number and Wages of IL Welfare Recipients")
## Sourcing:
plt.annotate('Source: IL IDES & IDHS', xy=(0.8,-0.20), xycoords="axes fraction")
plt.show()
```
### Directed Scatterplot
A directed scatterplot still uses one point for each year, but then uses the x-axis and the y-axis for variabes. In order to maintain the ordinal relationship, a line is drawn between the years. To do this in seaborn, we actually use sns.FacetGrid, which allows us to overlay different plots together. Specifically, it lets us overlay a scatterplot (`plt.scatter` and a line chart `plt.plot`).
### An Important Note on Graph Titles:
The title of a visualization occupies the most valuable real estate on the page. If nothing else, you can be reasonably sure a viewer will at least read the title and glance at your visualization. This is why you want to put thought into making a clear and effective title that acts as a **narrative** for your chart. Many novice visualizers default to an **explanatory** title, something like: "Income and Number of Recipients over Time (2005-2015)". This title is correct - it just isn't very useful. This is particularly true since any good graph will have explained what the visualization is through the axes and legends. Instead, use the title to reinforce and explain the core point of the visualization. It should answer the question "Why is this graph important?" and focus the viewer onto the most critical take-away.
```
cncted_scatter = sns.FacetGrid(data=yearly_avg_wages, size=7)
cncted_scatter.map(plt.scatter, 'num_recipients', 'avg_wages', color="#A72313")
cncted_scatter.map(plt.plot, 'num_recipients', 'avg_wages', color="#A72313")
cncted_scatter.set(title="Rising Wages of Welfare Recipients", xlabel="Number of Recipients", ylabel="Average Wages")
## Adding data labels:
for i, row, in yearly_avg_wages.iterrows():
plt.text(row["num_recipients"], row["avg_wages"], int(row["year"]))
## Sourcing:
plt.annotate('Source: IL IDES & IDHS', xy=(0.8,-0.10), xycoords="axes fraction")
plt.show()
```
### Exporting Completed Graphs
When you are satisfied with your visualization, you may want to save a a copy outside of your notebook. You can do this with `matplotlib`'s savefig function. You simply need to run:
plt.savefig("fileName.fileExtension")
The file extension is actually surprisingly important. Image formats like png and jpeg are actually **not ideal**. These file formats store your graph as a giant grid of pixels, which is space-efficient, but can't be edited later. Saving your visualizations instead as a PDF is strongly advised. PDFs are a type of vector image, which means all the component of the graph will be maintained.
With PDFs, you can later open the image in a program like Adobe Illustrator and make changes like the size or typeface of your text, move your legends, or adjust the colors of your visual encodings. All of this would be impossible with a png or jpeg.
```
cncted_scatter = sns.FacetGrid(data=yearly_avg_wages, size=7)
cncted_scatter.map(plt.scatter, 'num_recipients', 'avg_wages', color="#A72313")
cncted_scatter.map(plt.plot, 'num_recipients', 'avg_wages', color="#A72313")
cncted_scatter.set(title="Rising Wages of Welfare Recipients", xlabel="Number of Recipients", ylabel="Average Wages")
## Adding data labels:
for i, row, in yearly_avg_wages.iterrows():
plt.text(row["num_recipients"], row["avg_wages"], int(row["year"]))
## Sourcing:
plt.annotate('Source: IL IDES & IDHS', xy=(0.8,-0.10), xycoords="axes fraction")
plt.savefig('dscatter.png')
plt.savefig('dscatter.pdf')
```
## Exercises & Practice
- [Back to top](#Introduction)
### Excercise 1: Heatmap
Below, I query the database for the average sentence of men, broken out by their race and education level. I then format this data into a wider form using pandas - where each row corresponds to an education level and each column corresponds to a race. This grid is format that `seaborn`'s heatmap function is expecting. Understanding the relationship between your data and the resulting graph is an important aspect of data visualization - and you can really only master this with practice.
Query one of the tables again and create dataframe in the correct format, then pass that along to seaborn's heatmap function. Use the code you learned above to add a title, better axis labels, and data sourcing.
Note that the color map used here `viridis` is a scientifically derived color palette meant to be perceptually linear. The color maps `inferno`, `plasma` and `magama` also all meet this criteria.
#### More information:
* [seaborn heatmap documentation](http://seaborn.pydata.org/generated/seaborn.heatmap.html)
* [matplotlib color map documentation](http://matplotlib.org/users/colormap.html)
```
## Querying Average Jailtime by Race and Gender
select_string = "SELECT race, educlvl, avg(jailtime) as avg_jailtime"
select_string += " FROM ildoc.ildoc_admit"
select_string += " WHERE sex = 'M'"
select_string += " GROUP BY race, educlvl"
print(select_string)
jail = pd.read_sql(select_string, engine)
## Format the data for a heatmap:
jail = jail.pivot("educlvl", "race", "avg_jailtime")
jail = jail.round(0)
jail = jail.apply(pd.to_numeric)
print(jail)
## Create a heatmap, with annotations:
sns.heatmap(jail, annot=True, fmt='g', cmap="viridis")
plt.show()
## Enter your code for excercise 1 here:
```
### Exercise 2
Below, we pull two continuous variables from the Illinois Department of Employment Security, summed over each employer. See if you can pass this data to the sns.jointplot() function. I have filled in some of the arguments for you, while others need completion.
```
pd.read_sql("SELECT * FROM ada_class3.il_des_subset_2014q3 LIMIT 5;",engine)
## Querying Total Wages and Jobs by Employer
select_string = "SELECT name_legal, sum(total_wages) as agg_wages, sum(total_jobs) as agg_jobs"
select_string += " FROM ada_class3.il_des_subset_2014q3"
select_string += " GROUP BY name_legal"
print(select_string)
## Run SQL query:
employers = pd.read_sql(select_string, engine)
print(len(employers))
## Take a one percent sample to ease computational burden:
employers_lim = employers.sample(frac=0.01)
print(len(employers_lim))
## Fill in the arguments (x, y, data, kind) below to get the visualiztion to run.
sns.jointplot(x=, y=, data=, kind=, color="#137B80", marginal_kws={"bins":30})
plt.show()
```
### Exercise 3
Let's see if we can use seaborn's FacetGrid to create small multiple scatterplots. First you need to query a database and get at least one categorical variable and at least two continuous variables (floats).
Then try passing this data to the FacetGrid function from `seaborn` and the scatter function from `matplotlib`.
[FacetGrid Documentation](http://seaborn.pydata.org/examples/many_facets.html)
```
## Pseudo-code to get you started:
grid = sns.FacetGrid(dataframe, col = "categorical_var", hue="categorical_var", col_wrap=2)
grid.map(plt.scatter("x_var", "y_var"))
## Enter your code for excercise 3 here:
```
### Exercise 4:
Test your mettle. Check out the seaborn [data visualization gallery](http://seaborn.pydata.org/examples) and see if you can implement an interesting visualization.
## More Resources
* [A Thorough Comparison of Python's DataViz Modules](https://dsaber.com/2016/10/02/a-dramatic-tour-through-pythons-data-visualization-landscape-including-ggplot-and-altair)
* [Seaborn Documentation](http://seaborn.pydata.org)
* [Matplotlib Documentation](https://matplotlib.org)
* [Advanced Functionality in Seaborn](blog.insightdatalabs.com/advanced-functionality-in-seaborn)
## Other Python Visualization Libraries
* [Bokeh](http://bokeh.pydata.org)
* [Altair](https://altair-viz.github.io)
* [ggplot](http://ggplot.yhathq.com.com)
* [Plotly](https://plot.ly)
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Text classification with TensorFlow Lite model customization with TensorFlow 2.0
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/examples/blob/master/tensorflow_examples/lite/model_customization/demo/text_classification.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/examples/blob/master/tensorflow_examples/lite/model_customization/demo/text_classification.ipynb">
<img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
View source on GitHub</a>
</td>
</table>
The TensorFlow Lite model customization library simplifies the process of adapting and converting a TensorFlow neural-network model to particular input data when deploying this model for on-device ML applications.
This notebook shows an end-to-end example that utilizes this model customization library to illustrate the adaption and conversion of a commonly-used text classification model to classify movie reviews on a mobile device.
## Prerequisites
To run this example, we first need to install serveral required packages, including model customization package that in github [repo](https://github.com/tensorflow/examples).
```
!pip uninstall -q -y tensorflow google-colab grpcio
!pip install -q tf-nightly
!pip install -q git+https://github.com/tensorflow/examples
```
Import the required packages.
```
from __future__ import absolute_import, division, print_function, unicode_literals
import numpy as np
import os
import tensorflow as tf
assert tf.__version__.startswith('2')
from tensorflow_examples.lite.model_customization.core.data_util.text_dataloader import TextClassifierDataLoader
from tensorflow_examples.lite.model_customization.core.model_export_format import ModelExportFormat
from tensorflow_examples.lite.model_customization.core.task.model_spec import AverageWordVecModelSpec
import tensorflow_examples.lite.model_customization.core.task.text_classifier as text_classifier
```
## Simple End-to-End Example
Let's get some texts to play with this simple end-to-end example. You could replace it with your own text folders.
```
data_path = tf.keras.utils.get_file(
fname='aclImdb',
origin='http://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz',
untar=True)
```
The example just consists of 4 lines of code as shown below, each of which representing one step of the overall process.
1. Load train and test data specific to an on-device ML app.
```
train_data = TextClassifierDataLoader.from_folder(os.path.join(data_path, 'train'), class_labels=['pos', 'neg'])
test_data = TextClassifierDataLoader.from_folder(os.path.join(data_path, 'test'), shuffle=False)
```
2. Customize the TensorFlow model.
```
model = text_classifier.create(train_data)
```
3. Evaluate the model.
```
loss, acc = model.evaluate(test_data)
```
4. Export to TensorFlow Lite model.
```
model.export('movie_review_classifier.tflite', 'text_label.txt', 'vocab.txt')
```
After this simple 4 steps, we could further use TensorFlow Lite model file and label file in on-device applications like in [text classification](https://github.com/tensorflow/examples/tree/master/lite/examples/text_classification) reference app.
## Detailed Process
In above, we tried the simple end-to-end example. The following walks through the example step by step to show more detail.
### Step 1: Load Input Data Specific to an On-device ML App
The IMDB dataset contains 25000 movie reviews for training and 25000 movie reviews for testing from the [Internet Movie Database](https://www.imdb.com/). The dataset have two classes: positive and negative movie reviews.
Download the archive version of the dataset and untar it.
The IMDB dataset has the following directory structure:
<pre>
<b>aclImdb</b>
|__ <b>train</b>
|______ <b>pos</b>: [1962_10.txt, 2499_10.txt, ...]
|______ <b>neg</b>: [104_3.txt, 109_2.txt, ...]
|______ unsup: [12099_0.txt, 1424_0.txt, ...]
|__ <b>test</b>
|______ <b>pos</b>: [1384_9.txt, 191_9.txt, ...]
|______ <b>neg</b>: [1629_1.txt, 21_1.txt]
</pre>
Note that the text data under `train/unsup` folder are unlabeled documents for unsupervised learning and such data should be ignored in this tutorial.
```
data_path = tf.keras.utils.get_file(
fname='aclImdb',
origin='http://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz',
untar=True)
```
Use `TextClassifierDataLoader` to load data.
As for `from_folder()` method, it could load data from the folder. It assumes that the text data of the same class are in the same subdirectory and the subfolder name is the class name. Each text file contains one movie review sample.
Parameter `class_labels` is used to specify which subfolder should be considered. As for `train` folder, this parameter is used to skip `unsup` subfolder.
```
train_data = TextClassifierDataLoader.from_folder(os.path.join(data_path, 'train'), class_labels=['pos', 'neg'])
test_data = TextClassifierDataLoader.from_folder(os.path.join(data_path, 'test'), shuffle=False)
train_data, validation_data = train_data.split(0.9)
```
Take a glance at 25 training data.
```
for text, label in train_data.dataset.take(25):
print ("%s: %s"%(train_data.index_to_label[label.numpy()], text.numpy()))
```
### Step 2: Customize the TensorFlow Model
Create a custom text classifier model based on the loaded data. Currently, we only supports averging word embedding method.
```
model = text_classifier.create(train_data, validation_data=validation_data)
```
Have a look at the detailed model structure.
```
model.summary()
```
### Step 3: Evaluate the Customized Model
Evaluate the result of the model, get the loss and accuracy of the model.
Evaluate the loss and accuracy in `test_data`. If no data is given the results are evaluated on the data that's splitted in the `create` method.
```
loss, acc = model.evaluate(test_data)
```
### Step 4: Export to TensorFlow Lite Model
Convert the existing model to TensorFlow Lite model format that could be later used in on-device ML application. Meanwhile, save the text labels in label file and vocabulary in vocab file.
```
model.export('movie_review_classifier.tflite', 'text_label.txt', 'vocab.txt')
```
The TensorFlow Lite model file and label file could be used in [text classification](https://github.com/tensorflow/examples/tree/master/lite/examples/text_classification) reference app.
In detail, we could add `movie_review_classifier.tflite`, `text_label.txt` and `vocab.txt` in [assets](https://github.com/tensorflow/examples/tree/master/lite/examples/text_classification/android/app/src/main/assets) folder. Meanwhile, change the filenames in [code](https://github.com/tensorflow/examples/blob/master/lite/examples/text_classification/android/app/src/main/java/org/tensorflow/lite/examples/textclassification/TextClassificationClient.java#L43).
Here, we also demonstrate how to use the above files to run and evaluate the TensorFlow Lite model.
```
# Read TensorFlow Lite model from TensorFlow Lite file.
with tf.io.gfile.GFile('movie_review_classifier.tflite', 'rb') as f:
model_content = f.read()
# Read label names from label file.
with tf.io.gfile.GFile('text_label.txt', 'r') as f:
label_names = f.read().split('\n')
# Initialze TensorFlow Lite inpterpreter.
interpreter = tf.lite.Interpreter(model_content=model_content)
interpreter.allocate_tensors()
input_index = interpreter.get_input_details()[0]['index']
output = interpreter.tensor(interpreter.get_output_details()[0]["index"])
# Run predictions on each test data and calculate accuracy.
accurate_count = 0
for i, (text, label) in enumerate(test_data.dataset):
# Pre-processing should remain the same.
text, label = model.preprocess(text, label)
# Add batch dimension and convert to float32 to match with the model's input
# data format.
text = tf.expand_dims(text, 0).numpy()
text = tf.cast(text, tf.float32)
# Run inference.
interpreter.set_tensor(input_index, text)
interpreter.invoke()
# Post-processing: remove batch dimension and find the label with highest
# probability.
predict_label = np.argmax(output()[0])
# Get label name with label index.
predict_label_name = label_names[predict_label]
accurate_count += (predict_label == label.numpy())
accuracy = accurate_count * 1.0 / test_data.size
print('TensorFlow Lite model accuracy = %.4f' % accuracy)
```
Note that preprocessing for inference should be the same as training. Currently, preprocessing contains split the text to tokens by '\W', encode the tokens to ids, the pad the text with `pad_id` to have the length of `sentence_length`.
# Advanced Usage
The `create` function is the critical part of this library in which parameter `model_spec` defines the specification of the model, currently only `AverageWordVecModelSpec` is supported. The `create` function contains the following steps:
1. Split the data into training, validation, testing data according to parameter `validation_ratio` and `test_ratio`. The default value of `validation_ratio` and `test_ratio` are `0.1` and `0.1`.
2. Tokenize the text and select the top `num_words` frequency of words to generate the vocubulary. The default value of `num_words` in `AverageWordVecModelSpec` object is `10000`.
3. Encode the text string tokens to int ids.
4. Create the text classifier model. Currently, this library supports one model: average the word embedding of the text with RELU activation, then leverage softmax dense layer for classification. As for [Embedding layer](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Embedding), the input dimension is the size of the vocabulary, the output dimension is `AverageWordVecModelSpec` object's variable `wordvec_dim` which default value is `16`, the input length is `AverageWordVecModelSpec` object's variable `sentence_len` which default value is `256`.
5. Train the classifier model. The default epoch is `2` and the default batch size is `32`.
In this section, we describe several advanced topics, including adjusting the model, changing the training hyperparameters etc.
# Adjust the model
We could adjust the model infrastructure like variables `wordvec_dim`, `sentence_len` in `AverageWordVecModelSpec` class.
* `wordvec_dim`: Dimension of word embedding.
* `sentence_len`: length of sentence.
For example, we could train with larger `wordvec_dim`.
```
model = text_classifier.create(train_data, validation_data=validation_data, model_spec=AverageWordVecModelSpec(wordvec_dim=32))
```
## Change the training hyperparameters
We could also change the training hyperparameters like `epochs` and `batch_size` that could affect the model accuracy. For instance,
* `epochs`: more epochs could achieve better accuracy until converage but training for too many epochs may lead to overfitting.
* `batch_size`: number of samples to use in one training step.
For example, we could train with more epochs.
```
model = text_classifier.create(train_data, validation_data=validation_data, epochs=5)
```
Evaluate the newly retrained model with 5 training epochs.
```
loss, accuracy = model.evaluate(test_data)
```
| github_jupyter |
```
import numpy as np
import torch
import sklearn
import sklearn.datasets
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
def load_data():
N = 500
gq = sklearn.datasets.make_gaussian_quantiles(mean=None, cov=0.7,
n_samples=N, n_features=2,
n_classes=2, shuffle=True,
random_state=None)
return gq
gaussian_quantiles = load_data()
X, y = gaussian_quantiles
print(X[0])
print(y)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=42)
plt.figure(figsize=(9,6))
plt.scatter(X_train[:, 0], X_train[:, 1], c=y_train, marker='.', s=40, cmap=plt.cm.Spectral, label='training-samples')
plt.scatter(X_test[:, 0], X_test[:, 1], c=y_test, marker='X', s=40, cmap=plt.cm.Spectral, label='validation-samples')
plt.legend()
plt.show()
class NeuralNetwork:
def __init__(self, n_in, n_hidden, n_out):
self.n_x = n_in
self.n_h = n_hidden
self.n_y = n_out
self.W1 = torch.rand(self.n_h, self.n_x) * 0.01
self.b1 = torch.zeros(self.n_h, 1)
self.W2 = torch.rand(self.n_y, self.n_h) * 0.01
self.b2 = torch.zeros(self.n_y, 1)
def my_forward(self, torch_X):
self.Z1 = self.W1.matmul(torch_X.t()) + self.b1
self.A1 = torch.tanh(self.Z1)
self.Z2 = self.W2.matmul(self.A1) + self.b2
self.A2 = torch.sigmoid(self.Z2)
def my_backward(self, torch_X, torch_y):
m = torch_X.shape[0]
self.dZ2 = self.A2 - torch_y
self.dW2 = (1. / m) * torch.matmul(self.dZ2, self.A1.t())
self.db2 = (1. / m) * torch.sum(self.dZ2, dim=1, keepdim=True)
self.dZ1 = torch.mul(torch.matmul(self.W2.t(), self.dZ2), 1 - torch.pow(self.A1, 2))
self.dW1 = (1. / m) * torch.matmul(self.dZ1, torch_X)
self.db1 = (1. / m) * torch.sum(self.dZ1, dim=1, keepdim=True)
def train(self, training_X, training_y, validation_X, validation_y, epochs, learning_rate):
m = training_X.shape[0]
for e in range(epochs):
self.my_forward(training_X)
training_loss = -torch.sum(torch.mul(torch.log(self.A2), training_y) + torch.mul(torch.log(1-self.A2), (1 - training_y))) / m
self.my_backward(training_X, training_y)
self.W1 -= learning_rate * self.dW1
self.b1 -= learning_rate * self.db1
self.W2 -= learning_rate * self.dW2
self.b2 -= learning_rate * self.db2
if e % 250 == 0:
m = validation_X.shape[0]
self.my_forward(validation_X)
validation_loss = -torch.sum(torch.mul(torch.log(self.A2), validation_y) + torch.mul(torch.log(1-self.A2), (1 - validation_y))) / m
print("Training loss: {}".format(training_loss))
print("Validation loss: {}".format(validation_loss))
training_X = torch.from_numpy(X_train).float()
training_y = torch.from_numpy(y_train).float()
validation_X = torch.from_numpy(X_test).float()
validation_y = torch.from_numpy(y_test).float()
nn = NeuralNetwork(2, 10, 1)
nn.train(training_X, training_y, validation_X, validation_y, 2500, 0.5)
```
| github_jupyter |
```
import read_data
import pandas as pd
import numpy as np
from IPython import embed
from sklearn.decomposition import PCA
import matplotlib.pyplot as plt
import seaborn
import logging
import six
import scipy.stats
from sklearn.preprocessing import Imputer
df=pd.read_csv('dyl_ecoli_df.csv',usecols=['index', 'Week', 'Yearday', 'Monthday', 'Weekday',
'Month', 'Timestamp', 'Beach', 'Ecoli', 'Year'],index_col=[0],parse_dates=[6])
forecast_df=pd.read_csv('dyl_forecast_df.csv',usecols=['beach', 'time', 'precipIntensity', 'precipProbability',
'temperature', 'apparentTemperature', 'dewPoint', 'humidity',
'windSpeed', 'visibility', 'cloudCover', 'pressure', 'Breezy', 'Dry',
'Foggy', 'Humid', 'MostlyCloudy', 'PartlyCloudy', 'Overcast', 'Clear',
'Drizzle', 'DangerouslyWindy', 'Windy', 'HeavyRain', 'LightRain',
'Rain', 'windSin', 'windCos'],parse_dates=[1])
forecast_df['windCos']=forecast_df['windCos']*forecast_df['windSpeed']
forecast_df['windSin']=forecast_df['windSin']*forecast_df['windSpeed']
forecast_df=forecast_df.drop(['windSpeed'],axis=1)
df=df[df.Timestamp.dt.hour==0]
dfs=[]
for y in range(2006,2015):
dfs.append(pd.DataFrame(index=pd.DatetimeIndex(start=df.Timestamp[df.Year==y].min()-pd.Timedelta(days=10),freq='H',end=df.Timestamp[df.Year==y].max())));
timeindexed_df=pd.concat(dfs)
for beach in df.Beach.unique():
b=beach.replace(' ','').strip()
timeindexed_df['ecoli_'+b]=np.nan
timeindexed_df.loc[df.Timestamp[df.Beach==beach],'ecoli_'+b]=df.Ecoli[df.Beach==beach].values
for beach in df.Beach.unique():
b=beach.replace(' ','').strip()
sub_df=forecast_df[forecast_df.beach==beach]
for c in (set(forecast_df.columns)-set(['time','beach'])):
timeindexed_df[c+'_'+b]=np.nan
timeindexed_df.loc[sub_df.time,c+'_'+b]=sub_df[c].values
predictor_pcas=pd.DataFrame(index=timeindexed_df.index)
for c in (set(forecast_df.columns)-set(['time','beach'])):
forecast_pivot=forecast_df.pivot(index='time',columns='beach',values=c)
forecast_pivot.drop('39th',axis=1,inplace=True)
forecast_pivot=forecast_pivot[forecast_pivot.notnull().all(axis=1)]
forecast_pivot=forecast_pivot.loc[forecast_pivot.notnull().all(axis=1),:]
pca=PCA(n_components=6)
predictor_pcas=predictor_pcas.merge(pd.DataFrame(pca.fit_transform(forecast_pivot),index=forecast_pivot.index,columns=[c+'0',c+'1',c+'2',c+'3',c+'4',c+'5']),left_index=True,right_index=True,how='outer')
c='Ecoli'
ecoli_pivot=df.pivot(index='Timestamp',columns='Beach',values=c)
ecoli_pivot.drop('39th',axis=1,inplace=True)
ecoli_pivot=ecoli_pivot[ecoli_pivot.notnull().all(axis=1)]
ecoli_pivot=ecoli_pivot.loc[ecoli_pivot.notnull().all(axis=1),:]
pca=PCA(n_components=6)
predictor_pcas=predictor_pcas.merge(pd.DataFrame(pca.fit_transform(ecoli_pivot),index=ecoli_pivot.index,columns=[c+'0',c+'1',c+'2',c+'3',c+'4',c+'5']),left_index=True,right_index=True,how='outer')
pca_hit_means={}
pca_miss_means={}
pca_hit_stds={}
pca_miss_stds={}
pca_hit_counts={}
pca_miss_counts={}
# pca_hitmiss_kw={}
pca_hitmiss_mwu={}
pca_hitmiss_ranksum={}
all_columns=predictor_pcas.columns
empty_dt_df=pd.DataFrame(columns=all_columns,
index=pd.timedelta_range(start='8H', end='-24H',freq='-1H').append(pd.timedelta_range(start='-2 days', end='-10 days',freq='-1D')))
for beach in df.Beach.unique():
b=beach.replace(' ','').strip()
print(b)
sub_df=df[df.Beach==beach]
pca_hit_means[b]=empty_dt_df.copy()
pca_miss_means[b]=empty_dt_df.copy()
pca_hit_stds[b]=empty_dt_df.copy()
pca_miss_stds[b]=empty_dt_df.copy()
pca_hit_counts[b]=empty_dt_df.copy()
pca_miss_counts[b]=empty_dt_df.copy()
# pca_hitmiss_kw[b]=empty_dt_df.copy()
pca_hitmiss_mwu[b]=empty_dt_df.copy()
pca_hitmiss_ranksum[b]=empty_dt_df.copy()
hit_times=sub_df.Timestamp[sub_df.Ecoli>=235]
miss_times=sub_df.Timestamp[sub_df.Ecoli<235]
for dt in pd.timedelta_range(start='8H', end='-24H',freq='-1H').append(pd.timedelta_range(start='-2 days', end='-10 days',freq='-1D')):
shift_hit=hit_times+dt
shift_miss=miss_times+dt
pca_hit_means[b].loc[dt,:]=predictor_pcas.loc[shift_hit].mean();
pca_miss_means[b].loc[dt,:]=predictor_pcas.loc[shift_miss].mean();
pca_hit_stds[b].loc[dt,:]=predictor_pcas.loc[shift_hit].std();
pca_miss_stds[b].loc[dt,:]=predictor_pcas.loc[shift_miss].std();
pca_hit_counts[b].loc[dt,:]=predictor_pcas.loc[shift_hit].notnull().sum();
pca_miss_counts[b].loc[dt,:]=predictor_pcas.loc[shift_miss].notnull().sum();
for f in all_columns:
if ~np.isnan(predictor_pcas.loc[shift_hit,f].sum()):
try:
pca_hitmiss_mwu[b].loc[dt,f]=scipy.stats.mstats.mannwhitneyu(predictor_pcas.loc[shift_hit,f],predictor_pcas.loc[shift_miss,f]).pvalue+scipy.stats.mstats.mannwhitneyu(predictor_pcas.loc[shift_miss,f],predictor_pcas.loc[shift_hit,f]).pvalue;
pca_hitmiss_ranksum[b].loc[dt,f]=scipy.stats.ranksums(predictor_pcas.loc[shift_miss,f],predictor_pcas.loc[shift_hit,f]).pvalue;
# pca_hitmiss_kw[b].loc[dt,f]=scipy.stats.mstats.kruskalwallis(predictor_pcas.loc[shift_hit,f],predictor_pcas.loc[shift_miss,f]).pvalue;
except:
continue
predictors_df=df.copy();
for c in (set(forecast_df.columns)-set(['time','beach'])):
predictors_df[c+'0']=0
predictors_df[c+'1']=0
predictors_df[c+'A']=0
predictors_df['Ecoli0']=0
predictors_df['Ecoli1']=0
predictors_df['EcoliA']=0
predictors_df.reset_index(inplace=True)
for beach in predictors_df.Beach.unique():
beach_hits=predictors_df.Beach==beach
beach_index=predictors_df.index[beach_hits]
beach_times=predictors_df.loc[beach_hits,'Timestamp']
b=beach.replace(' ','').strip()
print(b)
for dt in pd.timedelta_range(start='8H', end='-24H',freq='-1H').append(pd.timedelta_range(start='-2 days', end='-10 days',freq='-1D')):
shift_times=beach_times+dt
for c in (set(forecast_df.columns)-set(['time','beach'])):
score=(predictor_pcas.loc[beach_times+dt,c+'0']-pca_miss_means[b].loc[dt,c+'0'])*(pca_hit_means[b].loc[dt,c+'0']-pca_miss_means[b].loc[dt,c+'0'])*(1-pca_hitmiss_ranksum[b].loc[dt,c+'0'])**100;
predictors_df.loc[beach_index[score.notnull()],c+'0']+=score[score.notnull()].values;
score=(predictor_pcas.loc[beach_times+dt,c+'1']-pca_miss_means[b].loc[dt,c+'1'])*(pca_hit_means[b].loc[dt,c+'1']-pca_miss_means[b].loc[dt,c+'1'])*(1-pca_hitmiss_ranksum[b].loc[dt,c+'1'])**100;
predictors_df.loc[beach_index[score.notnull()],c+'1']+=score[score.notnull()].values;
score=((predictor_pcas.loc[beach_times+dt,[c+'2',c+'3',c+'4',c+'5']]-pca_miss_means[b].loc[dt,[c+'2',c+'3',c+'4',c+'5']])*(pca_hit_means[b].loc[dt,[c+'2',c+'3',c+'4',c+'5']]-pca_miss_means[b].loc[dt,[c+'2',c+'3',c+'4',c+'5']])*(1-pca_hitmiss_ranksum[b].loc[dt,[c+'2',c+'3',c+'4',c+'5']])**100).sum(axis=1);
predictors_df.loc[beach_index[score.notnull()],c+'A']+=score[score.notnull()].values;
c='Ecoli'
if dt.days<0:
score=(predictor_pcas.loc[beach_times+dt,c+'0']-pca_miss_means[b].loc[dt,c+'0'])*(pca_hit_means[b].loc[dt,c+'0']-pca_miss_means[b].loc[dt,c+'0'])*(1-pca_hitmiss_ranksum[b].loc[dt,c+'0'])**100;
predictors_df.loc[beach_index[score.notnull()],c+'0']+=score[score.notnull()].values;
score=(predictor_pcas.loc[beach_times+dt,c+'1']-pca_miss_means[b].loc[dt,c+'1'])*(pca_hit_means[b].loc[dt,c+'1']-pca_miss_means[b].loc[dt,c+'1'])*(1-pca_hitmiss_ranksum[b].loc[dt,c+'1'])**100;
predictors_df.loc[beach_index[score.notnull()],c+'1']+=score[score.notnull()].values;
score=((predictor_pcas.loc[beach_times+dt,[c+'2',c+'3',c+'4',c+'5']]-pca_miss_means[b].loc[dt,[c+'2',c+'3',c+'4',c+'5']])*(pca_hit_means[b].loc[dt,[c+'2',c+'3',c+'4',c+'5']]-pca_miss_means[b].loc[dt,[c+'2',c+'3',c+'4',c+'5']])*(1-pca_hitmiss_ranksum[b].loc[dt,[c+'2',c+'3',c+'4',c+'5']])**100).sum(axis=1);
predictors_df.loc[beach_index[score.notnull()],c+'A']+=score[score.notnull()].values;
from sklearn.preprocessing import Imputer
import sklearn.ensemble as ens
import sklearn.metrics
%matplotlib inline
Fresh_run=predictors_df.copy()
# usingParams=['Year','Ecoli_geomean','precipIntensity','precipProbability','temperature','apparentTemperature','dewPoint','cloudCover','pressure','windSin','windCos','precipIntensity_pca','precipProbability_pca','temperature_pca','apparentTemperature_pca','dewPoint_pca','cloudCover_pca','pressure_pca','windCos_pca','dtemperature_pca','dapparentTemperature_pca','dvisibility_pca','dwindCos_pca']
# Fresh_run=ecoli_df.loc[:,usingParams].copy()
Fresh_run=Fresh_run.drop(['index','Timestamp','Beach'],1)
years=Fresh_run.Year.unique()
columns=Fresh_run.columns.drop(['Ecoli'])
speration=(predictors_df[predictors_df.Ecoli>=235].mean()-predictors_df[predictors_df.Ecoli<235].mean())/predictors_df[predictors_df.Ecoli<235].std()
predictor_columns=speration[speration>0.15].index
predictor_columns=predictor_columns.drop('Ecoli')
# Fresh_run.loc[Fresh_run[np.isinf(Fresh_run.precipIntensity)].index,'precipIntensity']=100;
# Fresh_run.loc[Fresh_run[np.isinf(Fresh_run.precipProbability)].index,'precipProbability']=100;
# imp = Imputer(missing_values='NaN', strategy='mean', axis=1)
cleaned_data = Fresh_run[columns]
E_levels=Fresh_run.Ecoli.as_matrix()
plt.figure(figsize=[12,12])
RF=list()
count=0;
predictions=list()
E_test=list()
legend=list()
for y in years:
print(y)
train_ind=(cleaned_data.Year != y).as_matrix()
test_ind=(cleaned_data.Year == y).as_matrix()
RF.append(ens.RandomForestClassifier(n_estimators=500,criterion='entropy',class_weight={True:.8,False:.2}))
RF[count]=RF[count].fit(cleaned_data.loc[train_ind,predictor_columns],E_levels[train_ind]>=235)
predictions.append(RF[count].predict_proba(cleaned_data.loc[test_ind,predictor_columns]))
E_test.append(E_levels[test_ind])
fpr, tpr, _ =sklearn.metrics.roc_curve(E_test[count]>=235,predictions[count][:,1])
plt.plot(fpr,tpr)
plt.hold(True)
count+=1
legend.append(y)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend(legend,loc=0)
plt.axis('equal')
coverage=list()
for count in range(0,9):
temp=predictions[count][:,1].copy()
temp=E_test[count][temp.argsort()]>235
temp=temp[::-1]
temp2=np.cumsum(temp)/np.arange(1,temp.size+1)
temp3=np.argwhere(temp2>0.45).max()
coverage.append(temp2[temp3]*(temp3+1)/temp.sum())
coverage
predictor_columns
speration[predictor_columns]
Fresh_run.to_csv('Ecoli_filtered_pcas.csv')
```
| github_jupyter |
```
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
pd.options.mode.chained_assignment = None
relevant_cols = ['Which round did you get your coop in?', 'Which sector was your first co-op in?']
software = class_df[relevant_cols].dropna()
software.head(100)
all = software
```
Software
```
software = software[software['Which round did you get your coop in?'].notna()]
software.head()
software['Which sector was your first co-op in?'] = software['Which sector was your first co-op in?'].str.split(';')
software = (software
.set_index(['Which round did you get your coop in?'])['Which sector was your first co-op in?']
.apply(pd.Series)
.stack()
.reset_index()
.drop('level_1', axis=1)
.rename(columns={0:'Which sector was your first co-op in?'}))
software.head()
software[software['Which sector was your first co-op in?'] != 'Unhired']
software = software[software['Which sector was your first co-op in?'] != 'Unhired']
software = software[software['Which round did you get your coop in?'] != 'Still Looking']
software.head()
software = software[software['Which sector was yourfirst co-op in?'] == 'Software']
software.head()
software['Number of people'] = software.groupby(['Which round did you get your coop in?'])['Which sector was your first co-op in?'].transform('count')
software = software.drop_duplicates(subset=['Which round did you get your coop in?', 'Which sector was your first co-op in?', 'Number of people'], keep='first')
software
software['Percentage of People'] = (software['Number of people'] / 98) * 100
software
```
Other
```
other = class_df[relevant_cols]
other.head()
other = other[other['Which round did you get your coop in?'].notna()]
other.head()
other['Which sector was your first co-op in?'] = other['Which sector was your first co-op in?'].str.split(';')
other = (other
.set_index(['Which round did you get your coop in?'])['Which sector was your first co-op in?']
.apply(pd.Series)
.stack()
.reset_index()
.drop('level_1', axis=1)
.rename(columns={0:'Which sector was your first co-op in?'}))
other.head()
other[other['Which sector was your first co-op in?'] != 'Unhired']
other = other[other['Which sector was your first co-op in?'] != 'Unhired']
other.head()
other = other[other['Which sector was your first co-op in?'] == 'Other']
other.head()
other['Number of people'] = other.groupby(['Which round did you get your coop in?'])['Which sector was your first co-op in?'].transform('count')
other = other.drop_duplicates(subset=['Which round did you get your coop in?', 'Which sector was your first co-op in?', 'Number of people'], keep='first')
other
other['Percentage of People'] = (other['Number of people'] / 98) * 100
other
```
hardware
```
hardware = class_df[relevant_cols]
hardware.head()
hardware = hardware[hardware['Which round did you get your coop in?'].notna()]
hardware.head()
hardware['Which sector was your first co-op in?'] = hardware['Which sector was your first co-op in?'].str.split(';')
hardware = (hardware
.set_index(['Which round did you get your coop in?'])['Which sector was your first co-op in?']
.apply(pd.Series)
.stack()
.reset_index()
.drop('level_1', axis=1)
.rename(columns={0:'Which sector was your first co-op in?'}))
hardware.head()
hardware[hardware['Which sector was your first co-op in?'] != 'Unhired']
hardware = hardware[hardware['Which sector was your first co-op in?'] != 'Unhired']
hardware.head()
hardware = hardware[hardware['Which sector was your first co-op in?'] == 'Hardware']
hardware.head()
hardware['Number of people'] = hardware.groupby(['Which round did you get your coop in?'])['Which sector was your first co-op in?'].transform('count')
hardware = hardware.drop_duplicates(subset=['Which round did you get your coop in?', 'Which sector was your first co-op in?', 'Number of people'], keep='first')
hardware
hardware['Percentage of People'] = (hardware['Number of people'] / 98) * 100
hardware
```
Product Management
```
prodmag = class_df[relevant_cols]
prodmag.head()
prodmag = prodmag[prodmag['Which round did you get your coop in?'].notna()]
prodmag.head()
prodmag['Which sector was your first co-op in?'] = prodmag['Which sector was your first co-op in?'].str.split(';')
prodmag = (prodmag
.set_index(['Which round did you get your coop in?'])['Which sector was your first co-op in?']
.apply(pd.Series)
.stack()
.reset_index()
.drop('level_1', axis=1)
.rename(columns={0:'Which sector was your first co-op in?'}))
prodmag.head()
prodmag = prodmag[prodmag['Which sector was your first co-op in?'] != 'Unhired']
prodmag.head()
prodmag = prodmag[prodmag['Which sector was your first co-op in?'] == 'Product Management']
prodmag.head()
prodmag['Number of people'] = prodmag.groupby(['Which round did you get your coop in?'])['Which sector was your first co-op in?'].transform('count')
prodmag = prodmag.drop_duplicates(subset=['Which round did you get your coop in?', 'Which sector was your first co-op in?', 'Number of people'], keep='first')
prodmag
prodmag['Percentage of People'] = (prodmag['Number of people'] / 98) * 100
prodmag
```
UI/UX
```
uiUx = class_df[relevant_cols]
uiUx.head()
uiUx = uiUx[uiUx['Which round did you get your coop in?'].notna()]
uiUx.head()
uiUx['Which sector was your first co-op in?'] = uiUx['Which sector was your first co-op in?'].str.split(';')
uiUx = (uiUx
.set_index(['Which round did you get your coop in?'])['Which sector was your first co-op in?']
.apply(pd.Series)
.stack()
.reset_index()
.drop('level_1', axis=1)
.rename(columns={0:'Which sector was your first co-op in?'}))
uiUx.head()
uiUx = uiUx[uiUx['Which sector was your first co-op in?'] != 'Unhired']
uiUx.head()
uiUx = uiUx[uiUx['Which sector was your first co-op in?'] == 'UI/UX']
uiUx.head()
uiUx['Number of people'] = uiUx.groupby(['Which round did you get your coop in?'])['Which sector was your first co-op in?'].transform('count')
uiUx = uiUx.drop_duplicates(subset=['Which round did you get your coop in?', 'Which sector was your first co-op in?', 'Number of people'], keep='first')
uiUx
uiUx['Percentage of People'] = (uiUx['Number of people'] / 98) * 100
uiUx
```
Product Design
```
prodDes = class_df[relevant_cols]
prodDes.head()
prodDes = prodDes[prodDes['Which round did you get your coop in?'].notna()]
prodDes.head()
prodDes['Which sector was your first co-op in?'] = prodDes['Which sector was your first co-op in?'].str.split(';')
prodDes = (prodDes
.set_index(['Which round did you get your coop in?'])['Which sector was your first co-op in?']
.apply(pd.Series)
.stack()
.reset_index()
.drop('level_1', axis=1)
.rename(columns={0:'Which sector was your first co-op in?'}))
prodDes.head()
prodDes = prodDes[prodDes['Which sector was your first co-op in?'] != 'Unhired']
prodDes.head()
prodDes = prodDes[prodDes['Which sector was your first co-op in?'] == 'Product Design']
prodDes.head()
prodDes['Number of people'] = prodDes.groupby(['Which round did you get your coop in?'])['Which sector was your first co-op in?'].transform('count')
prodDes = prodDes.drop_duplicates(subset=['Which round did you get your coop in?', 'Which sector was your first co-op in?', 'Number of people'], keep='first')
prodDes
prodDes['Percentage of People'] = (prodDes['Number of people'] / 98) * 100
prodDes
all_rounds = pd.DataFrame(columns=["Sector", "First Round", "Second Round", "Continuous Round", "Direct Offer"],
data=[["Software", 10.204082, 12.244898, 25.510204, 7.142857],
["Hardware", 0, 1.020408, 1.020408, 0],
["Product Management", 0, 2.040816, 2.040816, 1.020408],
["UI/UX", 3.061224, 5.102041, 4.081633, 3.061224],
["Product Design", 0, 0, 3.061224, 0],
["Other", 4.081633, 3.061224, 10.204082, 3.061224]])
sns.set(rc={'figure.figsize':(15, 8)})
ax = all_rounds.set_index('Sector').T.plot(kind='bar', stacked=True)
ax.set(ylim=(0, 50))
plt.title("CO-OP Round VS Sector")
plt.xlabel("Round", labelpad=15)
plt.ylabel("Percentage of people (%)", labelpad=15)
plt.xticks(rotation=0)
```
| github_jupyter |
# Springboard Logistic Regression Advanced Case Study
$$
\renewcommand{\like}{{\cal L}}
\renewcommand{\loglike}{{\ell}}
\renewcommand{\err}{{\cal E}}
\renewcommand{\dat}{{\cal D}}
\renewcommand{\hyp}{{\cal H}}
\renewcommand{\Ex}[2]{E_{#1}[#2]}
\renewcommand{\x}{{\mathbf x}}
\renewcommand{\v}[1]{{\mathbf #1}}
$$
This case study delves into the math behind logistic regression in a Python environment. We've adapted this case study from [Lab 5 in the CS109](https://github.com/cs109/2015lab5) course. Please feel free to check out the original lab, both for more exercises, as well as solutions.
We turn our attention to **classification**. Classification tries to predict, which of a small set of classes, an observation belongs to. Mathematically, the aim is to find $y$, a **label** based on knowing a feature vector $\x$. For instance, consider predicting gender from seeing a person's face, something we do fairly well as humans. To have a machine do this well, we would typically feed the machine a bunch of images of people which have been labelled "male" or "female" (the training set), and have it learn the gender of the person in the image from the labels and the *features* used to determine gender. Then, given a new photo, the trained algorithm returns us the gender of the person in the photo.
There are different ways of making classifications. One idea is shown schematically in the image below, where we find a line that divides "things" of two different types in a 2-dimensional feature space. The classification show in the figure below is an example of a maximum-margin classifier where construct a decision boundary that is far as possible away from both classes of points. The fact that a line can be drawn to separate the two classes makes the problem *linearly separable*. Support Vector Machines (SVM) are an example of a maximum-margin classifier.
<img src="images/onelinesplit.png" width="400" height="200">
```
%matplotlib inline
import numpy as np
import scipy as sp
import matplotlib as mpl
import matplotlib.cm as cm
from matplotlib.colors import ListedColormap
import matplotlib.pyplot as plt
import pandas as pd
pd.set_option('display.width', 500)
pd.set_option('display.max_columns', 100)
pd.set_option('display.notebook_repr_html', True)
import seaborn as sns
sns.set_style("whitegrid")
sns.set_context("poster")
import sklearn.model_selection
import warnings # For handling error messages.
# Don't worry about the following two instructions: they just suppress warnings that could occur later.
warnings.simplefilter(action="ignore", category=FutureWarning)
warnings.filterwarnings(action="ignore", module="scipy", message="^internal gelsd")
c0=sns.color_palette()[0]
c1=sns.color_palette()[1]
c2=sns.color_palette()[2]
cmap_light = ListedColormap(['#FFAAAA', '#AAFFAA', '#AAAAFF'])
cmap_bold = ListedColormap(['#FF0000', '#00FF00', '#0000FF'])
cm = plt.cm.RdBu
cm_bright = ListedColormap(['#FF0000', '#0000FF'])
def points_plot(ax, Xtr, Xte, ytr, yte, clf, mesh=True, colorscale=cmap_light,
cdiscrete=cmap_bold, alpha=0.1, psize=10, zfunc=False, predicted=False):
h = .02
X=np.concatenate((Xtr, Xte))
x_min, x_max = X[:, 0].min() - .5, X[:, 0].max() + .5
y_min, y_max = X[:, 1].min() - .5, X[:, 1].max() + .5
xx, yy = np.meshgrid(np.linspace(x_min, x_max, 100),
np.linspace(y_min, y_max, 100))
#plt.figure(figsize=(10,6))
if zfunc:
p0 = clf.predict_proba(np.c_[xx.ravel(), yy.ravel()])[:, 0]
p1 = clf.predict_proba(np.c_[xx.ravel(), yy.ravel()])[:, 1]
Z=zfunc(p0, p1)
else:
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
ZZ = Z.reshape(xx.shape)
if mesh:
plt.pcolormesh(xx, yy, ZZ, cmap=cmap_light, alpha=alpha, axes=ax)
if predicted:
showtr = clf.predict(Xtr)
showte = clf.predict(Xte)
else:
showtr = ytr
showte = yte
ax.scatter(Xtr[:, 0], Xtr[:, 1], c=showtr-1, cmap=cmap_bold,
s=psize, alpha=alpha,edgecolor="k")
# and testing points
ax.scatter(Xte[:, 0], Xte[:, 1], c=showte-1, cmap=cmap_bold,
alpha=alpha, marker="s", s=psize+10)
ax.set_xlim(xx.min(), xx.max())
ax.set_ylim(yy.min(), yy.max())
return ax,xx,yy
def points_plot_prob(ax, Xtr, Xte, ytr, yte, clf, colorscale=cmap_light,
cdiscrete=cmap_bold, ccolor=cm, psize=10, alpha=0.1):
ax,xx,yy = points_plot(ax, Xtr, Xte, ytr, yte, clf, mesh=False,
colorscale=colorscale, cdiscrete=cdiscrete,
psize=psize, alpha=alpha, predicted=True)
Z = clf.predict_proba(np.c_[xx.ravel(), yy.ravel()])[:, 1]
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=ccolor, alpha=.2, axes=ax)
cs2 = plt.contour(xx, yy, Z, cmap=ccolor, alpha=.6, axes=ax)
#plt.clabel(cs2, fmt = '%2.1f', colors = 'k', fontsize=14, axes=ax)
return ax
```
## A Motivating Example Using `sklearn`: Heights and Weights
We'll use a dataset of heights and weights of males and females to hone our understanding of classifiers. We load the data into a dataframe and plot it.
```
dflog = pd.read_csv("data/01_heights_weights_genders.csv")
dflog.head()
```
Remember that the form of data we will use always is
<img src="images/dataform.jpg" width="400" height="200">
with the "response" or "label" $y$ as a plain array of 0s and 1s for binary classification. Sometimes we will also see -1 and +1 instead. There are also *multiclass* classifiers that can assign an observation to one of $K > 2$ classes and the labe may then be an integer, but we will not be discussing those here.
`y = [1,1,0,0,0,1,0,1,0....]`.
<div class="span5 alert alert-info">
<h3>Checkup Exercise Set I</h3>
<ul>
<li> <b>Exercise:</b> Create a scatter plot of Weight vs. Height
<li> <b>Exercise:</b> Color the points differently by Gender
</ul>
</div>
```
_ = sns.scatterplot(x='Weight', y='Height', hue='Gender', data=dflog, linestyle = 'None', color = 'blue', alpha=0.25)
```
### Training and Test Datasets
When fitting models, we would like to ensure two things:
* We have found the best model (in terms of model parameters).
* The model is highly likely to generalize i.e. perform well on unseen data.
<br/>
<div class="span5 alert alert-success">
<h4>Purpose of splitting data into Training/testing sets</h4>
<ul>
<li> We built our model with the requirement that the model fit the data well. </li>
<li> As a side-effect, the model will fit <b>THIS</b> dataset well. What about new data? </li>
<ul>
<li> We wanted the model for predictions, right?</li>
</ul>
<li> One simple solution, leave out some data (for <b>testing</b>) and <b>train</b> the model on the rest </li>
<li> This also leads directly to the idea of cross-validation, next section. </li>
</ul>
</div>
First, we try a basic Logistic Regression:
* Split the data into a training and test (hold-out) set
* Train on the training set, and test for accuracy on the testing set
```
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
# Split the data into a training and test set.
Xlr, Xtestlr, ylr, ytestlr = train_test_split(dflog[['Height','Weight']].values,
(dflog.Gender == "Male").values,random_state=5)
clf = LogisticRegression()
# Fit the model on the trainng data.
clf.fit(Xlr, ylr)
# Print the accuracy from the testing data.
print(accuracy_score(clf.predict(Xtestlr), ytestlr))
```
### Tuning the Model
The model has some hyperparameters we can tune for hopefully better performance. For tuning the parameters of your model, you will use a mix of *cross-validation* and *grid search*. In Logistic Regression, the most important parameter to tune is the *regularization parameter* `C`. Note that the regularization parameter is not always part of the logistic regression model.
The regularization parameter is used to control for unlikely high regression coefficients, and in other cases can be used when data is sparse, as a method of feature selection.
You will now implement some code to perform model tuning and selecting the regularization parameter $C$.
We use the following `cv_score` function to perform K-fold cross-validation and apply a scoring function to each test fold. In this incarnation we use accuracy score as the default scoring function.
```
from sklearn.model_selection import KFold
from sklearn.metrics import accuracy_score
def cv_score(clf, x, y, score_func=accuracy_score):
result = 0
nfold = 5
for train, test in KFold(nfold).split(x): # split data into train/test groups, 5 times
clf.fit(x[train], y[train]) # fit
result += score_func(clf.predict(x[test]), y[test]) # evaluate score function on held-out data
return result / nfold # average
```
Below is an example of using the `cv_score` function for a basic logistic regression model without regularization.
```
clf = LogisticRegression()
score = cv_score(clf, Xlr, ylr)
print(round(score,4))
```
<div class="span5 alert alert-info">
<h3>Checkup Exercise Set II</h3>
<b>Exercise:</b> Implement the following search procedure to find a good model
<ul>
<li> You are given a list of possible values of `C` below
<li> For each C:
<ol>
<li> Create a logistic regression model with that value of C
<li> Find the average score for this model using the `cv_score` function **only on the training set** `(Xlr, ylr)`
</ol>
<li> Pick the C with the highest average score
</ul>
Your goal is to find the best model parameters based *only* on the training set, without showing the model test set at all (which is why the test set is also called a *hold-out* set).
</div>
```
#the grid of parameters to search over
Cs = [0.001, 0.1, 1, 10, 100]
highest_score = 0
best_c = 0
for C in Cs:
clf = LogisticRegression(C=C)
score = cv_score(clf, Xlr, ylr)
if score > highest_score:
highest_score = score
best_c = C
print("Best score is {}".format(round(highest_score,4)))
```
<div class="span5 alert alert-info">
<h3>Checkup Exercise Set III</h3>
**Exercise:** Now you want to estimate how this model will predict on unseen data in the following way:
<ol>
<li> Use the C you obtained from the procedure earlier and train a Logistic Regression on the training data
<li> Calculate the accuracy on the test data
</ol>
<p>You may notice that this particular value of `C` may or may not do as well as simply running the default model on a random train-test split. </p>
<ul>
<li> Do you think that's a problem?
<li> Why do we need to do this whole cross-validation and grid search stuff anyway?
</ul>
</div>
```
clf = LogisticRegression(C=C)
clf.fit(Xlr, ylr)
ypredlr = clf.predict(Xtestlr)
print(accuracy_score(ypredlr, ytestlr))
```
### Black Box Grid Search in `sklearn`
Scikit-learn, as with many other Python packages, provides utilities to perform common operations so you do not have to do it manually. It is important to understand the mechanics of each operation, but at a certain point, you will want to use the utility instead to save time...
<div class="span5 alert alert-info">
<h3>Checkup Exercise Set IV</h3>
<b>Exercise:</b> Use scikit-learn's [GridSearchCV](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html) tool to perform cross validation and grid search.
* Instead of writing your own loops above to iterate over the model parameters, can you use GridSearchCV to find the best model over the training set?
* Does it give you the same best value of `C`?
* How does this model you've obtained perform on the test set?</div>
```
from sklearn.model_selection import GridSearchCV
parameters = {
'C': [0.001, 0.1, 1, 10, 100]
}
grid = GridSearchCV(clf, parameters)
grid.fit(Xlr, ylr)
ypredlr = grid.predict(Xtestlr)
print("Accuracy score is {}".format(accuracy_score(ypredlr, ytestlr)))
print("Best tuned parameters are {}".format(grid.best_params_))
print("Best score is {}".format(grid.best_score_))
print("Best estimator is {}".format(grid.best_estimator_))
```
## A Walkthrough of the Math Behind Logistic Regression
### Setting up Some Demo Code
Let's first set some code up for classification that we will need for further discussion on the math. We first set up a function `cv_optimize` which takes a classifier `clf`, a grid of hyperparameters (such as a complexity parameter or regularization parameter) implemented as a dictionary `parameters`, a training set (as a samples x features array) `Xtrain`, and a set of labels `ytrain`. The code takes the traning set, splits it into `n_folds` parts, sets up `n_folds` folds, and carries out a cross-validation by splitting the training set into a training and validation section for each foldfor us. It prints the best value of the parameters, and retuens the best classifier to us.
```
def cv_optimize(clf, parameters, Xtrain, ytrain, n_folds=5):
gs = sklearn.model_selection.GridSearchCV(clf, param_grid=parameters, cv=n_folds)
gs.fit(Xtrain, ytrain)
print("BEST PARAMS", gs.best_params_)
best = gs.best_estimator_
return best
```
We then use this best classifier to fit the entire training set. This is done inside the `do_classify` function which takes a dataframe `indf` as input. It takes the columns in the list `featurenames` as the features used to train the classifier. The column `targetname` sets the target. The classification is done by setting those samples for which `targetname` has value `target1val` to the value 1, and all others to 0. We split the dataframe into 80% training and 20% testing by default, standardizing the dataset if desired. (Standardizing a data set involves scaling the data so that it has 0 mean and is described in units of its standard deviation. We then train the model on the training set using cross-validation. Having obtained the best classifier using `cv_optimize`, we retrain on the entire training set and calculate the training and testing accuracy, which we print. We return the split data and the trained classifier.
```
from sklearn.model_selection import train_test_split
def do_classify(clf, parameters, indf, featurenames, targetname, target1val, standardize=False, train_size=0.8):
subdf=indf[featurenames]
if standardize:
subdfstd=(subdf - subdf.mean())/subdf.std()
else:
subdfstd=subdf
X=subdfstd.values
y=(indf[targetname].values==target1val)*1
Xtrain, Xtest, ytrain, ytest = train_test_split(X, y, train_size=train_size)
clf = cv_optimize(clf, parameters, Xtrain, ytrain)
clf=clf.fit(Xtrain, ytrain)
training_accuracy = clf.score(Xtrain, ytrain)
test_accuracy = clf.score(Xtest, ytest)
print("Accuracy on training data: {:0.2f}".format(training_accuracy))
print("Accuracy on test data: {:0.2f}".format(test_accuracy))
return clf, Xtrain, ytrain, Xtest, ytest
```
## Logistic Regression: The Math
We could approach classification as linear regression, there the class, 0 or 1, is the target variable $y$. But this ignores the fact that our output $y$ is discrete valued, and futhermore, the $y$ predicted by linear regression will in general take on values less than 0 and greater than 1. Additionally, the residuals from the linear regression model will *not* be normally distributed. This violation means we should not use linear regression.
But what if we could change the form of our hypotheses $h(x)$ instead?
The idea behind logistic regression is very simple. We want to draw a line in feature space that divides the '1' samples from the '0' samples, just like in the diagram above. In other words, we wish to find the "regression" line which divides the samples. Now, a line has the form $w_1 x_1 + w_2 x_2 + w_0 = 0$ in 2-dimensions. On one side of this line we have
$$w_1 x_1 + w_2 x_2 + w_0 \ge 0,$$
and on the other side we have
$$w_1 x_1 + w_2 x_2 + w_0 < 0.$$
Our classification rule then becomes:
\begin{eqnarray*}
y = 1 &\mbox{if}& \v{w}\cdot\v{x} \ge 0\\
y = 0 &\mbox{if}& \v{w}\cdot\v{x} < 0
\end{eqnarray*}
where $\v{x}$ is the vector $\{1,x_1, x_2,...,x_n\}$ where we have also generalized to more than 2 features.
What hypotheses $h$ can we use to achieve this? One way to do so is to use the **sigmoid** function:
$$h(z) = \frac{1}{1 + e^{-z}}.$$
Notice that at $z=0$ this function has the value 0.5. If $z > 0$, $h > 0.5$ and as $z \to \infty$, $h \to 1$. If $z < 0$, $h < 0.5$ and as $z \to -\infty$, $h \to 0$. As long as we identify any value of $y > 0.5$ as 1, and any $y < 0.5$ as 0, we can achieve what we wished above.
This function is plotted below:
```
h = lambda z: 1. / (1 + np.exp(-z))
zs=np.arange(-5, 5, 0.1)
plt.plot(zs, h(zs), alpha=0.5);
```
So we then come up with our rule by identifying:
$$z = \v{w}\cdot\v{x}.$$
Then $h(\v{w}\cdot\v{x}) \ge 0.5$ if $\v{w}\cdot\v{x} \ge 0$ and $h(\v{w}\cdot\v{x}) \lt 0.5$ if $\v{w}\cdot\v{x} \lt 0$, and:
\begin{eqnarray*}
y = 1 &if& h(\v{w}\cdot\v{x}) \ge 0.5\\
y = 0 &if& h(\v{w}\cdot\v{x}) \lt 0.5.
\end{eqnarray*}
We will show soon that this identification can be achieved by minimizing a loss in the ERM framework called the **log loss** :
$$ R_{\cal{D}}(\v{w}) = - \sum_{y_i \in \cal{D}} \left ( y_i \log(h(\v{w}\cdot\v{x})) + ( 1 - y_i) \log(1 - h(\v{w}\cdot\v{x})) \right )$$
We will also add a regularization term:
$$ R_{\cal{D}}(\v{w}) = - \sum_{y_i \in \cal{D}} \left ( y_i \log(h(\v{w}\cdot\v{x})) + ( 1 - y_i) \log(1 - h(\v{w}\cdot\v{x})) \right ) + \frac{1}{C} \v{w}\cdot\v{w},$$
where $C$ is the regularization strength (equivalent to $1/\alpha$ from the Ridge case), and smaller values of $C$ mean stronger regularization. As before, the regularization tries to prevent features from having terribly high weights, thus implementing a form of feature selection.
How did we come up with this loss? We'll come back to that, but let us see how logistic regression works out.
```
dflog.head()
clf_l, Xtrain_l, ytrain_l, Xtest_l, ytest_l = do_classify(LogisticRegression(),
{"C": [0.01, 0.1, 1, 10, 100]},
dflog, ['Weight', 'Height'], 'Gender','Male')
plt.figure()
ax=plt.gca()
points_plot(ax, Xtrain_l, Xtest_l, ytrain_l, ytest_l, clf_l, alpha=0.2);
```
In the figure here showing the results of the logistic regression, we plot the actual labels of both the training(circles) and test(squares) samples. The 0's (females) are plotted in red, the 1's (males) in blue. We also show the classification boundary, a line (to the resolution of a grid square). Every sample on the red background side of the line will be classified female, and every sample on the blue side, male. Notice that most of the samples are classified well, but there are misclassified people on both sides, as evidenced by leakage of dots or squares of one color ontothe side of the other color. Both test and traing accuracy are about 92%.
### The Probabilistic Interpretaion
Remember we said earlier that if $h > 0.5$ we ought to identify the sample with $y=1$? One way of thinking about this is to identify $h(\v{w}\cdot\v{x})$ with the probability that the sample is a '1' ($y=1$). Then we have the intuitive notion that lets identify a sample as 1 if we find that the probabilty of being a '1' is $\ge 0.5$.
So suppose we say then that the probability of $y=1$ for a given $\v{x}$ is given by $h(\v{w}\cdot\v{x})$?
Then, the conditional probabilities of $y=1$ or $y=0$ given a particular sample's features $\v{x}$ are:
\begin{eqnarray*}
P(y=1 | \v{x}) &=& h(\v{w}\cdot\v{x}) \\
P(y=0 | \v{x}) &=& 1 - h(\v{w}\cdot\v{x}).
\end{eqnarray*}
These two can be written together as
$$P(y|\v{x}, \v{w}) = h(\v{w}\cdot\v{x})^y \left(1 - h(\v{w}\cdot\v{x}) \right)^{(1-y)} $$
Then multiplying over the samples we get the probability of the training $y$ given $\v{w}$ and the $\v{x}$:
$$P(y|\v{x},\v{w}) = P(\{y_i\} | \{\v{x}_i\}, \v{w}) = \prod_{y_i \in \cal{D}} P(y_i|\v{x_i}, \v{w}) = \prod_{y_i \in \cal{D}} h(\v{w}\cdot\v{x_i})^{y_i} \left(1 - h(\v{w}\cdot\v{x_i}) \right)^{(1-y_i)}$$
Why use probabilities? Earlier, we talked about how the regression function $f(x)$ never gives us the $y$ exactly, because of noise. This hold for classification too. Even with identical features, a different sample may be classified differently.
We said that another way to think about a noisy $y$ is to imagine that our data $\dat$ was generated from a joint probability distribution $P(x,y)$. Thus we need to model $y$ at a given $x$, written as $P(y|x)$, and since $P(x)$ is also a probability distribution, we have:
$$P(x,y) = P(y | x) P(x)$$
and can obtain our joint probability $P(x, y)$.
Indeed its important to realize that a particular training set can be thought of as a draw from some "true" probability distribution (just as we did when showing the hairy variance diagram). If for example the probability of classifying a test sample as a '0' was 0.1, and it turns out that the test sample was a '0', it does not mean that this model was necessarily wrong. After all, in roughly a 10th of the draws, this new sample would be classified as a '0'! But, of-course its more unlikely than its likely, and having good probabilities means that we'll be likely right most of the time, which is what we want to achieve in classification. And furthermore, we can quantify this accuracy.
Thus its desirable to have probabilistic, or at the very least, ranked models of classification where you can tell which sample is more likely to be classified as a '1'. There are business reasons for this too. Consider the example of customer "churn": you are a cell-phone company and want to know, based on some of my purchasing habit and characteristic "features" if I am a likely defector. If so, you'll offer me an incentive not to defect. In this scenario, you might want to know which customers are most likely to defect, or even more precisely, which are most likely to respond to incentives. Based on these probabilities, you could then spend a finite marketing budget wisely.
### Maximizing the Probability of the Training Set
Now if we maximize $P(y|\v{x},\v{w})$, we will maximize the chance that each point is classified correctly, which is what we want to do. While this is not exactly the same thing as maximizing the 1-0 training risk, it is a principled way of obtaining the highest probability classification. This process is called **maximum likelihood** estimation since we are maximising the **likelihood of the training data y**,
$$\like = P(y|\v{x},\v{w}).$$
Maximum likelihood is one of the corenerstone methods in statistics, and is used to estimate probabilities of data.
We can equivalently maximize
$$\loglike = \log{P(y|\v{x},\v{w})}$$
since the natural logarithm $\log$ is a monotonic function. This is known as maximizing the **log-likelihood**. Thus we can equivalently *minimize* a risk that is the negative of $\log(P(y|\v{x},\v{w}))$:
$$R_{\cal{D}}(h(x)) = -\loglike = -\log \like = -\log{P(y|\v{x},\v{w})}.$$
Thus
\begin{eqnarray*}
R_{\cal{D}}(h(x)) &=& -\log\left(\prod_{y_i \in \cal{D}} h(\v{w}\cdot\v{x_i})^{y_i} \left(1 - h(\v{w}\cdot\v{x_i}) \right)^{(1-y_i)}\right)\\
&=& -\sum_{y_i \in \cal{D}} \log\left(h(\v{w}\cdot\v{x_i})^{y_i} \left(1 - h(\v{w}\cdot\v{x_i}) \right)^{(1-y_i)}\right)\\
&=& -\sum_{y_i \in \cal{D}} \log\,h(\v{w}\cdot\v{x_i})^{y_i} + \log\,\left(1 - h(\v{w}\cdot\v{x_i}) \right)^{(1-y_i)}\\
&=& - \sum_{y_i \in \cal{D}} \left ( y_i \log(h(\v{w}\cdot\v{x})) + ( 1 - y_i) \log(1 - h(\v{w}\cdot\v{x})) \right )
\end{eqnarray*}
This is exactly the risk we had above, leaving out the regularization term (which we shall return to later) and was the reason we chose it over the 1-0 risk.
Notice that this little process we carried out above tells us something very interesting: **Probabilistic estimation using maximum likelihood is equivalent to Empiricial Risk Minimization using the negative log-likelihood**, since all we did was to minimize the negative log-likelihood over the training samples.
`sklearn` will return the probabilities for our samples, or for that matter, for any input vector set $\{\v{x}_i\}$, i.e. $P(y_i | \v{x}_i, \v{w})$:
```
clf_l.predict_proba(Xtest_l)
```
### Discriminative vs Generative Classifier
Logistic regression is what is known as a **discriminative classifier** as we learn a soft boundary between/among classes. Another paradigm is the **generative classifier** where we learn the distribution of each class. For more examples of generative classifiers, look [here](https://en.wikipedia.org/wiki/Generative_model).
Let us plot the probabilities obtained from `predict_proba`, overlayed on the samples with their true labels:
```
plt.figure()
ax = plt.gca()
points_plot_prob(ax, Xtrain_l, Xtest_l, ytrain_l, ytest_l, clf_l, psize=20, alpha=0.1);
```
Notice that lines of equal probability, as might be expected are stright lines. What the classifier does is very intuitive: if the probability is greater than 0.5, it classifies the sample as type '1' (male), otherwise it classifies the sample to be class '0'. Thus in the diagram above, where we have plotted predicted values rather than actual labels of samples, there is a clear demarcation at the 0.5 probability line.
Again, this notion of trying to obtain the line or boundary of demarcation is what is called a **discriminative** classifier. The algorithm tries to find a decision boundary that separates the males from the females. To classify a new sample as male or female, it checks on which side of the decision boundary the sample falls, and makes a prediction. In other words we are asking, given $\v{x}$, what is the probability of a given $y$, or, what is the likelihood $P(y|\v{x},\v{w})$?
| github_jupyter |
# Load Packages
```
import pandas as pd
import spacy
from spacy.tokens import DocBin
from tqdm.notebook import tqdm
import warnings
warnings.filterwarnings("ignore")
```
# Processing
Data source: https://www.kaggle.com/abhinavwalia95/entity-annotated-corpus
```
df = pd.read_csv('data/ner_dataset.csv')
df['Sentence #'] = df['Sentence #'].ffill()
df.head()
def get_spacy_file(df, sentences, train_test="train"):
nlp = spacy.blank("en") # load a new spacy model
db = DocBin() # create a DocBin obje
for i in tqdm(sentences):
df2 = df[df['Sentence #']==i]
word_length = df2['Word'].apply(lambda x: len(x)+1)
df2['End'] = word_length.cumsum()-1
df2['Start'] = df2['End']-word_length+1
# get content
content = ' '.join([df['Word'][j] for j in df2.index])
doc = nlp.make_doc(content)
# get entities
ents = []
df2 = df2[df2['Tag'] != 'O']
for index, row in df2.iterrows():
span = doc.char_span(row['Start'], row['End'], label=row['Tag'], alignment_mode="contract")
if span:
ents.append(span)
doc.ents = ents
db.add(doc)
db.to_disk(f"./{train_test}.spacy")
train_pct = 0.8
dev_pct = 0.1
sentences = df['Sentence #'].unique()
n = len(sentences)
n_train = int(train_pct*n)
n_dev = int(dev_pct*n)
train_sentences = df['Sentence #'].unique()[:n_train]
dev_sentences = df['Sentence #'].unique()[n_train:n_train+n_dev]
get_spacy_file(df, train_sentences, 'train')
get_spacy_file(df, dev_sentences, 'dev')
```
# Fill Config
```
!python3 -m spacy init fill-config base_config.cfg config.cfg
```
# Train Model
```
!python3 -m spacy train config.cfg --output ./output --paths.train ./train.spacy --paths.dev ./dev.spacy
```
# Prediction
```
nlp = spacy.load(R"./output/model-best")
type1_error, type2_error, n_words = 0, 0, 0
test_sentences = df['Sentence #'].unique()[n_train+n_dev:]
for i in tqdm(test_sentences):
df2 = df[df['Sentence #']==i]
sentence = ' '.join([df2['Word'][j] for j in df2.index])
df2 = df2[df2['Tag'] != 'O']
n_words += len(df2)
# truth
words = df2['Word'].values
tags = df2['Tag'].values
truths = []
for j in range(len(words)):
truths.append((words[j], tags[j]))
# predict
doc = nlp(sentence)
predictions = [(ent.text, ent.label_) for ent in doc.ents]
# type 1 error
for prediction in predictions:
if prediction not in truths:
type1_error += 1
# type 2 error
for truth in truths:
if truth not in predictions:
type2_error += 1
type1_error = type1_error/n_words
type2_error = type2_error/n_words
print('Type 1 error: {}'.format(type1_error))
print('Type 2 error: {}'.format(type2_error))
# visualize
df2 = df[df['Sentence #']==test_sentences[0]]
df2
sentence = ' '.join([df2['Word'][j] for j in df2.index])
doc = nlp(sentence)
spacy.displacy.render(doc, style="ent", jupyter=True)
```
| github_jupyter |
```
import os
import random
from IPython.display import display, HTML, Image
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import colors
%matplotlib inline
import seaborn as sns
import moranpycess
mario_blue = "#6485fb"
mario_red = "#d20709"
mario_purple = "#9b4682"
mario_brown = "#974a02"
mario_gold = "#f7bc31"
mario_dark_green = "#148817"
mario_light_green = "#6df930"
```
AUTHOR: Maciej_Bak
AFFILIATION: University_of_Basel
AFFILIATION: Swiss_Institute_of_Bioinformatics
CONTACT: wsciekly.maciek@gmail.com
CREATED: 12-08-2020
LICENSE: MIT
# <center>Moran Pycess: usecase tests</center>
## Moran Process
Population evolve according to the Moran Process with fitness-based selection: https://en.wikipedia.org/wiki/Moran_process
### Test 1: Stag Hunt
https://en.wikipedia.org/wiki/Stag_hunt
Birth Payoff Matrix:
$
B = \begin{pmatrix}
4 & 1\\
3 & 2
\end{pmatrix}
$
Death Payoff Matrix:
$
D = \begin{pmatrix}
1 & 1\\
1 & 1
\end{pmatrix}
$ (random selection)
```
# initiate Moran Process
size_list = [700, 300]
label_list = ["stag", "hare"]
BirthPayoffMatrix = np.array([[4, 1], [3, 2]])
DeathPayoffMatrix = np.array([[1, 1], [1, 1]])
mp = moranpycess.MoranProcess(
size_list=size_list,
label_list=label_list,
BirthPayoffMatrix=BirthPayoffMatrix,
DeathPayoffMatrix=DeathPayoffMatrix,
)
# simulate evolution
random.seed(0)
simulation1 = mp.simulate(generations=10000)
# plot the results
plt.figure(figsize=(14,6))
ax = plt.gca()
ax.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(1)
simulation1['stag__size'].plot(color=mario_red, linewidth=1.5, ax=ax, label="Stag")
simulation1['hare__size'].plot(color=mario_blue, linewidth=1.5, ax=ax, label="Hare")
ax.set_ylim([0,1000])
plt.xlabel('Generation', size=14)
plt.ylabel('# Individuals', size=14)
ax.tick_params(axis='both', which='major', labelsize=12)
ax.legend(loc=7, fontsize=20)
plt.show()
# plot the entropy
plt.figure(figsize=(14,6))
ax = plt.gca()
ax.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(1)
simulation1['Entropy'].plot(color="black", linewidth=1.5, ax=ax)
ax.set_ylim([0,1])
plt.xlabel('Generation', size=14)
plt.ylabel('')
ax.tick_params(axis='both', which='major', labelsize=12)
ax.legend(loc=4, fontsize=20)
plt.show()
```
### Test 2: Chicken
https://en.wikipedia.org/wiki/Chicken_(game)
Birth Payoff Matrix:
$
B = \begin{pmatrix}
3 & 2\\
4 & 0
\end{pmatrix}
$
Death Payoff Matrix:
$
D = \begin{pmatrix}
1 & 1\\
1 & 1
\end{pmatrix}
$ (random selection)
```
# initiate Moran Process
size_list = [10, 990]
label_list = ["swerve", "straight"]
BirthPayoffMatrix = np.array([[3, 2], [4, 0]])
DeathPayoffMatrix = np.array([[1, 1], [1, 1]])
mp = moranpycess.MoranProcess(
size_list=size_list,
label_list=label_list,
BirthPayoffMatrix=BirthPayoffMatrix,
DeathPayoffMatrix=DeathPayoffMatrix,
)
# simulate evolution
random.seed(0)
simulation2 = mp.simulate(generations=30000)
# plot the results
plt.figure(figsize=(14,6))
ax = plt.gca()
ax.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(1)
simulation2['swerve__size'].plot(color=mario_blue, linewidth=1.5, ax=ax, label="Swerve")
simulation2['straight__size'].plot(color=mario_red, linewidth=1.5, ax=ax, label="Straight")
ax.set_ylim([0,1000])
plt.xlabel('Generation', size=14)
plt.ylabel('# Individuals', size=14)
ax.tick_params(axis='both', which='major', labelsize=12)
ax.legend(loc=7, fontsize=20)
plt.show()
# plot the entropy
plt.figure(figsize=(14,6))
ax = plt.gca()
ax.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(1)
simulation2['Entropy'].plot(color="black", linewidth=1.5, ax=ax)
ax.set_ylim([0,1.1])
plt.xlabel('Generation', size=14)
plt.ylabel('')
ax.tick_params(axis='both', which='major', labelsize=12)
ax.legend(loc=4, fontsize=20)
plt.show()
```
### Test 3: Prisoners Dilemma
https://en.wikipedia.org/wiki/Prisoner%27s_dilemma
Birth Payoff Matrix:
$
B = \begin{pmatrix}
3 & 0\\
5 & 1
\end{pmatrix}
$
Death Payoff Matrix:
$
D = \begin{pmatrix}
1 & 1\\
1 & 1
\end{pmatrix}
$ (random selection)
```
# initiate Moran Process
size_list = [990, 10]
label_list = ["cooperate", "defect"]
BirthPayoffMatrix = np.array([[3, 0], [5, 1]])
DeathPayoffMatrix = np.array([[1, 1], [1, 1]])
mp = moranpycess.MoranProcess(
size_list=size_list,
label_list=label_list,
BirthPayoffMatrix=BirthPayoffMatrix,
DeathPayoffMatrix=DeathPayoffMatrix,
)
# simulate evolution
random.seed(0)
simulation3 = mp.simulate(generations=25000)
# plot the results
plt.figure(figsize=(14,6))
ax = plt.gca()
ax.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(1)
simulation3['cooperate__size'].plot(color=mario_blue, linewidth=1.5, ax=ax, label="Cooperate")
simulation3['defect__size'].plot(color=mario_red, linewidth=1.5, ax=ax, label="Defect")
ax.set_ylim([0,1000])
plt.xlabel('Generation', size=14)
plt.ylabel('# Individuals', size=14)
ax.tick_params(axis='both', which='major', labelsize=12)
ax.legend(loc=7, fontsize=20)
plt.show()
# plot the entropy
plt.figure(figsize=(14,6))
ax = plt.gca()
ax.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(1)
simulation3['Entropy'].plot(color="black", linewidth=1.5, ax=ax)
ax.set_ylim([0,1.1])
plt.xlabel('Generation', size=14)
plt.ylabel('')
ax.tick_params(axis='both', which='major', labelsize=12)
ax.legend(loc=4, fontsize=20)
plt.show()
```
### Test 4: Rock-Paper-Scissors
https://en.wikipedia.org/wiki/Rock_paper_scissors
Birth Payoff Matrix:
$
B = \begin{pmatrix}
10 & 0 & 20\\
20 & 10 & 0\\
0 & 20 & 10
\end{pmatrix}
$
Death Payoff Matrix:
$
D = \begin{pmatrix}
1 & 1 & 1\\
1 & 1 & 1\\
1 & 1 & 1
\end{pmatrix}
$ (random selection)
```
# initiate Moran Process
size_list = [333, 333, 333]
label_list = ["rock", "paper", "scissors"]
BirthPayoffMatrix = np.array([[10, 0, 20], [20, 10, 0], [0, 20, 10]])
DeathPayoffMatrix = np.array([[1, 1, 1], [1, 1, 1], [1, 1, 1]])
mp = moranpycess.MoranProcess(
size_list=size_list,
label_list=label_list,
BirthPayoffMatrix=BirthPayoffMatrix,
DeathPayoffMatrix=DeathPayoffMatrix,
)
# simulate evolution
random.seed(0)
simulation4 = mp.simulate(generations=30000)
# plot the results
plt.figure(figsize=(14,6))
ax = plt.gca()
ax.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(1)
simulation4['rock__size'].plot(color=mario_red, linewidth=1.5, ax=ax, label="Rock")
simulation4['paper__size'].plot(color=mario_blue, linewidth=1.5, ax=ax, label="Paper")
simulation4['scissors__size'].plot(color=mario_gold, linewidth=1.5, ax=ax, label="Scissors")
ax.set_ylim([0,700])
plt.xlabel('Generation', size=14)
plt.ylabel('# Individuals', size=14)
ax.tick_params(axis='both', which='major', labelsize=12)
ax.legend(loc=2, fontsize=20)
plt.show()
# plot the entropy
plt.figure(figsize=(14,6))
ax = plt.gca()
ax.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(1)
simulation4['Entropy'].plot(color="black", linewidth=1.5, ax=ax)
ax.set_ylim([1,2])
plt.xlabel('Generation', size=14)
plt.ylabel('')
ax.tick_params(axis='both', which='major', labelsize=12)
ax.legend(loc=4, fontsize=20)
plt.show()
```
### Publication Figure
Prepare a figure with all the simulations altogether:
```
fig, ax = plt.subplots(2, 2, figsize=(14,8))
plt.subplot(2, 2, 1)
ax1 = plt.gca()
ax1.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax1.spines[axis].set_linewidth(1)
simulation1['stag__size'].plot(color=mario_red, linewidth=1.5, ax=ax1, label="Stag")
simulation1['hare__size'].plot(color=mario_blue, linewidth=1.5, ax=ax1, label="Hare")
ax1.set_ylim([0,1000])
plt.title("A", size=18)
plt.xlabel('Generation', size=14)
plt.ylabel('# Individuals', size=14)
ax1.tick_params(axis='both', which='major', labelsize=12)
ax1.legend(loc=4, fontsize=10)
plt.subplot(2, 2, 2)
ax2 = plt.gca()
ax2.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax2.spines[axis].set_linewidth(1)
simulation2['swerve__size'].plot(color=mario_blue, linewidth=1.5, ax=ax2, label="Swerve")
simulation2['straight__size'].plot(color=mario_red, linewidth=1.5, ax=ax2, label="Straight")
ax2.set_ylim([0,1000])
plt.title("B", size=18)
plt.xlabel('Generation', size=14)
plt.ylabel('# Individuals', size=14)
ax2.tick_params(axis='both', which='major', labelsize=12)
ax2.legend(loc=4, fontsize=10)
plt.subplot(2, 2, 3)
ax3 = plt.gca()
ax3.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax3.spines[axis].set_linewidth(1)
simulation3['cooperate__size'].plot(color=mario_blue, linewidth=1.5, ax=ax3, label="Cooperate")
simulation3['defect__size'].plot(color=mario_red, linewidth=1.5, ax=ax3, label="Defect")
ax3.set_ylim([0,1000])
plt.title("C", size=18)
plt.xlabel('Generation', size=14)
plt.ylabel('# Individuals', size=14)
ax3.tick_params(axis='both', which='major', labelsize=12)
ax3.legend(loc=4, fontsize=10)
plt.subplot(2, 2, 4)
ax4 = plt.gca()
ax4.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax4.spines[axis].set_linewidth(1)
simulation4['rock__size'].plot(color=mario_red, linewidth=1.5, ax=ax4, label="Rock")
simulation4['paper__size'].plot(color=mario_blue, linewidth=1.5, ax=ax4, label="Paper")
simulation4['scissors__size'].plot(color=mario_gold, linewidth=1.5, ax=ax4, label="Scissors")
ax4.set_ylim([0,700])
plt.title("D", size=18)
plt.xlabel('Generation', size=14)
plt.ylabel('# Individuals', size=14)
ax4.tick_params(axis='both', which='major', labelsize=12)
ax4.legend(loc=4, fontsize=10)
sns.despine(ax=ax1)
sns.despine(ax=ax2)
sns.despine(ax=ax3)
sns.despine(ax=ax4)
plt.tight_layout()
plt.savefig(os.path.join("..", "images", "figure.png"), dpi=300)
plt.show()
```
## Moran Process 2D
Simulate Moran Process on a 2D population (Payoffs calculated based on the neighbourhood)
```
# settings for the MoranProcess2D (Prisoner's Dilemma)
size_list = [10000-9, 9]
label_list = ["A", "B"]
grid = np.full((100, 100), "A")
grid[40,40] = "B"
grid[40,41] = "B"
grid[40,42] = "B"
grid[41,40] = "B"
grid[41,41] = "B"
grid[41,42] = "B"
grid[42,40] = "B"
grid[42,41] = "B"
grid[42,42] = "B"
BirthPayoffMatrix = np.array([[5, 0], [15, 1]])
DeathPayoffMatrix = np.array([[1, 1], [1, 1]])
# initialize an instance of MoranProcess2D:
mp = moranpycess.MoranProcess2D(
size_list=size_list,
label_list=label_list,
grid=grid,
BirthPayoffMatrix=BirthPayoffMatrix,
DeathPayoffMatrix=DeathPayoffMatrix,
)
simulation2D_t0 = mp.curr_grid.copy()
# simulate evolution
random.seed(0)
mp.simulate(generations=50000)
simulation2D_t1 = mp.curr_grid.copy()
# initialize an instance of MoranProcess2D:
mp = moranpycess.MoranProcess2D(
size_list=size_list,
label_list=label_list,
grid=grid,
BirthPayoffMatrix=BirthPayoffMatrix,
DeathPayoffMatrix=DeathPayoffMatrix,
)
# simulate evolution
random.seed(0)
mp.simulate(generations=200000)
simulation2D_t2 = mp.curr_grid.copy()
# initialize an instance of MoranProcess2D:
mp = moranpycess.MoranProcess2D(
size_list=size_list,
label_list=label_list,
grid=grid,
BirthPayoffMatrix=BirthPayoffMatrix,
DeathPayoffMatrix=DeathPayoffMatrix,
)
# simulate evolution
random.seed(0)
mp.simulate(generations=500000)
simulation2D_t3 = mp.curr_grid.copy()
```
### Supplementary Figure 1a: 2D Prisoners Dilemma population snapshots
```
fig, ax = plt.subplots(2, 2, figsize=(8,8))
cmap = colors.ListedColormap([mario_red, mario_blue])
plt.subplot(2, 2, 1)
ax1 = plt.gca()
ax1.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax1.spines[axis].set_linewidth(1)
plt.imshow((simulation2D_t0 == "A").astype(float), cmap=cmap)
plt.title("A", size=18)
plt.ylabel('')
plt.yticks([])
plt.xlabel('')
plt.xticks([])
plt.subplot(2, 2, 2)
ax2 = plt.gca()
ax2.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax2.spines[axis].set_linewidth(1)
plt.imshow((simulation2D_t1 == "A").astype(float), cmap=cmap)
plt.title("B", size=18)
plt.ylabel('')
plt.yticks([])
plt.xlabel('')
plt.xticks([])
plt.subplot(2, 2, 3)
ax3 = plt.gca()
ax3.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax3.spines[axis].set_linewidth(1)
plt.imshow((simulation2D_t2 == "A").astype(float), cmap=cmap)
plt.title("C", size=18)
plt.ylabel('')
plt.yticks([])
plt.xlabel('')
plt.xticks([])
plt.subplot(2, 2, 4)
ax4 = plt.gca()
ax4.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax4.spines[axis].set_linewidth(1)
plt.imshow((simulation2D_t3 == "A").astype(float), cmap=cmap)
plt.title("D", size=18)
plt.ylabel('')
plt.yticks([])
plt.xlabel('')
plt.xticks([])
plt.tight_layout()
plt.savefig(os.path.join("..", "images", "supplementary_figure1a.png"), dpi=300)
plt.show()
```
### Supplementary Figure 1b: 2D Prisoners Dilemma growth curve
```
# initialize an instance of MoranProcess2D:
mp = moranpycess.MoranProcess2D(
size_list=size_list,
label_list=label_list,
grid=grid,
BirthPayoffMatrix=BirthPayoffMatrix,
DeathPayoffMatrix=DeathPayoffMatrix,
)
# simulate evolution
random.seed(0)
simulation2D = mp.simulate(generations=600000)
# plot the results
plt.figure(figsize=(14,8))
ax = plt.gca()
ax.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(1)
simulation2D['A__size'].plot(color=mario_blue, linewidth=1.5, ax=ax, label="Cooperate")
simulation2D['B__size'].plot(color=mario_red, linewidth=1.5, ax=ax, label="Defect")
ax.set_ylim([0,10100])
plt.xlabel('Generation', size=14)
plt.ylabel('# Individuals', size=14)
ax.tick_params(axis='both', which='major', labelsize=12)
ax.legend(loc=7, fontsize=20)
sns.despine(ax=ax)
plt.tight_layout()
plt.savefig(os.path.join("..", "images", "supplementary_figure1b.png"), dpi=300)
plt.show()
```
## Moran Process 3D
Simulate Moran Process on a 3D population (Payoffs calculated based on the neighbourhood)
```
# settings for the MoranProcess3D (Prisoner's Dilemma)
size_list = [8000-8, 8]
label_list = ["A", "B"]
grid = np.full((20, 20, 20), "A")
grid[9,9, 9] = "B"
grid[9,9, 10] = "B"
grid[9,10, 9] = "B"
grid[9,10, 10] = "B"
grid[10,9, 9] = "B"
grid[10,9, 10] = "B"
grid[10,10, 9] = "B"
grid[10,10, 10] = "B"
BirthPayoffMatrix = np.array([[5, 0], [15, 1]])
DeathPayoffMatrix = np.array([[1, 1], [1, 1]])
# initialize an instance of MoranProcess2D:
mp = moranpycess.MoranProcess3D(
size_list=size_list,
label_list=label_list,
grid=grid,
BirthPayoffMatrix=BirthPayoffMatrix,
DeathPayoffMatrix=DeathPayoffMatrix,
)
# simulate evolution
random.seed(0)
simulation3D = mp.simulate(generations=150000)
```
### Supplementary Figure 2: 3D Prisoners Dilemma growth curve
```
# plot the results
plt.figure(figsize=(14,8))
ax = plt.gca()
ax.tick_params(width=1)
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(1)
simulation3D['A__size'].plot(color=mario_blue, linewidth=1.5, ax=ax, label="Cooperate")
simulation3D['B__size'].plot(color=mario_red, linewidth=1.5, ax=ax, label="Defect")
#ax.set_ylim([0,10100])
plt.xlabel('Generation', size=14)
plt.ylabel('# Individuals', size=14)
ax.tick_params(axis='both', which='major', labelsize=12)
ax.legend(loc=7, fontsize=20)
sns.despine(ax=ax)
plt.tight_layout()
plt.savefig(os.path.join("..", "images", "supplementary_figure2.png"), dpi=300)
plt.show()
```
---
| github_jupyter |
This notebook was prepared by [Donne Martin](http://donnemartin.com). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# Challenge Notebook
## Problem: Implement an algorithm to determine if a string has all unique characters.
* [Constraints](#Constraints)
* [Test Cases](#Test-Cases)
* [Algorithm](#Algorithm)
* [Code](#Code)
* [Unit Test](#Unit-Test)
* [Solution Notebook](#Solution-Notebook)
## Constraints
* Can we assume the string is ASCII?
* Yes
* Note: Unicode strings could require special handling depending on your language
* Can we assume this is case sensitive?
* Yes
* Can we use additional data structures?
* Yes
* Can we assume this fits in memory?
* Yes
## Test Cases
* None -> False
* '' -> True
* 'foo' -> False
* 'bar' -> True
## Algorithm
Refer to the [Solution Notebook](http://nbviewer.ipython.org/github/donnemartin/interactive-coding-challenges/blob/master/arrays_strings/unique_chars/unique_chars_solution.ipynb). If you are stuck and need a hint, the solution notebook's algorithm discussion might be a good place to start.
## Code
```
class UniqueChars(object):
def has_unique_chars(self, string):
return ((string is not None)
and (len(set(string)) == len(string)))
#if string is None:
# return False
#return not(len(set(string)) < len(string))
# My recursive UniqueChars
class UniqueCharsRec(object):
def has_unique_chars(self, string):
if string is None:
return False
if string == '':
return True
if string.count(string[0]) > 1:
return False
return self.has_unique_chars(string[1:])
```
## Unit Test
**The following unit test is expected to fail until you solve the challenge.**
```
# %load test_unique_chars.py
import cProfile
from nose.tools import assert_equal
class TestUniqueChars(object):
def test_unique_chars(self, func):
assert_equal(func(None), False)
assert_equal(func(''), True)
assert_equal(func('foo'), False)
assert_equal(func('bar'), True)
print('Success: test_unique_chars')
def main():
test = TestUniqueChars()
unique_chars = UniqueChars()
# test.test_unique_chars(unique_chars.has_unique_chars)
cProfile.runctx('test.test_unique_chars(unique_chars.has_unique_chars)',
globals=globals(),
locals=locals())
try:
unique_chars_set = UniqueCharsSet()
test.test_unique_chars(unique_chars_set.has_unique_chars)
unique_chars_in_place = UniqueCharsInPlace()
test.test_unique_chars(unique_chars_in_place.has_unique_chars)
except NameError:
# Alternate solutions are only defined
# in the solutions file
pass
try:
unique_chars_rec = UniqueCharsRec()
cProfile.runctx('test.test_unique_chars(unique_chars_rec.has_unique_chars)',globals=globals(), locals=locals())
# test.test_unique_chars(unique_chars_rec.has_unique_chars)
except NameError:
pass
if __name__ == '__main__':
main()
```
## Solution Notebook
Review the [Solution Notebook](http://nbviewer.ipython.org/github/donnemartin/interactive-coding-challenges/blob/master/arrays_strings/unique_chars/unique_chars_solution.ipynb) for a discussion on algorithms and code solutions.
| github_jupyter |
```
####################################
#
# generateAdversarials: This code will load a pre-trained model, sample validation data
# and find adversarial inputs.
#
# Method: Let the input to the forcaster be X and the target be y
# Note that both X & y are real valued
# We find X' such that X' ~ X and |y'-y| >> 0
# We will adapt the optimisation method of Carlini & Wagner
# for the generation of the adversarials
#
# Author: Anurag Dwarakanath
###################################
import tensorflow as tf
from tensorflow.python.saved_model import tag_constants
import pandas
import numpy as np
import matplotlib.pyplot as plt
# Constants
DATAFILE_VALIDATE = "mock_kaggle_edit_validate.csv"
TRAINED_MODEL_PATH = 'savedModel'
TIME_STEPS = 10 # i.e. look at the past 10 days and forecast
NUMBER_OF_DAYS_TO_FORECAST = 1 # for now we will only forecast the next day's sales
BATCH_SIZE=100
LEARNING_RATE = 0.1
#Load the validation data
rawData = pandas.read_csv(DATAFILE_VALIDATE)
validationSales=rawData['sales']
#We need to normalise the data
MIN = 0
RANGE = 542
validationSalesNormalised = [(i-MIN)/RANGE for i in validationSales]
#Create the sequences
validationSalesSequences = np.zeros(shape=(len(validationSales)-TIME_STEPS - NUMBER_OF_DAYS_TO_FORECAST + 1, TIME_STEPS, 1))
validationSalesTargets = np.zeros(shape=(len(validationSales)-TIME_STEPS - NUMBER_OF_DAYS_TO_FORECAST + 1, NUMBER_OF_DAYS_TO_FORECAST))
for i in range(len(validationSales)-TIME_STEPS - NUMBER_OF_DAYS_TO_FORECAST + 1):
validationSalesSequences[i,:,0] = validationSalesNormalised[i:i+TIME_STEPS]
validationSalesTargets[i,:] = validationSalesNormalised[i+TIME_STEPS:i+TIME_STEPS+NUMBER_OF_DAYS_TO_FORECAST]
#create the data structure to hold the perturbations
perturbedSequences = np.zeros(shape=(len(validationSales)-TIME_STEPS - NUMBER_OF_DAYS_TO_FORECAST + 1, TIME_STEPS, 1))
perturbedForecasts = np.zeros(shape=(len(validationSales)-TIME_STEPS - NUMBER_OF_DAYS_TO_FORECAST + 1, NUMBER_OF_DAYS_TO_FORECAST))
originalForecasts = np.zeros(shape=(len(validationSales)-TIME_STEPS - NUMBER_OF_DAYS_TO_FORECAST + 1, NUMBER_OF_DAYS_TO_FORECAST))
inputSequenceLosses = np.zeros(shape=(len(validationSales)-TIME_STEPS - NUMBER_OF_DAYS_TO_FORECAST + 1, NUMBER_OF_DAYS_TO_FORECAST))
forecastLosses = np.zeros(shape=(len(validationSales)-TIME_STEPS - NUMBER_OF_DAYS_TO_FORECAST + 1, NUMBER_OF_DAYS_TO_FORECAST))
#We now load the pre-trained graph
tf.reset_default_graph()
with tf.Session() as sess:
perturbVariables=tf.get_variable(name='pVar', shape=(BATCH_SIZE, TIME_STEPS, 1), dtype=tf.float32)
#perturbVariables=tf.Variable(name='pVar', initial_value=np.zeros(size=(None, TIME_STEPS, 1)), dtype=tf.float32, validate_shape=False)
perturbedSequence = tf.math.square(x=perturbVariables) # We want the perturbed Sequence to always be positive.
actualValidationInputSequence = tf.placeholder(name='aInp', shape=(None, TIME_STEPS, 1), dtype=tf.float32)
#actualValidationTarget = tf.placeholder(name='aTar', shape=(None, NUMBER_OF_DAYS_TO_FORECAST), dtype=tf.float32)
sess.run(tf.variables_initializer([perturbVariables]))
print('Loading the model from:', TRAINED_MODEL_PATH)
tf.saved_model.loader.load(sess=sess, export_dir=TRAINED_MODEL_PATH, tags=[tag_constants.SERVING], input_map={'inputSequencePlaceholder:0':perturbedSequence})
#inputSequence = tf.get_default_graph().get_tensor_by_name('inputSequencePlaceholder:0')
forecast_normalisedScale = tf.get_default_graph().get_tensor_by_name('forecast_normalised_scale:0')
forecast_originalScale = tf.get_default_graph().get_tensor_by_name('forecast_original_scale:0')
targetForecast = tf.get_default_graph().get_tensor_by_name('targetPlaceholder:0')
minLoss = 1000
minLossSequenceID=-1
#loop through all the validation sequences
start=0
end=0
numIterations = np.int(np.ceil(len(validationSalesTargets)/BATCH_SIZE))
print(numIterations)
currentSequence = np.zeros(shape=(BATCH_SIZE, TIME_STEPS, 1))
for i in range(numIterations):
print('Starting Batch:', i)
start=i*BATCH_SIZE
if (start+BATCH_SIZE < len(validationSalesTargets)):
end=start+BATCH_SIZE
else:
end=len(validationSalesTargets)
#get the forecast for the current inputSeqeunce
currentSequence[0:end-start] = validationSalesSequences[start:end]
#initialise perturbVariables with the actual values received.
assignValue = tf.assign(ref=perturbVariables, value=currentSequence)
sess.run(assignValue)
#Get the forecasts for the actual values
tarFor = sess.run(forecast_originalScale)
print(forecast_originalScale)
print('current 2 Forecast for sequence number ', i, 'is: ', tarFor[0:2])
#random initialisation of perturbed Variables
#assignValue = tf.assign(ref=perturbVariables, value=currentSequence)# + np.random.normal(size=(1, TIME_STEPS, 1)))
#sess.run(assignValue)
#inputSequenceLoss = tf.nn.l2_loss(t=(perturbedSequence - currentSequence))
inputSequenceLoss = tf.math.reduce_sum(input_tensor=tf.math.square(perturbedSequence - currentSequence), axis=1)
#inputSequenceLoss = tf.math.reduce_max(input_tensor=tf.math.abs(perturbVar - currentSequence))
#forecastLoss = 1/(tf.losses.mean_squared_error(labels=forecast_originalScale, predictions=tarFor) + 0.00001)
#forecastLoss = -tf.math.log(tf.math.abs(forecast_originalScale - tarFor)/tf.math.abs(forecast_originalScale))
forecastLoss = tf.math.square(forecast_originalScale - 2*tarFor)
totalLoss = 1000* inputSequenceLoss + 1 * forecastLoss
optimizer = tf.train.AdamOptimizer(learning_rate=LEARNING_RATE)
trainStep = optimizer.minimize(totalLoss, var_list=[perturbVariables])
all_variables = tf.all_variables()
is_not_initialized = sess.run([tf.is_variable_initialized(var) for var in all_variables])
not_initialized_vars = [v for (v, f) in zip(all_variables, is_not_initialized) if not f]
sess.run(tf.variables_initializer(not_initialized_vars))
for optStep in range(200):
_, perturbedInput, forecastForPerturbedInput, inpSeqLoss, forLoss = sess.run([trainStep, perturbedSequence, forecast_originalScale, inputSequenceLoss, forecastLoss])
#save the values
perturbedSequences[start:end] = (perturbedInput[0:end-start] * RANGE) + MIN
perturbedForecasts[start:end] = forecastForPerturbedInput[0:end-start]
inputSequenceLosses[start:end] = inpSeqLoss[0:end-start]
forecastLosses[start:end] = forLoss[0:end-start]
originalForecasts[start:end] = tarFor[0:end-start]
if np.min(inpSeqLoss) < minLoss :
minLoss = np.min(inpSeqLoss)
minLossSequenceID = start+np.argmin(inpSeqLoss)
start+=BATCH_SIZE
print('Mininum sequence loss:', inputSequenceLosses[minLossSequenceID])
print('Perturbed sequence:',perturbedSequences[minLossSequenceID])
print('Original Sequence:', (validationSalesSequences[minLossSequenceID] * RANGE ) + MIN )
print('Original Forecast:', originalForecasts[minLossSequenceID])
print('Perturbed Forecast:', perturbedForecasts[minLossSequenceID])
print('Actual target:',(validationSalesTargets[minLossSequenceID]*RANGE) + MIN )
print('--------')
#Generating the maximum amount of perturbation needed to double the forecast.
# m will hold the maximum change needed in any time dimension for each of the input sequence
m = np.max(np.abs(perturbedSequences - (validationSalesSequences * RANGE ) + MIN), axis=1 )
# We now plot the histogram
plt.hist(m, bins=[1, 2, 3, 4, 5, 10, 20, 30], rwidth=0.9)
# That's all folks.
```
| github_jupyter |
```
import os
import numpy as np
import matplotlib.pyplot as plt
from keras.models import load_model
from keras.preprocessing.image import ImageDataGenerator
from panotti.datautils import build_dataset
```
# Class Distribution
```
import os
import glob
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
def get_class_freq(root_path, y='', title='', color=sns.xkcd_rgb["denim blue"], plot=True):
'''
Returns a dataframe of class frequency distribution when structured in the Keras ImageDataGenerator manner for classification
'''
walker = os.walk(root_path)
next(walker)
class_freq = dict()
for r, d, f in walker:
class_freq[r.split('/')[-1]] = len(f)
class_freq_df = pd.DataFrame.from_dict(
class_freq, orient='index', columns=['count'])
class_freq_df.reset_index(inplace=True)
class_freq_df.columns = [y, 'count']
class_freq_df.sort_values('count', axis=0, ascending=False, inplace=True)
if plot:
sns.catplot(x="count", y=y, kind="bar",
data=class_freq_df, color=color)
plt.title(title)
plt.show()
return class_freq_df
else:
return class_freq_df
get_class_freq('Preproc/Train/', y='Classes', title= 'Class Distribution for Training Data')
get_class_freq('Preproc/Test/', y='Classes', title= 'Class Distribution for Testing Data', color=sns.xkcd_rgb["dusty purple"])
```
# Confusion Matrix
```
weights_path = 'weights.hdf5'
model = load_model(weights_path)
melgram_path = glob.glob('Preproc/Test/*/*.npz')
test_mel = melgram_path[0]
with np.load(test_mel) as data:
melgram = data['melgram']
melgram.shape
X_test, Y_test, paths, class_names = build_dataset('Preproc/Test/')
X_test.shape
pred = model.predict(X_test)
from sklearn.metrics import confusion_matrix
confusion_1 = confusion_matrix(np.argmax(Y_test, axis = -1), np.argmax(pred, axis = -1))
NUM_LABELS = len(class_names)
f, axes = plt.subplots(1,1, figsize = (12,12))
axes.set_xlabel('Predicted')
axes.set_ylabel('Actual')
axes.grid(False)
axes.set_xticklabels(class_names, rotation = 90)
axes.set_yticklabels(class_names)
axes.set_yticks(list(range(NUM_LABELS)))
axes.set_xticks(list(range(NUM_LABELS)))
plt.imshow(confusion_1, cmap=plt.cm.Set2, interpolation='nearest')
for i, cas in enumerate(confusion_1):
for j, count in enumerate(cas):
if count > 0:
xoff = .07 * len(str(count))
plt.text(j-xoff, i+.2, int(count), fontsize=12, color='black')
print(round((1-(21/1120))*100, 2))
print(round((1-(13/1120))*100, 2))
```
| github_jupyter |
# TensorFlow Lattice estimators
In this tutorial, we will cover basics of TensorFlow Lattice estimators.
```
# import libraries
!pip install tensorflow_lattice
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow_lattice as tfl
import tempfile
from six.moves import urllib
```
# Synthetic dataset
Here we create a synthetic dataset.
```
%matplotlib inline
# Training dataset contains one feature, "distance".
train_features = {
'distance': np.array([1.0, 1.3, 1.5, 2.0, 2.1, 3.0,
4.0, 5.0, 1.3, 1.7, 2.5, 2.8,
4.7, 4.2, 3.5, 4.75, 5.2,
5.8, 5.9]) * 0.1,
}
train_labels = np.array([4.8, 4.9, 5.0, 5.0,
4.8, 3.3, 2.5, 2.0,
4.7, 4.6, 4.0, 3.2,
2.12, 2.1, 2.5, 2.2,
2.3, 2.34, 2.6])
plt.scatter(train_features['distance'], train_labels)
plt.xlabel('distance')
plt.ylabel('user hapiness')
# This function draws two plots.
# Firstly, we draw the scatter plot of `distance` vs. `label`.
# Secondly, we generate predictions from `estimator` distance ranges in
# [xmin, xmax].
def Plot(distance, label, estimator, xmin=0.0, xmax=10.0):
%matplotlib inline
test_features = {
'distance': np.linspace(xmin, xmax, num=100)
}
# Estimator accepts an input in the form of input_fn (callable).
# numpy_input_fn creates an input function that generates a dictionary where
# the key is a feaeture name ('distance'), and the value is a tensor with
# a shape [batch_size, 1].
test_input_fn = tf.estimator.inputs.numpy_input_fn(
x=test_features,
batch_size=1,
num_epochs=1,
shuffle=False)
# Estimator's prediction is 1d tensor with a shape [batch_size]. Since we
# set batch_size == 1 in the above, p['predictions'] will contain only one
# element in each batch, and we fetch this value by p['predictions'][0].
predictions = [p['predictions'][0]
for p in estimator.predict(input_fn=test_input_fn)]
# Plot estimator's response and (distance, label) scatter plot.
fig, ax = plt.subplots(1, 1)
ax.plot(test_features['distance'], predictions)
ax.scatter(distance, label)
plt.xlabel('distance')
plt.ylabel('user hapiness')
plt.legend(['prediction', 'data'])
```
# DNN Estimator
Now let us define feature columns and use DNN regressor to fit a model.
```
# Specify feature.
feature_columns = [
tf.feature_column.numeric_column('distance'),
]
# Define a neural network legressor.
# The first hidden layer contains 30 hidden units, and the second
# hidden layer contains 10 hidden units.
dnn_estimator = tf.estimator.DNNRegressor(
feature_columns=feature_columns,
hidden_units=[30, 10],
optimizer=tf.train.GradientDescentOptimizer(
learning_rate=0.01,
),
)
# Define training input function.
# mini-batch size is 10, and we iterate the dataset over
# 1000 times.
train_input_fn = tf.estimator.inputs.numpy_input_fn(
x=train_features,
y=train_labels,
batch_size=10,
num_epochs=1000,
shuffle=False)
tf.logging.set_verbosity(tf.logging.ERROR)
# Train this estimator
dnn_estimator.train(input_fn=train_input_fn)
# Response in [0.0, 1.0] range
Plot(train_features['distance'], train_labels, dnn_estimator, 0.0, 1.0)
# Now let's increase the prediction range to [0.0, 3.0]
# Note) In most machines, the prediction is going up.
# However, DNN training does not have a unique solution, so it's possible
# not to see this phenomenon.
Plot(train_features['distance'], train_labels, dnn_estimator, 0.0, 3.0)
```
# TensorFlow Lattice calibrated linear model
Let's use calibrated linear model to fit the data.
Since we only have one example, there's no reason to use a lattice.
```
# TensorFlow Lattice needs feature names to specify
# per-feature parameters.
feature_names = [fc.name for fc in feature_columns]
num_keypoints = 5
hparams = tfl.CalibratedLinearHParams(
feature_names=feature_names,
learning_rate=0.1,
num_keypoints=num_keypoints)
# input keypoint initializers.
# init_fns are dict of (feature_name, callable initializer).
keypoints_init_fns = {
'distance': lambda: tfl.uniform_keypoints_for_signal(num_keypoints,
input_min=0.0,
input_max=0.7,
output_min=-1.0,
output_max=1.0)}
non_monotnic_estimator = tfl.calibrated_linear_regressor(
feature_columns=feature_columns,
keypoints_initializers_fn=keypoints_init_fns,
hparams=hparams)
non_monotnic_estimator.train(input_fn=train_input_fn)
# The prediction goes up!
Plot(train_features['distance'], train_labels, non_monotnic_estimator, 0.0, 1.0)
# Declare distance as a decreasing monotonic input.
hparams.set_feature_param('distance', 'monotonicity', -1)
monotonic_estimator = tfl.calibrated_linear_regressor(
feature_columns=feature_columns,
keypoints_initializers_fn=keypoints_init_fns,
hparams=hparams)
monotonic_estimator.train(input_fn=train_input_fn)
# Now it's decreasing.
Plot(train_features['distance'], train_labels, monotonic_estimator, 0.0, 1.0)
# Even if the output range becomes larger, the prediction never goes up!
Plot(train_features['distance'], train_labels, monotonic_estimator, 0.0, 3.0)
```
| github_jupyter |
# Introduction to Jupyter
## BProf Python course
### June 25-29, 2018
#### Judit Ács
# Jupyter
- Jupyter - formally known as IPython Notebook is a web application that allows you to create and share documents with live code, equations, visualizations etc.
- Jupyter notebooks are JSON files with the extension `.ipynb`
- can be converted to HTML, PDF, LateX etc.
- can render images, tables, graphs, LateX equations
- large number of extensions
- `jupyter-vim-binding` is used in this lecture
- content is organized into cells
# Cell types
1. code cell: Python/R/Lua/etc. code
2. raw cell: raw text
3. markdown cell: formatted text using Markdown
# Code cell
```
print("Hello world")
```
The last command's output is displayed
```
2 + 3
3 + 4
```
This can be a tuple of multiple values
```
2 + 3, 3 + 4, "hello " + "world"
```
# Markdown cell
**This is in bold**
*This is in italics*
| This | is |
| --- | --- |
| a | table |
and is a pretty LateX equation:
$$
\mathbf{E}\cdot\mathrm{d}\mathbf{S} = \frac{1}{\varepsilon_0} \iiint_\Omega \rho \,\mathrm{d}V
$$
# Using Jupyter
## Command mode and edit mode
Jupyter has two modes: command mode and edit mode
1. Command mode: perform non-edit operations on selected cells (can select more than one cell)
- selected cells are marked blue
2. Edit mode: edit a single cell
- the cell being edited is marked green
### Switching between modes
1. Esc: Edit mode -> Command mode
2. Enter or double click: Command mode -> Edit mode
## Running cells
1. Ctrl + Enter: run cell
2. Shift + Enter: run cell and select next cell
3. Alt + Enter: run cell and insert new cell below
# Cell magic
Special commands can modify a single cell's behavior, for example
```
%%time
for x in range(1000000):
pass
%%timeit
x = 2
%%writefile hello.py
print("Hello world from BME")
```
For a complete list of magic commands:
```
%lsmagic
```
# Under the hood
- each notebook is run by its own _Kernel_ (Python interpreter)
- the kernel can interrupted or restarted through the Kernel menu
- **always** run `Kernel -> Restart & Run All` before submitting homework to make sure that your notebook behaves as expected
- all cells share a single namespace
- cells can be run in arbitrary order, execution count is helpful
```
print("this is run first")
print("this is run afterwords. Note the execution count on the left.")
```
## The input and output of code cells can be accessed
Previous output:
```
42
_
```
Next-previous output:
```
"first"
"second"
__
__
```
Next-next previous output:
```
___
_3
```
N-th output can also be accessed as a variable `_output_count`. This is only defined if the N-th cell had an output.
Here is a way to list all defined outputs (you will understand this piece of code in a few days):
```
list(filter(lambda x: x.startswith('_') and x[1:].isdigit(),
globals()))
```
## Inputs can be accessed similarly
Previous input:
```
_i
```
N-th input:
```
_i2
```
| github_jupyter |
# Behaviorial Cloning Project
[](http://www.udacity.com/drive)
**Behavioral Cloning Project**
The goals / steps of this project are the following:
* Read and explore dataset provide by Udacity
* Build, a convolution neural network in Keras that predicts steering angles from images
* Train and validate the model with a training and validation set
* Test that the model successfully drives around track one without leaving the road
* Summarize the results with a written report
[//]: # (Image References)
[image1]: ./examples/raw_image.png "raw_image"
[image2]: ./examples/steering.png "steering"
[image3]: ./examples/crop.png "crop Image"
[image4]: ./examples/bir.png "bir Image"
[image5]: ./examples/flip.png "flip Image"
[image6]: ./examples/model.png "model Image"
[image7]: ./examples/placeholder_small.png "Flipped Image"
---
## Read and explore dataset
I use the dataset provided by Udacity which is enough to train my model, so i did not record images myself.
If you want, you can use .[Udacity simulator](https://github.com/udacity/self-driving-car-sim) to generate your own dataset.
The images output from simulator is 160x320x3 dimensions. I randomly print out three images with three different positions.
![alt text][image1]
The dataset contains 8036 images. One pontential issue of this dataset is that most of the steering angles are close to zero.
![alt text][image2]
---
## Preprocess and augment dataset
### Data Preprecessing
**Cropping ane resizing images**
The top of images mostly capture sky and tress and hills and other elements that might be more distracting the model. Beside bottom portion of the image captures the hood of the car.
In order to focus on only the portion of the image that is useful for predicting a steering angle, i crop 55 pixels from the top and 25 pixels from the bottom.
![alt text][image3]
**Normalizing the data and mean centering the data**
* pixel_normalized = pixel / 127.5
* pixel_mean_centered = pixel / 127.5 - 1
### Data Augmentation
**Adjust the brightness of the images**
![alt text][image4]
**Flipping Images**
Flipping images and taking the opposite sign of the steering measurement
![alt text][image5]
**Using multiple cameras**
The simulator can capture three position images which are a center, right and left camera image. I can use these side camera images to increase the dataset. More importantly, it will be helpful to recover the car from being off the center.
In order to use these side cameras, I will add a small correction factor to these cameras.
* For the left camera: center steering angle + 0.25
* For the right camera: center steering angle - 0.25
---
## Build a convolution neural network to predicts steering angles
I implmented the end-to-end CNN model to predict the steering angles which is based on .[NVIDIA architecture](http://images.nvidia.com/content/tegra/automotive/images/2016/solutions/pdf/end-to-end-dl-using-px.pdf)
**My final model consisted of the following layers:**
* Optimizer: Adam
* Loss: Mean square error
* Batch size: 32
* Epoch: 3
![alt text][image6]
---
## Results
* .[Behavioral_Cloning.py](Behavioral_Cloning.py): project file
* .[drive.py](drive.py): drive a car in autonomous mode
* .[model.h5](model.h5): trained model to predict steering angles
* .[video.py](video.py): create video file
* .[run1.mp4](run1.mp4): a final result, drive the car on the track 1
---
## Potential shortcomings and future works
My model can drive car safely on the tack1, but the speed is limited. When the speed gets faster, the car will shake a bit from side to side.
### solutions:
* Since the dataset is unbalance and all the steering angle are cloesed to zero, i can try to balance this dataset which will help to make the car stable.
* Try to apply binary and color threshold techniques to preprocess the data captured by cameras. It will be helpful when the car drives under shadows.
| github_jupyter |
```
# find the dataset definition by name, for example dtu_yao (dtu_yao.py)
def find_dataset_def(dataset_name):
module_name = 'datasets.{}'.format(dataset_name)
module = importlib.import_module(module_name)
return getattr(module, "MVSDataset")
"""
Implementation of Pytorch layer primitives, such as Conv+BN+ReLU, differentiable warping layers,
and depth regression based upon expectation of an input probability distribution.
"""
import torch
import torch.nn as nn
import torch.nn.functional as F
class ConvBnReLU(nn.Module):
"""Implements 2d Convolution + batch normalization + ReLU"""
def __init__(
self,
in_channels: int,
out_channels: int,
kernel_size: int = 3,
stride: int = 1,
pad: int = 1,
dilation: int = 1,
) -> None:
"""initialization method for convolution2D + batch normalization + relu module
Args:
in_channels: input channel number of convolution layer
out_channels: output channel number of convolution layer
kernel_size: kernel size of convolution layer
stride: stride of convolution layer
pad: pad of convolution layer
dilation: dilation of convolution layer
"""
super(ConvBnReLU, self).__init__()
self.conv = nn.Conv2d(
in_channels, out_channels, kernel_size, stride=stride, padding=pad, dilation=dilation, bias=False
)
self.bn = nn.BatchNorm2d(out_channels)
def forward(self, x: torch.Tensor) -> torch.Tensor:
"""forward method"""
return F.relu(self.bn(self.conv(x)), inplace=True)
class ConvBnReLU3D(nn.Module):
"""Implements of 3d convolution + batch normalization + ReLU."""
def __init__(
self,
in_channels: int,
out_channels: int,
kernel_size: int = 3,
stride: int = 1,
pad: int = 1,
dilation: int = 1,
) -> None:
"""initialization method for convolution3D + batch normalization + relu module
Args:
in_channels: input channel number of convolution layer
out_channels: output channel number of convolution layer
kernel_size: kernel size of convolution layer
stride: stride of convolution layer
pad: pad of convolution layer
dilation: dilation of convolution layer
"""
super(ConvBnReLU3D, self).__init__()
self.conv = nn.Conv3d(
in_channels, out_channels, kernel_size, stride=stride, padding=pad, dilation=dilation, bias=False
)
self.bn = nn.BatchNorm3d(out_channels)
def forward(self, x: torch.Tensor) -> torch.Tensor:
"""forward method"""
return F.relu(self.bn(self.conv(x)), inplace=True)
class ConvBnReLU1D(nn.Module):
"""Implements 1d Convolution + batch normalization + ReLU."""
def __init__(
self,
in_channels: int,
out_channels: int,
kernel_size: int = 3,
stride: int = 1,
pad: int = 1,
dilation: int = 1,
) -> None:
"""initialization method for convolution1D + batch normalization + relu module
Args:
in_channels: input channel number of convolution layer
out_channels: output channel number of convolution layer
kernel_size: kernel size of convolution layer
stride: stride of convolution layer
pad: pad of convolution layer
dilation: dilation of convolution layer
"""
super(ConvBnReLU1D, self).__init__()
self.conv = nn.Conv1d(
in_channels, out_channels, kernel_size, stride=stride, padding=pad, dilation=dilation, bias=False
)
self.bn = nn.BatchNorm1d(out_channels)
def forward(self, x: torch.Tensor) -> torch.Tensor:
"""forward method"""
return F.relu(self.bn(self.conv(x)), inplace=True)
class ConvBn(nn.Module):
"""Implements of 2d convolution + batch normalization."""
def __init__(
self, in_channels: int, out_channels: int, kernel_size: int = 3, stride: int = 1, pad: int = 1
) -> None:
"""initialization method for convolution2D + batch normalization + ReLU module
Args:
in_channels: input channel number of convolution layer
out_channels: output channel number of convolution layer
kernel_size: kernel size of convolution layer
stride: stride of convolution layer
pad: pad of convolution layer
"""
super(ConvBn, self).__init__()
self.conv = nn.Conv2d(in_channels, out_channels, kernel_size, stride=stride, padding=pad, bias=False)
self.bn = nn.BatchNorm2d(out_channels)
def forward(self, x: torch.Tensor) -> torch.Tensor:
"""forward method"""
return self.bn(self.conv(x))
def differentiable_warping(
src_fea: torch.Tensor, src_proj: torch.Tensor, ref_proj: torch.Tensor, depth_samples: torch.Tensor
):
"""Differentiable homography-based warping, implemented in Pytorch.
Args:
src_fea: [B, C, H, W] source features, for each source view in batch
src_proj: [B, 4, 4] source camera projection matrix, for each source view in batch
ref_proj: [B, 4, 4] reference camera projection matrix, for each ref view in batch
depth_samples: [B, Ndepth, H, W] virtual depth layers
Returns:
warped_src_fea: [B, C, Ndepth, H, W] features on depths after perspective transformation
"""
batch, channels, height, width = src_fea.shape
num_depth = depth_samples.shape[1]
with torch.no_grad():
proj = torch.matmul(src_proj, torch.inverse(ref_proj))
rot = proj[:, :3, :3] # [B,3,3]
trans = proj[:, :3, 3:4] # [B,3,1]
y, x = torch.meshgrid(
[
torch.arange(0, height, dtype=torch.float32, device=src_fea.device),
torch.arange(0, width, dtype=torch.float32, device=src_fea.device),
]
)
y, x = y.contiguous(), x.contiguous()
y, x = y.view(height * width), x.view(height * width)
xyz = torch.stack((x, y, torch.ones_like(x))) # [3, H*W]
xyz = torch.unsqueeze(xyz, 0).repeat(batch, 1, 1) # [B, 3, H*W]
rot_xyz = torch.matmul(rot, xyz) # [B, 3, H*W]
rot_depth_xyz = rot_xyz.unsqueeze(2).repeat(1, 1, num_depth, 1) * depth_samples.view(
batch, 1, num_depth, height * width
) # [B, 3, Ndepth, H*W]
proj_xyz = rot_depth_xyz + trans.view(batch, 3, 1, 1) # [B, 3, Ndepth, H*W]
# avoid negative depth
negative_depth_mask = proj_xyz[:, 2:] <= 1e-3
proj_xyz[:, 0:1][negative_depth_mask] = float(width)
proj_xyz[:, 1:2][negative_depth_mask] = float(height)
proj_xyz[:, 2:3][negative_depth_mask] = 1.0
proj_xy = proj_xyz[:, :2, :, :] / proj_xyz[:, 2:3, :, :] # [B, 2, Ndepth, H*W]
proj_x_normalized = proj_xy[:, 0, :, :] / ((width - 1) / 2) - 1 # [B, Ndepth, H*W]
proj_y_normalized = proj_xy[:, 1, :, :] / ((height - 1) / 2) - 1
proj_xy = torch.stack((proj_x_normalized, proj_y_normalized), dim=3) # [B, Ndepth, H*W, 2]
grid = proj_xy
warped_src_fea = F.grid_sample(
src_fea,
grid.view(batch, num_depth * height, width, 2),
mode="bilinear",
padding_mode="zeros",
align_corners=True,
)
return warped_src_fea.view(batch, channels, num_depth, height, width)
def depth_regression(p: torch.Tensor, depth_values: torch.Tensor) -> torch.Tensor:
"""Implements per-pixel depth regression based upon a probability distribution per-pixel.
The regressed depth value D(p) at pixel p is found as the expectation w.r.t. P of the hypotheses.
Args:
p: probability volume [B, D, H, W]
depth_values: discrete depth values [B, D]
Returns:
result depth: expected value, soft argmin [B, 1, H, W]
"""
return torch.sum(p * depth_values.view(depth_values.shape[0], 1, 1), dim=1).unsqueeze(1)
def is_empty(x: torch.Tensor) -> bool:
return x.numel() == 0
from typing import Dict, List, Tuple
import torch
import torch.nn as nn
import torch.nn.functional as F
from .module import ConvBnReLU, depth_regression
from .patchmatch import PatchMatch
class FeatureNet(nn.Module):
"""Feature Extraction Network: to extract features of original images from each view"""
def __init__(self):
"""Initialize different layers in the network"""
super(FeatureNet, self).__init__()
self.conv0 = ConvBnReLU(3, 8, 3, 1, 1)
# [B,8,H,W]
self.conv1 = ConvBnReLU(8, 8, 3, 1, 1)
# [B,16,H/2,W/2]
self.conv2 = ConvBnReLU(8, 16, 5, 2, 2)
self.conv3 = ConvBnReLU(16, 16, 3, 1, 1)
self.conv4 = ConvBnReLU(16, 16, 3, 1, 1)
# [B,32,H/4,W/4]
self.conv5 = ConvBnReLU(16, 32, 5, 2, 2)
self.conv6 = ConvBnReLU(32, 32, 3, 1, 1)
self.conv7 = ConvBnReLU(32, 32, 3, 1, 1)
# [B,64,H/8,W/8]
self.conv8 = ConvBnReLU(32, 64, 5, 2, 2)
self.conv9 = ConvBnReLU(64, 64, 3, 1, 1)
self.conv10 = ConvBnReLU(64, 64, 3, 1, 1)
self.output1 = nn.Conv2d(64, 64, 1, bias=False)
self.inner1 = nn.Conv2d(32, 64, 1, bias=True)
self.inner2 = nn.Conv2d(16, 64, 1, bias=True)
self.output2 = nn.Conv2d(64, 32, 1, bias=False)
self.output3 = nn.Conv2d(64, 16, 1, bias=False)
def forward(self, x: torch.Tensor) -> Dict[int, torch.Tensor]:
"""Forward method
Args:
x: images from a single view, in the shape of [B, C, H, W]. Generally, C=3
Returns:
output_feature: a python dictionary contains extracted features from stage 1 to stage 3
keys are 1, 2, and 3
"""
output_feature: Dict[int, torch.Tensor] = {}
conv1 = self.conv1(self.conv0(x))
conv4 = self.conv4(self.conv3(self.conv2(conv1)))
conv7 = self.conv7(self.conv6(self.conv5(conv4)))
conv10 = self.conv10(self.conv9(self.conv8(conv7)))
output_feature[3] = self.output1(conv10)
intra_feat = F.interpolate(conv10, scale_factor=2.0, mode="bilinear", align_corners=False) + self.inner1(conv7)
del conv7
del conv10
output_feature[2] = self.output2(intra_feat)
intra_feat = F.interpolate(
intra_feat, scale_factor=2.0, mode="bilinear", align_corners=False) + self.inner2(conv4)
del conv4
output_feature[1] = self.output3(intra_feat)
del intra_feat
return output_feature
class Refinement(nn.Module):
"""Depth map refinement network"""
def __init__(self):
"""Initialize"""
super(Refinement, self).__init__()
# img: [B,3,H,W]
self.conv0 = ConvBnReLU(in_channels=3, out_channels=8)
# depth map:[B,1,H/2,W/2]
self.conv1 = ConvBnReLU(in_channels=1, out_channels=8)
self.conv2 = ConvBnReLU(in_channels=8, out_channels=8)
self.deconv = nn.ConvTranspose2d(
in_channels=8, out_channels=8, kernel_size=3, padding=1, output_padding=1, stride=2, bias=False
)
self.bn = nn.BatchNorm2d(8)
self.conv3 = ConvBnReLU(in_channels=16, out_channels=8)
self.res = nn.Conv2d(in_channels=8, out_channels=1, kernel_size=3, padding=1, bias=False)
def forward(
self, img: torch.Tensor, depth_0: torch.Tensor, depth_min: torch.Tensor, depth_max: torch.Tensor
) -> torch.Tensor:
"""Forward method
Args:
img: input reference images (B, 3, H, W)
depth_0: current depth map (B, 1, H//2, W//2)
depth_min: pre-defined minimum depth (B, )
depth_max: pre-defined maximum depth (B, )
Returns:
depth: refined depth map (B, 1, H, W)
"""
batch_size = depth_min.size()[0]
# pre-scale the depth map into [0,1]
depth = (depth_0 - depth_min.view(batch_size, 1, 1, 1)) / (depth_max - depth_min).view(batch_size, 1, 1, 1)
conv0 = self.conv0(img)
deconv = F.relu(self.bn(self.deconv(self.conv2(self.conv1(depth)))), inplace=True)
# depth residual
res = self.res(self.conv3(torch.cat((deconv, conv0), dim=1)))
del conv0
del deconv
depth = F.interpolate(depth, scale_factor=2.0, mode="nearest") + res
# convert the normalized depth back
return depth * (depth_max - depth_min).view(batch_size, 1, 1, 1) + depth_min.view(batch_size, 1, 1, 1)
class PatchmatchNet(nn.Module):
""" Implementation of complete structure of PatchmatchNet"""
def __init__(
self,
patchmatch_interval_scale: List[float] = [0.005, 0.0125, 0.025],
propagation_range: List[int] = [6, 4, 2],
patchmatch_iteration: List[int] = [1, 2, 2],
patchmatch_num_sample: List[int] = [8, 8, 16],
propagate_neighbors: List[int] = [0, 8, 16],
evaluate_neighbors: List[int] = [9, 9, 9],
) -> None:
"""Initialize modules in PatchmatchNet
Args:
patchmatch_interval_scale: depth interval scale in patchmatch module
propagation_range: propagation range
patchmatch_iteration: patchmatch iteration number
patchmatch_num_sample: patchmatch number of samples
propagate_neighbors: number of propagation neighbors
evaluate_neighbors: number of propagation neighbors for evaluation
"""
super(PatchmatchNet, self).__init__()
self.stages = 4
self.feature = FeatureNet()
self.patchmatch_num_sample = patchmatch_num_sample
num_features = [16, 32, 64]
self.propagate_neighbors = propagate_neighbors
self.evaluate_neighbors = evaluate_neighbors
# number of groups for group-wise correlation
self.G = [4, 8, 8]
for i in range(self.stages - 1):
patchmatch = PatchMatch(
propagation_out_range=propagation_range[i],
patchmatch_iteration=patchmatch_iteration[i],
patchmatch_num_sample=patchmatch_num_sample[i],
patchmatch_interval_scale=patchmatch_interval_scale[i],
num_feature=num_features[i],
G=self.G[i],
propagate_neighbors=self.propagate_neighbors[i],
evaluate_neighbors=evaluate_neighbors[i],
stage=i + 1,
)
setattr(self, f"patchmatch_{i+1}", patchmatch)
self.upsample_net = Refinement()
def forward(
self,
images: Dict[str, torch.Tensor],
proj_matrices: Dict[str, torch.Tensor],
depth_min: torch.Tensor,
depth_max: torch.Tensor,
) -> Tuple[torch.Tensor, torch.Tensor, Dict[int, List[torch.Tensor]]]:
"""Forward method for PatchMatchNet
Args:
images: different stages of images (B, 3, H, W) stored in the dictionary
proj_matrices: different stages of camera projection matrices (B, 4, 4) stored in the dictionary
depth_min: minimum virtual depth (B, )
depth_max: maximum virtual depth (B, )
Returns:
output tuple of PatchMatchNet, containing refined depthmap, depth patchmatch, and photometric confidence.
"""
imgs_0 = torch.unbind(images["stage_0"], 1)
del images
ref_image = imgs_0[0]
proj_mtx = {
0: torch.unbind(proj_matrices["stage_0"].float(), 1),
1: torch.unbind(proj_matrices["stage_1"].float(), 1),
2: torch.unbind(proj_matrices["stage_2"].float(), 1),
3: torch.unbind(proj_matrices["stage_3"].float(), 1)
}
del proj_matrices
assert len(imgs_0) == len(proj_mtx[0]), "Different number of images and projection matrices"
# step 1. Multi-scale feature extraction
features: List[Dict[int, torch.Tensor]] = []
for img in imgs_0:
output_feature = self.feature(img)
features.append(output_feature)
del imgs_0
ref_feature, src_features = features[0], features[1:]
depth_min = depth_min.float()
depth_max = depth_max.float()
# step 2. Learning-based patchmatch
depth = torch.empty(0)
depths: List[torch.Tensor] = []
score = torch.empty(0)
view_weights = torch.empty(0)
depth_patchmatch: Dict[int, List[torch.Tensor]] = {}
for stage in range(self.stages - 1, 0, -1):
src_features_l = [src_fea[stage] for src_fea in src_features]
ref_proj, src_projs = proj_mtx[stage][0], proj_mtx[stage][1:]
# Need conditional since TorchScript only allows "getattr" access with string literals
if stage == 3:
depths, _, view_weights = self.patchmatch_3(
ref_feature=ref_feature[stage],
src_features=src_features_l,
ref_proj=ref_proj,
src_projs=src_projs,
depth_min=depth_min,
depth_max=depth_max,
depth=depth,
view_weights=view_weights,
)
elif stage == 2:
depths, _, view_weights = self.patchmatch_2(
ref_feature=ref_feature[stage],
src_features=src_features_l,
ref_proj=ref_proj,
src_projs=src_projs,
depth_min=depth_min,
depth_max=depth_max,
depth=depth,
view_weights=view_weights,
)
elif stage == 1:
depths, score, _ = self.patchmatch_1(
ref_feature=ref_feature[stage],
src_features=src_features_l,
ref_proj=ref_proj,
src_projs=src_projs,
depth_min=depth_min,
depth_max=depth_max,
depth=depth,
view_weights=view_weights,
)
depth_patchmatch[stage] = depths
depth = depths[-1].detach()
if stage > 1:
# upsampling the depth map and pixel-wise view weight for next stage
depth = F.interpolate(depth, scale_factor=2.0, mode="nearest")
view_weights = F.interpolate(view_weights, scale_factor=2.0, mode="nearest")
del ref_feature
del src_features
# step 3. Refinement
depth = self.upsample_net(ref_image, depth, depth_min, depth_max)
if self.training:
return depth, torch.empty(0), depth_patchmatch
else:
num_depth = self.patchmatch_num_sample[0]
score_sum4 = 4 * F.avg_pool3d(
F.pad(score.unsqueeze(1), pad=(0, 0, 0, 0, 1, 2)), (4, 1, 1), stride=1, padding=0
).squeeze(1)
# [B, 1, H, W]
depth_index = depth_regression(
score, depth_values=torch.arange(num_depth, device=score.device, dtype=torch.float)
).long().clamp(0, num_depth - 1)
photometric_confidence = torch.gather(score_sum4, 1, depth_index)
photometric_confidence = F.interpolate(photometric_confidence, scale_factor=2.0, mode="nearest").squeeze(1)
return depth, photometric_confidence, depth_patchmatch
def patchmatchnet_loss(
depth_patchmatch: Dict[int, List[torch.Tensor]],
depth_gt: Dict[str, torch.Tensor],
mask: Dict[str, torch.Tensor],
) -> torch.Tensor:
"""Patchmatch Net loss function
Args:
depth_patchmatch: depth map predicted by patchmatch net
depth_gt: ground truth depth map
mask: mask for filter valid points
Returns:
loss: result loss value
"""
loss = 0
for i in range(0, 4):
mask_i = mask[f"stage_{i}"] > 0.5
gt_depth = depth_gt[f"stage_{i}"][mask_i]
for depth in depth_patchmatch[i]:
loss = loss + F.smooth_l1_loss(depth[mask_i], gt_depth, reduction="mean")
return loss
"""
PatchmatchNet uses the following main steps:
1. Initialization: generate random hypotheses;
2. Propagation: propagate hypotheses to neighbors;
3. Evaluation: compute the matching costs for all the hypotheses and choose best solutions.
"""
from typing import List, Tuple
import torch
import torch.nn as nn
import torch.nn.functional as F
from .module import ConvBnReLU3D, differentiable_warping, is_empty
class DepthInitialization(nn.Module):
"""Initialization Stage Class"""
def __init__(self, patchmatch_num_sample: int = 1) -> None:
"""Initialize method
Args:
patchmatch_num_sample: number of samples used in patchmatch process
"""
super(DepthInitialization, self).__init__()
self.patchmatch_num_sample = patchmatch_num_sample
def forward(
self,
min_depth: torch.Tensor,
max_depth: torch.Tensor,
height: int,
width: int,
depth_interval_scale: float,
device: torch.device,
depth: torch.Tensor = torch.empty(0),
) -> torch.Tensor:
"""Forward function for depth initialization
Args:
min_depth: minimum virtual depth, (B, )
max_depth: maximum virtual depth, (B, )
height: height of depth map
width: width of depth map
depth_interval_scale: depth interval scale
device: device on which to place tensor
depth: current depth (B, 1, H, W)
Returns:
depth_sample: initialized sample depth map by randomization or local perturbation (B, Ndepth, H, W)
"""
batch_size = min_depth.size()[0]
inverse_min_depth = 1.0 / min_depth
inverse_max_depth = 1.0 / max_depth
if is_empty(depth):
# first iteration of Patchmatch on stage 3, sample in the inverse depth range
# divide the range into several intervals and sample in each of them
patchmatch_num_sample = 48
# [B,Ndepth,H,W]
depth_sample = torch.rand(
size=(batch_size, patchmatch_num_sample, height, width), device=device
) + torch.arange(start=0, end=patchmatch_num_sample, step=1, device=device).view(
1, patchmatch_num_sample, 1, 1
)
depth_sample = inverse_max_depth.view(batch_size, 1, 1, 1) + depth_sample / patchmatch_num_sample * (
inverse_min_depth.view(batch_size, 1, 1, 1) - inverse_max_depth.view(batch_size, 1, 1, 1)
)
return 1.0 / depth_sample
elif self.patchmatch_num_sample == 1:
return depth.detach()
else:
# other Patchmatch, local perturbation is performed based on previous result
# uniform samples in an inversed depth range
depth_sample = (
torch.arange(-self.patchmatch_num_sample // 2, self.patchmatch_num_sample // 2, 1, device=device)
.view(1, self.patchmatch_num_sample, 1, 1).repeat(batch_size, 1, height, width).float()
)
inverse_depth_interval = (inverse_min_depth - inverse_max_depth) * depth_interval_scale
inverse_depth_interval = inverse_depth_interval.view(batch_size, 1, 1, 1)
depth_sample = 1.0 / depth.detach() + inverse_depth_interval * depth_sample
depth_clamped = []
del depth
for k in range(batch_size):
depth_clamped.append(
torch.clamp(depth_sample[k], min=inverse_max_depth[k], max=inverse_min_depth[k]).unsqueeze(0)
)
return 1.0 / torch.cat(depth_clamped, dim=0)
class Propagation(nn.Module):
""" Propagation module implementation"""
def __init__(self) -> None:
"""Initialize method"""
super(Propagation, self).__init__()
def forward(self, depth_sample: torch.Tensor, grid: torch.Tensor) -> torch.Tensor:
# [B,D,H,W]
"""Forward method of adaptive propagation
Args:
depth_sample: sample depth map, in shape of [batch, num_depth, height, width],
grid: 2D grid for bilinear gridding, in shape of [batch, neighbors*H, W, 2]
Returns:
propagate depth: sorted propagate depth map [batch, num_depth+num_neighbors, height, width]
"""
batch, num_depth, height, width = depth_sample.size()
num_neighbors = grid.size()[1] // height
propagate_depth_sample = F.grid_sample(
depth_sample[:, num_depth // 2, :, :].unsqueeze(1),
grid,
mode="bilinear",
padding_mode="border",
align_corners=False
).view(batch, num_neighbors, height, width)
return torch.sort(torch.cat((depth_sample, propagate_depth_sample), dim=1), dim=1)[0]
class Evaluation(nn.Module):
"""Evaluation module for adaptive evaluation step in Learning-based Patchmatch
Used to compute the matching costs for all the hypotheses and choose best solutions.
"""
def __init__(self, G: int = 8) -> None:
"""Initialize method`
Args:
G: the feature channels of input will be divided evenly into G groups
"""
super(Evaluation, self).__init__()
self.G = G
self.pixel_wise_net = PixelwiseNet(self.G)
self.softmax = nn.LogSoftmax(dim=1)
self.similarity_net = SimilarityNet(self.G)
def forward(
self,
ref_feature: torch.Tensor,
src_features: List[torch.Tensor],
ref_proj: torch.Tensor,
src_projs: List[torch.Tensor],
depth_sample: torch.Tensor,
grid: torch.Tensor,
weight: torch.Tensor,
view_weights: torch.Tensor = torch.empty(0),
is_inverse: bool = False
) -> Tuple[torch.Tensor, torch.Tensor, torch.Tensor]:
"""Forward method for adaptive evaluation
Args:
ref_feature: feature from reference view, (B, C, H, W)
src_features: features from (Nview-1) source views, (Nview-1) * (B, C, H, W), where Nview is the number of
input images (or views) of PatchmatchNet
ref_proj: projection matrix of reference view, (B, 4, 4)
src_projs: source matrices of source views, (Nview-1) * (B, 4, 4), where Nview is the number of input
images (or views) of PatchmatchNet
depth_sample: sample depth map, (B,Ndepth,H,W)
grid: grid, (B, evaluate_neighbors*H, W, 2)
weight: weight, (B,Ndepth,1,H,W)
view_weights: Tensor to store weights of source views, in shape of (B,Nview-1,H,W),
Nview-1 represents the number of source views
is_inverse: Flag for inverse depth regression
Returns:
depth_sample: expectation of depth sample, (B,H,W)
score: probability map, (B,Ndepth,H,W)
view_weights: optional, Tensor to store weights of source views, in shape of (B,Nview-1,H,W),
Nview-1 represents the number of source views
"""
batch, feature_channel, height, width = ref_feature.size()
device = ref_feature.device
num_depth = depth_sample.size()[1]
assert (
len(src_features) == len(src_projs)
), "Patchmatch Evaluation: Different number of images and projection matrices"
if not is_empty(view_weights):
assert (
len(src_features) == view_weights.size()[1]
), "Patchmatch Evaluation: Different number of images and view weights"
# Change to a tensor with value 1e-5
pixel_wise_weight_sum = 1e-5 * torch.ones((batch, 1, 1, height, width), dtype=torch.float32, device=device)
ref_feature = ref_feature.view(batch, self.G, feature_channel // self.G, 1, height, width)
similarity_sum = torch.zeros((batch, self.G, num_depth, height, width), dtype=torch.float32, device=device)
i = 0
view_weights_list = []
for src_feature, src_proj in zip(src_features, src_projs):
warped_feature = differentiable_warping(
src_feature, src_proj, ref_proj, depth_sample
).view(batch, self.G, feature_channel // self.G, num_depth, height, width)
# group-wise correlation
similarity = (warped_feature * ref_feature).mean(2)
# pixel-wise view weight
if is_empty(view_weights):
view_weight = self.pixel_wise_net(similarity)
view_weights_list.append(view_weight)
else:
# reuse the pixel-wise view weight from first iteration of Patchmatch on stage 3
view_weight = view_weights[:, i].unsqueeze(1) # [B,1,H,W]
i = i + 1
similarity_sum += similarity * view_weight.unsqueeze(1)
pixel_wise_weight_sum += view_weight.unsqueeze(1)
# aggregated matching cost across all the source views
similarity = similarity_sum.div_(pixel_wise_weight_sum) # [B, G, Ndepth, H, W]
# adaptive spatial cost aggregation
score = self.similarity_net(similarity, grid, weight) # [B, G, Ndepth, H, W]
# apply softmax to get probability
score = torch.exp(self.softmax(score))
if is_empty(view_weights):
view_weights = torch.cat(view_weights_list, dim=1) # [B,4,H,W], 4 is the number of source views
if is_inverse:
# depth regression: inverse depth regression
depth_index = torch.arange(0, num_depth, 1, device=device).view(1, num_depth, 1, 1)
depth_index = torch.sum(depth_index * score, dim=1)
inverse_min_depth = 1.0 / depth_sample[:, -1, :, :]
inverse_max_depth = 1.0 / depth_sample[:, 0, :, :]
depth_sample = inverse_max_depth + depth_index / (num_depth - 1) * (inverse_min_depth - inverse_max_depth)
depth_sample = 1.0 / depth_sample
else:
# depth regression: expectation
depth_sample = torch.sum(depth_sample * score, dim=1)
return depth_sample, score, view_weights.detach()
class PatchMatch(nn.Module):
"""Patchmatch module"""
def __init__(
self,
propagation_out_range: int = 2,
patchmatch_iteration: int = 2,
patchmatch_num_sample: int = 16,
patchmatch_interval_scale: float = 0.025,
num_feature: int = 64,
G: int = 8,
propagate_neighbors: int = 16,
evaluate_neighbors: int = 9,
stage: int = 3,
) -> None:
"""Initialize method
Args:
propagation_out_range: range of propagation out,
patchmatch_iteration: number of iterations in patchmatch,
patchmatch_num_sample: number of samples in patchmatch,
patchmatch_interval_scale: interval scale,
num_feature: number of features,
G: the feature channels of input will be divided evenly into G groups,
propagate_neighbors: number of neighbors to be sampled in propagation,
stage: number of stage,
evaluate_neighbors: number of neighbors to be sampled in evaluation,
"""
super(PatchMatch, self).__init__()
self.patchmatch_iteration = patchmatch_iteration
self.patchmatch_interval_scale = patchmatch_interval_scale
self.propa_num_feature = num_feature
# group wise correlation
self.G = G
self.stage = stage
self.dilation = propagation_out_range
self.propagate_neighbors = propagate_neighbors
self.evaluate_neighbors = evaluate_neighbors
# Using dictionary instead of Enum since TorchScript cannot recognize and export it correctly
self.grid_type = {"propagation": 1, "evaluation": 2}
self.depth_initialization = DepthInitialization(patchmatch_num_sample)
self.propagation = Propagation()
self.evaluation = Evaluation(self.G)
# adaptive propagation: last iteration on stage 1 does not have propagation,
# but we still define this for TorchScript export compatibility
self.propa_conv = nn.Conv2d(
in_channels=self.propa_num_feature,
out_channels=max(2 * self.propagate_neighbors, 1),
kernel_size=3,
stride=1,
padding=self.dilation,
dilation=self.dilation,
bias=True,
)
nn.init.constant_(self.propa_conv.weight, 0.0)
nn.init.constant_(self.propa_conv.bias, 0.0)
# adaptive spatial cost aggregation (adaptive evaluation)
self.eval_conv = nn.Conv2d(
in_channels=self.propa_num_feature,
out_channels=2 * self.evaluate_neighbors,
kernel_size=3,
stride=1,
padding=self.dilation,
dilation=self.dilation,
bias=True,
)
nn.init.constant_(self.eval_conv.weight, 0.0)
nn.init.constant_(self.eval_conv.bias, 0.0)
self.feature_weight_net = FeatureWeightNet(self.evaluate_neighbors, self.G)
def get_grid(
self, grid_type: int, batch: int, height: int, width: int, offset: torch.Tensor, device: torch.device
) -> torch.Tensor:
"""Compute the offset for adaptive propagation or spatial cost aggregation in adaptive evaluation
Args:
grid_type: type of grid - propagation (1) or evaluation (2)
batch: batch size
height: grid height
width: grid width
offset: grid offset
device: device on which to place tensor
Returns:
generated grid: in the shape of [batch, propagate_neighbors*H, W, 2]
"""
if grid_type == self.grid_type["propagation"]:
if self.propagate_neighbors == 4: # if 4 neighbors to be sampled in propagation
original_offset = [[-self.dilation, 0], [0, -self.dilation], [0, self.dilation], [self.dilation, 0]]
elif self.propagate_neighbors == 8: # if 8 neighbors to be sampled in propagation
original_offset = [
[-self.dilation, -self.dilation],
[-self.dilation, 0],
[-self.dilation, self.dilation],
[0, -self.dilation],
[0, self.dilation],
[self.dilation, -self.dilation],
[self.dilation, 0],
[self.dilation, self.dilation],
]
elif self.propagate_neighbors == 16: # if 16 neighbors to be sampled in propagation
original_offset = [
[-self.dilation, -self.dilation],
[-self.dilation, 0],
[-self.dilation, self.dilation],
[0, -self.dilation],
[0, self.dilation],
[self.dilation, -self.dilation],
[self.dilation, 0],
[self.dilation, self.dilation],
]
for i in range(len(original_offset)):
offset_x, offset_y = original_offset[i]
original_offset.append([2 * offset_x, 2 * offset_y])
else:
raise NotImplementedError
elif grid_type == self.grid_type["evaluation"]:
dilation = self.dilation - 1 # dilation of evaluation is a little smaller than propagation
if self.evaluate_neighbors == 9: # if 9 neighbors to be sampled in evaluation
original_offset = [
[-dilation, -dilation],
[-dilation, 0],
[-dilation, dilation],
[0, -dilation],
[0, 0],
[0, dilation],
[dilation, -dilation],
[dilation, 0],
[dilation, dilation],
]
elif self.evaluate_neighbors == 17: # if 17 neighbors to be sampled in evaluation
original_offset = [
[-dilation, -dilation],
[-dilation, 0],
[-dilation, dilation],
[0, -dilation],
[0, 0],
[0, dilation],
[dilation, -dilation],
[dilation, 0],
[dilation, dilation],
]
for i in range(len(original_offset)):
offset_x, offset_y = original_offset[i]
if offset_x != 0 or offset_y != 0:
original_offset.append([2 * offset_x, 2 * offset_y])
else:
raise NotImplementedError
else:
raise NotImplementedError
with torch.no_grad():
y_grid, x_grid = torch.meshgrid(
[
torch.arange(0, height, dtype=torch.float32, device=device),
torch.arange(0, width, dtype=torch.float32, device=device),
]
)
y_grid, x_grid = y_grid.contiguous().view(height * width), x_grid.contiguous().view(height * width)
xy = torch.stack((x_grid, y_grid)) # [2, H*W]
xy = torch.unsqueeze(xy, 0).repeat(batch, 1, 1) # [B, 2, H*W]
xy_list = []
for i in range(len(original_offset)):
original_offset_y, original_offset_x = original_offset[i]
offset_x = original_offset_x + offset[:, 2 * i, :].unsqueeze(1)
offset_y = original_offset_y + offset[:, 2 * i + 1, :].unsqueeze(1)
xy_list.append((xy + torch.cat((offset_x, offset_y), dim=1)).unsqueeze(2))
xy = torch.cat(xy_list, dim=2) # [B, 2, 9, H*W]
del xy_list
del x_grid
del y_grid
x_normalized = xy[:, 0, :, :] / ((width - 1) / 2) - 1
y_normalized = xy[:, 1, :, :] / ((height - 1) / 2) - 1
del xy
grid = torch.stack((x_normalized, y_normalized), dim=3) # [B, 9, H*W, 2]
del x_normalized
del y_normalized
return grid.view(batch, len(original_offset) * height, width, 2)
def forward(
self,
ref_feature: torch.Tensor,
src_features: List[torch.Tensor],
ref_proj: torch.Tensor,
src_projs: List[torch.Tensor],
depth_min: torch.Tensor,
depth_max: torch.Tensor,
depth: torch.Tensor,
view_weights: torch.Tensor = torch.empty(0),
) -> Tuple[List[torch.Tensor], torch.Tensor, torch.Tensor]:
"""Forward method for PatchMatch
Args:
ref_feature: feature from reference view, (B, C, H, W)
src_features: features from (Nview-1) source views, (Nview-1) * (B, C, H, W), where Nview is the number of
input images (or views) of PatchmatchNet
ref_proj: projection matrix of reference view, (B, 4, 4)
src_projs: source matrices of source views, (Nview-1) * (B, 4, 4), where Nview is the number of input
images (or views) of PatchmatchNet
depth_min: minimum virtual depth, (B,)
depth_max: maximum virtual depth, (B,)
depth: current depth map, (B,1,H,W) or None
view_weights: Tensor to store weights of source views, in shape of (B,Nview-1,H,W),
Nview-1 represents the number of source views
Returns:
depth_samples: list of depth maps from each patchmatch iteration, Niter * (B,1,H,W)
score: evaluted probabilities, (B,Ndepth,H,W)
view_weights: Tensor to store weights of source views, in shape of (B,Nview-1,H,W),
Nview-1 represents the number of source views
"""
score = torch.empty(0)
depth_samples = []
device = ref_feature.device
batch, _, height, width = ref_feature.size()
# the learned additional 2D offsets for adaptive propagation
propa_grid = torch.empty(0)
if self.propagate_neighbors > 0 and not (self.stage == 1 and self.patchmatch_iteration == 1):
# last iteration on stage 1 does not have propagation (photometric consistency filtering)
propa_offset = self.propa_conv(ref_feature).view(batch, 2 * self.propagate_neighbors, height * width)
propa_grid = self.get_grid(self.grid_type["propagation"], batch, height, width, propa_offset, device)
# the learned additional 2D offsets for adaptive spatial cost aggregation (adaptive evaluation)
eval_offset = self.eval_conv(ref_feature).view(batch, 2 * self.evaluate_neighbors, height * width)
eval_grid = self.get_grid(self.grid_type["evaluation"], batch, height, width, eval_offset, device)
# [B, evaluate_neighbors, H, W]
feature_weight = self.feature_weight_net(ref_feature.detach(), eval_grid)
depth_sample = depth
del depth
for iter in range(1, self.patchmatch_iteration + 1):
is_inverse = self.stage == 1 and iter == self.patchmatch_iteration
# first iteration on stage 3, random initialization (depth is empty), no adaptive propagation
# subsequent iterations, local perturbation based on previous result, [B,Ndepth,H,W]
depth_sample = self.depth_initialization(
min_depth=depth_min,
max_depth=depth_max,
height=height,
width=width,
depth_interval_scale=self.patchmatch_interval_scale,
device=device,
depth=depth_sample
)
# adaptive propagation
if self.propagate_neighbors > 0 and not (self.stage == 1 and iter == self.patchmatch_iteration):
# last iteration on stage 1 does not have propagation (photometric consistency filtering)
depth_sample = self.propagation(depth_sample=depth_sample, grid=propa_grid)
# weights for adaptive spatial cost aggregation in adaptive evaluation, [B,Ndepth,N_neighbors_eval,H,W]
weight = depth_weight(
depth_sample=depth_sample.detach(),
depth_min=depth_min,
depth_max=depth_max,
grid=eval_grid.detach(),
patchmatch_interval_scale=self.patchmatch_interval_scale,
neighbors=self.evaluate_neighbors,
) * feature_weight.unsqueeze(1)
weight = weight / torch.sum(weight, dim=2).unsqueeze(2) # [B,Ndepth,1,H,W]
# evaluation, outputs regressed depth map and pixel-wise view weights which will
# be used for subsequent iterations
depth_sample, score, view_weights = self.evaluation(
ref_feature=ref_feature,
src_features=src_features,
ref_proj=ref_proj,
src_projs=src_projs,
depth_sample=depth_sample,
grid=eval_grid,
weight=weight,
view_weights=view_weights,
is_inverse=is_inverse,
)
depth_sample = depth_sample.unsqueeze(1)
depth_samples.append(depth_sample)
return depth_samples, score, view_weights
class SimilarityNet(nn.Module):
"""Similarity Net, used in Evaluation module (adaptive evaluation step)
1. Do 1x1x1 convolution on aggregated cost [B, G, Ndepth, H, W] among all the source views,
where G is the number of groups
2. Perform adaptive spatial cost aggregation to get final cost (scores)
"""
def __init__(self, G: int) -> None:
"""Initialize method
Args:
G: the feature channels of input will be divided evenly into G groups
"""
super(SimilarityNet, self).__init__()
self.conv0 = ConvBnReLU3D(in_channels=G, out_channels=16, kernel_size=1, stride=1, pad=0)
self.conv1 = ConvBnReLU3D(in_channels=16, out_channels=8, kernel_size=1, stride=1, pad=0)
self.similarity = nn.Conv3d(in_channels=8, out_channels=1, kernel_size=1, stride=1, padding=0)
def forward(self, x1: torch.Tensor, grid: torch.Tensor, weight: torch.Tensor) -> torch.Tensor:
"""Forward method for SimilarityNet
Args:
x1: [B, G, Ndepth, H, W], where G is the number of groups, aggregated cost among all the source views with
pixel-wise view weight
grid: position of sampling points in adaptive spatial cost aggregation, (B, evaluate_neighbors*H, W, 2)
weight: weight of sampling points in adaptive spatial cost aggregation, combination of
feature weight and depth weight, [B,Ndepth,1,H,W]
Returns:
final cost: in the shape of [B,Ndepth,H,W]
"""
batch, G, num_depth, height, width = x1.size()
num_neighbors = grid.size()[1] // height
# [B,Ndepth,num_neighbors,H,W]
x1 = F.grid_sample(
input=self.similarity(self.conv1(self.conv0(x1))).squeeze(1),
grid=grid,
mode="bilinear",
padding_mode="border",
align_corners=False
).view(batch, num_depth, num_neighbors, height, width)
return torch.sum(x1 * weight, dim=2)
class FeatureWeightNet(nn.Module):
"""FeatureWeight Net: Called at the beginning of patchmatch, to calculate feature weights based on similarity of
features of sampling points and center pixel. The feature weights is used to implement adaptive spatial
cost aggregation.
"""
def __init__(self, neighbors: int = 9, G: int = 8) -> None:
"""Initialize method
Args:
neighbors: number of neighbors to be sampled
G: the feature channels of input will be divided evenly into G groups
"""
super(FeatureWeightNet, self).__init__()
self.neighbors = neighbors
self.G = G
self.conv0 = ConvBnReLU3D(in_channels=G, out_channels=16, kernel_size=1, stride=1, pad=0)
self.conv1 = ConvBnReLU3D(in_channels=16, out_channels=8, kernel_size=1, stride=1, pad=0)
self.similarity = nn.Conv3d(in_channels=8, out_channels=1, kernel_size=1, stride=1, padding=0)
self.output = nn.Sigmoid()
def forward(self, ref_feature: torch.Tensor, grid: torch.Tensor) -> torch.Tensor:
"""Forward method for FeatureWeightNet
Args:
ref_feature: reference feature map, [B,C,H,W]
grid: position of sampling points in adaptive spatial cost aggregation, (B, evaluate_neighbors*H, W, 2)
Returns:
weight based on similarity of features of sampling points and center pixel, [B,Neighbor,H,W]
"""
batch, feature_channel, height, width = ref_feature.size()
weight = F.grid_sample(
ref_feature, grid, mode="bilinear", padding_mode="border", align_corners=False
).view(batch, self.G, feature_channel // self.G, self.neighbors, height, width)
# [B,G,C//G,H,W]
ref_feature = ref_feature.view(batch, self.G, feature_channel // self.G, height, width).unsqueeze(3)
# [B,G,Neighbor,H,W]
weight = (weight * ref_feature).mean(2)
# [B,Neighbor,H,W]
return self.output(self.similarity(self.conv1(self.conv0(weight))).squeeze(1))
def depth_weight(
depth_sample: torch.Tensor,
depth_min: torch.Tensor,
depth_max: torch.Tensor,
grid: torch.Tensor,
patchmatch_interval_scale: float,
neighbors: int,
) -> torch.Tensor:
"""Calculate depth weight
1. Adaptive spatial cost aggregation
2. Weight based on depth difference of sampling points and center pixel
Args:
depth_sample: sample depth map, (B,Ndepth,H,W)
depth_min: minimum virtual depth, (B,)
depth_max: maximum virtual depth, (B,)
grid: position of sampling points in adaptive spatial cost aggregation, (B, neighbors*H, W, 2)
patchmatch_interval_scale: patchmatch interval scale,
neighbors: number of neighbors to be sampled in evaluation
Returns:
depth weight
"""
batch, num_depth, height, width = depth_sample.size()
inverse_depth_min = 1.0 / depth_min
inverse_depth_max = 1.0 / depth_max
# normalization
x = 1.0 / depth_sample
del depth_sample
x = (x - inverse_depth_max.view(batch, 1, 1, 1)) / (inverse_depth_min - inverse_depth_max).view(batch, 1, 1, 1)
x1 = F.grid_sample(
x, grid, mode="bilinear", padding_mode="border", align_corners=False
).view(batch, num_depth, neighbors, height, width)
del grid
# [B,Ndepth,N_neighbors,H,W]
x1 = torch.abs(x1 - x.unsqueeze(2)) / patchmatch_interval_scale
del x
# sigmoid output approximate to 1 when x=4
return torch.sigmoid(4.0 - 2.0 * x1.clamp(min=0, max=4)).detach()
class PixelwiseNet(nn.Module):
"""Pixelwise Net: A simple pixel-wise view weight network, composed of 1x1x1 convolution layers
and sigmoid nonlinearities, takes the initial set of similarities to output a number between 0 and 1 per
pixel as estimated pixel-wise view weight.
1. The Pixelwise Net is used in adaptive evaluation step
2. The similarity is calculated by ref_feature and other source_features warped by differentiable_warping
3. The learned pixel-wise view weight is estimated in the first iteration of Patchmatch and kept fixed in the
matching cost computation.
"""
def __init__(self, G: int) -> None:
"""Initialize method
Args:
G: the feature channels of input will be divided evenly into G groups
"""
super(PixelwiseNet, self).__init__()
self.conv0 = ConvBnReLU3D(in_channels=G, out_channels=16, kernel_size=1, stride=1, pad=0)
self.conv1 = ConvBnReLU3D(in_channels=16, out_channels=8, kernel_size=1, stride=1, pad=0)
self.conv2 = nn.Conv3d(in_channels=8, out_channels=1, kernel_size=1, stride=1, padding=0)
self.output = nn.Sigmoid()
def forward(self, x1: torch.Tensor) -> torch.Tensor:
"""Forward method for PixelwiseNet
Args:
x1: pixel-wise view weight, [B, G, Ndepth, H, W], where G is the number of groups
"""
# [B,1,H,W]
return torch.max(self.output(self.conv2(self.conv1(self.conv0(x1))).squeeze(1)), dim=1)[0].unsqueeze(1)
from typing import Any, Callable, Union, Dict
import numpy as np
import torchvision.utils as vutils
import torch
import torch.utils.tensorboard as tb
def print_args(args: Any) -> None:
"""Utilities to print arguments
Arsg:
args: arguments to pring out
"""
print("################################ args ################################")
for k, v in args.__dict__.items():
print("{0: <10}\t{1: <30}\t{2: <20}".format(k, str(v), str(type(v))))
print("########################################################################")
def make_nograd_func(func: Callable) -> Callable:
"""Utilities to make function no gradient
Args:
func: input function
Returns:
no gradient function wrapper for input function
"""
def wrapper(*f_args, **f_kwargs):
with torch.no_grad():
ret = func(*f_args, **f_kwargs)
return ret
return wrapper
def make_recursive_func(func: Callable) -> Callable:
"""Convert a function into recursive style to handle nested dict/list/tuple variables
Args:
func: input function
Returns:
recursive style function
"""
def wrapper(vars):
if isinstance(vars, list):
return [wrapper(x) for x in vars]
elif isinstance(vars, tuple):
return tuple([wrapper(x) for x in vars])
elif isinstance(vars, dict):
return {k: wrapper(v) for k, v in vars.items()}
else:
return func(vars)
return wrapper
@make_recursive_func
def tensor2float(vars: Any) -> float:
"""Convert tensor to float"""
if isinstance(vars, float):
return vars
elif isinstance(vars, torch.Tensor):
return vars.data.item()
else:
raise NotImplementedError("invalid input type {} for tensor2float".format(type(vars)))
@make_recursive_func
def tensor2numpy(vars: Any) -> np.ndarray:
"""Convert tensor to numpy array"""
if isinstance(vars, np.ndarray):
return vars
elif isinstance(vars, torch.Tensor):
return vars.detach().cpu().numpy().copy()
else:
raise NotImplementedError("invalid input type {} for tensor2numpy".format(type(vars)))
@make_recursive_func
def tocuda(vars: Any) -> Union[str, torch.Tensor]:
"""Convert tensor to tensor on GPU"""
if isinstance(vars, torch.Tensor):
return vars.cpu()
elif isinstance(vars, str):
return vars
else:
raise NotImplementedError("invalid input type {} for tocuda".format(type(vars)))
def save_scalars(logger: tb.SummaryWriter, mode: str, scalar_dict: Dict[str, Any], global_step: int) -> None:
"""Log values stored in the scalar dictionary
Args:
logger: tensorboard summary writer
mode: mode name used in writing summaries
scalar_dict: python dictionary stores the key and value pairs to be recorded
global_step: step index where the logger should write
"""
scalar_dict = tensor2float(scalar_dict)
for key, value in scalar_dict.items():
if not isinstance(value, (list, tuple)):
name = "{}/{}".format(mode, key)
logger.add_scalar(name, value, global_step)
else:
for idx in range(len(value)):
name = "{}/{}_{}".format(mode, key, idx)
logger.add_scalar(name, value[idx], global_step)
def save_images(logger: tb.SummaryWriter, mode: str, images_dict: Dict[str, Any], global_step: int) -> None:
"""Log images stored in the image dictionary
Args:
logger: tensorboard summary writer
mode: mode name used in writing summaries
images_dict: python dictionary stores the key and image pairs to be recorded
global_step: step index where the logger should write
"""
images_dict = tensor2numpy(images_dict)
def preprocess(name, img):
if not (len(img.shape) == 3 or len(img.shape) == 4):
raise NotImplementedError("invalid img shape {}:{} in save_images".format(name, img.shape))
if len(img.shape) == 3:
img = img[:, np.newaxis, :, :]
img = torch.from_numpy(img[:1])
return vutils.make_grid(img, padding=0, nrow=1, normalize=True, scale_each=True)
for key, value in images_dict.items():
if not isinstance(value, (list, tuple)):
name = "{}/{}".format(mode, key)
logger.add_image(name, preprocess(name, value), global_step)
else:
for idx in range(len(value)):
name = "{}/{}_{}".format(mode, key, idx)
logger.add_image(name, preprocess(name, value[idx]), global_step)
class DictAverageMeter:
"""Wrapper class for dictionary variables that require the average value"""
def __init__(self) -> None:
"""Initialization method"""
self.data: Dict[Any, float] = {}
self.count = 0
def update(self, new_input: Dict[Any, float]) -> None:
"""Update the stored dictionary with new input data
Args:
new_input: new data to update self.data
"""
self.count += 1
if len(self.data) == 0:
for k, v in new_input.items():
if not isinstance(v, float):
raise NotImplementedError("invalid data {}: {}".format(k, type(v)))
self.data[k] = v
else:
for k, v in new_input.items():
if not isinstance(v, float):
raise NotImplementedError("invalid data {}: {}".format(k, type(v)))
self.data[k] += v
def mean(self) -> Any:
"""Return the average value of values stored in self.data"""
return {k: v / self.count for k, v in self.data.items()}
def compute_metrics_for_each_image(metric_func: Callable) -> Callable:
"""A wrapper to compute metrics for each image individually"""
def wrapper(depth_est, depth_gt, mask, *args):
batch_size = depth_gt.shape[0]
print(batch_size)
# if batch_size < BATCH_SIZE:
# break
results = []
# compute result one by one
for idx in range(batch_size):
ret = metric_func(depth_est[idx], depth_gt[idx], mask[idx], *args)
results.append(ret)
return torch.stack(results).mean()
return wrapper
@make_nograd_func
@compute_metrics_for_each_image
def Thres_metrics(
depth_est: torch.Tensor, depth_gt: torch.Tensor, mask: torch.Tensor, thres: Union[int, float]
) -> torch.Tensor:
"""Return error rate for where absolute error is larger than threshold.
Args:
depth_est: estimated depth map
depth_gt: ground truth depth map
mask: mask
thres: threshold
Returns:
error rate: error rate of the depth map
"""
# if thres is int or float, then True
assert isinstance(thres, (int, float))
depth_est, depth_gt = depth_est[mask], depth_gt[mask]
errors = torch.abs(depth_est - depth_gt)
err_mask = errors > thres
return torch.mean(err_mask.float())
# NOTE: please do not use this to build up training loss
@make_nograd_func
@compute_metrics_for_each_image
def AbsDepthError_metrics(depth_est: torch.Tensor, depth_gt: torch.Tensor, mask: torch.Tensor) -> torch.Tensor:
"""Calculate average absolute depth error
Args:
depth_est: estimated depth map
depth_gt: ground truth depth map
mask: mask
"""
depth_est, depth_gt = depth_est[mask], depth_gt[mask]
return torch.mean((depth_est - depth_gt).abs())
"""Utilities for reading and writing images, depth maps, and auxiliary data (cams, pairs) from/to disk."""
import re
import struct
import sys
from typing import Dict, List, Tuple
import cv2
import numpy as np
from PIL import Image
def scale_to_max_dim(image: np.ndarray, max_dim: int) -> Tuple[np.ndarray, int, int]:
"""Scale image to specified max dimension
Args:
image: the input image in original size
max_dim: the max dimension to scale the image down to if smaller than the actual max dimension
Returns:
Tuple of scaled image along with original image height and width
"""
original_height = image.shape[0]
original_width = image.shape[1]
scale = max_dim / max(original_height, original_width)
if 0 < scale < 1:
width = int(scale * original_width)
height = int(scale * original_height)
image = cv2.resize(image, (width, height), interpolation=cv2.INTER_LINEAR)
return image, original_height, original_width
def read_image(filename: str, max_dim: int = -1) -> Tuple[np.ndarray, int, int]:
"""Read image and rescale to specified max dimension (if exists)
Args:
filename: image input file path string
max_dim: max dimension to scale down the image; keep original size if -1
Returns:
Tuple of scaled image along with original image height and width
"""
image = Image.open(filename)
# scale 0~255 to 0~1
np_image = np.array(image, dtype=np.float32) / 255.0
return scale_to_max_dim(np_image, max_dim)
def save_image(filename: str, image: np.ndarray) -> None:
"""Save images including binary mask (bool), float (0<= val <= 1), or int (as-is)
Args:
filename: image output file path string
image: output image array
"""
if image.dtype == bool:
image = image.astype(np.uint8) * 255
elif image.dtype == np.float32 or image.dtype == np.float64:
image = image * 255
image = image.astype(np.uint8)
else:
image = image.astype(np.uint8)
Image.fromarray(image).save(filename)
def read_image_dictionary(filename: str) -> Dict[int, str]:
"""Create image dictionary from file; useful for ETH3D dataset reading and conversion.
Args:
filename: input dictionary text file path
Returns:
Dictionary of image id (int) and corresponding image file name (string)
"""
image_dict: Dict[int, str] = {}
with open(filename) as f:
num_entries = int(f.readline().strip())
for _ in range(num_entries):
parts = f.readline().strip().split(' ')
image_dict[int(parts[0].strip())] = parts[1].strip()
return image_dict
def read_cam_file(filename: str) -> Tuple[np.ndarray, np.ndarray, np.ndarray]:
"""Read camera intrinsics, extrinsics, and depth values (min, max) from text file
Args:
filename: cam text file path string
Returns:
Tuple with intrinsics matrix (3x3), extrinsics matrix (4x4), and depth params vector (min and max) if exists
"""
with open(filename) as f:
lines = [line.rstrip() for line in f.readlines()]
# extrinsics: line [1,5), 4x4 matrix
extrinsics = np.fromstring(' '.join(lines[1:5]), dtype=np.float32, sep=' ').reshape((4, 4))
# intrinsics: line [7-10), 3x3 matrix
intrinsics = np.fromstring(' '.join(lines[7:10]), dtype=np.float32, sep=' ').reshape((3, 3))
# depth min and max: line 11
if len(lines) >= 12:
depth_params = np.fromstring(lines[11], dtype=np.float32, sep=' ')
else:
depth_params = np.empty(0)
return intrinsics, extrinsics, depth_params
def read_pair_file(filename: str) -> List[Tuple[int, List[int]]]:
"""Read image pairs from text file and output a list of tuples each containing the reference image ID and a list of
source image IDs
Args:
filename: pair text file path string
Returns:
List of tuples with reference ID and list of source IDs
"""
data = []
with open(filename) as f:
num_viewpoint = int(f.readline())
for _ in range(num_viewpoint):
# ref_view = int(f.readline().rstrip())
ref_view = int(f.readline().rstrip())
# print(ref_view)
# src_views = [int(x) for x in f.readline().rstrip().split()[1::2]]
src_views = [int(x) for x in f.readline().rstrip().split()[1::2]]
# print(src_views)
view_ids = [ref_view] + src_views[:2]
# print(view_ids)
if len(src_views) != 0:
data.append((ref_view, src_views))
return data
def read_map(path: str, max_dim: int = -1) -> np.ndarray:
""" Read a binary depth map from either PFM or Colmap (bin) format determined by the file extension and also scale
the map to the max dim if given
Args:
path: input depth map file path string
max_dim: max dimension to scale down the map; keep original size if -1
Returns:
Array of depth map values
"""
if path.endswith('.bin'):
in_map = read_bin(path)
elif path.endswith('.pfm'):
in_map, _ = read_pfm(path)
else:
raise Exception('Invalid input format; only pfm and bin are supported')
return scale_to_max_dim(in_map, max_dim)[0]
def save_map(path: str, data: np.ndarray) -> None:
"""Save binary depth or confidence maps in PFM or Colmap (bin) format determined by the file extension
Args:
path: output map file path string
data: map data array
"""
if path.endswith('.bin'):
save_bin(path, data)
elif path.endswith('.pfm'):
save_pfm(path, data)
else:
raise Exception('Invalid input format; only pfm and bin are supported')
def read_bin(path: str) -> np.ndarray:
"""Read a depth map from a Colmap .bin file
Args:
path: .pfm file path string
Returns:
data: array of shape (H, W, C) representing loaded depth map
"""
with open(path, 'rb') as fid:
width, height, channels = np.genfromtxt(fid, delimiter='&', max_rows=1,
usecols=(0, 1, 2), dtype=int)
fid.seek(0)
num_delimiter = 0
byte = fid.read(1)
while True:
if byte == b'&':
num_delimiter += 1
if num_delimiter >= 3:
break
byte = fid.read(1)
data = np.fromfile(fid, np.float32)
data = data.reshape((width, height, channels), order='F')
data = np.transpose(data, (1, 0, 2))
return data
def save_bin(filename: str, data: np.ndarray):
"""Save a depth map to a Colmap .bin file
Args:
filename: output .pfm file path string,
data: depth map to save, of shape (H,W) or (H,W,C)
"""
if data.dtype != np.float32:
raise Exception('Image data type must be float32.')
if len(data.shape) == 2:
height, width = data.shape
channels = 1
elif len(data.shape) == 3 and (data.shape[2] == 3 or data.shape[2] == 1):
height, width, channels = data.shape
else:
raise Exception('Image must have H x W x 3, H x W x 1 or H x W dimensions.')
with open(filename, 'w') as fid:
fid.write(str(width) + '&' + str(height) + '&' + str(channels) + '&')
with open(filename, 'ab') as fid:
if len(data.shape) == 2:
image_trans = np.transpose(data, (1, 0))
else:
image_trans = np.transpose(data, (1, 0, 2))
data_1d = image_trans.reshape(-1, order='F')
data_list = data_1d.tolist()
endian_character = '<'
format_char_sequence = ''.join(['f'] * len(data_list))
byte_data = struct.pack(endian_character + format_char_sequence, *data_list)
fid.write(byte_data)
def read_pfm(filename: str) -> Tuple[np.ndarray, float]:
"""Read a depth map from a .pfm file
Args:
filename: .pfm file path string
Returns:
data: array of shape (H, W, C) representing loaded depth map
scale: float to recover actual depth map pixel values
"""
file = open(filename, "rb") # treat as binary and read-only
header = file.readline().decode("utf-8").rstrip()
if header == "PF":
color = True
elif header == "Pf": # depth is Pf
color = False
else:
raise Exception("Not a PFM file.")
dim_match = re.match(r"^(\d+)\s(\d+)\s$", file.readline().decode("utf-8"))
if dim_match:
width, height = map(int, dim_match.groups())
else:
raise Exception("Malformed PFM header.")
scale = float(file.readline().rstrip())
if scale < 0: # little-endian
endian = "<"
scale = -scale
else:
endian = ">" # big-endian
data = np.fromfile(file, endian + "f")
shape = (height, width, 3) if color else (height, width, 1)
data = np.reshape(data, shape)
data = np.flipud(data)
file.close()
return data, scale
def save_pfm(filename: str, image: np.ndarray, scale: float = 1) -> None:
"""Save a depth map to a .pfm file
Args:
filename: output .pfm file path string,
image: depth map to save, of shape (H,W) or (H,W,C)
scale: scale parameter to save
"""
file = open(filename, "wb")
color = None
image = np.flipud(image)
if image.dtype.name != "float32":
raise Exception("Image dtype must be float32.")
if len(image.shape) == 3 and image.shape[2] == 3: # color image
color = True
elif len(image.shape) == 2 or len(image.shape) == 3 and image.shape[2] == 1: # greyscale
color = False
else:
raise Exception("Image must have H x W x 3, H x W x 1 or H x W dimensions.")
file.write("PF\n".encode("utf-8") if color else "Pf\n".encode("utf-8"))
file.write("{} {}\n".format(image.shape[1], image.shape[0]).encode("utf-8"))
endian = image.dtype.byteorder
if endian == "<" or endian == "=" and sys.byteorder == "little":
scale = -scale
file.write(("%f\n" % scale).encode("utf-8"))
image.tofile(file)
file.close()
import argparse
import os
import torch.nn as nn
import torch.nn.parallel
import torch.backends.cudnn as cudnn
from torch.utils.data import DataLoader
import time
# from datasets import find_dataset_def
# from models import *
# from utils import *
import sys
# from datasets.data_io import read_cam_file, read_pair_file, read_image, read_map, save_image, save_map
import cv2
from plyfile import PlyData, PlyElement
cudnn.benchmark = True
parser = argparse.ArgumentParser(description='Predict depth, filter, and fuse')
parser.add_argument('--model', default='PatchmatchNet', help='select model')
parser.add_argument('--dataset', default='eth3d', help='select dataset')
parser.add_argument('--testpath', help='testing data path')
parser.add_argument('--testlist', help='testing scan list')
parser.add_argument('--split', default='test', help='select data')
parser.add_argument('--batch_size', type=int, default=1, help='testing batch size')
parser.add_argument('--n_views', type=int, default=5, help='num of view')
parser.add_argument('--loadckpt', default=None, help='load a specific checkpoint')
parser.add_argument('--outdir', default='./outputs', help='output dir')
parser.add_argument('--display', action='store_true', help='display depth images and masks')
parser.add_argument('--patchmatch_iteration', nargs='+', type=int, default=[1, 2, 2],
help='num of iteration of patchmatch on stages 1,2,3')
parser.add_argument('--patchmatch_num_sample', nargs='+', type=int, default=[8, 8, 16],
help='num of generated samples in local perturbation on stages 1,2,3')
parser.add_argument('--patchmatch_interval_scale', nargs='+', type=float, default=[0.005, 0.0125, 0.025],
help='normalized interval in inverse depth range to generate samples in local perturbation')
parser.add_argument('--patchmatch_range', nargs='+', type=int, default=[6, 4, 2],
help='fixed offset of sampling points for propogation of patchmatch on stages 1,2,3')
parser.add_argument('--propagate_neighbors', nargs='+', type=int, default=[0, 8, 16],
help='num of neighbors for adaptive propagation on stages 1,2,3')
parser.add_argument('--evaluate_neighbors', nargs='+', type=int, default=[9, 9, 9],
help='num of neighbors for adaptive matching cost aggregation of adaptive evaluation on stages 1,2,3')
parser.add_argument('--geo_pixel_thres', type=float, default=1,
help='pixel threshold for geometric consistency filtering')
parser.add_argument('--geo_depth_thres', type=float, default=0.01,
help='depth threshold for geometric consistency filtering')
parser.add_argument('--photo_thres', type=float, default=0.8, help='threshold for photometric consistency filtering')
# parse arguments and check
args = parser.parse_args()
print("argv:", sys.argv[1:])
print_args(args)
# run MVS model to save depth maps
def save_depth():
# dataset, dataloader
mvs_dataset = find_dataset_def(args.dataset)
test_dataset = mvs_dataset(args.testpath, args.n_views)
image_loader = DataLoader(test_dataset, args.batch_size, shuffle=False, num_workers=4, drop_last=False)
# image_loader = DataLoader(test_dataset, args.batch_size, shuffle=False, drop_last=False)
# model
model = PatchmatchNet(
patchmatch_interval_scale=args.patchmatch_interval_scale,
propagation_range=args.patchmatch_range,
patchmatch_iteration=args.patchmatch_iteration,
patchmatch_num_sample=args.patchmatch_num_sample,
propagate_neighbors=args.propagate_neighbors,
evaluate_neighbors=args.evaluate_neighbors
)
model = nn.DataParallel(model)
model.cpu()
# load checkpoint file specified by args.loadckpt
print("loading model {}".format(args.loadckpt))
state_dict = torch.load(args.loadckpt,map_location=torch.device('cpu'))
model.load_state_dict(state_dict['model'], strict=False)
model.eval()
with torch.no_grad():
for batch_idx, sample in enumerate(image_loader):
# print(batch_idx)
start_time = time.time()
sample_cuda = tocuda(sample)
refined_depth, confidence, _ = model(sample_cuda["imgs"], sample_cuda["proj_matrices"],
sample_cuda["depth_min"], sample_cuda["depth_max"])
refined_depth = tensor2numpy(refined_depth)
confidence = tensor2numpy(confidence)
del sample_cuda
print('Iter {}/{}, time = {:.3f}'.format(batch_idx, len(image_loader), time.time() - start_time))
filenames = sample["filename"]
# save depth maps and confidence maps
for filename, depth_est, photometric_confidence in zip(filenames, refined_depth, confidence):
depth_filename = os.path.join(args.outdir, filename.format('depth_est', '.pfm'))
confidence_filename = os.path.join(args.outdir, filename.format('confidence', '.pfm'))
os.makedirs(depth_filename.rsplit('/', 1)[0], exist_ok=True)
os.makedirs(confidence_filename.rsplit('/', 1)[0], exist_ok=True)
# save depth maps
depth_est = np.squeeze(depth_est, 0)
save_map(depth_filename, depth_est)
# save confidence maps
save_map(confidence_filename, photometric_confidence)
# project the reference point cloud into the source view, then project back
def reproject_with_depth(depth_ref, intrinsics_ref, extrinsics_ref, depth_src, intrinsics_src, extrinsics_src):
width, height = depth_ref.shape[1], depth_ref.shape[0]
# step1. project reference pixels to the source view
# reference view x, y
x_ref, y_ref = np.meshgrid(np.arange(0, width), np.arange(0, height))
x_ref, y_ref = x_ref.reshape([-1]), y_ref.reshape([-1])
# reference 3D space
xyz_ref = np.matmul(np.linalg.inv(intrinsics_ref),
np.vstack((x_ref, y_ref, np.ones_like(x_ref))) * depth_ref.reshape([-1]))
# source 3D space
xyz_src = np.matmul(np.matmul(extrinsics_src, np.linalg.inv(extrinsics_ref)),
np.vstack((xyz_ref, np.ones_like(x_ref))))[:3]
# source view x, y
k_xyz_src = np.matmul(intrinsics_src, xyz_src)
xy_src = k_xyz_src[:2] / k_xyz_src[2:3]
# step2. reproject the source view points with source view depth estimation
# find the depth estimation of the source view
x_src = xy_src[0].reshape([height, width]).astype(np.float32)
y_src = xy_src[1].reshape([height, width]).astype(np.float32)
sampled_depth_src = cv2.remap(depth_src, x_src, y_src, interpolation=cv2.INTER_LINEAR)
# mask = sampled_depth_src > 0
# source 3D space
# NOTE that we should use sampled source-view depth_here to project back
xyz_src = np.matmul(np.linalg.inv(intrinsics_src),
np.vstack((xy_src, np.ones_like(x_ref))) * sampled_depth_src.reshape([-1]))
# reference 3D space
xyz_reprojected = np.matmul(np.matmul(extrinsics_ref, np.linalg.inv(extrinsics_src)),
np.vstack((xyz_src, np.ones_like(x_ref))))[:3]
# source view x, y, depth
depth_reprojected = xyz_reprojected[2].reshape([height, width]).astype(np.float32)
k_xyz_reprojected = np.matmul(intrinsics_ref, xyz_reprojected)
xy_reprojected = k_xyz_reprojected[:2] / k_xyz_reprojected[2:3]
x_reprojected = xy_reprojected[0].reshape([height, width]).astype(np.float32)
y_reprojected = xy_reprojected[1].reshape([height, width]).astype(np.float32)
return depth_reprojected, x_reprojected, y_reprojected, x_src, y_src
def check_geometric_consistency(depth_ref, intrinsics_ref, extrinsics_ref, depth_src, intrinsics_src, extrinsics_src,
geo_pixel_thres, geo_depth_thres):
width, height = depth_ref.shape[1], depth_ref.shape[0]
x_ref, y_ref = np.meshgrid(np.arange(0, width), np.arange(0, height))
depth_reprojected, x2d_reprojected, y2d_reprojected, x2d_src, y2d_src = reproject_with_depth(
depth_ref, intrinsics_ref, extrinsics_ref, depth_src, intrinsics_src, extrinsics_src)
# print(depth_ref.shape)
# print(depth_reprojected.shape)
# check |p_reproj-p_1| < 1
dist = np.sqrt((x2d_reprojected - x_ref) ** 2 + (y2d_reprojected - y_ref) ** 2)
# check |d_reproj-d_1| / d_1 < 0.01
# depth_ref = np.squeeze(depth_ref, 2)
depth_diff = np.abs(depth_reprojected - depth_ref)
relative_depth_diff = depth_diff / depth_ref
mask = np.logical_and(dist < geo_pixel_thres, relative_depth_diff < geo_depth_thres)
depth_reprojected[~mask] = 0
return mask, depth_reprojected, x2d_src, y2d_src
def filter_depth(
scan_folder, out_folder, plyfilename, geo_pixel_thres, geo_depth_thres, photo_thres, img_wh, geo_mask_thres):
# the pair file
pair_file = os.path.join(scan_folder, "pair.txt")
# for the final point cloud
vertexs = []
vertex_colors = []
pair_data = read_pair_file(pair_file)
# for each reference view and the corresponding source views
for ref_view, src_views in pair_data:
# load the reference image
ref_img, original_h, original_w = read_image(
os.path.join(scan_folder, 'images/{:0>8}.jpg'.format(ref_view)), max(img_wh))
ref_intrinsics, ref_extrinsics, _ = read_cam_file(
os.path.join(scan_folder, 'cams/{:0>8}_cam.txt'.format(ref_view)))[0:2]
# print([ref_intrinsics,ref_extrinsics])
ref_intrinsics[0] *= img_wh[0]/original_w
ref_intrinsics[1] *= img_wh[1]/original_h
# load the estimated depth of the reference view
ref_depth_est = read_map(os.path.join(out_folder, 'depth_est/{:0>8}.pfm'.format(ref_view)))
ref_depth_est = np.squeeze(ref_depth_est, 2)
# load the photometric mask of the reference view
confidence = read_map(os.path.join(out_folder, 'confidence/{:0>8}.pfm'.format(ref_view)))
photo_mask = confidence > photo_thres
photo_mask = np.squeeze(photo_mask, 2)
all_srcview_depth_ests = []
# compute the geometric mask
geo_mask_sum = 0
for src_view in src_views:
# camera parameters of the source view
_, original_h, original_w = read_image(
os.path.join(scan_folder, 'images/{:0>8}.jpg'.format(src_view)), max(img_wh))
src_intrinsics, src_extrinsics, _ = read_cam_file(
os.path.join(scan_folder, 'cams/{:0>8}_cam.txt'.format(src_view)))[0:2]
src_intrinsics[0] *= img_wh[0]/original_w
src_intrinsics[1] *= img_wh[1]/original_h
# the estimated depth of the source view
src_depth_est = read_map(os.path.join(out_folder, 'depth_est/{:0>8}.pfm'.format(src_view)))
geo_mask, depth_reprojected, _, _ = check_geometric_consistency(
ref_depth_est, ref_intrinsics, ref_extrinsics, src_depth_est, src_intrinsics, src_extrinsics,
geo_pixel_thres, geo_depth_thres)
geo_mask_sum += geo_mask.astype(np.int32)
all_srcview_depth_ests.append(depth_reprojected)
depth_est_averaged = (sum(all_srcview_depth_ests) + ref_depth_est) / (geo_mask_sum + 1)
geo_mask = geo_mask_sum >= geo_mask_thres
final_mask = np.logical_and(photo_mask, geo_mask)
os.makedirs(os.path.join(out_folder, "mask"), exist_ok=True)
save_image(os.path.join(out_folder, "mask/{:0>8}_photo.png".format(ref_view)), photo_mask)
save_image(os.path.join(out_folder, "mask/{:0>8}_geo.png".format(ref_view)), geo_mask)
save_image(os.path.join(out_folder, "mask/{:0>8}_final.png".format(ref_view)), final_mask)
print("processing {}, ref-view{:0>2}, geo_mask:{:3f} photo_mask:{:3f} final_mask: {:3f}".format(
scan_folder, ref_view, geo_mask.mean(), photo_mask.mean(), final_mask.mean()))
if args.display:
cv2.imshow('ref_img', ref_img[:, :, ::-1])
cv2.imshow('ref_depth', ref_depth_est)
cv2.imshow('ref_depth * photo_mask', ref_depth_est * photo_mask.astype(np.float32))
cv2.imshow('ref_depth * geo_mask', ref_depth_est * geo_mask.astype(np.float32))
cv2.imshow('ref_depth * mask', ref_depth_est * final_mask.astype(np.float32))
cv2.waitKey(1)
height, width = depth_est_averaged.shape[:2]
x, y = np.meshgrid(np.arange(0, width), np.arange(0, height))
valid_points = final_mask
x, y, depth = x[valid_points], y[valid_points], depth_est_averaged[valid_points]
color = ref_img[valid_points]
xyz_ref = np.matmul(np.linalg.inv(ref_intrinsics), np.vstack((x, y, np.ones_like(x))) * depth)
xyz_world = np.matmul(np.linalg.inv(ref_extrinsics), np.vstack((xyz_ref, np.ones_like(x))))[:3]
vertexs.append(xyz_world.transpose((1, 0)))
vertex_colors.append((color * 255).astype(np.uint8))
vertexs = np.concatenate(vertexs, axis=0)
vertex_colors = np.concatenate(vertex_colors, axis=0)
vertexs = np.array([tuple(v) for v in vertexs], dtype=[('x', 'f4'), ('y', 'f4'), ('z', 'f4')])
vertex_colors = np.array([tuple(v) for v in vertex_colors], dtype=[('red', 'u1'), ('green', 'u1'), ('blue', 'u1')])
vertex_all = np.empty(len(vertexs), vertexs.dtype.descr + vertex_colors.dtype.descr)
for prop in vertexs.dtype.names:
vertex_all[prop] = vertexs[prop]
for prop in vertex_colors.dtype.names:
vertex_all[prop] = vertex_colors[prop]
el = PlyElement.describe(vertex_all, 'vertex')
PlyData([el]).write(plyfilename)
print("saving the final model to", plyfilename)
if __name__ == '__main__':
# step1. save all the depth maps and the masks in outputs directory
save_depth()
# the size of image input for PatchmatchNet, maybe downsampled
img_wh = (640, 480)
# number of source images need to be consistent with in geometric consistency filtering
geo_mask_thres = 2
# step2. filter saved depth maps and reconstruct point cloud
filter_depth(args.testpath, args.outdir, os.path.join(args.outdir, 'custom.ply'), args.geo_pixel_thres,
args.geo_depth_thres, args.photo_thres, img_wh, geo_mask_thres)
```
| github_jupyter |
<img align="left" src="https://lever-client-logos.s3.amazonaws.com/864372b1-534c-480e-acd5-9711f850815c-1524247202159.png" width=200>
<br></br>
<br></br>
## *Data Science Unit 4 Sprint 3 Lesson 1*
# Recurrent Neural Networks and Long Short Term Memory (LSTM)
## _aka_ PREDICTING THE FUTURE!
<img src="https://media.giphy.com/media/l2JJu8U8SoHhQEnoQ/giphy.gif" width=480 height=356>
<br></br>
<br></br>
> "Yesterday's just a memory - tomorrow is never what it's supposed to be." -- Bob Dylan
Wish you could save [Time In A Bottle](https://www.youtube.com/watch?v=AnWWj6xOleY)? With statistics you can do the next best thing - understand how data varies over time (or any sequential order), and use the order/time dimension predictively.
A sequence is just any enumerated collection - order counts, and repetition is allowed. Python lists are a good elemental example - `[1, 2, 2, -1]` is a valid list, and is different from `[1, 2, -1, 2]`. The data structures we tend to use (e.g. NumPy arrays) are often built on this fundamental structure.
A time series is data where you have not just the order but some actual continuous marker for where they lie "in time" - this could be a date, a timestamp, [Unix time](https://en.wikipedia.org/wiki/Unix_time), or something else. All time series are also sequences, and for some techniques you may just consider their order and not "how far apart" the entries are (if you have particularly consistent data collected at regular intervals it may not matter).
## Recurrent Neural Networks
There's plenty more to "traditional" time series, but the latest and greatest technique for sequence data is recurrent neural networks. A recurrence relation in math is an equation that uses recursion to define a sequence - a famous example is the Fibonacci numbers:
$F_n = F_{n-1} + F_{n-2}$
For formal math you also need a base case $F_0=1, F_1=1$, and then the rest builds from there. But for neural networks what we're really talking about are loops:

The hidden layers have edges (output) going back to their own input - this loop means that for any time `t` the training is at least partly based on the output from time `t-1`. The entire network is being represented on the left, and you can unfold the network explicitly to see how it behaves at any given `t`.
Different units can have this "loop", but a particularly successful one is the long short-term memory unit (LSTM):

There's a lot going on here - in a nutshell, the calculus still works out and backpropagation can still be implemented. The advantage (ane namesake) of LSTM is that it can generally put more weight on recent (short-term) events while not completely losing older (long-term) information.
After enough iterations, a typical neural network will start calculating prior gradients that are so small they effectively become zero - this is the [vanishing gradient problem](https://en.wikipedia.org/wiki/Vanishing_gradient_problem), and is what RNN with LSTM addresses. Pay special attention to the $c_t$ parameters and how they pass through the unit to get an intuition for how this problem is solved.
So why are these cool? One particularly compelling application is actually not time series but language modeling - language is inherently ordered data (letters/words go one after another, and the order *matters*). [The Unreasonable Effectiveness of Recurrent Neural Networks](https://karpathy.github.io/2015/05/21/rnn-effectiveness/) is a famous and worth reading blog post on this topic.
For our purposes, let's use TensorFlow and Keras to train RNNs with natural language. Resources:
- https://github.com/keras-team/keras/blob/master/examples/imdb_lstm.py
- https://keras.io/layers/recurrent/#lstm
- http://adventuresinmachinelearning.com/keras-lstm-tutorial/
Note that `tensorflow.contrib` [also has an implementation of RNN/LSTM](https://www.tensorflow.org/tutorials/sequences/recurrent).
### RNN/LSTM Sentiment Classification with Keras
```
'''
#Trains an LSTM model on the IMDB sentiment classification task.
The dataset is actually too small for LSTM to be of any advantage
compared to simpler, much faster methods such as TF-IDF + LogReg.
**Notes**
- RNNs are tricky. Choice of batch size is important,
choice of loss and optimizer is critical, etc.
Some configurations won't converge.
- LSTM loss decrease patterns during training can be quite different
from what you see with CNNs/MLPs/etc.
'''
from __future__ import print_function
from tensorflow.keras.preprocessing import sequence
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Embedding
from tensorflow.keras.layers import LSTM
from tensorflow.keras.datasets import imdb
# adjust these to faster/more stable running
max_features = 20000
# cut texts after this number of words (among top max_features most common words)
#number of lines...
maxlen = 80
# ?
batch_size = 32
print('Loading data...')
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=max_features)
print(len(x_train), 'train sequences')
print(len(x_test), 'test sequences')
len(x_train)
print('Pad sequences (samples x time)')
x_train = sequence.pad_sequences(x_train, maxlen=maxlen)
x_test = sequence.pad_sequences(x_test, maxlen=maxlen)
print('x_train shape:', x_train.shape)
print('x_test shape:', x_test.shape)
# these have been encoded
x_train[0]
print('Build model...')
model = Sequential()
model.add(Embedding(max_features, 128))
model.add(LSTM(128, dropout=0.2, recurrent_dropout=0.2))
model.add(Dense(1, activation='sigmoid'))
# try using different optimizers and different optimizer configs
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
print('Train...')
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=1,
validation_data=(x_test, y_test))
score, acc = model.evaluate(x_test, y_test,
batch_size=batch_size)
print('Test score:', score)
print('Test accuracy:', acc)
```
### LSTM Text generation with Keras (bottom up) Generate text...
What else can we do with LSTMs? Since we're analyzing the *sequence*, we can do more than classify - we can *generate* text. I'ved pulled some news stories using [newspaper](https://github.com/codelucas/newspaper/).
This example is drawn from the Keras [documentation](https://keras.io/examples/lstm_text_generation/).
```
from tensorflow.keras.callbacks import LambdaCallback
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, LSTM
from tensorflow.keras.optimizers import RMSprop
import numpy as np
import random
import sys
import os
# produces: type=list
# this is the notation for subdirectory
data_files = os.listdir('./articles')
# Inspecting
#data_files
# for sagemaker...
# directed at current working directory
# anything generally local...anything uploaded
#data_files = os.listdir(os.curdir)
#data_files
```
['LS_DS_431_RNN_and_LSTM_Assignment.ipynb',
'articles',
'.ipynb_checkpoints',
'LS_DS_431_RNN_and_LSTM_Lecture.ipynb']
```
# we need to do some cleaning
# import everything, cycle though files, add to end of
# of text string
text = " "
for filename in data_files:
# this checks the filename
if filename[-3:] == 'txt':
path = f'{filename}'
# modified with different directory?
#with open(path, 'r') as data:
with open(os.path.abspath('./articles/')) as data:
# reads data object and adds to content object
content = data.read()
#updates text string
text = text + " " + content
print ('corpus length', len(text))
# Read in Data
# Encode Data as Chars
# Create the Sequence Data
# Specify x & y
# build the model: a single LSTM
def sample(preds, temperature=1.0):
# helper function to sample an index from a probability array
preds = np.asarray(preds).astype('float64')
preds = np.log(preds) / temperature
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
probas = np.random.multinomial(1, preds, 1)
return np.argmax(probas)
def on_epoch_end(epoch, _):
# Function invoked at end of each epoch. Prints generated text.
print()
print('----- Generating text after Epoch: %d' % epoch)
start_index = random.randint(0, len(text) - maxlen - 1)
for diversity in [0.2, 0.5, 1.0, 1.2]:
print('----- diversity:', diversity)
generated = ''
sentence = text[start_index: start_index + maxlen]
generated += sentence
print('----- Generating with seed: "' + sentence + '"')
sys.stdout.write(generated)
for i in range(400):
x_pred = np.zeros((1, maxlen, len(chars)))
for t, char in enumerate(sentence):
x_pred[0, t, char_indices[char]] = 1.
preds = model.predict(x_pred, verbose=0)[0]
next_index = sample(preds, diversity)
next_char = indices_char[next_index]
sentence = sentence[1:] + next_char
sys.stdout.write(next_char)
sys.stdout.flush()
print()
print_callback = LambdaCallback(on_epoch_end=on_epoch_end)
model.fit(x, y,
batch_size=128,
epochs=5,
callbacks=[print_callback])
```
| github_jupyter |
```
import sys
sys.path.append('..')
import torch
import numpy as np
import matplotlib.pyplot as plt
from lens import logic
torch.manual_seed(0)
np.random.seed(0)
# XOR problem
x_train = torch.tensor([
[0, 0],
[0, 1],
[1, 0],
[1, 1],
], dtype=torch.float)
y_train = torch.tensor([0, 1, 1, 0], dtype=torch.float).unsqueeze(1)
x_test = torch.tensor([
[0, 0.95],
[0, 0.9],
[0.05, 1],
[0.1, 0.8],
[0.45, 1],
[0, 0.35],
[0.95, 0.9],
[0.75, 0.2],
[0.75, 0.15],
], dtype=torch.float)
y_test = torch.tensor([1, 1, 1, 1, 1, 0, 0, 1, 1], dtype=torch.float).unsqueeze(1)
layers = [
torch.nn.Linear(x_train.size(1), 10),
torch.nn.LeakyReLU(),
torch.nn.Linear(10, 4),
torch.nn.LeakyReLU(),
torch.nn.Linear(4, 1),
torch.nn.Sigmoid(),
]
model = torch.nn.Sequential(*layers)
optimizer = torch.optim.Adam(model.parameters(), lr=0.01)
model.train()
need_pruning = True
for epoch in range(1000):
# forward pass
optimizer.zero_grad()
y_pred = model(x_train)
# Compute Loss
loss = torch.nn.functional.mse_loss(y_pred, y_train)
for module in model.children():
if isinstance(module, torch.nn.Linear):
loss += 0.001 * torch.norm(module.weight, 1)
# backward pass
loss.backward()
optimizer.step()
# compute accuracy
if epoch % 100 == 0:
y_pred_d = (y_pred > 0.5)
accuracy = (y_pred_d.eq(y_train).sum(dim=1) == y_train.size(1)).sum().item() / y_train.size(0)
print(f'Epoch {epoch}: train accuracy: {accuracy:.4f}')
```
# Decision boundaries
```
def plot_decision_bundaries(model, x, h=0.1, cmap='BrBG'):
x1_min, x1_max = x[:, 0].min() - 1, x[:, 0].max() + 1
x2_min, x2_max = x[:, 1].min() - 1, x[:, 1].max() + 1
xx1, xx2 = np.meshgrid(np.arange(x1_min, x1_max, h),
np.arange(x2_min, x2_max, h))
xx = torch.FloatTensor(np.c_[xx1.ravel(), xx2.ravel()])
Z = model(xx).detach().numpy()
Z = Z.reshape(xx1.shape)
plt.contourf(xx1, xx2, Z, alpha=0.2, cmap=cmap)
return
cmap = 'BrBG'
plt.figure(figsize=[8, 8])
for sample_id, (xin, yin) in enumerate(zip(x_train, y_train)):
output = model(xin)
explanation = logic.relu_nn.explain_local(model, x_train, y_train,
xin, yin, method='lime',
concept_names=['f1', 'f2'])
plt.subplot(2, 2, sample_id+1)
plt.title(f'INPUT={xin.detach().numpy()} - OUTPUT={output.detach().numpy()} \n Explanation: {explanation}')
plot_decision_bundaries(model, x_train, h=0.01)
plt.scatter(x_train[:, 0].detach().numpy(), x_train[:, 1].detach().numpy(), c=y_train.detach().numpy(), cmap=cmap)
plt.scatter(xin[0], xin[1], c='k', marker='x', s=100, cmap=cmap)
c = plt.Circle((xin[0], xin[1]), radius=0.2, edgecolor='k', fill=False, linestyle='--')
plt.gca().add_artist(c)
plt.xlim([-0.5, 1.5])
plt.ylim([-0.5, 1.5])
plt.tight_layout()
plt.show()
```
# Combine local explanations
```
for i, target_class in enumerate(range(2)):
global_explanation, _, _ = logic.relu_nn.combine_local_explanations(model, x_train,
y_train.squeeze(),
target_class=target_class,
method='lime')
accuracy, preds = logic.base.test_explanation(global_explanation, target_class, x_test, y_test)
final_formula = logic.base.replace_names(global_explanation, ['f1', 'f2'])
print(f'Class {target_class} - Global explanation: "{final_formula}" - Accuracy: {accuracy:.4f}')
```
| github_jupyter |

#Ejercicio: Algoritmo genético para optimizar un rotor o hélice, paso a paso
##El problema
A menudo, en ingeniería, cuando nos enfrentamos a un problema, no podemos resolver directamente o despejar la solución como en los problemas sencillos típicos de matemáticas o física clásica. Una manera muy típica en la que nos encontraremos los problemas es en la forma de simulación: tenemos una serie de parámetros y un modelo, y podemos simularlo para obtener sus características, pero sin tener ninguna fórmula explícita que relacione parámetros y resultados y que nos permita obtener una función inversa.
En este ejercicio, nos plantearemos un problema de ese tipo: tenemos una función que calcula las propiedades de una hélice en función de una serie de parámetros, pero no conocemos los cálculos que hace internamente. Para nosotros, es una caja negra.
Para optimizar, iremos recuperando las funciones del algoritmo genético que se vieron en la parte de teoría.
```
%matplotlib inline
import numpy as np # Trabajaremos con arrays
import matplotlib.pyplot as plt # Y vamos a pintar gráficos
from optrot.rotor import calcular_rotor # Esta función es la que vamos a usar para calcular el rotor
import random as random # Necesitaremos números aleatorios
```
Empecemos echando un ojo a la función del rotor, para ver qué vamos a necesitar y con qué parámetros vamos a trabajar.
```
help(calcular_rotor)
```
Podemos trazar unas cuantas curvas para observar qué pinta va a tener lo que saquemos. Por ejemplo, cómo cambian las características de la hélice dependiendo de la velocidad de vuelo, para una hélice de ejemplo que gira a uyna velocidad dada.
```
vel = np.linspace(0, 30, 100)
efic = np.zeros_like(vel)
T = np.zeros_like(vel)
P = np.zeros_like(vel)
mach = np.zeros_like(vel)
for i in range(len(vel)):
T[i], P[i], efic[i], mach[i] = calcular_rotor(130, vel[i], 0.5, 3)
plt.plot(vel, T)
plt.title('Tracción de la hélice')
plt.plot(vel, P)
plt.title('Potencia consumida')
plt.plot(vel, efic)
plt.title('Eficiencia de la hélice')
plt.plot(vel, mach)
plt.title('Mach en la punta de las palas')
```
##Definiendo el genoma
Definamos un individuo genérico: Cada individuo será un posible diseño del rotor, con unas características determinadas.
```
class Individual (object):
def __init__(self, genome):
self.genome = genome
self.traits = {}
self.performances = {}
self.fitness = 0
```
Nuestro rotor depende de varios parámetros, pero en general, buscaremos optimizar el valor de unos, mateniendo un valor controlado de otros. Por ejemplo, la velocidad de avance y la altitud normalmente las impondremos, ya que querremos optimizar para una velocidad y altura de vuelos dadas.
En nuestro algoritmo, usaremos como genoma los parámetros de optimización, y las variables circunstanciales las controlaremos a mano.
***Sugerencia*** (esta es una manera de organizar las variables, aunque puedes escoger otras)
Parámetros de optimización:
- omega (velocidad de rotación) (Entre 0 y 200 radianes/segundo)
- R (radio de la hélice) (Entre 0.1 y 2 metros)
- b (número de palas) (Entre 2 y 5 palas)
- theta0 (ángulo de paso colectivo) (Entre -0.26 y 0.26 radianes)(*se corresponde a -15 y 15 grados*)
- p (parámetro de torsión) (Entre -5 y 20 grados)
- cuerda (anchura de la pala) (Entre 0.01 y 0.2 metros)
Parámetros circunstanciales:
- vz (velocidad de vuelo)
- h (altura de vuelo)
Variables que se van a mantener
- ley de torsión (hiperbólica)
- formato de chord params: un solo número, para que la anchura sea constante a lo largo de la pala
```
15 * np.pi / 180
```
A continuación crearemos un diccionario de genes. En él iremos almacenando los nombres de los parámetros y la cantidad de bits que usaremos para definirlos. Cuantos más bits, más resolución
Ej: 1 bit : 2 valores, 2 bit : 4 valores, 10 bit : 1024 valores
```
#Completa este diccionario con las variables que hayas elegido y los bits que usarás
dict_genes = {
'omega' : 10,
'R': 10,
'b': 2
}
```
Ahora, crearemos una función que rellene estos genomas con datos aleatorios:
```
def generate_genome (dict_genes):
#Calculamos el número total de bits con un bucle que recorra el diccionario
n_bits = ?
#Generamos un array aletorio de 1 y 0 de esa longitud con numpy
genome = np.random.randint(0, 2, nbits)
#Transformamos el array en una lista antes de devolverlo
return list(genome)
# Podemos probar a usar nuestra función, para ver qué pinta tiene el ADN de un rotor:
generate_genome(dict_genes)
```
##Trabajando con el individuo
Ahora necesitamos una función que transforme esos genes a valores con sentido. Cada gen es un número binario cuyo valor estará entre 0 y 2 ^ n, siendo n el número de bits que hayamos escogido. Estas variables traducidas las guardaremos en otro diccionario, ya con su valor. Estos genes no están volando por ahí sueltos, sino que estarán guardados en el interior del individuo al que pertenezcan, por lo que la función deberá estar preparada para extraerlos del individuo, y guardar los resultados a su vez en el interior del individuo.
```
def calculate_traits (individual, dict_genes):
genome = individual.genome
integer_temporal_list = []
for gen in dict_genes: #Recorremos el diccionario de genes para ir traduciendo del binario
??? #Buscamos los bits que se corresponden al bit en cuestión
??? #Pasamos de lista binaria a número entero
integer_temporal_list.append(??) #Añadimos el entero a la lista
# Transformamos cada entero en una variable con sentido físico:
# Por ejemplo, si el entero de la variable Omega está entre 0 y 1023 (10bits),
# pero la variable Omega real estará entre 0 y 200 radianes por segundo:
omega = integer_temporal_list[0] * 200 / 1023
#del mismo modo, para R:
R = 0.1 + integer_temporal_list[1] * 1.9 / 1023 #Obtendremos un radio entre 0.1 y 2 metros
#El número de palas debe ser un entero, hay que tener cuidado:
b = integer_temporal_list[2] + 2 #(entre 2 y 5 palas)
#Continúa con el resto de variables que hayas elegido!
dict_traits = { #Aquí iremos guardando los traits, o parámetros
'omega' : omega,
'R': R
}
individual.traits = dict_traits #Por último, guardamos los traits en el individuo
```
El siguiente paso es usar estos traits(parámetros) para calcular las performances (características o desempeños) del motor. Aquí es donde entra el modelo del motor propiamente dicho.
```
def calculate_performances (individual):
dict_traits = individual.traits
#Nuestras circunstancias las podemos imponer aquí, o irlas pasando como argumento a la función
h = 2000 #Altitud de vuelo en metros
vz = 70 #velocidad de avance en m/s, unos 250 km/h
#Extraemos los traits del diccionario:
omega = dict_traits['omega']
R = dict_traits['R']
#... etc
T, P, efic, mach_punta = calcular_rotor(omega, vz, R, b, h...) #Introduce las variables que uses de parámetro.
# Consulta la ayuda para asegurarte de que usas el
# formato correcto!
dict_perfo = {
'T' : T, #Tracción de la hélice
'P' : P, #Potencia consumida por la hélice
'efic': efic, #Eficiencia propulsiva de la hélice
'mach_punta': mach_punta #Mach en la punta de las palas
}
individual.performances = dict_perfo
```
Comprobemos si todo funciona!
```
individuo = Individual(generate_genome(dict_genes))
calculate_traits(individuo, dict_genes)
calculate_performances(individuo)
print(individuo.traits)
print(individuo.performances)
```
El último paso que tenemos que realizar sobre el individuo es uno de los más críticos: Transformar las performances en un valor único (fitness) que con exprese cómo de bueno es con respecto al objetivo de optimización. La función de fitness puede ser función de parámetros(traits) y performances, dependiendo de qué queramos optimizar.
Por ejemplo, si buscáramos que tuviera la tracción máxima sin preocuparnos de nada más, el valor de fitnes sería simplemente igual al de T:
fitness = T
Si queremos imponer restricciones, por ejemplo, que la potencia sea menor a 1000 watios, se pueden añadir sentencias del tipo:
if P > 1000:
fitness -= 1000
Se puede hacer depender la fitness de varios parámetros de manera ponderada:
fitness = parámetro_importante * 10 + parámetro_poco_importante * 0.5
También se pueden combinar diferentes funciones no lineales:
fitness = parámetro_1 * parámetro_2 - parámetro_3 **2 * log(parámetro_4)
Ahora te toca ser creativo! Elige con qué objetivo quieres optimizar la hélice!
Sugerencias de posibles objetivos de optimización:
- Mínimo radio posible, manteniendo una tracción mínima de 30 Newtons
- Mínima potencia posible, máxima eficiencia, y mínimo radio posible en menor medida, manteniendo una tracción mínima de 40 Newtons y un mach en la punta de las palas de como mucho 0.7
- Mínima potencia posible y máxima eficiencia cuando vuela a 70 m/s, tracción mayor a 50 Newtons en el despegue (vz = 0), mínimo peso posible (calculado a partir del radio, número y anchura de las palas) (Puede que tengas que reescribir la función y el diccionario de performances!)
```
def calculate_fitness (individual):
dict_traits = individuo.traits
dict_performances = individuo.performances
fitness = ????? #Be Creative!
individual.fitness = fitness
```
Ya tenemos terminado todo lo que necesitamos a nivel de individuo!
## Que comiencen los Juegos!
Es hora de trabajar a nivel de algoritmo, y para ello, lo primero es crear una sociedad compuesta de individuos aleatorios. Definamos una función para ello.
```
def immigration (society, target_population, dict_genes):
while len(society) < target_population:
new_individual = Individual (generate_genome (dict_genes)) # Generamos un individuo aleatorio
calculate_traits (new_individual, dict_genes) # Calculamos sus traits
calculate_performances (new_individual) # Calculamos sus performances
calculate_fitness (new_individual) # Calculamos su fitness
society.append (new_individual) # Nuestro nuevo ciudadano está listo para unirse al grupo!
```
Ahora podemos crear nuestra sociedad:
```
society = []
immigration (society, 12, dict_genes) #12 por ejemplo, pueden ser los que sean
#Veamos qué pinta tienen los genes de la población
plt.matshow([individual.genome for individual in society], cmap=plt.cm.gray)
```
Ya tenemos nuestra pequeña sociedad, aumentémosla un poco más mezclando entre sí a los ciudadanos con mejores fitness! Vamos a extender nuestra población mezclando los genomas de otros individuos. Los individuos con mejor fitness es más probable que se reproduzcan. Además, en los nuevos individuos produciremos ligeras mutaciones aleatorias.
```
#This function was taken from Eli Bendersky's website
#It returns an index of a list called "weights",
#where the content of each element in "weights" is the probability of this index to be returned.
#For this function to be as fast as possible we need to pass it a list of weights in descending order.
def weighted_choice_sub(weights):
rnd = random.random() * sum(weights)
for i, w in enumerate(weights):
rnd -= w
if rnd < 0:
return i
def crossover (society, reproduction_rate, mutation_rate):
#First we create a list with the fitness values of every individual in the society
fitness_list = [individual.fitness for individual in society]
#We sort the individuals in the society in descending order of fitness.
society_sorted = [x for (y, x) in sorted(zip(fitness_list, society), key=lambda x: x[0], reverse=True)]
#We then create a list of relative probabilities in descending order,
#so that the fittest individual in the society has N times more chances to reproduce than the least fit,
#where N is the number of individuals in the society.
probability = [i for i in reversed(range(1,len(society_sorted)+1))]
#We create a list of weights with the probabilities of non-mutation and mutation
mutation = [1 - mutation_rate, mutation_rate]
#For every new individual to be created through reproduction:
for i in range (int(len(society) * reproduction_rate)):
#We select two parents randomly, using the list of probabilities in "probability".
father, mother = society_sorted[weighted_choice_sub(probability)], society_sorted[weighted_choice_sub(probability)]
#We randomly select two cutting points for the genome.
a, b = random.randrange(0, len(father.genome)), random.randrange(0, len(father.genome))
#And we create the genome of the child putting together the genome slices of the parents in the cutting points.
child_genome = father.genome[0:min(a,b)]+mother.genome[min(a,b):max(a,b)]+father.genome[max(a,b):]
#For every bit in the not-yet-born child, we generate a list containing
#1's in the positions where the genome must mutate (i.e. the bit must switch its value)
#and 0's in the positions where the genome must stay the same.
n = [weighted_choice_sub(mutation) for ii in range(len(child_genome))]
#This line switches the bits of the genome of the child that must mutate.
mutant_child_genome = [abs(n[i] - child_genome[i]) for i in range(len(child_genome))]
#We finally append the newborn individual to the society
newborn = Individual(mutant_child_genome)
calculate_traits (newborn, dict_genes)
calculate_performances (newborn)
calculate_fitness (newborn)
society.append(newborn)
```
Ahora que tenemos una sociedad extensa, es el momento de que actúe la selección "natural": Eliminaremos de la sociedad a los individuos con peor fitness hasta llegar a una población objetivo.
```
def tournament(society, target_population):
while len(society) > target_population:
fitness_list = [individual.fitness for individual in society]
society.pop(fitness_list.index(min(fitness_list)))
```
Ya tenemos nuestro algoritmo prácticamente terminado!
```
society = []
fitness_max = []
for generation in range(30):
immigration (society, 100, dict_genes) #Añade individuos aleatorios a la sociedad hasta tener 100
fitness_max += [max([individual.fitness for individual in society])]
tournament (society, 15) #Los hace competir hasta que quedan 15
crossover(society, 5, 0.05) #Los ganadores se reproducen hasta tener 75
plt.plot(fitness_max)
plt.title('Evolución del valor de fitness')
tournament (society, 1) #Buscamos el mejor de todos
winner = society[0]
print(winner.traits) #Comprobamos sus características
print(winner.performances)
```
Siro Moreno y Carlos Dorado, Aeropython, 20 de Noviembre de 2015
| github_jupyter |
# PyTorch
<img src="https://raw.githubusercontent.com/GokuMohandas/practicalAI/master/images/logo.png" width=150>
In this lesson we'll learn about PyTorch which is a machine learning library used to build dynamic neural networks. We'll learn about the basics, like creating and using Tensors, in this lesson but we'll be making models with it in the next lesson.
<img src="https://raw.githubusercontent.com/GokuMohandas/practicalAI/master/images/pytorch.png" width=300>
# Tensor basics
```
# Load PyTorch library
!pip3 install torch
import numpy as np
import torch
# Creating a zero tensor
x = torch.Tensor(3, 4)
print("Type: {}".format(x.type()))
print("Size: {}".format(x.shape))
print("Values: \n{}".format(x))
# Creating a random tensor
x = torch.randn(2, 3) # normal distribution (rand(2,3) -> uniform distribution)
print (x)
# Zero and Ones tensor
x = torch.zeros(2, 3)
print (x)
x = torch.ones(2, 3)
print (x)
# List → Tensor
x = torch.Tensor([[1, 2, 3],[4, 5, 6]])
print("Size: {}".format(x.shape))
print("Values: \n{}".format(x))
# NumPy array → Tensor
x = torch.from_numpy(np.random.rand(2, 3))
print("Size: {}".format(x.shape))
print("Values: \n{}".format(x))
# Changing tensor type
x = torch.Tensor(3, 4)
print("Type: {}".format(x.type()))
x = x.long()
print("Type: {}".format(x.type()))
```
# Tensor operations
```
# Addition
x = torch.randn(2, 3)
y = torch.randn(2, 3)
z = x + y
print("Size: {}".format(z.shape))
print("Values: \n{}".format(z))
# Dot product
x = torch.randn(2, 3)
y = torch.randn(3, 2)
z = torch.mm(x, y)
print("Size: {}".format(z.shape))
print("Values: \n{}".format(z))
# Transpose
x = torch.randn(2, 3)
print("Size: {}".format(x.shape))
print("Values: \n{}".format(x))
y = torch.t(x)
print("Size: {}".format(y.shape))
print("Values: \n{}".format(y))
# Reshape
z = x.view(3, 2)
print("Size: {}".format(z.shape))
print("Values: \n{}".format(z))
# Dangers of reshaping (unintended consequences)
x = torch.tensor([
[[1,1,1,1], [2,2,2,2], [3,3,3,3]],
[[10,10,10,10], [20,20,20,20], [30,30,30,30]]
])
print("Size: {}".format(x.shape))
print("Values: \n{}\n".format(x))
a = x.view(x.size(1), -1)
print("Size: {}".format(a.shape))
print("Values: \n{}\n".format(a))
b = x.transpose(0,1).contiguous()
print("Size: {}".format(b.shape))
print("Values: \n{}\n".format(b))
c = b.view(b.size(0), -1)
print("Size: {}".format(c.shape))
print("Values: \n{}".format(c))
# Dimensional operations
x = torch.randn(2, 3)
print("Values: \n{}".format(x))
y = torch.sum(x, dim=0) # add each row's value for every column
print("Values: \n{}".format(y))
z = torch.sum(x, dim=1) # add each columns's value for every row
print("Values: \n{}".format(z))
```
# Indexing, Splicing and Joining
```
x = torch.randn(3, 4)
print("x: \n{}".format(x))
print ("x[:1]: \n{}".format(x[:1]))
print ("x[:1, 1:3]: \n{}".format(x[:1, 1:3]))
# Select with dimensional indicies
x = torch.randn(2, 3)
print("Values: \n{}".format(x))
col_indices = torch.LongTensor([0, 2])
chosen = torch.index_select(x, dim=1, index=col_indices) # values from column 0 & 2
print("Values: \n{}".format(chosen))
row_indices = torch.LongTensor([0, 1])
chosen = x[row_indices, col_indices] # values from (0, 0) & (2, 1)
print("Values: \n{}".format(chosen))
# Concatenation
x = torch.randn(2, 3)
print("Values: \n{}".format(x))
y = torch.cat([x, x], dim=0) # stack by rows (dim=1 to stack by columns)
print("Values: \n{}".format(y))
```
# Gradients
```
# Tensors with gradient bookkeeping
x = torch.rand(3, 4, requires_grad=True)
y = 3*x + 2
z = y.mean()
z.backward() # z has to be scalar
print("Values: \n{}".format(x))
print("x.grad: \n", x.grad)
```
* $ y = 3x + 2 $
* $ z = \sum{y}/N $
* $ \frac{\partial(z)}{\partial(x)} = \frac{\partial(z)}{\partial(y)} \frac{\partial(y)}{\partial(x)} = \frac{1}{N} * 3 = \frac{1}{12} * 3 = 0.25 $
# CUDA tensors
```
# Is CUDA available?
print (torch.cuda.is_available())
```
If the code above return False, then go to `Runtime` → `Change runtime type` and select `GPU` under `Hardware accelerator`.
```
# Creating a zero tensor
x = torch.Tensor(3, 4).to("cpu")
print("Type: {}".format(x.type()))
# Creating a zero tensor
x = torch.Tensor(3, 4).to("cuda")
print("Type: {}".format(x.type()))
```
| github_jupyter |
<!--NAVIGATION-->
< [特征工程](05.04-Feature-Engineering.ipynb) | [目录](Index.ipynb) | [深入:线性回归](05.06-Linear-Regression.ipynb) >
<a href="https://colab.research.google.com/github/wangyingsm/Python-Data-Science-Handbook/blob/master/notebooks/05.05-Naive-Bayes.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
# In Depth: Naive Bayes Classification
# 深入:朴素贝叶斯分类
> The previous four sections have given a general overview of the concepts of machine learning.
In this section and the ones that follow, we will be taking a closer look at several specific algorithms for supervised and unsupervised learning, starting here with naive Bayes classification.
前面四个小节对机器学习的概念给出了概述。本节开始,我们会进入到有监督学习和无监督学习的一些特定算法当中,进行较深入的介绍。首先从本节的朴素贝叶斯分类开始。
> Naive Bayes models are a group of extremely fast and simple classification algorithms that are often suitable for very high-dimensional datasets.
Because they are so fast and have so few tunable parameters, they end up being very useful as a quick-and-dirty baseline for a classification problem.
This section will focus on an intuitive explanation of how naive Bayes classifiers work, followed by a couple examples of them in action on some datasets.
朴素贝叶斯模型是一组非常快和简单的分类算法,它们经常用来对高维度数据集进行分类处理。因为它们非常快和有一些可调的参数,它们最终成为了分类问题很好用的临时基线方法。本节会聚焦在对朴素贝叶斯分类器工作原理的直观介绍,然后会在不同的数据集上应用它作为例子。
## Bayesian Classification
## 贝叶斯分类
> Naive Bayes classifiers are built on Bayesian classification methods.
These rely on Bayes's theorem, which is an equation describing the relationship of conditional probabilities of statistical quantities.
In Bayesian classification, we're interested in finding the probability of a label given some observed features, which we can write as $P(L~|~{\rm features})$.
Bayes's theorem tells us how to express this in terms of quantities we can compute more directly:
朴素贝叶斯分类建立在贝叶斯分类方法的基础上。这些分类方法的基础是贝叶斯定理,这是一个用来描述统计理论中条件概率的等式。在贝叶斯分类中,我们感兴趣的是在给定观测特征数据上找到一个标签的概率,我们写做$P(L~|~{\rm features})$。贝叶斯定理告诉我们如何使用这些已知的特征量直接计算概率:
$$
P(L~|~{\rm features}) = \frac{P({\rm features}~|~L)P(L)}{P({\rm features})}
$$
> If we are trying to decide between two labels—let's call them $L_1$ and $L_2$—then one way to make this decision is to compute the ratio of the posterior probabilities for each label:
如果我们尝试在两个标签中去选择,假设我们称它们为$L_1$和$L_2$,那么做这个选择的一种方法是计算每一个标签的后验概率:
$$
\frac{P(L_1~|~{\rm features})}{P(L_2~|~{\rm features})} = \frac{P({\rm features}~|~L_1)}{P({\rm features}~|~L_2)}\frac{P(L_1)}{P(L_2)}
$$
> All we need now is some model by which we can compute $P({\rm features}~|~L_i)$ for each label.
Such a model is called a *generative model* because it specifies the hypothetical random process that generates the data.
Specifying this generative model for each label is the main piece of the training of such a Bayesian classifier.
The general version of such a training step is a very difficult task, but we can make it simpler through the use of some simplifying assumptions about the form of this model.
因此我们所需要的就是一个能够计算每一个标签的$P({\rm features}~|~L_i)$值的模型。这个模型被称为*生成模型*,因为它指定了产生数据的假设随机过程。对于训练贝叶斯分类器来说,为每个标签找到这样的通用模型是最主要的步骤。获得这种训练步骤的通用版本是很困难的,但是我们能够通过使用关于该模型的假设来简化这项任务。
> This is where the "naive" in "naive Bayes" comes in: if we make very naive assumptions about the generative model for each label, we can find a rough approximation of the generative model for each class, and then proceed with the Bayesian classification.
Different types of naive Bayes classifiers rest on different naive assumptions about the data, and we will examine a few of these in the following sections.
这就是“朴素贝叶斯”中的“朴素”的由来:如果我们对通用模型中的每个标签作出非常朴素的假设,我们就可以找到通用模型中每个标签的大概分布,然后进行贝叶斯分类。不同的朴素贝叶斯分类器取决于对数据不同的朴素假设上,我们在本节后续内容中会介绍它们中的一部分。
> We begin with the standard imports:
首先是需要用到的包:
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns; sns.set()
```
## Gaussian Naive Bayes
## 高斯朴素贝叶斯
> Perhaps the easiest naive Bayes classifier to understand is Gaussian naive Bayes.
In this classifier, the assumption is that *data from each label is drawn from a simple Gaussian distribution*.
Imagine that you have the following data:
朴素贝叶斯分类器中最容易理解的也许就是高斯朴素贝叶斯。这个分类器假定*每个标签的数据都服从简单正态分布*。例如你有如下数据:
```
from sklearn.datasets import make_blobs
X, y = make_blobs(100, 2, centers=2, random_state=2, cluster_std=1.5)
plt.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='RdBu');
```
> One extremely fast way to create a simple model is to assume that the data is described by a Gaussian distribution with no covariance between dimensions.
This model can be fit by simply finding the mean and standard deviation of the points within each label, which is all you need to define such a distribution.
The result of this naive Gaussian assumption is shown in the following figure:
创建一个简单模型的最快速方法就是假定数据服从一个两个维度之间没有协方差的正态分布。这个模型可以通过简单的寻找每个标签中点的均值和标准差来拟合,你只需要定义这个分布即可。高斯朴素假设的结果显示在下图中:

[附录中生成图像的代码](06.00-Figure-Code.ipynb#Gaussian-Naive-Bayes)
> The ellipses here represent the Gaussian generative model for each label, with larger probability toward the center of the ellipses.
With this generative model in place for each class, we have a simple recipe to compute the likelihood $P({\rm features}~|~L_1)$ for any data point, and thus we can quickly compute the posterior ratio and determine which label is the most probable for a given point.
上图中的椭圆表示每个标签的高斯生成模型,越接近椭圆中心位置具有越大的概率。有了每个分类的生成模型后,我们就能简单的计算每一个点的概率$P({\rm features}~|~L_1)$,也就是后验概率,然后找到哪个标签在给定数据点上具有最大的概率。
> This procedure is implemented in Scikit-Learn's ``sklearn.naive_bayes.GaussianNB`` estimator:
这个过程在Scikit-Learn中实现成了`sklearn.naive_bayes.GaussianNB`评估器:
```
from sklearn.naive_bayes import GaussianNB
model = GaussianNB()
model.fit(X, y);
```
> Now let's generate some new data and predict the label:
现在让我们创建一些新数据,然后预测标签:
```
rng = np.random.RandomState(0)
Xnew = [-6, -14] + [14, 18] * rng.rand(2000, 2)
ynew = model.predict(Xnew)
```
> Now we can plot this new data to get an idea of where the decision boundary is:
下面我们将新数据点绘制在图上,你能看到分类判定的边界位置:
```
plt.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='RdBu')
lim = plt.axis()
plt.scatter(Xnew[:, 0], Xnew[:, 1], c=ynew, s=20, cmap='RdBu', alpha=0.1)
plt.axis(lim);
```
> We see a slightly curved boundary in the classifications—in general, the boundary in Gaussian naive Bayes is quadratic.
我们看到分类之间的边界是有点弯曲的,因为通常来说,高斯朴素贝叶斯的边界是二次曲线。
> A nice piece of this Bayesian formalism is that it naturally allows for probabilistic classification, which we can compute using the ``predict_proba`` method:
这种贝叶斯分类方法的一个好处是它天然支持概率分类,我们可以通过`predict_proba`计算每个分类的概率:
```
yprob = model.predict_proba(Xnew)
yprob[-8:].round(2)
```
> The columns give the posterior probabilities of the first and second label, respectively.
If you are looking for estimates of uncertainty in your classification, Bayesian approaches like this can be a useful approach.
上面结果中的两列分别给出了两个标签的后验概率。如果你在寻找你分类中的不确定性的话,贝叶斯方法能提供有效的判断依据。
> Of course, the final classification will only be as good as the model assumptions that lead to it, which is why Gaussian naive Bayes often does not produce very good results.
Still, in many cases—especially as the number of features becomes large—this assumption is not detrimental enough to prevent Gaussian naive Bayes from being a useful method.
当然最终分类结果最多只能达到模型的假定情况,这表明高斯朴素贝叶斯方法常常不会产生非常好的结果。但是在很多情况下,特别是当特征数量变得很大时,这个假定并不会导致高斯朴素贝叶斯方法完全失去意义。
## Multinomial Naive Bayes
## 多项式朴素贝叶斯
> The Gaussian assumption just described is by no means the only simple assumption that could be used to specify the generative distribution for each label.
Another useful example is multinomial naive Bayes, where the features are assumed to be generated from a simple multinomial distribution.
The multinomial distribution describes the probability of observing counts among a number of categories, and thus multinomial naive Bayes is most appropriate for features that represent counts or count rates.
前面描述的高斯假设不是唯一的简单假设可以用来为每个标签产生生成分布。另一个有用的方法是多项式朴素贝叶斯,这个方法假定数据的特征是从一个简单的多项式分布中生成的。多项式分布描述了在一些分组中观察到的计数的概率,因此多项式朴素贝叶斯对于表达计数或计数的比例之类的特征是最合适的。
> The idea is precisely the same as before, except that instead of modeling the data distribution with the best-fit Gaussian, we model the data distribuiton with a best-fit multinomial distribution.
这里的原理和前面是一样的,只是不是使用正态分布来拟合数据模型,而是使用多项式分布来拟合数据模型。
### Example: Classifying Text
### 例子:分类文字
> One place where multinomial naive Bayes is often used is in text classification, where the features are related to word counts or frequencies within the documents to be classified.
We discussed the extraction of such features from text in [Feature Engineering](05.04-Feature-Engineering.ipynb); here we will use the sparse word count features from the 20 Newsgroups corpus to show how we might classify these short documents into categories.
多项式朴素贝叶斯经常被用到的场合是文字分类,因为这个场景下的特征是单词的计数或者文档中单词出现的频率。我们在[特征工程](05.04-Feature-Engineering.ipynb)一节中介绍过在文本中提取这样的特征的方法;这里我们会使用20个新闻组的语料库提取出来的稀疏单词计数特征来展示将这些短文档分类的方法。
> Let's download the data and take a look at the target names:
让我们下载这个数据然后查看一下目标分类的名称:
```
from sklearn.datasets import fetch_20newsgroups
data = fetch_20newsgroups()
data.target_names
```
> For simplicity here, we will select just a few of these categories, and download the training and testing set:
这里为了简化,我们仅选择其中部分分类,然后载入训练集和测试集:
```
categories = ['talk.religion.misc', 'soc.religion.christian',
'sci.space', 'comp.graphics']
train = fetch_20newsgroups(subset='train', categories=categories)
test = fetch_20newsgroups(subset='test', categories=categories)
```
> Here is a representative entry from the data:
下面展示部分数据:
```
print(train.data[5])
```
> In order to use this data for machine learning, we need to be able to convert the content of each string into a vector of numbers.
For this we will use the TF-IDF vectorizer (discussed in [Feature Engineering](05.04-Feature-Engineering.ipynb)), and create a pipeline that attaches it to a multinomial naive Bayes classifier:
为了要将这个数据集应用到机器学习上,我们需要将数据中的每个字符串内容转换为数字的向量。我们使用TF-IDF来实现向量化(参见[特征工程](05.04-Feature-Engineering.ipynb)),然后创建一个管道操作将一个多项式朴素贝叶斯分类器连接进来:
```
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.naive_bayes import MultinomialNB
from sklearn.pipeline import make_pipeline
model = make_pipeline(TfidfVectorizer(), MultinomialNB())
```
> With this pipeline, we can apply the model to the training data, and predict labels for the test data:
我们可以将这个管道应用到训练集上,然后在测试集上去进行标签预测:
```
model.fit(train.data, train.target)
labels = model.predict(test.data)
```
> Now that we have predicted the labels for the test data, we can evaluate them to learn about the performance of the estimator.
For example, here is the confusion matrix between the true and predicted labels for the test data:
有了对测试数据预测的标签之后,我们可以对评估器的性能作出判断。例如下面展示了预测标签和实际标签之间的混淆矩阵:
```
from sklearn.metrics import confusion_matrix
mat = confusion_matrix(test.target, labels)
sns.heatmap(mat.T, square=True, annot=True, fmt='d', cbar=False,
xticklabels=train.target_names, yticklabels=train.target_names)
plt.xlabel('true label')
plt.ylabel('predicted label');
```
> Evidently, even this very simple classifier can successfully separate space talk from computer talk, but it gets confused between talk about religion and talk about Christianity.
This is perhaps an expected area of confusion!
从上图看出,即便是这么简单的分类器也能成功的将宇宙学讨论和计算机科学讨论内容区分开,但是它在将宗教讨论和基督教讨论区分的时候遇到了困难。因为可能这是一个本来就容易混淆的领域。
> The very cool thing here is that we now have the tools to determine the category for *any* string, using the ``predict()`` method of this pipeline.
Here's a quick utility function that will return the prediction for a single string:
我们现在有了一个模型来对*任何*字符串进行分类检测了,非常酷对不对,只需要在这个管道对象上使用`predict()`方法即可。下面我们创建一个简单的工具函数来对任何字符串输入返回标签预测的输出结果:
```
def predict_category(s, train=train, model=model):
pred = model.predict([s])
return train.target_names[pred[0]]
```
> Let's try it out:
赶快来试一下:
```
predict_category('sending a payload to the ISS')
predict_category('discussing islam vs atheism')
predict_category('determining the screen resolution')
```
> Remember that this is nothing more sophisticated than a simple probability model for the (weighted) frequency of each word in the string; nevertheless, the result is striking.
Even a very naive algorithm, when used carefully and trained on a large set of high-dimensional data, can be surprisingly effective.
请记住这里做的事情仅是对字符串中每个单词的(加权)出现频率生成了一个概率模型而已;然而结果却令人惊奇。即使非常朴素的算法,只要小心使用,并且在一个大规模的高维度数据集上进行训练的话,也能非常有效。
## When to Use Naive Bayes
## 何时使用朴素贝叶斯方法
> Because naive Bayesian classifiers make such stringent assumptions about data, they will generally not perform as well as a more complicated model.
That said, they have several advantages:
> - They are extremely fast for both training and prediction
- They provide straightforward probabilistic prediction
- They are often very easily interpretable
- They have very few (if any) tunable parameters
因为朴素贝叶斯分类器对数据进行了如此严格的假设,它们通常不会比其他复杂的模型更加有效。朴素贝叶斯方法有下面几个优点:
- 它们非常快,无论是在训练还是预测中
- 它们提供了直接的概率预测
- 它们通常很容易解释
- 它们有很少的可调参数
> These advantages mean a naive Bayesian classifier is often a good choice as an initial baseline classification.
If it performs suitably, then congratulations: you have a very fast, very interpretable classifier for your problem.
If it does not perform well, then you can begin exploring more sophisticated models, with some baseline knowledge of how well they should perform.
这些特点导致朴素贝叶斯分类器经常被作为初始化的基线分类标准。如果它性能很好,恭喜:你的问题已经有了一个非常快速很容易解释的分类模型了。如果它的性能不如人意,那么你可以开始尝试更加复杂的模型,然后将朴素贝叶斯分类器的性能结果作为标准来对新的模型进行评判。
> Naive Bayes classifiers tend to perform especially well in one of the following situations:
> - When the naive assumptions actually match the data (very rare in practice)
- For very well-separated categories, when model complexity is less important
- For very high-dimensional data, when model complexity is less important
朴素贝叶斯分类器在下面的一些情况下通常能够特别良好的工作:
- 当朴素假定能够拟合数据时(实践中非常少见)
- 对于数据本身分类就已经很清晰的情况,此时模型复杂度并不十分重要
- 对于数据维度非常多的情况,此时模型复杂度并不十分重要
> The last two points seem distinct, but they actually are related: as the dimension of a dataset grows, it is much less likely for any two points to be found close together (after all, they must be close in *every single dimension* to be close overall).
This means that clusters in high dimensions tend to be more separated, on average, than clusters in low dimensions, assuming the new dimensions actually add information.
For this reason, simplistic classifiers like naive Bayes tend to work as well or better than more complicated classifiers as the dimensionality grows: once you have enough data, even a simple model can be very powerful.
后两点看起来是独立的因素,但是实际上它们是关联的:当数据集的维度增加时,两个数据点非常接近的情况是非常少见的(毕竟它们要在*每个维度*都接近才能互相接近)。这意味着高纬度中的分类相对于低维度数据,如果新增的维度确实增加了数据的信息量(特征)的话,高维度数据点会倾向于出现在更不同的位置。因此像朴素贝叶斯这样的简单分类器在数据维度增加情况下可能会比复杂分类器工作的更好:一旦你有了足够的数据,哪怕是简单的模型也能非常强大。
<!--NAVIGATION-->
< [特征工程](05.04-Feature-Engineering.ipynb) | [目录](Index.ipynb) | [深入:线性回归](05.06-Linear-Regression.ipynb) >
<a href="https://colab.research.google.com/github/wangyingsm/Python-Data-Science-Handbook/blob/master/notebooks/05.05-Naive-Bayes.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
| github_jupyter |
**Download** (right-click, save target as ...) this page as a jupyterlab notebook from:
[Lab7](https://atomickitty.ddns.net:8000/user/sensei/files/engr-1330-webroot/engr-1330-webbook/ctds-psuedocourse/docs/8-Labs/Lab6/Lab7_Dev.ipynb?_xsrf=2%7C1b4d47c3%7C0c3aca0c53606a3f4b71c448b09296ae%7C1623531240)
___
# <font color=darkred>Laboratory 7: Numpy for Bread! </font>
```
# Preamble script block to identify host, user, and kernel
import sys
! hostname
! whoami
print(sys.executable)
print(sys.version)
print(sys.version_info)
```
## Full name:
## R#:
## Title of the notebook:
## Date:
___
 <br>
## <font color=purple>Numpy</font>
Numpy is the core library for scientific computing in Python. It provides a high-performance multidimensional array object, and tools for working with these arrays. The library’s name is short for “Numeric Python” or “Numerical Python”. If you are curious about NumPy, this cheat sheet is recommended:
https://s3.amazonaws.com/assets.datacamp.com/blog_assets/Numpy_Python_Cheat_Sheet.pdf
### <font color=purple>Arrays</font>
A numpy array is a grid of values, all of the same type, and is indexed by a tuple of nonnegative integers. The number of dimensions is the rank of the array; the shape of an array is a tuple of integers giving the size of the array along each dimension. In other words, an array contains information about the raw data, how to locate an element and how to interpret an element.To make a numpy array, you can just use the np.array() function. All you need to do is pass a list to it. Don’t forget that, in order to work with the np.array() function, you need to make sure that the numpy library is present in your environment.
If you want to read more about the differences between a Python list and NumPy array, this link is recommended:
https://webcourses.ucf.edu/courses/1249560/pages/python-lists-vs-numpy-arrays-what-is-the-difference
___
### Example- 1D Arrays
__Let's create a 1D array from the 2000s (2000-2009):__
```
import numpy as np #First, we need to impoty "numpy"
mylist = [2000,2001,2002,2003,2004,2005,2006,2007,2008,2009] #Create a list of the years
print(mylist) #Check how it looks
np.array(mylist) #Define it as a numpy array
```
___
### Example- n-Dimensional Arrays
__Let's create a 5x2 array from the 2000s (2000-2009):__
```
myotherlist = [[2000,2001],[2002,2003],[2004,2005],[2006,2007],[2008,2009]] #Since I want a 5x2 array, I should group the years two by two
print(myotherlist) #See how it looks as a list
np.array(myotherlist) #See how it looks as a numpy array
```
### <font color=purple>Arrays Arithmetic</font>
Once you have created the arrays, you can do basic Numpy operations. Numpy offers a variety of operations applicable on arrays. From basic operations such as summation, subtraction, multiplication and division to more advanced and essential operations such as matrix multiplication and other elementwise operations. In the examples below, we will go over some of these:
___
### Example- 1D Array Arithmetic
- Define a 1D array with [0,12,24,36,48,60,72,84,96]
- Multiple all elements by 2
- Take all elements to the power of 2
- Find the maximum value of the array and its position
- Find the minimum value of the array and its position
- Define another 1D array with [-12,0,12,24,36,48,60,72,84]
- Find the summation and subtraction of these two arrays
- Find the multiplication of these two arrays
```
import numpy as np #import numpy
Array1 = np.array([0,12,24,36,48,60,72,84,96]) #Step1: Define Array1
print(Array1)
print(Array1*2) #Step2: Multiple all elements by 2
print(Array1**2) #Step3: Take all elements to the power of 2
print(np.power(Array1,2)) #Another way to do the same thing, by using a function in numpy
print(np.max(Array1)) #Step4: Find the maximum value of the array
print(np.argmax(Array1)) ##Step4: Find the postition of the maximum value
print(np.min(Array1)) #Step5: Find the minimum value of the array
print(np.argmin(Array1)) ##Step5: Find the postition of the minimum value
Array2 = np.array([-12,0,12,24,36,48,60,72,84]) #Step6: Define Array2
print(Array2)
print(Array1+Array2) #Step7: Find the summation of these two arrays
print(Array1-Array2) #Step7: Find the subtraction of these two arrays
print(Array1*Array2) #Step8: Find the multiplication of these two arrays
```
___
### Example- n-Dimensional Array Arithmetic
- Define a 2x2 array with [5,10,15,20]
- Define another 2x2 array with [3,6,9,12]
- Find the summation and subtraction of these two arrays
- Find the minimum number in the multiplication of these two arrays
- Find the position of the maximum in the multiplication of these two arrays
- Find the mean of the multiplication of these two arrays
- Find the mean of the first row of the multiplication of these two arrays
```
import numpy as np #import numpy
Array1 = np.array([[5,10],[15,20]]) #Step1: Define Array1
print(Array1)
Array2 = np.array([[3,6],[9,12]]) #Step2: Define Array2
print(Array2)
print(Array1+Array2) #Step3: Find the summation
print(Array1-Array2) #Step3: Find the subtraction
MultArray = Array1@Array2 #Step4: To perform a typical matrix multiplication (or matrix product)
MultArray1 = Array1.dot(Array2) #Step4: Another way To perform a matrix multiplication
print(MultArray)
print(MultArray1)
print(np.min(MultArray)) #Step4: Find the minimum value of the multiplication
print(np.argmax(MultArray)) ##Step5: Find the postition of the maximum value
print(np.mean(MultArray)) ##Step6: Find the mean of the multiplication of these two arrays
print(np.mean(MultArray[0,:])) ##Step7: Find the mean of the first row of the multiplication of these two arrays
```
___
### <font color=purple>Arrays Comparison</font>
Comparing two NumPy arrays determines whether they are equivalent by checking if every element at each corresponding index are the same.
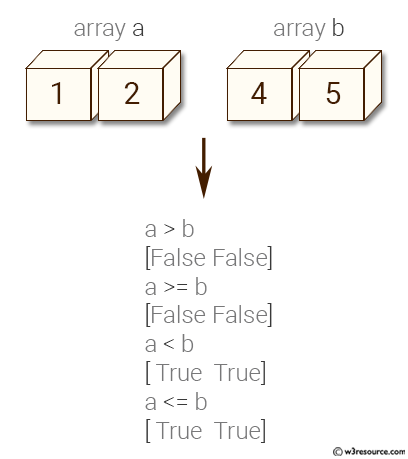
___
### Example- 1D Array Comparison
- Define a 1D array with [1.0,2.5,3.4,7,7]
- Define another 1D array with [5.0/5.0,5.0/2,6.8/2,21/3,14/2]
- Compare and see if the two arrays are equal
- Define another 1D array with [6,1.4,2.2,7.5,7]
- Compare and see if the first array is greater than or equal to the third array
```
import numpy as np #import numpy
Array1 = np.array([1.0,2.5,3.4,7,7]) #Step1: Define Array1
print(Array1)
Array2 = np.array([5.0/5.0,5.0/2,6.8/2,21/3,14/2]) #Step2: Define Array1
print(Array2)
print(np.equal(Array1, Array2)) #Step3: Compare and see if the two arrays are equal
Array3 = np.array([6,1.4,2.2,7.5,7]) #Step4: Define Array3
print(Array3)
print(np.greater_equal(Array1, Array3)) #Step3: Compare and see if the two arrays are equal
```
___
### <font color=purple>Arrays Manipulation</font>
numpy.copy() allows us to create a copy of an array. This is particularly useful when we need to manipulate an array while keeping an original copy in memory.
The numpy.delete() function returns a new array with sub-arrays along an axis deleted. Let's have a look at the examples.
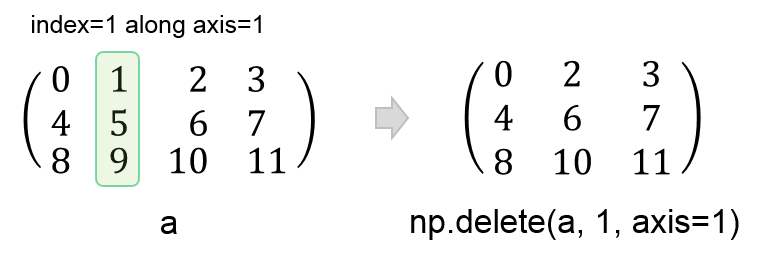

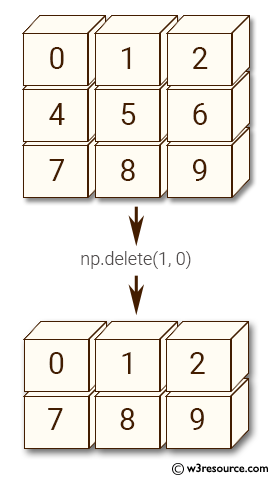
___
### Example- Copying and Deleting Arrays and Elements
- Define a 1D array, named "x" with [1,2,3]
- Define "y" so that "y=x"
- Define "z" as a copy of "x"
- Discuss the difference between y and z
- Delete the second element of x
```
import numpy as np #import numpy
x = np.array([1,2,3]) #Step1: Define x
print(x)
y = x #Step2: Define y as y=x
print(y)
z = np.copy(x) #Step3: Define z as a copy of x
print(z)
# For Step4: They look similar but check this out:
x[1] = 8 # If we change x ...
print(x)
print(y)
print(z)
# By modifying x, y changes but z remains as a copy of the initial version of x.
x = np.delete(x, 1) #Step5: Delete the second element of x
print(x)
```
___
### <font color=purple>Sorting Arrays</font>
Sorting means putting elements in an ordered sequence. Ordered sequence is any sequence that has an order corresponding to elements, like numeric or alphabetical, ascending or descending. If you use the sort() method on a 2-D array, both arrays will be sorted.

___
### Example- Sorting 1D Arrays
__Define a 1D array as ['FIFA 2020','Red Dead Redemption','Fallout','GTA','NBA 2018','Need For Speed'] and print it out. Then, sort the array alphabetically.__
```
import numpy as np #import numpy
games = np.array(['FIFA 2020','Red Dead Redemption','Fallout','GTA','NBA 2018','Need For Speed'])
print(games)
print(np.sort(games))
```
___
### Example- Sorting n-Dimensional Arrays
__Define a 3x3 array with 17,-6,2,86,-12,0,0,23,12 and print it out. Then, sort the array.__
```
import numpy as np #import numpy
a = np.array([[17,-6,2],[86,-12,0],[0,23,12]])
print(a)
print ("Along columns : \n", np.sort(a,axis = 0) ) #This will be sorting in each column
print ("Along rows : \n", np.sort(a,axis = 1) ) #This will be sorting in each row
print ("Sorting by default : \n", np.sort(a) ) #Same as above
print ("Along None Axis : \n", np.sort(a,axis = None) ) #This will be sorted like a 1D array
```
___
### <font color=purple>Partitioning (Slice) Arrays</font>
Slicing in python means taking elements from one given index to another given index.
We can do slicing like this: [start:end].
We can also define the step, like this: [start:end:step].
If we don't pass start its considered 0
If we don't pass end its considered length of array in that dimension
If we don't pass step its considered 1

___
### Example- Slicing 1D Arrays
__Define a 1D array as [1,3,5,7,9], slice out the [3,5,7] and print it out.__
```
import numpy as np #import numpy
a = np.array([1,3,5,7,9]) #Define the array
print(a)
aslice = a[1:4] #slice the [3,5,7]
print(aslice) #print it out
```
___
### Example- Slicing n-Dimensional Arrays
__Define a 5x5 array with "Superman, Batman, Jim Hammond, Captain America, Green Arrow, Aquaman, Wonder Woman, Martian Manhunter, Barry Allen, Hal Jordan, Hawkman, Ray Palmer, Spider Man, Thor, Hank Pym, Solar, Iron Man, Dr. Strange, Daredevil, Ted Kord, Captian Marvel, Black Panther, Wolverine, Booster Gold, Spawn " and print it out. Then:__
- Slice the first column and print it out
- Slice the third row and print it out
- Slice 'Wolverine' and print it out
- Slice a 3x3 array with 'Wonder Woman, Ray Palmer, Iron Man, Martian Manhunter, Spider Man, Dr. Strange, Barry Allen, Thor, Daredevil'
```
import numpy as np #import numpy
Superheroes = np.array([['Superman', 'Batman', 'Jim Hammond', 'Captain America', 'Green Arrow'],
['Aquaman', 'Wonder Woman', 'Martian Manhunter', 'Barry Allen', 'Hal Jordan'],
['Hawkman', 'Ray Palmer', 'Spider Man', 'Thor', 'Hank Pym'],
['Solar', 'Iron Man', 'Dr. Strange', 'Daredevil', 'Ted Kord'],
['Captian Marvel', 'Black Panther', 'Wolverine', 'Booster Gold', 'Spawn']])
print(Superheroes) #Step1
print(Superheroes[:,0])
print(Superheroes[2,:])
print(Superheroes[4,2])
print(Superheroes[1:4,1:4])
```
___
## <font color=orange>This is a Numpy Cheat Sheet- similar to the one you had on top of this notebook!</font>

### Check out this link for more: <br>
https://blog.finxter.com/collection-10-best-numpy-cheat-sheets-every-python-coder-must-own/
___
 <br>
*Here are some of the resources used for creating this notebook:*
- Johnson, J. (2020). Python Numpy Tutorial (with Jupyter and Colab). Retrieved September 15, 2020, from https://cs231n.github.io/python-numpy-tutorial/ <br>
- Willems, K. (2019). (Tutorial) Python NUMPY Array TUTORIAL. Retrieved September 15, 2020, from https://www.datacamp.com/community/tutorials/python-numpy-tutorial?utm_source=adwords_ppc <br>
- Willems, K. (2017). NumPy Cheat Sheet: Data Analysis in Python. Retrieved September 15, 2020, from https://www.datacamp.com/community/blog/python-numpy-cheat-sheet <br>
- W3resource. (2020). NumPy: Compare two given arrays. Retrieved September 15, 2020, from https://www.w3resource.com/python-exercises/numpy/python-numpy-exercise-28.php <br>
*Here are some great reads on this topic:*
- __"Python NumPy Tutorial"__ available at *https://www.geeksforgeeks.org/python-numpy-tutorial/<br>
- __"What Is NumPy?"__ a collection of blogs, available at *https://realpython.com/tutorials/numpy/ <br>
- __"Look Ma, No For-Loops: Array Programming With NumPy"__ by __Brad Solomon__ available at *https://realpython.com/numpy-array-programming/ <br>
- __"The Ultimate Beginner’s Guide to NumPy"__ by __Anne Bonner__ available at *https://towardsdatascience.com/the-ultimate-beginners-guide-to-numpy-f5a2f99aef54 <br>
*Here are some great videos on these topics:*
- __"Learn NUMPY in 5 minutes - BEST Python Library!"__ by __Python Programmer__ available at *https://www.youtube.com/watch?v=xECXZ3tyONo <br>
- __"Python NumPy Tutorial for Beginners"__ by __freeCodeCamp.org__ available at *https://www.youtube.com/watch?v=QUT1VHiLmmI <br>
- __"Complete Python NumPy Tutorial (Creating Arrays, Indexing, Math, Statistics, Reshaping)"__ by __Keith Galli__ available at *https://www.youtube.com/watch?v=GB9ByFAIAH4 <br>
- __"Python NumPy Tutorial | NumPy Array | Python Tutorial For Beginners | Python Training | Edureka"__ by __edureka!__ available at *https://www.youtube.com/watch?v=8JfDAm9y_7s <br>
___
 <br>
## Exercise: Python List vs. Numpy Arrays? <br>
### What are some differences between Python lists and Numpy arrays?
#### * Make sure to cite any resources that you may use.

| github_jupyter |
# Practical example
## Importing the relevant libraries
```
import numpy as np
import pandas as pd
import statsmodels.api as sm
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
import seaborn as sns
sns.set()
```
## Loading the raw data
```
data = pd.read_csv('1.04. Real-life example.csv')
data.head()
```
## Preprocessing
### Exploring the descriptive statistics of the variables
```
data.describe(include='all')
```
### Before deleting rows, drop all features we are going to drop:
- **Mileage** - because of high correlation with Year, and Year helps more than Mileage
- **Body** - doesn't help much (coefficients are pretty low)
- **Engine Type** - doesn't help much (coefficients are pretty low)
```
data.drop(['Engine Type','Body','Mileage'],axis=1, inplace=True)
data.describe(include='all')
```
### Dealing with missing values
```
data.isnull().sum()
data = data.dropna(axis=0)
data.describe(include='all')
```
### Exploring the PDFs
```
sns.distplot(data['Price'])
```
### Dealing with outliers
```
q = data['Price'].quantile(0.99)
data = data[data['Price']<q]
data.describe(include='all')
sns.distplot(data['Price'])
sns.distplot(data['EngineV'])
data = data[data['EngineV']<6.5]
sns.distplot(data['EngineV'])
sns.distplot(data['Year'])
q = data['Year'].quantile(0.01)
data = data[data['Year']>q]
sns.distplot(data['Year'])
data = data.reset_index(drop=True)
data.describe(include='all')
```
## Checking the OLS assumptions
```
f, (ax1, ax2) = plt.subplots(1, 2, sharey=True, figsize =(15,3))
ax1.scatter(data['Year'],data['Price'])
ax1.set_title('Price and Year')
ax2.scatter(data['EngineV'],data['Price'])
ax2.set_title('Price and EngineV')
plt.show()
sns.distplot(data['Price'])
```
### Relaxing the assumptions
```
log_price = np.log(data['Price'])
data['log_price'] = log_price
data = data.drop(['Price'],axis=1)
data
f, (ax1, ax2) = plt.subplots(1, 2, sharey=True, figsize =(15,3))
ax1.scatter(data['Year'],data['log_price'])
ax1.set_title('Log Price and Year')
ax2.scatter(data['EngineV'],data['log_price'])
ax2.set_title('Log Price and EngineV')
plt.show()
```
### Multicollinearity
```
data.columns.values
from statsmodels.stats.outliers_influence import variance_inflation_factor
variables = data[['Year','EngineV']]
vif = pd.DataFrame()
vif["VIF"] = [variance_inflation_factor(variables.values, i) for i in range(variables.shape[1])]
vif["features"] = variables.columns
vif
```
### Remove rare Model values
```
counts = data['Model'].value_counts()
max_count = counts.max()
index = -1
axes = counts.axes
for count in counts:
index += 1
if count < (0.04 * max_count):
data['Model'].replace(axes[0][index], "Other", inplace=True)
data.describe(include='all')
```
### Combine Brand and Model together
```
data['Brand_Model'] = data['Brand'] + '_' + data['Model']
data.drop(['Brand', 'Model'], axis=1, inplace=True)
data.describe(include='all')
```
## Create dummy variables
```
data = pd.get_dummies(data, drop_first=True)
data.head()
```
## Linear regression model
### Declare the inputs and the targets
```
targets = data['log_price']
inputs = data.drop(['log_price'],axis=1)
```
### Scale the data
```
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(inputs)
inputs_scaled = scaler.transform(inputs)
```
### Train Test Split
```
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(inputs_scaled, targets, test_size=0.2, random_state=365)
```
### Create the regression
```
reg = LinearRegression()
reg.fit(x_train,y_train)
y_hat = reg.predict(x_train)
plt.scatter(y_train, y_hat)
plt.xlabel('Targets (y_train)',size=18)
plt.ylabel('Predictions (y_hat)',size=18)
plt.xlim(6,13)
plt.ylim(6,13)
plt.show()
sns.distplot(y_train - y_hat)
plt.title("Residuals PDF", size=18)
reg.score(x_train,y_train)
```
### Finding the weights and bias
```
reg.intercept_
reg.coef_
reg_summary = pd.DataFrame(inputs.columns.values, columns=['Features'])
reg_summary['Weights'] = reg.coef_
reg_summary
```
## Testing
```
y_hat_test = reg.predict(x_test)
plt.scatter(y_test, y_hat_test, alpha=0.2)
plt.xlabel('Targets (y_test)',size=18)
plt.ylabel('Predictions (y_hat_test)',size=18)
plt.xlim(6,13)
plt.ylim(6,13)
plt.show()
df_pf = pd.DataFrame(np.exp(y_hat_test), columns=['Prediction'])
df_pf.head()
df_pf['Target'] = np.exp(y_test)
df_pf
y_test = y_test.reset_index(drop=True)
y_test.head()
df_pf['Target'] = np.exp(y_test)
df_pf
df_pf['Residual'] = df_pf['Target'] - df_pf['Prediction']
df_pf['Difference%'] = np.absolute(df_pf['Residual']/df_pf['Target']*100)
df_pf
df_pf.describe()
pd.options.display.max_rows = 999
pd.set_option('display.float_format', lambda x: '%.2f' % x)
df_pf.sort_values(by=['Difference%'])
```
## Main other conclusions:
- Getting Adjusted R2 above .91-.92 is overfitting:
- the coefficients are huge
- testing fails with bad results
- Not doing Log function on Price was not shown to be a good idea
- Year is extremely important in the predictions of the model. Not including it lowers R2 significantly
- Model is also extremely important in the predictions of the model. Not including it lowers R2 significantly
- Playing with outliers didn't help much (making them higher/lower, taking from an high/low end that wsan't done in the lecture), most outliers were left after optimization to be ~0.01 percent in the direction as presented in the lecture
- Brand is an important features, there are models that are very rare (and might not be known from training), and Brands differ greatly in price.
- However, there are a lot collinearity between results (some was shown in the course, and some is intuition that needs to be proven):
- Year and Mileage - high
- Brand and Model - very high
- Year and Model - unknown, presumed high
- Mileage and Model - unknown, presumed average
- Solutions for collinearity:
- Not including Year and Mileage together, only one of them. Empirically Year wins
- Not including Brand and Model - however, to make results better, they were made into one feature with their values concatenated
- Following these 2 changes, the coefficients come out decent (and not too high / low which shows overfitting, and including 2 correlated variables)
- It is possible that because of Year and Model are still very correlated, one of them will need to be removed to improve coefficients, and VIFs. This will make the predictions drastically worse.
- In addition, because Model has a lot of rare values, it was attempted to group into Other all rare fields
- It is very possible that with the given data, and using only Linear Regression, it is not possible to create both low collinearity / good coefficients, and good predictions
- Adjusted R2 and R2 gives very close values due to large sample size. Therefore left using R2 and not Adjusted R2.
- Most of the outliers / removing null values remove too big of percentage of the dataset (~10%), needs to be better resolved by putting values instead of deleting rows
| github_jupyter |
The following additional libraries are needed to run this
notebook. Note that running on Colab is experimental, please report a Github
issue if you have any problem.
```
!pip install d2l==0.14.3
```
# Deep Convolutional Neural Networks (AlexNet)
:label:`sec_alexnet`
Although CNNs were well known
in the computer vision and machine learning communities
following the introduction of LeNet,
they did not immediately dominate the field.
Although LeNet achieved good results on early small datasets,
the performance and feasibility of training CNNs
on larger, more realistic datasets had yet to be established.
In fact, for much of the intervening time between the early 1990s
and the watershed results of 2012,
neural networks were often surpassed by other machine learning methods,
such as support vector machines.
For computer vision, this comparison is perhaps not fair.
That is although the inputs to convolutional networks
consist of raw or lightly-processed (e.g., by centering) pixel values, practitioners would never feed raw pixels into traditional models.
Instead, typical computer vision pipelines
consisted of manually engineering feature extraction pipelines.
Rather than *learn the features*, the features were *crafted*.
Most of the progress came from having more clever ideas for features,
and the learning algorithm was often relegated to an afterthought.
Although some neural network accelerators were available in the 1990s,
they were not yet sufficiently powerful to make
deep multichannel, multilayer CNNs
with a large number of parameters.
Moreover, datasets were still relatively small.
Added to these obstacles, key tricks for training neural networks
including parameter initialization heuristics,
clever variants of stochastic gradient descent,
non-squashing activation functions,
and effective regularization techniques were still missing.
Thus, rather than training *end-to-end* (pixel to classification) systems,
classical pipelines looked more like this:
1. Obtain an interesting dataset. In early days, these datasets required expensive sensors (at the time, 1 megapixel images were state-of-the-art).
2. Preprocess the dataset with hand-crafted features based on some knowledge of optics, geometry, other analytic tools, and occasionally on the serendipitous discoveries of lucky graduate students.
3. Feed the data through a standard set of feature extractors such as the SIFT (scale-invariant feature transform) :cite:`Lowe.2004`, the SURF (speeded up robust features) :cite:`Bay.Tuytelaars.Van-Gool.2006`, or any number of other hand-tuned pipelines.
4. Dump the resulting representations into your favorite classifier, likely a linear model or kernel method, to train a classifier.
If you spoke to machine learning researchers,
they believed that machine learning was both important and beautiful.
Elegant theories proved the properties of various classifiers.
The field of machine learning was thriving, rigorous, and eminently useful. However, if you spoke to a computer vision researcher,
you would hear a very different story.
The dirty truth of image recognition, they would tell you,
is that features, not learning algorithms, drove progress.
Computer vision researchers justifiably believed
that a slightly bigger or cleaner dataset
or a slightly improved feature-extraction pipeline
mattered far more to the final accuracy than any learning algorithm.
## Learning Representations
Another way to cast the state of affairs is that
the most important part of the pipeline was the representation.
And up until 2012 the representation was calculated mechanically.
In fact, engineering a new set of feature functions, improving results, and writing up the method was a prominent genre of paper.
SIFT :cite:`Lowe.2004`,
SURF :cite:`Bay.Tuytelaars.Van-Gool.2006`,
HOG (histograms of oriented gradient) :cite:`Dalal.Triggs.2005`,
[bags of visual words](https://en.wikipedia.org/wiki/Bag-of-words_model_in_computer_vision)
and similar feature extractors ruled the roost.
Another group of researchers,
including Yann LeCun, Geoff Hinton, Yoshua Bengio,
Andrew Ng, Shun-ichi Amari, and Juergen Schmidhuber,
had different plans.
They believed that features themselves ought to be learned.
Moreover, they believed that to be reasonably complex,
the features ought to be hierarchically composed
with multiple jointly learned layers, each with learnable parameters.
In the case of an image, the lowest layers might come
to detect edges, colors, and textures.
Indeed,
Alex Krizhevsky, Ilya Sutskever, and Geoff Hinton
proposed a new variant of a CNN,
*AlexNet*,
that achieved excellent performance in the 2012 ImageNet challenge.
AlexNet was named after Alex Krizhevsky,
the first author of the breakthrough ImageNet classification paper :cite:`Krizhevsky.Sutskever.Hinton.2012`.
Interestingly in the lowest layers of the network,
the model learned feature extractors that resembled some traditional filters.
:numref:`fig_filters` is reproduced from the AlexNet paper :cite:`Krizhevsky.Sutskever.Hinton.2012`
and describes lower-level image descriptors.

:width:`400px`
:label:`fig_filters`
Higher layers in the network might build upon these representations
to represent larger structures, like eyes, noses, blades of grass, and so on.
Even higher layers might represent whole objects
like people, airplanes, dogs, or frisbees.
Ultimately, the final hidden state learns a compact representation
of the image that summarizes its contents
such that data belonging to different categories be separated easily.
While the ultimate breakthrough for many-layered CNNs
came in 2012, a core group of researchers had dedicated themselves
to this idea, attempting to learn hierarchical representations of visual data
for many years.
The ultimate breakthrough in 2012 can be attributed to two key factors.
### Missing Ingredient: Data
Deep models with many layers require large amounts of data
in order to enter the regime
where they significantly outperform traditional methods
based on convex optimizations (e.g., linear and kernel methods).
However, given the limited storage capacity of computers,
the relative expense of sensors,
and the comparatively tighter research budgets in the 1990s,
most research relied on tiny datasets.
Numerous papers addressed the UCI collection of datasets,
many of which contained only hundreds or (a few) thousands of images
captured in unnatural settings with low resolution.
In 2009, the ImageNet dataset was released,
challenging researchers to learn models from 1 million examples,
1000 each from 1000 distinct categories of objects.
The researchers, led by Fei-Fei Li, who introduced this dataset
leveraged Google Image Search to prefilter large candidate sets
for each category and employed
the Amazon Mechanical Turk crowdsourcing pipeline
to confirm for each image whether it belonged to the associated category.
This scale was unprecedented.
The associated competition, dubbed the ImageNet Challenge
pushed computer vision and machine learning research forward,
challenging researchers to identify which models performed best
at a greater scale than academics had previously considered.
### Missing Ingredient: Hardware
Deep learning models are voracious consumers of compute cycles.
Training can take hundreds of epochs, and each iteration
requires passing data through many layers of computationally-expensive
linear algebra operations.
This is one of the main reasons why in the 1990s and early 2000s,
simple algorithms based on the more-efficiently optimized
convex objectives were preferred.
*Graphical processing units* (GPUs) proved to be a game changer
in making deep learning feasible.
These chips had long been developed for accelerating
graphics processing to benefit computer games.
In particular, they were optimized for high throughput $4 \times 4$ matrix-vector products, which are needed for many computer graphics tasks.
Fortunately, this math is strikingly similar
to that required to calculate convolutional layers.
Around that time, NVIDIA and ATI had begun optimizing GPUs
for general computing operations,
going as far as to market them as *general-purpose GPUs* (GPGPU).
To provide some intuition, consider the cores of a modern microprocessor
(CPU).
Each of the cores is fairly powerful running at a high clock frequency
and sporting large caches (up to several megabytes of L3).
Each core is well-suited to executing a wide range of instructions,
with branch predictors, a deep pipeline, and other bells and whistles
that enable it to run a large variety of programs.
This apparent strength, however, is also its Achilles heel:
general-purpose cores are very expensive to build.
They require lots of chip area,
a sophisticated support structure
(memory interfaces, caching logic between cores,
high-speed interconnects, and so on),
and they are comparatively bad at any single task.
Modern laptops have up to 4 cores,
and even high-end servers rarely exceed 64 cores,
simply because it is not cost effective.
By comparison, GPUs consist of $100 \sim 1000$ small processing elements
(the details differ somewhat between NVIDIA, ATI, ARM and other chip vendors),
often grouped into larger groups (NVIDIA calls them warps).
While each core is relatively weak,
sometimes even running at sub-1GHz clock frequency,
it is the total number of such cores that makes GPUs orders of magnitude faster than CPUs.
For instance, NVIDIA's recent Volta generation offers up to 120 TFlops per chip for specialized instructions
(and up to 24 TFlops for more general-purpose ones),
while floating point performance of CPUs has not exceeded 1 TFlop to date.
The reason for why this is possible is actually quite simple:
first, power consumption tends to grow *quadratically* with clock frequency.
Hence, for the power budget of a CPU core that runs 4 times faster (a typical number),
you can use 16 GPU cores at $1/4$ the speed,
which yields $16 \times 1/4 = 4$ times the performance.
Furthermore, GPU cores are much simpler
(in fact, for a long time they were not even *able*
to execute general-purpose code),
which makes them more energy efficient.
Last, many operations in deep learning require high memory bandwidth.
Again, GPUs shine here with buses that are at least 10 times as wide as many CPUs.
Back to 2012. A major breakthrough came
when Alex Krizhevsky and Ilya Sutskever
implemented a deep CNN
that could run on GPU hardware.
They realized that the computational bottlenecks in CNNs,
convolutions and matrix multiplications,
are all operations that could be parallelized in hardware.
Using two NVIDIA GTX 580s with 3GB of memory,
they implemented fast convolutions.
The code [cuda-convnet](https://code.google.com/archive/p/cuda-convnet/)
was good enough that for several years
it was the industry standard and powered
the first couple years of the deep learning boom.
## AlexNet
AlexNet, which employed an 8-layer CNN,
won the ImageNet Large Scale Visual Recognition Challenge 2012
by a phenomenally large margin.
This network showed, for the first time,
that the features obtained by learning can transcend manually-designed features, breaking the previous paradigm in computer vision.
The architectures of AlexNet and LeNet are very similar,
as :numref:`fig_alexnet` illustrates.
Note that we provide a slightly streamlined version of AlexNet
removing some of the design quirks that were needed in 2012
to make the model fit on two small GPUs.

:label:`fig_alexnet`
The design philosophies of AlexNet and LeNet are very similar,
but there are also significant differences.
First, AlexNet is much deeper than the comparatively small LeNet5.
AlexNet consists of eight layers: five convolutional layers,
two fully-connected hidden layers, and one fully-connected output layer. Second, AlexNet used the ReLU instead of the sigmoid
as its activation function.
Let us delve into the details below.
### Architecture
In AlexNet's first layer, the convolution window shape is $11\times11$.
Since most images in ImageNet are more than ten times higher and wider
than the MNIST images,
objects in ImageNet data tend to occupy more pixels.
Consequently, a larger convolution window is needed to capture the object.
The convolution window shape in the second layer
is reduced to $5\times5$, followed by $3\times3$.
In addition, after the first, second, and fifth convolutional layers,
the network adds maximum pooling layers
with a window shape of $3\times3$ and a stride of 2.
Moreover, AlexNet has ten times more convolution channels than LeNet.
After the last convolutional layer there are two fully-connected layers
with 4096 outputs.
These two huge fully-connected layers produce model parameters of nearly 1 GB.
Due to the limited memory in early GPUs,
the original AlexNet used a dual data stream design,
so that each of their two GPUs could be responsible
for storing and computing only its half of the model.
Fortunately, GPU memory is comparatively abundant now,
so we rarely need to break up models across GPUs these days
(our version of the AlexNet model deviates
from the original paper in this aspect).
### Activation Functions
Besides, AlexNet changed the sigmoid activation function to a simpler ReLU activation function. On one hand, the computation of the ReLU activation function is simpler. For example, it does not have the exponentiation operation found in the sigmoid activation function.
On the other hand, the ReLU activation function makes model training easier when using different parameter initialization methods. This is because, when the output of the sigmoid activation function is very close to 0 or 1, the gradient of these regions is almost 0, so that backpropagation cannot continue to update some of the model parameters. In contrast, the gradient of the ReLU activation function in the positive interval is always 1. Therefore, if the model parameters are not properly initialized, the sigmoid function may obtain a gradient of almost 0 in the positive interval, so that the model cannot be effectively trained.
### Capacity Control and Preprocessing
AlexNet controls the model complexity of the fully-connected layer
by dropout (:numref:`sec_dropout`),
while LeNet only uses weight decay.
To augment the data even further, the training loop of AlexNet
added a great deal of image augmentation,
such as flipping, clipping, and color changes.
This makes the model more robust and the larger sample size effectively reduces overfitting.
We will discuss data augmentation in greater detail in :numref:`sec_image_augmentation`.
```
from d2l import torch as d2l
import torch
from torch import nn
net = nn.Sequential(
# Here, we use a larger 11 x 11 window to capture objects. At the same
# time, we use a stride of 4 to greatly reduce the height and width of the
# output. Here, the number of output channels is much larger than that in
# LeNet
nn.Conv2d(1, 96, kernel_size=11, stride=4, padding=1), nn.ReLU(),
nn.MaxPool2d(kernel_size=3, stride=2),
# Make the convolution window smaller, set padding to 2 for consistent
# height and width across the input and output, and increase the number of
# output channels
nn.Conv2d(96, 256, kernel_size=5, padding=2), nn.ReLU(),
nn.MaxPool2d(kernel_size=3, stride=2),
# Use three successive convolutional layers and a smaller convolution
# window. Except for the final convolutional layer, the number of output
# channels is further increased. Pooling layers are not used to reduce the
# height and width of input after the first two convolutional layers
nn.Conv2d(256, 384, kernel_size=3, padding=1), nn.ReLU(),
nn.Conv2d(384, 384, kernel_size=3, padding=1), nn.ReLU(),
nn.Conv2d(384, 256, kernel_size=3, padding=1), nn.ReLU(),
nn.MaxPool2d(kernel_size=3, stride=2),
nn.Flatten(),
# Here, the number of outputs of the fully-connected layer is several
# times larger than that in LeNet. Use the dropout layer to mitigate
# overfitting
nn.Linear(6400, 4096), nn.ReLU(),
nn.Dropout(p=0.5),
nn.Linear(4096, 4096), nn.ReLU(),
nn.Dropout(p=0.5),
# Output layer. Since we are using Fashion-MNIST, the number of classes is
# 10, instead of 1000 as in the paper
nn.Linear(4096, 10))
```
We construct a single-channel data example with both height and width of 224 to observe the output shape of each layer. It matches the AlexNet architecture in :numref:`fig_alexnet`.
```
X = torch.randn(1, 1, 224, 224)
for layer in net:
X=layer(X)
print(layer.__class__.__name__,'Output shape:\t',X.shape)
```
## Reading the Dataset
Although AlexNet is trained on ImageNet in the paper, we use Fashion-MNIST here
since training an ImageNet model to convergence could take hours or days
even on a modern GPU.
One of the problems with applying AlexNet directly on Fashion-MNIST
is that its images have lower resolution ($28 \times 28$ pixels)
than ImageNet images.
To make things work, we upsample them to $224 \times 224$
(generally not a smart practice,
but we do it here to be faithful to the AlexNet architecture).
We perform this resizing with the `resize` argument in the `d2l.load_data_fashion_mnist` function.
```
batch_size = 128
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size, resize=224)
```
## Training
Now, we can start training AlexNet.
Compared with LeNet in :numref:`sec_lenet`,
the main change here is the use of a smaller learning rate
and much slower training due to the deeper and wider network,
the higher image resolution, and the more costly convolutions.
```
lr, num_epochs = 0.01, 10
d2l.train_ch6(net, train_iter, test_iter, num_epochs, lr)
```
## Summary
* AlexNet has a similar structure to that of LeNet, but uses more convolutional layers and a larger parameter space to fit the large-scale ImageNet dataset.
* Today AlexNet has been surpassed by much more effective architectures but it is a key step from shallow to deep networks that are used nowadays.
* Although it seems that there are only a few more lines in AlexNet's implementation than in LeNet, it took the academic community many years to embrace this conceptual change and take advantage of its excellent experimental results. This was also due to the lack of efficient computational tools.
* Dropout, ReLU, and preprocessing were the other key steps in achieving excellent performance in computer vision tasks.
## Exercises
1. Try increasing the number of epochs. Compared with LeNet, how are the results different? Why?
1. AlexNet may be too complex for the Fashion-MNIST dataset.
1. Try simplifying the model to make the training faster, while ensuring that the accuracy does not drop significantly.
1. Design a better model that works directly on $28 \times 28$ images.
1. Modify the batch size, and observe the changes in accuracy and GPU memory.
1. Analyze computational performance of AlexNet.
1. What is the dominant part for the memory footprint of AlexNet?
1. What is the dominant part for computation in AlexNet?
1. How about memory bandwidth when computing the results?
1. Apply dropout and ReLU to LeNet-5. Does it improve? How about preprocessing?
[Discussions](https://discuss.d2l.ai/t/76)
| github_jupyter |
import sys
import nltk
import sklearn
import pandas
import numpy
print('Python: {}'.format(sys.version))
print('NLTK: {}'.format(nltk.__version__))
print('Scikit-learn: {}'.format(sklearn.__version__))
print('Pandas: {}'.format(pandas.__version__))
print('Numpy: {}'.format(numpy.__version__))
```
import pandas as pd
import numpy as np
df = pd.read_table('SMSSPamCollection', header=None, encoding='utf-8')
print(df.info())
print(df.head())
classes = df[0]
print(classes.value_counts())
from sklearn.preprocessing import LabelEncoder
encoder = LabelEncoder()
Y = encoder.fit_transform(classes)
print(Y[:10])
text_messages = df[1]
print(text_messages[:10])
processed = text_messages.str.replace(r'^.+@[^\.].*\.[a-z]{2,}$',
'emailaddress')
print(processed[:10])
processed = processed.str.replace(r'^http\://[a-zA-Z0-9\-\.]+\.[a-zA-Z]{2,3}(/\S*)?$',
'webaddress')
processed = processed.str.replace(r'£|\$', 'moneysymb')
processed = processed.str.replace(r'^\(?[\d]{3}\)?[\s-]?[\d]{3}[\s-]?[\d]{4}$',
'phonenumbr')
processed = processed.str.replace(r'\d+(\.\d+)?', 'numbr')
processed = processed.str.replace(r'[^\w\d\s]', ' ')
processed = processed.str.replace(r'\s+', ' ')
processed = processed.str.replace(r'^\s+|\s+?$', '')
processed = processed.str.lower()
print(processed)
from nltk.corpus import stopwords
stop_words = set(stopwords.words('english'))
from nltk.corpus import stopwords
stop_words = set(stopwords.words('english'))
processed = processed.apply(lambda x: ' '.join(term for term in x.split() if term not in stop_words))
import nltk
nltk.download('stopwords')
nltk.download('stopwords')
stop_words = set(stopwords.words('english'))
processed = processed.apply(lambda x: ' '.join(
term for term in x.split() if term not in stop_words))
ps = nltk.PorterStemmer()
processed = processed.apply(lambda x: ' '.join(
ps.stem(term) for term in x.split()))
print (processed)
from nltk.tokenize import word_tokenize
# create bag-of-words
all_words = []
for message in processed:
words = word_tokenize(message)
for w in words:
all_words.append(w)
all_words = nltk.FreqDist(all_words)
print(all_words)
nltk.download(punkt)
nltk.download('punkt')
all_words = []
for message in processed:
words = word_tokenize(message)
for w in words:
all_words.append(w)
all_words = nltk.FreqDist(all_words)
print(all_words)
print('Number of words: {}'.format(len(all_words)))
print('Most common words: {}'.format(all_words.most_common(20)))
word_features = list(all_words.keys())[:1500]
print(word_features)
def find_features(message):
words = word_tokenize(message)
features = {}
for word in word_features:
features[word] = (word in words)
return features
features = find_features(processed[1])
for key, value in features.items():
if value == True:
print (key)
messages = list(zip(processed, Y))
seed = 1
np.random.seed = seed
np.random.shuffle(messages)
featuresets = [(find_features(text), label) for (text, label) in messages]
from sklearn import model_selection
training, testing = model_selection.train_test_split(featuresets, test_size = 0.25, random_state=seed)
print(len(training))
print(len(testing))
from nltk.classify.scikitlearn import SklearnClassifier
from sklearn.svm import SVC
model = SklearnClassifier(SVC(kernel = 'linear'))
model.train(training)
accuracy = nltk.classify.accuracy(model, testing)*100
print("SVC Accuracy: {}".format(accuracy))
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression, SGDClassifier
from sklearn.naive_bayes import MultinomialNB
from sklearn.svm import SVC
from sklearn.metrics import classification_report, accuracy_score, confusion_matrix
names = ["K Nearest Neighbors", "Decision Tree", "Random Forest", "Logistic Regression", "SGD Classifier",
"Naive Bayes", "SVM Linear"]
classifiers = [
KNeighborsClassifier(),
DecisionTreeClassifier(),
RandomForestClassifier(),
LogisticRegression(),
SGDClassifier(max_iter = 100),
MultinomialNB(),
SVC(kernel = 'linear')
]
models = zip(names, classifiers)
for name, model in models:
nltk_model = SklearnClassifier(model)
nltk_model.train(training)
accuracy = nltk.classify.accuracy(nltk_model, testing)*100
print("{} Accuracy: {}".format(name, accuracy))
from sklearn.ensemble import VotingClassifier
names = ["K Nearest Neighbors", "Decision Tree", "Random Forest", "Logistic Regression", "SGD Classifier",
"Naive Bayes", "SVM Linear"]
classifiers = [
KNeighborsClassifier(),
DecisionTreeClassifier(),
RandomForestClassifier(),
LogisticRegression(),
SGDClassifier(max_iter = 100),
MultinomialNB(),
SVC(kernel = 'linear')
]
models = list(zip(names, classifiers))
nltk_ensemble = SklearnClassifier(VotingClassifier(estimators = models, voting = 'hard', n_jobs = -1))
nltk_ensemble.train(training)
accuracy = nltk.classify.accuracy(nltk_model, testing)*100
print("Voting Classifier: Accuracy: {}".format(accuracy))
txt_features, labels = zip(*testing)
prediction = nltk_ensemble.classify_many(txt_features)
print(classification_report(labels, prediction))
pd.DataFrame(
confusion_matrix(labels, prediction),
index = [['actual', 'actual'], ['ham', 'spam']],
columns = [['predicted', 'predicted'], ['ham', 'spam']])
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from sklearn import neighbors, datasets
import seaborn as sns
import pandas as pd
from sklearn import neighbors, datasets
from sklearn.tree import DecisionTreeClassifier
from sklearn import neighbors, datasets
from helpers import plotPairwiseDecisionTrees
```
# Lecture 9: Kaggle 2 kNN, Decision Trees, Random Forest, Ensemble Learning
## 11/6/18
### Hosted by and maintained by the [Statistics Undergraduate Students Association (SUSA)](https://susa.berkeley.edu). Updated and maintained by [Ajay Raj](mailto:araj@berkeley.edu)
#### Originally authored by [Calvin Chen](mailto:chencalvin99@berkeley.edu), [Michelle Hao](mailto:mhao@berkeley.edu), and [Patrick Chao](mailto:prc@berkeley.edu).
## Classification
Decision trees rely a series of yes or no questions to make a decision on which class an input point falls under. You've seen decision trees your entire life. Here's one made by Rosa Choe, a beloved member of the club that graduated last semester.
<img src='pictures/meme.png' width=40%>
Meme credit to $\text{Rosa Choe}^{\text{TM}}$.
As we can see from the tree above, we are able to answer a yes or no question at every step, and depending on our answer, we either went one way or another through the tree. They are very synonymous to flowcharts, but we'll go into more intricacies with decision trees later on.
Now let's apply this to the data science setting for a classification task. In particular, you're given a data point $X = \begin{bmatrix} X_1 & X_2 & ... & X_k \end{bmatrix}$, and you want to assign it a class $c$. We've seen examples of this before: logistic regression from last week tries to assign a class $c \in \{0, 1\}$ for each data point by predicting $\mathbb{P}(X = 1)$.
For a decision tree to work for this, we want to look at $X$, ask yes-no questions about its features, and assign it to a class.
<a id='dataset'></a>
### The Dataset
<img src='pictures/iris.jpg' width="250" height="250">
<center> Image from: A Complete Guide to K-Nearest-Neighbors by Zakka </center>
The dataset we'll be using is the [Iris Flower Dataset](https://archive.ics.uci.edu/ml/datasets/Iris). It contains a series of observations on three species of Iris (Iris setosa, Iris virginica, and Iris verisolor). Each observation contains four features: the *petal length, petal width, sepal length, and sepal width*. The **question** we're asking today is: can we predict the species of Iris from its *petal length, petal width, sepal length, and sepal width*.
```
#importing the data
iris = datasets.load_iris()
iris = pd.DataFrame(data= np.c_[iris['data'], iris['target']],
columns= ['Sepal Length', 'Sepal Width','Petal Length','Petal Width'] + ['species'])
#y contains the correct classifications (0, 1, 2 for each type of Iris)
Y = iris["species"]
```
#### Summarize the dataset
```
iris.describe()
# Let’s now take a look at the number of instances (rows) that
# belong to each class. We can view this as an absolute count.
iris.groupby('species').size()
```
#### Dividing the Dataframe into Feature and Labels
```
feature_columns = ['Sepal Length', 'Sepal Width','Petal Length','Petal Width']
X = iris[feature_columns].values
Y = iris['species'].values
# Alternative way of selecting features and labels arrays:
# X = dataset.iloc[:, 1:5].values
# y = dataset.iloc[:, 5].values
```
#### Splitting the Data into Train and Test Sets
```
#Splitting dataset into training and test
from sklearn.model_selection import train_test_split
X_train_iris, X_test_iris, Y_train_iris, Y_test_iris = train_test_split(X, Y, test_size = 0.2, random_state = 0)
```
An example decision tree to solve this **classification** task could look as follows:
<img src='pictures/Example Decision Tree.png'>
Let's see how this decision tree fares on our training data.
```
class TreeNode:
def __init__(self, left=None, right=None, split_fn=None, leaf_evaluate=None):
self.left = left
self.right = right
self.split_fn = split_fn
self.leaf_evaluate = leaf_evaluate
def is_leaf(self):
return self.left == None and self.right == None
def evaluate(self, X_i):
if self.is_leaf():
return self.leaf_evaluate()
if self.split_fn(X_i):
return self.left.evaluate(X_i)
else:
return self.right.evaluate(X_i)
class Leaf(TreeNode):
def __init__(self, label):
TreeNode.__init__(self, leaf_evaluate=lambda: label)
def accuracy(y_pred, y_true):
return (y_pred == y_true).sum() / y_true.shape[0]
def predict(X, tree):
if len(X.shape) == 1:
X = X.reshape(1, -1)
preds = np.zeros(X.shape[0])
for i in range(X.shape[0]):
preds[i] = tree.evaluate(X[i])
return preds
root = TreeNode(
split_fn=lambda X_i: X_i[0] > 5,
left=TreeNode(
split_fn=lambda X_i: X_i[2] > 3,
left=Leaf(0),
right=Leaf(2)
),
right=TreeNode(
split_fn=lambda X_i: X_i[3] > 3,
left=Leaf(0),
right=Leaf(2)
)
)
preds = predict(X_train_iris, root)
preds[:5]
accuracy(preds, Y_train_iris)
```
This decision tree is horrible! Maybe it's because we didn't try to train it on our data.
### Training a Decision Tree
The question is now: how do we choose how to make the splits? The answer, of course, comes from our training data.
To make things a little simpler, let's just examine the first ten data points.
```
X_train_small = X_train_iris[:10]
Y_train_small = Y_train_iris[:10]
X_train_small
Y_train_small
```
Say that our first split is based on sepal length (the first feature).
```
sl_0 = X_train_small[Y_train_small == 0][:, 0]
sl_1 = X_train_small[Y_train_small == 1][:, 0]
sl_2 = X_train_small[Y_train_small == 2][:, 0]
sl_0
sl_1
sl_2
```
For our decision tree, how should we split on sepal length?
Just based on our training data, if we split on (Sepal Length > 6), we've isolated all irises that are class 2 (iris verisolor).
<img src='pictures/Simple Decision Tree.png'/>
I decided make the "No" choice "Iris virginica". Why do you think that is the case?
```
simple = TreeNode(
split_fn=lambda X_i: X_i[0] > 6,
left=Leaf(2),
right=Leaf(1)
)
preds = predict(X_train_small, simple)
preds
accuracy(preds, Y_train_small)
```
Pretty good! Now let's try to come up with a programmatic way of doing this.
The intuition behind making a good decision tree is optimizing our questions (or different steps in the decision tree) to be able *to split up the data into as different categories as possible*. For example in the iris case, we would like to find a split where we may separate the various irises as much as possible.
This idea of "splitting" to separate our irises the most introduces the idea of **entropy**. We minimize the entropy, or randomness in each split section of the data.
<a id='entropy'></a>
### Entropy
To begin, let's first define what entropy is. In the context of machine learning, entropy is **the measure of disorder within a set** or the **amount of surprise**.
Let's take a look at our training data, and the feature we chose to split on, **sepal length**.
```
sepal_length = X_train_small[:, 0]
sepal_length
Y_train_small
```
After we split on (Sepal Length > 6), we divided our data into two halves.
```
yes = Y_train_small[sepal_length > 6]
no = Y_train_small[sepal_length <= 6]
yes
no
```
Let's consider a different split: (Sepal Length > 5.5).
```
yes_bad = Y_train_small[sepal_length > 5.5]
no_bad = Y_train_small[sepal_length <= 5.5]
yes_bad
no_bad
```
Which split was better? The first, because once we made the split, *we were more sure of what class we should predict*. How can we quantify this?
The mathematical definition of entropy is:
$$H(\textbf{p}) = -\sum_i p_i \cdot \log(p_i)$$
where $H(\textbf{p})$ is equal to the total entropy of the data set, and $p_i$ is equal to the probability of something occurring.
**Something to note:** When you calculate the entropy on a subset with data points that are all the same class, you run into a mathematical error, which is because $\log_{2}(0)$ cannot be calculated. So, as an alternative to calculating $\log_{2}(0)$, we can bring in the following limit instead (meaning our terms with $p_i = 0$ become 0:
$$\lim _{p\to 0+}p\log(p)=0$$
A great visualization for different entropies is as follows:
<img src='pictures/Entropy.png' width='50%'>
Let's say $Pr(X = 1)$ is the probability that you flips a heads, where heads is represented by $1$ and tails is represented by $0$. From this, we can see that the y-value, $H(X)$ (or calculated entropy), is at a minimum when the chance of flipping a heads is $0$ or $1$, but is at a maximum when the chance of flipping a heads is $0.5$. In other words, the data subset is the most random when there is an equal probability of all classes, and minimized when there are probabilites of classes that are equal to $0$.
When we look at a set of y-values, the entropy is:
$$\sum_{\text{class $c_i$}} -\left(\text{proportion of $c_i$'s}\right) \cdot \log \left(\text{proportion of $c_i$'s}\right)$$
```
def H(y):
def proportion(val, y):
return (y == val).sum() / len(y)
unique = set(y)
return sum(-1 * proportion(val, y) * np.log2(proportion(val, y)) for val in unique)
```
Let's see how this comes into play in our splits.
```
original_entropy = H(Y_train_small)
original_entropy
```
In our good split, our entropies were:
```
H(yes), H(no)
```
In our bad split, our entropies were:
```
H(yes_bad), H(no_bad)
```
Clearly, the first split was better, because we reduced entropy the most.
To combine these statistics together for one measure, we'll take the **weighted average**, weighting by the sizes of the two sets.
```
def weighted_entropy(yes, no):
total_size = len(yes) + len(no)
return (len(yes) / total_size) * H(yes) + (len(no) / total_size) * H(no)
H(Y_train_small)
weighted_entropy(yes, no)
weighted_entropy(yes_bad, no_bad)
```
This is huge! We now have a way to choose our splits for our decision tree:
**Find the best split value (of each feature) that reduces our entropy from the original set the most!**
### Training
```
from scipy.stats import mode
def train(X_train, Y_train, max_depth=None):
if len(Y_train) == 0:
return Leaf(0)
if len(set(Y_train)) == 1 or max_depth == 1:
return Leaf(mode(Y_train).mode)
def split_weighted_entropy(feature_idx, feature_value):
feature = X_train[:, feature_idx]
yes = Y_train[feature > feature_value]
no = Y_train[feature <= feature_value]
return weighted_entropy(yes, no)
splits = np.zeros(X_train.shape)
for feature_idx in range(X_train.shape[1]):
for i, feature_value in enumerate(X_train[:, feature_idx]): # try to split on each X-value
splits[i, feature_idx] = split_weighted_entropy(feature_idx, feature_value)
max_idxs = X_train.argmax(axis=0)
for col, max_idx in enumerate(max_idxs):
splits[max_idx, col] = float('inf')
i = np.argmin(splits)
best_feature_idx = i % splits.shape[1]
best_feature_value = X_train[i // splits.shape[1], best_feature_idx]
yes = X_train[:, best_feature_idx] > best_feature_value
no = X_train[:, best_feature_idx] <= best_feature_value
tree = TreeNode(
split_fn=lambda X_i: X_i[best_feature_idx] > best_feature_value,
left=train(X_train[yes], Y_train[yes], max_depth=max_depth - 1 if max_depth is not None else None),
right=train(X_train[no], Y_train[no], max_depth=max_depth - 1 if max_depth is not None else None)
)
return tree
tree = train(X_train_iris, Y_train_iris)
preds = predict(X_train_iris, tree)
accuracy(preds, Y_train_iris)
```
Whoa! We have a model that performs at 100% training accuracy! Let's see what happens when we try the model on the validation set.
```
preds = predict(X_test_iris, tree)
accuracy(preds, Y_test_iris)
```
We're doing significantly worse, so we're probably overfitting.
### Regression
How can we use decision trees to perform regression?
When we decide to make a leaf, take the mean/median of the points that are left, instead of the mode.
### Titanic
```
titanic_train = pd.read_csv('titanic/train.csv')
titanic_test = pd.read_csv('titanic/test.csv')
titanic_survived = titanic_train['Survived']
titanic_train = titanic_train.drop('Survived', axis=1)
full_data = pd.concat((titanic_train, titanic_test), sort=False)
def feature_engineering(df):
df = df.drop(['Name', 'PassengerId', 'Age', 'Ticket'], axis=1)
df['Embarked'] = df['Embarked'].fillna(df['Embarked'].value_counts().idxmax()) # fill with Southampton
df['Deck'] = df['Cabin'].apply(lambda s: s[0] if s is not np.nan else 'U') # U for unassigned
df['Fare'] = np.log(df['Fare'] + 1)
df = df.drop('Cabin', axis=1)
df = pd.get_dummies(df, columns=['Pclass', 'Embarked', 'Sex', 'Deck'])
df = df.drop('Deck_T', axis=1)
return df
full_data_cleaned = feature_engineering(full_data)
full_data_cleaned.head()
titanic_train_cleaned = full_data_cleaned.iloc[:titanic_train.shape[0]]
titanic_test_cleaned = full_data_cleaned.iloc[titanic_train.shape[0]:]
X = titanic_train_cleaned.values
y = titanic_survived.values
X_train_titanic, X_valid_titanic, y_train_titanic, y_valid_titanic = train_test_split(X, y, test_size=0.2, random_state=42)
tree = train(X_train_titanic, y_train_titanic.astype(int), max_depth=10) # for computation sake
preds = predict(X_train_titanic, tree)
accuracy(preds, y_train_titanic)
preds = predict(X_valid_titanic, tree)
accuracy(preds, y_valid_titanic)
```
We're clearly overfitting. Let's investigate why this is the case.
In a decision tree, if we group each of the training points into the leaves that they would be classified as, a **pure leaf** is one that points all of the same task.
The first fact to recognize is that it is always possible to get 100% training accuracy using a decision tree. How? Make every data point a leaf / make every leaf **pure**.
<img src="pictures/DecisionTreeError.png" width="60%">
Image from http://www.cs.cornell.edu/courses/cs4780/2017sp/lectures/lecturenote17.html
What is wrong with making every leaf pure?
It doesn't generalize well to test points. The **decision boundaries** become too complicated.
### Controlling Overfitting
There are many ways to control overfitting in decision trees: today we'll talk about **max-depth**.
The **max-depth** parameter allows your tree to be cut off at a certain depth, which reduces overfitting by making the decision boundaries simpler.
```
tree = train(X_train_titanic, y_train_titanic.astype(int), max_depth=4)
preds = predict(X_train_titanic, tree)
accuracy(preds, y_train_titanic)
preds = predict(X_valid_titanic, tree)
accuracy(preds, y_valid_titanic)
```
#### Cross-Validation
**Max-depth** is called a *hyperparameter* in your model (a parameter for your training, but one that cannot be learned from the training data). There are many ways to find the optimal values for these *hyperparameters*. Today we'll be discussing one, **$k$-fold cross-validation**.
<img src="pictures/k-fold.png"/>
In this procedure, we isolate $\frac{1}{k}$ of the data as the temporary test set, train on the other data, and evaluate the model that is trained. We do this for each of the $k$ possibilities of test set, and look at the mean accuracy.
For each choice of hyperparameter, look at the mean accuracy over all possible folds and choose the choice that has the highest mean accuracy. Why does this do better than just trying all choices on one validation set: it makes the model more **robust**, and less dependent on one validation set.
```
# 5-Fold Cross Validation
from sklearn.model_selection import KFold
kf = KFold(n_splits=5)
max_depths = [6, 8, 10]
cv_scores = np.zeros(len(max_depths))
for i, max_depth in enumerate(max_depths):
print('Training max_depth =', max_depth, end='\t')
scores = np.zeros(5)
for j, (train_index, test_index) in enumerate(kf.split(X_train_titanic)):
train_X = X_train_titanic[train_index]
train_y = y_train_titanic[train_index]
valid_X = X_train_titanic[test_index]
valid_y = y_train_titanic[test_index]
tree = train(train_X, train_y.astype(int), max_depth=max_depth)
preds = predict(valid_X, tree)
scores[j] = accuracy(preds, valid_y)
cv_scores[i] = scores.mean()
print('accuracy = ', cv_scores[i])
```
Below are a few examples of different decision trees. Let's continue to investigate the effect of the **max-depth** parameter.
```
# From http://scikit-learn.org/stable/auto_examples/tree/plot_iris.html
# Max Depth 2
plotPairwiseDecisionTrees(2)
# Max Depth 4
plotPairwiseDecisionTrees(4)
# No Max Depth
plotPairwiseDecisionTrees()
```
**Questions for understanding:**
> 1. What do the colored points and regions represent?
> 2. What are some trends as the trees get deeper?
> 3. How do the decision boundaries change with depth?
As we can see, the deeper our tree goes, the higher the variance is within the tree, as the decision tree is tailored towards our training data, and could be completely different had we just added/removed a couple data points. However, it also has low bias, as it won't consistently classify certain data points incorrectly (it's too precise!)
Now, we'd love to have a decision tree that had both low bias and low variance, but it seems like it's a tradeoff for one or the other. So, it'd be ideal to get the best of both worlds, and get low bias and low variance. But how?
Idea: **What if we got more models trained on our dataset?**
This idea of training more models on our training set introduces the idea of **ensemble learning**, which we will go into further in the next section, and help us solve our dilemna of wanting both low bias and low variance!
<a id='ensemble_learning'></a>
# Ensemble Learning
<img src='pictures/elephant.jpeg' width="700" height="700">
This is the fable of the blind men and the elephant. Each individual is correct in their own right, however together their descriptions paint a much more accurate picture.
We have discussed notions of bias and variance. To refresh these concepts again,
**Bias** is how well the average model would perform if you trained models on many data sets.
**Variance** is how different the models you would obtain if you trained models on many data sets.
In practice, we only have one data set. If we train a model on this dataset, we would like to minimize both bias and variance. However, we can use some techniques to try to get the best of both worlds.
Since bias is talking about how well the average model performs, and variance is about how varied the different models are, we can attempt to minimizing both of these by considering an *average model*.
Consider the following analogy.
<img src='pictures/weather.jpg' width="700" height="700">
We would like to predict the weather tomorrow. Perhaps we have $3$ separate sources for weather, Channel $4$ on TV, a online website, and the iPhone weather application.
We may expect that a better estimate for the weather tomorrow is actually the average of all these estimates. Perhaps the different sources all have their own methods and data for creating a prediction, thus taking the average pools together all their resources into a more powerful estimator.
The important gain of this approach is our improvement in variance. Keep in mind this is mentioning how different would another similar estimator be. While a single source may have high variation, such as an online website, we would expect another averaged weather amalgamation would be similar. If we considered Channel $5$ predictions, a different online website, and the Android weather application, we would not expect as much variation between their predictions since we already took the average of multiple sources.
Thus, one technique to improve the quality of a model is to *train multiple models on the same data and pool their predictions*. This is known as **ensembling**.
<a id='bootstrapping_and_bagging'></a>
# Bootstrapping and Bagging
An important idea often used in data science is **bootstrapping**. This is a method to generate more samples of data, despite the fact that we only have a single dataset.
**Bootstrapping**: We take the original dataset of size $N$ and draw $M$ samples with replacement of size $N$.
For example, if we would like to estimate the average height of people in the U.S., we may take a sample of $1000$ people and average their heights. However, this does not tell us much about the data other than the average. We pooled together $1000$ data points into a single value, but there is much more information available.
What we can do is draw many samples with replacement of size $1000$, and compute the average heights of these. This mimics as if we had many dataset, and we have many average heights. Then we can compute a distribute of average heights we would have collected, and from this we can determine how good our estimate is. By the Central Limit theorem, this distribution of bootstrapped statistics should approach the normal distribution.
However, we are not limited to just calculating the average. We may calculate the median, standard deviation, or any other statistic of each bootstrapped sample. Furthermore, we can even create a model for each sample! This allows us to utilize the notion of training many models on the same dataset.
If we create many models and then aggregate our predictions, this is known as **bagging** (bootstrapp aggregating). Thus we may create many separate models that all are trained on separate data to obtain new predictions and better results.
The purpose of bagging is to decrease the variance of our model. Since we essentially take many models together in parallel, we avoid overfitting and decreasing variance.
<a id='random_forest'></a>
# Random Forest
A single decision tree often results in poor predictions, as they are very simple and lack much complexity. Just cutting the feature space into separate regions does not perform very well, as these strict linear constraints prevent complex boundaries.
However, if we include many decision trees and create a **random forest**, we obtain drastically better results.
The idea of a random forest is quite simple, take many decision trees and output their average guess.
<center>
<img src="pictures/RandomForest.png" width="60%">
Image from https://medium.com/@williamkoehrsen/random-forest-simple-explanation-377895a60d2d
</center>
In the case of classification, we take the most popular guess.
In the case of regression, we take some form of the average guess.
Now, let's consider this exact setup. What would happen if we created many decision trees and took the most popular guess?
In practice, we could obtain the same decision tree over and over. This is because there is some optimal set of splitting values in the dataset to minimize entropy, even with different sets of data. Perhaps we have one feature that works very well in splitting the data, and it is always utilized as the first split in the tree. Then all decision trees end up looking quite similar, despite our efforts in bagging.
A solution to this problem is feature bagging. We may also select a subset of features for each tree to train on, thus each feature has a chance to be split on.
<center>
<img src="pictures/RandomForestPipeline.jpg" width="40%">
Image from https://sites.google.com/site/rajhansgondane2506/publications
</center>
In summary, we begin with a dataset $\mathcal{D}$ of size $N$.
1. We bootstrap the data so that we have $M$ new datasets $d_1,\ldots, d_M$ drawn with replacement of size $N$ from $\mathcal{D}$.
2. Select a subset of features $f_i$ for each new dataset $d_i$.
3. Fit a decision tree for each $d_i$ with features $f_i$.
Now to predict, we take the input data and feed it through each decision tree to get an output. Then we can take the most popular vote or the average output as the output of our model, based on the type of problem we are attempting to solve.
```
from sklearn.ensemble import RandomForestClassifier
model = RandomForestClassifier(n_estimators=200, max_depth=10)
model.fit(X_train_titanic, y_train_titanic)
accuracy(model.predict(X_valid_titanic), y_valid_titanic)
```
<a id='boosting'></a>
# Boosting
We have mentioned bagging as a method of decreasing variance, but what about bias? There are also techniques to do this, namely **boosting**. This is a very popular technique in Kaggle competitions and most models that win competitions utilize huge ensembles of boosted random forests.
The exact implementation of boosting is out of scope for this discussion, but the main idea is to *fit your models sequentially rather than in parallel in bagging*.
In random forest, we take many samples of our data, and fit separate decision trees to each one. We account for similar decision trees by feature bagging as well. However, many of these decision trees will end up predicting similar things and essentially only reduce variance.
The key idea of boosting is to **emphasize the specific data points that we fail on**. Rather than trying to improve the prediction by considering the problem from different angles (e.g. new datasets from bagging or new features from feature bagging), consider where we predict incorrectly and attempt to improve our predictions from there.
<center>
<img src="pictures/Boosting.png" width="70%">
Image from https://quantdare.com/what-is-the-difference-between-bagging-and-boosting/
</center>
This is similar to real life. If you would like to learn a new musical piece, it is more beneficial to practice the specific part that is challenging, rather than playing the entire piece from the start every time you mess up. By boosting our model, we attempt to place greater emphasis on the samples that we consistently misclassify.
For further reading, we highly recommend the following resources for explanations for boosting:
https://quantdare.com/what-is-the-difference-between-bagging-and-boosting/
https://medium.com/mlreview/gradient-boosting-from-scratch-1e317ae4587d
Spectacular visualizations of decision trees and boosting:
http://arogozhnikov.github.io/2016/06/24/gradient_boosting_explained.html
There are many forms of boosting in practice. Popular ones include: Adaboost, GradientBoost, and XGBoost. XGBoost is famous (or infamous) for excelling at Kaggle competitions, most winning solutions contain XGBoost (if not many). We encourage you to look at the links above (especially the vis
| github_jupyter |
<a href="https://colab.research.google.com/github/RaviGprec/Machine-Learning/blob/master/nlp_text_classification_part_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#Text Classification
Reference:
[https://github.com/miguelfzafra/Latest-News-Classifier](https://github.com/miguelfzafra/Latest-News-Classifier)
```
import pickle
import pandas as pd
import nltk
from nltk.corpus import stopwords
from nltk.tokenize import punkt
from nltk.corpus.reader import wordnet
from nltk.stem import WordNetLemmatizer
#from sklearn.feature_extraction.text import TfidfVectorizer
import requests
from bs4 import BeautifulSoup
import numpy as np
import plotly.graph_objs as go
import re
import pandas as pd
import matplotlib.pyplot as plt
import pickle
import seaborn as sns
sns.set_style("whitegrid")
# Code for hiding seaborn warnings
import warnings
warnings.filterwarnings("ignore")
import pickle
#df = pd.read_csv("News_dataset.csv",sep = ";", encoding = "utf-8")
#df.head()
with open("News_dataset.pickle", 'rb') as data:
df = pickle.load(data)
df.head()
df.loc[1]['Content']
with open("best_svc.pickle","rb") as model:
svc_model = pickle.load(model)
with open("tfidf.pickle","rb") as data:
tfidf = pickle.load(data)
import nltk
from nltk.corpus import stopwords
nltk.download("stopwords")
from nltk.stem import WordNetLemmatizer
nltk.download('wordnet')
punctuation_signs = list("?:!.,;")
stop_words = list(stopwords.words('english'))
def create_features_from_df(df):
df['Content_Parsed_1'] = df['Content'].str.replace("\r", " ")
df['Content_Parsed_1'] = df['Content_Parsed_1'].str.replace("\n", " ")
df['Content_Parsed_1'] = df['Content_Parsed_1'].str.replace(" ", " ")
df['Content_Parsed_1'] = df['Content_Parsed_1'].str.replace('"', '')
df['Content_Parsed_2'] = df['Content_Parsed_1'].str.lower()
df['Content_Parsed_3'] = df['Content_Parsed_2']
for punct_sign in punctuation_signs:
df['Content_Parsed_3'] = df['Content_Parsed_3'].str.replace(punct_sign, '')
df['Content_Parsed_4'] = df['Content_Parsed_3'].str.replace("'s", "")
wordnet_lemmatizer = WordNetLemmatizer()
nrows = len(df)
lemmatized_text_list = []
for row in range(0, nrows):
# Create an empty list containing lemmatized words
lemmatized_list = []
# Save the text and its words into an object
text = df.loc[row]['Content_Parsed_4']
text_words = text.split(" ")
# Iterate through every word to lemmatize
for word in text_words:
lemmatized_list.append(wordnet_lemmatizer.lemmatize(word, pos="v"))
# Join the list
lemmatized_text = " ".join(lemmatized_list)
# Append to the list containing the texts
lemmatized_text_list.append(lemmatized_text)
df['Content_Parsed_5'] = lemmatized_text_list
df['Content_Parsed_6'] = df['Content_Parsed_5']
for stop_word in stop_words:
regex_stopword = r"\b" + stop_word + r"\b"
df['Content_Parsed_6'] = df['Content_Parsed_6'].str.replace(regex_stopword, '')
df = df['Content_Parsed_6']
df = df.rename(columns={'Content_Parsed_6': 'Content_Parsed'})
# TF-IDF
features = tfidf.transform(df).toarray()
return features
category_codes = {
'business': 0,
'entertainment': 1,
'politics': 2,
'sport': 3,
'tech': 4,
'other':5
}
def get_category_name(category_id):
for category, id_ in category_codes.items():
if id_ == category_id:
return category
def predict_from_features(features):
# Obtain the highest probability of the predictions for each article
predictions_proba = svc_model.predict_proba(features).max(axis=1)
# Predict using the input model
predictions_pre = svc_model.predict(features)
# Replace prediction with 6 if associated cond. probability less than threshold
predictions = []
for prob, cat in zip(predictions_proba, predictions_pre):
if prob > .65:
predictions.append(cat)
else:
predictions.append(5)
# Return result
categories = [get_category_name(x) for x in predictions]
return categories
def complete_df(df, categories):
df['Prediction'] = categories
return df
def get_news():
news_contents = "In the men's freestyle, Rahul Aware (61kg) had clinched his first career Ranking Series title with a tactical 4-1 victory over Munir Aktas of Turkey. Utkarsh Kale had won bronze in the same category."
df_features = pd.DataFrame(
{'Content': [news_contents]
})
return df_features
df_features = get_news()
features = create_features_from_df(df_features)
predictions = predict_from_features(features)
predictions
```
| github_jupyter |
<a href="https://colab.research.google.com/github/EXYNOS-999/ACM-ICPC-Preparation/blob/master/Corona_EDA_viz.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Mathematical Simulation of nCOV Transmission Model
---
---
We are also investigating the use of a deterministic SIR metapopulation transmission model of infection within and between major Chinese cities to confirmed cases of 2019-nCoV in Chinese cities and cases.
reported in other countries; to study th epidemiological and population dynamics occurring
over the same time scale. (edited)
We define and study an open stochastic SIR (Susceptible – Infected –
Removed) model on a graph in order to describe the spread of an epidemic on a
cattle trade network with epidemiological and demographic dynamics occurring over
the same time scale.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
data=pd.read_csv("/content/2019_nCoV_data.csv")
data.info()
data['Last Update'] = pd.to_datetime(data['Last Update'])
data.info()
type(date)
usa=data[data['Country']=='US']
mchina=data[data['Country']=='Mainland China']
mchina=data[data['Country']=='China']
mchina.sort_values(['Last Update'])
hk=data[data['Country']=='Hong Kong']
hk
grouped_country=data.groupby("Country")[['Last Update','Confirmed', 'Deaths', 'Recovered']]
grouped_country['Country'].reset_index()
china=data[(data['Country']=='China') |( data['Country']=='Mainland China')|( data['Country']=='Hong Kong')]
china['Country'].replace('Mainland China','Chinese Sub',inplace=True)
china['Country'].replace('Hong Kong','Chinese Sub',inplace=True)
Chinese=china.groupby('Last Update')['Confirmed','Deaths','Recovered'].sum().reset_index()
Chinese.to_csv('sample_data/coronokimaakiankh.csv',date_format='%Y-%m-%d %H:%M:%S')
x=Chinese[['Recovered','Deaths']]
y=Chinese[['Confirmed']]
sns.regplot(np.arange(0,10,1),Chinese['Confirmed'],data=Chinese)
import statsmodels.api as sm
from sklearn.svm import SVR
model=SVR(kernel='linear',C=0.5)
model.fit(x.values,y.values)
from sklearn.preprocessing import PolynomialFeatures
from sklearn.svm import SVC
svc=SVC()
pf=PolynomialFeatures(degree=3,include_bias=True)
x_poly=pf.fit_transform(x.values)
svc.fit(x_poly, y.values)
y_poly_pred = svc.predict(x_poly)
res
```
Slight errors in prediction yet to be improved
```
plt.suptitle("Predicted in red ,Actual in blue ")
lines=plt.plot(np.arange(0,10,1),res,c="red")
sns.regplot(np.arange(0,10,1),Chinese['Confirmed'],data=Chinese)
y.values
res = mod.fit(maxiter=1000, disp=False)
print(res.summary())
data
```
No correlation between variables ,thus more feature engineering is required
```
sns.heatmap(data.corr())
```
# Features
- Who Reports
- Seafood wholesale market
- Population Density
- Stock Market Price
- Geographical Conditions
- Visits to Outbreak zone
- Wild Animals in China
- Base Reproduction Rate
- Incubation Period
- Medical Personal Deployed
- Realtime Social Media Sentiment
- Analyis of flights
- Analysis of wuhan metro data
- Highway Traffic using Google Sattelite API
- Attendance in Schools
- Medical/Army Personal Deployement
# Note: Subject to change
# *Investigating the possibility of Transfer Learning for prediction.
```
!pip install prophet
from fbprophet import Prophet
Chinese.columns=['ds','y','Deaths','Recovered']
chinese=Chinese.drop(columns=['Recovered','Deaths'])
m = Prophet()
m.fit(chinese)
future = m.make_future_dataframe(periods=7,include_history=True)
future.tail()
forecast = m.predict(future)
forecast[['ds', 'yhat', 'yhat_lower', 'yhat_upper']].tail()
m.plot_components(forecast)
res=m.predict(future)
res
res.iloc[10]
m.plot(res)
```
# CLEANED NEW PART
```
data=pd.read_csv("/content/china.csv")
data.columns=['DROPPER','Provinces','Country','Last Update' ,'Confirmed' ,'Deaths' ,'Recovered']
data=data.drop(columns=['DROPPER','Provinces','Country','Deaths','Recovered'])
data.columns=['ds','y']
from fbprophet import Prophet
m = Prophet()
m.fit(naan)
future = m.make_future_dataframe(periods=2,freq='d')
future.tail()
forecast = m.predict(future)
forecast[['ds', 'yhat', 'yhat_lower', 'yhat_upper']].tail()
m.plot_components(forecast)
res=m.predict(future)
res.groupby('ds')['yhat'].sum().reset_index()
m.plot(res)
```
| github_jupyter |
<a id="topD"></a>
# Downloading COS Data
# Learning Goals
<font size="5"> This Notebook is designed to walk the user (<em>you</em>) through: <b>Downloading existing Cosmic Origins Spectrograph (<em>COS</em>) data from the online archive</b></font>
**1. [Using the web browser interface](#mastD)**
\- 1.1. [The Classic HST Web Search](#mastD)
\- 1.2. [Searching for a Series of Observations on the Classic Web Search](#WebSearchSeriesD)
\- 1.3. [The MAST Portal](#mastportD)
\- 1.4. [Searching for a Series of Observations on the MAST Portal](#mastportSeriesD)
**2. [Using the `Python` module `Astroquery`](#astroqueryD)**
\- 2.1. [Searching for a single source with Astroquery](#Astroquery1D)
\- 2.2. [Narrowing Search with Observational Parameters](#NarrowSearchD)
\- 2.3. [Choosing and Downloading Data Products](#dataprodsD)
\- 2.4. [Using astroquery to find data on a series of sources](#Astroquery2D)
## Choosing how to access the data
**This Notebook explains three methods of accessing COS data hosted by the STScI Mikulski Archive for Space Telescopes (MAST).**
You may read through all three, or you may wish to focus on a particular method which best suits your needs.
**Please use the table below to determine which section on which to focus.**
||The [Classic HST Search (Web Interface)](https://archive.stsci.edu/hst/search.php)|The [MAST Portal (Web Interface)](https://mast.stsci.edu/portal/Mashup/Clients/Mast/Portal.html)|The [`Astroquery` (`Python` Interface)](http://astroquery.readthedocs.io/)|
|-|-|-|-|
||- User-friendly point-and-click searching|- Very user-friendly point-and-click searching|- Requires a bit of `Python` experience|
||- Advanced **mission-specific** search parameters, including: central wavelength, detector, etc.|- Lacks some mission-specific search parameters|- Allows for programmatic searching and downloads|
||- Can be difficult to download the data if not on the STScI network|- Easy to download selected data|- Best for large datasets|
|||||
|***Use this method if...*** |*...You're unfamiliar with `Python` and need to search for data by cenwave*|*...You're exploring the data and you don't need to search by cenwave*|*...You know `Python` and have an idea of what data you're looking for, or you have a lot of data*|
|***Described in...***|*[Section 1.1](#mastD)*|*[Section 1.3](#mastportD)*|*[Section 2.1](#astroqueryD)*|
*Note* that these are only recommendations, and you may prefer another option. For most purposes, the writer of this tutorial recommends the `Astroquery` `Python` interface, unless you are not at all comfortable using python or doing purely exploratory work.
<!-- *You may review Section 1 or 2 independently or together.* -->
<!-- *The web search (Section 1) is generally better for introductory users and exploratory use, while the `astroquery` method (Section 2) is easier for those with some `python` experience.* -->
# 0. Introduction
**The Cosmic Origins Spectrograph ([*COS*](https://www.nasa.gov/content/hubble-space-telescope-cosmic-origins-spectrograph)) is an ultraviolet spectrograph on-board the Hubble Space Telescope([*HST*](https://www.stsci.edu/hst/about)) with capabilities in the near ultraviolet (*NUV*) and far ultraviolet (*FUV*).**
**This tutorial aims to prepare you to access the existing COS data of your choice by walking you through downloading a processed spectrum, as well as various calibration files obtained with COS.**
- For an in-depth manual to working with COS data and a discussion of caveats and user tips, see the [COS Data Handbook](https://hst-docs.stsci.edu/display/COSDHB/).
- For a detailed overview of the COS instrument, see the [COS Instrument Handbook](https://hst-docs.stsci.edu/display/COSIHB/).
<font size="5"> We will define a few directories in which to place our data.</font>
And to create new directories, we'll import `pathlib.Path`:
```
#Import for: working with system paths
from pathlib import Path
# This will be an important directory for the Notebook, where we save data
data_dir = Path('./data/')
data_dir.mkdir(exist_ok=True)
```
<a id="downloadD"></a>
# 1. Downloading the data through the browser interface
One can search for COS data from both a browser-based Graphical User Interface (*gui*) and a `Python` interface. This Section (1) will examine two web interfaces. [Section 2](#astroqueryD) will explain the `Python` interface.
*Note, there are other, more specialized ways to query the mast API not discussed in this Notebook. An in-depth MAST API tutorial can be found [here](https://mast.stsci.edu/api/v0/MastApiTutorial.html).*
<a id="mastD"></a>
## 1.1 The Classic HST Web Search
**A browser gui for searching *specifically* through [HST archival data can be found here](http://archive.stsci.edu/hst/search.php). We will be discussing *this* HST search in the section below.**
As of September, 2021, two other portals also allow access to the same data:
* A newer HST-specific search page ([here](https://mast.stsci.edu/search/hst/ui/#/)). Most downloading difficulties have been solved with this new site, and upcoming versions of this tutorial will focus on its use.
* A more general MAST gui, which also allows access to data from other telescopes such as TESS, but does not offer all HST-specific search parameters. We will discuss this interface in [Section 1.3](#mastportD).
The search page of the HST interface is laid out as in fig. 1.1:
### Fig 1.1
<center><img src=./figures/Mast_hst_searchformQSO.png width ="900" title="MAST Archive search form for a COS data query"> </center>
where here we have indicated we would like to find all archival science data from the **COS far-ultraviolet (FUV) configuration**, taken with any grating while looking at Quasi-Stellar Objects (QSO) within a 3 arcminute radius of (1hr:37':40", +33d 09m 32s). The output columns we have selected to see are visible in the bottom left of Fig 1.1.
Note that if you have a list of coordinates, Observation ID(s), etc. for a series of targets you can click on the "File Upload Form" and attach your list of OBSIDs or identifying features. Then specify which type of data your list contains using the "File Contents" drop-down menu.
Figure 1.2 shows the results of our search shown in Fig 1.1.
### Fig 1.2
<center><img src=figures/QSO_MastSearchRes.png width ="900" title="MAST Archive search results for a COS data query"> </center>
**We now choose our dataset.**
We rather arbitrarily select **`LCXV13050`** because of its long exposure time, taken under an observing program described as:
> "Project AMIGA: Mapping the Circumgalactic Medium of Andromeda"
This is a Quasar known as [3C48](http://simbad.u-strasbg.fr/simbad/sim-basic?Ident=3c48&submit=SIMBAD+search), one of the first quasars discovered.
Clicking on the dataset, we are taken to a page displaying a preview spectrum (Fig 1.3).
### Fig 1.3
<center><img src=./figures/QSOPreviewSpec.png width ="900" title="MAST Archive preview spectrum of LCXV13050"> </center>
We now return to the [search page](http://archive.stsci.edu/hst/search.php) and enter in LCXV13050 under "Dataset" with no other parameters set. Clicking "search", now we see a single-rowed table with *just* our dataset, and the option to download datasets. We mark the row we wish to download and click "Submit marked data for retrieval from STDADS". See Fig 1.4.
### Fig 1.4
<center><img src =figures/LCXV13050_res.png width ="900" title="MAST Archive dataset overview of LCXV13050"> </center>
Now we see a page like in Fig 1.5,
where we can either sign in with STScI credentials, or simply provide our email to proceed without credentials. Generally, you may proceed anonymously, unless you are retrieving proprietary data to which you have access. Next, make sure to select "Deliver the data to the Archive staging area". Click "Send Retrieval Request to ST-DADS" and you will receive an email with instructions on downloading the data.
### Fig 1.5
<center><img src =figures/DownloadOptions.png width ="900" title="Download Options for LCXV13050"> </center>
Now the data is "staged" on a MAST server, and you need to download it to your local computer.
### Downloading the staged data
We demonstrate three methods of downloading your staged data:
1. If your terminal supports it, you may [use the `wget` tool](#wgetDLD).
2. However if that does not work, we recommend [using a secure ftp client application](#download_ftps_cyduckD).
3. Finally, if you would instead like to download *staged data* programmatically, you may [use the Python `ftplib` package](#download_ftps_funcD), as described [here](https://archive.stsci.edu/ftp.html) in STScI's documentation of the MAST FTP Service. For your convenience, we have built the `download_anonymous_staged_data` function below, which will download anonymously staged data via ftps.
<a id=wgetDLD></a>
#### Downloading the staged data with `wget`
**If you are connected to the STScI network, either in-person or via a virtual private network (VPN), you should use the `wget` command as in the example below:**
`wget -r --ftp-user=anonymous --ask-password ftps://archive.stsci.edu/stage/anonymous/anonymous<directory_number> --directory-prefix=<data_dir>`
where `directory_number` is the number at the end of the anonymous path specified in the email you received from MAST and `data_dir` is the local directory where you want the downloaded data.
You will be prompted for a password. Type in the email address you used, then press enter/return.
Now all the data will be downloaded into a subdirectory of data_dir: `"./archive.stsci.edu/stage/anonymous/anonymous<directory_number>/"`
<a id=download_ftps_cyduckD></a>
#### Downloading the staged data with a secure ftp client application (`CyberDuck`)
CyberDuck is an application which allows you to securely access data stored on another machine using ftps. To download your staged data using Cyberduck, first download the [Cyberduck](https://cyberduck.io) application (*free, with a recommended donation*). Next, open a new browser window (Safari, Firefox, and Google Chrome have all been shown to work,) and type in the following web address: `ftps://archive.stsci.edu/stage/anonymous<directory_number>`, where `directory_number` is the number at the end of the anonymous path specified in the email you received from MAST. For example, if the email specifies:
> "The data can be found in the directory... /stage/anonymous/anonymous42822"
then this number is **42822**
Your browser will attempt to redirect to the CyberDuck application. Allow it to "Open CyberDuck.app", and CyberDuck should open a finder window displaying your files. Select whichever files you want to download by highlighting them (command-click or control-click) then right click one of the highlighted files, and select "Download To". This will bring up a file browser allowing you to save the selected files to wherever you wish on your local computer.
<a id=download_ftps_funcD></a>
#### Downloading the staged data with `ftps`
To download anonymously staged data programmatically with ftps, you may run the `download_anonymous_staged_data` function as shown here:
```python
download_anonymous_staged_data(email_used="my_email@stsci.edu", directory_number=80552, outdir="./here_is_where_I_want_the_data")
```
Which results in:
```
Downloading lcxv13050_x1dsum1.fits
Done
...
...
Downloading lcxv13gxq_flt_b.fits
Done
```
```
import ftplib
def download_anonymous_staged_data(email_used, directory_number, outdir = "./data/ftps_download/", verbose=True):
"""
A direct implementation of the MAST FTP Service webpage's ftplib example code.
Downloads anonymously staged data from the MAST servers via ftps.
Inputs:
email_used (str): the email address used to stage the data
directory_number (str or int): The number at the end of the anonymous filepath. i.e. if the email you received includes:
"The data can be found in the directory... /stage/anonymous/anonymous42822",
then this number is 42822
outdir (str): Path to where the file will download locally.
verbose (bool): If True, prints name of each file downloaded.
"""
ftps = ftplib.FTP_TLS('archive.stsci.edu') # Set up connection
ftps.login(user="anonymous", passwd=email_used) # Login with anonymous credentials
ftps.prot_p() # Add protection to the connection
ftps.cwd(f"stage/anonymous/anonymous{directory_number}")
filenames = ftps.nlst()
outdir = Path(outdir) # Set up the output directory as a path
outdir.mkdir(exist_ok=True)
for filename in filenames: # Loop through all the staged files
if verbose:
print("Downloading " + filename)
with open(outdir / filename, 'wb') as fp: # Download each file locally
ftps.retrbinary('RETR {}'.format(filename), fp.write)
if verbose:
print(" Done")
```
<font size="5"> <b>Well Done making it this far!</b></font>
Attempt the exercise below for some extra practice.
### Exercise 1: *Searching the archive for TRAPPIST-1 data*
[TRAPPIST-1](https://en.wikipedia.org/wiki/TRAPPIST-1) is a cool red dwarf with a multiple-exoplanet system.
- Find its coordinates using the [SIMBAD Basic Search](http://simbad.u-strasbg.fr/simbad/sim-fbasic).
- Use those coordinates in the [HST web search](https://archive.stsci.edu/hst/search.php) or the [MAST portal](https://mast.stsci.edu/portal/Mashup/Clients/Mast/Portal.html) to find all COS exposures of the system.
- Limit the search terms to find the COS dataset taken in the COS far-UV configuration with the grating G130M.
**What is the dataset ID, and how long was the exposure?**
Place your answer in the cell below.
```
# Your answer here
```
<a id=WebSearchSeriesD></a>
## 1.2. Searching for a Series of Observations on the Classic HST Web Search
Now let's try using the web interface's [file upload form](http://archive.stsci.edu/hst/search.php?form=fuf) to search for a series of observations by their dataset IDs. We're going to look for three observations of the same object, the white dwarf WD1057+719, taken with three different COS gratings. Two are in the FUV and one in the NUV. The dataset IDs are
- LDYR52010
- LBNM01040
- LBBD04040
So that we have an example list of datasets to input to the web search, we make a comma-separated-value txt file with these three obs_ids, and save it as `obsId_list.txt`.
```
obsIdList = ['LDYR52010','LBNM01040','LBBD04040'] # The three observation IDs we want to gather
obsIdList_length = len(obsIdList)
with open('./obsId_list.txt', 'w') as f: # Open up this new file in "write" mode
for i, item in enumerate(obsIdList): # We want a newline after each obs_id except the last one
if i < obsIdList_length - 1:
f.writelines(item + "," + '\n')
if i == obsIdList_length - 1: # Make sure we don't end the file with a blank line (below)
f.writelines(item)
```
Then we link to this file under the **Local File Name** browse menu on the file upload form. We must set the **File Contents** term to Data ID, as that is the identifier we have provided in our file, and we change the **delimiter** to a comma.
Because we are searching by Dataset ID, we don't need to specify any additional parameters to narrow down the data.
### Fig 1.6
<center><img src =figures/FUF_search.png width ="900" title="File Upload Search Form"> </center>
**We now can access all the datasets, as shown in Fig. 1.7:**
### Fig 1.7
<center><img src =figures/FUF_res.png width ="900" title="File Upload Search Results"> </center>
Now, to download all of the relevant files, we can check the **mark** box for all of them, and again hit "Submit marked data for retrieval from STDADS". This time, we want to retrieve **all the calibration files** associated with each dataset, so we check the following boxes:
- Uncalibrated
- Calibrated
- Used Reference Files
(*See Fig. 1.8*)
### Fig 1.8
<center><img src =./figures/DownloadOptions_FUF.png width ="900" title="Download Options for multiple datasets"> </center>
The procedure from here is the same described above in Section 1.1. Now, when we download the staged data, we obtain multiple subdirectories with each dataset separated.
<a id = mastportD></a>
## 1.3. The MAST Portal
STScI hosts another web-based gui for accessing data, the [MAST Portal](https://mast.stsci.edu/portal/Mashup/Clients/Mast/Portal.html). This is a newer interface which hosts data from across many missions and allows the user to visualize the target in survey images, take quick looks at spectra or lightcurves, and manage multiple search tabs at once. Additionally, it handles downloads in a slightly more beginner-friendly manner than the current implementation of the Classic HST Search. This guide will only cover the basics of accessing COS data through the MAST Portal; you can find more in-depth documentation in the form of helpful video guides on the [MAST YouTube Channel](https://www.youtube.com/user/STScIMAST).
**Let's find the same data we found in Section 1.1, on the QSO 3C48:**
Navigate to the MAST Portal at <https://mast.stsci.edu/portal/Mashup/Clients/Mast/Portal.html>, and you will be greeted by a screen where the top looks like Fig. 1.9.
### Fig 1.9
<center><img src =figures/mastp_top.png width ="900" title="Top of MAST Portal Home"> </center>
Click on "Advanced Search" (boxed in red in Fig. 1.9). This will open up a new search tab, as shown in Fig. 1.10:
### Fig 1.10
<center><img src =figures/mastp_adv.png width ="900" title="The advanced search tab"> </center>
Fig 1.10 (above) shows the default search fields which appear. Depending on what you are looking for, these may or may not be the most helpful search fields. By unchecking some of the fields which we are not interested in searching by right now (boxed in green), and then entering the parameter values by which to narrow the search into each parameter's box, we generate Fig. 1.11 (below). One of the six fields (Mission) by which we are narrowing is boxed in a dashed blue line. The list of applied filters is boxed in red. A dashed pink box at the top left indicates that 2 records were found matching all of these parameters. To its left is an orange box around the "Search" button to press to bring up the list of results
Here we are searching by:
|**Search Parameter**|**Value**|
|-|-|
|Mission|HST|
|Instrument|COS/FUV|
|Filters|G160M|
|Target Name|3C48|
|Observation ID|LCXV\* (*the star is a "wild card" value, so the search will find any file whose `obs_id` begins with LCXV*)|
|Product Type|spectrum|
### Fig 1.11
<center><img src =figures/mastp_adv_2.png width ="900" title="The advanced search tab with some selections"> </center>
Click the "Search" button (boxed in orange), and you will be brought to a page resembling Fig. 1.12.
### Fig 1.12
<center><img src =figures/mastp_res1.png width ="900" title="Results of MAST Portal search"> </center>
<font size="4"> <b>Above, in Fig 1.12</b>:</font>
- The yellow box to the right shows the AstroView panel, where you can interactively explore the area around your target:
- click and drag to pan around
- scroll to zoom in/out
- The dashed-blue box highlights additional filters you can use to narrow your search results.
- The red box highlights a button you can click with *some* spectral datasets to pull up an interactive spectrum.
- The green box highlights the "Mark" checkboxes for each dataset.
- The black circle highlights the single dataset download button:
- **If you only need to download one or two datasets, you may simply click this button for each dataset**
- Clicking the single dataset download button will attempt to open a "pop-up" window, which you must allow in order to download the file. Some browsers will require you to manually allow pop-ups.
<a id="mastportSeriesD"></a>
## 1.4. Searching for a Series of Observations on the MAST Portal
<font size="4"> <b>To download multiple datasets</b>:</font>
The MAST portal acts a bit like an online shopping website, where you add your *data products* to the checkout *cart*/*basket*, then open up your cart to *checkout* and download the files.
Using the checkboxes, mark all the datasets you wish to download (in this case, we'll download both LCXV13040 and LCXV13050). Then, click the "Add data products to Download Basket" button (circled in a dashed-purple line), which will take you to a "Download Basket" screen resembling Fig 1.13:
### Fig 1.13
<center><img src =figures/mastp_cart2.png width ="900" title="MAST Portal Download Basket"> </center>
Each dataset contains *many* files, most of which are calibration files or intermediate processing files. You may or may not want some of these intermediate files in addition to the final product file.
In the leftmost "Filters" section of the Download Basket page, you can narrow which files will be downloaded (boxed in red).
By default, only the **minimum recommended products** (*mrp*) will be selected. In the case of most COS data, this will be the final spectrum `x1dsum` file and association `asn` file for each dataset. The mrp files for the first dataset (`LCXV13040`) are highlighted in yellow. These two mrp filetypes are fine for our purposes here; however if you want to download files associated with specific exposures, or any calibration files or intermediate files, you can select those you wish to download with the checkboxes in the file tree system (boxed in dashed-green).
**For this tutorial, we simply select "Minimum Recommended Products" at the top left. With this box checked, all of the folders representing individual exposures are no longer visible.**
Check the box labelled "HST" to select all files included by the filters, and click the "Download Selected Items" button at the top right (dashed-black circle). This will bring up a small window asking you what format to download your files as. For datasets smaller than several Gigabytes, the `Zip` format will do fine. Click Download, and a pop-up window will try to open to download the files. If no download begins, make sure to enable this particular pop-up, or allow pop-ups on the MAST page.
**Your files should now be downloaded as a compressed `Zip` folder.**
If you need help uncompressing the `Zip`ped files, check out these links for: [Windows](https://support.microsoft.com/en-us/windows/zip-and-unzip-files-8d28fa72-f2f9-712f-67df-f80cf89fd4e5) and [Mac](https://support.apple.com/guide/mac-help/zip-and-unzip-files-and-folders-on-mac-mchlp2528/mac). There are numerous ways to do this on Linux, however we have not vetted them.
<a id = astroqueryD></a>
# 2. The Python Package `astroquery.mast`
Another way to search for and download archived datasets is from within `Python` using the module [`astroquery.mast`](https://astroquery.readthedocs.io/en/latest/mast/mast.html). We will import one of this module's key submodules: `Observations`.
*Please note* that the canonical source of information on this package is the [`astroquery` docs](https://astroquery.readthedocs.io/en/latest/) - please look there for the most up-to-date instructions.
## We will import the following packages:
- `astroquery.mast`'s submodule `Observations` for finding and downloading data from the [MAST](https://mast.stsci.edu/portal/Mashup/Clients/Mast/Portal.html) archive
- `csv`'s submodule `reader` for reading in/out from a csv file of source names.
```
# Downloading data from archive
from astroquery.mast import Observations
# Reading in multiple source names from a csv file
from csv import reader
```
<a id=Astroquery1D></a>
## 2.1. Searching for a single source with Astroquery
There are *many* options for searching the archive with astroquery, but we will begin with a very general search using the coordinates we found for WD1057+719 in the last section to find the dataset with the longest exposure time using the COS/FUV mode through the G160M filter. We could also search by object name to have it resolved to a set of coordinates, with the function `Observations.query_object(objectname = '3C48')`.
- Our coordinates were: (11:00:34.126 +71:38:02.80).
- We can search these coordinates as sexagesimal coordinates, or convert them to decimal degrees.
```
query_1 = Observations.query_object("11:00:34.126 +71:38:02.80", radius="5 sec")
```
This command has generated a table of objects called **"query_1"**. We can see what information we have on the objects in the table by printing its *`keys`*, and see how many objects are in the table with `len(query_1)`.
```
print(f"We have table information on {len(query_1)} observations in the following categories/columns:\n")
q1_keys = (query_1.keys())
q1_keys
```
<a id=NarrowSearchD></a>
## 2.2. Narrowing Search with Observational Parameters
Now we narrow down a bit with some additional parameters and sort by exposure time.
The parameter limits we add to the search are:
- *Only look for sources in the coordinate range between right ascension 165 to 166 degrees and declination +71 to +72 degrees*
- *Only find observations in the UV*
- *Only find observations taken with the COS instrument (either in its FUV or NUV configuration).*
- *Only find spectrographic observations*
- *Only find observations made using the COS grating "G160M"*
```
query_2 = Observations.query_criteria(s_ra=[165., 166.], s_dec=[+71.,+72.],
wavelength_region="UV", instrument_name=["COS/NUV","COS/FUV"],
dataproduct_type = "spectrum", filters = 'G160M')
# Next lines simplifies the columns of data we see to some useful data we will look at right now
limq2 = query_2['obsid','obs_id', 'target_name', 'dataproduct_type', 'instrument_name',
'project', 'filters', 'wavelength_region', 't_exptime']
sort_order = query_2.argsort('t_exptime') # This is the index list in order of exposure time, increasing
print(limq2[sort_order])
chosenObs = limq2[sort_order][-1] # Grab the last value of the sorted list
print(f"\n\nThe longest COS/FUV exposure with the G160M filter is: \n\n{chosenObs}")
```
<font size="5">Caution! </font>
<img src=./figures/warning.png width ="60" title="CAUTION">
Please note that these queries are `Astropy` tables and do not always respond as expected for other data structures like `Pandas DataFrames`. For instance, the first way of filtering a table shown below is correct, but the second will consistently produce the *wrong result*. You *must* search and filter these tables by masking them, as in the first example below.
```
# Searching a table generated with a query
## First, correct way using masking
mask = (query_1['obs_id'] == 'lbbd01020') # NOTE, obs_id must be lower-case
print("Correct way yields: \n" , query_1[mask]['obs_id'],"\n\n")
# Second INCORRECT way
print("Incorrect way yields: \n" , query_1['obs_id' == 'LBBD01020']['obs_id'], "\nwhich is NOT what we're looking for!")
```
<a id=dataprodsD></a>
## 2.3. Choosing and Downloading Data Products
**Now we can choose and download our data products from the archive dataset.**
We will first generate a list of data products in the dataset: `product_list`. This will generate a large list, but we will only show the first 10 values.
```
product_list = Observations.get_product_list(chosenObs)
product_list[:10] #Not the whole dataset, just first 10 lines/observations
```
Now, we will download *just the* **minimum recommended products** (*mrp*) which are the fully calibrated spectrum (denoted by the suffix `_x1d` or here `x1dsum`) and the association file (denoted by the suffix `_asn`). We do this by setting the parameter `mrp_only` to True. The association file contains no data, but rather the metadata explaining which exposures produced the `x1dsum` dataset. The `x1dsum` file is the final product summed across all of the [fixed pattern noise positions](https://hst-docs.stsci.edu/cosdhb/chapter-1-cos-overview/1-1-instrument-capabilities-and-design#id-1.1InstrumentCapabilitiesandDesign-GratingOffset(FP-POS)GratingOffsetPositions(FP-POS)) (`FP-POS`). The `x1d` and `x1dsum<n>` files are intermediate spectra. Much more information can be found in the [COS Instrument Handbook](https://hst-docs.stsci.edu/display/COSIHB/).
We would set `mrp_only` to False, if we wanted to download ***all*** the data from the observation, including:
- support files such as the spacecraft's pointing data over time (`jit` files).
- intermediate data products such as calibrated TIME-TAG data (`corrtag` or `corrtag_a`/`corrtag_b` files) and extracted 1-dimensional spectra averaged over exposures with a specific `FP-POS` value (`x1dsum<n>` files).
<img src=./figures/warning.png width ="60" title="CAUTION">
However, use caution with downloading all files, as in this case, setting `mrp_only` to False results in the transfer of **7 Gigabytes** of data, which can take a long time to transfer and eat away at your computer's storage! In general, only download the files you need. On the other hand, often researchers will download only the raw data, so that they can process it for themselves. Since here we only need the final `x1dsum` and `asn` files, we only need to download 2 Megabytes.
```
downloads = Observations.download_products(product_list, download_dir=str(data_dir) , extension='fits', mrp_only=True, cache=False)
```
### Exercise 2: *Download the raw counts data on TRAPPIST-1*
In the previous exercise, we found an observation COS took on TRAPPIST-1 system. In case you skipped Exercise 1, the observation's Dataset ID is `LDLM40010`.
Use `Astroquery.mast` to download the raw `TIME-TAG` data, rather than the x1d spectra files. See the [COS Data Handbook Ch. 2](https://hst-docs.stsci.edu/cosdhb/chapter-2-cos-data-files/2-4-cos-data-products) for details on TIME-TAG data files. Make sure to get the data from both segments of the FUV detector (i.e. both `RAWTAG_A` and `RAWTAG_B` files). If you do this correctly, there should be five data files for each detector segment.
*Note that some of the obs_id may appear in the table as slightly different, i.e.: ldlm40alq and ldlm40axq, rather than ldlm40010. The main obs_id they fall under is still ldlm40010, and this will still work as a search term. They are linked together by the association file described here in section 2.3.*
```
# Your answer here
```
<a id=Astroquery2D></a>
## 2.4. Using astroquery to find data on a series of sources
In this case, we'll look for COS data around several bright globular clusters:
- Omega Centauri
- M5
- M13
- M15
- M53
We will first write a comma-separated-value (csv) file `objectname_list.csv` listing these sources by their common name. This is a bit redundant here, as we will immediately read back in what we have written; however it is done here to deliberately teach both sides of the writing/reading process, and as many users will find themselves with a csv sourcelist they must search.
```
sourcelist = ['omega Centauri', 'M5', 'M13', 'M15', 'M53'] # The 5 sources we want to look for
sourcelist_length = len(sourcelist) # measures the length of the list for if statements below
with open('./objectname_list.csv', 'w') as f: # Open this new file in "write" mode
for i, item in enumerate(sourcelist): # We want a comma after each source name except the last one
if i < sourcelist_length - 1:
f.writelines(item + ",")
if i == sourcelist_length - 1: # No comma after the last entry
f.writelines(item)
with open('./objectname_list.csv', 'r', newline = '') as csvFile: # Open the file we just wrote in "read" mode
objList = list(reader(csvFile, delimiter = ','))[0] # This is the exact same list as `sourcelist`!
print("The input csv file contained the following sources:\n", objList)
globular_cluster_queries = {} # Make a dictionary, where each source name (i.e. "M15") corresponds to a list of its observations with COS
for obj in objList: # each "obj" is a source name
query_x = Observations.query_criteria(objectname = obj, radius = "5 min", instrument_name=['COS/FUV', 'COS/NUV']) # query the area in +/- 5 arcminutes
globular_cluster_queries[obj] = (query_x) # add this entry to the dictionary
globular_cluster_queries # show the dictionary
```
**Excellent! You've now done the hardest part - finding and downloading the right data.** From here, it's generally straightforward to read in and plot the spectrum. We recommend you look into our tutorial on [Viewing a COS Spectrum](https://github.com/spacetelescope/notebooks/blob/master/notebooks/COS/ViewData/ViewData.ipynb).
## Congratulations! You finished this Notebook!
### There are more COS data walkthrough Notebooks on different topics. You can find them [here](https://spacetelescope.github.io/COS-Notebooks/).
---
## About this Notebook
**Author:** Nat Kerman <nkerman@stsci.edu>
**Updated On:** 2021-10-29
> *This tutorial was generated to be in compliance with the [STScI style guides](https://github.com/spacetelescope/style-guides) and would like to cite the [Jupyter guide](https://github.com/spacetelescope/style-guides/blob/master/templates/example_notebook.ipynb) in particular.*
## Citations
If you use `astropy`, `matplotlib`, `astroquery`, or `numpy` for published research, please cite the
authors. Follow these links for more information about citations:
* [Citing `astropy`/`numpy`/`matplotlib`](https://www.scipy.org/citing.html)
* [Citing `astroquery`](https://astroquery.readthedocs.io/en/latest/)
---
[Top of Page](#topD)
<img style="float: right;" src="https://raw.githubusercontent.com/spacetelescope/notebooks/master/assets/stsci_pri_combo_mark_horizonal_white_bkgd.png" alt="Space Telescope Logo" width="200px"/>
<br></br>
<br></br>
<br></br>
## Exercise Solutions:
Note, that for many of these, there are multiple ways to get an answer.
**We will import:**
- numpy to handle array functions
- astropy.table Table for creating tidy tables of the data
```
# Manipulating arrays
import numpy as np
# Reading in data
from astropy.table import Table
## Ex. 1 solution:
dataset_id_ = 'LDLM40010'
exptime_ = 12403.904
print(f"The TRAPPIST-1 COS data is in dataset {dataset_id_}, taken with an exosure time of {exptime_}")
## Ex. 2 solution:
query_3 = Observations.query_criteria(obs_id = 'LDLM40010',
wavelength_region="UV", instrument_name="COS/FUV", filters = 'G130M')
product_list2 = Observations.get_product_list(query_3)
rawRowsA = np.where(product_list2['productSubGroupDescription'] == "RAWTAG_A")
rawRowsB = np.where(product_list2['productSubGroupDescription'] == "RAWTAG_B")
rawRows = np.append(rawRowsA,rawRowsB)
!mkdir ./data/Ex2/
downloads2 = Observations.download_products(product_list2[rawRows], download_dir=str(data_dir/'Ex2/') , extension='fits', mrp_only=False, cache=True)
downloads3 = Observations.download_products(product_list2, download_dir=str(data_dir/'Ex2/') , extension='fits', mrp_only=True, cache=True)
asn_data = Table.read('./data/Ex2/mastDownload/HST/ldlm40010/ldlm40010_asn.fits', hdu = 1)
print(asn_data)
```
| github_jupyter |
# Generating counterfactuals for multi-class classification and regression models
This notebook will demonstrate how the DiCE library can be used for multiclass classification and regression for scikit-learn models.
You can use any method ("random", "kdtree", "genetic"), just specific it in the method argument in the initialization step. The rest of the code is completely identical.
For demonstration, we will be using the genetic algorithm for CFs.
```
%load_ext autoreload
%autoreload 2
import dice_ml
from dice_ml import Dice
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler, OneHotEncoder
import pandas as pd
```
We will use sklearn's internal datasets to demonstrate DiCE's features in this notebook
```
outcome_name = 'target'
# Function to process sklearn's internal datasets
def sklearn_to_df(sklearn_dataset):
df = pd.DataFrame(sklearn_dataset.data, columns=sklearn_dataset.feature_names)
df[outcome_name] = pd.Series(sklearn_dataset.target)
return df
```
## Multiclass Classification
For multiclass classification, we will use sklearn's Iris dataset. This data set consists of 3 different types of irises’ (Setosa, Versicolour, and Virginica) petal and sepal length. More information at https://scikit-learn.org/stable/datasets/toy_dataset.html#iris-plants-dataset
```
from sklearn.datasets import load_iris
df_iris = sklearn_to_df(load_iris())
df_iris.head()
df_iris.info()
continuous_features_iris = df_iris.drop(outcome_name, axis=1).columns.tolist()
target = df_iris[outcome_name]
# Split data into train and test
from sklearn.model_selection import train_test_split
from sklearn.compose import ColumnTransformer
from sklearn.ensemble import RandomForestClassifier
datasetX = df_iris.drop(outcome_name, axis=1)
x_train, x_test, y_train, y_test = train_test_split(datasetX,
target,
test_size = 0.2,
random_state=0,
stratify=target)
categorical_features = x_train.columns.difference(continuous_features_iris)
# We create the preprocessing pipelines for both numeric and categorical data.
numeric_transformer = Pipeline(steps=[
('scaler', StandardScaler())])
categorical_transformer = Pipeline(steps=[
('onehot', OneHotEncoder(handle_unknown='ignore'))])
transformations = ColumnTransformer(
transformers=[
('num', numeric_transformer, continuous_features_iris),
('cat', categorical_transformer, categorical_features)])
# Append classifier to preprocessing pipeline.
# Now we have a full prediction pipeline.
clf_iris = Pipeline(steps=[('preprocessor', transformations),
('classifier', RandomForestClassifier())])
model_iris = clf_iris.fit(x_train, y_train)
d_iris = dice_ml.Data(dataframe=df_iris,
continuous_features=continuous_features_iris,
outcome_name=outcome_name)
# We provide the type of model as a parameter (model_type)
m_iris = dice_ml.Model(model=model_iris, backend="sklearn", model_type='classifier')
exp_genetic_iris = Dice(d_iris, m_iris, method="genetic")
```
As we can see below, all the target values will lie in the desired class
```
# Single input
query_instances_iris = x_train[2:3]
genetic_iris = exp_genetic_iris.generate_counterfactuals(query_instances_iris, total_CFs=7, desired_class = 2)
genetic_iris.visualize_as_dataframe()
# Multiple queries can be given as input at once
query_instances_iris = x_train[17:19]
genetic_iris = exp_genetic_iris.generate_counterfactuals(query_instances_iris, total_CFs=7, desired_class = 2)
genetic_iris.visualize_as_dataframe(show_only_changes=True)
```
# Regression
For regression, we will use sklearn's boston dataset. This dataset contains boston house-prices. More information at https://scikit-learn.org/stable/datasets/toy_dataset.html#boston-house-prices-dataset
```
from sklearn.datasets import load_boston
df_boston = sklearn_to_df(load_boston())
df_boston.head()
df_boston.info()
continuous_features_boston = df_boston.drop(outcome_name, axis=1).columns.tolist()
target = df_boston[outcome_name]
from sklearn.model_selection import train_test_split
from sklearn.compose import ColumnTransformer
from sklearn.ensemble import RandomForestRegressor
# Split data into train and test
datasetX = df_boston.drop(outcome_name, axis=1)
x_train, x_test, y_train, y_test = train_test_split(datasetX,
target,
test_size = 0.2,
random_state=0)
categorical_features = x_train.columns.difference(continuous_features_boston)
# We create the preprocessing pipelines for both numeric and categorical data.
numeric_transformer = Pipeline(steps=[
('scaler', StandardScaler())])
categorical_transformer = Pipeline(steps=[
('onehot', OneHotEncoder(handle_unknown='ignore'))])
transformations = ColumnTransformer(
transformers=[
('num', numeric_transformer, continuous_features_boston),
('cat', categorical_transformer, categorical_features)])
# Append classifier to preprocessing pipeline.
# Now we have a full prediction pipeline.
regr_boston = Pipeline(steps=[('preprocessor', transformations),
('regressor', RandomForestRegressor())])
model_boston = regr_boston.fit(x_train, y_train)
d_boston = dice_ml.Data(dataframe=df_boston, continuous_features=continuous_features_boston, outcome_name=outcome_name)
# We provide the type of model as a parameter (model_type)
m_boston = dice_ml.Model(model=model_boston, backend="sklearn", model_type='regressor')
exp_genetic_boston = Dice(d_boston, m_boston, method="genetic")
```
As we can see below, all the target values will lie in the desired range
```
# Multiple queries can be given as input at once
query_instances_boston = x_train[2:3]
genetic_boston = exp_genetic_boston.generate_counterfactuals(query_instances_boston,
total_CFs=2,
desired_range=[30, 45])
genetic_boston.visualize_as_dataframe(show_only_changes=True)
# Multiple queries can be given as input at once
query_instances_boston = x_train[17:19]
genetic_boston = exp_genetic_boston.generate_counterfactuals(query_instances_boston, total_CFs=4, desired_range=[40, 50])
genetic_boston.visualize_as_dataframe(show_only_changes=True)
```
| github_jupyter |
```
%matplotlib inline
import pyfits
import numpy as np
import matplotlib
matplotlib.rcParams['font.size'] = 15
from matplotlib import pyplot as plt
import sys
sys.path.append('../../')
import photPack2
from astropy.time import Time
import emcee
## Read in raw data, organize
rawch1 = np.genfromtxt('thirdPLD/wasp6_channel1.ascii')
rawch2 = np.genfromtxt('thirdPLD/wasp6_channel2.ascii')
# rawch1 = np.genfromtxt('thirdPLD/wasp6_channel1_binned.ascii')
# rawch2 = np.genfromtxt('thirdPLD/wasp6_channel2_binned.ascii')
ch1 = {}
ch2 = {}
for rawdata, output in zip([rawch1, rawch2], [ch1, ch2]):
for i, key, offset in zip(range(3), ['t', 'f', 'e'], [2450000.0, 0.0, 0.0]):
output[key] = rawdata[:,i] + offset
for ch in [ch1, ch2]:
ch['e'] = np.zeros_like(ch['f']) + np.std(ch['f'][int(0.66*len(ch['f'])):])
## Double check that time offset is what I think it is, by subtracting the start time
## of the observations shown on the Spitzer Heritage Archive to the start time of the data:
# print Time('2013-01-21 08:23:00', format='iso', scale='utc').jd - ch1['t'].min() + 0.00195
def autocorr(x):
result = np.correlate(x, x, mode='full')
return result[result.size/2:]
import sys
sys.path.append('/astro/users/bmmorris/Downloads/Fast_MA')
from ext_func.rsky import rsky
from ext_func.occultquad import occultquad
def get_lc(aRs, i, t0, q1, q2, p0, F0, e, w, period, eps, t):
'''
e - eccentricity
aRs - "a over R-star"
i - inclination angle in radians
u1, u2 - quadratic limb-darkening coeffs
p0 - planet to star radius ratio
w - argument of periapse
period - period
t0 - midtransit (JD)
eps - minimum eccentricity for Kepler's equation
t - time array
'''
u1 = 2*np.sqrt(q1)*q2
u2 = np.sqrt(q1)*(1 - 2*q2)
r_s = 1.0
npoints = len(t)
#calculates separation of centers between the planet and the star
z0 = rsky(e, aRs, i, r_s, w, period, t0, eps, t)
#returns limb darkened model lightcurve
mu_c = occultquad(z0, u1, u2, p0, npoints)
return F0*mu_c
def T14b2aRsi(P, T14, b):
'''
Convert from duration and impact param to a/Rs and inclination
'''
i = np.arccos( ( (P/np.pi)*np.sqrt(1 - b**2)/(T14*b) )**-1 )
aRs = b/np.cos(i)
return aRs, i
def aRsi2T14b(P, aRs, i):
b = aRs*np.cos(i)
T14 = (P/np.pi)*np.sqrt(1-b**2)/aRs
return T14, b
def reparameterized_lc(T14, b, t0, q1, q2, p0, F0, e, w, period, eps, t):
'''
Reparameterization of the transit light curve in `get_lc()` with
duration (first-to-fourth contact) instead of a/R* and impact
parameter instead of inclination
'''
aRs, i = T14b2aRsi(period, T14, b)
return get_lc(aRs, i, t0, q1, q2, p0, F0, e, w, period, eps, t)
from scipy import optimize
aOverRs = 1./0.0932 # Jord`an et al 2013
RpOverRs = 0.1404 # Jord`an et al 2013
eccentricity = 0.0 # Husnoo 2012
inclination = 88.47*np.pi/180
q1 = 0.00001
q2 = 0.2
periapse = np.pi/2 # To match e=0, from Husnoo 2012
period = 3.36100239 # Nikolov 2015 #3.361006
mineccentricity = 1.0e-7
t0_roughfit = 2456918.8793039066
Nbins = 8
Nlightcurves = Nbins + 2
lastp = 0
mosfire_meantimediff = airmass = ch1['t']
ch1_meantimediff = np.median(np.diff(ch1['t']))
ch2_meantimediff = np.median(np.diff(ch2['t']))
#mosfire_exptime = np.median(exposuredurs)/(60*60*24) # convert to units of days from seconds
def fine_lc(T, b, t0, q1, q2, p0, F0, e, w, period, eps, t, meantimediff):
new_t = np.linspace(t.min() - 2*meantimediff, t.max() + 2*meantimediff, 5*len(t))
#return new_t, get_lc(aRs, i, t0, q1, q2, p0, F0, e, w, period, eps, new_t)
return new_t, reparameterized_lc(T, b, t0, q1, q2, p0, F0, e, w, period, eps, new_t)
def binned_lc(T, b, t0_roughfit, q1, q2, RpOverRs, F0, am, eccentricity,
periapse, period, eps, t, meantimediff, airmassvector=airmass):
new_t, finemodel = fine_lc(T, b, t0_roughfit, q1, q2, RpOverRs,
F0, eccentricity, periapse, period, eps, t, meantimediff)
exptime = t[1] - t[0]
timebinedges = np.sort(np.concatenate([t - 0.5*exptime, t + 0.5*exptime]))
d = np.digitize(new_t, timebinedges)
binned_model = np.array([np.mean(finemodel[d == i]) for i in range(1, 2*len(t), 2)])
if airmassvector is None:
return binned_model
else:
return binned_model*(1 + (airmassvector - 1)/am)
def genmodel(parameters, Nbins=Nbins):
# mosfiremodel = np.zeros_like(lightcurve)
listparams = parameters.tolist()
# for eachbin in xrange(Nbins):
# mosfirelcparams = listparams[0:3] + listparams[4:6] + \
# [parameters[10+eachbin], parameters[20+eachbin], np.exp(parameters[30+eachbin]), eccentricity, \
# periapse, period, 1e-7, times, mosfire_meantimediff] # Fixed params
# mosfiremodel[:,eachbin] = binned_lc(*mosfirelcparams)
spitzeram = [np.e] # placeholder argument, ignored
ch1lcparams = listparams[0:2] + [parameters[3]] + listparams[6:8] + \
listparams[18:19] + listparams[28:29] + spitzeram + \
[eccentricity, periapse, period, 1e-7, ch1['t'], ch1_meantimediff]
ch2lcparams = listparams[0:2] + [parameters[3]] + listparams[8:10] + \
listparams[19:20] + listparams[29:30] + spitzeram + \
[eccentricity, periapse, period, 1e-7, ch2['t'], ch2_meantimediff]
ch1model = binned_lc(*ch1lcparams, airmassvector=None)
ch2model = binned_lc(*ch2lcparams, airmassvector=None)
return ch1model, ch2model
#return mosfiremodel, ch1model, ch2model
spitzwhitekernelall_params = np.load('/local/tmp/mosfire/longchains/mosfirespitzer/max_lnp_params_201503040921.npy')
spitzwhitekernelall_params[:2] = aRsi2T14b(period, spitzwhitekernelall_params[0], spitzwhitekernelall_params[1])
ch1model, ch2model = genmodel(spitzwhitekernelall_params)
fig, ax = plt.subplots(2, 2, figsize=(16,12), sharey='row')
ax[0, 0].plot(ch1['t'], ch1['f'] - ch1model,',')
ax[0, 1].plot(ch2['t'], ch2['f'] - ch2model,',')
a1 = autocorr(ch1['f'] - ch1model)[20:]
a2 = autocorr(ch2['f'] - ch2model)
ax[1, 0].plot(a1)
ax[1, 1].plot(a2)
testrs = np.arange(len(a1))
expsquared = lambda r, sig: np.exp(-0.5*r**2/sig**2)
cosine = lambda r, p: np.cos(2*np.pi*r/p)
def kernel(params, r=testrs):
amp, p, sig = params
return amp*expsquared(r, sig)*cosine(r,p)
def errfunc(params, y):
return kernel(params) - y
initP = [0.02, 75, 10000]
bestp = optimize.leastsq(errfunc, initP, args=(a1))[0]
print bestp
ax[1, 0].plot(kernel(bestp))
ax[1, 0].set_ylim([-0.1, 0.1])
ax[1, 0].set_xlim([-10, 600])
ax[1, 1].set_xlim([-10, 600])
ax[0, 0].set_title('ch1')
ax[0, 1].set_title('ch2')
ax[0, 0].set_ylabel('residuals')
ax[1, 0].set_ylabel('autocorrelation')
fig.savefig('secondpld.png')
plt.show()
np.mean(np.diff(ch1['t']))*24*60*60
```
## Bin the data
The chains from Sarah take too long to compute with George. They have 2 second exposures -- who needs that? Bin them down!
```
%matplotlib inline
# Borrow guts from here
def binned_lc(T, b, t0_roughfit, q1, q2, RpOverRs, F0, am, eccentricity,
periapse, period, eps, t, meantimediff, airmassvector=airmass):
new_t, finemodel = fine_lc(T, b, t0_roughfit, q1, q2, RpOverRs,
F0, eccentricity, periapse, period, eps, t, meantimediff)
exptime = t[1] - t[0]
timebinedges = np.sort(np.concatenate([t - 0.5*exptime, t + 0.5*exptime]))
d = np.digitize(new_t, timebinedges)
binned_model = np.array([np.mean(finemodel[d == i]) for i in range(1, 2*len(t), 2)])
if airmassvector is None:
return binned_model
else:
return binned_model*(1 + (airmassvector - 1)/am)
print np.std(np.diff(ch1['t']))*24*60*60
# Mean time difference between points is 2 seconds. Change this to 10 seconds.
t = ch1['t']
f = ch1['f']
def bindata(t, f, binfactor = 6):
originalexptime = t[1] - t[0]
binned_t = np.linspace(t.min()+(binfactor/2)*originalexptime,
t.max()-(binfactor/2)*originalexptime, len(t)/binfactor)
exptime = binned_t[1] - binned_t[0]
timebinedges = np.sort(np.concatenate([binned_t - 0.5*exptime, binned_t + 0.5*exptime]))
d = np.digitize(t, timebinedges)
binned_f = np.array([np.mean(f[d == i]) for i in range(1, 2*len(binned_t), 2)])
return binned_t, binned_f
ch1binned_t, ch1binned_f = bindata(ch1['t'], ch1['f'])
ch2binned_t, ch2binned_f = bindata(ch2['t'], ch2['f'])
print 'Binned by factor: {0}'.format(len(f)/float(len(binned_data)))
print 'New # data points: {0}'.format(len(binned_data))
print 'new apparent exposure time: {0}'.format(np.mean(np.diff(binned_t))*24*60*60)
plt.plot(ch1['t'], ch1['f'], '.')
plt.plot(ch1binned_t, ch1binned_f, 'o')
#plt.xlim([binned_t[100], binned_t[150]])
plt.show()
plt.plot(ch2['t'], ch2['f'], '.')
plt.plot(ch2binned_t, ch2binned_f, 'o')
#plt.xlim([binned_t[100], binned_t[150]])
plt.show()
binned_ch1 = [ch1binned_t, ch1binned_f]
binned_ch2 = [ch2binned_t, ch2binned_f]
fnames = ['wasp6_channel1_binned.ascii', 'wasp6_channel2_binned.ascii']
channels = [binned_ch1, binned_ch2]
for fname, channel in zip(fnames, channels):
with open('thirdPLD/'+fname, 'w') as f:
time, flux = channel
for i in range(len(time)):
f.write('{0} {1} {2}\n'.format(time[i]-2450000.0, flux[i], 0.01))
```
| github_jupyter |
```
import matplotlib.pyplot as plt
import pickle
import numpy as np
from scipy.stats import skewnorm
from ll_xy import lonlat_to_xy
from scipy.stats import linregress
import datetime
import string
import cartopy
import tools
import cartopy.crs as ccrs
import pandas as pd
pd.set_option("mode.chained_assignment", None)
dep_l = pickle.load(open('../pickles/line_depths_dict.p','rb'))
statistics = pickle.load(open('../pickles/statistics.p','rb'))
def compare_magna(df,ax=None,impose_y = False,anno=False,season='all'):
if impose_y: df = df[df['yc'] < 0]
bw = 5
bin_edges = np.arange(0,81,bw)
mean_magna = np.nanmean(df['DepthCm'])
##################################################
bc, ft = tools.depth_distribution_from_depth(mean_magna,bin_edges,statistics=statistics['all'])
hist, bin_edges = np.histogram(df['DepthCm'],bins=bin_edges, density=True)
rmse = np.round(np.sqrt(np.mean(np.square(ft-(hist*bw)))),decimals=2)
if ax == None: fig, ax = plt.subplots(1,1)
if anno:
epoch = df['datetime_ref_1970'].iloc[0]
dt = datetime.datetime.fromtimestamp(epoch)
month = dt.date().month; day = dt.date().day
ax.annotate(text=f'{day}/{month}',xy=(0.95,0.95),ha='right',va='top',xycoords='axes fraction',fontsize='xx-large')
ax.step([0]+list(bc+bw/2),[0]+list(ft),color='r',)
ax.step([0]+list(bc+bw/2), [0]+list(hist*bw),color='b')
alpha=0.5
t = ax.bar(bc, hist*bw,
width=bw,
alpha=alpha,
color='b',
label='Transect')
m = ax.bar(bc, ft,
width=bw,
alpha=alpha,
color='r',
label='Model')
ax.legend(loc='center right')
return(bc, hist*bw, ft)
path = '../MOSAiC_transects/PS122-2_22-92-ANJA_38_Nloop-20200130-UTC-0.csv'
df = pd.read_csv(path)
compare_magna(df, ax=None, impose_y=True,anno=True,season='winter')
MOSAiC_dir = '../MOSAiC_transects'
paths = [f'{MOSAiC_dir}/PS122-1_4-1-magnaprobe-transect-20191024-PS122-1_4-1-20191023-UTC-6.csv',
f'{MOSAiC_dir}/PS122-1_5-27-magnaprobe-transect-20191031-PS122-1_5-27-20191030-UTC-6.csv',
f'{MOSAiC_dir}/PS122-1_6-50-magnaprobe-transect-20191107-PS122-1_6-50-20191106-UTC-6.csv',
f'{MOSAiC_dir}/PS122-1_7-62-magnaprobe-transect-20191114-PS122-1_7-62-20191114-UTC-6.csv',
f'{MOSAiC_dir}/PS122-1_8-58-magnaprobe-transect-20191121-PS122-1_8-58-20191121-UTC-6.csv',
f'{MOSAiC_dir}/PS122-1_9-54-magnaprobe-transect-20191128-PS122-1_9-54-20191128-UTC-6.csv',
f'{MOSAiC_dir}/PS122-1_10-59-magnaprobe-transect-20191205-PS122-1_10-59-20191204-UTC-6.csv',
f'{MOSAiC_dir}/PS122-2_16-83-Katrin_0-20191219-UTC-6.csv',
f'{MOSAiC_dir}/PS122-2_18-80-ANJA_25-20200102-UTC-0.csv',
f'{MOSAiC_dir}/PS122-2_19-110-ANJA_28-20200109-UTC-0.csv',
f'{MOSAiC_dir}/PS122-2_20-95-ANJA_33_cleanN-20200116-UTC-0.csv',
f'{MOSAiC_dir}/PS122-2_22-92-ANJA_38_Nloop-20200130-UTC-0.csv',
f'{MOSAiC_dir}/PS122-2_23-62-ANJA_40_Nloop-20200206-UTC-0.csv',
f'{MOSAiC_dir}/PS122-2_25-119-ANJA_43_Nloop-20200220-UTC-0.csv',
]
per_cm = r'cm$^{-1}$'
fig, axs = plt.subplots(5,3,figsize=(8,15))
thin_snow_fit, thin_snow_obs = [], []
for path, ax in zip(paths, list(axs.reshape(-1))[:2]+list(axs.reshape(-1))[3:]):
df = pd.read_csv(path)
bc, obs, fit = compare_magna(df, ax=ax, impose_y=True,anno=True)
ax.set_ylim(0,0.3)
thin_snow_fit.append(np.mean(fit[0:2]))
thin_snow_obs.append(np.mean(obs[0:2]))
for ax in list(axs.reshape(-1))[-3:]:
ax.set_xlabel('Depth (cm)', fontsize='x-large')
for ax in list(axs.reshape(-1))[::3]:
ax.set_ylabel(f'Probability', fontsize='x-large') # This is probability, not prob dens!
ax = axs.reshape(-1)[2]
for path in paths:
df = pd.read_csv(path)
df = df[df['yc'] < 0]
ax.plot(df['xc'],df['yc'])
ax.set_xlabel('x (m)',fontsize='x-large')
ax.set_ylabel('y (m)',fontsize='x-large',rotation=270,labelpad=15)
ax.yaxis.tick_right()
ax.xaxis.tick_top()
ax.yaxis.set_label_position("right")
ax.xaxis.set_label_position("top")
for counter, ax in enumerate(axs.reshape(-1)):
if counter %3 != 0:
ax.set_yticklabels([])
ax = axs.reshape(-1)[1]
l1 = ax.bar(range(1),range(1),color='b',alpha=0.5)
l2 = ax.bar(range(1),range(1),color='r',alpha=0.5)
ax.legend(handles=[l1,l2],labels=['Transect','NP Model'],
bbox_to_anchor=(0.5,1.25),
loc='center',fontsize='xx-large')
plt.subplots_adjust(wspace=0.08)
plt.savefig('../figures/fig4.png', bbox_inches='tight',dpi=500)
plt.savefig('/home/robbie/Dropbox/Apps/Overleaf/sub-km-snow-depth-dist/fig4.png', bbox_inches='tight',dpi=500)
plt.show()
transect_count = 0
stds = []
mean = []
t_lengths = []
norms = []
dates = []
for key in dep_l:
station = dep_l[key]
for date in station:
transect_count += 1
series = np.array(list(station[date]))
n_series = (series - np.nanmean(series))
std = np.nanstd(n_series)
ns_series = n_series/std
stds.append(std)
mean.append(np.nanmean(series))
norms.append(ns_series)
dates.append(date)
t_lengths.append(len(series[~np.isnan(series)]))
md = []
ms = []
normed = []
for path in paths:
df = pd.read_csv(path)
mean_depth = np.nanmean(df['DepthCm'])
stdv_depth = np.nanstd(df['DepthCm'])
md.append(mean_depth)
ms.append(stdv_depth)
normed.append( (np.array(df['DepthCm']) - mean_depth)/stdv_depth )
stats = np.linalg.lstsq(np.array(mean)[:,np.newaxis],stds)
magna_stats = np.linalg.lstsq(np.array(md)[:,np.newaxis],ms)
magna_linear_prediction = np.array([np.min(md),np.max(md)])*magna_stats[0][0]
fig, (ax1, ax2) = plt.subplots(1,2,figsize=(8,4))
ax1.scatter(md,ms, label='Magnaprobe\nTransects')
linear_prediction = np.array([np.min(md),np.max(md)])*statistics['all']['stats']
ax1.plot([np.min(md),np.max(md)], linear_prediction, color='r', label = 'NP Fit')
# linear_prediction = np.array([np.min(md),np.max(md)])*statistics['winter']['stats']
# ax1.plot([np.min(md),np.max(md)], linear_prediction, color='r', label = 'NP Winter Fit', ls = '--')
ax1.plot([np.min(md),np.max(md)], magna_linear_prediction, color='#1f77b4', label='MOSAiC Fit')
ax1.set_ylabel('Transect Depth\nStandard Deviation (cm)', fontsize='x-large')
ax1.set_xlabel('Transect Mean Depth (cm)', fontsize='x-large')
all_norms = np.concatenate(normed)
bin_edges = np.arange(-3,4,0.5)
bin_centres = np.array([x+0.25 for x in bin_edges[:-1]])
season = 'winter'
fit = skewnorm.pdf(bin_centres,
statistics[season]['a'],
statistics[season]['loc'],
statistics[season]['scale'])
ax2.bar(x=bin_centres,height=fit,width=0.5,alpha=0.5,color='r', label='NP Model')
hist, bc = np.histogram(all_norms, bin_edges,density=True)
ax2.bar(bin_centres, hist,alpha=0.5,width=0.5, label = 'MOSAiC')
bw=0.5
ax2.step([0]+list(bin_centres+bw/2),[0]+list(fit),color='r',alpha=0.5)
ax2.step([0]+list(bin_centres+bw/2), [0]+list(hist),color='b',alpha=0.5)
ax2.set_ylabel(r'Probability Density', fontsize='x-large')
ax2.set_xlabel('Number of Standard\nDeviations From Mean', fontsize='x-large')
ax2.yaxis.tick_right()
ax2.yaxis.set_label_position("right")
print(np.sum(fit)*bw)
print(np.sum(hist)*bw)
ax1.legend()
ax2.legend()
ax1.annotate(text='(a)', fontsize='xx-large', xy=(-0.1,1.05), xycoords= 'axes fraction')
ax2.annotate(text='(b)', fontsize='xx-large', xy=(-0.1,1.05), xycoords= 'axes fraction')
plt.savefig('../figures/fig3.png', bbox_inches='tight',dpi=500)
plt.savefig('/home/robbie/Dropbox/Apps/Overleaf/sub-km-snow-depth-dist/fig3.png', bbox_inches='tight',dpi=500)
plt.plot(thin_snow_fit,marker='o')
plt.plot(thin_snow_obs,marker='o')
errors = np.array(thin_snow_fit) - np.array(thin_snow_obs)
np.mean(errors[:7]), np.mean(thin_snow_obs[:7]),np.mean(thin_snow_fit[:7])
np.mean(errors[7:]), np.mean(thin_snow_obs[7:]),np.mean(thin_snow_fit[7:])
import os
def plot_magna_track(path):
df = pd.read_csv(path)
plt.plot(df['xc'],df['yc'])
plt.show()
magna_dir = '../MOSAiC_transects'
magna_files = os.listdir(magna_dir)
for file in magna_files:
path = magna_dir + '/' + file
print(path)
plot_magna_track(path)
```
| github_jupyter |
## Systematic modelling of surface deformation at active volcanoes
### ethz-02-03-01
This application takes Surface displacement retrieved with DInSAR at active volcanoes to retrieve a first orde estimate of the volume change in the subsurface.
### <a name="service">Service definition
```
service = dict([('title', 'Systematic modelling of surface deformation at active volcanoes'),
('abstract', 'Systematic modelling of surface deformation at active volcanoes'),
('id', 'ewf-ethz-02-03-01')])
coordinates = dict([('id', 'coordinates'),
('title', 'coordinates'),
('abstract', 'Approx Coordinates fo the co-seismic signal (Lat, Lon)'),
('value', '31.614,130.658')])
buffer_aoi = dict([('id', 'buffer_aoi'),
('title', 'buffer_aoi'),
('abstract', 'Buffer AOI (degrees)'),
('value', '0.075')])
downsampling = dict([('id', 'downsampling'),
('title', 'downsampling'),
('abstract', 'Downsampling for speed (0.05-1)'),
('value', '0.2')])
los_angle = dict([('id', 'los_angle'),
('title', 'los_angle'),
('abstract', 'LOS angles of the satellite (incidence33-43 for S1, azimuth, +15 Descending, -15 Ascending)'),
('value', '40,-15')])
_T2Username = dict([('id', '_T2Username'),
('title', 'T2Username'),
('abstract', 'Terradue username'),
('value', '')])
_T2ApiKey = dict([('id', '_T2ApiKey'),
('title', 'T2ApiKey'),
('abstract', 'Terradue api_key'),
('value', '')])
```
### Runtime parameter definition
**Input identifier**
Product identifier
```
input_identifier = ['C858B8C24E1B7ED261F1B0D9AF04A39DB11B1C82']
```
**Input reference**
Catalogue reference
```
input_references = ['https://catalog.terradue.com/better-ethz-02-01-01/search?uid=C858B8C24E1B7ED261F1B0D9AF04A39DB11B1C82']
```
**Data path**
This path defines where the data is staged-in.
```
data_path = '/workspace/data'
```
#### Import the packages required for processing the data
```
import os
import sys
import subprocess
sys.path.append('/application/notebook/libexec/')
sys.path.append(os.getcwd())
from ellip_helpers import create_metadata
import gdal
import cioppy
ciop = cioppy.Cioppy()
```
### Product check
```
creds = '{}:{}'.format(_T2Username['value'],_T2ApiKey['value'])
search = ciop.search(end_point=input_references[0],
params=[('do', 'terradue')],
output_fields='enclosure, startdate, enddate, wkt',
model='EOP',
creds=creds)[0]
enclosure = search['enclosure']
product = os.path.basename(enclosure)
product
product_path = os.path.join(data_path, product)
print 'Searching:', product_path
if os.path.isfile(product_path):
print "Product {} Retrieved".format(product)
else:
raise(Exception("Product {} with reference {} not found in data path {}".format(product, input_references, data_path)))
ciop.copy(product_path, '.')
```
### <a name="workflow">Workflow
```
if 'LD_LIBRARY_PATH' not in os.environ.keys():
os.environ['LD_LIBRARY_PATH'] = '/opt/v94/runtime/glnxa64:/opt/v94/bin/glnxa64:/opt/v94/sys/os/glnxa64:/opt/v94/extern/bin/glnxa64'
else:
os.environ['LD_LIBRARY_PATH'] = '/opt/v94/runtime/glnxa64:/opt/v94/bin/glnxa64:/opt/v94/sys/os/glnxa64:/opt/v94/extern/bin/glnxa64:' + os.environ['LD_LIBRARY_PATH']
import run_inverse_model
```
#### Creating the input_modeling file
```
with open('input_modeling.txt', 'wb') as file:
file.write('{}\n'.format(product))
file.write('{} {}\n'.format(coordinates['value'].split(',')[0], coordinates['value'].split(',')[1]))
file.write('{}\n'.format(buffer_aoi['value']))
file.write('{}\n'.format(downsampling['value']))
file.write('{} {}\n'.format(los_angle['value'].split(',')[0], los_angle['value'].split(',')[1]))
with open('input_modeling.txt') as file:
print file.read()
command = 'import run_inverse_model; mr = run_inverse_model.initialize(); mr.run_inverse_model(\"input_modeling.txt\", nargout=0)'
options = ['python',
'-c',
command,
]
print options
p = subprocess.Popen(options,
stdout=subprocess.PIPE,
stdin=subprocess.PIPE,
stderr=subprocess.PIPE)
res, err = p.communicate()
print(res)
print(err)
```
#### Read the geo_transform and the projection from the input
```
src = gdal.Open(product_path)
geo_transform = src.GetGeoTransform()
projection = src.GetProjection()
src.FlushCache()
os.listdir('./')
```
#### Get the output file list
```
os.remove(product)
#os.remove("input_modeling.txt")
```
#### Create the geo_transform and the projection file txt
```
metadata = {'geo_transform' : geo_transform,
'projection' : projection}
with open('metadata.txt', 'wb') as file:
file.write(str(metadata))
file.close()
with open('metadata.txt') as f:
print f.read()
os.listdir('./')
output_file = list()
for file in os.listdir('.'):
if '.mat' in file or '.txt' in file:
output_file.append(file)
output_file
```
for file in output_file:
ds = gdal.Open(file, gdal.OF_UPDATE)
ds.SetGeoTransform(geo_transform)
ds.SetProjection(projection)
ds.FlushCache()
print gdal.Info(file)
```
metadata = dict()
metadata['startdate'] = search['startdate']
metadata['enddate'] = search['enddate']
metadata['wkt'] = search['wkt']
metadata
for file in output_file:
print os.path.splitext(file)[0]
metadata['identifier'] = os.path.splitext(file)[0]
metadata['title'] = metadata['identifier']
create_metadata(metadata, metadata['identifier'])
os.listdir('./')
```
### License
This work is licenced under a [Attribution-ShareAlike 4.0 International License (CC BY-SA 4.0)](http://creativecommons.org/licenses/by-sa/4.0/)
YOU ARE FREE TO:
* Share - copy and redistribute the material in any medium or format.
* Adapt - remix, transform, and built upon the material for any purpose, even commercially.
UNDER THE FOLLOWING TERMS:
* Attribution - You must give appropriate credit, provide a link to the license, and indicate if changes were made. You may do so in any reasonable manner, but not in any way that suggests the licensor endorses you or your use.
* ShareAlike - If you remix, transform, or build upon the material, you must distribute your contributions under the same license as the original.
| github_jupyter |
```
import re
import urllib
import json
import subprocess
import itertools
import pandas as pd
from bs4 import BeautifulSoup
from multiprocessing import Pool
def get_schools(county, year, grade):
"""Get all the schools in a county for a year and grade"""
url = "https://app.azdhs.gov/IDRReportStats/Home/GetSchoolTable?{0}"
query = {
'bRegex': 'false',
'bRegex_0': 'false',
'bRegex_1': 'false',
'bRegex_2': 'false',
'bRegex_3': 'false',
'bRegex_4': 'false',
'bRegex_5': 'false',
'bRegex_6': 'false',
'bRegex_7': 'false',
'bRegex_8': 'false',
'bSearchable_0': 'false',
'bSearchable_1': 'true',
'bSearchable_2': 'false',
'bSearchable_3': 'false',
'bSearchable_4': 'false',
'bSearchable_5': 'false',
'bSearchable_6': 'true',
'bSearchable_7': 'true',
'bSearchable_8': 'false',
'iColumns': '9',
'iDisplayLength': '2000',
'iDisplayStart': '0',
'mDataProp_0': 'SCHOOL_YEAR',
'mDataProp_1': 'SCHOOL_NAME',
'mDataProp_2': 'SCHOOL_TYPE',
'mDataProp_3': 'SCHOOL_GRADE',
'mDataProp_4': 'ENROLLED',
'mDataProp_5': 'ADDRESS',
'mDataProp_6': 'CITY',
'mDataProp_7': 'ZIP',
'mDataProp_8': 'COUNTY',
'sColumns': ',,,,,,,,',
'sEcho': '1',
'selectedCounty': county,
'selectedGrade': grade,
'selectedYear': year,
}
with subprocess.Popen(['curl', url.format(urllib.parse.urlencode(query))], stdout=subprocess.PIPE) as proc:
schools = json.loads(proc.communicate()[0].decode())['aaData']
return schools
def get_data_from_table(table):
"""Put the html table into a dictionary"""
soup = BeautifulSoup(table, 'html5lib')
data = {
'school type': {'SCHOOL_TYPE': 'N/A'},
'enrolled': {'ENROLLED': 'N/A'},
'medical': {'PCT_MEDICAL_EXEMPT': 'N/A'},
'personal': {'PCT_PBE': 'N/A'},
'every': {'PCT_PBE_EXEMPT_ALL': 'N/A'},
'does': {'HAS_NURSE': 'N/A'},
'nurse type': {'NURSE_TYPE': ''},
'dtap': {'PCT_IMMUNE_DTAP': 'N/A', 'PCT_EXEMPT_DTAP': 'N/A', 'PCT_COMPLIANCE_DTAP': 'N/A'},
'tdap': {'PCT_IMMUNE_TDAP': 'N/A', 'PCT_EXEMPT_TDAP': 'N/A', 'PCT_COMPLIANCE_TDAP': 'N/A'},
'mcv': {'PCT_IMMUNE_MVMVC': 'N/A', 'PCT_EXEMPT_MVMVC': 'N/A', 'PCT_COMPLIANCE_MVMVC': 'N/A'},
'polio': {'PCT_IMMUNE_POLIO': 'N/A', 'PCT_EXEMPT_POLIO': 'N/A', 'PCT_COMPLIANCE_POLIO': 'N/A'},
'mmr': {'PCT_IMMUNE_MMR': 'N/A', 'PCT_EXEMPT_MMR': 'N/A', 'PCT_COMPLIANCE_MMR': 'N/A'},
'hep b': {'PCT_IMMUNE_HEPB': 'N/A', 'PCT_EXEMPT_HEPB': 'N/A', 'PCT_COMPLIANCE_HEPB': 'N/A'},
'hep a': {'PCT_IMMUNE_HEPA': 'N/A', 'PCT_EXEMPT_HEPA': 'N/A', 'PCT_COMPLIANCE_HEPA': 'N/A'},
'hib': {'PCT_IMMUNE_HIB': 'N/A', 'PCT_EXEMPT_HIB': 'N/A', 'PCT_COMPLIANCE_HIB': 'N/A'},
'var': {'PCT_IMMUNE_VAR': 'N/A', 'PCT_EXEMPT_VAR': 'N/A', 'PCT_COMPLIANCE_VAR': 'N/A'},
}
for row in soup.find_all('div', {'class': 'row'}):
key = None
children = list(row.children)
if len(children) <= 1:
continue
key = children[1].text.lower()
for k in data.keys():
if re.search(k, key):
break
else:
continue
cols = data[k]
col_names = list(cols.keys())
index = 0
for child in children[2:]:
try:
text = child.text.lower()
except:
continue
cols[col_names[index]] = text
index += 1
if index == len(col_names):
break
data[k] = cols
return data
def get_school_data(school_name, address, grade, year, county, zipcode, city):
params = {
'paramSelectedAddress': address,
'paramSelectedCity': city,
'paramSelectedGrade': grade,
'paramSelectedSchool': school_name,
'paramSelectedYear': year,
}
cmnd = [
'curl',
'-d',
"{0}".format(urllib.parse.urlencode(params)),
"https://app.azdhs.gov/IDRReportStats/Home/GetSchoolSpecifications",
]
with subprocess.Popen(cmnd, stdout=subprocess.PIPE) as proc:
table = proc.communicate()[0].decode()
try:
data = {
'School': str(school_name),
'Grade': str(grade),
'Address': str(address),
'School Year': str(year),
'Zipcode': str(zipcode),
'County': str(county),
'City': str(city),
}
table_data = get_data_from_table(table)
for value in table_data.values():
data.update(value)
return data
except:
print(f'Failed: {county}, {year}, {grade}, {school}')
raise
def to_csv(vaccines_df):
def create_file_name(n):
return '_'.join(n).replace('-', '_') + '.csv'
columns = {
'Sixth': [
'SCHOOL_NAME',
'SCHOOL_TYPE',
'SCHOOL_ADDRESS_ONE',
'CITY',
'COUNTY',
'ZIP_CODE',
'HAS_NURSE',
'NURSE_TYPE',
'ENROLLED',
'PCT_IMMUNE_DTAP',
'PCT_EXEMPT_DTAP',
'PCT_COMPLIANCE_DTAP',
'PCT_IMMUNE_TDAP',
'PCT_EXEMPT_TDAP',
'PCT_COMPLIANCE_TDAP',
'PCT_IMMUNE_MVMVC',
'PCT_EXEMPT_MVMVC',
'PCT_COMPLIANCE_MVMVC',
'PCT_IMMUNE_POLIO',
'PCT_EXEMPT_POLIO',
'PCT_COMPLIANCE_POLIO',
'PCT_IMMUNE_MMR',
'PCT_EXEMPT_MMR',
'PCT_COMPLIANCE_MMR',
'PCT_IMMUNE_HEPB',
'PCT_EXEMPT_HEPB',
'PCT_COMPLIANCE_HEPB',
'PCT_IMMUNE_VAR',
'PCT_EXEMPT_VAR',
'PCT_COMPLIANCE_VAR',
'PCT_PBE',
'PCT_MEDICAL_EXEMPT',
'PCT_PBE_EXEMPT_ALL',
],
'Childcare': [
'SCHOOL_NAME',
'SCHOOL_TYPE',
'SCHOOL_ADDRESS_ONE',
'CITY',
'COUNTY',
'ZIP_CODE',
'HAS_NURSE',
'NURSE_TYPE',
'ENROLLED',
'PCT_IMMUNE_DTAP',
'PCT_EXEMPT_DTAP',
'PCT_COMPLIANCE_DTAP',
'PCT_IMMUNE_POLIO',
'PCT_EXEMPT_POLIO',
'PCT_COMPLIANCE_POLIO',
'PCT_IMMUNE_MMR',
'PCT_EXEMPT_MMR',
'PCT_COMPLIANCE_MMR',
'PCT_IMMUNE_HIB',
'PCT_EXEMPT_HIB',
'PCT_COMPLIANCE_HIB',
'PCT_IMMUNE_HEPA',
'PCT_EXEMPT_HEPA',
'PCT_COMPLIANCE_HEPA',
'PCT_IMMUNE_HEPB',
'PCT_EXEMPT_HEPB',
'PCT_COMPLIANCE_HEPB',
'PCT_IMMUNE_VAR',
'PCT_EXEMPT_VAR',
'PCT_COMPLIANCE_VAR',
'PCT_PBE',
'PCT_MEDICAL_EXEMPT',
'PCT_PBE_EXEMPT_ALL'
],
'Kindergarten': [
'SCHOOL_NAME',
'SCHOOL_TYPE',
'SCHOOL_ADDRESS_ONE',
'CITY',
'COUNTY',
'ZIP_CODE',
'HAS_NURSE',
'NURSE_TYPE',
'ENROLLED',
'PCT_IMMUNE_DTAP',
'PCT_EXEMPT_DTAP',
'PCT_COMPLIANCE_DTAP',
'PCT_IMMUNE_POLIO',
'PCT_EXEMPT_POLIO',
'PCT_COMPLIANCE_POLIO',
'PCT_IMMUNE_MMR',
'PCT_EXEMPT_MMR',
'PCT_COMPLIANCE_MMR',
'PCT_IMMUNE_HEPB',
'PCT_EXEMPT_HEPB',
'PCT_COMPLIANCE_HEPB',
'PCT_IMMUNE_VAR',
'PCT_EXEMPT_VAR',
'PCT_COMPLIANCE_VAR',
'PCT_PBE',
'PCT_MEDICAL_EXEMPT',
'PCT_PBE_EXEMPT_ALL'
]
}
group_by = ['Grade', 'School Year']
for name, group in vaccines_df.groupby(group_by):
grade, year = name
cols = columns[grade]
df = pd.DataFrame(group)[cols]
df.sort_values(by=['SCHOOL_NAME'], inplace=True)
df.reset_index(drop=True, inplace=True)
df.to_csv(create_file_name(name), index=False)
grades = ['Childcare', 'Kindergarten', 'Sixth']
years = ['2010-2011', '2011-2012', '2012-2013', '2013-2014', '2014-2015', '2015-2016', '2016-2017']
counties = [
'Apache',
'Cochise',
'Coconino',
'Gila',
'Graham',
'Greenlee',
'La Paz',
'Maricopa',
'Mohave',
'Navajo',
'Pima',
'Pinal',
'Santa Cruz',
'Yavapai',
'Yuma',
]
with Pool(processes=7) as pool:
all_schools = pool.starmap(get_schools, itertools.product(counties, years, grades))
schools = [school for school_list in all_schools for school in school_list]
args = []
for school in schools:
school_name = school['SCHOOL_NAME']
address = school['ADDRESS']
grade = school['SCHOOL_GRADE']
year = school['SCHOOL_YEAR']
county = school['COUNTY']
zipcode = school['ZIP']
city = school['CITY']
args.append((school_name, address, grade, year, county, zipcode, city))
with Pool(processes=7) as pool:
vaccines = pool.starmap(get_school_data, args)
vaccines_df = pd.DataFrame(vaccines)
names = {
'Address': 'SCHOOL_ADDRESS_ONE',
'City': 'CITY',
'County': 'COUNTY',
'School': 'SCHOOL_NAME',
'Zipcode': 'ZIP_CODE'
}
vaccines_df.rename(index=str, columns=names, inplace=True)
to_csv(vaccines_df)
```
| github_jupyter |
```
!git clone https://github.com/broadinstitute/raman_classifier_challenge.git
import pandas as pd
import numpy as np
from scipy import stats
import matplotlib.pyplot as plt
import seaborn as sns
df= pd.read_csv('raman_classifier_challenge/data/raman_data.csv')
df.describe()
df.dtypes
len(df)
df.head()
# Dataset contains almost equal samples of all class
#df.hist(figsize = (200,200))
condition_tmpdf= df.iloc[:,0]
features_tmpdf= df.iloc[:,1:]
features_tmpdf.boxplot()
#filtering corresponding condition values
condition_tmpdf= condition_tmpdf[(np.abs(stats.zscore(features_tmpdf)) < 4).all(axis=1)]
#removing outliers from features
features_tmpdf= features_tmpdf[(np.abs(stats.zscore(features_tmpdf)) < 4).all(axis=1)]
mlist=[]
for i in features_tmpdf.columns:
if float(i) <=1800.0 and float(i) >=800.0:
mlist.append(i)
print(mlist)
print(len(mlist))
features_tmpdf= features_tmpdf[mlist]
features_tmpdf
```
#### I was trying averaging the features, didnt work
```
# mlist=[[] for i in range(40)]
# for i in features_tmpdf.columns:
# ind= (int(float(i)/100))
# mlist[ind].append(i)
# #mlist
# f_tmpdf= pd.DataFrame()
# for i in mlist:
# if i:
# colname= str(int(float(i[0])/100))
# ar= features_tmpdf[i].mean(axis=1)
# features_tmpdf= features_tmpdf.drop(features_tmpdf[i], axis=1)
# f_tmpdf[colname] = ar
# features_tmpdf= f_tmpdf
# features_tmpdf.head()
#features with pretty much outliers removed
features_tmpdf.boxplot()
tmerg= pd.concat([condition_tmpdf, features_tmpdf], axis=1)
tmerg
#I was hoping each concencetration would have different area for spectrometry reading but it was not so.
cdf= condition_tmpdf.copy()
cdf= cdf.replace('0mM', 'red')
cdf= cdf.replace('0.1mM', 'green')
cdf= cdf.replace('0.5mM', 'blue')
cdf= cdf.replace('1mM', 'yellow')
features_tmpdf.T.iloc[:,:].plot(color=cdf)
```
## scale features
```
from sklearn import preprocessing
x = features_tmpdf.values
min_max_scaler = preprocessing.MinMaxScaler()
x_scaled = min_max_scaler.fit_transform(x)
scaled_features_tmpdf= pd.DataFrame(x_scaled)
scaled_features_tmpdf
```
## Encode Classes
```
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import OneHotEncoder
x= condition_tmpdf.values
label_encoder = LabelEncoder()
integer_encoded = label_encoder.fit_transform(x)
print(integer_encoded[:5])
onehot_encoder = OneHotEncoder(sparse=False)
integer_encoded = integer_encoded.reshape(len(integer_encoded), 1)
onehot_encoded = onehot_encoder.fit_transform(integer_encoded)
print(onehot_encoded[:5])
onehot_condition_tmpdf= onehot_encoded
# verifying if both are same length
print(len(onehot_condition_tmpdf))
print(len(scaled_features_tmpdf))
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import chi2
from sklearn.preprocessing import MinMaxScaler
X_norm = MinMaxScaler().fit_transform(features_tmpdf)
chi_selector = SelectKBest(chi2, k=30)
chi_selector.fit(X_norm, integer_encoded)
chi_support = chi_selector.get_support()
chi_feature = features_tmpdf.loc[:,chi_support].columns.tolist()
print(str(len(chi_feature)), 'selected features')
chi2_features_tmpdf= features_tmpdf.loc[:, chi_support]
```
## PCA
```
from sklearn.decomposition import PCA
pca = PCA(n_components=30)
principalComponents = pca.fit_transform(scaled_features_tmpdf)
principalDf = pd.DataFrame(data = principalComponents)
print(principalDf.head())
print(len(condition_tmpdf))
# finalDf = pd.concat([principalDf, condition_tmpdf], axis = 1)
finalDf= principalDf.copy()
finalDf['condition']= condition_tmpdf
print(finalDf.head())
```
#### Let's see first what amount of variance does each PC explain.
```
print(pca.explained_variance_ratio_)
```
PC1 explains 97.3% and PC2 1.4%. Together, if we keep PC1 and PC2 only, they explain 98.7%. Now lets look at the important features.
```
print(abs( pca.components_ ))
#Selecting PC1
yaxis= pca.components_[0]
from collections import OrderedDict
od = OrderedDict()
for i in range(len(yaxis)):
od[i]= yaxis[i]
od2= OrderedDict(sorted(od.items(), key=lambda t: t[1], reverse=True))
```
#### The ordered dict below contains all the features sorted in order of their importance, like feature 299th (feature indexing starts from 0) has the most importance.
```
print(od2)
```
Here, pca.components_ has shape [n_components, n_features]. Thus, by looking at the PC1 (First Principal Component) which is the first row: [0.04295346 0.04492502 0.04503862 ... 0.02696454 0.02666117 0.02636576] we can conclude that feature 299, 91 and 305 are the most important.
```
#Selecting PC2
yaxis= pca.components_[1]
od = OrderedDict()
for i in range(len(yaxis)):
od[i]= yaxis[i]
od2= OrderedDict(sorted(od.items(), key=lambda t: t[1], reverse=True))
print(od2)
```
##### Similarly for PC2, feature at 481, 482, 480th column have the most importance
```
len(principalDf)
```
### Creating a features dictionary so it will be easy to traverse features when training a model.
```
featuredict={
'pca':{
'X':principalDf
},
'chi2':{
'X':chi2_features_tmpdf
}
}
```
## Train Test Split
```
from sklearn.model_selection import train_test_split
```
#### As the number of rows are less, I only took 10% of them for the test set and 90% for train set.
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(scaled_features_tmpdf, onehot_condition_tmpdf, test_size=0.1, random_state=42)
X_train2, X_test2, y_train2, y_test2 = train_test_split(scaled_features_tmpdf, integer_encoded, test_size=0.1, random_state=42)
X_train3, X_test3, y_train3, y_test3 = train_test_split(principalDf, integer_encoded, test_size=0.1, random_state=42)
X_train4, X_test4, y_train4, y_test4 = train_test_split(principalDf, onehot_condition_tmpdf, test_size=0.1, random_state=42)
```
## SVM
achieved 66% accuracy
```
featuredict['chi2']['X']
from sklearn import svm
#linear kernel works the best. I also tried changing C and gamma but no improvrmrnt
for i in featuredict.keys():
print(i)
X= featuredict[i]['X']
X_train, X_test, y_train, y_test = train_test_split(X, integer_encoded, test_size=0.1, random_state=42)
# print(X_train)
clf = svm.SVC(kernel='linear')
clf.fit(X_train, y_train)
print(clf.score(X_test, y_test))
clf = svm.SVC(kernel='rbf')
clf.fit(X_train3, y_train)
print(clf.score(X_test, y_test))
clf = svm.SVC(kernel='poly')
clf.fit(X_train, y_train)
print(clf.score(X_test, y_test))
print("===============================")
```
61% accuracy with ridge classifier
```
from pandas.core.common import flatten
from sklearn.metrics import accuracy_score, classification_report
from sklearn.linear_model import RidgeClassifier
for i in featuredict.keys():
print(i)
X= featuredict[i]['X']
X_train, X_test, y_train, y_test = train_test_split(X, integer_encoded, test_size=0.1, random_state=42)
clf = RidgeClassifier().fit(X_train, y_train)
print(clf.score(X_test, y_test))
y_pred= clf.predict(X_test)
print(classification_report(y_test, y_pred))
print("==================================================")
```
## Random Forest
77.8% accuracy
```
from sklearn.ensemble import RandomForestClassifier
for i in featuredict.keys():
print(i)
X= featuredict[i]['X']
X_train, X_test, y_train, y_test = train_test_split(X, integer_encoded, test_size=0.1, random_state=42)
rfc = RandomForestClassifier(max_depth=10, criterion = 'entropy', random_state = 0)
#Fit model on the training Data
rfc.fit(X_train, y_train)
print(rfc.score(X_test, y_test))
print("==============================")
# Build Neural Network
from keras.layers import Dense
from keras import Sequential
from keras.constraints import maxnorm
from keras.layers import Dropout
from keras.optimizers import SGD
from keras import optimizers
from keras import metrics
from keras.losses import CategoricalCrossentropy, KLDivergence, SparseCategoricalCrossentropy
hidden_units=512
learning_rate=0.0002 #Learning rate was quite optimal
hidden_layer_act='relu'
output_layer_act='softmax'
no_epochs=1000 #Increasing The epochs would overfit
bsize = 16 #Batch Size Of 128
def create_network(optimizer='rmsprop', init_mode='uniform', lossfns='categorical_crossentropy'):
model = Sequential()
model.add(Dense(hidden_units, kernel_initializer=init_mode, input_shape=(30, ), activation=hidden_layer_act))
model.add(Dropout(0.1))
model.add(Dense(256, kernel_initializer=init_mode, activation=hidden_layer_act))
model.add(Dropout(0.1))
model.add(Dense(128, kernel_initializer=init_mode, activation=hidden_layer_act))
model.add(Dense(64, kernel_initializer=init_mode, activation=hidden_layer_act))
model.add(Dense(4, kernel_initializer=init_mode, activation=output_layer_act))
model.compile(loss=lossfns, optimizer=optimizer, metrics = ["accuracy"])#metrics.categorical_accuracy
return model
# model.fit(X_train, y_train, epochs=no_epochs, batch_size= bsize, verbose=2)
# model.evaluate(x=X_test, y=y_test, batch_size=bsize)
from keras.wrappers.scikit_learn import KerasClassifier
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import accuracy_score, confusion_matrix, precision_recall_fscore_support
# Wrap Keras model so it can be used by scikit-learn
neural_network = KerasClassifier(build_fn=create_network,
epochs=no_epochs,
batch_size=bsize,
verbose=0)
from keras.losses import CategoricalCrossentropy, KLDivergence, SparseCategoricalCrossentropy
```
### After googling a bit I finalized 3 popular loss functions for Multi Class Classification
##### Multi-Class Cross-Entropy Loss
##### Sparse Multiclass Cross-Entropy Loss
##### Kullback Leibler Divergence Loss
```
from sklearn.model_selection import GridSearchCV
# define the grid search parameters
init_mode = ['uniform', 'lecun_uniform', 'normal', 'zero',
'glorot_normal', 'glorot_uniform', 'he_normal', 'he_uniform']
lossfns = ['categorical_crossentropy', 'kl_divergence', 'sparse_categorical_crossentropy']
param_grid = dict(init_mode=init_mode, lossfns=lossfns)
grid = GridSearchCV(estimator=neural_network, param_grid=param_grid, n_jobs=-1, cv=3)
for i in featuredict.keys():
print(i)
X= featuredict[i]['X']
X_train, X_test, y_train, y_test = train_test_split(X, onehot_condition_tmpdf, test_size=0.1, random_state=42)
grid_result = grid.fit(X_train, y_train)
# print results
print(f'Best Accuracy for {grid_result.best_score_} using {grid_result.best_params_}')
means = grid_result.cv_results_['mean_test_score']
stds = grid_result.cv_results_['std_test_score']
params = grid_result.cv_results_['params']
for mean, stdev, param in zip(means, stds, params):
print(f' mean={mean:.4}, std={stdev:.4} using {param}')
```
We see that the best results are obtained from the model using he_uniform initialization and kl_divergence loss function, which is close to 76%.
```
from sklearn.metrics import accuracy_score, confusion_matrix, precision_recall_fscore_support
# we choose the initializers that came at the top in our previous cross-validation!!
init_mode = ['he_uniform']
lossfns = ['categorical_crossentropy']
batches = [16, 32 ]
epochs = [100, 300, 500]
model_init_batch_epoch_CV = KerasClassifier(build_fn=create_network, verbose=0)
# grid search for initializer, batch size and number of epochs
param_grid = dict(epochs=epochs, batch_size=batches, init_mode=init_mode)
grid = GridSearchCV(estimator=model_init_batch_epoch_CV,
param_grid=param_grid,
cv=3)
for i in featuredict.keys():
print(i)
X= featuredict[i]['X']
X_train, X_test, y_train, y_test = train_test_split(X, onehot_condition_tmpdf, test_size=0.1, random_state=42)
grid_result = grid.fit(X_train, y_train)
# print results
print(f'Best Accuracy for {grid_result.best_score_:.4} using {grid_result.best_params_}')
means = grid_result.cv_results_['mean_test_score']
stds = grid_result.cv_results_['std_test_score']
params = grid_result.cv_results_['params']
for mean, stdev, param in zip(means, stds, params):
print(f'mean={mean:.4}, std={stdev:.4} using {param}')
```
#### using the required parameters which gave good accuracy, now doing kfold using those parameters.
```
# Wrap Keras model so it can be used by scikit-learn
neural_network = KerasClassifier(build_fn=create_network,
epochs=300,
batch_size=32,
verbose=0)
X_train, X_test, y_train, y_test = train_test_split(featuredict['pca']['X'], onehot_condition_tmpdf, test_size=0.1, random_state=42)
out= cross_val_score(neural_network, X_train, y_train, cv=10)
out.mean()
neural_network.fit(X_train, y_train)
y_pred=neural_network.predict(X_test)
y_test_inv=onehot_encoder.inverse_transform(y_test)
print(classification_report(y_test_inv, y_pred))
cm= confusion_matrix(y_test_inv, y_pred)
labels= label_encoder.inverse_transform([0,1,2,3])
# Transform to df for easier plotting
cm_df = pd.DataFrame(cm,
index = labels,
columns = labels)
plt.figure(figsize=(5.5,4))
sns.heatmap(cm_df, annot=True)
plt.title('NN \nAccuracy:{0:.3f}'.format(accuracy_score(y_test_inv, y_pred)))
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.show()
```
### The confusion matrix won't tell much because there are very less entries and out of those some labels are selected more. However we can see using KFold that our accuracy average is around 75-77%. Random Forest also gave similar results. Machine learning models scale with data, so I think with more data the performance of my model will give more promising results. Now 100-200 rows are very few generally to train deeplearning models.
| github_jupyter |
# EJERCICIO 7
A partir de análisis clínicos y de la edad y el sexo de pacientes de una clínica ubicada en el noreste de Andhra Pradesh, India, se intentará obtener un clasificador automático que sirva para diagnosticar a pacientes con problemas de hígado.
Para esto, se recabaron muestras de ocho análisis distintos realizados a 579 pacientes que, junto con su edad y sexo, se dividieron en dos grupos: 414 de ellos diagnosticados con problemas de hígado por expertos en el área mientras que los 165 restantes fueron señalados como exentos de ese problema.
Los 11 atributos que constituyen una muestra son los
indicados en la tabla de la derecha. Todos son atributos son
valores numéricos continuos a excepción del atributo “Sexo”,
en donde el valor 1 representa “HOMBRE” y el valor 2
representa “MUJER”, y del atributo “Diagnóstico”, donde el valor 1 representa “CON PROBLEMA DE HÍGADO” mientras que el valor 2 representa “SIN PROBLEMA DE HÍGADO”.
Utilice perceptrones o una red neuronal artificial (según resulte más conveniente). Informe el motivo por el que se eligió el tipo de clasificador. Detalle la arquitectura y los parámetros usados en su entrenamiento (según corresponda). Documente todos los intentos realizados.
Para el entrenamiento emplee sólo el 90% de las muestras disponibles de cada tipo. Informe la matriz de confusión que produce el mejor clasificador obtenido al evaluarlo con las muestras de entrenamiento e indique la matriz que ese clasificador produce al usarlo sobre el resto de las muestras reservadas para prueba.
$$
\begin{array}{|c|c|}
\hline 1 & Edad \\
\hline 2 & Sexo \\
\hline 3 & Bilirrubina Total \\
\hline 4 & Bilirrubina Directa \\
\hline 5 & Fosfatasa Alcalina \\
\hline 6 & Alanina Aminotransferasa \\
\hline 7 & Aspartato Aminotransferasa \\
\hline 8 & Proteínas Total \\
\hline 9 & Albúmina \\
\hline 10 & Relación Albúmina/Globulina \\
\hline 11 & Diagnóstico (valor a predecir) \\
\hline
\end{array}
$$
```
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
import mpld3
%matplotlib inline
mpld3.enable_notebook()
from cperceptron import Perceptron
from cbackpropagation import ANN #, Identidad, Sigmoide
import patrones as magia
def progreso(ann, X, T, y=None, n=-1, E=None):
if n % 20 == 0:
print("Pasos: {0} - Error: {1:.32f}".format(n, E))
def progresoPerceptron(perceptron, X, T, n):
y = perceptron.evaluar(X)
incorrectas = (T != y).sum()
print("Pasos: {0}\tIncorrectas: {1}\n".format(n, incorrectas))
#Cargo datos
higado = np.load('higado.npy')
#muestras = higado[:, :-1]
muestras = np.hstack((higado[:,0].reshape(-1,1), higado[:,2:-1]))
# Atributo 'sexo':
# muestras[:, 1] == 1 --> HOMBRE
# muestras[:, 1] == 2 --> MUJER
diagnostico = (higado[:, -1] != 2).astype(np.int8)
# diagnostico == 1 --> CON PROBLEMA DE HÍGADO
# diagnostico == 0 --> SIN PROBLEMA DE HÍGADO
#Armo Patrones
clases, patronesEnt, patronesTest = magia.generar_patrones(magia.escalar(muestras),diagnostico,90)
X, T = magia.armar_patrones_y_salida_esperada(clases,patronesEnt)
Xtest, Ttest = magia.armar_patrones_y_salida_esperada(clases,patronesTest)
```
## Intento con Perceptrones, pero no funciona
```
# Esto es para poder usar Cython y que sea mas rapido
TT = T[:,0].copy(order='C')
TT = TT.astype(np.int8)
#Entrenamiento
p1 = Perceptron(X.shape[1])
p1.reiniciar()
I1 = p1.entrenar(X, TT, max_pasos=100000, callback=progresoPerceptron, frecuencia_callback=50000)
print("Pasos:{0}".format(I1))
#Evaluo
print("Errores:{0} de {1}\n".format((p1.evaluar(Xtest) != Ttest[:,0]).sum(), Ttest.shape[0]))
```
## Ahora intento con BackPropagation
```
# Crea la red neuronal
ocultas = 20 #2,3,5,10,20
entradas = X.shape[1]
salidas = T.shape[1]
ann = ANN(entradas, ocultas, salidas)
ann.reiniciar()
#Entreno
E, n = ann.entrenar_rprop(X, T, min_error=0, max_pasos=100000, callback=progreso, frecuencia_callback=50000)
print("\nRed entrenada en {0} pasos con un error de {1:.32f}".format(n, E))
Y = (ann.evaluar(Xtest) >= 0.5).astype(np.float32)
magia.matriz_de_confusion(Ttest,Y)
```
| github_jupyter |
# Cheat Sheet: Writing Python 2-3 compatible code
- **Copyright (c):** 2013-2015 Python Charmers Pty Ltd, Australia.
- **Author:** Ed Schofield.
- **Licence:** Creative Commons Attribution.
A PDF version is here: http://python-future.org/compatible_idioms.pdf
This notebook shows you idioms for writing future-proof code that is compatible with both versions of Python: 2 and 3. It accompanies Ed Schofield's talk at PyCon AU 2014, "Writing 2/3 compatible code". (The video is here: <http://www.youtube.com/watch?v=KOqk8j11aAI&t=10m14s>.)
Minimum versions:
- Python 2: 2.6+
- Python 3: 3.3+
## Setup
The imports below refer to these ``pip``-installable packages on PyPI:
import future # pip install future
import builtins # pip install future
import past # pip install future
import six # pip install six
The following scripts are also ``pip``-installable:
futurize # pip install future
pasteurize # pip install future
See http://python-future.org and https://pythonhosted.org/six/ for more information.
## Essential syntax differences
### print
```
# Python 2 only:
print 'Hello'
# Python 2 and 3:
print('Hello')
```
To print multiple strings, import ``print_function`` to prevent Py2 from interpreting it as a tuple:
```
# Python 2 only:
print 'Hello', 'Guido'
# Python 2 and 3:
from __future__ import print_function # (at top of module)
print('Hello', 'Guido')
# Python 2 only:
print >> sys.stderr, 'Hello'
# Python 2 and 3:
from __future__ import print_function
print('Hello', file=sys.stderr)
# Python 2 only:
print 'Hello',
# Python 2 and 3:
from __future__ import print_function
print('Hello', end='')
```
### Raising exceptions
```
# Python 2 only:
raise ValueError, "dodgy value"
# Python 2 and 3:
raise ValueError("dodgy value")
```
Raising exceptions with a traceback:
```
# Python 2 only:
traceback = sys.exc_info()[2]
raise ValueError, "dodgy value", traceback
# Python 3 only:
raise ValueError("dodgy value").with_traceback()
# Python 2 and 3: option 1
from six import reraise as raise_
# or
from future.utils import raise_
traceback = sys.exc_info()[2]
raise_(ValueError, "dodgy value", traceback)
# Python 2 and 3: option 2
from future.utils import raise_with_traceback
raise_with_traceback(ValueError("dodgy value"))
```
Exception chaining (PEP 3134):
```
# Setup:
class DatabaseError(Exception):
pass
# Python 3 only
class FileDatabase:
def __init__(self, filename):
try:
self.file = open(filename)
except IOError as exc:
raise DatabaseError('failed to open') from exc
# Python 2 and 3:
from future.utils import raise_from
class FileDatabase:
def __init__(self, filename):
try:
self.file = open(filename)
except IOError as exc:
raise_from(DatabaseError('failed to open'), exc)
# Testing the above:
try:
fd = FileDatabase('non_existent_file.txt')
except Exception as e:
assert isinstance(e.__cause__, IOError) # FileNotFoundError on Py3.3+ inherits from IOError
```
### Catching exceptions
```
# Python 2 only:
try:
...
except ValueError, e:
...
# Python 2 and 3:
try:
...
except ValueError as e:
...
```
### Division
Integer division (rounding down):
```
# Python 2 only:
assert 2 / 3 == 0
# Python 2 and 3:
assert 2 // 3 == 0
```
"True division" (float division):
```
# Python 3 only:
assert 3 / 2 == 1.5
# Python 2 and 3:
from __future__ import division # (at top of module)
assert 3 / 2 == 1.5
```
"Old division" (i.e. compatible with Py2 behaviour):
```
# Python 2 only:
a = b / c # with any types
# Python 2 and 3:
from past.utils import old_div
a = old_div(b, c) # always same as / on Py2
```
### Long integers
Short integers are gone in Python 3 and ``long`` has become ``int`` (without the trailing ``L`` in the ``repr``).
```
# Python 2 only
k = 9223372036854775808L
# Python 2 and 3:
k = 9223372036854775808
# Python 2 only
bigint = 1L
# Python 2 and 3
from builtins import int
bigint = int(1)
```
To test whether a value is an integer (of any kind):
```
# Python 2 only:
if isinstance(x, (int, long)):
...
# Python 3 only:
if isinstance(x, int):
...
# Python 2 and 3: option 1
from builtins import int # subclass of long on Py2
if isinstance(x, int): # matches both int and long on Py2
...
# Python 2 and 3: option 2
from past.builtins import long
if isinstance(x, (int, long)):
...
```
### Octal constants
```
0644 # Python 2 only
0o644 # Python 2 and 3
```
### Backtick repr
```
`x` # Python 2 only
repr(x) # Python 2 and 3
```
### Metaclasses
```
class BaseForm(object):
pass
class FormType(type):
pass
# Python 2 only:
class Form(BaseForm):
__metaclass__ = FormType
pass
# Python 3 only:
class Form(BaseForm, metaclass=FormType):
pass
# Python 2 and 3:
from six import with_metaclass
# or
from future.utils import with_metaclass
class Form(with_metaclass(FormType, BaseForm)):
pass
```
## Strings and bytes
### Unicode (text) string literals
If you are upgrading an existing Python 2 codebase, it may be preferable to mark up all string literals as unicode explicitly with ``u`` prefixes:
```
# Python 2 only
s1 = 'The Zen of Python'
s2 = u'きたないのよりきれいな方がいい\n'
# Python 2 and 3
s1 = u'The Zen of Python'
s2 = u'きたないのよりきれいな方がいい\n'
```
The ``futurize`` and ``python-modernize`` tools do not currently offer an option to do this automatically.
If you are writing code for a new project or new codebase, you can use this idiom to make all string literals in a module unicode strings:
```
# Python 2 and 3
from __future__ import unicode_literals # at top of module
s1 = 'The Zen of Python'
s2 = 'きたないのよりきれいな方がいい\n'
```
See http://python-future.org/unicode_literals.html for more discussion on which style to use.
### Byte-string literals
```
# Python 2 only
s = 'This must be a byte-string'
# Python 2 and 3
s = b'This must be a byte-string'
```
To loop over a byte-string with possible high-bit characters, obtaining each character as a byte-string of length 1:
```
# Python 2 only:
for bytechar in 'byte-string with high-bit chars like \xf9':
...
# Python 3 only:
for myint in b'byte-string with high-bit chars like \xf9':
bytechar = bytes([myint])
# Python 2 and 3:
from builtins import bytes
for myint in bytes(b'byte-string with high-bit chars like \xf9'):
bytechar = bytes([myint])
```
As an alternative, ``chr()`` and ``.encode('latin-1')`` can be used to convert an int into a 1-char byte string:
```
# Python 3 only:
for myint in b'byte-string with high-bit chars like \xf9':
char = chr(myint) # returns a unicode string
bytechar = char.encode('latin-1')
# Python 2 and 3:
from builtins import bytes, chr
for myint in bytes(b'byte-string with high-bit chars like \xf9'):
char = chr(myint) # returns a unicode string
bytechar = char.encode('latin-1') # forces returning a byte str
```
### basestring
```
# Python 2 only:
a = u'abc'
b = 'def'
assert (isinstance(a, basestring) and isinstance(b, basestring))
# Python 2 and 3: alternative 1
from past.builtins import basestring # pip install future
a = u'abc'
b = b'def'
assert (isinstance(a, basestring) and isinstance(b, basestring))
# Python 2 and 3: alternative 2: refactor the code to avoid considering
# byte-strings as strings.
from builtins import str
a = u'abc'
b = b'def'
c = b.decode()
assert isinstance(a, str) and isinstance(c, str)
# ...
```
### unicode
```
# Python 2 only:
templates = [u"blog/blog_post_detail_%s.html" % unicode(slug)]
# Python 2 and 3: alternative 1
from builtins import str
templates = [u"blog/blog_post_detail_%s.html" % str(slug)]
# Python 2 and 3: alternative 2
from builtins import str as text
templates = [u"blog/blog_post_detail_%s.html" % text(slug)]
```
### StringIO
```
# Python 2 only:
from StringIO import StringIO
# or:
from cStringIO import StringIO
# Python 2 and 3:
from io import BytesIO # for handling byte strings
from io import StringIO # for handling unicode strings
```
## Imports relative to a package
Suppose the package is:
mypackage/
__init__.py
submodule1.py
submodule2.py
and the code below is in ``submodule1.py``:
```
# Python 2 only:
import submodule2
# Python 2 and 3:
from . import submodule2
# Python 2 and 3:
# To make Py2 code safer (more like Py3) by preventing
# implicit relative imports, you can also add this to the top:
from __future__ import absolute_import
```
## Dictionaries
```
heights = {'Fred': 175, 'Anne': 166, 'Joe': 192}
```
### Iterating through ``dict`` keys/values/items
Iterable dict keys:
```
# Python 2 only:
for key in heights.iterkeys():
...
# Python 2 and 3:
for key in heights:
...
```
Iterable dict values:
```
# Python 2 only:
for value in heights.itervalues():
...
# Idiomatic Python 3
for value in heights.values(): # extra memory overhead on Py2
...
# Python 2 and 3: option 1
from builtins import dict
heights = dict(Fred=175, Anne=166, Joe=192)
for key in heights.values(): # efficient on Py2 and Py3
...
# Python 2 and 3: option 2
from builtins import itervalues
# or
from six import itervalues
for key in itervalues(heights):
...
```
Iterable dict items:
```
# Python 2 only:
for (key, value) in heights.iteritems():
...
# Python 2 and 3: option 1
for (key, value) in heights.items(): # inefficient on Py2
...
# Python 2 and 3: option 2
from future.utils import viewitems
for (key, value) in viewitems(heights): # also behaves like a set
...
# Python 2 and 3: option 3
from future.utils import iteritems
# or
from six import iteritems
for (key, value) in iteritems(heights):
...
```
### dict keys/values/items as a list
dict keys as a list:
```
# Python 2 only:
keylist = heights.keys()
assert isinstance(keylist, list)
# Python 2 and 3:
keylist = list(heights)
assert isinstance(keylist, list)
```
dict values as a list:
```
# Python 2 only:
heights = {'Fred': 175, 'Anne': 166, 'Joe': 192}
valuelist = heights.values()
assert isinstance(valuelist, list)
# Python 2 and 3: option 1
valuelist = list(heights.values()) # inefficient on Py2
# Python 2 and 3: option 2
from builtins import dict
heights = dict(Fred=175, Anne=166, Joe=192)
valuelist = list(heights.values())
# Python 2 and 3: option 3
from future.utils import listvalues
valuelist = listvalues(heights)
# Python 2 and 3: option 4
from future.utils import itervalues
# or
from six import itervalues
valuelist = list(itervalues(heights))
```
dict items as a list:
```
# Python 2 and 3: option 1
itemlist = list(heights.items()) # inefficient on Py2
# Python 2 and 3: option 2
from future.utils import listitems
itemlist = listitems(heights)
# Python 2 and 3: option 3
from future.utils import iteritems
# or
from six import iteritems
itemlist = list(iteritems(heights))
```
## Custom class behaviour
### Custom iterators
```
# Python 2 only
class Upper(object):
def __init__(self, iterable):
self._iter = iter(iterable)
def next(self): # Py2-style
return self._iter.next().upper()
def __iter__(self):
return self
itr = Upper('hello')
assert itr.next() == 'H' # Py2-style
assert list(itr) == list('ELLO')
# Python 2 and 3: option 1
from builtins import object
class Upper(object):
def __init__(self, iterable):
self._iter = iter(iterable)
def __next__(self): # Py3-style iterator interface
return next(self._iter).upper() # builtin next() function calls
def __iter__(self):
return self
itr = Upper('hello')
assert next(itr) == 'H' # compatible style
assert list(itr) == list('ELLO')
# Python 2 and 3: option 2
from future.utils import implements_iterator
@implements_iterator
class Upper(object):
def __init__(self, iterable):
self._iter = iter(iterable)
def __next__(self): # Py3-style iterator interface
return next(self._iter).upper() # builtin next() function calls
def __iter__(self):
return self
itr = Upper('hello')
assert next(itr) == 'H'
assert list(itr) == list('ELLO')
```
### Custom ``__str__`` methods
```
# Python 2 only:
class MyClass(object):
def __unicode__(self):
return 'Unicode string: \u5b54\u5b50'
def __str__(self):
return unicode(self).encode('utf-8')
a = MyClass()
print(a) # prints encoded string
# Python 2 and 3:
from future.utils import python_2_unicode_compatible
@python_2_unicode_compatible
class MyClass(object):
def __str__(self):
return u'Unicode string: \u5b54\u5b50'
a = MyClass()
print(a) # prints string encoded as utf-8 on Py2
```
### Custom ``__nonzero__`` vs ``__bool__`` method:
```
# Python 2 only:
class AllOrNothing(object):
def __init__(self, l):
self.l = l
def __nonzero__(self):
return all(self.l)
container = AllOrNothing([0, 100, 200])
assert not bool(container)
# Python 2 and 3:
from builtins import object
class AllOrNothing(object):
def __init__(self, l):
self.l = l
def __bool__(self):
return all(self.l)
container = AllOrNothing([0, 100, 200])
assert not bool(container)
```
## Lists versus iterators
### xrange
```
# Python 2 only:
for i in xrange(10**8):
...
# Python 2 and 3: forward-compatible
from builtins import range
for i in range(10**8):
...
# Python 2 and 3: backward-compatible
from past.builtins import xrange
for i in xrange(10**8):
...
```
### range
```
# Python 2 only
mylist = range(5)
assert mylist == [0, 1, 2, 3, 4]
# Python 2 and 3: forward-compatible: option 1
mylist = list(range(5)) # copies memory on Py2
assert mylist == [0, 1, 2, 3, 4]
# Python 2 and 3: forward-compatible: option 2
from builtins import range
mylist = list(range(5))
assert mylist == [0, 1, 2, 3, 4]
# Python 2 and 3: option 3
from future.utils import lrange
mylist = lrange(5)
assert mylist == [0, 1, 2, 3, 4]
# Python 2 and 3: backward compatible
from past.builtins import range
mylist = range(5)
assert mylist == [0, 1, 2, 3, 4]
```
### map
```
# Python 2 only:
mynewlist = map(f, myoldlist)
assert mynewlist == [f(x) for x in myoldlist]
# Python 2 and 3: option 1
# Idiomatic Py3, but inefficient on Py2
mynewlist = list(map(f, myoldlist))
assert mynewlist == [f(x) for x in myoldlist]
# Python 2 and 3: option 2
from builtins import map
mynewlist = list(map(f, myoldlist))
assert mynewlist == [f(x) for x in myoldlist]
# Python 2 and 3: option 3
try:
import itertools.imap as map
except ImportError:
pass
mynewlist = list(map(f, myoldlist)) # inefficient on Py2
assert mynewlist == [f(x) for x in myoldlist]
# Python 2 and 3: option 4
from future.utils import lmap
mynewlist = lmap(f, myoldlist)
assert mynewlist == [f(x) for x in myoldlist]
# Python 2 and 3: option 5
from past.builtins import map
mynewlist = map(f, myoldlist)
assert mynewlist == [f(x) for x in myoldlist]
```
### imap
```
# Python 2 only:
from itertools import imap
myiter = imap(func, myoldlist)
assert isinstance(myiter, iter)
# Python 3 only:
myiter = map(func, myoldlist)
assert isinstance(myiter, iter)
# Python 2 and 3: option 1
from builtins import map
myiter = map(func, myoldlist)
assert isinstance(myiter, iter)
# Python 2 and 3: option 2
try:
import itertools.imap as map
except ImportError:
pass
myiter = map(func, myoldlist)
assert isinstance(myiter, iter)
```
### zip, izip
As above with ``zip`` and ``itertools.izip``.
### filter, ifilter
As above with ``filter`` and ``itertools.ifilter`` too.
## Other builtins
### File IO with open()
```
# Python 2 only
f = open('myfile.txt')
data = f.read() # as a byte string
text = data.decode('utf-8')
# Python 2 and 3: alternative 1
from io import open
f = open('myfile.txt', 'rb')
data = f.read() # as bytes
text = data.decode('utf-8') # unicode, not bytes
# Python 2 and 3: alternative 2
from io import open
f = open('myfile.txt', encoding='utf-8')
text = f.read() # unicode, not bytes
```
### reduce()
```
# Python 2 only:
assert reduce(lambda x, y: x+y, [1, 2, 3, 4, 5]) == 1+2+3+4+5
# Python 2 and 3:
from functools import reduce
assert reduce(lambda x, y: x+y, [1, 2, 3, 4, 5]) == 1+2+3+4+5
```
### raw_input()
```
# Python 2 only:
name = raw_input('What is your name? ')
assert isinstance(name, str) # native str
# Python 2 and 3:
from builtins import input
name = input('What is your name? ')
assert isinstance(name, str) # native str on Py2 and Py3
```
### input()
```
# Python 2 only:
input("Type something safe please: ")
# Python 2 and 3
from builtins import input
eval(input("Type something safe please: "))
```
Warning: using either of these is **unsafe** with untrusted input.
### file()
```
# Python 2 only:
f = file(pathname)
# Python 2 and 3:
f = open(pathname)
# But preferably, use this:
from io import open
f = open(pathname, 'rb') # if f.read() should return bytes
# or
f = open(pathname, 'rt') # if f.read() should return unicode text
```
### execfile()
```
# Python 2 only:
execfile('myfile.py')
# Python 2 and 3: alternative 1
from past.builtins import execfile
execfile('myfile.py')
# Python 2 and 3: alternative 2
exec(compile(open('myfile.py').read()))
# This can sometimes cause this:
# SyntaxError: function ... uses import * and bare exec ...
# See https://github.com/PythonCharmers/python-future/issues/37
```
### unichr()
```
# Python 2 only:
assert unichr(8364) == '€'
# Python 3 only:
assert chr(8364) == '€'
# Python 2 and 3:
from builtins import chr
assert chr(8364) == '€'
```
### intern()
```
# Python 2 only:
intern('mystring')
# Python 3 only:
from sys import intern
intern('mystring')
# Python 2 and 3: alternative 1
from past.builtins import intern
intern('mystring')
# Python 2 and 3: alternative 2
from six.moves import intern
intern('mystring')
# Python 2 and 3: alternative 3
from future.standard_library import install_aliases
install_aliases()
from sys import intern
intern('mystring')
# Python 2 and 3: alternative 2
try:
from sys import intern
except ImportError:
pass
intern('mystring')
```
### apply()
```
args = ('a', 'b')
kwargs = {'kwarg1': True}
# Python 2 only:
apply(f, args, kwargs)
# Python 2 and 3: alternative 1
f(*args, **kwargs)
# Python 2 and 3: alternative 2
from past.builtins import apply
apply(f, args, kwargs)
```
### chr()
```
# Python 2 only:
assert chr(64) == b'@'
assert chr(200) == b'\xc8'
# Python 3 only: option 1
assert chr(64).encode('latin-1') == b'@'
assert chr(0xc8).encode('latin-1') == b'\xc8'
# Python 2 and 3: option 1
from builtins import chr
assert chr(64).encode('latin-1') == b'@'
assert chr(0xc8).encode('latin-1') == b'\xc8'
# Python 3 only: option 2
assert bytes([64]) == b'@'
assert bytes([0xc8]) == b'\xc8'
# Python 2 and 3: option 2
from builtins import bytes
assert bytes([64]) == b'@'
assert bytes([0xc8]) == b'\xc8'
```
### cmp()
```
# Python 2 only:
assert cmp('a', 'b') < 0 and cmp('b', 'a') > 0 and cmp('c', 'c') == 0
# Python 2 and 3: alternative 1
from past.builtins import cmp
assert cmp('a', 'b') < 0 and cmp('b', 'a') > 0 and cmp('c', 'c') == 0
# Python 2 and 3: alternative 2
cmp = lambda(x, y): (x > y) - (x < y)
assert cmp('a', 'b') < 0 and cmp('b', 'a') > 0 and cmp('c', 'c') == 0
```
### reload()
```
# Python 2 only:
reload(mymodule)
# Python 2 and 3
from imp import reload
reload(mymodule)
```
## Standard library
### dbm modules
```
# Python 2 only
import anydbm
import whichdb
import dbm
import dumbdbm
import gdbm
# Python 2 and 3: alternative 1
from future import standard_library
standard_library.install_aliases()
import dbm
import dbm.ndbm
import dbm.dumb
import dbm.gnu
# Python 2 and 3: alternative 2
from future.moves import dbm
from future.moves.dbm import dumb
from future.moves.dbm import ndbm
from future.moves.dbm import gnu
# Python 2 and 3: alternative 3
from six.moves import dbm_gnu
# (others not supported)
```
### commands / subprocess modules
```
# Python 2 only
from commands import getoutput, getstatusoutput
# Python 2 and 3
from future import standard_library
standard_library.install_aliases()
from subprocess import getoutput, getstatusoutput
```
### subprocess.check_output()
```
# Python 2.7 and above
from subprocess import check_output
# Python 2.6 and above: alternative 1
from future.moves.subprocess import check_output
# Python 2.6 and above: alternative 2
from future import standard_library
standard_library.install_aliases()
from subprocess import check_output
```
### collections: Counter, OrderedDict, ChainMap
```
# Python 2.7 and above
from collections import Counter, OrderedDict, ChainMap
# Python 2.6 and above: alternative 1
from future.backports import Counter, OrderedDict, ChainMap
# Python 2.6 and above: alternative 2
from future import standard_library
standard_library.install_aliases()
from collections import Counter, OrderedDict, ChainMap
```
### StringIO module
```
# Python 2 only
from StringIO import StringIO
from cStringIO import StringIO
# Python 2 and 3
from io import BytesIO
# and refactor StringIO() calls to BytesIO() if passing byte-strings
```
### http module
```
# Python 2 only:
import httplib
import Cookie
import cookielib
import BaseHTTPServer
import SimpleHTTPServer
import CGIHttpServer
# Python 2 and 3 (after ``pip install future``):
import http.client
import http.cookies
import http.cookiejar
import http.server
```
### xmlrpc module
```
# Python 2 only:
import DocXMLRPCServer
import SimpleXMLRPCServer
# Python 2 and 3 (after ``pip install future``):
import xmlrpc.server
# Python 2 only:
import xmlrpclib
# Python 2 and 3 (after ``pip install future``):
import xmlrpc.client
```
### html escaping and entities
```
# Python 2 and 3:
from cgi import escape
# Safer (Python 2 and 3, after ``pip install future``):
from html import escape
# Python 2 only:
from htmlentitydefs import codepoint2name, entitydefs, name2codepoint
# Python 2 and 3 (after ``pip install future``):
from html.entities import codepoint2name, entitydefs, name2codepoint
```
### html parsing
```
# Python 2 only:
from HTMLParser import HTMLParser
# Python 2 and 3 (after ``pip install future``)
from html.parser import HTMLParser
# Python 2 and 3 (alternative 2):
from future.moves.html.parser import HTMLParser
```
### urllib module
``urllib`` is the hardest module to use from Python 2/3 compatible code. You may like to use Requests (http://python-requests.org) instead.
```
# Python 2 only:
from urlparse import urlparse
from urllib import urlencode
from urllib2 import urlopen, Request, HTTPError
# Python 3 only:
from urllib.parse import urlparse, urlencode
from urllib.request import urlopen, Request
from urllib.error import HTTPError
# Python 2 and 3: easiest option
from future.standard_library import install_aliases
install_aliases()
from urllib.parse import urlparse, urlencode
from urllib.request import urlopen, Request
from urllib.error import HTTPError
# Python 2 and 3: alternative 2
from future.standard_library import hooks
with hooks():
from urllib.parse import urlparse, urlencode
from urllib.request import urlopen, Request
from urllib.error import HTTPError
# Python 2 and 3: alternative 3
from future.moves.urllib.parse import urlparse, urlencode
from future.moves.urllib.request import urlopen, Request
from future.moves.urllib.error import HTTPError
# or
from six.moves.urllib.parse import urlparse, urlencode
from six.moves.urllib.request import urlopen
from six.moves.urllib.error import HTTPError
# Python 2 and 3: alternative 4
try:
from urllib.parse import urlparse, urlencode
from urllib.request import urlopen, Request
from urllib.error import HTTPError
except ImportError:
from urlparse import urlparse
from urllib import urlencode
from urllib2 import urlopen, Request, HTTPError
```
### Tkinter
```
# Python 2 only:
import Tkinter
import Dialog
import FileDialog
import ScrolledText
import SimpleDialog
import Tix
import Tkconstants
import Tkdnd
import tkColorChooser
import tkCommonDialog
import tkFileDialog
import tkFont
import tkMessageBox
import tkSimpleDialog
# Python 2 and 3 (after ``pip install future``):
import tkinter
import tkinter.dialog
import tkinter.filedialog
import tkinter.scolledtext
import tkinter.simpledialog
import tkinter.tix
import tkinter.constants
import tkinter.dnd
import tkinter.colorchooser
import tkinter.commondialog
import tkinter.filedialog
import tkinter.font
import tkinter.messagebox
import tkinter.simpledialog
import tkinter.ttk
```
### socketserver
```
# Python 2 only:
import SocketServer
# Python 2 and 3 (after ``pip install future``):
import socketserver
```
### copy_reg, copyreg
```
# Python 2 only:
import copy_reg
# Python 2 and 3 (after ``pip install future``):
import copyreg
```
### configparser
```
# Python 2 only:
from ConfigParser import ConfigParser
# Python 2 and 3 (after ``pip install future``):
from configparser import ConfigParser
```
### queue
```
# Python 2 only:
from Queue import Queue, heapq, deque
# Python 2 and 3 (after ``pip install future``):
from queue import Queue, heapq, deque
```
### repr, reprlib
```
# Python 2 only:
from repr import aRepr, repr
# Python 2 and 3 (after ``pip install future``):
from reprlib import aRepr, repr
```
### UserDict, UserList, UserString
```
# Python 2 only:
from UserDict import UserDict
from UserList import UserList
from UserString import UserString
# Python 3 only:
from collections import UserDict, UserList, UserString
# Python 2 and 3: alternative 1
from future.moves.collections import UserDict, UserList, UserString
# Python 2 and 3: alternative 2
from six.moves import UserDict, UserList, UserString
# Python 2 and 3: alternative 3
from future.standard_library import install_aliases
install_aliases()
from collections import UserDict, UserList, UserString
```
### itertools: filterfalse, zip_longest
```
# Python 2 only:
from itertools import ifilterfalse, izip_longest
# Python 3 only:
from itertools import filterfalse, zip_longest
# Python 2 and 3: alternative 1
from future.moves.itertools import filterfalse, zip_longest
# Python 2 and 3: alternative 2
from six.moves import filterfalse, zip_longest
# Python 2 and 3: alternative 3
from future.standard_library import install_aliases
install_aliases()
from itertools import filterfalse, zip_longest
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
from tqdm import tqdm
```
### Classes
#### Model class
```
class Model:
"""This class is used to store information about the model
Function Parameters stores voltages and conductances: Ek, gk, Ena, gna, Eleak, gleak
Function Cable stores cable parameters: a, rL, L, Cm
Function Initial_values stores initial values: n, m, h, V
Functions for calculating alpha and beta are named by the following rule:
alpha_n -> an, beta_n -> bn
"""
def Parameters(self, Ek, gk, Ena, gna, Eleak, gleak):
"""This function is used to store voltages and conductances
Parameters: Ek, gk, Ena, gna, Eleak, gleak
stores voltages in V, condctances in S/m2
"""
self.Ek = Ek
self.gk = gk
self.Ena = Ena
self.gna = gna
self.Eleak = Eleak
self.gleak = gleak
def Cable(self, a, rL, L, Cm):
"""This function is used to store cable parameters
a = axon radius, in meters
rL = intracellular resistivity, in Om m
L = compartment length, in meters
Cm = membrane capacitance, in F
"""
self.a = a
self.rL = rL
self.L = L
self.Cm = Cm
self.A = 2 * np.pi * a * L
self.g = a / (2 * rL * L * L)
def Myelin(self, ranvier, interval, length, toggle):
"""This function defines how many compartments the model has and which ones are myelinated
ranvier = length of Ranvier node
interval = interval between beggining of nodes
length = length of axon
all parameters are in meters
function returns variable Mask, an M length array
"""
self.M = int(length / self.L)
R = int(ranvier / self.L)
I = int(interval / self.L)
self.Mask = np.zeros(self.M)
if toggle == True:
for i in range(self.M):
if i % I >= R: self.Mask[i] = 1;
def Initial_values(self, n0, m0, h0, V0):
"""This function is used to store the initial values
Values: n0, m0, h0, V0
"""
self.n = n0
self.m = m0
self.h = h0
self.V = V0
# Alpha and Beta
def an(V):
return np.divide(10000*(V + 0.055), 1 - np.exp(-100*(V + 0.055)));
def bn(V):
return 125 * np.exp(-12.5*(V + 0.065));
def am(V):
return np.divide(100000*(V + 0.040), 1 - np.exp(-100*(V + 0.040)));
def bm(V):
return 4000 * np.exp(-55.6*(V + 0.065));
def ah(V):
return 70 * np.exp(-50*(V + 0.065));
def bh(V):
return np.divide(1000, 1 + np.exp(-100*(V +0.035)));
```
#### Results
```
class Results:
"""This class is used to store the results after running the HH function
Inputs: n, m, h, V, Ik, Ina
"""
def __init__(self, n, m, h, V):
self.n = n
self.m = m
self.h = h
self.V = V
```
### Functions
#### Crank-Nicholson
```
def C_N(dt, M, Cm, g, A, gm, I, Ie, V):
"""This function calculates dV using the Crank-Nicholson method
dt = timestep, in seconds
M = number of compartments
Cm = membrane capacitance, in F
g = resistive coupling, in S
A = compartment surface area
gm = sum of gi
I = sum of gi * Ei
Ie = current input
V = potential
Function uses the method described in Chapter 6.6B from Dayan and Abbott, 2005
returns one variable, V_new
"""
z = 0.5 # Crank-Nicholson
### Helper variables ###
b = np.zeros(M)
c = np.zeros(M)
d = np.zeros(M)
f = np.zeros(M)
b[1:M] = g * z * dt / Cm[1:M]
d[0:M-1] = g * z * dt / Cm[0:M-1]
c = -gm*z*dt/Cm - b - d
f = (I + Ie/A)/Cm * z * dt + c * V
for i in range(M-1):
f[i+1] += b[i+1] * V[i]
f[i] += d[i] * V[i+1]
f = f*2 # getting rid of z
### Forward prop ###
c1 = np.zeros(M)
f1 = np.zeros(M)
c1[0] = c[0]
f1[0] = f[0]
for i in range(M-1):
c1[i+1] = c[i+1] + b[i+1] * d[i] / (1-c1[i])
f1[i+1] = f[i+1] + b[i+1] * f1[i] / (1-c1[i])
### Backprop ###
dV = np.zeros(M)
dV[M-1] = f1[M-1] / (1-c1[M-1])
for i in range(M-2, -1, -1):
dV[i] = (d[i] * dV[i+1] + f1[i]) / (1-c1[i])
return V + dV
```
#### Hodgkin-Huxley function
```
def HH(tmax, dt, model, Ie):
"""Calculates the dynamics of the Hodgkin-Huxley model
Input:
tmax = maximum time, in s
dt = time step, in s
model = model parameters(g and E), cable parameters(a, rL, L, Cm) and initial values(n, m, h, V), class Model
model also contains the functions needed to calculate alpha and beta
Ie = current input, in A
Output:
"""
# Creating the arrays
N = int(tmax / dt)
M = model.M
n = np.zeros((M, N+1))
m = np.zeros((M, N+1))
h = np.zeros((M, N+1))
V = np.zeros((M, N+1))
gk = np.zeros(M)
gna = np.zeros(M)
gl = np.zeros(M)
Cm = np.zeros(M)
gm = np.zeros(M)
I = np.zeros(M)
# Initial values
n[:, 0] = model.n
m[:, 0] = model.m
h[:, 0] = model.h
V[:, 0] = model.V
Cm[:] = model.Cm
for j in range(M):
if model.Mask[j] == 1: Cm[j] /= 50;
for i in range(N):
### Membrane conductance ###
# Potassium
n[:, i+1] = n[:, i] + (model.an(V[:, i]))*(1 - n[:, i])*dt - (model.bn(V[:, i]))*n[:, i]*dt;
gk[:] = model.gk * np.power(n[:, i], 4)
# Sodium
m[:, i+1] = m[:, i] + (model.am(V[:, i]))*(1 - m[:, i])*dt - (model.bm(V[:, i]))*m[:, i]*dt;
h[:, i+1] = h[:, i] + (model.ah(V[:, i]))*(1 - h[:, i])*dt - (model.bh(V[:, i]))*h[:, i]*dt;
gna[:] = model.gna * np.power(m[:, i], 3) * h[:, i]
# Leak
gl[:] = model.gleak
### Voltage ###
gm = gk + gna + gl
I = gk * model.Ek + gna * model.Ena + gl * model.Eleak
for j in range(M):
if model.Mask[j] == 1: gm[j] /= 5000; I[j] /= 5000; # Myelin increases resistance by 5000
V[:, i+1] = C_N(dt=dt, M=M, Cm=Cm, g=model.g, A=model.A, gm=gm, I=I, Ie=Ie[:, i], V=V[:, i])
# Storing results
results = Results(n=n, m=m, h=h, V=V)
return results
# Time variables
tmax = 0.01 # s
dt = 0.00001 # s
N = int(tmax / dt)+1
t = np.linspace(0, tmax, N)
```
#### Multiple compartments
```
model_myelin = Model
model_myelin.Parameters(self=model_myelin, Ek=-0.077, gk=360, Ena=0.050, gna=1200, Eleak=-0.054387, gleak=3)
model_myelin.Initial_values(self=model_myelin, n0=0.3177, m0=0.0529, h0=0.5961, V0=-0.065)
model_myelin.Cable(self=model_myelin, a=238e-06, rL=0.354, L=1e-06, Cm=0.01)
model_myelin.Myelin(self=model_myelin, ranvier=2e-06, interval=1e-03, length=5e-03, toggle=False)
```
### 3. Initiate an action potential on one end of the axon by injecting a current in the terminal compartment.
### 4. Determine the action potential propagation velocity as a function of the axon radius.
```
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(1, 1, 1)
Ie = np.zeros((model_myelin.M, N))
Ie[0,0:40] = 400e-12 #initiate current injection
sample = np.linspace(1,2,2,dtype=int)
apt = np.zeros(len(sample)) #action potential time
apd = np.zeros(len(sample)) #action potential distance
a = np.zeros(len(sample)) #axon radius
for i in tqdm(sample):
a[i-1] = i * 5e-06
model_myelin.Cable(self=model_myelin, a=a[i-1], rL=0.354, L=1e-06, Cm=0.01)
results_m = HH(tmax=tmax, dt=dt, model=model_myelin, Ie=Ie)
ax.plot(t, results_m.V[0],label='%r'%a[i-1])
ax.legend(title='axon radius')
apt[i-1] = np.argmax(results_m.V[0])
apd[i-1] = results_m.V[0][int(apt[i-1])]
#print(apt)
#print(apd)
fig = plt.figure(figsize=(10, 5))
ax = fig.add_subplot(1, 1, 1)
ax.plot(a,apd/(apt*dt))
ax.set_ylabel('action potential propagation velocity []')
ax.set_xlabel('axon radius [µm]')
ax.grid()
plt.show()
```
| github_jupyter |
```
%matplotlib inline
```
# Simple 1D Kernel Density Estimation
This example uses the :class:`sklearn.neighbors.KernelDensity` class to
demonstrate the principles of Kernel Density Estimation in one dimension.
The first plot shows one of the problems with using histograms to visualize
the density of points in 1D. Intuitively, a histogram can be thought of as a
scheme in which a unit "block" is stacked above each point on a regular grid.
As the top two panels show, however, the choice of gridding for these blocks
can lead to wildly divergent ideas about the underlying shape of the density
distribution. If we instead center each block on the point it represents, we
get the estimate shown in the bottom left panel. This is a kernel density
estimation with a "top hat" kernel. This idea can be generalized to other
kernel shapes: the bottom-right panel of the first figure shows a Gaussian
kernel density estimate over the same distribution.
Scikit-learn implements efficient kernel density estimation using either
a Ball Tree or KD Tree structure, through the
:class:`sklearn.neighbors.KernelDensity` estimator. The available kernels
are shown in the second figure of this example.
The third figure compares kernel density estimates for a distribution of 100
samples in 1 dimension. Though this example uses 1D distributions, kernel
density estimation is easily and efficiently extensible to higher dimensions
as well.
```
# Author: Jake Vanderplas <jakevdp@cs.washington.edu>
#
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import norm
from sklearn.neighbors import KernelDensity
#----------------------------------------------------------------------
# Plot the progression of histograms to kernels
np.random.seed(1)
N = 20
X = np.concatenate((np.random.normal(0, 1, int(0.3 * N)),
np.random.normal(5, 1, int(0.7 * N))))[:, np.newaxis]
X_plot = np.linspace(-5, 10, 1000)[:, np.newaxis]
bins = np.linspace(-5, 10, 10)
fig, ax = plt.subplots(2, 2, sharex=True, sharey=True)
fig.subplots_adjust(hspace=0.05, wspace=0.05)
# histogram 1
ax[0, 0].hist(X[:, 0], bins=bins, fc='#AAAAFF', normed=True)
ax[0, 0].text(-3.5, 0.31, "Histogram")
# histogram 2
ax[0, 1].hist(X[:, 0], bins=bins + 0.75, fc='#AAAAFF', normed=True)
ax[0, 1].text(-3.5, 0.31, "Histogram, bins shifted")
# tophat KDE
kde = KernelDensity(kernel='tophat', bandwidth=0.75).fit(X)
log_dens = kde.score_samples(X_plot)
ax[1, 0].fill(X_plot[:, 0], np.exp(log_dens), fc='#AAAAFF')
ax[1, 0].text(-3.5, 0.31, "Tophat Kernel Density")
# Gaussian KDE
kde = KernelDensity(kernel='gaussian', bandwidth=0.75).fit(X)
log_dens = kde.score_samples(X_plot)
ax[1, 1].fill(X_plot[:, 0], np.exp(log_dens), fc='#AAAAFF')
ax[1, 1].text(-3.5, 0.31, "Gaussian Kernel Density")
for axi in ax.ravel():
axi.plot(X[:, 0], np.zeros(X.shape[0]) - 0.01, '+k')
axi.set_xlim(-4, 9)
axi.set_ylim(-0.02, 0.34)
for axi in ax[:, 0]:
axi.set_ylabel('Normalized Density')
for axi in ax[1, :]:
axi.set_xlabel('x')
#----------------------------------------------------------------------
# Plot all available kernels
X_plot = np.linspace(-6, 6, 1000)[:, None]
X_src = np.zeros((1, 1))
fig, ax = plt.subplots(2, 3, sharex=True, sharey=True)
fig.subplots_adjust(left=0.05, right=0.95, hspace=0.05, wspace=0.05)
def format_func(x, loc):
if x == 0:
return '0'
elif x == 1:
return 'h'
elif x == -1:
return '-h'
else:
return '%ih' % x
for i, kernel in enumerate(['gaussian', 'tophat', 'epanechnikov',
'exponential', 'linear', 'cosine']):
axi = ax.ravel()[i]
log_dens = KernelDensity(kernel=kernel).fit(X_src).score_samples(X_plot)
axi.fill(X_plot[:, 0], np.exp(log_dens), '-k', fc='#AAAAFF')
axi.text(-2.6, 0.95, kernel)
axi.xaxis.set_major_formatter(plt.FuncFormatter(format_func))
axi.xaxis.set_major_locator(plt.MultipleLocator(1))
axi.yaxis.set_major_locator(plt.NullLocator())
axi.set_ylim(0, 1.05)
axi.set_xlim(-2.9, 2.9)
ax[0, 1].set_title('Available Kernels')
#----------------------------------------------------------------------
# Plot a 1D density example
N = 100
np.random.seed(1)
X = np.concatenate((np.random.normal(0, 1, int(0.3 * N)),
np.random.normal(5, 1, int(0.7 * N))))[:, np.newaxis]
X_plot = np.linspace(-5, 10, 1000)[:, np.newaxis]
true_dens = (0.3 * norm(0, 1).pdf(X_plot[:, 0])
+ 0.7 * norm(5, 1).pdf(X_plot[:, 0]))
fig, ax = plt.subplots()
ax.fill(X_plot[:, 0], true_dens, fc='black', alpha=0.2,
label='input distribution')
for kernel in ['gaussian', 'tophat', 'epanechnikov']:
kde = KernelDensity(kernel=kernel, bandwidth=0.5).fit(X)
log_dens = kde.score_samples(X_plot)
ax.plot(X_plot[:, 0], np.exp(log_dens), '-',
label="kernel = '{0}'".format(kernel))
ax.text(6, 0.38, "N={0} points".format(N))
ax.legend(loc='upper left')
ax.plot(X[:, 0], -0.005 - 0.01 * np.random.random(X.shape[0]), '+k')
ax.set_xlim(-4, 9)
ax.set_ylim(-0.02, 0.4)
plt.show()
```
| github_jupyter |
# Train a CNN Model for MNIST
This script here is to train a CNN model with 2 convolutional layers each with a pooling layer and a 2 fully-connected layers. The variables that would be needed for inference later have been added to tensorflow collections in this script.
- The MNIST dataset should be placed under a folder named 'MNIST_data' in the same directory as this script.
- The outputs of this script are tensorflow checkpoint models in a folder called 'models' in the same directory.
```
import tensorflow as tf
import numpy as np
import os
#import MNIST dataset
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets('MNIST_data', one_hot=True)
sess = tf.InteractiveSession() #initialize session
#input placeholders
with tf.name_scope('x'):
x = tf.placeholder(tf.float32, shape=[None, 784], name='x')
y_ = tf.placeholder(tf.float32, shape=[None, 10])
#function definitions
def weight_variable(shape, name):
initial = tf.truncated_normal(shape, stddev=0.1, name=name)
return tf.Variable(initial)
def bias_variable(shape, name):
initial = tf.constant(0.1, shape=shape, name=name)
return tf.Variable(initial)
def conv2d(x, W, name):
return tf.nn.conv2d(x, W, strides=[1, 1, 1, 1], padding='SAME', name=name)
def max_pool_2x2(x,name):
return tf.nn.max_pool(x, ksize=[1, 2, 2, 1],
strides=[1, 2, 2, 1], padding='SAME', name=name)
W_conv1 = weight_variable([5, 5, 1, 32], name='W_C1')
b_conv1 = bias_variable([32], name='B_C1')
x_image = tf.reshape(x, [-1,28,28,1]) #vectorize the image
h_conv1 = tf.nn.relu(conv2d(x_image, W_conv1 , name='conv_1') + b_conv1)
h_pool1 = max_pool_2x2(h_conv1, name='pool_1')
W_conv2 = weight_variable([5, 5, 32, 64], name='W_C2')
b_conv2 = bias_variable([64], name='B_C2')
h_conv2 = tf.nn.relu(conv2d(h_pool1, W_conv2, name='conv_2') + b_conv2)
h_pool2 = max_pool_2x2(h_conv2, name='pool_2')
W_fc1 = weight_variable([7 * 7 * 64, 1024], name='W_FC1')
b_fc1 = bias_variable([1024], name='B_FC1')
feature_vector = tf.reshape(h_pool2, [-1, 7*7*64])
h_fc1 = tf.nn.relu(tf.matmul(feature_vector, W_fc1) + b_fc1, name='FC_1')
W_fc2 = weight_variable([1024, 10], name='W_FC2')
b_fc2 = bias_variable([10], name='B_FC2')
with tf.name_scope('logits'):
logits = tf.add(tf.matmul(h_fc1, W_fc2), b_fc2, name='logits')
y = tf.nn.softmax(logits, name='softmax_prediction')
cross_entropy = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(labels=y_, logits=logits))
train_step = tf.train.AdamOptimizer(1e-4).minimize(cross_entropy)
correct_prediction = tf.equal(tf.argmax(logits,1), tf.argmax(y_,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32), name='accuracy')
# we only need these two to make inference using the trained model
tf.add_to_collection("logits", logits)
tf.add_to_collection("x", x)
sess.run(tf.global_variables_initializer())
saver = tf.train.Saver(tf.global_variables())
for i in range(500):
batch = mnist.train.next_batch(100)
if i%100 == 0:
train_acc = accuracy.eval(feed_dict={
x:batch[0], y_:batch[1]
})
print("Step %d, training accuracy %g"%(i, train_acc))
train_step.run(feed_dict={x:batch[0], y_:batch[1]})
current_dir = os.getcwd() #get the current working directory
saver.save(sess, current_dir + '/model/mnist.ckpt') #save the model in the specified directory
print("Training is finished.")
```
| github_jupyter |
# Laboratorio 2: Armado de un esquema de aprendizaje automático
En el laboratorio final se espera que puedan poner en práctica los conocimientos adquiridos en el curso, trabajando con un conjunto de datos de clasificación.
El objetivo es que se introduzcan en el desarrollo de un esquema para hacer tareas de aprendizaje automático: selección de un modelo, ajuste de hiperparámetros y evaluación.
El conjunto de datos a utilizar está en `./data/loan_data.csv`. Si abren el archivo verán que al principio (las líneas que empiezan con `#`) describen el conjunto de datos y sus atributos (incluyendo el atributo de etiqueta o clase).
Se espera que hagan uso de las herramientas vistas en el curso. Se espera que hagan uso especialmente de las herramientas brindadas por `scikit-learn`.
```
import numpy as np
import pandas as pd
# TODO: Agregar las librerías que hagan falta
from sklearn.model_selection import train_test_split
```
## Carga de datos y división en entrenamiento y evaluación
La celda siguiente se encarga de la carga de datos (haciendo uso de pandas). Estos serán los que se trabajarán en el resto del laboratorio.
```
dataset = pd.read_csv("./data/loan_data.csv", comment="#")
# División entre instancias y etiquetas
X, y = dataset.iloc[:, 1:], dataset.TARGET
# división entre entrenamiento y evaluación
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
```
Documentación:
- https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html
## Ejercicio 1: Descripción de los Datos y la Tarea
Responder las siguientes preguntas:
1. ¿De qué se trata el conjunto de datos?
2. ¿Cuál es la variable objetivo que hay que predecir? ¿Qué significado tiene?
3. ¿Qué información (atributos) hay disponible para hacer la predicción?
4. ¿Qué atributos imagina ud. que son los más determinantes para la predicción?
**No hace falta escribir código para responder estas preguntas.**
## Ejercicio 2: Predicción con Modelos Lineales
En este ejercicio se entrenarán modelos lineales de clasificación para predecir la variable objetivo.
Para ello, deberán utilizar la clase SGDClassifier de scikit-learn.
Documentación:
- https://scikit-learn.org/stable/modules/sgd.html
- https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.SGDClassifier.html
### Ejercicio 2.1: SGDClassifier con hiperparámetros por defecto
Entrenar y evaluar el clasificador SGDClassifier usando los valores por omisión de scikit-learn para todos los parámetros. Únicamente **fijar la semilla aleatoria** para hacer repetible el experimento.
Evaluar sobre el conjunto de **entrenamiento** y sobre el conjunto de **evaluación**, reportando:
- Accuracy
- Precision
- Recall
- F1
- matriz de confusión
### Ejercicio 2.2: Ajuste de Hiperparámetros
Seleccionar valores para los hiperparámetros principales del SGDClassifier. Como mínimo, probar diferentes funciones de loss, tasas de entrenamiento y tasas de regularización.
Para ello, usar grid-search y 5-fold cross-validation sobre el conjunto de entrenamiento para explorar muchas combinaciones posibles de valores.
Reportar accuracy promedio y varianza para todas las configuraciones.
Para la mejor configuración encontrada, evaluar sobre el conjunto de **entrenamiento** y sobre el conjunto de **evaluación**, reportando:
- Accuracy
- Precision
- Recall
- F1
- matriz de confusión
Documentación:
- https://scikit-learn.org/stable/modules/grid_search.html
- https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html
## Ejercicio 3: Árboles de Decisión
En este ejercicio se entrenarán árboles de decisión para predecir la variable objetivo.
Para ello, deberán utilizar la clase DecisionTreeClassifier de scikit-learn.
Documentación:
- https://scikit-learn.org/stable/modules/tree.html
- https://scikit-learn.org/stable/modules/tree.html#tips-on-practical-use
- https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html
- https://scikit-learn.org/stable/auto_examples/tree/plot_unveil_tree_structure.html
### Ejercicio 3.1: DecisionTreeClassifier con hiperparámetros por defecto
Entrenar y evaluar el clasificador DecisionTreeClassifier usando los valores por omisión de scikit-learn para todos los parámetros. Únicamente **fijar la semilla aleatoria** para hacer repetible el experimento.
Evaluar sobre el conjunto de **entrenamiento** y sobre el conjunto de **evaluación**, reportando:
- Accuracy
- Precision
- Recall
- F1
- matriz de confusión
### Ejercicio 3.2: Ajuste de Hiperparámetros
Seleccionar valores para los hiperparámetros principales del DecisionTreeClassifier. Como mínimo, probar diferentes criterios de partición (criterion), profundidad máxima del árbol (max_depth), y cantidad mínima de samples por hoja (min_samples_leaf).
Para ello, usar grid-search y 5-fold cross-validation sobre el conjunto de entrenamiento para explorar muchas combinaciones posibles de valores.
Reportar accuracy promedio y varianza para todas las configuraciones.
Para la mejor configuración encontrada, evaluar sobre el conjunto de **entrenamiento** y sobre el conjunto de **evaluación**, reportando:
- Accuracy
- Precision
- Recall
- F1
- matriz de confusión
Documentación:
- https://scikit-learn.org/stable/modules/grid_search.html
- https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html
### Ejercicio 3.3: Inspección del Modelo
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
```
### Indepedent Component Analysis
```
class icaDemo:
def __init__(self,N):
self.N = N
def remmean(self,sig):
newVec = np.zeros(sig.shape)
meanVal = np.mean(sig,axis=1)
newVec = sig-np.transpose(np.tile(meanVal,(self.N,1)))
return newVec, meanVal
def demoSig(self):
v = np.linspace(0,self.N-1,self.N)
sinArr = np.sin(v/2); s1 = np.std(sinArr)
funArr = ((np.remainder(v,23)-11)/9)**5; s2 = np.std(funArr)
sawtooth = (np.remainder(v,27)-13)/9; s3 = np.std(sawtooth)
uni = np.random.uniform(0,1,self.N)
impul = ((uni<.5)*2-1)*np.log(uni); s4 = np.std(impul)
sig = np.vstack((sinArr/s1,funArr/s2,sawtooth/s3,impul/s4))
sig, mean = self.remmean(sig)
Aorig = np.random.uniform(0,1,(sig.shape[0],sig.shape[0]))
mixedsig = np.matmul(Aorig,sig)
return sig, mixedsig
def pcamat(self,mixedsig,Eig1,Eig_1):
oldDimension = mixedsig.shape[0]
D,E = np.linalg.eigh(np.cov(mixedsig))
return E,D
def whitenv(self,mixed_sig,E,D):
whiteMat = np.matmul(np.linalg.inv(np.diag(D)**.5),np.transpose(E))
dewhiteMat = np.matmul(E,np.diag(D)**.5)
whitesig = np.matmul(whiteMat,mixed_sig)
return whitesig, whiteMat, dewhiteMat
def fpica(self,whitesig,whiteMat,dewhiteMat):
vecSize, numSamples = whitesig.shape
B = np.zeros((vecSize,vecSize))
iteration = 1; numFailures = 0; epsilon = 1e-4
while iteration <= vecSize:
w = np.random.normal(0,1,(vecSize,1))
w -= np.matmul(np.matmul(B,np.transpose(B)),w)
w /= np.linalg.norm(w)
wOld = np.zeros(w.shape); wOld2 = np.zeros(w.shape)
i = 1; gabba = 1; maxIter = 1000;
while i<=maxIter+gabba:
w -= np.matmul(np.matmul(B,np.transpose(B)),w)
w /= np.linalg.norm(w)
if (np.linalg.norm(w-wOld)<epsilon) or (np.linalg.norm(w+wOld)<epsilon):
numFailures = 0;
B[:,iteration-1]=np.transpose(w)
if iteration==1:
A = np.matmul(dewhiteMat,w)
W = np.matmul(np.transpose(w),whiteMat)
else:
A = np.concatenate((A,np.matmul(dewhiteMat,w)),axis=1)
W = np.concatenate((W,np.matmul(np.transpose(w),whiteMat)))
break
wOld2 = wOld; wOld = w;
w=np.matmul(whitesig,np.matmul(np.transpose(whitesig),w)**3)/numSamples-3*w
w /= np.linalg.norm(w)
i += 1
iteration += 1
return A, W
def fastica(self):
sigTrue, mixedsig = self.demoSig()
mixedsig,mixedmean = self.remmean(mixedsig)
Dim, NumOfSample = mixedsig.shape
E,D = self.pcamat(mixedsig,1,4)
whitesig,whiteMat,dewhiteMat = self.whitenv(mixedsig,E,D)
A,W = self.fpica(whitesig,whiteMat,dewhiteMat)
icasig1 = np.matmul(W,mixedsig)
icasig2 = np.tile(np.matmul(W,np.transpose(mixedmean)),(self.N,1)).transpose()
icasig = icasig1+icasig2
return icasig, A, W
demo = icaDemo(500)
sigTrue, mixedsig = demo.demoSig()
```
#### Latent Indepedent Signals
```
for i in range(4):
plt.subplot(4,1,i+1); plt.plot(sigTrue[i,:500])
```
#### Mixed Observation Signals
```
for i in range(4):
plt.subplot(4,1,i+1); plt.plot(mixedsig[i,:500])
```
#### ICA estimate Signals
```
icasig, A, W = demo.fastica()
for i in range(4):
plt.subplot(4,1,i+1); plt.plot(icasig[i,:500])
```
| github_jupyter |
```
import opendatasets as od
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import random
import seaborn as sns
from sklearn.metrics import mean_squared_error
pd.options.display.float_format = '{:.5f}'.format
dataset_url = "https://www.kaggle.com/c/new-york-city-taxi-fare-prediction/overview"
od.download(dataset_url)
data_dir = "D:/Internship/Taxi Fare nyc/new-york-city-taxi-fare-prediction"
df = pd.read_csv("new-york-city-taxi-fare-prediction/train.csv")
df
col_name = list(df.columns)
col_name
n = len(df)
s = round(n * 0.1)
skip = sorted(random.sample(range(1, n+1), n-s))
df = pd.read_csv("new-york-city-taxi-fare-prediction/train.csv", skiprows=skip)
df
df = df.drop(["key"], axis=1)
df
lat_values = []
for i in range(-90,91):
lat_values.append(i)
lon_values = []
for j in range(-180, 181):
lon_values.append(j)
```
the range of latitude is -90 to +90
the range of longitude is -180 to +180
Hence, any value greater than mentioned value must be dropped
```
def remove_outliers(df):
return df[(df['fare_amount'] >= 1.) &
(df['fare_amount'] <= 500.) &
(df['pickup_longitude'] >= -75) &
(df['pickup_longitude'] <= -72) &
(df['dropoff_longitude'] >= -75) &
(df['dropoff_longitude'] <= -72) &
(df['pickup_latitude'] >= 40) &
(df['pickup_latitude'] <= 42) &
(df['dropoff_latitude'] >=40) &
(df['dropoff_latitude'] <= 42) &
(df['passenger_count'] >= 1) &
(df['passenger_count'] <= 6)]
df = remove_outliers(df)
df
df.describe()
df.info()
sns.boxplot(data = df, y = "fare_amount")
df["pickup_datetime"] = pd.to_datetime(df["pickup_datetime"])
df
df_index = df.set_index("pickup_datetime")
df_index
df_index["fare_amount"].plot(figsize = (25, 5))
```
Using Haversine formula
* a = sin²(Δφ/2) + cos φ1 ⋅ cos φ2 ⋅ sin²(Δλ/2)
* c = 2 ⋅ atan2( √a, √(1−a) )
* d = R ⋅ c
math module expects a single number, using numpy for providing series as input
```
# Haversine Formula
from math import radians, cos, sin, asin, sqrt
def haversine_dist(lon1, lat1, lon2, lat2):
lat1 = np.radians(lat1)
lat2 = np.radians(lat2)
lon1 = np.radians(lon1)
lon2 = np.radians(lon2)
# Haversine formula
dlat = lat2 - lat1
dlon = lon2 - lon1
a = np.sin(dlat / 2.0)** 2 + np.cos(lat1) * np.cos(lat2) * np.sin(dlon / 2.0) ** 2
c = 2 * np.arcsin(np.sqrt(a))
r = 6371 # radius of earth in km
distance = c * r
return distance
def add_distance(df):
df["trip_distance"] = haversine_dist(df["pickup_longitude"], df["pickup_latitude"],
df["dropoff_longitude"], df["dropoff_latitude"])
add_distance(df)
df
df["trip_distance"].max()
max_df = df[df["trip_distance"] == 178.81596366391366]
max_df
test_df = pd.read_csv("new-york-city-taxi-fare-prediction/test.csv", parse_dates=["pickup_datetime"])
test_df
test_df = test_df.drop(["key"], axis = 1)
test_df
add_distance(test_df)
test_df
test_df.info()
df.pickup_datetime.min(), df.pickup_datetime.max()
sns.set(rc={"figure.figsize":(10, 7)})
sns.histplot(data = df, x = "trip_distance", kde=True)
sns.set(rc={"figure.figsize":(8, 6)})
sns.histplot(data = df, x = "fare_amount", kde=True)
sns.set(rc={"figure.figsize":(15, 7)})
sns.violinplot(data = df, x = df["trip_distance"])
sns.set(rc={"figure.figsize":(15, 7)})
sns.boxplot(data = df, x = df["trip_distance"])
sns.set(rc={"figure.figsize":(15, 7)})
sns.boxplot(data = df, x = df["fare_amount"])
sns.set(rc={"figure.figsize":(15, 7)})
sns.violinplot(data = df, x = df["fare_amount"])
from sklearn.model_selection import train_test_split
train_df, val_df = train_test_split(df, test_size=0.2, random_state=10)
len(train_df), len(val_df)
train_df.isna().sum(), val_df.isna().sum()
train_df, val_df = train_df.dropna(), val_df.dropna()
train_df
sns.set(rc={"figure.figsize":(8, 6)})
sns.scatterplot(y = train_df["fare_amount"], x = train_df["passenger_count"])
test_df
x = train_df.iloc[:, 6:]
x
x_col = list(x.columns)
y_col = "fare_amount"
x_train = train_df[x_col]
x_train
y_train = train_df[y_col]
y_train
```
### Validation Data
```
x_val = val_df[x_col]
x_val
y_val = val_df[y_col]
y_val
```
Test Data
```
x_test = test_df[x_col]
x_test
x_train[x_col] = x_train[x_col].astype(float)
x_train.info()
```
Modelling
Linear Regression
```
from sklearn.linear_model import LinearRegression
model_lr = LinearRegression()
model_lr.fit(x_train, y_train)
y_lr_train = model_lr.predict(x_train)
y_lr_train
y_lr_val = model_lr.predict(x_val)
y_lr_val
train_rmse = mean_squared_error(y_val, y_lr_val, squared=False) # returns rmse value when set to false
train_rmse
from sklearn.metrics import mean_absolute_error
val_mae = mean_absolute_error(y_val, y_lr_val)
val_mae
from sklearn.tree import DecisionTreeRegressor
dt_regressor = DecisionTreeRegressor(random_state=0)
dt_regressor.fit(x_train, y_train)
y_pred_dt = dt_regressor.predict(x_val)
y_pred_dt
dt_rmse = mean_squared_error(y_val, y_pred_dt, squared=False) # returns rmse value when set to false
dt_rmse
train_df.describe()
```
Feature Engineering
```
def date_time_extract(df, col):
df[col + "_year"] = df[col].dt.year
df[col + "_month"] = df[col].dt.month
df[col + "_day"] = df[col].dt.day
df[col + "_weekday"] = df[col].dt.weekday
df[col + "_time"] = df[col].dt.time
return date_time_extract
date_time_extract(train_df, "pickup_datetime")
date_time_extract(val_df, "pickup_datetime")
date_time_extract(test_df, "pickup_datetime")
train_df
train_df.info()
sns.countplot(data = train_df, x = "pickup_datetime_month")
sns.scatterplot(data = train_df, x = "pickup_datetime_month", y = "fare_amount")
sns.scatterplot(data = train_df, x = "trip_distance", y = "fare_amount", hue = "pickup_datetime_month")
# some senario where distance traveled is larger with respect to price
train_df.query("trip_distance >= 100 and fare_amount <= 10")
train_df.query("trip_distance <= 10 and fare_amount >= 100")
# removing data where trip price is very high compared to distance traveled
train_df = train_df.drop(train_df[(train_df.trip_distance <= 10) & (train_df.fare_amount >= 100)].index)
train_df
train_df.columns[2:6]
train_df = train_df.drop(columns=['pickup_longitude', 'pickup_latitude',
'dropoff_longitude', 'dropoff_latitude'])
train_df
train_df.columns
train_df[['fare_amount', 'passenger_count', 'trip_distance']].describe()
fig, axes = plt.subplots(2, 1, figsize = (15, 9), sharey = True)
fig.suptitle("Box and Violin Plot for Fare Amount")
sns.boxplot(ax = axes[0], data = train_df, x = train_df["fare_amount"])
axes[0].set_title("Box Plot")
#
sns.violinplot(ax = axes[1], data = train_df, x = train_df["fare_amount"])
axes[1].set_title("Violin Plot")
fig, axes = plt.subplots(2, 1, figsize = (15, 9), sharey = True)
fig.suptitle("Box and Violin Plot for Trip Distance")
sns.boxplot(ax = axes[0], data = train_df, x = train_df["trip_distance"])
axes[0].set_title("Box Plot")
#
sns.violinplot(ax = axes[1], data = train_df, x = train_df["trip_distance"])
axes[1].set_title("Violin Plot")
```
The dataset is positively Skewed, have to drop outliers
```
fig, axes = plt.subplots(2, 1, figsize = (15, 9), sharey = True)
fig.suptitle("Box and Violin Plot for Trip Distance")
sns.boxplot(ax = axes[0], data = test_df, x = test_df["trip_distance"])
axes[0].set_title("Box Plot")
#
sns.violinplot(ax = axes[1], data = test_df, x = test_df["trip_distance"])
axes[1].set_title("Violin Plot")
train_df = train_df.drop(train_df[(train_df.trip_distance <= 10) & (train_df.fare_amount >= 100)].index)
train_df
train_df
x_train = train_df.iloc[:, 2:4]
x_train
y_train = train_df.iloc[:, 0:1]
y_train
from sklearn.preprocessing import normalize
x_arr = np.array(x_train[["passenger_count", "trip_distance"]])
train_df_norm = normalize(x_arr, axis = 0)
train_df_norm
from sklearn.preprocessing import MinMaxScaler
trans = MinMaxScaler()
train_df_norm = trans.fit_transform(x_train)
train_df_norm
train_df_norm = pd.DataFrame(train_df_norm, columns=x_train.columns)
train_df_norm
fig, axes = plt.subplots(2, 1, figsize = (15, 9), sharey = True)
fig.suptitle("Box and Violin Plot for Trip Distance")
sns.boxplot(ax = axes[0], data = train_df_norm, x = train_df_norm["trip_distance"])
axes[0].set_title("Box Plot")
#
sns.violinplot(ax = axes[1], data = train_df_norm, x = train_df_norm["trip_distance"])
axes[1].set_title("Violin Plot")
print(f"The standard deviation for trip distance is {np.std(train_df['trip_distance'])}")
print(f"The mean for trip distance is {np.mean(train_df['trip_distance'])}")
train_df["pickup_datetime_month"].value_counts()
```
## ML Using Standardizing Data
```
from sklearn.ensemble import RandomForestRegressor
model2 = RandomForestRegressor(max_depth=10, n_jobs=-1, random_state=42, n_estimators=50)
model2.fit(train_df_norm, y_train)
x_val = trans.transform(x_val)
x_val
y_pred_rfr = model2.predict(x_val)
y_pred_rfr
train_rmse = mean_squared_error(y_val, y_pred_rfr, squared=False) # returns rmse value when set to false
train_rmse
```
### Using Standard Scalar
```
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
x_train = sc.fit_transform(x_train)
x_train
x_val = sc.transform(x_val)
x_val
x_test_std = sc.transform(x_test)
x_test_std
from sklearn.ensemble import RandomForestRegressor
model2 = RandomForestRegressor(max_depth=10, n_jobs=-1, random_state=42, n_estimators=50)
model2.fit(x_train, y_train)
y_pred_rfr = model2.predict(x_val)
y_pred_rfr
train_rmse = mean_squared_error(y_val, y_pred_rfr, squared=False) # returns rmse value when set to false
train_rmse
def predict_and_submit(model, fname):
test_preds = model2.predict(x_test_std)
sub_df = pd.read_csv(data_dir+'/sample_submission.csv')
sub_df['fare_amount'] = test_preds
sub_df.to_csv(fname, index=None)
return sub_df
predict_and_submit(model2, "random_forest_regressor.csv")
from xgboost import XGBRegressor
model3 = XGBRegressor(random_state=42, n_jobs=-1, objective='reg:squarederror')
model3.fit(x_train, y_train)
y_pred_xgboost = model3.predict(x_val)
y_pred_xgboost
val_rmse = mean_squared_error(y_val, y_pred_xgboost, squared=False) # returns rmse value when set to false
val_rmse
predict_and_submit(model3, "xgboost_regressor.csv")
import matplotlib.pyplot as plt
def test_params(ModelClass, **params):
"""Trains a model with the given parameters and returns training & validation RMSE"""
model = ModelClass(**params).fit(x_train, y_train)
train_rmse = mean_squared_error(model.predict(x_train), y_train, squared=False)
val_rmse = mean_squared_error(model.predict(x_val), y_val, squared=False)
return train_rmse, val_rmse
def test_param_and_plot(ModelClass, param_name, param_values, **other_params):
"""Trains multiple models by varying the value of param_name according to param_values"""
train_errors, val_errors = [], []
for value in param_values:
params = dict(other_params)
params[param_name] = value
train_rmse, val_rmse = test_params(ModelClass, **params)
train_errors.append(train_rmse)
val_errors.append(val_rmse)
plt.figure(figsize=(10,6))
plt.title('Overfitting curve: ' + param_name)
plt.plot(param_values, train_errors, 'b-o')
plt.plot(param_values, val_errors, 'r-o')
plt.xlabel(param_name)
plt.ylabel('RMSE')
plt.legend(['Training', 'Validation'])
best_params = {
'random_state': 42,
'n_jobs': -1,
'objective': 'reg:squarederror'
}
%%time
test_param_and_plot(XGBRegressor, 'n_estimators', [100, 250, 500], **best_params)
best_params['n_estimators'] = 100
test_param_and_plot(XGBRegressor, 'max_depth', [3, 4, 5], **best_params)
%%time
test_param_and_plot(XGBRegressor, 'learning_rate', [0.05, 0.1, 0.25], **best_params)
```
| github_jupyter |
# Introduction to Numpy
NumPy is the fundamental package for scientific computing
in Python. It is a Python library that provides a multidimensional array
object. In this course, we will be using NumPy for linear algebra.
If you are interested in learning more about NumPy, you can find the user
guide and reference at https://docs.scipy.org/doc/numpy/index.html
Let's first import the NumPy package
```
import numpy as np # we commonly use the np abbreviation when referring to numpy
```
## Creating Numpy Arrays
New arrays can be made in several ways. We can take an existing list and convert it to a numpy array:
```
a = np.array([1,2,3])
```
There are also functions for creating arrays with ones and zeros
```
np.zeros((2,2))
np.ones((3,2))
```
## Accessing Numpy Arrays
You can use the common square bracket syntax for accessing elements
of a numpy array
```
A = np.arange(9).reshape(3,3)
print(A)
print(A[0].shape) # Access the first row of A
print(A[0, 1]) # Access the second item of the first row
print(A[:, 1]) # Access the second column
```
## Operations on Numpy Arrays
You can use the operations '*', '**', '\', '+' and '-' on numpy arrays and they operate elementwise.
```
a = np.array([[1,2],
[2,3]])
b = np.array([[4,5],
[6,7]])
print(a + b)
print(a - b)
print(a * b)
print(a / b)
print(a**2)
```
There are also some commonly used function
For example, you can sum up all elements of an array
```
print(a)
print(np.sum(a))
```
Or sum along the first dimension
```
np.sum(a, axis=0)
```
There are many other functions in numpy, and some of them **will be useful**
for your programming assignments. As an exercise, check out the documentation
for these routines at https://docs.scipy.org/doc/numpy/reference/routines.html
and see if you can find the documentation for `np.sum` and `np.reshape`.
## Linear Algebra
In this course, we use the numpy arrays for linear algebra.
We usually use 1D arrays to represent vectors and 2D arrays to represent
matrices
```
A = np.array([[2,4],
[6,8]])
```
You can take transposes of matrices with `A.T`
```
print('A\n', A)
print('A.T\n', A.T)
```
Note that taking the transpose of a 1D array has **NO** effect.
```
a = np.ones(3)
print(a)
print(a.shape)
print(a.T)
print(a.T.shape)
```
But it does work if you have a 2D array of shape (3,1)
```
a = np.ones((3,1))
print(a)
print(a.shape)
print(a.T)
print(a.T.shape)
```
### Dot product
We can compute the dot product between two vectors with np.dot
```
x = np.array([1,2,3])
y = np.array([4,5,6])
np.dot(x, y)
```
We can compute the matrix-matrix product, matrix-vector product too. In Python 3, this is conveniently expressed with the @ syntax
```
A = np.eye(3) # You can create an identity matrix with np.eye
B = np.random.randn(3,3)
x = np.array([1,2,3])
# Matrix-Matrix product
A @ B
# Matrix-vector product
A @ x
```
Sometimes, we might want to compute certain properties of the matrices. For example, we might be interested in a matrix's determinant, eigenvalues/eigenvectors. Numpy ships with the `numpy.linalg` package to do
these things on 2D arrays (matrices).
```
from numpy import linalg
# This computes the determinant
linalg.det(A)
# This computes the eigenvalues and eigenvectors
eigenvalues, eigenvectors = linalg.eig(A)
print("The eigenvalues are\n", eigenvalues)
print("The eigenvectors are\n", eigenvectors)
```
## Miscellaneous
### Time your code
One tip that is really useful is to use the magic commannd `%time` to time the execution time of your function.
```
%time np.abs(A)
```
| github_jupyter |
```
%load_ext autoreload
from __future__ import print_function, division
%autoreload
import copy, math, os, pickle, time, pandas as pd, numpy as np, scipy.stats as ss
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import average_precision_score, roc_auc_score, accuracy_score, f1_score
import torch, torch.utils.data as utils, torch.nn as nn, torch.nn.functional as F, torch.optim as optim
from torch.autograd import Variable
from torch.nn.parameter import Parameter
DATA_FILEPATH = '/scratch/mmd/mimic_data/final/grouping_5/all_hourly_data.h5'
RAW_DATA_FILEPATH = '/scratch/mmd/mimic_data/final/nogrouping_5/all_hourly_data.h5'
GAP_TIME = 6 # In hours
WINDOW_SIZE = 24 # In hours
SEED = 1
ID_COLS = ['subject_id', 'hadm_id', 'icustay_id']
GPU = '2'
os.environ['CUDA_VISIBLE_DEVICES'] = GPU
np.random.seed(SEED)
torch.manual_seed(SEED)
class DictDist():
def __init__(self, dict_of_rvs): self.dict_of_rvs = dict_of_rvs
def rvs(self, n):
a = {k: v.rvs(n) for k, v in self.dict_of_rvs.items()}
out = []
for i in range(n): out.append({k: vs[i] for k, vs in a.items()})
return out
class Choice():
def __init__(self, options): self.options = options
def rvs(self, n): return [self.options[i] for i in ss.randint(0, len(self.options)).rvs(n)]
%%time
data_full_lvl2 = pd.read_hdf(DATA_FILEPATH, 'vitals_labs')
data_full_raw = pd.read_hdf(RAW_DATA_FILEPATH, 'vitals_labs')
statics = pd.read_hdf(DATA_FILEPATH, 'patients')
data_full_lvl2.head()
data_full_raw.head()
statics.head()
def simple_imputer(df):
idx = pd.IndexSlice
df = df.copy()
if len(df.columns.names) > 2: df.columns = df.columns.droplevel(('label', 'LEVEL1', 'LEVEL2'))
df_out = df.loc[:, idx[:, ['mean', 'count']]]
icustay_means = df_out.loc[:, idx[:, 'mean']].groupby(ID_COLS).mean()
df_out.loc[:,idx[:,'mean']] = df_out.loc[:,idx[:,'mean']].groupby(ID_COLS).fillna(
method='ffill'
).groupby(ID_COLS).fillna(icustay_means).fillna(0)
df_out.loc[:, idx[:, 'count']] = (df.loc[:, idx[:, 'count']] > 0).astype(float)
df_out.rename(columns={'count': 'mask'}, level='Aggregation Function', inplace=True)
is_absent = (1 - df_out.loc[:, idx[:, 'mask']])
hours_of_absence = is_absent.cumsum()
time_since_measured = hours_of_absence - hours_of_absence[is_absent==0].fillna(method='ffill')
time_since_measured.rename(columns={'mask': 'time_since_measured'}, level='Aggregation Function', inplace=True)
df_out = pd.concat((df_out, time_since_measured), axis=1)
df_out.loc[:, idx[:, 'time_since_measured']] = df_out.loc[:, idx[:, 'time_since_measured']].fillna(100)
df_out.sort_index(axis=1, inplace=True)
return df_out
Ys = statics[statics.max_hours > WINDOW_SIZE + GAP_TIME][['mort_hosp', 'mort_icu', 'los_icu']]
Ys['los_3'] = Ys['los_icu'] > 3
Ys['los_7'] = Ys['los_icu'] > 7
Ys.drop(columns=['los_icu'], inplace=True)
Ys.astype(float)
lvl2, raw = [df[
(df.index.get_level_values('icustay_id').isin(set(Ys.index.get_level_values('icustay_id')))) &
(df.index.get_level_values('hours_in') < WINDOW_SIZE)
] for df in (data_full_lvl2, data_full_raw)]
raw.columns = raw.columns.droplevel(level=['label', 'LEVEL1', 'LEVEL2'])
train_frac, dev_frac, test_frac = 0.7, 0.1, 0.2
lvl2_subj_idx, raw_subj_idx, Ys_subj_idx = [df.index.get_level_values('subject_id') for df in (lvl2, raw, Ys)]
lvl2_subjects = set(lvl2_subj_idx)
assert lvl2_subjects == set(Ys_subj_idx), "Subject ID pools differ!"
assert lvl2_subjects == set(raw_subj_idx), "Subject ID pools differ!"
np.random.seed(SEED)
subjects, N = np.random.permutation(list(lvl2_subjects)), len(lvl2_subjects)
N_train, N_dev, N_test = int(train_frac * N), int(dev_frac * N), int(test_frac * N)
train_subj = subjects[:N_train]
dev_subj = subjects[N_train:N_train + N_dev]
test_subj = subjects[N_train+N_dev:]
[(lvl2_train, lvl2_dev, lvl2_test), (raw_train, raw_dev, raw_test), (Ys_train, Ys_dev, Ys_test)] = [
[df[df.index.get_level_values('subject_id').isin(s)] for s in (train_subj, dev_subj, test_subj)] \
for df in (lvl2, raw, Ys)
]
idx = pd.IndexSlice
lvl2_means, lvl2_stds = lvl2_train.loc[:, idx[:,'mean']].mean(axis=0), lvl2_train.loc[:, idx[:,'mean']].std(axis=0)
raw_means, raw_stds = raw_train.loc[:, idx[:,'mean']].mean(axis=0), raw_train.loc[:, idx[:,'mean']].std(axis=0)
lvl2_train.loc[:, idx[:,'mean']] = (lvl2_train.loc[:, idx[:,'mean']] - lvl2_means)/lvl2_stds
lvl2_dev.loc[:, idx[:,'mean']] = (lvl2_dev.loc[:, idx[:,'mean']] - lvl2_means)/lvl2_stds
lvl2_test.loc[:, idx[:,'mean']] = (lvl2_test.loc[:, idx[:,'mean']] - lvl2_means)/lvl2_stds
raw_train.loc[:, idx[:,'mean']] = (raw_train.loc[:, idx[:,'mean']] - raw_means)/raw_stds
raw_dev.loc[:, idx[:,'mean']] = (raw_dev.loc[:, idx[:,'mean']] - raw_means)/raw_stds
raw_test.loc[:, idx[:,'mean']] = (raw_test.loc[:, idx[:,'mean']] - raw_means)/raw_stds
raw_train, raw_dev, raw_test, lvl2_train, lvl2_dev, lvl2_test = [
simple_imputer(df) for df in (raw_train, raw_dev, raw_test, lvl2_train, lvl2_dev, lvl2_test)
]
raw_flat_train, raw_flat_dev, raw_flat_test, lvl2_flat_train, lvl2_flat_dev, lvl2_flat_test = [
df.pivot_table(index=['subject_id', 'hadm_id', 'icustay_id'], columns=['hours_in']) for df in (
raw_train, raw_dev, raw_test, lvl2_train, lvl2_dev, lvl2_test
)
]
for df in lvl2_train, lvl2_dev, lvl2_test, raw_train, raw_dev, raw_test: assert not df.isnull().any().any()
```
### Task Prediction
#### Hyperparams
```
N = 15
LR_dist = DictDist({
'C': Choice(np.geomspace(1e-3, 1e3, 10000)),
'penalty': Choice(['l1', 'l2']),
'solver': Choice(['liblinear', 'lbfgs']),
'max_iter': Choice([100, 500])
})
np.random.seed(SEED)
LR_hyperparams_list = LR_dist.rvs(N)
for i in range(N):
if LR_hyperparams_list[i]['solver'] == 'lbfgs': LR_hyperparams_list[i]['penalty'] = 'l2'
RF_dist = DictDist({
'n_estimators': ss.randint(50, 500),
'max_depth': ss.randint(2, 10),
'min_samples_split': ss.randint(2, 75),
'min_samples_leaf': ss.randint(1, 50),
})
np.random.seed(SEED)
RF_hyperparams_list = RF_dist.rvs(N)
GRU_D_dist = DictDist({
'cell_size': ss.randint(50, 75),
'hidden_size': ss.randint(65, 95),
'learning_rate': ss.uniform(2e-3, 1e-1),
'num_epochs': ss.randint(15, 150),
'patience': ss.randint(3, 7),
'batch_size': ss.randint(35, 65),
'early_stop_frac': ss.uniform(0.05, 0.1),
'seed': ss.randint(1, 10000),
})
np.random.seed(SEED)
GRU_D_hyperparams_list = GRU_D_dist.rvs(N)
def run_basic(model, hyperparams_list, X_flat_train, X_flat_dev, X_flat_test, target):
best_s, best_hyperparams = -np.Inf, None
for i, hyperparams in enumerate(hyperparams_list):
print("On sample %d / %d (hyperparams = %s)" % (i+1, len(hyperparams_list), repr((hyperparams))))
M = model(**hyperparams)
M.fit(X_flat_train, Ys_train[target])
s = roc_auc_score(Ys_dev[target], M.predict_proba(X_flat_dev)[:, 1])
if s > best_s:
best_s, best_hyperparams = s, hyperparams
print("New Best Score: %.2f @ hyperparams = %s" % (100*best_s, repr((best_hyperparams))))
return run_only_final(model, best_hyperparams, X_flat_train, X_flat_dev, X_flat_test, target)
def run_only_final(model, best_hyperparams, X_flat_train, X_flat_dev, X_flat_test, target):
best_M = model(**best_hyperparams)
best_M.fit(pd.concat((X_flat_train, X_flat_dev)), pd.concat((Ys_train, Ys_dev))[target])
y_true = Ys_test[target]
y_score = best_M.predict_proba(X_flat_test)[:, 1]
y_pred = best_M.predict(X_flat_test)
auc = roc_auc_score(y_true, y_score)
auprc = average_precision_score(y_true, y_score)
acc = accuracy_score(y_true, y_pred)
F1 = f1_score(y_true, y_pred)
return best_M, best_hyperparams, auc, auprc, acc, F1
```
### Sklearn
```
RESULTS_PATH = '/scratch/mmd/extraction_baselines-sklearn.pkl'
with open(RESULTS_PATH, mode='rb') as f: results = pickle.load(f)
RERUN = True
for model_name, model, hyperparams_list in [
('RF', RandomForestClassifier, RF_hyperparams_list), ('LR', LogisticRegression, LR_hyperparams_list)
]:
if model_name not in results: results[model_name] = {}
for t in ['mort_icu', 'los_3']:
if t not in results[model_name]: results[model_name][t] = {}
for n, X_flat_train, X_flat_dev, X_flat_test in (
('lvl2', lvl2_flat_train, lvl2_flat_dev, lvl2_flat_test),
('raw', raw_flat_train, raw_flat_dev, raw_flat_test)
):
if n in results[model_name][t]:
print("Finished model %s on target %s with representation %s" % (model_name, t, n))
if RERUN:
h = results[model_name][t][n][1]
results[model_name][t][n] = run_only_final(model, h, X_flat_train, X_flat_dev, X_flat_test, t)
print("Final results for model %s on target %s with representation %s" % (model_name, t, n))
print(results[model_name][t][n][2:])
with open(RESULTS_PATH, mode='wb') as f: pickle.dump(results, f)
continue
print("Running model %s on target %s with representation %s" % (model_name, t, n))
results[model_name][t][n] = run_basic(
model, hyperparams_list, X_flat_train, X_flat_dev, X_flat_test, t
)
print("Final results for model %s on target %s with representation %s" % (model_name, t, n))
print(results[model_name][t][n][2:])
with open(RESULTS_PATH, mode='wb') as f: pickle.dump(results, f)
np.random.seed(SEED+1)
LR_hyperparams_list_2 = LR_dist.rvs(45)
for i in range(45):
if LR_hyperparams_list_2[i]['solver'] == 'lbfgs': LR_hyperparams_list_2[i]['penalty'] = 'l2'
results_2 = {}
results_2_PATH = '/scratch/mmd/extraction_baselines-sklearn_LR_2_runs.pkl'
for model_name, model, hyperparams_list in [
# ('RF', RandomForestClassifier, RF_hyperparams_list),
('LR', LogisticRegression, LR_hyperparams_list_2)
]:
if model_name not in results_2: results_2[model_name] = {}
for t in ['mort_icu', 'los_3']:
if t not in results_2[model_name]: results_2[model_name][t] = {}
for n, X_flat_train, X_flat_dev, X_flat_test in (
('lvl2', lvl2_flat_train, lvl2_flat_dev, lvl2_flat_test),
# ('raw', raw_flat_train, raw_flat_dev, raw_flat_test)
):
if n in results_2[model_name][t]:
print("Finished model %s on target %s with representation %s" % (model_name, t, n))
if RERUN:
h = results_2[model_name][t][n][1]
results_2[model_name][t][n] = run_only_final(model, h, X_flat_train, X_flat_dev, X_flat_test, t)
print("Final results_2 for model %s on target %s with representation %s" % (model_name, t, n))
print(results_2[model_name][t][n][2:])
with open(results_2_PATH, mode='wb') as f: pickle.dump(results_2, f)
continue
print("Running model %s on target %s with representation %s" % (model_name, t, n))
results_2[model_name][t][n] = run_basic(
model, hyperparams_list, X_flat_train, X_flat_dev, X_flat_test, t
)
print("Final results_2 for model %s on target %s with representation %s" % (model_name, t, n))
print(results_2[model_name][t][n][2:])
with open(results_2_PATH, mode='wb') as f: pickle.dump(results_2, f)
for model_name, model, hyperparams_list in [
# ('RF', RandomForestClassifier, RF_hyperparams_list),
('LR', LogisticRegression, LR_hyperparams_list_2)
]:
if model_name not in results_2: results_2[model_name] = {}
for t in ['mort_icu', 'los_3']:
if t not in results_2[model_name]: results_2[model_name][t] = {}
for n, X_flat_train, X_flat_dev, X_flat_test in (
# ('lvl2', lvl2_flat_train, lvl2_flat_dev, lvl2_flat_test),
('raw', raw_flat_train, raw_flat_dev, raw_flat_test),
):
if n in results_2[model_name][t]:
print("Finished model %s on target %s with representation %s" % (model_name, t, n))
if RERUN:
h = results_2[model_name][t][n][1]
results_2[model_name][t][n] = run_only_final(model, h, X_flat_train, X_flat_dev, X_flat_test, t)
print("Final results_2 for model %s on target %s with representation %s" % (model_name, t, n))
print(results_2[model_name][t][n][2:])
with open(results_2_PATH, mode='wb') as f: pickle.dump(results_2, f)
continue
print("Running model %s on target %s with representation %s" % (model_name, t, n))
results_2[model_name][t][n] = run_basic(
model, hyperparams_list, X_flat_train, X_flat_dev, X_flat_test, t
)
print("Final results_2 for model %s on target %s with representation %s" % (model_name, t, n))
print(results_2[model_name][t][n][2:])
with open(results_2_PATH, mode='wb') as f: pickle.dump(results_2, f)
for model_name, model, hyperparams_list in [
('RF', RandomForestClassifier, RF_hyperparams_list), ('LR', LogisticRegression, LR_hyperparams_list)
]:
if model_name not in results: results[model_name] = {}
for t in ['mort_hosp', 'los_7']:
if t not in results[model_name]: results[model_name][t] = {}
for n, X_flat_train, X_flat_dev, X_flat_test in (
('lvl2', lvl2_flat_train, lvl2_flat_dev, lvl2_flat_test),
('raw', raw_flat_train, raw_flat_dev, raw_flat_test)
):
if n in results[model_name][t]:
print("Finished model %s on target %s with representation %s" % (model_name, t, n))
if RERUN:
h = results[model_name][t][n][1]
results[model_name][t][n] = run_only_final(model, h, X_flat_train, X_flat_dev, X_flat_test, t)
print("Final results for model %s on target %s with representation %s" % (model_name, t, n))
print(results[model_name][t][n][2:])
with open(RESULTS_PATH, mode='wb') as f: pickle.dump(results, f)
continue
print("Running model %s on target %s with representation %s" % (model_name, t, n))
results[model_name][t][n] = run_basic(
model, hyperparams_list, X_flat_train, X_flat_dev, X_flat_test, t
)
print("Final results for model %s on target %s with representation %s" % (model_name, t, n))
print(results[model_name][t][n][2:])
with open(RESULTS_PATH, mode='wb') as f: pickle.dump(results, f)
```
| github_jupyter |
# **3D-RCAN**
---
<font size = 4>3D-RCAN is a neural network capable of image restoration from corrupted bio-images, first released in 2020 by [Chen *et al.* in biorXiv](https://www.biorxiv.org/content/10.1101/2020.08.27.270439v1).
<font size = 4> **This particular notebook enables restoration of 3D dataset. If you are interested in restoring 2D dataset, you should use the CARE 2D notebook instead.**
---
<font size = 4>*Disclaimer*:
<font size = 4>This notebook is part of the Zero-Cost Deep-Learning to Enhance Microscopy project (https://github.com/HenriquesLab/DeepLearning_Collab/wiki). Jointly developed by the Jacquemet (link to https://cellmig.org/) and Henriques (https://henriqueslab.github.io/) laboratories.
<font size = 4>This notebook is largely based on the following paper:
<font size = 4>**Three-dimensional residual channel attention networks denoise and sharpen fluorescence microscopy image volumes**, by Chen *et al.* published in bioRxiv in 2020 (https://www.biorxiv.org/content/10.1101/2020.08.27.270439v1)
<font size = 4>And source code found in: https://github.com/AiviaCommunity/3D-RCAN
<font size = 4>We provide a dataset for the training of this notebook as a way to test its functionalities but the training and test data of the restoration experiments is also available from the authors of the original paper [here](https://www.dropbox.com/sh/hieldept1x476dw/AAC0pY3FrwdZBctvFF0Fx0L3a?dl=0).
<font size = 4>**Please also cite this original paper when using or developing this notebook.**
# **How to use this notebook?**
---
<font size = 4>Video describing how to use our notebooks are available on youtube:
- [**Video 1**](https://www.youtube.com/watch?v=GzD2gamVNHI&feature=youtu.be): Full run through of the workflow to obtain the notebooks and the provided test datasets as well as a common use of the notebook
- [**Video 2**](https://www.youtube.com/watch?v=PUuQfP5SsqM&feature=youtu.be): Detailed description of the different sections of the notebook
---
###**Structure of a notebook**
<font size = 4>The notebook contains two types of cell:
<font size = 4>**Text cells** provide information and can be modified by douple-clicking the cell. You are currently reading the text cell. You can create a new text by clicking `+ Text`.
<font size = 4>**Code cells** contain code and the code can be modfied by selecting the cell. To execute the cell, move your cursor on the `[ ]`-mark on the left side of the cell (play button appears). Click to execute the cell. After execution is done the animation of play button stops. You can create a new coding cell by clicking `+ Code`.
---
###**Table of contents, Code snippets** and **Files**
<font size = 4>On the top left side of the notebook you find three tabs which contain from top to bottom:
<font size = 4>*Table of contents* = contains structure of the notebook. Click the content to move quickly between sections.
<font size = 4>*Code snippets* = contain examples how to code certain tasks. You can ignore this when using this notebook.
<font size = 4>*Files* = contain all available files. After mounting your google drive (see section 1.) you will find your files and folders here.
<font size = 4>**Remember that all uploaded files are purged after changing the runtime.** All files saved in Google Drive will remain. You do not need to use the Mount Drive-button; your Google Drive is connected in section 1.2.
<font size = 4>**Note:** The "sample data" in "Files" contains default files. Do not upload anything in here!
---
###**Making changes to the notebook**
<font size = 4>**You can make a copy** of the notebook and save it to your Google Drive. To do this click file -> save a copy in drive.
<font size = 4>To **edit a cell**, double click on the text. This will show you either the source code (in code cells) or the source text (in text cells).
You can use the `#`-mark in code cells to comment out parts of the code. This allows you to keep the original code piece in the cell as a comment.
#**0. Before getting started**
---
<font size = 4> For CARE to train, **it needs to have access to a paired training dataset**. This means that the same image needs to be acquired in the two conditions (for instance, low signal-to-noise ratio and high signal-to-noise ratio) and provided with indication of correspondence.
<font size = 4> Therefore, the data structure is important. It is necessary that all the input data are in the same folder and that all the output data is in a separate folder. The provided training dataset is already split in two folders called "Training - Low SNR images" (Training_source) and "Training - high SNR images" (Training_target). Information on how to generate a training dataset is available in our Wiki page: https://github.com/HenriquesLab/ZeroCostDL4Mic/wiki
<font size = 4>**We strongly recommend that you generate extra paired images. These images can be used to assess the quality of your trained model (Quality control dataset)**. The quality control assessment can be done directly in this notebook.
<font size = 4> **Additionally, the corresponding input and output files need to have the same name**.
<font size = 4> Please note that you currently can **only use .tif files!**
<font size = 4> You can also provide a folder that contains the data that you wish to analyse with the trained network once all training has been performed.
<font size = 4>Here's a common data structure that can work:
* Experiment A
- **Training dataset**
- Low SNR images (Training_source)
- img_1.tif, img_2.tif, ...
- High SNR images (Training_target)
- img_1.tif, img_2.tif, ...
- **Quality control dataset**
- Low SNR images
- img_1.tif, img_2.tif
- High SNR images
- img_1.tif, img_2.tif
- **Data to be predicted**
- **Results**
---
<font size = 4>**Important note**
<font size = 4>- If you wish to **Train a network from scratch** using your own dataset (and we encourage everyone to do that), you will need to run **sections 1 - 4**, then use **section 5** to assess the quality of your model and **section 6** to run predictions using the model that you trained.
<font size = 4>- If you wish to **Evaluate your model** using a model previously generated and saved on your Google Drive, you will only need to run **sections 1 and 2** to set up the notebook, then use **section 5** to assess the quality of your model.
<font size = 4>- If you only wish to **run predictions** using a model previously generated and saved on your Google Drive, you will only need to run **sections 1 and 2** to set up the notebook, then use **section 6** to run the predictions on the desired model.
---
# **1. Initialise the Colab session**
---
## **1.1. Check for GPU access**
---
By default, the session should be using Python 3 and GPU acceleration, but it is possible to ensure that these are set properly by doing the following:
<font size = 4>Go to **Runtime -> Change the Runtime type**
<font size = 4>**Runtime type: Python 3** *(Python 3 is programming language in which this program is written)*
<font size = 4>**Accelator: GPU** *(Graphics processing unit)*
```
#@markdown ##Run this cell to check if you have GPU access
%tensorflow_version 1.x
import tensorflow as tf
if tf.test.gpu_device_name()=='':
print('You do not have GPU access.')
print('Did you change your runtime ?')
print('If the runtime setting is correct then Google did not allocate a GPU for your session')
print('Expect slow performance. To access GPU try reconnecting later')
else:
print('You have GPU access')
!nvidia-smi
```
## **1.2. Mount your Google Drive**
---
<font size = 4> To use this notebook on the data present in your Google Drive, you need to mount your Google Drive to this notebook.
<font size = 4> Play the cell below to mount your Google Drive and follow the link. In the new browser window, select your drive and select 'Allow', copy the code, paste into the cell and press enter. This will give Colab access to the data on the drive.
<font size = 4> Once this is done, your data are available in the **Files** tab on the top left of notebook.
```
#@markdown ##Play the cell to connect your Google Drive to Colab
#@markdown * Click on the URL.
#@markdown * Sign in your Google Account.
#@markdown * Copy the authorization code.
#@markdown * Enter the authorization code.
#@markdown * Click on "Files" site on the right. Refresh the site. Your Google Drive folder should now be available here as "drive".
# mount user's Google Drive to Google Colab.
from google.colab import drive
drive.mount('/content/gdrive')
```
# **2. Install 3D-RCAN and dependencies**
---
## **2.1. Install key dependencies**
---
<font size = 4>
```
Notebook_version = ['1.12']
#@markdown ##Install 3D-RCAN and dependencies
!git clone https://github.com/AiviaCommunity/3D-RCAN
import os
!pip install q keras==2.2.5
!pip install colorama; sys_platform=='win32'
!pip install jsonschema
!pip install numexpr
!pip install tqdm>=4.41.0
%tensorflow_version 1.x
#Here, we install libraries which are not already included in Colab.
!pip install tifffile # contains tools to operate tiff-files
!pip install wget
!pip install fpdf
!pip install memory_profiler
%load_ext memory_profiler
```
## **2.2. Restart your runtime**
---
<font size = 4>
**<font size = 4> Here you need to restart your runtime to load the newly installed dependencies**
<font size = 4> Click on "Runtime" ---> "Restart Runtime"
## **2.3. Load key dependencies**
---
<font size = 4>
```
Notebook_version = ['1.11.1']
#@markdown ##Load key dependencies
!pip install q keras==2.2.5
#Here, we import and enable Tensorflow 1 instead of Tensorflow 2.
%tensorflow_version 1.x
import tensorflow
import tensorflow as tf
print(tensorflow.__version__)
print("Tensorflow enabled.")
# ------- Variable specific to 3D-RCAN -------
# ------- Common variable to all ZeroCostDL4Mic notebooks -------
import numpy as np
from matplotlib import pyplot as plt
import urllib
import os, random
import shutil
import zipfile
from tifffile import imread, imsave
import time
import sys
import wget
from pathlib import Path
import pandas as pd
import csv
from glob import glob
from scipy import signal
from scipy import ndimage
from skimage import io
from sklearn.linear_model import LinearRegression
from skimage.util import img_as_uint
import matplotlib as mpl
from skimage.metrics import structural_similarity
from skimage.metrics import peak_signal_noise_ratio as psnr
from astropy.visualization import simple_norm
from skimage import img_as_float32
from skimage.util import img_as_ubyte
from tqdm import tqdm
from fpdf import FPDF, HTMLMixin
from datetime import datetime
from pip._internal.operations.freeze import freeze
import subprocess
# For sliders and dropdown menu and progress bar
from ipywidgets import interact
import ipywidgets as widgets
# Colors for the warning messages
class bcolors:
WARNING = '\033[31m'
W = '\033[0m' # white (normal)
R = '\033[31m' # red
#Disable some of the tensorflow warnings
import warnings
warnings.filterwarnings("ignore")
print("Libraries installed")
# Check if this is the latest version of the notebook
Latest_notebook_version = pd.read_csv("https://raw.githubusercontent.com/HenriquesLab/ZeroCostDL4Mic/master/Colab_notebooks/Latest_ZeroCostDL4Mic_Release.csv")
print('Notebook version: '+Notebook_version[0])
strlist = Notebook_version[0].split('.')
Notebook_version_main = strlist[0]+'.'+strlist[1]
if Notebook_version_main == Latest_notebook_version.columns:
print("This notebook is up-to-date.")
else:
print(bcolors.WARNING +"A new version of this notebook has been released. We recommend that you download it at https://github.com/HenriquesLab/ZeroCostDL4Mic/wiki")
def pdf_export(trained = False, augmentation = False, pretrained_model = False):
class MyFPDF(FPDF, HTMLMixin):
pass
pdf = MyFPDF()
pdf.add_page()
pdf.set_right_margin(-1)
pdf.set_font("Arial", size = 11, style='B')
Network = '3D-RCAN'
day = datetime.now()
datetime_str = str(day)[0:16]
Header = 'Training report for '+Network+' model ('+model_name+')\nDate and Time: '+datetime_str
pdf.multi_cell(180, 5, txt = Header, align = 'L')
# add another cell
if trained:
training_time = "Training time: "+str(hour)+ "hour(s) "+str(mins)+"min(s) "+str(round(sec))+"sec(s)"
pdf.cell(190, 5, txt = training_time, ln = 1, align='L')
pdf.ln(1)
Header_2 = 'Information for your materials and method:'
pdf.cell(190, 5, txt=Header_2, ln=1, align='L')
all_packages = ''
for requirement in freeze(local_only=True):
all_packages = all_packages+requirement+', '
#print(all_packages)
#Main Packages
main_packages = ''
version_numbers = []
for name in ['tensorflow','numpy','Keras']:
find_name=all_packages.find(name)
main_packages = main_packages+all_packages[find_name:all_packages.find(',',find_name)]+', '
#Version numbers only here:
version_numbers.append(all_packages[find_name+len(name)+2:all_packages.find(',',find_name)])
cuda_version = subprocess.run('nvcc --version',stdout=subprocess.PIPE, shell=True)
cuda_version = cuda_version.stdout.decode('utf-8')
cuda_version = cuda_version[cuda_version.find(', V')+3:-1]
gpu_name = subprocess.run('nvidia-smi',stdout=subprocess.PIPE, shell=True)
gpu_name = gpu_name.stdout.decode('utf-8')
gpu_name = gpu_name[gpu_name.find('Tesla'):gpu_name.find('Tesla')+10]
#print(cuda_version[cuda_version.find(', V')+3:-1])
#print(gpu_name)
shape = io.imread(Training_source+'/'+os.listdir(Training_source)[1]).shape
dataset_size = len(os.listdir(Training_source))
text = 'The '+Network+' model was trained from scratch for '+str(number_of_epochs)+' epochs (image dimensions: '+str(shape)+', using the '+Network+' ZeroCostDL4Mic notebook (v '+Notebook_version[0]+') (von Chamier & Laine et al., 2020). Key python packages used include tensorflow (v '+version_numbers[0]+'), Keras (v '+version_numbers[2]+'), numpy (v '+version_numbers[1]+'), cuda (v '+cuda_version+'). The training was accelerated using a '+gpu_name+'GPU.'
pdf.set_font('')
pdf.set_font_size(10.)
pdf.multi_cell(190, 5, txt = text, align='L')
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.ln(1)
pdf.cell(28, 5, txt='Augmentation: ', ln=0)
pdf.set_font('')
if augmentation:
aug_text = 'The dataset was augmented by'
if Rotation:
aug_text = aug_text+'\n- rotation'
if Flip:
aug_text = aug_text+'\n- flipping'
else:
aug_text = 'No augmentation was used for training.'
pdf.multi_cell(190, 5, txt=aug_text, align='L')
pdf.set_font('Arial', size = 11, style = 'B')
pdf.ln(1)
pdf.cell(180, 5, txt = 'Parameters', align='L', ln=1)
pdf.set_font('')
pdf.set_font_size(10.)
if Use_Default_Advanced_Parameters:
pdf.cell(200, 5, txt='Default Advanced Parameters were enabled')
pdf.cell(200, 5, txt='The following parameters were used for training:')
pdf.ln(1)
html = """
<table width=40% style="margin-left:0px;">
<tr>
<th width = 50% align="left">Parameter</th>
<th width = 50% align="left">Value</th>
</tr>
<tr>
<td width = 50%>number_of_epochs</td>
<td width = 50%>{0}</td>
</tr>
<tr>
<td width = 50%>number_of_steps</td>
<td width = 50%>{1}</td>
</tr>
<tr>
<td width = 50%>percentage_validation</td>
<td width = 50%>{2}</td>
</tr>
<tr>
<td width = 50%>num_residual_groups</td>
<td width = 50%>{3}</td>
</tr>
<tr>
<td width = 50%>num_residual_blocks</td>
<td width = 50%>{4}</td>
</tr>
<tr>
<td width = 50%>num_channels</td>
<td width = 50%>{5}</td>
</tr>
<tr>
<td width = 50%>channel_reduction</td>
<td width = 50%>{6}</td>
</tr>
</table>
""".format(number_of_epochs,number_of_steps, percentage_validation, num_residual_groups, num_residual_blocks, num_channels, channel_reduction)
pdf.write_html(html)
#pdf.multi_cell(190, 5, txt = text_2, align='L')
pdf.set_font("Arial", size = 11, style='B')
pdf.ln(1)
pdf.cell(190, 5, txt = 'Training Dataset', align='L', ln=1)
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(32, 5, txt= 'Training_source:', align = 'L', ln=0)
pdf.set_font('')
pdf.multi_cell(170, 5, txt = Training_source, align = 'L')
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(30, 5, txt= 'Training_target:', align = 'L', ln=0)
pdf.set_font('')
pdf.multi_cell(170, 5, txt = Training_target, align = 'L')
#pdf.cell(190, 5, txt=aug_text, align='L', ln=1)
pdf.ln(1)
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(22, 5, txt= 'Model Path:', align = 'L', ln=0)
pdf.set_font('')
pdf.multi_cell(170, 5, txt = model_path+'/'+model_name, align = 'L')
pdf.ln(1)
pdf.cell(60, 5, txt = 'Example Training pair', ln=1)
pdf.ln(1)
exp_size = io.imread('/content/TrainingDataExample_3D_RCAN.png').shape
pdf.image('/content/TrainingDataExample_3D_RCAN.png', x = 11, y = None, w = round(exp_size[1]/8), h = round(exp_size[0]/8))
pdf.ln(1)
ref_1 = 'References:\n - ZeroCostDL4Mic: von Chamier, Lucas & Laine, Romain, et al. "ZeroCostDL4Mic: an open platform to simplify access and use of Deep-Learning in Microscopy." BioRxiv (2020).'
pdf.multi_cell(190, 5, txt = ref_1, align='L')
ref_2 = '- 3D-RCAN: Chen et al. "Three-dimensional residual channel attention networks denoise and sharpen fluorescence microscopy image volumes." bioRxiv 2020 https://www.biorxiv.org/content/10.1101/2020.08.27.270439v1'
pdf.multi_cell(190, 5, txt = ref_2, align='L')
pdf.ln(3)
reminder = 'Important:\nRemember to perform the quality control step on all newly trained models\nPlease consider depositing your training dataset on Zenodo'
pdf.set_font('Arial', size = 11, style='B')
pdf.multi_cell(190, 5, txt=reminder, align='C')
if trained:
pdf.output(model_path+'/'+model_name+'/'+model_name+"_training_report.pdf")
else:
pdf.output('/content/'+model_name+"_training_report.pdf")
def qc_pdf_export():
class MyFPDF(FPDF, HTMLMixin):
pass
pdf = MyFPDF()
pdf.add_page()
pdf.set_right_margin(-1)
pdf.set_font("Arial", size = 11, style='B')
Network = '3D RCAN'
day = datetime.now()
datetime_str = str(day)[0:10]
Header = 'Quality Control report for '+Network+' model ('+QC_model_name+')\nDate: '+datetime_str
pdf.multi_cell(180, 5, txt = Header, align = 'L')
all_packages = ''
for requirement in freeze(local_only=True):
all_packages = all_packages+requirement+', '
pdf.set_font('')
pdf.set_font('Arial', size = 11, style = 'B')
pdf.ln(2)
pdf.cell(190, 5, txt = 'Development of Training Losses', ln=1, align='L')
exp_size = io.imread(full_QC_model_path+'/Quality Control/QC_example_data.png').shape
if os.path.exists(full_QC_model_path+'/Quality Control/lossCurvePlots.png'):
pdf.image(full_QC_model_path+'/Quality Control/lossCurvePlots.png', x = 11, y = None, w = round(exp_size[1]/10), h = round(exp_size[0]/13))
else:
pdf.set_font('')
pdf.set_font('Arial', size=10)
# pdf.ln(3)
pdf.multi_cell(190, 5, txt='You can see these curves in the notebook.')
pdf.ln(3)
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.ln(3)
pdf.cell(80, 5, txt = 'Example Quality Control Visualisation', ln=1)
pdf.ln(1)
exp_size = io.imread(full_QC_model_path+'/Quality Control/QC_example_data.png').shape
pdf.image(full_QC_model_path+'/Quality Control/QC_example_data.png', x = 16, y = None, w = round(exp_size[1]/10), h = round(exp_size[0]/10))
pdf.ln(1)
pdf.set_font('')
pdf.set_font('Arial', size = 11, style = 'B')
pdf.ln(1)
pdf.cell(180, 5, txt = 'Quality Control Metrics', align='L', ln=1)
pdf.set_font('')
pdf.set_font_size(10.)
pdf.ln(1)
html = """
<body>
<font size="7" face="Courier New" >
<table width=97% style="margin-left:0px;">"""
with open(full_QC_model_path+'/Quality Control/QC_metrics_'+QC_model_name+'.csv', 'r') as csvfile:
metrics = csv.reader(csvfile)
header = next(metrics)
image = header[0]
slice_n = header[1]
mSSIM_PvsGT = header[2]
mSSIM_SvsGT = header[3]
NRMSE_PvsGT = header[4]
NRMSE_SvsGT = header[5]
PSNR_PvsGT = header[6]
PSNR_SvsGT = header[7]
header = """
<tr>
<th width = 9% align="left">{0}</th>
<th width = 4% align="left">{1}</th>
<th width = 15% align="center">{2}</th>
<th width = 14% align="left">{3}</th>
<th width = 15% align="center">{4}</th>
<th width = 14% align="left">{5}</th>
<th width = 15% align="center">{6}</th>
<th width = 14% align="left">{7}</th>
</tr>""".format(image,slice_n,mSSIM_PvsGT,mSSIM_SvsGT,NRMSE_PvsGT,NRMSE_SvsGT,PSNR_PvsGT,PSNR_SvsGT)
html = html+header
for row in metrics:
image = row[0]
slice_n = row[1]
mSSIM_PvsGT = row[2]
mSSIM_SvsGT = row[3]
NRMSE_PvsGT = row[4]
NRMSE_SvsGT = row[5]
PSNR_PvsGT = row[6]
PSNR_SvsGT = row[7]
cells = """
<tr>
<td width = 9% align="left">{0}</td>
<td width = 4% align="center">{1}</td>
<td width = 15% align="center">{2}</td>
<td width = 14% align="center">{3}</td>
<td width = 15% align="center">{4}</td>
<td width = 14% align="center">{5}</td>
<td width = 15% align="center">{6}</td>
<td width = 14% align="center">{7}</td>
</tr>""".format(image,slice_n,str(round(float(mSSIM_PvsGT),3)),str(round(float(mSSIM_SvsGT),3)),str(round(float(NRMSE_PvsGT),3)),str(round(float(NRMSE_SvsGT),3)),str(round(float(PSNR_PvsGT),3)),str(round(float(PSNR_SvsGT),3)))
html = html+cells
html = html+"""</body></table>"""
pdf.write_html(html)
pdf.ln(1)
pdf.set_font('')
pdf.set_font_size(10.)
ref_1 = 'References:\n - ZeroCostDL4Mic: von Chamier, Lucas & Laine, Romain, et al. "ZeroCostDL4Mic: an open platform to simplify access and use of Deep-Learning in Microscopy." bioRxiv (2020).'
pdf.multi_cell(190, 5, txt = ref_1, align='L')
ref_2 = '- Three-dimensional residual channel attention networks denoise and sharpen fluorescence microscopy image volumes, by Chen et al. bioRxiv (2020)'
pdf.multi_cell(190, 5, txt = ref_2, align='L')
pdf.ln(3)
reminder = 'To find the parameters and other information about how this model was trained, go to the training_report.pdf of this model which should be in the folder of the same name.'
pdf.set_font('Arial', size = 11, style='B')
pdf.multi_cell(190, 5, txt=reminder, align='C')
pdf.output(full_QC_model_path+'/Quality Control/'+QC_model_name+'_QC_report.pdf')
!pip freeze > requirements.txt
```
# **3. Select your parameters and paths**
---
## **3.1. Setting main training parameters**
---
<font size = 4>
<font size = 5> **Paths for training, predictions and results**
<font size = 4>**`Training_source:`, `Training_target`:** These are the paths to your folders containing the Training_source (Low SNR images) and Training_target (High SNR images or ground truth) training data respecively. To find the paths of the folders containing the respective datasets, go to your Files on the left of the notebook, navigate to the folder containing your files and copy the path by right-clicking on the folder, **Copy path** and pasting it into the right box below.
<font size = 4>**`model_name`:** Use only my_model -style, not my-model (Use "_" not "-"). Do not use spaces in the name. Avoid using the name of an existing model (saved in the same folder) as it will be overwritten.
<font size = 4>**`model_path`**: Enter the path where your model will be saved once trained (for instance your result folder).
<font size = 5>**Training Parameters**
<font size = 4>**`number of epochs`:**Input how many epochs (rounds) the network will be trained. Preliminary results can already be observed after a few (10-30) epochs, but a full training should run for 100-300 epochs. Evaluate the performance after training (see 5.). **Default value: 30**
<font size = 4>**`number_of_steps`:** Define the number of training steps by epoch. By default this parameter is calculated so that each patch is seen at least once per epoch. **Default value: 256**
<font size = 5>**Advanced Parameters - experienced users only**
<font size = 4>**`percentage_validation`:** Input the percentage of your training dataset you want to use to validate the network during the training. **Default value: 10**
<font size = 4>**`num_residual_groups`:** Number of residual groups in RCAN. **Default value: 5**
<font size = 4>**If you get an Out of memory (OOM) error during the training, manually decrease the num_residual_groups value until the OOM error disappear.**
<font size = 4>**`num_residual_blocks`:** Number of residual channel attention blocks in each residual group in RCAN. **Default value: 3**
<font size = 4>**`num_channels`:** Number of feature channels in RCAN. **Default value: 32**
<font size = 4>**`channel_reduction`:** Channel reduction ratio for channel attention. **Default value: 8**
```
#@markdown ###Path to training images:
# base folder of GT and low images
base = "/content"
# low SNR images
Training_source = "" #@param {type:"string"}
lowfile = Training_source+"/*.tif"
# Ground truth images
Training_target = "" #@param {type:"string"}
GTfile = Training_target+"/*.tif"
# model name and path
#@markdown ###Name of the model and path to model folder:
model_name = "" #@param {type:"string"}
model_path = "" #@param {type:"string"}
# create the training data file into model_path folder.
training_data = model_path+"/my_training_data.npz"
# other parameters for training.
#@markdown ###Training Parameters
#@markdown Number of epochs:
number_of_epochs = 30#@param {type:"number"}
number_of_steps = 256#@param {type:"number"}
#@markdown ###Advanced Parameters
Use_Default_Advanced_Parameters = True #@param {type:"boolean"}
#@markdown ###If not, please input:
percentage_validation = 10 #@param {type:"number"}
num_residual_groups = 5 #@param {type:"number"}
num_residual_blocks = 3 #@param {type:"number"}
num_channels = 32 #@param {type:"number"}
channel_reduction = 8 #@param {type:"number"}
if (Use_Default_Advanced_Parameters):
print("Default advanced parameters enabled")
percentage_validation = 10
num_residual_groups = 5
num_channels = 32
num_residual_blocks = 3
channel_reduction = 8
percentage = percentage_validation/100
full_model_path = model_path+'/'+model_name
#here we check that no model with the same name already exist, if so print a warning
if os.path.exists(model_path+'/'+model_name):
print(bcolors.WARNING +"!! WARNING: "+model_name+" already exists and will be deleted in the following cell !!")
print(bcolors.WARNING +"To continue training "+model_name+", choose a new model_name here, and load "+model_name+" in section 3.3"+W)
# Here we disable pre-trained model by default (in case the next cell is not ran)
Use_pretrained_model = False
# Here we disable data augmentation by default (in case the cell is not ran)
Use_Data_augmentation = False
#Load one randomly chosen training source file
random_choice=random.choice(os.listdir(Training_source))
x = imread(Training_source+"/"+random_choice)
# Here we check that the input images are stacks
if len(x.shape) == 3:
print("Image dimensions (z,y,x)",x.shape)
if not len(x.shape) == 3:
print(bcolors.WARNING +"Your images appear to have the wrong dimensions. Image dimension",x.shape)
#Find image Z dimension and select the mid-plane
Image_Z = x.shape[0]
mid_plane = int(Image_Z / 2)+1
#Find image XY dimension
Image_Y = x.shape[1]
Image_X = x.shape[2]
# Here we split the data between training and validation
# Here we count the number of files in the training target folder
Filelist = os.listdir(Training_target)
number_files = len(Filelist)
File_for_validation = int((number_files)/percentage_validation)+1
#Here we split the training dataset between training and validation
# Everything is copied in the /Content Folder
Training_source_temp = "/content/training_source"
if os.path.exists(Training_source_temp):
shutil.rmtree(Training_source_temp)
os.makedirs(Training_source_temp)
Training_target_temp = "/content/training_target"
if os.path.exists(Training_target_temp):
shutil.rmtree(Training_target_temp)
os.makedirs(Training_target_temp)
Validation_source_temp = "/content/validation_source"
if os.path.exists(Validation_source_temp):
shutil.rmtree(Validation_source_temp)
os.makedirs(Validation_source_temp)
Validation_target_temp = "/content/validation_target"
if os.path.exists(Validation_target_temp):
shutil.rmtree(Validation_target_temp)
os.makedirs(Validation_target_temp)
list_source = os.listdir(os.path.join(Training_source))
list_target = os.listdir(os.path.join(Training_target))
#Move files into the temporary source and target directories:
for f in os.listdir(os.path.join(Training_source)):
shutil.copy(Training_source+"/"+f, Training_source_temp+"/"+f)
for p in os.listdir(os.path.join(Training_target)):
shutil.copy(Training_target+"/"+p, Training_target_temp+"/"+p)
list_source_temp = os.listdir(os.path.join(Training_source_temp))
list_target_temp = os.listdir(os.path.join(Training_target_temp))
#Here we move images to be used for validation
for i in range(File_for_validation):
name = list_source_temp[i]
shutil.move(Training_source_temp+"/"+name, Validation_source_temp+"/"+name)
shutil.move(Training_target_temp+"/"+name, Validation_target_temp+"/"+name)
#Load one randomly chosen training target file
os.chdir(Training_target)
y = imread(Training_target+"/"+random_choice)
f=plt.figure(figsize=(16,8))
plt.subplot(1,2,1)
plt.imshow(x[mid_plane], norm=simple_norm(x[mid_plane], percent = 99), interpolation='nearest')
plt.axis('off')
plt.title('Low SNR image (single Z plane)');
plt.subplot(1,2,2)
plt.imshow(y[mid_plane], norm=simple_norm(y[mid_plane], percent = 99), interpolation='nearest')
plt.axis('off')
plt.title('High SNR image (single Z plane)');
plt.savefig('/content/TrainingDataExample_3D_RCAN.png',bbox_inches='tight',pad_inches=0)
```
## **3.2. Data augmentation**
---
<font size = 4>Data augmentation can improve training progress by amplifying differences in the dataset. This can be useful if the available dataset is small since, in this case, it is possible that a network could quickly learn every example in the dataset (overfitting), without augmentation. Augmentation is not necessary for training and if your training dataset is large you should disable it.
<font size = 4> **However, data augmentation is not a magic solution and may also introduce issues. Therefore, we recommend that you train your network with and without augmentation, and use the QC section to validate that it improves overall performances.**
<font size = 4>Data augmentation is performed here by rotating the training images in the XY-Plane and flipping them along X-Axis.
<font size = 4>**The flip option alone will double the size of your dataset, rotation will quadruple and both together will increase the dataset by a factor of 8.**
```
Use_Data_augmentation = False #@param{type:"boolean"}
#@markdown Select this option if you want to use augmentation to increase the size of your dataset
#@markdown **Rotate each image 3 times by 90 degrees.**
Rotation = False #@param{type:"boolean"}
#@markdown **Flip each image once around the x axis of the stack.**
Flip = False #@param{type:"boolean"}
#@markdown **Would you like to save your augmented images?**
Save_augmented_images = False #@param {type:"boolean"}
Saving_path = "" #@param {type:"string"}
if not Save_augmented_images:
Saving_path= "/content"
def rotation_aug(Source_path, Target_path, flip=False):
Source_images = os.listdir(Source_path)
Target_images = os.listdir(Target_path)
for image in Source_images:
source_img = io.imread(os.path.join(Source_path,image))
target_img = io.imread(os.path.join(Target_path,image))
# Source Rotation
source_img_90 = np.rot90(source_img,axes=(1,2))
source_img_180 = np.rot90(source_img_90,axes=(1,2))
source_img_270 = np.rot90(source_img_180,axes=(1,2))
# Target Rotation
target_img_90 = np.rot90(target_img,axes=(1,2))
target_img_180 = np.rot90(target_img_90,axes=(1,2))
target_img_270 = np.rot90(target_img_180,axes=(1,2))
# Add a flip to the rotation
if flip == True:
source_img_lr = np.fliplr(source_img)
source_img_90_lr = np.fliplr(source_img_90)
source_img_180_lr = np.fliplr(source_img_180)
source_img_270_lr = np.fliplr(source_img_270)
target_img_lr = np.fliplr(target_img)
target_img_90_lr = np.fliplr(target_img_90)
target_img_180_lr = np.fliplr(target_img_180)
target_img_270_lr = np.fliplr(target_img_270)
#source_img_90_ud = np.flipud(source_img_90)
# Save the augmented files
# Source images
io.imsave(Saving_path+'/augmented_source/'+image,source_img)
io.imsave(Saving_path+'/augmented_source/'+os.path.splitext(image)[0]+'_90.tif',source_img_90)
io.imsave(Saving_path+'/augmented_source/'+os.path.splitext(image)[0]+'_180.tif',source_img_180)
io.imsave(Saving_path+'/augmented_source/'+os.path.splitext(image)[0]+'_270.tif',source_img_270)
# Target images
io.imsave(Saving_path+'/augmented_target/'+image,target_img)
io.imsave(Saving_path+'/augmented_target/'+os.path.splitext(image)[0]+'_90.tif',target_img_90)
io.imsave(Saving_path+'/augmented_target/'+os.path.splitext(image)[0]+'_180.tif',target_img_180)
io.imsave(Saving_path+'/augmented_target/'+os.path.splitext(image)[0]+'_270.tif',target_img_270)
if flip == True:
io.imsave(Saving_path+'/augmented_source/'+os.path.splitext(image)[0]+'_lr.tif',source_img_lr)
io.imsave(Saving_path+'/augmented_source/'+os.path.splitext(image)[0]+'_90_lr.tif',source_img_90_lr)
io.imsave(Saving_path+'/augmented_source/'+os.path.splitext(image)[0]+'_180_lr.tif',source_img_180_lr)
io.imsave(Saving_path+'/augmented_source/'+os.path.splitext(image)[0]+'_270_lr.tif',source_img_270_lr)
io.imsave(Saving_path+'/augmented_target/'+os.path.splitext(image)[0]+'_lr.tif',target_img_lr)
io.imsave(Saving_path+'/augmented_target/'+os.path.splitext(image)[0]+'_90_lr.tif',target_img_90_lr)
io.imsave(Saving_path+'/augmented_target/'+os.path.splitext(image)[0]+'_180_lr.tif',target_img_180_lr)
io.imsave(Saving_path+'/augmented_target/'+os.path.splitext(image)[0]+'_270_lr.tif',target_img_270_lr)
def flip(Source_path, Target_path):
Source_images = os.listdir(Source_path)
Target_images = os.listdir(Target_path)
for image in Source_images:
source_img = io.imread(os.path.join(Source_path,image))
target_img = io.imread(os.path.join(Target_path,image))
source_img_lr = np.fliplr(source_img)
target_img_lr = np.fliplr(target_img)
io.imsave(Saving_path+'/augmented_source/'+image,source_img)
io.imsave(Saving_path+'/augmented_source/'+os.path.splitext(image)[0]+'_lr.tif',source_img_lr)
io.imsave(Saving_path+'/augmented_target/'+image,target_img)
io.imsave(Saving_path+'/augmented_target/'+os.path.splitext(image)[0]+'_lr.tif',target_img_lr)
if Use_Data_augmentation:
if os.path.exists(Saving_path+'/augmented_source'):
shutil.rmtree(Saving_path+'/augmented_source')
os.mkdir(Saving_path+'/augmented_source')
if os.path.exists(Saving_path+'/augmented_target'):
shutil.rmtree(Saving_path+'/augmented_target')
os.mkdir(Saving_path+'/augmented_target')
print("Data augmentation enabled")
print("Data augmentation in progress....")
if Rotation == True:
rotation_aug(Training_source_temp,Training_target_temp,flip=Flip)
elif Rotation == False and Flip == True:
flip(Training_source_temp,Training_target_temp)
print("Done")
if not Use_Data_augmentation:
print(bcolors.WARNING+"Data augmentation disabled")
```
# **4. Train the network**
---
## **4.1. Prepare the training data and model for training**
---
<font size = 4>Here, we use the information from 3. to build the model and convert the training data into a suitable format for training.
```
#@markdown ##Create the model and dataset objects
# --------------------- Here we delete the model folder if it already exist ------------------------
if os.path.exists(model_path+'/'+model_name):
print(bcolors.WARNING +"!! WARNING: Model folder already exists and has been removed !!" + W)
shutil.rmtree(model_path+'/'+model_name)
print("Preparing the config file...")
if Use_Data_augmentation == True:
Training_source_temp = Saving_path+'/augmented_source'
Training_target_temp = Saving_path+'/augmented_target'
# Here we prepare the JSON file
import json
# Config file for 3D-RCAN
dictionary ={
"epochs": number_of_epochs,
"steps_per_epoch": number_of_steps,
"num_residual_groups": num_residual_groups,
"training_data_dir": {"raw": Training_source_temp,
"gt": Training_target_temp},
"validation_data_dir": {"raw": Validation_source_temp,
"gt": Validation_target_temp},
"num_channels": num_channels,
"num_residual_blocks": num_residual_blocks,
"channel_reduction": channel_reduction
}
json_object = json.dumps(dictionary, indent = 4)
with open("/content/config.json", "w") as outfile:
outfile.write(json_object)
# Export pdf summary of training parameters
pdf_export(augmentation = Use_Data_augmentation)
print("Done")
```
## **4.2. Start Training**
---
<font size = 4>When playing the cell below you should see updates after each epoch (round). Network training can take some time.
<font size = 4>* **CRITICAL NOTE:** Google Colab has a time limit for processing (to prevent using GPU power for datamining). Training time must be less than 12 hours! If training takes longer than 12 hours, please decrease the number of epochs or number of patches. Another way circumvent this is to save the parameters of the model after training and start training again from this point.
<font size = 4>Once training is complete, the trained model is automatically saved on your Google Drive, in the **model_path** folder that was selected in Section 3. It is however wise to download the folder from Google Drive as all data can be erased at the next training if using the same folder.
```
#@markdown ##Start Training
start = time.time()
# Start Training
!python /content/3D-RCAN/train.py -c /content/config.json -o "$full_model_path"
print("Training, done.")
if os.path.exists(model_path+"/"+model_name+"/Quality Control"):
shutil.rmtree(model_path+"/"+model_name+"/Quality Control")
os.makedirs(model_path+"/"+model_name+"/Quality Control")
# Displaying the time elapsed for training
dt = time.time() - start
mins, sec = divmod(dt, 60)
hour, mins = divmod(mins, 60)
print("Time elapsed:",hour, "hour(s)",mins,"min(s)",round(sec),"sec(s)")
#Create a pdf document with training summary
pdf_export(trained = True, augmentation = Use_Data_augmentation)
```
# **5. Evaluate your model**
---
<font size = 4>This section allows the user to perform important quality checks on the validity and generalisability of the trained model.
<font size = 4>**We highly recommend to perform quality control on all newly trained models.**
```
# model name and path
#@markdown ###Do you want to assess the model you just trained ?
Use_the_current_trained_model = True #@param {type:"boolean"}
#@markdown ###If not, please provide the path to the model folder:
QC_model_folder = "" #@param {type:"string"}
#Here we define the loaded model name and path
QC_model_name = os.path.basename(QC_model_folder)
QC_model_path = os.path.dirname(QC_model_folder)
if (Use_the_current_trained_model):
QC_model_name = model_name
QC_model_path = model_path
full_QC_model_path = QC_model_path+'/'+QC_model_name+'/'
if os.path.exists(full_QC_model_path):
print("The "+QC_model_name+" network will be evaluated")
else:
W = '\033[0m' # white (normal)
R = '\033[31m' # red
print(R+'!! WARNING: The chosen model does not exist !!'+W)
print('Please make sure you provide a valid model path and model name before proceeding further.')
```
## **5.1. Inspection of the loss function**
---
<font size = 4>First, it is good practice to evaluate the training progress by comparing the training loss with the validation loss. The latter is a metric which shows how well the network performs on a subset of unseen data which is set aside from the training dataset. For more information on this, see for example [this review](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6381354/) by Nichols *et al.*
<font size = 4>**Training loss** describes an error value after each epoch for the difference between the model's prediction and its ground-truth target.
<font size = 4>**Validation loss** describes the same error value between the model's prediction on a validation image and compared to it's target.
<font size = 4>During training both values should decrease before reaching a minimal value which does not decrease further even after more training. Comparing the development of the validation loss with the training loss can give insights into the model's performance.
<font size = 4>Decreasing **Training loss** and **Validation loss** indicates that training is still necessary and increasing the `number_of_epochs` is recommended. Note that the curves can look flat towards the right side, just because of the y-axis scaling. The network has reached convergence once the curves flatten out. After this point no further training is required. If the **Validation loss** suddenly increases again an the **Training loss** simultaneously goes towards zero, it means that the network is overfitting to the training data. In other words the network is remembering the exact patterns from the training data and no longer generalizes well to unseen data. In this case the training dataset has to be increased.
```
#@markdown ##Play the cell to show a plot of training errors vs. epoch number
%load_ext tensorboard
%tensorboard --logdir "$full_QC_model_path"
```
## **5.2. Error mapping and quality metrics estimation**
---
<font size = 4>This section will display SSIM maps and RSE maps as well as calculating total SSIM, NRMSE and PSNR metrics for all the images provided in the "Source_QC_folder" and "Target_QC_folder" !
<font size = 4>**1. The SSIM (structural similarity) map**
<font size = 4>The SSIM metric is used to evaluate whether two images contain the same structures. It is a normalized metric and an SSIM of 1 indicates a perfect similarity between two images. Therefore for SSIM, the closer to 1, the better. The SSIM maps are constructed by calculating the SSIM metric in each pixel by considering the surrounding structural similarity in the neighbourhood of that pixel (currently defined as window of 11 pixels and with Gaussian weighting of 1.5 pixel standard deviation, see our Wiki for more info).
<font size=4>**mSSIM** is the SSIM value calculated across the entire window of both images.
<font size=4>**The output below shows the SSIM maps with the mSSIM**
<font size = 4>**2. The RSE (Root Squared Error) map**
<font size = 4>This is a display of the root of the squared difference between the normalized predicted and target or the source and the target. In this case, a smaller RSE is better. A perfect agreement between target and prediction will lead to an RSE map showing zeros everywhere (dark).
<font size =4>**NRMSE (normalised root mean squared error)** gives the average difference between all pixels in the images compared to each other. Good agreement yields low NRMSE scores.
<font size = 4>**PSNR (Peak signal-to-noise ratio)** is a metric that gives the difference between the ground truth and prediction (or source input) in decibels, using the peak pixel values of the prediction and the MSE between the images. The higher the score the better the agreement.
<font size=4>**The output below shows the RSE maps with the NRMSE and PSNR values.**
```
#@markdown ##Choose the folders that contain your Quality Control dataset
Source_QC_folder = "" #@param{type:"string"}
Target_QC_folder = "" #@param{type:"string"}
path_metrics_save = QC_model_path+'/'+QC_model_name+'/Quality Control/'
path_QC_prediction = path_metrics_save+'Prediction'
# Create a quality control/Prediction Folder
if os.path.exists(path_QC_prediction):
shutil.rmtree(path_QC_prediction)
os.makedirs(path_QC_prediction)
# Perform the predictions
print("Restoring images...")
!python /content/3D-RCAN/apply.py -m "$full_QC_model_path" -i "$Source_QC_folder" -o "$path_QC_prediction"
print("Done...")
def normalize(x, pmin=3, pmax=99.8, axis=None, clip=False, eps=1e-20, dtype=np.float32):
"""This function is adapted from Martin Weigert"""
"""Percentile-based image normalization."""
mi = np.percentile(x,pmin,axis=axis,keepdims=True)
ma = np.percentile(x,pmax,axis=axis,keepdims=True)
return normalize_mi_ma(x, mi, ma, clip=clip, eps=eps, dtype=dtype)
def normalize_mi_ma(x, mi, ma, clip=False, eps=1e-20, dtype=np.float32):#dtype=np.float32
"""This function is adapted from Martin Weigert"""
if dtype is not None:
x = x.astype(dtype,copy=False)
mi = dtype(mi) if np.isscalar(mi) else mi.astype(dtype,copy=False)
ma = dtype(ma) if np.isscalar(ma) else ma.astype(dtype,copy=False)
eps = dtype(eps)
try:
import numexpr
x = numexpr.evaluate("(x - mi) / ( ma - mi + eps )")
except ImportError:
x = (x - mi) / ( ma - mi + eps )
if clip:
x = np.clip(x,0,1)
return x
def norm_minmse(gt, x, normalize_gt=True):
"""This function is adapted from Martin Weigert"""
"""
normalizes and affinely scales an image pair such that the MSE is minimized
Parameters
----------
gt: ndarray
the ground truth image
x: ndarray
the image that will be affinely scaled
normalize_gt: bool
set to True of gt image should be normalized (default)
Returns
-------
gt_scaled, x_scaled
"""
if normalize_gt:
gt = normalize(gt, 0.1, 99.9, clip=False).astype(np.float32, copy = False)
x = x.astype(np.float32, copy=False) - np.mean(x)
#x = x - np.mean(x)
gt = gt.astype(np.float32, copy=False) - np.mean(gt)
#gt = gt - np.mean(gt)
scale = np.cov(x.flatten(), gt.flatten())[0, 1] / np.var(x.flatten())
return gt, scale * x
# Open and create the csv file that will contain all the QC metrics
with open(path_metrics_save+'QC_metrics_'+QC_model_name+".csv", "w", newline='') as file:
writer = csv.writer(file)
# Write the header in the csv file
writer.writerow(["File name","Slice #","Prediction v. GT mSSIM","Input v. GT mSSIM", "Prediction v. GT NRMSE", "Input v. GT NRMSE", "Prediction v. GT PSNR", "Input v. GT PSNR"])
# These lists will be used to collect all the metrics values per slice
file_name_list = []
slice_number_list = []
mSSIM_GvP_list = []
mSSIM_GvS_list = []
NRMSE_GvP_list = []
NRMSE_GvS_list = []
PSNR_GvP_list = []
PSNR_GvS_list = []
# These lists will be used to display the mean metrics for the stacks
mSSIM_GvP_list_mean = []
mSSIM_GvS_list_mean = []
NRMSE_GvP_list_mean = []
NRMSE_GvS_list_mean = []
PSNR_GvP_list_mean = []
PSNR_GvS_list_mean = []
# Let's loop through the provided dataset in the QC folders
for thisFile in os.listdir(Source_QC_folder):
if not os.path.isdir(os.path.join(Source_QC_folder, thisFile)):
print('Running QC on: '+thisFile)
test_GT_stack = io.imread(os.path.join(Target_QC_folder, thisFile))
test_source_stack = io.imread(os.path.join(Source_QC_folder,thisFile))
test_prediction_stack_raw = io.imread(os.path.join(path_metrics_save+"Prediction/",thisFile))
test_prediction_stack = test_prediction_stack_raw[:, 1, :, :]
n_slices = test_GT_stack.shape[0]
# Calculating the position of the mid-plane slice
z_mid_plane = int(n_slices / 2)+1
img_SSIM_GTvsPrediction_stack = np.zeros((n_slices, test_GT_stack.shape[1], test_GT_stack.shape[2]))
img_SSIM_GTvsSource_stack = np.zeros((n_slices, test_GT_stack.shape[1], test_GT_stack.shape[2]))
img_RSE_GTvsPrediction_stack = np.zeros((n_slices, test_GT_stack.shape[1], test_GT_stack.shape[2]))
img_RSE_GTvsSource_stack = np.zeros((n_slices, test_GT_stack.shape[1], test_GT_stack.shape[2]))
for z in range(n_slices):
# -------------------------------- Normalising the dataset --------------------------------
test_GT_norm, test_source_norm = norm_minmse(test_GT_stack[z], test_source_stack[z], normalize_gt=True)
test_GT_norm, test_prediction_norm = norm_minmse(test_GT_stack[z], test_prediction_stack[z], normalize_gt=True)
# -------------------------------- Calculate the SSIM metric and maps --------------------------------
# Calculate the SSIM maps and index
index_SSIM_GTvsPrediction, img_SSIM_GTvsPrediction = structural_similarity(test_GT_norm, test_prediction_norm, data_range=1.0, full=True, gaussian_weights=True, use_sample_covariance=False, sigma=1.5)
index_SSIM_GTvsSource, img_SSIM_GTvsSource = structural_similarity(test_GT_norm, test_source_norm, data_range=1.0, full=True, gaussian_weights=True, use_sample_covariance=False, sigma=1.5)
#Calculate ssim_maps
img_SSIM_GTvsPrediction_stack[z] = img_as_float32(img_SSIM_GTvsPrediction, force_copy=False)
img_SSIM_GTvsSource_stack[z] = img_as_float32(img_SSIM_GTvsSource, force_copy=False)
# -------------------------------- Calculate the NRMSE metrics --------------------------------
# Calculate the Root Squared Error (RSE) maps
img_RSE_GTvsPrediction = np.sqrt(np.square(test_GT_norm - test_prediction_norm))
img_RSE_GTvsSource = np.sqrt(np.square(test_GT_norm - test_source_norm))
# Calculate SE maps
img_RSE_GTvsPrediction_stack[z] = img_as_float32(img_RSE_GTvsPrediction, force_copy=False)
img_RSE_GTvsSource_stack[z] = img_as_float32(img_RSE_GTvsSource, force_copy=False)
# Normalised Root Mean Squared Error (here it's valid to take the mean of the image)
NRMSE_GTvsPrediction = np.sqrt(np.mean(img_RSE_GTvsPrediction))
NRMSE_GTvsSource = np.sqrt(np.mean(img_RSE_GTvsSource))
# Calculate the PSNR between the images
PSNR_GTvsPrediction = psnr(test_GT_norm,test_prediction_norm,data_range=1.0)
PSNR_GTvsSource = psnr(test_GT_norm,test_source_norm,data_range=1.0)
writer.writerow([thisFile, str(z),str(index_SSIM_GTvsPrediction),str(index_SSIM_GTvsSource),str(NRMSE_GTvsPrediction),str(NRMSE_GTvsSource), str(PSNR_GTvsPrediction), str(PSNR_GTvsSource)])
# Collect values to display in dataframe output
slice_number_list.append(z)
mSSIM_GvP_list.append(index_SSIM_GTvsPrediction)
mSSIM_GvS_list.append(index_SSIM_GTvsSource)
NRMSE_GvP_list.append(NRMSE_GTvsPrediction)
NRMSE_GvS_list.append(NRMSE_GTvsSource)
PSNR_GvP_list.append(PSNR_GTvsPrediction)
PSNR_GvS_list.append(PSNR_GTvsSource)
if (z == z_mid_plane): # catch these for display
SSIM_GTvsP_forDisplay = index_SSIM_GTvsPrediction
SSIM_GTvsS_forDisplay = index_SSIM_GTvsSource
NRMSE_GTvsP_forDisplay = NRMSE_GTvsPrediction
NRMSE_GTvsS_forDisplay = NRMSE_GTvsSource
# If calculating average metrics for dataframe output
file_name_list.append(thisFile)
mSSIM_GvP_list_mean.append(sum(mSSIM_GvP_list)/len(mSSIM_GvP_list))
mSSIM_GvS_list_mean.append(sum(mSSIM_GvS_list)/len(mSSIM_GvS_list))
NRMSE_GvP_list_mean.append(sum(NRMSE_GvP_list)/len(NRMSE_GvP_list))
NRMSE_GvS_list_mean.append(sum(NRMSE_GvS_list)/len(NRMSE_GvS_list))
PSNR_GvP_list_mean.append(sum(PSNR_GvP_list)/len(PSNR_GvP_list))
PSNR_GvS_list_mean.append(sum(PSNR_GvS_list)/len(PSNR_GvS_list))
# ----------- Change the stacks to 32 bit images -----------
img_SSIM_GTvsSource_stack_32 = img_as_float32(img_SSIM_GTvsSource_stack, force_copy=False)
img_SSIM_GTvsPrediction_stack_32 = img_as_float32(img_SSIM_GTvsPrediction_stack, force_copy=False)
img_RSE_GTvsSource_stack_32 = img_as_float32(img_RSE_GTvsSource_stack, force_copy=False)
img_RSE_GTvsPrediction_stack_32 = img_as_float32(img_RSE_GTvsPrediction_stack, force_copy=False)
# ----------- Saving the error map stacks -----------
io.imsave(path_metrics_save+'SSIM_GTvsSource_'+thisFile,img_SSIM_GTvsSource_stack_32)
io.imsave(path_metrics_save+'SSIM_GTvsPrediction_'+thisFile,img_SSIM_GTvsPrediction_stack_32)
io.imsave(path_metrics_save+'RSE_GTvsSource_'+thisFile,img_RSE_GTvsSource_stack_32)
io.imsave(path_metrics_save+'RSE_GTvsPrediction_'+thisFile,img_RSE_GTvsPrediction_stack_32)
#Averages of the metrics per stack as dataframe output
pdResults = pd.DataFrame(file_name_list, columns = ["File name"])
pdResults["Prediction v. GT mSSIM"] = mSSIM_GvP_list_mean
pdResults["Input v. GT mSSIM"] = mSSIM_GvS_list_mean
pdResults["Prediction v. GT NRMSE"] = NRMSE_GvP_list_mean
pdResults["Input v. GT NRMSE"] = NRMSE_GvS_list_mean
pdResults["Prediction v. GT PSNR"] = PSNR_GvP_list_mean
pdResults["Input v. GT PSNR"] = PSNR_GvS_list_mean
# All data is now processed saved
Test_FileList = os.listdir(Source_QC_folder) # this assumes, as it should, that both source and target are named the same way
plt.figure(figsize=(20,20))
# Currently only displays the last computed set, from memory
# Target (Ground-truth)
plt.subplot(3,3,1)
plt.axis('off')
img_GT = io.imread(os.path.join(Target_QC_folder, Test_FileList[-1]))
# Calculating the position of the mid-plane slice
z_mid_plane = int(img_GT.shape[0] / 2)+1
plt.imshow(img_GT[z_mid_plane], norm=simple_norm(img_GT[z_mid_plane], percent = 99))
plt.title('Target (slice #'+str(z_mid_plane)+')')
# Source
plt.subplot(3,3,2)
plt.axis('off')
img_Source = io.imread(os.path.join(Source_QC_folder, Test_FileList[-1]))
plt.imshow(img_Source[z_mid_plane], norm=simple_norm(img_Source[z_mid_plane], percent = 99))
plt.title('Source (slice #'+str(z_mid_plane)+')')
#Prediction
plt.subplot(3,3,3)
plt.axis('off')
img_Prediction_raw = io.imread(os.path.join(path_metrics_save+'Prediction/', Test_FileList[-1]))
img_Prediction = img_Prediction_raw[:, 1, :, :]
plt.imshow(img_Prediction[z_mid_plane], norm=simple_norm(img_Prediction[z_mid_plane], percent = 99))
plt.title('Prediction (slice #'+str(z_mid_plane)+')')
#Setting up colours
cmap = plt.cm.CMRmap
#SSIM between GT and Source
plt.subplot(3,3,5)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_SSIM_GTvsSource = io.imread(os.path.join(path_metrics_save, 'SSIM_GTvsSource_'+Test_FileList[-1]))
imSSIM_GTvsSource = plt.imshow(img_SSIM_GTvsSource[z_mid_plane], cmap = cmap, vmin=0, vmax=1)
plt.colorbar(imSSIM_GTvsSource,fraction=0.046, pad=0.04)
plt.title('Target vs. Source',fontsize=15)
plt.xlabel('mSSIM: '+str(round(SSIM_GTvsS_forDisplay,3)),fontsize=14)
plt.ylabel('SSIM maps',fontsize=20, rotation=0, labelpad=75)
#SSIM between GT and Prediction
plt.subplot(3,3,6)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_SSIM_GTvsPrediction = io.imread(os.path.join(path_metrics_save, 'SSIM_GTvsPrediction_'+Test_FileList[-1]))
imSSIM_GTvsPrediction = plt.imshow(img_SSIM_GTvsPrediction[z_mid_plane], cmap = cmap, vmin=0,vmax=1)
plt.colorbar(imSSIM_GTvsPrediction,fraction=0.046, pad=0.04)
plt.title('Target vs. Prediction',fontsize=15)
plt.xlabel('mSSIM: '+str(round(SSIM_GTvsP_forDisplay,3)),fontsize=14)
#Root Squared Error between GT and Source
plt.subplot(3,3,8)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_RSE_GTvsSource = io.imread(os.path.join(path_metrics_save, 'RSE_GTvsSource_'+Test_FileList[-1]))
imRSE_GTvsSource = plt.imshow(img_RSE_GTvsSource[z_mid_plane], cmap = cmap, vmin=0, vmax = 1)
plt.colorbar(imRSE_GTvsSource,fraction=0.046,pad=0.04)
plt.title('Target vs. Source',fontsize=15)
plt.xlabel('NRMSE: '+str(round(NRMSE_GTvsS_forDisplay,3))+', PSNR: '+str(round(PSNR_GTvsSource,3)),fontsize=14)
#plt.title('Target vs. Source PSNR: '+str(round(PSNR_GTvsSource,3)))
plt.ylabel('RSE maps',fontsize=20, rotation=0, labelpad=75)
#Root Squared Error between GT and Prediction
plt.subplot(3,3,9)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_RSE_GTvsPrediction = io.imread(os.path.join(path_metrics_save, 'RSE_GTvsPrediction_'+Test_FileList[-1]))
imRSE_GTvsPrediction = plt.imshow(img_RSE_GTvsPrediction[z_mid_plane], cmap = cmap, vmin=0, vmax=1)
plt.colorbar(imRSE_GTvsPrediction,fraction=0.046,pad=0.04)
plt.title('Target vs. Prediction',fontsize=15)
plt.xlabel('NRMSE: '+str(round(NRMSE_GTvsP_forDisplay,3))+', PSNR: '+str(round(PSNR_GTvsPrediction,3)),fontsize=14)
plt.savefig(full_QC_model_path+'/Quality Control/QC_example_data.png',bbox_inches='tight',pad_inches=0)
print('-----------------------------------')
print('Here are the average scores for the stacks you tested in Quality control. To see values for all slices, open the .csv file saved in the Quality Control folder.')
pdResults.head()
#Make a pdf summary of the QC results
qc_pdf_export()
```
# **6. Using the trained model**
---
<font size = 4>In this section the unseen data is processed using the trained model (in section 4). First, your unseen images are uploaded and prepared for prediction. After that your trained model from section 4 is activated and finally saved into your Google Drive.
## **6.1. Generate prediction(s) from unseen dataset**
---
<font size = 4>The current trained model (from section 4.2) can now be used to process images. If you want to use an older model, untick the **Use_the_current_trained_model** box and enter the name and path of the model to use. Predicted output images are saved in your **Result_folder** folder as restored image stacks (ImageJ-compatible TIFF images).
<font size = 4>**`Data_folder`:** This folder should contain the images that you want to use your trained network on for processing.
<font size = 4>**`Result_folder`:** This folder will contain the predicted output images.
```
#@markdown ##Provide the path to your dataset and to the folder where the prediction will be saved, then play the cell to predict output on your unseen images.
Data_folder = "" #@param {type:"string"}
Result_folder = "" #@param {type:"string"}
# model name and path
#@markdown ###Do you want to use the current trained model?
Use_the_current_trained_model = True #@param {type:"boolean"}
#@markdown ###If not, please provide the path to the model folder:
Prediction_model_folder = "" #@param {type:"string"}
#Here we find the loaded model name and parent path
Prediction_model_name = os.path.basename(Prediction_model_folder)
Prediction_model_path = os.path.dirname(Prediction_model_folder)
if (Use_the_current_trained_model):
print("Using current trained network")
Prediction_model_name = model_name
Prediction_model_path = model_path
full_Prediction_model_path = Prediction_model_path+'/'+Prediction_model_name+'/'
if os.path.exists(full_Prediction_model_path):
print("The "+Prediction_model_name+" network will be used.")
else:
W = '\033[0m' # white (normal)
R = '\033[31m' # red
print(R+'!! WARNING: The chosen model does not exist !!'+W)
print('Please make sure you provide a valid model path and model name before proceeding further.')
print("Restoring images...")
!python /content/3D-RCAN/apply.py -m "$full_Prediction_model_path" -i "$Data_folder" -o "$Result_folder"
print("Images saved into the result folder:", Result_folder)
#Display an example
random_choice=random.choice(os.listdir(Data_folder))
x = imread(Data_folder+"/"+random_choice)
z_mid_plane = int(x.shape[0] / 2)+1
@interact
def show_results(file=os.listdir(Data_folder), z_plane=widgets.IntSlider(min=0, max=(x.shape[0]-1), step=1, value=z_mid_plane)):
x = imread(Data_folder+"/"+file)
y_raw = imread(Result_folder+"/"+file)
y = y_raw[:, 1, :, :]
f=plt.figure(figsize=(16,8))
plt.subplot(1,2,1)
plt.imshow(x[z_plane], norm=simple_norm(x[z_plane], percent = 99), interpolation='nearest')
plt.axis('off')
plt.title('Noisy Input (single Z plane)');
plt.subplot(1,2,2)
plt.imshow(y[z_plane], norm=simple_norm(y[z_plane], percent = 99), interpolation='nearest')
plt.axis('off')
plt.title('Prediction (single Z plane)');
```
## **6.2. Download your predictions**
---
<font size = 4>**Store your data** and ALL its results elsewhere by downloading it from Google Drive and after that clean the original folder tree (datasets, results, trained model etc.) if you plan to train or use new networks. Please note that the notebook will otherwise **OVERWRITE** all files which have the same name.
#**Thank you for using 3D-RCAN!**
| github_jupyter |
# Nosy Bagging Duelling Prioritised Replay Double Deep Q Learning - A simple ambulance dispatch point allocation model
## Reinforcement learning introduction
### RL involves:
* Trial and error search
* Receiving and maximising reward (often delayed)
* Linking state -> action -> reward
* Must be able to sense something of their environment
* Involves uncertainty in sensing and linking action to reward
* Learning -> improved choice of actions over time
* All models find a way to balance best predicted action vs. exploration
### Elements of RL
* *Environment*: all observable and unobservable information relevant to us
* *Observation*: sensing the environment
* *State*: the perceived (or perceivable) environment
* *Agent*: senses environment, decides on action, receives and monitors rewards
* *Action*: may be discrete (e.g. turn left) or continuous (accelerator pedal)
* *Policy* (how to link state to action; often based on probabilities)
* *Reward signal*: aim is to accumulate maximum reward over time
* *Value function* of a state: prediction of likely/possible long-term reward
* *Q*: prediction of likely/possible long-term reward of an *action*
* *Advantage*: The difference in Q between actions in a given state (sums to zero for all actions)
* *Model* (optional): a simulation of the environment
### Types of model
* *Model-based*: have model of environment (e.g. a board game)
* *Model-free*: used when environment not fully known
* *Policy-based*: identify best policy directly
* *Value-based*: estimate value of a decision
* *Off-policy*: can learn from historic data from other agent
* *On-policy*: requires active learning from current decisions
## Duelling Deep Q Networks for Reinforcement Learning
Q = The expected future rewards discounted over time. This is what we are trying to maximise.
The aim is to teach a network to take the current state observations and recommend the action with greatest Q.
Duelling is very similar to Double DQN, except that the policy net splits into two. One component reduces to a single value, which will model the state *value*. The other component models the *advantage*, the difference in Q between different actions (the mean value is subtracted from all values, so that the advtantage always sums to zero). These are aggregated to produce Q for each action.
<img src="./images/duelling_dqn.png" width="500"/>
Q is learned through the Bellman equation, where the Q of any state and action is the immediate reward achieved + the discounted maximum Q value (the best action taken) of next best action, where gamma is the discount rate.
$$Q(s,a)=r + \gamma.maxQ(s',a')$$
## Key DQN components
<img src="./images/dqn_components.png" width="700"/>
## General method for Q learning:
Overall aim is to create a neural network that predicts Q. Improvement comes from improved accuracy in predicting 'current' understood Q, and in revealing more about Q as knowledge is gained (some rewards only discovered after time).
<img src="./images/dqn_process.png" width="600|"/>
Target networks are used to stabilise models, and are only updated at intervals. Changes to Q values may lead to changes in closely related states (i.e. states close to the one we are in at the time) and as the network tries to correct for errors it can become unstable and suddenly lose signficiant performance. Target networks (e.g. to assess Q) are updated only infrequently (or gradually), so do not have this instability problem.
## Training networks
Double DQN contains two networks. This ammendment, from simple DQN, is to decouple training of Q for current state and target Q derived from next state which are closely correlated when comparing input features.
The *policy network* is used to select action (action with best predicted Q) when playing the game.
When training, the predicted best *action* (best predicted Q) is taken from the *policy network*, but the *policy network* is updated using the predicted Q value of the next state from the *target network* (which is updated from the policy network less frequently). So, when training, the action is selected using Q values from the *policy network*, but the the *policy network* is updated to better predict the Q value of that action from the *target network*. The *policy network* is copied across to the *target network* every *n* steps (e.g. 1000).
<img src="./images/dqn_training.png" width="700|"/>
## Bagging (Bootstrap Aggregation)
Each network is trained from the same memory, but have different starting weights and are trained on different bootstrap samples from that memory. In this example actions are chosen randomly from each of the networks (an alternative could be to take the most common action recommended by the networks, or an average output). This bagging method may also be used to have some measure of uncertainty of action by looking at the distribution of actions recommended from the different nets. Bagging may also be used to aid exploration during stages where networks are providing different suggested action.
<img src="./images/bagging.png" width="800|"/>
## Noisy layers
Noisy layers are an alternative to epsilon-greedy exploration (here, we leave the epsilon-greedy code in the model, but set it to reduce to zero immediately after the period of fully random action choice).
For every weight in the layer we have a random value that we draw from the normal distribution. This random value is used to add noise to the output. The parameters for the extent of noise for each weight, sigma, are stored within the layer and get trained as part of the standard back-propogation.
A modification to normal nosiy layers is to use layers with ‘factorized gaussian noise’. This reduces the number of random numbers to be sampled (so is less computationally expensive). There are two random vectors, one with the size of the input, and the other with the size of the output. A random matrix is created by calculating the outer product of the two vectors.
## Prioritised replay
In standard DQN samples are taken randomly from the memory (replay buffer). In *prioritised replay* samples are taken in proportion to their loss when training the network; where the network has the greatest error in predicting the target valur of a state/action, then those samples will be sampled more frequently (which will reduce the error in the network until the sample is not prioritised). In other words, the training focuses more heavenly on samples it gets most wrong, and spends less time training on samples that it can acurately predict already.
This priority may also be used as a weight for training the network, but this i snot implemented here; we use loss just for sampling.
When we use the loss for priority we add a small value (1e-5) t the loss. This avoids any sample having zero priority (and never having a chance of being sampled). For frequency of sampling we also raise the loss to the power of 'alpha' (default value of 0.6). Smaller values of alpha will compress the differences between samples, making the priority weighting less significant in the frequency of sampling.
The memory stores the priority/loss of state/action/Next_state/reward, and this is particular to each network, so we create a separate memory for each network.
## References
Double DQN:
van Hasselt H, Guez A, Silver D. (2015) Deep Reinforcement Learning with Double Q-learning. arXiv:150906461 http://arxiv.org/abs/1509.06461
Bagging:
Osband I, Blundell C, Pritzel A, et al. (2016) Deep Exploration via Bootstrapped DQN. arXiv:160204621 http://arxiv.org/abs/1602.04621
Noisy networks:
Fortunato M, Azar MG, Piot B, et al. (2019) Noisy Networks for Exploration. arXiv:170610295 http://arxiv.org/abs/1706.10295
Prioritised replay:
Schaul T, Quan J, Antonoglou I, et al (2016). Prioritized Experience Replay. arXiv:151105952 http://arxiv.org/abs/1511.05952
Code for the nosiy layers comes from:
Lapan, M. (2020). Deep Reinforcement Learning Hands-On: Apply modern RL methods to practical problems of chatbots, robotics, discrete optimization, web automation, and more, 2nd Edition. Packt Publishing.
## Code structure
<img src="./images/dqn_program_structure.png" width="700|"/>
```
################################################################################
# 1 Import packages #
################################################################################
from amboworld.environment import Env
import math
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import random
import torch
import torch.nn as nn
import torch.optim as optim
from torch.nn import functional as F
# Use a double ended queue (deque) for memory
# When memory is full, this will replace the oldest value with the new one
from collections import deque
# Supress all warnings (e.g. deprecation warnings) for regular use
import warnings
warnings.filterwarnings("ignore")
################################################################################
# 2 Define model parameters #
################################################################################
# Set whether to display on screen (slows model)
DISPLAY_ON_SCREEN = False
# Discount rate of future rewards
GAMMA = 0.99
# Learing rate for neural network
LEARNING_RATE = 0.003
# Maximum number of game steps (state, action, reward, next state) to keep
MEMORY_SIZE = 10000000
# Sample batch size for policy network update
BATCH_SIZE = 5
# Number of game steps to play before starting training (all random actions)
REPLAY_START_SIZE = 50000
# Number of steps between policy -> target network update
SYNC_TARGET_STEPS = 1000
# Exploration rate (epsilon) is probability of choosing a random action
EXPLORATION_MAX = 1.0
EXPLORATION_MIN = 0.0
# Reduction in epsilon with each game step
EXPLORATION_DECAY = 0.0
# Training episodes
TRAINING_EPISODES = 50
# Set number of parallel networks
NUMBER_OF_NETS = 5
# Results filename
RESULTS_NAME = 'bagging_pr_noisy_d3qn'
# SIM PARAMETERS
RANDOM_SEED = 42
SIM_DURATION = 5000
NUMBER_AMBULANCES = 3
NUMBER_INCIDENT_POINTS = 1
INCIDENT_RADIUS = 2
NUMBER_DISPTACH_POINTS = 25
AMBOWORLD_SIZE = 50
INCIDENT_INTERVAL = 60
EPOCHS = 2
AMBO_SPEED = 60
AMBO_FREE_FROM_HOSPITAL = False
################################################################################
# 3 Define DQN (Duelling Deep Q Network) class #
# (Used for both policy and target nets) #
################################################################################
"""
Code for nosiy layers comes from:
Lapan, M. (2020). Deep Reinforcement Learning Hands-On: Apply modern RL methods
to practical problems of chatbots, robotics, discrete optimization,
web automation, and more, 2nd Edition. Packt Publishing.
"""
class NoisyLinear(nn.Linear):
"""
Noisy layer for network.
For every weight in the layer we have a random value that we draw from the
normal distribution.Paraemters for the noise, sigma, are stored within the
layer and get trained as part of the standard back-propogation.
'register_buffer' is used to create tensors in the network that are not
updated during back-propogation. They are used to create normal
distributions to add noise (multiplied by sigma which is a paramater in the
network).
"""
def __init__(self, in_features, out_features,
sigma_init=0.017, bias=True):
super(NoisyLinear, self).__init__(
in_features, out_features, bias=bias)
w = torch.full((out_features, in_features), sigma_init)
self.sigma_weight = nn.Parameter(w)
z = torch.zeros(out_features, in_features)
self.register_buffer("epsilon_weight", z)
if bias:
w = torch.full((out_features,), sigma_init)
self.sigma_bias = nn.Parameter(w)
z = torch.zeros(out_features)
self.register_buffer("epsilon_bias", z)
self.reset_parameters()
def reset_parameters(self):
std = math.sqrt(3 / self.in_features)
self.weight.data.uniform_(-std, std)
self.bias.data.uniform_(-std, std)
def forward(self, input):
self.epsilon_weight.normal_()
bias = self.bias
if bias is not None:
self.epsilon_bias.normal_()
bias = bias + self.sigma_bias * \
self.epsilon_bias.data
v = self.sigma_weight * self.epsilon_weight.data + self.weight
return F.linear(input, v, bias)
class NoisyFactorizedLinear(nn.Linear):
"""
NoisyNet layer with factorized gaussian noise. This reduces the number of
random numbers to be sampled (so less computationally expensive). There are
two random vectors. One with the size of the input, and the other with the
size of the output. A random matrix is create by calculating the outer
product of the two vectors.
'register_buffer' is used to create tensors in the network that are not
updated during back-propogation. They are used to create normal
distributions to add noise (multiplied by sigma which is a paramater in the
network).
"""
def __init__(self, in_features, out_features,
sigma_zero=0.4, bias=True):
super(NoisyFactorizedLinear, self).__init__(
in_features, out_features, bias=bias)
sigma_init = sigma_zero / math.sqrt(in_features)
w = torch.full((out_features, in_features), sigma_init)
self.sigma_weight = nn.Parameter(w)
z1 = torch.zeros(1, in_features)
self.register_buffer("epsilon_input", z1)
z2 = torch.zeros(out_features, 1)
self.register_buffer("epsilon_output", z2)
if bias:
w = torch.full((out_features,), sigma_init)
self.sigma_bias = nn.Parameter(w)
def forward(self, input):
self.epsilon_input.normal_()
self.epsilon_output.normal_()
func = lambda x: torch.sign(x) * torch.sqrt(torch.abs(x))
eps_in = func(self.epsilon_input.data)
eps_out = func(self.epsilon_output.data)
bias = self.bias
if bias is not None:
bias = bias + self.sigma_bias * eps_out.t()
noise_v = torch.mul(eps_in, eps_out)
v = self.weight + self.sigma_weight * noise_v
return F.linear(input, v, bias)
class DQN(nn.Module):
"""Deep Q Network. Udes for both policy (action) and target (Q) networks."""
def __init__(self, observation_space, action_space):
"""Constructor method. Set up neural nets."""
# nerurones per hidden layer = 2 * max of observations or actions
neurons_per_layer = 2 * max(observation_space, action_space)
# Set starting exploration rate
self.exploration_rate = EXPLORATION_MAX
# Set up action space (choice of possible actions)
self.action_space = action_space
# First layerswill be common to both Advantage and value
super(DQN, self).__init__()
self.feature = nn.Sequential(
nn.Linear(observation_space, neurons_per_layer),
nn.ReLU()
)
# Advantage has same number of outputs as the action space
self.advantage = nn.Sequential(
NoisyFactorizedLinear(neurons_per_layer, neurons_per_layer),
nn.ReLU(),
NoisyFactorizedLinear(neurons_per_layer, action_space)
)
# State value has only one output (one value per state)
self.value = nn.Sequential(
nn.Linear(neurons_per_layer, neurons_per_layer),
nn.ReLU(),
nn.Linear(neurons_per_layer, 1)
)
def act(self, state):
"""Act either randomly or by redicting action that gives max Q"""
# Act randomly if random number < exploration rate
if np.random.rand() < self.exploration_rate:
action = random.randrange(self.action_space)
else:
# Otherwise get predicted Q values of actions
q_values = self.forward(torch.FloatTensor(state))
# Get index of action with best Q
action = np.argmax(q_values.detach().numpy()[0])
return action
def forward(self, x):
x = self.feature(x)
advantage = self.advantage(x)
value = self.value(x)
action_q = value + advantage - advantage.mean()
return action_q
################################################################################
# 4 Define policy net training function #
################################################################################
def optimize(policy_net, target_net, memory):
"""
Update model by sampling from memory.
Uses policy network to predict best action (best Q).
Uses target network to provide target of Q for the selected next action.
"""
# Do not try to train model if memory is less than reqired batch size
if len(memory) < BATCH_SIZE:
return
# Reduce exploration rate (exploration rate is stored in policy net)
policy_net.exploration_rate *= EXPLORATION_DECAY
policy_net.exploration_rate = max(EXPLORATION_MIN,
policy_net.exploration_rate)
# Sample a random batch from memory
batch = memory.sample(BATCH_SIZE)
for state, action, reward, state_next, terminal, index in batch:
state_action_values = policy_net(torch.FloatTensor(state))
# Get target Q for policy net update
if not terminal:
# For non-terminal actions get Q from policy net
expected_state_action_values = policy_net(torch.FloatTensor(state))
# Detach next state values from gradients to prevent updates
expected_state_action_values = expected_state_action_values.detach()
# Get next state action with best Q from the policy net (double DQN)
policy_next_state_values = policy_net(torch.FloatTensor(state_next))
policy_next_state_values = policy_next_state_values.detach()
best_action = np.argmax(policy_next_state_values[0].numpy())
# Get target net next state
next_state_action_values = target_net(torch.FloatTensor(state_next))
# Use detach again to prevent target net gradients being updated
next_state_action_values = next_state_action_values.detach()
best_next_q = next_state_action_values[0][best_action].numpy()
updated_q = reward + (GAMMA * best_next_q)
expected_state_action_values[0][action] = updated_q
else:
# For termal actions Q = reward (-1)
expected_state_action_values = policy_net(torch.FloatTensor(state))
# Detach values from gradients to prevent gradient update
expected_state_action_values = expected_state_action_values.detach()
# Set Q for all actions to reward (-1)
expected_state_action_values[0] = reward
# Set net to training mode
policy_net.train()
# Reset net gradients
policy_net.optimizer.zero_grad()
# calculate loss
loss_v = nn.MSELoss()(state_action_values, expected_state_action_values)
# Backpropogate loss
loss_v.backward()
# Update replay buffer (add 1e-5 to loss to avoid zero priority with no
# chance of being sampled).
loss_numpy = loss_v.data.numpy()
memory.update_priorities(index, loss_numpy + 1e-5)
# Update network gradients
policy_net.optimizer.step()
return
################################################################################
# 5 Define prioritised replay memory class #
################################################################################
class NaivePrioritizedBuffer():
"""
Based on code from https://github.com/higgsfield/RL-Adventure
Each sample (state, action, reward, next_state, done) has an associated
priority, which is the loss from training the policy network. The priority
is used to adjust the frequency of sampling.
"""
def __init__(self, capacity=MEMORY_SIZE, prob_alpha=0.6):
self.prob_alpha = prob_alpha
self.capacity = capacity
self.buffer = []
self.pos = 0
self.priorities = np.zeros((capacity,), dtype=np.float32)
def remember(self, state, action, reward, next_state, done):
"""
Add sample (state, action, reward, next_state, done) to memory, or
replace oldest sample if memory full"""
max_prio = self.priorities.max() if self.buffer else 1.0
if len(self.buffer) < self.capacity:
# Add new sample when room in memory
self.buffer.append((state, action, reward, next_state, done))
else:
# Replace sample when memory full
self.buffer[self.pos] = (state, action, reward, next_state, done)
# Set maximum priority present
self.priorities[self.pos] = max_prio
# Increment replacement position
self.pos = (self.pos + 1) % self.capacity
def sample(self, batch_size, beta=0.4):
# Get priorities
if len(self.buffer) == self.capacity:
prios = self.priorities
else:
prios = self.priorities[:self.pos]
# Raise priorities by the square of 'alpha'
# (lower alpha compresses differences)
probs = prios ** self.prob_alpha
# Normlaise priorities
probs /= probs.sum()
# Sample using priorities for relative sampling frequency
indices = np.random.choice(len(self.buffer), batch_size, p=probs)
samples = [self.buffer[idx] for idx in indices]
# Add index to sample (used to update priority after getting new loss)
batch = []
for index, sample in enumerate(samples):
sample = list(sample)
sample.append(indices[index])
batch.append(sample)
return batch
def update_priorities(self, index, priority):
"""Update sample priority with new loss"""
self.priorities[index] = priority
def __len__(self):
return len(self.buffer)
################################################################################
# 6 Define results plotting function #
################################################################################
def plot_results(run, exploration, score, mean_call_to_arrival,
mean_assignment_to_arrival):
"""Plot and report results at end of run"""
# Set up chart (ax1 and ax2 share x-axis to combine two plots on one graph)
fig = plt.figure(figsize=(6,6))
ax1 = fig.add_subplot(111)
ax2 = ax1.twinx()
# Plot results
lns1 = ax1.plot(
run, exploration, label='exploration', color='g', linestyle=':')
lns2 = ax2.plot(run, mean_call_to_arrival,
label='call to arrival', color='r')
lns3 = ax2.plot(run, mean_assignment_to_arrival,
label='assignment to arrival', color='b', linestyle='--')
# Get combined legend
lns = lns1 + lns2 + lns3
labs = [l.get_label() for l in lns]
ax1.legend(lns, labs, loc='upper center', bbox_to_anchor=(0.5, -0.1), ncol=3)
# Set axes
ax1.set_xlabel('run')
ax1.set_ylabel('exploration')
ax2.set_ylabel('Response time')
filename = 'output/' + RESULTS_NAME +'.png'
plt.savefig(filename, dpi=300)
plt.show()
################################################################################
# 7 Main program #
################################################################################
def qambo():
"""Main program loop"""
############################################################################
# 8 Set up environment #
############################################################################
# Set up game environemnt
sim = Env(
random_seed = RANDOM_SEED,
duration_incidents = SIM_DURATION,
number_ambulances = NUMBER_AMBULANCES,
number_incident_points = NUMBER_INCIDENT_POINTS,
incident_interval = INCIDENT_INTERVAL,
number_epochs = EPOCHS,
number_dispatch_points = NUMBER_DISPTACH_POINTS,
incident_range = INCIDENT_RADIUS,
max_size = AMBOWORLD_SIZE,
ambo_kph = AMBO_SPEED,
ambo_free_from_hospital = AMBO_FREE_FROM_HOSPITAL
)
# Get number of observations returned for state
observation_space = sim.observation_size
# Get number of actions possible
action_space = sim.action_number
############################################################################
# 9 Set up policy and target nets #
############################################################################
# Set up policy and target neural nets
policy_nets = [DQN(observation_space, action_space)
for i in range(NUMBER_OF_NETS)]
target_nets = [DQN(observation_space, action_space)
for i in range(NUMBER_OF_NETS)]
best_nets = [DQN(observation_space, action_space)
for i in range(NUMBER_OF_NETS)]
# Set optimizer, copy weights from policy_net to target, and
for i in range(NUMBER_OF_NETS):
# Set optimizer
policy_nets[i].optimizer = optim.Adam(
params=policy_nets[i].parameters(), lr=LEARNING_RATE)
# Copy weights from policy -> target
target_nets[i].load_state_dict(policy_nets[i].state_dict())
# Set target net to eval rather than training mode
target_nets[i].eval()
############################################################################
# 10 Set up memory #
############################################################################
# Set up memomry
memory = [NaivePrioritizedBuffer() for i in range(NUMBER_OF_NETS)]
############################################################################
# 11 Set up + start training loop #
############################################################################
# Set up run counter and learning loop
run = 0
all_steps = 0
continue_learning = True
best_reward = -np.inf
# Set up list for results
results_run = []
results_exploration = []
results_score = []
results_mean_call_to_arrival = []
results_mean_assignment_to_arrival = []
# Continue repeating games (episodes) until target complete
while continue_learning:
########################################################################
# 12 Play episode #
########################################################################
# Increment run (episode) counter
run += 1
########################################################################
# 13 Reset game #
########################################################################
# Reset game environment and get first state observations
state = sim.reset()
# Reset total reward and rewards list
total_reward = 0
rewards = []
# Reshape state into 2D array with state obsverations as first 'row'
state = np.reshape(state, [1, observation_space])
# Continue loop until episode complete
while True:
####################################################################
# 14 Game episode loop #
####################################################################
####################################################################
# 15 Get action #
####################################################################
# Get actions to take (use evalulation mode)
actions = []
for i in range(NUMBER_OF_NETS):
policy_nets[i].eval()
actions.append(policy_nets[i].act(state))
# Randomly choose an action from net actions
random_index = random.randint(0, NUMBER_OF_NETS - 1)
action = actions[random_index]
####################################################################
# 16 Play action (get S', R, T) #
####################################################################
# Act
state_next, reward, terminal, info = sim.step(action)
total_reward += reward
# Update trackers
rewards.append(reward)
# Reshape state into 2D array with state observations as first 'row'
state_next = np.reshape(state_next, [1, observation_space])
# Update display if needed
if DISPLAY_ON_SCREEN:
sim.render()
####################################################################
# 17 Add S/A/R/S/T to memory #
####################################################################
# Record state, action, reward, new state & terminal
for i in range(NUMBER_OF_NETS):
memory[i].remember(state, action, reward, state_next, terminal)
# Update state
state = state_next
####################################################################
# 18 Check for end of episode #
####################################################################
# Actions to take if end of game episode
if terminal:
# Get exploration rate
exploration = policy_nets[0].exploration_rate
# Clear print row content
clear_row = '\r' + ' ' * 79 + '\r'
print(clear_row, end='')
print(f'Run: {run}, ', end='')
print(f'Exploration: {exploration: .3f}, ', end='')
average_reward = np.mean(rewards)
print(f'Average reward: {average_reward:4.1f}, ', end='')
mean_assignment_to_arrival = np.mean(info['assignment_to_arrival'])
print(f'Mean assignment to arrival: {mean_assignment_to_arrival:4.1f}, ', end='')
mean_call_to_arrival = np.mean(info['call_to_arrival'])
print(f'Mean call to arrival: {mean_call_to_arrival:4.1f}, ', end='')
demand_met = info['fraction_demand_met']
print(f'Demand met {demand_met:0.3f}')
# Add to results lists
results_run.append(run)
results_exploration.append(exploration)
results_score.append(total_reward)
results_mean_call_to_arrival.append(mean_call_to_arrival)
results_mean_assignment_to_arrival.append(mean_assignment_to_arrival)
# Save model if best reward
total_reward = np.sum(rewards)
if total_reward > best_reward:
best_reward = total_reward
# Copy weights to best net
for i in range(NUMBER_OF_NETS):
best_nets[i].load_state_dict(policy_nets[i].state_dict())
################################################################
# 18b Check for end of learning #
################################################################
if run == TRAINING_EPISODES:
continue_learning = False
# End episode loop
break
####################################################################
# 19 Update policy net #
####################################################################
# Avoid training model if memory is not of sufficient length
if len(memory[0]) > REPLAY_START_SIZE:
# Update policy net
for i in range(NUMBER_OF_NETS):
optimize(policy_nets[i], target_nets[i], memory[i])
################################################################
# 20 Update target net periodically #
################################################################
# Use load_state_dict method to copy weights from policy net
if all_steps % SYNC_TARGET_STEPS == 0:
for i in range(NUMBER_OF_NETS):
target_nets[i].load_state_dict(
policy_nets[i].state_dict())
############################################################################
# 21 Learning complete - plot and save results #
############################################################################
# Target reached. Plot results
plot_results(results_run, results_exploration, results_score,
results_mean_call_to_arrival, results_mean_assignment_to_arrival)
# SAVE RESULTS
run_details = pd.DataFrame()
run_details['run'] = results_run
run_details['exploration '] = results_exploration
run_details['mean_call_to_arrival'] = results_mean_call_to_arrival
run_details['mean_assignment_to_arrival'] = results_mean_assignment_to_arrival
filename = 'output/' + RESULTS_NAME + '.csv'
run_details.to_csv(filename, index=False)
############################################################################
# Test best model #
############################################################################
print()
print('Test Model')
print('----------')
for i in range(NUMBER_OF_NETS):
best_nets[i].eval()
best_nets[i].exploration_rate = 0
# Set up results dictionary
results = dict()
results['call_to_arrival'] = []
results['assign_to_arrival'] = []
results['demand_met'] = []
# Replicate model runs
for run in range(30):
# Reset game environment and get first state observations
state = sim.reset()
state = np.reshape(state, [1, observation_space])
# Continue loop until episode complete
while True:
# Get actions to take (use evalulation mode)
actions = []
for i in range(NUMBER_OF_NETS):
actions.append(best_nets[i].act(state))
# Randomly choose an action from net actions
random_index = random.randint(0, NUMBER_OF_NETS - 1)
action = actions[random_index]
# Act
state_next, reward, terminal, info = sim.step(action)
# Reshape state into 2D array with state observations as first 'row'
state_next = np.reshape(state_next, [1, observation_space])
# Update state
state = state_next
if terminal:
print(f'Run: {run}, ', end='')
mean_assignment_to_arrival = np.mean(info['assignment_to_arrival'])
print(f'Mean assignment to arrival: {mean_assignment_to_arrival:4.1f}, ', end='')
mean_call_to_arrival = np.mean(info['call_to_arrival'])
print(f'Mean call to arrival: {mean_call_to_arrival:4.1f}, ', end='')
demand_met = info['fraction_demand_met']
print(f'Demand met: {demand_met:0.3f}')
# Add to results
results['call_to_arrival'].append(mean_call_to_arrival)
results['assign_to_arrival'].append(mean_assignment_to_arrival)
results['demand_met'].append(demand_met)
# End episode loop
break
results = pd.DataFrame(results)
filename = './output/results_' + RESULTS_NAME +'.csv'
results.to_csv(filename, index=False)
print()
print(results.describe())
return run_details
######################## MODEL ENTRY POINT #####################################
# Run model and return last run results
last_run = qambo()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
```
# 샘플 펭귄 데이터셋 읽기
- Python에서 데이터셋은 주로 pandas의 DataFrame이란 데이터 형식으로 처리함
- DataFrame 변수는 주로 df라는 이름으로 선언됨
```
df = sns.load_dataset('penguins') ## 샘플 데이터셋을 읽어서 df로 선언
df.head(10) ## 첫 10개 행 확인
df.tail(5) ## 마지막 5개 행 확인
df.info() ## 데이터 형식 확인, 결측값 제외 값 개수 확인
## cf. object : int, string, list 등 모든 종류의 데이터
df.describe() ## 데이터의 각 열 별 평균, 표준편차, 최솟값, 중앙값, 최댓값
df.describe(include='all') ## object 데이터에 대해서도 묘사
print('평균')
print(df.mean())
print('\n중앙값')
print(df.median())
print('\n최댓값')
print(df.max())
print('\n최솟값')
print(df.min())
```
# 데이터 가시화
```
## 여러 그래프 그리기 1 : plt.subplots
fig, ax = plt.subplots(nrows=1, ncols=3, figsize=(15,5))
sns.countplot(x='island', data=df, ax=ax[0])
sns.countplot(x='species', data=df, ax=ax[1])
sns.countplot(x='sex', data=df, ax=ax[2])
plt.show()
## 여러 그래프 그리기 2 : plt.subplot
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(2, 1, 1) ## nrows, ncols, index
sns.countplot(x='island', data=df, ax=ax)
ax = fig.add_subplot(2, 2, 3) ## nrows, ncols, index
sns.countplot(x='species', data=df, ax=ax)
ax = fig.add_subplot(2, 2, 4) ## nrows, ncols, index
sns.countplot(x='sex', data=df, ax=ax)
plt.show()
sns.countplot(x='island',hue='species',data=df)
plt.show()
```
# 데이터 슬라이싱
```
print('COLUMN', df.columns) ## 데이터셋의 columns(열)
print('INDEX',df.index) ## 데이터셋의 row(행)은 index로 사용함
# cf. df.index는 int값이 아닐 수도 있음
## 특정 행 추출 : df.loc, df.iloc
print(df.loc[339]) ## index의 이름이 339인 데이터 출력
print(df.iloc[-5]) ## 마지막에서 5번째 index의 데이터 출력
print(df.loc[339:350:2])
print(df.iloc[339:350:2])
## 복수의 행 추출 : list 사용
indices = [5,10,15,20,25]
df.loc[indices]
## 이 때, loc[] 안에 int를 넣으면 Series가 출력, list를 넣으면 DataFrame이 출력됨
## Series는 데이터의 한 열 또는 행을 표현하기 위한 데이터 형식
## 특정 열 추출 : 4가지 방법
print(df.species)
print(df['species'])
print(df.loc[:,'species']) ## 모든 index에 대해 species column 출력
print(df.iloc[:, 0]) ## 0번째 column인 species 출력
## 복수의 열 추출 : list 사용
cols = ['species', 'island', 'bill_length_mm']
print(df.loc[:,cols])
print(df[cols])
cols = [0, 1, 2]
print(df.iloc[:,cols])
### 열 및 행 추출
indices = [5, 10]
cols = ['species', 'bill_length_mm']
df.loc[indices, cols]
## 데이터 필터링
print(df['species']=='Adelie') ## species가 Adelie인 index는 True로, 나머지는 False로
df.loc[df['species']=='Adelie'] ## species가 Adelie인 index만 출력
## species가 Adelie인 데이터의 bill_length_mm와 bill_depth_mm 값만 출력
df.loc[df['species']=='Adelie', ['bill_length_mm', 'bill_depth_mm']]
```
# 결측값 매꾸기
```
# 각 값이 NaN인지 확인
df.isna()
# 각 열 별로 NaN값이 존재하는지 확인
print(df.isna().any())
# 각 행 별로 NaN값이 존재하는지 확인
df.isna().any(axis=1)
## NaN값이 존재하는 행만 출력
df.loc[df.isna().any(axis=1)]
## NaN값 채우기 : fillna
df2 = df.fillna(df.mean())
df2
df.loc[0]
df[df['species']=='Adelie']
```
| github_jupyter |

> **Copyright (c) 2021 CertifAI Sdn. Bhd.**<br>
<br>
This program is part of OSRFramework. You can redistribute it and/or modify
<br>it under the terms of the GNU Affero General Public License as published by
<br>the Free Software Foundation, either version 3 of the License, or
<br>(at your option) any later version.
<br>
<br>This program is distributed in the hope that it will be useful
<br>but WITHOUT ANY WARRANTY; without even the implied warranty of
<br>MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
<br>GNU Affero General Public License for more details.
<br>
<br>You should have received a copy of the GNU Affero General Public License
<br>along with this program. If not, see <http://www.gnu.org/licenses/>.
<br>
# Introduction
In this notebook, we are going to build neural machine translation (NMT) using Transformer with pytorch. This NMT could translate English to French.
Let's get started.

# What will we accomplish?
Steps to implement neural machine translation using Transformer with Pytorch:
> Step 1: Load and preprocess dataset
> Step 2: Building transformer architecture
> Step 3: Train the transformer model
> Step 4: Test the trained model
# Notebook Content
* [Load Dataset](#Load-Dataset)
* [Tokenization](#Tokenization)
* [Preprocessing](#Preprocessing)
* [Train-Test Split](#Train-Test-Split)
* [TabularDataset](#TabularDataset)
* [BucketIterator](#BucketIterator)
* [Custom Iterator](#Custom-Iterator)
* [Dive Deep into Transformer](#Dive-Deep-into-Transformer)
* [Embedding](#Embedding)
* [Positional Encoding](#Positional-Encoding)
* [Masking](#Masking)
* [Input Masks](#Input-Masks)
* [Target Sequence Masks](#Target-Sequence-Masks)
* [Multi-Headed Attention](#Multi-Headed-Attention)
* [Attention](#Attention)
* [Feed-Forward Network](#Feed-Forward-Network)
* [Normalisation](#Normalisation)
* [Building Transformer](#Building-Transformer)
* [EncoderLayer](#EncoderLayer)
* [DecoderLayer](#DecoderLayer)
* [Encoder](#Encoder)
* [Decoder](#Decoder)
* [Transformer](#Transformer)
* [Training the Model](#Training-the-Model)
* [Testing the Model](#Testing-the-Model)
# Load Dataset
The dataset we used is [parallel corpus French-English](http://www.statmt.org/europarl/v7/fr-en.tgz) dataset from [European Parliament Proceedings Parallel Corpus (1996–2011)](http://www.statmt.org/europarl/). This dataset contains 15 years of write-ups from E.U. proceedings, weighing in at 2,007,724 sentences, and 50,265,039 words. You should found the dataset in the `datasets` folder, else you may download it [here](http://www.statmt.org/europarl/v7/fr-en.tgz). You will have the following files after unzipping the downloaded file:
1. europarl-v7.fr-en.en
2. europarl-v7.fr-en.fr

Now we are going to load the dataset for preprocessing.
```
europarl_en = open('../../../resources/day_11/fr-en/europarl-v7.fr-en.en', encoding='utf-8').read().split('\n')
europarl_fr = open('../../../resources/day_11/fr-en/europarl-v7.fr-en.fr', encoding='utf-8').read().split('\n')
```
# Tokenization
The first job we need done is to **create a tokenizer for each language**. This is a function that will split the text into separate words and assign them unique numbers (indexes). This number will come into play later when we discuss embeddings.

He we will tokenize the text using **Torchtext** and **Spacy** together. Spacy is a library that has been specifically built to take sentences in various languages and split them into different tokens (see [here](https://spacy.io/) for more information). Without Spacy, Torchtext defaults to a simple .split(' ') method for tokenization. This is much less nuanced than Spacy’s approach, which will also split words like “don’t” into “do” and “n’t”, and much more.
```
import spacy
import torchtext
import torch
import numpy as np
from torchtext.legacy.data import Field, BucketIterator, TabularDataset
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# !python -m spacy download fr_core_news_lg
# !python -m spacy download en_core_web_lg
en = spacy.load('en_core_web_lg')
fr = spacy.load('fr_core_news_lg')
def tokenize_en(sentence):
return [tok.text for tok in en.tokenizer(sentence)]
def tokenize_fr(sentence):
return [tok.text for tok in fr.tokenizer(sentence)]
EN_TEXT = Field(tokenize=tokenize_en)
FR_TEXT = Field(tokenize=tokenize_fr, init_token = "<sos>", eos_token = "<eos>")
```
# Preprocessing
The best way to work with Torchtext is to turn your data into **spreadsheet format**, no matter the original format of your data file. This is due to the incredible versatility of the **Torchtext TabularDataset** function, which creates datasets from spreadsheet formats. So first to turn our data into an appropriate CSV file.
```
import pandas as pd
raw_data = {'English' : [line for line in europarl_en], 'French': [line for line in europarl_fr]}
df = pd.DataFrame(raw_data, columns=["English", "French"])
# Remove very long sentences and sentences where translations are not of roughly equal length
df['eng_len'] = df['English'].str.count(' ')
df['fr_len'] = df['French'].str.count(' ')
df = df.query('fr_len < 80 & eng_len < 80')
df = df.query('fr_len < eng_len * 1.5 & fr_len * 1.5 > eng_len')
```
## Train-Test Split
Now we are going to split the data into train set and test set. Fortunately Sklearn and Torchtext together make this process incredibly easy.
```
from sklearn.model_selection import train_test_split
# Create train and validation set
train, test = train_test_split(df, test_size=0.1)
train.to_csv("../../../resources/day_11/train.csv", index=False)
test.to_csv("../../../resources/day_11/test.csv", index=False)
```
This creates a train and test csv each with two columns (English, French), where each row contains an English sentence in the 'English' column, and its French translation in the 'French' column.
## TabularDataset
Calling the magic `TabularDataset.splits` then returns a train and test dataset with the respective data loaded into them, processed(/tokenized) according to the fields we defined earlier.
```
# Associate the text in the 'English' column with the EN_TEXT field, # and 'French' with FR_TEXT
data_fields = [('English', EN_TEXT), ('French', FR_TEXT)]
train, test = TabularDataset.splits(path='../../../resources/day_11', train='train.csv', validation='test.csv',
format='csv', fields=data_fields)
```
Processing a few million words can take a while so grab a cup of tea here…
```
FR_TEXT.build_vocab(train, test)
EN_TEXT.build_vocab(train, test)
```
To see what numbers the tokens have been assigned and vice versa in each field, we can use `self.vocab.stoi` and `self.vocab.itos`.
```
print(EN_TEXT.vocab.stoi['the'])
print(EN_TEXT.vocab.itos[11])
```
## BucketIterator
**BucketIterator** Defines an iterator that batches examples of similar lengths together.
It minimizes amount of padding needed while producing freshly shuffled batches for each new epoch. See pool for the bucketing procedure used.
```
train_iter = BucketIterator(train, batch_size=20, sort_key=lambda x: len(x.French), shuffle=True)
```
The `sort_key` dictates how to form each batch. The lambda function tells the iterator to try and find sentences of the **same length** (meaning more of the matrix is filled with useful data and less with padding).
```
batch = next(iter(train_iter))
print(batch.English)
print("Number of columns:", len(batch))
```
In each batch, sentences have been transposed so they are descending vertically (important: we will need to transpose these again to work with transformer). **Each index represents a token (word)**, and **each column represents a sentence**. We have 20 columns, as 20 was the batch_size we specified.
You might notice all the ‘1’s and think which incredibly common word is this the index for? Well the ‘1’ is not of course a word, but purely **padding**. While Torchtext is brilliant, it’s `sort_key` based batching leaves a little to be desired. Often the sentences aren’t of the same length at all, and you end up feeding a lot of padding into your network (as you can see with all the 1s in the last figure). We will solve this by implementing our own iterator.
## Custom Iterator
The custom iterator is built in reference to the code from http://nlp.seas.harvard.edu/2018/04/03/attention.html. Feel free to explore yourself to have more understanding about `MyIterator` class.
```
from torchtext.legacy import data
global max_src_in_batch, max_tgt_in_batch
def batch_size_fn(new, count, sofar):
"Keep augmenting batch and calculate total number of tokens + padding."
global max_src_in_batch, max_tgt_in_batch
if count == 1:
max_src_in_batch = 0
max_tgt_in_batch = 0
max_src_in_batch = max(max_src_in_batch, len(new.English))
max_tgt_in_batch = max(max_tgt_in_batch, len(new.French) + 2)
src_elements = count * max_src_in_batch
tgt_elements = count * max_tgt_in_batch
return max(src_elements, tgt_elements)
class MyIterator(data.Iterator):
def create_batches(self):
if self.train:
def pool(d, random_shuffler):
for p in data.batch(d, self.batch_size * 100):
p_batch = data.batch(
sorted(p, key=self.sort_key),
self.batch_size, self.batch_size_fn)
for b in random_shuffler(list(p_batch)):
yield b
self.batches = pool(self.data(), self.random_shuffler)
else:
self.batches = []
for b in data.batch(self.data(), self.batch_size, self.batch_size_fn):
self.batches.append(sorted(b, key=self.sort_key))
train_iter = MyIterator(train, batch_size= 64, device=device, repeat=False,
sort_key= lambda x: (len(x.English), len(x.French)),
batch_size_fn=batch_size_fn, train=True, shuffle=True)
```
# Dive Deep into Transformer

The diagram above shows the overview of the Transformer model. The inputs to the encoder will be the **English** sentence, and the 'Outputs' from the decoder will be the **French** sentence.
## Embedding
Embedding words has become standard practice in NMT, feeding the network with far more information about words than a one hot encoding would.

```
from torch import nn
class Embedder(nn.Module):
def __init__(self, vocab_size, embedding_dimension):
super().__init__()
self.embed = nn.Embedding(vocab_size, embedding_dimension)
def forward(self, x):
return self.embed(x)
```
When each word is fed into the network, this code will perform a look-up and retrieve its embedding vector. These vectors will then be learnt as a parameters by the model, adjusted with each iteration of gradient descent.
## Positional Encoding
In order for the model to make sense of a sentence, it needs to know two things about each word: what does the **word mean**? And what is its **position** in the sentence?
The embedding vector for each word will **learn the meaning**, so now we need to input something that tells the network about the word’s position.
*Vasmari et al* answered this problem by using these functions to create a constant of position-specific values:


This constant is a 2D matrix. Pos refers to the order in the sentence, and $i$ refers to the position along the embedding vector dimension. Each value in the pos/i matrix is then worked out using the equations above.

```
import math
class PositionalEncoder(nn.Module):
def __init__(self, d_model, max_seq_len = 200, dropout = 0.1):
super().__init__()
self.d_model = d_model
self.dropout = nn.Dropout(dropout)
# Create constant 'pe' matrix with values dependant on pos and i
pe = torch.zeros(max_seq_len, d_model)
for pos in range(max_seq_len):
for i in range(0, d_model, 2):
pe[pos, i] = \
math.sin(pos / (10000 ** ((2 * i)/d_model)))
pe[pos, i + 1] = \
math.cos(pos / (10000 ** ((2 * (i + 1))/d_model)))
pe = pe.unsqueeze(0)
self.register_buffer('pe', pe)
def forward(self, x):
# Make embeddings relatively larger
x = x * math.sqrt(self.d_model)
# Add constant to embedding
seq_len = x.size(1)
pe = Variable(self.pe[:,:seq_len], requires_grad=False)
if x.is_cuda:
pe.cuda()
x = x + pe
return self.dropout(x)
```
`PositionalEncoder` lets us add the **positional encoding to the embedding vector**, providing information about structure to the model.
The reason we increase the embedding values before addition is to make the positional encoding relatively smaller. This means the original meaning in the embedding vector won’t be lost when we add them together.
## Masking
Masking plays an important role in the transformer. It serves two purposes:
* In the encoder and decoder: To zero attention outputs wherever there is just padding in the input sentences.
* In the decoder: To prevent the decoder ‘peaking’ ahead at the rest of the translated sentence when predicting the next word.

### Input Masks
```
batch = next(iter(train_iter))
input_seq = batch.English.transpose(0,1)
input_pad = EN_TEXT.vocab.stoi['<pad>']
# creates mask with 0s wherever there is padding in the input
input_msk = (input_seq != input_pad).unsqueeze(1)
```
### Target Sequence Masks
```
from torch.autograd import Variable
target_seq = batch.French.transpose(0,1)
target_pad = FR_TEXT.vocab.stoi['<pad>']
target_msk = (target_seq != target_pad).unsqueeze(1)
```
The initial input into the decoder will be the **target sequence** (the French translation). The way the decoder predicts each output word is by making use of all the encoder outputs and the French sentence only up until the point of each word its predicting.
Therefore we need to prevent the first output predictions from being able to see later into the sentence. For this we use the `nopeak_mask`.
```
# Get seq_len for matrix
size = target_seq.size(1)
nopeak_mask = np.triu(np.ones((1, size, size)), k=1).astype('uint8')
nopeak_mask = Variable(torch.from_numpy(nopeak_mask) == 0).cuda()
print(nopeak_mask)
target_msk = target_msk & nopeak_mask
def create_masks(src, trg):
src_pad = EN_TEXT.vocab.stoi['<pad>']
trg_pad = FR_TEXT.vocab.stoi['<pad>']
src_mask = (src != src_pad).unsqueeze(-2)
if trg is not None:
trg_mask = (trg != trg_pad).unsqueeze(-2)
# Get seq_len for matrix
size = trg.size(1)
np_mask = nopeak_mask(size)
if device.type == 'cuda':
np_mask = np_mask.cuda()
trg_mask = trg_mask & np_mask
else:
trg_mask = None
return src_mask, trg_mask
def nopeak_mask(size):
np_mask = np.triu(np.ones((1, size, size)), k=1).astype('uint8')
np_mask = Variable(torch.from_numpy(np_mask) == 0)
return np_mask
```
If we later apply this mask to the attention scores, the values wherever the input is ahead will not be able to contribute when calculating the outputs.
## Multi-Headed Attention
Once we have our embedded values (with positional encodings) and our masks, we can start building the layers of our model.
Here is an overview of the multi-headed attention layer:

In multi-headed attention layer, each **input is split into multiple heads** which allows the network to simultaneously attend to different subsections of each embedding.
$V$, $K$ and $Q$ stand for ***key***, ***value*** and ***query***. These are terms used in attention functions. In the case of the Encoder, $V$, $K$ and $Q$ will simply be identical copies of the embedding vector (plus positional encoding). They will have the dimensions `Batch_size` * `seq_len` * `embedding_dimension`.
In multi-head attention we split the embedding vector into $N$ heads, so they will then have the dimensions `batch_size` * `N` * `seq_len` * (`embedding_dimension` / `N`).
This final dimension (`embedding_dimension` / `N`) we will refer to as $d_k$.
```
class MultiHeadAttention(nn.Module):
def __init__(self, heads, d_model, dropout = 0.1):
super().__init__()
self.d_model = d_model
self.d_k = d_model // heads
self.h = heads
self.q_linear = nn.Linear(d_model, d_model)
self.v_linear = nn.Linear(d_model, d_model)
self.k_linear = nn.Linear(d_model, d_model)
self.dropout = nn.Dropout(dropout)
self.out = nn.Linear(d_model, d_model)
def forward(self, q, k, v, mask=None):
bs = q.size(0)
# Perform linear operation and split into h heads
k = self.k_linear(k).view(bs, -1, self.h, self.d_k)
q = self.q_linear(q).view(bs, -1, self.h, self.d_k)
v = self.v_linear(v).view(bs, -1, self.h, self.d_k)
# Transpose to get dimensions bs * h * sl * d_model
k = k.transpose(1,2)
q = q.transpose(1,2)
v = v.transpose(1,2)
# Calculate attention using function we will define next
scores = attention(q, k, v, self.d_k, mask, self.dropout)
# Concatenate heads and put through final linear layer
concat = scores.transpose(1,2).contiguous()\
.view(bs, -1, self.d_model)
output = self.out(concat)
return output
```
## Attention
The equation below is the attention formula with retrieved from [Attention is All You Need](https://arxiv.org/abs/1706.03762) paper and it does a good job at explaining each step.


Each arrow in the diagram reflects a part of the equation.
Initially we must **multiply** $Q$ by the transpose of $K$. This is then scaled by **dividing the output by the square root** of $d_k$.
A step that’s not shown in the equation is the masking operation. Before we perform **Softmax**, we apply our mask and hence reduce values where the input is padding (or in the decoder, also where the input is ahead of the current word). Another step not shown is **dropout**, which we will apply after Softmax.
Finally, the last step is doing a **dot product** between the result so far and $V$.
```
import torch.nn.functional as F
def attention(q, k, v, d_k, mask=None, dropout=None):
scores = torch.matmul(q, k.transpose(-2, -1)) / math.sqrt(d_k)
if mask is not None:
mask = mask.unsqueeze(1)
scores = scores.masked_fill(mask == 0, -1e9)
scores = F.softmax(scores, dim=-1)
if dropout is not None:
scores = dropout(scores)
output = torch.matmul(scores, v)
return output
```
## Feed-Forward Network
The feed-forward layer just consists of two linear operations, with a **relu** and **dropout** operation in between them. It simply deepens our network, employing linear layers to **analyse patterns** in the attention layers output.

```
class FeedForward(nn.Module):
def __init__(self, d_model, d_ff=2048, dropout = 0.1):
super().__init__()
# We set d_ff as a default to 2048
self.linear_1 = nn.Linear(d_model, d_ff)
self.dropout = nn.Dropout(dropout)
self.linear_2 = nn.Linear(d_ff, d_model)
def forward(self, x):
x = self.dropout(F.relu(self.linear_1(x)))
x = self.linear_2(x)
return x
```
## Normalisation
Normalisation is highly important in deep neural networks. It prevents the range of values in the layers changing too much, meaning the model **trains faster** and has **better ability to generalise**.

We will be normalising our results between each layer in the encoder/decoder.
```
class Norm(nn.Module):
def __init__(self, d_model, eps = 1e-6):
super().__init__()
self.size = d_model
# create two learnable parameters to calibrate normalisation
self.alpha = nn.Parameter(torch.ones(self.size))
self.bias = nn.Parameter(torch.zeros(self.size))
self.eps = eps
def forward(self, x):
norm = self.alpha * (x - x.mean(dim=-1, keepdim=True)) / (x.std(dim=-1, keepdim=True) + self.eps) + self.bias
return norm
```
# Building Transformer
Let’s have another look at the over-all architecture and start building:

**One last Variable**: If you look at the diagram closely you can see a $N_x$ next to the encoder and decoder architectures. In reality, the encoder and decoder in the diagram above represent one layer of an encoder and one of the decoder. $N$ is the variable for the **number of layers** there will be. Eg. if `N=6`, the data goes through six encoder layers (with the architecture seen above), then these outputs are passed to the decoder which also consists of six repeating decoder layers.
We will now build `EncoderLayer` and `DecoderLayer` modules with the architecture shown in the model above. Then when we build the encoder and decoder we can define how many of these layers to have.
## EncoderLayer
```
# build an encoder layer with one multi-head attention layer and one feed-forward layer
class EncoderLayer(nn.Module):
def __init__(self, d_model, heads, dropout = 0.1):
super().__init__()
self.norm_1 = Norm(d_model)
self.norm_2 = Norm(d_model)
self.attn = MultiHeadAttention(heads, d_model)
self.ff = FeedForward(d_model)
self.dropout_1 = nn.Dropout(dropout)
self.dropout_2 = nn.Dropout(dropout)
def forward(self, x, mask):
x2 = self.norm_1(x)
x = x + self.dropout_1(self.attn(x2,x2,x2,mask))
x2 = self.norm_2(x)
x = x + self.dropout_2(self.ff(x2))
return x
```
## DecoderLayer
```
# build a decoder layer with two multi-head attention layers and one feed-forward layer
class DecoderLayer(nn.Module):
def __init__(self, d_model, heads, dropout=0.1):
super().__init__()
self.norm_1 = Norm(d_model)
self.norm_2 = Norm(d_model)
self.norm_3 = Norm(d_model)
self.dropout_1 = nn.Dropout(dropout)
self.dropout_2 = nn.Dropout(dropout)
self.dropout_3 = nn.Dropout(dropout)
self.attn_1 = MultiHeadAttention(heads, d_model)
self.attn_2 = MultiHeadAttention(heads, d_model)
self.ff = FeedForward(d_model).cuda()
def forward(self, x, e_outputs, src_mask, trg_mask):
x2 = self.norm_1(x)
x = x + self.dropout_1(self.attn_1(x2, x2, x2, trg_mask))
x2 = self.norm_2(x)
x = x + self.dropout_2(self.attn_2(x2, e_outputs, e_outputs, src_mask))
x2 = self.norm_3(x)
x = x + self.dropout_3(self.ff(x2))
return x
```
We can then build a convenient cloning function that can generate multiple layers:
```
import copy
def get_clones(module, N):
return nn.ModuleList([copy.deepcopy(module) for i in range(N)])
```
## Encoder
```
class Encoder(nn.Module):
def __init__(self, vocab_size, d_model, N, heads):
super().__init__()
self.N = N
self.embed = Embedder(vocab_size, d_model)
self.pe = PositionalEncoder(d_model)
self.layers = get_clones(EncoderLayer(d_model, heads), N)
self.norm = Norm(d_model)
def forward(self, src, mask):
x = self.embed(src)
x = self.pe(x)
for i in range(N):
x = self.layers[i](x, mask)
return self.norm(x)
```
## Decoder
```
class Decoder(nn.Module):
def __init__(self, vocab_size, d_model, N, heads):
super().__init__()
self.N = N
self.embed = Embedder(vocab_size, d_model)
self.pe = PositionalEncoder(d_model)
self.layers = get_clones(DecoderLayer(d_model, heads), N)
self.norm = Norm(d_model)
def forward(self, trg, e_outputs, src_mask, trg_mask):
x = self.embed(trg)
x = self.pe(x)
for i in range(self.N):
x = self.layers[i](x, e_outputs, src_mask, trg_mask)
return self.norm(x)
```
## Transformer
```
class Transformer(nn.Module):
def __init__(self, src_vocab, trg_vocab, d_model, N, heads):
super().__init__()
self.encoder = Encoder(src_vocab, d_model, N, heads)
self.decoder = Decoder(trg_vocab, d_model, N, heads)
self.out = nn.Linear(d_model, trg_vocab)
def forward(self, src, trg, src_mask, trg_mask):
e_outputs = self.encoder(src, src_mask)
d_output = self.decoder(trg, e_outputs, src_mask, trg_mask)
output = self.out(d_output)
return output
```
**Note**: We don't perform softmax on the output as this will be handled automatically by our loss function.
# Training the Model
With the transformer built, all that remains is to train on the dataset. The coding part is done, but be prepared to wait for about 2 days for this model to start converging! However, in this session, we only perform minimal epoch to train the model and you may try to use more epoch during your self-study.
Let’s define some parameters first:
```
embedding_dimension = 512
heads = 4
N = 6
src_vocab = len(EN_TEXT.vocab)
trg_vocab = len(FR_TEXT.vocab)
model = Transformer(src_vocab, trg_vocab, embedding_dimension, N, heads)
if device.type == 'cuda':
model.cuda()
for p in model.parameters():
if p.dim() > 1:
nn.init.xavier_uniform_(p)
# This code is very important! It initialises the parameters with a
# range of values that stops the signal fading or getting too big.
optim = torch.optim.Adam(model.parameters(), lr=0.0001, betas=(0.9, 0.98), eps=1e-9)
```
Now we’re good to train the transformer model
```
import time
import torch
def train_model(epochs, print_every=100, timelimit=None):
model.train()
start = time.time()
temp = start
total_loss = 0
min_loss = float('inf')
for epoch in range(epochs):
for i, batch in enumerate(train_iter):
src = batch.English.transpose(0,1)
trg = batch.French.transpose(0,1)
# the French sentence we input has all words except
# the last, as it is using each word to predict the next
trg_input = trg[:, :-1]
# the words we are trying to predict
targets = trg[:, 1:].contiguous().view(-1)
# create function to make masks using mask code above
src_mask, trg_mask = create_masks(src, trg_input)
preds = model(src, trg_input, src_mask, trg_mask)
ys = trg[:, 1:].contiguous().view(-1)
optim.zero_grad()
loss = F.cross_entropy(preds.view(-1, preds.size(-1)), ys, ignore_index=target_pad)
loss.backward()
optim.step()
total_loss += loss.data.item()
if (i + 1) % print_every == 0:
loss_avg = total_loss / print_every
duration = (time.time() - start) // 60
print("time = %dm, epoch %d, iter = %d, loss = %.3f, %ds per %d iters" %
(duration, epoch + 1, i + 1, loss_avg, time.time() - temp, print_every))
if loss_avg < min_loss:
min_loss = loss_avg
torch.save(model, "model/training.model")
print("Current best model saved", "loss =", loss_avg)
if (timelimit and duration >= timelimit):
break
total_loss = 0
temp = time.time()
# train_model(1, timelimit=300)
torch.load("model/pretrained.model")
```
# Testing the Model
We can use the below function to translate sentences. We can feed it sentences directly from our batches, or input custom strings.
The translator works by running a loop. We start off by encoding the English sentence. We then feed the decoder the `<sos>` token index and the encoder outputs. The decoder makes a prediction for the first word, and we add this to our decoder input with the sos token. We rerun the loop, getting the next prediction and adding this to the decoder input, until we reach the `<eos>` token letting us know it has finished translating.
```
def translate(model, src, max_len = 80, custom_string=False):
model.eval()
if custom_string == True:
src = tokenize_en(src)
src = Variable(torch.LongTensor([[EN_TEXT.vocab.stoi[tok] for tok in src]])).cuda()
src_mask = (src != input_pad).unsqueeze(-2)
e_outputs = model.encoder(src, src_mask)
outputs = torch.zeros(max_len).type_as(src.data)
outputs[0] = torch.LongTensor([FR_TEXT.vocab.stoi['<sos>']])
for i in range(1, max_len):
trg_mask = np.triu(np.ones((1, i, i)), k=1).astype('uint8')
trg_mask = Variable(torch.from_numpy(trg_mask) == 0).cuda()
out = model.out(model.decoder(outputs[:i].unsqueeze(0), e_outputs, src_mask, trg_mask))
out = F.softmax(out, dim=-1)
val, ix = out[:, -1].data.topk(1)
outputs[i] = ix[0][0]
if ix[0][0] == FR_TEXT.vocab.stoi['<eos>']:
break
return ' '.join([FR_TEXT.vocab.itos[ix] for ix in outputs[:i]])
translate(model, "How're you my friend?", custom_string=True)
```
# Contributors
**Author**
<br>Chee Lam
# References
1. [How to Code The Transformer in Pytorch](https://towardsdatascience.com/how-to-code-the-transformer-in-pytorch-24db27c8f9ec#b0ed)
2. [How to Use TorchText for Neural Machine Translation](https://towardsdatascience.com/how-to-use-torchtext-for-neural-machine-translation-plus-hack-to-make-it-5x-faster-77f3884d95)
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.