code stringlengths 2.5k 150k | kind stringclasses 1 value |
|---|---|
```
from selenium import webdriver
from selenium.webdriver.common.desired_capabilities import DesiredCapabilities
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as EC
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.common.by import By
from selenium.common.exceptions import TimeoutException
import cv2
import pickle
import dlib
import face_recognition as fr
import matplotlib.pyplot as plt
import re
import glob
import json
from urllib.request import urlopen
import time
import os
```
3 ~ 17번까지 한국 연예인
# 연예인 이름 목록
### 목록 크롤링
```
driver = webdriver.Chrome()
name_list_url = 'https://namu.wiki/w/%EC%97%B0%EC%98%88%EC%9D%B8/%EB%B3%B8%EB%AA%85%20%EB%B0%8F%20%EC%98%88%EB%AA%85'
driver.get(name_list_url)
def preprocess(name):
name = re.sub('\(.+\)', '', name)
name = name.strip()
return name
name_list = []
error_list = []
for i in range(3,18):
class_ = driver.find_elements_by_class_name('wiki-heading-content')[i]
raw_name_list = class_.find_elements_by_css_selector('ul.wiki-list li')
for raw_name in raw_name_list:
split_name = raw_name.text.split('→')
stage_name = preprocess(split_name[0])
try:
real_name = re.search('[가-힣]+', split_name[1])[0].strip()
except Exception as e:
error_list.append(raw_name.text)
continue
new_name = stage_name + ' ' + real_name
name_list.append(new_name)
print(f'{len(name_list)} persons returned')
print(f'{len(error_list)} persons got error')
print(error_list)
```
### 전처리
오타로 인해 다른 이름과 다르게 표시가 된 이름들 수정
```
stage_name = preprocess(error_list[0].split('-')[0])
real_name = re.search('[가-힣]+', error_list[0].split('-')[1])[0]
print(stage_name, real_name)
name_list.append(stage_name + ' ' + real_name)
stage_name = preprocess(error_list[1].split('→')[0])
real_name = re.search('[가-힣A-Za-z ]+', error_list[1].split('→')[1])[0].strip()
print(stage_name, real_name)
name_list.append(stage_name + ' ' + real_name)
stage_name = preprocess(error_list[2].split('→')[0])
real_name = re.search('[가-힣A-Za-z ]+', error_list[2].split('→')[-1])[0].strip()
print(stage_name, real_name)
name_list.append(stage_name + ' ' + real_name)
```
### 목록 이름 저장
```
name_list = name_list[6:]
with open('celebrity/celebrity_name.pkl', 'wb') as f:
pickle.dump(name_list, f)
with open('celebrity/celebrity_name.pkl', 'rb') as f:
celebrity_name_list = pickle.load(f)
len(celebrity_name_list)
```
# 얼굴 크롤링 간 조건
다양한 얼굴 중 임베딩에 가장 적합한 이미지를 크롤링하기위해 조건 설정
- 얼굴 탐지 및 Embedding이 가능한 이미지
- 썬글라스 착용 시 제외
- 마스크 착용 시 제외
- 얼굴 각도가 좌,우로 돌아간 경우 제외
- 이미지 크기가 340x340보다 작으면 제외
- 제외되는 경우가 너무 많아서 크기 수정
### 썬글라스 탐지
눈 주위의 픽셀값을 이용해 썬글라스 착용 여부 탐지
- 왼쪽 : Landmark 36 ~ 41
- 오른쪽 : Landmakr 42 ~ 47
- luminance를 이용해 밝기 측정
```
example = fr.load_image_file('celebrity/GRAY 이성화.jpg')
example_show = example.copy()
landmarks = fr.face_landmarks(example)
landmarks = []
for k, v in fr.face_landmarks(example)[0].items():
landmarks.extend(v)
for number, landmark in enumerate(landmarks):
cv2.circle(example_show, landmark, 2, (0, 255, 255), -1)
plt.imshow(example_show)
def get_brightness_around_eye(image):
try:
landmarks = []
for k, v in fr.face_landmarks(image)[0].items():
landmarks.extend(v)
left1, right1 = landmarks[36][0], landmarks[39][0]
top1, bottom1 = landmarks[37][1], landmarks[41][1]
left2, right2 = landmarks[42][0], landmarks[45][0]
top2, bottom2 = landmarks[43][1], landmarks[47][1]
image[top1:bottom1, left1:right1] = np.nan
image[top2:bottom2, left2:right2] = np.nan
left_glass = image[top1-10:bottom1+10, left1-5:right1+5]
right_glass = image[top2-10:bottom2+10, left2-5:right2+5]
luminance_left = np.nanmean(0.2126*left_glass[:,:,0] + 0.7152*left_glass[:,:,1] + 0.0722*left_glass[:,:,2])
luminance_right = np.nanmean(0.2126*right_glass[:,:,0] + 0.7152*right_glass[:,:,1] + 0.0722*right_glass[:,:,2])
#luminance_left2 = np.nanmean(left_glass[:,:,0] + left_glass[:,:,1] + left_glass[:,:,2])
#luminance_right2 = np.nanmean(right_glass[:,:,0] + right_glass[:,:,1] + right_glass[:,:,2])
return luminance_left, luminance_right
except:
return None
```
### 마스크 착용 탐지
다양한 얼굴 중 마스크 착용 시 제외
- 입 주변의 픽셀값을 이용해 마스크 착용 여부 탐지
- Lanmark 2, 6, 11, 14 사용
```
# 랜드마크 2, 6, 11, 14
def get_brightness_around_mouse(image):
landmarks = []
for k, v in fr.face_landmarks(image)[0].items():
landmarks.extend(v)
left1, right1 = landmarks[6][0], landmarks[11][0]
top1, bottom1 = landmarks[2][1], landmarks[6][1]
mask = image[top1:bottom1, left1:right1]
mask_luminance = np.nanmean(0.2126*mask[:,:,0] + 0.7152*mask[:,:,1] + 0.0722*mask[:,:,2])
return mask_luminance
```
### 얼굴 각도 측정
얼굴이 좌,우로 돌아간 정도를 측정해 정면이 아닐 시 제외
- 왼쪽과 오른쪽 볼의 길이차를 이용해 회전 정도 측정
- Landmark 2, 14, 30 사용
```
def ratio_of_face_rotate(image):
landmarks = []
for k, v in fr.face_landmarks(image)[0].items():
landmarks.extend(v)
left = np.linalg.norm(np.array(landmarks[30]) - np.array(landmarks[2]))
right = np.linalg.norm(np.array(landmarks[30]) - np.array(landmarks[14]))
ratio = min(left, right) / max(left, right)
return ratio
```
### 이미지 사이즈 조건
행 : 404.16
열 : 340.0
# 이미지 크롤링
크롤링 간 사용하는 유용함 함수는 따로 정의
### 이미지 저장
사이트로부터 얻은 이미지의 URL을 이용해 이미지 다운로드
```
def save_image_from_url(image_url):
with urlopen(image_url.get_attribute('src')) as f:
with open(f'celebrity/{name}.jpg', 'wb') as file_name:
img = f.read()
file_name.write(img)
return img, file_name
```
### 이미지 다운로드
```
delay=2
error_name = []
driver = webdriver.Chrome()
for name in name_list:
driver.get(f'https://search.naver.com/search.naver?where=image&sm=tab_jum&query={name}')
time.sleep(0.2)
try:
myElem = WebDriverWait(driver, delay).until(EC.presence_of_element_located((By.TAG_NAME, 'section')))
driver.find_element_by_tag_name('html').send_keys(Keys.END)
image_url_list = driver.find_elements_by_css_selector('div.tile_item._item img._image._listImage')[:50]
is_succeed = 0
for image_url in image_url_list:
img, file_name = save_image_from_url(image_url)
error_message = ''
try:
image = fr.load_image_file(file_name.name)
if (image.shape[0] < 200) or (image.shape[1] < 200):
print(name, 'Too Small')
continue
locations = fr.face_locations(image)
if len(locations) != 1:
print(name, 'No Face')
continue
top, right, bottom, left = locations[0]
face_cropped = image[top:bottom, left:right]
if (face_cropped.shape[0] < 30) or (face_cropped.shape[1] < 30):
print(name, 'Cropped Too Small')
continue
face_embedding = fr.face_encodings(face_cropped)
if len(face_embedding) != 1:
print(name, 'No Embedding')
continue
ratio = ratio_of_face_rotate(image)
if ratio < 0.85:
print(name, 'Rotated')
continue
left_eye, right_eye = get_brightness_around_eye(image)
if (left_eye < 60) & (right_eye < 60):
print(name, 'Maybe Sunglasses')
continue
mask = get_brightness_around_mouse(image)
if mask > 220:
print(name, 'Maybe Mask?')
is_succeed = 1
print(name, 'Succeed')
break
except Exception as ex:
print(name, 'failed detection', ex)
continue
if is_succeed == 0:
os.remove(file_name.name)
error_name.append(name)
print(name, 'collecting Failed')
except Exception as ex:
print(name, ex)
error_name.append(name)
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
x = np.array([[0, 1, 2], [0, 1, 2]])
y = np.array([[0, 0, 0], [1, 1, 1]])
plt.plot(x, y, color='red', marker='.', linestyle='')
plt.grid(True)
plt.show()
import numpy as np
import matplotlib.pyplot as plt
x = np.linspace(0, 100, 100)
y = np.linspace(0, 100, 100)
X, Y = np.meshgrid(x, y)
plt.plot(X, Y, color='red', marker='.', linestyle='')
plt.grid(True)
plt.show()
import numpy as np
import matplotlib.pyplot as plt
x = np.arange(1, 10, 0.02)
y = np.arange(1, 10, 0.02)
X, Y = np.meshgrid(x, y)
plt.plot(X, Y, color='red')
plt.grid(True)
plt.show()
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
import numpy as np
def plot_decision_regions(X_train, y_train, X_test, y_test, classifier):
X = np.concatenate((X_train, X_test))
y = np.concatenate((y_train, y_test))
# 随机选择所有的
markers = ('s', 'x', 'o', '^', 'v')
# 随机选择所有的
colors = ('red', 'blue', 'lightgreen', 'gray', 'cyan')
# 每个类别的颜色还有marker
cmap = ListedColormap(colors[:len(np.unique(y))])
x1_min, x1_max = X[:, 0].min() - 1, X[:, 0].max() + 1
x2_min, x2_max = X[:, 1].min() - 1, X[:, 1].max() + 1
XX, YY = np.meshgrid(np.arange(x1_min, x1_max, 0.02),
np.arange(x2_min, x2_max, 0.02))
z = classifier.predict(np.array([XX.ravel(), YY.ravel()]).T)
Z = z.reshape(XX.shape)
plt.contourf(XX, YY, Z, alpha=0.3, cmap=cmap)
plt.xlim(XX.min(), XX.max())
plt.ylim(YY.min(), YY.max())
for i, c in enumerate(np.unique(y_train)):
plt.scatter(x=X_train[y_train==c, 0],
y=X_train[y_train==c, 1],
alpha=0.8,
c=colors[i],
marker=markers[i],
label=c,
edgecolor='black')
plt.scatter(X_test[:, 0],
X_test[:, 1],
c='',
edgecolor='black',
alpha=1.0,
linewidths=1,
marker='o',
s=100,
label='test set')
plt.xlabel('petal lenght [standardized]')
plt.ylabel('petal width [standardized]')
plt.legend(loc='upper left')
plt.tight_layout()
plt.show()
import pandas as pd
df = pd.read_csv('./wine.data', header=None)
X, y = df.iloc[:, 1:3].values, df.iloc[:, 0].values
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, stratify=y)
print(X_train.shape, X_test.shape, y_train.shape, y_test.shape)
from sklearn.neighbors import KNeighborsClassifier
knc = KNeighborsClassifier()
knc.fit(X_train, y_train)
plot_decision_regions(X_train, y_train, X_test, y_test, knc)
from sklearn import datasets
iris = datasets.load_iris()
X = iris.data[:, [2, 3]]
y = iris.target
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, stratify=y)
from sklearn.preprocessing import StandardScaler
ss = StandardScaler()
ss.fit(X_train)
X_train_std = ss.transform(X_train)
X_test_std = ss.transform(X_test)
from sklearn.neighbors import KNeighborsClassifier
knc = KNeighborsClassifier()
knc.fit(X_train_std, y_train)
print('score: ', knc.score(X_test_std, y_test))
plot_decision_regions(X_train_std, y_train, X_test_std, y_test, knc)
```
| github_jupyter |
# Using SageMaker Neo to Compile a Tensorflow U-Net Model
[SageMaker Neo](https://aws.amazon.com/sagemaker/neo/) makes it easy to compile pre-trained TensorFlow models and build an inference optimized container without the need for any custom model serving or inference code.
<img src="https://paperswithcode.com/media/methods/Screen_Shot_2020-07-07_at_9.08.00_PM_rpNArED.png" align="center" style="padding: 8px;width:500px;">
[U-Net](https://paperswithcode.com/method/u-net) is an architecture for semantic segmentation. It's a popular model for biological images including Ultrasound, Microscopy, CT, MRI and more.
In this example, we will show how deploy a pre-trained U-Net model to a SageMaker Endpoint with Neo compilation using the [SageMaker Python SDK](https://github.com/aws/sagemaker-python-sdk), and then use the models to perform inference requests. We also provide a performance comparison so you can see the benefits of model compilation.
## Setup
First, we need to ensure we have SageMaker Python SDK 1.x and Tensorflow 1.15.x. Then, import necessary Python packages.
```
!pip install -U --quiet --upgrade "sagemaker"
!pip install -U --quiet "tensorflow==1.15.3"
import tarfile
import numpy as np
import sagemaker
import time
from sagemaker.utils import name_from_base
```
Next, we'll get the IAM execution role and a few other SageMaker specific variables from our notebook environment, so that SageMaker can access resources in your AWS account later in the example.
```
from sagemaker import get_execution_role
from sagemaker.session import Session
role = get_execution_role()
sess = Session()
region = sess.boto_region_name
bucket = sess.default_bucket()
```
SageMaker [Neo supports Tensorflow 1.15.x](https://docs.amazonaws.cn/en_us/sagemaker/latest/dg/neo-supported-cloud.html). Check your version of Tensorflow to prevent downstream framework errors.
```
import tensorflow as tf
print(tf.__version__) # This notebook runs on TensorFlow 1.15.x or earlier
```
## Download U-Net Model
The SageMaker Neo TensorFlow Serving Container works with any model stored in TensorFlow's [SavedModel format](https://www.tensorflow.org/guide/saved_model). This could be the output of your own training job or a model trained elsewhere. For this example, we will use a pre-trained version of the U-Net model based on this [repo](https://github.com/kamalkraj/DATA-SCIENCE-BOWL-2018).
```
model_name = 'unet_medical'
export_path = 'export'
model_archive_name = 'unet-medical.tar.gz'
model_archive_url = 'https://sagemaker-neo-artifacts.s3.us-east-2.amazonaws.com/{}'.format(model_archive_name)
!wget {model_archive_url}
```
The pre-trained model and its artifacts are saved in a compressed tar file (.tar.gz) so unzip first with:
```
!tar -xvzf unet-medical.tar.gz
```
After downloading the model, we can inspect it using TensorFlow's ``saved_model_cli`` command. In the command output, you should see
```
MetaGraphDef with tag-set: 'serve' contains the following SignatureDefs:
signature_def['serving_default']:
...
```
The command output should also show details of the model inputs and outputs.
```
import os
model_path = os.path.join(export_path, 'Servo/1')
!saved_model_cli show --all --dir {model_path}
```
Next we need to create a model archive file containing the exported model.
## Upload the model archive file to S3
We now have a suitable model archive ready in our notebook. We need to upload it to S3 before we can create a SageMaker Model that. We'll use the SageMaker Python SDK to handle the upload.
```
model_data = Session().upload_data(path=model_archive_name, key_prefix='model')
print('model uploaded to: {}'.format(model_data))
```
## Create a SageMaker Model and Endpoint
Now that the model archive is in S3, we can create an unoptimized Model and deploy it to an
Endpoint.
```
from sagemaker.tensorflow.serving import Model
instance_type = 'ml.c4.xlarge'
framework = "TENSORFLOW"
framework_version = "1.15.3"
sm_model = Model(model_data=model_data, framework_version=framework_version,role=role)
uncompiled_predictor = sm_model.deploy(initial_instance_count=1, instance_type=instance_type)
```
## Make predictions using the endpoint
The endpoint is now up and running, and ready to handle inference requests. The `deploy` call above returned a `predictor` object. The `predict` method of this object handles sending requests to the endpoint. It also automatically handles JSON serialization of our input arguments, and JSON deserialization of the prediction results.
We'll use this sample image:
<img src="https://sagemaker-neo-artifacts.s3.us-east-2.amazonaws.com/cell-4.png" align="left" style="padding: 8px;">
```
sample_img_fname = 'cell-4.png'
sample_img_url = 'https://sagemaker-neo-artifacts.s3.us-east-2.amazonaws.com/{}'.format(sample_img_fname)
!wget {sample_img_url}
# read the image file into a tensor (numpy array)
import cv2
image = cv2.imread(sample_img_fname)
original_shape = image.shape
import matplotlib.pyplot as plt
plt.imshow(image, cmap='gray', interpolation='none')
plt.show()
image = np.resize(image, (256, 256, 3))
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = np.asarray(image)
image = np.expand_dims(image, axis=0)
start_time = time.time()
# get a prediction from the endpoint
# the image input is automatically converted to a JSON request.
# the JSON response from the endpoint is returned as a python dict
result = uncompiled_predictor.predict(image)
print("Prediction took %.2f seconds" % (time.time() - start_time))
# show the predicted segmentation image
cutoff = 0.4
segmentation_img = np.squeeze(np.asarray(result['predictions'])) > cutoff
segmentation_img = segmentation_img.astype(np.uint8)
segmentation_img = np.resize(segmentation_img, (original_shape[0], original_shape[1]))
plt.imshow(segmentation_img, "gray")
plt.show()
```
## Uncompiled Predictor Performance
```
shape_input = np.random.rand(1, 256, 256, 3)
uncompiled_results = []
for _ in range(100):
start = time.time()
uncompiled_predictor.predict(image)
uncompiled_results.append((time.time() - start) * 1000)
print("\nPredictions for un-compiled model: \n")
print('\nP95: ' + str(np.percentile(uncompiled_results, 95)) + ' ms\n')
print('P90: ' + str(np.percentile(uncompiled_results, 90)) + ' ms\n')
print('P50: ' + str(np.percentile(uncompiled_results, 50)) + ' ms\n')
print('Average: ' + str(np.average(uncompiled_results)) + ' ms\n')
```
## Compile model using SageMaker Neo
```
# Replace the value of data_shape below and
# specify the name & shape of the expected inputs for your trained model in JSON
# Note that -1 is replaced with 1 for the batch size placeholder
data_shape = {'inputs':[1, 224, 224, 3]}
instance_family = 'ml_c4'
compilation_job_name = name_from_base('medical-tf-Neo')
# output path for compiled model artifact
compiled_model_path = 's3://{}/{}/output'.format(bucket, compilation_job_name)
optimized_estimator = sm_model.compile(target_instance_family=instance_family,
input_shape=data_shape,
job_name=compilation_job_name,
role=role,
framework=framework.lower(),
framework_version=framework_version,
output_path=compiled_model_path
)
```
## Create Optimized Endpoint
```
optimized_predictor = optimized_estimator.deploy(initial_instance_count = 1, instance_type = instance_type)
start_time = time.time()
# get a prediction from the endpoint
# the image input is automatically converted to a JSON request.
# the JSON response from the endpoint is returned as a python dict
result = optimized_predictor.predict(image)
print("Prediction took %.2f seconds" % (time.time() - start_time))
```
## Compiled Predictor Performance
```
compiled_results = []
test_input = {"instances": np.asarray(shape_input).tolist()}
#Warmup inference.
optimized_predictor.predict(image)
# Inferencing 100 times.
for _ in range(100):
start = time.time()
optimized_predictor.predict(image)
compiled_results.append((time.time() - start) * 1000)
print("\nPredictions for compiled model: \n")
print('\nP95: ' + str(np.percentile(compiled_results, 95)) + ' ms\n')
print('P90: ' + str(np.percentile(compiled_results, 90)) + ' ms\n')
print('P50: ' + str(np.percentile(compiled_results, 50)) + ' ms\n')
print('Average: ' + str(np.average(compiled_results)) + ' ms\n')
```
## Performance Comparison
Here we compare inference speed up provided by SageMaker Neo. P90 is 90th percentile latency. We add this because it represents the tail of the latency distribution (worst case). More information on latency percentiles [here](https://blog.bramp.net/post/2018/01/16/measuring-percentile-latency/).
```
p90 = np.percentile(uncompiled_results, 90) / np.percentile(compiled_results, 90)
p50 = np.percentile(uncompiled_results, 50) / np.percentile(compiled_results, 50)
avg = np.average(uncompiled_results) / np.average(compiled_results)
print("P90 Speedup: %.2f" % p90)
print("P50 Speedup: %.2f" % p50)
print("Average Speedup: %.2f" % avg)
```
## Additional Information
## Cleaning up
To avoid incurring charges to your AWS account for the resources used in this tutorial, you need to delete the SageMaker Endpoint.
```
uncompiled_predictor.delete_endpoint()
optimized_predictor.delete_endpoint()
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib as mpl
%matplotlib inline
plt.style.use("ggplot")
plt.rcParams["font.size"] = 13
mpl.rcParams["font.family"] = "Osaka"
np.random.seed(seed=1)
```
# Data Preprocessing
```
# read and rename data
df = pd.read_csv("AggDemand.csv", header = None)
df.columns = ["price", "quantity","weight", "horsepower", "airconditioner", "ownership"]
df.head()
# define variables
p = np.log(df["price"].values)
q = df["quantity"].values
num = len(df)
# explanatory variables
x = np.stack([np.ones(num),
np.log(df["weight"].values),
np.log(df["horsepower"].values),
df["airconditioner"].values],
axis = 1)
# Define market share and num of consumers
MS = 50000000 # total market
ns = 1000 # num of consumers
s0 = (MS - np.sum(q))/MS # market share of outside option
sj = q/MS # market share of each goods
# Berry's inversion
# see slides
delta = np.log(sj) - np.log(s0) # \delta =y = X'B
```
# OLS
```
# analytical solution
%timeit np.dot(np.dot(np.linalg.inv(np.dot(x.T, x)), x.T), delta)
np.dot(np.dot(np.linalg.inv(np.dot(x.T, x)), x.T), delta)
# Another solution method provides same solution and faster
%timeit np.linalg.solve(np.dot(x.T, x), np.dot(x.T, delta))
np.linalg.solve(np.dot(x.T, x), np.dot(x.T, delta))
```
# BLP
```
# initial guess
delta_init = np.log(sj) - np.log(s0)
# coefficients
sigma =np.array((0.5, 0.5))
nu = np.random.randn(ns,2)
nu = nu * sigma # For vectorization
mu_mat = np.dot(x[:,1:3], nu.T)
delta_mat = np.tile(delta_init, (ns, 1)).T
delta_mat.shape
exp_util = np.exp(delta_mat + mu_mat)
denom = np.sum(exp_util, axis = 0) + 1
choiceprob = exp_util/denom
PredMS = (1/ns) * np.sum(choiceprob, axis= 1)
delta_update = delta_init + np.log(sj) - np.log(PredMS)
# iteration settings
max_iter = 100000
tol = 1e-6
# Contraction mapping
for i in range(max_iter):
delta_init = delta_update
delta_mat = np.tile(delta_init, (ns, 1)).T
exp_util = np.exp(delta_mat + mu_mat)
denom = np.sum(exp_util, axis = 0) + 1
choiceprob = exp_util/denom
dist = delta_update - delta_init
PredMS = (1/ns) * np.sum(choiceprob, axis= 1)
delta_update = delta_init + np.log(sj) - np.log(PredMS)
err = (delta_update - delta_init)
err2 = np.dot(err, err)
if err2 < tol:
break
if i == max_iter -1:
print("Iteration doesn't converge...")
```
# Check solution
```
ans = np.stack([PredMS,
sj], axis=1)
pd.DataFrame(ans).head()
```
| github_jupyter |
# Example of Comparing All Implemented Outlier Detection Models
**[PyOD](https://github.com/yzhao062/pyod)** is a comprehensive **Python toolkit** to **identify outlying objects** in
multivariate data with both unsupervised and supervised approaches.
The model covered in this example includes:
1. Linear Models for Outlier Detection:
1. **PCA: Principal Component Analysis** use the sum of
weighted projected distances to the eigenvector hyperplane
as the outlier outlier scores)
2. **MCD: Minimum Covariance Determinant** (use the mahalanobis distances
as the outlier scores)
3. **OCSVM: One-Class Support Vector Machines**
2. Proximity-Based Outlier Detection Models:
1. **LOF: Local Outlier Factor**
2. **CBLOF: Clustering-Based Local Outlier Factor**
3. **kNN: k Nearest Neighbors** (use the distance to the kth nearest
neighbor as the outlier score)
4. **Median kNN** Outlier Detection (use the median distance to k nearest
neighbors as the outlier score)
5. **HBOS: Histogram-based Outlier Score**
3. Probabilistic Models for Outlier Detection:
1. **ABOD: Angle-Based Outlier Detection**
4. Outlier Ensembles and Combination Frameworks
1. **Isolation Forest**
2. **Feature Bagging**
3. **LSCP**
Corresponding file could be found at /examples/compare_all_models.py
```
from __future__ import division
from __future__ import print_function
import os
import sys
from time import time
# temporary solution for relative imports in case pyod is not installed
# if pyod is installed, no need to use the following line
sys.path.append(
os.path.abspath(os.path.join(os.path.dirname("__file__"), '..')))
import numpy as np
from numpy import percentile
import matplotlib.pyplot as plt
import matplotlib.font_manager
# Import all models
from pyod.models.abod import ABOD
from pyod.models.cblof import CBLOF
from pyod.models.feature_bagging import FeatureBagging
from pyod.models.hbos import HBOS
from pyod.models.iforest import IForest
from pyod.models.knn import KNN
from pyod.models.lof import LOF
from pyod.models.mcd import MCD
from pyod.models.ocsvm import OCSVM
from pyod.models.pca import PCA
from pyod.models.lscp import LSCP
# Define the number of inliers and outliers
n_samples = 200
outliers_fraction = 0.25
clusters_separation = [0]
# Compare given detectors under given settings
# Initialize the data
xx, yy = np.meshgrid(np.linspace(-7, 7, 100), np.linspace(-7, 7, 100))
n_inliers = int((1. - outliers_fraction) * n_samples)
n_outliers = int(outliers_fraction * n_samples)
ground_truth = np.zeros(n_samples, dtype=int)
ground_truth[-n_outliers:] = 1
# initialize a set of detectors for LSCP
detector_list = [LOF(n_neighbors=5), LOF(n_neighbors=10), LOF(n_neighbors=15),
LOF(n_neighbors=20), LOF(n_neighbors=25), LOF(n_neighbors=30),
LOF(n_neighbors=35), LOF(n_neighbors=40), LOF(n_neighbors=45),
LOF(n_neighbors=50)]
# Show the statics of the data
print('Number of inliers: %i' % n_inliers)
print('Number of outliers: %i' % n_outliers)
print('Ground truth shape is {shape}. Outlier are 1 and inliers are 0.\n'.format(shape=ground_truth.shape))
print(ground_truth)
random_state = np.random.RandomState(42)
# Define nine outlier detection tools to be compared
classifiers = {
'Angle-based Outlier Detector (ABOD)':
ABOD(contamination=outliers_fraction),
'Cluster-based Local Outlier Factor (CBLOF)':
CBLOF(contamination=outliers_fraction,
check_estimator=False, random_state=random_state),
'Feature Bagging':
FeatureBagging(LOF(n_neighbors=35),
contamination=outliers_fraction,
random_state=random_state),
'Histogram-base Outlier Detection (HBOS)': HBOS(
contamination=outliers_fraction),
'Isolation Forest': IForest(contamination=outliers_fraction,
random_state=random_state),
'K Nearest Neighbors (KNN)': KNN(
contamination=outliers_fraction),
'Average KNN': KNN(method='mean',
contamination=outliers_fraction),
'Local Outlier Factor (LOF)':
LOF(n_neighbors=35, contamination=outliers_fraction),
'Minimum Covariance Determinant (MCD)': MCD(
contamination=outliers_fraction, random_state=random_state),
'One-class SVM (OCSVM)': OCSVM(contamination=outliers_fraction),
'Principal Component Analysis (PCA)': PCA(
contamination=outliers_fraction, random_state=random_state),
'Locally Selective Combination (LSCP)': LSCP(
detector_list, contamination=outliers_fraction,
random_state=random_state)
}
# Show all detectors
for i, clf in enumerate(classifiers.keys()):
print('Model', i + 1, clf)
# Fit the models with the generated data and
# compare model performances
for i, offset in enumerate(clusters_separation):
np.random.seed(42)
# Data generation
X1 = 0.3 * np.random.randn(n_inliers // 2, 2) - offset
X2 = 0.3 * np.random.randn(n_inliers // 2, 2) + offset
X = np.r_[X1, X2]
# Add outliers
X = np.r_[X, np.random.uniform(low=-6, high=6, size=(n_outliers, 2))]
# Fit the model
plt.figure(figsize=(15, 12))
for i, (clf_name, clf) in enumerate(classifiers.items()):
print(i + 1, 'fitting', clf_name)
# fit the data and tag outliers
clf.fit(X)
scores_pred = clf.decision_function(X) * -1
y_pred = clf.predict(X)
threshold = percentile(scores_pred, 100 * outliers_fraction)
n_errors = (y_pred != ground_truth).sum()
# plot the levels lines and the points
Z = clf.decision_function(np.c_[xx.ravel(), yy.ravel()]) * -1
Z = Z.reshape(xx.shape)
subplot = plt.subplot(3, 4, i + 1)
subplot.contourf(xx, yy, Z, levels=np.linspace(Z.min(), threshold, 7),
cmap=plt.cm.Blues_r)
a = subplot.contour(xx, yy, Z, levels=[threshold],
linewidths=2, colors='red')
subplot.contourf(xx, yy, Z, levels=[threshold, Z.max()],
colors='orange')
b = subplot.scatter(X[:-n_outliers, 0], X[:-n_outliers, 1], c='white',
s=20, edgecolor='k')
c = subplot.scatter(X[-n_outliers:, 0], X[-n_outliers:, 1], c='black',
s=20, edgecolor='k')
subplot.axis('tight')
subplot.legend(
[a.collections[0], b, c],
['learned decision function', 'true inliers', 'true outliers'],
prop=matplotlib.font_manager.FontProperties(size=10),
loc='lower right')
subplot.set_xlabel("%d. %s (errors: %d)" % (i + 1, clf_name, n_errors))
subplot.set_xlim((-7, 7))
subplot.set_ylim((-7, 7))
plt.subplots_adjust(0.04, 0.1, 0.96, 0.94, 0.1, 0.26)
plt.suptitle("Outlier detection")
plt.show()
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/JavaScripts/Image/Polynomial.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/JavaScripts/Image/Polynomial.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/JavaScripts/Image/Polynomial.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as emap
except:
import geemap as emap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Satellite`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py#L13) can be added using the `Map.add_basemap()` function.
```
Map = emap.Map(center=[40,-100], zoom=4)
Map.add_basemap('ROADMAP') # Add Google Map
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
# Applies a non-linear contrast enhancement to a MODIS image using
# function -0.2 + 2.4x - 1.2x^2.
# Load a MODIS image and apply the scaling factor.
img = ee.Image('MODIS/006/MOD09GA/2012_03_09') \
.select(['sur_refl_b01', 'sur_refl_b04', 'sur_refl_b03']) \
.multiply(0.0001)
# Apply the polynomial enhancement.
adj = img.polynomial([-0.2, 2.4, -1.2])
Map.setCenter(-107.24304, 35.78663, 8)
Map.addLayer(img, {'min': 0, 'max': 1}, 'original')
Map.addLayer(adj, {'min': 0, 'max': 1}, 'adjusted')
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
```
#NOTE: This must be the first call in order to work properly!
from deoldify import device
from deoldify.device_id import DeviceId
#choices: CPU, GPU0...GPU7
device.set(device=DeviceId.GPU0)
from deoldify.visualize import *
plt.style.use('dark_background')
import warnings
warnings.filterwarnings("ignore", category=UserWarning, message=".*?Your .*? set is empty.*?")
colorizer = get_video_colorizer()
```
# Instructions
### source_url
Type in a url hosting a video from YouTube, Imgur, Twitter, Reddit, Vimeo, etc. Many sources work! GIFs also work. Full list here: https://ytdl-org.github.io/youtube-dl/supportedsites.html NOTE: If you want to use your own video, you can set source_url to None and just upload the file to video/source/ in Jupyter. Just make sure that the file_name parameter matches the file you uploaded.
### file_name
Name this whatever sensible file name you want (minus extension)! It should actually exist in video/source if source_url=None
### render_factor
The default value of 21 has been carefully chosen and should work -ok- for most scenarios (but probably won't be the -best-). This determines resolution at which the color portion of the video is rendered. Lower resolution will render faster, and colors also tend to look more vibrant. Older and lower quality film in particular will generally benefit by lowering the render factor. Higher render factors are often better for higher quality videos and inconsistencies (flashy render) will generally be reduced, but the colors may get slightly washed out.
### file_name_ext
There's no reason to changes this.
### result_path
Ditto- don't change.
### How to Download a Copy
Simply shift+right click on the displayed video and click "Save video as..."!
## Pro Tips
1. If a video takes a long time to render and you're wondering how well the frames will actually be colorized, you can preview how well the frames will be rendered at each render_factor by using the code at the bottom. Just stop the video rendering by hitting the stop button on the cell, then run that bottom cell under "See how well render_factor values perform on a frame here". It's not perfect and you may still need to experiment a bit especially when it comes to figuring out how to reduce frame inconsistency. But it'll go a long way in narrowing down what actually works.
## Troubleshooting
The video player may wind up not showing up, in which case- make sure to wait for the Jupyter cell to complete processing first (the play button will stop spinning). Then follow these alternative download instructions
1. In the menu to the left, click Home icon.
2. By default, rendered video will be in /video/result/
If a video you downloaded doesn't play, it's probably because the cell didn't complete processing and the video is in a half-finished state.
If you get a 'CUDA out of memory' error, you probably have the render_factor too high. The max is 44 on 11GB video cards.
## Colorize!!
```
#NOTE: Max is 44 with 11GB video cards. 21 is a good default
render_factor=21
#NOTE: Make source_url None to just read from file at ./video/source/[file_name] directly without modification
source_url='https://twitter.com/silentmoviegifs/status/1116751583386034176'
file_name = 'DogShy1926'
file_name_ext = file_name + '.mp4'
result_path = None
if source_url is not None:
result_path = colorizer.colorize_from_url(source_url, file_name_ext, render_factor=render_factor)
else:
result_path = colorizer.colorize_from_file_name(file_name_ext, render_factor=render_factor)
show_video_in_notebook(result_path)
```
## See how well render_factor values perform on a frame here
```
for i in range(10,45,2):
colorizer.vis.plot_transformed_image('video/bwframes/' + file_name + '/00001.jpg', render_factor=i, display_render_factor=True, figsize=(8,8))
```
| github_jupyter |
# Unit 5 - Financial Planning
```
# Initial imports
import os
import requests
import pandas as pd
from dotenv import load_dotenv
import alpaca_trade_api as tradeapi
from MCForecastTools import MCSimulation
import json
%matplotlib inline
# Load .env enviroment variables
load_dotenv()
True
```
## Part 1 - Personal Finance Planner
### Collect Crypto Prices Using the `requests` Library
```
# Set current amount of crypto assets
my_btc = 1.2
my_eth = 5.3
# Crypto API URLs
btc_url = "https://api.alternative.me/v2/ticker/Bitcoin/?convert=CAD"
eth_url = "https://api.alternative.me/v2/ticker/Ethereum/?convert=CAD"
# Fetch current BTC price
btc_price = requests.get(btc_url)
btc_price = btc_price.json()["data"]["1"]["quotes"]["USD"]["price"]
# Fetch current ETH price
eth_price = requests.get(eth_url)
eth_price = eth_price.json()["data"]["1027"]["quotes"]["CAD"]["price"]
# Compute current value of my crpto
my_btc_value =btc_price * my_btc
my_eth_value = eth_price * my_eth
# Print current crypto wallet balance
print(f"The current value of your {my_btc} BTC is ${my_btc_value:0.2f}")
print(f"The current value of your {my_eth} ETH is ${my_eth_value:0.2f}")
```
### Collect Investments Data Using Alpaca: `SPY` (stocks) and `AGG` (bonds)
```
# Set current amount of shares
my_agg = 200
my_spy = 50
# Set Alpaca API key and secret
alpaca_api_key= os.getenv("ALPACA_API_KEY")
alpaca_secret_key= os.getenv("ALPACA_SECRET_KEY")
# Create the Alpaca API object
api = tradeapi.REST(
alpaca_api_key,
alpaca_secret_key,
api_version="v2"
)
# Format current date as ISO format
current_date = pd.Timestamp("2020-05-01", tz="America/New_York").isoformat()
# Set the tickers
tickers = ["AGG", "SPY"]
# Set timeframe to '1D' for Alpaca API
timeframe = "1D"
# Get current closing prices for SPY and AGG
# (use a limit=1000 parameter to call the most recent 1000 days of data)
current_date = api.get_barset(
tickers,
timeframe,
start = current_date,
limit = 1000,
).df
# Preview DataFrame
current_date.head()
# Pick AGG and SPY close prices
agg_close_price = current_date["AGG"]["close"][0]
spy_close_price = current_date["SPY"]["close"][0]
# Print AGG and SPY close prices
print(f"Current AGG closing price: ${agg_close_price}")
print(f"Current SPY closing price: ${spy_close_price}")
# Compute the current value of shares
agg_share_value = agg_close_price * my_agg
spy_share_value = spy_close_price * my_spy
# Print current value of shares
print(f"The current value of your {my_spy} SPY shares is ${spy_share_value:0.2f}")
print(f"The current value of your {my_agg} AGG shares is ${agg_share_value:0.2f}")
```
### Savings Health Analysis
```
# Set monthly household income
monthly_income = 12000
# Consolidate financial assets data
crypto_price = btc_price + eth_price
stock_price = agg_share_value + spy_share_value
data = [crypto_price, stock_price]
# Create savings DataFrame
savings_df = pd.DataFrame(data, columns= ["Amount"], index= ["Shares", "Crypto"])
savings_df
# Display savings DataFrame
display(savings_df)
# Plot savings pie chart
savings_df.plot.pie(y="Amount", title= "Personal Savings")
# Set ideal emergency fund
emergency_fund = monthly_income * 3
# Calculate total amount of savings
total_savings = savings_df.sum()
total_savings = total_savings["Amount"]
savings_goal = total_savings - emergency_fund
# Validate saving health
if total_savings > emergency_fund:
print("Congratulations! You have enough money in your emergency fund")
elif total_savings == emergency_fund:
print("Congratulations on reaching your financial goal!")
else:
print(f"You are ${total_savings-emergency_fund} from reaching your goal")
```
## Part 2 - Retirement Planning
### Monte Carlo Simulation
```
# Set start and end dates of five years back from today.
# Sample results may vary from the solution based on the time frame chosen
start_date = pd.Timestamp('2016-05-01', tz='America/New_York').isoformat()
end_date = pd.Timestamp('2021-05-01', tz='America/New_York').isoformat()
# Get 5 years' worth of historical data for SPY and AGG
# (use a limit=1000 parameter to call the most recent 1000 days of data)
df_stock_data = api.get_barset(
tickers,
timeframe,
start = start_date,
end = end_date,
limit = 1000
).df.dropna()
# Display sample data
df_stock_data.head()
# Configuring a Monte Carlo simulation to forecast 30 years cumulative returns
MC_forecast = MCSimulation(
portfolio_data = df_stock_data,
num_simulation = 500,
num_trading_days = 252*30
)
# Printing the simulation input data
MC_forecast.portfolio_data.head()
# Running a Monte Carlo simulation to forecast 30 years cumulative returns
MC_forecast.calc_cumulative_return()
# Plot simulation outcomes
plot = MC_forecast.plot_simulation()
# Plot probability distribution and confidence intervals
plot = MC_forecast.plot_distribution()
```
### Retirement Analysis
```
# Fetch summary statistics from the Monte Carlo simulation results
summary_stats = MC_forecast.summarize_cumulative_return()
# Print summary statistics
print(summary_stats)
```
### Calculate the expected portfolio return at the `95%` lower and upper confidence intervals based on a `$20,000` initial investment.
```
# Set initial investment
initial_investment = 20000
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $20,000
ci_lower = round(summary_stats[8]*initial_investment,2)
ci_upper = round(summary_stats[9]*initial_investment,2)
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 30 years will end within in the range of"
f" ${ci_lower} and ${ci_upper}")
```
### Calculate the expected portfolio return at the `95%` lower and upper confidence intervals based on a `50%` increase in the initial investment.
```
# Set initial investment
initial_investment = 20000 * 1.5
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $30,000
ci_lower = round(summary_stats[8]*initial_investment*1.5,2)
ci_upper = round(summary_stats[9]*initial_investment*1.5,2)
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 30 years will end within in the range of"
f" ${ci_lower} and ${ci_upper}")
```
## Optional Challenge - Early Retirement
### Five Years Retirement Option
```
# Configuring a Monte Carlo simulation to forecast 5 years cumulative returns
MC_forecast5 = MCSimulation(
portfolio_data = df_stock_data,
num_simulation = 500,
num_trading_days = 252*5
)
# Running a Monte Carlo simulation to forecast 5 years cumulative returns
MC_forecast5.calc_cumulative_return()
# Plot simulation outcomes
plot5 = MC_forecast5.plot_simulation()
# Plot probability distribution and confidence intervals
plot5 = MC_forecast5.plot_distribution()
# Fetch summary statistics from the Monte Carlo simulation results
summary_stats5 = MC_forecast5.summarize_cumulative_return()
# Print summary statistics
print(summary_stats5)
# Set initial investment
initial_investment = 10000
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $60,000
ci_lower_five = round(summary_stats5[8]*initial_investment,2)
ci_upper_five = round(summary_stats5[9]*initial_investment,2)
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 5 years will end within in the range of"
f" ${ci_lower_five} and ${ci_upper_five}")
```
### Ten Years Retirement Option
```
# Configuring a Monte Carlo simulation to forecast 10 years cumulative returns
MC_forecast10 = MCSimulation(
portfolio_data = df_stock_data,
num_simulation = 500,
num_trading_days = 252*10
)
# Running a Monte Carlo simulation to forecast 10 years cumulative returns
MC_forecast10.calc_cumulative_return()
# Plot simulation outcomes
plot10 = MC_forecast10.plot_simulation()
# Plot probability distribution and confidence intervals
plot10 = MC_forecast10.plot_distribution()
# Fetch summary statistics from the Monte Carlo simulation results
summary_stats10 = MC_forecast10.summarize_cumulative_return()
# Print summary statistics
print(summary_stats10)
# Set initial investment
initial_investment = 30000
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $60,000
ci_lower_ten = round(summary_stats10[8]*initial_investment,2)
ci_upper_ten = round(summary_stats10[9]*initial_investment,2)
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 10 years will end within in the range of"
f" ${ci_lower_ten} and ${ci_upper_ten}")
```
| github_jupyter |
```
%matplotlib inline
%load_ext autoreload
%autoreload 2
import numpy as np
import matplotlib.pyplot as plt
import pylab
pylab.rcParams['figure.figsize'] = (10.0, 8.0)
from torch.utils.data import DataLoader
import torch
import os
import torchvision.transforms as transforms
from faster_rcnn.utils.datasets.voc.voc import VOCDetection
from faster_rcnn.utils.datasets.adapter import convert_data
from faster_rcnn.utils.display.images import imshow, result_show
from faster_rcnn.utils.datasets.merge import VOCMerge
root = '/data/data'
ds = VOCMerge(root, 'train', dataset_name='tmp')
val_ds = VOCMerge(root, 'val', dataset_name='tmp')
print(len(val_ds))
batch_size = 1
train_data_loader = DataLoader(ds, batch_size=batch_size, shuffle=True, collate_fn=convert_data, num_workers=0, drop_last=True)
val_data_loader = DataLoader(val_ds, batch_size=batch_size, collate_fn=convert_data, num_workers=0, drop_last=True)
categories = ds.classes
print(len(categories))
from faster_rcnn.faster_rcnn import FastRCNN
net = FastRCNN(categories, debug=False)
net.load_state_dict(torch.load('./checkpoints/faster_model.pkl'))
import random, string
def randomword(length):
letters = string.ascii_lowercase
return ''.join(random.choice(letters) for i in range(length))
with torch.no_grad():
for i, data in enumerate(val_data_loader):
file_name = randomword(10)
print file_name + ".txt"
if not data:
continue
batch_tensor, im_info, batch_boxes, batch_boxes_index, _ = data
pred_boxes, scores, classes, rois, im_data = net.detect_blob(batch_tensor, im_info, 0.2)
for k in range(batch_tensor.shape[0]):
with open(os.path.join('./evaluate/ground-truth/', file_name + ".txt"), "w") as f:
for box in batch_boxes:
f.write("%s %d %d %d %d\n" % (categories[int(box[4])] , box[0],box[1],box[2],box[3]))
with open(os.path.join('./evaluate/predicted/', file_name + ".txt"), "w") as f:
for box in zip(classes, scores, pred_boxes):
f.write("%s %.2f %d %d %d %d\n" % (box[0], box[1], box[2][0], box[2][1],box[2][2],box[2][3]))
# imshow(batch_tensor[k], predict_boxes=batch_boxes[ np.where(batch_boxes_index == k )])
# result_show(im_data[0], pred_boxes, classes, scores)
with torch.no_grad():
result = net.detect("./test_im/146.jpg", thr=0.7)
if result:
pred_boxes, scores, classes, rois, im_data = result
if(len(pred_boxes)):
result_show(im_data[0], pred_boxes, classes, scores)
```
| github_jupyter |
# Instructions
Implement a PyTorch dataset for keypoint detection.
Read about custom datasets here:
* https://jdhao.github.io/2017/10/23/pytorch-load-data-and-make-batch/
Image augmentation is an important part of deep learning pipelines. It artificially increases your training sample by generating transformed versions of images.
<img src="static/imgaug.jpg" alt="Drawing" style="width: 600px;"/>
You can read about it here:
* https://blog.keras.io/building-powerful-image-classification-models-using-very-little-data.html
* https://github.com/aleju/imgaug
You should implement the following augmentations:
* randomly fliping left and right
* randomly fliping up and down
* randomly translating by up to 4 pixels
* randomly rotating the image by 180 degrees
* randomly scaling the image from 1.0 to 1.5
Apart from reading images and augmenting, the loader is also cropping the input image by using outputs of the localizer network (bounding box coordinates).
# Your Solution
Your solution function should be called solution. In this case we leave it for consistency but you don't need to do anything with it.
CONFIG is a dictionary with all parameters that you want to pass to your solution function.
```
def solution():
return DatasetAligner
class DatasetAligner(Dataset):
def __init__(self, X, y, crop_coordinates, img_dirpath, augmentation, target_size, bins_nr):
super().__init__()
self.X = X.reset_index(drop=True)
self.y = y.reset_index(drop=True)
self.crop_coordinates = crop_coordinates
self.img_dirpath = img_dirpath
self.target_size = target_size
self.bins_nr = bins_nr
self.augmentation = augmentation
def load_image(self, img_name):
"""
Read image from disk to numpy array
"""
return img_array
def __len__(self):
"""
Determine the length of the dataset
"""
return length
def __getitem__(self, index):
"""
This method should take the image filepath at X[index] and targets at y[index] and
preprocess them. Use your aligner_preprocessing function.
Xi_tensor: is a torch.FloatTensor for image
yi_tensors: is a torch.LongTensor for targets it's shape should be 1 x k where k is the number of outputs
"""
return Xi_tensor, yi_tensors
def aligner_preprocessing(img, target, crop_coordinates, augmentation, *, org_size, target_size, bins_nr):
"""
Run augmentations and transformations on image and target
"""
processed_image, processed_target = crop_image_and_adjust_target(img, target, crop_coordinates)
if augmentation:
"""
Run augmentations on Image (and target if needed)
"""
"""
Transform coordinates to bin numbers as explained below and normalize the image
"""
processed_target = bin_quantizer(processed_target, (height, width), bins_nr)
processed_image = normalize_img(processed_image)
return processed_image, processed_target
def crop_image_and_adjust_target(img, target, crop_coordinates):
"""
crop image by using localization network predictions.
Remember to adjust the keypoint positions to the cropped image
"""
return cropped_image, adjusted_target
def bin_quantizer(coordinates, shape, bins_nr):
"""
Quantize the height and width and transform coordinates to bin numbers
"""
return binned_coordinates
def normalize_img(img):
mean = [0.28201905, 0.37246801, 0.42341868]
std = [0.13609867, 0.12380088, 0.13325344]
"""
Normalize Image
"""
return normalized_img
```
| github_jupyter |
```
import os
path_parent = os.path.dirname(os.getcwd())
os.chdir(path_parent)
from ggmodel_dev import GraphModel
GAS_nodes = {
'GASCAPACITY': {
'type': 'input',
'name': 'Gas cumulative capacity',
'unit':'GW',
},
'GASCAPFACTOR': {
'type': 'input',
'name': 'Gas capacity factor',
'unit':'%'
},
'GASHOURS':{
'type': 'input',
'name': 'Gas generation time',
'unit': 'hours'
},
'GASEFFICIENCY':{
'type': 'input',
'name': 'Gas conversion efficiency',
'unit':'1'
},
'GASFUEL':{
'type': 'variable',
'name': 'Fuel required for gas generation',
'unit': 'TWh',
'computation': lambda GASCAPACITY, GASCAPFACTOR, GASHOURS, GASEFFICIENCY, **kwargs: GASCAPACITY * GASCAPFACTOR * GASHOURS / GASEFFICIENCY
},
'GASEMFACTORi':{
'type': 'input',
'name': 'Gas emission factors',
'unit': 'Mt CO2eq / TWh'
},
'GASEMISSIONSi':{
'type': 'variable',
'name': 'C02, CH4, NO2 emissions',
'unit': 'Mt CO2eq',
'computation': lambda GASEMFACTORi, GASFUEL, **kwargs: GASEMFACTORi * GASFUEL
},
'GASLANDREQ':{
'type': 'input',
'name': 'land requirement per GW of gas plant',
'unit': 'ha / GW'
},
'GASLAND':{
'type': 'variable',
'name': 'Gas land requirement',
'unit': 'ha',
'computation': lambda GASLANDREQ, GASFUEL, **kwargs: GASLANDREQ * GASCAPACITY
}
}
GraphModel(GAS_nodes).draw()
COAL_nodes = {
'COALCAPACITY':{
'type': 'input',
'name': 'Coal cumulative capacity',
'unit':'GW',
},
'COALGENERATION':{
'type': 'variable',
'name': 'Coal generation',
'unit': 'TWh',
'computation': lambda COALCAPTGENERATION, COALUTILGENERATION, OWNUSEFRAC, **kwargs: COALCAPTGENERATION + COALUTILGENERATION * (1 - OWNUSEFRAC)
},
'OWNUSEFRAC':{
'type': 'input',
'name': 'Fraction of energy for own use',
'unit': '1',
},
'COALEFF':{
'type': 'input',
'name': 'Coal thermal conversion efficiency',
'unit': '1'
},
'COALFUEL':{
'type': 'variable',
'name': 'Fuel required for coal generation',
'unit': 'TWh',
'computation': lambda COALEFF, COALGENERATION, **kwargs: COALEFF * COALGENERATION
},
'COALEMISSIONSi':{
'type': 'variable',
'name': 'C02, CH4, NO2 emissions',
'unit': 'Mt CO2eq',
'computation': lambda COALFUEL, COALEMFACTORi, **kwargs: COALFUEL * COALEMFACTORi
},
'COALLANDREQ':{
'type': 'input',
'name': 'land requirement per GW of coal plant',
'unit': 'ha / GW'
},
'COALEMFACTORi':{
'type': 'input',
'name': 'Gas emission factors',
'unit': 'Mt CO2eq / TWh'
},
'COALLAND':{
'type': 'variable',
'name': 'Coal plant land requirement',
'unit': 'ha',
'computation': lambda COALCAPACITY, COALLANDREQ, **kwargs: COALCAPACITY * COALLANDREQ
},
'COALUTILCAPFACTOR':{
'type': 'input',
'name': 'Coal plant capacity factor utility',
'unit': '1',
},
'COALCAPTCAPFACTOR':{
'type': 'input',
'name': 'Coal plant capacity factor captive',
'unit': '1',
},
'COALHOURS':{
'type': 'input',
'name': 'Coal generation time',
'unit': 'hours'
},
'COALUTILGENERATION':{
'type': 'variable',
'name': 'Coal generation for utilities',
'unit': 'TWh',
'computation': lambda COALHOURS, COALUTILCAPFACTOR, COALCAPACITY, **kwargs: COALHOURS * COALUTILCAPFACTOR * COALCAPACITY
},
'COALCAPTGENERATION':{
'type': 'variable',
'name': 'Coal generation for captive',
'unit': 'TWh',
'computation': lambda COALHOURS, COALCAPTCAPFACTOR, COALCAPACITY, **kwargs: COALHOURS * COALCAPTCAPFACTOR * COALCAPACITY
},
}
GraphModel(COAL_nodes).draw()
SOLAR_nodes = {
'SOLARCAPACITY':{
'type': 'input',
'name': 'Solar cumulative capacity',
'unit':'GW',
},
'SOLARCAPFACTOR': {
'type': 'input',
'name': 'Solar capacity factor',
'unit':'%'
},
'SOLARHOURS':{
'type': 'input',
'name': 'Solar generation time',
'unit':'hours'
},
'SOLARGENERATION':{
'type': 'variable',
'name': 'Solar generation',
'unit': 'TWh',
'computation': lambda SOLARHOURS, SOLARCAPFACTOR, SOLARCAPACITY, **kwargs: SOLARHOURS * SOLARCAPFACTOR * SOLARCAPACITY
},
'SOLARLANDREQ':{
'type': 'input',
'name': 'land requirement per GW of solar plant',
'unit': 'ha / GW'
},
'SOLARLAND':{
'type': 'variable',
'name': 'Solar PV land requirement',
'unit': 'ha',
'computation': lambda SOLARCAPACITY, SOLARLANDREQ, **kwargs: SOLARCAPACITY * SOLARLANDREQ
}
}
GraphModel(SOLAR_nodes).draw()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import yfinance as yf
from fbprophet import Prophet
from fbprophet.plot import add_changepoints_to_plot
import multiprocessing as mp
from datetime import date,timedelta
import time as t
import matplotlib.pyplot as plt
import defs
from sklearn.metrics import mean_squared_error, r2_score, mean_absolute_error
%matplotlib inline
plt.rcParams['figure.figsize']=(20,10)
plt.style.use('ggplot')
market_dfset = {}
dfset = {}
modelset = {}
futureset = {}
forecastset = {}
figureset = {}
legendset = {}
tickers = []
manager = mp.Manager()
tickers = manager.list()
#get file object
f = open("list", "r")
while(True):
#read next line
line = f.readline()
#if line is empty, you are done with all lines in the file
if not line:
break
#you can access the line
tickers.append(line.strip())
#close file
f.close()
market_dfset=manager.dict()
modelset=manager.dict()
forecastset=manager.dict()
p = {}
for ticker in tickers:
p[ticker]= mp.Process(target=defs.run_prophet,args=(tickers,ticker,market_dfset,modelset,forecastset))
l = len(tickers)
c = mp.cpu_count()
for i in range(0, l, c):
for j in range(0,c):
if (i+j<l):
p[tickers[i+j]].start()
for j in range(0,c):
if (i+j<l): p[tickers[i+j]].join()
for ticker in tickers:
figureset[ticker] = market_dfset[ticker]["Close"].plot()
legendset[ticker]=figureset[ticker].legend() #get the legend
legendset[ticker].get_texts()[0].set_text(ticker) #change the legend text
plt.show()
for ticker in tickers:
print(ticker)
print(forecastset[ticker][['ds', 'yhat', 'yhat_lower', 'yhat_upper']].tail(n=30))
figure=modelset[ticker].plot(forecastset[ticker])
fig = modelset[ticker].plot(forecastset[ticker])
a = add_changepoints_to_plot(fig.gca(), modelset[ticker], forecastset[ticker])
figure2=modelset[ticker].plot_components(forecastset[ticker])
plt.show()
print('\n')
```
# Plotting the forecast
```
two_yearset = {}
```
With the data that we have, it is hard to see how good/bad the forecast (blue line) is compared to the actual data (black dots). Let's take a look at the last 800 data points (~2 years) of forecast vs actual without looking at the future forecast (because we are just interested in getting a visual of the error between actual vs forecast).
```
for ticker in tickers:
two_yearset[ticker] = forecastset[ticker].set_index('ds').join(market_dfset[ticker])
two_yearset[ticker] = two_yearset[ticker][['Close', 'yhat', 'yhat_upper', 'yhat_lower' ]].dropna().tail(800)
two_yearset[ticker]['yhat']=np.exp(two_yearset[ticker].yhat)
two_yearset[ticker]['yhat_upper']=np.exp(two_yearset[ticker].yhat_upper)
two_yearset[ticker]['yhat_lower']=np.exp(two_yearset[ticker].yhat_lower)
two_yearset[ticker].tail()
figureset[ticker]=two_yearset[ticker][['Close', 'yhat']].plot()
figureset[ticker].plot(two_yearset[ticker].yhat_upper, color='black', linestyle=':', alpha=0.5)
figureset[ticker].plot(two_yearset[ticker].yhat_lower, color='black', linestyle=':', alpha=0.5)
figureset[ticker].set_title('Actual (Orange) vs Forecasted Upper & Lower Confidence (Black)')
figureset[ticker].set_ylabel('Price')
figureset[ticker].set_xlabel('Date')
legendset[ticker]=figureset[ticker].legend() #get the legend
legendset[ticker].get_texts()[0].set_text(ticker) #change the legend text
plt.show()
two_years_AE_set = {}
for ticker in tickers:
two_years_AE_set[ticker] = (two_yearset[ticker].yhat - two_yearset[ticker].Close)
print(ticker)
print(two_years_AE_set[ticker].describe())
print("R2 score: ",r2_score(two_yearset[ticker].Close, two_yearset[ticker].yhat))
print("MSE score: ",mean_squared_error(two_yearset[ticker].Close, two_yearset[ticker].yhat))
print("MAE score: ",mean_absolute_error(two_yearset[ticker].Close, two_yearset[ticker].yhat))
print('\n')
full_dfset = {}
for ticker in tickers:
full_dfset[ticker] = forecastset[ticker].set_index('ds').join(market_dfset[ticker])
full_dfset[ticker]['yhat']=np.exp(full_dfset[ticker]['yhat'])
for ticker in tickers:
fig, ax1 = plt.subplots()
n=365
ax1.plot(full_dfset[ticker].tail(n).Close)
ax1.plot(full_dfset[ticker].tail(n).yhat, color='black', linestyle=':')
ax1.fill_between(full_dfset[ticker].tail(n).index, np.exp(full_dfset[ticker]['yhat_upper'].tail(n)), np.exp(full_dfset[ticker]['yhat_lower'].tail(n)), alpha=0.5, color='darkgray')
ax1.set_title(ticker)
ax1.set_ylabel('Price')
ax1.set_xlabel('Date')
plt.show()
n=365 + 15
d=30
fig2, ax2 = plt.subplots()
ax2.plot(full_dfset[ticker][full_dfset[ticker].last_valid_index()-pd.DateOffset(n, 'D'):full_dfset[ticker].last_valid_index()-pd.DateOffset(n-d, 'D')].Close)
ax2.plot(full_dfset[ticker][full_dfset[ticker].last_valid_index()-pd.DateOffset(n, 'D'):full_dfset[ticker].last_valid_index()-pd.DateOffset(n-d, 'D')].yhat, color='black', linestyle=':')
ax2.fill_between(full_dfset[ticker][full_dfset[ticker].last_valid_index()-pd.DateOffset(n, 'D'):full_dfset[ticker].last_valid_index()-pd.DateOffset(n-d, 'D')].index, np.exp(full_dfset[ticker][full_dfset[ticker].last_valid_index()-pd.DateOffset(n, 'D'):full_dfset[ticker].last_valid_index()-pd.DateOffset(n-d, 'D')]['yhat_upper']), np.exp(full_dfset[ticker][full_dfset[ticker].last_valid_index()-pd.DateOffset(n, 'D'):full_dfset[ticker].last_valid_index()-pd.DateOffset(n-d, 'D')]['yhat_lower']), alpha=0.5, color='darkgray')
ax2.set_title(ticker)
ax2.set_ylabel('Price')
ax2.set_xlabel('Date')
pd.merge(full_dfset[ticker]['yhat'].tail(n=365), pd.merge(np.exp(full_dfset[ticker]['yhat_upper'].tail(n=365)), np.exp(full_dfset[ticker]['yhat_lower'].tail(n=365)),on='ds'),on='ds').to_csv(ticker+'-365Days.csv')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/ausafahmed08/Anonymous/blob/main/titanic_train.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from google.colab import files
file = files.upload()
df = pd.read_csv('train.csv')
print(df.shape)
df.head(5)
df.dropna().sum()
df1 = df.drop(['PassengerId','Name','Ticket','Cabin','Embarked'], axis = 1)
df1.head()
(df1['Sex'])
sex_set = set(df1['Sex'])
sex_set
sibsp_set = set(df1['SibSp'])
sibsp_set
parch_set = set(df1['Parch'])
parch_set
df.info()
df1['Sex'] = df1['Sex'].map({'female': 0, 'male': 1}).astype(int)
print(df1.head)
df
data = df.dropna()
data
df1
data1 = df1.dropna()
data1
# returns all rows from index 1(which is pclass)
X = data1.iloc[:,1:]
X
X = data1.iloc[:,1:].values
X
Y = data1.iloc[:,0]
Y
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size = 0.25, random_state = 0) #only test_size taken as the train_size will adjust accordingly
print(X_train.shape)
print(X_test.shape)
print(X_test)
print(X_train)
print(y_train)
print(y_test)
```
Logistic Regression
```
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
print(X_train)
print(X_test)
X_trainf = sc.fit(X_train)
print(X_trainf)
fit_t = X_trainf.transform(X_train)
print(fit_t)
X_testf = sc.fit(X_test)
print(X_testf)
from sklearn.linear_model import LogisticRegression
model = LogisticRegression(random_state = 0 , max_iter =100 , verbose =0) #max iterations
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
plt.figure(figsize=(18,4));
plt.scatter(data['Age'] , data['Survived'] , color = 'green' , marker ='+')
plt.xlabel("Age")
from sklearn.metrics import accuracy_score
print("Accuracy of Model is {0}%".format(accuracy_score(y_test,y_pred)*100))
```
Support Vector Machine
```
from sklearn import svm
model = svm.SVC(kernel='linear' ,probability= True) #prob set true because it gives each class a score.
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
from sklearn.metrics import accuracy_score
print("Accuracy of Model is {0}%".format(accuracy_score(y_test,y_pred)*100))
from sklearn import svm
model1 = svm.SVC(kernel='linear')
model2 = svm.SVC(kernel='rbf')
model3 = svm.SVC(gamma = 0.001)
model4 = svm.SVC(gamma = 0.001,C = 1)
model1.fit(X_train,y_train)
model2.fit(X_train,y_train)
model3.fit(X_train,y_train)
model4.fit(X_train,y_train)
y_predModel1 = model1.predict(X_test)
y_predModel2 = model2.predict(X_test)
y_predModel3 = model3.predict(X_test)
y_predModel4 = model4.predict(X_test)
print("Accuracy of the Model 1: {0}%".format(accuracy_score(y_test, y_predModel1)*100))
print("Accuracy of the Model 2: {0}%".format(accuracy_score(y_test, y_predModel2)*100))
print("Accuracy of the Model 3: {0}%".format(accuracy_score(y_test, y_predModel3)*100))
print("Accuracy of the Model 4: {0}%".format(accuracy_score(y_test, y_predModel4)*100))
```
Random Forest
```
n_estimators = [int(x) for x in np.linspace(start = 10, stop = 100, num = 10)]
n_estimators
n_estimators = [int(x) for x in np.linspace(start = 10, stop = 100, num = 10)]
max_features = ['auto', 'sqrt']
max_depth = [int(x) for x in np.linspace(start = 1, stop = 10, num =2)]
min_samples_split = [int(x) for x in np.linspace(start = 1, stop = 10, num = 2)]
min_samples_leaf = [int(x) for x in np.linspace(start = 1, stop = 5, num = 2)]
bootstrap = [True, False]
param_grid = {'n_estimators': n_estimators,
'max_features': max_features,
'max_depth': max_depth,
'min_samples_split': min_samples_split,
'min_samples_leaf': min_samples_leaf,
'bootstrap': bootstrap}
print(param_grid)
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
rf_Model = RandomForestClassifier(random_state=0)
rf_Grid = GridSearchCV(estimator = rf_Model, param_grid = param_grid, cv = 3, verbose=2, n_jobs = 4)
rf_Grid.fit(X_train, y_train)
rf_Grid.best_params_
gc = RandomForestClassifier(random_state = 0, bootstrap = True , max_depth=10 , max_features= 'auto' , min_samples_leaf= 1, min_samples_split= 10, n_estimators= 90)
gc.fit(X_train , y_train)
pred = gc.predict(X_test)
acc = accuracy_score(y_test , pred)
acc
y_pred = rf_Grid.predict(X_test)
from sklearn.metrics import accuracy_score
print("Accuracy of the Model: {0}%".format(accuracy_score(y_test, y_pred)*100))
```
Ensemble methods (BAGGING)
```
from sklearn.ensemble import BaggingClassifier
from sklearn.neighbors import KNeighborsClassifier
bagging = BaggingClassifier(KNeighborsClassifier(), random_state=0, max_samples=0.5, max_features=0.5)
bagging.fit(X_train, y_train)
pred2 = bagging.predict(X_test)
from sklearn.metrics import accuracy_score
acc2 = accuracy_score(y_test , pred2)
acc2
```
Boosting
```
#from sklearn.datasets import make_hastie_10_2
from sklearn.ensemble import GradientBoostingClassifier
clf = GradientBoostingClassifier(n_estimators=100, learning_rate=1.0, max_depth=1, random_state=0)
clf.fit(X_train, y_train)
#clf.score(X_test, y_test)
pred3 = clf.predict(X_test)
from sklearn.metrics import accuracy_score
acc3 = accuracy_score(y_test , pred3)
acc3
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Keras overview
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/guide/keras/overview"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/keras/overview.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/keras/overview.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/keras/overview.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
This guide gives you the basics to get started with Keras. It's a 10-minute read.
## Import tf.keras
`tf.keras` is TensorFlow's implementation of the
[Keras API specification](https://keras.io). This is a high-level
API to build and train models that includes first-class support for
TensorFlow-specific functionality, such as [eager execution](../eager.ipynb),
`tf.data` pipelines, and [Estimators](../estimator.ipynb).
`tf.keras` makes TensorFlow easier to use without sacrificing flexibility and
performance.
To get started, import `tf.keras` as part of your TensorFlow program setup:
```
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
from tensorflow import keras
```
`tf.keras` can run any Keras-compatible code, but keep in mind:
* The `tf.keras` version in the latest TensorFlow release might not be the same
as the latest `keras` version from PyPI. Check `tf.keras.__version__`.
* When [saving a model's weights](./save_and_serialize.ipynb), `tf.keras` defaults to the
[checkpoint format](../checkpoint.ipynb). Pass `save_format='h5'` to
use HDF5 (or pass a filename that ends in `.h5`).
## Build a simple model
### Sequential model
In Keras, you assemble *layers* to build *models*. A model is (usually) a graph
of layers. The most common type of model is a stack of layers: the
`tf.keras.Sequential` model.
To build a simple, fully-connected network (i.e. multi-layer perceptron):
```
from tensorflow.keras import layers
model = tf.keras.Sequential()
# Adds a densely-connected layer with 64 units to the model:
model.add(layers.Dense(64, activation='relu'))
# Add another:
model.add(layers.Dense(64, activation='relu'))
# Add a softmax layer with 10 output units:
model.add(layers.Dense(10, activation='softmax'))
```
You can find a complete, short example of how to use Sequential models [here](https://www.tensorflow.org/tutorials/quickstart/beginner).
To learn about building more advanced models than Sequential models, see:
- [Guide to the Keras Functional API](./functional.ipynb)
- [Guide to writing layers and models from scratch with subclassing](./custom_layers_and_models.ipynb)
### Configure the layers
There are many `tf.keras.layers` available. Most of them share some common constructor
arguments:
* `activation`: Set the activation function for the layer. This parameter is
specified by the name of a built-in function or as a callable object. By
default, no activation is applied.
* `kernel_initializer` and `bias_initializer`: The initialization schemes
that create the layer's weights (kernel and bias). This parameter is a name or
a callable object. This defaults to the `"Glorot uniform"` initializer.
* `kernel_regularizer` and `bias_regularizer`: The regularization schemes
that apply the layer's weights (kernel and bias), such as L1 or L2
regularization. By default, no regularization is applied.
The following instantiates `tf.keras.layers.Dense` layers using constructor
arguments:
```
# Create a sigmoid layer:
layers.Dense(64, activation='sigmoid')
# Or:
layers.Dense(64, activation=tf.keras.activations.sigmoid)
# A linear layer with L1 regularization of factor 0.01 applied to the kernel matrix:
layers.Dense(64, kernel_regularizer=tf.keras.regularizers.l1(0.01))
# A linear layer with L2 regularization of factor 0.01 applied to the bias vector:
layers.Dense(64, bias_regularizer=tf.keras.regularizers.l2(0.01))
# A linear layer with a kernel initialized to a random orthogonal matrix:
layers.Dense(64, kernel_initializer='orthogonal')
# A linear layer with a bias vector initialized to 2.0s:
layers.Dense(64, bias_initializer=tf.keras.initializers.Constant(2.0))
```
## Train and evaluate
### Set up training
After the model is constructed, configure its learning process by calling the
`compile` method:
```
model = tf.keras.Sequential([
# Adds a densely-connected layer with 64 units to the model:
layers.Dense(64, activation='relu', input_shape=(32,)),
# Add another:
layers.Dense(64, activation='relu'),
# Add a softmax layer with 10 output units:
layers.Dense(10, activation='softmax')])
model.compile(optimizer=tf.keras.optimizers.Adam(0.01),
loss='categorical_crossentropy',
metrics=['accuracy'])
```
`tf.keras.Model.compile` takes three important arguments:
* `optimizer`: This object specifies the training procedure. Pass it optimizer
instances from the `tf.keras.optimizers` module, such as
`tf.keras.optimizers.Adam` or
`tf.keras.optimizers.SGD`. If you just want to use the default parameters, you can also specify optimizers via strings, such as `'adam'` or `'sgd'`.
* `loss`: The function to minimize during optimization. Common choices include
mean square error (`mse`), `categorical_crossentropy`, and
`binary_crossentropy`. Loss functions are specified by name or by
passing a callable object from the `tf.keras.losses` module.
* `metrics`: Used to monitor training. These are string names or callables from
the `tf.keras.metrics` module.
* Additionally, to make sure the model trains and evaluates eagerly, you can make sure to pass `run_eagerly=True` as a parameter to compile.
The following shows a few examples of configuring a model for training:
```
# Configure a model for mean-squared error regression.
model.compile(optimizer=tf.keras.optimizers.Adam(0.01),
loss='mse', # mean squared error
metrics=['mae']) # mean absolute error
# Configure a model for categorical classification.
model.compile(optimizer=tf.keras.optimizers.RMSprop(0.01),
loss=tf.keras.losses.CategoricalCrossentropy(),
metrics=[tf.keras.metrics.CategoricalAccuracy()])
```
### Train from NumPy data
For small datasets, use in-memory [NumPy](https://www.numpy.org/)
arrays to train and evaluate a model. The model is "fit" to the training data
using the `fit` method:
```
import numpy as np
data = np.random.random((1000, 32))
labels = np.random.random((1000, 10))
model.fit(data, labels, epochs=10, batch_size=32)
```
`tf.keras.Model.fit` takes three important arguments:
* `epochs`: Training is structured into *epochs*. An epoch is one iteration over
the entire input data (this is done in smaller batches).
* `batch_size`: When passed NumPy data, the model slices the data into smaller
batches and iterates over these batches during training. This integer
specifies the size of each batch. Be aware that the last batch may be smaller
if the total number of samples is not divisible by the batch size.
* `validation_data`: When prototyping a model, you want to easily monitor its
performance on some validation data. Passing this argument—a tuple of inputs
and labels—allows the model to display the loss and metrics in inference mode
for the passed data, at the end of each epoch.
Here's an example using `validation_data`:
```
import numpy as np
data = np.random.random((1000, 32))
labels = np.random.random((1000, 10))
val_data = np.random.random((100, 32))
val_labels = np.random.random((100, 10))
model.fit(data, labels, epochs=10, batch_size=32,
validation_data=(val_data, val_labels))
```
### Train from tf.data datasets
Use the [Datasets API](../data.ipynb) to scale to large datasets
or multi-device training. Pass a `tf.data.Dataset` instance to the `fit`
method:
```
# Instantiates a toy dataset instance:
dataset = tf.data.Dataset.from_tensor_slices((data, labels))
dataset = dataset.batch(32)
model.fit(dataset, epochs=10)
```
Since the `Dataset` yields batches of data, this snippet does not require a `batch_size`.
Datasets can also be used for validation:
```
dataset = tf.data.Dataset.from_tensor_slices((data, labels))
dataset = dataset.batch(32)
val_dataset = tf.data.Dataset.from_tensor_slices((val_data, val_labels))
val_dataset = val_dataset.batch(32)
model.fit(dataset, epochs=10,
validation_data=val_dataset)
```
### Evaluate and predict
The `tf.keras.Model.evaluate` and `tf.keras.Model.predict` methods can use NumPy
data and a `tf.data.Dataset`.
Here's how to *evaluate* the inference-mode loss and metrics for the data provided:
```
# With Numpy arrays
data = np.random.random((1000, 32))
labels = np.random.random((1000, 10))
model.evaluate(data, labels, batch_size=32)
# With a Dataset
dataset = tf.data.Dataset.from_tensor_slices((data, labels))
dataset = dataset.batch(32)
model.evaluate(dataset)
```
And here's how to *predict* the output of the last layer in inference for the data provided,
as a NumPy array:
```
result = model.predict(data, batch_size=32)
print(result.shape)
```
For a complete guide on training and evaluation, including how to write custom training loops from scratch, see the [guide to training and evaluation](./train_and_evaluate.ipynb).
## Build complex models
### The Functional API
The `tf.keras.Sequential` model is a simple stack of layers that cannot
represent arbitrary models. Use the
[Keras functional API](./functional.ipynb)
to build complex model topologies such as:
* Multi-input models,
* Multi-output models,
* Models with shared layers (the same layer called several times),
* Models with non-sequential data flows (e.g. residual connections).
Building a model with the functional API works like this:
1. A layer instance is callable and returns a tensor.
2. Input tensors and output tensors are used to define a `tf.keras.Model`
instance.
3. This model is trained just like the `Sequential` model.
The following example uses the functional API to build a simple, fully-connected
network:
```
inputs = tf.keras.Input(shape=(32,)) # Returns an input placeholder
# A layer instance is callable on a tensor, and returns a tensor.
x = layers.Dense(64, activation='relu')(inputs)
x = layers.Dense(64, activation='relu')(x)
predictions = layers.Dense(10, activation='softmax')(x)
```
Instantiate the model given inputs and outputs.
```
model = tf.keras.Model(inputs=inputs, outputs=predictions)
# The compile step specifies the training configuration.
model.compile(optimizer=tf.keras.optimizers.RMSprop(0.001),
loss='categorical_crossentropy',
metrics=['accuracy'])
# Trains for 5 epochs
model.fit(data, labels, batch_size=32, epochs=5)
```
### Model subclassing
Build a fully-customizable model by subclassing `tf.keras.Model` and defining
your own forward pass. Create layers in the `__init__` method and set them as
attributes of the class instance. Define the forward pass in the `call` method.
Model subclassing is particularly useful when
[eager execution](../eager.ipynb) is enabled, because it allows the forward pass
to be written imperatively.
Note: if you need your model to *always* run imperatively, you can set `dynamic=True` when calling the `super` constructor.
> Key Point: Use the right API for the job. While model subclassing offers
flexibility, it comes at a cost of greater complexity and more opportunities for
user errors. If possible, prefer the functional API.
The following example shows a subclassed `tf.keras.Model` using a custom forward
pass that does not have to be run imperatively:
```
class MyModel(tf.keras.Model):
def __init__(self, num_classes=10):
super(MyModel, self).__init__(name='my_model')
self.num_classes = num_classes
# Define your layers here.
self.dense_1 = layers.Dense(32, activation='relu')
self.dense_2 = layers.Dense(num_classes, activation='sigmoid')
def call(self, inputs):
# Define your forward pass here,
# using layers you previously defined (in `__init__`).
x = self.dense_1(inputs)
return self.dense_2(x)
```
Instantiate the new model class:
```
model = MyModel(num_classes=10)
# The compile step specifies the training configuration.
model.compile(optimizer=tf.keras.optimizers.RMSprop(0.001),
loss='categorical_crossentropy',
metrics=['accuracy'])
# Trains for 5 epochs.
model.fit(data, labels, batch_size=32, epochs=5)
```
### Custom layers
Create a custom layer by subclassing `tf.keras.layers.Layer` and implementing
the following methods:
* `__init__`: Optionally define sublayers to be used by this layer.
* `build`: Create the weights of the layer. Add weights with the `add_weight`
method.
* `call`: Define the forward pass.
* Optionally, a layer can be serialized by implementing the `get_config` method
and the `from_config` class method.
Here's an example of a custom layer that implements a `matmul` of an input with
a kernel matrix:
```
class MyLayer(layers.Layer):
def __init__(self, output_dim, **kwargs):
self.output_dim = output_dim
super(MyLayer, self).__init__(**kwargs)
def build(self, input_shape):
# Create a trainable weight variable for this layer.
self.kernel = self.add_weight(name='kernel',
shape=(input_shape[1], self.output_dim),
initializer='uniform',
trainable=True)
def call(self, inputs):
return tf.matmul(inputs, self.kernel)
def get_config(self):
base_config = super(MyLayer, self).get_config()
base_config['output_dim'] = self.output_dim
return base_config
@classmethod
def from_config(cls, config):
return cls(**config)
```
Create a model using your custom layer:
```
model = tf.keras.Sequential([
MyLayer(10),
layers.Activation('softmax')])
# The compile step specifies the training configuration
model.compile(optimizer=tf.keras.optimizers.RMSprop(0.001),
loss='categorical_crossentropy',
metrics=['accuracy'])
# Trains for 5 epochs.
model.fit(data, labels, batch_size=32, epochs=5)
```
Learn more about creating new layers and models from scratch with subclassing in the [Guide to writing layers and models from scratch](./custom_layers_and_models.ipynb).
## Callbacks
A callback is an object passed to a model to customize and extend its behavior
during training. You can write your own custom callback, or use the built-in
`tf.keras.callbacks` that include:
* `tf.keras.callbacks.ModelCheckpoint`: Save checkpoints of your model at
regular intervals.
* `tf.keras.callbacks.LearningRateScheduler`: Dynamically change the learning
rate.
* `tf.keras.callbacks.EarlyStopping`: Interrupt training when validation
performance has stopped improving.
* `tf.keras.callbacks.TensorBoard`: Monitor the model's behavior using
[TensorBoard](https://tensorflow.org/tensorboard).
To use a `tf.keras.callbacks.Callback`, pass it to the model's `fit` method:
```
callbacks = [
# Interrupt training if `val_loss` stops improving for over 2 epochs
tf.keras.callbacks.EarlyStopping(patience=2, monitor='val_loss'),
# Write TensorBoard logs to `./logs` directory
tf.keras.callbacks.TensorBoard(log_dir='./logs')
]
model.fit(data, labels, batch_size=32, epochs=5, callbacks=callbacks,
validation_data=(val_data, val_labels))
```
<a name='save_and_restore'></a>
## Save and restore
<a name="weights_only"></a>
### Save just the weights values
Save and load the weights of a model using `tf.keras.Model.save_weights`:
```
model = tf.keras.Sequential([
layers.Dense(64, activation='relu', input_shape=(32,)),
layers.Dense(10, activation='softmax')])
model.compile(optimizer=tf.keras.optimizers.Adam(0.001),
loss='categorical_crossentropy',
metrics=['accuracy'])
# Save weights to a TensorFlow Checkpoint file
model.save_weights('./weights/my_model')
# Restore the model's state,
# this requires a model with the same architecture.
model.load_weights('./weights/my_model')
```
By default, this saves the model's weights in the
[TensorFlow checkpoint](../checkpoint.ipynb) file format. Weights can
also be saved to the Keras HDF5 format (the default for the multi-backend
implementation of Keras):
```
# Save weights to a HDF5 file
model.save_weights('my_model.h5', save_format='h5')
# Restore the model's state
model.load_weights('my_model.h5')
```
### Save just the model configuration
A model's configuration can be saved—this serializes the model architecture
without any weights. A saved configuration can recreate and initialize the same
model, even without the code that defined the original model. Keras supports
JSON and YAML serialization formats:
```
# Serialize a model to JSON format
json_string = model.to_json()
json_string
import json
import pprint
pprint.pprint(json.loads(json_string))
```
Recreate the model (newly initialized) from the JSON:
```
fresh_model = tf.keras.models.model_from_json(json_string)
```
Serializing a model to YAML format requires that you install `pyyaml` *before you import TensorFlow*:
```
yaml_string = model.to_yaml()
print(yaml_string)
```
Recreate the model from the YAML:
```
fresh_model = tf.keras.models.model_from_yaml(yaml_string)
```
Caution: Subclassed models are not serializable because their architecture is
defined by the Python code in the body of the `call` method.
### Save the entire model in one file
The entire model can be saved to a file that contains the weight values, the
model's configuration, and even the optimizer's configuration. This allows you
to checkpoint a model and resume training later—from the exact same
state—without access to the original code.
```
# Create a simple model
model = tf.keras.Sequential([
layers.Dense(10, activation='softmax', input_shape=(32,)),
layers.Dense(10, activation='softmax')
])
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
model.fit(data, labels, batch_size=32, epochs=5)
# Save entire model to a HDF5 file
model.save('my_model.h5')
# Recreate the exact same model, including weights and optimizer.
model = tf.keras.models.load_model('my_model.h5')
```
Learn more about saving and serialization for Keras models in the guide to [save and serialize models](./save_and_serialize.ipynb).
<a name="eager_execution"></a>
## Eager execution
[Eager execution](../eager.ipynb) is an imperative programming
environment that evaluates operations immediately. This is not required for
Keras, but is supported by `tf.keras` and useful for inspecting your program and
debugging.
All of the `tf.keras` model-building APIs are compatible with eager execution.
And while the `Sequential` and functional APIs can be used, eager execution
especially benefits *model subclassing* and building *custom layers*—the APIs
that require you to write the forward pass as code (instead of the APIs that
create models by assembling existing layers).
See the [eager execution guide](../eager.ipynb) for
examples of using Keras models with custom training loops and `tf.GradientTape`.
You can also find a complete, short example [here](https://www.tensorflow.org/tutorials/quickstart/advanced).
## Distribution
### Multiple GPUs
`tf.keras` models can run on multiple GPUs using
`tf.distribute.Strategy`. This API provides distributed
training on multiple GPUs with almost no changes to existing code.
Currently, `tf.distribute.MirroredStrategy` is the only supported
distribution strategy. `MirroredStrategy` does in-graph replication with
synchronous training using all-reduce on a single machine. To use
`distribute.Strategy`s , nest the optimizer instantiation and model construction and compilation in a `Strategy`'s `.scope()`, then
train the model.
The following example distributes a `tf.keras.Model` across multiple GPUs on a
single machine.
First, define a model inside the distributed strategy scope:
```
strategy = tf.distribute.MirroredStrategy()
with strategy.scope():
model = tf.keras.Sequential()
model.add(layers.Dense(16, activation='relu', input_shape=(10,)))
model.add(layers.Dense(1, activation='sigmoid'))
optimizer = tf.keras.optimizers.SGD(0.2)
model.compile(loss='binary_crossentropy', optimizer=optimizer)
model.summary()
```
Next, train the model on data as usual:
```
x = np.random.random((1024, 10))
y = np.random.randint(2, size=(1024, 1))
x = tf.cast(x, tf.float32)
dataset = tf.data.Dataset.from_tensor_slices((x, y))
dataset = dataset.shuffle(buffer_size=1024).batch(32)
model.fit(dataset, epochs=1)
```
For more information, see the [full guide on Distributed Training in TensorFlow](../distributed_training.ipynb).
| github_jupyter |
The k-means clustering algorithm represents each cluster by its corresponding cluster centroid. The algorithm partition the input data into *k* disjoint clusters by iteratively applying the following two steps:
1. Form *k* clusters by assigning each instance to its nearest centroid.
2. Recompute the centroid of each cluster.
In this section, we perform k-means clustering on a toy example of movie ratings dataset. We first create the dataset as follows.
```
import pandas as pd
ratings = [['john',5,5,2,1],['mary',4,5,3,2],['bob',4,4,4,3],['lisa',2,2,4,5],['lee',1,2,3,4],['harry',2,1,5,5]]
titles = ['user','Jaws','Star Wars','Exorcist','Omen']
movies = pd.DataFrame(ratings,columns=titles)
movies
```
In this example dataset, the first 3 users liked action movies (Jaws and Star Wars) while the last 3 users enjoyed horror movies (Exorcist and Omen). Our goal is to apply k-means clustering on the users to identify groups of users with similar movie preferences.
The example below shows how to apply k-means clustering (with k=2) on the movie ratings data. We must remove the "user" column first before applying the clustering algorithm. The cluster assignment for each user is displayed as a dataframe object.
```
from sklearn import cluster
data = movies.drop('user',axis=1)
k_means = cluster.KMeans(n_clusters=2, max_iter=50, random_state=1)
k_means.fit(data)
labels = k_means.labels_
pd.DataFrame(labels, index=movies.user, columns=['Cluster ID'])
```
The k-means clustering algorithm assigns the first three users to one cluster and the last three users to the second cluster. The results are consistent with our expectation. We can also display the centroid for each of the two clusters.
```
centroids = k_means.cluster_centers_
pd.DataFrame(centroids,columns=data.columns)
```
Observe that cluster 0 has higher ratings for the horror movies whereas cluster 1 has higher ratings for action movies. The cluster centroids can be applied to other users to determine their cluster assignments.
```
import numpy as np
testData = np.array([[4,5,1,2],[3,2,4,4],[2,3,4,1],[3,2,3,3],[5,4,1,4]])
labels = k_means.predict(testData)
labels = labels.reshape(-1,1)
usernames = np.array(['paul','kim','liz','tom','bill']).reshape(-1,1)
cols = movies.columns.tolist()
cols.append('Cluster ID')
newusers = pd.DataFrame(np.concatenate((usernames, testData, labels), axis=1),columns=cols)
newusers
```
To determine the number of clusters in the data, we can apply k-means with varying number of clusters from 1 to 6 and compute their corresponding sum-of-squared errors (SSE) as shown in the example below. The "elbow" in the plot of SSE versus number of clusters can be used to estimate the number of clusters.
```
import matplotlib.pyplot as plt
%matplotlib inline
numClusters = [1,2,3,4,5,6]
SSE = []
for k in numClusters:
k_means = cluster.KMeans(n_clusters=k)
k_means.fit(data)
SSE.append(k_means.inertia_)
plt.plot(numClusters, SSE)
plt.xlabel('Number of Clusters')
plt.ylabel('SSE')
```
### Question 1
Complete the codes below to perform a K-mean clustering on the Iris dataset
```
import matplotlib.pyplot as plt
from sklearn import datasets
from sklearn.cluster import KMeans
import sklearn.metrics as sm
import pandas as pd
import numpy as np
%matplotlib inline
df = pd.read_csv('data/iris.csv', header=None)
df.columns = ['sepal length', 'sepal width', 'petal length', 'petal width', 'class']
print (df.head())
print (df.shape)
X = df.drop('class', axis=1)
y_text = df['class']
```
The iris dataset has three classes: Iris-setosa, Iris-virginica, and Iris-versicolor. The class labels need to be converted to numeric form for Kmean.
```
y = pd.Series(range(0,y_text.size), index = range(0, y_text.size))
for i in range(0,y_text.size):
if (y_text[i] == '______'):
y[i] = 0;
elif (y_text[i] == ______):
y[i] = 1;
else:
y[i] = 2;
model = KMeans(n_clusters=__)
model.fit(X)
plt.figure(figsize=(14,7))
# Create a colormap (how many colors?)
colormap = np.random.rand(__,)
# Plot the Original Classifications
plt.subplot(1, 2, 1)
plt.scatter(X['petal length'], X['petal width'], c=colormap[y], s=40)
plt.title('Real Classification')
predY = np.choose(model.labels_, [2, 0, 1]).astype(np.int64)
# Plot the Models Classifications
plt.subplot(1, 2, 2)
plt.scatter(X['petal length'], X['petal width'], c=colormap[predY], s=40)
plt.title('K Mean Classification')
```
Next, we examine examples of applying hierarchical clustering to the vertebrate dataset used in Module 6 (Classification). Specifically, we illustrate the results of using 3 hierarchical clustering algorithms provided by the Python scipy library: (1) single link (MIN), (2) complete link (MAX), and (3) group average. Other hierarchical clustering algorithms provided by the library include centroid-based and Ward's method.
```
import pandas as pd
data = pd.read_csv('data/vertebrate.csv',header='infer')
data
from scipy.cluster import hierarchy
import matplotlib.pyplot as plt
%matplotlib inline
names = data['Name']
Y = data['Class']
X = data.drop(['Name','Class'],axis=1)
Z = hierarchy.linkage(X.as_matrix(), 'single')
dn = hierarchy.dendrogram(Z,labels=names.tolist(),orientation='right')
Z = hierarchy.linkage(X.as_matrix(), 'complete')
dn = hierarchy.dendrogram(Z,labels=names.tolist(),orientation='right')
Z = hierarchy.linkage(X.as_matrix(), 'average')
dn = hierarchy.dendrogram(Z,labels=names.tolist(),orientation='right')
```
In density-based clustering, we identify the individual clusters as high-density regions that are separated by regions of low density. DBScan is one of the most popular density based clustering algorithms. In DBScan, data points are classified into 3 types---core points, border points, and noise points---based on the density of their local neighborhood. The local neighborhood density is defined according to 2 parameters: radius of neighborhood size (eps) and minimum number of points in the neighborhood (min_samples).
```
import pandas as pd
data = pd.read_csv('data/chameleon.data', delimiter=' ', names=['x','y'])
data.plot.scatter(x='x',y='y')
data
```
We apply the DBScan clustering algorithm on the data by setting the neighborhood radius (eps) to 15.5 and minimum number of points (min_samples) to be 5. The clusters are assigned to IDs between 0 to 8 while the noise points are assigned to a cluster ID equals to -1.
```
from sklearn.cluster import DBSCAN
db = DBSCAN(eps=15.5, min_samples=5).fit(data)
core_samples_mask = np.zeros_like(db.labels_, dtype=bool)
core_samples_mask[db.core_sample_indices_] = True
labels = pd.DataFrame(db.labels_,columns=['Cluster ID'])
result = pd.concat((data,labels), axis=1)
result.plot.scatter(x='x',y='y',c='Cluster ID', colormap='jet')
```
One of the main limitations of the k-means clustering algorithm is its tendency to seek for globular-shaped clusters. Thus, it does not work when applied to datasets with arbitrary-shaped clusters or when the cluster centroids overlapped with one another. Spectral clustering can overcome this limitation by exploiting properties of the similarity graph to overcome such limitations. To illustrate this, consider the following two-dimensional datasets.
```
import pandas as pd
data1 = pd.read_csv('data/2d_data.txt', delimiter=' ', names=['x','y'])
data2 = pd.read_csv('data/elliptical.txt', delimiter=' ', names=['x','y'])
fig, (ax1,ax2) = plt.subplots(nrows=1, ncols=2, figsize=(12,5))
data1.plot.scatter(x='x',y='y',ax=ax1)
data2.plot.scatter(x='x',y='y',ax=ax2)
```
Below, we demonstrate the results of applying k-means to the datasets (with k=2).
```
from sklearn import cluster
k_means = cluster.KMeans(n_clusters=2, max_iter=50, random_state=1)
k_means.fit(data1)
labels1 = pd.DataFrame(k_means.labels_,columns=['Cluster ID'])
result1 = pd.concat((data1,labels1), axis=1)
k_means2 = cluster.KMeans(n_clusters=2, max_iter=50, random_state=1)
k_means2.fit(data2)
labels2 = pd.DataFrame(k_means2.labels_,columns=['Cluster ID'])
result2 = pd.concat((data2,labels2), axis=1)
fig, (ax1,ax2) = plt.subplots(nrows=1, ncols=2, figsize=(12,5))
result1.plot.scatter(x='x',y='y',c='Cluster ID',colormap='jet',ax=ax1)
ax1.set_title('K-means Clustering')
result2.plot.scatter(x='x',y='y',c='Cluster ID',colormap='jet',ax=ax2)
ax2.set_title('K-means Clustering')
```
The plots above show the poor performance of k-means clustering. Next, we apply spectral clustering to the datasets. Spectral clustering converts the data into a similarity graph and applies the normalized cut graph partitioning algorithm to generate the clusters. In the example below, we use the Gaussian radial basis function as our affinity (similarity) measure. Users need to tune the kernel parameter (gamma) value in order to obtain the appropriate clusters for the given dataset.
```
from sklearn import cluster
import pandas as pd
spectral = cluster.SpectralClustering(n_clusters=2,random_state=1,affinity='rbf',gamma=5000)
spectral.fit(data1)
labels1 = pd.DataFrame(spectral.labels_,columns=['Cluster ID'])
result1 = pd.concat((data1,labels1), axis=1)
spectral2 = cluster.SpectralClustering(n_clusters=2,random_state=1,affinity='rbf',gamma=100)
spectral2.fit(data2)
labels2 = pd.DataFrame(spectral2.labels_,columns=['Cluster ID'])
result2 = pd.concat((data2,labels2), axis=1)
fig, (ax1,ax2) = plt.subplots(nrows=1, ncols=2, figsize=(12,5))
result1.plot.scatter(x='x',y='y',c='Cluster ID',colormap='jet',ax=ax1)
ax1.set_title('Spectral Clustering')
result2.plot.scatter(x='x',y='y',c='Cluster ID',colormap='jet',ax=ax2)
ax2.set_title('Spectral Clustering')
```
## Question 2
This is a free-form exercise. You are to use the *titanic3.xls* data and perform a K-mean clustering analysis on the data. In data mining, and in data science in general, we always want to *let the data do the talking*. Use one or more of the techniques above to analyze this data, and draw some conclusions from your analysis. Think about a story you want to tell based on the Titanic data.
| github_jupyter |
```
import collections
from collections import defaultdict
import sys
import json
import random
from jsmin import jsmin
from io import StringIO
import numpy as np
import copy
import importlib
from functools import partial
import math
import os
import compress_pickle
# script_n = os.path.basename(__file__).split('.')[0]
script_n = 'mf_combination_representation_210412'
sys.path.insert(0, '/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc')
import my_plot
importlib.reload(my_plot)
from my_plot import MyPlotData
fname = ('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/'\
'mf_grc_model/input_graph_210407_all.gz')
input_graph = compress_pickle.load(fname)
# z_min = 19800
# z_max = 29800
z_min = 19800
z_max = 29800
# GrCs are fully reconstructed and proofread from 90k to 150k
x_min = 105*1000*4
x_max = 135*1000*4
pair_reps = defaultdict(int)
mfs_within_box = set()
for mf_id, mf in input_graph.mfs.items():
rosette_loc_size = {}
mf.get_all_mf_locs_size(rosette_loc_size)
for rosette_loc, size in rosette_loc_size.items():
x, y, z = rosette_loc
if x < x_min or x > x_max:
continue
if z < z_min or z > z_max:
continue
mfs_within_box.add(mf_id)
import itertools
grcs_edge_count = defaultdict(int)
def get_prob_2share(in_graph, count_within_box=True):
shares = defaultdict(lambda: defaultdict(int))
# processed = set()
# total_n_pairs = 0
# hist = defaultdict(int)
# n = 0
counted_grcs = 0
for grc_i_id in in_graph.grcs:
grc_i = in_graph.grcs[grc_i_id]
if count_within_box:
mf_ids = set([mf[0] for mf in grc_i.edges if mf[0] in mfs_within_box])
else:
mf_ids = set([mf[0] for mf in grc_i.edges if mf[0]])
mf_ids = sorted(list(mf_ids))
if len(mf_ids):
grcs_edge_count[grc_i_id] = len(mf_ids)
# print(mf_ids)
for i in range(1, len(mf_ids)+1):
for combination in itertools.combinations(mf_ids, i):
shares[i][combination] += 1
# print(combination)
return shares
# if len(rosettes_i) == 0:
# continue
# if len(rosettes_i) == 1:
# shares[1][mf_ids[0]]
# continue
# for grc_j_id in in_graph.grcs:
# if grc_i_id == grc_j_id:
# continue
# if unique_count and (grc_i_id, grc_j_id) in processed:
# continue
# processed.add((grc_i_id, grc_j_id))
# processed.add((grc_j_id, grc_i_id))
# grc_j = in_graph.grcs[grc_j_id]
# common_rosettes = set([mf[0] for mf in grc_j.edges])
# common_rosettes = common_rosettes & rosettes_i
# hist[len(common_rosettes)] += 1
# for k in hist:
# # fix 0 datapoint plots
# if hist[k] == 0:
# hist[k] = 1
# if return_counted:
# return hist, counted_grcs
# else:
# return hist
hist_data = get_prob_2share(input_graph)
# print(hist_data)
# n_grcs = len(input_graph.grcs)
def calc_distribution(hist):
dist = defaultdict(lambda: defaultdict(int))
dist_raw = defaultdict(list)
for k in hist:
for v in hist[k]:
val = hist[k][v]
dist[k][val] += 1
dist_raw[k].append(val)
return dist, dist_raw
dist, dist_raw = calc_distribution(hist_data)
for k in sorted(hist_data.keys()):
pairs = list(hist_data[k].items())
pairs.sort(key=lambda x: x[1])
print(k)
for pair in pairs:
print(pair)
for share in dist:
print(share)
for v in sorted(dist[share].keys()):
print(f'{v}: {dist[share][v]}')
print(len(mfs_within_box))
print(len(grcs_edge_count))
s = 0
for k, v in grcs_edge_count.items():
if v >= 2:
s += 1
print(s)
# shuffle the graph and see how it affects distribution
shuffled_shares = defaultdict(lambda: defaultdict(int))
mfs_within_box = list(mfs_within_box)
for grc_id, count in grcs_edge_count.items():
mf_ids = []
for i in range(count):
mf_ids.append(random.choice(mfs_within_box))
mf_ids = list(set(mf_ids))
for i in range(1, len(mf_ids)+1):
for combination in itertools.combinations(mf_ids, i):
shuffled_shares[i][combination] += 1
shuffle_dist, shuffle_dist_raw = calc_distribution(shuffled_shares)
for share in shuffle_dist:
print(share)
for v in sorted(shuffle_dist[share].keys()):
print(f'{v}: {shuffle_dist[share][v]}')
```
| github_jupyter |
# 更多字符串和特殊方法
- 前面我们已经学了类,在Python中还有一些特殊的方法起着非常重要的作用,这里会介绍一些特殊的方法和运算符的重载,以及使用特殊方法设计类
## str 类
- 一个str对象是不可变的,也就是说,一旦创建了这个字符串,那么它的内容在认为不改变的情况下是不会变的
- s1 = str()
- s2 = str('welcome to Python')
## 创建两个对象,分别观察两者id
- id为Python内存地址
## 处理字符串的函数
- len
- max
- min
- 字符串一切是按照ASCII码值进行比较
## 下角标运算符 []
- 一个字符串是一个字符序列,可以通过索引进行访问
- 观察字符串是否是一个可迭代序列 \__iter__
```
for i in 'sss':
print (i)
```
## 切片 [start: end]
- start 默认值为0
- end 默认值为-1
```
w = 'jokerisabadman'
w[-8:-4] #前闭后开
```
## 链接运算符 + 和复制运算符 *
- \+ 链接多个字符串,同时''.join()也是
- \* 复制多个字符串
```
'a' + 'b'.join('cdefg')
'a' * 10 #不可以和浮点型的相乘
'a' * False
```
## in 和 not in 运算符
- in :判断某个字符是否在字符串内
- not in :判断某个字符是否不在字符串内
- 返回的是布尔值
```
w = 'jokerisabadman'
'ker' in w #整体进行匹配
def find_():
str.find(str, beg=0, end=len(string))
import string
s='nihao,shijie'
t='nihao'
result = string.find(s,t)!=-1
print result
True
result = string.rfind(s,t)!=-1
print result
True
```
## 比较字符串
- ==, !=, >=, <=, >, <
- 依照ASCII码值进行比较
## 测试字符串

- 注意:
> - isalnum() 中是不能包含空格,否则会返回False
```
def number1():
n = str(input('输入密码:'))
if n.isalnum() is True :
if n.isalpha() is True :
if n.islower() ia True :
print('可以')
else :
print('至少一个大小写')
else :
print('不符合')
else :
print('至少一个字符')
number1()
a = 'aaA111'
count1 = 0
count2 = 0
count3 = 0
for i in a:
if i.islower() is True:
count1 +=1
if i.isupper() is True:
count2 +=1
if i.isdigit() is True:
count3 +=1
else:
if count1 == 0:
print('密码必须含有小写字母')
if count2 == 0:
print('密码必须含有大写')
if count3 == 0:
print('密码必须含有数字')
if count1 !=0 and count2 !=0 and count3 !=0:
print('密码设置成功')
print(count1,count2,count3)
```
## 搜索子串

```
b = '你们都是小天才'
b.endswith('你们')
b.startswith('你们都是')
b = '你们都是小天都才' #从右向左查询
b.rfind('都')
path = 'G:\yun2kaifang.txt'
'a,b,c'.split(',')
```
## 转换字符串

```
'asdk'.capitalize()
'sFKSMFdgd'.lower()
'dsadasASIFBH'.upper()
'old,new'.replace('old','new')
import random,string #调用random、string模块
def yanzheng():
src_digits = string.digits #string_数字
src_uppercase = string.ascii_uppercase #string_大写字母
src_lowercase = string.ascii_lowercase #string_小写字母
digits_num = random.randint(1,6)
uppercase_num = random.randint(1,8-digits_num-1)
lowercase_num = 8 - (digits_num + uppercase_num) #生成字符串
password = random.sample(src_digits,digits_num) + random.sample(src_uppercase,uppercase_num) + random.sample(src_lowercase,lowercase_num)#打乱字符串
random.shuffle(password) #列表转字符串
new_password = ''.join(password)
print(new_password)
n = str(input('输入验证码:'))
if n == new_password.lower():
print('输入正确!')
elif n == new_password.upper():
print('输入正确!')
else :
print('输入不正确')
yanzheng()
```
## 删除字符串

```
a = ' adaspk oasdopa apodjaspjd spajda jdaposda @#$$$#@!! '
b = a.replace(' ',' ')
b
```
## 格式化字符串

```
'a'.center(10,'@')
'a'.ljust(10,'$')
'a'.rjust(10,'#')
```
## EP:
- 1

- 2
随机参数100个数字,将www.baidu.com/?page=进行拼接
```
'a b'.isalpha
```
## Python高级使用方法 -- 字符串
- 我们经常使用的方法实际上就是调用Python的运算重载

# Homework
- 1

```
def safe():
n = str(input('请按照ddd-dd-dddd的格式输入一个社会安全号码:'))
if :
print('Valid SSN')
else :
print('Invalid SSN')
'456-88-8456'.split('-')
```
- 2

- 3

- 4

- 5

- 6

- 7

- 8

- 9

| github_jupyter |
<table width="100%"> <tr>
<td style="background-color:#ffffff;">
<a href="https://qsoftware.lu.lv/index.php/qworld/" target="_blank"><img src="../images/qworld.jpg" width="35%" align="left"> </a></td>
<td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
prepared by Abuzer Yakaryilmaz (<a href="http://qworld.lu.lv/index.php/qlatvia/" target="_blank">QLatvia</a>)
</td>
</tr></table>
<table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
$ \newcommand{\bra}[1]{\langle #1|} $
$ \newcommand{\ket}[1]{|#1\rangle} $
$ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
$ \newcommand{\dot}[2]{ #1 \cdot #2} $
$ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
$ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
$ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
$ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
$ \newcommand{\mypar}[1]{\left( #1 \right)} $
$ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
$ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
$ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
$ \newcommand{\onehalf}{\frac{1}{2}} $
$ \newcommand{\donehalf}{\dfrac{1}{2}} $
$ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
$ \newcommand{\vzero}{\myvector{1\\0}} $
$ \newcommand{\vone}{\myvector{0\\1}} $
$ \newcommand{\vhadamardzero}{\myvector{ \sqrttwo \\ \sqrttwo } } $
$ \newcommand{\vhadamardone}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
$ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
$ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
$ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
$ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
$ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
$ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
<h2> <font color="blue"> Solutions for </font>Coin Flip: A Probabilistic Bit</h2>
<a id="task1"></a>
<h3> Task 1: Simulating FairCoin in Python</h3>
Flip a fair coin 100 times. Calculate the total number of heads and tails, and then check the ratio of the number of heads and the number of tails.
Do the same experiment 1000 times.
Do the same experiment 10,000 times.
Do the same experiment 100,000 times.
Do your results get close to the ideal case (the numbers of heads and tails are equal)?
<h3>Solution</h3>
```
from random import randrange
for experiment in [100,1000,10000,100000]:
heads = tails = 0
for i in range(experiment):
if randrange(2) == 0: heads = heads + 1
else: tails = tails + 1
print("experiment:",experiment)
print("heads =",heads," tails = ",tails)
print("the ratio of #heads/#tails is",(round(heads/tails,4)))
print() # empty line
```
<a id="task2"></a>
<h3> Task 2: Simulating BiasedCoin in Python</h3>
Flip the following biased coin 100 times. Calcuate the total numbers of heads and tails, and then check the ratio of the number of heads and the number of tails.
$
BiasedCoin = \begin{array}{c|cc} & \mathbf{Head} & \mathbf{Tail} \\ \hline \mathbf{Head} & 0.6 & 0.6 \\ \mathbf{Tail} & 0.4 & 0.4 \end{array}
$
Do the same experiment 1000 times.
Do the same experiment 10,000 times.
Do the same experiment 100,000 times.
Do your results get close to the ideal case $ \mypar{ \dfrac{ \mbox{# of heads} }{ \mbox{# of tails} } = \dfrac{0.6}{0.4} = 1.50000000 } $?
<h3>Solution</h3>
```
from random import randrange
# let's pick a random number between {0,1,...,99}
# it is expected to be less than 60 with probability 0.6
# and greater than or equal to 60 with probability 0.4
for experiment in [100,1000,10000,100000]:
heads = tails = 0
for i in range(experiment):
if randrange(100) <60: heads = heads + 1 # with probability 0.6
else: tails = tails + 1 # with probability 0.4
print("experiment:",experiment)
print("heads =",heads," tails = ",tails)
print("the ratio of #heads/#tails is",(round(heads/tails,4)))
print() # empty line
```
<a id="task3"></a>
<h3> Task 3</h3>
Write a function to implement the described biased coin,
The inputs are integers $ N >0 $ and $ 0 \leq B < N $.
The output is either "Heads" or "Tails".
<h3>Solution</h3>
```
def biased_coin(N,B):
from random import randrange
random_number = randrange(N)
if random_number < B:
return "Heads"
else:
return "Tails"
```
<a id="task4"></a>
<h3> Task 4</h3>
We use the biased coin described in Task 3.
(You may use the function given in the solution.)
We pick $ N $ as 101.
Our task is to determine the value of $ B $ experimentially without checking its value directly.
Flip the (same) biased coin 500 times, collect the statistics, and then guess the bias.
Compare your guess with the actual bias by calculating the error (the absolute value of the difference).
<h3>Solution</h3>
```
def biased_coin(N,B):
from random import randrange
random_number = randrange(N)
if random_number < B:
return "Heads"
else:
return "Tails"
from random import randrange
N = 101
B = randrange(100)
total_tosses = 500
the_number_of_heads = 0
for i in range(total_tosses):
if biased_coin(N,B) == "Heads":
the_number_of_heads = the_number_of_heads + 1
my_guess = the_number_of_heads/total_tosses
real_bias = B/N
error = abs(my_guess-real_bias)/real_bias*100
print("my guess is",my_guess)
print("real bias is",real_bias)
print("error (%) is",error)
```
| github_jupyter |
# Introduction to Deep Learning with PyTorch
In this notebook, you'll get introduced to [PyTorch](http://pytorch.org/), a framework for building and training neural networks. PyTorch in a lot of ways behaves like the arrays you love from Numpy. These Numpy arrays, after all, are just tensors. PyTorch takes these tensors and makes it simple to move them to GPUs for the faster processing needed when training neural networks. It also provides a module that automatically calculates gradients (for backpropagation!) and another module specifically for building neural networks. All together, PyTorch ends up being more coherent with Python and the Numpy/Scipy stack compared to TensorFlow and other frameworks.
## Neural Networks
Deep Learning is based on artificial neural networks which have been around in some form since the late 1950s. The networks are built from individual parts approximating neurons, typically called units or simply "neurons." Each unit has some number of weighted inputs. These weighted inputs are summed together (a linear combination) then passed through an activation function to get the unit's output.
<img src="assets/simple_neuron.png" width=400px>
Mathematically this looks like:
$$
\begin{align}
y &= f(w_1 x_1 + w_2 x_2 + b) \\
y &= f\left(\sum_i w_i x_i +b \right)
\end{align}
$$
With vectors this is the dot/inner product of two vectors:
$$
h = \begin{bmatrix}
x_1 \, x_2 \cdots x_n
\end{bmatrix}
\cdot
\begin{bmatrix}
w_1 \\
w_2 \\
\vdots \\
w_n
\end{bmatrix}
$$
## Tensors
It turns out neural network computations are just a bunch of linear algebra operations on *tensors*, a generalization of matrices. A vector is a 1-dimensional tensor, a matrix is a 2-dimensional tensor, an array with three indices is a 3-dimensional tensor (RGB color images for example). The fundamental data structure for neural networks are tensors and PyTorch (as well as pretty much every other deep learning framework) is built around tensors.
<img src="assets/tensor_examples.svg" width=600px>
With the basics covered, it's time to explore how we can use PyTorch to build a simple neural network.
```
# First, import PyTorch
import torch
def activation(x):
""" Sigmoid activation function
Arguments
---------
x: torch.Tensor
"""
return 1/(1+torch.exp(-x))
### Generate some data
torch.manual_seed(7) # Set the random seed so things are predictable
# Features are 5 random normal variables
features = torch.randn((1, 5))
# True weights for our data, random normal variables again
weights = torch.randn_like(features)
# and a true bias term
bias = torch.randn((1, 1))
```
Above I generated data we can use to get the output of our simple network. This is all just random for now, going forward we'll start using normal data. Going through each relevant line:
`features = torch.randn((1, 5))` creates a tensor with shape `(1, 5)`, one row and five columns, that contains values randomly distributed according to the normal distribution with a mean of zero and standard deviation of one.
`weights = torch.randn_like(features)` creates another tensor with the same shape as `features`, again containing values from a normal distribution.
Finally, `bias = torch.randn((1, 1))` creates a single value from a normal distribution.
PyTorch tensors can be added, multiplied, subtracted, etc, just like Numpy arrays. In general, you'll use PyTorch tensors pretty much the same way you'd use Numpy arrays. They come with some nice benefits though such as GPU acceleration which we'll get to later. For now, use the generated data to calculate the output of this simple single layer network.
> **Exercise**: Calculate the output of the network with input features `features`, weights `weights`, and bias `bias`. Similar to Numpy, PyTorch has a [`torch.sum()`](https://pytorch.org/docs/stable/torch.html#torch.sum) function, as well as a `.sum()` method on tensors, for taking sums. Use the function `activation` defined above as the activation function.
```
## Calculate the output of this network using the weights and bias tensors
def metwork_op(features,weights,bias):
return activation(torch.sum(torch.mm(features,weights)))
```
You can do the multiplication and sum in the same operation using a matrix multiplication. In general, you'll want to use matrix multiplications since they are more efficient and accelerated using modern libraries and high-performance computing on GPUs.
Here, we want to do a matrix multiplication of the features and the weights. For this we can use [`torch.mm()`](https://pytorch.org/docs/stable/torch.html#torch.mm) or [`torch.matmul()`](https://pytorch.org/docs/stable/torch.html#torch.matmul) which is somewhat more complicated and supports broadcasting. If we try to do it with `features` and `weights` as they are, we'll get an error
```python
>> torch.mm(features, weights)
---------------------------------------------------------------------------
RuntimeError Traceback (most recent call last)
<ipython-input-13-15d592eb5279> in <module>()
----> 1 torch.mm(features, weights)
RuntimeError: size mismatch, m1: [1 x 5], m2: [1 x 5] at /Users/soumith/minicondabuild3/conda-bld/pytorch_1524590658547/work/aten/src/TH/generic/THTensorMath.c:2033
```
As you're building neural networks in any framework, you'll see this often. Really often. What's happening here is our tensors aren't the correct shapes to perform a matrix multiplication. Remember that for matrix multiplications, the number of columns in the first tensor must equal to the number of rows in the second column. Both `features` and `weights` have the same shape, `(1, 5)`. This means we need to change the shape of `weights` to get the matrix multiplication to work.
**Note:** To see the shape of a tensor called `tensor`, use `tensor.shape`. If you're building neural networks, you'll be using this method often.
There are a few options here: [`weights.reshape()`](https://pytorch.org/docs/stable/tensors.html#torch.Tensor.reshape), [`weights.resize_()`](https://pytorch.org/docs/stable/tensors.html#torch.Tensor.resize_), and [`weights.view()`](https://pytorch.org/docs/stable/tensors.html#torch.Tensor.view).
* `weights.reshape(a, b)` will return a new tensor with the same data as `weights` with size `(a, b)` sometimes, and sometimes a clone, as in it copies the data to another part of memory.
* `weights.resize_(a, b)` returns the same tensor with a different shape. However, if the new shape results in fewer elements than the original tensor, some elements will be removed from the tensor (but not from memory). If the new shape results in more elements than the original tensor, new elements will be uninitialized in memory. Here I should note that the underscore at the end of the method denotes that this method is performed **in-place**. Here is a great forum thread to [read more about in-place operations](https://discuss.pytorch.org/t/what-is-in-place-operation/16244) in PyTorch.
* `weights.view(a, b)` will return a new tensor with the same data as `weights` with size `(a, b)`.
I usually use `.view()`, but any of the three methods will work for this. So, now we can reshape `weights` to have five rows and one column with something like `weights.view(5, 1)`.
> **Exercise**: Calculate the output of our little network using matrix multiplication.
```
## Calculate the output of this network using matrix multiplication
print(features.shape)
c=features.reshape(5,1)
print(c)
print(weights.shape)
print(metwork_op(c,weights,bias))
```
### Stack them up!
That's how you can calculate the output for a single neuron. The real power of this algorithm happens when you start stacking these individual units into layers and stacks of layers, into a network of neurons. The output of one layer of neurons becomes the input for the next layer. With multiple input units and output units, we now need to express the weights as a matrix.
<img src='assets/multilayer_diagram_weights.png' width=450px>
The first layer shown on the bottom here are the inputs, understandably called the **input layer**. The middle layer is called the **hidden layer**, and the final layer (on the right) is the **output layer**. We can express this network mathematically with matrices again and use matrix multiplication to get linear combinations for each unit in one operation. For example, the hidden layer ($h_1$ and $h_2$ here) can be calculated
$$
\vec{h} = [h_1 \, h_2] =
\begin{bmatrix}
x_1 \, x_2 \cdots \, x_n
\end{bmatrix}
\cdot
\begin{bmatrix}
w_{11} & w_{12} \\
w_{21} &w_{22} \\
\vdots &\vdots \\
w_{n1} &w_{n2}
\end{bmatrix}
$$
The output for this small network is found by treating the hidden layer as inputs for the output unit. The network output is expressed simply
$$
y = f_2 \! \left(\, f_1 \! \left(\vec{x} \, \mathbf{W_1}\right) \mathbf{W_2} \right)
$$
```
### Generate some data
torch.manual_seed(7) # Set the random seed so things are predictable
# Features are 3 random normal variables
features = torch.randn((1, 3))
# Define the size of each layer in our network
n_input = features.shape[1] # Number of input units, must match number of input features
n_hidden = 2 # Number of hidden units
n_output = 1 # Number of output units
# Weights for inputs to hidden layer
W1 = torch.randn(n_input, n_hidden)
# Weights for hidden layer to output layer
W2 = torch.randn(n_hidden, n_output)
# and bias terms for hidden and output layers
B1 = torch.randn((1, n_hidden))
B2 = torch.randn((1, n_output))
```
> **Exercise:** Calculate the output for this multi-layer network using the weights `W1` & `W2`, and the biases, `B1` & `B2`.
```
## Your solution here
print(features.shape)
print(W1.shape)
print(W2.shape)
print(B1.shape)
c=activation(torch.add(torch.mm(features,W1),B1))
d=activation(torch.add(torch.mm(c,W2),B2))
print(c)
print(d)
```
If you did this correctly, you should see the output `tensor([[ 0.3171]])`.
The number of hidden units a parameter of the network, often called a **hyperparameter** to differentiate it from the weights and biases parameters. As you'll see later when we discuss training a neural network, the more hidden units a network has, and the more layers, the better able it is to learn from data and make accurate predictions.
## Numpy to Torch and back
Special bonus section! PyTorch has a great feature for converting between Numpy arrays and Torch tensors. To create a tensor from a Numpy array, use `torch.from_numpy()`. To convert a tensor to a Numpy array, use the `.numpy()` method.
```
import numpy as np
a = np.random.rand(4,3)
a
b = torch.from_numpy(a)
b
b.numpy()
```
The memory is shared between the Numpy array and Torch tensor, so if you change the values in-place of one object, the other will change as well.
```
# Multiply PyTorch Tensor by 2, in place
b.mul_(2)
# Numpy array matches new values from Tensor
a
```
| github_jupyter |
# Unit 5 - Financial Planning
```
# Initial imports
import os
import requests
import pandas as pd
from dotenv import load_dotenv
import alpaca_trade_api as tradeapi
from MCForecastTools import MCSimulation
%matplotlib inline
# Load .env enviroment variables
load_dotenv()
```
## Part 1 - Personal Finance Planner
### Collect Crypto Prices Using the `requests` Library
```
# Set current amount of crypto assets
my_btc = 1.2
my_eth = 5.3
# Crypto API URLs
btc_url = "https://api.alternative.me/v2/ticker/Bitcoin/?convert=CAD"
eth_url = "https://api.alternative.me/v2/ticker/Ethereum/?convert=CAD"
# Fetch current BTC price
btc_response_data = requests.get(btc_url).json()
btc_response_data
# Fetch current ETH price
eth_response_data = requests.get(eth_url).json()
eth_response_data
# Compute current value of my crypto
btc_current_value = btc_response_data['data']['1']['quotes']['USD']['price']
eth_current_value = eth_response_data['data']['1027']['quotes']['USD']['price']
# Print current crypto wallet balance
print(f"The current value of your {my_btc} BTC is ${btc_current_value:0.2f}") #0.2f prints two decimals on a float
print(f"The current value of your {my_eth} ETH is ${eth_current_value:0.2f}")
```
### Collect Investments Data Using Alpaca: `SPY` (stocks) and `AGG` (bonds)
```
# Set current amount of shares
my_agg = 200
my_spy = 50
# Set Alpaca API key and secret
alpaca_api_key = os.getenv("ALPACA_API_KEY")
alpaca_secret_key = os.getenv("ALPACA_SECRET_KEY")
alpaca_end_point = os.getenv("ALPACA_END_POINT")
# Create the Alpaca API object
alpaca = tradeapi.REST(
alpaca_api_key,
alpaca_secret_key,
api_version="v2",
base_url= alpaca_end_point)
# Format current date as ISO format
today = pd.Timestamp("2021-10-8", tz="America/New_York").isoformat()
# Set the tickers
tickers = ["AGG", "SPY"]
# Set timeframe to '1D' for Alpaca API
timeframe = "1D"
# Get current closing prices for SPY and AGG
df_portfolio = alpaca.get_barset(
tickers,
timeframe,
start = today,
end = today,
limit = 1000
).df
# Preview DataFrame
df_portfolio
# Pick AGG and SPY close prices
agg_close_price = float(df_portfolio["AGG"]["close"])
spy_close_price = float(df_portfolio["SPY"]["close"])
# Print AGG and SPY close prices
print(f"Current AGG closing price: ${agg_close_price}")
print(f"Current SPY closing price: ${spy_close_price}")
# Compute the current value of shares
my_spy_value = my_spy * spy_close_price
my_agg_value = my_agg * agg_close_price
# Print current value of shares
print(f"The current value of your {my_spy} SPY shares is ${my_spy_value:0.2f}")
print(f"The current value of your {my_agg} AGG shares is ${my_agg_value:0.2f}")
```
### Savings Health Analysis
```
# Set monthly household income
monthly_income = 12000
# Consolidate financial assets data
crypto_assets = btc_current_value + eth_current_value #adding together all crypto
stock_bond_assets = my_agg_value + my_agg_value #adding together all stocks and bonds
value_data = {
'amount': [crypto_assets, stock_bond_assets],
'asset': ["crypto", "shares"]
}
# Create savings DataFrame
df_savings = pd.DataFrame(value_data).set_index('asset')
# Display savings DataFrame
display(df_savings)
# Plot savings pie chart
pie_chart = df_savings.plot.pie(y= 'amount', title="Savings Portfolio Composition", figsize=(5,5))
# Set ideal emergency fund
emergency_fund = monthly_income * 3
# Calculate total amount of savings
total_savings = df_savings['amount'].sum()
# Validate saving health
#If total savings are greater than the emergency fund, display a message congratulating the person for having enough money in this fund.
if total_savings > emergency_fund:
print(f"Congratulations! You have enough money in your emergency fund.")
#If total savings are equal to the emergency fund, display a message congratulating the person on reaching this financial goal.
elif total_savings == emergency_fund:
print(f"You have reached your financial goal!")
#If total savings are less than the emergency fund, display a message showing how many dollars away the person is from reaching the goal.
else:
print(f"You're ${emergency_fund - total_savings:0.2f} away from achieving your financial goal. Keep saving!")
```
## Part 2 - Retirement Planning
### Monte Carlo Simulation
```
# Set start and end dates of five years back from today.
# Sample results may vary from the solution based on the time frame chosen
start_date1 = pd.Timestamp('2016-10-08', tz='America/New_York').isoformat()
end_date1 = pd.Timestamp('2019-10-08', tz='America/New_York').isoformat()
start_date2 = pd.Timestamp('2019-10-09', tz='America/New_York').isoformat()
end_date2 = pd.Timestamp('2021-10-08', tz='America/New_York').isoformat()
# Get 5 years' worth of historical data for SPY and AGG
# (use a limit=1000 parameter to call the most recent 1000 days of data)
#The 5 year time period is greater than 1000 days. I will use two dataframes to obtain the data from the Alpaca API and then concatenate them.
df_portfolio_3_year = alpaca.get_barset(
tickers,
timeframe,
start = start_date1,
end = end_date1,
limit = 1000
).df
df_portfolio_2_year = alpaca.get_barset(
tickers,
timeframe,
start = start_date2,
end = end_date2,
limit = 1000
).df
# Concatenate dataframes
df_portfolio_5_year = pd.concat((df_portfolio_3_year,df_portfolio_2_year),axis="rows", join="inner", sort=True).dropna()
# Display sample data
df_portfolio_5_year.head()
# Configuring a Monte Carlo simulation to forecast 30 years cumulative returns
# Set number of simulations
num_sims = 500
MC_portfolio = MCSimulation(
portfolio_data = df_portfolio_5_year,
weights = [0.6,0.4],
num_simulation = num_sims,
num_trading_days = 252*30
)
# Running a Monte Carlo simulation to forecast 30 years cumulative returns
MC_portfolio.calc_cumulative_return()
# Plot simulation outcomes
line_plot = MC_portfolio.plot_simulation()
# Plot probability distribution and confidence intervals
dist_plot = MC_portfolio.plot_distribution()
```
### Retirement Analysis
```
# Fetch summary statistics from the Monte Carlo simulation results
summary_tbl = MC_portfolio.summarize_cumulative_return()
# Print summary statistics
print(summary_tbl)
```
### Calculate the expected portfolio return at the `95%` lower and upper confidence intervals based on a `$20,000` initial investment.
```
# Set initial investment
initial_investment = 20000
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $20,000
ci_lower = round(summary_tbl[8]*initial_investment,2)
ci_upper = round(summary_tbl[9]*initial_investment,2)
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 30 years will end within in the range of"
f" ${ci_lower} and ${ci_upper}")
```
### Calculate the expected portfolio return at the `95%` lower and upper confidence intervals based on a `50%` increase in the initial investment.
```
# Set initial investment
initial_investment = 20000 * 1.5
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $30,000
ci_lower = round(summary_tbl[8]*initial_investment,2)
ci_upper = round(summary_tbl[9]*initial_investment,2)
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 30 years will end within the range of"
f" ${ci_lower} and ${ci_upper}")
```
## Optional Challenge - Early Retirement
### Five Years Retirement Option
```
# Configuring a Monte Carlo simulation to forecast 5 years cumulative returns
#Set number of simulations
num_sims = 500
MC_portfolio_5y = MCSimulation(
portfolio_data = df_portfolio_5_year,
weights = [0.8,0.2], #changed weights of stocks vs. bonds for a more aggressive/riskier portfolio
num_simulation = num_sims,
num_trading_days = 252*5
)
# Running a Monte Carlo simulation to forecast 5 years cumulative returns
MC_portfolio_5y.calc_cumulative_return()
# Plot simulation outcomes
line_plot_5y = MC_portfolio_5y.plot_simulation()
# Plot probability distribution and confidence intervals
dist_plot_5y = MC_portfolio_5y.plot_distribution()
# Fetch summary statistics from the Monte Carlo simulation results
summary_tbl_5y = MC_portfolio_5y.summarize_cumulative_return()
# Print summary statistics
print(summary_tbl_5y)
# Set initial investment
initial_investment = 20000 #same initial investment as the 30 year retirement plan
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $30,000
ci_lower_5y = round(summary_tbl_5y[8]*initial_investment,2)
ci_upper_5y = round(summary_tbl_5y[9]*initial_investment,2)
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 5 years will end within in the range of"
f" ${ci_lower_5y} and ${ci_upper_5y}")
```
### Ten Years Retirement Option
```
# Configuring a Monte Carlo simulation to forecast 10 years cumulative returns
#Set number of simulations
num_sims = 500
MC_portfolio_10y = MCSimulation(
portfolio_data = df_portfolio_5_year,
weights = [0.8,0.2], #changed weights of stocks vs. bonds for a more aggressive/riskier portfolio
num_simulation = num_sims,
num_trading_days = 252*10
)
# Running a Monte Carlo simulation to forecast 10 years cumulative returns
MC_portfolio_10y.calc_cumulative_return()
# Plot simulation outcomes
line_plot_10y = MC_portfolio_10y.plot_simulation()
# Plot probability distribution and confidence intervals
dist_plot_10y = MC_portfolio_10y.plot_distribution()
# Fetch summary statistics from the Monte Carlo simulation results
summary_tbl_10y = MC_portfolio_10y.summarize_cumulative_return()
# Print summary statistics
print(summary_tbl_10y)
# Set initial investment
initial_investment = 20000 #same initial investment as the 30 year retirement plan
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $30,000
ci_lower_10y = round(summary_tbl_10y[8]*initial_investment,2)
ci_upper_10y = round(summary_tbl_10y[9]*initial_investment,2)
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 10 years will end within in the range of"
f" ${ci_lower_10y} and ${ci_upper_10y}")
```
| github_jupyter |
<a href="https://colab.research.google.com/github/fidelis-eng/it-cert-automation-practice/blob/master/C2/W1/assignment/C2W1_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Week 1: Using CNN's with the Cats vs Dogs Dataset
Welcome to the 1st assignment of the course! This week, you will be using the famous `Cats vs Dogs` dataset to train a model that can classify images of dogs from images of cats. For this, you will create your own Convolutional Neural Network in Tensorflow and leverage Keras' image preprocessing utilities.
You will also create some helper functions to move the images around the filesystem so if you are not familiar with the `os` module be sure to take a look a the [docs](https://docs.python.org/3/library/os.html).
Let's get started!
```
import os
import zipfile
import random
import shutil
import tensorflow as tf
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from shutil import copyfile
import matplotlib.pyplot as plt
```
Download the dataset from its original source by running the cell below.
Note that the `zip` file that contains the images is unzipped under the `/tmp` directory.
```
# If the URL doesn't work, visit https://www.microsoft.com/en-us/download/confirmation.aspx?id=54765
# And right click on the 'Download Manually' link to get a new URL to the dataset
# Note: This is a very large dataset and will take some time to download
!wget --no-check-certificate \
"https://download.microsoft.com/download/3/E/1/3E1C3F21-ECDB-4869-8368-6DEBA77B919F/kagglecatsanddogs_3367a.zip" \
-O "/tmp/cats-and-dogs.zip"
local_zip = '/tmp/cats-and-dogs.zip'
zip_ref = zipfile.ZipFile(local_zip, 'r')
zip_ref.extractall('/tmp')
zip_ref.close()
```
Now the images are stored within the `/tmp/PetImages` directory. There is a subdirectory for each class, so one for dogs and one for cats.
```
source_path = '/tmp/PetImages'
source_path_dogs = os.path.join(source_path, 'Dog')
source_path_cats = os.path.join(source_path, 'Cat')
# os.listdir returns a list containing all files under the given path
print(f"There are {len(os.listdir(source_path_dogs))} images of dogs.")
print(f"There are {len(os.listdir(source_path_cats))} images of cats.")
```
**Expected Output:**
```
There are 12501 images of dogs.
There are 12501 images of cats.
```
You will need a directory for cats-v-dogs, and subdirectories for training
and testing. These in turn will need subdirectories for 'cats' and 'dogs'. To accomplish this, complete the `create_train_test_dirs` below:
```
# Define root directory
root_dir = '/tmp/cats-v-dogs'
# Empty directory to prevent FileExistsError is the function is run several times
if os.path.exists(root_dir):
shutil.rmtree(root_dir)
# GRADED FUNCTION: create_train_test_dirs
def create_train_test_dirs(root_path):
### START CODE HERE
# HINT:
# Use os.makedirs to create your directories with intermediate subdirectories
# Don't hardcode the paths. Use os.path.join to append the new directories to the root_path parameter
dir = ['training', 'testing', 'training/cats', 'testing/cats', 'training/dogs', 'testing/dogs']
for index,f in enumerate(dir):
path = os.path.join(root_path, f)
os.makedirs(path)
### END CODE HERE
try:
create_train_test_dirs(root_path=root_dir)
except FileExistsError:
print("You should not be seeing this since the upper directory is removed beforehand")
# Test your create_train_test_dirs function
for rootdir, dirs, files in os.walk(root_dir):
for subdir in dirs:
print(os.path.join(rootdir, subdir))
```
**Expected Output (directory order might vary):**
``` txt
/tmp/cats-v-dogs/training
/tmp/cats-v-dogs/testing
/tmp/cats-v-dogs/training/cats
/tmp/cats-v-dogs/training/dogs
/tmp/cats-v-dogs/testing/cats
/tmp/cats-v-dogs/testing/dogs
```
Code the `split_data` function which takes in the following arguments:
- SOURCE: directory containing the files
- TRAINING: directory that a portion of the files will be copied to (will be used for training)
- TESTING: directory that a portion of the files will be copied to (will be used for testing)
- SPLIT SIZE: to determine the portion
The files should be randomized, so that the training set is a random sample of the files, and the test set is made up of the remaining files.
For example, if `SOURCE` is `PetImages/Cat`, and `SPLIT` SIZE is .9 then 90% of the images in `PetImages/Cat` will be copied to the `TRAINING` dir
and 10% of the images will be copied to the `TESTING` dir.
All images should be checked before the copy, so if they have a zero file length, they will be omitted from the copying process. If this is the case then your function should print out a message such as `"filename is zero length, so ignoring."`. **You should perform this check before the split so that only non-zero images are considered when doing the actual split.**
Hints:
- `os.listdir(DIRECTORY)` returns a list with the contents of that directory.
- `os.path.getsize(PATH)` returns the size of the file
- `copyfile(source, destination)` copies a file from source to destination
- `random.sample(list, len(list))` shuffles a list
```
# GRADED FUNCTION: split_data
def split_data(SOURCE, TRAINING, TESTING, SPLIT_SIZE):
### START CODE HERE
splitSize = 0.0
for f in os.listdir(SOURCE):
splitSize +=1.0
splitSize = (splitSize * SPLIT_SIZE) - SPLIT_SIZE
files = []
sumdata = 0
for f in os.listdir(SOURCE):
if os.path.getsize(SOURCE + f) <= 0:
print('{} is zero length, so ignoring.'.format(f))
continue
elif splitSize > sumdata:
files.append(f)
sumdata +=1
else:
copyfile(SOURCE+f, TESTING+f)
sumdata +=1
random.sample(files, len(files))
for f in files:
copyfile(SOURCE+f, TRAINING+f)
### END CODE HERE
# Test your split_data function
# Define paths
CAT_SOURCE_DIR = "/tmp/PetImages/Cat/"
DOG_SOURCE_DIR = "/tmp/PetImages/Dog/"
TRAINING_DIR = "/tmp/cats-v-dogs/training/"
TESTING_DIR = "/tmp/cats-v-dogs/testing/"
TRAINING_CATS_DIR = os.path.join(TRAINING_DIR, "cats/")
TESTING_CATS_DIR = os.path.join(TESTING_DIR, "cats/")
TRAINING_DOGS_DIR = os.path.join(TRAINING_DIR, "dogs/")
TESTING_DOGS_DIR = os.path.join(TESTING_DIR, "dogs/")
# Empty directories in case you run this cell multiple times
if len(os.listdir(TRAINING_CATS_DIR)) > 0:
for file in os.scandir(TRAINING_CATS_DIR):
os.remove(file.path)
if len(os.listdir(TRAINING_DOGS_DIR)) > 0:
for file in os.scandir(TRAINING_DOGS_DIR):
os.remove(file.path)
if len(os.listdir(TESTING_CATS_DIR)) > 0:
for file in os.scandir(TESTING_CATS_DIR):
os.remove(file.path)
if len(os.listdir(TESTING_DOGS_DIR)) > 0:
for file in os.scandir(TESTING_DOGS_DIR):
os.remove(file.path)
# Define proportion of images used for training
split_size = .9
# Run the function
# NOTE: Messages about zero length images should be printed out
split_data(CAT_SOURCE_DIR, TRAINING_CATS_DIR, TESTING_CATS_DIR, split_size)
split_data(DOG_SOURCE_DIR, TRAINING_DOGS_DIR, TESTING_DOGS_DIR, split_size)
# Check that the number of images matches the expected output
print(f"\n\nThere are {len(os.listdir(TRAINING_CATS_DIR))} images of cats for training")
print(f"There are {len(os.listdir(TRAINING_DOGS_DIR))} images of dogs for training")
print(f"There are {len(os.listdir(TESTING_CATS_DIR))} images of cats for testing")
print(f"There are {len(os.listdir(TESTING_DOGS_DIR))} images of dogs for testing")
```
**Expected Output:**
```
666.jpg is zero length, so ignoring.
11702.jpg is zero length, so ignoring.
```
```
There are 11250 images of cats for training
There are 11250 images of dogs for training
There are 1250 images of cats for testing
There are 1250 images of dogs for testing
```
Now that you have successfully organized the data in a way that can be easily fed to Keras' `ImageDataGenerator`, it is time for you to code the generators that will yield batches of images, both for training and validation. For this, complete the `train_val_generators` function below.
Something important to note is that the images in this dataset come in a variety of resolutions. Luckily, the `flow_from_directory` method allows you to standarize this by defining a tuple called `target_size` that will be used to convert each image to this target resolution. **For this exercise, use a `target_size` of (150, 150)**.
**Note:** So far, you have seen the term `testing` being used a lot for referring to a subset of images within the dataset. In this exercise, all of the `testing` data is actually being used as `validation` data. This is not very important within the context of the task at hand but it is worth mentioning to avoid confusion.
```
# GRADED FUNCTION: train_val_generators
def train_val_generators(TRAINING_DIR, VALIDATION_DIR):
### START CODE HERE
# Instantiate the ImageDataGenerator class (don't forget to set the rescale argument)
train_datagen = ImageDataGenerator(rescale=(1./255))
# Pass in the appropiate arguments to the flow_from_directory method
train_generator = train_datagen.flow_from_directory(directory=TRAINING_DIR,
batch_size=20,
class_mode='binary',
target_size=(150, 150))
# Instantiate the ImageDataGenerator class (don't forget to set the rescale argument)
validation_datagen = ImageDataGenerator(rescale=(1./255))
# Pass in the appropiate arguments to the flow_from_directory method
validation_generator = validation_datagen.flow_from_directory(directory=VALIDATION_DIR,
batch_size=20,
class_mode='binary',
target_size=(150, 150))
### END CODE HERE
return train_generator, validation_generator
# Test your generators
train_generator, validation_generator = train_val_generators(TRAINING_DIR, TESTING_DIR)
```
**Expected Output:**
```
Found 22498 images belonging to 2 classes.
Found 2500 images belonging to 2 classes.
```
One last step before training is to define the architecture of the model that will be trained.
Complete the `create_model` function below which should return a Keras' `Sequential` model.
Aside from defining the architecture of the model, you should also compile it so make sure to use a `loss` function that is compatible with the `class_mode` you defined in the previous exercise, which should also be compatible with the output of your network. You can tell if they aren't compatible if you get an error during training.
**Note that you should use at least 3 convolution layers to achieve the desired performance.**
```
from tensorflow.keras.optimizers import RMSprop
# GRADED FUNCTION: create_model
def create_model():
# DEFINE A KERAS MODEL TO CLASSIFY CATS V DOGS
# USE AT LEAST 3 CONVOLUTION LAYERS
### START CODE HERE
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(32, (3,3), activation='relu', input_shape=(150,150,3)),
tf.keras.layers.MaxPooling2D((2,2)),
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D((2,2)),
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D((2,2)),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(optimizer=RMSprop(learning_rate=0.001),
loss='binary_crossentropy',
metrics=['accuracy'])
### END CODE HERE
return model
model = create_model()
model.summary()
```
Now it is time to train your model!
**Note:** You can ignore the `UserWarning: Possibly corrupt EXIF data.` warnings.
```
# Get the untrained model
model = create_model()
# Train the model
# Note that this may take some time.
history = model.fit(train_generator,
epochs=15,
verbose=1,
validation_data=validation_generator)
```
Once training has finished, you can run the following cell to check the training and validation accuracy achieved at the end of each epoch.
**To pass this assignment, your model should achieve a training accuracy of at least 95% and a validation accuracy of at least 80%**. If your model didn't achieve these thresholds, try training again with a different model architecture and remember to use at least 3 convolutional layers.
```
#-----------------------------------------------------------
# Retrieve a list of list results on training and test data
# sets for each training epoch
#-----------------------------------------------------------
acc=history.history['accuracy']
val_acc=history.history['val_accuracy']
loss=history.history['loss']
val_loss=history.history['val_loss']
epochs=range(len(acc)) # Get number of epochs
#------------------------------------------------
# Plot training and validation accuracy per epoch
#------------------------------------------------
plt.plot(epochs, acc, 'r', "Training Accuracy")
plt.plot(epochs, val_acc, 'b', "Validation Accuracy")
plt.title('Training and validation accuracy')
plt.show()
print("")
#------------------------------------------------
# Plot training and validation loss per epoch
#------------------------------------------------
plt.plot(epochs, loss, 'r', "Training Loss")
plt.plot(epochs, val_loss, 'b', "Validation Loss")
plt.show()
```
You will probably encounter that the model is overfitting, which means that it is doing a great job at classifying the images in the training set but struggles with new data. This is perfectly fine and you will learn how to mitigate this issue in the upcoming week.
Before downloading this notebook and closing the assignment, be sure to also download the `history.pkl` file which contains the information of the training history of your model. You can download this file by running the cell below:
```
def download_history():
import pickle
from google.colab import files
with open('history.pkl', 'wb') as f:
pickle.dump(history.history, f)
files.download('history.pkl')
download_history()
```
You will also need to submit this notebook for grading. To download it, click on the `File` tab in the upper left corner of the screen then click on `Download` -> `Download .ipynb`. You can name it anything you want as long as it is a valid `.ipynb` (jupyter notebook) file.
**Congratulations on finishing this week's assignment!**
You have successfully implemented a convolutional neural network that classifies images of cats and dogs, along with the helper functions needed to pre-process the images!
**Keep it up!**
| github_jupyter |
```
from datetime import datetime, timezone
import pandas as pd
import langdetect
import json
lang_code = pd.read_json('ISO-639-1-language.json')
def get_language(text=None):
langs = []
if text:
langs = [{
"name": lang_code.set_index('code').loc[lang.lang, 'name'],
"code": lang.lang.upper()
} for lang in langdetect.detect_langs(text)]
return langs
def get_software(data):
software = [dict(
name=lang.get('node', {}).get('name', ''),
version="",
library=(
[] if lang.get('node', {}).get('name', '') not in ['R', 'Python'] else
data.get('r_libs', []) if lang.get('node', {}).get('name', '') == 'R' else
data.get('py_libs', []))
) for lang in data.get('languages', [])]
return software
def has_content(node):
if isinstance(node, dict):
unode = {}
for key, value in node.items():
v = has_content(value)
if isinstance(v, int) or v: # This is placed here to prevent the `published` parameter from being removed in case `published` is set to 0.
unode[key] = v
elif isinstance(node, list):
unode = []
for element in node:
v = has_content(element)
if isinstance(v, int) or v: # This is placed here to prevent the `published` parameter from being removed in case `published` is set to 0.
unode.append(v)
else: # str, int, float <for now, assume no other container data types, e.g., tuples.>
unode = node
return unode
import pymongo
mongo = pymongo.MongoClient(port=27018)
db = mongo['nlp']
collection = db['github-bq']
collection.create_index([('readme', pymongo.TEXT)], name='readme_text_idx')
# collection.create_index(('description', pymongo.TEXT), name='description_text_idx')
```
# Fill the script template
```
def build_template_for_github(data, overwrite='no', published=1, repositoryid='central', cleanup=False):
title_idno = data['_id'].replace('/', '_')
tp_template = dict(
repositoryid=repositoryid,
published=published,
overwrite=overwrite,
doc_desc=dict(
title="",
idno="",
producers=[
dict(
name= "GitHub Bot",
abbr="",
affiliation="",
role="bot"
)
],
prod_date=datetime.now().strftime('%d %B %Y'),
version=""
),
project_desc=dict(
title_statement=dict(
idno=title_idno,
title=data.get('description', data.get('name', '')),
sub_title="",
alternate_title="",
translated_title=""
),
production_date=[
# pd.to_datetime(data['repo_created_at']).strftime('%d %B %Y')
pd.to_datetime(data['repo_created_at']).strftime('%B %Y')
],
geographic_units=[
dict(
name="",
code="",
type=""
)
],
authoring_entity=[
dict(
name=data['owner'],
role="owner",
affiliation=data.get('homepage_url', ''),
abbreviation="",
email=""
)
],
contributors=[
dict(
name="",
role="",
affiliation="",
abbreviation="",
email="",
url=""
)
],
curators= [
dict(
name= "",
role= "",
affiliation= "",
abbreviation= "",
email= "",
url= ""
)
],
abstract=data.get('readme', data.get('description', data.get('name', ''))),
keywords=[
dict(
name="",
vocabulary="",
uri=""
)
],
themes=[
dict(
name="",
vocabulary="",
uri=""
)
],
topics=[
dict(
id="",
name="",
parent_id="",
vocabulary="",
uri=""
)
],
disciplines=[
dict(
name="",
vocabulary="",
uri=""
)
],
output_types=[
dict(
type="",
description="",
uri="",
doi=""
)
],
repository_uri=[
dict(
name=data['_id'],
type="Github",
uri=f"https://github.com/{data['_id']}"
)
],
project_website=[
data.get('homepage_url', '')
],
version_statement=dict(
version="latest",
version_date=pd.to_datetime(data['repo_updated_at']).strftime('%d %B %Y'),
version_resp="",
version_notes="Latest update"
),
language=get_language(data.get('readme', data.get('description', data.get('name', 'english')))),
methods=[
dict(
name="",
note=""
)
],
software=get_software(data),
technology_environment="",
technology_requirements="",
reproduction_instructions="",
license=[
dict(
name=data.get('license_info', ''),
uri=""
)
],
review_process=[
dict(
submission_date="",
reviewer="",
review_status="",
approval_authority="",
approval_date=""
)
],
disclaimer="",
confidentiality="",
citation_requirement="",
datasets=[
dict(
name="",
idno="",
note="",
access_type="",
uri=""
)
],
sponsors=[
dict(
name="",
abbr="",
role="",
grant_no=""
)
],
acknowledgements=[
dict(
name="",
affiliation="",
role=""
)
],
related_projects=[
dict(
name="",
uri="",
note=""
)
],
contacts=[
dict(
name="",
affiliation="",
uri="",
phone=""
)
],
scripts=[
dict(
file_name="",
title="",
authors=[
dict(
name="",
abbr="",
role=""
)
],
date="",
format="",
software= "",
description= "",
methods= "",
dependencies= "",
instructions= "",
source_code_repo= "",
notes= ""
)
]
)
)
if cleanup:
tp_template = has_content(tp_template)
return tp_template
```
# Get data from the database
```
collection.count_documents({'$text': {'$search': '"economic"'}, 'readme': {'$exists': True}})
%%time
keywords = ['economic', 'nutrition', 'income inequality', 'agriculture', 'climate change', 'poverty', 'fragility', 'refugee']
payloads = []
for kw in keywords:
for data in collection.find({'$text': {'$search': f'"{kw}"'}, 'readme': {'$exists': True}}):
payloads.append(build_template_for_github(data, overwrite='yes', cleanup=True))
with open('github_nada_data.json', 'w') as fl:
json.dump(payloads, fl)
# climate_dataset = collection.find({'$text': {'$search': '"climate change"'}, 'readme': {'$exists': True}})
# poverty_dataset = collection.find({'$text': {'$search': '"poverty"'}, 'readme': {'$exists': True}})
# nutrition_dataset = collection.find({'$text': {'$search': '"nutrition"'}, 'readme': {'$exists': True}})
# refugee_dataset = collection.find({'$text': {'$search': '"refugee"'}, 'readme': {'$exists': True}})
# fragility_dataset = collection.find({'$text': {'$search': '"fragility"'}, 'readme': {'$exists': True}})
# agriculture_dataset = collection.find({'$text': {'$search': '"agriculture"'}, 'readme': {'$exists': True}})
# income_dataset = collection.find({'$text': {'$search': '"income inequality"'}, 'readme': {'$exists': True}})
# economics_dataset = collection.find({'$text': {'$search': '"economics"'}, 'readme': {'$exists': True}})
# data = collection.find_one({'primary_language': 'Python', '$text': {'$search': '"climate change"'}})
# data = collection.find_one({'primary_language': 'Python', '$text': {'$search': 'poverty'}})
# tp_template = build_template_for_github(data, cleanup=True)
# tp_template['doc_desc']
# template = {
# "repositoryid": 'central',
# "published": 1,
# "overwrite": "yes",
# "doc_desc": {
# # "title": "",
# "idno": "",
# "producers": [
# {
# "name": "GitHub Bot",
# "abbr": "",
# "affiliation": "",
# "role": "bot"
# }
# ],
# "prod_date": datetime.now().strftime('%d %B %Y'),
# "version": "1.0"
# },
# "project_desc": {
# "title_statement": {
# "idno": data['_id'].replace('/', '_'),
# "title": data.get('description', data.get('name', '')),
# "sub_title": "",
# "alternate_title": "",
# "translated_title": ""
# },
# "production_date": [
# datetime.now().strftime('%B %Y')
# ],
# "authoring_entity": [
# {
# "name": data['owner'],
# "role": "owner",
# "affiliation": data.get('homepage_url', ''),
# }
# ],
# "abstract": data.get('readme', data.get('description', data.get('name', ''))),
# "repository_uri": [
# {
# "name": data['_id'],
# "type": "GitHub",
# "uri": f"https://github.com/{data['_id']}"
# }
# ],
# "project_website": [
# data.get('homepage_url', '')
# ],
# "version_statement": {
# "version": "latest",
# "version_date": pd.to_datetime(data['repo_updated_at']).strftime('%d %B %Y'),
# "version_resp": "",
# "version_notes": "Latest update"
# },
# "language": get_language(data.get('readme', data.get('description', data.get('name', 'english')))),
# "software": [
# {
# "name": lang.get('node', {}).get('name', ''),
# "version": "",
# "library": (
# [] if lang.get('node', {}).get('name', '') not in ['R', 'Python'] else
# data.get('r_libs', []) if lang.get('node', {}).get('name', '') == 'R' else
# data.get('py_libs', []))
# } for lang in data.get('languages', [])
# ],
# "license": [
# {
# "name": data.get('license_info', ''),
# "uri": ""
# }
# ],
# }
# }
```
# Post to API
```
idno = data['project_desc']['title_statement']['idno'].replace('/', '_')
headers = {'X-API-KEY': '<API_KEY>'}
api_url = 'http://dev.ihsn.org/nada/index.php/api/datasets/create/script/'
response = requests.post(api_url + idno, headers=headers, json=template)
# collection.find_one({'primary_language': 'R', '$text': {'$search': 'poverty'}})
# collection.find({'$text': {'$search': "\"climate change\""}})
# data = {
# "_id": "00tau/skyline-addon-easyqc",
# "description": "Add-on script for performing easy quality control tasks within Skyline",
# "fork_count": 0,
# "insertion_date": "2019-11-24T04:36:07.963844+00:00",
# "languages": [
# {
# "node": {
# "name": "R"
# }
# }
# ],
# "last_updated_date": "2019-11-24T04:36:07.963844+00:00",
# "license_info": "GNU General Public License v3.0",
# "name": "skyline-addon-easyqc",
# "owner": "00tau",
# "primary_language": "R",
# "py_libs": [],
# "r_libs": [
# "chron",
# "ggplot2",
# "plyr"
# ],
# "readme": "# Start using easyQC for statistical process and quality control in mass spectrometry workflows\n\n## Introduction\n\nThe program `easyQC` is an external tool for statistical process and quality\ncontrol in mass spectrometry workflows that integrates nicely in the [Skyline\nTargeted Proteomics\nEnvironment](https://skyline.gs.washington.edu/labkey/project/home/software/Skyline/begin.view).\n\n## Feature list at a glance\n\n- Automatically sorts your data by date and time, and orders your observations\n with the most recent on the right. (\"What? Does this mean I don't need to\n sort my data manually, as it is the case for some other software tools out\n there?\", \"Yes.\")\n- Dynamically adapts to custom report templates. (See details below.)\n- Flow charts for single peptides can optionally be grouped together by their\n common protein accession. (See details below.)\n- Plots are generated in a nice page layout, ready for printing.\n- Observations are colour-coded by a beneficial four-colour-code. This makes\n it particularly easy to detect deviations from the norm.\n- Has a built in outlier detection, which provides you with useful robust\n features. (See details below.)\n- Plot as _many_ flow charts for as _many_ peptides as you like.\n\n## How to cite this software\n\nThe [Harvard UoB format]\n(http://lrweb.beds.ac.uk/guides/a-guide-to-referencing/cite_computer_program)\nsuggests to cite this software in the following fashion:\n\n Möbius, T.W. and Malchow, S. (2014) easyQC: Statistical Process and Quality\n Control in Mass Spectrometry Workflows (Version 1.0) [Computer program].\n Available at: http://00tau.github.io/skyline-addon-easyqc/ (Accessed 03.\n April, 2014)\n\nThank you for using (and citing) this software.\n\n## Installation using the skyline GUI\n\nSimply follow the GUI-clicking adventure by successively clicking on `Tools ->\nExternal Tools -> External Tool Store`. In the appearing list select (click\non) `easyQC`. You will be promoted for the path to `Rscript`, which needs to\nbe installed on you system.\n\nWe have realised that since the introduction of \"Live Reports\" in new Versions\nof Skyline, the import of new templates might fail. If this is the case for\nyou, make sure two switch off \"Live Reports\", restart Skyline, and try the\ninstallation again.\n\nThe underlying code-base of `easyQC` relies on the R-packages\n[ggplot2](http://ggplot2.org/), [plyr](http://plyr.had.co.nz/) and\n[chron](http://cran.r-project.org/web/packages/chron/index.html). Fortunately,\nall these packages are hosted on [CRAN](http://cran.r-project.org/), and should\nautomatically be installed into your R-environment, when installing `easyQC` in\nSkyline. If, for some reasons, this should not be the case for you, make sure\nthese three packages are installed in your R-environment.\n\n## Description\n\nThe software comes with an exemplary report template called `easyQC`. We\nrecommend to just go with this template, but feel free to create your own. The\nabsolute necessary fields your template should contain are:\n`PeptideModifiedSequence` and `PrecursorMz`. These two fields are used as\nidentifiers for your peptides, and, thus, all other fields should uniquely be\nidentifiable by these two. Optionally, the field `ProteinName` can be added to\nyour template.\n\nBy default, the flow charts of ten peptides are grouped together into one plot\neach. If your report template also contains the associated protein accession\nof each peptide, namely the field `ProteinName`, then all peptides which belong\nto the same protein accession are grouped into one plot.\n\nBefore the calculation of the mean and standard deviations of each flow chart,\nthe software will do some outlier detection of your data, namely [Grubbs' test\nfor outliers](http://en.wikipedia.org/wiki/Grubbs%27_test_for_outliers) will be\napplied. Observations which are classified as outliers by this test are\ndiscarded in the estimation of the mean and standard deviations. This gives\nthe estimated means and standard deviations some desirable\n[robust](http://en.wikipedia.org/wiki/Robust_statistics) features.\n\n## You can also use easyQC as a stand-alone command line program\n\nOn Linux, you simply need to add the directory in which you have cloned\n`easyQC`'s repository to your path. Also make sure that `easyQC.r` is\nexecutable.\n\n```\n% git clone https://github.com/00tau/skyline-addon-easyqc.git\n% cd skyline-addonn-easyqc\n% chmod +x easyQC.r\n% PATH=$(pwd):$PATH\n```\n\nThe synopsis is as follows:\n\n```\neasyQC.r [OPTIONS] REPORTFILE\n```\n\nWhere `OPTIONS` is either `verbose` or noting. For example, to produce some\nquality control plots from a file `some-report-file.csv` that has been\ngenerated by Skyline via some report template (e.g. the template `easyQC.skyr`\nshould come in mind here), run either one of the following two code lines from\nthe command line.\n\n```\n% easyQC.r some-report-file.csv\n% easyQC.r verbose some-report-file.csv\n```\n\nThis will produce a file `some-report-file.pdf` with all the plots you need.\n\nYou what to install the most recent and latest version in Skyline\n-----------------------------------------------------------------\n\nIf for some reasons, you are interested in installing the latest GitHub-version\n(or any other version of this software that is available on GitHub), the\nrepository contains a convenient Makefile that will create the necessary files\nfor the installation process for you. Simply type:\n\n```\n% make\n```\n\nThis will create a `easyQC.zip` file which contains the needed install scripts\nfor Skyline. Now, just follow your Skyline-GUI.\n\nAuthors\n-------\n\nThomas W. D. Möbius (Maintainer, R-programming), Sebastian Malchow (Skyline wizard)\n",
# "repo_created_at": "2014-02-25T15:26:30Z",
# "repo_id": "MDEwOlJlcG9zaXRvcnkxNzE3NzYxOQ==",
# "repo_updated_at": "2014-04-04T14:56:54Z",
# "stargazers": 0,
# "topics": [],
# "watchers": 1
# }
# collection.create_index([('readme', pymongo.TEXT)], name='readme_text_idx')
# collection.create_index([('description', pymongo.TEXT)], name='description_text_idx')
# template = {
# "repositoryid": 'central',
# "published": 1,
# "overwrite": "yes",
# "doc_desc": {
# # "title": "",
# "idno": "",
# "producers": [
# {
# "name": "GitHub Bot",
# "abbr": "",
# "affiliation": "",
# "role": "bot"
# }
# ],
# "prod_date": datetime.now().strftime('%B %Y'),
# "version": ""
# },
# "project_desc": {
# "title_statement": {
# "idno": data['_id'].replace('/', '_'),
# "title": data.get('description', data.get('name', '')),
# "sub_title": "",
# "alternate_title": "",
# "translated_title": ""
# },
# "production_date": [
# datetime.now().strftime('%B %Y')
# ],
# # "geographic_units": [
# # {
# # "name": "",
# # "code": "",
# # "type": ""
# # }
# # ],
# "authoring_entity": [
# {
# "name": data['owner'],
# "role": "owner",
# "affiliation": data.get('homepage_url', ''),
# # "abbreviation": null,
# # "email": null
# }
# ],
# # "contributors": [
# # {
# # "name": "string",
# # "role": "string",
# # "affiliation": "string",
# # "abbreviation": null,
# # "email": null,
# # "url": null
# # }
# # ],
# # "curators": [
# # {
# # "name": "string",
# # "role": "string",
# # "affiliation": "string",
# # "abbreviation": null,
# # "email": null,
# # "url": null
# # }
# # ],
# "abstract": data.get('readme', data.get('description', data.get('name', ''))),
# # "keywords": [
# # {
# # "name": "string",
# # "vocabulary": "string",
# # "uri": "string"
# # }
# # ],
# # "themes": [
# # {
# # "name": "string",
# # "vocabulary": "string",
# # "uri": "string"
# # }
# # ],
# # "topics": [
# # {
# # "id": "string",
# # "name": "string",
# # "parent_id": "string",
# # "vocabulary": "string",
# # "uri": "string"
# # }
# # ],
# # "disciplines": [
# # {
# # "name": "string",
# # "vocabulary": "string",
# # "uri": "string"
# # }
# # ],
# # "output_types": [
# # {
# # "type": "string",
# # "description": "string",
# # "uri": "string",
# # "doi": "string"
# # }
# # ],
# "repository_uri": [
# {
# "name": data['_id'],
# "type": "GitHub",
# "uri": f"https://github.com/{data['_id']}"
# }
# ],
# "project_website": [
# data.get('homepage_url', '')
# ],
# "version_statement": {
# "version": "latest",
# "version_date": pd.to_datetime(data['repo_updated_at']).strftime('%d %B %Y'),
# "version_resp": "",
# "version_notes": "Latest update"
# },
# "language": get_language(data.get('readme', data.get('description', data.get('name', 'english')))),
# # "methods": [
# # {
# # "name": "string",
# # "note": "string"
# # }
# # ],
# "software": [
# {
# "name": lang.get('node', {}).get('name', ''),
# "version": "",
# "library": (
# [] if lang.get('node', {}).get('name', '') not in ['R', 'Python'] else
# data.get('r_libs', []) if lang.get('node', {}).get('name', '') == 'R' else
# data.get('py_libs', []))
# } for lang in data.get('languages', [])
# ],
# # "technology_environment": "string",
# # "technology_requirements": "string",
# # "reproduction_instructions": "string",
# "license": [
# {
# "name": data.get('license_info', ''),
# "uri": ""
# }
# ],
# # "review_process": [
# # {
# # "submission_date": "string",
# # "reviewer": "string",
# # "review_status": "string",
# # "approval_authority": "string",
# # "approval_date": "string"
# # }
# # ],
# # "disclaimer": "string",
# # "confidentiality": "string",
# # "citation_requirement": "string",
# # "datasets": [
# # {
# # "name": "string",
# # "idno": "string",
# # "note": "string",
# # "access_type": "string",
# # "uri": "string"
# # }
# # ],
# # "sponsors": [
# # {
# # "name": "string",
# # "abbr": "string",
# # "role": "string",
# # "grant_no": "string"
# # }
# # ],
# # "acknowledgements": [
# # {
# # "name": "string",
# # "affiliation": "string",
# # "role": "string"
# # }
# # ],
# # "related_projects": [
# # {
# # "name": "string",
# # "uri": "string",
# # "note": "string"
# # }
# # ],
# # "contacts": [
# # {
# # "name": "string",
# # "affiliation": "string",
# # "uri": "string",
# # "phone": "string"
# # }
# # ],
# # "scripts": [
# # {
# # "file_name": "string",
# # "title": "string",
# # "authors": [
# # {
# # "name": "string",
# # "abbr": "string",
# # "role": "string"
# # }
# # ],
# # "date": "string",
# # "format": "string",
# # "software": "string",
# # "description": "string",
# # "methods": "string",
# # "dependencies": "string",
# # "instructions": "string",
# # "source_code_repo": "string",
# # "notes": "string"
# # }
# # ]
# }
# }
```
| github_jupyter |
# Starbucks Capstone Challenge
### Introduction
This data set contains simulated data that mimics customer behavior on the Starbucks rewards mobile app. Once every few days, Starbucks sends out an offer to users of the mobile app. An offer can be merely an advertisement for a drink or an actual offer such as a discount or BOGO (buy one get one free). Some users might not receive any offer during certain weeks.
Not all users receive the same offer, and that is the challenge to solve with this data set.
Your task is to combine transaction, demographic and offer data to determine which demographic groups respond best to which offer type. This data set is a simplified version of the real Starbucks app because the underlying simulator only has one product whereas Starbucks actually sells dozens of products.
Every offer has a validity period before the offer expires. As an example, a BOGO offer might be valid for only 5 days. You'll see in the data set that informational offers have a validity period even though these ads are merely providing information about a product; for example, if an informational offer has 7 days of validity, you can assume the customer is feeling the influence of the offer for 7 days after receiving the advertisement.
You'll be given transactional data showing user purchases made on the app including the timestamp of purchase and the amount of money spent on a purchase. This transactional data also has a record for each offer that a user receives as well as a record for when a user actually views the offer. There are also records for when a user completes an offer.
Keep in mind as well that someone using the app might make a purchase through the app without having received an offer or seen an offer.
### Example
To give an example, a user could receive a discount offer buy 10 dollars get 2 off on Monday. The offer is valid for 10 days from receipt. If the customer accumulates at least 10 dollars in purchases during the validity period, the customer completes the offer.
However, there are a few things to watch out for in this data set. Customers do not opt into the offers that they receive; in other words, a user can receive an offer, never actually view the offer, and still complete the offer. For example, a user might receive the "buy 10 dollars get 2 dollars off offer", but the user never opens the offer during the 10 day validity period. The customer spends 15 dollars during those ten days. There will be an offer completion record in the data set; however, the customer was not influenced by the offer because the customer never viewed the offer.
### Cleaning
This makes data cleaning especially important and tricky.
You'll also want to take into account that some demographic groups will make purchases even if they don't receive an offer. From a business perspective, if a customer is going to make a 10 dollar purchase without an offer anyway, you wouldn't want to send a buy 10 dollars get 2 dollars off offer. You'll want to try to assess what a certain demographic group will buy when not receiving any offers.
### Final Advice
Because this is a capstone project, you are free to analyze the data any way you see fit. For example, you could build a machine learning model that predicts how much someone will spend based on demographics and offer type. Or you could build a model that predicts whether or not someone will respond to an offer. Or, you don't need to build a machine learning model at all. You could develop a set of heuristics that determine what offer you should send to each customer (i.e., 75 percent of women customers who were 35 years old responded to offer A vs 40 percent from the same demographic to offer B, so send offer A).
# Overview
Starbucks Corporation is an American multinational chain of coffeehouses and roastery reserves headquartered in Seattle, Washington.
Like many other other companies, Starbucks has offers of various types.
They give offers depending on demographics, so not all customers receive those offers, again depending on their demographics.
In this notebook, I am going to take through analysis that I have done about the startbuck datasets.
Most importantly, I will perform a machine learning algorithm to help predict the offer type given for a client
# Problem statement
The goal of this project if to predict the amount that a customers will spend, depending on the demographics and the orders.
1. Clean the data
2. Create a classifier
3. Evaluate the model
4. Improve the model using an other algorithm
5. Re-evaluate the model
The final algorithm should be able to make accurate prediction about the type of offer that a client should receive.
# Metrics
The most common metric that is used in classification models is the **ACCURACY** and that is what we are going to use to evaluate our model.
Accuracy works fine in this case because we have a quite balance datasets for the three classes we are predicting for.
For instance if we had a model with two classes A and B and A represent 92% of the dataset and B 8%, computing the model's accuracy would easily give us an accuracy around 92%.
In those cases we could have used other methods like Logarithmic Loss confusion matrix.
# Data Exploration
### Data Sets
The data is contained in three files:
* portfolio.json - containing offer ids and meta data about each offer (duration, type, etc.)
* profile.json - demographic data for each customer
* transcript.json - records for transactions, offers received, offers viewed, and offers completed
Here is the schema and explanation of each variable in the files:
**portfolio.json**
* id (string) - offer id
* offer_type (string) - type of offer ie BOGO, discount, informational
* difficulty (int) - minimum required spend to complete an offer
* reward (int) - reward given for completing an offer
* duration (int) - time for offer to be open, in days
* channels (list of strings)
**profile.json**
* age (int) - age of the customer
* became_member_on (int) - date when customer created an app account
* gender (str) - gender of the customer (note some entries contain 'O' for other rather than M or F)
* id (str) - customer id
* income (float) - customer's income
**transcript.json**
* event (str) - record description (ie transaction, offer received, offer viewed, etc.)
* person (str) - customer id
* time (int) - time in hours since start of test. The data begins at time t=0
* value - (dict of strings) - either an offer id or transaction amount depending on the record
**Note:** If you are using the workspace, you will need to go to the terminal and run the command `conda update pandas` before reading in the files. This is because the version of pandas in the workspace cannot read in the transcript.json file correctly, but the newest version of pandas can. You can access the termnal from the orange icon in the top left of this notebook.
You can see how to access the terminal and how the install works using the two images below. First you need to access the terminal:
<img src="pic1.png"/>
Then you will want to run the above command:
<img src="pic2.png"/>
Finally, when you enter back into the notebook (use the jupyter icon again), you should be able to run the below cell without any errors.
```
import pandas as pd
import numpy as np
import math
import json
from matplotlib import pyplot as plt
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.ensemble import RandomForestRegressor
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report, confusion_matrix, accuracy_score
from sklearn.tree import DecisionTreeClassifier
from sklearn.preprocessing import MinMaxScaler
% matplotlib inline
# read in the json files
portfolio = pd.read_json('data/portfolio.json', orient='records', lines=True)
profile = pd.read_json('data/profile.json', orient='records', lines=True)
transcript = pd.read_json('data/transcript.json', orient='records', lines=True)
profile.head()
profile.describe()
profile.hist(bins = 100)
transcript.head()
transcript.describe()
transcript.hist(bins = 100)
portfolio.head()
portfolio.hist(bins = 20)
# Finding the number of unique clients in this data set
print(len(profile['id'].unique()))
# Finding the unique offers in the data set
print(len(portfolio['id'].unique()))
# average difficulty to complete an offer
portfolio['difficulty'].mean()
# average rewards for offers
portfolio['reward'].mean()
# Unique events
transcript['event'].unique()
# Unique offer type in the data set
portfolio['offer_type'].unique()
# Average duration for the offers
portfolio['duration'].mean()
```
# Exploratory visualizations
Below are visuals that can help us better understand this dataset
```
# Distribution of the ages
# Creating histogram
fig, ax = plt.subplots(figsize =(10, 7))
ax.hist(profile['age'], bins = 20)
# Show plot
plt.xlabel('Ages')
plt.ylabel('Counts')
plt.title('Distribution of ages')
plt.show()
transcript.shape[0]
# visualize count per event
x_label = transcript['event'].unique()
y_label = transcript['event'].value_counts().values
# plotting a bar chart
plt.bar(x_label, y_label, width = 0.8, color = ['red', 'green'])
# naming the x-axis
plt.xlabel('Offer types')
# naming the y-axis
plt.ylabel('Counts')
# plot title
plt.title('Offer status')
# function to show the plot
plt.show()
profile['gender'].value_counts()
# Visualize gender repartition
# Creating plot
labels = ['Female', 'Male', 'Other']
data = profile['gender'].value_counts().values
fig = plt.figure(figsize =(10, 7))
plt.pie(data, labels = labels)
# show plot
plt.show()
```
# Data Preprocessing
The below function helps us put together a dataframe that we will use for the training and testing our datasets.
We have three datasets and we want to combine them into one.
This is what we want the final dataframe to look like:
Event, Time, Offer_id, Amount, Reward, Age_group, Gender,Income
```
profile.head(1)
portfolio.head(1)
transcript.head(1)
transcript['value'].unique
```
We want to be able to contruct a dataframe that has all the listed columns to be to train.
The transcript dataframe will be the main the start.
The dataframe has offer_id marked as values, and we first want to be able to extract those from the value column
```
# Extracting offer_id from the transcript df
transcript['offer_id'] = ''
transcript['amount'] = 0
transcript['reward'] = 0
for idx, row in transcript.iterrows():
for k in row['value']:
if k == 'offer_id' or k == 'offer id':
transcript.at[idx, 'offer_id']= row['value'][k]
if k == 'amount':
transcript.at[idx, 'amount']= row['value'][k]
if k == 'reward':
transcript.at[idx, 'reward']= row['value'][k]
transcript.head()
# We then drop the values column since we don't need it anymore
transcript = transcript.drop(['value'] , axis =1)
transcript.head()
# Now that we have the id, let's get the offer type
def extract_offer_type(offer_id):
'''
offer_id : the offer id you want to find the type for
'''
try:
offer_type = portfolio[portfolio['id'] == offer_id]['offer_type'].values[0]
return offer_type
except:
offer_type = 'NA'
return offer_type
transcript['offer_type'] = transcript.apply(lambda x: extract_offer_type(x['offer_id']), axis=1)
```
There above, we said that we wanted to have an age group column, which does not exist, so let create it
```
# Creating the age group group column
profile['age_groups'] = pd.cut(profile.age, bins=[0, 12, 18, 21, 64, 200],
labels=['children', 'infant', 'Teens', 'adults', 'elderly'])
# dropping the age columns
profile = profile.drop(['age'], axis = 1)
profile.head()
```
Now that we have our dataframe almost ready, let's start combinining them to create the final dataframe
```
# We are predicting for the pffer type so we do not want row that do not have the offer_id
transcript = transcript[transcript['offer_id'] != 'NA']
def add_gender(profile_id):
gender = profile[profile['id'] == profile_id]['gender'].values[0]
return gender
transcript['gender'] = transcript.apply(lambda x: add_gender(x['person']), axis=1)
def add_income(profile_id):
income = profile[profile['id'] == profile_id]['income'].values[0]
return income
transcript['income'] = transcript.apply(lambda x: add_income(x['person']), axis=1)
def add_age_group(profile_id):
age_group = profile[profile['id'] == profile_id]['age_groups'].values[0]
return age_group
transcript['age_group'] = transcript.apply(lambda x: add_age_group(x['person']), axis=1)
transcript.head()
#Let's now transform our features into classes
labels_event = transcript['event'].astype('category').cat.categories.tolist()
replace_map_comp_event = {'event' : {k: v for k,v in zip(labels_event,list(range(1,len(labels_event)+1)))}}
print(replace_map_comp_event)
labels_offer_id = transcript['offer_id'].astype('category').cat.categories.tolist()
replace_map_comp_offer_id = {'offer_id' : {k: v for k,v in zip(labels_offer_id,list(range(1,len(labels_offer_id)+1)))}}
print(replace_map_comp_offer_id)
labels_age_group = transcript['age_group'].astype('category').cat.categories.tolist()
replace_map_comp_age_group = {'age_group' : {k: v for k,v in zip(labels_age_group,list(range(1,len(labels_age_group)+1)))}}
print(replace_map_comp_age_group)
labels_gender = transcript['gender'].astype('category').cat.categories.tolist()
replace_map_comp_gender = {'gender' : {k: v for k,v in zip(labels_gender,list(range(1,len(labels_gender)+1)))}}
print(replace_map_comp_gender)
# Now let tranform the target which is the offer type col
labels_offer_type = transcript['offer_type'].astype('category').cat.categories.tolist()
replace_map_comp_offer_type = {'offer_type' : {k: v for k,v in zip(labels_offer_type,list(range(1,len(labels_offer_type)+1)))}}
print(replace_map_comp_offer_type)
# For all the features transformed up there, we will turn them into numerical values
transcript.replace(replace_map_comp_event, inplace=True)
transcript.replace(replace_map_comp_offer_id, inplace=True)
transcript.replace(replace_map_comp_age_group, inplace=True)
transcript.replace(replace_map_comp_gender, inplace=True)
transcript.replace(replace_map_comp_offer_type, inplace=True)
# Here are the X and Y that we want to use to predict
y = transcript['offer_type']
X = transcript.drop(['person', 'offer_type'], axis = 1)
X['gender'].fillna(X['gender'].mode()[0], inplace=True)
X['income'].fillna(X['income'].mean(), inplace=True)
X.head()
sum(X['gender'].isnull())
```
Before we train , it is important that we scale some of the columns
The following cell does that
```
scaler = MinMaxScaler()
to_normalize = ['time', 'amount', 'reward', 'income']
X[to_normalize] = scaler.fit_transform(X[to_normalize])
X
```
# Implementation / refinement
In this part we will start with a radom forest classifier
```
# Let's split our data into training and testing sets
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
# Implementing Random forest
Rand_model = RandomForestRegressor(n_estimators = 100, random_state = 42)
Rand_model.fit(X_train, y_train)
y_pred_rand = Rand_model.predict(X_test)
# Let's now try a linear regression model
Linear_model = LogisticRegression()
Linear_model.fit(X_train, y_train)
y_Linear_model = Linear_model.predict(X_test)
# Last we will try a decision tree model
Decision_model = DecisionTreeClassifier()
Decision_model.fit(X_train, y_train)
y_Decision_model = Decision_model.predict(X_test)
```
# Evaluating the model
```
len(y_Decision_model)
# reference: https://towardsdatascience.com/random-forest-in-python-24d0893d51c0
def pred_score(model):
pred = model.predict(X_test)
# Calculate the absolute errors
errors = abs(pred - y_test)
# Calculate mean absolute percentage error
mape = 100 * (errors / y_test)
accuracy = 100 - np.mean(mape)
return round(accuracy, 2)
# SIde by side comparison
models = [Decision_model, Rand_model, Linear_model]
model_names = [type(n).__name__ for n in models]
tr_accuracy = [x.score(X_train, y_train)*100 for x in models]
pred_accuracy = [pred_score(y) for y in models]
results = [tr_accuracy, pred_accuracy]
results_df = pd.DataFrame(results, columns = model_names, index=['Training Accuracy', 'Predicting Accuracy'])
results_df
```
# Model Improvement
```
# improving the model takes too long to run, you will need around 3 hours to run the improvment depending on your pc's performances
# Let's tune our linear regression model to see it we get better results
max_iter = [100, 120, 140, 160, 180, 200, 220]
C = [1.0,1.5,2.0,2.5,3.0,3.5,4.0]
dual = [True, False]
parameters = dict(dual = dual, max_iter = max_iter, C = C)
imp_model = LogisticRegression(random_state=42)
grid = GridSearchCV(estimator = imp_model, param_grid = parameters, cv = 3, n_jobs = -1)
grid_result = grid.fit(X_train, y_train)
print(f'Best Score: {grid_result.best_score_}')
print(f'Best params: {grid_result.best_params_}')
```
# Model Validation + Justification
From the above result, we can see that all the model predict good, the DecisionTreeClassifier and the RandomForestRegressor both predict 100% accurate on the train and the test data set, while in the other hand, the Logistic regression predicts 80 for the train and 92 for the test.
Taking one of the two models that predict 100% might lead us to overfitting, so to avoid that we are going to keep the Logistic regression, because event if it is predicting 100% accurate, the accuracy is still good and we can avoid overfitting
# Reflection
On the most important and I would say interresting part of the project was cleaning the data for the training.
We had three different datasets that we wanted to combine and that was difficult but interesting.
Also I playing around with different algoriths was fun.
I could see how great certain algorithms are doing, and also how bad some other were doing.
I finaly chose to go with the regression algorith, given that we wanted to avoid overfitting.
We can as well improve this algorithm by using gridsearch.
Given that gridsearch is great at finding the perfect parameters, we would probabibly have better result since this regression algorithm is using default parameters.
# Conlusion
We wanted from the begining the create a machine learning model that can well predict the offer to be given to a customer, given its demographics.
and we were able to do so.
The advantage of this that we are able to predict, what should be given to a customer, therefore increase the chances of having a customer react to an offer.
What type of offer would you like to receive?
| github_jupyter |
# Project: No-Show Appointments
## Table of Contents
<ul>
<li><a href="#intro">INTRODUCTION</a></li>
<li><a href="#wrangling">DATA WRANGLING</a></li>
<li><a href="#gathering">Gathering</a></li>
<li><a href="#assessing">Assessing</a></li>
<li><a href="#cleaning">Cleaning</a></li>
<li><a href="#da">DATA ANALYSIS</a></li>
<li><a href="#conclusions">CONCLUSIONS</a></li>
</ul>
<a id='intro'></a>
## Introduction
```
# import all packages and set plots to be embedded inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
%matplotlib inline
```
<a id='wrangling'></a>
## Data Wrangling
<a id='gathering'></a>
### Gathering
```
df = pd.read_csv('noshowappointments-kagglev2-may-2016.csv')
```
<a id='assessing'></a>
### Assessing
#### Assessing of the whole dataframe
```
df.head()
df.tail()
df.shape
df.info()
df.describe()
df.isna().sum()
df.duplicated().sum()
```
#### Assessing of concrete variables
```
df.groupby('Gender')['AppointmentID'].count()
print(df['ScheduledDay'].min())
print(df['ScheduledDay'].max())
print(df['AppointmentDay'].min())
print(df['AppointmentDay'].max())
#Age disbribution overview
df['Age'].hist()
plt.title('Distribution of Patients Age');
plt.xlabel('Age')
plt.ylabel('Number of patients');
df['No-show'].value_counts()/df.shape[0]
df['Neighbourhood'].value_counts()
```
<a id='cleaning'></a>
### Cleaning
#### Define
Drop duplicated registers
#### Code
```
df.duplicated().sum()
df.drop_duplicates(inplace=True)
```
#### Test
```
df.duplicated().sum()
```
#### Define
Drop registers whose response variable (No-show) is Null
#### Code
```
df['No-show'].isna().sum()
df=df[df['No-show'].isna()==False]
```
#### Test
```
df['No-show'].isna().sum()
df.shape
df['No-show'].value_counts()
```
#### Define
Format No-show variable with Yes=1 and No=0 values and convert it to number
#### Code
```
df.loc[df['No-show'] == 'No','No-show'] = 0
df.loc[df['No-show'] == 'Yes','No-show'] = 1
df['No-show']=df['No-show'].astype(int)
```
#### Test
```
df['No-show'].value_counts()
```
#### Define
Conver to datetime the ScheduledDay and AppointmentDay columns.
#### Code
```
df['ScheduledDay'] = pd.to_datetime(df['ScheduledDay'])
df['AppointmentDay'] = pd.to_datetime(df['AppointmentDay'])
```
#### Test
```
df.info()
print(df['ScheduledDay'].min())
print(df['ScheduledDay'].max())
print(df['AppointmentDay'].min())
print(df['AppointmentDay'].max())
```
#### Define
Drop registers whose Scheduled day is later than the Appointment day. These registers are wrong and might distort the analysis
#### Code
```
df['diff_App_Sch_Day']=df['AppointmentDay'].dt.date-df['ScheduledDay'].dt.date
df['diff_App_Sch_Day']=df['diff_App_Sch_Day']/np.timedelta64(1,'D')
df=df[df['diff_App_Sch_Day']>=0]
```
#### Test
```
df['diff_App_Sch_Day'].min()
```
#### Define
Drop registers with Age < 0
#### Code
```
(df['Age']<0).sum()
df=df[df['Age']>=0]
```
#### Test
```
(df['Age']<0).sum()
```
<a id='da'></a>
## Data Analysis
```
df['No-show'].value_counts()*100/df.shape[0]
#Definition of function for plotting a bar chart
def plot_barchart(locations,measurement,labels,p_title,p_xlabel,p_ylabel,orientation='v'):
'''
Plot bar chart based on provided parameters. Horizontal or vertical orientation can be chosen.
INPUT:
locations - range of locations where labels are placed
measurement - list of numerical values
labels - list of strings with the categorical values
p_title - bar chart title
p_xlabel - x axis label of bar chart
p_ylabel - y axis label of bar chart
orientation - bar chart orientation ('h'=horizontal or 'v'=vertical). Default value = 'v' (vertical)
OUTPUT:
None
'''
if orientation == 'h':
plt.barh(locations,measurement,tick_label=labels)
else:
plt.bar(locations,measurement,tick_label=labels)
plt.title(p_title)
plt.xlabel(p_xlabel)
plt.ylabel(p_ylabel)
return None
```
### I. Does the time between Scheduled and Appointment dates have an effect on the attendance rate?
```
#Average days for each No-show value
days_avg=df.groupby('No-show')['diff_App_Sch_Day'].mean()
days_avg
#Variables and parameters are defined for plotting the bar chart
locations_a =np.arange(len(days_avg)) #set the locations according to how we want to show the data
measurement_a = days_avg
labels_a = ['Show','No-show'] #labels are created
title_a = 'Averaged days between the Scheduled and Appointment dates'
xlabel_a = 'Averaged days'
ylabel_a = ''
orientation_a = 'h'
#Plotting average days for each No-show value
plot_barchart(locations_a,measurement_a,labels_a,title_a,xlabel_a,ylabel_a,orientation_a)
#Both No-show values are shown in a histogram
bins_n = 20
g = sb.FacetGrid(data=df,hue='No-show',height=5)
g.map(plt.hist,'diff_App_Sch_Day',bins=bins_n, alpha=0.6)
```
#### Conclusion:
Patients that did not show up booked the appointment 1 week, on average, earlier than patients that showed up.
### II. How does patients' age impact on absenteeism?
```
#Age range is created
df['Age_Range']='0-9'
df.loc[df['Age']>=10,'Age_Range'] = '10-19'
df.loc[df['Age']>=20,'Age_Range'] = '20-29'
df.loc[df['Age']>=30,'Age_Range'] = '30-39'
df.loc[df['Age']>=40,'Age_Range'] = '40-49'
df.loc[df['Age']>=50,'Age_Range'] = '50-59'
df.loc[df['Age']>=60,'Age_Range'] = '60-69'
df.loc[df['Age']>=70,'Age_Range'] = '70-79'
df.loc[df['Age']>=80,'Age_Range'] = '>=80'
#No-show mean is calculated (multiplied by 100 to get percentages)
Age_Range_App = df.groupby('Age_Range')['No-show'].mean()*100
Age_Range_App
#Variables and parameters are defined for plotting the bar chart
locations_a =np.arange(len(Age_Range_App)) #set the locations according to how we want to show the data
measurement_a = Age_Range_App
labels_a = Age_Range_App.index #labels are created
title_a = 'Percentage of patients not showing up at the medical appointment by age'
xlabel_a = 'Patients Age'
ylabel_a = 'No-show average (%)'
orientation_a = 'v'
#Percentage of patiens not showing up is represented in a bar chart
plot_barchart(locations_a,measurement_a,labels_a,title_a,xlabel_a,ylabel_a,orientation_a)
```
#### Conclusion:
Younger generations have lower attendance rate than older ones.
### III. Is the patients' location an important aspect to consider regarding the attendance?
```
#No-show mean is calculated by neighbourhood
Neigh_NoShow = df.groupby('Neighbourhood')['No-show'].mean()*100
#Appointment count percentage is calculated by neighbourhood
Neigh_CountWeight = df.groupby('Neighbourhood')['AppointmentID'].count()*100/df.shape[0]
#Index is reset
Neigh_NoShow=Neigh_NoShow.reset_index()
Neigh_CountWeight=Neigh_CountWeight.reset_index()
#Both dataframes are merged and columns renamed
Neighbourhood_Stats = Neigh_CountWeight.merge(Neigh_NoShow)
Neighbourhood_Stats=Neighbourhood_Stats.rename(columns={"AppointmentID":"AppointCount (%)","No-show":"No-show (%)"})
Neighbourhood_Stats.head()
#Represent both variables in a scatter plot
plt.scatter(Neighbourhood_Stats['No-show (%)'],Neighbourhood_Stats['AppointCount (%)'])
#Represent both variables in a scatter plot (zoomed in)
plt.scatter(Neighbourhood_Stats['No-show (%)'],Neighbourhood_Stats['AppointCount (%)'])
plt.xlim([10,30])
plt.ylim([1,8]);
Neighbourhood_Stats['AppointCount (%)'].median()
```
Awareness campaigns can be done on neighbourhoods whose:
- number of appointments > median AND
- have the highest No-show rate
```
#Select neighbourhoods with number of appointments > median
Neigh_mostApp = Neighbourhood_Stats[Neighbourhood_Stats['AppointCount (%)']>Neighbourhood_Stats['AppointCount (%)'].median()]
#Neighbourhoods with highest No-show rate
Neigh_mostApp[['Neighbourhood','No-show (%)']].sort_values(by='No-show (%)',ascending=False).head(5)
#Neighbourhoods with lowest No-show rate
Neigh_mostApp[['Neighbourhood','No-show (%)']].sort_values(by='No-show (%)',ascending=False).tail(5)
```
#### Conclusion:
There is a gap of up to 13 percentual points between the neighbourhoods with highest and lowest absenteeism rate once we have filtered out the neighbourhoods with the lowest appointments.
Awareness campaign could be launch in the neighbourhoods with the highest absenteeism rate.
<a id='conclusions'></a>
## Conclusions
The analysis showed that the sooner the patients scheduled the appointment the more likely they were for not showing up at the doctor's office. Taking a close look at the patients' profile, I learnt that younger generations are more prone than older ones to not showing up, and also that the attendance rate might drastically vary between different patient' locations.
| github_jupyter |
<img src="https://upload.wikimedia.org/wikipedia/commons/4/47/Logo_UTFSM.png" width="200" alt="utfsm-logo" align="left"/>
# MAT281
### Aplicaciones de la Matemática en la Ingeniería
## Módulo 04
## Laboratorio Clase 04: Métricas y selección de modelos
### Instrucciones
* Completa tus datos personales (nombre y rol USM) en siguiente celda.
* La escala es de 0 a 4 considerando solo valores enteros.
* Debes _pushear_ tus cambios a tu repositorio personal del curso.
* Como respaldo, debes enviar un archivo .zip con el siguiente formato `mXX_cYY_lab_apellido_nombre.zip` a alonso.ogueda@gmail.com, debe contener todo lo necesario para que se ejecute correctamente cada celda, ya sea datos, imágenes, scripts, etc.
* Se evaluará:
- Soluciones
- Código
- Que Binder esté bien configurado.
- Al presionar `Kernel -> Restart Kernel and Run All Cells` deben ejecutarse todas las celdas sin error.
__Nombre__: Simón Masnú
__Rol__: 201503026-K
En este laboratorio utilizaremos el conjunto de datos _Abolone_.
**Recuerdo**
La base de datos contiene mediciones a 4177 abalones, donde las mediciones posibles son sexo ($S$), peso entero $W_1$, peso sin concha $W_2$, peso de visceras $W_3$, peso de concha $W_4$, largo ($L$), diametro $D$, altura $H$, y el número de anillos $A$.
```
import pandas as pd
import numpy as np
abalone = pd.read_csv(
"data/abalone.data",
header=None,
names=["sex", "length", "diameter", "height", "whole_weight", "shucked_weight", "viscera_weight", "shell_weight", "rings"]
)
abalone_data = (
abalone.assign(sex=lambda x: x["sex"].map({"M": 1, "I": 0, "F": -1}))
.loc[lambda x: x.drop(columns="sex").gt(0).all(axis=1)]
.astype(np.float)
)
abalone_data.head()
```
#### Modelo A
Consideramos 9 parámetros, llamados $\alpha_i$, para el siguiente modelo:
$$ \log(A) = \alpha_0 + \alpha_1 W_1 + \alpha_2 W_2 +\alpha_3 W_3 +\alpha_4 W_4 + \alpha_5 S + \alpha_6 \log L + \alpha_7 \log D+ \alpha_8 \log H$$
```
def train_model_A(data):
y = np.log(data.loc[:, "rings"].values.ravel())
X = (
data.assign(
intercept=1.,
length=lambda x: x["length"].apply(np.log),
diameter=lambda x: x["diameter"].apply(np.log),
height=lambda x: x["height"].apply(np.log),
)
.loc[: , ["intercept", "whole_weight", "shucked_weight", "viscera_weight", "shell_weight", "sex", "length", "diameter", "height"]]
.values
)
coeffs = np.linalg.lstsq(X, y, rcond=None)[0]
return coeffs
def test_model_A(data, coeffs):
X = (
data.assign(
intercept=1.,
length=lambda x: x["length"].apply(np.log),
diameter=lambda x: x["diameter"].apply(np.log),
height=lambda x: x["height"].apply(np.log),
)
.loc[: , ["intercept", "whole_weight", "shucked_weight", "viscera_weight", "shell_weight", "sex", "length", "diameter", "height"]]
.values
)
ln_anillos = np.dot(X, coeffs)
return np.exp(ln_anillos)
```
#### Modelo B
Consideramos 6 parámetros, llamados $\beta_i$, para el siguiente modelo:
$$ \log(A) = \beta_0 + \beta_1 W_1 + \beta_2 W_2 +\beta_3 W_3 +\beta W_4 + \beta_5 \log( L D H ) $$
```
def train_model_B(data):
y = np.log(data.loc[:, "rings"].values.ravel())
X = (
data.assign(
intercept=1.,
ldh=lambda x: (x["length"] * x["diameter"] * x["height"]).apply(np.log),
)
.loc[: , ["intercept", "whole_weight", "shucked_weight", "viscera_weight", "shell_weight", "ldh"]]
.values
)
coeffs = np.linalg.lstsq(X, y, rcond=None)[0]
return coeffs
def test_model_B(data, coeffs):
X = (
data.assign(
intercept=1.,
ldh=lambda x: (x["length"] * x["diameter"] * x["height"]).apply(np.log),
)
.loc[: , ["intercept", "whole_weight", "shucked_weight", "viscera_weight", "shell_weight", "ldh"]]
.values
)
ln_anillos = np.dot(X, coeffs)
return np.exp(ln_anillos)
```
#### Modelo C
Consideramos 12 parámetros, llamados $\theta_i^{k}$, con $k \in \{M, F, I\}$, para el siguiente modelo:
Si $S=male$:
$$ \log(A) = \theta_0^M + \theta_1^M W_2 + \theta_2^M W_4 + \theta_3^M \log( L D H ) $$
Si $S=female$
$$ \log(A) = \theta_0^F + \theta_1^F W_2 + \theta_2^F W_4 + \theta_3^F \log( L D H ) $$
Si $S=indefined$
$$ \log(A) = \theta_0^I + \theta_1^I W_2 + \theta_2^I W_4 + \theta_3^I \log( L D H ) $$
```
def train_model_C(data):
df = (
data.assign(
intercept=1.,
ldh=lambda x: (x["length"] * x["diameter"] * x["height"]).apply(np.log),
)
.loc[: , ["intercept", "shucked_weight", "shell_weight", "ldh", "sex", "rings"]]
)
coeffs_dict = {}
for sex, df_sex in df.groupby("sex"):
X = df_sex.drop(columns=["sex", "rings"])
y = np.log(df_sex["rings"].values.ravel())
coeffs_dict[sex] = np.linalg.lstsq(X, y, rcond=None)[0]
return coeffs_dict
def test_model_C(data, coeffs_dict):
df = (
data.assign(
intercept=1.,
ldh=lambda x: (x["length"] * x["diameter"] * x["height"]).apply(np.log),
)
.loc[: , ["intercept", "shucked_weight", "shell_weight", "ldh", "sex", "rings"]]
)
pred_dict = {}
for sex, df_sex in df.groupby("sex"):
X = df_sex.drop(columns=["sex", "rings"])
ln_anillos = np.dot(X, coeffs_dict[sex])
pred_dict[sex] = np.exp(ln_anillos)
return pred_dict
```
### 1. Split Data (1 pto)
Crea dos dataframes, uno de entrenamiento (80% de los datos) y otro de test (20% restante de los datos) a partir de `abalone_data`.
_Hint:_ `sklearn.model_selection.train_test_split` funciona con dataframes!
```
from sklearn.model_selection import train_test_split
abalone_train, abalone_test = train_test_split(abalone_data, test_size=0.20,random_state=42)
abalone_train.head()
```
### 2. Entrenamiento (1 pto)
Utilice las funciones de entrenamiento definidas más arriba con tal de obtener los coeficientes para los datos de entrenamiento. Recuerde que para el modelo C se retorna un diccionario donde la llave corresponde a la columna `sex`.
```
coeffs_A = train_model_A(abalone_train)
coeffs_B = train_model_B(abalone_train)
coeffs_C = train_model_C(abalone_train)
```
### 3. Predicción (1 pto)
Con los coeficientes de los modelos realize la predicción utilizando el conjunto de test. El resultado debe ser un array de shape `(835, )` por lo que debes concatenar los resultados del modelo C.
**Hint**: Usar `np.concatenate`.
```
y_pred_A = test_model_A(abalone_test,coeffs_A)
y_pred_B = test_model_B(abalone_test,coeffs_B)
y_pred_C = np.concatenate([test_model_C(abalone_test,coeffs_C)[-1],test_model_C(abalone_test,coeffs_C)[0],test_model_C(abalone_test,coeffs_C)[1]])
```
### 4. Cálculo del error (1 pto)
Se utilizará el Error Cuadrático Medio (MSE) que se define como
$$\textrm{MSE}(y,\hat{y}) =\dfrac{1}{n}\sum_{t=1}^{n}\left | y_{t}-\hat{y}_{t}\right |^2$$
Defina una la función `MSE` y el vectores `y_test_A`, `y_test_B` e `y_test_C` para luego calcular el error para cada modelo.
**Ojo:** Nota que al calcular el error cuadrático medio se realiza una resta elemento por elemento, por lo que el orden del vector es importante, en particular para el modelo que separa por `sex`.
```
def MSE(y_real, y_pred):
return sum(np.absolute(y_real-y_pred)**2)/len(y_real)
y_test_A = abalone_test.loc[:,'rings']
y_test_B = abalone_test.loc[:,'rings']
y_test_C = np.concatenate([abalone_test[ abalone_test['sex'] == -1].loc[:,'rings']
,abalone_test[ abalone_test['sex'] == 0].loc[:,'rings']
,abalone_test[ abalone_test['sex'] == 1].loc[:,'rings']]
,axis=None) #perdon por el Hard-Coding
error_A = MSE(y_test_A,y_pred_A)
error_B = MSE(y_test_B,y_pred_B)
error_C = MSE(y_test_C,y_pred_C)
print(f"Error modelo A: {error_A:.2f}")
print(f"Error modelo B: {error_B:.2f}")
print(f"Error modelo C: {error_C:.2f}")
```
**¿Cuál es el mejor modelo considerando esta métrica?**
El mejor modelo considerando como métrica el `MSE` es el modelo **B**.
| github_jupyter |
## Recognized Formats
```
# 1 (Array of Integers parameter)
from beakerx import *
TableDisplay({"a":100, "b":200, "c":300})
%%groovy
// expected result of cell 1
Image("../../resources/img/python/tableAPI/cell1_case1.png")
# 2 (2D Array of Integers parameter)
TableDisplay([{"a":1}, {"a":10, "b":20}])
%%groovy
// expected result of cell 2
Image("../../resources/img/python/tableAPI/cell2_case1.png")
# 3 (Array of Decimals parameter)
TableDisplay({"a":1/10, "b":1/20, "c":0.33})
%%groovy
// expected result of cell 3
Image("../../resources/img/python/tableAPI/cell3_case1.png")
# 4 (2D Array of Decimals parameter)
TableDisplay([{"a":1/10}, {"a":1/100, "b":3.12345}])
%%groovy
// expected result of cell 4
Image("../../resources/img/python/tableAPI/cell4_case1.png")
# 5 (Array of Strings parameter)
TableDisplay({"a":'string aaa', "b":'string bbb', "c":'string ccc'})
%%groovy
// expected result of cell 5
Image("../../resources/img/python/tableAPI/cell5_case1.png")
# 6 (2D Array of Strings parameter)
TableDisplay([{"a":'a'}, {"a":'1a', "b":'2b'}])
%%groovy
// expected result of cell 6
Image("../../resources/img/python/tableAPI/cell6_case1.png")
# 7 (Array of Integer Arrays parameter)
TableDisplay({"a":[1, 2, 3], "b":[10, 20, 30], "c":[100, 200, 300]})
%%groovy
// expected result of cell 7
Image("../../resources/img/python/tableAPI/cell7_case1.png")
# 8 (2D Array of Integer Arrays parameter)
TableDisplay([
{"a":[1, 2, 3]},
{"a":[10, 20, 30], "b":[100, 200, 300]}])
%%groovy
// expected result of cell 8
Image("../../resources/img/python/tableAPI/cell8_case1.png")
# 9 (2D Array of Integer,Decimal,String,Array Arrays parameter)
row1 = {"a":100, "b":200, "c":300}
row2 = {"a":1/10, "b":1/20, "c":0.33}
row3 = {"a":'a a a', "b":'b b b', "c":'c c c'}
row4 = {"a":[1, 2, 3], "b":[10, 20, 30], "c":[100, 200, 300]}
TableDisplay([row1, row2, row3, row4])
%%groovy
// expected result of cell 9
Image("../../resources/img/python/tableAPI/cell9_case1.png")
# 10 ([Integer,Decimal,String,Array] parameter)
TableDisplay({"a":100, "b":1/20, "c":'c c c', "d":[100, 200, 300]})
%%groovy
// expected result of cell 10
Image("../../resources/img/python/tableAPI/cell10_case1.png")
# 11 (2D Arrays of [Integer,Decimal,String,Array] parameter)
row1 = {"a":10, "b":1/10, "c":'c', "d":[100, 200]}
row2 = {"a":100, "b":1/20, "c":'c c c', "d":[100, 200, 300]}
TableDisplay([row1, row2])
%%groovy
// expected result of cell 11
Image("../../resources/img/python/tableAPI/cell11_case1.png")
# 12 (numbers as name of Array keys (Array parameter))
TableDisplay({10:20, 1/10:1/20, 'c':'c c c', '[100, 200]':[100, 200, 300]})
%%groovy
// expected result of cell 12
Image("../../resources/img/python/tableAPI/cell12_case1.png")
# 13 (numbers as name of Array keys (2D Array parameter)
row1 = {40:40, 1/40:1/40, 'c':'c'}
row2 = {40:20, 1/40:1/20, 'c':'c c c', '[100, 200]':[100, 200, 300]}
TableDisplay([row1, row2])
%%groovy
// expected result of cell 13
Image("../../resources/img/python/tableAPI/cell13_case1.png")
```
# Python API for Table Display
In addition to APIs for creating and formatting BeakerX's interactive table widget, the Python runtime configures pandas to display tables with the interactive widget instead of static HTML.
```
# 14
import pandas as pd
from beakerx import *
df = pd.DataFrame({"runway": ["24", "36L"]})
TableDisplay(df)
%%groovy
// expected result of cell 14
Image("../../resources/img/python/tableAPI/cell14_case1.png")
import pandas as pd
from beakerx import *
from beakerx.object import beakerx
#Display mode: TableDisplay Widget
beakerx.pandas_display_table()
pd.DataFrame(data=np.zeros((5,5)), index=pd.Int64Index([0, 1, 2, 3, 4], dtype='int64'))
# 15
pd.read_csv('../../resources/data/pd_index.csv', index_col=0)
%%groovy
// expected result of cell 15
Image("../../resources/img/python/tableAPI/cell15_case1.png")
# 16
pd.read_csv('../../resources/data/interest-rates-small.csv')
%%groovy
// expected result of cell 16
Image("../../resources/img/python/tableAPI/cell16_case1.png")
```
# Alignment
```
# 16_2
colNames = ["default aligment", "right aligment", "left aligment","center aligment"]
row1 = [123, 123, 123, 123]
row2 = [231, 231, 231, 231]
row3 = [312, 312, 312, 312]
table162 = TableDisplay(pd.DataFrame([row1, row2, row3], columns=colNames))
table162.setAlignmentProviderForColumn('left aligment', TableDisplayAlignmentProvider.LEFT_ALIGNMENT)
table162.setAlignmentProviderForColumn('center aligment', TableDisplayAlignmentProvider.CENTER_ALIGNMENT)
table162.setAlignmentProviderForColumn('right aligment', TableDisplayAlignmentProvider.RIGHT_ALIGNMENT)
table162
%%groovy
// expected result of cell 16_2
Image("../../resources/img/python/tableAPI/cell162_case1.png")
# 16_3
colNames = ["string column", "int column 1", "double column","int column 2"]
row1 = ["123", 123, 12.3, 123]
row2 = ["231", 231, 23.1, 231]
row3 = ["312", 312, 31.2, 312]
table163 = TableDisplay(pd.DataFrame([row1, row2, row3], columns=colNames))
table163.setAlignmentProviderForType(ColumnType.String, TableDisplayAlignmentProvider.CENTER_ALIGNMENT)
table163.setAlignmentProviderForType(ColumnType.Double, TableDisplayAlignmentProvider.LEFT_ALIGNMENT)
table163
%%groovy
// expected result of cell 16_3
Image("../../resources/img/python/tableAPI/cell163_case1.png")
```
# Bar Charts Renderer
```
# 17
table2 = TableDisplay(pd.read_csv('../../resources/data/interest-rates-small.csv'))
table2.setRendererForType(ColumnType.Double, TableDisplayCellRenderer.getDataBarsRenderer(True))
#use the false parameter to hide value
table2.setRendererForColumn("y10", TableDisplayCellRenderer.getDataBarsRenderer(False))
table2
%%groovy
// expected result of cell 17
Image("../../resources/img/python/tableAPI/cell17_case1.png")
```
# Formatting
```
# 18
df3 = pd.read_csv('../../resources/data/interest-rates-small.csv')
df3['time'] = df3['time'].str.slice(0,19).astype('datetime64[ns]')
table3 = TableDisplay(df3)
table3.setStringFormatForTimes(TimeUnit.DAYS)
table3.setStringFormatForType(ColumnType.Double, TableDisplayStringFormat.getDecimalFormat(2,3))
table3.setStringFormatForColumn("m3", TableDisplayStringFormat.getDecimalFormat(0, 0))
table3
%%groovy
// expected result of cell 18
Image("../../resources/img/python/tableAPI/cell18_case1.png")
```
## HTML format
HTML format allows markup and styling of the cell's content. Interactive JavaScript is not supported however.
```
# 18_2
table182 = TableDisplay({'x': '<em style="color:red">italic red</em>',
'y': '<b style="color:blue">bold blue</b>',
'z': 'strings without markup work fine too'})
table182.setStringFormatForColumn("Value", TableDisplayStringFormat.getHTMLFormat())
table182
%%groovy
// expected result of cell 18_2
Image("../../resources/img/python/tableAPI/cell182_case1.png")
```
# Column Visibility and Placement
```
# 19
table4 = TableDisplay(pd.read_csv('../../resources/data/interest-rates-small.csv'))
#freeze a column
table4.setColumnFrozen("y1", True)
#hide a column
table4.setColumnVisible("y30", False)
table4
%%groovy
// expected result of cell 19
Image("../../resources/img/python/tableAPI/cell19_case1.png")
# 20
table5 = TableDisplay(pd.read_csv('../../resources/data/interest-rates-small.csv'))
#Columns in the list will be shown in the provided order. Columns not in the list will be hidden.
table5.setColumnOrder(["m3", "y1", "y10", "time", "y2"])
table5
%%groovy
// expected result of cell 20
Image("../../resources/img/python/tableAPI/cell20_case1.png")
```
# HeatmapHighlighter
```
# 21
colNames = ["m3","y30","time","y5","y7","spread"]
row1 = [7.8981, 8.2586, "1990-01-30 19:00:00.000 -0500", 8.1195, 8.1962, 0.3086]
row2 = [2.0021, 8.5037, "1990-02-27 19:00:00.000 -0500", 8.4247, 8.4758, 0.4711]
row3 = [2.0021, 3.5037, "1990-05-27 19:00:00.000 -0500", 1.4247, 4.4758, 5.4711]
table6 = TableDisplay(pd.DataFrame([row1, row2, row3], columns=colNames))
table6.addCellHighlighter(TableDisplayCellHighlighter.getHeatmapHighlighter("m3", TableDisplayCellHighlighter.FULL_ROW))
table6
%%groovy
// expected result of cell 21
Image("../../resources/img/python/tableAPI/cell21_case1.png")
# 21_2
table6.removeAllCellHighlighters()
%%groovy
// expected result of cell 21_2
Image("../../resources/img/python/tableAPI/cell21_case2.png")
```
# UniqueEntriesHighlighter
```
# 22
table22 = TableDisplay(pd.read_csv('../../resources/data/interest-rates-small.csv'))
table22.addCellHighlighter(TableDisplayCellHighlighter.getUniqueEntriesHighlighter("m3"))
table22
%%groovy
// expected result of cell 22
Image("../../resources/img/python/tableAPI/cell22_case1.png")
# 23
table23 = TableDisplay(pd.read_csv('../../resources/data/interest-rates-small.csv'))
table23.addCellHighlighter(TableDisplayCellHighlighter.getHeatmapHighlighter("m3", HighlightStyle.SINGLE_COLUMN))
table23.addCellHighlighter(TableDisplayCellHighlighter.getHeatmapHighlighter("y10", TableDisplayCellHighlighter.SINGLE_COLUMN))
table23.addCellHighlighter(TableDisplayCellHighlighter.getUniqueEntriesHighlighter("m6", HighlightStyle.SINGLE_COLUMN))
table23
%%groovy
// expected result of cell 23
Image("../../resources/img/python/tableAPI/cell23_case1.png")
```
# FontSize
```
# 24
table24 = TableDisplay(pd.read_csv('../../resources/data/interest-rates-small.csv'))
table24.setDataFontSize(10)
table24.setHeaderFontSize(16)
table24
%%groovy
// expected result of cell 24
Image("../../resources/img/python/tableAPI/cell24_case1.png")
# 25
table25 = TableDisplay(pd.read_csv('../../resources/data/interest-rates-small.csv'))
table25.setHeadersVertical(True)
table25
%%groovy
// expected result of cell 25
Image("../../resources/img/python/tableAPI/cell25_case1.png")
```
# ColorProvider
```
# 26
mapListColorProvider = [
{"a": 1, "b": 2, "c": 3},
{"a": 4, "b": 5, "c": 6},
{"a": 7, "b": 8, "c": 5}
]
table26 = TableDisplay(mapListColorProvider)
colors = [
[Color.LIGHT_GRAY, Color.GRAY, Color.RED],
[Color.DARK_GREEN, Color.ORANGE, Color.RED],
[Color.MAGENTA, Color.BLUE, Color.BLACK]
]
def color_provider(row, column, table):
return colors[row][column]
table26.setFontColorProvider(color_provider)
table26
%%groovy
// expected result of cell 26
Image("../../resources/img/python/tableAPI/cell26_case1.png")
```
# ToolTip
```
# 27
table27 = TableDisplay(pd.read_csv('../../resources/data/interest-rates-small.csv'))
def config_tooltip(row, column, table):
return "The value is: " + str(table.values[row][column])
table27.setToolTip(config_tooltip)
table27
```
| github_jupyter |
# CDR3 entropy in shared and unshared clonotypes
Starting with unique cross-subject clonotype datasets, computes per-position entropy for CDR3s in unshared or shared (found in at least 6 of 10 samples) clonotypes.
The following Python packages are required:
* numpy
* pandas
and can be installed by running `pip install numpy pandas`
```
from __future__ import print_function
from collections import Counter
import os
import subprocess as sp
import sys
import numpy as np
import pandas as pd
```
## Get sequences
The raw dataset (unique cross-subject clonotypes) is too large to be included in this Github repo. Instead, a compressed archive containing all of the required data can be downloaded [**HERE**](http://burtonlab.s3.amazonaws.com/GRP_github_data/dedup_10-subject_pools.tar.gz). Decompressing the archive in the `./data` directory will allow the following code blocks to run without modification.
***NOTE:*** *The required data files are relatively large (~30GB in total), so ensure adequate storage space is available before downloading.*
```
def get_sequences(seq_files):
all_seqs = {}
for seq_type in seq_files.keys():
seq_file = seq_files[seq_type]
seqs = {'shared': {i: [] for i in range(7, 15)},
'unshared': {i: [] for i in range(7, 15)}}
with open(seq_file) as f:
for line in f:
sline = line.strip().split()
if not sline:
continue
try:
c = int(sline[0])
if c == 1:
s = 'unshared'
elif c in range(6, 11):
s = 'shared'
else:
continue
aa = sline[3]
l = len(aa)
if l not in range(7, 15):
continue
seqs[s][l].append(aa)
except IndexError:
continue
all_seqs[seq_type] = seqs
# downselect sequences so that shared and unshared pools are the same size
selected_seqs = {t: {'shared': {}, 'unshared': {}} for t in all_seqs.keys()}
for seq_type in ['observed', 'subject-specific synthetic']:
for length in range(7, 15):
num_seqs = min([len(all_seqs[seq_type][t][length]) for t in ['shared', 'unshared']])
for shared_type in ['shared', 'unshared']:
s = all_seqs[seq_type][shared_type][length]
if len(s) > num_seqs:
s = np.random.choice(s, size=num_seqs, replace=False)
selected_seqs[seq_type][shared_type][length] = s
return all_seqs, selected_seqs
files = {'observed': './data/dedup_10-subject_pools/10-subject_dedup_pool_with-counts.txt',
'subject-specific synthetic': './data/dedup_10-subject_pools/10-sample_dedup_pool_synthetic_subject-specific-models_with-counts.txt'}
all_seqs, seq_dict = get_sequences(files)
```
## Compute shared/unshared CDR3 entropy
```
def calculate_entropies(seq_dict, seq_type):
edata = []
for s in seq_dict.keys():
for l in seq_dict[s].keys():
seqs = seq_dict[s][l]
for residues in list(zip(*seqs))[3:-3]:
e = entropy(residues)
edata.append({'sample': '{} ({})'.format(seq_type, s), 'seq_type': seq_type,
'Shannon entropy': e, 'CDR3 length': l, 'shared': s})
return edata
def entropy(residues):
n_residues = len(residues)
if n_residues <= 1:
return 0.
counts = np.asarray(Counter(residues).values(), dtype=np.float64)
probs = counts[np.nonzero(counts)] / n_residues
n_classes = len(probs)
if n_classes <= 1:
return 0.
return - np.sum(probs * np.log(probs)) / np.log(n_classes)
entropy_data = []
print('Getting sequences...')
for seq_type in seq_dict.keys():
print(seq_type)
entropies = calculate_entropies(seq_dict[seq_type], seq_type)
entropy_data += entropies
entropy_df = pd.DataFrame(entropy_data)
entropy_df.to_csv('./data/per-position_shannon_entropies.csv')
```
## Shared CDR3 sequence properties
```
shared_seqs = []
for n in range(6, 11):
shared_seqs += seqs[n]
```
| github_jupyter |
```
#from preprocess import *
#standard module
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
# import sklearn
from sklearn import linear_model
from sklearn.neural_network import MLPClassifier
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.preprocessing import PolynomialFeatures
from sklearn import neighbors
from sklearn.metrics import f1_score
from sklearn.metrics import precision_score
from sklearn.metrics import accuracy_score
from scipy.spatial.distance import squareform
from scipy.stats import rankdata
from matplotlib.backends.backend_pdf import PdfPages
from sklearn.neural_network import MLPRegressor
sns.set_context("paper", font_scale=1.5, rc={"lines.linewidth": 2.5})
import sys
sys.path.append("../../tools/")
from preprocess import *
```
## Simple machine learning model
```
#load data
alldata_15G=np.loadtxt('../../mddata/15grid_shuffled.dat')
alldata = alldata_15G
```
Linear Regression
```
def linear_models_with_regularizations(X_train, X_test, y_train, y_test, alpha_ridge, alpha_lasso):
"""
Parameters
--------------
X_train, X_test: numpy matrix
y_train, y_test: numpy array
ridge: boolean
set ridge = True for including Ridge regression
Return: float
r2_score
"""
logTrans = False
if logTrans is True:
y_test = np.log(y_test)
y_train = np.log(y_train)
regr = linear_model.LinearRegression()
regr.fit(X_train, y_train)
y_pred_regr = regr.predict(X_test)
#accuracy_score(Y_test, Y_pred)
# The coefficients
#print('Coefficients: \n', regr.coef_)
print("Mean squared error Linear Regression: %.2f" % mean_squared_error(y_test, y_pred_regr))
# Explained variance score: 1 is perfect prediction
#ac1 = r2_score(y_test, y_pred)
print("RMSE: %lf" %np.sqrt(np.sum(np.square(y_test-y_pred_regr))/len(y_test)))
print('r2_score: %.2f' % r2_score(y_test, y_pred_regr))
ysorted= np.sort(y_test)
xx = np.linspace(ysorted[0], ysorted[-1], len(ysorted))
plt.plot(xx, xx, 'r')
plt.plot(y_pred_regr, y_test, 'bo', alpha=0.5)
plt.xlabel('Predicted yield stress') #change the name here stress/strain
plt.ylabel('True yield stress')
plt.title('OLS with polynomial degree=2')
#plt.ylim(0, 1.2)
#plt.xlim(0, 1.2)
#plt.show()
#yy = y_test.reshape((len(y_test), 1))
plt.show()
ridge = linear_model.Ridge(alpha=alpha_ridge)
ridge.fit(X_train, y_train)
y_pred_ridge=ridge.predict(X_test)
#accuracy_score(Y_test, Y_pred)
# The coefficients
#print('Coefficients: \n', clf.coef_)
print("Mean squared error Ridge Regression: %.2f" % mean_squared_error(y_test, y_pred_ridge))
# Explained variance score: 1 is perfect prediction
print("RMSE: %lf" %np.sqrt(np.sum(np.square(y_test-y_pred_ridge))/len(y_test)))
print('r2_score: %.2f' % r2_score(y_test, y_pred_ridge))
#ac_ridge = r2_score(y_test, y_pred)
#plt.plot(y_pred, y_test, 'bo', alpha=0.5)
#plt.xlabel('y_test (fracture strain)')
#plt.ylabel('y_pred (fracture strain)')
#plt.title('Ridge Regression')
lasso = linear_model.Lasso(alpha=alpha_lasso)
lasso.fit(X_train, y_train)
y_pred_lasso=lasso.predict(X_test)
#accuracy_score(Y_test, Y_pred)
# The coefficients
#print('Coefficients: \n', clf.coef_)
print("Mean squared error LASSO: %.2f" % mean_squared_error(y_test, y_pred_lasso))
# Explained variance score: 1 is perfect prediction
print("RMSE: %lf" %np.sqrt(np.sum(np.square(y_test-y_pred_lasso))/len(y_test)))
print('r2_score: %.2f' % r2_score(y_test, y_pred_lasso))
#ac_lasso = r2_score(y_test, y_pred)
#plt.plot(y_test, y_pred, 'o')
#plt.xlabel('y_test (fracture strain)')
#plt.ylabel('y_pred (fracture strain)')
#plt.title('LASSO Regression')
#plt.show()
return y_pred_regr, y_pred_ridge, y_pred_lasso, regr.coef_, ridge.coef_, lasso.coef_
```
## Training
You can choose how many features to train
```
#split data into training and test set
sns.set_context("paper", font_scale=1.5, rc={"lines.linewidth": .5})
#np.random.shuffle(alldata)
x, y=create_matrix(alldata, False, 2, 0.3, 15)
x = (x-.5)*2
X_train, X_valid, X_test, y_train, y_valid, y_test = split_data(x, y, 0.8, 0.2)
#choose polynomial degrees
poly = PolynomialFeatures(2, interaction_only=True, include_bias=True)
#poly = PolynomialFeatures(2)
X_train2 = poly.fit_transform(X_train)
print("Number of features: %d" %len(X_train2[0]))
X_test2 = poly.fit_transform(X_test)
#linear_models(X_train2, X_test2, y_train, y_test, ridge=True)
#y_train = (y_train-0.45937603178269587)/0.22056868516982353
#y_test = (y_test-0.45937603178269587)/0.22056868516982353
alpha=0.1
y_pred_regr, y_pred_ridge, y_pred_lasso, coef_regr, coef_ridge, coef_lasso = linear_models_with_regularizations(X_train2, X_test2, y_train, y_test, 10, 0.1)
#split data into training and test set
sns.set_context("paper", font_scale=1.5, rc={"lines.linewidth": .5})
#np.random.shuffle(alldata)
x, y=create_matrix(alldata, False, 0, 0.3, 15)
x = (x-.5)*2
X_train, X_valid, X_test, y_train, y_valid, y_test = split_data(x, y, 0.8, 0.2)
#choose polynomial degrees
poly = PolynomialFeatures(3, interaction_only=True, include_bias=True)
#poly = PolynomialFeatures(2)
X_train2 = poly.fit_transform(X_train)
print("Number of features: %d" %len(X_train2[0]))
X_test2 = poly.fit_transform(X_test)
#linear_models(X_train2, X_test2, y_train, y_test, ridge=True)
#y_train = (y_train-0.45937603178269587)/0.22056868516982353
#y_test = (y_test-0.45937603178269587)/0.22056868516982353
alpha=0.1
y_pred_regr, y_pred_ridge, y_pred_lasso, coef_regr, coef_ridge, coef_lasso = linear_models_with_regularizations(X_train2, X_test2, y_train, y_test, 10, 0.1)
def NN_regressor(alldata, hl, obj, transform):
nn_regr = MLPRegressor(solver='lbfgs', alpha=1e-2, hidden_layer_sizes=hl, activation='relu', random_state=1)
#sorted_data = alldata[alldata[:,15].argsort()] #index 18 prob bad design, small -> goode design
np.random.shuffle(alldata)
#0nly fit top 20%
#sorted_data = sorted_data[int(0.8*len(sorted_data)):]
#np.random.shuffle(sorted_data)
#cutoff = sorted_data[int(len(alldata)/2), 17]
#x, y=create_matrix(sorted_data, True, 2, 30, NCcell_x*NCcell_y)
x, y=create_matrix(alldata, False, obj, 0.375, 15)
X_train, X_valid, X_test, y_train, y_valid, y_test = split_data(x, y, 0.8, 0.2)
#poly = PolynomialFeatures(1, interaction_only=True, include_bias=False)
#poly = PolynomialFeatures(interaction_only=True)
#X_train2 = X_train
#poly.fit_transform(X_train)
#x2 = poly.fit_transform(x)
#print("Number of features: %d" %len(X_train2[0]))
#X_test2 = poly.fit_transform(X_test)
if (transform is True):
poly = PolynomialFeatures(2, interaction_only=True, include_bias=False)
#poly = PolynomialFeatures(interaction_only=True)
X_train2 = poly.fit_transform(X_train)
#x2 = poly.fit_transform(x)
#print("Number of features: %d" %len(X_train2[0]))
X_test2 = poly.fit_transform(X_test)
else:
X_train2 = X_train
X_test2 = X_test
nn_regr.fit(X_train2, y_train)
y_pred_nn= nn_regr.predict(X_test2)
ysorted= np.sort(y_test)
xx = np.linspace(ysorted[0], ysorted[-1], len(ysorted))
plt.plot(xx, xx, 'r')
plt.plot(y_pred_nn, y_test, 'bo', alpha=0.5)
plt.xlabel('Predicted yield stress')
plt.ylabel('True yield strain')
plt.title('Neural Network')
print("Mean squared error: %lf" % mean_squared_error(y_test, y_pred_nn))
print("RMSE: %lf" %np.sqrt(np.sum(np.square(y_test-y_pred_nn))/len(y_test)))
# Explained variance score: 1 is perfect prediction
print('r2_score: %.2f' % r2_score(y_test, y_pred_nn))
return hl[0], np.sqrt(np.sum(np.square(y_test-y_pred_nn))/len(y_test)), r2_score(y_test, y_pred_nn), y_test, y_pred_nn
hl, rmse, ac, y_test, y_pred=NN_regressor(alldata, (1024, ), 0, False)
```
| github_jupyter |
# Upgrade to rclone-based Storage Initializer - secret format intuition
In this documentation page we provide an example upgrade path from kfserving-based to rclone-based storage initializer. This is required due to the fact that secret format expected by these two storage initializers is different.
Storage initializers are used by Seldon's pre-packaged model servers to download models binaries.
As it is explained in the [SC 1.8 upgrading notes](https://docs.seldon.io/projects/seldon-core/en/latest/reference/upgrading.html#upgrading-to-1-8) the [seldonio/rclone-storage-initializer](https://github.com/SeldonIO/seldon-core/tree/master/components/rclone-storage-initializer) became default storage initializer in v1.8.0. However, it is still possible to run with kfserving-based Storage Initializer as documented [here](https://docs.seldon.io/projects/seldon-core/en/latest/servers/kfserving-storage-initializer.html).
In this tutorial we will show how to upgrade your configuration to new Storage Initializer with focus on getting the new format of a required secret right.
Read more:
- [Prepackaged Model Servers documentation page](https://docs.seldon.io/projects/seldon-core/en/latest/servers/overview.html)
- [SC 1.8 upgrading notes](https://docs.seldon.io/projects/seldon-core/en/latest/reference/upgrading.html#upgrading-to-1-8)
- [Example upgrade path to use rclone-based storage initializer globally](https://docs.seldon.io/projects/seldon-core/en/latest/examples/global-rclone-upgrade.html)
## Prerequisites
* A kubernetes cluster with kubectl configured
* mc client
* curl
## Steps in this tutorial
* Copy iris model from GCS into in-cluster minio and configure old-style storage initializer secret
* Deploy SKlearn Pre-Packaged server using kfserving storage initializer
* Discuss upgrading procedure and tips how to test new secret format
* Deploy Pre-packaged model server using rclone storage initializer
## Setup Seldon Core
Use the setup notebook to [Setup Cluster](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Setup-Cluster) with [Ambassador Ingress](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Ambassador) and [Install Seldon Core](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Install-Seldon-Core).
## Setup MinIO
Use the provided [notebook](https://docs.seldon.io/projects/seldon-core/en/latest/examples/minio_setup.html) to install Minio in your cluster and configure `mc` CLI tool.
## Copy iris model into local MinIO
```
%%bash
mc config host add gcs https://storage.googleapis.com "" ""
mc mb minio-seldon/sklearn/iris/ -p
mc cp gcs/seldon-models/sklearn/iris/model.joblib minio-seldon/sklearn/iris/
mc cp gcs/seldon-models/sklearn/iris/metadata.yaml minio-seldon/sklearn/iris/
%%bash
mc ls minio-seldon/sklearn/iris/
```
## Deploy SKLearn Server with kfserving-storage-initializer
First we deploy the model using kfserving-storage-initializer. This is using the default Storage Initializer for pre Seldon Core v1.8.0.
```
%%writefile sklearn-iris-kfserving.yaml
apiVersion: v1
kind: Secret
metadata:
name: seldon-kfserving-secret
type: Opaque
stringData:
AWS_ACCESS_KEY_ID: minioadmin
AWS_SECRET_ACCESS_KEY: minioadmin
AWS_ENDPOINT_URL: http://minio.minio-system.svc.cluster.local:9000
USE_SSL: "false"
---
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: sklearn-iris-kfserving
spec:
predictors:
- name: default
replicas: 1
graph:
name: classifier
implementation: SKLEARN_SERVER
modelUri: s3://sklearn/iris
envSecretRefName: seldon-kfserving-secret
storageInitializerImage: gcr.io/kfserving/storage-initializer:v0.4.0
!kubectl apply -f sklearn-iris-kfserving.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=sklearn-iris-kfserving -o jsonpath='{.items[0].metadata.name}')
%%bash
curl -s -X POST -H 'Content-Type: application/json' \
-d '{"data":{"ndarray":[[5.964, 4.006, 2.081, 1.031]]}}' \
http://localhost:8003/seldon/seldon/sklearn-iris-kfserving/api/v1.0/predictions | jq .
```
## Preparing rclone-compatible secret
The [rclone](https://rclone.org/)-based storage initializer expects one to define a new secret. General documentation credentials hadling can be found [here](https://docs.seldon.io/projects/seldon-core/en/latest/servers/overview.html#handling-credentials) with constantly updated examples of tested configurations.
If we do not have yet an example for Cloud Storage solution that you are using, please, consult the relevant page on [RClone documentation](https://rclone.org/#providers).
### Preparing seldon-rclone-secret
Knowing format of required format of the secret we can create it now
```
%%writefile seldon-rclone-secret.yaml
apiVersion: v1
kind: Secret
metadata:
name: seldon-rclone-secret
type: Opaque
stringData:
RCLONE_CONFIG_S3_TYPE: s3
RCLONE_CONFIG_S3_PROVIDER: minio
RCLONE_CONFIG_S3_ENV_AUTH: "false"
RCLONE_CONFIG_S3_ACCESS_KEY_ID: minioadmin
RCLONE_CONFIG_S3_SECRET_ACCESS_KEY: minioadmin
RCLONE_CONFIG_S3_ENDPOINT: http://minio.minio-system.svc.cluster.local:9000
!kubectl apply -f seldon-rclone-secret.yaml
```
### Testing seldon-rclone-secret
Before deploying SKLearn server one can test directly using the rclone-storage-initializer image
```
%%writefile rclone-pod.yaml
apiVersion: v1
kind: Pod
metadata:
name: rclone-pod
spec:
containers:
- name: rclone
image: seldonio/rclone-storage-initializer:1.11.0-dev
command: [ "/bin/sh", "-c", "--", "sleep 3600"]
envFrom:
- secretRef:
name: seldon-rclone-secret
!kubectl apply -f rclone-pod.yaml
! kubectl exec -it rclone-pod -- rclone ls s3:sklearn
! kubectl exec -it rclone-pod -- rclone copy s3:sklearn .
! kubectl exec -it rclone-pod -- sh -c "ls iris/"
```
Once we tested that secret format is correct we can delete the pod
```
!kubectl delete -f rclone-pod.yaml
```
## Deploy SKLearn Server with rclone-storage-initializer
```
%%writefile sklearn-iris-rclone.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: sklearn-iris-rclone
spec:
predictors:
- name: default
replicas: 1
graph:
name: classifier
implementation: SKLEARN_SERVER
modelUri: s3://sklearn/iris
envSecretRefName: seldon-rclone-secret
storageInitializerImage: seldonio/rclone-storage-initializer:1.11.0-dev
!kubectl apply -f sklearn-iris-rclone.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=sklearn-iris-rclone -o jsonpath='{.items[0].metadata.name}')
%%bash
curl -s -X POST -H 'Content-Type: application/json' \
-d '{"data":{"ndarray":[[5.964, 4.006, 2.081, 1.031]]}}' \
http://localhost:8003/seldon/seldon/sklearn-iris-rclone/api/v1.0/predictions | jq .
```
## Cleanup
```
%%bash
kubectl delete -f sklearn-iris-rclone.yaml
kubectl delete -f sklearn-iris-kfserving.yaml
```
| github_jupyter |
## Calculate null distribution from median Cell Painting scores with same sample size as L1000
Code modified from @adeboyeML
```
import os
import pathlib
import pandas as pd
import numpy as np
from collections import defaultdict
from pycytominer import feature_select
from statistics import median
import random
from scipy import stats
import pickle
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
np.warnings.filterwarnings('ignore', category=np.VisibleDeprecationWarning)
np.random.seed(42)
# Load common compounds
common_file = pathlib.Path(
"..", "..", "..", "6.paper_figures", "data", "significant_compounds_by_threshold_both_assays.tsv.gz"
)
common_df = pd.read_csv(common_file, sep="\t")
common_compounds = common_df.compound.unique()
print(len(common_compounds))
cp_level4_path = "cellpainting_lvl4_cpd_replicate_datasets"
df_level4 = pd.read_csv(os.path.join(cp_level4_path, 'cp_level4_cpd_replicates_subsample.csv.gz'),
compression='gzip',low_memory = False)
print(df_level4.shape)
df_level4.head()
df_cpd_med_scores = pd.read_csv(os.path.join(cp_level4_path, 'cpd_replicate_median_scores_subsample.csv'))
df_cpd_med_scores = df_cpd_med_scores.set_index('cpd').rename_axis(None, axis=0).copy()
# Subset to common compound measurements
df_cpd_med_scores = df_cpd_med_scores.loc[df_cpd_med_scores.index.isin(common_compounds), :]
print(df_cpd_med_scores.shape)
df_cpd_med_scores.head()
def get_cpds_replicates(df, df_lvl4):
"""
This function returns all replicates id/names found in each compound
and in all doses(1-6)
"""
dose_list = list(set(df_lvl4['Metadata_dose_recode'].unique().tolist()))[1:7]
replicates_in_all = []
cpds_replicates = {}
for dose in dose_list:
rep_list = []
df_doses = df_lvl4[df_lvl4['Metadata_dose_recode'] == dose].copy()
for cpd in df.index:
replicate_names = df_doses[df_doses['pert_iname'] == cpd]['replicate_name'].values.tolist()
rep_list += replicate_names
if cpd not in cpds_replicates:
cpds_replicates[cpd] = [replicate_names]
else:
cpds_replicates[cpd] += [replicate_names]
replicates_in_all.append(rep_list)
return replicates_in_all, cpds_replicates
replicates_in_all, cpds_replicates = get_cpds_replicates(df_cpd_med_scores, df_level4)
def get_replicates_classes_per_dose(df, df_lvl4, cpds_replicates):
"""
This function gets all replicates ids for each distinct
no_of_replicates (i.e. number of replicates per cpd) class per dose (1-6)
Returns replicate_class_dict dictionary, with no_of_replicate classes as the keys,
and all the replicate_ids for each no_of_replicate class as the values
"""
df['replicate_id'] = list(cpds_replicates.values())
dose_list = list(set(df_lvl4['Metadata_dose_recode'].unique().tolist()))[1:7]
replicate_class_dict = {}
for dose in dose_list:
for size in df['no_of_replicates'].unique():
rep_lists = []
for idx in range(df[df['no_of_replicates'] == size].shape[0]):
rep_ids = df[df['no_of_replicates'] == size]['replicate_id'].values.tolist()[idx][dose-1]
rep_lists += rep_ids
if size not in replicate_class_dict:
replicate_class_dict[size] = [rep_lists]
else:
replicate_class_dict[size] += [rep_lists]
return replicate_class_dict
cpd_replicate_class_dict = get_replicates_classes_per_dose(df_cpd_med_scores, df_level4, cpds_replicates)
cpd_replicate_class_dict.keys()
def check_similar_replicates(replicates, dose, cpd_dict):
"""This function checks if two replicates are of the same compounds"""
for x in range(len(replicates)):
for y in range(x+1, len(replicates)):
for kys in cpd_dict:
if all(i in cpd_dict[kys][dose-1] for i in [replicates[x], replicates[y]]):
return True
return False
def get_random_replicates(all_replicates, no_of_replicates, dose, replicates_ids, cpd_replicate_dict):
"""
This function return a list of random replicates that are not of the same compounds
or found in the current cpd's size list
"""
while (True):
random_replicates = random.sample(all_replicates, no_of_replicates)
if not (any(rep in replicates_ids for rep in random_replicates) &
(check_similar_replicates(random_replicates, dose, cpd_replicate_dict))):
break
return random_replicates
def get_null_distribution_replicates(
cpd_replicate_class_dict,
dose_list,
replicates_lists,
cpd_replicate_dict,
rand_num = 1000
):
"""
This function returns a null distribution dictionary, with no_of_replicates(replicate class)
as the keys and 1000 lists of randomly selected replicate combinations as the values
for each no_of_replicates class per DOSE(1-6)
"""
random.seed(1903)
null_distribution_reps = {}
for dose in dose_list:
for replicate_class in cpd_replicate_class_dict:
replicates_ids = cpd_replicate_class_dict[replicate_class][dose-1]
replicate_list = []
for idx in range(rand_num):
start_again = True
while (start_again):
rand_cpds = get_random_replicates(replicates_lists[dose-1], replicate_class, dose,
replicates_ids, cpd_replicate_dict)
if rand_cpds not in replicate_list:
start_again = False
replicate_list.append(rand_cpds)
if replicate_class not in null_distribution_reps:
null_distribution_reps[replicate_class] = [replicate_list]
else:
null_distribution_reps[replicate_class] += [replicate_list]
return null_distribution_reps
len(cpds_replicates.keys())
dose_list = list(set(df_level4['Metadata_dose_recode'].unique().tolist()))[1:7]
null_distribution_replicates = get_null_distribution_replicates(
cpd_replicate_class_dict, dose_list, replicates_in_all, cpds_replicates
)
def save_to_pickle(null_distribution, path, file_name):
"""This function saves the null distribution replicates ids into a pickle file"""
if not os.path.exists(path):
os.mkdir(path)
with open(os.path.join(path, file_name), 'wb') as handle:
pickle.dump(null_distribution, handle, protocol=pickle.HIGHEST_PROTOCOL)
#save the null_distribution_moa to pickle
save_to_pickle(null_distribution_replicates, cp_level4_path, 'null_distribution_subsample.pickle')
##load the null_distribution_moa from pickle
with open(os.path.join(cp_level4_path, 'null_distribution_subsample.pickle'), 'rb') as handle:
null_distribution_replicates = pickle.load(handle)
def assert_null_distribution(null_distribution_reps, dose_list):
"""
This function assert that each of the list in the 1000 lists of random replicate
combination (per dose) for each no_of_replicate class are distinct with no duplicates
"""
duplicates_reps = {}
for dose in dose_list:
for keys in null_distribution_reps:
null_dist = null_distribution_reps[keys][dose-1]
for reps in null_dist:
dup_reps = []
new_list = list(filter(lambda x: x != reps, null_dist))
if (len(new_list) != len(null_dist) - 1):
dup_reps.append(reps)
if dup_reps:
if keys not in duplicates_reps:
duplicates_reps[keys] = [dup_reps]
else:
duplicates_reps[keys] += [dup_reps]
return duplicates_reps
duplicate_replicates = assert_null_distribution(null_distribution_replicates, dose_list)
duplicate_replicates ##no duplicates
def calc_null_dist_median_scores(df, dose_num, replicate_lists):
"""
This function calculate the median of the correlation
values for each list in the 1000 lists of random replicate
combination for each no_of_replicate class per dose
"""
df_dose = df[df['Metadata_dose_recode'] == dose_num].copy()
df_dose = df_dose.set_index('replicate_name').rename_axis(None, axis=0)
df_dose.drop(['Metadata_broad_sample', 'Metadata_pert_id', 'Metadata_dose_recode',
'Metadata_Plate', 'Metadata_Well', 'Metadata_broad_id', 'Metadata_moa',
'broad_id', 'pert_iname', 'moa'],
axis = 1, inplace = True)
median_corr_list = []
for rep_list in replicate_lists:
df_reps = df_dose.loc[rep_list].copy()
reps_corr = df_reps.astype('float64').T.corr(method = 'pearson').values
median_corr_val = median(list(reps_corr[np.triu_indices(len(reps_corr), k = 1)]))
median_corr_list.append(median_corr_val)
return median_corr_list
def get_null_dist_median_scores(null_distribution_cpds, dose_list, df):
"""
This function calculate the median correlation scores for all
1000 lists of randomly combined compounds for each no_of_replicate class
across all doses (1-6)
"""
null_distribution_medians = {}
for key in null_distribution_cpds:
median_score_list = []
for dose in dose_list:
replicate_median_scores = calc_null_dist_median_scores(df, dose, null_distribution_cpds[key][dose-1])
median_score_list.append(replicate_median_scores)
null_distribution_medians[key] = median_score_list
return null_distribution_medians
null_distribution_medians = get_null_dist_median_scores(null_distribution_replicates, dose_list, df_level4)
def compute_dose_median_scores(null_dist_medians, dose_list):
"""
This function align median scores per dose, and return a dictionary,
with keys as dose numbers and values as all median null distribution/non-replicate correlation
scores for each dose
"""
median_scores_per_dose = {}
for dose in dose_list:
median_list = []
for keys in null_distribution_medians:
dose_median_list = null_distribution_medians[keys][dose-1]
median_list += dose_median_list
median_scores_per_dose[dose] = median_list
return median_scores_per_dose
dose_null_medians = compute_dose_median_scores(null_distribution_medians, dose_list)
#save the null_distribution_medians_per_dose to pickle
save_to_pickle(dose_null_medians, cp_level4_path, 'null_dist_medians_per_dose_subsample.pickle')
def get_p_value(median_scores_list, df, dose_name, cpd_name):
"""
This function calculate the p-value from the
null_distribution median scores for each compound
"""
actual_med = df.loc[cpd_name, dose_name]
p_value = np.sum(median_scores_list >= actual_med) / len(median_scores_list)
return p_value
def get_moa_p_vals(null_dist_median, dose_list, df_med_values):
"""
This function returns a dict, with compounds as the keys and the compound's
p-values for each dose (1-6) as the values
"""
null_p_vals = {}
for key in null_dist_median:
df_replicate_class = df_med_values[df_med_values['no_of_replicates'] == key]
for cpd in df_replicate_class.index:
dose_p_values = []
for num in dose_list:
dose_name = 'dose_' + str(num)
cpd_p_value = get_p_value(null_dist_median[key][num-1], df_replicate_class, dose_name, cpd)
dose_p_values.append(cpd_p_value)
null_p_vals[cpd] = dose_p_values
sorted_null_p_vals = {key:value for key, value in sorted(null_p_vals.items(), key=lambda item: item[0])}
return sorted_null_p_vals
null_p_vals = get_moa_p_vals(null_distribution_medians, dose_list, df_cpd_med_scores)
df_null_p_vals = pd.DataFrame.from_dict(null_p_vals, orient='index', columns = ['dose_' + str(x) for x in dose_list])
df_null_p_vals['no_of_replicates'] = df_cpd_med_scores['no_of_replicates']
df_null_p_vals.head(10)
def save_to_csv(df, path, file_name):
"""saves dataframes to csv"""
if not os.path.exists(path):
os.mkdir(path)
df.to_csv(os.path.join(path, file_name), index = False)
save_to_csv(df_null_p_vals.reset_index().rename({'index':'cpd'}, axis = 1), cp_level4_path,
'cpd_replicate_p_values_subsample.csv')
cpd_summary_file = pathlib.Path(cp_level4_path, 'cpd_replicate_p_values_melted_subsample.csv')
dose_recode_info = {
'dose_1': '0.04 uM', 'dose_2':'0.12 uM', 'dose_3':'0.37 uM',
'dose_4': '1.11 uM', 'dose_5':'3.33 uM', 'dose_6':'10 uM'
}
# Melt the p values
cpd_score_summary_pval_df = (
df_null_p_vals
.reset_index()
.rename(columns={"index": "compound"})
.melt(
id_vars=["compound", "no_of_replicates"],
value_vars=["dose_1", "dose_2", "dose_3", "dose_4", "dose_5", "dose_6"],
var_name="dose",
value_name="p_value"
)
)
cpd_score_summary_pval_df.dose = cpd_score_summary_pval_df.dose.replace(dose_recode_info)
# Melt the median matching scores
cpd_score_summary_df = (
df_cpd_med_scores
.reset_index()
.rename(columns={"index": "compound"})
.melt(
id_vars=["compound", "no_of_replicates"],
value_vars=["dose_1", "dose_2", "dose_3", "dose_4", "dose_5", "dose_6"],
var_name="dose",
value_name="matching_score"
)
)
cpd_score_summary_df.dose = cpd_score_summary_df.dose.replace(dose_recode_info)
summary_df = (
cpd_score_summary_pval_df
.merge(cpd_score_summary_df, on=["compound", "no_of_replicates", "dose"], how="inner")
.assign(
assay="Cell Painting",
normalization="spherized",
category="subsampled"
)
)
summary_df.to_csv(cpd_summary_file, sep="\t", index=False)
print(summary_df.shape)
summary_df.head()
```
| github_jupyter |
```
# hide
%load_ext autoreload
%autoreload 2
from nbdev import *
# default_exp navi_widget
```
# Navi Widget
```
#exporti
from ipywidgets import Button, IntSlider, HBox, Layout
import warnings
from typing import Callable
#exporti
class NaviGUI(HBox):
def __init__(self, max_im_number: int = 0):
self._im_number_slider = IntSlider(
min=0,
max=max_im_number,
value=0,
description='Image Nr.'
)
self._prev_btn = Button(description='< Previous',
layout=Layout(width='auto'))
self._next_btn = Button(description='Next >',
layout=Layout(width='auto'))
super().__init__(children=[self._prev_btn, self._im_number_slider, self._next_btn],
layout=Layout(display='flex', flex_flow='row wrap', align_items='center'))
#exporti
class NaviLogic:
"""
Acts like an intermediator between GUI and its interactions
"""
def __init__(self, gui: NaviGUI):
self._gui = gui
def slider_updated(self, change: dict):
self._gui._index = change['new']
self.set_slider_value(change['new'])
def set_slider_value(self, index: int):
self._gui._im_number_slider.value = index
def set_slider_max(self, max_im_number: int):
self._gui._im_number_slider.max = max_im_number
def _increment_state_index(self, index: int):
max_im_number = self._gui._max_im_num
safe_index = (self._gui._index + index) % max_im_number
self._gui._index = (safe_index + max_im_number) % max_im_number
self.set_slider_value(self._gui._index)
def check_im_num(self, max_im_number: int):
if not hasattr(self._gui, '_im_number_slider'):
return
self._gui._im_number_slider.max = max_im_number - 1
#export
class Navi(NaviGUI):
"""
Represents simple navigation module with slider.
on_navi_clicked: callable
A callback that runs after every navigation
change. The callback should have, as a
parameter the navi's index.
"""
def __init__(self, max_im_num: int = 1, on_navi_clicked: Callable = None):
super().__init__(max_im_num)
self._max_im_num = max_im_num
self.on_navi_clicked = on_navi_clicked
self._index = 0
self.model = NaviLogic(gui=self)
self._listen_next_click()
self._listen_prev_click()
self._listen_slider_changes()
@property
def index(self) -> int:
return self._index
@index.setter
def index(self, value: int):
self.model.set_slider_value(value)
self._index = value
self._external_call()
@property
def max_im_num(self) -> int:
return self._max_im_num
@max_im_num.setter
def max_im_num(self, value: int):
self.model.set_slider_max(value - 1)
self._max_im_num = value
def _next_clicked(self, *args):
self.model._increment_state_index(1)
def _slider_updated(self, value: dict):
self.model.slider_updated(value)
self._external_call()
def _prev_clicked(self, *args):
self.model._increment_state_index(-1)
def _listen_slider_changes(self):
self._im_number_slider.observe(
self._slider_updated, names='value'
)
def _listen_next_click(self):
self._next_btn.on_click(self._next_clicked)
def _listen_prev_click(self):
self._prev_btn.on_click(self._prev_clicked)
def _external_call(self):
if self.on_navi_clicked:
self.on_navi_clicked(self._index)
else:
warnings.warn(
"Navi callable was not defined."
"The navigation will not trigger any action!"
)
# it start navi with slider index at 0
navi = Navi(6)
assert navi._im_number_slider.value == 0
# it changes state if slider.value changes
navi._im_number_slider.value = 2
assert navi._index == 2
# it changes state and slider.value if button is clicked
navi._next_btn.click()
assert navi._index == 3
assert navi._im_number_slider.value == 3
navi._prev_btn.click()
assert navi._index == 2
assert navi._im_number_slider.value == 2
# it changes slider.max if navi changes max im num
navi.max_im_num = 6
assert navi._im_number_slider.max == 5
# it changes slider.index if navi changes its index
navi.index = 3
assert navi._im_number_slider.value == 3
# testing callback
callback_index = 0
def increment_callback(index):
global callback_index
callback_index = index
navi.on_navi_clicked = increment_callback
navi._next_btn.click()
assert callback_index == 4
navi._prev_btn.click()
assert callback_index == 3
navi.index = 2
assert callback_index == 2
navi
#hide
from nbdev.export import notebook2script
notebook2script()
```
| github_jupyter |
<table width="100%">
<tr style="border-bottom:solid 2pt #009EE3">
<td class="header_buttons">
<a href="generation_of_time_axis.zip" download><img src="../../images/icons/download.png" alt="biosignalsnotebooks | download button"></a>
</td>
<td class="header_buttons">
<a href="https://mybinder.org/v2/gh/biosignalsplux/biosignalsnotebooks/mybinder_complete?filepath=biosignalsnotebooks_environment%2Fcategories%2FPre-Process%2Fgeneration_of_time_axis.dwipynb" target="_blank"><img src="../../images/icons/program.png" alt="biosignalsnotebooks | binder server" title="Be creative and test your solutions !"></a>
</td>
<td></td>
<td class="header_icons">
<a href="../MainFiles/biosignalsnotebooks.ipynb"><img src="../../images/icons/home.png" alt="biosignalsnotebooks | home button"></a>
</td>
<td class="header_icons">
<a href="../MainFiles/contacts.ipynb"><img src="../../images/icons/contacts.png" alt="biosignalsnotebooks | contacts button"></a>
</td>
<td class="header_icons">
<a href="https://github.com/biosignalsplux/biosignalsnotebooks" target="_blank"><img src="../../images/icons/github.png" alt="biosignalsnotebooks | github button"></a>
</td>
<td class="header_logo">
<img src="../../images/ost_logo.png" alt="biosignalsnotebooks | project logo">
</td>
</tr>
</table>
<link rel="stylesheet" href="../../styles/theme_style.css">
<!--link rel="stylesheet" href="../../styles/header_style.css"-->
<link rel="stylesheet" href="https://cdnjs.cloudflare.com/ajax/libs/font-awesome/4.7.0/css/font-awesome.min.css">
<table width="100%">
<tr>
<td id="image_td" width="15%" class="header_image_color_4"><div id="image_img" class="header_image_4"></div></td>
<td class="header_text"> Generation of a time axis (conversion of samples into seconds) </td>
</tr>
</table>
<div id="flex-container">
<div id="diff_level" class="flex-item">
<strong>Difficulty Level:</strong> <span class="fa fa-star checked"></span>
<span class="fa fa-star checked"></span>
<span class="fa fa-star"></span>
<span class="fa fa-star"></span>
<span class="fa fa-star"></span>
</div>
<div id="tag" class="flex-item-tag">
<span id="tag_list">
<table id="tag_list_table">
<tr>
<td class="shield_left">Tags</td>
<td class="shield_right" id="tags">pre-process☁time☁conversion</td>
</tr>
</table>
</span>
<!-- [OR] Visit https://img.shields.io in order to create a tag badge-->
</div>
</div>
All electrophysiological signals, collected by PLUX acquisition systems, are, in its essence, time series.
Raw data contained in the generated .txt, .h5 and .edf files consists in samples and each sample value is in a raw value with 8 or 16 bits that needs to be converted to a physical unit by the respective transfer function.
PLUX has examples of conversion rules for each sensor (in separate .pdf files), which may be accessed at <a href="http://biosignalsplux.com/en/learn/documentation">"Documentation>>Sensors" section <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></a> of <strong><span class="color2">biosignalsplux</span></strong> website.
<img src="../../images/pre-process/generation_of_time_axis/sensors_section.gif">
Although each file returned by <strong><span class="color2">OpenSignals</span></strong> contains a sequence number linked to each sample, giving a notion of "time order" and that can be used as x axis, working with real time units is, in many occasions, more intuitive.
So, in the present **<span class="color5">Jupyter Notebook</span>** is described how to associate a time axis to an acquired signal, taking into consideration the number of acquired samples and the respective sampling rate.
<hr>
<p class="steps">1 - Importation of the needed packages </p>
```
# Package dedicated to download files remotely
from wget import download
# Package used for loading data from the input text file and for generation of a time axis
from numpy import loadtxt, linspace
# Package used for loading data from the input h5 file
import h5py
# biosignalsnotebooks own package.
import biosignalsnotebooks as bsnb
```
<p class="steps"> A - Text Files</p>
<p class="steps">A1 - Load of support data inside .txt file (described in a <span class="color5">Jupyter Notebook</span> entitled <a href="../Load/open_txt.ipynb"><strong> "Load acquired data from .txt file" <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></strong></a>) </p>
```
# Download of the text file followed by content loading.
txt_file_url = "https://drive.google.com/uc?export=download&id=1m7E7PnKLfcd4HtOASH6vRmyBbCmIEkLf"
txt_file = download(txt_file_url, out="download_file_name.txt")
txt_file = open(txt_file, "r")
# [Internal code for overwrite file if already exists]
import os
import shutil
txt_file.close()
if os.path.exists("download_file_name.txt"):
shutil.move(txt_file.name,"download_file_name.txt")
txt_file = "download_file_name.txt"
txt_file = open(txt_file, "r")
```
<p class="steps">A2 - Load of acquisition samples (in this case from the third column of the text file - list entry 2)</p>
```
txt_signal = loadtxt(txt_file)[:, 2]
```
<p class="steps">A3 - Determination of the number of acquired samples</p>
```
# Number of acquired samples
nbr_samples_txt = len(txt_signal)
from sty import fg, rs
print(fg(98,195,238) + "\033[1mNumber of samples (.txt file):\033[0m" + fg.rs + " " + str(nbr_samples_txt))
```
<p class="steps"> B - H5 Files</p>
<p class="steps">B1 - Load of support data inside .h5 file (described in the <span class="color5">Jupyter Notebook</span> entitled <a href="../Load/open_h5.ipynb"><strong> "Load acquired data from .h5 file"<img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></strong></a>) </p>
```
# Download of the .h5 file followed by content loading.
h5_file_url = "https://drive.google.com/uc?export=download&id=1UgOKuOMvHTm3LlQ_e7b6R_qZL5cdL4Rv"
h5_file = download(h5_file_url, out="download_file_name.h5")
h5_object = h5py.File(h5_file)
# [Internal code for overwrite file if already exists]
import os
import shutil
h5_object.close()
if os.path.exists("download_file_name.h5"):
shutil.move(h5_file,"download_file_name.h5")
h5_file = "download_file_name.h5"
h5_object = h5py.File(h5_file)
```
<p class="steps">B2 - Load of acquisition samples inside .h5 file</p>
```
# Device mac-address.
mac_address = list(h5_object.keys())[0]
# Access to signal data acquired by the device identified by "mac_address" in "channel_1".
h5_signal = list(h5_object.get(mac_address).get("raw").get("channel_1"))
```
<p class="steps">B3 - Determination of the number of acquired samples</p>
```
# Number of acquired samples
nbr_samples_h5 = len(h5_signal)
print(fg(232,77,14) + "\033[1mNumber of samples (.h5 file):\033[0m" + fg.rs + " " + str(nbr_samples_h5))
```
As it can be seen, the number of samples is equal for both file types.
```
print(fg(98,195,238) + "\033[1mNumber of samples (.txt file):\033[0m" + fg.rs + " " + str(nbr_samples_txt))
print(fg(232,77,14) + "\033[1mNumber of samples (.h5 file):\033[0m" + fg.rs + " " + str(nbr_samples_h5))
```
So, we can simplify and reduce the number of variables:
```
nbr_samples = nbr_samples_txt
```
Like described in the Notebook intro, for generating a time-axis it is needed the <strong><span class="color4">number of acquired samples</span></strong> and the <strong><span class="color7">sampling rate</span></strong>.
Currently the only unknown parameter is the <strong><span class="color7">sampling rate</span></strong>, which can be easily accessed for .txt and .h5 files as described in <a href="../Load/signal_loading_preparatory_steps.ipynb" target="_blank">"Signal Loading - Working with File Header"<img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></a>.
For our acquisition the sampling rate is:
```
sampling_rate = 1000 # Hz
```
<p class="steps">AB4 - Determination of acquisition time in seconds</p>
```
# Conversion between sample number and seconds
acq_time = nbr_samples / sampling_rate
print ("Acquisition Time: " + str(acq_time) + " s")
```
<p class="steps">AB5 - Creation of the time axis (between 0 and 417.15 seconds) through <span class="color4">linspace</span> function</p>
```
time_axis = linspace(0, acq_time, nbr_samples)
print ("Time-Axis: \n" + str(time_axis))
```
<p class="steps">AB6 - Plot of the acquired signal (first 10 seconds) with the generated time-axis</p>
```
bsnb.plot(time_axis[:10*sampling_rate], txt_signal[:10*sampling_rate])
```
*This procedure can be automatically done by **generate_time** function in **conversion** module of **<span class="color2">biosignalsnotebooks</span>** package*
```
time_axis_auto = bsnb.generate_time(h5_file_url)
from numpy import array
print ("Time-Axis returned by generateTime function:")
print (array(time_axis_auto))
```
Time is a really important "dimension" in our daily lives and particularly on signal processing analysis. Without a time "anchor" like <strong><span class="color7">sampling rate</span></strong> it is very difficult to link the acquired digital data with real events.
Concepts like "temporal duration" or "time rate" become meaningless, being more difficult to take adequate conclusions.
However, as can be seen, a researcher in possession of the data to process and a single parameter (sampling rate) can easily generate a time-axis, following the demonstrated procedure.
<strong><span class="color7">We hope that you have enjoyed this guide. </span><span class="color2">biosignalsnotebooks</span><span class="color4"> is an environment in continuous expansion, so don't stop your journey and learn more with the remaining <a href="../MainFiles/biosignalsnotebooks.ipynb">Notebooks <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></a></span></strong> !
<hr>
<table width="100%">
<tr>
<td class="footer_logo">
<img src="../../images/ost_logo.png" alt="biosignalsnotebooks | project logo [footer]">
</td>
<td width="40%" style="text-align:left">
<a href="../MainFiles/aux_files/biosignalsnotebooks_presentation.pdf" target="_blank">☌ Project Presentation</a>
<br>
<a href="https://github.com/biosignalsplux/biosignalsnotebooks" target="_blank">☌ GitHub Repository</a>
<br>
<a href="https://pypi.org/project/biosignalsnotebooks/" target="_blank">☌ How to install biosignalsnotebooks Python package ?</a>
<br>
<a href="https://www.biosignalsplux.com/notebooks/Categories/MainFiles/signal_samples.ipynb">☌ Signal Library</a>
</td>
<td width="40%" style="text-align:left">
<a href="https://www.biosignalsplux.com/notebooks/Categories/MainFiles/biosignalsnotebooks.ipynb">☌ Notebook Categories</a>
<br>
<a href="https://www.biosignalsplux.com/notebooks/Categories/MainFiles/by_diff.ipynb">☌ Notebooks by Difficulty</a>
<br>
<a href="https://www.biosignalsplux.com/notebooks/Categories/MainFiles/by_signal_type.ipynb">☌ Notebooks by Signal Type</a>
<br>
<a href="https://www.biosignalsplux.com/notebooks/Categories/MainFiles/by_tag.ipynb">☌ Notebooks by Tag</a>
</td>
</tr>
</table>
```
from biosignalsnotebooks.__notebook_support__ import css_style_apply
css_style_apply()
%%html
<script>
// AUTORUN ALL CELLS ON NOTEBOOK-LOAD!
require(
['base/js/namespace', 'jquery'],
function(jupyter, $) {
$(jupyter.events).on("kernel_ready.Kernel", function () {
console.log("Auto-running all cells-below...");
jupyter.actions.call('jupyter-notebook:run-all-cells-below');
jupyter.actions.call('jupyter-notebook:save-notebook');
});
}
);
</script>
```
| github_jupyter |
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
```
# Unsupervised Learning Part 2 -- Clustering
Clustering is the task of gathering samples into groups of similar
samples according to some predefined similarity or distance (dissimilarity)
measure, such as the Euclidean distance.
<img width="60%" src='figures/clustering.png'/>
In this section we will explore a basic clustering task on some synthetic and real-world datasets.
Here are some common applications of clustering algorithms:
- Compression for data reduction
- Summarizing data as a reprocessing step for recommender systems
- Similarly:
- grouping related web news (e.g. Google News) and web search results
- grouping related stock quotes for investment portfolio management
- building customer profiles for market analysis
- Building a code book of prototype samples for unsupervised feature extraction
Let's start by creating a simple, 2-dimensional, synthetic dataset:
```
from sklearn.datasets import make_blobs
X, y = make_blobs(random_state=42)
X.shape
plt.figure(figsize=(8, 8))
plt.scatter(X[:, 0], X[:, 1])
```
In the scatter plot above, we can see three separate groups of data points and we would like to recover them using clustering -- think of "discovering" the class labels that we already take for granted in a classification task.
Even if the groups are obvious in the data, it is hard to find them when the data lives in a high-dimensional space, which we can't visualize in a single histogram or scatterplot.
Now we will use one of the simplest clustering algorithms, K-means.
This is an iterative algorithm which searches for three cluster
centers such that the distance from each point to its cluster is
minimized. The standard implementation of K-means uses the Euclidean distance, which is why we want to make sure that all our variables are measured on the same scale if we are working with real-world datastets. In the previous notebook, we talked about one technique to achieve this, namely, standardization.
<br/>
<div class="alert alert-success">
<b>Question</b>:
<ul>
<li>
what would you expect the output to look like?
</li>
</ul>
</div>
```
from sklearn.cluster import KMeans
kmeans = KMeans(n_clusters=3, random_state=42)
```
We can get the cluster labels either by calling fit and then accessing the
``labels_`` attribute of the K means estimator, or by calling ``fit_predict``.
Either way, the result contains the ID of the cluster that each point is assigned to.
```
labels = kmeans.fit_predict(X)
labels
np.all(y == labels)
```
Let's visualize the assignments that have been found
```
plt.figure(figsize=(8, 8))
plt.scatter(X[:, 0], X[:, 1], c=labels)
```
Compared to the true labels:
```
plt.figure(figsize=(8, 8))
plt.scatter(X[:, 0], X[:, 1], c=y)
```
Here, we are probably satisfied with the clustering results. But in general we might want to have a more quantitative evaluation. How about comparing our cluster labels with the ground truth we got when generating the blobs?
```
from sklearn.metrics import confusion_matrix, accuracy_score
print('Accuracy score:', accuracy_score(y, labels))
print(confusion_matrix(y, labels))
np.mean(y == labels)
```
<div class="alert alert-success">
<b>EXERCISE</b>:
<ul>
<li>
After looking at the "True" label array y, and the scatterplot and `labels` above, can you figure out why our computed accuracy is 0.0, not 1.0, and can you fix it?
</li>
</ul>
</div>
Even though we recovered the partitioning of the data into clusters perfectly, the cluster IDs we assigned were arbitrary,
and we can not hope to recover them. Therefore, we must use a different scoring metric, such as ``adjusted_rand_score``, which is invariant to permutations of the labels:
```
from sklearn.metrics import adjusted_rand_score
adjusted_rand_score(y, labels)
```
One of the "short-comings" of K-means is that we have to specify the number of clusters, which we often don't know *apriori*. For example, let's have a look what happens if we set the number of clusters to 2 in our synthetic 3-blob dataset:
```
kmeans = KMeans(n_clusters=2, random_state=42)
labels = kmeans.fit_predict(X)
plt.figure(figsize=(8, 8))
plt.scatter(X[:, 0], X[:, 1], c=labels)
kmeans.cluster_centers_
```
#### The Elbow Method
The Elbow method is a "rule-of-thumb" approach to finding the optimal number of clusters. Here, we look at the cluster dispersion for different values of k:
```
distortions = []
for i in range(1, 11):
km = KMeans(n_clusters=i,
random_state=0)
km.fit(X)
distortions.append(km.inertia_)
plt.plot(range(1, 11), distortions, marker='o')
plt.xlabel('Number of clusters')
plt.ylabel('Distortion')
plt.show()
```
Then, we pick the value that resembles the "pit of an elbow." As we can see, this would be k=3 in this case, which makes sense given our visual expection of the dataset previously.
**Clustering comes with assumptions**: A clustering algorithm finds clusters by making assumptions with samples should be grouped together. Each algorithm makes different assumptions and the quality and interpretability of your results will depend on whether the assumptions are satisfied for your goal. For K-means clustering, the model is that all clusters have equal, spherical variance.
**In general, there is no guarantee that structure found by a clustering algorithm has anything to do with what you were interested in**.
We can easily create a dataset that has non-isotropic clusters, on which kmeans will fail:
```
plt.figure(figsize=(12, 12))
n_samples = 1500
random_state = 170
X, y = make_blobs(n_samples=n_samples, random_state=random_state)
# Incorrect number of clusters
y_pred = KMeans(n_clusters=2, random_state=random_state).fit_predict(X)
plt.subplot(221)
plt.scatter(X[:, 0], X[:, 1], c=y_pred)
plt.title("Incorrect Number of Blobs")
# Anisotropicly distributed data
transformation = [[0.60834549, -0.63667341], [-0.40887718, 0.85253229]]
X_aniso = np.dot(X, transformation)
y_pred = KMeans(n_clusters=3, random_state=random_state).fit_predict(X_aniso)
plt.subplot(222)
plt.scatter(X_aniso[:, 0], X_aniso[:, 1], c=y_pred)
plt.title("Anisotropicly Distributed Blobs")
# Different variance
X_varied, y_varied = make_blobs(n_samples=n_samples,
cluster_std=[1.0, 2.5, 0.5],
random_state=random_state)
y_pred = KMeans(n_clusters=3, random_state=random_state).fit_predict(X_varied)
plt.subplot(223)
plt.scatter(X_varied[:, 0], X_varied[:, 1], c=y_pred)
plt.title("Unequal Variance")
# Unevenly sized blobs
X_filtered = np.vstack((X[y == 0][:500], X[y == 1][:100], X[y == 2][:10]))
y_pred = KMeans(n_clusters=3,
random_state=random_state).fit_predict(X_filtered)
plt.subplot(224)
plt.scatter(X_filtered[:, 0], X_filtered[:, 1], c=y_pred)
plt.title("Unevenly Sized Blobs")
```
## Some Notable Clustering Routines
The following are two well-known clustering algorithms.
- `sklearn.cluster.KMeans`: <br/>
The simplest, yet effective clustering algorithm. Needs to be provided with the
number of clusters in advance, and assumes that the data is normalized as input
(but use a PCA model as preprocessor).
- `sklearn.cluster.MeanShift`: <br/>
Can find better looking clusters than KMeans but is not scalable to high number of samples.
- `sklearn.cluster.DBSCAN`: <br/>
Can detect irregularly shaped clusters based on density, i.e. sparse regions in
the input space are likely to become inter-cluster boundaries. Can also detect
outliers (samples that are not part of a cluster).
- `sklearn.cluster.AffinityPropagation`: <br/>
Clustering algorithm based on message passing between data points.
- `sklearn.cluster.SpectralClustering`: <br/>
KMeans applied to a projection of the normalized graph Laplacian: finds
normalized graph cuts if the affinity matrix is interpreted as an adjacency matrix of a graph.
- `sklearn.cluster.Ward`: <br/>
Ward implements hierarchical clustering based on the Ward algorithm,
a variance-minimizing approach. At each step, it minimizes the sum of
squared differences within all clusters (inertia criterion).
Of these, Ward, SpectralClustering, DBSCAN and Affinity propagation can also work with precomputed similarity matrices.
<img src="figures/cluster_comparison.png" width="900">
<div class="alert alert-success">
<b>EXERCISE: digits clustering</b>:
<ul>
<li>
Perform K-means clustering on the digits data, searching for ten clusters.
Visualize the cluster centers as images (i.e. reshape each to 8x8 and use
``plt.imshow``) Do the clusters seem to be correlated with particular digits? What is the ``adjusted_rand_score``?
</li>
<li>
Visualize the projected digits as in the last notebook, but this time use the
cluster labels as the color. What do you notice?
</li>
</ul>
</div>
```
from sklearn.datasets import load_digits
digits = load_digits()
# ...
# %load solutions/08B_digits_clustering.py
```
| github_jupyter |
# Import tools
```
# built-in utilities
import copy
import os
import time
import datetime
# data tools
import numpy as np
# pytorch
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader, TensorDataset
import torchvision
from torchvision import datasets, models, transforms
from torch.utils.tensorboard import SummaryWriter
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# visualization
import matplotlib.pyplot as plt
%matplotlib inline
```
# Load data
```
# define data transformer
transformation = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))]
)
## download data
# train
trainDataset = datasets.MNIST(
"data", train=True, transform=transformation, download=True
)
# test
testDataset = datasets.MNIST(
"data", train=False, transform=transformation, download=True
)
## load data
trainDataLoader = torch.utils.data.DataLoader(trainDataset, batch_size=128, shuffle=True)
testDataLoader = torch.utils.data.DataLoader(testDataset, batch_size=128, shuffle=True)
# plot sample image
def plotSample(image):
image = image.numpy()
mean = 0.1307
std = 0.3081
image = (mean * image) + std
plt.imshow(image, cmap="gray")
sample = iter(trainDataLoader.dataset.data)
plotSample(next(sample))
```
# Question 1
## Question 1, Part 1
```
# set model architecture
class fcNet(nn.Module):
def __init__(self):
super(fcNet, self).__init__()
self.inputLayer = nn.Linear(784, 1024)
self.fullyConnected1 = nn.Linear(1024, 1024)
self.fullyConnected2 = nn.Linear(1024, 1024)
self.fullyConnected3 = nn.Linear(1024, 1024)
self.fullyConnected4 = nn.Linear(1024, 1024)
self.outputLayer = nn.Linear(1024, 10)
def forward(self, x):
x = F.relu(self.inputLayer(x))
x = F.relu(self.fullyConnected1(x))
x = F.relu(self.fullyConnected2(x))
x = F.relu(self.fullyConnected3(x))
x = F.relu(self.fullyConnected4(x))
x = F.log_softmax(self.outputLayer(x), dim=1)
return x
# set input kwargs as object attributes
class ParamConfig:
def __init__(self, **kwargs):
for key, value in kwargs.items():
setattr(self, key, value)
# configure all necessary parameters
modelParams = ParamConfig(
model = fcNet,
optimizer = torch.optim.Adam,
criterion = F.nll_loss,
trainDataLoader = torch.utils.data.DataLoader(trainDataset, batch_size=128, shuffle=True),
testDataLoader = torch.utils.data.DataLoader(testDataset, batch_size=128, shuffle=True),
cuda = True if torch.cuda.is_available() else False,
device = torch.device("cuda" if torch.cuda.is_available() else "cpu"),
seed = 0,
lr = 0.001,
epochs = 25,
saveModel = True,
)
class PyTorchTrainer:
"""
"""
def __init__(self, config):
self.globaliter = 0
# data loaders
self.trainDataLoader = config.trainDataLoader
self.testDataLoader = config.testDataLoader
# random seed settings
self.seed = config.seed
torch.manual_seed(self.seed)
# device settings
self.cuda = config.cuda
self.device = config.device
# model training settings
self.model = config.model().to(self.device)
self.lr = config.lr
self.epochs = config.epochs
self.optimizer = config.optimizer(self.model.parameters(), lr=self.lr)
self.criterion = config.criterion
# save model
self.saveModel = config.saveModel
# statistics
self.trainLoss = []
self.testLoss = []
self.testAccuracy = []
def train(self, epoch):
# set model to train mode
self.model.train()
print("*" * 80)
# iterate through batches
for batchIdx, (data, target) in enumerate(self.trainDataLoader):
self.globaliter += 1
# reshape data as needed and send data to GPU if available
data = data.reshape(-1, 28*28).to(self.device)
target = target.to(self.device)
# zero out gradients
self.optimizer.zero_grad()
# generate predictiona
preds = self.model(data)
# calculate loss given current predictions vs. ground truth
loss = self.criterion(preds, target)
# back propagate error and optimize weights
loss.backward()
self.optimizer.step()
# capture batch loss
self.trainLoss.append(loss)
if batchIdx % 100 == 0:
print("Train Epoch: {} | Batch: {} [Processed {}/{} ({:.0f}%)]\tLoss: {:.6f}".format(
epoch, batchIdx, batchIdx * len(data), len(self.trainDataLoader.dataset),
100. * batchIdx / len(self.trainDataLoader), loss.item()))
print()
def test(self, epoch):
# set model to eval mode
self.model.eval()
testLoss = 0
correct = 0
# turn off gradients
with torch.no_grad():
# iterate through batches
for batchIdx, (data, target) in enumerate(self.testDataLoader):
# reshape data as needed and send data to GPU if available
data = data.reshape(-1, 28*28).to(self.device)
target = target.to(self.device)
# generate predictiona
preds = self.model(data)
# calculate loss given current predictions vs. ground truth
testLoss = self.criterion(preds, target).item()
preds = preds.argmax(dim=1, keepdim=True)
# capture count of correct answers
correct += preds.eq(target.view_as(preds)).sum().item()
# capture batch loss
self.testLoss.append(testLoss)
# overall epoch loss and accuracy
testLoss /= len(self.testDataLoader.dataset)
accuracy = 100. * correct / len(self.testDataLoader.dataset)
# capture batch loss
self.testAccuracy.append(accuracy)
print('Test set: Average loss: {:.4f}, Accuracy: {}/{} ({:.2f}%)\n\n'.format(
testLoss, correct, len(self.testDataLoader.dataset), accuracy))
### fit model
# instantiate model object
trainer = PyTorchTrainer(config=modelParams)
# iterate fitting procedure over specified epoch count
for epoch in range(1, trainer.epochs + 1):
trainer.train(epoch)
trainer.test(epoch)
# save model
if trainer.saveModel:
if not os.path.isdir("models/"):
os.mkdir("models/")
PATH = "models/_hw3_q1_baseline.pt"
torch.save(trainer.model.state_dict(), PATH)
# plot test accuracy
fig, ax = plt.subplots(figsize=(20,10), facecolor="white")
ax.plot(trainer.testAccuracy)
plt.title("Test set accuracy")
plt.show()
```
## Question 1, Part 3
```
# load model
PATH = "models/_hw3_q1_baseline.pt"
model = fcNet().to(device)
model.load_state_dict(torch.load(PATH))
model.eval()
# SVD - input layer
wIn = model.inputLayer.weight
uIn, sIn, vIn = torch.svd(wIn)
print("SVD - input layer")
print("*"*40 + "\n")
print("w shape: {}".format(wIn.shape))
print()
print("u shape: {}".format(uIn.shape))
print("s shape: {}".format(sIn.shape))
print("v shape: {}".format(vIn.shape))
wInHat = torch.mm(uIn, torch.mm(sIn.diag(), vIn.t()))
print()
print("w hat shape: {}".format(wInHat.shape))
# SVD - Fully connected 1
wFc1 = model.fullyConnected1.weight
uFc1, sFc1, vFc1 = torch.svd(wFc1)
print("SVD - Fully connected 1")
print("*"*40 + "\n")
print("w shape: {}".format(wFc1.shape))
print()
print("u shape: {}".format(uFc1.shape))
print("s shape: {}".format(sFc1.shape))
print("v shape: {}".format(vFc1.shape))
wFc1Hat = torch.mm(uFc1, torch.mm(sFc1.diag(), vFc1.t()))
print()
print("w hat shape: {}".format(wFc1Hat.shape))
# SVD - Fully connected 2
wFc2 = model.fullyConnected2.weight
uFc2, sFc2, vFc2 = torch.svd(wFc2)
print("SVD - Fully connected 2")
print("*"*40 + "\n")
print("w shape: {}".format(wFc2.shape))
print()
print("u shape: {}".format(uFc2.shape))
print("s shape: {}".format(sFc2.shape))
print("v shape: {}".format(vFc2.shape))
wFc2Hat = torch.mm(uFc2, torch.mm(sFc2.diag(), vFc2.t()))
print()
print("w hat shape: {}".format(wFc2Hat.shape))
# SVD - Fully connected 3
wFc3 = model.fullyConnected3.weight
uFc3, sFc3, vFc3 = torch.svd(wFc3)
print("SVD - Fully connected 3")
print("*"*40 + "\n")
print("w shape: {}".format(wFc3.shape))
print()
print("u shape: {}".format(uFc3.shape))
print("s shape: {}".format(sFc3.shape))
print("v shape: {}".format(vFc3.shape))
wFc3Hat = torch.mm(uFc3, torch.mm(sFc3.diag(), vFc3.t()))
print()
print("w hat shape: {}".format(wFc3Hat.shape))
# SVD - Fully connected 4
wFc4 = model.fullyConnected4.weight
uFc4, sFc4, vFc4 = torch.svd(wFc4)
print("SVD - Fully connected 4")
print("*"*40 + "\n")
print("w shape: {}".format(wFc4.shape))
print()
print("u shape: {}".format(uFc4.shape))
print("s shape: {}".format(sFc4.shape))
print("v shape: {}".format(vFc4.shape))
wFc4Hat = torch.mm(uFc4, torch.mm(sFc4.diag(), vFc4.t()))
print()
print("w hat shape: {}".format(wFc4Hat.shape))
```
## Question 1, Part 4
```
D = 10
compressExample = torch.mm(uIn[:,:D], torch.mm(sIn[:D].diag(), vIn[:,:D].t()))
print("compressed matrix shape: {}".format(compressExample.shape))
```
## Question 1, Part 5
do one feed forward in a network for each value of D
```
# load model
PATH = "models/_hw3_q1_baseline.pt"
model = fcNet().to(device)
model.load_state_dict(torch.load(PATH))
model.eval()
# load model
PATH = "models/_hw3_q1_baseline.pt"
model = fcNet().to(device)
model.load_state_dict(torch.load(PATH))
model.eval()
scores = []
Ds = [10, 20, 50, 100, 200, 784]
model.eval()
for D in Ds:
model.inputLayer.weight.data = torch.mm(uIn[:,:D], torch.mm(sIn[:D].diag(), vIn[:,:D].t()))
model.fullyConnected1.weight.data = torch.mm(uFc1[:,:D], torch.mm(sFc1[:D].diag(), vFc1[:,:D].t()))
model.fullyConnected2.weight.data = torch.mm(uFc2[:,:D], torch.mm(sFc2[:D].diag(), vFc2[:,:D].t()))
model.fullyConnected3.weight.data = torch.mm(uFc3[:,:D], torch.mm(sFc3[:D].diag(), vFc3[:,:D].t()))
model.fullyConnected4.weight.data = torch.mm(uFc4[:,:D], torch.mm(sFc4[:D].diag(), vFc4[:,:D].t()))
correct = 0
# iterate through batches
for batchIdx, (data, target) in enumerate(testDataLoader):
# reshape data as needed and send data to GPU if available
data = data.reshape(-1, 28*28).to("cuda")
target = target.to("cuda")
# generate predictiona
preds = model(data)
# calculate loss given current predictions vs. ground truth
preds = preds.argmax(dim=1, keepdim=True)
# capture count of correct answers
correct += preds.eq(target.view_as(preds)).sum().item()
# overall epoch loss and accuracy
accuracy = 100. * correct / len(testDataLoader.dataset)
print("D value: {} | Accuracy = {}".format(D, accuracy))
```
## Question 1, Part 6
train the model and update the weights
```
D = 20
uIn, vIn = uIn[:,:D], torch.mm(sIn[:D].diag(), vIn[:,:D].t())
uFc1, vFc1 = uFc1[:,:D], torch.mm(sFc1[:D].diag(), vFc1[:,:D].t())
uFc2, vFc2 = uFc2[:,:D], torch.mm(sFc2[:D].diag(), vFc2[:,:D].t())
uFc3, vFc3 = uFc3[:,:D], torch.mm(sFc3[:D].diag(), vFc3[:,:D].t())
uFc4, vFc4 = uFc4[:,:D], torch.mm(sFc4[:D].diag(), vFc4[:,:D].t())
print(uIn.shape)
print(vIn.shape)
print()
print(uFc1.shape)
print(vFc1.shape)
print()
print(uFc2.shape)
print(vFc2.shape)
print()
print(uFc3.shape)
print(vFc3.shape)
print()
print(uFc4.shape)
print(vFc4.shape)
print()
# dont forget about the bias
# 10 layers?
# set model architecture
class fcNetCompressed(nn.Module):
def __init__(self):
super(fcNetCompressed, self).__init__()
self.inputLayer_V = nn.Linear(784, 20)
self.inputLayer_U = nn.Linear(20, 1024)
self.fullyConnected1_U = nn.Linear(1024, 20)
self.fullyConnected1_V = nn.Linear(20, 1024)
self.fullyConnected2_U = nn.Linear(1024, 20)
self.fullyConnected2_V = nn.Linear(20, 1024)
self.fullyConnected3_U = nn.Linear(1024, 20)
self.fullyConnected3_V = nn.Linear(20, 1024)
self.fullyConnected4_U = nn.Linear(1024, 20)
self.fullyConnected4_V = nn.Linear(20, 1024)
self.outputLayer = nn.Linear(1024, 10)
def forward(self, x):
x = F.relu(self.inputLayer_V(x))
x = F.relu(self.inputLayer_U(x))
x = F.relu(self.fullyConnected1_U(x))
x = F.relu(self.fullyConnected1_V(x))
x = F.relu(self.fullyConnected2_U(x))
x = F.relu(self.fullyConnected2_V(x))
x = F.relu(self.fullyConnected3_U(x))
x = F.relu(self.fullyConnected3_V(x))
x = F.relu(self.fullyConnected4_U(x))
x = F.relu(self.fullyConnected4_V(x))
x = F.log_softmax(self.outputLayer(x), dim=1)
return x
model = fcNetCompressed()
### initialize weights and biases
# input weights and bias units
model.inputLayer_V.weight.data = vIn
model.inputLayer_U.weight.data = uIn
model.inputLayer_V.bias.data = torch.zeros_like(model.inputLayer_V.bias.data)
model.inputLayer_U.bias.data = torch.zeros_like(model.inputLayer_U.bias.data)
# input weights and bias units
model.fullyConnected1_U.weight.data = uFc1
model.fullyConnected1_V.weight.data = vFc1
model.fullyConnected1_U.bias.data = torch.zeros_like(model.fullyConnected1_U.bias.data)
model.fullyConnected1_V.bias.data = torch.zeros_like(model.fullyConnected1_V.bias.data)
# input weights and bias units
model.fullyConnected2_U.weight.data = uFc2
model.fullyConnected2_V.weight.data = vFc2
model.fullyConnected2_U.bias.data = torch.zeros_like(model.fullyConnected2_U.bias.data)
model.fullyConnected2_V.bias.data = torch.zeros_like(model.fullyConnected2_V.bias.data)
# input weights and bias units
model.fullyConnected3_U.weight.data = uFc3
model.fullyConnected3_V.weight.data = vFc3
model.fullyConnected3_U.bias.data = torch.zeros_like(model.fullyConnected3_U.bias.data)
model.fullyConnected3_V.bias.data = torch.zeros_like(model.fullyConnected3_V.bias.data)
# input weights and bias units
model.fullyConnected4_U.weight.data = uFc4
model.fullyConnected4_V.weight.data = vFc4
model.fullyConnected4_U.bias.data = torch.zeros_like(model.fullyConnected4_U.bias.data)
model.fullyConnected4_V.bias.data = torch.zeros_like(model.fullyConnected4_V.bias.data)
# set input kwargs as object attributes
class ParamConfig:
def __init__(self, **kwargs):
for key, value in kwargs.items():
setattr(self, key, value)
# configure all necessary parameters
modelParams = ParamConfig(
model = fcNetCompressed,
optimizer = torch.optim.Adam,
criterion = F.nll_loss,
trainDataLoader = torch.utils.data.DataLoader(trainDataset, batch_size=128, shuffle=True),
testDataLoader = torch.utils.data.DataLoader(testDataset, batch_size=128, shuffle=True),
cuda = True if torch.cuda.is_available() else False,
device = torch.device("cuda" if torch.cuda.is_available() else "cpu"),
seed = 0,
lr = 0.0001,
epochs = 100,
saveModel = True,
)
### fit model
# instantiate model object
trainer = PyTorchTrainer(config=modelParams)
# iterate fitting procedure over specified epoch count
for epoch in range(1, trainer.epochs + 1):
trainer.train(epoch)
trainer.test(epoch)
# save model
if trainer.saveModel:
if not os.path.isdir("models/"):
os.mkdir("models/")
PATH = "models/_hw3_q1_compressed.pt"
torch.save(trainer.model.state_dict(), PATH)
# plot test accuracy
fig, ax = plt.subplots(figsize=(20,10), facecolor="white")
ax.plot(trainer.testAccuracy)
plt.title("Test set accuracy - SVD")
plt.show()
```
| github_jupyter |
# Clustering Sprint Challenge
Objectives:
* Describe two clustering algorithms
* Create k clusters with the k-Means algorithm
* Compare/contrast the performance of the two algorithms on two datasets
### 1. Describe two different clustering algorithms
There are many clustering algorithms with profoundly different implementations. Their objective is the same - to identify groups in unlabeled data.
Fill out the below python objects.
```
# Clustering algorithm 1:
algorithm_one_name = "K Means"
algorithm_one_description = "K centroids are initialized, randomly or through sampling \
\nThen loop through the following 2 steps: \
\n1. Each point is assigned to the nearest centroid \
\n2. New centroids are calculated by taking the means of the assigned points \
\nClusters found minimize within-cluster sum of squares, or 'inertia' \
\nWorks best when clusters are convex and isotropic\n"
# Clustering algorithm 2:
algorithm_two_name = "Spectral Clustering"
algorithm_two_description = "An affinity matrix is first computed \
\nIt contains some sort of pairwise distance/similarity measure \
\nThe matrix is then factored through eigendecomposition \
\nThe eigenvectors corresponding to the lowest nonzero eigenvalues are then selected \
\nTogether, they make up a lower dimensional feature space \
\nThe data is projected onto the lower dimension, and K Means is performed \
\nOther standard clustering algorithms are also acceptable \
\nUseful when clusters are non-convex"
print(algorithm_one_name)
print(algorithm_one_description)
print(algorithm_two_name)
print(algorithm_two_description)
```
### 2. Create k clusters with k-Means algorithm
```
# Import libraries
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.cluster import KMeans, SpectralClustering
# Dataset
set1 = pd.read_csv('https://www.dropbox.com/s/zakq7e0r8n1tob9/clustering_set1.csv?raw=1', index_col=0)
set1.head()
plt.scatter(set1['x'], set1['y']);
```
There appear to be 2 clusters.
```
# Create kmeans object
model = KMeans(n_clusters=2)
# Fit kmeans object to data
model.fit(set1.as_matrix())
# Print location of clusters learned by kmeans object
centroids = model.cluster_centers_
print('Cluster Centroids:\n' + str(centroids))
plt.scatter(set1['x'], set1['y'])
plt.plot(centroids[:,0], centroids[:,1], 'ro');
```
### 3. Compare/contrast the performance of your two algorithms with two datasets
```
# Second dataset
set2 = pd.read_csv('https://www.dropbox.com/s/zakq7e0r8n1tob9/clustering_set2.csv?raw=1', index_col=0)
set2.head()
plt.scatter(set2['x'], set2['y']);
```
The data seems to be the same as in part 1.
The clusters are mostly convex, meaning that given two points in the cluster, the points on the line connecting them are likely to also be in the cluster. They are also isotropic (the same in any direction), since they cover about 8 units of distance in both the x and y directions, and appear circular.
Because of this, I expect K means to perform well. Spectral clustering should also perform well, but wont be too useful, especially given that the clusters are linearly separable in the first place. In fact, because it discards information during the projection onto a lower dimension, it may even perform worse.
```
n_clusters=2
model1 = KMeans(n_clusters)
model2 = SpectralClustering(n_clusters)
model1.fit(set2.as_matrix())
model2.fit(set2.as_matrix())
plt.scatter(set2['x'], set2['y'], c=model1.labels_, cmap='coolwarm')
plt.title('K Means Clustering');
plt.scatter(set2['x'], set2['y'], c=model2.labels_, cmap='coolwarm')
plt.title('Spectral Clustering');
```
Interestingly, Spectral Clustering labeled some of the outlying points as part of the wrong cluster. This may have something to do with the information lost when projecting onto a lower dimension. Aside from this, both algorithms performed similarly, as expected.
| github_jupyter |
## MAGIC DL3 spectrum with gammapy
author:
* Cosimo Nigro (cosimonigro2@gmail.com)
```
#setup
%matplotlib inline
import matplotlib.pyplot as plt
#plt.style.use('ggplot')
# check package versions
import numpy as np
import astropy
import regions
import sherpa
import os
print('numpy:', np.__version__)
print('astropy', astropy.__version__)
print('regions', regions.__version__)
print('sherpa', sherpa.__version__)
import gammapy
gammapy.version.version
# units and coordinates from astropy
import astropy.units as u
from astropy.io import fits
from astropy.coordinates import SkyCoord, Angle
from astropy.table import Table
# regions
from regions import CircleSkyRegion
# Datastore is the interface to the index of observations
import gammapy.data as data # to many functions to import
# The background method
import gammapy.background as background
# Some utility function to deal with energy axis (log binning with a unit etc)
from gammapy.utils.energy import EnergyBounds
# The class performing the extraction and extraction results
from gammapy.spectrum import SpectrumExtraction, SpectrumObservation, SpectrumFit, SpectrumResult
# Spectral models
from gammapy.spectrum.models import LogParabola
# Utilities for flux points plotting
from gammapy.spectrum import FluxPoints, SpectrumEnergyGroupMaker, FluxPointEstimator
# A class to deal with exclusion regions
from gammapy.image import SkyImage
from gammapy.extern.pathlib import Path
```
# defining the DataStore object
```
MAGIC_DIR = '../data/magic'
datastore = data.DataStore.from_dir(MAGIC_DIR)
obs_ids = [5029747,5029748]
datastore.info()
obs_list = datastore.obs_list(obs_ids)
```
## defining Target and Exclusion regions
The next step is to define a signal extraction region, also known as **on region**. This is realized with the CircleSkyRegion object. Since we have generated the IRFs with Hadronness and (especially) $\theta^2$ cuts constant over all the enrgy range, **set** a on region radius equal to your $\theta^2$ cut.
```
# fetch the cooordinate of the object we are wobbling around from the fits file
RA_OBJ = 83.63
DEC_OBJ = 22.01
target_position = SkyCoord(ra=RA_OBJ, dec=DEC_OBJ, unit='deg', frame='icrs') # coordinates of the Crab
on_region_radius = Angle('0.141 deg') # we use 0.02 theta2 cut to generate the files
on_region = CircleSkyRegion(center=target_position, radius=on_region_radius)
```
for defining the **exclusion region** we will use a predefined mask that is in **gammapy-extra**, you should download this repository because it contains supplementary material sometimes required, so:
``` git clone https://github.com/gammapy/gammapy-extra.git```
``` export GAMMAPY_EXTRA="/home/wherever/you/have/downloaded/it/"```
```
EXCLUSION_FILE = '$GAMMAPY_EXTRA/datasets/exclusion_masks/tevcat_exclusion.fits'
allsky_mask = SkyImage.read(EXCLUSION_FILE)
exclusion_mask = allsky_mask.cutout(
position=on_region.center,
size=Angle('6 deg'),
)
```
## Estimate background
We will manually perform a background estimate by placing reflected regions around the pointing position and looking at the source statistics. This will result in a gammapy.background.BackgroundEstimate that serves as input for other classes in gammapy.
```
background_estimator = background.ReflectedRegionsBackgroundEstimator(
obs_list=obs_list,
on_region=on_region,
exclusion_mask = exclusion_mask)
background_estimator.run()
print(background_estimator.result[0])
# we plot it!
plt.figure(figsize=(6,6))
background_estimator.plot()
```
## Source statistic
Next we're going to look at the overall source statistics in our signal region. For more info about what debug plots you can create check out the **ObservationSummary** class.
```
stats = []
for obs, bkg in zip(obs_list, background_estimator.result):
stats.append(data.ObservationStats.from_obs(obs, bkg))
obs_summary = data.ObservationSummary(stats)
fig = plt.figure(figsize=(10,6))
ax1=fig.add_subplot(121)
obs_summary.plot_excess_vs_livetime(ax=ax1)
ax2=fig.add_subplot(122)
obs_summary.plot_significance_vs_livetime(ax=ax2)
```
## Spectrum extraction
Here we will perform the spectrum extraction and obtain the MAGIC Crab spectrum in FITS file.
We use the same binning in E-reco e_true tahat was used for exporting the IRFs
```
MAGIC_reco_bins_edges = [5.54354939e-03, 7.53565929e-03, 1.02436466e-02, 1.39247665e-02,
1.89287203e-02, 2.57308770e-02, 3.49774320e-02, 4.75467958e-02,
6.46330407e-02, 8.78593369e-02, 1.19432151e-01, 1.62350858e-01,
2.20692676e-01, 3.00000000e-01, 4.07806917e-01, 5.54354939e-01,
7.53565929e-01, 1.02436466e+00, 1.39247665e+00, 1.89287203e+00,
2.57308770e+00, 3.49774320e+00, 4.75467958e+00, 6.46330407e+00,
8.78593369e+00, 1.19432151e+01, 1.62350858e+01, 2.20692676e+01,
3.00000000e+01, 4.07806917e+01, 5.54354939e+01] * u.TeV
MAGIC_true_bins_edges = [5.54354939e-03, 8.59536081e-03, 1.33272426e-02, 2.06640999e-02,
3.20400130e-02, 4.96785457e-02, 7.70273690e-02, 1.19432151e-01,
1.85181435e-01, 2.87126737e-01, 4.45194537e-01, 6.90281154e-01,
1.07029182e+00, 1.65950436e+00, 2.57308770e+00, 3.98961308e+00,
6.18595803e+00, 9.59142554e+00, 1.48716566e+01, 2.30587383e+01,
3.57529376e+01, 7.91911869e+01] * u.TeV
extraction = SpectrumExtraction(obs_list=obs_list,
bkg_estimate=background_estimator.result,
containment_correction=False,
e_reco = MAGIC_reco_bins_edges,
e_true = MAGIC_true_bins_edges
)
extraction.run()
# gammapy diagnostic plots
extraction.observations[0].peek()
```
## fit spectrum
We will now fit the spectrum with a log parabola
```
model = LogParabola(amplitude = 3.80*1e-11 * u.Unit('cm-2 s-1 TeV-1'),
reference = 1 * u.Unit('TeV'),
alpha = 2.47 * u.Unit(''),
beta = 0.24 * u.Unit(''))
fit = SpectrumFit(obs_list = extraction.observations, model = model)
fit.fit()
fit.est_errors()
fit_result = fit.result
ax0, ax1 = fit_result[0].plot(figsize=(8,8))
ax0.set_ylim(0, 20)
print(fit_result[0])
```
## Compute flux points
Here we compute the flux points after the fit, we have to define a stack of observation (through ```observations.stack()```) in order to repeat the fit in each energy band delimited by the bin.
```
ebounds = MAGIC_true_bins_edges[6:-4]
stacked_obs = extraction.observations.stack()
seg = SpectrumEnergyGroupMaker(obs=stacked_obs)
#seg.compute_range_safe()
seg.compute_groups_fixed(ebounds=ebounds)
print(seg.groups)
fpe = FluxPointEstimator(
obs=stacked_obs,
groups=seg.groups,
model=fit_result[0].model,
)
fpe.compute_points()
print(fpe.flux_points.table)
print('\n\n E_true binning \n\n e_max [TeV] e_min [TeV]')
for i in range(len(ebounds)-1):
print(ebounds[i+1].value, ebounds[i].value)
# Have to set flux unit here because flux points default unit is ph / ... which raises a unit conversion error
fpe.flux_points.plot(flux_unit = 'cm-2 s-1 TeV-1')
from gammapy.spectrum import CrabSpectrum
crab_hess_ecpl = CrabSpectrum('hess_ecpl')
crab_magic_lp = CrabSpectrum('magic_lp')
spectrum_result = SpectrumResult(
points=fpe.flux_points,
model=fit_result[0].model,
)
print (spectrum_result.flux_point_residuals)
ax0, ax1 = spectrum_result.plot(
energy_range=fit.result[0].fit_range,
energy_power=2, flux_unit='TeV-1 cm-2 s-1',
fig_kwargs=dict(figsize = (8,8)),
point_kwargs=dict(color='navy', marker='o', linewidth = 1.3, capsize = 3., label = 'MAGIC_DL3 + gammapy')
)
# plot HESS and MAGIC reference spectrum
crab_hess_ecpl.model.plot([0.4,50] * u.TeV, ax=ax0, energy_power=2, flux_unit='TeV-1 cm-2 s-1',
ls ='-.', lw=2, color='dodgerblue', label = 'HESS A&A 457 (2006)')
crab_magic_lp.model.plot([0.05,30] * u.TeV, ax=ax0, energy_power=2, flux_unit='TeV-1 cm-2 s-1',
ls ='-.', lw=2, color='darkorange', label = 'MAGIC JHEAP 5-6 (2015)')
ax0.yaxis.grid(True, linewidth = 0.4)
ax0.xaxis.grid(True, linewidth = 0.4)
ax1.yaxis.grid(True, linewidth = 0.4)
ax1.xaxis.grid(True, linewidth = 0.4)
ax0.legend(loc = 0, numpoints = 1, prop={'size':10})
ax0.set_ylabel(r'E$^2$ d$\phi$/dE [TeV cm$^{-2}$ s$^{-1}]$', size=13.)
ax0.set_xlim(0.01, 100)
ax0.set_ylim(1e-12, 1e-9)
plt.savefig('DL3_gammapy_final_comparison_same_binning.png')
```
| github_jupyter |
**[Pandas Home Page](https://www.kaggle.com/learn/pandas)**
---
# Introduction
The first step in most data analytics projects is reading the data file. In this exercise, you'll create Series and DataFrame objects, both by hand and by reading data files.
Run the code cell below to load libraries you will need (including code to check your answers).
```
import pandas as pd
pd.set_option('max_rows', 5)
from learntools.core import binder; binder.bind(globals())
from learntools.pandas.creating_reading_and_writing import *
print("Setup complete.")
```
# Exercises
## 1.
In the cell below, create a DataFrame `fruits` that looks like this:

```
# Your code goes here. Create a dataframe matching the above diagram and assign it to the variable fruits.
fruits = pd.DataFrame({'Apples': [30], 'Bananas': [21]})
# Check your answer
q1.check()
fruits
#q1.hint()
#q1.solution()
```
## 2.
Create a dataframe `fruit_sales` that matches the diagram below:

```
# Your code goes here. Create a dataframe matching the above diagram and assign it to the variable fruit_sales.
fruit_sales = pd.DataFrame({'Apples': [35, 41], 'Bananas': [21, 34]}, index = ['2017 Sales', '2018 Sales'])
# Check your answer
q2.check()
fruit_sales
#q2.hint()
#q2.solution()
```
## 3.
Create a variable `ingredients` with a Series that looks like:
```
Flour 4 cups
Milk 1 cup
Eggs 2 large
Spam 1 can
Name: Dinner, dtype: object
```
```
ingredients = pd.Series(['4 cups', '1 cup', '2 large', '1 can'], index = ['Flour', 'Milk', 'Eggs', 'Spam'], name = "Dinner")
# Check your answer
q3.check()
ingredients
#q3.hint()
#q3.solution()
```
## 4.
Read the following csv dataset of wine reviews into a DataFrame called `reviews`:

The filepath to the csv file is `../input/wine-reviews/winemag-data_first150k.csv`. The first few lines look like:
```
,country,description,designation,points,price,province,region_1,region_2,variety,winery
0,US,"This tremendous 100% varietal wine[...]",Martha's Vineyard,96,235.0,California,Napa Valley,Napa,Cabernet Sauvignon,Heitz
1,Spain,"Ripe aromas of fig, blackberry and[...]",Carodorum Selección Especial Reserva,96,110.0,Northern Spain,Toro,,Tinta de Toro,Bodega Carmen Rodríguez
```
```
reviews = pd.read_csv("../input/wine-reviews/winemag-data_first150k.csv", index_col=0)
# Check your answer
q4.check()
reviews
#q4.hint()
#q4.solution()
```
## 5.
Run the cell below to create and display a DataFrame called `animals`:
```
animals = pd.DataFrame({'Cows': [12, 20], 'Goats': [22, 19]}, index=['Year 1', 'Year 2'])
animals
```
In the cell below, write code to save this DataFrame to disk as a csv file with the name `cows_and_goats.csv`.
```
# Your code goes here
animals.to_csv('cows_and_goats.csv')
# Check your answer
q5.check()
#q5.hint()
#q5.solution()
```
# Keep going
Move on to learn about **[indexing, selecting and assigning](https://www.kaggle.com/residentmario/indexing-selecting-assigning)**.
---
**[Pandas Home Page](https://www.kaggle.com/learn/pandas)**
*Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum) to chat with other Learners.*
| github_jupyter |
```
import os
from os.path import abspath, join
from pathlib import Path
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import plotly.express as px
from PIL import Image
from skimage import io
from sklearn.preprocessing import MinMaxScaler
import cv2
from tqdm import tqdm
from skimage import measure
# Read in raw data file
# Read in mask
raw = io.imread(abspath("../data/N-2016-2156-1_hBN220215_11038-38401_component_data.tif"))
# Read in tiff
mask = io.imread(abspath("../data/N-2016-2156-1_hBN220215_11038-38401_mask.tif"))
# Read in post-MCMicro data
data = pd.read_csv(abspath("../data/mcmicro_quants/mesmer-N-2016-2156-1_hBN220215_11038-38401_cell.csv"), index_col="CellID")
raw.shape
print(raw.shape)
raw.shape[2]*0.5*0.1
def get_scale_bar_fos(im_dim, scale, um):
return um/(im_dim[2]* scale)
scale_bar_fos = get_scale_bar_fos(raw.shape, 0.5, 50)
scale_bar_fos
# Define a min max scaler function so the final image doesn't look ugly
def min_max_scaler(im):
return ((im-im.min())/(im.max()-im.min()))
# Select one channel and min max scale it
DAPI = raw[0]
DAPI = min_max_scaler(DAPI)
NCAM1 = raw[3]
NCAM1 = min_max_scaler(NCAM1)
SOX10 = raw[5]
SOX10 = min_max_scaler(SOX10)
CD68 = raw[2]
CD68 = min_max_scaler(CD68)
def grayscale2rgbcolor(im, color=(1, 1, 1)):
# Convert gray scale image to RGB
r = np.multiply(im, color[0], casting="unsafe")
r = np.where(r>1, 1, r)
g = np.multiply(im, color[1], casting="unsafe")
g = np.where(g>1, 1, g)
b = np.multiply(im, color[2], casting="unsafe")
b = np.where(b>1, 1, b)
# Stack channels
bgr = np.stack([b, g, r], axis=-1)
return np.uint8(255*bgr)
# Get channels colored
DAPI_im = grayscale2rgbcolor(DAPI, color = (0.0, 0.2, 1.5)) # Blue
NCAM1_im = grayscale2rgbcolor(NCAM1, color = (1.5, 1.5, 0.3)) # Yellow
SOX10_im = grayscale2rgbcolor(SOX10, color = (2.0, 0.1, 0.1)) # red
CD68_im = grayscale2rgbcolor(CD68, color = (0.0, 2, 0.2)) # green
# Blend channels together
DAPI_CD68 = cv2.addWeighted(DAPI_im, 1, CD68_im, 1, 0.0)
NCAM1_SOX10 = cv2.addWeighted(SOX10_im, 1, NCAM1_im, 1, 0.0)
DAPI_NCAM1_SOX10 = cv2.addWeighted(DAPI_im, 1, NCAM1_SOX10, 1, 0.0)
# Get contours for each cell in the mask
# Calculate and draw contours over masked channel
# # Get range of cells
min = mask[np.nonzero(mask)].min()
max = mask[np.nonzero(mask)].max()
print(max-min)
contours = {}
for i in tqdm(range(min, max+1)):
try:
y, x = measure.find_contours(mask==i, 0.8)[0].T
contours[i] = (y, x)
#ax.plot(x, y, linewidth=3, alpha=0.5, color=color[i])
except IndexError:
pass
```
## DAPI
```
# IN THE CASE OF DAPI WE DO THE SAME PROCESS BUT NO CUTOFF WAS APPLIED
marker = "DAPI"
cutoff = 0
color = data[marker].apply(lambda x: "cyan" if x >= cutoff else "red")
%%time
fig, ax = plt.subplots(1, 1, figsize=(10, 10))
ax.matshow(DAPI_im[:,:,::-1])
for c, (y, x) in contours.items():
ax.plot(x, y, linewidth=0.5, alpha=0.6, color=color[c], rasterized=True)
plt.tight_layout()
ax.axhline(y=50, xmin=0.03, xmax=0.03+scale_bar_fos, linewidth=4, color="w",alpha=0.8, linestyle="-")
ax.axes.get_xaxis().set_ticks([])
ax.axes.get_yaxis().set_ticks([])
plt.savefig(abspath("../out/overlay-DAPI.pdf"), dpi=600)
plt.savefig(abspath("../out/overlay-DAPI.png"), dpi=600)
plt.close()
```
## CD68
```
# Get flags
marker = "CD68"
cutoff = 1.756449425220487
color = data[marker].apply(lambda x: "cyan" if x >= cutoff else "red")
%%time
fig, ax = plt.subplots(1, 1, figsize=(10, 10))
ax.matshow(DAPI_CD68[:,:,::-1])
for c, (y, x) in contours.items():
ax.plot(x, y, linewidth=0.5, alpha=0.8, color=color[c], rasterized=True)
ax.axhline(y=50, xmin=0.03, xmax=0.03+scale_bar_fos, linewidth=4, color="w",alpha=0.8, linestyle="-")
plt.tight_layout()
ax.axes.get_xaxis().set_ticks([])
ax.axes.get_yaxis().set_ticks([])
plt.savefig(abspath("../out/overlay-CD68.pdf"), dpi=600)
plt.savefig(abspath("../out/overlay-CD68.png"), dpi=600)
plt.close()
```
## NCAM1 SOX10
```
# Get flags
marker = "NCAM1"
cutoff = 1
flag_ncam1 = data[marker]>=cutoff
marker = "SOX10"
cutoff = 1
flag_sox10 = data[marker]>=cutoff
double_positives = flag_ncam1 & flag_sox10
color = double_positives.apply(lambda x: "cyan" if x else "red")
%%time
fig, ax = plt.subplots(1, 1, figsize=(10, 10))
ax.matshow(DAPI_NCAM1_SOX10[:,:,::-1])
for c, (y, x) in contours.items():
ax.plot(x, y, linewidth=0.5, alpha=0.8, color=color[c], rasterized=True)
ax.axhline(y=50, xmin=0.03, xmax=0.03+scale_bar_fos, linewidth=4, color="w",alpha=0.8, linestyle="-")
plt.tight_layout()
ax.axes.get_xaxis().set_ticks([])
ax.axes.get_yaxis().set_ticks([])
plt.savefig(abspath("../out/overlay-NCAM1SOX10.pdf"), dpi=600)
plt.savefig(abspath("../out/overlay-NCAM1SOX10.png"), dpi=600)
plt.close()
```
## Using a different image for CD138
```
# Read in mask
raw = io.imread(abspath("../data/N-2020-845_Restain_hBN220216_5129-53809_component_data.tif"))
# Read in tiff
mask = io.imread(abspath("../data/N-2020-845_Restain_hBN220216_5129-53809_mask.tif"))
# Read in post-MCMicro data
data = pd.read_csv(abspath("../data/mcmicro_quants/mesmer-N-2020-845_Restain_hBN220216_5129-53809_cell.csv"), index_col="CellID")
plt.matshow(raw[0])
scale_bar_fos = get_scale_bar_fos(raw.shape, 0.5, 100)
scale_bar_fos
DAPI = raw[0]
DAPI = min_max_scaler(DAPI)
CD138 = raw[4]
CD138 = min_max_scaler(CD138)
DAPI_im = grayscale2rgbcolor(DAPI, color = (0.0, 0.2, 1.5)) # Blue
CD138_im = grayscale2rgbcolor(CD138, color = (0.0, 2, 2)) # Cyan
DAPI_CD138 = cv2.addWeighted(DAPI_im, 1, CD138_im, 1, 0.0)
plt.matshow(DAPI_CD138[:,:,::-1])
def slice_image(im, mask , sy=(0, -1), sx=(0, -1)):
# Subsetting image
mask = mask[sy[0]:sy[1], sx[0]:sx[1]]
im = im[sy[0]:sy[1], sx[0]:sx[1]]
return im, mask
im, mk = slice_image(DAPI_CD138, mask,sy=(250,500), sx=(1250,1500))
print(im.shape)
print(mk.shape)
# zoom rate
scale_bar_fos = get_scale_bar_fos(im.shape, 0.5, 50)
scale_bar_fos = scale_bar_fos/100
scale_bar_fos
# # Get range of cells
min = mk[np.nonzero(mk)].min()
max = mk[np.nonzero(mk)].max()
print(max-min)
contours = {}
for i in tqdm(range(min, max+1)):
try:
y, x = measure.find_contours(mk==i, 0.8)[0].T
contours[i] = (y, x)
#ax.plot(x, y, linewidth=3, alpha=0.5, color=color[i])
except IndexError:
pass
# Get flags
marker = "CD138"
cutoff = 1.6312074160575833
color = data[marker].apply(lambda x: "cyan" if x >= cutoff else "red")
%%time
fig, ax = plt.subplots(1, 1, figsize=(10, 10))
ax.matshow(im[:, :, ::-1])
for c, (y, x) in contours.items():
ax.plot(x, y, linewidth=0.5, alpha=0.8, color=color[c], rasterized=True)
ax.axhline(y=5, xmin=0.03, xmax=0.03+scale_bar_fos, linewidth=4, color="w",alpha=0.8, linestyle="-")
plt.tight_layout()
ax.axes.get_xaxis().set_ticks([])
ax.axes.get_yaxis().set_ticks([])
plt.savefig(abspath("../out/overlay-CD138.pdf"), dpi=600)
plt.savefig(abspath("../out/overlay-CD138.png"), dpi=600)
plt.close()
# Get flags
marker = "DAPI"
cutoff = 0
color = data[marker].apply(lambda x: "cyan" if x >= cutoff else "red")
%%time
fig, ax = plt.subplots(1, 1, figsize=(10, 10))
ax.matshow(im[:, :, ::-1])
for c, (y, x) in contours.items():
ax.plot(x, y, linewidth=0.5, alpha=0.8, color=color[c], rasterized=True)
ax.axhline(y=5, xmin=0.03, xmax=0.03+scale_bar_fos, linewidth=4, color="w",alpha=0.8, linestyle="-")
plt.tight_layout()
ax.axes.get_xaxis().set_ticks([])
ax.axes.get_yaxis().set_ticks([])
plt.savefig(abspath("../out/overlay-DAPI2.pdf"), dpi=600)
plt.savefig(abspath("../out/overlay-DAPI2.png"), dpi=600)
plt.close()
```
| github_jupyter |
## Installing dependencies
```
!pip install simpletransformers datasets tqdm pandas
```
##Loading data from huggingface(optional)
```
import pandas as pd
from datasets import load_dataset
from tqdm import tqdm
dataset = load_dataset('tapaco', 'en')
def process_tapaco_dataset(dataset, out_file):
tapaco = []
# The dataset has only train split.
for data in tqdm(dataset["train"]):
keys = data.keys()
tapaco.append([data[key] for key in keys])
tapaco_df = pd.DataFrame(
data=tapaco,
columns=[
"language",
"lists",
"paraphrase",
"paraphrase_set_id",
"sentence_id",
"tags",
],
)
tapaco_df.to_csv(out_file, sep="\t", index=None)
return tapaco_df
tapaco_df = process_tapaco_dataset(dataset,"tapaco_huggingface.csv")
tapaco_df.head()
```
## Preprocessing TaPaCo for training(optional)
```
import pandas as pd
from tqdm import tqdm
tapaco_df = pd.read_csv("tapaco_huggingface.csv",sep="\t")
def generate_tapaco_paraphrase_dataset(dataset, out_file):
dataset_df = dataset[["paraphrase", "paraphrase_set_id"]]
non_single_labels = (
dataset_df["paraphrase_set_id"]
.value_counts()[dataset_df["paraphrase_set_id"].value_counts() > 1]
.index.tolist()
)
tapaco_df_sorted = dataset_df.loc[
dataset_df["paraphrase_set_id"].isin(non_single_labels)
]
tapaco_paraphrases_dataset = []
for paraphrase_set_id in tqdm(tapaco_df_sorted["paraphrase_set_id"].unique()):
id_wise_paraphrases = tapaco_df_sorted[
tapaco_df_sorted["paraphrase_set_id"] == paraphrase_set_id
]
len_id_wise_paraphrases = (
id_wise_paraphrases.shape[0]
if id_wise_paraphrases.shape[0] % 2 == 0
else id_wise_paraphrases.shape[0] - 1
)
for ix in range(0, len_id_wise_paraphrases, 2):
current_phrase = id_wise_paraphrases.iloc[ix][0]
for count_ix in range(ix + 1, ix + 2):
next_phrase = id_wise_paraphrases.iloc[ix + 1][0]
tapaco_paraphrases_dataset.append([current_phrase, next_phrase])
tapaco_paraphrases_dataset_df = pd.DataFrame(
tapaco_paraphrases_dataset, columns=["Text", "Paraphrase"]
)
tapaco_paraphrases_dataset_df.to_csv(out_file, sep="\t", index=None)
return tapaco_paraphrases_dataset_df
dataset_df = generate_tapaco_paraphrase_dataset(tapaco_df,"tapaco_paraphrases_dataset.csv")
dataset_df.head()
```
## Load already preprocessed version of TaPaCo
```
!wget https://github.com/hetpandya/paraphrase-datasets-pretrained-models/raw/main/datasets/tapaco/tapaco_paraphrases_dataset.csv
import pandas as pd
dataset_df = pd.read_csv("tapaco_paraphrases_dataset.csv",sep="\t")
```
##Model Training
```
from simpletransformers.t5 import T5Model
from sklearn.model_selection import train_test_split
import sklearn
dataset_df.columns = ["input_text","target_text"]
dataset_df["prefix"] = "paraphrase"
train_data,test_data = train_test_split(dataset_df,test_size=0.1)
train_data
test_data
args = {
"reprocess_input_data": True,
"overwrite_output_dir": True,
"max_seq_length": 256,
"num_train_epochs": 4,
"num_beams": None,
"do_sample": True,
"top_k": 50,
"top_p": 0.95,
"use_multiprocessing": False,
"save_steps": -1,
"save_eval_checkpoints": True,
"evaluate_during_training": False,
'adam_epsilon': 1e-08,
'eval_batch_size': 6,
'fp_16': False,
'gradient_accumulation_steps': 16,
'learning_rate': 0.0003,
'max_grad_norm': 1.0,
'n_gpu': 1,
'seed': 42,
'train_batch_size': 6,
'warmup_steps': 0,
'weight_decay': 0.0
}
model = T5Model("t5","t5-small", args=args)
model.train_model(train_data, eval_data=test_data, use_cuda=True,acc=sklearn.metrics.accuracy_score)
```
##Loading Trained Model & Prediction Using Trained Model
```
from simpletransformers.t5 import T5Model
from pprint import pprint
import os
root_dir = os.getcwd()
trained_model_path = os.path.join(root_dir,"outputs")
args = {
"overwrite_output_dir": True,
"max_seq_length": 256,
"max_length": 50,
"top_k": 50,
"top_p": 0.95,
"num_return_sequences": 5,
}
trained_model = T5Model("t5",trained_model_path,args=args)
prefix = "paraphrase"
pred = trained_model.predict([f"{prefix}: The house will be cleaned by me every Saturday."])
pprint(pred)
```
| github_jupyter |
```
%matplotlib inline
```
Autograd: Automatic Differentiation
===================================
Central to all neural networks in PyTorch is the ``autograd`` package.
Let’s first briefly visit this, and we will then go to training our
first neural network.
The ``autograd`` package provides automatic differentiation for all operations
on Tensors. It is a define-by-run framework, which means that your backprop is
defined by how your code is run, and that every single iteration can be
different.
Let us see this in more simple terms with some examples.
Tensor
--------
``torch.Tensor`` is the central class of the package. If you set its attribute
``.requires_grad`` as ``True``, it starts to track all operations on it. When
you finish your computation you can call ``.backward()`` and have all the
gradients computed automatically. The gradient for this tensor will be
accumulated into ``.grad`` attribute.
To stop a tensor from tracking history, you can call ``.detach()`` to detach
it from the computation history, and to prevent future computation from being
tracked.
To prevent tracking history (and using memory), you can also wrap the code block
in ``with torch.no_grad():``. This can be particularly helpful when evaluating a
model because the model may have trainable parameters with `requires_grad=True`,
but for which we don't need the gradients.
There’s one more class which is very important for autograd
implementation - a ``Function``.
``Tensor`` and ``Function`` are interconnected and build up an acyclic
graph, that encodes a complete history of computation. Each tensor has
a ``.grad_fn`` attribute that references a ``Function`` that has created
the ``Tensor`` (except for Tensors created by the user - their
``grad_fn is None``).
If you want to compute the derivatives, you can call ``.backward()`` on
a ``Tensor``. If ``Tensor`` is a scalar (i.e. it holds a one element
data), you don’t need to specify any arguments to ``backward()``,
however if it has more elements, you need to specify a ``gradient``
argument that is a tensor of matching shape.
```
import torch
torch.cuda.device_count()
```
Create a tensor and set requires_grad=True to track computation with it
```
x = torch.ones(2, 2, requires_grad=True)
print(x)
```
Do an operation of tensor:
```
y = x + 2
print(y)
```
``y`` was created as a result of an operation, so it has a ``grad_fn``.
```
print(y.grad_fn)
y.shape
```
Do more operations on y
```
z = y * y * 3
out = z.mean()
print(z)
print('\n----\n')
print(out)
```
``.requires_grad_( ... )`` changes an existing Tensor's ``requires_grad``
flag in-place. The input flag defaults to ``False`` if not given.
```
a = torch.randn(2, 2)
a = ((a * 3) / (a - 1))
print(a.requires_grad)
a.requires_grad_(True)
print(a.requires_grad)
b = (a * a).sum()
print(b.grad_fn)
```
Gradients
---------
Let's backprop now
Because ``out`` contains a single scalar, ``out.backward()`` is
equivalent to ``out.backward(torch.tensor(1))``.
```
out.backward()
```
print gradients d(out)/dx
```
print(x.grad)
```
You should have got a matrix of ``4.5``. Let’s call the ``out``
*Tensor* “$o$”.
We have that $o = \frac{1}{4}\sum_i z_i$,
$z_i = 3(x_i+2)^2$ and $z_i\bigr\rvert_{x_i=1} = 27$.
Therefore,
$\frac{\partial o}{\partial x_i} = \frac{3}{2}(x_i+2)$, hence
$\frac{\partial o}{\partial x_i}\bigr\rvert_{x_i=1} = \frac{9}{2} = 4.5$.
You can do many crazy things with autograd!
```
x = torch.randn(3, requires_grad=True)
y = x * 2
while y.data.norm() < 1000:
y = y * 2
print(y)
gradients = torch.tensor([0.1, 1.0, 0.0001], dtype=torch.float)
y.backward(gradients)
print(x.grad)
```
You can also stop autograd from tracking history on Tensors
with ``.requires_grad=True`` by wrapping the code block in
``with torch.no_grad()``:
```
print(x.requires_grad)
print((x ** 2).requires_grad)
with torch.no_grad():
print((x ** 2).requires_grad)
```
**Read Later:**
Documentation of ``autograd`` and ``Function`` is at
http://pytorch.org/docs/autograd
| github_jupyter |
```
import numpy as np
import h5py
import matplotlib.pyplot as plt
# Helper function to help read the h5 files.
def simple_read_data(fileName):
print(fileName)
hf = h5py.File('{}.h5'.format(fileName), 'r')
# We'll return a dictionary object.
results = {}
results['rs_glob_acc'] = np.array(hf.get('rs_glob_acc')[:])
results['rs_train_acc'] = np.array(hf.get('rs_train_acc')[:])
results['rs_train_loss'] = np.array(hf.get('rs_train_loss')[:])
# 3D array: Read as [number of times, number of epochs, number of users].
results['perUserAccs'] = np.array(hf.get('perUserAccs'))
return results
# Define the global directory path.
directoryPath = '/home/adgdri/pFedMe/results/'
```
### Datasplit-1
```
fileNames = [
# TODO: Enter the filenames of the experimental results over the different runs.
]
# Get the number of users.
numUsers = int(fileNames[0].split('u')[0].split('_')[-1])
avgPersAcc = []
perUserAcc = np.zeros((1, numUsers))
for fileName in fileNames:
ob = simple_read_data(directoryPath + fileName)
avgPersAcc.append( ob['rs_glob_acc'][-1] )
# Take the per user accuracy from the last epoch.
perUserAcc += ob['perUserAccs'][:,-1, :]
# Average out over the different runs.
perUserAcc /= len(fileNames)
print ('----------------------------------------')
print ('\n Average accuracies across all the users over different runs : %s' % avgPersAcc)
print ('\n Average accuracy across all the different runs : %f.' % np.mean(avgPersAcc) )
print ('\n Average per user accuracy averaged over different runs: \n %s.' % np.round(perUserAcc.T, 4))
print ('\n Per-user averaged accuracy: \n %f.' % np.mean(np.round(perUserAcc.T, 4)))
```
### Datasplit-2
```
fileNames = [
# TODO: Enter the filenames of the experimental results over the different runs.
]
# Get the number of users.
numUsers = int(fileNames[0].split('u')[0].split('_')[-1])
avgPersAcc = []
perUserAcc = np.zeros((1, numUsers))
for fileName in fileNames:
ob = simple_read_data(directoryPath + fileName)
avgPersAcc.append( ob['rs_glob_acc'][-1] )
# Take the per user accuracy from the last epoch.
perUserAcc += ob['perUserAccs'][:,-1, :]
# Average out over the different runs.
perUserAcc /= len(fileNames)
print ('----------------------------------------')
print ('\n Average accuracies across all the users over different runs : %s' % avgPersAcc)
print ('\n Average accuracy across all the different runs : %f.' % np.mean(avgPersAcc) )
print ('\n Average per user accuracy averaged over different runs: \n %s.' % np.round(perUserAcc.T, 4))
print ('\n Per-user averaged accuracy: \n %f.' % np.mean(np.round(perUserAcc.T, 4)))
```
### Datasplit-3
```
fileNames = [
# TODO: Enter the filenames of the experimental results over the different runs.
]
# Get the number of users.
numUsers = int(fileNames[0].split('u')[0].split('_')[-1])
avgPersAcc = []
perUserAcc = np.zeros((1, numUsers))
for fileName in fileNames:
ob = simple_read_data(directoryPath + fileName)
avgPersAcc.append( ob['rs_glob_acc'][-1] )
# Take the per user accuracy from the last epoch.
perUserAcc += ob['perUserAccs'][:,-1, :]
# Average out over the different runs.
perUserAcc /= len(fileNames)
print ('----------------------------------------')
print ('\n Average accuracies across all the users over different runs : %s' % avgPersAcc)
print ('\n Average accuracy across all the different runs : %f.' % np.mean(avgPersAcc) )
print ('\n Average per user accuracy averaged over different runs: \n %s.' % np.round(perUserAcc.T, 4))
print ('\n Per-user averaged accuracy: \n %f.' % np.mean(np.round(perUserAcc.T, 4)))
```
| github_jupyter |
# Abstract
I am acting as an NBA consultant; based on the 3-year performances of players before becoming Free Agents and the contract they ended up signing (calculated per year), I want to predict how this year's free agents will do (the 2018-19 season is only 1-2 games from being over). Teams targeting certain free agents in the Summer of '19 will be able to use this to determine who they want to target as their main pursuit.
# Obtain the Data
First I will scrape data from basketball-reference.com that has player's individual statitstics per season from 2008-2009 to 2017-18 seasons. This will contain data with various player statistics that I can use as my features in the model. Also from basketball-reference.com I will collect rookie lists to make sure I don't have outliers since 3-year performances are being combined.
I will also scrape free agent lists from 2011 to 2018 seasons that will help me filter out non-impending free agents. This is available on spotrac.com.
Finally, I will collect salary information for players from 2008-2009 to 2017-2018 from HoopsHype.com. This data will help me collect the target variable information.
```
# %%writefile ../src/data/make_dataset.py
# imports
import pandas as pd
import time
from datetime import datetime
import os
import pickle
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.common.action_chains import ActionChains
from selenium.webdriver.common.by import By
from selenium.common.exceptions import NoSuchElementException
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as EC
from selenium.webdriver.support.select import Select
def check_exists(driver,classname):
try:
driver.find_element_by_class_name(classname)
except NoSuchElementException:
return False
return True
def initialize_selenium(URL):
# initialize selenium
chromedriver = "/Applications/chromedriver"
os.environ["webdriver.chrome.driver"] = chromedriver
driver = webdriver.Chrome(chromedriver)
driver.get(URL)
return driver
# Generate dictionary to store our data per year
def data_to_dict(years):
"""
Generate Dictionary that will store our data per year in this format:
Key (Year): Value (Data)
years: int indicating how many years of data will be stored
"""
data = {}
CURRENT_YEAR = int(datetime.now().year)
years_label = range(CURRENT_YEAR-1,CURRENT_YEAR-years,-1)
return years_label, data
def download_salary_data(URL,years):
years_label, data = data_to_dict(years)
driver = initialize_selenium(URL)
print(list(years_label))
for i in years_label:
time.sleep(2)
df = pd.read_html(driver.current_url)[0]
data[i]=df
years = driver.find_element_by_class_name("salaries-team-selector-top")
years.click()
year = driver.find_element_by_link_text(str(i-1)+"/"+str(i-2000).zfill(2))
year.click()
driver.quit()
return data
def download_rookie_data(URL, years):
years_label, data = data_to_dict(years)
driver = initialize_selenium(URL)
wait = WebDriverWait(driver, 10)
for i in years_label:
df = pd.read_html(driver.current_url)[0]
df.columns=df.columns.droplevel()
df = df[['Player']]
data[i]=df
prev_year = driver.find_element_by_css_selector("a.button2.prev")
prev_year.click()
time.sleep(10)
driver.quit()
return data
def download_player_data(URL, years, type_data):
years_label, data = data_to_dict(years)
driver = initialize_selenium(URL)
wait = WebDriverWait(driver, 10)
# get to the current season stats, this may have changed
tab = driver.find_elements_by_id("header_leagues")
hover = ActionChains(driver).move_to_element(tab[0])
hover.perform()
wait.until(EC.visibility_of_element_located((By.LINK_TEXT, type_data))).click()
for i in years_label:
prev_year = driver.find_element_by_css_selector("a.button2.prev")
prev_year.click()
time.sleep(10)
df = pd.read_html(driver.current_url)[0]
df = df[df.Rk != 'Rk']
data[i]=df
driver.quit()
return data
def download_fa_data(URL):
data={}
driver = initialize_selenium(URL)
for i in range(2018,2010,-1):
years = Select(driver.find_element_by_name('year'))
years.select_by_visible_text(str(i))
submit = driver.find_element_by_class_name('go')
submit.click()
time.sleep(10)
df = pd.read_html(driver.current_url)[0]
data[i]=df
driver.quit()
return data
def save_dataset(data,filename):
with open(filename, 'wb') as w:
pickle.dump(data,w)
def run():
"""
Executes a set of helper functions that download data from one or more sources
and saves those datasets to the data/raw directory.
"""
data_fa = download_fa_data("https://www.spotrac.com/nba/free-agents/")
data_reg = download_player_data("https://www.basketball-reference.com", 12, "Per G")
data_adv = download_player_data("https://www.basketball-reference.com", 12, "Advanced")
data_salary = download_salary_data("https://hoopshype.com/salaries/players/", 12)
data_rookie = download_rookie_data("https://www.basketball-reference.com/leagues/NBA_2018_rookies.html", 12)
save_dataset(data_fa, "data/raw/freeagents2.pickle")
save_dataset(data_reg, "data/raw/regstats2.pickle")
save_dataset(data_adv, "data/raw/advstats2.pickle")
save_dataset(data_salary, "data/raw/salaries2.pickle")
save_dataset(data_rookie, "data/raw/rookies2.pickle")
# run()
```
# Scrub the Data
## Salary List Data
Here are the thing's that were fixed going through the Salary List Data:
1. Fix some headers (0 -> Rank, 1-> Name, 2-> Team, 3-> Salary)
2. Remove rows with those header labels, as they were repeated in the website tables
3. Add year column for when the lists are aggregated into a single dataframe
Then the lists were aggregated into a single salary dataframe. Afterwards, these were fixed:
1. Change Salary format (remove $ and commas)
2. Split position from the name into a new column
3. Change Salary datatype to int
4. Remove Rk column; it's not significant
## Player Stats Data
### Before Aggregation of Regular and Advanced Stats Year-wise
Some things I notice:
- Both: There are players who were traded mid-season that have appeared as rows of both teams, and a total season. I want to keep the cumulative total row (Tm assigned to TOT) and get rid of partial team stats.
- Both: It will be useful to again add a Year column for after I aggregate each lists into a single dataframe. This can be done after this step
After taking care of these, I combined regular stats and advanced stats for the same year into a single dataframe. Now the dataframes are by year (with both regular and advanced stats)
### Before Aggregation of Yearly stats into a single dataframe
Now we have combined stats for each type of stats. Some basic cleaning can be done before all years are combined into one giant dataframe:
1. Some columns can be eliminated (Two "unnamed" arbitrary empty columns were on the website when I scraped. Rk_x and Rk_y were arbitrary rankings done by alphabetic order and is insiginficant as well)
2. We can remove one of the MP (Minutes Played) columns; there was a conflict during dataframe merge because regular stats data compiled minutes played as per game average, whereas the Advance stats data compiled minutes played as season total. I will remove MP_y.
3. Add the year of the player stat here in a column called 'Year'.
After adding them all together, I changed some datatypes for better processing.
- Player, Position, Tm, Year -> unchanged
- Age, G, GS -> int
- Everything else -> floats
## Rookies list and using it to remove from player stats
Rookies have no previous year's stats (because they were in college or overseas), so we cannot use their data. Therefore we must identify the rookies for each year and remove them from the stats list.
There are some weird "Player" that shows up. Also some null values got picked up when it was scraped. We can remove those rows ("Player" is part of the table header that got repeated on Basketball Reference). We can also add a Year column that will be useful to identify which year the rookies belong (just like our other lists). After that we can concatenate the dataframes into a single one, and merge with the stats to remove them from the stats dataframe.
## Adding FA information
Now we want to filter out the data for all the FA's on our list, but before we do that, we must look at one thing: the way naming is done differently between the salaries list (from ESPN.com) and the stats list (from Basketball-Reference.com). I noticed two weird things:
1. Suffixes are missing (notably Jr.) in Basketball-Reference list
2. Also players that go by initials (i.e. J.J. Redick in Basketball-Reference) are missing periods in the other list (i.e. JJ Redick in ESPN).
### Some column conflicts happened on the merge:
- Tm: I will use Tm_x as that was from the salary data. It is the team that paid the player on the season following (2018 salary is for the 2018-19 season, while 2018 stats is for the 2017-18 season)
- Pos: I will also use Pos_x although position ambiguity is really on the data collector's hands; some players can be either guard position, or either forward position, or some could be SG/SF. There is no real definition on positions now, as NBA is becoming more positionless and a guard is able to do what forwards used to, and vice versa. Even some centers handle the ball like a guard!
## Missing Data after merging all the dataframes:
Here are the possibilities based on some research:
1. Some did not have stats because they were out of the NBA (not playing basketball entirely or overseas). These players should also be removed from the considerations.
2. Some players are missing a couple stats only (way to treat those datapoints will be explored in the next section).
After taking care of all of these, I pickle'd the dataframe and move onto the next section.
```
# %%writefile ../src/features/build_features.py
# imports
import re
import os
import pickle
import pandas as pd
import numpy as np
from functools import reduce
# Remove rookies from stats
def remove_rookies(stats, rookies):
COLS = ['Player','Year']
no_rookies = stats.merge(rookies, indicator=True, how='outer')
no_rookies = no_rookies[no_rookies['_merge'] == 'left_only']
del no_rookies['_merge']
save_dataset(no_rookies,"../data/interim/norookiestats.pickle")
return no_rookies
# Merge stats with salaries
def salary_merge(salaries, no_rookies):
salaries['Player'] = salaries['Player'].map(lambda x: x.replace(' Jr.',""))
no_rookies['Player'] = no_rookies['Player'].map(lambda x: x.replace('.',""))
data_all = pd.merge(salaries,no_rookies, on=['Player','Year'], how='left')
# Remove unnecessary columns that happened on merge conflict
# data_all.rename(columns={'Tm_x': 'Tm','Pos_x':'Pos','MP_x':'MP'}, inplace=True)
# Drop players that have too many missing stat information
playerinfo =['Player','Tm','Salary','Year','Pos']
rest = data_all.columns.difference(playerinfo)
played = data_all.dropna(thresh=20)
return played
# Merge the FA list into the stats list
def FA_merge(played, freeagents):
FA_check = played.merge(freeagents, indicator=True, how='left')
played["FA"] = FA_check["_merge"]
played["FA"] = played["FA"].str.replace("left_only",'No').replace("both","Yes")
played = played[~played['FA'].isnull()]
# I chose to fill Null values with 0
played = played.fillna(0)
return played
# Accumulate stats of 3 past seasons and update the list with it
def accumulate_stats(played,stats):
totallist=['MP', 'FG', 'FGA','3P', '3PA', '2P',
'2PA', 'FT', 'FTA', 'ORB', 'DRB',
'TRB', 'AST', 'STL', 'BLK', 'TOV', 'PF', 'PTS']
for i in range(played.shape[0]):
curr = played.iloc[i].Player
curryear = played.iloc[i].Year
years = [curryear, curryear-1, curryear-2]
threeyrs = stats[(stats.Player == curr) & (stats.Year.isin(years))]
if threeyrs.shape[0] > 1:
print("Update row "+str(i))
for stat in totallist:
played.iloc[i, played.columns.get_loc(stat)] = (reduce((lambda x, y: x + y),
[k*v for k,v in zip(threeyrs[stat],threeyrs.G)])) #/ threeyrs.G.sum()
played.iloc[i, played.columns.get_loc('G')] = threeyrs.G.sum()#/len(threeyrs)
played.iloc[i, played.columns.get_loc('GS')] = threeyrs.GS.sum()#/len(threeyrs)
return played
# Clean salaries data
def clean_salaries_dataset(path, filename):
money = pickle.load(open(path+"/"+filename, "rb"))
combined={}
for k,v in money.items():
calendar = str(k)+"/"+str(k-1999).zfill(2)
print(calendar)
temp = v[['Player',calendar]]
temp["Year"] = k
temp = temp.rename(columns={calendar:"Salary"})
combined[k]=temp
salaries = reduce(lambda x,y:pd.concat([x,y]),[v for k,v in combined.items()])
salaries["Salary"] = salaries["Salary"].str.replace('$','').str.replace(',','')
salaries.Salary = salaries.Salary.astype(int)
return salaries
# Clean stats (regular and advanced) data
def clean_stats_dataset(path, filename1, filename2):
stats = pickle.load(open(path+"/"+filename2, "rb"))
advs = pickle.load(open(path+"/"+filename1, "rb"))
for i,j in stats.items():
temp = j
temp['total'] = (temp['Tm'] == 'TOT')
temp = temp.sort_values('total', ascending=False).drop_duplicates(['Player','Age']).drop('total', 1)
stats[i]=temp
for i,j in advs.items():
temp = j
temp['total'] = (temp['Tm'] == 'TOT')
temp = temp.sort_values('total', ascending=False).drop_duplicates(['Player','Age']).drop('total', 1)
advs[i]=temp
combined={}
for (a1,b1),(a2,b2) in zip(stats.items(),advs.items()):
df = b1.merge(b2, how="inner",on=["Player","Age","Pos","Tm","G"])#,"MP"])
combined[a1]=df.sort_values("Player")
print("Stats Row for "+str(a1)+": "+str(b1.shape[0])
+", Adv Row for "+str(a2)+": "+str(b2.shape[0])+", After combined: "+str(df.shape[0]))
for k,v in combined.items():
v=v.drop(['Rk_x','Unnamed: 19','Unnamed: 24', 'Rk_y','MP_y'], axis=1)
v['Year'] = k
combined[k]=v
combined_stats = reduce(lambda x,y:pd.concat([x,y]),[v for k,v in combined.items() if k != 2019 or k != 2008])
combined_stats = combined_stats.reset_index(drop=True);
unchanged = ['Player','Pos','Tm','Year']
intlist = ['Age','G','GS']
floatlist= combined_stats.columns.difference(unchanged+intlist)
combined_stats[intlist] = combined_stats[intlist].astype(int)
combined_stats[floatlist] = combined_stats[floatlist].astype(float)
combined_stats.rename(columns={'MP_x':'MP'}, inplace=True)
return combined_stats
# Clean rookies data
def clean_rookies_dataset(path, filename):
rookies = pickle.load(open(path+"/"+filename, "rb"))
combined_rookies = pd.DataFrame()
for v,k in rookies.items():
temp = rookies[v][rookies[v].Player != 'Player']
temp = temp[~(temp.Player.isnull())]
temp['Year']=v
combined_rookies = pd.concat([combined_rookies,temp])
return combined_rookies
# Clean FA data
def clean_fa_dataset(path, filename):
freeagents = pickle.load(open(path+"/"+filename, "rb"))
FAS={}
for k,v in freeagents.items():
v.columns=[re.sub(r"Player.+","Player",col) for col in v.columns]
v.columns=[re.sub(r"\d+ Cap Hit","Cap Hit",col) for col in v.columns]
v["Year"] = k
FAS[k]=v
freeagents = reduce(lambda x,y:pd.concat([x,y]),[v for k,v in FAS.items() if k != 2019])
freeagents = freeagents[['Player','Year']]
return freeagents
# Build overall dataset
def build_dataset(salaries, stats, rookies, freeagents):
no_rookies = remove_rookies(stats, rookies)
played = salary_merge(salaries, no_rookies)
players = FA_merge(played, freeagents)
return accumulate_stats(players,stats)
# dump file to pickle
def save_features(data,filename):
with open(filename,"wb") as writer:
pickle.dump(data,writer)
def run():
"""
Executes a set of helper functions that read files from data/raw, cleans them,
and converts the data into a design matrix that is ready for modeling.
"""
salaries = clean_salaries_dataset('data/raw', "salaries2.pickle")
stats = clean_stats_dataset('data/raw', "advstats2.pickle", "regstats2.pickle")
rookies = clean_rookies_dataset('data/raw','rookies2.pickle')
freeagents = clean_fa_dataset('data/raw','freeagents2.pickle')
save_dataset(salaries, "data/interim/salaries2.pickle")
save_dataset(stats, "data/interim/stats2.pickle")
save_dataset(rookies, "data/interim/rookies2.pickle")
save_dataset(freeagents, "data/interim/fa2.pickle")
full_data = build_dataset(salaries, stats, rookies, freeagents)
save_features(full_data,'data/processed/data2.pickle')
run()
```
*Before moving on to exploratory analysis, write down some notes about challenges encountered while working with this data that might be helpful for anyone else (including yourself) who may work through this later on.*
# Explore the Data
## What am I looking for:
Now that we have our data, here are some of the things I looked for:
1. Eliminate some features that I know are collinear with the others (by basketball stats definition)
2. Look at correlations of the features with the target variable (using Correlation Matrix, statsmodel, and determine which ones are to be looked at more than others.
3.
### Eliminating some features:
Before we proceed, we can eliminate some of the columns as they can be defined in terms of the other.
$FG = 2P+3P$
$TRB = ORB+DRB$
$TRB\% = ORB\% + DRB\%$
$WS = OWS+DWS$
$BPM = OBPM + DBPM$
$WS/48 = \frac{WS}{48 min}$
We can eliminate at least 6 columns to make things easier.
### Look at correlations:
I took a look at correlation matrix from the remaining features as well as statsmodel to see which features were stronger in correlation with the target than others.
### Look at relationships
I also took a look at the features relationship with the target variable via pairplots to notice any strong relationships.
```
# %%writefile ../src/visualization/visualize.py
import os
import pickle
import pandas as pd
import numpy as np
import re
import seaborn as sns
import matplotlib.pyplot as plt
import scipy.stats as stats
import statsmodels.api as sm
import statsmodels.formula.api as smf
import patsy
def load_features(filename):
return pickle.load(open(filename, "rb"))
# Looks at the OLS stats on statsmodel and outputs them to txt file
def ols_stats(data,features,directory,num):
y, X= patsy.dmatrices(features, data, return_type = "dataframe")
model = sm.OLS(y,X)
fit = model.fit()
fit.summary()
text_file = open(directory+"OLS_report_"+str(num)+".txt", "w")
text_file.write(fit.summary().as_text())
text_file.close()
# plots correlation heatmap
def corr_map(data,features,directory):
plt.figure(figsize=(15,10))
sns.heatmap((data[features]).corr()
, xticklabels=['SALARY','G','GS','MP','OBPM','FGA','_3PA','_2PA','OWS','DWS','ORBPCT'
,'VORP','USGPCT','DBPM','STLPCT','BLKPCT']
, yticklabels=['SALARY','G','GS','MP','OBPM','FGA','_3PA','_2PA','OWS','DWS','ORBPCT',
'VORP','USGPCT','DBPM','STLPCT','BLKPCT'])
plt.xticks(np.arange(0.5, 16, step=1), rotation=30);
plt.title("Correlation of Selected Player Statistics", size=30);
plt.savefig(directory+"heatmap.png", dpi=400);
def eliminate_stats(data,directory):
columns=['Salary', 'Year', 'Pos', 'Age', 'G', 'GS', 'MP','FGA', 'FG%', '3P', '3PA', '3P%', '2P',
'2PA', '2P%', 'eFG%', 'FT','FTA', 'FT%', 'ORB', 'DRB', 'AST', 'STL', 'BLK', 'TOV', 'PF',
'PTS', 'PER', 'TS%', '3PAr', 'FTr', 'ORB%', 'DRB%', 'AST%','STL%', 'BLK%', 'TOV%', 'USG%', 'OWS',
'DWS', 'OBPM','DBPM', 'VORP', 'FA']
df_sm2 = data[columns]
df_sm2.columns = df_sm2.columns.str.replace("%","PCT").str.upper()
df_sm2.columns = df_sm2.columns.str.replace("2","_2").str.replace("3","_3")
del df_sm2["POS"]
del df_sm2["FA"]
# Some stats to look at on Statsmodel
ols_stats(df_sm2,
"""SALARY ~ G + GS + MP + OBPM + FTA + FT + FGA + _3P+_3PA+_2P + _2PA + ORB+ AST + TOV + STL + DRB +
PTS + PER + OWS + DWS + VORP + USGPCT + FGPCT + _3PPCT + _2PPCT + EFGPCT + FTPCT + BLK + PF + TSPCT
+_3PAR + FTR + ORBPCT + DRBPCT + ASTPCT + STLPCT + BLKPCT + TOVPCT + DBPM""",directory,1)
ols_stats(df_sm2, """SALARY ~ G + GS + MP + OBPM + FGA + _3PA + _2PA + OWS + DWS + VORP + USGPCT +
ORBPCT + STLPCT + BLKPCT + DBPM""",directory,2)
return data, columns
# plots
def generate_charts(data, columns, directory):
plt.hist(data['Salary']);
plt.savefig(directory+"salary_hist.png");
plt.hist(np.cbrt(data['Salary']));
plt.savefig(directory+"salary_hist_cbrt.png");
#Corr heatmap
corr_map(data,['Salary','G','GS','MP','OBPM','FGA','3PA','2PA','OWS','DWS','ORB%',
'VORP','USG%','DBPM','STL%','BLK%'],directory)
for feature in columns:
pp = sns.pairplot(data=data,y_vars=['Salary'],x_vars=[feature]);
pp.savefig(directory+feature+"_correlation.png");
def run():
"""
Executes a set of helper functions that read files from data/processed,
calculates descriptive statistics for the population, and plots charts
that visualize interesting relationships between features.
"""
data = load_features('../data/processed/data2.pickle')
data, columns = eliminate_stats(data, '../reports/')
generate_charts(data, columns, '../reports/figures/')
run()
```
*What did you learn? What relationships do you think will be most helpful as you build your model?*
# Model the Data
## Linear Regression
First I ran linear regression on the 13 features that I selected from looking at the statistics in the exploration part. After doing so, I tried improving the R2 score while not reducing the adjusted R2 for adding more features. This was done by adding square terms and interaction terms between the features.
Then I went back and looked at some of the features I took out that I thought were useful in adding back. I also tracked to make sure the adjusted R2 did not drop (while R2 goes up). Doing it and then adding additional square and interaction terms, I was able to come up with what I thought was the best model.
## Linear Regression vs. Ridge Regression
With the best model I came up with by linear regression, I decided to implement a Linear vs. Ridge Regression (with scaling). I thought Ridge regularization was better than Lasso because I had already removed needless features and therefore Lasso wouldn't really help with improving the model (Lasso will kill off features that won't help with the model while Ridge will smooth things out.)
This was also the stage in which I implemented Cross-validation. Like in Train-Test split, where I had to group the 2018 stats together in the test set, when doing the CV-train split, I made sure to keep a year's worth of stat as the validation set, and 3 years previous stats data as the training set. This limited how random the CV-train split could be but it also prevented leakage of future knowledge in affecting the model training.
Overall, the Ridge regression showed better in terms of cross validation error mean.
## Minimizing the error by adjusting alpha value
The last step was then to minimize the error by adjusting the alpha value of the ridge regression. I ran cross validation training on alpha values ranging from 0.01 to 100 and found 8.21 to be the optimized alpha value. With that I trained on the whole training set and tested on the test set to come to my final prediction and error.
```
## %%writefile ../src/models/train_model.py
# imports
import pickle
import pandas as pd
import numpy as np
import itertools
from sklearn.linear_model import LinearRegression, Ridge, Lasso
from sklearn.model_selection import train_test_split, KFold,cross_val_score
from sklearn.metrics import r2_score, mean_absolute_error, mean_squared_error
from sklearn.preprocessing import PolynomialFeatures, StandardScaler
def load_data(filename):
return pickle.load(open(filename, "rb"))
def save_data(data, filename):
with open(filename,"wb") as writer:
pickle.dump(data,writer)
def column_ops(data,columns):
data2 = data[columns]
data2.columns = data2.columns.str.replace("%","PCT").str.upper()
data2.columns = data2.columns.str.replace("2","_2").str.replace("3","_3")
return data2
def add_squares(data,features):
for fea in features:
data[fea+"2"] = data[fea]**2
return data
def interactions(data,features):
for item in itertools.combinations(features,2):
data[item[0]+item[1]] = data[item[0]]*data[item[1]]
return data
def other_terms(data,original):
data["_2PA"] = original["2PA"]
data["_2PA2"] = original["2PA"]**2
data["FGA"] = original["FGA"]
data["FGA2"] = data["FGA"]**2
data["DWS"] = original["DWS"]
data["DWS2"] = data["DWS"]**2
data["BLK"] = original["BLK"]
data["FT"] = original["FT"]
return data
def lr_test(dataset, target, itr):
test = dataset[(dataset.YEAR == 2018)]
train = dataset[dataset.YEAR != 2018]
test = test.drop('YEAR', axis=1)
train = train.drop('YEAR', axis=1)
X_test, y_test = pd.get_dummies(test[test.columns[1:]]), test[target]
X_train, y_train = pd.get_dummies(train[train.columns[1:]]), np.cbrt(train[target])
# Linear Regression
lr = LinearRegression()
lr.fit(X_train, y_train)
r2 = lr.score(X_train, y_train)
N = X_train.shape[0]
p = X_train.shape[1]
adjusted_r2 = 1-((1-r2**2)*(N-1)/(N-p-1))
print(f'Linear Regression R2 score: {r2:.6f}')
print(f'Adjusted R2 Score: {adjusted_r2: .6f}')
# Predictions
y_pred = lr.predict(X_test)
print("\nLR Test Data R2 score: ", r2_score(y_test,y_pred**3))
print("LR Test MAE: ", mean_absolute_error(y_test,y_pred**3))
print("Median 2018 NBA Salary: ", y_test.median())
print("Mean 2018 NBA Salary: ", y_test.mean())
print("Median predicted Salary: ", np.median(y_pred**3))
plt.figure(figsize=(10,10))
sns.regplot(y_pred**3, y_test)
ax = plt.gca()
locs, labels = plt.xticks()
ax.set_xticklabels(['0','0','10','20','30','40','50','60','70','80'])
ax.set_yticklabels(['0','0','20', '40', '60', '80'])
plt.title("2018 NBA Salary Prediction", size=30)
plt.xlabel("Prediction Salary (Million $)", size=20)
plt.ylabel("Actual Salary (Million $)", size=20)
plt.savefig("../reports/figures/LR_pred_"+str(itr)+".png")
save_data(lr,"../models/lr"+str(itr)+".pickle")
return y_pred, lr
def linear_vs_ridge(dataset):
# Split out the y (keep the Year on both for test/train split)
X,y = dataset.drop('SALARY',axis=1), dataset[['YEAR','SALARY']]
# Test/Train split
X,X_test = X[X.YEAR != 2018],X[X.YEAR==2018]
y,y_test = y[y.YEAR != 2018],y[y.YEAR==2018]
# Drop the Year column on Test data first
y_test = y_test.drop('YEAR',axis=1)
X_test = X_test.drop('YEAR',axis=1)
LR_r2s, LM_REG_r2s = [],[]
LR_MAES, LM_REG_MAES = [],[]
# CV Split
for i in range(2017,2013,-1):
years = [i-j for j in range(1,4)]
X_val,y_val = X[X.YEAR == i], y[y.YEAR == i]
X_val = X_val.drop('YEAR', axis=1)
y_val = y_val.drop('YEAR', axis=1)
X_train, y_train = X[X.YEAR.isin(years)], y[y.YEAR.isin(years)]
X_train = X_train.drop('YEAR', axis=1)
y_train = y_train.drop('YEAR', axis=1)
# Set dummies
X_val,X_test,X_train = pd.get_dummies(X_val), pd.get_dummies(X_test), pd.get_dummies(X_train)
#simple linear regression
lm = LinearRegression()
lm_reg = Ridge(alpha=1)
lm.fit(X_train, y_train)
LR_r2s.append(lm.score(X_val, y_val))
y_pred = lm.predict(X_val)
LR_MAES.append(mean_absolute_error(y_pred,y_val))
#ridge with feature scaling
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_val_scaled = scaler.transform(X_val)
lm_reg.fit(X_train_scaled, y_train)
LM_REG_r2s.append(lm_reg.score(X_val_scaled, y_val))
y_pred_reg = lm_reg.predict(X_val_scaled)
LM_REG_MAES.append(mean_absolute_error(y_pred_reg,y_val))
mean_MAE_LR = np.mean(LR_MAES)
mean_MAE_REG = np.mean(LM_REG_MAES)
print("Average R2 Score of Linear Regression: ", np.mean(LR_r2s))
print("Average MAE of Linear Regression: ", mean_MAE_LR)
print("Average R2 Score of Ridge Regression w/ Scaler: ", np.mean(LM_REG_r2s))
print("Average MAE of Ridge Regression: ", mean_MAE_REG)
# Test on the whole training set
y = y.drop('YEAR',axis=1)
X = X.drop('YEAR',axis=1)
X = pd.get_dummies(X)
X_te = scaler.transform(X)
if mean_MAE_REG < mean_MAE_LR:
print("\nRidge Regression performs better")
pred = lm_reg.predict(X_te)
else:
print("\nLinear Regression performs better")
pred = lm.predict(X)
print("MAE on the whole training set: ", mean_absolute_error(pred,y))
X_test_te = scaler.transform(X_test)
pred_test = lm_reg.predict(X_test_te)
pred_ln = lm.predict(X_test)
print("MAE on the test set: ", mean_absolute_error(pred_test,y_test))
return lm_reg, lm, pred_test, y_test
def ridge_alpha_test(dataset):
# Split out the y (keep the Year on both for test/train split)
X,y = dataset.drop('SALARY',axis=1), dataset[['YEAR','SALARY']]
# Test/Train split
X,X_test = X[X.YEAR != 2018],X[X.YEAR==2018]
y,y_test = y[y.YEAR != 2018],y[y.YEAR==2018]
# Drop the Year column on Test data first
y_test = y_test.drop('YEAR',axis=1)
X_test = X_test.drop('YEAR',axis=1)
alphalist = 10**(np.linspace(-2,2,200))
err_vec_val = np.zeros(len(alphalist))
for k,curr_alpha in enumerate(alphalist):
MAES = []
for i in range(2017,2013,-1):
# Set CV/Train set split
years = [i-j for j in range(1,4)]
X_val, y_val = X[X.YEAR == i], y[y.YEAR == i]
X_val = X_val.drop('YEAR', axis=1)
y_val = y_val.drop('YEAR', axis=1)
X_train, y_train = X[X.YEAR.isin(years)], y[y.YEAR.isin(years)]
X_train = X_train.drop('YEAR', axis=1)
y_train = np.cbrt(y_train.drop('YEAR', axis=1))
# Set dummies
X_val,X_test,X_train = pd.get_dummies(X_val), pd.get_dummies(X_test), pd.get_dummies(X_train)
values = []
# note the use of a new sklearn utility: Pipeline to pack
# multiple modeling steps into one fitting process
steps = [('standardize', StandardScaler()),
('ridge', Ridge(alpha = curr_alpha))]
pipe = Pipeline(steps)
pipe.fit(X_train.values, y_train)
val_set_pred = pipe.predict(X_val.values)
MAES.append(mean_absolute_error(y_val, val_set_pred**3))
err_vec_val[k] = np.mean(MAES)
min_mae = err_vec_val.min()
min_alpha = alphalist[np.argmin(err_vec_val)]
print("Minimum MAE is: ", min_mae)
print("When alpha is: ", min_alpha)
# Retrain on full training set
X = X.drop('YEAR', axis=1)
y = y.drop('YEAR', axis=1)
steps_train = [('standardize', StandardScaler()),
('ridge', Ridge(alpha = min_alpha))]
pipe_train = Pipeline(steps)
pipe_train.fit(X.values, np.cbrt(y))
test_pred = pipe_train.predict(X_test.values)
test_mae = mean_absolute_error(y_test, test_pred**3)
print("Test MAE: ", test_mae)
return pipe_train, test_pred, y_test
def run():
"""
Executes a set of helper functions that read files from data/processed,
calculates descriptive statistics for the population, and plots charts
that visualize interesting relationships between features.
"""
original = ['Salary','Year','G','GS','MP','3P','ORB','AST','TOV','DRB','PTS','VORP','USG%','STL%','DBPM', 'TOV%']
features = ['G','GS', 'MP','_3P','ORB','AST','TOV','DRB','PTS','VORP','USGPCT','STLPCT','DBPM', 'TOVPCT']
new = ['_2PA','FGA','DWS','BLK','FT']
data = load_data('../data/processed/data2.pickle')
data2 = column_ops(data,original)
y_pred,lr = lr_test(data2,"SALARY",1)
data3 = add_squares(data2,features)
y_pred, lr2 = lr_test(data3,"SALARY",2)
data4 = interactions(data3,features)
y_pred, lr3 = lr_test(data4,"SALARY",3)
data5 = other_terms(data4,data)
y_pred, lr4 = lr_test(data5,"SALARY",4)
data6 = interactions(data5,new)
y_pred, lr5 = lr_test(data6,"SALARY",5)
data7 = interactions2(data6,features,new)
y_pred, lr6 = lr_test(data7,"SALARY",6)
lm_reg_cv, lm_cv, pred_test, y_test = linear_vs_ridge(data7)
lm_reg_cv, test, y_test = ridge_alpha_test(data7)
y_test["PRED"] = test**3
y_test["Player"] = data[data.Year==2018]["Player"]
y_test.columns
y_test["DIFF"] = (y_test.SALARY- y_test.PRED)
plt.figure(figsize=(10,10))
sns.regplot("PRED","SALARY", data= y_test);
ax = plt.gca()
ax.set_xticklabels(['0','0','5','10','15','20','25']);
ax.set_yticklabels(['0','0','10', '20', '30', '40']);
ax.tick_params(labelsize=15)
plt.title("2018 NBA Salary Prediction", size=30);
plt.xlabel("Prediction Salary (Million $)", size=20);
plt.ylabel("Actual Salary (Million $)", size=20, fontfamily='sans-serif');
plt.savefig("../reports/figures/prediction2.png", dpi=400)
plt.figure(figsize=(10,10))
sns.scatterplot("PRED","DIFF", data= y_test);
ax = plt.gca()
ax.set_xticklabels(['0','0','5','10','15','20','25']);
ax.set_yticklabels(['0','-15','-10', '-5', '0', '5', '10', '15', '20','25']);
ax.tick_params(labelsize=15)
plt.title("2018 NBA Salary Prediction Error", size=30);
plt.xlabel("Prediction Salary (Million $)", size=20);
plt.ylabel("Pred. Error (Million $)", size=20);
plt.savefig("../reports/figures/residual.png", dpi=400)
```
_Write down any thoughts you may have about working with these algorithms on this data. What other ideas do you want to try out as you iterate on this pipeline?_
# Interpret the Model
The Model produced a MAE of 4.4 million \\$, which seems to be bad considering the average salary is 10 million \\$. However throughout modeling this I've realized that there could be many other possible features I can scrape and add to make this model a much better product.
They are:
- Social effects: Popularity of a player determined by twitter mentions, or instagram followers
- Personal effects: player's love for a particular city, their family information, injury history
- Financial effects: NBA contract structure (team, player-wise)
| github_jupyter |
# Evaluation Metric Testing
The point of this notebook is to walk through an evaluation metric taken from one of the kernels posted on [Kaggle](https://www.kaggle.com/wcukierski/example-metric-implementation) to ensure that was it was functioning correctly and gain a deeper undertanding of the [IoU](https://www.kaggle.com/c/data-science-bowl-2018#evaluation) metric.
```
!pwd
import skimage
import importlib
import numpy as np
from scipy import stats
from sklearn.metrics import confusion_matrix
import matplotlib.pyplot as plt
import matplotlib as mpl
import pandas as pd
%matplotlib inline
from utils import imaging
from utils import evaluate
importlib.reload(imaging)
importlib.reload(evaluate)
```
# Notes on method below
When we are calculating the the intersection of objects between images we use a 2D histogram of the two images. This takes the two images flattened and compares the pixel values at each location. It reutrns an $\mathbf{n}$ $\times$ $\mathbf{m}$ matrix where $\mathbf{n}$ is the number of true objects and $\mathbf{m}$ is the number of predicted objects. The values of this matrix are counts of the paired pixel values between the two images. So if row 1, column 2 = 50, this means a pixel value of 1 in the true image was given a value of 2 in the predicted image.
$\textbf{Note:}$ It doesn't matter if the pixel value of the predicted mask is different than the ground truth mask (unless it is 0). All we care about is that a predict object has pixels that overlaps with a ground truth object.
## Evaluate a single image
Evaluate a single image to check the result of our evaluation metric is reasonable.
```
image_id = '0a7d30b252359a10fd298b638b90cb9ada3acced4e0c0e5a3692013f432ee4e9'
gt_path = imaging.get_path('output_train_1_lab_gt')
seg_path = imaging.get_path('output_train_1_lab_seg')
gt_image_1 = skimage.io.imread(gt_path + image_id + '.png' )
seg_image_1 = skimage.io.imread(seg_path + image_id + '.png' )
f, axarr = plt.subplots(1,2,figsize=(15,15))
axarr[0].imshow(gt_image_1)
axarr[0].set_title('Ground Truth')
axarr[1].imshow(seg_image_1)
axarr[1].set_title('Segmented')
```
# Evaluate test cases
We examine how the evaluation metric peforms in a few scenarios.
1. Perfect overlap with nonmatching class labels.
2. Not predicting one the ground truth objects (False Negative)
3. How a 50% overlap performs with a threshold of .5
4. Two predicted objects that lay over the ground truth object.
```
n = 1000 # matrices will be nxn
gt = np.zeros((n,n))
gt[300:700,300:700] = 1
gt[800:850,800:850] = 2
t1 = np.zeros((n,n))
t2 = np.zeros((n,n))
t3 = np.zeros((n,n))
t4 = np.zeros((n,n))
# perfect prediction
t1[300:700,300:700] = 2
t1[800:850,800:850] = 1
# different labels
t2[300:700,300:700] = 20
# 50% overlap
t3[300:700,500:900] = 1
# Having to small sub regions where the 1 truth region is.
t4[300:500,300:700] = 1 # creating first small sub region
t4[500:700,300:700] = 2 # creating second small sub region
test_cases = [t1,t2,t3,t4]
f, axarr = plt.subplots(1,5,figsize=(15,15))
axarr[0].imshow(gt)
axarr[0].set_title('gt')
axarr[1].imshow(t1)
axarr[1].set_title('t1')
axarr[2].imshow(t2)
axarr[2].set_title('t2')
axarr[3].imshow(t3)
axarr[3].set_title('t3')
axarr[4].imshow(t4)
axarr[4].set_title('t4')
f.tight_layout()
```
## Test case 1
```
evaluate.evaluate_image(gt, t1)
```
As we would hope, inverting the labels has no effect on the evaluation.
## Test case 2
```
evaluate.evaluate_image(gt, t2)
```
Since there is one correctly predicted object and 1 missed sobject, so 1 TP and 1 FN, the average precision is .5 as expected.
## Test case 3
```
evaluate.evaluate_image(gt, t3)
```
The object has a 50$\%$ overlap with the ground truth object so the IoU is .3, which gives 0 TPs for all thresholds used.
## Test case 4
```
evaluate.evaluate_image(gt, t4)
```
Neither predicted object has an IoU with the ground truth object that satisifies any of the thresholds so there are 0 TPs for each threshold.
## Evaluate all images
Evaluate all images in stage 1 to test the `evaluate_images` function and see the distribution of scores.
```
scores = evaluate.evaluate_images(stage_num=1)
scores.head()
f, axarr = plt.subplots(1,2,figsize=(15,5))
axarr[0].hist(scores.score, bins=50)
axarr[0].set_title('Histogram of scores')
axarr[0].set_xlabel('score')
axarr[0].set_ylabel('# of images')
axarr[1].boxplot(scores.score, 0, 'rs', 0)
axarr[1].set_title('Box plot of scores')
axarr[1].set_xlabel('score')
f.tight_layout()
```
| github_jupyter |
# Ejercicios Programación científica en Python: Numpy
```
# Importamos las librerias que vamos a usar en este NoteBook:
import numpy as np
# Para las representaciones gráficas de los ejercicios 5 y 6.
%matplotlib inline
import matplotlib.pyplot as plt
```
**Ejer 1.** A partir del array que se muestra a continuación, generar un nuevo array que contenga la 2º y 4º fila.
`[[1, 6, 11],
[2, 7, 12],
[3, 8, 13],
[4, 9, 14],
[5, 10, 15]]`
```
mi_array = np.array([[1, 6, 11],
[2, 7, 12],
[3, 8, 13],
[4, 9, 14],
[5, 10, 15]])
nuevo_array = mi_array[[1,3], :]
print(nuevo_array)
```
**Ejer 2**. Divide cada columna del array `a` elemento a elemento de acuerdo con el contenido del array `b`:
```
a = np.arange(25).reshape(5, 5)
b = np.array([1., 5, 10, 15, 20])
print(a)
print(b)
# Suma por dimensiones: eje vertical
a_sum = a.sum(axis=0, dtype=np.float)
result = a_sum/b
print(result)
```
**Ejer 3**. Genera un array de dimensión 10 x 3 con números aleatorios en el intervalo [0, 1]. Para cada fila elije el número más cercano a 0.5. Haz uso de la indexación elegante para seleccionar los elementos.
```
np.random.seed(3) # Fijamos semilla
rand_array = np.random.rand(30).reshape(10, 3)
print(rand_array, '\r\n')
value = 0.5
idx = np.abs(rand_array - value).argmin(axis=1)
print(idx)
# Expected: [0, 0, 1, 2, 0, 2, 1, 1, 0, 1]
```
**Ejer 4**. Construye una tabla, como la que se muestra en la figura, que agrupa las tablas de multiplicar del 1 al 10, utilizando expresiones generadoras y la biblioteca NumPy. En las columnas está representado el multiplicando y en las filas el multiplicador, de modo que cada celda contiene el valor de multiplicando * multiplicador.

```
multiplicando = np.arange(1,11)
multiplicador = np.arange(1,11)
tabla = np.empty(shape=(10, 10), dtype=int)
for num in multiplicando:
mult_row = np.multiply(num, multiplicador)
print(mult_row)
np.append(tabla, [mult_row], axis=0)
print('\r\n')
print(tabla)
```
**Ejer 5**. El dataset *president_heights.csv* recoge información sobre la estatura de los presidentes de US. Sabiendo que la información que se almacena en cada columna es:
* name: nombre del presidente (2º columna)
* height(cm): altura (3º columna)
A partir de la lectura de este dataset en un array de NumPy mediante la función *genfromtxt()* ([puede consultar aquí la documentación](https://docs.scipy.org/doc/numpy/reference/generated/numpy.genfromtxt.html)), conteste a las siguientes preguntas:
* Calcule la media de las alturas
* Calcule la desviación estandar de las alturas
* Calcule la mediana
* Muestre la altura mínima y la máxima
* Muestre las alturas ordenadas de menor a mayor
* Represente en un histograma la distribución de alturas de los presidentes de US. Sugerencia: utilice la función plt.hist(), ([puede consultar aquí la documentación](https://matplotlib.org/api/pyplot_api.html)).
```
datos = np.genfromtxt('data/president_heights.csv', delimiter=',')
alturas = datos[:,2]
alturas = alturas[1:]
alturas
print("Media: ", alturas.mean())
print("Desviación típica: ", alturas.std())
print("Mediana: ", np.median(alturas))
print("Altura mínima: ", np.min(alturas))
print("Altura máxima: ", np.max(alturas))
# Alturas ordenadas de menor a mayor
sorted_alturas = np.sort(alturas)
print(sorted_alturas)
plt.hist(alturas)
plt.title("Distribución de alturas de los presidentes de EE.UU")
plt.xlabel("altura (cm)")
plt.ylabel("valor")
```
**Ejer 6**. El dataset *populations.txt* contiene información sobre la población de liebres, linces y zanahorias en el norte de Canadá durante 20 años. A partir de estos datos ([documentación de la función que se usa para la lectura del fichero](https://docs.scipy.org/doc/numpy/reference/generated/numpy.loadtxt.html)), calcula y muestra por pantalla:
* La media y la desviación estandar de cada especie.
* ¿En qué año se dió la mayor población de cada especie?
* ¿Qué especie tuvo la mayor población en cada año? (Sugerencia: utilizar argsort & indexación elegante)
* Los dos años en los que cada especie tuvo la menor población. (Sugerencia: utiizar argsort & indexación elegante)
```
datos_popul = np.loadtxt('data/populations.txt')
año, liebres, linces, zanahorias = datos_popul.T # separamos las columnas en variables
plt.axes([0.2, 0.1, 0.5, 0.8])
plt.plot(año, liebres, año, linces, año, zanahorias)
plt.legend(('Liebre', 'Lince', 'Zanahoria'), loc=(1.05, 0.5))
especies = np.array([liebres, linces, zanahorias])
for especie in especies:
print("Media: ", especie.mean())
print("Desviación típica: ", especie.std())
print(datos_popul)
print(np.max(datos_popul[:, 1]))
```
**Ejer 7**. El dataset *airquality.csv* contiene información sobre la calidad del aire de una determinada ciudad para todos los día de los meses comprendidos desde mayo hasta septiembre. Sabiendo que las variables que contiene dicho archivo son:
* Ozone: medidas de ozono (2º columna)
* Solar.R: medidas de radiación solar (3º columna)
* Wind: medidas de viento (4º columna)
* Temp: medidas de temperatura (5º columna)
* Month: meses (6º columna)
* Day: días (7º columna)
Conteste a las siguientes preguntas:
* Calcula la media de ozono durante esos meses. ¿El resultado obtenido era el esperado? ¿Encuentras alguna explicación que justifique el valor obtenido?
* Calcula la media de radiación solar durante esos meses. ¿El resultado obtenido era el esperado? ¿Encuentras alguna explicación que justifique el valor obtenido?
* Calcula la media de viento durante esos meses.
* Calcula la media de temperatura durante esos meses.
* Identifica el mes más caluroso.
* Identifica el mes y día que hizo menos viento.
| github_jupyter |
```
import os
import sys
data_dir = "/home/ec2-user/pwp-summer-2019/master_thesis_nhh_2019/processed_data/"
raw_dir = "/home/ec2-user/pwp-summer-2019/master_thesis_nhh_2019/raw_data/"
import pandas as pd
import numpy as np
import random
import math
pd.set_option('display.max_columns', 999)
```
### Function for splitting the data sets based on formation-distribution
```
# Inspired by: https://stackoverflow.com/questions/56872664/complex-dataset-split-stratifiedgroupshufflesplit
def StratifiedGroupShuffleSplit(
df_main,
train_proportion=0.6,
val_proportion = 0.3,
hparam_mse_wgt = 0.1,
df_group="title",
y_var="formation_2",
norm_keys=['gr','tvd','rdep'],
seed = 42
):
np.random.seed(seed) # Set seed
df_main.index = range(len(df_main)) # Create unique index for each observation in order to reindex
df_main = df_main.reindex(np.random.permutation(df_main.index)) # Shuffle dataset
# Create empty train, val and test datasets
df_train = pd.DataFrame()
df_val = pd.DataFrame()
df_test = pd.DataFrame()
hparam_mse_wgt = hparam_mse_wgt # Must be between 0 and 1
assert(0 <= hparam_mse_wgt <= 1)
train_proportion = train_proportion # Must be between 0 and 1
assert(0 <= train_proportion <= 1)
val_proportion = val_proportion # Must be between 0 and 1
assert(0 <= val_proportion <= 1)
test_proportion = 1-train_proportion-val_proportion # Remaining in test proportion
assert(0 <= test_proportion <= 1)
# Group the data set
subject_grouped_df_main = df_main.groupby([df_group], sort=False, as_index=False)
# Find the proportion of the total for each category
category_grouped_df_main = df_main.groupby(y_var).count()[[df_group]]/len(df_main)*100
# Functoin for calculating MSE
def calc_mse_loss(df):
# Find the proportion of the total for each category in the specific data set
grouped_df = df.groupby(y_var).count()[[df_group]]/len(df)*100
# Merge the data set above with the original proportion for each category
df_temp = category_grouped_df_main.join(grouped_df, on = y_var, how = 'left', lsuffix = '_main')
# Fill NA
df_temp.fillna(0, inplace=True)
# Square the difference
df_temp['diff'] = (df_temp[df_group+'_main'] - df_temp[df_group])**2
# Mean of the squared difference
mse_loss = np.mean(df_temp['diff'])
return mse_loss
# Initialize the train/val/test set
# First three wells are assigned to train/val/test
i = 0
for well, group in subject_grouped_df_main:
group = group.sort_index()
if (i < 3):
if (i == 0):
df_train = df_train.append(pd.DataFrame(group), ignore_index=True)
i += 1
continue
elif (i == 1):
df_val = df_val.append(pd.DataFrame(group), ignore_index=True)
i += 1
continue
else:
df_test = df_test.append(pd.DataFrame(group), ignore_index=True)
i += 1
continue
# Caluclate the difference between previous dataset and the one in the loop
mse_loss_diff_train = calc_mse_loss(df_train) - calc_mse_loss(df_train.append(pd.DataFrame(group),
ignore_index=True))
mse_loss_diff_val = calc_mse_loss(df_val) - calc_mse_loss(df_val.append(pd.DataFrame(group),
ignore_index=True))
mse_loss_diff_test = calc_mse_loss(df_test) - calc_mse_loss(df_test.append(pd.DataFrame(group),
ignore_index=True))
# Calculate the total lenght so far
total_records = df_train.title.nunique() + df_val.title.nunique() + df_test.title.nunique()
# Calculate how far much much is left before the goal is reached
len_diff_train = (train_proportion - (df_train.title.nunique()/total_records))
len_diff_val = (val_proportion - (df_val.title.nunique()/total_records))
len_diff_test = (test_proportion - (df_test.title.nunique()/total_records))
len_loss_diff_train = len_diff_train * abs(len_diff_train)
len_loss_diff_val = len_diff_val * abs(len_diff_val)
len_loss_diff_test = len_diff_test * abs(len_diff_test)
loss_train = (hparam_mse_wgt * mse_loss_diff_train) + ((1-hparam_mse_wgt) * len_loss_diff_train)
loss_val = (hparam_mse_wgt * mse_loss_diff_val) + ((1-hparam_mse_wgt) * len_loss_diff_val)
loss_test = (hparam_mse_wgt * mse_loss_diff_test) + ((1-hparam_mse_wgt) * len_loss_diff_test)
# Assign to either train, val or test
if (max(loss_train,loss_val,loss_test) == loss_train):
df_train = df_train.append(pd.DataFrame(group), ignore_index=True)
elif (max(loss_train,loss_val,loss_test) == loss_val):
df_val = df_val.append(pd.DataFrame(group), ignore_index=True)
else:
df_test = df_test.append(pd.DataFrame(group), ignore_index=True)
i += 1
return df_train, df_val, df_test
```
### Function for setting up the LSTM data set
```
# Inspired by:
# https://github.com/blasscoc/LinkedInArticles/blob/master/WellFaciesLSTM/LSTM%20Facies%20Competition.ipynb
from sklearn.preprocessing import OneHotEncoder
def chunk(x, y, num_chunks, size=61, random=True):
rng = x.shape[0] - size
if random:
indx = np.int_(
np.random.rand(num_chunks) * rng) + size//2
else:
indx = np.arange(0,rng,1) + size//2
Xwords = np.array([[x[i-size//2:i+size//2+1,:]
for i in indx]])
ylabel = np.array([y[i] for i in indx])
return Xwords[0,...], ylabel
def _num_pad(size, batch_size):
return (batch_size - np.mod(size, batch_size))
def setup_lstm_stratify(df,
df_group='title',
batch_size=128,
wvars=['gr','tvd','rdep'],
y_var = 'formation',
win=9,
n_val=39
):
df = df.fillna(0)
df_grouped = df.groupby([df_group], sort=False, as_index=False)
df_x = []
df_y = []
for key,val in df_grouped:
val = val.copy()
_x = val[wvars].values
_y = val[y_var].values
__x, __y = chunk(_x, _y, 400, size=win, random=False)
df_x.extend(__x)
df_y.extend(__y)
df_x = np.array(df_x)
df_y = np.array(df_y)
#One Hot Encoding
enc = OneHotEncoder(sparse=False, categories=[range(n_val)])
df_y = enc.fit_transform(np.atleast_2d(df_y).T)
df_x = df_x.transpose(0,2,1)
# pad to batch size
num_pad = _num_pad(df_x.shape[0], batch_size)
df_x = np.pad(df_x, ((0,num_pad),(0,0),(0,0)), mode='edge')
df_y = np.pad(df_y, ((0,num_pad), (0,0)), mode='edge')
return df_x, df_y
```
### Data generator for feeding the LSTM model
```
import numpy as np
import keras
class DataGenerator(keras.utils.Sequence):
'Generates data for Keras'
def __init__(self, df_x, df_y, batch_size=128):
'Initialization'
self.df_x = df_x
self.df_y = df_y
self.batch_size = batch_size
self.indexes = np.arange(len(self.df_x))
def __len__(self):
'Denotes the number of batches per epoch'
return int(np.floor(len(self.df_x) / self.batch_size))
def __getitem__(self, index):
'Generate one batch of data'
# Generate indexes of the batch
index_epoch = self.indexes[index*self.batch_size:(index+1)*self.batch_size]
return (self.df_x[index_epoch], self.df_y[index_epoch])
```
### Class for feature engineering and cleaning the data sets
```
class feature_engineering:
def __init__(self,df, above_below_variables, num_shifts, cols_to_remove,thresh,
log_variables, y_variable, outlier_values, var1_ratio = 'gr'
):
#Variables:
self.original_df = df
self.df = df
self.above_below_variables = above_below_variables
self.y_variable = y_variable
self.num_shifts = num_shifts
self.cols_to_remove = cols_to_remove
self.thresh = thresh
self.log_variables = log_variables
self.var1_ratio = var1_ratio
self.outlier_values = outlier_values
self.var2_ratio = log_variables
def log_values(self):
'Calculates both the log values'
for variable in self.log_variables:
self.df[variable] = np.log(self.df[variable])
#self.df = self.df.drop(self.log_variables,axis = 1)
def above_below(self):
'Add the value above and below for each column in variables'
for var in self.above_below_variables:
self.df[var+'_above'] = self.df.groupby('title')[var].shift(self.num_shifts)
self.df[var+'_below'] = self.df.groupby('title')[var].shift(self.num_shifts)
drop = [self.above_below_variables[0]+'_above', self.above_below_variables[0]+'_below']
for i in drop:
self.df = self.df.dropna(subset=[i])
def var_ratio(self):
'Generate the ratio of GR divided by specified variables'
for var in self.var2_ratio:
self.df[self.var1_ratio + '_' + var] = (self.df[self.var1_ratio]/self.df[var])
self.df[self.var1_ratio + '_' + var].loc[self.df[self.var1_ratio + '_' + var] == float('Inf')] = 0
self.df[self.var1_ratio + '_' + var].loc[self.df[self.var1_ratio + '_' + var] == -float('Inf')] = 0
def cleaning(self):
'Remove certain formations, rows with a lot of NAs and make y_variabl categorical'
self.df = self.df.drop(self.cols_to_remove,axis = 1)
self.df = self.df.dropna(thresh=self.thresh) #thresh= 12
self.df = self.df[np.isfinite(self.df['tvd'])]
self.df = self.df[(self.df.formation != 'water depth')]
self.df[self.y_variable] = self.df[self.y_variable].astype('category')
self.df = self.df[self.df[self.y_variable].cat.codes != -1]
self.df.reset_index(inplace=True, drop = True)
def xyz(self):
'Lat/Long for ML purposes'
self.df['x'] = np.cos(self.df['lat']) * np.cos(self.df['long'])
self.df['y'] = np.cos(self.df['lat']) * np.sin(self.df['long'])
self.df['z'] = np.sin(self.df['lat'])
self.df = self.df.drop(['lat','long'],axis = 1)
def single_pt_haversine(self,degrees=True):
"""
'Single-point' Haversine: Calculates the great circle distance
between a point on Earth and the (0, 0) lat-long coordinate
"""
r = 6371 # Earth's radius (km). Have r = 3956 if you want miles
# Convert decimal degrees to radians
if degrees:
lat, lng = map(math.radians, [self.df.lat, self.df.lng])
# 'Single-point' Haversine formula
a = math.sin(lat/2)**2 + math.cos(lat) * math.sin(lng/2)**2
d = 2 * r * math.asin(math.sqrt(a))
self.df['well_distance'] = [self.single_pt_haversine(x, y) for x, y in zip(lat, long)]
def drop_new_values(self):
'NAs are introduced when we calculate above and below. This function removes them'
drop = ["gr_above", "gr_below"]
for i in drop:
self.df = self.df.dropna(subset=[i])
self.df = self.df
def remove_outliers(self):
for key,value in self.outlier_values.items():
self.df = self.df[self.df[key] <= value]
self.df = self.df[self.df[key] >= 0]
def done(self):
'Return the self.df set and a dictionary of formations and their corresponding number'
self.remove_outliers()
self.remove_outliers()
self.log_values()
self.above_below()
self.cleaning()
self.xyz()
return self.df
```
### Visualizations
```
formation_colors = ['#d96c6c', '#ffe680', '#336633','#4d5766', '#cc99c9',
'#733939', '#f2eeb6', '#739978', '#333366', '#cc669c',
'#f2b6b6', '#8a8c69', '#66ccb8', '#bfbfff', '#733950',
'#b27159', '#c3d96c', '#336663', '#69698c', '#33262b',
'#bfa38f', '#2d3326', '#1a3133', '#8f66cc', '#99737d',
'#736256', '#65b359', '#73cfe6', '#673973', '#f2ba79',
'#bef2b6', '#86aab3', '#554359', '#8c6c46', '#465943',
'#73b0e6', '#ff80f6', '#4c3b26']
group_colors = ['#ff4400', '#cc804e', '#e5b800', '#403300', '#4da63f',
'#133328', '#00cad9', '#005fb3', '#0000f2', '#292259',
'#d052d9', '#33131c', '#ff6176']
def plot_well_comparison(df, well_index, formation_colors, group_colors, model_name = None, save = False):
#df['group_2'] = df['group'].astype('category').cat.codes
logs = df.loc[df["title"] == df.title.unique()[well_index]]
#logs = logs.sort_values(by='tvd')
cluster_predicted_formation=np.repeat(np.expand_dims(logs['predicted'].values,1), 100, 1)
cluster_actual_formation=np.repeat(np.expand_dims(logs['formation_2'].values,1), 100, 1)
cluster_predicted_group=np.repeat(np.expand_dims(logs['predicted_group'].values,1), 100, 1)
cluster_actual_group=np.repeat(np.expand_dims(logs['group_2'].values,1), 100, 1)
cmap_formation = colors.ListedColormap(formation_colors)
bounds_formation = [l for l in range(n_formation+1)]
norm_formation = colors.BoundaryNorm(bounds_formation, cmap_formation.N)
cmap_group = colors.ListedColormap(group_colors)
bounds_group = [l for l in range(n_group+1)]
norm_group = colors.BoundaryNorm(bounds_group, cmap_group.N)
#ztop=logs.tvd.min(); zbot=logs.tvd.max()
f, ax = plt.subplots(nrows=1, ncols=4, figsize=(8, 12))
im1=ax[0].imshow(cluster_predicted_formation, interpolation='none', aspect='auto',
cmap=cmap_formation,vmin=0,vmax=37)#, norm = norm_formation)
im2=ax[1].imshow(cluster_actual_formation, interpolation='none', aspect='auto',
cmap=cmap_formation,vmin=0,vmax=37)#, norm = norm_formation)
im3=ax[2].imshow(cluster_predicted_group, interpolation='none', aspect='auto',
cmap=cmap_group,vmin=0,vmax=12)#, norm = norm_group)
im4=ax[3].imshow(cluster_actual_group, interpolation='none', aspect='auto',
cmap=cmap_group,vmin=0,vmax=12)#, norm = norm_group)
ax[0].set_xlabel('Predicted formations')
ax[1].set_xlabel('Actual formations')
ax[2].set_xlabel('Predicted groups')
ax[3].set_xlabel('Actual groups')
ax[0].set_yticklabels([])
ax[1].set_yticklabels([]); ax[2].set_yticklabels([]); ax[3].set_yticklabels([])
f.suptitle('Well: %s'%logs.iloc[0]['title'], fontsize=14,y=0.91)
if save:
plt.savefig(fig_dir+'prediction_'+model_name+'_'+'well_'+str(well_index)+'.png')
plt.show()
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import make_axes_locatable
import matplotlib.colors as colors
def plot_well_logs(df, well_index, formation_colors, save = False):
df['group_2'] = df['group'].map(group_dictionary)
logs = df.loc[df["title"] == df.title.unique()[well_index]]
#logs = logs.sort_values(by='tvd')
cmap_formation = colors.ListedColormap(formation_colors)
cmap_group = colors.ListedColormap(group_colors1)
ztop=logs.tvd.min(); zbot=logs.tvd.max()
cluster=np.repeat(np.expand_dims(logs['formation_2'].values,1), 100, 1)
cluster_2=np.repeat(np.expand_dims(logs['group_2'].values,1), 100, 1)
f, ax = plt.subplots(nrows=1, ncols=8, figsize=(12, 16))
ax[0].plot(logs.gr, logs.tvd, '-g')
ax[1].plot(logs.rdep, logs.tvd, '-')
ax[2].plot(logs.rmed, logs.tvd, '-', color='r')
ax[3].plot(logs.dt, logs.tvd, '-', color='0.5')
ax[4].plot(logs.nphi, logs.tvd, '-', color='y')
ax[5].plot(logs.rhob, logs.tvd, '-', color='c')
im1=ax[6].imshow(cluster, interpolation='none', aspect='auto',
cmap=cmap_formation,vmin=1,vmax=37)
im2=ax[7].imshow(cluster_2, interpolation='none', aspect='auto',
cmap=cmap_group,vmin=1,vmax=12)
for i in range(len(ax)-2):
ax[i].set_ylim(ztop,zbot)
ax[i].invert_yaxis()
ax[i].grid()
ax[i].locator_params(axis='x', nbins=3)
ax[0].set_xlabel("gr")
ax[0].set_xlim(logs.gr.min(),logs.gr.max()+10)
ax[1].set_xlabel("rdep")
ax[1].set_xlim(logs.rdep.min(),logs.rdep.max()+0.5)
ax[2].set_xlabel("rmed")
ax[2].set_xlim(logs.rmed.min(),logs.rmed.max()+0.5)
ax[3].set_xlabel("dt")
ax[3].set_xlim(logs.dt.min(),logs.dt.max()+0.5)
ax[4].set_xlabel("nphi")
ax[5].set_xlim(logs.nphi.min(),logs.nphi.max()+0.5)
ax[5].set_xlabel("rhob")
ax[5].set_xlim(logs.rhob.min(),logs.rhob.max()+0.5)
ax[6].set_xlabel('Formations')
ax[7].set_xlabel('Group')
ax[1].set_yticklabels([]); ax[2].set_yticklabels([]); ax[3].set_yticklabels([]);ax[4].set_yticklabels([])
ax[5].set_yticklabels([]); ax[6].set_yticklabels([]); ax[7].set_yticklabels([])
f.suptitle('Well: %s'%logs.iloc[0]['title'], fontsize=14,y=0.91)
if save:
plt.savefig(fig_dir+'well_'+str(well_index)+'.01.png')
plt.show()
```
| github_jupyter |
# Fictional Army - Filtering and Sorting
### Introduction:
This exercise was inspired by this [page](http://chrisalbon.com/python/)
Special thanks to: https://github.com/chrisalbon for sharing the dataset and materials.
### Step 1. Import the necessary libraries
```
import pandas as pd
```
### Step 2. This is the data given as a dictionary
```
# Create an example dataframe about a fictional army
raw_data = {'regiment': ['Nighthawks', 'Nighthawks', 'Nighthawks', 'Nighthawks', 'Dragoons', 'Dragoons', 'Dragoons', 'Dragoons', 'Scouts', 'Scouts', 'Scouts', 'Scouts'],
'company': ['1st', '1st', '2nd', '2nd', '1st', '1st', '2nd', '2nd','1st', '1st', '2nd', '2nd'],
'deaths': [523, 52, 25, 616, 43, 234, 523, 62, 62, 73, 37, 35],
'battles': [5, 42, 2, 2, 4, 7, 8, 3, 4, 7, 8, 9],
'size': [1045, 957, 1099, 1400, 1592, 1006, 987, 849, 973, 1005, 1099, 1523],
'veterans': [1, 5, 62, 26, 73, 37, 949, 48, 48, 435, 63, 345],
'readiness': [1, 2, 3, 3, 2, 1, 2, 3, 2, 1, 2, 3],
'armored': [1, 0, 1, 1, 0, 1, 0, 1, 0, 0, 1, 1],
'deserters': [4, 24, 31, 2, 3, 4, 24, 31, 2, 3, 2, 3],
'origin': ['Arizona', 'California', 'Texas', 'Florida', 'Maine', 'Iowa', 'Alaska', 'Washington', 'Oregon', 'Wyoming', 'Louisana', 'Georgia']}
```
### Step 3. Create a dataframe and assign it to a variable called army.
#### Don't forget to include the columns names in the order presented in the dictionary ('regiment', 'company', 'deaths'...) so that the column index order is consistent with the solutions. If omitted, pandas will order the columns alphabetically.
```
army = pd.DataFrame(data=raw_data)
army
```
### Step 4. Set the 'origin' colum as the index of the dataframe
```
army
```
### Step 5. Print only the column veterans
```
army["veterans"]
```
### Step 6. Print the columns 'veterans' and 'deaths'
```
army[["veterans", "deaths"]]
```
### Step 7. Print the name of all the columns.
```
army.columns
```
### Step 8. Select the 'deaths', 'size' and 'deserters' columns from Maine and Alaska
```
army.loc[["Maine", "Alaska"], ["deaths", "size", "deserters"]]
```
### Step 9. Select the rows 3 to 7 and the columns 3 to 6
```
army.iloc[2:7, 2:6]
```
### Step 10. Select every row after the fourth row and all columns
```
army.iloc[4:, :]
```
### Step 11. Select every row up to the 4th row and all columns
```
army.iloc[:4, :]
```
### Step 12. Select the 3rd column up to the 7th column
```
army.iloc[:, 2:7]
```
### Step 13. Select rows where df.deaths is greater than 50
```
army[army.deaths > 50]
```
### Step 14. Select rows where df.deaths is greater than 500 or less than 50
```
army[(army.deaths > 500) | (army.deaths < 50)]
```
### Step 15. Select all the regiments not named "Dragoons"
```
army[army["regiment"] != "Dragoons"]
```
### Step 16. Select the rows called Texas and Arizona
```
army.loc[["Texas", "Arizona"]]
```
### Step 17. Select the third cell in the row named Arizona
```
army.loc[["Arizona"]].iloc[:,2]
```
### Step 18. Select the third cell down in the column named deaths
```
army.loc[:, ["deaths"]].iloc[2]
```
| github_jupyter |
This notebook was prepared by [Donne Martin](http://donnemartin.com). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# Challenge Notebook
## Problem: Determine if a linked list is a palindrome.
* [Constraints](#Constraints)
* [Test Cases](#Test-Cases)
* [Algorithm](#Algorithm)
* [Code](#Code)
* [Unit Test](#Unit-Test)
* [Solution Notebook](#Solution-Notebook)
## Constraints
* Can we assume this is a non-circular, singly linked list?
* Yes
* Is a single character or number a palindrome?
* No
* Can we assume we already have a linked list class that can be used for this problem?
* Yes
* Can we use additional data structures?
* Yes
* Can we assume this fits in memory?
* Yes
## Test Cases
* Empty list -> False
* Single element list -> False
* Two or more element list, not a palindrome -> False
* General case: Palindrome with even length -> True
* General case: Palindrome with odd length -> True
## Algorithm
Refer to the [Solution Notebook](http://nbviewer.ipython.org/github/donnemartin/interactive-coding-challenges/blob/master/linked_lists/palindrome/palindrome_solution.ipynb). If you are stuck and need a hint, the solution notebook's algorithm discussion might be a good place to start.
## Code
```
# %load ../linked_list/linked_list.py
class Node(object):
def __init__(self, data, next=None):
self.next = next
self.data = data
def __str__(self):
return self.data
class LinkedList(object):
def __init__(self, head=None):
self.head = head
def __len__(self):
curr = self.head
counter = 0
while curr is not None:
counter += 1
curr = curr.next
return counter
def insert_to_front(self, data):
if data is None:
return None
node = Node(data, self.head)
self.head = node
return node
def append(self, data):
if data is None:
return None
node = Node(data)
if self.head is None:
self.head = node
return node
curr_node = self.head
while curr_node.next is not None:
curr_node = curr_node.next
curr_node.next = node
return node
def find(self, data):
if data is None:
return None
curr_node = self.head
while curr_node is not None:
if curr_node.data == data:
return curr_node
curr_node = curr_node.next
return None
def delete(self, data):
if data is None:
return
if self.head is None:
return
if self.head.data == data:
self.head = self.head.next
return
prev_node = self.head
curr_node = self.head.next
while curr_node is not None:
if curr_node.data == data:
prev_node.next = curr_node.next
return
prev_node = curr_node
curr_node = curr_node.next
def delete_alt(self, data):
if data is None:
return
if self.head is None:
return
curr_node = self.head
if curr_node.data == data:
curr_node = curr_node.next
return
while curr_node.next is not None:
if curr_node.next.data == data:
curr_node.next = curr_node.next.next
return
curr_node = curr_node.next
def print_list(self):
curr_node = self.head
while curr_node is not None:
print(curr_node.data)
curr_node = curr_node.next
def get_all_data(self):
data = []
curr_node = self.head
while curr_node is not None:
data.append(curr_node.data)
curr_node = curr_node.next
return data
def get_middle(self):
print(f'inside get middle')
data = []
slow = self.head #by the time fast reaches to the end of linked list ie None, slow would reach middle
fast = self.head.next
while fast and fast.next:
slow = slow.next
fast = fast.next.next
return slow.data
def merge_linkedlists(self,head1,head2):
print(f'inside merge')
head=head1
print(id(head))
while(head2):
print(f'inside while')
list1=head1
list2=head2
temp1=list1.next
temp2=list2.next
list1.next=list2
list2.next=temp1
head1=temp1
head2=temp2
return head
def rotate(self,size):
if size==0:
return None
print(f'inside rotate')
rotate1=self.head
temp=self.head
for i in range(size):
print(i)
print(f'data is {rotate1.data}')
previous=rotate1
rotate1=rotate1.next
self.head=rotate1
print(self.head)
print(id(self.head))
previous.next=None
print(id(self.head))
forward=self.head
while(forward.next):
print(f'inside forward')
print(f'data is {forward.data}')
forward=forward.next
print(f'exited while')
print(f'data is {forward.data}')
forward.next=temp
class MyLinkedList(LinkedList):
def is_palindrome(self):
# TODO: Implement me
l=self.__len__()
if not l or l==1 or l==2:
return False
list1=self.get_all_data()
print(f'list1 is {list1}')
listrev=list1[::-1]
if list1==listrev and list1:
return True
return False
```
## Unit Test
**The following unit test is expected to fail until you solve the challenge.**
```
# %load test_palindrome.py
from nose.tools import assert_equal
class TestPalindrome(object):
def test_palindrome(self):
print('Test: Empty list')
linked_list = MyLinkedList()
assert_equal(linked_list.is_palindrome(), False)
print('Test: Single element list')
head = Node(1)
linked_list = MyLinkedList(head)
assert_equal(linked_list.is_palindrome(), False)
print('Test: Two element list, not a palindrome')
linked_list.append(2)
assert_equal(linked_list.is_palindrome(), False)
print('Test: General case: Palindrome with even length')
head = Node('a')
linked_list = MyLinkedList(head)
linked_list.append('b')
linked_list.append('b')
linked_list.append('a')
assert_equal(linked_list.is_palindrome(), True)
print('Test: General case: Palindrome with odd length')
head = Node(1)
linked_list = MyLinkedList(head)
linked_list.append(2)
linked_list.append(3)
linked_list.append(2)
linked_list.append(1)
assert_equal(linked_list.is_palindrome(), True)
print('Success: test_palindrome')
head = Node(1)
linked_list = MyLinkedList(head)
print(id(linked_list.head))
print(linked_list.head.data)
linked_list.append(3)
linked_list.append(5)
linked_list.append(7)
linked_list.append(9)
head = Node(2)
linked_list1 = MyLinkedList(head)
linked_list1.append(4)
linked_list1.append(6)
linked_list1.append(8)
linked_list.head=linked_list.merge_linkedlists(linked_list.head,linked_list1.head)
print(id(linked_list.head))
print(linked_list.get_all_data())
#print(linked_list.head.next.next.next.data)
#passing heads of both linked lists which are going to be merged
print(f'mid element is {linked_list.get_middle()}')
head = Node('a')
linked_list = MyLinkedList(head)
linked_list.append('b')
linked_list.append('c')
linked_list.append('d')
print(linked_list.get_all_data())
linked_list.rotate(2)
print(linked_list.get_all_data())
def main():
test = TestPalindrome()
test.test_palindrome()
if __name__ == '__main__':
main()
```
## Solution Notebook
Review the [Solution Notebook](http://nbviewer.ipython.org/github/donnemartin/interactive-coding-challenges/blob/master/linked_lists/palindrome/palindrome_solution.ipynb) for a discussion on algorithms and code solutions.
| github_jupyter |
# Soft Pseudo-labels
```
! rsync -a /kaggle/input/mmdetection-v280/mmdetection /
! pip install /kaggle/input/mmdetection-v280/src/mmpycocotools-12.0.3/mmpycocotools-12.0.3/
! pip install /kaggle/input/hpapytorchzoo/pytorch_zoo-master/
! pip install /kaggle/input/hpacellsegmentation/HPA-Cell-Segmentation/
! pip install /kaggle/input/iterative-stratification/iterative-stratification-master/
! cp -r /kaggle/input/kgl-humanprotein-data/kgl_humanprotein_data /
! cp -r /kaggle/input/humanpro/kgl_humanprotein /
import sys
sys.path.append('/kgl_humanprotein/')
import os
import time
from pathlib import Path
import shutil
import zipfile
import functools
import multiprocessing
import numpy as np
import pandas as pd
import cv2
from sklearn.model_selection import KFold,StratifiedKFold
from iterstrat.ml_stratifiers import MultilabelStratifiedKFold
import torch
from torch.backends import cudnn
from torch.utils.data import Dataset, DataLoader, RandomSampler, SequentialSampler
from torch.nn import DataParallel
import matplotlib.pyplot as plt
from tqdm import tqdm
from kgl_humanprotein.utils.common_util import *
from kgl_humanprotein.config.config import *
from kgl_humanprotein.data_process import *
from kgl_humanprotein.datasets.tool import image_to_tensor
from kgl_humanprotein.datasets.protein_dataset import *
from kgl_humanprotein.networks.imageclsnet import init_network
from kgl_humanprotein.layers.loss import *
from kgl_humanprotein.layers.scheduler import *
from kgl_humanprotein.utils.augment_util import train_multi_augment2
from kgl_humanprotein.utils.log_util import Logger
from kgl_humanprotein.run.train import *
%cd /kaggle
dir_data = Path('/kaggle/input')
dir_mdata = Path('/kaggle/mdata')
# set cuda visible device
gpu_id = '0'
os.environ['CUDA_VISIBLE_DEVICES'] = gpu_id
cudnn.benchmark = True
# set random seeds
torch.manual_seed(0)
torch.cuda.manual_seed_all(0)
np.random.seed(0)
```
# Data
```
def load_subset5_raw():
'''
Samples of a few under-represented single-labels from the external dataset.
With each channel's maximum value.
'''
pth = Path('/kaggle/input/humanpro-train-cells-subset5/humanpro_train_cells_subset5/train/train.feather')
df = pd.read_feather(pth)
df['subset'] = 5
return df
def load_subsets_raw():
'''
Samples from the competition dataset.
With each channel's maximum value.
'''
df_0to4 = pd.read_feather('/kaggle/input/humanpro-raw-meta-channel-max/train.feather')
df_5 = load_subset5_raw()
df = pd.concat([df_0to4, df_5], axis=0, ignore_index=True)
return df
def load_pseudo_raw():
'''
Multi-label samples from the competition dataset.
With hard pseudo-labels predicted by `humanpro-classifier-crop`.
'''
return pd.read_feather('/kaggle/input/humanpro-data-multilabel-cells-meta/train.feather')
def seperate_single_multi_labels(df):
is_singlelabel = df.Target.apply(lambda o: len(o.split('|')) == 1)
df_sgl = df[is_singlelabel].reset_index(drop=True)
df_mul = df[~is_singlelabel].reset_index(drop=True)
return df_sgl, df_mul
def replace_multi_with_pseudo(df_orig_multi, df_pseudo):
'''
Replace the `Target` in `df_orig_multi` with the pseudo-labels in `df_pseudo`.
'''
df = pd.merge(df_orig_multi, df_pseudo[['Id', 'Target']], left_on='Id', right_on='Id', how='inner')
df.rename({'Target_x':'original_target', 'Target_y':'Target'}, axis=1, inplace=True)
return df
def sort_target_labels(target):
labels = sorted(set(int(label) for label in target.split('|')))
target = '|'.join(str(label) for label in labels)
return target
%%time
df_orig = load_subsets_raw()
df_orig['Target'] = df_orig.Target.apply(sort_target_labels)
df_orig_sgl, df_orig_mul = seperate_single_multi_labels(df_orig)
df_orig.shape, df_orig_sgl.shape, df_orig_mul.shape
df_cells = df_orig_mul[df_orig_mul.subset == 5].reset_index(drop=True)
# del df_orig, df_orig_sgl, df_orig_mul
dir_mdata_raw = dir_mdata/'raw'
dir_mdata_raw.mkdir(exist_ok=True, parents=True)
df_cells.to_feather(dir_mdata_raw/'train.feather')
```
## Data preview
```
nsample = 10
df = df_cells.sample(nsample).reset_index(drop=True)
ncols = 4
nrows = (nsample - 1) // ncols + 1
fig, axs = plt.subplots(nrows=nrows, ncols=ncols, figsize=(4*ncols, 4*nrows))
axs = axs.flatten()
for ax in axs:
ax.axis('off')
for ax, (_, r) in zip(axs, df.iterrows()):
dir_img = (dir_data
/f'humanpro-train-cells-subset{r.subset}'
/f'humanpro_train_cells_subset{r.subset}'/'train'/'images_384')
img = load_RGBY_image(dir_img, r.Id)
ax.imshow(img[...,[0,3,2]])
ax.set_title(f"Subset{r.subset} | {r.Target} {r[['max_red', 'max_green', 'max_blue', 'max_yellow']].values}")
plt.tight_layout()
del df
```
## Filter samples
```
df_cells = pd.read_feather(dir_mdata/'raw'/'train.feather')
len(df_cells)
# For testing, just take a few samples
n_sample = len(df_cells)
df_cells = df_cells.sample(n_sample).reset_index(drop=True)
df_cells.to_feather(dir_mdata_raw/'train.feather')
```
## One-hot encode labels
```
%%time
generate_meta(dir_mdata, 'train.feather')
```
## Dataset, DataLoader
```
img_size = 384
crop_size = 256
batch_size = 64
workers = 3
pin_memory = True
valid_file = Path(dir_mdata/'meta'/'train_meta.feather')
in_channels = 4
assert valid_file.exists()
valid_dataset = ProteinDataset(dir_data, valid_file, img_size=img_size, is_trainset=True,
return_label=True, in_channels=in_channels, transform=None,
crop_size=crop_size, random_crop=False)
valid_loader = DataLoader(valid_dataset, sampler=SequentialSampler(valid_dataset),
batch_size=batch_size, drop_last=False,
num_workers=workers, pin_memory=pin_memory)
```
# Model
```
arch = 'class_densenet121_dropout'
num_classes = len(LABEL_NAME_LIST)
resume = Path('/kaggle/input/humanpro-classifier-crop/results/models/'
'external_crop256_focal_slov_hardlog_class_densenet121_dropout_i384_aug2_5folds/'
'fold0/final.pth')
model_params = {}
model_params['architecture'] = arch
model_params['num_classes'] = num_classes
model_params['in_channels'] = in_channels
model = init_network(model_params)
model = DataParallel(model)
model.to(DEVICE)
checkpoint = torch.load(resume)
model.module.load_state_dict(checkpoint['state_dict'])
```
# Loss
```
criterion = FocalSymmetricLovaszHardLogLoss().to(DEVICE)
focal_loss = FocalLoss().to(DEVICE)
```
# Predict
```
from torch.autograd import Variable
import torch.nn.functional as F
from sklearn.metrics import f1_score
def validate(valid_loader, model, criterion, epoch, focal_loss, threshold=0.5):
batch_time = AverageMeter()
losses = AverageMeter()
accuracy = AverageMeter()
# switch to evaluate mode
model.eval()
probs_list = []
labels_list = []
logits_list = []
loss_list = []
acc_list = []
end = time.time()
for it, iter_data in tqdm(enumerate(valid_loader, 0)):
images, labels, indices = iter_data
images = Variable(images.to(DEVICE))
labels = Variable(labels.to(DEVICE))
outputs = model(images)
loss = criterion(outputs, labels, epoch=epoch)
logits = outputs
probs = F.sigmoid(logits)
acc = multi_class_acc(probs, labels)
probs_list.append(probs.cpu().detach().numpy())
labels_list.append(labels.cpu().detach().numpy())
logits_list.append(logits.cpu().detach().numpy())
loss_list.append(loss.item())
acc_list.append(acc.item())
losses.update(loss.item())
accuracy.update(acc.item())
# measure elapsed time
batch_time.update(time.time() - end)
end = time.time()
probs = np.vstack(probs_list)
y_true = np.vstack(labels_list)
logits = np.vstack(logits_list)
loss = np.vstack(loss_list)
acc = np.vstack(acc_list)
valid_focal_loss = focal_loss.forward(torch.from_numpy(logits), torch.from_numpy(y_true))
y_pred = probs > threshold
kaggle_score = f1_score(y_true, y_pred, average='macro')
# return losses.avg, accuracy.avg, valid_focal_loss, kaggle_score
return losses.avg, accuracy.avg, valid_focal_loss, kaggle_score, probs, y_true, logits, loss, acc
%%time
with torch.no_grad():
(valid_loss, valid_acc, valid_focal_loss, kaggle_score,
probs, y_true, logits, loss, acc) = validate(valid_loader, model, criterion, 0, focal_loss)
36033 / 10 * 2.06 / 60**2
```
## Plot predicted probabilities
```
def plt_prob_target(prob, target):
'''
Red inidicates the image-level labels
'''
class_id = np.arange(len(prob))
prob_target = np.zeros_like(prob)
prob_target[target==1] = prob[target==1].copy()
prob_others = prob.copy()
prob_others[target==1] = 0
fig, ax = plt.subplots()
ax.bar(class_id, prob_others)
ax.bar(class_id, prob_target, color='red')
ax.set_ylabel('Probability')
ax.set_xlabel('Class')
ax.set_title(f'Target {np.where(target==1)[0]}')
return fig, ax
idx = 8
fig, ax = plt_prob_target(probs[idx], y_true[idx])
def get_softlabels(probs, y_true, keep_neg=False):
labels = probs.copy()
labels[y_true != 1] = 0
if keep_neg:
labels[:,-1] = probs[:,-1].copy()
return labels
```
# Write out softlabels
```
softlabels = get_softlabels(probs, y_true)
df_cells_softlabel_subset5 = df_cells.copy()
df_cells_softlabel_subset5[LABEL_NAME_LIST] = softlabels
idx = 8
fig, ax = plt_prob_target(softlabels[idx], y_true[idx])
df_cells_softlabel_subset5.iloc[8]
df_cells_softlabel = pd.read_feather('/kaggle/input/humanpro-data-soft-pseudolabel/df_cells_softlabel.feather')
# "13|13" were mis-identified as multi-labels before, so these need to be removed.
df_cells_softlabel['Target'] = df_cells_softlabel.Target.apply(sort_target_labels)
df_cells_softlabel_sgl, df_cells_softlabel_mul = seperate_single_multi_labels(df_cells_softlabel)
df_cells_softlabel = pd.concat([df_cells_softlabel_mul, df_cells_softlabel_subset5], axis=0, ignore_index=True)
df_cells_softlabel.subset.unique(), df_cells_softlabel_mul.shape, df_cells_softlabel_subset5.shape, df_cells_softlabel.shape, df_orig_mul.shape
df_cells_softlabel.to_feather('/kaggle/working/df_cells_softlabel.feather')
```
# Write out softlabels with `Negative` always non-zero
```
softlabels_neg = get_softlabels(probs, y_true, keep_neg=True)
df_cells_softlabel_neg_subset5 = df_cells.copy()
df_cells_softlabel_neg_subset5[LABEL_NAME_LIST] = softlabels_neg
idx = 3
fig, ax = plt_prob_target(softlabels_neg[idx], y_true[idx])
df_cells_softlabel_neg_subset5.iloc[3]
df_cells_softlabel_neg = pd.read_feather('/kaggle/input/humanpro-data-soft-pseudolabel/df_cells_softlabel_neg.feather')
# Targets like "13|13" were mis-identified as multi-labels before, so these need to be removed.
df_cells_softlabel_neg['Target'] = df_cells_softlabel_neg.Target.apply(sort_target_labels)
df_cells_softlabel_neg_sgl, df_cells_softlabel_neg_mul = seperate_single_multi_labels(df_cells_softlabel_neg)
df_cells_softlabel_neg = pd.concat([df_cells_softlabel_neg_mul, df_cells_softlabel_neg_subset5], axis=0, ignore_index=True)
df_cells_softlabel_neg.subset.unique(), df_cells_softlabel_neg_mul.shape, df_cells_softlabel_neg_subset5.shape, df_cells_softlabel_neg.shape, df_orig_mul.shape
df_cells_softlabel_neg.to_feather('/kaggle/working/df_cells_softlabel_neg.feather')
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from doctest import testmod as run_doctest
pd.set_option('display.max_rows', 1000)
CONFIRMED = 'https://raw.githubusercontent.com/CSSEGISandData/COVID-19' \
'/master/csse_covid_19_data/csse_covid_19_time_series' \
'/time_series_covid19_confirmed_global.csv'
DEATHS = 'https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master' \
'/csse_covid_19_data/csse_covid_19_time_series' \
'/time_series_covid19_deaths_global.csv'
RECOVERED = 'https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_recovered_global.csv'
confirmed = pd.read_csv(CONFIRMED)
deaths = pd.read_csv(DEATHS)
recovered = pd.read_csv(RECOVERED)
def _get(df, column, country):
"""
>>> _get(confirmed, 'Confirmed', 'Poland').loc['2021-07-01']
Confirmed 2880010
Name: 2021-07-01 00:00:00, dtype: int32
>>> _get(confirmed, 'Confirmed', 'India').loc['2021-07-01']
Confirmed 30458251
Name: 2021-07-01 00:00:00, dtype: int32
"""
if country:
q = df['Country/Region'] == country
df = df[q]
df = df.transpose()[4:].sum(axis='columns')
df = pd.DataFrame(df).astype(np.int32)
df.columns = [column]
df.index = pd.to_datetime(c.index)
return df
def covid19(country=None):
"""
>>> covid19('Poland').loc['2021-07-01']
Confirmed 2880010
Deaths 75044
Recovered 2651906
Name: 2021-07-01 00:00:00, dtype: int32
>>> covid19('India').loc['2021-07-01']
Confirmed 30458251
Deaths 400312
Recovered 29548302
Name: 2021-07-01 00:00:00, dtype: int32
>>> covid19('France').loc['2021-07-01']
Confirmed 5839967
Deaths 111298
Recovered 405572
Name: 2021-07-01 00:00:00, dtype: int32
>>> covid19().loc['2021-07-01']
Confirmed 182641987
Deaths 3956360
Recovered 120005935
Name: 2021-07-01 00:00:00, dtype: int32
"""
return pd.concat((
_get(confirmed, 'Confirmed', country),
_get(deaths, 'Deaths', country),
_get(recovered, 'Recovered', country),
), axis='columns')
run_doctest()
poland = covid19('Poland')
germany = covid19('Germany')
india = covid19('India')
france = covid19('France')
china = covid19('China')
world = covid19()
def ratio(df, since='2021-01-01', until='2021-07-27'):
return df.loc[since:until, 'Confirmed'] / df.loc[since:until, 'Recovered']
ratio(poland, '2021-07', '2021-07').plot()
ratio(india).plot()
```
| github_jupyter |
```
import torch
import torchvision
import torchvision.transforms as transforms
import torch.optim as optim
import numpy as np
import time
from prune import *
#TODO
n_epochs = 3
batch_size_train = 64
batch_size_test = 1000
learning_rate = 0.01
momentum = 0.5
log_interval = 10
random_seed = 1
torch.backends.cudnn.enabled = False
torch.manual_seed(random_seed)
train_loader = torch.utils.data.DataLoader(
torchvision.datasets.MNIST('/files/', train=True, download=True,
transform=torchvision.transforms.Compose([
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize(
(0.1307,), (0.3081,))
])),
batch_size=batch_size_train, shuffle=True)
test_loader = torch.utils.data.DataLoader(
torchvision.datasets.MNIST('/files/', train=False, download=True,
transform=torchvision.transforms.Compose([
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize(
(0.1307,), (0.3081,))
])),
batch_size=batch_size_test, shuffle=True)
import torch.nn as nn
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 10, kernel_size=5)
self.conv2 = nn.Conv2d(10, 20, kernel_size=5)
self.conv2_drop = nn.Dropout2d()
self.fc1 = nn.Linear(320, 50)
self.fc2 = nn.Linear(50, 10)
def forward(self, x):
x = F.relu(F.max_pool2d(self.conv1(x), 2))
x = F.relu(F.max_pool2d(self.conv2_drop(self.conv2(x)), 2))
x = x.view(-1, 320)
x = F.relu(self.fc1(x))
x = F.dropout(x, training=self.training)
x = self.fc2(x)
return F.log_softmax(x)
network = Net()
optimizer = optim.SGD(network.parameters(), lr=learning_rate,
momentum=momentum)
train_losses = []
train_counter = []
test_losses = []
test_counter = [i*len(train_loader.dataset) for i in range(n_epochs + 1)]
def train(epoch,locked_masks,network):
#network=Net()
network.train()
for prune_step in range(5):
if prune_step > 0:
print('Start Pruning')
prune(network,locked_masks, prune_random=False, prune_weight=True, prune_bias=False, ratio=0.5,
threshold=None, threshold_bias=None, function=None, function_bias=None, prune_across_layers=True)
print('Done Pruning')
#correct(test_loader,network)
prune_diag(network,locked_masks)
test(network)
for batch_idx, (data, target) in enumerate(train_loader):
optimizer.zero_grad()
output = network(data)
loss = F.nll_loss(output, target)
loss.backward()
network = prune_grad(network,locked_masks) #zeros gradients of the pruned weights
#for n, w in network.named_parameters():
# if w.grad is not None and n in locked_masks:
# w.grad[locked_masks[n]] = 0
optimizer.step()
if batch_idx % log_interval == 0:
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
epoch, batch_idx * len(data), len(train_loader.dataset),
100. * batch_idx / len(train_loader), loss.item()))
train_losses.append(loss.item())
train_counter.append(
(batch_idx*64) + ((epoch-1)*len(train_loader.dataset)))
torch.save(network.state_dict(), './results/model.pth')
torch.save(optimizer.state_dict(), './results/optimizer.pth')
test(network)
def test(network):
network.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for data, target in test_loader:
output = network(data)
test_loss += F.nll_loss(output, target, size_average=False).item()
pred = output.data.max(1, keepdim=True)[1]
correct += pred.eq(target.data.view_as(pred)).sum()
test_loss /= len(test_loader.dataset)
test_losses.append(test_loss)
print('\nTest set: Avg. loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)\n'.format(
test_loss, correct, len(test_loader.dataset),
100. * correct / len(test_loader.dataset)))
#initialize locked masks:
locked_masks = {n: torch.zeros(w.size(), dtype=torch.bool) for n, w in network.named_parameters()}
for n, w in network.named_parameters():
print(n)
print(w.size())
for epoch in range(1, n_epochs + 1):
train(epoch,locked_masks,network)
# test()
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(net.parameters(), lr=0.005, momentum=0.9)
#initialize locked masks:
locked_masks = {n: torch.zeros(w.size(), dtype=torch.bool) for n, w in net.named_parameters()}
#start time
start=time.time()
for prune_step in range(5):
if prune_step > 0:
print('Start Pruning')
prune(net,locked_masks, prune_random=False, prune_weight=True, prune_bias=False, ratio=0.75,
threshold=None, threshold_bias=None, function=None, function_bias=None, prune_across_layers=True)
print('Done Pruning')
correct(test_loader,net)
prune_diag(net,locked_masks)
#print('prune diag time: ',time.time()-s2)
for epoch in range(4): # loop over the dataset multiple times
running_loss = 0.0
for i, data in enumerate(train_loader,0):
# get the inputs; data is a list of [inputs, labels]
inputs, labels = data
inputs = inputs.view(inputs.shape[0], -1)
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + prune_grad + optimize
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
net = prune_grad(net,locked_masks) #zeros gradients of the pruned weights
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 200 == 199: # print every 200 mini-batches
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / 200))
running_loss = 0.0
correct(test_loader,net)
print('Finished Training')
print('time: ',time.time()-start)
for n, w in network.named_parameters():
#print(len(w.data.numpy().shape))
print(len(w.shape))
print(w)
```
| github_jupyter |
Copyright (c) Microsoft Corporation. Licensed under the MIT license.
# Train and Deploy a model using Feast
In this notebook we show how to:
1. access a feature store
1. discover features in the feature store
1. train a model using the offline store (using the feast function `get_historical_features()`)
1. use the feast `materialize()` function to push features from the offline store to an online store (redis)
1. Deploy the model to an Azure ML endpoint where the features are consumed from the online store (feast function `get_online_features()`)
## Connect to Feature store
Below we create a Feast repository config, which accesses the registry.db file and also provides the credentials to the offline and online storage.
__You need to update the feature_repo/feature_store.yaml file so that the registry path points to your blob location__
```
import os
from feast import FeatureStore
from azureml.core import Workspace
# access key vault to get secrets
ws = Workspace.from_config()
kv = ws.get_default_keyvault()
os.environ['SQL_CONN']=kv.get_secret("FEAST-SQL-CONN")
os.environ['REDIS_CONN']=kv.get_secret("FEAST-REDIS-CONN")
# connect to feature store
fs = FeatureStore("./feature_repo")
```
### List the feature views
Below lists the registered feature views.
```
fs.list_feature_views()
```
## Load features into a pandas dataframe
Below you load the features from the feature store into a pandas data frame.
```
sql_job = fs.get_historical_features(
entity_df="SELECT * FROM orders",
features=[
"driver_stats:conv_rate",
"driver_stats:acc_rate",
"driver_stats:avg_daily_trips",
"customer_profile:current_balance",
"customer_profile:avg_passenger_count",
"customer_profile:lifetime_trip_count",
],
)
training_df = sql_job.to_df()
training_df.head()
```
## Train a model and capture metrics with MLFlow
```
import mlflow
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from azureml.core import Workspace
# connect to your workspace
ws = Workspace.from_config()
# create experiment and start logging to a new run in the experiment
experiment_name = "order_model"
# set up MLflow to track the metrics
mlflow.set_tracking_uri(ws.get_mlflow_tracking_uri())
mlflow.set_experiment(experiment_name)
mlflow.sklearn.autolog()
training_df = training_df.dropna()
X = training_df[['conv_rate', 'acc_rate', 'avg_daily_trips',
'current_balance', 'avg_passenger_count','lifetime_trip_count' ]].dropna()
y = training_df['order_is_success']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
clf = RandomForestClassifier(n_estimators=10)
# train the model
with mlflow.start_run() as run:
clf.fit(X_train, y_train)
```
## Prepare for deployment
### Register model and the feature registry
```
# register the model
model_uri = "runs:/{}/model".format(run.info.run_id)
model = mlflow.register_model(model_uri, "order_model")
```
### `materialize()` data into the online store (redis)
```
from datetime import datetime, timedelta
end_date = datetime.now()
start_date = end_date - timedelta(days=365)
fs.materialize(start_date=start_date, end_date=end_date)
```
## Set up deployment configuration
__Note: You will need to set up a service principal (SP) and add that SP to your blob storage account as a *Storage Blob Data Contributor* role to authenticate to the storage containing the feast registry file.__
Once you have set up the SP, populate the `AZURE_CLIENT_ID`, `AZURE_TENANT_ID`, `AZURE_CLIENT_SECRET` environment variables below.
```
from azureml.core.environment import Environment
from azureml.core.webservice import AciWebservice
from azureml.core import Workspace
ws = Workspace.from_config()
keyvault = ws.get_default_keyvault()
# create deployment config i.e. compute resources
aciconfig = AciWebservice.deploy_configuration(
cpu_cores=1,
memory_gb=1,
description="orders service using feast",
)
# get registered environment
env = Environment.from_conda_specification("feast-env", "model_service_env.yml")
# again ensure that the scoring environment has access to the registry file
env.environment_variables = {
"FEAST_SQL_CONN": fs.config.offline_store.connection_string,
"FEAST_REDIS_CONN": fs.config.online_store.connection_string,
"FEAST_REGISTRY_BLOB": fs.config.registry.path,
"AZURE_CLIENT_ID": "",
"AZURE_TENANT_ID": "",
"AZURE_CLIENT_SECRET": ""
}
```
## Deploy model
Next, you deploy the model to Azure Container Instance. Please note that this may take approximately 10 minutes.
```
import uuid
from azureml.core.model import InferenceConfig
from azureml.core.environment import Environment
from azureml.core.model import Model
# get the registered model
model = Model(ws, "order_model")
# create an inference config i.e. the scoring script and environment
inference_config = InferenceConfig(
entry_script="score.py",
environment=env,
source_directory="src"
)
# deploy the service
service_name = "orders-service" + str(uuid.uuid4())[:4]
service = Model.deploy(
workspace=ws,
name=service_name,
models=[model],
inference_config=inference_config,
deployment_config=aciconfig,
)
service.wait_for_deployment(show_output=True)
```
## Test service
Below you test the service. The first score takes a while as the feast registry file is downloaded from blob. Subsequent runs will be faster as feast uses a local cache for the registry.
```
import json
input_payload = json.dumps({"driver":50521, "customer_id":20265})
service.run(input_data=input_payload)
```
| github_jupyter |
```
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import os
import fasttext
from sklearn import cluster
from sklearn import metrics
from sklearn.feature_extraction.text import TfidfVectorizer
import re
import string
import numpy as np
from nltk.corpus import stopwords
from pymystem3 import Mystem
from collections import Counter
data_loc = '/home/cherepaha/Projects/made/3sem/realtime_news/news_data/'
news_path = os.path.join(data_loc, 'social_network_news.csv')
stats_path = os.path.join(data_loc, 'social_network_stats.csv')
rss_path = os.path.join(data_loc, 'news.csv')
news = pd.read_csv(news_path)
stats = pd.read_csv(stats_path)
rss = pd.read_csv(rss_path)
news.head()
stats.head()
stat_fields = ['comments', 'likes', 'reposts', 'views']
group_stats = stats.groupby(['post_id', 'social_network']).agg(
**{f'{field}_max': (field, max) for field in stat_fields},
**{f'{field}_min': (field, min) for field in stat_fields}
)
group_stats.head()
```
# Per post visualization
```
post_id = 2573845
sn = 'vk'
post_dynamic = stats[(stats.post_id == post_id) & (stats.social_network == sn)]
for f in stat_fields:
post_dynamic.plot('id', f)
```
# Resource agg
Графики по какому-то домену: распределения лайков, вьюшек, в каком канале (вк или тг), средние по дням и т.п..
```
resource_stats = stats.merge(news[['source_name', 'post_id', 'date']], on='post_id', how='inner')
resource_stats = resource_stats.sort_values(stat_fields, ascending=False)
resource_stats.drop_duplicates('post_id', inplace=True)
domain = 'ria.ru'
# domain = 'svpressa.ru'
domain_stats = resource_stats[resource_stats.source_name == domain]
plt.figure(figsize=(15, 5))
for i, f in enumerate(stat_fields):
ax = plt.subplot(1, len(stat_fields), i + 1)
left = 0
right = domain_stats[f].quantile(0.9)
plt_kwargs = dict(
ax=ax,
bins=10,
alpha=0.5,
range=(left, right)
)
domain_stats[domain_stats.social_network == 'vk'].hist(f, **plt_kwargs, label='vk')
if domain_stats[domain_stats.social_network == 'tg'][f].sum():
domain_stats[domain_stats.social_network == 'tg'].hist(f, **plt_kwargs, label='tg')
plt.legend()
date_dynamic = resource_stats.groupby(['social_network', 'source_name', 'date']).agg(**{f: (f, sum) for f in stat_fields})
date_dynamic = date_dynamic.reset_index()
```
П.С. тут думал сделать сравнение между доменами, но их много и график получается нечитабельным
```
social_network = 'vk'
domain = 'iz.ru'
for f in stat_fields:
data_filter = (date_dynamic['social_network'] == social_network) & (date_dynamic['source_name'] == domain)
date_dynamic[data_filter].plot('date', f)
```
# Clusterization
Идея: найти похожие новости и сравнить лайки, вьюшки и т.д.
```
model = fasttext.load_model('cc.ru.300.bin')
stopwords
emoji_pattern = re.compile(pattern = "["
u"\U0001F600-\U0001F64F" # emoticons
u"\U0001F300-\U0001F5FF" # symbols & pictographs
u"\U0001F680-\U0001F6FF" # transport & map symbols
u"\U0001F1E0-\U0001F1FF" # flags (iOS)
"]+", flags = re.UNICODE)
rus_stopwords = stopwords.words('russian')
mystem = Mystem()
def prepare_text(text):
text = re.sub(r'(https?://\S+)', ' ', text)
text = re.sub(r'\n', ' ', text)
text = re.sub(r'\d', ' ', text)
text = re.sub(emoji_pattern, ' ', text)
text = re.sub(r'\s+', ' ', text)
text = text.translate(str.maketrans('', '', string.punctuation)).lower()
tokens = mystem.lemmatize(text.lower())
tokens = [token for token in tokens if token not in rus_stopwords\
and token != " "
]
text = " ".join(tokens)
text = re.sub(r'\n', ' ', text)
text = re.sub(r'\s+', ' ', text)
return text
date = '2021-12-18'
news_text = news[news.date == date][['post_id', 'text', 'social_network']]
news_text.dropna(inplace=True)
news_text.loc[:, 'text'] = news_text['text'].apply(prepare_text)
news_text = news_text[news_text.text.apply(len) > 1]
tfidf = TfidfVectorizer(smooth_idf=True)
tfidf.fit(news_text.text)
def embed_text(text):
text = set(text.split())
text = sum(model[token] for token in text) / len(text)
return text
def embed_tfidf_text(text):
tf = Counter(text.split())
text_emb = [model[token] * tf[token] * tfidf.idf_[tfidf.vocabulary_[token]]
for token in tf
if token in tfidf.vocabulary_]
text_emb = sum(text_emb) / len(text.split())
return text_emb
# text_emb = news_text['text'].apply(lambda x: model[x])
text_emb = news_text['text'].apply(embed_tfidf_text)
text_emb = pd.DataFrame(text_emb.to_list(), )
ax = plt.subplot(1, 1, 1)
news_text[news_text.social_network == 'vk'].text.apply(len).hist(ax=ax, range=(0, 1000))
news_text[news_text.social_network == 'tg'].text.apply(len).hist(ax=ax, range=(0, 1000))
(text_emb**2).sum(axis=1).hist()
%%time
text_clusters = cluster.AgglomerativeClustering(
n_clusters=None,
affinity='cosine',
linkage='complete',
distance_threshold=0.2,
).fit_predict(text_emb)
# text_clusters = cluster.DBSCAN(metric='cosine', eps=0.03).fit_predict(text_emb)
# length = np.sqrt((text_emb**2).sum(axis=1))[:,None]
# text_clusters = cluster.KMeans(n_clusters=50, n_jobs=-1).fit_predict(text_emb / length)
news_text['cluster'] = text_clusters
metrics.silhouette_score(text_emb, text_clusters, metric='cosine')
cluster_stat = news_text.groupby('cluster').count()
cluster_stat.sort_values('post_id', ascending=False)
cluster_stat.post_id.hist(bins=20)
cluster_stat[cluster_stat.post_id > 10]
news_text[news_text.cluster == 55].values
```
Вывод: Нужна хорошая кластеризация (Можно еще попробовать knn)
# RSS join
Идея: подтягивать текст новости из вк/тг
```
def preproc_link(link_set):
link_set = link_set[1:-1].split(',')
return link_set
news_link = news[['link', 'text']].dropna(subset=['link'])
news_link.loc[:, 'link'] = news_link.link.apply(preproc_link)
news_link = news_link.explode('link')
news_link = news_link[news_link.link.apply(len) > 0]
rss_vk_merged = rss[['date', 'content', 'title', 'source_url', 'source_name']].merge(
news_link,
left_on='source_url',
right_on='link'
)
rss_vk_merged.shape
rss.shape
```
Слишком маленькое пересечение
| github_jupyter |
```
import pandas as pd
import tensorflow as tf
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
# 준비된 수술 환자 데이터를 불러들입니다.
Data_set = pd.read_csv("sample_data/pima-indians-diabetes.csv",header=None)
data_vale=Data_set.values
# 환자의 기록과 수술 결과를 X와 Y로 구분하여 저장합니다.
X = data_vale[:,0:8]
Y = data_vale[:,8:9]
# 전체 데이터에서 학습 데이터와 테스트 데이터(0.2)로 구분
X_train1, X_test, Y_train1, Y_test = train_test_split(X, Y, test_size=0.3,shuffle=True)
X_train, X_valid, Y_train, Y_valid = train_test_split(X_train1, Y_train1, test_size=0.2,shuffle=True)
# 딥러닝 구조를 결정합니다(모델을 설정하고 실행하는 부분입니다).
input_Layer = tf.keras.layers.Input(shape=(8,))
x = tf.keras.layers.Dense(20, activation='sigmoid')(input_Layer)
Out_Layer= tf.keras.layers.Dense(1, activation='sigmoid')(x)
model = tf.keras.models.Model(inputs=[input_Layer], outputs=[Out_Layer])
model.summary()
# 딥러닝을 실행합니다.
loss=tf.keras.losses.binary_crossentropy
optimizer = tf.keras.optimizers.SGD(learning_rate=0.0005)
metrics=tf.keras.metrics.binary_accuracy
model.compile(loss=loss, optimizer=optimizer, metrics=[metrics])
result=model.fit(X_train, Y_train, epochs=1000, batch_size=300, validation_data=(X_valid,Y_valid))
print(result.history.keys())
### history에서 loss와 val_loss의 key를 가지는 값들만 추출
loss = result.history['loss']
val_loss = result.history['val_loss']
### loss와 val_loss를 그래프화
epochs = range(1, len(loss) + 1)
plt.subplot(211) ## 2x1 개의 그래프 중에 1번째
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
### history에서 binary_accuracy와 val_binary_accuracy key를 가지는 값들만 추출
rmse = result.history['binary_accuracy']
val_rmse = result.history['val_binary_accuracy']
epochs = range(1, len(rmse) + 1)
### binary_accuracy와 val_binary_accuracy key를 그래프화
plt.subplot(212) ## 2x1 개의 그래프 중에 2번째
plt.plot(epochs, rmse, 'ro', label='Training binary_accuracy')
plt.plot(epochs, val_rmse, 'r', label='Validation binary_accuracy')
plt.title('Training and validation binary_accuracy')
plt.xlabel('Epochs')
plt.ylabel('binary_accuracy')
plt.legend()
# 결과를 출력합니다.
print("-----")
print(model.evaluate(X_test, Y_test))
print("\n Accuracy: %.4f" % (model.evaluate(X_test, Y_test)[1]))
plt.show()
# 준비된 수술 환자 데이터를 불러들입니다.
Data_set = pd.read_csv("sample_data/pima-indians-diabetes.csv",header=None)
data_vale=Data_set.values
# 환자의 기록과 수술 결과를 X와 Y로 구분하여 저장합니다.
X = data_vale[:,0:8]
Y = data_vale[:,8:9]
# 전체 데이터에서 학습 데이터와 테스트 데이터(0.2)로 구분
X_train1, X_test, Y_train1, Y_test = train_test_split(X, Y, test_size=0.3,shuffle=True)
X_train, X_valid, Y_train, Y_valid = train_test_split(X_train1, Y_train1, test_size=0.2,shuffle=True)
# 딥러닝 구조를 결정합니다(모델을 설정하고 실행하는 부분입니다).
input_Layer = tf.keras.layers.Input(shape=(8,))
x = tf.keras.layers.Dense(20, activation='sigmoid')(input_Layer)
Out_Layer= tf.keras.layers.Dense(1, activation='sigmoid')(x)
model = tf.keras.models.Model(inputs=[input_Layer], outputs=[Out_Layer])
model.summary()
# 딥러닝을 실행합니다.
loss=tf.keras.losses.binary_crossentropy
optimizer = tf.keras.optimizers.SGD(learning_rate=0.0005)
metrics=tf.keras.metrics.binary_accuracy
model.compile(loss=loss, optimizer=optimizer, metrics=[metrics])
result=model.fit(X_train, Y_train, epochs=1000, batch_size=300, validation_data=(X_valid,Y_valid))
print(result.history.keys())
### history에서 loss와 val_loss의 key를 가지는 값들만 추출
loss = result.history['loss']
val_loss = result.history['val_loss']
### loss와 val_loss를 그래프화
epochs = range(1, len(loss) + 1)
plt.subplot(211) ## 2x1 개의 그래프 중에 1번째
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
### history에서 binary_accuracy와 val_binary_accuracy key를 가지는 값들만 추출
rmse = result.history['binary_accuracy']
val_rmse = result.history['val_binary_accuracy']
epochs = range(1, len(rmse) + 1)
### binary_accuracy와 val_binary_accuracy key를 그래프화
plt.subplot(212) ## 2x1 개의 그래프 중에 2번째
plt.plot(epochs, rmse, 'ro', label='Training binary_accuracy')
plt.plot(epochs, val_rmse, 'r', label='Validation binary_accuracy')
plt.title('Training and validation binary_accuracy')
plt.xlabel('Epochs')
plt.ylabel('binary_accuracy')
plt.legend()
plt.show()
# 결과를 출력합니다.
print("-----")
print(model.evaluate(X_test, Y_test))
print("\n Accuracy: %.4f" % (model.evaluate(X_test, Y_test)[1]))
```
| github_jupyter |
# Data Acquisition
```
import pandas as pd
from datetime import datetime
def revertStrToDate(date_str_list):
datelist = []
for date_str in date_str_list:
date_temp = datetime.strptime(date_str, '%m/%d/%y').date()
datelist.append(date_temp)
return datelist
us_confirm_data = pd.read_csv('data/RAW_us_confirmed_cases.csv')
mask_mandates_data = pd.read_csv('data/U.S._State_and_Territorial_Public_Mask_Mandates_From_April_10__2020_through_August_15__2021_by_County_by_Day.csv')
mask_use_data = pd.read_csv('data/mask-use-by-county.csv')
us_confirm_data.head(10)
mask_mandates_data.head(10)
mask_use_data.head(10)
```
# Data Cleaning
```
county_confirm_data = pd.DataFrame(us_confirm_data[us_confirm_data['Admin2'] =='Essex'])
county_confirm_data = pd.DataFrame(county_confirm_data[us_confirm_data['Province_State'] =='Massachusetts'])
county_confirm_data
county_confirm_data = county_confirm_data.drop(['Province_State', 'Admin2','UID','iso2' ,'iso3', 'code3','FIPS','Country_Region','Lat','Long_','Combined_Key'], axis=1)
date = list(county_confirm_data.columns)
datelist1 = revertStrToDate(date)
cum_cases = county_confirm_data.values.tolist()[0]
con_cases = []
i = 0
for i in range(len(cum_cases) - 1):
con = cum_cases[i + 1] - cum_cases[i]
con_cases.append(con)
i = i + 1
con_cases.append(0)
new_cases = []
i = 0
for i in range(len(con_cases) - 1):
new = con_cases[i + 1] - con_cases[i]
new_cases.append(new)
i = i + 1
new_cases.append(0)
county_confirm = pd.DataFrame(datelist1, columns=['date'])
county_confirm.insert(loc=1, column='confirmed_cases', value = con_cases)
county_confirm.insert(loc=2, column='new_cases', value = new_cases)
county_mask_mandate_data = pd.DataFrame(mask_mandates_data[mask_mandates_data['County_Name'].str.contains('Essex')])
county_mask_mandate_data = pd.DataFrame(county_mask_mandate_data[county_mask_mandate_data['State_Tribe_Territory']=='MA'])
county_mask_mandate_data.head()
county_mask_mandate_data = county_mask_mandate_data.drop(['State_Tribe_Territory','County_Name','FIPS_State','FIPS_County','order_code', 'Source_of_Action','URL','Citation'], axis = 1)
county_mask_mandate_data = county_mask_mandate_data.fillna(0)
county_mask_mandate_data['Face_Masks_Required_in_Public'] = county_mask_mandate_data['Face_Masks_Required_in_Public'].replace({'No': 0, 'Yes': 1})
county_mask_mandate_data.head()
date2 = county_mask_mandate_data['date'].tolist()
cvt_date = []
for temp in date2:
d = temp[0:-4] + temp[-2:]
cvt_date.append(d)
datelist2 = revertStrToDate(cvt_date)
county_mask_mandate_data['date'] = datelist2
county_mask_use_data = pd.DataFrame(mask_use_data[mask_use_data['COUNTYFP'] == 25009])
county_mask_use_data
county_confirm.head(10)
county_mask_mandate_data.head(10)
print(county_mask_mandate_data['Face_Masks_Required_in_Public'].tolist())
mask_require = pd.DataFrame(county_mask_mandate_data[county_mask_mandate_data['Face_Masks_Required_in_Public'] ==1])
mask_require.head()
mask_require.tail()
county_covid_df = county_confirm
county_covid_df = county_covid_df.set_index('date')
```
# Visualization
```
import matplotlib.pyplot as plt
from matplotlib.dates import date2num
# Plot the visualization
plt.figure(figsize=(14,8))
plt.title('Covid Confirmed Cases in Essex, MA from Feb,2020 to Oct,2021')
plt.plot(county_covid_df['confirmed_cases'],label = 'confirmed cases')
plt.axvspan(date2num(datetime(2020,5,6)), date2num(datetime(2021,5,28)),
label="Mask Require",color="green", alpha=0.2)
plt.xlabel('Year')
plt.ylabel('Confirmed Cases')
plt.legend()
plt.savefig('covid confirmed cases vs time.png')
plt.show
```
| github_jupyter |
# Monte Carlo Localization
千葉工業大学 上田 隆一
(c) 2017 Ryuichi Ueda
This software is released under the MIT License, see LICENSE.
## はじめに
このコードは、移動ロボットの自己位置推定に使われるMonte Carlo Localization(MCL)のサンプルです。
## コードの流れ
確率的な考え方で自分の位置を推定するロボットは、自身の姿勢(位置と向き)について確証は持たず、常に自身の姿勢について曖昧な表現をします。 MCLでは、ロボットが自身の姿勢を複数の候補点で表現します。ロボットは複数の候補点のうちのどれか、あるいは候補点が分布している範囲のどこかに自身が存在するという考え(自我)を持つことになります。
候補点は、空間に浮かぶ粒という意味で「パーティクル」(粒子)と呼ばれたり、ロボットが居そうなところから候補点を一つ選んだという意味で「サンプル」(標本)と呼ばれたりします。
パーティクルの分布は、ロボットが移動や観測を行うごとに、ロボットの姿勢をよりよく推定できるように更新されます。通常、ロボットが移動するとパーティクスの分布は広がり、観測すると狭くなります。
## ヘッダ
ヘッダです。計算に対してはnumpy以外、特に変わったものは使いません。
```
%matplotlib inline
import numpy as np
from copy import copy
import math, random
import matplotlib.pyplot as plt # for plotting data
from matplotlib.patches import Ellipse # for drawing
```
## 二次元のガウス分布を表現するクラス
尤度の計算と描画に使用します。
```
class Gaussian2D:
# 共分散行列、中心の座標を属性に持つ
def __init__(self,sigma_x = 1.0, sigma_y = 1.0, cov_xy = 0.0,mu_x = 0.0, mu_y = 0.0):
self.cov = np.array([[sigma_x**2,cov_xy],[cov_xy,sigma_y**2]])
self.mean = np.array([mu_x,mu_y]).T
# ガウス分布の移動
def shift(self,delta,angle):
ca = math.cos(angle)
sa = math.sin(angle)
rot = np.array([[ca,sa],[-sa,ca]])
self.cov = rot.dot(self.cov).dot(rot.T)
self.mean = self.mean + delta
# 密度の算出
def value(self, pos):
delta = pos - self.mean
numerator = math.exp(-0.5 * (delta.T).dot(np.linalg.inv(self.cov)).dot(delta))
denominator = 2 * math.pi * math.sqrt(np.linalg.det(self.cov))
return numerator / denominator
```
## ランドマークの管理クラス
配列でもいいのですが、描画の関係で一つのクラスで複数のランドマークの位置を管理するという方式にしています。ランドマークの位置は、世界座標系のx,y座標で表されます。このランドマークをロボットが見ると、どのランドマークか(ランドマークのID)、ロボットからの距離と見える方角の3つの値が雑音付きで得られると仮定します。ランドマークのIDは、self.positionsのリストでの位置(0,1,2,...)とします。
```
class Landmarks:
def __init__(self,array):
self.positions = array
def draw(self):
xs = [ e[0] for e in self.positions]
ys = [ e[1] for e in self.positions]
plt.scatter(xs,ys,s=300,marker="*",label="landmarks",color="orange")
```
## 1回のランドマーク計測を管理するクラス
ロボットの真の姿勢とランドマークの位置情報をコンストラクタで受けて、センサの返す値をシミュレートします。
```
class Observation:
def __init__(self,robot_pos, landmark,lid):
# センサの有効範囲の設定
self.sensor_max_range = 1.0
self.sensor_min_range = 0.1
self.sensor_max_angle = math.pi / 2
self.sensor_min_angle = - math.pi /2
# ランドマークのIDを保存しておく属性。ランドマークがセンサの有効範囲にないとNoneのまま
self.lid = None
# 真の位置の情報をセットする。ロボットの真の姿勢はシミュレーション用でロボットは知らないという前提。
# 真のランドマークの位置は、ロボットは知っているのでこのインスタンスの属性として保存します。
rx,ry,rt = robot_pos
self.true_lx,self.true_ly = landmark
# ロボットからランドマークまでの距離の真値を算出
distance = math.sqrt((rx-self.true_lx)**2 + (ry-self.true_ly)**2)
if distance > self.sensor_max_range or distance < self.sensor_min_range:
return
# ロボットからランドマークがどの方向に見えるか真値を算出
direction = math.atan2(self.true_ly-ry, self.true_lx-rx) - rt
if direction > math.pi: direction -= 2*math.pi
if direction < -math.pi: direction += 2*math.pi
if direction > self.sensor_max_angle or direction < self.sensor_min_angle:
return
# 真値に混入する雑音の大きさ(標準偏差)を設定
sigma_distance = distance * 0.1 # 距離に対して10%の標準偏差
sigma_direction = math.pi * 3 / 180 # ランドマークの方向に対して3degの標準偏差
# 雑音を混ぜてセンサの値とする
self.distance = random.gauss(distance, sigma_distance)
self.direction = random.gauss(direction, sigma_direction)
# ロボット座標系での共分散行列を作っておく。あとで尤度を計算するときに使用
# x方向が奥行きで、sigma_distanceを標準偏差に設定。y方向がロボットから見て横方向の誤差で、距離*sin(3[deg])となる。
self.error_ellipse = Gaussian2D(sigma_x = sigma_distance, sigma_y = self.distance * math.sin(sigma_direction) , cov_xy = 0.0)
self.lid = lid
# 尤度の計算(遅い実装です。)
# パーティクルの姿勢とランドマークの計測値からランドマークの位置を推定し、その位置に誤差楕円を置き、
# ランドマークの真の位置が誤差楕円からどれだけ外れているかを確率密度関数の密度として返します。
# この計算はもっと簡略化できますが、描画の関係でこういう手順を踏んでいます。
# 簡略な方法: パーティクルの姿勢とランドマークの真の位置から、想定されるランドマークの距離・方向を算出し、
# 実際の距離・方向とそれぞれ比較する方法。距離の誤差の傾向、方向の誤差の傾向をそれぞれ1次元のガウス分布で表現し、
# それぞれを独立して計算して尤度を算出し、掛け算する。
def likelihood(self,particle_pos):
# パーティクルの姿勢と、このインスタンスに保存されているセンサの値から、ランドマークの位置を求める
rx, ry, rt = particle_pos
proposed_lx = rx + self.distance * math.cos(rt + self.direction)
proposed_ly = ry + self.distance * math.sin(rt + self.direction)
# このインスタンスに保存されている共分散行列を、計算されたランドマークの位置に移し、パーティクルの向きに合わせて共分散行列を回転
e = copy(self.error_ellipse)
e.shift(np.array([proposed_lx, proposed_ly]).T, rt + self.direction)
# そのままガウス分布の計算式から密度(尤度)を返します。
return e.value(np.array([self.true_lx,self.true_ly]).T)
# 描画用
def ellipse(self,robot_pos):
rx, ry, rt = robot_pos[0], robot_pos[1], robot_pos[2]
proposed_lx = rx + self.distance * math.cos(rt + self.direction)
proposed_ly = ry + self.distance * math.sin(rt + self.direction)
e = copy(self.error_ellipse)
e.shift(np.array([proposed_lx, proposed_ly]).T, rt + self.direction)
# 固有ベクトルを二つ求めて、それぞれの大きさを求めて楕円を作り、幅を計算した方の固有ベクトルの向きに楕円を回転すると誤差楕円になります。
eigen = np.linalg.eig(e.cov)
v1 = eigen[0][0] * eigen[1][0]
v2 = eigen[0][1] * eigen[1][1]
v1_direction = math.atan2(v1[1],v1[0])
elli = Ellipse([proposed_lx, proposed_ly],width=math.sqrt(np.linalg.norm(v1)),height=math.sqrt(np.linalg.norm(v2)),angle=v1_direction/3.14*180)
elli.set_alpha(0.2)
return elli
# 描画用
def draw(self,sp,robot_pos):
sp.add_artist(self.ellipse(robot_pos))
```
### ランドマークを3つ環境に置く
```
actual_landmarks = Landmarks(np.array([[-0.5,0.0],[0.5,0.0],[0.0,0.5]]))
actual_landmarks.draw()
```
### パーティクルのクラスとパーティクルフィルタのクラス
```
# パーティクルのクラス。単なる構造体
class Particle:
def __init__(self,x,y,t,w):
self.pos = np.array([x,y,t])
self.w = w
# パーティクルフィルタのクラス
class ParticleFilter:
# この実装ではコンストラクタはパーティクルの個数だけを引数にとる
def __init__(self,num):
# 空のパーティクルのリストを作って一つずつ追加していく(実装がベタ)
self.particles = []
for i in range(num):
self.particles.append(Particle(0.0,0.0,0.0,1.0/num)) # パーティクルは重みを持つ。全パーティクルの重みの合計は1。1つのパーティクルの重みは1/個数
# ロボットが動いたときにパーティクルを動かすためのメソッド
# 引数の「motion」はメソッドで、ロボットの移動を再現するためのもの。
# ロボットは自身がどのように動作するとどう姿勢が変化するかを知っており、このメソッドがその知識となる。
def moveParticles(self,fw,rot,motion):
self.resampling() # このメソッドについては後述
# パーティクルごとに移動した後の姿勢を計算し、姿勢を更新する。
for p in self.particles:
after = motion(p.pos,fw,rot)
p.pos = after
# リサンプリングのためのメソッド。
# リサンプリングは、重みがごく少数のパーティクルに偏ることを防ぐための措置で、近似していない理論上の数式では出現しない。
def resampling(self):
num = len(self.particles) # numはパーティクルの個数
ws = [e.w for e in self.particles] # 重みのリストを作る
print(sum(ws))
if sum(ws) < 1e-100: #重みの和がゼロに丸め込まれるとサンプリングできなくなるので小さな数を足しておく
ws = [e + 1e-100 for e in ws]
ps = random.choices(self.particles, weights=ws, k=num) # パーティクルのリストから、weightsのリストの重みに比例した確率で、num個選ぶ
self.particles = [Particle(*e.pos,1.0/num) for e in ps] # 選んだリストからパーティクルを取り出し、パーティクルの姿勢から重み1/numの新しいパーティクルを作成
# 描画用
def draw(self,c="blue",lbl="particles"):
xs = [p.pos[0] for p in self.particles]
ys = [p.pos[1] for p in self.particles]
vxs = [math.cos(p.pos[2]) for p in self.particles]
vys = [math.sin(p.pos[2]) for p in self.particles]
plt.quiver(xs,ys,vxs,vys,color=c,label=lbl,alpha=0.7)
```
### ロボットを表現するクラス
ロボットはランドマークを観測して1ステップ進んで・・・を繰り返します。
```
class Robot:
def __init__(self,x,y,rad):
random.seed()
# actual_poses: ロボットの姿勢の真値を1ステップごとに記録したもの
# (ロボットのクラス内にいるけどロボットはこの情報を使えない)
self.actual_poses = [np.array([x,y,rad])]
# パーティクルフィルタの準備(パーティクル数30個)
self.pf = ParticleFilter(30)
# ロボットの動作をシミュレートするメソッド。シミュレーションだけでなく、ロボットがパーティクルを移動するときにも用いる。
# つまり実機に実装する場合もこのメソッドが必要となる。雑音の度合いは事前に計測するか、
# ざっくり決めてフィルタのロバスト性に頼る。
def motion(self, pos, fw, rot):
# fwだけ前進してその後rotだけ回転。雑音を混入させる
actual_fw = random.gauss(fw,fw/10) #進む距離に対して標準偏差10%の雑音を混入
dir_error = random.gauss(0.0, math.pi / 180.0 * 3.0) # 前進方向がヨレる雑音を標準偏差3[deg]で混入
px, py, pt = pos
# 移動後の位置を算出
x = px + actual_fw * math.cos(pt + dir_error)
y = py + actual_fw * math.sin(pt + dir_error)
# 雑音込みの回転各を算出。rotに対して標準偏差10%の雑音を混ぜる
actual_rot = random.gauss(rot,rot/10)
t = pt + dir_error + actual_rot # さらにヨレの分の角度を足す
return np.array([x,y,t])
# ロボットが動くときに呼び出すメソッド。ロボットの位置の更新とパーティクルの位置の更新
def move(self,fw,rot):
self.actual_poses.append(self.motion(self.actual_poses[-1],fw,rot))
self.pf.moveParticles(fw,rot,self.motion)
# ロボットがランドマーク観測するときに呼び出すメソッド
def observation(self,landmarks):
obss = []
for i,landmark in enumerate(landmarks.positions): # 3つあるランドマークを1つずつ観測
obss.append(Observation(self.actual_poses[-1],landmark,i))
obss = list(filter(lambda e : e.lid != None, obss)) # 観測データのないものを除去
# 重みに尤度をかける
for obs in obss:
for p in self.pf.particles:
p.w *= obs.likelihood(p.pos)
# 描画用に観測のリストを返す
return obss
# 描画用
def draw(self,sp,observations):
for obs in observations:
for p in self.pf.particles:
obs.draw(sp,p.pos)
self.pf.draw()
xs = [e[0] for e in self.actual_poses]
ys = [e[1] for e in self.actual_poses]
vxs = [math.cos(e[2]) for e in self.actual_poses]
vys = [math.sin(e[2]) for e in self.actual_poses]
plt.quiver(xs,ys,vxs,vys,color="red",label="actual robot motion")
```
## 描画用の関数
説明は割愛。
```
def draw(i,observations):
fig = plt.figure(i,figsize=(8, 8))
sp = fig.add_subplot(111, aspect='equal')
sp.set_xlim(-1.0,1.0)
sp.set_ylim(-0.5,1.5)
robot.draw(sp,observations)
actual_landmarks.draw()
plt.legend()
```
## シミュレーションの実行
図の説明:
* 赤の矢印: 真の姿勢
* 星: ランドマークの位置
* 青の矢印: パーティクルの姿勢
* 楕円: ランドマークの観測値と各パーティクルの姿勢からランドマークの位置を計算したものと、その位置の曖昧さを表す共分散行列
```
robot = Robot(0,0,0) # ロボットを原点に
# 観測、描画、移動の繰り返し
for i in range(0,18):
obss = robot.observation(actual_landmarks)
draw(i,obss)
robot.move(0.2,math.pi / 180.0 * 20)
```
| github_jupyter |
# DSA simulations
```
% matplotlib inline
%config InlineBackend.figure_format = 'retina'
%load_ext line_profiler
%load_ext autoreload
%autoreload 2
from __future__ import division
import numpy as np
import glob, os
import matplotlib.pyplot as plt
import matplotlib
matplotlib.rcParams['figure.dpi'] = 2.5 * matplotlib.rcParams['figure.dpi']
import astropy
from astropy.time import Time
import enterprise
from enterprise.pulsar import Pulsar
import enterprise_extensions
from enterprise_extensions import models, model_utils
import libstempo as T2, libstempo.toasim as LT, libstempo.plot as LP
from ephem import Ecliptic, Equatorial
datadir = '../partim_no_noise/'
def figsize(scale):
fig_width_pt = 513.17 #469.755 # Get this from LaTeX using \the\textwidth
inches_per_pt = 1.0/72.27 # Convert pt to inch
golden_mean = (np.sqrt(5.0)-1.0)/2.0 # Aesthetic ratio (you could change this)
fig_width = fig_width_pt*inches_per_pt*scale # width in inches
fig_height = fig_width*golden_mean # height in inches
fig_size = [fig_width,fig_height]
return fig_size
#plt.rcParams.update(plt.rcParamsDefault)
params = {'backend': 'pdf',
'axes.labelsize': 10,
'lines.markersize': 4,
'font.size': 10,
'xtick.major.size':6,
'xtick.minor.size':3,
'ytick.major.size':6,
'ytick.minor.size':3,
'xtick.major.width':0.5,
'ytick.major.width':0.5,
'xtick.minor.width':0.5,
'ytick.minor.width':0.5,
'lines.markeredgewidth':1,
'axes.linewidth':1.2,
'legend.fontsize': 7,
'xtick.labelsize': 10,
'ytick.labelsize': 10,
'savefig.dpi':200,
'path.simplify':True,
'font.family': 'serif',
'font.serif':'Times',
'text.latex.preamble': [r'\usepackage{amsmath}',r'\usepackage{amsbsy}',
r'\DeclareMathAlphabet{\mathcal}{OMS}{cmsy}{m}{n}'],
'text.usetex':True,
'figure.figsize': figsize(0.5)}
plt.rcParams.update(params)
```
## Useful functions
```
import pandas as pd
# Get (equatorial) position of pulsars from name
def pos_from_name(name):
pos = []
for p in name:
if '+' in p:
tmp = p.split('+')
raj = '{0}:{1}:00.00'.format(tmp[0][1:3],tmp[0][3:])
decj = '+{0}:{1}:00.00'.format(tmp[1][:2],tmp[1][2:])
else:
tmp = p.split('-')
raj = '{0}:{1}:00.00'.format(tmp[0][1:3],tmp[0][3:])
decj = '-{0}:{1}:00.00'.format(tmp[1][:2],tmp[1][2:])
eq = Equatorial(raj, decj)
if 'B' in p: epoch = '1950'
elif 'J' in p: epoch= '2000'
ec = Ecliptic(eq, epoch=str(epoch))
pos.append([float(eq.ra), float(eq.dec)])
return np.array(pos)
# Get name of pulsar from (equatorial) position
def name_from_pos(pos):
name = []
for p in pos:
eq = Equatorial(p[0], p[1])
ra = str(eq.ra)
dec = str(eq.dec)
#
if float(ra.split(':')[0]) < 10:
ra = '0' + ''.join(ra.split(':')[:2])
else:
ra = ''.join(ra.split(':')[:2])
#
if np.abs(float(dec.split(':')[0])) < 10:
dec = ':'.join(['-0'+dec.split(':')[0][1:],dec.split(':')[1]])
else:
dec = ':'.join(dec.split(':')[:2])
if float(dec.split(':')[0]) < 0:
dec = ''.join(dec.split(':')[:2])
elif float(dec.split(':')[0]) > 0:
dec = '+' + ''.join(dec.split(':')[:2])
else:
dec = ''.join(dec.split(':')[:2])
#
name.append('J' + ra + dec)
return np.array(name)
# Draw new random positions based on a sample
def invtran_sample(pos, size):
x, y = pos[:,0], pos[:,1]
hist, xbin, ybin = np.histogram2d(x, y, bins=(10, 10))
xbinc = xbin[:-1] + np.diff(xbin)/2.0
ybinc = ybin[:-1] + np.diff(ybin)/2.0
cdf = np.cumsum(hist.ravel())
cdf = cdf / cdf[-1]
values = np.random.rand(size)
value_bins = np.searchsorted(cdf, values)
x_idx, y_idx = np.unravel_index(value_bins,
(len(xbinc),
len(ybinc)))
delta_x = np.diff(xbin)[0]
delta_y = np.diff(ybin)[0]
if size == 1:
random_from_cdf = [xbinc[x_idx][0] + np.random.uniform(-delta_x/2.0, delta_x/2.0),
ybinc[y_idx][0] + np.random.uniform(-delta_y/2.0, delta_y/2.0)]
else:
random_from_cdf = np.column_stack((xbinc[x_idx] + np.random.uniform(-delta_x/2.0, delta_x/2.0),
ybinc[y_idx] + np.random.uniform(-delta_y/2.0, delta_y/2.0)))
return random_from_cdf
def year2mjd(year):
# rounds to nearest year
return float(Time("{}-01-01T00:00:00".format(str(int(np.rint(year)))),
format='isot').mjd)
def mjd2year(mjd):
return float(Time(mjd, format='mjd').decimalyear)
```
## Process data
```
data = pd.read_csv('../data/v2/RMSonlyvsTime2018-DSA2000-LoseAO.csv',header=0,skip_blank_lines=True,)
# Correcting for mistake in spreadsheet
for ii,name in data.iterrows():
print name
#if name.Observatory == 'GBT' and name.PSR == 'NEWPSR' and name.Epoch2 >= 2025:
# data.iloc[ii,6] = name.RMS3
# data.iloc[ii,7] = np.nan
# data.iloc[ii,8] = np.nan
# names of all real gbt and ao pulsars
real_pulsars_gbt = np.array([name.PSR for ii,name in data.iterrows() if 'NEWPSR' not in name.PSR and name.Observatory=='GBT'])
real_pulsars_ao = np.array([name.PSR for ii,name in data.iterrows() if 'NEWPSR' not in name.PSR and name.Observatory=='AO'])
# names of all fake gbt and ao pulsars (these are all NEWPSR)
fake_pulsars_gbt = np.array([name.PSR for ii,name in data.iterrows() if 'NEWPSR' in name.PSR and name.Observatory=='GBT'])
fake_pulsars_ao = np.array([name.PSR for ii,name in data.iterrows() if 'NEWPSR' in name.PSR and name.Observatory=='AO'])
# Get equatorial positions (ra, dec...in radians) from names
pos_pulsars_gbt = pos_from_name(real_pulsars_gbt)
pos_pulsars_ao = pos_from_name(real_pulsars_ao)
# Generate new fake pulsar positions
fakepos_pulsars_gbt = np.array([invtran_sample(pos_pulsars_gbt, size=1) for ii in range(len(fake_pulsars_gbt))])
fakepos_pulsars_ao = np.array([invtran_sample(pos_pulsars_ao, size=1) for ii in range(len(fake_pulsars_ao))])
# Generate new fake pulsar names
fake_pulsars_gbt = name_from_pos(fakepos_pulsars_gbt)
fake_pulsars_ao = name_from_pos(fakepos_pulsars_ao)
# Make copy of data frame and replace NEWPSR with new names
data_copy = data.copy(deep=True)
ct_gbt = 0
ct_ao = 0
for ii,name in data_copy.iterrows():
# New GBT pulsars
if name.Observatory == 'GBT' and name.PSR=='NEWPSR':
#tmp_pos = invtran_sample(pos_pulsars_gbt, size=1)
#tmp_name = name_from_pos(tmp_pos)
#data_copy.iloc[ii,0] = tmp_name[0]
data_copy.iloc[ii,0] = fake_pulsars_gbt[ct_gbt]
ct_gbt += 1
# New AO pulsars
if name.Observatory == 'AO' and name.PSR=='NEWPSR':
#tmp_pos = invtran_sample(pos_pulsars_ao, size=1)
#tmp_name = name_from_pos(tmp_pos)
#data_copy.iloc[ii,0] = tmp_name[0]
data_copy.iloc[ii,0] = fake_pulsars_ao[ct_ao]
ct_ao += 1
# Create new data columns for RAJ and DECJ
data_copy = data_copy.assign(RAJ=pd.Series(np.concatenate([pos_pulsars_gbt[:,0],
fakepos_pulsars_gbt[:,0],
pos_pulsars_ao[:,0],
fakepos_pulsars_ao[:,0]])).values)
data_copy = data_copy.assign(DECJ=pd.Series(np.concatenate([pos_pulsars_gbt[:,1],
fakepos_pulsars_gbt[:,1],
pos_pulsars_ao[:,1],
fakepos_pulsars_ao[:,1]])).values)
```
### Dealing with Burning Dumpster sim
```
data = pd.read_csv('../data/v2/RMSonlyvsTime2018-DSA2000-BurningDumpster.csv',header=0,skip_blank_lines=True,)
data_copy = data.copy(deep=True)
final_data = pd.read_csv('../data/v2/status_quo.csv')
data_copy.PSR = final_data.PSR
data_copy = data_copy.assign(RAJ=final_data.RAJ)
data_copy = data_copy.assign(DECJ=final_data.DECJ)
# Write to csv
data_copy.to_csv('../data/v2/burning_dumpster.csv')
```
## Creating par files
```
sim_type = 'status_quo' # 'status_quo', 'lose_ao', or 'burning_dumpster'
# Read from csv
data_copy = pd.read_csv('../data/v2/{}.csv'.format(sim_type))
for ii,name in data_copy.iterrows():
# Get coordinates for par file
tmp = Equatorial(data_copy.iloc[ii].RAJ, data_copy.iloc[ii].DECJ, epoch='2000')
# Set PEPOCHs to be 5 years in
rms = np.array([name.RMS1, name.RMS2, name.RMS3,
name.RMS4, name.RMS5])
rms[rms=='gap'] = np.inf
rms = np.array(rms,dtype=float)
epoch = np.array([name.Epoch1, name.Epoch2, name.Epoch3,
name.Epoch4, name.Epoch5])
start = epoch[np.where(~np.isnan(rms))[0][0]]
with open('../data/template.par', 'r') as fil:
pardata = fil.read()
with open('../data/v2/par/' + name.PSR + '.par', 'w') as filnew:
for line in pardata.split('\n'):
if 'PSR' in line:
print >>filnew, '\t\t'.join([line.split()[0], name.PSR])
elif 'RAJ' in line:
print >>filnew, '\t\t'.join([line.split()[0], str(tmp.ra),
line.split()[2], line.split()[3]])
elif 'DECJ' in line:
print >>filnew, '\t\t'.join([line.split()[0], str(tmp.dec),
line.split()[2], line.split()[3]])
elif 'PEPOCH' in line or 'POSEPOCH' in line or 'DMEPOCH' in line:
print >>filnew, '\t\t'.join([line.split()[0], str(int(year2mjd(start+5)))])
else:
print >>filnew, line
print name.PSR, start
```
## Creating tim files
```
sim_type = 'burning_dumpster' # 'status_quo', 'lose_ao', or 'burning_dumptser'
# Read from csv
data_copy = pd.read_csv('../data/v2/{}.csv'.format(sim_type))
dsa_sims = []
start_data = []
for jj,name in data_copy.iterrows():
psrname = name.PSR
rms = np.array([name.RMS1, name.RMS2, name.RMS3,
name.RMS4, name.RMS5])
rms[rms=='gap'] = np.inf
rms = np.array(rms,dtype=float)
epoch = np.array([name.Epoch1, name.Epoch2, name.Epoch3,
name.Epoch4, name.Epoch5])
### Start and End year
start_yr = epoch[np.where(~np.isnan(rms))[0][0]]
start_yr_mjd = year2mjd(start_yr)
#
end_yr = 2045
end_yr_mjd = year2mjd(end_yr)
### Spacing and obstimes
spacing = 365.25 / 20.0 # days between observations
#
obstimes = np.arange(start_yr_mjd, end_yr_mjd, spacing)
# removing data gaps
for kk,rmss in enumerate(rms):
if np.isinf(rmss):
mask = np.logical_and(obstimes >= year2mjd(epoch[kk]),
obstimes <= year2mjd(epoch[kk+1]))
obstimes = obstimes[~mask]
### Segmenting obstimes based on hardware/telescope switches
stops = list(epoch[np.where(~np.isnan(rms))[0]]) + [end_yr]
stops = [year2mjd(yr) for yr in stops]
errors = list(rms[np.where(~np.isnan(rms))[0]])
### Masking sections of data based on these stops
masks = []
for kk,stop in enumerate(stops):
if kk < len(stops)-1:
masks.append(np.logical_and(obstimes >= stops[kk],
obstimes <= stops[kk+1]))
### Applying RMS errors
toa_errs = np.ones_like(obstimes)
for kk,mask in enumerate(masks):
toa_errs[mask] *= float(errors[kk])
### Make fake dataset
par = '../data/v2/par/' + psrname + '.par'
dsa_sims.append(LT.fakepulsar(parfile=par, obstimes=obstimes,
toaerr=toa_errs,
observatory=name.Observatory.lower()))
# white noise
LT.add_efac(dsa_sims[jj])
# save .tim
dsa_sims[jj].savetim('../data/v2/tim_{}/'.format(sim_type)
+ dsa_sims[jj].name + '.tim')
###
start_data.append([psrname, start_yr, start_yr_mjd])
print psrname, par, start_yr_mjd, end_yr_mjd, len(stops), len(masks), len(errors)
start_data = np.array(start_data)
start_data[start_data[:,1].argsort()]
fil = open('sims_psr_startdata_{}.txt'.format(sim_type),'w')
for line in start_data[start_data[:,1].argsort()]:
print >>fil, line[0], line[1], line[2]
fil.close()
```
# Read In And Check Pulsars
```
import enterprise
from enterprise.pulsar import Pulsar
from enterprise.signals import parameter
from enterprise.signals import white_signals
from enterprise.signals import gp_signals
from enterprise.signals import signal_base
import enterprise_extensions
from enterprise_extensions import models, model_utils
psr1 = Pulsar('../data/v2/par/J2234+0944.par', '../data/v2/tim_status_quo/J2234+0944.tim', ephem='DE436')
psr2 = Pulsar('../data/v2/par/J2234+0944.par', '../data/v2/tim_lose_ao/J2234+0944.tim', ephem='DE436')
psr3 = Pulsar('../data/v2/par/J2234+0944.par', '../data/v2/tim_burning_dumpster/J2234+0944.tim', ephem='DE436')
plt.errorbar([mjd2year(p) for p in psr1.toas/86400.0],
psr1.residuals/1e-6, psr1.toaerrs/1e-6,
alpha=0.3, fmt='.')
#plt.errorbar([mjd2year(p) for p in psr2.toas/86400.0],
# psr2.residuals/1e-6, psr2.toaerrs/1e-6,
# alpha=0.3, fmt='.')
plt.errorbar([mjd2year(p) for p in psr3.toas/86400.0],
psr3.residuals/1e-6, psr3.toaerrs/1e-6,
alpha=0.3, fmt='.')
plt.xlabel(r'Year')
plt.ylabel(r'Residuals [$\mu$s]')
plt.title(psr1.name)
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
from IPython.display import Audio
```
# Import Modules
```
from services import load_credentials, get_database, get_watson_service, reset_services
vcap = load_credentials()
reset_services()
from watson_developer_cloud import LanguageTranslationV2, TextToSpeechV1, NaturalLanguageUnderstandingV1
```
# Database
```
db, client = get_database('test')
client.all_dbs()
[doc for doc in db]
db.all_docs(include_docs=True)
db.create_document({'name' : 'Manolo'})
db.create_document({'name' : 'Pedro'})
list(map(lambda doc: doc['name'], db))
db.delete()
```
# Translator
```
translator = get_watson_service('language_translator')
translator.get_identifiable_languages()
translator.identify('Je ne sais pas francais')['languages'][0]
s = translator.translate('I am very happy', source='en', target='ja')
s
translator.translate(s, source='ja', target='en')
s = translator.translate('I am on top of the world', model_id='en-es-conversational')
s
translator.translate(s, model_id='es-en-conversational')
```
# Text to Speech
```
text_to_speech = get_watson_service('text_to_speech')
audioEN = text_to_speech.synthesize('Hello world!', accept='audio/ogg',
voice="en-GB_KateVoice")
Audio(audioEN)
[x['name'] for x in text_to_speech.voices()['voices']]
audioES = text_to_speech.synthesize('Hola Mundo', accept='audio/ogg',
voice="es-ES_LauraVoice")
Audio(audioES)
```
# Speech to Text
```
speech_to_text = get_watson_service('speech_to_text')
speech_to_text.recognize(
audioEN, content_type='audio/ogg')
speech_to_text.recognize(
audioEN, content_type='audio/ogg', timestamps=True,
word_confidence=True)
[(x['language'], x['name']) for x in speech_to_text.models()['models'] ]
speech_to_text.recognize(
audioES, content_type='audio/ogg', model='es-ES_BroadbandModel', timestamps=True,
word_confidence=True, )
```
# Natural Language Understanding
```
import watson_developer_cloud.natural_language_understanding.features.v1 as features
nlu = get_watson_service('natural-language-understanding')
nlu.analyze(text='this is my experimental text. Bruce Banner is the Hulk and Bruce Wayne is BATMAN! Superman fears not Banner, but Wayne.',
features=[features.Entities(), features.Keywords()])
nlu.analyze(text='I love you', features=[features.Emotion(), features.Sentiment()], language='en')
nlu.analyze(text='Lucia odia a Margarita', features=[features.Entities(), features.Sentiment()], language='es')
text='One day, there was a ferret in the Wellness Room (at Petco, the Wellness Room is where animals who are ill or having any type of issue are kept in order to receive special attention and any medical care needed.) He was by himself and he dooked, a lot. “Dooking” is how ferrets “talk.” Cats meow. Dogs bark. Birds chirp. Ferrets dook. It almost sounds like a chicken clucking. In talking with the General Manager, we decided that the ferret was lonely, so I made it a point to go visit him in the Wellness Room whenever possible. This ferret had a skin infection and was receiving oral medications to help clear up the infection. Well, the day finally came when he finished all of his medication and was ready to join the others on the sales floor. However, I couldn’t give him up; we had bonded during our daily visits. I’d grown accustomed to him being there in the Wellness Room, waiting for me each morning; greeting me with enthusiasm that grew each day he got better. So, I bought him and named him Gizmo since his dooking reminded me of Gizmo the Gremlin. As I was getting ready to take Gizmo home, I happened to walk past the habitat on the sales floor (I’m sure I was just heading to the ladies room, yeah, that’s it). Well, low and behold, there was a single, little female ferret all by her lonesome. How could I pack up Gizmo and just walk past her and leave her by herself? Especially knowing that ferrets do better in pairs or multiples, especially when raised together or introduced correctly. Welcome Mink to the family.'
nlu.analyze(text=text, features=[features.Entities(), features.Keywords(), features.Sentiment(),
features.Emotion(), features.Relations(), features.Concepts()])
```
# Visual Recognition
```
visual_recognition = get_watson_service('watson_vision_combined')
url = "http://weknowyourdreams.com/images/volcano/volcano-05.jpg"
visual_recognition.classify(images_url=url)
url="http://creator.keepcalmandcarryon.com/kcp-preview/NQOxtJXW"
visual_recognition.recognize_text(images_url=url)
url = "https://fthmb.tqn.com/TbYcG1ZT3HMQNs2APucg94XyunE=/400x0/filters:no_upscale()/about/overlyattached-5900fa4a3df78c54563e3d8b.jpg"
visual_recognition.detect_faces(images_url=url)
```
| github_jupyter |
```
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.layers import Dense, Input, GlobalMaxPooling1D
from keras.layers import Conv1D, MaxPooling1D, Embedding
from keras.models import Model
from sklearn.metrics import roc_auc_score
MAX_SEQUENCE_LENGTH = 100
MAX_VOCAB_SIZE = 20000
EMBEDDING_DIM = 100
VALIDATION_SPLIT = 0.2
BATCH_SIZE = 128
EPOCHS = 10
word2vec = {}
with open(os.path.join('./data/glove.6B.%sd.txt' % EMBEDDING_DIM)) as f:
for line in f:
values = line.split()
word = values[0]
vec = np.asarray(values[1:], dtype='float32')
word2vec[word] = vec
print("Found %s word vectors" % len(word2vec))
train = pd.read_csv('./data/train.csv')
train.head()
train.info()
sentences = train["comment_text"].fillna("DUMMY_VALUE").values
possible_labels = ["toxic", "severe_toxic", "obscene", "threat", "insult", "identity_hate"]
targets = train[possible_labels].values
# convert the sentences (strings) into integers
tokenizer = Tokenizer(num_words=MAX_VOCAB_SIZE)
tokenizer.fit_on_texts(sentences)
sequences = tokenizer.texts_to_sequences(sentences)
print("max sequence length", max(len(s) for s in sequences))
print("min sequence length", min(len(s) for s in sequences))
s = sorted(len(s) for s in sequences)
print("median sequence length", s[len(s) // 2])
# get word -> integer mapping
word2idx = tokenizer.word_index
print("Unique tokens", len(word2idx))
# pad sequences so that we get a N x T matrix
data = pad_sequences(sequences, maxlen=MAX_SEQUENCE_LENGTH)
print(data.shape)
# prepare embedding matrix
num_words = min(MAX_VOCAB_SIZE, len(word2idx) + 1)
embedding_matrix = np.zeros((num_words, EMBEDDING_DIM))
for word, i in word2idx.items():
if i < MAX_VOCAB_SIZE:
embedding_vector = word2vec.get(word)
if embedding_vector is not None:
# words not found in embedding index will be all zeros
embedding_matrix[i] = embedding_vector
# load pre-trained word embeddings into an Embedding layer
# note that we set trainable = False
embedding_layer = Embedding(
num_words,
EMBEDDING_DIM,
weights=[embedding_matrix],
input_length=MAX_SEQUENCE_LENGTH,
trainable=False
)
input_ = Input(shape=(MAX_SEQUENCE_LENGTH,))
x = embedding_layer(input_)
x = Conv1D(128, 3, activation='relu')(x)
x = MaxPooling1D(3)(x)
x = Conv1D(128, 3, activation='relu')(x)
x = MaxPooling1D(3)(x)
x = Conv1D(128, 3, activation='relu')(x)
x = GlobalMaxPooling1D()(x)
x = Dense(128, activation='relu')(x)
output = Dense(len(possible_labels), activation='sigmoid')(x)
model = Model(input_, output)
model.compile(
loss='binary_crossentropy',
optimizer='rmsprop',
metrics=['accuracy']
)
r = model.fit(
data, targets, batch_size=BATCH_SIZE, epochs=EPOCHS,
validation_split=VALIDATION_SPLIT
)
# plot some data
plt.plot(r.history['loss'], label='loss')
plt.plot(r.history['val_loss'], label='val_loss')
plt.legend()
plt.show()
# accuracies
plt.plot(r.history['acc'], label='acc')
plt.plot(r.history['val_acc'], label='val_acc')
plt.legend()
plt.show()
p = model.predict(data)
aucs = []
for j in range(6):
auc = roc_auc_score(targets[:,j], p[:,j])
aucs.append(auc)
print(np.mean(aucs))
```
| github_jupyter |
# Visualizing time-resolved LFP-Spiking Analysis of CRCNS PFC2 Dataset
```
%load_ext autoreload
%autoreload 2
import sys
sys.path.append('/Users/rdgao/Documents/code/research/spectralCV/')
sys.path.append('/Users/rdgao/Documents/code/research/neurodsp/')
%matplotlib inline
# imports
import numpy as np
import scipy as sp
import matplotlib.pyplot as plt
import neurodsp as ndsp
from scv_funcs import lfpca
from scv_funcs import utils
import pandas as pd
import pandas.plotting as pdplt
CKEYS = plt.rcParams['axes.prop_cycle'].by_key()['color']
font = {'family' : 'arial',
'weight' : 'regular',
'size' : 13}
import matplotlib
matplotlib.rc('font', **font)
def compute_pairwise_corr(spkstats, lfpstats, sub_inds, corr_type='spearman', log_power=False, plot_matrix=False):
fr,isi_cv = spkstats[sub_inds,0], spkstats[sub_inds,2]
if log_power:
pwr = lfpstats[sub_inds,:,0]
else:
pwr = np.log10(lfpstats[sub_inds,:,0])
scv = lfpstats[sub_inds,:,1]
ks_stat = lfpstats[sub_inds,:,2]
num_freqs = pwr.shape[1]
rho = np.zeros((5,5,num_freqs))
for freq in range(num_freqs):
df = pd.DataFrame(np.array([fr, isi_cv, pwr[:,freq], scv[:,freq], ks_stat[:,freq]]).T, columns=labels)
corr = df.corr(method='spearman').as_matrix()
rho[:,:,freq] = corr
if plot_matrix:
axes = pdplt.scatter_matrix(df, alpha=0.2, figsize=(7, 7), diagonal='kde');
for i, j in zip(*plt.np.triu_indices_from(axes, k=1)):
# label with correlation coefficient
axes[i, j].annotate("%.3f" %corr[i,j], (0.8, 0.8), xycoords='axes fraction', ha='center', va='center')
return rho
def plot_corrs(corr_mat,f1,f2,labels,freq=None,YL=(-0.5,0.5)):
if freq is None:
plt.plot(corr_mat[f1,f2,:,:].T)
else:
plt.plot(corr_mat[f1,f2,freq,:].T)
num_chan = corr_mat.shape[-1]
re_div = num_chan/3.*2.-0.5 # region division
plt.xticks([re_div], ["PFC | CA1"])
plt.ylim(YL)
plt.plot([re_div,re_div], plt.ylim(), 'k--', alpha=0.5)
plt.plot(plt.xlim(), [0,0], 'r--', alpha=0.5)
plt.title(labels[f1]+' : '+labels[f2])
# load LFP data
basefolder ='/Users/rdgao/Documents/data/CRCNS/pfc2/EE.049/EE.049'
lfp_file = basefolder+ '_LFP.mat'
lfp_struct = sp.io.loadmat(lfp_file, squeeze_me=True)
data = lfp_struct['lfp'][:96,:]
t = lfp_struct['t']
fs = lfp_struct['fs'] #1250
# load spike data
spike_file = basefolder + '_Behavior.mat'
spike_struct = sp.io.loadmat(spike_file, squeeze_me=True)
cell_info = spike_struct['spikeph']
spike_ind = spike_struct['spikeind']
spike_t = spike_struct['spiket'] # 20kHz
# organize spikes into cells, shanks, and areas
# spkt_c, spkt_sh, spkt_re = organize_spikes(spike_t, spike_ind, cell_info)
# _, cells_per_shank = np.unique(cell_info[:,1], return_counts=True)
nperseg= 1250
noverlap= int(nperseg/2)
N_skip=25
filt_bands = [(4,12), (24,40), (150,200)]
#filt_bands = [(0,4),(6,12),(14,20),(30,40),(50,70),(70,100),(150,200)]
```
# Load results
```
data_loaded = np.load('../results/pfc2/EE049_filt.npz')
t_win = data_loaded['t_win']
spkstats_cell = data_loaded['spkstats_cell']
spkstats_shank = data_loaded['spkstats_shank']
spkstats_re = data_loaded['spkstats_re']
lfpstats_rw = data_loaded['lfpstats_rw']
lfpstats_summary = data_loaded['lfpstats_summary']
sigpower_all = data_loaded['sigpower_all']
filt_bands = data_loaded['filt_bands']
# get task and rest indices
rest_inds = []
task_inds = []
for i in range(spike_struct['SessionNP'].shape[0]):
rest_inds.append(np.where(np.logical_and(t_win >= spike_struct['SessionNP'][i,0],t_win <= spike_struct['SessionNP'][i,1]))[0])
task_inds.append(np.where(np.logical_and(t_win >= spike_struct['SessionNP'][i,1],t_win <= spike_struct['SessionNP'][i,2]))[0])
rest_inds = np.concatenate(rest_inds)
task_inds = np.concatenate(task_inds)
```
# Visualizing results
### Computing correlations
```
corr_labels = ['Per Shank', 'All PFC Cells', 'All CA1 Cells']
labels = ['Cell_FR','Cell_ISICV','LFP_PWR','LFP_SCV', 'LFP_KS']
sub_inds = np.arange(len(t_win))[2:-2:] # all indices
sub_inds = task_inds
shank_corr = []
pfc_corr = []
ca1_corr = []
for sh in range(spkstats_shank.shape[0]):
# 2 LFP channels per shank
for chan in range(2):
shank_corr.append(compute_pairwise_corr(spkstats_shank[sh], lfpstats_rw[sh*2+chan], sub_inds))
pfc_corr.append(compute_pairwise_corr(spkstats_re[0], lfpstats_rw[sh*2+chan], sub_inds))
ca1_corr.append(compute_pairwise_corr(spkstats_re[1], lfpstats_rw[sh*2+chan], sub_inds))
# features, features, freq, shank
shank_corr = np.stack(shank_corr, axis=3)
pfc_corr = np.stack(pfc_corr, axis=3)
ca1_corr = np.stack(ca1_corr, axis=3)
all_corr = [shank_corr,pfc_corr,ca1_corr]
plt.figure(figsize=(18,9))
for freq in range(lfpstats_rw.shape[2]):
for ind in range(3):
rho, pv = utils.corrcoefp(lfpstats_rw[:,sub_inds,freq,ind])
plt.subplot(3,7,ind*7+freq+1)
utils.corr_plot(rho,bounds=[-1,1])
plt.xticks([])
plt.yticks([])
plt.title(labels[ind+2]+'%i-%iHz'%(filt_bands[freq][0],filt_bands[freq][1]) )
plt.tight_layout()
# FR-ISI
plt.figure(figsize=(15,8))
for ind in range(3):
plt.subplot(2,3,ind+1)
plot_corrs(all_corr[ind],f1=0,f2=1,labels=labels, YL=(-0.5, 0.8))
plt.ylabel(corr_labels[ind])
plt.legend(filt_bands)
plt.subplot(2,3,4)
plot_corrs(all_corr[0],f1=2,f2=3,labels=labels, YL=(-1, 0.8))
plt.subplot(2,3,5)
plot_corrs(all_corr[0],f1=2,f2=4,labels=labels, YL=(-1, 0.8))
plt.subplot(2,3,6)
plot_corrs(all_corr[0],f1=3,f2=4,labels=labels, YL=(-1, 0.8))
plt.tight_layout()
plot_freq = [0,1,2,3,4,5,6]
plt.figure(figsize=(18,10))
for corr_ind in range(3):
for i,f1 in enumerate([0,1]):
for j,f2 in enumerate([2,3,4]):
plt.subplot(3,6,i*3+j+1+corr_ind*6)
plot_corrs(all_corr[corr_ind],f1,f2,labels, freq=plot_freq)
plt.yticks([])
plt.subplot(3,6,1+corr_ind*6)
plt.ylabel(corr_labels[corr_ind], fontsize=18)
plt.yticks([-0.5,0.5])
# plt.legend(filt_bands[plot_freq])
plt.tight_layout()
```
| github_jupyter |
# Look-Aside Cache for MongoDB
### This is a sample notebook for using Aerospike as a read/look-aside cache
- This notebook demonstrates the use of Aerospike as a cache using Mongo as another primary datastore
- It is required to run Mongo as a separte container using `docker run --name some-mongo -d mongo:latest`
To test: Run the `get_data(key, value)` method once - to fetch from Mongo and populate Aerospike
Another run will fetch the data from Aerospike cache
#### Ensure that the Aerospike Database is running
```
!asd >& /dev/null
!pgrep -x asd >/dev/null && echo "Aerospike database is running!" || echo "**Aerospike database is not running!**"
```
#### Import all dependencies
```
import aerospike
import pymongo
from pymongo import MongoClient
import sys
```
## Configure the clients
The configuration is for
- Aerospike database running on port 3000 of localhost (IP 127.0.0.1) which is the default.
- Mongo running in a separate container whose IP can be found by `docker inspect <containerid> | grep -i ipaddress`
Modify config if your environment is different (Aerospike database running on a different host or different port).
```
# Define a few constants
AEROSPIKE_HOST = "0.0.0.0"
AEROSPIKE_PORT = 3000
AEROSPIKE_NAMESPACE = "test"
AEROSPIKE_SET = "demo"
MONGO_HOST = "172.17.0.3"
MONGO_PORT = 27017
MONGO_DB = "test-database"
MONGO_COLLECTION = "test-collection"
#Aerospike configuration
aero_config = {
'hosts': [ (AEROSPIKE_HOST, AEROSPIKE_PORT) ]
}
try:
aero_client = aerospike.client(aero_config).connect()
except:
print("Failed to connect to the cluster with", aero_config['hosts'])
sys.exit(1)
print("Connected to Aerospike")
#Mongo configuration
try:
mongo_client = MongoClient(MONGO_HOST, MONGO_PORT)
print("Connected to Mongo")
except:
print("Failed to connect to Mongo")
sys.exit(1)
```
#### Store data in Mongo and clear the keys in Aerospike if any
```
db = mongo_client[MONGO_DB]
collection = db[MONGO_COLLECTION]
def store_data(data_id, data):
m_data = {data_id: data}
collection.drop()
aero_key = ('test', 'demo', data_id)
#aero_client.remove(aero_key)
post_id = collection.insert_one(m_data)
store_data("key", "value")
```
#### Fetch the data. In this instance we are using a simple key value pair.
If the data exists in the cache it is returned, if not data is read from Mongo, put in the cache and then returned
```
def get_data(data_id, data):
aero_key = (AEROSPIKE_NAMESPACE, AEROSPIKE_SET, data_id)
#aero_client.remove(aero_key)
data_check = aero_client.exists(aero_key)
if data_check[1]:
(key, metadata, record) = aero_client.get(aero_key)
print("Data retrieved from Aerospike cache")
print("Record::: {} {}".format(data_id, record['value']))
else:
mongo_data = collection.find_one({data_id: data})
print("Data not present in Aerospike cache, retrieved from mongo {}".format(mongo_data))
aero_client.put(aero_key, {'value': mongo_data[data_id]})
get_data("key", "value")
```
| github_jupyter |
# Freedom of Information Law Opinions
-----
## Table of Contents
[Download the Data](#Data)
- [Data Preprocessing](#Prepare-Data-Structures)
[Non-Negative Matrix Factorization](#Non-Negative-Matrix-Factorization)
- [Parameter Tuning](#Tuning-the-Parameters)
- [Topic Generation](#Topics)
[Investigate Topics](#Evaluating-the-Topic-Determinations)
- [Data Visualization](#Visualizing-Topics)
- [Noise Reduction](#Reducing-Noise)
[Topic and Document Fingerprints](#Topic-and-Document-Fingerprints)
- [Finding Similar Documents](#Find-Documents-with-Matching-Topic-Fingerprints)
## Topic Modeling
While the Committee sorts their important Opinions into major content topics, like "Abstention from Voting" or "Tape Recorders, Use of," there are other topics contained in those documents—topics that could be useful to lawyers or citizens or politicians to understand and group together. An informed researcher pouring over the documents could come up with a dozen topics within each document. In this sense, we can think of documents as a recipe with topics as its ingredients:
> **Advisory Opinion 8762 Recipe**
> 1 cup describing the Audit Committee
> 2 cups clarifying the Statute of Limitations
> 1.5 tablespoons quoting §260-a of the Education Law
> 1 teaspoon invoking *French v. Board of Education*
But there really aren't that many experts on New York State Freedom of Information Law Advisory Committee Opinions. And unfortunately, none of them are available for hundreds of hours of document classification. Instead, we can computationally infer these topics by looking at the kinds of words used in an Opinion and then comparing those word choices to the words used in all of the Opinions. By doing this over and over again, and applying different statistics to the results, one can generate latent topics, see how similar two Opinions are to one another, or determine how likely a new Opinion is to fit into one of our topics.
With enough data, we could determine how likely a given request is to get a favorable Opinion, or which court cases are most likely to help an appeal. But first, let's discover topics in the already available Opinions from the Committee.
------------
If you want to follow along and didn't run the webscraping notebook to generate the data, you can get the data by running the following Data code cell:
<div style="text-align: right"> { [Table of Contents](#Table-of-Contents) }</div>
## Data
```
import requests
import pickle
# Finding and defining your home directory
home_dir = !echo $HOME
home = home_dir[0] +'/'
#downloading the pickled data and writing it to a file in your home directory
url = 'http://cpanel.ischool.illinois.edu/~adryden3/FOILAdvisoryDecisiondataWithText.pickle'
r = requests.get(url, allow_redirects=True)
with open(home + 'FOILdata.pickle', 'wb') as f:
f.write(r.content)
#reading the data and unpickeling it into a variable
with open(home + 'FOILdata.pickle', 'rb') as g:
opinions = pickle.load(g)
#remove the data_dict related to this data's provenance
if 'data_dictionary' in opinions:
opinions.pop('data_dictionary')
#set up the notebook and required libraries
% matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings("ignore")
```
-----
<div style="text-align: right"> { [Table of Contents](#Table-of-Contents) }</div>
## Prepare Data Structures
The pickled file includes a clean-text version of the Advisory Opinions from the web scraping notebook. From these, we generate a list of Opinions and two indexes to help us interpret the results. The indexes represent the keywords or topics that Advisory Committee members applied to the Opinions.
-----
```
#create indexes to use in dataframes later
human_topics = []
human_topic_number = []
#generate a list of the plain-text opinions
#skip any opinions where the full text isn't available
opinions_list = []
number = -1
for key in opinions:
number+=1
human_topics.append(key)
if len(opinions[key][3])==0:
number-=1
human_topics.pop()
else:
for i in range(len(opinions[key][3])):
opinions_list.append(opinions[key][3][i])
human_topic_number.append(number)
```
-----
<div style="text-align: right"> { [Table of Contents](#Table-of-Contents) }</div>
### Generate TF:IDF
The first machine learning process we are going to use looks at documents as a collection of words, and we serve it that collection as a list of terms, or matrix, and their frequency. To account for the fact that some words are more common than others, the frequency statistic for a term in a document is increased the more times it is mentioned in the document, and it is decreased the more times it is mentioned in the corpus. This way, rare but meaningful terms aren't drowned out by more common ones.
Words that appear in more than 90% of the documents aren't considered at all, and common or functional words that don't independently encode a lot of information ('the', 'it', 'for') are also removed.
-----
```
# Use TD-IDF on opinions corpus
from sklearn.feature_extraction.text import TfidfVectorizer
cv = TfidfVectorizer(stop_words = 'english',
lowercase=True,
min_df=1,
max_df=0.9,
max_features=5000)
opinion_vector = cv.fit_transform(opinions_list)
```
-----
<div style="text-align: right"> { [Table of Contents](#Table-of-Contents) }</div>
## Non-Negative Matrix Factorization
Matrix factorization is like numerical factorization (the factors of 12 are the numbers that can be multiplied to equal 12: 2x6, 3x4). This algorithm takes the TF-IDF term matrices that we produced and yeilds the matrix factors, significantly reducing their dimensions in the process. This factoring has an inherent clustering effect because it computationally reduces documents to component relationships among their terms.
### Tuning the Parameters
Unsupervised learning techniques can do a remarkable amount of work, but they can't always do everything. Here, we must indicate how many topics to find in the corpus. This is one of the most important aspects of this type of data analysis and usually requires some degree of domain knowledge and some experimetation. Different domains will inherently have different topic ranges, but you don't want to ask for so many topics that each document is its own topic, nor do you want to get topics that are so generic that they don't reveal anything meaningful about documents and clusters. For this domain, I felt 60 topics was appropriate because the corpus is made up of very specific requests, and therefore has a lot of granular detail that can be explored. In addition, the tool this supports privileges returing relevant information with little noise.
-----
```
# Compute topics by using NMF
from sklearn.decomposition import NMF
#select number of topics
num_opinion_topics = 60
#fit the model to the TFIDF vectors
nmf_opinion = NMF(n_components = num_opinion_topics, max_iter = 1000).fit(opinion_vector)
# Function to compute and display the top topic terms
def get_topics(cv, model):
# Number of terms per topic to display
max_topics = 10
# Number of terms per topic to retain
max_labels = 5
topics = []
feature_names = cv.get_feature_names()
# Iterate through the matrix components
for idx, topic in enumerate(model.components_):
# First we sort the terms in descending order ([::-1])
# And then retiain only the top terms
top_topics_idx = topic.argsort()[::-1][:max_topics]
top_topics = [feature_names[jdx] for jdx in top_topics_idx]
# Now extract out the terms themselves and display
top_features = " ".join(top_topics)
print('Topic {0:2d}: {1}'.format(idx, top_features))
topics.append(", ".join(top_topics[:max_labels]))
return(topics)
```
<div style="text-align: right"> { [Table of Contents](#Table-of-Contents) }</div>
### Topics
Topics are described by their representative terms. Here we have our 60 topics and their top 10 terms. Scanning through this list is a great way to understand how well the algorithm is performing and to gain insights into other processesing that can improve results. For example, numbers and number digit combinations likely represent references to laws or codes, which will inspire us use Named Entity Recognition tecniques to link documents which reference the same regulations or court cases, as these multi-word references would get jumbled in the Bag-of-Words approach used by our topic model.
-----
```
#print the topics and their top 10 terms
nmf_opinion_topics = get_topics(cv, nmf_opinion)
```
-----
<div style="text-align: right"> { [Table of Contents](#Table-of-Contents) }</div>
### Understanding Topic Terms
Looking at a random selection of topics, we can quickly see that the algorithm has generated fairly coherent topics at a level of granularity appropriate for our purposes.
>Topic 0: duly quorum number majority persons officers meeting duty total power
>Topic 14: tape recording use devices recorders davidson unobtrusive detract deliberative court
>Topic 51: clerk minutes town verbatim accurate 30 account determine consist prepare
Now let's look at how the algorithm applied these topics to our documents and to the clusters tagged by humans. In order to do so, we need to normalize the distributions so that we can read them as percentages and preserve the data. We then combine and average the topic distributions for Opinions that had the same human applied topics.
Quickly examining a few documents and their topics compared to the human-provided topic will help us understand if the application will be successful. To do so, let's look at a document that was classified by our model as including topic 14.
-----
```
from sklearn.preprocessing import normalize
#transforma and normalize the data;
#using the l1-norm the document topic probabilites sum to unity
td_opinion = nmf_opinion.transform(opinion_vector)
td_opinion_norm = normalize(td_opinion, norm='l1', axis=1)
#generate a dataframe with the data and add a column with the index of the
#human applied topic
df_opinion = pd.DataFrame(td_opinion_norm, columns=nmf_opinion_topics).round(2)
df_opinion.fillna(value=0, inplace=True)
df_opinion['label'] = pd.Series(human_topic_number)
#Now we group the human labeled opinions together and average their topic distributions
df_opinion_labeled = df_opinion.groupby('label').mean().round(2)
df_opinion_labeled['Human Label'] = pd.Series(human_topics, dtype='category')
#summary of all of the records labeled Tape Recorder, Use of at Executive Session
display_series = pd.Series(df_opinion_labeled.iloc[195])
print("Human Category: ", display_series[60])
print("\nNMF Distribution of Topics: ")
#show the top 5 topics the model applied to the human topic
display_series[:60].sort_values(ascending=False)[:5]
```
---
<div style="text-align: right"> { [Table of Contents](#Table-of-Contents) }</div>
## Evaluating the Topic Determinations
So, the topic model looked at the Opinions classified by humans as "Tape Recorders, Use of at Executive Session" and determined that their number one topic was "tape, recording, use, . . ". Which is about as close as we could ask for. However, the topic model also tells us that these Opinions are also about records, education and confidentiality. To get an idea of how accurate the model is, let's take a look at one of those documents and highlight some key terms.
---
<div id="mainContent" class="alert alert-block alert-warning">
<p>
</p>
<!-- InstanceBeginEditable name="Content" -->
<p>
OML-AO-05384
<br/>
December 27, 2013
</p>
<p>
</p>
<p>
<u>
The staff of the Committee on Open Government is authorized to issue advisory opinions. The ensuing staff advisory opinion is based solely upon the facts presented in your correspondence.
</u>
</p>
<p>
Dear :
</p>
<p>
This is in response to your request for an advisory opinion regarding application of the Open Meetings Law to executive sessions of the Oppenheim-Ephratah-St. Johnsville Central School District, and in particular, a rule that would prohibit a board member from <strong style="color: red">tape recording</strong> discussions held in executive session. It is our opinion that a rule prohibiting one board member from recording discussions held in executive session without consent of the board would be reasonable.
</p>
<p>
Initially, this will confirm that many believe recordings of executive session discussions are not desirable because copies of such <strong style="color: green">records</strong> may be sought pursuant to FOIL, at which point the agency would have to determine whether the <strong style="color: green">records</strong> were required to be made available, in whole or in part. As you correctly point out, it has long been the opinion of this office that notes taken and recordings made during executive sessions are “<strong style="color: green">records</strong>” subject to the Freedom of Information Law, the contents of which would determine rights of access. Further, a <strong style="color: red">tape recording</strong> of an executive session may be subject to subpoena or discovery in the context of litigation. Disclosure in that kind of situation may place a public body at a disadvantage should litigation arise relative to a topic that has been appropriately discussed behind closed doors.
</p>
<p>
Although in our opinion they are not prohibited by statute, surreptitious recordings of executive sessions made by one school board member resulted in a decision from the Commissioner of <strong style="color: blue">Education</strong> essentially warning members that such behavior would result in removal from the board. See,
<u>
Application of Nett and Raby
</u>
(No. 15315, October 24, 2005).
</p>
<p>
As indicated, you “began <strong style="color: red">taping</strong> executive sessions to assure compliance with the laws governing executive session,” and it is your way of taking notes. In that regard, we are aware that on perhaps many occasions, discussions that are appropriate for executive session evolve into those that are not.
</p>
<p>
This will confirm that there is no statute that deals directly with the <strong style="color: red">taping</strong> of executive sessions. Several judicial decisions have dealt with the ability to use recording devices at open meetings, and although those decisions do not refer to the <strong style="color: red">taping</strong> of executive sessions, they are likely pertinent to the matter. Perhaps the leading decision concerning the use of <strong style="color: red">tape recorders</strong> at meetings, a unanimous decision of the Appellate Division, involved the invalidation of a resolution adopted by a board of <strong style="color: blue">education</strong> prohibiting the use of tape recorders at its meetings (
<u>
Mitchell v. Board of <strong style="color: blue">Education</strong> of Garden City School District
</u>
, 113 AD 2d 924 [1985]). In so holding, the Court stated that:
</p>
<blockquote>
<p>
“While <strong style="color: blue">Education</strong> Law sec. 1709(1) authorizes a board of <strong style="color: blue">education</strong> to adopt by-laws and rules for its government and operations, this authority is not unbridled. Irrational and unreasonable rules will not be sanctioned. Moreover, Public Officers Law sec. 107(1) specifically provides that 'the court shall have the power, in its discretion, upon good cause shown, to declare any action *** taken in violation of [the Open Meetings Law], void in whole or in part.' Because we find that a prohibition against the use of unobtrusive recording goal of a fully informed citizenry, we accordingly affirm the judgement annulling the resolution of the respondent board of <strong style="color: blue">education</strong>” (id. at 925).
</p>
</blockquote>
<p>
Authority to tape record meetings in accordance with the above decision is now set forth in the Open Meetings Law; however, §103(d)(i) pertains only to those meetings that are open to the public.
</p>
<p>
While there are no decisions that deal with the use of <strong style="color: red">tape recorders</strong> during executive sessions, we believe that the principle in determining that issue is the same as that stated above, i.e., that a board may establish reasonable rules governing the use of <strong style="color: red">tape recorders</strong> at executive sessions.
</p>
<p>
Unlike an open meeting, when comments are conveyed with the public present, an executive session is generally held in order that the public cannot be aware of the details of the deliberative process. When an issue focuses upon a particular individual, the rationale for permitting the holding of an executive session generally involves an intent to protect personal privacy, coupled with an intent to enable the members of a public body to express their opinions freely. As previously mentioned, <strong style="color: red">tape recording</strong> executive sessions may result in unforeseen and potentially damaging consequences.
</p>
<p>
In short, we are suggesting that <strong style="color: red">tape recording</strong> an executive session could potentially defeat the purpose of holding an executive session. Accordingly, it is our opinion that it would be reasonable for a board of <strong style="color: blue">education</strong>, based on its authority to adopt rules to govern its own proceedings conferred by §1709 of the <strong style="color: blue">Education</strong> Law, to prohibit a member from using a tape recorder at an executive session absent the consent of a majority of the board.
</p>
<p>
Should you have further questions, please contact me directly.
</p>
<p>
Sincerely,
<br/>
</p>
<p>
Camille S. Jobin-Davis
<br/>
Assistant Director
<br/>
CSJ:mm
<br/>
c: Board of <strong style="color: blue">Education</strong>
</p>
<!-- InstanceEndEditable -->
<!-- end #mainContent -->
<script>
document.write("OML-AO-"+DocName);
</script>
</div>
---
<div style="text-align: right"> { [Table of Contents](#Table-of-Contents) }</div>
It is easy to see at a glance that "education" and "records" are emportant contexts for this Opinion. And the algorithm accuratly identifies that "confidentiality" despite few vocabulary words directly about confidentiality. So the algorithm seems to have reproduced the essential qualities of the human determined topic while describing and quantifying other topics as well. But the model is still a little noisy. We only took the top 5 topics, but there were more, and those could weaken other signals and increase processing resources down the road, so we will clean them out later in the notebook.
---
---
<div style="text-align: right"> { [Table of Contents](#Table-of-Contents) }</div>
## Visualizing Topics
We can use a heatmap to visually explore the whole topic space and look for tendencies. There are a total of 507 human labels, and for each one the model developed a topic distribution. In this heatmap, the dark spots indicate where the model found predominantly one single topic for a human label. Had the model and the humans come up with the exact same results, there would be 507 dark purple dots. Inspecting this can give us a rough idea of how diffuse the topics are.
---
```
import seaborn as sns
df = df_opinion.drop('label', axis=1)
fig, ax = plt.subplots(1, 1, figsize = (20, 15))
hm = sns.heatmap(df_opinion_labeled.drop('Human Label', axis=1).transpose(), xticklabels=5)
hm.axes.set_title('Topics per Human Label', fontsize=50)
hm.axes.set_xlabel('Human Label Number', fontsize=40)
hm.axes.set_ylabel('NMF Topics', fontsize=40)
hm.axes.set_yticklabels(range(60))
sns.set(font_scale=1)
```
---
So far it looks like we have a good deal of agreement while still generating more nuance than a single lable. Because the data was loaded in alphebetical order by human topic, we should be able to confirm this tendency by looking at the topic distributions of each individual document in a heatmap.
---
```
df = df_opinion.drop('label', axis=1)
fig, ax = plt.subplots(1, 1, figsize = (20, 15))
hm = sns.heatmap(df, yticklabels=10)
hm.axes.set_title('Corpus Topic Map', fontsize=50)
hm.axes.set_xlabel('Topic Words', fontsize=40)
hm.axes.set_ylabel('Document Number', fontsize=40)
sns.set(font_scale=1)
```
<div style="text-align: right"> { [Table of Contents](#Table-of-Contents) }</div>
## Reducing Noise
---
Topics that account for less than 7% of a document don't contribute very much to our understanding, and they may reduce the signal form more important topics, so we will eliminate any values below .07. Our applications can take advantage of algorithms that optimally handle sparse data, so this cleaning will also imporove performance.
We can also use it as an opportunity to see measure the health of the model. If any documents lost more than 50% of their signal from this cleaning, it means that the model wasn't able to find a good latent topic for the Opinion. That could just be an anomalous Opinion, but if we see it happening a lot then it means we need to reevaluate the model.
---
```
#these are all of the opinions and their distribution over the latent topics
df_graph = pd.DataFrame(td_opinion_norm).round(2)
df_graph.fillna(value=0, inplace=True)
#to eliminate noise from the data, we drop anything less than .07 from the topic distribution
df_graph.mask(df_graph<.06, 0, inplace=True)
#to make sure we didn't have a lot of data that was primarily made up these noisy fragments
#we sum each opinion's remaining values and filter out any that are below 60
sum_series = df_graph.transpose().sum()
sum_series.where(sum_series<.6).dropna()
```
---
Only 1 was below the threshold we set, and only 6 were close to it. At this point we can inspect those documents, but no matter what we should save these results in the data cleaning log before we renormalize the values. After we apply l1 normalization, topics per Opinion will again sum to 1 with the noise eliminated. Now we can evaluate the concentration of topics by sorting the Opinions by topic, starting with the topics with the highest cumulative score in the corpus. To the degree that the topics are concentrated, this will cause dark clumps to appear in the heatmap.
---
```
df_graph = pd.DataFrame(normalize(df_graph, norm='l1', axis=1).round(2))
fig, ax = plt.subplots(1, 1, figsize = (20, 15))
hm = sns.heatmap(df_graph.sort_values([i for i in df_graph.sum().sort_values(ascending=False).index], ascending=False),
xticklabels=2, yticklabels=10)
hm.axes.set_title('Opinions Ordered by Corpus Topic Weight', fontsize=40)
hm.axes.set_xlabel('Topic Number', fontsize=40)
hm.axes.set_ylabel('Document Number', fontsize=40)
sns.set(font_scale=1)
```
---
We can further inspect a particular topic by sorting on that topic. This can help us see connections among differnt topics, and give us ideas about how to detect them automatically.
---
```
fig, ax = plt.subplots(1, 1, figsize = (20, 15))
hm = sns.heatmap(df_graph.sort_values([10], ascending=False)[:75],
xticklabels=2)
hm.axes.set_title('Top 75 Opinions for Topic 10', fontsize=30)
hm.axes.set_xlabel('Topic Number', fontsize=20)
hm.axes.set_ylabel('Document Number', fontsize=20)
sns.set(font_scale=1)
```
---
<div style="text-align: right"> { [Table of Contents](#Table-of-Contents) }</div>
## Topic and Document Fingerprints
We can now use these topic and document distributions to represent the documents and topics within a database. This allows us to conduct powerful searches or document comparsions quickly using vector arithmetic.
---
```
fig, (ax1, ax2, ax3) = plt.subplots(3,1, figsize = (20, 9))
cbar_ax = fig.add_axes([.91,.1,.015,.75])
plt.subplots_adjust(hspace = 2)
hm1 = sns.heatmap(df_graph.transpose()[:1],
xticklabels=10, ax=ax1, cbar_ax=cbar_ax)
hm1.axes.set_title('Topic Fingerprint (Distribution in Corpus)', fontsize=20)
hm1.axes.set_xlabel('Documents', fontsize=20)
hm1.axes.set_ylabel('Topic 0', fontsize=20)
sns.set(font_scale=1)
hm2 = sns.heatmap(df_graph.iloc[[50]],
xticklabels=2, ax=ax2,cbar_ax=cbar_ax)
hm2.axes.set_title('Document Fingerprint (Topic Distribution)', fontsize=20)
hm2.axes.set_xlabel('Topics', fontsize=20)
hm2.axes.set_ylabel('Document 50', fontsize=20)
sns.set(font_scale=1)
hm3 = sns.heatmap(df_graph.iloc[[50,930]],
xticklabels=2, ax=ax3,cbar_ax=cbar_ax)
hm3.axes.set_title('Compare Documents (Topic Distributions)', fontsize=20)
hm3.axes.set_xlabel('Topics', fontsize=20)
hm3.axes.set_ylabel('Document Number', fontsize=20)
sns.set(font_scale=1)
```
## Find Documents with Matching Topic Fingerprints
We can compare fingerprints using a technique called Cosine Similarity. Essentially, we measure the distance between two topic distributions as if they were lines passing trough the topic space. This function takes an Opinion and finds the Opinion in the corpus with the most similar topic distribution and displays them both.
```
from numpy import dot
from numpy.linalg import norm
def find_similar(df, index):
top_score = 0
match = 0
a = df.iloc[index]
for i in range(len(df)):
if i == index:
pass
else:
b = df.iloc[i]
cos_sim = dot(a, b)/(norm(a)*norm(b))
if cos_sim > top_score:
match = i
top_score = cos_sim
fig, (ax1,ax2) = plt.subplots(2,1, figsize = (20,6))
cbar_ax = fig.add_axes([.91,.1,.015,.75])
plt.subplots_adjust(hspace = .1)
hm1 = sns.heatmap(df.iloc[[index]], xticklabels = False, ax=ax1, cbar_ax = cbar_ax)
hm1.axes.set_title('Target + Match', fontsize=25)
hm1.axes.set_ylabel('Target Opinion', fontsize=15)
hm2 = sns.heatmap(df.iloc[[match]], xticklabels = 2, ax=ax2, cbar_ax = cbar_ax)
hm2.axes.set_xlabel('Topic Number', fontsize=20)
hm2.axes.set_ylabel('Match Opinion', fontsize=15)
find_similar(df_graph, 330)
```
| github_jupyter |
# Evaluating a machine learning model
> There are three ways to evaluate Scikit-Learn models/estimators.
1. Estimator `score` method
2. The `scoring` parameter
3. Problem-specific metric functions
```
# Standard imports
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
## 1. Evaluating a model with the score method
```
heart_disease = pd.read_csv("https://raw.githubusercontent.com/CongLiu-CN/zero-to-mastery-ml/master/data/heart-disease.csv")
heart_disease.head()
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
np.random.seed(2020)
X = heart_disease.drop("target", axis=1)
y = heart_disease["target"]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
clf = RandomForestClassifier()
clf.fit(X_train, y_train)
clf.score(X_train, y_train)
clf.score(X_test, y_test)
```
Let's do the same but for regression...
```
# Import Boston housing dataset
from sklearn.datasets import load_boston
boston = load_boston()
boston;
# Turn boston into a pandas dataframe
boston_df = pd.DataFrame(boston["data"], columns=boston["feature_names"])
boston_df["target"] = pd.Series(boston["target"])
boston_df.head()
from sklearn.ensemble import RandomForestRegressor
np.random.seed(2020)
# Create the data
X = boston_df.drop("target", axis=1)
y = boston_df["target"]
# Split into training and test sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
# Instantiate and fit model
model = RandomForestRegressor().fit(X_train, y_train)
model.score(X_test, y_test)
```
## 2. Evaluating a model using the `scoring` parameter
```
from sklearn.model_selection import cross_val_score
from sklearn.ensemble import RandomForestClassifier
np.random.seed(2020)
X = heart_disease.drop("target", axis=1)
y = heart_disease["target"]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
clf = RandomForestClassifier()
clf.fit(X_train, y_train);
clf.score(X_test, y_test)
cross_val_score(clf, X, y)
```
What does Cross-validation do under the hood?

The `sv`=5 by default, but we can change it, for exapmle, to 10:
```
cross_val_score(clf, X, y, cv=10)
np.random.seed(2020)
# Single training and test split score
clf_single_score = clf.score(X_test, y_test)
# Take the mean of 5-fold cross-validation score
clf_cross_val_score = np.mean(cross_val_score(clf, X, y))
# Compare the two
clf_single_score, clf_cross_val_score
```
Default scoring parameter of classifier = mean accuracy
`clf.score()`
```
# Scoring parameter set to None by default
cross_val_score(clf, X, y, cv=5, scoring=None)
```
So Why do we use cross-validation score?
To avoid getting lucky scores.
## 3. Classification model evaluation metrics
1. Accuracy
2. Area under ROC curve
3. Confusion matrix
4. Classification report
**3.1 Accuracy**
```
heart_disease.head()
from sklearn.model_selection import cross_val_score
from sklearn.ensemble import RandomForestClassifier
np.random.seed(2020)
X = heart_disease.drop("target", axis=1)
y = heart_disease["target"]
clf = RandomForestClassifier()
cross_val_score = cross_val_score(clf, X, y)
np.mean(cross_val_score)
print(f"Heart Disease Classifier Cross-Validated Accuracy: {np.mean(cross_val_score) *100:.2f}%")
```
**3.2 Area under the receiver operating characteristic curve (AUC/ROC)**
* Area under curve (AUC)
* ROC curve
ROC curves are a comparison of a model's true positive rate (tpr) versus a model's false positive rate (fpr).
* True positive = model predicts 1 when truth is 1
* False positive = model predicts 1 when truth is 0
* True negative = model predicts 0 when truth is 0
* False negative = model predicts 1 when truth is 1
```
# Split data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
from sklearn.metrics import roc_curve
# Fit the classifier
clf.fit(X_train, y_train)
# Make predictions with probabilities
y_probs = clf.predict_proba(X_test)
y_probs[:10]
y_probs_positive = y_probs[:, 1]
y_probs_positive[:10]
# Caculate fpr, tpr and thresholds
fpr, tpr, thresholds = roc_curve(y_test, y_probs_positive)
# Check the false positive rates
fpr
# Create a function for plotting ROC curves
def plot_roc_curve(fpr, tpr):
"""
Plots a ROC curve given the false positive rate (fpr)
and true positive rate (tpr) of a model.
"""
# Plot roc curve
plt.plot(fpr, tpr, color="orange", label="ROC")
# Plot line with no predictive power (baseline)
plt.plot([0, 1], [0, 1], color="darkblue", linestyle="--", label="Guessing")
# Customize the plot
plt.xlabel("False positive rate (fpr)")
plt.ylabel("True positive rate (tpr)")
plt.title("Receiver Operating Characteristic (ROC) Curve")
plt.legend()
plt.show()
plot_roc_curve(fpr, tpr)
from sklearn.metrics import roc_auc_score
roc_auc_score(y_test, y_probs_positive)
# Plot perfect ROC curve and AUC score
fpr, tpr, thresholds = roc_curve(y_test, y_test)
plot_roc_curve(fpr, tpr)
# Perfect AUC score
roc_auc_score(y_test, y_test)
```
**3.3 Confusion Matrix**
A confusion matrix is a quick way to compare the labels a model predicts and the actual labels it was supposed to predict.
In essence, giving you an idea of where the model is getting confused.
```
from sklearn.metrics import confusion_matrix
y_preds = clf.predict(X_test)
confusion_matrix(y_test, y_preds)
# Visualize confusion matrix with pd.crosstab()
pd.crosstab(y_test,
y_preds,
rownames=["Actual Label"],
colnames=["Predicted Labels"])
# How to install a conda package into the current environment from a Jupyter Notebook
import sys
!conda install --yes --prefix {sys.prefix} seaborn
# Make our confusion matrix more visual with Seaborn's heatmap()
import seaborn as sns
# Set the font scale
sns.set(font_scale=1.5)
# Create a confusion matrix
conf_mat = confusion_matrix(y_test, y_preds)
# Plot it using Seaborn
sns.heatmap(conf_mat);
def plot_conf_mat(conf_mat):
"""
Plot a confusion matrix using Seaborn's heatmap().
"""
fig, ax = plt.subplots(figsize=(3,3))
ax = sns.heatmap(conf_mat,
annot=True, # Annotate the boxes with conf_mat info
cbar=False)
plt.xlabel("True label")
plt.ylabel("Predicted label");
plot_conf_mat(conf_mat)
```
**3.4 Classification Report**
```
from sklearn.metrics import classification_report
print(classification_report(y_test, y_preds))
# Where precision and recall become valuable
disease_true = np.zeros(10000)
disease_true[0] = 1 # only one positive case
disease_preds = np.zeros(10000) # model predicts every case as 0
pd.DataFrame(classification_report(disease_true,
disease_preds,
output_dict=True))
```
To summarize classification metrics:
* **Accuracy** is a good measure to start with if all classes are balanced (e.g. same amount of samples which are labelled with 0 or 1).
* **Precision** and **recall** become more important when classes are imbalanced.
* If false positive predictions are worse than false negatives, aim for higher precision.
* If false negative predictions are worse than false positives, aim for higher recall.
* **F1-score** is a combination of precision and recall.
## 4. Regression model evaluation metrics
Model evaluation metrics documentation - https://scikit-learn.org/stable/modules/model_evaluation.html
1. R^2 (pronounced r-squared) or coefficient of determination.
2. Mean absolute error (MAE)
3. Mean squared error (MSE)
**4.1 R^2**
What R-squared does:
Compares your models predictions to the mean of the target. Values can range from negative infinity (a very pool model) to 1.
For example, if all your model does is predict the mean of the targets, its R^2 value would be 0. But if your model perfectly predicts a range of numbers, its R^2 values would be 1.
```
from sklearn.ensemble import RandomForestRegressor
np.random.seed(2020)
X = boston_df.drop("target", axis=1)
y = boston_df["target"]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
model = RandomForestRegressor()
model.fit(X_train, y_train);
model.score(X_test, y_test)
from sklearn.metrics import r2_score
# Fill an array with y_test mean
y_test_mean = np.full(len(y_test), y_test.mean())
y_test.mean()
r2_score(y_test, y_test_mean)
r2_score(y_test, y_test)
```
**4.2 Mean absolute error**
MAE is the average of the absolute differences between predictions and actual values.
It gives an idea of how wrong your model's predictions are.
```
# Mean absolute error
from sklearn.metrics import mean_absolute_error
y_preds = model.predict(X_test)
mae = mean_absolute_error(y_test, y_preds)
mae
df = pd.DataFrame(data={"actual values": y_test,
"predicted values": y_preds})
df["differences"] = df["predicted values"] - df["actual values"]
df
```
**4.3 Mean squared error (MSE)**
```
# Mean squared error
from sklearn.metrics import mean_squared_error
y_preds = model.predict(X_test)
mse = mean_squared_error(y_test, y_preds)
mse
# Calculate MSE by hand
squared = np.square(df["differences"])
squared.mean()
```
**4.4 Finally, using the `scoring` parameter**
```
from sklearn.model_selection import cross_val_score
from sklearn.ensemble import RandomForestClassifier
np.random.seed(2020)
X = heart_disease.drop("target", axis=1)
y = heart_disease["target"]
clf = RandomForestClassifier()
np.random.seed(2020)
cv_acc = cross_val_score(clf, X, y)
cv_acc
# Cross-validated accuracy
print(f'The cross-validated accuracy is: {np.mean(cv_acc)*100:.2f}%')
np.random.seed(2020)
cv_acc = cross_val_score(clf, X, y, scoring="accuracy")
print(f'The cross-validated accuracy is: {np.mean(cv_acc)*100:.2f}%')
# Precision
cv_precision = cross_val_score(clf, X, y, scoring="precision")
np.mean(cv_precision)
# Recall
cv_recall = cross_val_score(clf, X, y, scoring="recall")
np.mean(cv_recall)
cv_f1 = cross_val_score(clf, X, y, scoring="f1")
np.mean(cv_f1)
```
How about the regression model?
```
from sklearn.model_selection import cross_val_score
from sklearn.ensemble import RandomForestRegressor
np.random.seed(2020)
X = boston_df.drop("target", axis=1)
y = boston_df["target"]
model = RandomForestRegressor()
np.random.seed(2020)
cv_r2 = cross_val_score(model, X, y, scoring=None)
cv_r2
np.random.seed(2020)
cv_r2 = cross_val_score(model, X, y, scoring="r2")
cv_r2
# Mean absolute error
cv_mae = cross_val_score(model, X, y, scoring="neg_mean_absolute_error")
cv_mae
# Mean sqaured error
cv_mse = cross_val_score(model, X, y, scoring="neg_mean_squared_error")
cv_mse
```
## Using different evaluation metrics as Scikit-Learn functions
**Classification evaluation functions**
```
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
np.random.seed(2020)
X = heart_disease.drop("target", axis=1)
y = heart_disease["target"]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
clf = RandomForestClassifier()
clf.fit(X_train, y_train)
# Make some predictions
y_preds = clf.predict(X_test)
# Evaluate the classifier
print("Classifier metrics on the test set")
print(f"Accuracy: {accuracy_score(y_test, y_preds)*100:.2f}%")
print(f"Precision: {precision_score(y_test, y_preds)*100:.2f}%")
print(f"Recall: {recall_score(y_test, y_preds)*100:.2f}%")
print(f"F1: {f1_score(y_test, y_preds)*100:.2f}%")
```
**Regression evaluation functions**
```
from sklearn.metrics import r2_score, mean_absolute_error, mean_squared_error
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import train_test_split
np.random.seed(2020)
X = boston_df.drop("target", axis=1)
y = boston_df["target"]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
model = RandomForestRegressor()
model.fit(X_train, y_train)
# Make predictions using our regression model
y_preds = model.predict(X_test)
# Evaluate the regression model
print("Regression model metrics on the test set")
print(f"R^2: {r2_score(y_test, y_preds)}")
print(f"MAE: {mean_absolute_error(y_test, y_preds)}")
print(f"MSE: {mean_squared_error(y_test, y_preds)}")
```
| github_jupyter |
```
import os
import sys
import logging
import json
import tensorflow_model_analysis as tfma
import tfx
from tfx.proto import example_gen_pb2, transform_pb2, trainer_pb2
from tfx.orchestration import pipeline, data_types
from tfx.dsl.components.base import executor_spec
from tfx.components.trainer import executor as trainer_executor
from tfx.extensions.google_cloud_ai_platform.trainer import (
executor as ai_platform_trainer_executor,
)
from tfx.extensions.google_cloud_big_query.example_gen.component import (
BigQueryExampleGen,
)
from tfx.components import StatisticsGen
from tfx.components import (
StatisticsGen,
ExampleValidator,
Transform,
Trainer,
Evaluator,
Pusher,
)
from tfx.dsl.components.common.importer import Importer
from tfx.dsl.components.common.resolver import Resolver
from tfx.dsl.experimental import latest_artifacts_resolver
from tfx.dsl.experimental import latest_blessed_model_resolver
from ml_metadata.proto import metadata_store_pb2
from src.tfx_pipelines import config
from src.tfx_pipelines import components as custom_components
from src.common import features, datasource_utils
#RAW_SCHEMA_DIR = "src/raw_schema"
#TRANSFORM_MODULE_FILE = "src/preprocessing/transformations.py"
#TRAIN_MODULE_FILE = "src/model_training/runner.py"
def create_pipeline(
pipeline_root: str,
num_epochs: data_types.RuntimeParameter,
batch_size: data_types.RuntimeParameter,
learning_rate: data_types.RuntimeParameter,
hidden_units: data_types.RuntimeParameter,
metadata_connection_config: metadata_store_pb2.ConnectionConfig = None,
):
local_executor_spec = executor_spec.ExecutorClassSpec(
trainer_executor.GenericExecutor
)
caip_executor_spec = executor_spec.ExecutorClassSpec(
ai_platform_trainer_executor.GenericExecutor
)
# Hyperparameter generation.
hyperparams_gen = custom_components.hyperparameters_gen(
num_epochs=num_epochs,
batch_size=batch_size,
learning_rate=learning_rate,
hidden_units=hidden_units,
).with_id("HyperparamsGen")
# Get train source query.
train_sql_query = datasource_utils.get_training_source_query(
config.PROJECT,
config.REGION,
config.DATASET_DISPLAY_NAME,
ml_use="UNASSIGNED",
limit=int(config.TRAIN_LIMIT),
)
train_output_config = example_gen_pb2.Output(
split_config=example_gen_pb2.SplitConfig(
splits=[
example_gen_pb2.SplitConfig.Split(
name="train", hash_buckets=int(config.NUM_TRAIN_SPLITS)
),
example_gen_pb2.SplitConfig.Split(
name="eval", hash_buckets=int(config.NUM_EVAL_SPLITS)
),
]
)
)
# Train example generation.
train_example_gen = BigQueryExampleGen(
query=train_sql_query,
output_config=train_output_config,
).with_id("TrainDataGen")
# Get test source query.
test_sql_query = datasource_utils.get_training_source_query(
config.PROJECT,
config.REGION,
config.DATASET_DISPLAY_NAME,
ml_use="TEST",
limit=int(config.TEST_LIMIT),
)
test_output_config = example_gen_pb2.Output(
split_config=example_gen_pb2.SplitConfig(
splits=[
example_gen_pb2.SplitConfig.Split(name="test", hash_buckets=1),
]
)
)
# Test example generation.
test_example_gen = BigQueryExampleGen(
query=test_sql_query,
output_config=test_output_config,
).with_id("TestDataGen")
# Schema importer.
schema_importer = Importer(
source_uri=RAW_SCHEMA_DIR,
artifact_type=tfx.types.standard_artifacts.Schema,
).with_id("SchemaImporter")
# Statistics generation.
#statistics_gen = StatisticsGen(examples=train_example_gen.outputs['examples']).with_id("StatisticsGen")
statistics_gen = tfx.components.StatisticsGen(examples=train_example_gen.outputs['examples']).with_id("StatisticsGen")
# Example validation.
example_validator = ExampleValidator(
#statistics=statistics_gen.outputs.statistics,
statistics=statistics_gen.outputs['statistics'],
schema=schema_importer.outputs['result'],
).with_id("ExampleValidator")
# Data transformation.
transform = Transform(
examples=train_example_gen.outputs['examples'],
schema=schema_importer.outputs['result'],
module_file=TRANSFORM_MODULE_FILE,
splits_config=transform_pb2.SplitsConfig(
analyze=["train"], transform=["train", "eval"]
),
).with_id("DataTransformer")
# Add dependency from example_validator to transform.
transform.add_upstream_node(example_validator)
# Get the latest model to warmstart
warmstart_model_resolver = Resolver(
strategy_class=latest_artifacts_resolver.LatestArtifactsResolver,
latest_model=tfx.types.Channel(type=tfx.types.standard_artifacts.Model),
).with_id("WarmstartModelResolver")
# Model training.
trainer = Trainer(
custom_executor_spec=local_executor_spec
if config.TRAINING_RUNNER == "local"
else caip_executor_spec,
module_file=TRAIN_MODULE_FILE,
transformed_examples=transform.outputs['transformed_examples'],
schema=schema_importer.outputs['result'],
# base_model=warmstart_model_resolver.outputs.latest_model,
transform_graph=transform.outputs['transform_graph'],
train_args=trainer_pb2.TrainArgs(num_steps=0),
eval_args=trainer_pb2.EvalArgs(num_steps=None),
hyperparameters=hyperparams_gen.outputs['hyperparameters'],
).with_id("ModelTrainer")
# Get the latest blessed model (baseline) for model validation.
baseline_model_resolver = Resolver(
strategy_class=latest_blessed_model_resolver.LatestBlessedModelResolver,
model=tfx.types.Channel(type=tfx.types.standard_artifacts.Model),
model_blessing=tfx.types.Channel(
type=tfx.types.standard_artifacts.ModelBlessing
),
).with_id("BaselineModelResolver")
# Prepare evaluation config.
eval_config = tfma.EvalConfig(
model_specs=[
tfma.ModelSpec(
signature_name="serving_tf_example",
label_key=features.TARGET_FEATURE_NAME,
prediction_key="probabilities",
)
],
slicing_specs=[
tfma.SlicingSpec(),
],
metrics_specs=[
tfma.MetricsSpec(
metrics=[
tfma.MetricConfig(class_name="ExampleCount"),
tfma.MetricConfig(
class_name="BinaryAccuracy",
threshold=tfma.MetricThreshold(
value_threshold=tfma.GenericValueThreshold(
lower_bound={"value": float(config.ACCURACY_THRESHOLD)}
),
# Change threshold will be ignored if there is no
# baseline model resolved from MLMD (first run).
change_threshold=tfma.GenericChangeThreshold(
direction=tfma.MetricDirection.HIGHER_IS_BETTER,
absolute={"value": -1e-10},
),
),
),
]
)
],
)
# Model evaluation.
evaluator = Evaluator(
examples=test_example_gen.outputs['examples'],
example_splits=["test"],
model=trainer.outputs['model'],
# baseline_model=baseline_model_resolver.outputs.model,
eval_config=eval_config,
schema=schema_importer.outputs['result'],
).with_id("ModelEvaluator")
exported_model_location = os.path.join(
config.MODEL_REGISTRY_URI, config.MODEL_DISPLAY_NAME
)
push_destination = tfx.proto.pusher_pb2.PushDestination(
filesystem=tfx.proto.pusher_pb2.PushDestination.Filesystem(
base_directory=exported_model_location
)
)
# Push custom model to model registry.
pusher = Pusher(
model=trainer.outputs['model'],
model_blessing=evaluator.outputs['blessing'],
push_destination=push_destination,
).with_id("ModelPusher")
# Upload custom trained model to Vertex AI.
explanation_config = json.dumps(features.generate_explanation_config())
vertex_model_uploader = custom_components.vertex_model_uploader(
project=config.PROJECT,
region=config.REGION,
model_display_name=config.MODEL_DISPLAY_NAME,
pushed_model_location=exported_model_location,
serving_image_uri=config.SERVING_IMAGE_URI,
explanation_config=explanation_config,
).with_id("VertexUploader")
pipeline_components = [
hyperparams_gen,
train_example_gen,
test_example_gen,
statistics_gen,
schema_importer,
example_validator,
transform,
# warmstart_model_resolver,
trainer,
# baseline_model_resolver,
evaluator,
pusher,
]
if int(config.UPLOAD_MODEL):
pipeline_components.append(vertex_model_uploader)
# Add dependency from pusher to aip_model_uploader.
vertex_model_uploader.add_upstream_node(pusher)
logging.info(
f"Pipeline components: {[component.id for component in pipeline_components]}"
)
beam_pipeline_args = config.BEAM_DIRECT_PIPELINE_ARGS
if config.BEAM_RUNNER == "DataflowRunner":
beam_pipeline_args = config.BEAM_DATAFLOW_PIPELINE_ARGS
logging.info(f"Beam pipeline args: {beam_pipeline_args}")
return pipeline.Pipeline(
pipeline_name=config.PIPELINE_NAME,
pipeline_root=pipeline_root,
components=pipeline_components,
beam_pipeline_args=beam_pipeline_args,
metadata_connection_config=metadata_connection_config,
enable_cache=int(config.ENABLE_CACHE),
)
import os
from kfp.v2.google.client import AIPlatformClient
from tfx.orchestration import data_types
from tfx.orchestration.kubeflow.v2 import kubeflow_v2_dag_runner
#from src.tfx_pipelines import config, training_pipeline, prediction_pipeline
#from src.model_training import defaults
def compile_training_pipeline1(pipeline_definition_file):
pipeline_root = os.path.join(
config.ARTIFACT_STORE_URI,
config.PIPELINE_NAME,
)
managed_pipeline = training_pipeline.create_pipeline(
pipeline_root=pipeline_root,
num_epochs=data_types.RuntimeParameter(
name="num_epochs",
default=defaults.NUM_EPOCHS,
ptype=int,
),
batch_size=data_types.RuntimeParameter(
name="batch_size",
default=defaults.BATCH_SIZE,
ptype=int,
),
learning_rate=data_types.RuntimeParameter(
name="learning_rate",
default=defaults.LEARNING_RATE,
ptype=float,
),
hidden_units=data_types.RuntimeParameter(
name="hidden_units",
default=",".join(str(u) for u in defaults.HIDDEN_UNITS),
ptype=str,
),
)
runner = kubeflow_v2_dag_runner.KubeflowV2DagRunner(
config=kubeflow_v2_dag_runner.KubeflowV2DagRunnerConfig(
default_image=config.TFX_IMAGE_URI
),
output_filename=pipeline_definition_file,
)
return runner.run(managed_pipeline, write_out=True)
def compile_prediction_pipeline(pipeline_definition_file):
pipeline_root = os.path.join(
config.ARTIFACT_STORE_URI,
config.PIPELINE_NAME,
)
managed_pipeline = prediction_pipeline.create_pipeline(
pipeline_root=pipeline_root,
)
runner = kubeflow_v2_dag_runner.KubeflowV2DagRunner(
config=kubeflow_v2_dag_runner.KubeflowV2DagRunnerConfig(
default_image=config.TFX_IMAGE_URI
),
output_filename=pipeline_definition_file,
)
return runner.run(managed_pipeline, write_out=True)
def submit_pipeline(pipeline_definition_file):
pipeline_client = AIPlatformClient(project_id=config.PROJECT, region=config.REGION)
pipeline_client.create_run_from_job_spec(pipeline_definition_file)
#from src.tfx_pipelines import runner
pipeline_definition_file = f'{config.PIPELINE_NAME}.json'
print(pipeline_definition_file)
#pipeline_definition = compile_training_pipeline(pipeline_definition_file)
def compile_training_pipeline12(pipeline_definition_file):
print("ravi")
pipeline_root = os.path.join(
config.ARTIFACT_STORE_URI,
config.PIPELINE_NAME,
)
managed_pipeline = create_pipeline(
pipeline_root=pipeline_root,
num_epochs=data_types.RuntimeParameter(
name="num_epochs",
default=defaults.NUM_EPOCHS,
ptype=int,
),
batch_size=data_types.RuntimeParameter(
name="batch_size",
default=defaults.BATCH_SIZE,
ptype=int,
),
learning_rate=data_types.RuntimeParameter(
name="learning_rate",
default=defaults.LEARNING_RATE,
ptype=float,
),
hidden_units=data_types.RuntimeParameter(
name="hidden_units",
default=",".join(str(u) for u in defaults.HIDDEN_UNITS),
ptype=str,
),
)
runner = kubeflow_v2_dag_runner.KubeflowV2DagRunner(
config=kubeflow_v2_dag_runner.KubeflowV2DagRunnerConfig(
default_image=config.TFX_IMAGE_URI
),
output_filename=pipeline_definition_file,
)
return runner.run(managed_pipeline, write_out=True)
pipeline_definition=compile_training_pipeline12(pipeline_definition_file)
pipeline_definition['pipelineSpec']['deploymentSpec']['executors']['ModelPusher_executor']['container']['image']='gcr.io/aiops-industrialization/chicago-taxi-tips:latest'
pipeline_definition['pipelineSpec']['deploymentSpec']['executors']['ModelPusher_executor']['container']['image']
pipeline_definition_file
#PIPELINES_STORE = f"gs://{BUCKET}/{DATASET_DISPLAY_NAME}/compiled_pipelines/"
PIPELINES_STORE='gs://aiops-industrialization-bucket-ravi/chicago-taxi-tips/compiled_pipelines/'
!gsutil cp {pipeline_definition_file} {PIPELINES_STORE}
PROJECT = 'aiops-industrialization' # Change to your project id.
REGION = 'us-central1' # Change to your region.
BUCKET = 'aiops-industrialization-bucket-ravi' # Change to your bucket name.
SERVICE_ACCOUNT = "175728527123-compute@developer.gserviceaccount.com"
from kfp.v2.google.client import AIPlatformClient
pipeline_client = AIPlatformClient(
project_id=PROJECT, region=REGION)
job = pipeline_client.create_run_from_job_spec(
job_spec_path=pipeline_definition_file,
parameter_values={
'learning_rate': 0.003,
'batch_size': 512,
'hidden_units': '128,128',
'num_epochs': 30,
}
)
```
| github_jupyter |
# Result Explorer Notebook for ccsn Project
## Overview
This notebook allows you to explore the results of `ccsn` model one-zone nucleosynthesis calculations performed by the Model-runner. Results can be organized in different directories (folders) managed by the Model-Runner.
The results are lists of how much of each isotope was produced or destroyed in the ccsn model calculation.
If things don't work make sure `Python 3` is displayed in the upper right corner of this Tab. If not go there and select it, and then try again.
## Instructions Step 1: Load Modules
Execute the following cell. This will load the modules for graphing and analysing (we are using the [NuGridPy](https://nugrid.github.io/NuGridPy/) Python package).
```
%pylab ipympl
from nugridpy import ppn
from nugridpy import utils
from nugridpy import utils as ut
from nugridpy import ascii_table as at
#### section to include at the beginnning of your notebook
#### that will suppress output of unnecessary information
import os
from contextlib import contextmanager
@contextmanager
def redirect_stdout(new_target):
old_target, sys.stdout = sys.stdout, new_target
try:
yield new_target
finally:
sys.stdout = old_target
def get_devnull():
#return open(os.devnull, "w")
return open('log_stuff.txt', "w") #where all the stuff goes you don't want to see
####
```
## Instructions Step 2: Load Data
Edit the directory (folder) name (the text within `' '` in the first line) and then execute the following cell to load the data.
You need to specify the directory (folder) that contains the results of the calculation you want to explore. Discuss with your group and the Model-runner what directory (folder) should be chosen (you can see the options on top of the list in the file browser on the left).
You create two Python data instances per case directory. Each of them can make different plots. In the following cell you specify the case directory in the string variable `case_directory` once, and then both instances are created using the same directory name.
```
case_directory = 'case_4'
pa1 = ppn.abu_vector(case_directory)
px1 = ppn.xtime(case_directory)
last_cycle = len(pa1.files)-1
```
## Instructions Step 3: Visualize Results on the Chart of Nuclides
Executing the following cell creates a chart of nuclides with a color coding that indicates the abundance of each isotope created. Stable isotopes are the thick lined squares. The number indicates the mass number of the isotope. Each row is an element indicated by the element number or proton number Z. This graph helps you identify the most abundant isotopes that have been produced.
Before you execute the next cell make sure the first argument of `pa1.abu_chart()` is the number of the cycle or time step you want to visualize. For example, if you want to visualize the abundances produced at the end of the calculation - then the number should be the highest cycle number (the range of cycle numbers is displayed as output when you load the data in the cell above). Using an earlier cycle number shows the abundances at an earlier time in the network evolution.
```
ifig=1;close(ifig);figure(ifig)
plot_cycle = int(last_cycle/2)
pa1.abu_chart(plot_cycle,ifig=ifig,ilabel=True,imlabel=True,imlabel_fontsize=5,boxstable=True,\
#plotaxis=[6.5,50.5,10.5,40.5], lbound=(-10, 0))
plotaxis=[30,80,25,60], lbound=(-10, 0))
```
## Instructions Step 3: Obtain the abundance number of a specific isotope of interest
As part of your project you will have to write down the exact produced abundance of a specific isotope your group is interested in, for example to determine the factor by which the abundance changed compared to a previous calculation. These are the steps that allow you to do that.
1. Go to the file explorer on the left and double click on the directory (folder) that contains the results of the calculation you want to analyze (the same directory (folder)) you used for the chart of nuclides plot.
2. You should see a long list of output files whose names start with `iso_massf...`. The file name also contains a number. Find the file with the highest number - this will contain the abundances at the end of the calculation (a file is created for each step of the calculation). You can make the file browser window wider by sliding the boundary to the right so you can see the full filenames.
3. Double click on the file with the largest number in the name to open it in a new Tab.
4. You see a long list of isotopes and their abundances. The last two columns contain element name and mass number to identify the isotope. The number in the column before is the abundance in scientific notation. Scroll down or use the Find feature (`Command - F` on Mac, `CTRL - F` on windows) to find the isotope you are interested in.
5. Once you are done, click on your group folder in the folder name on top of the file browser (example Group1 or Group2 or Group3) - this should be the name just before the results folder, separated by `/`. This will get you back to the folder (directory) you came from so you can select a different result folder.
Note the notation of the isotopes is always 5 characters with the first two characters denoting the element symbol in all upper case, and the last three characters the mass number. Element symbol is left justified and mass number is right justified with unused characters being blanks. You need to use that format when searching (type exactly 5 characters).
## Read in and plot temperature and density as functions of time from `trajectory.input` file used in the xrb model case calculation
```
# read in trajectory used in the nova model case calculation
traj_name = 'trajectory.input'
traj = at.readTable(filename=case_directory+'/'+traj_name, datatype='trajectory')
#at.readTable?
print ('trajectory head attributes:',traj.hattrs)
print ('trajectory data columns:',traj.dcols)
year_to_minutes = 365.2422*1440
tmin = year_to_minutes*traj.get('time')
t9 = traj.get('T')
rho = traj.get('rho')
# this is a function that makes a two-axis plot
def two_scales(ax1, time, data1, data2, c1, c2, ls1, ls2, xlbl, ylbl, y2lbl, fsize):
"""
Parameters
----------
ax1 : axis
Axis to put two scales on
time : array-like
x-axis values for both datasets
data1: array-like
Data for left hand scale
data2 : array-like
Data for right hand scale
c1 : color
Color for line 1
c2 : color
Color for line 2
ls1 : linestyle
style for line 1
ls2 : linestyle
style for line 2
xlbl: string
xlabel
ylbl: string
ylabel
y2lbl: string
label of the 2nd y axis
fsize: font size for axis labels
Returns
-------
ax1 : axis
Original axis
ax2 : axis
New twin axis
"""
ax2 = ax1.twinx()
ax1.plot(time, data1, color=c1, linestyle=ls1)
ax1.set_xlabel(xlbl,fontsize=fsize)
ax1.set_ylabel(ylbl,fontsize=fsize)
ax2.plot(time, data2, color=c2, linestyle=ls2)
ax2.set_ylabel(y2lbl,fontsize=fsize)
return ax1, ax2
# Change color of each axis
def color_y_axis(ax, color):
"""Color your axes."""
for t in ax.get_yticklabels():
t.set_color(color)
return None
# make a plot of trajectory used in the nova model case calculation
fig, ax = subplots()
# select color and style for plotting lines
im1 = 8
a1,b1,c1,d1=utils.linestylecb(im1)
im2 = im1+1
a2,b2,c2,d2=utils.linestylecb(im2)
ax1, ax2 = two_scales(ax, tmin, t9, rho, c1, c2, a1, a2,\
'time (min)', '$T_9$',\
'$\\rho\ (\mathrm{g\,cm}^{-3})$', 12)
color_y_axis(ax1, c1)
color_y_axis(ax2, c2)
ax3 = ax1.twiny()
ax3.set_xticks([])
t9max_cycle = argmax(t9)
title('trajectory for ccsn gamma-p model '+case_directory+', $T_9$ has maximum at cycle '+str(t9max_cycle))
tight_layout()
show()
# for this cell to work xrb case run has to output reaction rate fluxes!!!
# otherwise, skip this cell
# use here the cycle with the maximum T9
flux_cycle = plot_cycle # t9max_cycle
flux_file = case_directory+'/flux_'+str(flux_cycle).zfill(5)+'.DAT'
%cp "$flux_file" ./
pa1.flux_solo(flux_cycle,lbound=(-8,0),prange=8,plotaxis=[6.5,37.5,10.5,32.5],profile='neutron',which_flux=0)
tight_layout()
show()
# plot decayed elemental abundances from the ccsn model case for plot_cycle
ifig=5;close(ifig);figure(ifig)
sol_ab = 'iniab2.0E-02GN93.ppn' # file with solar abundances
Z_range = [11,71]
plot_cycle = last_cycle
with get_devnull() as devnull, redirect_stdout(devnull):
pa1.elemental_abund(plot_cycle,ref_filename=sol_ab,zrange=Z_range,ylim=[-2.0,4.0],\
label='cycle '+str(plot_cycle), colour='blue',plotlines='--', plotlabels=True, mark='.')
# to compare with other model run uncomment the following two raws
# pa1a.elemental_abund(plot_cycle,ref_filename=sol_ab,zrange=Z_range,ylim=[-4,5],\
# label='cycle '+str(plot_cycle), colour='red',plotlines=':', plotlabels=True, mark='.')
grid(False)
xlim(Z_range[0],Z_range[-1])
hlines(0,Z_range[0],Z_range[-1],linestyles='dotted')
ylabel('$\log_{10}\,X_i/X_\odot$')
title('decayed elemental abundances for ccsn gamma-p model '+case_directory+', cycle '+str(plot_cycle))
show()
```
## Read in solar isotopic abundances
```
# these are the solar abundances used in nova sims
f = open(sol_ab, 'r')
sol_iso_z=[]
sol_iso=[]
sol_iso_name = []
sol_iso_a = []
sol_iso_abu=[]
for line in f:
n = len(line.split())
if n == 3:
sol_iso = line.split()[1]
if sol_iso == 'PROT':
sol_iso_name.append('h')
sol_iso_a.append(1)
sol_iso_z.append(int(line.split()[0]))
sol_iso_abu.append(float(line.split()[2]))
else:
sol_iso_name.append(sol_iso[0:2])
sol_iso_a.append(int(sol_iso[2:5]))
sol_iso_z.append(int(line.split()[0]))
sol_iso_abu.append(float(line.split()[2]))
if n == 4:
sol_iso_z.append(int(line.split()[0]))
sol_iso_name.append(line.split()[1])
sol_iso_a.append(int(line.split()[2]))
sol_iso_abu.append(float(line.split()[3]))
f.close()
n_iso_sol = len(sol_iso_z)
# read in undecayed isotopic abundances from the xrb model case for the plot_cycle
ppn_file = case_directory+'/'+'iso_massf'+str(plot_cycle).zfill(5)+'.DAT'
print (ppn_file)
f1=open(ppn_file)
lines=f1.readlines()
f1.close()
massfrac=[]
A=[]
Z=[]
element = []
AI = []
for k in range(len(lines)):
# skip header
if k<7:
continue
line=lines[k]
Z.append(line[6:12].strip()) # Z
A.append(line[13:17].strip()) # A float
massfrac.append(line[24:35].strip()) # massf
element.append(line[37:39].strip()) # element (execept NEUT (first) and PROT (second))
AI.append(line[39:43].strip()) # A integer
n_iso_ppn = len(A)
element[0] = 'n'
AI[0] = '1'
element[1] = 'H'
AI[1] ='1'
element[n_iso_ppn-2] = 'ALm'
AI[n_iso_ppn-2] ='26'
element[n_iso_ppn-1] = 'KRm'
AI[n_iso_ppn-1] ='85'
iso_z_ppn = np.linspace(0,0,n_iso_ppn)
iso_a_ppn = np.linspace(0,0,n_iso_ppn)
iso_name_ppn = [" " for x in range(n_iso_ppn)]
iso_abu_ppn = np.linspace(0,0,n_iso_ppn)
for i in range(n_iso_ppn):
iso_name_ppn[i] = element[i]
iso_a_ppn[i] = float(A[i])
iso_z_ppn[i] = float(Z[i])
iso_abu_ppn[i] = float(massfrac[i])
# plot isotopic composition for the xrb model case
ifig=6;close(ifig);fig=figure(ifig)
size=8
fig.canvas.layout.height = str(0.8*size)+'in' # This is a hack to prevent ipympl
fig.canvas.layout.width = str(1.2*size)+'in' # to adjust horizontal figure size
z1 = 1; z2 = 195
for z in range(z1,z2):
a_plot_ppn = []
y_plot_ppn = []
for i in range(n_iso_ppn):
if int(iso_z_ppn[i]) == z:
for k in range(n_iso_sol):
if sol_iso_z[k] == z and sol_iso_a[k] == iso_a_ppn[i]:
a_plot_ppn.append(sol_iso_a[k])
y_plot_ppn.append(log10(iso_abu_ppn[i]/sol_iso_abu[k]))
if len(a_plot_ppn) > 0:
text(a_plot_ppn[argmax(y_plot_ppn)],max(y_plot_ppn),ut.get_el_from_z(z),\
horizontalalignment='center',verticalalignment='bottom',fontsize=10)
if len(a_plot_ppn) > 1:
a_plot_ppn, y_plot_ppn = (list(t) for t in zip(*sorted(zip(a_plot_ppn, y_plot_ppn))))
plot(a_plot_ppn,y_plot_ppn,'--')
plot(a_plot_ppn,y_plot_ppn,'bo',markersize=3)
xmin = z1; xmax = z2
hlines(0,xmin,xmax,linestyles='dotted')
xlim(xmin,xmax)
ylim(-3,5.0)
xlabel('mass number')
ylabel('$\log_{10}\,X_i/X_\odot$')
title('undecayed isotopic abundances for ccsn gamma-p model '+case_directory+', cycle '+str(plot_cycle))
show()
```
| github_jupyter |
# Predicting Student Admissions with Neural Networks in Keras
In this notebook, we predict student admissions to graduate school at UCLA based on three pieces of data:
- GRE Scores (Test)
- GPA Scores (Grades)
- Class rank (1-4)
The dataset originally came from here: http://www.ats.ucla.edu/
## Loading the data
To load the data and format it nicely, we will use two very useful packages called Pandas and Numpy. You can read on the documentation here:
- https://pandas.pydata.org/pandas-docs/stable/
- https://docs.scipy.org/
```
# Importing pandas and numpy
import pandas as pd
import numpy as np
# Reading the csv file into a pandas DataFrame
data = pd.read_csv('student_data.csv')
# Printing out the first 10 rows of our data
data[:10]
```
## Plotting the data
First let's make a plot of our data to see how it looks. In order to have a 2D plot, let's ingore the rank.
```
# Importing matplotlib
import matplotlib.pyplot as plt
# Function to help us plot
def plot_points(data):
X = np.array(data[["gre","gpa"]])
y = np.array(data["admit"])
admitted = X[np.argwhere(y==1)]
rejected = X[np.argwhere(y==0)]
plt.scatter([s[0][0] for s in rejected], [s[0][1] for s in rejected], s = 25, color = 'red', edgecolor = 'k')
plt.scatter([s[0][0] for s in admitted], [s[0][1] for s in admitted], s = 25, color = 'cyan', edgecolor = 'k')
plt.xlabel('Test (GRE)')
plt.ylabel('Grades (GPA)')
# Plotting the points
plot_points(data)
plt.show()
```
Roughly, it looks like the students with high scores in the grades and test passed, while the ones with low scores didn't, but the data is not as nicely separable as we hoped it would. Maybe it would help to take the rank into account? Let's make 4 plots, each one for each rank.
```
# Separating the ranks
data_rank1 = data[data["rank"]==1]
data_rank2 = data[data["rank"]==2]
data_rank3 = data[data["rank"]==3]
data_rank4 = data[data["rank"]==4]
# Plotting the graphs
plot_points(data_rank1)
plt.title("Rank 1")
plt.show()
plot_points(data_rank2)
plt.title("Rank 2")
plt.show()
plot_points(data_rank3)
plt.title("Rank 3")
plt.show()
plot_points(data_rank4)
plt.title("Rank 4")
plt.show()
```
This looks more promising, as it seems that the lower the rank, the higher the acceptance rate. Let's use the rank as one of our inputs. In order to do this, we should one-hot encode it.
## One-hot encoding the rank
For this, we'll use the `get_dummies` function in pandas.
```
# Make dummy variables for rank
one_hot_data = pd.concat([data, pd.get_dummies(data['rank'], prefix='rank')], axis=1)
# Drop the previous rank column
one_hot_data = one_hot_data.drop('rank', axis=1)
# Print the first 10 rows of our data
one_hot_data[:10]
```
## Scaling the data
The next step is to scale the data. We notice that the range for grades is 1.0-4.0, whereas the range for test scores is roughly 200-800, which is much larger. This means our data is skewed, and that makes it hard for a neural network to handle. Let's fit our two features into a range of 0-1, by dividing the grades by 4.0, and the test score by 800.
```
# Copying our data
processed_data = one_hot_data[:]
# Scaling the columns
processed_data['gre'] = processed_data['gre']/800
processed_data['gpa'] = processed_data['gpa']/4.0
processed_data[:10]
```
## Splitting the data into Training and Testing
In order to test our algorithm, we'll split the data into a Training and a Testing set. The size of the testing set will be 10% of the total data.
```
sample = np.random.choice(processed_data.index, size=int(len(processed_data)*0.9), replace=False)
train_data, test_data = processed_data.iloc[sample], processed_data.drop(sample)
print("Number of training samples is", len(train_data))
print("Number of testing samples is", len(test_data))
print(train_data[:10])
print(test_data[:10])
```
## Splitting the data into features and targets (labels)
Now, as a final step before the training, we'll split the data into features (X) and targets (y).
Also, in Keras, we need to one-hot encode the output. We'll do this with the `to_categorical function`.
```
import keras
# Separate data and one-hot encode the output
# Note: We're also turning the data into numpy arrays, in order to train the model in Keras
features = np.array(train_data.drop('admit', axis=1))
targets = np.array(keras.utils.to_categorical(train_data['admit'], 2))
features_test = np.array(test_data.drop('admit', axis=1))
targets_test = np.array(keras.utils.to_categorical(test_data['admit'], 2))
print(features[:10])
print(targets[:10])
```
## Defining the model architecture
Here's where we use Keras to build our neural network.
```
# Imports
import numpy as np
from keras.models import Sequential
from keras.layers.core import Dense, Dropout, Activation
from keras.optimizers import SGD
from keras.utils import np_utils
# Building the model
model = Sequential()
model.add(Dense(128, activation='relu', input_shape=(6,)))
model.add(Dropout(.2))
model.add(Dense(64, activation='relu'))
model.add(Dropout(.1))
model.add(Dense(2, activation='softmax'))
# Compiling the model
model.compile(loss = 'categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
model.summary()
```
## Training the model
```
# Training the model
model.fit(features, targets, epochs=200, batch_size=100, verbose=0)
```
## Scoring the model
```
# Evaluating the model on the training and testing set
score = model.evaluate(features, targets)
print("\n Training Accuracy:", score[1])
score = model.evaluate(features_test, targets_test)
print("\n Testing Accuracy:", score[1])
```
## Challenge: Play with the parameters!
You can see that we made several decisions in our training. For instance, the number of layers, the sizes of the layers, the number of epochs, etc.
It's your turn to play with parameters! Can you improve the accuracy? The following are other suggestions for these parameters. We'll learn the definitions later in the class:
- Activation function: relu and sigmoid
- Loss function: categorical_crossentropy, mean_squared_error
- Optimizer: rmsprop, adam, ada
| github_jupyter |
# Fighting Game AI
## Introduction
Do you know about Pokemon? Well this turn-based fighting game is similar to Pokemon, but we do not send out monsters to duke it out. Instead, the battle is between humans. Since the actions that can be taken by both players are the same, there is some form of game theory involved and we can determine the Nash equilibrium of the game through the use of Monte Carlo Tree Search (MCTS).
Players start with 100 health. For each turn, players can choose one of four actions available:
1. <b>Attack</b> <i>{DAMAGE SKILL}</i> (opponent is damaged for 8-12 health)
2. <b>Heal</b> <i>{HEAL SKILL}</i> (self is healed for 7-10 health)
3. <b>Power Up</b> <i>{BUFF SKILL}</i> (effectiveness of damage skills increase by 25%, to a maximum of 100%)
4. <b>Superspeed</b> <i>{BUFF SKILL}</i> (chance of dodging a damage skill or sudden death effect increases by 5%, to a maximum of 100%)
Starting from the 11th turn, sudden death will begin, and the players will be hit for 2 * (number_of_turns_passed - 10) health at the end of their turn. After turn 50, if both players have not fainted yet, the game is a draw. Players are able to dodge the effects of sudden death as well.
For buff skills, we will use the example of the skill "power up" to show how effects can stacked. When the player has used "power up" twice, the effectiveness of damage skills is increased by 50%.
You can edit the various variables to see how the sequence of actions that lead to the best result would differ.
```
!pip install mcts_simple
import numpy as np
import random
from gym import Env
from gym.spaces import Discrete, Box
class Skill:
def __init__(self, player):
self.name = None
self.type = None
self.description = None
self.player = player
def use(self, opponent):
pass
def damage(self, opponent, damage):
if random.random() < opponent.dodge_rate:
if self.player.output:
print(f"Player {opponent.name} dodged the incoming attack.")
else:
original_health = opponent.health
opponent.health = max(opponent.health - int(damage * self.player.damage_multiplier), 0)
if self.player.output:
print(f"Player {opponent.name} is hit for {original_health - opponent.health} damage by the skill {self.name}.")
def heal(self, damage):
original_health = self.player.health
self.player.health = min(self.player.health + damage, 100)
if self.player.output:
print(f"Player {self.player.name} is healed for {self.player.health - original_health} damage using the skill {self.name}.")
def increase_damage(self, multiplier):
original_damage_multiplier = self.player.damage_multiplier
self.player.damage_multiplier = max(min(self.player.damage_multiplier + multiplier, 2), 0)
if self.player.output:
print(f"Player {self.player.name}'s damage has increased by {self.player.damage_multiplier - original_damage_multiplier:.0%} using the skill {self.name}.")
def increase_dodge(self, dodge_rate):
original_dodge_rate = self.player.dodge_rate
self.player.dodge_rate = max(min(self.player.dodge_rate + dodge_rate, 1), 0)
if self.player.output:
print(f"Player {self.player.name}'s dodge rate has increased by {self.player.dodge_rate - original_dodge_rate:.0%} using the skill {self.name}.")
class Attack(Skill):
def __init__(self, player):
self.name = "Attack"
self.type = "Damage"
self.description = "Opponent is hit for 8-12 health."
self.player = player
def use(self, opponent):
self.damage(opponent, random.randint(8, 12))
class Heal(Skill):
def __init__(self, player):
self.name = "Heal"
self.type = "Heal"
self.description = "Player is healed for 7-10 health."
self.player = player
def use(self, opponent):
self.heal(random.randint(7, 10))
class PowerUp(Skill):
def __init__(self, player):
self.name = "Power Up"
self.type = "Buff"
self.description = "Effectiveness of damage skills increases by 25%. Damage multiplier can be increased to a maximum of 100%."
self.player = player
def use(self, opponent):
self.increase_damage(0.25)
class Superspeed(Skill):
def __init__(self, player):
self.name = "Superspeed"
self.type = "Buff"
self.description = "Chance of dodging damage skills or effect from sudden death increases by 5%. Dodge rate can be increased to a maximum of 100%."
self.player = player
def use(self, opponent):
self.increase_dodge(0.05)
class Player:
def __init__(self, number, player_name, output = False):
self.number = number
self.name = player_name
self.skills = [Attack(self), Heal(self), PowerUp(self), Superspeed(self)]
self.health = 100
self.damage_multiplier = 1
self.dodge_rate = 0
self.output = output
def has_fainted(self):
return self.health <= 0
class FightingGameEnv(Env): # Multi agent environment
def __init__(self, output = False):
# Output
self.output = output
# Players
self.player_1 = Player(1, "1", self.output)
self.player_2 = Player(2, "2", self.output)
self.agents = [self.player_1, self.player_2]
self.agent_mapping = dict(zip(self.agents, list(range(len(self.agents)))))
self.agent_selection = self.agents[0]
# Actions
self.action_spaces = {agent: Discrete(4) for agent in self.agents}
# Observations {}: set of int, []: range of float
# <<<player: {1, 2}, turn_number: {1, ..., 50}, sudden_death: {0, 1}, sudden_death_damage: {0, ..., 80},
# player_1_health: {0, ..., 100}, player_2_health: {0, ..., 100}, player_1_damage_multiplier: [1, 2],
# player_2_damage_multiplier: [1, 2], player_1_dodge_rate: [0, 1], player_2_dodge_rate: [0, 1]>>>
self.observation_spaces = {agent: Box(np.array([1., 1., 0., 0., 0., 0., 1., 1., 0., 0.], dtype = np.float64), np.array([1., 50., 1., 80., 100., 100., 2., 2., 1., 1.], dtype = np.float64), dtype = np.float64) for agent in self.agents}
# Parameters
self.turn_number = 1
self.episode_length = 50
self.sudden_death_damage = 2
# Things to return
self.observations = {agent: self.get_state() for agent in self.agents}
self.actions = {agent: None for agent in self.agents}
self.rewards = {agent: 0 for agent in self.agents} # DO NOT RETURN THIS
self.cumulative_rewards = {agent: 0 for agent in self.agents}
self.dones = {agent: False for agent in self.agents}
self.infos = {agent: {} for agent in self.agents}
def render(self, mode = "human"):
# Output
if self.output:
print("Turn:", self.turn_number)
print(f"Player {self.agents[0].name} health: {self.agents[0].health}")
print(f"Player {self.agents[0].name} damage increase: {self.agents[0].damage_multiplier - 1:.0%}")
print(f"Player {self.agents[0].name} dodge rate: {self.agents[0].dodge_rate:.0%}")
print(f"Player {self.agents[1].name} health: {self.agents[1].health}")
print(f"Player {self.agents[1].name} damage increase: {self.agents[1].damage_multiplier - 1:.0%}")
print(f"Player {self.agents[1].name} dodge rate: {self.agents[1].dodge_rate:.0%}")
def get_state(self):
return np.array([self.agent_mapping[self.agent_selection] + 1,
self.turn_number,
1 if self.turn_number > 10 else 0,
self.sudden_death_damage * max(self.turn_number - 10, 0),
self.player_1.health,
self.player_2.health,
self.player_1.damage_multiplier,
self.player_2.damage_multiplier,
self.player_1.dodge_rate,
self.player_2.dodge_rate],
dtype = np.float64)
def reset(self):
self.__init__(self.output) # reset classes made
def step(self, action):
# Agent's action
self.actions[self.agent_selection] = action
# Reset rewards
for agent in self.rewards:
self.rewards[self.agent_selection] = 0
self.cumulative_rewards[self.agent_selection] = 0
# Track previous health
self_health = self.agent_selection.health
opponent_health = self.agents[self.agent_mapping[self.agent_selection] ^ 1].health
# Agent takes action
self.agent_selection.skills[action].use(self.agents[self.agent_mapping[self.agent_selection] ^ 1])
self.infos[self.agent_selection]["action"] = action # logging purposes
# Sudden death
if not self.agents[self.agent_mapping[self.agent_selection] ^ 1].has_fainted(): # if opponent has not fainted
sudden_death_damage = self.sudden_death_damage * max(self.turn_number - 10, 0)
if sudden_death_damage:
if random.random() < self.agent_selection.dodge_rate:
if self.output:
print(f"Player {self.agent_selection.name} dodged the sudden death effect.")
else:
temp_health = self.agent_selection.health
self.agent_selection.health = max(self.agent_selection.health - sudden_death_damage, 0)
if self.output:
print(f"Player {self.agent_selection.name} has been hit for {temp_health - self.agent_selection.health} health by sudden death!")
# Calculate rewards
if self.agent_selection.has_fainted():
self.rewards[self.agent_selection] -= 1
self.rewards[self.agents[self.agent_mapping[self.agent_selection] ^ 1]] += 1
elif self.agents[self.agent_mapping[self.agent_selection] ^ 1].has_fainted():
self.rewards[self.agent_selection] += 1
self.rewards[self.agents[self.agent_mapping[self.agent_selection] ^ 1]] -= 1
# Determine episode completion
if self.agent_mapping[self.agent_selection] == 0: # PLAYER 1
self.dones = {agent: self.player_1.has_fainted() or self.player_2.has_fainted() for agent in self.agents}
elif self.agent_mapping[self.agent_selection] == 1: # PLAYER 2
self.turn_number += 1
self.dones = {agent: self.turn_number >= self.episode_length or self.player_1.has_fainted() or self.player_2.has_fainted() for agent in self.agents} # check for turn number only applies at end of second player's turn
# Selects the next agent
self.agent_selection = self.agents[self.agent_mapping[self.agent_selection] ^ 1]
# Next agent's observation
self.observations[self.agent_selection] = self.get_state()
# Update rewards
for agent, reward in self.rewards.items():
self.cumulative_rewards[agent] += reward
# Output next line
if self.output:
print()
from mcts_simple import Game
from copy import deepcopy
class FightingGame(Game):
def __init__(self, output = False):
self.env = FightingGameEnv(output)
self.prev_env = None
def render(self):
self.env.render()
def get_state(self):
return tuple(self.env.get_state())
def number_of_players(self):
return len(self.env.agents)
def current_player(self):
return self.env.agent_selection.name
def possible_actions(self):
return [str(i) for i in range(4)]
def take_action(self, action):
if action not in self.possible_actions():
raise RuntimeError("Action taken is invalid.")
action = int(action)
self.prev_env = deepcopy(self.env)
self.env.step(action)
def delete_last_action(self):
if self.prev_env is None:
raise RuntimeError("No last action to delete.")
if self.env.output:
raise RuntimeError("Output to terminal should be disabled using output = False when deleting last action.")
self.env = self.prev_env
self.prev_env = None
def has_outcome(self):
return True in self.env.dones.values()
def winner(self):
if not self.has_outcome():
raise RuntimeError("winner() cannot be called when outcome is undefined.")
if self.env.player_2.has_fainted() or self.env.player_1.health > self.env.player_2.health:
return self.env.player_1.name
elif self.env.player_1.has_fainted() or self.env.player_2.health > self.env.player_1.health:
return self.env.player_2.name
else:
return None
## This example shows how Open loop MCTS deals with uncertainty ###
from mcts_simple import OpenLoopMCTS, OpenLoopUCT
# Export trained MCTS
print("Export trained MCTS")
mcts = OpenLoopMCTS(FightingGame(output = False))
mcts.run(iterations = 50000)
mcts._export("FightingGame_MCTS.json")
print()
# Import trained MCTS
print("Import trained MCTS")
mcts = OpenLoopMCTS(FightingGame(output = True))
mcts._import("FightingGame_MCTS.json")
mcts.self_play(activation = "best")
print()
# Export trained UCT
print("Export trained UCT")
uct = OpenLoopUCT(FightingGame(output = False))
uct.run(iterations = 100000)
uct._export("FightingGame_UCT.json")
print()
# Import trained UCT
print("Import trained UCT")
uct = OpenLoopUCT(FightingGame(output = True))
uct._import("FightingGame_UCT.json")
uct.self_play(activation = "best")
print()
# Play with UCT agent
print("Play with UCT agent")
uct = OpenLoopUCT(FightingGame(output = True))
uct._import("FightingGame_UCT.json")
uct.play_with_human(activation = "linear")
print()
```
| github_jupyter |
# 判断爬取下来的url是哪种类型的url
```
CSDN_SEEDS_URL = "https://blog.csdn.net/"
REGEX_CSDN_USER_MY_URL = "http[s]*://my\\.csdn\\.net/\\w+"
REGEX_CSDN_USER_BLOG_URL = "http[s]*://blog\\.csdn\\.net/\\w+"
REGEX_CSDN_BLOG_LIST_URL = "http[s]*://blog\\.csdn\\.net/\\w+/article/list/\\d+\\?"
REGEX_CSDN_BLOG_URL = "http[s]*://blog\\.csdn\\.net/\\w+/article/details/\\w+"
with open("./urls.txt", "r", encoding="utf-8") as f:
urls = f.readlines()
urls[0:5]
import re
from tqdm import tqdm
USER_MY_URL = []
USER_BLOG_URL = []
BLOG_LIST_URL = []
BLOG_URL = []
# 其实也可以简单一点就是一种是博客url,一种是非博客url
for url in tqdm(urls):
if re.match(REGEX_CSDN_BLOG_URL, url) != None:
BLOG_URL.append(url)
elif re.match(REGEX_CSDN_BLOG_LIST_URL, url) != None:
BLOG_LIST_URL.append(url)
elif re.match(REGEX_CSDN_USER_MY_URL, url) != None:
USER_MY_URL.append(url)
elif re.match(REGEX_CSDN_USER_BLOG_URL, url) != None:
USER_BLOG_URL.append(url)
print(len(USER_MY_URL))
print(len(USER_BLOG_URL))
print(len(BLOG_LIST_URL))
print(len(BLOG_URL))
```
# 测试GEN的效果
```
! pip install gne -i https://pypi.tuna.tsinghua.edu.cn/simple
with open('./test.html', 'r', encoding="utf-8") as f:
html = f.read()
print(html)
extractor = GeneralNewsExtractor().extractor.extract(html, with_body_html=True)
from gne import GeneralNewsExtractor
extractor = GeneralNewsExtractor()
result = extractor.extract(html)
print(result)
result["content"]
```
# 测试Python Goose
```
! pip install goose-extractor i https://pypi.tuna.tsinghua.edu.cn/simple
import goose
g = goose.Goose()
article = g.extract(raw_html=html)
print(article.title.encode('gbk', 'ignore'))
print(article.meta_description.encode('gbk', 'ignore'))
print(article.cleaned_text.encode('gbk', 'ignore'))
```
# 使用xpath提取content
```
# 提取content
'//div[contains(@class, "content") and not(contains(@class, "comment" or ""))]'
```
# 测试html_extractor
```
from Engine.html_extractor import MainContent
extractor = MainContent()
url = "https://blog.csdn.net/hihell/article/details/121012464"
with open('./test.html', 'r', encoding="utf-8") as f:
html = f.read()
title,content = extractor.extract(url, html)
print(title, "\n" , "*"*40)
print(content)
```
# 判断url是否为内容界面
而不是列表界面或者主页又或者用户页等
```
# 去除http域名这些
# 粗略地统计以下url中会含有地关键词,后面可以尝试以下神经网络
import jieba
import nltk
from tqdm import tqdm
import re
import pandas as pd
with open('./csdn_urls.txt', 'r', encoding="utf-8") as f:
csdn_urls = f.readlines()
with open('./juejin_urls.txt', 'r', encoding="utf-8") as f:
juejin_urls = f.readlines()
with open('./sg_urls.txt', 'r', encoding="utf-8") as f:
sg_urls = f.readlines()
urls = sg_urls + csdn_urls + juejin_urls
print(len(urls))
dic_words = {}
# nltk不能区分\
# for url in tqdm(urls):
# text = nltk.word_tokenize(url)
# for word in text:
# dic_words[word] = dic_words.get(word, 0) + 1
# for url in tqdm(urls):
# text = jieba.cut(url)
# for word in text:
# dic_words[word] = dic_words.get(word, 0) + 1
for url in tqdm(urls):
text = re.findall('[a-z]+', url.lower())
for word in text:
dic_words[word] = dic_words.get(word, 0) + 1
df = pd.DataFrame.from_dict(dic_words, orient="index")
print(df.sort_values(by=[0],na_position='last'))
df.to_csv("url_keywords.csv")
# 这类词加分减分
# 可以把这些url分词排序然后认为判断加入消极还是积极
NEG_WORDS = ['user', 'list', 'authors', 'comment','writers','blogs']
POS_WORDS = ['article', 'blog', 'details']
# 参考https://help.aliyun.com/document_detail/65096.html
# 还要记住匹配的时候不能区分大小写,同时匹配的时候也仅仅需要匹配url的最后四位就可以了
# 这类词一票否决
FILE_WORDS = ['gif','png','bmp','jpeg','jpg', 'svg',
'mp3','wma','flv','mp4','wmv','ogg','avi',
'doc','docx','xls','xlsx','ppt','pptx','txt','pdf',
'zip','exe','tat','ico','css','js','swf','apk','m3u8','ts']
# 还有就是如果包含很长一串数字的一般都是内容界面
# 不应该是各种文件名的后缀
# 连续数字的匹配
import re
test_url = "https://blog.csdn.net/ITF_001?utm_source=feed"
test_str = "982374589234789"
test_str1 = "sdfdsfsdfsdf232sdfsdf"
rule1 = "[0-9]"*8
print(re.match(rule1, test_str, flags=0))
```
# 将爬取下来的相对URL链接转换为绝对链接
```
from urllib import parse
page_url = 'http://fcg.gxepb.gov.cn/ztzl/hjwfbgt/'
new_relative_url = 'http://fcg.gxepb.gov.cn/hjzf/xzcf/201811/t20181102_46347.html'
new_absolute_url = '../../hjzf/xzcf/201811/t20181102_46347.html'
new_full_url1 = parse.urljoin(page_url, new_relative_url)
new_full_url2 = parse.urljoin(page_url, new_absolute_url)
print(new_full_url1)
print(new_full_url2)
```
# 测试url_parser
```
from Engine.url_parser import is_static_url
with open('./csdn_urls.txt', 'r', encoding="utf-8") as f:
csdn_urls = f.readlines()
with open('./juejin_urls.txt', 'r', encoding="utf-8") as f:
juejin_urls = f.readlines()
with open('./sg_urls.txt', 'r', encoding="utf-8") as f:
sg_urls = f.readlines()
urls = sg_urls + csdn_urls + juejin_urls
print(len(urls))
for i in range(8000):
if is_static_url(urls[i]):
print(urls[i])
```
# 测试gerapy_auto_extractor
```
import sys
sys.path.append("C:/My_app/code/咻Search/Engine")
from gerapy_auto_extractor.classifiers.list import is_list
from gerapy_auto_extractor.classifiers.detail import is_detail
with open("./test.html", 'r', encoding='utf-8') as f:
html= f.read()
# 这个两个可以用
print(is_detail(html,threshold=0.3))
print(is_list(html,threshold=0.9))
import sys
sys.path.append("C:/My_app/code/咻Search/Engine")
from gerapy_auto_extractor.extractors.content import extract_content
from gerapy_auto_extractor.extractors.datetime import extract_datetime
from gerapy_auto_extractor.extractors.list import extract_list
from gerapy_auto_extractor.extractors.title import extract_title
with open("./test.html", 'r', encoding='utf-8') as f:
html= f.read()
# print(extract_title(html))
# print(extract_list(html))
print(extract_datetime(html))
# 这个效果极其垃圾
# content_html = extract_content(html)
{"hah":"hello","D": 123}.get("hah","")
print("zhenghui" or "郑辉")
{"hah":"hello","D": 123}.get("hah","")
```
# 测试查出mysql数据
```
import pandas as pd
import pymysql
from config import MYSQL_HOST, MYSQL_DBNAME, MYSQL_USER, MYSQL_PASSWORD
# 连接数据库
# 加上charset='utf8',避免 'latin-1' encoding 报错等问题
conn = pymysql.connect(host=MYSQL_HOST, user=MYSQL_USER, passwd=MYSQL_PASSWORD,
db=MYSQL_DBNAME, charset='utf8')
# 创建cursor
cursor_blogs = conn.cursor()
cursor_list = conn.cursor()
sql_blogs = 'SELECT page_url, urls FROM search_blogs;'
sql_list = 'SELECT page_url, urls FROM search_blogs;'
# 执行sql语句
cursor_blogs.execute(sql_blogs)
cursor_list.execute(sql_list)
# 获取数据库列表信息
col_blogs = cursor_blogs.description
col_list = cursor_list.description
# 获取全部查询信息
re_blogs = cursor_blogs.fetchall()
re_list = cursor_list.fetchall()
# 获取一行信息
# re = cursor.fetchone()
# 获取的信息默认为tuple类型,将columns转换成DataFrame类型
columns_blogs = pd.DataFrame(list(col_blogs))
# 将数据转换成DataFrame类型,并匹配columns
df_blogs = pd.DataFrame(list(re_blogs), columns=columns_blogs[0])
columns_list = pd.DataFrame(list(col_blogs))
# 将数据转换成DataFrame类型,并匹配columns
df_list = pd.DataFrame(list(re_blogs), columns=columns_list[0])
import ast
blogs_index = [url[0] for url in re_blogs]
list_index = [ast.literal_eval(url[1]) for url in re_blogs]
list_index[0][0]
import numpy as np
def urls2G():
'''
将数据库中urls的关系转化为图
'''
# 连接数据库
# 加上charset='utf8',避免 'latin-1' encoding 报错等问题
conn = pymysql.connect(host=MYSQL_HOST, user=MYSQL_USER, passwd=MYSQL_PASSWORD,
db=MYSQL_DBNAME, charset='utf8')
# 创建cursor
cursor_blogs = conn.cursor()
cursor_list = conn.cursor()
sql_blogs = 'SELECT page_url, urls FROM search_blogs;'
sql_list = 'SELECT page_url, urls FROM search_blogs;'
# 执行sql语句
cursor_blogs.execute(sql_blogs)
cursor_list.execute(sql_list)
# 获取全部查询信息
re_blogs = cursor_blogs.fetchall()
re_list = cursor_list.fetchall()
# 将获取的元组信息转换为图
blogs_index = [url[0] for url in re_blogs]
blogs_point = [ast.literal_eval(url[1]) for url in re_blogs]
list_index = [url[0] for url in re_list]
list_point = [ast.literal_eval(url[1]) for url in re_list]
indexs = blogs_index + list_index
points = blogs_point + list_point
G = np.zeros((len(indexs), len(indexs)))
for i, index in enumerate(indexs):
# 依次判断包含的url是是否在爬取过的列表中,有些广告之类的链接页会包含,但没爬取
for p_url in points[i]:
try:
p_index = indexs.index(p_url)
except:
p_index = -1
if p_index != -1:
G[i][p_index] = 1
return G
urls2G()
```
| github_jupyter |
Please find torch implementation of this notebook here: https://colab.research.google.com/github/probml/pyprobml/blob/master/notebooks/book1/15/gru_torch.ipynb
<a href="https://colab.research.google.com/github/probml/probml-notebooks/blob/main/notebooks-d2l/gru_jax.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Gated Recurrent Units
We show how to implement GRUs from scratch.
Based on sec 9.1 of http://d2l.ai/chapter_recurrent-modern/gru.html
This uses code from the [basic RNN colab](https://colab.research.google.com/github/probml/pyprobml/blob/master/notebooks/rnn_torch.ipynb).
```
import jax.numpy as jnp
import matplotlib.pyplot as plt
import math
from IPython import display
import jax
try:
import flax.linen as nn
except ModuleNotFoundError:
%pip install flax
import flax.linen as nn
from flax import jax_utils
try:
import optax
except ModuleNotFoundError:
%pip install optax
import optax
import collections
import re
import random
import os
import requests
import hashlib
import time
import functools
random.seed(0)
rng = jax.random.PRNGKey(0)
!mkdir figures # for saving plots
```
# Data
As data, we use the book "The Time Machine" by H G Wells,
preprocessed using the code in [this colab](https://colab.research.google.com/github/probml/pyprobml/blob/master/notebooks/text_preproc_torch.ipynb).
```
class SeqDataLoader:
"""An iterator to load sequence data."""
def __init__(self, batch_size, num_steps, use_random_iter, max_tokens):
if use_random_iter:
self.data_iter_fn = seq_data_iter_random
else:
self.data_iter_fn = seq_data_iter_sequential
self.corpus, self.vocab = load_corpus_time_machine(max_tokens)
self.batch_size, self.num_steps = batch_size, num_steps
def __iter__(self):
return self.data_iter_fn(self.corpus, self.batch_size, self.num_steps)
class Vocab:
"""Vocabulary for text."""
def __init__(self, tokens=None, min_freq=0, reserved_tokens=None):
if tokens is None:
tokens = []
if reserved_tokens is None:
reserved_tokens = []
# Sort according to frequencies
counter = count_corpus(tokens)
self.token_freqs = sorted(counter.items(), key=lambda x: x[1], reverse=True)
# The index for the unknown token is 0
self.unk, uniq_tokens = 0, ["<unk>"] + reserved_tokens
uniq_tokens += [token for token, freq in self.token_freqs if freq >= min_freq and token not in uniq_tokens]
self.idx_to_token, self.token_to_idx = [], dict()
for token in uniq_tokens:
self.idx_to_token.append(token)
self.token_to_idx[token] = len(self.idx_to_token) - 1
def __len__(self):
return len(self.idx_to_token)
def __getitem__(self, tokens):
if not isinstance(tokens, (list, tuple)):
return self.token_to_idx.get(tokens, self.unk)
return [self.__getitem__(token) for token in tokens]
def to_tokens(self, indices):
if not isinstance(indices, (list, tuple)):
return self.idx_to_token[indices]
return [self.idx_to_token[index] for index in indices]
def tokenize(lines, token="word"):
"""Split text lines into word or character tokens."""
if token == "word":
return [line.split() for line in lines]
elif token == "char":
return [list(line) for line in lines]
else:
print("ERROR: unknown token type: " + token)
def count_corpus(tokens):
"""Count token frequencies."""
# Here `tokens` is a 1D list or 2D list
if len(tokens) == 0 or isinstance(tokens[0], list):
# Flatten a list of token lists into a list of tokens
tokens = [token for line in tokens for token in line]
return collections.Counter(tokens)
def seq_data_iter_random(corpus, batch_size, num_stepsz):
"""Generate a minibatch of subsequences using random sampling."""
# Start with a random offset (inclusive of `num_steps - 1`) to partition a
# sequence
corpus = corpus[random.randint(0, num_steps - 1) :]
# Subtract 1 since we need to account for labels
num_subseqs = (len(corpus) - 1) // num_steps
# The starting indices for subsequences of length `num_steps`
initial_indices = list(range(0, num_subseqs * num_steps, num_steps))
# In random sampling, the subsequences from two adjacent random
# minibatches during iteration are not necessarily adjacent on the
# original sequence
random.shuffle(initial_indices)
def data(pos):
# Return a sequence of length `num_steps` starting from `pos`
return corpus[pos : pos + num_steps]
num_batches = num_subseqs // batch_size
for i in range(0, batch_size * num_batches, batch_size):
# Here, `initial_indices` contains randomized starting indices for
# subsequences
initial_indices_per_batch = initial_indices[i : i + batch_size]
X = [data(j) for j in initial_indices_per_batch]
Y = [data(j + 1) for j in initial_indices_per_batch]
yield jnp.array(X), jnp.array(Y)
def seq_data_iter_sequential(corpus, batch_size, num_steps):
"""Generate a minibatch of subsequences using sequential partitioning."""
# Start with a random offset to partition a sequence
offset = random.randint(0, num_steps)
num_tokens = ((len(corpus) - offset - 1) // batch_size) * batch_size
Xs = jnp.array(corpus[offset : offset + num_tokens])
Ys = jnp.array(corpus[offset + 1 : offset + 1 + num_tokens])
Xs, Ys = Xs.reshape(batch_size, -1), Ys.reshape(batch_size, -1)
num_batches = Xs.shape[1] // num_steps
for i in range(0, num_steps * num_batches, num_steps):
X = Xs[:, i : i + num_steps]
Y = Ys[:, i : i + num_steps]
yield X, Y
def download(name, cache_dir=os.path.join("..", "data")):
"""Download a file inserted into DATA_HUB, return the local filename."""
assert name in DATA_HUB, f"{name} does not exist in {DATA_HUB}."
url, sha1_hash = DATA_HUB[name]
os.makedirs(cache_dir, exist_ok=True)
fname = os.path.join(cache_dir, url.split("/")[-1])
if os.path.exists(fname):
sha1 = hashlib.sha1()
with open(fname, "rb") as f:
while True:
data = f.read(1048576)
if not data:
break
sha1.update(data)
if sha1.hexdigest() == sha1_hash:
return fname # Hit cache
print(f"Downloading {fname} from {url}...")
r = requests.get(url, stream=True, verify=True)
with open(fname, "wb") as f:
f.write(r.content)
return fname
def read_time_machine():
"""Load the time machine dataset into a list of text lines."""
with open(download("time_machine"), "r") as f:
lines = f.readlines()
return [re.sub("[^A-Za-z]+", " ", line).strip().lower() for line in lines]
def load_corpus_time_machine(max_tokens=-1):
"""Return token indices and the vocabulary of the time machine dataset."""
lines = read_time_machine()
tokens = tokenize(lines, "char")
vocab = Vocab(tokens)
# Since each text line in the time machine dataset is not necessarily a
# sentence or a paragraph, flatten all the text lines into a single list
corpus = [vocab[token] for line in tokens for token in line]
if max_tokens > 0:
corpus = corpus[:max_tokens]
return corpus, vocab
def load_data_time_machine(batch_size, num_steps, use_random_iter=False, max_tokens=10000):
"""Return the iterator and the vocabulary of the time machine dataset."""
data_iter = SeqDataLoader(batch_size, num_steps, use_random_iter, max_tokens)
return data_iter, data_iter.vocab
DATA_HUB = dict()
DATA_URL = "http://d2l-data.s3-accelerate.amazonaws.com/"
DATA_HUB["time_machine"] = (DATA_URL + "timemachine.txt", "090b5e7e70c295757f55df93cb0a180b9691891a")
batch_size, num_steps = 32, 35
train_iter, vocab = load_data_time_machine(batch_size, num_steps)
```
# Creating the model from scratch
Initialize the parameters.
```
def get_params(vocab_size, num_hiddens, init_rng):
num_inputs = num_outputs = vocab_size
def normal(shape, rng):
return jax.random.normal(rng, shape=shape) * 0.01
def three(rng):
return (normal((num_inputs, num_hiddens), rng), normal((num_hiddens, num_hiddens), rng), jnp.zeros(num_hiddens))
update_rng, reset_rng, hidden_rng, out_rng = jax.random.split(init_rng, num=4)
W_xz, W_hz, b_z = three(update_rng) # Update gate parameters
W_xr, W_hr, b_r = three(reset_rng) # Reset gate parameters
W_xh, W_hh, b_h = three(hidden_rng) # Candidate hidden state parameters
# Output layer parameters
W_hq = normal((num_hiddens, num_outputs), out_rng)
b_q = jnp.zeros(num_outputs)
params = [W_xz, W_hz, b_z, W_xr, W_hr, b_r, W_xh, W_hh, b_h, W_hq, b_q]
return params
```
Initial state is an array of zeros of size (batch-size, num-hiddens)
```
def init_gru_state(batch_size, num_hiddens):
return (jnp.zeros((batch_size, num_hiddens)),)
```
Forward function
```
@jax.jit
def gru(params, state, inputs):
W_xz, W_hz, b_z, W_xr, W_hr, b_r, W_xh, W_hh, b_h, W_hq, b_q = params
(H,) = state
outputs = []
for X in inputs:
Z = jax.nn.sigmoid((X @ W_xz) + (H @ W_hz) + b_z)
R = jax.nn.sigmoid((X @ W_xr) + (H @ W_hr) + b_r)
H_tilda = jnp.tanh((X @ W_xh) + ((R * H) @ W_hh) + b_h)
H = Z * H + (1 - Z) * H_tilda
Y = H @ W_hq + b_q
outputs.append(Y)
return jnp.concatenate(outputs, axis=0), (H,)
# Make the model class
# Input X to apply is (B,T) matrix of integers (from vocab encoding).
# We transpose this to (T,B) then one-hot encode to (T,B,V), where V is vocab.
# The result is passed to the forward function.
# (We define the forward function as an argument, so we can change it later.)
class RNNModelScratch:
"""A RNN Model implemented from scratch."""
def __init__(self, vocab_size, num_hiddens, get_params, init_state, forward_fn):
self.vocab_size, self.num_hiddens = vocab_size, num_hiddens
self.init_state, self.get_params = init_state, get_params
self.forward_fn = forward_fn
def apply(self, params, state, X):
X = jax.nn.one_hot(X.T, num_classes=self.vocab_size)
return self.forward_fn(params, state, X)
def begin_state(self, batch_size):
return self.init_state(batch_size, self.num_hiddens)
def init_params(self, rng):
return self.get_params(self.vocab_size, self.num_hiddens, rng)
```
# Training and prediction
```
@jax.jit
def grad_clipping(grads, theta):
"""Clip the gradient."""
def grad_update(grads):
return jax.tree_map(lambda g: g * theta / norm, grads)
norm = jnp.sqrt(sum(jax.tree_util.tree_leaves(jax.tree_map(lambda x: jnp.sum(x**2), grads))))
# Update gradient if norm > theta
# This is jax.jit compatible
grads = jax.lax.cond(norm > theta, grad_update, lambda g: g, grads)
return grads
class Animator:
"""For plotting data in animation."""
def __init__(
self,
xlabel=None,
ylabel=None,
legend=None,
xlim=None,
ylim=None,
xscale="linear",
yscale="linear",
fmts=("-", "m--", "g-.", "r:"),
nrows=1,
ncols=1,
figsize=(3.5, 2.5),
):
# Incrementally plot multiple lines
if legend is None:
legend = []
display.set_matplotlib_formats("svg")
self.fig, self.axes = plt.subplots(nrows, ncols, figsize=figsize)
if nrows * ncols == 1:
self.axes = [
self.axes,
]
# Use a lambda function to capture arguments
self.config_axes = lambda: set_axes(self.axes[0], xlabel, ylabel, xlim, ylim, xscale, yscale, legend)
self.X, self.Y, self.fmts = None, None, fmts
def add(self, x, y):
# Add multiple data points into the figure
if not hasattr(y, "__len__"):
y = [y]
n = len(y)
if not hasattr(x, "__len__"):
x = [x] * n
if not self.X:
self.X = [[] for _ in range(n)]
if not self.Y:
self.Y = [[] for _ in range(n)]
for i, (a, b) in enumerate(zip(x, y)):
if a is not None and b is not None:
self.X[i].append(a)
self.Y[i].append(b)
self.axes[0].cla()
for x, y, fmt in zip(self.X, self.Y, self.fmts):
self.axes[0].plot(x, y, fmt)
self.config_axes()
display.display(self.fig)
display.clear_output(wait=True)
class Timer:
"""Record multiple running times."""
def __init__(self):
self.times = []
self.start()
def start(self):
"""Start the timer."""
self.tik = time.time()
def stop(self):
"""Stop the timer and record the time in a list."""
self.times.append(time.time() - self.tik)
return self.times[-1]
def avg(self):
"""Return the average time."""
return sum(self.times) / len(self.times)
def sum(self):
"""Return the sum of time."""
return sum(self.times)
def cumsum(self):
"""Return the accumulated time."""
return jnp.array(self.times).cumsum().tolist()
class Accumulator:
"""For accumulating sums over `n` variables."""
def __init__(self, n):
self.data = [0.0] * n
def add(self, *args):
self.data = [a + float(b) for a, b in zip(self.data, args)]
def reset(self):
self.data = [0.0] * len(self.data)
def __getitem__(self, idx):
return self.data[idx]
def set_axes(axes, xlabel, ylabel, xlim, ylim, xscale, yscale, legend):
"""Set the axes for matplotlib."""
axes.set_xlabel(xlabel)
axes.set_ylabel(ylabel)
axes.set_xscale(xscale)
axes.set_yscale(yscale)
axes.set_xlim(xlim)
axes.set_ylim(ylim)
if legend:
axes.legend(legend)
axes.grid()
@jax.jit
def sgd(params, grads, lr, batch_size):
"""Minibatch stochastic gradient descent."""
params = jax.tree_map(lambda p, g: p - lr * g / batch_size, params, grads)
return params
@jax.jit
def train_step(apply_fn, loss_fn, params, state, X, Y):
def loss(params, state, X, Y):
y = Y.T.reshape(-1) # (B,T) -> (T,B)
y_hat, state = apply_fn(params, state, X)
y_hat = y_hat.reshape(-1, y_hat.shape[-1])
y_one_hot = jax.nn.one_hot(y, num_classes=y_hat.shape[-1])
return loss_fn(y_hat, y_one_hot).mean(), state
grad_fn = jax.value_and_grad(loss, has_aux=True)
(l, state), grads = grad_fn(params, state, X, Y)
grads = grad_clipping(grads, 1)
return l, state, grads
def train_epoch(net, params, train_iter, loss, updater, use_random_iter):
state, timer = None, Timer()
metric = Accumulator(2) # Sum of training loss, no. of tokens
if isinstance(updater, optax.GradientTransformation):
updater_state = updater.init(params)
# Convert to jax Partial functions for jax.jit compatibility
apply_fn = jax.tree_util.Partial(net.apply)
loss_fn = jax.tree_util.Partial(loss)
for X, Y in train_iter:
if state is None or use_random_iter:
# Initialize `state` when either it is the first iteration or
# using random sampling
state = net.begin_state(batch_size=X.shape[0])
l, state, grads = train_step(apply_fn, loss_fn, params, state, X, Y)
if isinstance(updater, optax.GradientTransformation):
updates, updater_state = updater.update(grads, updater_state)
params = optax.apply_updates(params, updates)
else:
# batch_size=1 since the `mean` function has been invoked
params = updater(params, grads, batch_size=1)
metric.add(l * Y.size, Y.size)
return params, math.exp(metric[0] / metric[1]), metric[1] / timer.stop()
def train(net, params, train_iter, vocab, lr, num_epochs, use_random_iter=False):
loss = optax.softmax_cross_entropy
animator = Animator(xlabel="epoch", ylabel="perplexity", legend=["train"], xlim=[10, num_epochs])
# Initialize
if isinstance(net, nn.Module):
updater = optax.sgd(lr)
else:
updater = lambda params, grads, batch_size: sgd(params, grads, lr, batch_size)
num_preds = 50
predict_ = lambda prefix: predict(prefix, num_preds, net, params, vocab)
# Train and predict
for epoch in range(num_epochs):
params, ppl, speed = train_epoch(net, params, train_iter, loss, updater, use_random_iter)
if (epoch + 1) % 10 == 0:
# Prediction takes time on the flax model
# print(predict_('time traveller'))
animator.add(epoch + 1, [ppl])
device = jax.default_backend()
print(f"perplexity {ppl:.1f}, {speed:.1f} tokens/sec on {device}")
print(predict_("time traveller"))
print(predict_("traveller"))
return params
def predict(prefix, num_preds, net, params, vocab):
"""Generate new characters following the `prefix`."""
state = net.begin_state(batch_size=1)
outputs = [vocab[prefix[0]]]
get_input = lambda: jnp.array([outputs[-1]]).reshape((1, 1))
for y in prefix[1:]: # Warm-up period
_, state = net.apply(params, state, get_input())
outputs.append(vocab[y])
for _ in range(num_preds): # Predict `num_preds` steps
y, state = net.apply(params, state, get_input())
y = y.reshape(-1, y.shape[-1])
outputs.append(int(y.argmax(axis=1).reshape(1)))
return "".join([vocab.idx_to_token[i] for i in outputs])
random.seed(0)
vocab_size, num_hiddens = len(vocab), 256
num_epochs, lr = 500, 1
model = RNNModelScratch(len(vocab), num_hiddens, get_params, init_gru_state, gru)
params = model.init_params(rng)
params = train(model, params, train_iter, vocab, lr, num_epochs)
class GRU(nn.Module):
@functools.partial(
nn.transforms.scan, variable_broadcast="params", in_axes=0, out_axes=0, split_rngs={"params": False}
)
@nn.compact
def __call__(self, state, x):
return nn.GRUCell()(state, x)
@staticmethod
def initialize_carry(rng, batch_dims, size):
return nn.GRUCell.initialize_carry(rng, batch_dims, size)
class RNNModel(nn.Module):
"""The RNN model."""
rnn: nn.Module
vocab_size: int
num_hiddens: int
bidirectional: bool = False
def setup(self):
# If the RNN is bidirectional (to be introduced later),
# `num_directions` should be 2, else it should be 1.
if not self.bidirectional:
self.num_directions = 1
else:
self.num_directions = 2
@nn.compact
def __call__(self, state, inputs):
X = jax.nn.one_hot(inputs.T, num_classes=self.vocab_size)
state, Y = self.rnn(state, X)
output = nn.Dense(self.vocab_size)(Y)
return output, state
def begin_state(self, batch_size=1):
# Use fixed random key since default state init fn is just `zeros`.
return self.rnn.initialize_carry(jax.random.PRNGKey(0), (batch_size,), num_hiddens)
random.seed(0)
gru_layer = GRU()
model = RNNModel(rnn=gru_layer, vocab_size=len(vocab), num_hiddens=num_hiddens)
initial_state = model.begin_state(batch_size)
params = model.init(rng, initial_state, jnp.ones([batch_size, num_steps]))
params = train(model, params, train_iter, vocab, lr, num_epochs)
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Datasets/Terrain/alos_global_dsm.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/Terrain/alos_global_dsm.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=Datasets/Terrain/alos_global_dsm.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/Terrain/alos_global_dsm.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geehydro](https://github.com/giswqs/geehydro). The **geehydro** Python package builds on the [folium](https://github.com/python-visualization/folium) package and implements several methods for displaying Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, `Map.centerObject()`, and `Map.setOptions()`.
The magic command `%%capture` can be used to hide output from a specific cell.
```
# %%capture
# !pip install earthengine-api
# !pip install geehydro
```
Import libraries
```
import ee
import folium
import geehydro
```
Authenticate and initialize Earth Engine API. You only need to authenticate the Earth Engine API once. Uncomment the line `ee.Authenticate()`
if you are running this notebook for this first time or if you are getting an authentication error.
```
# ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
This step creates an interactive map using [folium](https://github.com/python-visualization/folium). The default basemap is the OpenStreetMap. Additional basemaps can be added using the `Map.setOptions()` function.
The optional basemaps can be `ROADMAP`, `SATELLITE`, `HYBRID`, `TERRAIN`, or `ESRI`.
```
Map = folium.Map(location=[40, -100], zoom_start=4)
Map.setOptions('HYBRID')
```
## Add Earth Engine Python script
```
dataset = ee.Image('JAXA/ALOS/AW3D30_V1_1')
elevation = dataset.select('AVE')
elevationVis = {
'min': 0.0,
'max': 4000.0,
'palette': ['0000ff', '00ffff', 'ffff00', 'ff0000', 'ffffff'],
}
Map.setCenter(136.85, 37.37, 4)
Map.addLayer(elevation, elevationVis, 'Elevation')
```
## Display Earth Engine data layers
```
Map.setControlVisibility(layerControl=True, fullscreenControl=True, latLngPopup=True)
Map
```
| github_jupyter |
```
import json
import pandas as pd
import matplotlib.pyplot as plt
from xgboost import XGBRegressor
from sklearn.model_selection import train_test_split, KFold
from sklearn.metrics import mean_squared_error as MSE, r2_score
import math
import time
from xgboost import plot_importance
all_zones_df = pd.read_csv("../data/scsb_all_zones.csv")
zone_25_df = pd.read_csv("../data/scsb_zone_25.csv")
zone_26_df = pd.read_csv("../data/scsb_zone_26.csv")
zone_27_df = pd.read_csv("../data/scsb_zone_27.csv")
month_dependant_variables = ['jan_dist','feb_dist','mar_dist','apr_dist','may_dist','jun_dist','jul_dist','aug_dist','sep_dist','oct_dist','nov_dist','dec_dist']
data = zone_25_df
dependant_variable = 'mean_annual_runoff'
start_time = time.time()
# features_df = data[['annual_precipitation', 'drainage_area', 'median_elevation', \
# 'average_slope', 'glacial_coverage', 'potential_evapo_transpiration', dependant_variable]]
# zone 25 independant variables
# features_df = data[['annual_precipitation', 'glacial_coverage', 'potential_evapo_transpiration', dependant_variable]]
# zone 26 independant variables
# features_df = data[['median_elevation', 'glacial_coverage', 'annual_precipitation', 'potential_evapo_transpiration', dependant_variable]]
# zone 27 independant variables
# features_df = data[['median_elevation', 'annual_precipitation', 'average_slope', dependant_variable]]
features_df = data[['median_elevation', 'solar_exposure', 'glacial_coverage', 'annual_precipitation', 'potential_evapo_transpiration', 'average_slope', 'drainage_area', dependant_variable]]
X = features_df.drop([dependant_variable], axis=1)
y = features_df.get(dependant_variable)
X = X.tail(30)
y = y.tail(30)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.10, random_state=42)
# One Time Split
xgb = XGBRegressor(random_state=42)
xgb.fit(X_train, y_train)
xgb_pred = xgb.predict(X_test)
print('Score XGBoost Regressor', xgb.score(X_test, y_test))
plot_importance(xgb, title='One Time Split')
# plt.bar(range(len(xgb.feature_importances_)), xgb.feature_importances_)
# plt.show()
# KFold Validation
folds = 10
best_model = {}
max_r2, min_r2, acc_r2 = 0, math.inf, []
kfold = KFold(n_splits=folds, shuffle=True, random_state=42)
for train_index, test_index in kfold.split(X, y):
# print("--- %s seconds ---" % (time.time() - start_time))
print("TRAIN:", train_index, "TEST:", test_index)
X_train , X_test = X.iloc[train_index,:],X.iloc[test_index,:]
y_train , y_test = y[train_index] , y[test_index]
model = XGBRegressor(random_state=42)
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
score = model.score(X_test, y_test)
acc_r2.append(score)
if score > max_r2:
max_r2 = score
best_model = model
if score < min_r2:
min_r2 = score
avg_r2_score = sum(acc_r2)/folds
print('r2 of each fold - {}'.format(acc_r2))
print('R2 values:')
print("min:", min_r2)
print('avg: {}'.format(avg_r2_score))
print("max:", max_r2)
print("Feature Importances:", best_model.feature_importances_)
plot_importance(best_model, title='KFold Validation')
# plt.bar(range(len(best_model.feature_importances_)), best_model.feature_importances_)
plt.show()
```
| github_jupyter |
# "Decision Tree implementation in python"
> "Decision Tree implementation in python"
- toc: true
- comments: true
- categories: [machine learning]
- search_exclude: true
# Decision Tree
Predictive model in the tree (Acyclic Graph) form that maps inputs to its target value from root to leaf.
> Note: Greedy + Top down + Recursive partitioning
**Example**
[](https://mermaid-js.github.io/mermaid-live-editor/#/edit/eyJjb2RlIjoiZ3JhcGggVERcblx0QVtwZXRhbF93aWR0aCA-IDEuNzVdIC0tPiB8bm98IEIoSXJpcyB2ZXJzaWNvbG9yKVxuXHRBIC0tPiB8eWVzfCBDKElyaXMgVmVyZ2luaWNhKVxuXG4iLCJtZXJtYWlkIjp7InRoZW1lIjoiZGVmYXVsdCJ9LCJ1cGRhdGVFZGl0b3IiOmZhbHNlfQ)
petal_width : feature
Iris versicolor, Iris Verginica: labels
# How do we construct tree
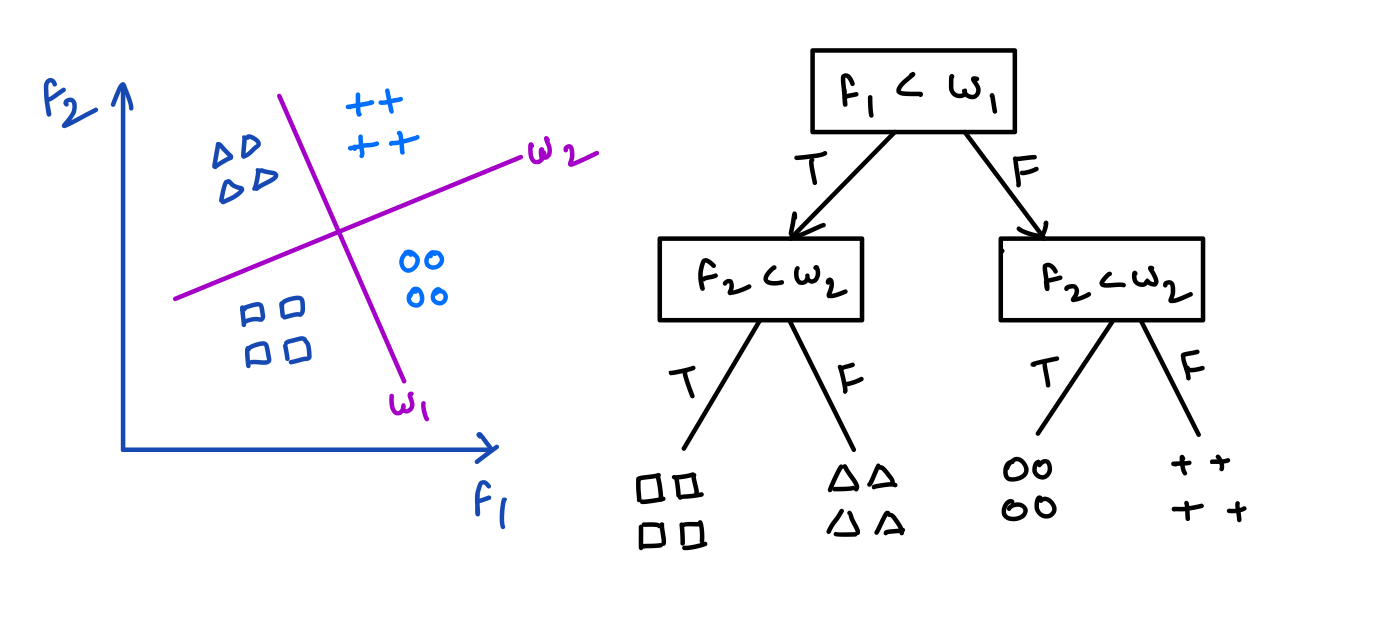
# when good choice ?
- output is discrete
- no large data
- noise in data
- categories or classes are disjoint
# Types
[](https://mermaid-js.github.io/mermaid-live-editor/#/edit/eyJjb2RlIjoiZ3JhcGggVERcblx0QVtEZWNpc2lvbiBUcmVlXSAtLT4gQihDbGFzc2lmaWNhdGlvbiB0cmVlKVxuXHRBIC0tPiBDKFJlZ3Jlc3Npb24gdHJlZSlcblxuIiwibWVybWFpZCI6eyJ0aGVtZSI6ImRlZmF1bHQifSwidXBkYXRlRWRpdG9yIjpmYWxzZX0)
Regression trees are used when dependent variable is continous. Classification trees are used when dependent variable is categorical.
# Family of decision tree algo's
- ID3
- c4.5
- c5.0
- CART (uses Gini index to create splits)
# Impurity functions
Measure how pure a label set of leaf nodes.
- Entropy based Measure
- Gini Measure
## Entropy
Entropy is a measure of disorder in the dataset (or) how much varied the data is.
Lets say we have dataset of N items .... $S=\left \{ \left ( \mathbf{x}_1,y_1 \right ),\dots,\left ( \mathbf{x}_n,y_n \right ) \right \}$
These N items should be divided into c categories or classes represented by $y_i$
$y_i\in\left \{ 1,\dots,c \right \}$, where $c$ is the number of classes
The entropy of our dataset is given by equation...
$H=-\sum_{i=1}^{C} p\left(x_{i}\right) \log _{2} p\left(x_{i}\right)$
where $p\left(x_{i}\right)$ = ratios of elements of each label $i$ in our dataset.
$C$ = No of class items in our data.
The value of entropy always lies between 0 to 1. 1 being worst and 0 being best.
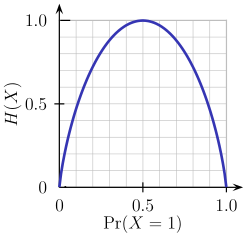
Let's see with an example.
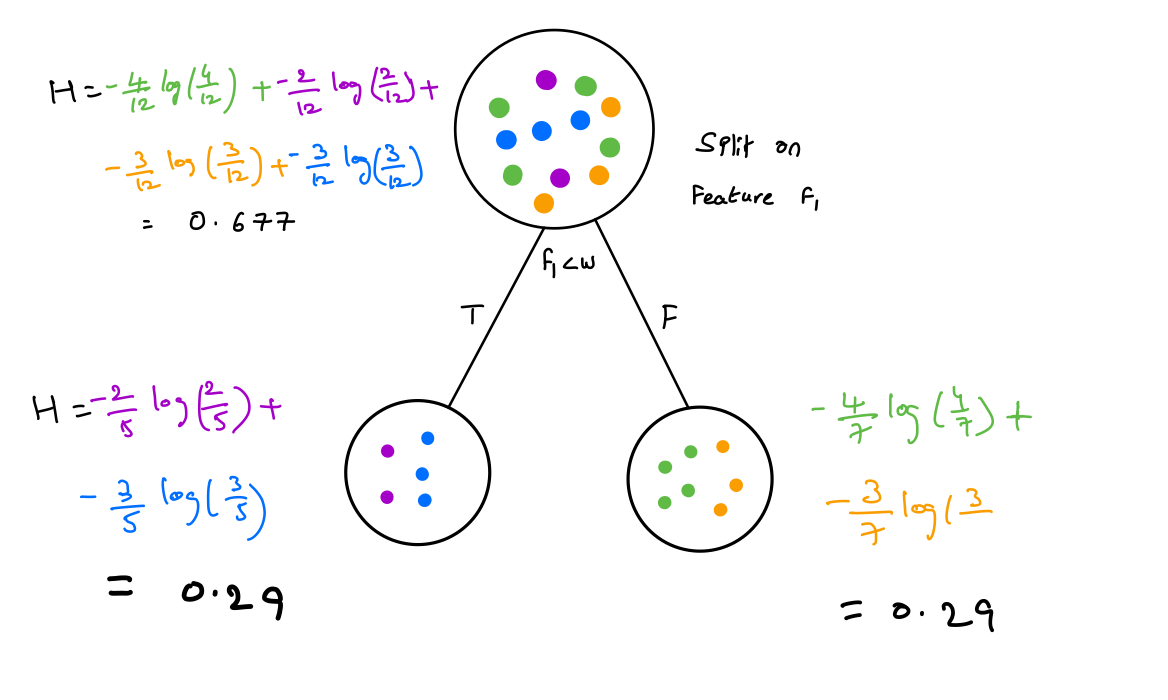
## Information Gain(IG)
Measure of decrease in disorder with the help of splitting the original dataset.
In other words, its the difference between parent node impurity and weighted child node impurity
Based on this value, we split the node and build decision tree.
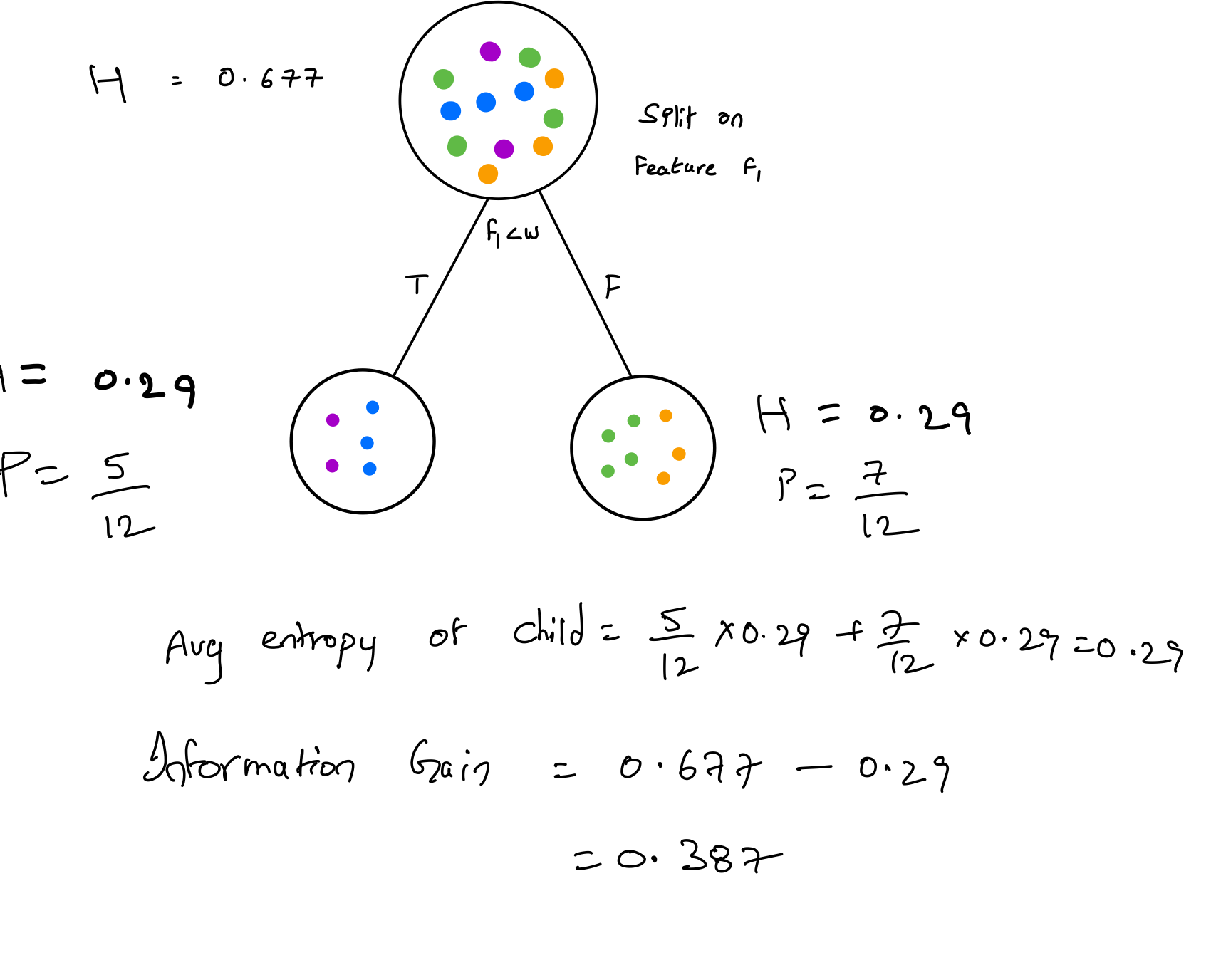
## Gini Impurity
$G=\sum_{i=1}^{C} p_i *(1-p_i)$
(or)
$G = 1-\sum_{i=1}^{C}\left(p_{i}\right)^{2}$
where
C : no of classes (or) target variables
($p_i$) is the probability of class i in a node
# Avoid overfitting Pruning & constrains
- Technique that reduces size of tree by removing sub tree's that provide little value to classification
- reduces overfitting chance
- setting constrains on controlling depth
- max_depth
- min_samples_leaf: min num of samples a leaf can have to avoid furthur splitting
- max_leaf_nodes: limits total leaves in a tree
# Comparision of decision tree algorithms

# How it works
Lets see how one of the algorithm(ID3) which is used to calculate decision tree's work.
[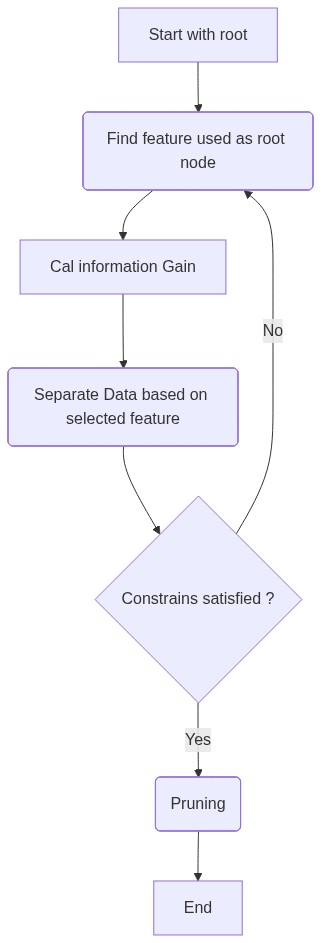](https://mermaid-js.github.io/mermaid-live-editor/#/edit/eyJjb2RlIjoiZ3JhcGggVERcblx0QVtTdGFydCB3aXRoIHJvb3RdIC0tPiBCKEZpbmQgZmVhdHVyZSB1c2VkIGFzIHJvb3Qgbm9kZSlcblx0QiAtLT4gQyh0ZXN0IGF0dHJpYnV0ZSlcbiAgICBDW0NhbCBpbmZvcm1hdGlvbiBHYWluXSAgLS0-IEQoU2VwYXJhdGUgRGF0YSBiYXNlZCBvbiBzZWxlY3RlZCBmZWF0dXJlKVxuICAgIEQgLS0-IENDe0NvbnN0cmFpbnMgc2F0aXNmaWVkID99XG4gICAgQ0MgLS0-IHxOb3wgQlxuICAgIENDIC0tPiB8WWVzfCBQKFBydW5pbmcpXG4gICAgUCAtLT4gRW5kXG5cbiIsIm1lcm1haWQiOnsidGhlbWUiOiJkZWZhdWx0In0sInVwZGF0ZUVkaXRvciI6ZmFsc2V9)
# Implementation in python
```
# calculate entropy
import numpy as np
from collections import Counter
def entropy(y):
histo = np.bincount(y)
ps = histo/len(y)
entropy = -np.sum([p * np.log2(p) for p in ps if p > 0])
# Define decision node
class DecisionNode:
def __init__(self, feature=None, threshold=None, left=None, right=None, *, value=None):
self.feature = feature
self.threshold = threshold
self.left = left
self.right = right
self.value = value
def isLeafNode(self):
return self.value is not None
# Decision tree class
class DecisionTree:
def __init__(self, min_sample_split=2, max_depth=100, n_features=None):
self.min_sample_split = min_sample_split
self.max_depth = max_depth
self.n_features = n_features
self.root = None
def fit(self, X, y):
self.root = self.grow_tree(X, y)
def grow_tree(self, X, y, depth=0):
n_samples, n_features = X.shape
n_labels = len(np.unique(y))
# stopping criteria
if(depth >= self.max_depth or n_labels == 1 or n_samples < self.min_sample_split):
leaf_value = self.most_common_label(y)
return DecisionNode(value=leaf_value)
features_idxs = np.random.choice(n_features, self.n_features, replace=False)
# greedy search
best_feature, best_threshold = self.best_criteria(X, y , features_idxs)
left_idxs, right_idxs = self.split(X[:, best_feature], best_threshold)
left = self.grow_tree(X[left_idxs, :], y[left_idxs], depth+1)
right = self.grow_tree(X[right_idxs, :], y[right_idxs], depth+1)
return Node(best_feature, best_threshold, left, right)
def predict(self, X):
# traverse tree
return np.array([self._traverse_tree(x) for x in X], self.root)
def _traverse_tree(self, x, node):
if node.isLeafNode():
return node.value
if x[node.feature_idx] <= node.thresholds:
return self._traverse_tree(x, node.left)
return self._traverse_tree(x, node.right)
def most_common_label(self, y):
counter = Counter(y)
most_common = counter.most_common(1)[0]
return most_common
def best_criteria(self, X, y, feature_idxs):
best_gain = -1
split_idx, split_threshold = None, None
for feature_idx in feature_idxs:
X_column = X[:, feature_idx]
thresholds = np.unique(X_column)
for threshold in thresholds:
gain = self.information_gain(y, X_column, threshold)
if gain > best_gain:
best_gain = gain
split_idx = feature_idx
split_threshold = threshold
return split_idx, split_threshold
def information_gain(self, y, X_column, split_threshold):
parent_entropy = entropy(y)
# generate splits
left_idxs, right_idxs = self.split(X_column, split_threshold)
if len(left_idxs) == 0 or len(right_idxs) == 0:
return 0
# weighted avg child E
n = len(y)
n_l, n_r = len(left_idxs), len(right_idxs)
e_l, e_r = entropy(y[left_idxs], entropy[y[right_idxs]])
child_entropy = (n_l/n) * e_l + (n_r/n) * e_r
# return ig
ig = parent_entropy - child_entropy
return ig
def split(self, X_column, split_threshold):
left_idxs = np.argwhere(X_column <= split_threshold).flatten()
right_idxs = np.argwhere(X_column > split_threshold).flatten()
return left_idxs, right_idxs
```
# References
- [U of waterloo](https://www.youtube.com/watch?v=E4HFVAjhQWQ&feature=youtu.be)
- [Ytube](https://www.youtube.com/watch?v=Bqi7EFFvNOg&t=5s)
- An Overview of Classification Algorithm in Data mining
Sampath K. Kumar, Panneerselvam Kiruthika
| github_jupyter |
```
from __future__ import division
import random
import pprint
import sys
import time
import numpy as np
from optparse import OptionParser
import pickle
from keras import backend as K
from keras.optimizers import Adam, SGD, RMSprop
from keras.layers import Input
from keras.models import Model
from rcnn import config, data_generators
from rcnn import losses as losses
import rcnn.roi_helpers as roi_helpers
from keras.utils import generic_utils
from keras.layers import TimeDistributed, Lambda
import tensorflow as tf
from rcnn.clstm import clstm
sess = tf.Session()
K.set_session(sess)
sys.setrecursionlimit(40000)
parser = OptionParser()
video_path = './videos'
annotation_path = './annotations'
num_rois = 32
num_epochs = 2000
config_filename = 'config.pickle'
output_weight_path = './model_frcnn.hdf5'
input_weight_path = None
from rcnn.video_parser import get_data
C = config.Config()
C.use_horizontal_flips = False
C.use_vertical_flips = False
C.rot_90 = False
C.model_path = output_weight_path
C.num_rois = int(num_rois)
from rcnn import simple_nn as nn
C.network = 'simple_nn'
# check if weight path was passed via command line
if input_weight_path:
C.base_net_weights = input_weight_path
all_videos, classes_count, class_mapping = get_data(video_path, annotation_path)
if 'bg' not in classes_count:
classes_count['bg'] = 0
class_mapping['bg'] = len(class_mapping)
C.class_mapping = class_mapping
inv_map = {v: k for k, v in class_mapping.items()}
print('Training images per class:')
pprint.pprint(classes_count)
print('Num classes (including bg) = {}'.format(len(classes_count)))
config_output_filename = config_filename
with open(config_output_filename, 'wb') as config_f:
pickle.dump(C,config_f)
print('Config has been written to {}, and can be loaded when testing to ensure correct results'.format(config_output_filename))
random.shuffle(all_videos)
num_imgs = len(all_videos)
#train_videos = [s for s in all_videos if s['imageset'] == 'trainval']
#val_videos = [s for s in all_videos if s['imageset'] == 'test']
train_videos = all_videos
val_videos = all_videos
print('Num train samples {}'.format(len(train_videos)))
print('Num val samples {}'.format(len(val_videos)))
data_gen_train = data_generators.video_streamer(train_videos, classes_count, C, nn.get_img_output_length, K.image_dim_ordering(), mode='train')
data_gen_val = data_generators.video_streamer(val_videos, classes_count, C, nn.get_img_output_length,K.image_dim_ordering(), mode='val')
input_shape_img = (None, None, None, 3)
num_anchors = len(C.anchor_box_scales) * len(C.anchor_box_ratios)
video_input = tf.placeholder(tf.float32, [None,None,None,None,3])
rpn_target_cls = tf.placeholder(tf.float32, [None,None,None,None,2*num_anchors])
rpn_target_reg = tf.placeholder(tf.float32, [None,None,None,None,2*num_anchors*4])
#roi_input = Input(shape=(None, None, 4))
nb_clstm_filter = 40
def time_broadcast(f, x):
shape = tf.shape(x)
num_videos, num_frames, w, h, c = [shape[i] for i in range(5)]
time_flat = tf.reshape(x, [-1, w,h,c])
y = f(time_flat)
shape = tf.shape(y)
_, w, h, c = [shape[i] for i in range(4)]
y = tf.reshape(y, [num_videos, num_frames, w, h, c])
return y
def build_shared(video_input):
with tf.name_scope('shared_layers'):
base = nn.nn_base(trainable=True)
shared_layers = time_broadcast(base, video_input)
num_channels = 64
shared_layers = clstm(shared_layers,num_channels,nb_clstm_filter,3)
return shared_layers
shared = build_shared(video_input)
def build_rpn(x):
with tf.name_scope('RPN'):
shape = tf.shape(shared)
num_videos, num_frames, w, h, c = [shape[i] for i in range(5)]
c = nb_clstm_filter
time_flat = tf.reshape(x, [-1, w,h,c])
y_cls, y_reg, _ = nn.rpn(num_anchors)(time_flat)
shape = tf.shape(y_cls)
_, w, h, c = [shape[i] for i in range(4)]
y_cls = tf.reshape(y_cls, [num_videos, num_frames, w, h, c])
y_reg = tf.reshape(y_reg, [num_videos, num_frames, w, h, c*4])
return y_cls, y_reg
rpn = build_rpn(shared)
#classifier = nn.classifier(shared_layers, roi_input, C.num_rois, nb_classes=len(classes_count), trainable=True)
#model_rpn = Model(img_input, rpn[:2])
#model_classifier = Model([img_input, roi_input], classifier)
# this is a model that holds both the RPN and the classifier, used to load/save weights for the models
#model_all = Model([img_input, roi_input], rpn[:2] + classifier)
optimizer = tf.train.AdamOptimizer(0.001)
rpn_loss = losses.rpn_loss_regr(num_anchors)(rpn_target_reg, rpn[1]) \
+ losses.rpn_loss_cls(num_anchors)(rpn_target_cls, rpn[0])
rpn_train_op = optimizer.minimize(rpn_loss)
def run_rpn(X, Y):
sess.run(rpn_train_op, {video_input: X, rpn_target_cls: Y[0], rpn_target_reg: Y[1]})
#model_rpn.compile(optimizer=optimizer, loss=[losses.rpn_loss_cls(num_anchors), losses.rpn_loss_regr(num_anchors)])
#model_classifier.compile(optimizer=optimizer_classifier, loss=[losses.class_loss_cls, losses.class_loss_regr(len(classes_count)-1)], metrics={'dense_class_{}'.format(len(classes_count)): 'accuracy'})
#model_all.compile(optimizer='sgd', loss='mae')
epoch_length = 1000
num_epochs = int(num_epochs)
iter_num = 0
losses = np.zeros((epoch_length, 5))
rpn_accuracy_rpn_monitor = []
rpn_accuracy_for_epoch = []
start_time = time.time()
best_loss = np.Inf
class_mapping_inv = {v: k for k, v in class_mapping.items()}
print('Starting training')
vis = True
from keras.layers import Convolution2D
Convolution2D()
import rcnn
import rcnn.data_augment as data_augment
img_data_aug, x_img = data_augment.augment(all_videos[0][0], C, False)
img_data_aug
x_img.shape
all_videos[0][0]
from rcnn.data_generators import get_new_img_size, calc_rpn, get_anchor
import cv2
anc = get_anchor(all_videos[0][0], 2, C, lambda x,y: [x,y], 'tf', mode='val')
C.
cls, reg = calc_rpn(C, all_videos[0][0], 320, 320, 320, 320, lambda x,y: [x,y])
all_videos[0][0]
X.shape
anc[0][0].shape, anc[1][0][0].shape
from matplotlib import pyplot as plt
plt.imshow(anc[0][0])
plt.show()
plt.imshow(anc[1][0][0].sum(axis=-1))#[:30, :30])
plt.show()
from __future__ import absolute_import
import numpy as np
import cv2
import random
import copy
from . import data_augment
import threading
import itertools
def union(au, bu, area_intersection):
area_a = (au[2] - au[0]) * (au[3] - au[1])
area_b = (bu[2] - bu[0]) * (bu[3] - bu[1])
area_union = area_a + area_b - area_intersection
return area_union
def intersection(ai, bi):
x = max(ai[0], bi[0])
y = max(ai[1], bi[1])
w = min(ai[2], bi[2]) - x
h = min(ai[3], bi[3]) - y
if w < 0 or h < 0:
return 0
return w*h
def iou(a, b):
# a and b should be (x1,y1,x2,y2)
if a[0] >= a[2] or a[1] >= a[3] or b[0] >= b[2] or b[1] >= b[3]:
return 0.0
area_i = intersection(a, b)
area_u = union(a, b, area_i)
return float(area_i) / float(area_u + 1e-6)
def get_new_img_size(width, height, img_min_side=600):
if width <= height:
f = float(img_min_side) / width
resized_height = int(f * height)
resized_width = img_min_side
else:
f = float(img_min_side) / height
resized_width = int(f * width)
resized_height = img_min_side
return resized_width, resized_height
class SampleSelector:
def __init__(self, class_count):
# ignore classes that have zero samples
self.classes = [b for b in class_count.keys() if class_count[b] > 0]
self.class_cycle = itertools.cycle(self.classes)
self.curr_class = next(self.class_cycle)
def skip_sample_for_balanced_class(self, img_data):
class_in_img = False
for bbox in img_data['bboxes']:
cls_name = bbox['class']
if cls_name == self.curr_class:
class_in_img = True
self.curr_class = next(self.class_cycle)
break
if class_in_img:
return False
else:
return True
def calc_rpn(C, img_data, width, height, resized_width, resized_height, img_length_calc_function):
downscale = float(C.rpn_stride)
anchor_sizes = C.anchor_box_scales
anchor_ratios = C.anchor_box_ratios
num_anchors = len(anchor_sizes) * len(anchor_ratios)
# calculate the output map size based on the network architecture
(output_width, output_height) = img_length_calc_function(resized_width, resized_height)
n_anchratios = len(anchor_ratios)
# initialise empty output objectives
y_rpn_overlap = np.zeros((output_height, output_width, num_anchors))
y_is_box_valid = np.zeros((output_height, output_width, num_anchors))
y_rpn_regr = np.zeros((output_height, output_width, num_anchors * 4))
num_bboxes = len(img_data['bboxes'])
num_anchors_for_bbox = np.zeros(num_bboxes).astype(int)
best_anchor_for_bbox = -1*np.ones((num_bboxes, 4)).astype(int)
best_iou_for_bbox = np.zeros(num_bboxes).astype(np.float32)
best_x_for_bbox = np.zeros((num_bboxes, 4)).astype(int)
best_dx_for_bbox = np.zeros((num_bboxes, 4)).astype(np.float32)
# get the GT box coordinates, and resize to account for image resizing
gta = np.zeros((num_bboxes, 4))
for bbox_num, bbox in enumerate(img_data['bboxes']):
# get the GT box coordinates, and resize to account for image resizing
gta[bbox_num, 0] = bbox['x1'] * (resized_width / float(width))
gta[bbox_num, 1] = bbox['x2'] * (resized_width / float(width))
gta[bbox_num, 2] = bbox['y1'] * (resized_height / float(height))
gta[bbox_num, 3] = bbox['y2'] * (resized_height / float(height))
# rpn ground truth
for anchor_size_idx in range(len(anchor_sizes)):
for anchor_ratio_idx in range(n_anchratios):
anchor_x = anchor_sizes[anchor_size_idx] * anchor_ratios[anchor_ratio_idx][0]
anchor_y = anchor_sizes[anchor_size_idx] * anchor_ratios[anchor_ratio_idx][1]
for ix in range(output_width):
# x-coordinates of the current anchor box
x1_anc = downscale * (ix + 0.5) - anchor_x / 2
x2_anc = downscale * (ix + 0.5) + anchor_x / 2
# ignore boxes that go across image boundaries
if x1_anc < 0 or x2_anc > resized_width:
continue
for jy in range(output_height):
# y-coordinates of the current anchor box
y1_anc = downscale * (jy + 0.5) - anchor_y / 2
y2_anc = downscale * (jy + 0.5) + anchor_y / 2
# ignore boxes that go across image boundaries
if y1_anc < 0 or y2_anc > resized_height:
continue
# bbox_type indicates whether an anchor should be a target
bbox_type = 'neg'
# this is the best IOU for the (x,y) coord and the current anchor
# note that this is different from the best IOU for a GT bbox
best_iou_for_loc = 0.0
for bbox_num in range(num_bboxes):
# get IOU of the current GT box and the current anchor box
curr_iou = iou([gta[bbox_num, 0], gta[bbox_num, 2], gta[bbox_num, 1], gta[bbox_num, 3]], [x1_anc, y1_anc, x2_anc, y2_anc])
# calculate the regression targets if they will be needed
if curr_iou > best_iou_for_bbox[bbox_num] or curr_iou > C.rpn_max_overlap:
cx = (gta[bbox_num, 0] + gta[bbox_num, 1]) / 2.0
cy = (gta[bbox_num, 2] + gta[bbox_num, 3]) / 2.0
cxa = (x1_anc + x2_anc)/2.0
cya = (y1_anc + y2_anc)/2.0
tx = (cx - cxa) / (x2_anc - x1_anc)
ty = (cy - cya) / (y2_anc - y1_anc)
tw = np.log((gta[bbox_num, 1] - gta[bbox_num, 0]) / (x2_anc - x1_anc))
th = np.log((gta[bbox_num, 3] - gta[bbox_num, 2]) / (y2_anc - y1_anc))
if img_data['bboxes'][bbox_num]['class'] != 'bg':
# all GT boxes should be mapped to an anchor box, so we keep track of which anchor box was best
if curr_iou > best_iou_for_bbox[bbox_num]:
best_anchor_for_bbox[bbox_num] = [jy, ix, anchor_ratio_idx, anchor_size_idx]
best_iou_for_bbox[bbox_num] = curr_iou
best_x_for_bbox[bbox_num,:] = [x1_anc, x2_anc, y1_anc, y2_anc]
best_dx_for_bbox[bbox_num,:] = [tx, ty, tw, th]
# we set the anchor to positive if the IOU is >0.7 (it does not matter if there was another better box, it just indicates overlap)
if curr_iou > C.rpn_max_overlap:
bbox_type = 'pos'
num_anchors_for_bbox[bbox_num] += 1
# we update the regression layer target if this IOU is the best for the current (x,y) and anchor position
if curr_iou > best_iou_for_loc:
best_iou_for_loc = curr_iou
best_regr = (tx, ty, tw, th)
# if the IOU is >0.3 and <0.7, it is ambiguous and no included in the objective
if C.rpn_min_overlap < curr_iou < C.rpn_max_overlap:
# gray zone between neg and pos
if bbox_type != 'pos':
bbox_type = 'neutral'
# turn on or off outputs depending on IOUs
if bbox_type == 'neg':
y_is_box_valid[jy, ix, anchor_ratio_idx + n_anchratios * anchor_size_idx] = 1
y_rpn_overlap[jy, ix, anchor_ratio_idx + n_anchratios * anchor_size_idx] = 0
elif bbox_type == 'neutral':
y_is_box_valid[jy, ix, anchor_ratio_idx + n_anchratios * anchor_size_idx] = 0
y_rpn_overlap[jy, ix, anchor_ratio_idx + n_anchratios * anchor_size_idx] = 0
elif bbox_type == 'pos':
y_is_box_valid[jy, ix, anchor_ratio_idx + n_anchratios * anchor_size_idx] = 1
y_rpn_overlap[jy, ix, anchor_ratio_idx + n_anchratios * anchor_size_idx] = 1
start = 4 * (anchor_ratio_idx + n_anchratios * anchor_size_idx)
y_rpn_regr[jy, ix, start:start+4] = best_regr
# we ensure that every bbox has at least one positive RPN region
for idx in range(num_anchors_for_bbox.shape[0]):
if num_anchors_for_bbox[idx] == 0:
# no box with an IOU greater than zero ...
if best_anchor_for_bbox[idx, 0] == -1:
continue
y_is_box_valid[
best_anchor_for_bbox[idx,0], best_anchor_for_bbox[idx,1], best_anchor_for_bbox[idx,2] + n_anchratios *
best_anchor_for_bbox[idx,3]] = 1
y_rpn_overlap[
best_anchor_for_bbox[idx,0], best_anchor_for_bbox[idx,1], best_anchor_for_bbox[idx,2] + n_anchratios *
best_anchor_for_bbox[idx,3]] = 1
start = 4 * (best_anchor_for_bbox[idx,2] + n_anchratios * best_anchor_for_bbox[idx,3])
y_rpn_regr[
best_anchor_for_bbox[idx,0], best_anchor_for_bbox[idx,1], start:start+4] = best_dx_for_bbox[idx, :]
y_rpn_overlap = np.transpose(y_rpn_overlap, (2, 0, 1))
y_rpn_overlap = np.expand_dims(y_rpn_overlap, axis=0)
y_is_box_valid = np.transpose(y_is_box_valid, (2, 0, 1))
y_is_box_valid = np.expand_dims(y_is_box_valid, axis=0)
y_rpn_regr = np.transpose(y_rpn_regr, (2, 0, 1))
y_rpn_regr = np.expand_dims(y_rpn_regr, axis=0)
pos_locs = np.where(np.logical_and(y_rpn_overlap[0, :, :, :] == 1, y_is_box_valid[0, :, :, :] == 1))
neg_locs = np.where(np.logical_and(y_rpn_overlap[0, :, :, :] == 0, y_is_box_valid[0, :, :, :] == 1))
num_pos = len(pos_locs[0])
# one issue is that the RPN has many more negative than positive regions, so we turn off some of the negative
# regions. We also limit it to 256 regions.
num_regions = 256
if len(pos_locs[0]) > num_regions/2:
val_locs = random.sample(range(len(pos_locs[0])), len(pos_locs[0]) - num_regions/2)
y_is_box_valid[0, pos_locs[0][val_locs], pos_locs[1][val_locs], pos_locs[2][val_locs]] = 0
num_pos = num_regions/2
if len(neg_locs[0]) + num_pos > num_regions:
val_locs = random.sample(range(len(neg_locs[0])), len(neg_locs[0]) - num_pos)
y_is_box_valid[0, neg_locs[0][val_locs], neg_locs[1][val_locs], neg_locs[2][val_locs]] = 0
y_rpn_cls = np.concatenate([y_is_box_valid, y_rpn_overlap], axis=1)
y_rpn_regr = np.concatenate([np.repeat(y_rpn_overlap, 4, axis=1), y_rpn_regr], axis=1)
return np.copy(y_rpn_cls), np.copy(y_rpn_regr)
class threadsafe_iter:
"""Takes an iterator/generator and makes it thread-safe by
serializing call to the `next` method of given iterator/generator.
"""
def __init__(self, it):
self.it = it
self.lock = threading.Lock()
def __iter__(self):
return self
def next(self):
with self.lock:
return next(self.it)
def threadsafe_generator(f):
"""A decorator that takes a generator function and makes it thread-safe.
"""
def g(*a, **kw):
return threadsafe_iter(f(*a, **kw))
return g
def get_anchor_gt(all_img_data, class_count, C, img_length_calc_function, backend, mode='train'):
# The following line is not useful with Python 3.5, it is kept for the legacy
# all_img_data = sorted(all_img_data)
while True:
for img_data in all_img_data:
try:
# read in image, and optionally add augmentation
if mode == 'train':
img_data_aug, x_img = data_augment.augment(img_data, C, augment=True)
else:
img_data_aug, x_img = data_augment.augment(img_data, C, augment=False)
(width, height) = (img_data_aug['width'], img_data_aug['height'])
(rows, cols, _) = x_img.shape
assert cols == width
assert rows == height
# get image dimensions for resizing
(resized_width, resized_height) = get_new_img_size(width, height, C.im_size)
# resize the image so that smalles side is length = 600px
x_img = cv2.resize(x_img, (resized_width, resized_height), interpolation=cv2.INTER_CUBIC)
try:
y_rpn_cls, y_rpn_regr = calc_rpn(C, img_data_aug, width, height, resized_width, resized_height, img_length_calc_function)
except:
continue
# Zero-center by mean pixel, and preprocess image
x_img = x_img[:,:, (2, 1, 0)] # BGR -> RGB
x_img = x_img.astype(np.float32)
x_img[:, :, 0] -= C.img_channel_mean[0]
x_img[:, :, 1] -= C.img_channel_mean[1]
x_img[:, :, 2] -= C.img_channel_mean[2]
x_img /= C.img_scaling_factor
x_img = np.transpose(x_img, (2, 0, 1))
x_img = np.expand_dims(x_img, axis=0)
y_rpn_regr[:, y_rpn_regr.shape[1]//2:, :, :] *= C.std_scaling
if backend == 'tf':
x_img = np.transpose(x_img, (0, 2, 3, 1))
y_rpn_cls = np.transpose(y_rpn_cls, (0, 2, 3, 1))
y_rpn_regr = np.transpose(y_rpn_regr, (0, 2, 3, 1))
return np.copy(x_img), [np.copy(y_rpn_cls), np.copy(y_rpn_regr)], img_data_aug
except Exception as e:
print(e)
continue
Y[1].shape
init = tf.global_variables_initializer()
sess.run(init)
run_rpn(X, Y)
print('Now gonna start training!')
for epoch_num in range(num_epochs):
progbar = generic_utils.Progbar(epoch_length)
print('Epoch {}/{}'.format(epoch_num + 1, num_epochs))
while True:
try:
if len(rpn_accuracy_rpn_monitor) == epoch_length and C.verbose:
mean_overlapping_bboxes = float(sum(rpn_accuracy_rpn_monitor))/len(rpn_accuracy_rpn_monitor)
rpn_accuracy_rpn_monitor = []
print('Average number of overlapping bounding boxes from RPN = {} for {} previous iterations'.format(mean_overlapping_bboxes, epoch_length))
if mean_overlapping_bboxes == 0:
print('RPN is not producing bounding boxes that overlap the ground truth boxes. Check RPN settings or keep training.')
print('Now gonna generate data!')
X, Y, img_data = next(data_gen_train)
print('Now gonna run train op!')
#loss_rpn = model_rpn.train_on_batch(X, Y)
run_rpn(X,Y)
print('Success!')
#P_rpn = model_rpn.predict_on_batch(X)
R = roi_helpers.rpn_to_roi(P_rpn[0], P_rpn[1], C, K.image_dim_ordering(), use_regr=True, overlap_thresh=0.7, max_boxes=300)
# note: calc_iou converts from (x1,y1,x2,y2) to (x,y,w,h) format
X2, Y1, Y2, IouS = roi_helpers.calc_iou(R, img_data, C, class_mapping)
if X2 is None:
rpn_accuracy_rpn_monitor.append(0)
rpn_accuracy_for_epoch.append(0)
continue
neg_samples = np.where(Y1[0, :, -1] == 1)
pos_samples = np.where(Y1[0, :, -1] == 0)
if len(neg_samples) > 0:
neg_samples = neg_samples[0]
else:
neg_samples = []
if len(pos_samples) > 0:
pos_samples = pos_samples[0]
else:
pos_samples = []
rpn_accuracy_rpn_monitor.append(len(pos_samples))
rpn_accuracy_for_epoch.append((len(pos_samples)))
use_detector = False
if use_detector: #for first runs, do not use detection model
if C.num_rois > 1:
if len(pos_samples) < C.num_rois//2:
selected_pos_samples = pos_samples.tolist()
else:
selected_pos_samples = np.random.choice(pos_samples, C.num_rois//2, replace=False).tolist()
try:
selected_neg_samples = np.random.choice(neg_samples, C.num_rois - len(selected_pos_samples), replace=False).tolist()
except:
selected_neg_samples = np.random.choice(neg_samples, C.num_rois - len(selected_pos_samples), replace=True).tolist()
sel_samples = selected_pos_samples + selected_neg_samples
else:
# in the extreme case where num_rois = 1, we pick a random pos or neg sample
selected_pos_samples = pos_samples.tolist()
selected_neg_samples = neg_samples.tolist()
if np.random.randint(0, 2):
sel_samples = random.choice(neg_samples)
else:
sel_samples = random.choice(pos_samples)
loss_class = model_classifier.train_on_batch([X, X2[:, sel_samples, :]], [Y1[:, sel_samples, :], Y2[:, sel_samples, :]])
losses[iter_num, 0] = loss_rpn[1]
losses[iter_num, 1] = loss_rpn[2]
if use_detector:
losses[iter_num, 2] = loss_class[1]
losses[iter_num, 3] = loss_class[2]
losses[iter_num, 4] = loss_class[3]
iter_num += 1
if use_detector:
progbar.update(iter_num, [('rpn_cls', np.mean(losses[:iter_num, 0])), ('rpn_regr', np.mean(losses[:iter_num, 1])), ('detector_cls', np.mean(losses[:iter_num, 2])), ('detector_regr', np.mean(losses[:iter_num, 3]))])
else:
progbar.update(iter_num, [('rpn_cls', np.mean(losses[:iter_num, 0])), ('rpn_regr', np.mean(losses[:iter_num, 1]))])
if iter_num == epoch_length:
loss_rpn_cls = np.mean(losses[:, 0])
loss_rpn_regr = np.mean(losses[:, 1])
if use_detector:
loss_class_cls = np.mean(losses[:, 2])
loss_class_regr = np.mean(losses[:, 3])
class_acc = np.mean(losses[:, 4])
mean_overlapping_bboxes = float(sum(rpn_accuracy_for_epoch)) / len(rpn_accuracy_for_epoch)
rpn_accuracy_for_epoch = []
if C.verbose:
print('Mean number of bounding boxes from RPN overlapping ground truth boxes: {}'.format(mean_overlapping_bboxes))
print('Classifier accuracy for bounding boxes from RPN: {}'.format(class_acc))
print('Loss RPN classifier: {}'.format(loss_rpn_cls))
print('Loss RPN regression: {}'.format(loss_rpn_regr))
if use_detector:
print('Loss Detector classifier: {}'.format(loss_class_cls))
print('Loss Detector regression: {}'.format(loss_class_regr))
print('Elapsed time: {}'.format(time.time() - start_time))
if not use_detector:
loss_class_cls = 0
loss_class_regr = 0
curr_loss = loss_rpn_cls + loss_rpn_regr + loss_class_cls + loss_class_regr
iter_num = 0
start_time = time.time()
if curr_loss < best_loss:
if C.verbose:
print('Total loss decreased from {} to {}, saving weights'.format(best_loss,curr_loss))
best_loss = curr_loss
model_all.save_weights(C.model_path)
break
except Exception as e:
print('Exception: {}'.format(e))
continue
print('Training complete, exiting.')
X, Y, data = next(data_gen)
Y[1].shape
from matplotlib import pyplot as plt
plt.imshow(X[0][7])
plt.show()
plt.imshow(Y[1][0][7].sum(axis=-1))
plt.show()
from keras.layers import TimeDistributed
d = get_data('./videos/', './annotations/')
d[0][0]
len(d[0])
```
| github_jupyter |
# Runtime ≈ 1 minute
# This notebook completes the process of wrangling the text for EDA and other future analyses.
# The processing is the following order:
* Scispacy - Acronyms
* General Cleaning
* Spacy - Lemmatization
```
try:
from google.colab import drive
drive.mount('./drive/')
%cd drive/My \ Drive/Text_Summarization
except:
print("No Colab Environment")
import json
import pandas as pd
import numpy as np
import re
import seaborn as sns
import matplotlib
matplotlib.rcParams["figure.figsize"] = (20, 7)
```
# Load Data
```
with open("../Data/raw/Telehealth_article_texts.txt") as f:
#Skip header
for i in range(4):
next(f)
corpus = f.read()
dict_articles = json.loads(corpus.replace("\n",""))
df_articles = pd.DataFrame.from_dict(dict_articles,orient="index",columns=["Content"]).reset_index().drop(columns=["index"])
df_articles.head()
df_metadata = pd.read_excel("../Data/raw/Metadata_telehealth_article_key_2.25.xlsx",sheet_name="Tied_to_Notebook",index_col="Index")
df_metadata.head()
#Ensure both Indexes are of same type before merge
assert df_metadata.index.dtype == df_articles.index.dtype
#Merge dataframes
df_metadata = df_metadata.merge(df_articles,left_index=True,right_index=True,how="left")
df_metadata.head()
JournalCrosswalk = pd.read_excel('../Data/raw/JournalTitles.xlsx')
JournalCrosswalk.set_index('Journal', inplace=True)
JournalCrosswalk.head()
df_metadata.dtypes
df_metadata["Content_Length"] = df_metadata["Content"].apply(lambda text: len(text))
df_metadata["Abstract_Length"] = df_metadata["Abstract"].apply(lambda text: len(str(text)))
df_metadata["Parsed_Keywords"] = df_metadata["Keywords"].apply(lambda keywords: str(keywords).replace("\n\n"," ").split()[1:])
df_metadata["Parsed_Keywords_Length"] = df_metadata["Parsed_Keywords"].apply(lambda text: len(text))
df_metadata["Journal Title"] = df_metadata["Journal Title"].replace('Psychological Servies', 'Psychological Services')
subfield = []
for i in df_metadata['Journal Title']:
#print(i)
try:
if i == 'Clinical Psychology: Science and Practice': # Journal is missing from Crosswalk
subfield.append('Clinical & Counseling Psychology')
else:
subfield.append(JournalCrosswalk['Journal Subfield'][i])
except:
subfield.append('No Match')
df_metadata["Subfield"] = subfield
#Have an idea of reference amount per document
df_metadata["et_al_Count"] = df_metadata["Content"].apply(lambda text: len(list(re.finditer("et al",text))))
df_metadata.head()
#troubleshooting
#emental health
#df_metadata["Content"].iloc[2][4633:5000]
#df_metadata["Clean_Content"].iloc[2][4633:5000]
```
## Acronyms - Include as Vocabulary for Paper
```
#Sci Spacy
#!pip install scispacy
#!pip install https://s3-us-west-2.amazonaws.com/ai2-s2-scispacy/releases/v0.4.0/en_core_sci_sm-0.4.0.tar.gz
#Spacy org
#!pip install spacy
#!python3 -m spacy download en_core_web_sm
#!python3 -m spacy download en_core_web_md
```
## Source: https://youtu.be/2_HSKDALwuw?t=708
## Abbreviation Detector Works by:
## 1. Finding Parentheses
## 2. Look up to 10 words behind the bracket
## 3. Greedily choose definition: Look for words next to each other, that in the right order start with the letters in the acronym
```
#ScispaCy: Fast and Robust Models for Biomedical Natural Language Processing: https://www.semanticscholar.org/paper/ScispaCy%3A-Fast-and-Robust-Models-for-Biomedical-Neumann-King/de28ec1d7bd38c8fc4e8ac59b6133800818b4e29
#https://github.com/allenai/SciSpaCy
import spacy
from scispacy.abbreviation import AbbreviationDetector
nlp = spacy.load("en_core_web_md")
nlp.add_pipe("abbreviation_detector")
nlp.Defaults.stop_words |= {"PRON","ll","ve","eg"}
corpus = df_metadata["Content"]
docs = list(nlp.pipe(corpus,disable=["ner","parser","textcat"]))
abrv_dict = dict()
for index,doc in enumerate(docs):
for abrv in doc._.abbreviations:
if str(abrv) not in abrv_dict.keys():
abrv_dict[str(abrv)] = {"LongForm":str(abrv._.long_form),"Document":{index}}
else:
abrv_dict[str(abrv)]["Document"].add(index)
#print(f"{abrv} \t ({abrv.start}, {abrv.end}) {abrv._.long_form} \t Document: {index}")
#Source https://stackoverflow.com/questions/22281059/set-object-is-not-json-serializable
def set_default(obj):
if isinstance(obj, set):
return list(obj)
raise TypeError
#Write original abbriviation dictionary
with open("../references/abbreviation_table.json","w") as f:
json.dump(abrv_dict, f, indent = 4,default=set_default)
#Remove misidentified abbreviations
with open("../references/Incorrect_abbrev.json", "r") as f:
misidentified_abrv = json.load(f)
for key in misidentified_abrv.keys():
abrv_dict.pop(key)
#Correct LongForm of abbreviations
with open("../references/Abbreviation_corrections.json", "r") as f:
correction_abrv = json.load(f)
for key in abrv_dict.keys():
if key in correction_abrv.keys():
abrv_dict[key]["LongForm"] = correction_abrv[key]
#Add abbreviations
with open("../references/Add_to_abbreviation_table.json", "r") as f:
add_abrv = json.load(f)
for key in add_abrv.keys():
abrv_dict[key] = add_abrv[key]
with open("../references/abbreviation_table_processed.json","w") as f:
json.dump(abrv_dict, f, indent = 4, default=set_default)
abrv_dict["PTSD"]["LongForm"]
df_abrv = (pd.read_json("../references/abbreviation_table_processed.json")
.T
.reset_index()
.rename(columns={"index":"Term"}))
df_abrv.head()
df_abrv[df_abrv["Term"] == "PTSD"].head()
#Validation
df_abrv[df_abrv["Term"] == "NYH"]
```
## Clean Data
1. Lowercase
2. Remove Punctuation
3. White Spaces
```
documents_tokens = []
for index,doc in enumerate(docs):
document_tokens = []
for token in doc:
#removes stopwords and punct
if not token.is_stop and not token.is_punct:
if str(token) in abrv_dict.keys():
document_tokens.append(abrv_dict[str(token)]["LongForm"])#Replace short-form with long-form
else:
document_tokens.append(token.lemma_)
documents_tokens.append(" ".join(document_tokens))
df_metadata["Stopwords_Lemma_Longform_Clean_Content"] = documents_tokens
df_metadata.head()
def unwanted_tokens(text):
docuemnt = text
remove_ngrams = ["large image page new","image page new window", "page new window Download","image page new",
"page new window","new window Download","image page","large image","1TABLES figurestablefigure thumbnailtable",
"FIGUREStable","DOWNLOAD","Download","et al"]
for ngram in remove_ngrams:
docuemnt = docuemnt.replace(ngram,"")
return docuemnt
df_metadata["Clean_Content"] = (df_metadata["Stopwords_Lemma_Longform_Clean_Content"].apply(lambda text: unwanted_tokens(text)))
df_metadata.head()
#Add to clean function: Different than a dash ord(8207) compared to 45 for normal dash
print(ord("-"),ord("—"))
from yellowbrick.text import DispersionPlot
import sklearn.metrics
try:
#Troubleshooting tokens to remove
dispersion_text = [doc.split() for doc in df_metadata["Clean_Content"]]
other_words = [token.split() for token in ['et al']]
other_words_1D = np.unique(np.concatenate(other_words).reshape(-1))
target_words = other_words_1D
#Create the visualizer and draw the plot
visualizer = DispersionPlot(target_words,ignore_case=False)
_ = visualizer.fit(dispersion_text)
except:
print("No words found")
df_metadata["Classification"] = (df_metadata["Date Published"].apply(lambda pub_date: "Covid"
if pub_date >= 2020 else "Pre-Covid" ))
df_metadata.to_csv("../Data/processed/Telehealth.csv",index=False)
```
# Trouble Shooting
## Dispersion Plot
```
from yellowbrick.text import DispersionPlot
import sklearn.metrics
try:
#Troubleshooting tokens to remove
dispersion_text = [doc.split() for doc in df_metadata["Clean_Content"]]
other_words = [token.split() for token in ['kbinformation','binformation']]
other_words_1D = np.unique(np.concatenate(other_words).reshape(-1))
target_words = other_words_1D
#Create the visualizer and draw the plot
visualizer = DispersionPlot(target_words,ignore_case=False)
_ = visualizer.fit(dispersion_text)
except:
print("No Words to be found")
```
| github_jupyter |
<div style="text-align:center;">
<h1 style="font-size: 50px; margin: 0px; margin-bottom: 5px;">Initial-Core-Final Mass Relation Plot</h1>
<h2 style="margin:0px; margin-bottom: 5px;">COMPAS methods paper Figure 8</h2>
<p style="text-align:center;">A notebook for reproducing the initial-core-final mass relation plot in the COMPAS methods paper.</p>
</div>
<img src="https://compas.science/images/COMPAS_CasA.png" style="width:50%; display:block; margin:auto; margin-bottom:20px">
```
import numpy as np
import h5py as h5
import matplotlib.pyplot as plt
import astropy.constants as consts
import matplotlib
import astropy.units as u
# make the plots pretty
%config InlineBackend.figure_format = 'retina'
plt.rc('font', family='serif')
fs = 24
params = {'legend.fontsize': fs,
'axes.labelsize': fs,
'xtick.labelsize':0.7*fs,
'ytick.labelsize':0.7*fs}
plt.rcParams.update(params)
```
# Get the stellar types
First we can import the stellar types array to use the same colour palette as the other plots.
```
import sys
sys.path.append("../")
from stellar_types import stellar_types
```
# Get the data
```
def get_COMPAS_vars(file, group, var_list):
if isinstance(var_list, str):
return file[group][var_list][...]
else:
return [file[group][var][...] for var in var_list]
```
# Top panel: Solar metallicity default prescription
```
def core_remnant_mass_comparison(file, fig=None, ax=None, show=True):
if fig is None or ax is None:
fig, ax = plt.subplots(figsize=(10, 8))
with h5.File(file, "r") as compas:
m_ZAMS, m_final, m_co_core_atCO, Z, stellar_type = get_COMPAS_vars(compas, "SSE_System_Parameters",
["Mass@ZAMS",
"Mass",
"Mass_CO_Core@CO",
"Metallicity",
"Stellar_Type"])
# only plot things for solar metallicity
solar = Z == 0.01416
uni_types = np.unique(stellar_type[solar])
# annotate the plot with the stellar types
for i in range(len(uni_types)):
ax.annotate(stellar_types[uni_types[i]]["short"], xy=(0.02, 0.93 - 0.05 * i),
xycoords="axes fraction", color=plt.get_cmap("tab10")(i / 10), fontsize=0.7*fs, weight="bold")
for i in range(len(uni_types)):
# plot the final white dwarf mass for WDs
if uni_types[i] in [10, 11, 12]:
ax.loglog(m_ZAMS[solar][stellar_type[solar] == uni_types[i]],
m_final[solar][stellar_type[solar] == uni_types[i]],
lw=3, color=plt.get_cmap("tab10")(i / 10))
# plot CO core mass at CO formation for NSs and BHs
elif uni_types[i] in [13, 14]:
ax.loglog(m_ZAMS[solar][stellar_type[solar] == uni_types[i]],
m_co_core_atCO[solar][stellar_type[solar] == uni_types[i]],
lw=3, color=plt.get_cmap("tab10")(i / 10))
# annotate with solar metallicity
ax.annotate(r"$Z = Z_{\rm \odot}$", xy=(0.97, 0.04), xycoords="axes fraction", fontsize=0.7*fs, ha="right")
ax.set_ylabel(r"Core Mass $[\rm M_{\odot}]$")
if show:
plt.show()
return fig, ax
```
# Middle panel: Illustrate effect of metallicity on core masses
```
def remnant_mass_across_metallicity(file, fig=None, ax=None, show=True):
if fig is None or ax is None:
fig, ax = plt.subplots(figsize=(10, 8))
with h5.File(file, "r") as compas:
m_ZAMS, m_final, m_co_core_atCO, Z, stellar_type = get_COMPAS_vars(compas, "SSE_System_Parameters",
["Mass@ZAMS",
"Mass",
"Mass_CO_Core@CO",
"Metallicity",
"Stellar_Type"])
# create an inset axis for the linear version
inset_ax = ax.inset_axes([0.05, 0.55, 0.53, 0.425])
inset_ax.tick_params(labelsize=0.5*fs)
# plot three different metallicities
for Z_match, style in [(0.01, "-"), (0.001, "--"), (0.0001, "dotted")]:
matching_Z = Z == Z_match
uni_types = np.unique(stellar_type[matching_Z])
for i in range(len(uni_types)):
# plot in same way as top panel
matching_type = stellar_type[matching_Z] == uni_types[i]
y_quantity = m_final if uni_types[i] in [10, 11, 12] else m_co_core_atCO
ax.loglog(m_ZAMS[matching_Z][matching_type], y_quantity[matching_Z][matching_type],
lw=2, linestyle=style, markevery=25, color=plt.get_cmap("tab10")(i / 10),
label=r"$Z = {{{}}}$".format(Z_match) if i == len(uni_types) - 1 else None)
# for black holes also plot in the inset axis
if uni_types[i] == 14:
inset_ax.plot(m_ZAMS[matching_Z][matching_type], y_quantity[matching_Z][matching_type],
lw=2, linestyle=style, markevery=25, color=plt.get_cmap("tab10")(i / 10))
ax.legend(loc="lower right", fontsize=0.7 * fs)
ax.set_ylabel(r"Core Mass $[\rm M_{\odot}]$")
if show:
plt.show()
return fig, ax
```
# Bottom panel: Demonstrate how remnant mass prescriptions differ at solar metallicity
```
def remnant_mass_prescription_comparison(prescriptions, fig=None, ax=None, show=True):
if fig is None or ax is None:
fig, ax = plt.subplots(figsize=(10, 8))
for file, style, label in prescriptions:
with h5.File("COMPAS_Output_{}/COMPAS_Output_{}.h5".format(file, file), "r") as compas:
m_ZAMS, m_final, Z, stellar_type = get_COMPAS_vars(compas, "SSE_System_Parameters",
["Mass@ZAMS",
"Mass",
"Metallicity",
"Stellar_Type"])
solar = Z == 0.01416
uni_types = np.unique(stellar_type[solar])
for i in range(len(uni_types)):
# only plot the NSs and BHs
if uni_types[i] >= 13:
# use scatter points for the Mandel & Mueller prescription
if file == "MM20":
ax.scatter(m_ZAMS[solar][stellar_type[solar] == uni_types[i]],
m_final[solar][stellar_type[solar] == uni_types[i]],
s=0.2, alpha=0.5, color=plt.get_cmap("tab10")(i / 10),
label=label if i == len(uni_types) - 1 else None)
# use lines for the other ones
else:
ax.loglog(m_ZAMS[solar][stellar_type[solar] == uni_types[i]],
m_final[solar][stellar_type[solar] == uni_types[i]],
lw=2, linestyle=style, color=plt.get_cmap("tab10")(i / 10),
label=label if i == len(uni_types) - 1 else None, zorder=10)
ax.set_xscale("log")
ax.set_yscale("log")
leg = ax.legend(fontsize=0.7 * fs, loc="lower right", markerscale=25, title=r"$Z = Z_{\rm \odot}$")
leg.get_title().set_fontsize(0.7 * fs)
ax.set_ylabel(r"Remnant Mass $[\rm M_{\odot}]$")
if show:
plt.show()
return fig, ax
```
# Create the whole plot!
```
fig, axes = plt.subplots(3, figsize=(10, 24))
fig, axes[0] = core_remnant_mass_comparison("COMPAS_Output_default/COMPAS_Output_default.h5",
fig=fig, ax=axes[0], show=False)
fig, axes[1] = remnant_mass_across_metallicity("COMPAS_Output_default/COMPAS_Output_default.h5",
fig=fig, ax=axes[1], show=False)
fig, axes[2] = remnant_mass_prescription_comparison([("default", "dotted", "Fryer+2012 Delayed"),
("rapid", "--", "Fryer+2012 Rapid"),
("MM20", None, "Mandel & Mueller 2020")],
fig=fig, ax=axes[2], show=False)
for ax, xticks, yticks in zip(axes,
[[0.1, 1, 10, 100], [0.1, 1, 10, 100], [10.0, 100.0]],
[[0.1, 1, 10], [0.1, 1, 10], [1.0, 10.0]]):
ax.set_xticks(xticks)
ax.get_xaxis().set_major_formatter(matplotlib.ticker.ScalarFormatter())
ax.set_xlabel(r"Initial Mass $[\rm M_{\rm \odot}]$")
ax.set_yticks(yticks)
ax.get_yaxis().set_major_formatter(matplotlib.ticker.ScalarFormatter())
ax.tick_params(which="major", length=7)
ax.tick_params(which="minor", length=4)
plt.savefig("initial_core_final_mass_relations.pdf", format="pdf", bbox_inches="tight")
plt.show()
```
| github_jupyter |
# SageMaker Inference Pipeline with Scikit Learn and Linear Learner
ISO20022 pacs.008 inference pipeline notebook. This notebook uses training dataset to perform model training. It uses SageMaker Linear Learner to train a model. The problem is defined to be a `binary classification` problem of accepting or rejecting a pacs.008 message.
Amazon SageMaker provides a very rich set of [builtin algorithms](https://docs.aws.amazon.com/sagemaker/latest/dg/algorithms-choose.html) for model training and development. This notebook uses [Amazon SageMaker Linear Learner Algorithm](https://docs.aws.amazon.com/sagemaker/latest/dg/linear-learner.html) on training dataset to perform model training. The Amazon SageMaker linear learner algorithm provides a solution for both classification and regression problems. With the SageMaker algorithm, you can simultaneously explore different training objectives and choose the best solution from a validation set. You can also explore a large number of models and choose the best. The best model optimizes either of the following:
* Continuous objectives, such as mean square error, cross entropy loss, absolute error (regression models).
* Discrete objectives suited for classification, such as F1 measure, precision, recall, or accuracy (classification models).
ML Model development is an iterative process with several tasks that data scientists go through to produce an effective model that can solve business problem. The process typically involves:
* Data exploration and analysis
* Feature engineering
* Model development
* Model training and tuning
* Model deployment
We provide the accompanying notebook [pacs008_xgboost_local.ipynb](./pacs008_xgboost_local.ipynb) which demonstrates data exploration, analysis and feature engineering, focussing on text feature engineering. This notebook uses the results of analysis in [pacs008_xgboost_local.ipynb](./pacs008_xgboost_local.ipynb) to create a feature engineering pipeline using [SageMaker Inference Pipeline](https://docs.aws.amazon.com/sagemaker/latest/dg/inference-pipelines.html).
Here we define the ML problem to be a `binary classification` problem, that of predicting if a pacs.008 XML message with be processed sucessfully or lead to exception process. The predicts `Success` i.e. 1 or `Failure` i.e. 0.
**Feature Engineering**
Data pre-processing and featurizing the dataset by incorporating standard techniques or prior knowledge is a standard mechanism to make dataset meaningful for training. Once data has been pre-processed and transformed, it can be finally used to train an ML model using an algorithm. However, when the trained model is used for processing real time or batch prediction requests, the model receives data in a format which needs to be pre-processed (e.g. featurized) before it can be passed to the algorithm. In this notebook, we will demonstrate how you can build your ML Pipeline leveraging the Sagemaker Scikit-learn container and SageMaker XGBoost algorithm. After a model is trained, we deploy the Pipeline (Data preprocessing and XGBoost) as an **Inference Pipeline** behind a **single Endpoint** for real time inference and for **batch inferences** using Amazon SageMaker Batch Transform.
We use pacs.008 xml element `<InstrForNxtAgt><InstrInf>TEXT</InstrForNxtAgt></InstrInf>` to perform feature engineer i.e featurize text into new numeric features that can be used in making prodictions.
Since we featurize `InstrForNxtAgt` to numeric representations during training, we have to pre-processs to transform text into numeric features before using the trained model to make predictions.
**Inference Pipeline**
The diagram below shows how Amazon SageMaker Inference Pipeline works. It is used to deploy multi-container endpoints.

**Inference Endpoint**
The diagram below shows the places in the cross-border payment message flow where a call to ML inference endpoint can be injected to get inference from the ML model. The inference result can be used to take additional actions, including corrective actions before sending the message downstream.

**Further Reading:**
For information on Amazon SageMaker Linear Learner algorithm and SageMaker Inference Pipeline visit the following references:
[SageMaker Linear Learner Algorithm](https://docs.aws.amazon.com/sagemaker/latest/dg/linear-learner.html)
[SageMaker Inference Pipeline](https://docs.aws.amazon.com/sagemaker/latest/dg/inference-pipelines.html)
## Basic Setup
In this step we do basic setup needed for rest of the notebook:
* Amazon SageMaker API client using boto3
* Amazon SageMaker session object
* AWS region
* AWS IAM role
```
import os
import boto3
import sagemaker
from sagemaker import get_execution_role
sm_client = boto3.Session().client('sagemaker')
sm_session = sagemaker.Session()
region = boto3.session.Session().region_name
role = get_execution_role()
print ("Notebook is running with assumed role {}".format (role))
print("Working with AWS services in the {} region".format(region))
```
### Provide S3 Bucket Name
```
# Working directory for the notebook
WORKDIR = os.getcwd()
BASENAME = os.path.dirname(WORKDIR)
print(f"WORKDIR: {WORKDIR}")
print(f"BASENAME: {BASENAME}")
# Create a directory storing local data
iso20022_data_path = 'iso20022-data'
if not os.path.exists(iso20022_data_path):
# Create a new directory because it does not exist
os.makedirs(iso20022_data_path)
# Store all prototype assets in this bucket
s3_bucket_name = 'iso20022-prototype-t3'
s3_bucket_uri = 's3://' + s3_bucket_name
# Prefix for all files in this prototype
prefix = 'iso20022'
pacs008_prefix = prefix + '/pacs008'
raw_data_prefix = pacs008_prefix + '/raw-data'
labeled_data_prefix = pacs008_prefix + '/labeled-data'
training_data_prefix = pacs008_prefix + '/training-data'
training_headers_prefix = pacs008_prefix + '/training-headers'
test_data_prefix = pacs008_prefix + '/test-data'
training_job_output_prefix = pacs008_prefix + '/training-output'
print(f"Training data with headers will be uploaded to {s3_bucket_uri + '/' + training_headers_prefix}")
print(f"Training data will be uploaded to {s3_bucket_uri + '/' + training_data_prefix}")
print(f"Test data will be uploaded to {s3_bucket_uri + '/' + test_data_prefix}")
print(f"Training job output will be stored in {s3_bucket_uri + '/' + training_job_output_prefix}")
labeled_data_location = s3_bucket_uri + '/' + labeled_data_prefix
training_data_w_headers_location = s3_bucket_uri + '/' + training_headers_prefix
training_data_location = s3_bucket_uri + '/' + training_data_prefix
test_data_location = s3_bucket_uri + '/' + test_data_prefix
print(f"Raw labeled data location = {labeled_data_location}")
print(f"Training data with headers location = {training_data_w_headers_location}")
print(f"Training data location = {training_data_location}")
print(f"Test data location = {test_data_location}")
```
## Prepare Training Dataset
1. Select training dataset from raw labeled dataset.
1. Split labeled dataset to training and test datasets.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import string
from sklearn.model_selection import train_test_split
from sklearn import ensemble, metrics, model_selection, naive_bayes
color = sns.color_palette()
%matplotlib inline
```
### Download raw labeled dataset
```
# Download labeled raw dataset from S3
s3_client = boto3.client('s3')
s3_client.download_file(s3_bucket_name, labeled_data_prefix + '/labeled_data.csv', 'iso20022-data/labeled_data.csv')
# Read the train and test dataset and check the top few lines ##
labeled_raw_df = pd.read_csv("iso20022-data/labeled_data.csv")
labeled_raw_df.head()
```
### Select features for training
```
# Training features
fts=[
'y_target',
'Document_FIToFICstmrCdtTrf_CdtTrfTxInf_Dbtr_PstlAdr_Ctry',
'Document_FIToFICstmrCdtTrf_CdtTrfTxInf_Cdtr_PstlAdr_Ctry',
'Document_FIToFICstmrCdtTrf_CdtTrfTxInf_RgltryRptg_DbtCdtRptgInd',
'Document_FIToFICstmrCdtTrf_CdtTrfTxInf_RgltryRptg_Authrty_Ctry',
'Document_FIToFICstmrCdtTrf_CdtTrfTxInf_RgltryRptg_Dtls_Cd',
'Document_FIToFICstmrCdtTrf_CdtTrfTxInf_InstrForNxtAgt_InstrInf',
]
# New data frame with selected features
selected_df = labeled_raw_df[fts]
selected_df.head()
# Rename columns
selected_df = selected_df.rename(columns={
'Document_FIToFICstmrCdtTrf_CdtTrfTxInf_Dbtr_PstlAdr_Ctry': 'Dbtr_PstlAdr_Ctry',
'Document_FIToFICstmrCdtTrf_CdtTrfTxInf_Cdtr_PstlAdr_Ctry': 'Cdtr_PstlAdr_Ctry',
'Document_FIToFICstmrCdtTrf_CdtTrfTxInf_RgltryRptg_DbtCdtRptgInd': 'RgltryRptg_DbtCdtRptgInd',
'Document_FIToFICstmrCdtTrf_CdtTrfTxInf_RgltryRptg_Authrty_Ctry': 'RgltryRptg_Authrty_Ctry',
'Document_FIToFICstmrCdtTrf_CdtTrfTxInf_RgltryRptg_Dtls_Cd': 'RgltryRptg_Dtls_Cd',
'Document_FIToFICstmrCdtTrf_CdtTrfTxInf_InstrForNxtAgt_InstrInf': 'InstrForNxtAgt',
})
selected_df.head()
from sklearn.preprocessing import LabelEncoder
# Assign Pandas data types.
categorical_fts=[
'Dbtr_PstlAdr_Ctry',
'Cdtr_PstlAdr_Ctry',
'RgltryRptg_DbtCdtRptgInd',
'RgltryRptg_Authrty_Ctry',
'RgltryRptg_Dtls_Cd'
]
integer_fts=[
]
numeric_fts=[
]
text_fts=[
# Leave text as object
# 'InstrForNxtAgt'
]
# Categorical features to categorical data type.
for col in categorical_fts:
selected_df[col] = selected_df[col].astype(str).astype('category')
# Integer features to int64 data type.
for col in integer_fts:
selected_df[col] = selected_df[col].astype(str).astype('int64')
# Numeric features to float64 data type.
for col in numeric_fts:
selected_df[col] = selected_df[col].astype(str).astype('float64')
# Text features to string data type.
for col in text_fts:
selected_df[col] = selected_df[col].astype(str).astype('string')
label_encoder = LabelEncoder()
selected_df['y_target'] = label_encoder.fit_transform(selected_df['y_target'])
selected_df.dtypes
selected_df.info()
selected_df
X_train_df, X_test_df, y_train_df, y_test_df = train_test_split(selected_df, selected_df['y_target'], test_size=0.20, random_state=299, shuffle=True)
print("Number of rows in train dataset : ",X_train_df.shape[0])
print("Number of rows in test dataset : ",X_test_df.shape[0])
X_train_df
X_test_df
## Save training and test datasets to CSV
train_data_w_headers_output_path = 'iso20022-data/train_data_w_headers.csv'
print(f'Saving training data with headers to {train_data_w_headers_output_path}')
X_train_df.to_csv(train_data_w_headers_output_path, index=False)
train_data_output_path = 'iso20022-data/train_data.csv'
print(f'Saving training data without headers to {train_data_output_path}')
X_train_df.to_csv(train_data_output_path, header=False, index=False)
test_data_output_path = 'iso20022-data/test_data.csv'
print(f'Saving test data without headers to {test_data_output_path}')
X_test_df.to_csv(test_data_output_path, header=False, index=False)
```
### Upload training and test datasets to S3 for training
```
train_input_data_location = sm_session.upload_data(
path=train_data_w_headers_output_path,
bucket=s3_bucket_name,
key_prefix=training_headers_prefix,
)
print(f'Uploaded traing data with headers to: {train_input_data_location}')
train_input_data_location = sm_session.upload_data(
path=train_data_output_path,
bucket=s3_bucket_name,
key_prefix=training_data_prefix,
)
print(f'Uploaded data without headers to: {train_input_data_location}')
test_input_data_location = sm_session.upload_data(
path=test_data_output_path,
bucket=s3_bucket_name,
key_prefix=test_data_prefix,
)
print(f'Uploaded data without headers to: {test_input_data_location}')
```
# Feature Engineering
## Create a Scikit-learn script to train with <a class="anchor" id="create_sklearn_script"></a>
To run Scikit-learn on Sagemaker `SKLearn` Estimator with a script as an entry point. The training script is very similar to a training script you might run outside of SageMaker, but you can access useful properties about the training environment through various environment variables, such as:
* SM_MODEL_DIR: A string representing the path to the directory to write model artifacts to. These artifacts are uploaded to S3 for model hosting.
* SM_OUTPUT_DIR: A string representing the filesystem path to write output artifacts to. Output artifacts may include checkpoints, graphs, and other files to save, not including model artifacts. These artifacts are compressed and uploaded to S3 to the same S3 prefix as the model artifacts.
Supposing two input channels, 'train' and 'test', were used in the call to the Chainer estimator's fit() method, the following will be set, following the format SM_CHANNEL_[channel_name]:
* SM_CHANNEL_TRAIN: A string representing the path to the directory containing data in the 'train' channel
* SM_CHANNEL_TEST: Same as above, but for the 'test' channel.
A typical training script loads data from the input channels, configures training with hyperparameters, trains a model, and saves a model to model_dir so that it can be hosted later. Hyperparameters are passed to your script as arguments and can be retrieved with an argparse.ArgumentParser instance.
### Create SageMaker Scikit Estimator <a class="anchor" id="create_sklearn_estimator"></a>
To run our Scikit-learn training script on SageMaker, we construct a `sagemaker.sklearn.estimator.sklearn` estimator, which accepts several constructor arguments:
* __entry_point__: The path to the Python script SageMaker runs for training and prediction.
* __role__: Role ARN
* __framework_version__: Scikit-learn version you want to use for executing your model training code.
* __train_instance_type__ *(optional)*: The type of SageMaker instances for training. __Note__: Because Scikit-learn does not natively support GPU training, Sagemaker Scikit-learn does not currently support training on GPU instance types.
* __sagemaker_session__ *(optional)*: The session used to train on Sagemaker.
```
from sagemaker.sklearn.estimator import SKLearn
preprocessing_job_name = 'pacs008-preprocessor-ll'
print('data preprocessing job name: ' + preprocessing_job_name)
FRAMEWORK_VERSION = "0.23-1"
source_dir = "../sklearn-transformers"
script_file = "pacs008_sklearn_featurizer.py"
sklearn_preprocessor = SKLearn(
entry_point=script_file,
source_dir=source_dir,
role=role,
framework_version=FRAMEWORK_VERSION,
instance_type="ml.c4.xlarge",
sagemaker_session=sm_session,
base_job_name=preprocessing_job_name,
)
sklearn_preprocessor.fit({"train": train_input_data_location})
```
### Batch transform our training data <a class="anchor" id="preprocess_train_data"></a>
Now that our proprocessor is properly fitted, let's go ahead and preprocess our training data. Let's use batch transform to directly preprocess the raw data and store right back into s3.
```
# Define a SKLearn Transformer from the trained SKLearn Estimator
transformer = sklearn_preprocessor.transformer(
instance_count=1,
instance_type="ml.m5.xlarge",
assemble_with="Line",
accept="text/csv",
)
# Preprocess training input
transformer.transform(train_input_data_location, content_type="text/csv")
print("Waiting for transform job: " + transformer.latest_transform_job.job_name)
transformer.wait()
preprocessed_train = transformer.output_path
```
# Train a Linear Learner Model
## Fit a LinearLearner Model with the preprocessed data <a class="anchor" id="training_model"></a>
Let's take the preprocessed training data and fit a LinearLearner Model. Sagemaker provides prebuilt algorithm containers that can be used with the Python SDK. The previous Scikit-learn job preprocessed the labeled raw pacs.008 dataset into useable training data that we can now use to fit a binary classifier Linear Learner model.
For more on Linear Learner see: https://docs.aws.amazon.com/sagemaker/latest/dg/linear-learner.html
```
from sagemaker.image_uris import retrieve
ll_image = retrieve("linear-learner", boto3.Session().region_name)
# Set job name
training_job_name = 'pacs008-ll-training'
print('Linear Learner training job name: ' + training_job_name)
# S3 bucket for storing model artifacts
training_job_output_location = s3_bucket_uri + '/' + training_job_output_prefix + '/ll_model'
ll_estimator = sagemaker.estimator.Estimator(
ll_image,
role,
instance_count=1,
instance_type="ml.m4.2xlarge",
volume_size=20,
max_run=3600,
input_mode="File",
output_path=training_job_output_location,
sagemaker_session=sm_session,
base_job_name=training_job_name,
)
# binary_classifier_model_selection_criteria: accuracy is default
# - accuracy | f_beta | precision_at_target_recall |recall_at_target_precision | loss_function
# feature_dim=auto, # auto or actual number, default is auto
# epochs=15, default is 15
# learning_rate=auto or actual number 0.05 or 0.005
# loss=logistic | auto |hinge_loss, default is logistic
# mini_batch_size=32, default is 1000
# num_models=auto, or a number
# optimizer=auto or sgd | adam | rmsprop
ll_estimator.set_hyperparameters(
predictor_type="binary_classifier",
binary_classifier_model_selection_criteria="accuracy",
epochs=15,
mini_batch_size=32)
ll_train_data = sagemaker.inputs.TrainingInput(
preprocessed_train, # set after preprocessing job completes
distribution="FullyReplicated",
content_type="text/csv",
s3_data_type="S3Prefix",
)
data_channels = {"train": ll_train_data}
ll_estimator.fit(inputs=data_channels, logs=True)
```
# Serial Inference Pipeline with Scikit preprocessor and Linear Learner <a class="anchor" id="serial_inference"></a>
## Set up the inference pipeline <a class="anchor" id="pipeline_setup"></a>
Setting up a Machine Learning pipeline can be done with the Pipeline Model. This sets up a list of models in a single endpoint. We configure our pipeline model with the fitted Scikit-learn inference model (data preprocessing/feature engineering model) and the fitted Linear Learner model. Deploying the model follows the standard ```deploy``` pattern in the SageMaker Python SDK.
```
from sagemaker.model import Model
from sagemaker.pipeline import PipelineModel
import boto3
from time import gmtime, strftime
timestamp_prefix = strftime("%Y-%m-%d-%H-%M-%S", gmtime())
# The two SageMaker Models: one for data preprocessing, and second for inference
scikit_learn_inferencee_model = sklearn_preprocessor.create_model()
linear_learner_model = ll_estimator.create_model()
model_name = "pacs008-ll-inference-pipeline-" + timestamp_prefix
endpoint_name = "pacs008-ll-inference-pipeline-ep-" + timestamp_prefix
sm_model = PipelineModel(
name=model_name, role=role, models=[scikit_learn_inferencee_model, linear_learner_model]
)
sm_model.deploy(initial_instance_count=1, instance_type="ml.c4.xlarge", endpoint_name=endpoint_name)
```
### Store Model Name and Endpoint Name in Notebook Magic Store
These notebook magic store values are used in the example batch transform notebook.
```
%store model_name
%store endpoint_name
```
## Make a request to our pipeline endpoint <a class="anchor" id="pipeline_inference_request"></a>
The diagram below shows the places in the cross-border payment message flow where a call to ML inference endpoint can be injected to get inference from the ML model. The inference result can be used to take additional actions, including corrective actions before sending the message downstream.

Here we just grab the first line from the test data (you'll notice that the inference python script is very particular about the ordering of the inference request data). The ```ContentType``` field configures the first container, while the ```Accept``` field configures the last container. You can also specify each container's ```Accept``` and ```ContentType``` values using environment variables.
We make our request with the payload in ```'text/csv'``` format, since that is what our script currently supports. If other formats need to be supported, this would have to be added to the ```output_fn()``` method in our entry point. Note that we set the ```Accept``` to ```application/json```, since Linear Learner does not support ```text/csv``` ```Accept```. The inference output in this case is trying to predict `Success` or `Failure` of ISO20022 pacs.008 payment message using only the subset of message XML elements in the message i.e. features on which model was trained.
```
from sagemaker.predictor import Predictor
from sagemaker.serializers import CSVSerializer
# payload_1, expect: Failure
#payload_1 = "US,GB,,,,/SVC/It is to be delivered in three days. Greater than three days penalty add 2bp per day"
payload_1 = "MX,GB,,,,/SVC/It is to be delivered in four days. Greater than four days penalty add 2bp per day"
# payload_2, expect: Success
payload_2 = "MX,GB,,,,"
#payload_2 = "US,IE,,,,/TRSY/Treasury Services Platinum Customer"
# payload_3, expect: Failure
payload_3 = "TH,US,,,,/SVC/It is to be delivered in four days. Greater than four days penalty add 2bp per day"
#payload_3 = "CA,US,,,,/SVC/It is to be delivered in three days. Greater than three days penalty add 2bp per day"
# payload_4, expect: Success
payload_4 = "IN,CA,DEBT,IN,00.P0006,"
# payload_5, expect: Success
payload_5 = "IE,IN,CRED,IN,0,/REG/15.X0003 FDI in Transportation"
# Failure
payload_5 = "IE,IN,CRED,IN,0,/REG/15.X0009 FDI in Agriculture "
# Failure
payload_5 = "IE,IN,CRED,IN,0,/REG/15.X0004 retail"
# payload_6, expect: Failure
payload_6 = "IE,IN,CRED,IN,0,/REG/99.C34698"
#payload_6 = "MX,IE,,,,/TRSY/eweweww"
endpoint_name = 'pacs008-ll-inference-pipeline-ep-2021-11-25-00-58-52'
predictor = Predictor(
endpoint_name=endpoint_name, sagemaker_session=sm_session, serializer=CSVSerializer()
)
print(f"1. Expect Failure i.e. 0, {predictor.predict(payload_1)}")
print(f"2. Expect Success i.e. 1, {predictor.predict(payload_2)}")
print(f"3. Expect Failure i.e. 0, {predictor.predict(payload_3)}")
print(f"4. Expect Success i.e. 1, {predictor.predict(payload_4)}")
print(f"5. Expect Success i.e. 1, {predictor.predict(payload_5)}")
print(f"6. Expect Failure i.e. 0, {predictor.predict(payload_6)}")
```
# Delete Endpoint
Once we are finished with the endpoint, we clean up the resources!
```
sm_client = sm_session.boto_session.client("sagemaker")
sm_client.delete_endpoint(EndpointName=endpoint_name)
```
| github_jupyter |
# Machine Learning Pipeline - Model Training
In this notebook, we pick up the transformed datasets and the selected variables that we saved in the previous notebooks.
# Reproducibility: Setting the seed
With the aim to ensure reproducibility between runs of the same notebook, but also between the research and production environment, for each step that includes some element of randomness, it is extremely important that we **set the seed**.
```
# to handle datasets
import pandas as pd
import numpy as np
# for plotting
import matplotlib.pyplot as plt
# to save the model
import joblib
# to build the model
from sklearn.linear_model import Lasso
# to evaluate the model
from sklearn.metrics import mean_squared_error, r2_score
# to visualise al the columns in the dataframe
pd.pandas.set_option('display.max_columns', None)
# load the train and test set with the engineered variables
# we built and saved these datasets in a previous notebook.
# If you haven't done so, go ahead and check the previous notebooks (step 2)
# to find out how to create these datasets
X_train = pd.read_csv('xtrain.csv')
X_test = pd.read_csv('xtest.csv')
X_train.head()
# load the target (remember that the target is log transformed)
y_train = pd.read_csv('ytrain.csv')
y_test = pd.read_csv('ytest.csv')
y_train.head()
# load the pre-selected features
# ==============================
# we selected the features in the previous notebook (step 3)
# if you haven't done so, go ahead and visit the previous notebook
# to find out how to select the features
features = pd.read_csv('selected_features.csv')
features = features['0'].to_list()
# display final feature set
features
# reduce the train and test set to the selected features
X_train = X_train[features]
X_test = X_test[features]
```
### Regularised linear regression: Lasso
Remember to set the seed.
```
# set up the model
# remember to set the random_state / seed
lin_model = Lasso(alpha=0.001, random_state=0)
# train the model
lin_model.fit(X_train, y_train)
# evaluate the model:
# ====================
# remember that we log transformed the output (SalePrice)
# in our feature engineering notebook (step 2).
# In order to get the true performance of the Lasso
# we need to transform both the target and the predictions
# back to the original house prices values.
# We will evaluate performance using the mean squared error and
# the root of the mean squared error and r2
# make predictions for train set
pred = lin_model.predict(X_train)
# determine mse, rmse and r2
print('train mse: {}'.format(int(
mean_squared_error(np.exp(y_train), np.exp(pred)))))
print('train rmse: {}'.format(int(
mean_squared_error(np.exp(y_train), np.exp(pred), squared=False))))
print('train r2: {}'.format(
r2_score(np.exp(y_train), np.exp(pred))))
print()
# make predictions for test set
pred = lin_model.predict(X_test)
# determine mse, rmse and r2
print('test mse: {}'.format(int(
mean_squared_error(np.exp(y_test), np.exp(pred)))))
print('test rmse: {}'.format(int(
mean_squared_error(np.exp(y_test), np.exp(pred), squared=False))))
print('test r2: {}'.format(
r2_score(np.exp(y_test), np.exp(pred))))
print()
print('Average house price: ', int(np.exp(y_train).median()))
# let's evaluate our predictions respect to the real sale price
plt.scatter(y_test, lin_model.predict(X_test))
plt.xlabel('True House Price')
plt.ylabel('Predicted House Price')
plt.title('Evaluation of Lasso Predictions')
```
We can see that our model is doing a pretty good job at estimating house prices.
```
y_test.reset_index(drop=True)
# let's evaluate the distribution of the errors:
# they should be fairly normally distributed
y_test.reset_index(drop=True, inplace=True)
preds = pd.Series(lin_model.predict(X_test))
preds
# let's evaluate the distribution of the errors:
# they should be fairly normally distributed
errors = y_test['SalePrice'] - preds
errors.hist(bins=30)
plt.show()
```
The distribution of the errors follows quite closely a gaussian distribution. That suggests that our model is doing a good job as well.
### Feature importance
```
# Finally, just for fun, let's look at the feature importance
importance = pd.Series(np.abs(lin_model.coef_.ravel()))
importance.index = features
importance.sort_values(inplace=True, ascending=False)
importance.plot.bar(figsize=(18,6))
plt.ylabel('Lasso Coefficients')
plt.title('Feature Importance')
```
## Save the Model
```
# we are happy to our model, so we save it to be able
# to score new data
joblib.dump(lin_model, 'linear_regression.joblib')
```
# Additional Resources
## Feature Engineering
- [Feature Engineering for Machine Learning](https://www.udemy.com/course/feature-engineering-for-machine-learning/?referralCode=A855148E05283015CF06) - Online Course
- [Packt Feature Engineering Cookbook](https://www.packtpub.com/data/python-feature-engineering-cookbook) - Book
- [Feature Engineering for Machine Learning: A comprehensive Overview](https://trainindata.medium.com/feature-engineering-for-machine-learning-a-comprehensive-overview-a7ad04c896f8) - Article
- [Practical Code Implementations of Feature Engineering for Machine Learning with Python](https://towardsdatascience.com/practical-code-implementations-of-feature-engineering-for-machine-learning-with-python-f13b953d4bcd) - Article
## Feature Selection
- [Feature Selection for Machine Learning](https://www.udemy.com/course/feature-selection-for-machine-learning/?referralCode=186501DF5D93F48C4F71) - Online Course
- [Feature Selection for Machine Learning: A comprehensive Overview](https://trainindata.medium.com/feature-selection-for-machine-learning-a-comprehensive-overview-bd571db5dd2d) - Article
## Machine Learning
- [Best Resources to Learn Machine Learning](https://trainindata.medium.com/find-out-the-best-resources-to-learn-machine-learning-cd560beec2b7) - Article
- [Machine Learning with Imbalanced Data](https://www.udemy.com/course/machine-learning-with-imbalanced-data/?referralCode=F30537642DA57D19ED83) - Online Course
| github_jupyter |
```
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Vertex client library: Custom training tabular regression model with pipeline for online prediction with training pipeline
<table align="left">
<td>
<a href="https://colab.research.google.com/github/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/gapic/custom/showcase_custom_tabular_regression_online_pipeline.ipynb">
<img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
</a>
</td>
<td>
<a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/gapic/custom/showcase_custom_tabular_regression_online_pipeline.ipynb">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
</table>
<br/><br/><br/>
## Overview
This tutorial demonstrates how to use the Vertex client library for Python to train and deploy a custom tabular regression model for online prediction, using a training pipeline.
### Dataset
The dataset used for this tutorial is the [Boston Housing Prices dataset](https://www.cs.toronto.edu/~delve/data/boston/bostonDetail.html). The version of the dataset you will use in this tutorial is built into TensorFlow. The trained model predicts the median price of a house in units of 1K USD.
### Objective
In this tutorial, you create a custom model from a Python script in a Google prebuilt Docker container using the Vertex client library, and then do a prediction on the deployed model by sending data. You can alternatively create custom models using `gcloud` command-line tool or online using Google Cloud Console.
The steps performed include:
- Create a Vertex custom job for training a model.
- Create a `TrainingPipeline` resource.
- Train a TensorFlow model with the `TrainingPipeline` resource.
- Retrieve and load the model artifacts.
- View the model evaluation.
- Upload the model as a Vertex `Model` resource.
- Deploy the `Model` resource to a serving `Endpoint` resource.
- Make a prediction.
- Undeploy the `Model` resource.
### Costs
This tutorial uses billable components of Google Cloud (GCP):
* Vertex AI
* Cloud Storage
Learn about [Vertex AI
pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
Calculator](https://cloud.google.com/products/calculator/)
to generate a cost estimate based on your projected usage.
## Installation
Install the latest version of Vertex client library.
```
import os
import sys
# Google Cloud Notebook
if os.path.exists("/opt/deeplearning/metadata/env_version"):
USER_FLAG = "--user"
else:
USER_FLAG = ""
! pip3 install -U google-cloud-aiplatform $USER_FLAG
```
Install the latest GA version of *google-cloud-storage* library as well.
```
! pip3 install -U google-cloud-storage $USER_FLAG
```
### Restart the kernel
Once you've installed the Vertex client library and Google *cloud-storage*, you need to restart the notebook kernel so it can find the packages.
```
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
```
## Before you begin
### GPU runtime
*Make sure you're running this notebook in a GPU runtime if you have that option. In Colab, select* **Runtime > Change Runtime Type > GPU**
### Set up your Google Cloud project
**The following steps are required, regardless of your notebook environment.**
1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
3. [Enable the Vertex APIs and Compute Engine APIs.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component)
4. [The Google Cloud SDK](https://cloud.google.com/sdk) is already installed in Google Cloud Notebook.
5. Enter your project ID in the cell below. Then run the cell to make sure the
Cloud SDK uses the right project for all the commands in this notebook.
**Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
```
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
! gcloud config set project $PROJECT_ID
```
#### Region
You can also change the `REGION` variable, which is used for operations
throughout the rest of this notebook. Below are regions supported for Vertex. We recommend that you choose the region closest to you.
- Americas: `us-central1`
- Europe: `europe-west4`
- Asia Pacific: `asia-east1`
You may not use a multi-regional bucket for training with Vertex. Not all regions provide support for all Vertex services. For the latest support per region, see the [Vertex locations documentation](https://cloud.google.com/vertex-ai/docs/general/locations)
```
REGION = "us-central1" # @param {type: "string"}
```
#### Timestamp
If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append onto the name of resources which will be created in this tutorial.
```
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
```
### Authenticate your Google Cloud account
**If you are using Google Cloud Notebook**, your environment is already authenticated. Skip this step.
**If you are using Colab**, run the cell below and follow the instructions when prompted to authenticate your account via oAuth.
**Otherwise**, follow these steps:
In the Cloud Console, go to the [Create service account key](https://console.cloud.google.com/apis/credentials/serviceaccountkey) page.
**Click Create service account**.
In the **Service account name** field, enter a name, and click **Create**.
In the **Grant this service account access to project** section, click the Role drop-down list. Type "Vertex" into the filter box, and select **Vertex Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
Click Create. A JSON file that contains your key downloads to your local environment.
Enter the path to your service account key as the GOOGLE_APPLICATION_CREDENTIALS variable in the cell below and run the cell.
```
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# If on Google Cloud Notebook, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
%env GOOGLE_APPLICATION_CREDENTIALS ''
```
### Create a Cloud Storage bucket
**The following steps are required, regardless of your notebook environment.**
When you submit a custom training job using the Vertex client library, you upload a Python package
containing your training code to a Cloud Storage bucket. Vertex runs
the code from this package. In this tutorial, Vertex also saves the
trained model that results from your job in the same bucket. You can then
create an `Endpoint` resource based on this output in order to serve
online predictions.
Set the name of your Cloud Storage bucket below. Bucket names must be globally unique across all Google Cloud projects, including those outside of your organization.
```
BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
```
**Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
```
! gsutil mb -l $REGION $BUCKET_NAME
```
Finally, validate access to your Cloud Storage bucket by examining its contents:
```
! gsutil ls -al $BUCKET_NAME
```
### Set up variables
Next, set up some variables used throughout the tutorial.
### Import libraries and define constants
#### Import Vertex client library
Import the Vertex client library into our Python environment.
```
import time
from google.cloud.aiplatform import gapic as aip
from google.protobuf import json_format
from google.protobuf.json_format import MessageToJson, ParseDict
from google.protobuf.struct_pb2 import Struct, Value
```
#### Vertex constants
Setup up the following constants for Vertex:
- `API_ENDPOINT`: The Vertex API service endpoint for dataset, model, job, pipeline and endpoint services.
- `PARENT`: The Vertex location root path for dataset, model, job, pipeline and endpoint resources.
```
# API service endpoint
API_ENDPOINT = "{}-aiplatform.googleapis.com".format(REGION)
# Vertex location root path for your dataset, model and endpoint resources
PARENT = "projects/" + PROJECT_ID + "/locations/" + REGION
```
#### CustomJob constants
Set constants unique to CustomJob training:
- Dataset Training Schemas: Tells the `Pipeline` resource service the task (e.g., classification) to train the model for.
```
CUSTOM_TASK_GCS_PATH = (
"gs://google-cloud-aiplatform/schema/trainingjob/definition/custom_task_1.0.0.yaml"
)
```
#### Hardware Accelerators
Set the hardware accelerators (e.g., GPU), if any, for training and prediction.
Set the variables `TRAIN_GPU/TRAIN_NGPU` and `DEPLOY_GPU/DEPLOY_NGPU` to use a container image supporting a GPU and the number of GPUs allocated to the virtual machine (VM) instance. For example, to use a GPU container image with 4 Nvidia Telsa K80 GPUs allocated to each VM, you would specify:
(aip.AcceleratorType.NVIDIA_TESLA_K80, 4)
For GPU, available accelerators include:
- aip.AcceleratorType.NVIDIA_TESLA_K80
- aip.AcceleratorType.NVIDIA_TESLA_P100
- aip.AcceleratorType.NVIDIA_TESLA_P4
- aip.AcceleratorType.NVIDIA_TESLA_T4
- aip.AcceleratorType.NVIDIA_TESLA_V100
Otherwise specify `(None, None)` to use a container image to run on a CPU.
*Note*: TF releases before 2.3 for GPU support will fail to load the custom model in this tutorial. It is a known issue and fixed in TF 2.3 -- which is caused by static graph ops that are generated in the serving function. If you encounter this issue on your own custom models, use a container image for TF 2.3 with GPU support.
```
if os.getenv("IS_TESTING_TRAIN_GPU"):
TRAIN_GPU, TRAIN_NGPU = (
aip.AcceleratorType.NVIDIA_TESLA_K80,
int(os.getenv("IS_TESTING_TRAIN_GPU")),
)
else:
TRAIN_GPU, TRAIN_NGPU = (aip.AcceleratorType.NVIDIA_TESLA_K80, 1)
if os.getenv("IS_TESTING_DEPOLY_GPU"):
DEPLOY_GPU, DEPLOY_NGPU = (
aip.AcceleratorType.NVIDIA_TESLA_K80,
int(os.getenv("IS_TESTING_DEPOLY_GPU")),
)
else:
DEPLOY_GPU, DEPLOY_NGPU = (None, None)
```
#### Container (Docker) image
Next, we will set the Docker container images for training and prediction
- TensorFlow 1.15
- `gcr.io/cloud-aiplatform/training/tf-cpu.1-15:latest`
- `gcr.io/cloud-aiplatform/training/tf-gpu.1-15:latest`
- TensorFlow 2.1
- `gcr.io/cloud-aiplatform/training/tf-cpu.2-1:latest`
- `gcr.io/cloud-aiplatform/training/tf-gpu.2-1:latest`
- TensorFlow 2.2
- `gcr.io/cloud-aiplatform/training/tf-cpu.2-2:latest`
- `gcr.io/cloud-aiplatform/training/tf-gpu.2-2:latest`
- TensorFlow 2.3
- `gcr.io/cloud-aiplatform/training/tf-cpu.2-3:latest`
- `gcr.io/cloud-aiplatform/training/tf-gpu.2-3:latest`
- TensorFlow 2.4
- `gcr.io/cloud-aiplatform/training/tf-cpu.2-4:latest`
- `gcr.io/cloud-aiplatform/training/tf-gpu.2-4:latest`
- XGBoost
- `gcr.io/cloud-aiplatform/training/xgboost-cpu.1-1`
- Scikit-learn
- `gcr.io/cloud-aiplatform/training/scikit-learn-cpu.0-23:latest`
- Pytorch
- `gcr.io/cloud-aiplatform/training/pytorch-cpu.1-4:latest`
- `gcr.io/cloud-aiplatform/training/pytorch-cpu.1-5:latest`
- `gcr.io/cloud-aiplatform/training/pytorch-cpu.1-6:latest`
- `gcr.io/cloud-aiplatform/training/pytorch-cpu.1-7:latest`
For the latest list, see [Pre-built containers for training](https://cloud.google.com/vertex-ai/docs/training/pre-built-containers).
- TensorFlow 1.15
- `gcr.io/cloud-aiplatform/prediction/tf-cpu.1-15:latest`
- `gcr.io/cloud-aiplatform/prediction/tf-gpu.1-15:latest`
- TensorFlow 2.1
- `gcr.io/cloud-aiplatform/prediction/tf2-cpu.2-1:latest`
- `gcr.io/cloud-aiplatform/prediction/tf2-gpu.2-1:latest`
- TensorFlow 2.2
- `gcr.io/cloud-aiplatform/prediction/tf2-cpu.2-2:latest`
- `gcr.io/cloud-aiplatform/prediction/tf2-gpu.2-2:latest`
- TensorFlow 2.3
- `gcr.io/cloud-aiplatform/prediction/tf2-cpu.2-3:latest`
- `gcr.io/cloud-aiplatform/prediction/tf2-gpu.2-3:latest`
- XGBoost
- `gcr.io/cloud-aiplatform/prediction/xgboost-cpu.1-2:latest`
- `gcr.io/cloud-aiplatform/prediction/xgboost-cpu.1-1:latest`
- `gcr.io/cloud-aiplatform/prediction/xgboost-cpu.0-90:latest`
- `gcr.io/cloud-aiplatform/prediction/xgboost-cpu.0-82:latest`
- Scikit-learn
- `gcr.io/cloud-aiplatform/prediction/sklearn-cpu.0-23:latest`
- `gcr.io/cloud-aiplatform/prediction/sklearn-cpu.0-22:latest`
- `gcr.io/cloud-aiplatform/prediction/sklearn-cpu.0-20:latest`
For the latest list, see [Pre-built containers for prediction](https://cloud.google.com/vertex-ai/docs/predictions/pre-built-containers)
```
if os.getenv("IS_TESTING_TF"):
TF = os.getenv("IS_TESTING_TF")
else:
TF = "2-1"
if TF[0] == "2":
if TRAIN_GPU:
TRAIN_VERSION = "tf-gpu.{}".format(TF)
else:
TRAIN_VERSION = "tf-cpu.{}".format(TF)
if DEPLOY_GPU:
DEPLOY_VERSION = "tf2-gpu.{}".format(TF)
else:
DEPLOY_VERSION = "tf2-cpu.{}".format(TF)
else:
if TRAIN_GPU:
TRAIN_VERSION = "tf-gpu.{}".format(TF)
else:
TRAIN_VERSION = "tf-cpu.{}".format(TF)
if DEPLOY_GPU:
DEPLOY_VERSION = "tf-gpu.{}".format(TF)
else:
DEPLOY_VERSION = "tf-cpu.{}".format(TF)
TRAIN_IMAGE = "gcr.io/cloud-aiplatform/training/{}:latest".format(TRAIN_VERSION)
DEPLOY_IMAGE = "gcr.io/cloud-aiplatform/prediction/{}:latest".format(DEPLOY_VERSION)
print("Training:", TRAIN_IMAGE, TRAIN_GPU, TRAIN_NGPU)
print("Deployment:", DEPLOY_IMAGE, DEPLOY_GPU, DEPLOY_NGPU)
```
#### Machine Type
Next, set the machine type to use for training and prediction.
- Set the variables `TRAIN_COMPUTE` and `DEPLOY_COMPUTE` to configure the compute resources for the VMs you will use for for training and prediction.
- `machine type`
- `n1-standard`: 3.75GB of memory per vCPU.
- `n1-highmem`: 6.5GB of memory per vCPU
- `n1-highcpu`: 0.9 GB of memory per vCPU
- `vCPUs`: number of \[2, 4, 8, 16, 32, 64, 96 \]
*Note: The following is not supported for training:*
- `standard`: 2 vCPUs
- `highcpu`: 2, 4 and 8 vCPUs
*Note: You may also use n2 and e2 machine types for training and deployment, but they do not support GPUs*.
```
if os.getenv("IS_TESTING_TRAIN_MACHINE"):
MACHINE_TYPE = os.getenv("IS_TESTING_TRAIN_MACHINE")
else:
MACHINE_TYPE = "n1-standard"
VCPU = "4"
TRAIN_COMPUTE = MACHINE_TYPE + "-" + VCPU
print("Train machine type", TRAIN_COMPUTE)
if os.getenv("IS_TESTING_DEPLOY_MACHINE"):
MACHINE_TYPE = os.getenv("IS_TESTING_DEPLOY_MACHINE")
else:
MACHINE_TYPE = "n1-standard"
VCPU = "4"
DEPLOY_COMPUTE = MACHINE_TYPE + "-" + VCPU
print("Deploy machine type", DEPLOY_COMPUTE)
```
# Tutorial
Now you are ready to start creating your own custom model and training for Boston Housing.
## Set up clients
The Vertex client library works as a client/server model. On your side (the Python script) you will create a client that sends requests and receives responses from the Vertex server.
You will use different clients in this tutorial for different steps in the workflow. So set them all up upfront.
- Model Service for `Model` resources.
- Pipeline Service for training.
- Endpoint Service for deployment.
- Job Service for batch jobs and custom training.
- Prediction Service for serving.
```
# client options same for all services
client_options = {"api_endpoint": API_ENDPOINT}
def create_model_client():
client = aip.ModelServiceClient(client_options=client_options)
return client
def create_pipeline_client():
client = aip.PipelineServiceClient(client_options=client_options)
return client
def create_endpoint_client():
client = aip.EndpointServiceClient(client_options=client_options)
return client
def create_prediction_client():
client = aip.PredictionServiceClient(client_options=client_options)
return client
clients = {}
clients["model"] = create_model_client()
clients["pipeline"] = create_pipeline_client()
clients["endpoint"] = create_endpoint_client()
clients["prediction"] = create_prediction_client()
for client in clients.items():
print(client)
```
## Train a model
There are two ways you can train a custom model using a container image:
- **Use a Google Cloud prebuilt container**. If you use a prebuilt container, you will additionally specify a Python package to install into the container image. This Python package contains your code for training a custom model.
- **Use your own custom container image**. If you use your own container, the container needs to contain your code for training a custom model.
## Prepare your custom job specification
Now that your clients are ready, your first step is to create a Job Specification for your custom training job. The job specification will consist of the following:
- `worker_pool_spec` : The specification of the type of machine(s) you will use for training and how many (single or distributed)
- `python_package_spec` : The specification of the Python package to be installed with the pre-built container.
### Prepare your machine specification
Now define the machine specification for your custom training job. This tells Vertex what type of machine instance to provision for the training.
- `machine_type`: The type of GCP instance to provision -- e.g., n1-standard-8.
- `accelerator_type`: The type, if any, of hardware accelerator. In this tutorial if you previously set the variable `TRAIN_GPU != None`, you are using a GPU; otherwise you will use a CPU.
- `accelerator_count`: The number of accelerators.
```
if TRAIN_GPU:
machine_spec = {
"machine_type": TRAIN_COMPUTE,
"accelerator_type": TRAIN_GPU,
"accelerator_count": TRAIN_NGPU,
}
else:
machine_spec = {"machine_type": TRAIN_COMPUTE, "accelerator_count": 0}
```
### Prepare your disk specification
(optional) Now define the disk specification for your custom training job. This tells Vertex what type and size of disk to provision in each machine instance for the training.
- `boot_disk_type`: Either SSD or Standard. SSD is faster, and Standard is less expensive. Defaults to SSD.
- `boot_disk_size_gb`: Size of disk in GB.
```
DISK_TYPE = "pd-ssd" # [ pd-ssd, pd-standard]
DISK_SIZE = 200 # GB
disk_spec = {"boot_disk_type": DISK_TYPE, "boot_disk_size_gb": DISK_SIZE}
```
### Define the worker pool specification
Next, you define the worker pool specification for your custom training job. The worker pool specification will consist of the following:
- `replica_count`: The number of instances to provision of this machine type.
- `machine_spec`: The hardware specification.
- `disk_spec` : (optional) The disk storage specification.
- `python_package`: The Python training package to install on the VM instance(s) and which Python module to invoke, along with command line arguments for the Python module.
Let's dive deeper now into the python package specification:
-`executor_image_spec`: This is the docker image which is configured for your custom training job.
-`package_uris`: This is a list of the locations (URIs) of your python training packages to install on the provisioned instance. The locations need to be in a Cloud Storage bucket. These can be either individual python files or a zip (archive) of an entire package. In the later case, the job service will unzip (unarchive) the contents into the docker image.
-`python_module`: The Python module (script) to invoke for running the custom training job. In this example, you will be invoking `trainer.task.py` -- note that it was not neccessary to append the `.py` suffix.
-`args`: The command line arguments to pass to the corresponding Pythom module. In this example, you will be setting:
- `"--model-dir=" + MODEL_DIR` : The Cloud Storage location where to store the model artifacts. There are two ways to tell the training script where to save the model artifacts:
- direct: You pass the Cloud Storage location as a command line argument to your training script (set variable `DIRECT = True`), or
- indirect: The service passes the Cloud Storage location as the environment variable `AIP_MODEL_DIR` to your training script (set variable `DIRECT = False`). In this case, you tell the service the model artifact location in the job specification.
- `"--epochs=" + EPOCHS`: The number of epochs for training.
- `"--steps=" + STEPS`: The number of steps (batches) per epoch.
- `"--distribute=" + TRAIN_STRATEGY"` : The training distribution strategy to use for single or distributed training.
- `"single"`: single device.
- `"mirror"`: all GPU devices on a single compute instance.
- `"multi"`: all GPU devices on all compute instances.
- `"--param-file=" + PARAM_FILE`: The Cloud Storage location for storing feature normalization values.
```
JOB_NAME = "custom_job_" + TIMESTAMP
MODEL_DIR = "{}/{}".format(BUCKET_NAME, JOB_NAME)
if not TRAIN_NGPU or TRAIN_NGPU < 2:
TRAIN_STRATEGY = "single"
else:
TRAIN_STRATEGY = "mirror"
EPOCHS = 20
STEPS = 100
PARAM_FILE = BUCKET_NAME + "/params.txt"
DIRECT = True
if DIRECT:
CMDARGS = [
"--model-dir=" + MODEL_DIR,
"--epochs=" + str(EPOCHS),
"--steps=" + str(STEPS),
"--distribute=" + TRAIN_STRATEGY,
"--param-file=" + PARAM_FILE,
]
else:
CMDARGS = [
"--epochs=" + str(EPOCHS),
"--steps=" + str(STEPS),
"--distribute=" + TRAIN_STRATEGY,
"--param-file=" + PARAM_FILE,
]
worker_pool_spec = [
{
"replica_count": 1,
"machine_spec": machine_spec,
"disk_spec": disk_spec,
"python_package_spec": {
"executor_image_uri": TRAIN_IMAGE,
"package_uris": [BUCKET_NAME + "/trainer_boston.tar.gz"],
"python_module": "trainer.task",
"args": CMDARGS,
},
}
]
```
### Examine the training package
#### Package layout
Before you start the training, you will look at how a Python package is assembled for a custom training job. When unarchived, the package contains the following directory/file layout.
- PKG-INFO
- README.md
- setup.cfg
- setup.py
- trainer
- \_\_init\_\_.py
- task.py
The files `setup.cfg` and `setup.py` are the instructions for installing the package into the operating environment of the Docker image.
The file `trainer/task.py` is the Python script for executing the custom training job. *Note*, when we referred to it in the worker pool specification, we replace the directory slash with a dot (`trainer.task`) and dropped the file suffix (`.py`).
#### Package Assembly
In the following cells, you will assemble the training package.
```
# Make folder for Python training script
! rm -rf custom
! mkdir custom
# Add package information
! touch custom/README.md
setup_cfg = "[egg_info]\n\ntag_build =\n\ntag_date = 0"
! echo "$setup_cfg" > custom/setup.cfg
setup_py = "import setuptools\n\nsetuptools.setup(\n\n install_requires=[\n\n 'tensorflow_datasets==1.3.0',\n\n ],\n\n packages=setuptools.find_packages())"
! echo "$setup_py" > custom/setup.py
pkg_info = "Metadata-Version: 1.0\n\nName: Boston Housing tabular regression\n\nVersion: 0.0.0\n\nSummary: Demostration training script\n\nHome-page: www.google.com\n\nAuthor: Google\n\nAuthor-email: aferlitsch@google.com\n\nLicense: Public\n\nDescription: Demo\n\nPlatform: Vertex"
! echo "$pkg_info" > custom/PKG-INFO
# Make the training subfolder
! mkdir custom/trainer
! touch custom/trainer/__init__.py
```
#### Task.py contents
In the next cell, you write the contents of the training script task.py. I won't go into detail, it's just there for you to browse. In summary:
- Get the directory where to save the model artifacts from the command line (`--model_dir`), and if not specified, then from the environment variable `AIP_MODEL_DIR`.
- Loads Boston Housing dataset from TF.Keras builtin datasets
- Builds a simple deep neural network model using TF.Keras model API.
- Compiles the model (`compile()`).
- Sets a training distribution strategy according to the argument `args.distribute`.
- Trains the model (`fit()`) with epochs specified by `args.epochs`.
- Saves the trained model (`save(args.model_dir)`) to the specified model directory.
- Saves the maximum value for each feature `f.write(str(params))` to the specified parameters file.
```
%%writefile custom/trainer/task.py
# Single, Mirror and Multi-Machine Distributed Training for Boston Housing
import tensorflow_datasets as tfds
import tensorflow as tf
from tensorflow.python.client import device_lib
import numpy as np
import argparse
import os
import sys
tfds.disable_progress_bar()
parser = argparse.ArgumentParser()
parser.add_argument('--model-dir', dest='model_dir',
default=os.getenv('AIP_MODEL_DIR'), type=str, help='Model dir.')
parser.add_argument('--lr', dest='lr',
default=0.001, type=float,
help='Learning rate.')
parser.add_argument('--epochs', dest='epochs',
default=20, type=int,
help='Number of epochs.')
parser.add_argument('--steps', dest='steps',
default=100, type=int,
help='Number of steps per epoch.')
parser.add_argument('--distribute', dest='distribute', type=str, default='single',
help='distributed training strategy')
parser.add_argument('--param-file', dest='param_file',
default='/tmp/param.txt', type=str,
help='Output file for parameters')
args = parser.parse_args()
print('Python Version = {}'.format(sys.version))
print('TensorFlow Version = {}'.format(tf.__version__))
print('TF_CONFIG = {}'.format(os.environ.get('TF_CONFIG', 'Not found')))
# Single Machine, single compute device
if args.distribute == 'single':
if tf.test.is_gpu_available():
strategy = tf.distribute.OneDeviceStrategy(device="/gpu:0")
else:
strategy = tf.distribute.OneDeviceStrategy(device="/cpu:0")
# Single Machine, multiple compute device
elif args.distribute == 'mirror':
strategy = tf.distribute.MirroredStrategy()
# Multiple Machine, multiple compute device
elif args.distribute == 'multi':
strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy()
# Multi-worker configuration
print('num_replicas_in_sync = {}'.format(strategy.num_replicas_in_sync))
def make_dataset():
# Scaling Boston Housing data features
def scale(feature):
max = np.max(feature)
feature = (feature / max).astype(np.float)
return feature, max
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.boston_housing.load_data(
path="boston_housing.npz", test_split=0.2, seed=113
)
params = []
for _ in range(13):
x_train[_], max = scale(x_train[_])
x_test[_], _ = scale(x_test[_])
params.append(max)
# store the normalization (max) value for each feature
with tf.io.gfile.GFile(args.param_file, 'w') as f:
f.write(str(params))
return (x_train, y_train), (x_test, y_test)
# Build the Keras model
def build_and_compile_dnn_model():
model = tf.keras.Sequential([
tf.keras.layers.Dense(128, activation='relu', input_shape=(13,)),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(1, activation='linear')
])
model.compile(
loss='mse',
optimizer=tf.keras.optimizers.RMSprop(learning_rate=args.lr))
return model
NUM_WORKERS = strategy.num_replicas_in_sync
# Here the batch size scales up by number of workers since
# `tf.data.Dataset.batch` expects the global batch size.
BATCH_SIZE = 16
GLOBAL_BATCH_SIZE = BATCH_SIZE * NUM_WORKERS
with strategy.scope():
# Creation of dataset, and model building/compiling need to be within
# `strategy.scope()`.
model = build_and_compile_dnn_model()
# Train the model
(x_train, y_train), (x_test, y_test) = make_dataset()
model.fit(x_train, y_train, epochs=args.epochs, batch_size=GLOBAL_BATCH_SIZE)
model.save(args.model_dir)
```
#### Store training script on your Cloud Storage bucket
Next, you package the training folder into a compressed tar ball, and then store it in your Cloud Storage bucket.
```
! rm -f custom.tar custom.tar.gz
! tar cvf custom.tar custom
! gzip custom.tar
! gsutil cp custom.tar.gz $BUCKET_NAME/trainer_boston.tar.gz
```
## Train the model using a `TrainingPipeline` resource
Now start training of your custom training job using a training pipeline on Vertex. To train the your custom model, do the following steps:
1. Create a Vertex `TrainingPipeline` resource for the `Dataset` resource.
2. Execute the pipeline to start the training.
### Create a `TrainingPipeline` resource
You may ask, what do we use a pipeline for? We typically use pipelines when the job (such as training) has multiple steps, generally in sequential order: do step A, do step B, etc. By putting the steps into a pipeline, we gain the benefits of:
1. Being reusable for subsequent training jobs.
2. Can be containerized and ran as a batch job.
3. Can be distributed.
4. All the steps are associated with the same pipeline job for tracking progress.
#### The `training_pipeline` specification
First, you need to describe a pipeline specification. Let's look into the *minimal* requirements for constructing a `training_pipeline` specification for a custom job:
- `display_name`: A human readable name for the pipeline job.
- `training_task_definition`: The training task schema.
- `training_task_inputs`: A dictionary describing the requirements for the training job.
- `model_to_upload`: A dictionary describing the specification for the (uploaded) Vertex custom `Model` resource.
- `display_name`: A human readable name for the `Model` resource.
- `artificat_uri`: The Cloud Storage path where the model artifacts are stored in SavedModel format.
- `container_spec`: This is the specification for the Docker container that will be installed on the `Endpoint` resource, from which the custom model will serve predictions.
```
from google.protobuf import json_format
from google.protobuf.struct_pb2 import Value
MODEL_NAME = "custom_pipeline-" + TIMESTAMP
PIPELINE_DISPLAY_NAME = "custom-training-pipeline" + TIMESTAMP
training_task_inputs = json_format.ParseDict(
{"workerPoolSpecs": worker_pool_spec}, Value()
)
pipeline = {
"display_name": PIPELINE_DISPLAY_NAME,
"training_task_definition": CUSTOM_TASK_GCS_PATH,
"training_task_inputs": training_task_inputs,
"model_to_upload": {
"display_name": PIPELINE_DISPLAY_NAME + "-model",
"artifact_uri": MODEL_DIR,
"container_spec": {"image_uri": DEPLOY_IMAGE},
},
}
print(pipeline)
```
#### Create the training pipeline
Use this helper function `create_pipeline`, which takes the following parameter:
- `training_pipeline`: the full specification for the pipeline training job.
The helper function calls the pipeline client service's `create_pipeline` method, which takes the following parameters:
- `parent`: The Vertex location root path for your `Dataset`, `Model` and `Endpoint` resources.
- `training_pipeline`: The full specification for the pipeline training job.
The helper function will return the Vertex fully qualified identifier assigned to the training pipeline, which is saved as `pipeline.name`.
```
def create_pipeline(training_pipeline):
try:
pipeline = clients["pipeline"].create_training_pipeline(
parent=PARENT, training_pipeline=training_pipeline
)
print(pipeline)
except Exception as e:
print("exception:", e)
return None
return pipeline
response = create_pipeline(pipeline)
```
Now save the unique identifier of the training pipeline you created.
```
# The full unique ID for the pipeline
pipeline_id = response.name
# The short numeric ID for the pipeline
pipeline_short_id = pipeline_id.split("/")[-1]
print(pipeline_id)
```
### Get information on a training pipeline
Now get pipeline information for just this training pipeline instance. The helper function gets the job information for just this job by calling the the job client service's `get_training_pipeline` method, with the following parameter:
- `name`: The Vertex fully qualified pipeline identifier.
When the model is done training, the pipeline state will be `PIPELINE_STATE_SUCCEEDED`.
```
def get_training_pipeline(name, silent=False):
response = clients["pipeline"].get_training_pipeline(name=name)
if silent:
return response
print("pipeline")
print(" name:", response.name)
print(" display_name:", response.display_name)
print(" state:", response.state)
print(" training_task_definition:", response.training_task_definition)
print(" training_task_inputs:", dict(response.training_task_inputs))
print(" create_time:", response.create_time)
print(" start_time:", response.start_time)
print(" end_time:", response.end_time)
print(" update_time:", response.update_time)
print(" labels:", dict(response.labels))
return response
response = get_training_pipeline(pipeline_id)
```
# Deployment
Training the above model may take upwards of 20 minutes time.
Once your model is done training, you can calculate the actual time it took to train the model by subtracting `end_time` from `start_time`. For your model, you will need to know the fully qualified Vertex Model resource identifier, which the pipeline service assigned to it. You can get this from the returned pipeline instance as the field `model_to_deploy.name`.
```
while True:
response = get_training_pipeline(pipeline_id, True)
if response.state != aip.PipelineState.PIPELINE_STATE_SUCCEEDED:
print("Training job has not completed:", response.state)
model_to_deploy_id = None
if response.state == aip.PipelineState.PIPELINE_STATE_FAILED:
raise Exception("Training Job Failed")
else:
model_to_deploy = response.model_to_upload
model_to_deploy_id = model_to_deploy.name
print("Training Time:", response.end_time - response.start_time)
break
time.sleep(60)
print("model to deploy:", model_to_deploy_id)
if not DIRECT:
MODEL_DIR = MODEL_DIR + "/model"
model_path_to_deploy = MODEL_DIR
```
## Load the saved model
Your model is stored in a TensorFlow SavedModel format in a Cloud Storage bucket. Now load it from the Cloud Storage bucket, and then you can do some things, like evaluate the model, and do a prediction.
To load, you use the TF.Keras `model.load_model()` method passing it the Cloud Storage path where the model is saved -- specified by `MODEL_DIR`.
```
import tensorflow as tf
model = tf.keras.models.load_model(MODEL_DIR)
```
## Evaluate the model
Now let's find out how good the model is.
### Load evaluation data
You will load the Boston Housing test (holdout) data from `tf.keras.datasets`, using the method `load_data()`. This will return the dataset as a tuple of two elements. The first element is the training data and the second is the test data. Each element is also a tuple of two elements: the feature data, and the corresponding labels (median value of owner-occupied home).
You don't need the training data, and hence why we loaded it as `(_, _)`.
Before you can run the data through evaluation, you need to preprocess it:
x_test:
1. Normalize (rescaling) the data in each column by dividing each value by the maximum value of that column. This will replace each single value with a 32-bit floating point number between 0 and 1.
```
import numpy as np
from tensorflow.keras.datasets import boston_housing
(_, _), (x_test, y_test) = boston_housing.load_data(
path="boston_housing.npz", test_split=0.2, seed=113
)
def scale(feature):
max = np.max(feature)
feature = (feature / max).astype(np.float32)
return feature
# Let's save one data item that has not been scaled
x_test_notscaled = x_test[0:1].copy()
for _ in range(13):
x_test[_] = scale(x_test[_])
x_test = x_test.astype(np.float32)
print(x_test.shape, x_test.dtype, y_test.shape)
print("scaled", x_test[0])
print("unscaled", x_test_notscaled)
```
### Perform the model evaluation
Now evaluate how well the model in the custom job did.
```
model.evaluate(x_test, y_test)
```
## Upload the model for serving
Next, you will upload your TF.Keras model from the custom job to Vertex `Model` service, which will create a Vertex `Model` resource for your custom model. During upload, you need to define a serving function to convert data to the format your model expects. If you send encoded data to Vertex, your serving function ensures that the data is decoded on the model server before it is passed as input to your model.
### How does the serving function work
When you send a request to an online prediction server, the request is received by a HTTP server. The HTTP server extracts the prediction request from the HTTP request content body. The extracted prediction request is forwarded to the serving function. For Google pre-built prediction containers, the request content is passed to the serving function as a `tf.string`.
The serving function consists of two parts:
- `preprocessing function`:
- Converts the input (`tf.string`) to the input shape and data type of the underlying model (dynamic graph).
- Performs the same preprocessing of the data that was done during training the underlying model -- e.g., normalizing, scaling, etc.
- `post-processing function`:
- Converts the model output to format expected by the receiving application -- e.q., compresses the output.
- Packages the output for the the receiving application -- e.g., add headings, make JSON object, etc.
Both the preprocessing and post-processing functions are converted to static graphs which are fused to the model. The output from the underlying model is passed to the post-processing function. The post-processing function passes the converted/packaged output back to the HTTP server. The HTTP server returns the output as the HTTP response content.
One consideration you need to consider when building serving functions for TF.Keras models is that they run as static graphs. That means, you cannot use TF graph operations that require a dynamic graph. If you do, you will get an error during the compile of the serving function which will indicate that you are using an EagerTensor which is not supported.
## Get the serving function signature
You can get the signatures of your model's input and output layers by reloading the model into memory, and querying it for the signatures corresponding to each layer.
When making a prediction request, you need to route the request to the serving function instead of the model, so you need to know the input layer name of the serving function -- which you will use later when you make a prediction request.
```
loaded = tf.saved_model.load(model_path_to_deploy)
serving_input = list(
loaded.signatures["serving_default"].structured_input_signature[1].keys()
)[0]
print("Serving function input:", serving_input)
```
### Upload the model
Use this helper function `upload_model` to upload your model, stored in SavedModel format, up to the `Model` service, which will instantiate a Vertex `Model` resource instance for your model. Once you've done that, you can use the `Model` resource instance in the same way as any other Vertex `Model` resource instance, such as deploying to an `Endpoint` resource for serving predictions.
The helper function takes the following parameters:
- `display_name`: A human readable name for the `Endpoint` service.
- `image_uri`: The container image for the model deployment.
- `model_uri`: The Cloud Storage path to our SavedModel artificat. For this tutorial, this is the Cloud Storage location where the `trainer/task.py` saved the model artifacts, which we specified in the variable `MODEL_DIR`.
The helper function calls the `Model` client service's method `upload_model`, which takes the following parameters:
- `parent`: The Vertex location root path for `Dataset`, `Model` and `Endpoint` resources.
- `model`: The specification for the Vertex `Model` resource instance.
Let's now dive deeper into the Vertex model specification `model`. This is a dictionary object that consists of the following fields:
- `display_name`: A human readable name for the `Model` resource.
- `metadata_schema_uri`: Since your model was built without an Vertex `Dataset` resource, you will leave this blank (`''`).
- `artificat_uri`: The Cloud Storage path where the model is stored in SavedModel format.
- `container_spec`: This is the specification for the Docker container that will be installed on the `Endpoint` resource, from which the `Model` resource will serve predictions. Use the variable you set earlier `DEPLOY_GPU != None` to use a GPU; otherwise only a CPU is allocated.
Uploading a model into a Vertex Model resource returns a long running operation, since it may take a few moments. You call response.result(), which is a synchronous call and will return when the Vertex Model resource is ready.
The helper function returns the Vertex fully qualified identifier for the corresponding Vertex Model instance upload_model_response.model. You will save the identifier for subsequent steps in the variable model_to_deploy_id.
```
IMAGE_URI = DEPLOY_IMAGE
def upload_model(display_name, image_uri, model_uri):
model = {
"display_name": display_name,
"metadata_schema_uri": "",
"artifact_uri": model_uri,
"container_spec": {
"image_uri": image_uri,
"command": [],
"args": [],
"env": [{"name": "env_name", "value": "env_value"}],
"ports": [{"container_port": 8080}],
"predict_route": "",
"health_route": "",
},
}
response = clients["model"].upload_model(parent=PARENT, model=model)
print("Long running operation:", response.operation.name)
upload_model_response = response.result(timeout=180)
print("upload_model_response")
print(" model:", upload_model_response.model)
return upload_model_response.model
model_to_deploy_id = upload_model(
"boston-" + TIMESTAMP, IMAGE_URI, model_path_to_deploy
)
```
### Get `Model` resource information
Now let's get the model information for just your model. Use this helper function `get_model`, with the following parameter:
- `name`: The Vertex unique identifier for the `Model` resource.
This helper function calls the Vertex `Model` client service's method `get_model`, with the following parameter:
- `name`: The Vertex unique identifier for the `Model` resource.
```
def get_model(name):
response = clients["model"].get_model(name=name)
print(response)
get_model(model_to_deploy_id)
```
## Deploy the `Model` resource
Now deploy the trained Vertex custom `Model` resource. This requires two steps:
1. Create an `Endpoint` resource for deploying the `Model` resource to.
2. Deploy the `Model` resource to the `Endpoint` resource.
### Create an `Endpoint` resource
Use this helper function `create_endpoint` to create an endpoint to deploy the model to for serving predictions, with the following parameter:
- `display_name`: A human readable name for the `Endpoint` resource.
The helper function uses the endpoint client service's `create_endpoint` method, which takes the following parameter:
- `display_name`: A human readable name for the `Endpoint` resource.
Creating an `Endpoint` resource returns a long running operation, since it may take a few moments to provision the `Endpoint` resource for serving. You call `response.result()`, which is a synchronous call and will return when the Endpoint resource is ready. The helper function returns the Vertex fully qualified identifier for the `Endpoint` resource: `response.name`.
```
ENDPOINT_NAME = "boston_endpoint-" + TIMESTAMP
def create_endpoint(display_name):
endpoint = {"display_name": display_name}
response = clients["endpoint"].create_endpoint(parent=PARENT, endpoint=endpoint)
print("Long running operation:", response.operation.name)
result = response.result(timeout=300)
print("result")
print(" name:", result.name)
print(" display_name:", result.display_name)
print(" description:", result.description)
print(" labels:", result.labels)
print(" create_time:", result.create_time)
print(" update_time:", result.update_time)
return result
result = create_endpoint(ENDPOINT_NAME)
```
Now get the unique identifier for the `Endpoint` resource you created.
```
# The full unique ID for the endpoint
endpoint_id = result.name
# The short numeric ID for the endpoint
endpoint_short_id = endpoint_id.split("/")[-1]
print(endpoint_id)
```
### Compute instance scaling
You have several choices on scaling the compute instances for handling your online prediction requests:
- Single Instance: The online prediction requests are processed on a single compute instance.
- Set the minimum (`MIN_NODES`) and maximum (`MAX_NODES`) number of compute instances to one.
- Manual Scaling: The online prediction requests are split across a fixed number of compute instances that you manually specified.
- Set the minimum (`MIN_NODES`) and maximum (`MAX_NODES`) number of compute instances to the same number of nodes. When a model is first deployed to the instance, the fixed number of compute instances are provisioned and online prediction requests are evenly distributed across them.
- Auto Scaling: The online prediction requests are split across a scaleable number of compute instances.
- Set the minimum (`MIN_NODES`) number of compute instances to provision when a model is first deployed and to de-provision, and set the maximum (`MAX_NODES) number of compute instances to provision, depending on load conditions.
The minimum number of compute instances corresponds to the field `min_replica_count` and the maximum number of compute instances corresponds to the field `max_replica_count`, in your subsequent deployment request.
```
MIN_NODES = 1
MAX_NODES = 1
```
### Deploy `Model` resource to the `Endpoint` resource
Use this helper function `deploy_model` to deploy the `Model` resource to the `Endpoint` resource you created for serving predictions, with the following parameters:
- `model`: The Vertex fully qualified model identifier of the model to upload (deploy) from the training pipeline.
- `deploy_model_display_name`: A human readable name for the deployed model.
- `endpoint`: The Vertex fully qualified endpoint identifier to deploy the model to.
The helper function calls the `Endpoint` client service's method `deploy_model`, which takes the following parameters:
- `endpoint`: The Vertex fully qualified `Endpoint` resource identifier to deploy the `Model` resource to.
- `deployed_model`: The requirements specification for deploying the model.
- `traffic_split`: Percent of traffic at the endpoint that goes to this model, which is specified as a dictionary of one or more key/value pairs.
- If only one model, then specify as **{ "0": 100 }**, where "0" refers to this model being uploaded and 100 means 100% of the traffic.
- If there are existing models on the endpoint, for which the traffic will be split, then use `model_id` to specify as **{ "0": percent, model_id: percent, ... }**, where `model_id` is the model id of an existing model to the deployed endpoint. The percents must add up to 100.
Let's now dive deeper into the `deployed_model` parameter. This parameter is specified as a Python dictionary with the minimum required fields:
- `model`: The Vertex fully qualified model identifier of the (upload) model to deploy.
- `display_name`: A human readable name for the deployed model.
- `disable_container_logging`: This disables logging of container events, such as execution failures (default is container logging is enabled). Container logging is typically enabled when debugging the deployment and then disabled when deployed for production.
- `dedicated_resources`: This refers to how many compute instances (replicas) that are scaled for serving prediction requests.
- `machine_spec`: The compute instance to provision. Use the variable you set earlier `DEPLOY_GPU != None` to use a GPU; otherwise only a CPU is allocated.
- `min_replica_count`: The number of compute instances to initially provision, which you set earlier as the variable `MIN_NODES`.
- `max_replica_count`: The maximum number of compute instances to scale to, which you set earlier as the variable `MAX_NODES`.
#### Traffic Split
Let's now dive deeper into the `traffic_split` parameter. This parameter is specified as a Python dictionary. This might at first be a tad bit confusing. Let me explain, you can deploy more than one instance of your model to an endpoint, and then set how much (percent) goes to each instance.
Why would you do that? Perhaps you already have a previous version deployed in production -- let's call that v1. You got better model evaluation on v2, but you don't know for certain that it is really better until you deploy to production. So in the case of traffic split, you might want to deploy v2 to the same endpoint as v1, but it only get's say 10% of the traffic. That way, you can monitor how well it does without disrupting the majority of users -- until you make a final decision.
#### Response
The method returns a long running operation `response`. We will wait sychronously for the operation to complete by calling the `response.result()`, which will block until the model is deployed. If this is the first time a model is deployed to the endpoint, it may take a few additional minutes to complete provisioning of resources.
```
DEPLOYED_NAME = "boston_deployed-" + TIMESTAMP
def deploy_model(
model, deployed_model_display_name, endpoint, traffic_split={"0": 100}
):
if DEPLOY_GPU:
machine_spec = {
"machine_type": DEPLOY_COMPUTE,
"accelerator_type": DEPLOY_GPU,
"accelerator_count": DEPLOY_NGPU,
}
else:
machine_spec = {
"machine_type": DEPLOY_COMPUTE,
"accelerator_count": 0,
}
deployed_model = {
"model": model,
"display_name": deployed_model_display_name,
"dedicated_resources": {
"min_replica_count": MIN_NODES,
"max_replica_count": MAX_NODES,
"machine_spec": machine_spec,
},
"disable_container_logging": False,
}
response = clients["endpoint"].deploy_model(
endpoint=endpoint, deployed_model=deployed_model, traffic_split=traffic_split
)
print("Long running operation:", response.operation.name)
result = response.result()
print("result")
deployed_model = result.deployed_model
print(" deployed_model")
print(" id:", deployed_model.id)
print(" model:", deployed_model.model)
print(" display_name:", deployed_model.display_name)
print(" create_time:", deployed_model.create_time)
return deployed_model.id
deployed_model_id = deploy_model(model_to_deploy_id, DEPLOYED_NAME, endpoint_id)
```
## Make a online prediction request
Now do a online prediction to your deployed model.
### Get test item
You will use an example out of the test (holdout) portion of the dataset as a test item.
```
test_item = x_test[0]
test_label = y_test[0]
print(test_item.shape)
```
### Send the prediction request
Ok, now you have a test data item. Use this helper function `predict_data`, which takes the parameters:
- `data`: The test data item as a numpy 1D array of floating point values.
- `endpoint`: The Vertex fully qualified identifier for the `Endpoint` resource where the `Model` resource was deployed.
- `parameters_dict`: Additional parameters for serving.
This function uses the prediction client service and calls the `predict` method with the parameters:
- `endpoint`: The Vertex fully qualified identifier for the `Endpoint` resource where the `Model` resource was deployed.
- `instances`: A list of instances (data items) to predict.
- `parameters`: Additional parameters for serving.
To pass the test data to the prediction service, you package it for transmission to the serving binary as follows:
1. Convert the data item from a 1D numpy array to a 1D Python list.
2. Convert the prediction request to a serialized Google protobuf (`json_format.ParseDict()`)
Each instance in the prediction request is a dictionary entry of the form:
{input_name: content}
- `input_name`: the name of the input layer of the underlying model.
- `content`: The data item as a 1D Python list.
Since the `predict()` service can take multiple data items (instances), you will send your single data item as a list of one data item. As a final step, you package the instances list into Google's protobuf format -- which is what we pass to the `predict()` service.
The `response` object returns a list, where each element in the list corresponds to the corresponding image in the request. You will see in the output for each prediction:
- `predictions` -- the predicated median value of a house in units of 1K USD.
```
def predict_data(data, endpoint, parameters_dict):
parameters = json_format.ParseDict(parameters_dict, Value())
# The format of each instance should conform to the deployed model's prediction input schema.
instances_list = [{serving_input: data.tolist()}]
instances = [json_format.ParseDict(s, Value()) for s in instances_list]
response = clients["prediction"].predict(
endpoint=endpoint, instances=instances, parameters=parameters
)
print("response")
print(" deployed_model_id:", response.deployed_model_id)
predictions = response.predictions
print("predictions")
for prediction in predictions:
print(" prediction:", prediction)
predict_data(test_item, endpoint_id, None)
```
## Undeploy the `Model` resource
Now undeploy your `Model` resource from the serving `Endpoint` resoure. Use this helper function `undeploy_model`, which takes the following parameters:
- `deployed_model_id`: The model deployment identifier returned by the endpoint service when the `Model` resource was deployed to.
- `endpoint`: The Vertex fully qualified identifier for the `Endpoint` resource where the `Model` is deployed to.
This function calls the endpoint client service's method `undeploy_model`, with the following parameters:
- `deployed_model_id`: The model deployment identifier returned by the endpoint service when the `Model` resource was deployed.
- `endpoint`: The Vertex fully qualified identifier for the `Endpoint` resource where the `Model` resource is deployed.
- `traffic_split`: How to split traffic among the remaining deployed models on the `Endpoint` resource.
Since this is the only deployed model on the `Endpoint` resource, you simply can leave `traffic_split` empty by setting it to {}.
```
def undeploy_model(deployed_model_id, endpoint):
response = clients["endpoint"].undeploy_model(
endpoint=endpoint, deployed_model_id=deployed_model_id, traffic_split={}
)
print(response)
undeploy_model(deployed_model_id, endpoint_id)
```
# Cleaning up
To clean up all GCP resources used in this project, you can [delete the GCP
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
Otherwise, you can delete the individual resources you created in this tutorial:
- Dataset
- Pipeline
- Model
- Endpoint
- Batch Job
- Custom Job
- Hyperparameter Tuning Job
- Cloud Storage Bucket
```
delete_dataset = True
delete_pipeline = True
delete_model = True
delete_endpoint = True
delete_batchjob = True
delete_customjob = True
delete_hptjob = True
delete_bucket = True
# Delete the dataset using the Vertex fully qualified identifier for the dataset
try:
if delete_dataset and "dataset_id" in globals():
clients["dataset"].delete_dataset(name=dataset_id)
except Exception as e:
print(e)
# Delete the training pipeline using the Vertex fully qualified identifier for the pipeline
try:
if delete_pipeline and "pipeline_id" in globals():
clients["pipeline"].delete_training_pipeline(name=pipeline_id)
except Exception as e:
print(e)
# Delete the model using the Vertex fully qualified identifier for the model
try:
if delete_model and "model_to_deploy_id" in globals():
clients["model"].delete_model(name=model_to_deploy_id)
except Exception as e:
print(e)
# Delete the endpoint using the Vertex fully qualified identifier for the endpoint
try:
if delete_endpoint and "endpoint_id" in globals():
clients["endpoint"].delete_endpoint(name=endpoint_id)
except Exception as e:
print(e)
# Delete the batch job using the Vertex fully qualified identifier for the batch job
try:
if delete_batchjob and "batch_job_id" in globals():
clients["job"].delete_batch_prediction_job(name=batch_job_id)
except Exception as e:
print(e)
# Delete the custom job using the Vertex fully qualified identifier for the custom job
try:
if delete_customjob and "job_id" in globals():
clients["job"].delete_custom_job(name=job_id)
except Exception as e:
print(e)
# Delete the hyperparameter tuning job using the Vertex fully qualified identifier for the hyperparameter tuning job
try:
if delete_hptjob and "hpt_job_id" in globals():
clients["job"].delete_hyperparameter_tuning_job(name=hpt_job_id)
except Exception as e:
print(e)
if delete_bucket and "BUCKET_NAME" in globals():
! gsutil rm -r $BUCKET_NAME
```
| github_jupyter |
### Dependences
```
import sys
sys.path.append("../")
import math
from tqdm import tqdm
import numpy as np
import tensorflow as tf
from PIL import Image
import matplotlib.pyplot as plt
from IPython.display import clear_output
from lib.models.LinkNet import LinkNet
import lib.utils as utils
import IPython.display as ipd
```
### Loading experiment data
```
#set experiment ID
EXP_ID = "LinkNet"
utils.create_experiment_folders(EXP_ID)
utils.load_experiment_data()
```
### Model instantiation
```
model = LinkNet()
model.build((None,128,128,1))
print(model.summary())
```
### Loading Dataset
```
train_x = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/X_train.npy", mmap_mode='c')
train_y = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/y_train.npy", mmap_mode='c')
qtd_traning = train_x.shape
print("Loaded",qtd_traning, "samples")
valid_x_1 = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/X_val.npy", mmap_mode='c')
valid_y_1 = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/y_val.npy", mmap_mode='c')
qtd_traning = valid_x_1.shape
print("Loaded",qtd_traning, "samples")
```
### Dataset Normalization and Batches split
```
value = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/scale_and_shift.npy", mmap_mode='c')
print(value)
SHIFT_VALUE_X, SHIFT_VALUE_Y, SCALE_VALUE_X, SCALE_VALUE_Y = value[0], value[1], value[2], value[3]
# SHIFT_VALUE_X, SHIFT_VALUE_Y, SCALE_VALUE_X, SCALE_VALUE_Y = utils.get_shift_scale_maxmin(train_x, train_y, valid_x_1, valid_y_1)
mini_batch_size = 58
num_train_minibatches = math.floor(train_x.shape[0]/mini_batch_size)
num_val_minibatches = math.floor(valid_x_1.shape[0]/mini_batch_size)
print("train_batches:", num_train_minibatches, "valid_batches:", num_val_minibatches)
```
### Metrics
```
#default tf.keras metrics
train_loss = tf.keras.metrics.Mean(name='train_loss')
```
### Set Loss and load model weights
```
loss_object = tf.keras.losses.MeanSquaredError()
optimizer = tf.keras.optimizers.Adam(learning_rate=0.001)
#get last saved epoch index and best result in validation step
CURRENT_EPOCH, BEST_VALIDATION = utils.get_model_last_data()
if CURRENT_EPOCH > 0:
print("Loading last model state in epoch", CURRENT_EPOCH)
model.load_weights(utils.get_exp_folder_last_epoch())
print("Best validation result was PSNR=", BEST_VALIDATION)
```
### Training
```
@tf.function
def train_step(patch_x, patch_y):
with tf.GradientTape() as tape:
predictions = model(patch_x)
loss = loss_object(patch_y, predictions)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
train_loss(loss)
def valid_step(valid_x, valid_y, num_val_minibatches, mini_batch_size):
valid_mse = tf.keras.metrics.MeanSquaredError(name='train_mse')
valid_custom_metrics = utils.CustomMetric()
for i in tqdm(range(num_val_minibatches)):
data_x = valid_x[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_y = valid_y[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_x = tf.convert_to_tensor(data_x, dtype=tf.float32)
data_y = tf.convert_to_tensor(data_y, dtype=tf.float32)
data_x = ((data_x+SHIFT_VALUE_X)/SCALE_VALUE_X)+CONST_GAMA
data_y = ((data_y+SHIFT_VALUE_Y)/SCALE_VALUE_Y)+CONST_GAMA
predictions = model(data_x)
valid_mse(data_y, predictions)
predictions = predictions.numpy()
data_y = data_y.numpy()
#feed the metric evaluator
valid_custom_metrics.feed(data_y, predictions)
#get metric results
psnr, nrmse = valid_custom_metrics.result()
valid_mse_result = valid_mse.result().numpy()
valid_custom_metrics.reset_states()
valid_mse.reset_states()
return psnr, nrmse, valid_mse_result
MAX_EPOCHS = 100
EVAL_STEP = 1
CONST_GAMA = 0.001
for epoch in range(CURRENT_EPOCH, MAX_EPOCHS):
#TRAINING
print("TRAINING EPOCH", epoch)
for k in tqdm(range(0, num_train_minibatches)):
seismic_x = train_x[k * mini_batch_size : k * mini_batch_size + mini_batch_size]
seismic_y = train_y[k * mini_batch_size : k * mini_batch_size + mini_batch_size]
seismic_x = tf.convert_to_tensor(seismic_x, dtype=tf.float32)
seismic_y = tf.convert_to_tensor(seismic_y, dtype=tf.float32)
seismic_x = ((seismic_x+SHIFT_VALUE_X)/SCALE_VALUE_X)+CONST_GAMA
seismic_y = ((seismic_y+SHIFT_VALUE_Y)/SCALE_VALUE_Y)+CONST_GAMA
train_step(seismic_x, seismic_y)
#VALIDATION
if epoch%EVAL_STEP == 0:
clear_output()
print("VALIDATION EPOCH", epoch)
#saving last epoch model
model.save_weights(utils.get_exp_folder_last_epoch(), save_format='tf')
#valid with set 1
print("Validation set")
psnr_1, nrmse_1, mse_1 = valid_step(valid_x_1, valid_y_1, num_val_minibatches, mini_batch_size)
#valid with set 2
#print("Validation set 2")
#psnr_2, nrmse_2, mse_2 = valid_step(valid_x_2, valid_y_2, num_val_minibatches, mini_batch_size)
psnr_2, nrmse_2, mse_2 = 0, 0, 0
#valid with set 3
#print("Validation set 3")
#psnr_3, nrmse_3, mse_3 = valid_step(valid_x_3, valid_y_3, num_val_minibatches, mini_batch_size)
psnr_3, nrmse_3, mse_3 = 0, 0, 0
utils.update_chart_data(epoch=epoch, train_mse=train_loss.result().numpy(),
valid_mse=[mse_1,mse_2,mse_3], psnr=[psnr_1,psnr_2,psnr_3], nrmse=[nrmse_1,nrmse_2, nrmse_3])
utils.draw_chart()
#saving best validation model
if psnr_1 > BEST_VALIDATION:
BEST_VALIDATION = psnr_1
model.save_weights(utils.get_exp_folder_best_valid(), save_format='tf')
train_loss.reset_states()
utils.draw_chart()
#experimentos results
print(utils.get_experiment_results())
#load best model
model.load_weights(utils.get_exp_folder_best_valid())
CONST_GAMA = 0.001
# valid_x_1 = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/X_val.npy", mmap_mode='c')
# valid_y_1 = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/y_val.npy", mmap_mode='c')
qtd_traning = valid_x_1.shape
print("Loaded",qtd_traning, "samples")
# #normalization
# test_x = utils.shift_and_normalize(test_x, SHIFT_VALUE_X, SCALE_VALUE_X)
# test_y = utils.shift_and_normalize(test_y, SHIFT_VALUE_Y, SCALE_VALUE_Y)
#batches
num_val_minibatches = math.floor(valid_x_1.shape[0]/mini_batch_size)
# test_batches = utils.random_mini_batches(test_x, test_y, None, None, 8, seed=0)
#metrics
val_mse = tf.keras.metrics.MeanSquaredError(name='val_mse')
val_custom_metrics = utils.CustomMetric()
import json
f = open('/home/arthursrr/Documentos/Audio_Inpainting/Datasets/idx_genders_val.json', "r")
idx_gen = json.loads(f.read())
for k in idx_gen:
for i in tqdm(idx_gen[k]):
data_x = valid_x_1[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_y = valid_y_1[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_x = tf.convert_to_tensor(data_x, dtype=tf.float32)
data_y = tf.convert_to_tensor(data_y, dtype=tf.float32)
data_x = ((data_x+SHIFT_VALUE_X)/SCALE_VALUE_X)+CONST_GAMA
data_y = ((data_y+SHIFT_VALUE_Y)/SCALE_VALUE_Y)+CONST_GAMA
predictions = model(data_x)
val_mse(data_y, predictions)
predictions = predictions.numpy()
data_y = data_y.numpy()
#feed the metric evaluator
val_custom_metrics.feed(data_y, predictions)
#get metric results
psnr, nrmse = val_custom_metrics.result()
val_mse_result = val_mse.result().numpy()
val_custom_metrics.reset_states()
val_mse.reset_states()
print(k ,"\nPSNR:", psnr,"\nNRMSE:", nrmse)
# Closing file
f.close()
```
## Test
```
#load best model
model.load_weights(utils.get_exp_folder_best_valid())
CONST_GAMA = 0.001
test_x = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/X_test.npy", mmap_mode='c')
test_y = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/y_test.npy", mmap_mode='c')
qtd_traning = test_x.shape
print("Loaded",qtd_traning, "samples")
# #normalization
# test_x = utils.shift_and_normalize(test_x, SHIFT_VALUE_X, SCALE_VALUE_X)
# test_y = utils.shift_and_normalize(test_y, SHIFT_VALUE_Y, SCALE_VALUE_Y)
#batches
num_test_minibatches = math.floor(test_x.shape[0]/mini_batch_size)
# test_batches = utils.random_mini_batches(test_x, test_y, None, None, 8, seed=0)
#metrics
test_mse = tf.keras.metrics.MeanSquaredError(name='train_mse')
test_custom_metrics = utils.CustomMetric()
#test
for i in tqdm(range(num_test_minibatches)):
data_x = test_x[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_y = test_y[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_x = tf.convert_to_tensor(data_x, dtype=tf.float32)
data_y = tf.convert_to_tensor(data_y, dtype=tf.float32)
data_x = ((data_x+SHIFT_VALUE_X)/SCALE_VALUE_X)+CONST_GAMA
data_y = ((data_y+SHIFT_VALUE_Y)/SCALE_VALUE_Y)+CONST_GAMA
predictions = model(data_x)
test_mse(data_y, predictions)
predictions = predictions.numpy()
data_y = data_y.numpy()
#feed the metric evaluator
test_custom_metrics.feed(data_y, predictions)
#just show the first example of each batch until 5
# print("Spatial domain: X - Y - PREDICT - DIFF")
# plt.imshow(np.hstack((data_x[0,:,:,0], data_y[0,:,:,0], predictions[0,:,:,0], np.abs(predictions[0,:,:,0]-seismic_y[0,:,:,0]))) , cmap='Spectral', vmin=0, vmax=1)
# plt.axis('off')
# plt.pause(0.1)
#ATENÇÃO!!
#predictions = inv_shift_and_normalize(predictions, SHIFT_VALUE_Y, SCALE_VALUE_Y)
#np.save(outfile_path, predictions)
#get metric results
psnr, nrmse = test_custom_metrics.result()
test_mse_result = test_mse.result().numpy()
test_custom_metrics.reset_states()
test_mse.reset_states()
print("PSNR:", psnr,"\nNRMSE", nrmse)
#load best model
model.load_weights(utils.get_exp_folder_best_valid())
CONST_GAMA = 0.001
test_x = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/X_test.npy", mmap_mode='c')
test_y = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/y_test.npy", mmap_mode='c')
qtd_traning = test_x.shape
print("Loaded",qtd_traning, "samples")
# #normalization
# test_x = utils.shift_and_normalize(test_x, SHIFT_VALUE_X, SCALE_VALUE_X)
# test_y = utils.shift_and_normalize(test_y, SHIFT_VALUE_Y, SCALE_VALUE_Y)
#batches
num_test_minibatches = math.floor(test_x.shape[0]/mini_batch_size)
# test_batches = utils.random_mini_batches(test_x, test_y, None, None, 8, seed=0)
#metrics
test_mse = tf.keras.metrics.MeanSquaredError(name='test_mse')
test_custom_metrics = utils.CustomMetric()
import json
f = open('/home/arthursrr/Documentos/Audio_Inpainting/Datasets/idx_genders_test.json', "r")
idx_gen = json.loads(f.read())
for k in idx_gen:
for i in tqdm(idx_gen[k]):
data_x = test_x[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_y = test_y[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_x = tf.convert_to_tensor(data_x, dtype=tf.float32)
data_y = tf.convert_to_tensor(data_y, dtype=tf.float32)
data_x = ((data_x+SHIFT_VALUE_X)/SCALE_VALUE_X)+CONST_GAMA
data_y = ((data_y+SHIFT_VALUE_Y)/SCALE_VALUE_Y)+CONST_GAMA
predictions = model(data_x)
test_mse(data_y, predictions)
predictions = predictions.numpy()
data_y = data_y.numpy()
#feed the metric evaluator
test_custom_metrics.feed(data_y, predictions)
#get metric results
psnr, nrmse = test_custom_metrics.result()
test_mse_result = test_mse.result().numpy()
test_custom_metrics.reset_states()
test_mse.reset_states()
print(k ,"\nPSNR:", psnr,"\nNRMSE:", nrmse)
# Closing file
f.close()
def griffin_lim(S, frame_length=256, fft_length=255, stride=64):
'''
TensorFlow implementation of Griffin-Lim
Based on https://github.com/Kyubyong/tensorflow-exercises/blob/master/Audio_Processing.ipynb
'''
S = tf.expand_dims(S, 0)
S_complex = tf.identity(tf.cast(S, dtype=tf.complex64))
y = tf.signal.inverse_stft(S_complex, frame_length, stride, fft_length=fft_length)
for i in range(1000):
est = tf.signal.stft(y, frame_length, stride, fft_length=fft_length)
angles = est / tf.cast(tf.maximum(1e-16, tf.abs(est)), tf.complex64)
y = tf.signal.inverse_stft(S_complex * angles, frame_length, stride, fft_length=fft_length)
return tf.squeeze(y, 0)
model.load_weights(utils.get_exp_folder_best_valid())
test_x = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/X_test.npy", mmap_mode='c')
test_y = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/y_test.npy", mmap_mode='c')
qtd_traning = test_x.shape
print("Loaded",qtd_traning, "samples")
#batches
num_test_minibatches = math.floor(test_x.shape[0]/mini_batch_size)
#metrics
test_mse = tf.keras.metrics.MeanSquaredError(name='test_mse')
test_custom_metrics = utils.CustomMetric()
i=5000
CONST_GAMA = 0.001
data_x = test_x[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_y = test_y[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_x = tf.convert_to_tensor(data_x, dtype=tf.float32)
data_norm = ((data_x+SHIFT_VALUE_X)/SCALE_VALUE_X)+CONST_GAMA
predictions = model(data_norm)
predictions = utils.inv_shift_and_normalize(predictions, SHIFT_VALUE_Y, SCALE_VALUE_Y)
predictions
audio_pred = None
for i in range(0, 58):
if i==0:
audio_pred = predictions[i,:,:,0]
else:
audio_pred = np.concatenate((audio_pred, predictions[i,:,:,0]), axis=0)
audio_pred.shape
audio_corte = None
for i in range(0, 58):
if i==0:
audio_corte = data_x[i,:,:,0]
else:
audio_corte = np.concatenate((audio_corte, data_x[i,:,:,0]), axis=0)
audio_corte.shape
audio_original = None
for i in range(0, 58):
if i==0:
audio_original = data_y[i,:,:,0]
else:
audio_original = np.concatenate((audio_original, data_y[i,:,:,0]), axis=0)
audio_original.shape
wave_original = griffin_lim(audio_original, frame_length=256, fft_length=255, stride=64)
ipd.Audio(wave_original, rate=16000)
wave_corte = griffin_lim(audio_corte, frame_length=256, fft_length=255, stride=64)
ipd.Audio(wave_corte, rate=16000)
wave_pred = griffin_lim(audio_pred, frame_length=256, fft_length=255, stride=64)
ipd.Audio(wave_pred, rate=16000)
import soundfile as sf
sf.write('x.wav', wave_corte, 16000, subtype='PCM_16')
sf.write('pred.wav', wave_pred, 16000, subtype='PCM_16')
```
| github_jupyter |
#JRW, HW4 Solution, DATSCI W261, October 2015
```
!mkdir Data
!curl -L https://www.dropbox.com/s/vbalm3yva2rr86m/Consumer_Complaints.csv?dl=0 -o Data/Consumer_Complaints.csv
!curl -L https://www.dropbox.com/s/zlfyiwa70poqg74/ProductPurchaseData.txt?dl=0 -o Data/ProductPurchaseData.txt
!curl -L https://www.dropbox.com/s/6129k2urvbvobkr/topUsers_Apr-Jul_2014_1000-words.txt?dl=0 -o Data/topUsers_Apr-Jul_2014_1000-words.txt
!curl -L https://www.dropbox.com/s/w4oklbsoqefou3b/topUsers_Apr-Jul_2014_1000-words_summaries.txt?dl=0 -o Data/topUsers_Apr-Jul_2014_1000-words_summaries.txt
!curl -L https://kdd.ics.uci.edu/databases/msweb/anonymous-msweb.data.gz -o Data/anonymous-msweb.data.gz
!gunzip Data/anonymous-msweb.data.gz
ls Data
```
##HW 4.0: MRJob short answer responses
####What is mrjob? How is it different from Hadoop MapReduce?
>Mrjob is a Python package for running Hadoop streaming jobs.
Mrjob is a python-based framework that assists you in submitting your job
to the Hadoop job tracker and in running each individual step under Hadoop Streaming.
Hadoop is a general software implementation for MapReduce programming
and the MapReduce execution framework.
####What are the mapper_final(), combiner_final(), reducer_final() methods? When are they called?
>These methods run a user-defined action. They are called
as part of the life cycle (init, main, and final) of
the mapper, combiner, and reducer methods once
they have processed all input and have completed execution.
##HW4.1: Serialization short answer responses
####What is serialization in the context of mrjob or Hadoop?
>Serialization is the process of turning structured objects into a byte stream.
In the context of Hadoop, serialization is leveraged for compression
to reduce network and disk loads. Contrast this to mrjob, where serialization
is leveraged to convienentally pass structured objects
between mapper, reducer, etc. methods.
####When is it used in these frameworks?
>These frameworks accept and use a variety of serializations
for input, output, and internal transmissions of data.
####What is the default serialization mode for input and output for mrjob?
>For input, the default serialization mode is raw text (RawValueProtocol),
and for output (and internal), the default mode is JSON format (JSONProtocol).
##HW 4.2: Preprocessing logfiles on a single node
```
!head -50 Data/anonymous-msweb.data
import re
open("anonymous-msweb-preprocessed.data", "w").close
custID = "NA"
with open("Data/anonymous-msweb.data", "r") as IF:
for line in IF:
line = line.strip()
data = re.split(",",line)
if data[0] == "C":
custID = data[1]
custID = re.sub("\"","",custID)
if data[0] == "V" and not custID == "NA":
with open("anonymous-msweb-preprocessed.data", "a") as OF:
OF.writelines(line+","+"C"+","+custID+"\n")
!head -10 anonymous-msweb-preprocessed.data
!wc -l anonymous-msweb-preprocessed.data
```
##HW 4.4: Find the most frequent visitor of each page using mrjob and the output of 4.2. In this output please include the webpage URL, webpage ID and visitor ID.
```
%%writefile mostFrequentVisitors.py
#!/usr/bin/python
from mrjob.job import MRJob
from mrjob.step import MRStep
from mrjob.protocol import RawValueProtocol
import re
import operator
class mostFrequentVisitors(MRJob):
OUTPUT_PROTOCOL = RawValueProtocol
URLs = {}
def steps(self):
return [MRStep(
mapper = self.mapper,
combiner = self.combiner,
reducer_init = self.reducer_init,
reducer = self.reducer
)]
def mapper(self, _, line):
data = re.split(",",line)
pageID = data[1]
custID = data[4]
yield pageID,{custID:1}
def combiner(self,pageID,visits):
allVisits = {}
for visit in visits:
for custID in visit.keys():
allVisits.setdefault(custID,0)
allVisits[custID] += visit[custID]
yield pageID,allVisits
def reducer_init(self):
with open("anonymous-msweb.data", "r") as IF:
for line in IF:
try:
line = line.strip()
data = re.split(",",line)
URL = data[4]
pageID = data[1]
self.URLs[pageID] = URL
except IndexError:
pass
def reducer(self,pageID,visits):
allVisits = {}
for visit in visits:
for custID in visit.keys():
allVisits.setdefault(custID,0)
allVisits[custID] += visit[custID]
custID = max(allVisits.items(), key=operator.itemgetter(1))[0]
yield None,self.URLs[pageID]+","+pageID+","+custID+","+str(allVisits[custID])
if __name__ == '__main__':
mostFrequentVisitors.run()
!chmod +x mostFrequentVisitors.py
!python mostFrequentVisitors.py anonymous-msweb-preprocessed.data --file Data/anonymous-msweb.data > mostFrequentVisitors.txt
```
####Check on output
Note that in the output below, the number of visits (col. 4)
for each frequent visitor of each web page is 1.
So, no page had > 1 visits by any individual,
and technically every visitor was a most frequent visitor.
```
!head -25 mostFrequentVisitors.txt
```
##HW 4.5: K-means clustering of Twitter users with 1,000 words as features
###MRJob class for 1k dimensional k-means clustering used in parts (A-D)
```
%%writefile kMeans.py
#!/usr/bin/env python
from mrjob.job import MRJob
from mrjob.step import MRStep
import re
class kMeans(MRJob):
def steps(self):
return [MRStep(
mapper_init = self.mapper_init,
mapper = self.mapper,
combiner = self.combiner,
reducer = self.reducer
)]
## mapper_init is responsible for reading in the centroids.
def mapper_init(self):
self.centroid_points = [map(float,s.split('\n')[0].split(',')) for s in open("centroids.txt").readlines()]
## mapper is responsible for finding the centroid
## that is closest to the user (line), and then
## passing along the closest centroid's idx with the user vector as:
## (k,v) = (idx,[users,1,vector])
## where 'users' initially is a singleton vector, [ID]
def mapper(self, _, datstr):
total = 0
data = re.split(',',datstr)
ID = data[0]
code = int(data[1])
users = [ID]
codes = [0,0,0,0]
codes[code] = 1
coords = [float(data[i+3])/float(data[2]) for i in range(1000)]
for coord in coords:
total += coord
minDist = 0
IDX = -1
for idx in range(len(self.centroid_points)):
centroid = self.centroid_points[idx]
dist = 0
for ix in range(len(coords)):
dist += (centroid[ix]-coords[ix])**2
dist = dist ** 0.5
if minDist:
if dist < minDist:
minDist = dist
IDX = idx
else:
minDist = dist
IDX = idx
yield (IDX,[users,1,coords,codes])
## combiner takes the mapper output and aggregates (sum) by idx-key
def combiner(self,IDX,data):
N = 0
sumCoords = [0*num for num in range(1000)]
sumCodes = [0,0,0,0]
users = []
for line in data:
users.extend(line[0])
N += line[1]
coords = line[2]
codes = line[3]
sumCoords = [sumCoords[i]+coords[i] for i in range(len(sumCoords))]
sumCodes = [sumCodes[i]+codes[i] for i in range(len(sumCodes))]
yield (IDX,[users,N,sumCoords,sumCodes])
## reducer finishes aggregating all mapper outputs
## and then takes the means by idx-key.
def reducer(self,IDX,data):
N = 0
sumCoords = [0*num for num in range(1000)]
sumCodes = [0,0,0,0]
users = []
for line in data:
users.extend(line[0])
N += line[1]
coords = line[2]
codes = line[3]
sumCoords = [sumCoords[i]+coords[i] for i in range(len(sumCoords))]
sumCodes = [sumCodes[i]+codes[i] for i in range(len(sumCodes))]
centroid = [sumCoords[i]/N for i in range(len(sumCoords))]
yield (IDX,[users,N,centroid,sumCodes])
if __name__ == '__main__':
kMeans.run()
%%writefile kMeans_driver.py
#!/usr/bin/env python
from numpy import random
from kMeans import kMeans
import re,sys
mr_job = kMeans(args=["topUsers_Apr-Jul_2014_1000-words.txt","--file","centroids.txt"])
thresh = 0.0001
scriptName,part = sys.argv
## only stop when distance is below thresh for all centroids
def stopSignal(k,thresh,newCentroids,oldCentroids):
stop = 1
for i in range(k):
dist = 0
for j in range(len(newCentroids[i])):
dist += (newCentroids[i][j] - oldCentroids[i][j]) ** 2
dist = dist ** 0.5
if (dist > thresh):
stop = 0
break
return stop
##################################################################################
# Use four centroids from the coding
##################################################################################
def startCentroidsA():
k = 4
centroids = []
for i in range(k):
rndpoints = random.sample(1000)
total = sum(rndpoints)
centroid = [pt/total for pt in rndpoints]
centroids.append(centroid)
return centroids
###################################################################################
###################################################################################
## Geneate random initial centroids around the global aggregate
###################################################################################
def startCentroidsBC(k):
counter = 0
for line in open("topUsers_Apr-Jul_2014_1000-words_summaries.txt").readlines():
if counter == 2:
data = re.split(",",line)
globalAggregate = [float(data[i+3])/float(data[2]) for i in range(1000)]
counter += 1
## perturb the global aggregate for the four initializations
centroids = []
for i in range(k):
rndpoints = random.sample(1000)
peturpoints = [rndpoints[n]/10+globalAggregate[n] for n in range(1000)]
centroids.append(peturpoints)
total = 0
for j in range(len(centroids[i])):
total += centroids[i][j]
for j in range(len(centroids[i])):
centroids[i][j] = centroids[i][j]/total
return centroids
###################################################################################
##################################################################################
# Use four centroids from the coding
##################################################################################
def startCentroidsD():
k = 4
centroids = []
counter = 0
for line in open("topUsers_Apr-Jul_2014_1000-words_summaries.txt").readlines():
if counter and counter > 1:
data = re.split(",",line)
coords = [float(data[i+3])/float(data[2]) for i in range(1000)]
centroids.append(coords)
counter += 1
return centroids
###################################################################################
if part == "A":
k = 4
centroids = startCentroidsA()
if part == "B":
k = 2
centroids = startCentroidsBC(k)
if part == "C":
k = 4
centroids = startCentroidsBC(k)
if part == "D":
k = 4
centroids = startCentroidsD()
## the totals for each user type
numType = [752,91,54,103]
numType = [float(numType[i]) for i in range(4)]
with open("centroids.txt", 'w+') as f:
for centroid in centroids:
centroid = [str(coord) for coord in centroid]
f.writelines(",".join(centroid) + "\n")
iterate = 0
stop = 0
clusters = ["NA" for i in range(k)]
N = ["NA" for i in range(k)]
while(not stop):
with mr_job.make_runner() as runner:
runner.run()
oldCentroids = centroids[:]
clusterPurities = []
for line in runner.stream_output():
key,value = mr_job.parse_output_line(line)
clusters[key] = value[0]
N[key] = value[1]
centroids[key] = value[2]
sumCodes = value[3]
clusterPurities.append(float(max(sumCodes))/float(sum(sumCodes)))
## update the centroids
with open("centroids.txt", 'w+') as f:
for centroid in centroids:
centroid = [str(coord) for coord in centroid]
f.writelines(",".join(centroid) + "\n")
print str(iterate+1)+","+",".join(str(purity) for purity in clusterPurities)
stop = stopSignal(k,thresh,centroids,oldCentroids)
if not iterate:
stop = 0
iterate += 1
!chmod +x kMeans.py kMeans_driver.py
```
####Run k-means for parts A-D
```
!./kMeans_driver.py A > purities-A.txt
!./kMeans_driver.py B > purities-B.txt
!./kMeans_driver.py C > purities-C.txt
!./kMeans_driver.py D > purities-D.txt
####Plot cluster purity output
from matplotlib import pyplot as plot
import numpy as np
import re
%matplotlib inline
k = 4
plt.figure(figsize=(15, 15))
## function loads data from any of the 4 initializations
def loadData(filename):
purities = {}
f = open(filename, 'r')
for line in f:
line = line.strip()
data = re.split(",",line)
iterations.append(int(data[0]))
i = 0
for i in range(len(data)):
if i:
purities.setdefault(i,[])
purities[i].append(float(data[i]))
return purities
## load purities for initialization A
purities = {}
purities = loadData("purities-A.txt")
iterations = [i+1 for i in range(len(purities[1]))]
## plot purities for initialization A
plot.subplot(2,2,1)
plot.axis([0.25, max(iterations)+0.25,0.45, 1.01])
plot.plot(iterations,purities[1],'b',lw=2)
plot.plot(iterations,purities[2],'r',lw=2)
plot.plot(iterations,purities[3],'g',lw=2)
plot.plot(iterations,purities[4],'black',lw=2)
plot.ylabel('Purity',fontsize=15)
plot.title("A",fontsize=20)
plot.grid(True)
## load purities for initialization A
purities = {}
purities = loadData("purities-B.txt")
iterations = [i+1 for i in range(len(purities[1]))]
## plot purities for initialization B
plot.subplot(2,2,2)
plot.axis([0.25, max(iterations)+0.25,0.45, 1.01])
plot.plot(iterations,purities[1],'b',lw=2)
plot.plot(iterations,purities[2],'r',lw=2)
plot.title("B",fontsize=20)
plot.grid(True)
## load purities for initialization C
purities = {}
purities = loadData("purities-C.txt")
iterations = [i+1 for i in range(len(purities[1]))]
## plot purities for initialization C
plot.subplot(2,2,3)
plot.axis([0.25, max(iterations)+0.25,0.45, 1.01])
plot.plot(iterations,purities[1],'b',lw=2)
plot.plot(iterations,purities[2],'r',lw=2)
plot.plot(iterations,purities[3],'g',lw=2)
plot.plot(iterations,purities[4],'black',lw=2)
plot.xlabel('Iteration',fontsize=15)
plot.ylabel('Purity',fontsize=15)
plot.title("C",fontsize=20)
plot.grid(True)
## load purities for initialization D
purities = {}
purities = loadData("purities-D.txt")
iterations = [i+1 for i in range(len(purities[1]))]
## plot purities for initialization D
plot.subplot(2,2,4)
plot.axis([0.25, max(iterations)+0.25,0.45, 1.01])
plot.plot(iterations,purities[1],'b',lw=2)
plot.plot(iterations,purities[2],'r',lw=2)
plot.plot(iterations,purities[3],'g',lw=2)
plot.plot(iterations,purities[4],'black',lw=2)
plot.xlabel('Iteration',fontsize=15)
plot.title("D",fontsize=20)
plot.grid(True)
```
###Discussion
As a general note, our comparison of initializations must be 'taken with a grain of salt,'
as A,B, and C all incorporate some randomization into initialization,
leading to results that will vary from run to run.
If we wished to compare the randomization initializations with greater confidence,
it would be best to run our experiment a number of times,
recording purities at convergence, the numbers of iterations before convergence, etc.,
and summarizing these results across runs.
The above being said, in the printout above we can see across those runs with k=4 (A,B, and D),
that the 'trained' centroid initializations (D) succeeded in converging in the fewest iterations.
In addition, we can see that when D is compared to all other initializations,
the top three (most pure) clusters are generally purer (>90%) after convergence,
indicating that the labeling of users accompanying the data are likely meaningful.
Of all of the initializations, we can see that B performs the worst with regard to
cluster purity. However, this not exactly a fair comparison, as the purities of 2 clusters
with 4 labels will necessarily be low---it is not possible to isolate all user types!
However, if we wished to see the value of this initialization
in the context of the dataset, we could take note of whether
the two most human classes (0 and 3) are clustered together,
and separate from the two most automated classes (1 and 2),
which is likely the case, but would require keeping track of the
number of each user type present in each cluster (at convergence).
Looking closer at D, we can see that one of the clusters is very non-pure,
indicating that some users were discordant with respect to their classes,
forming a mixed cluster in terms of word-frequency.
However, we must be careful not to mislead ourselves while interpreting these results,
as we do not know which user class dominates each cluster. Noting that approximately
75% of all users are labeled as human, it is possible for all clusters to be dominated
by the human class. This however, is not the case, and could be made clear through further analysis,
plotting the numbers of each type present in each cluster at convergence
(which would also inform us of the fact that the least pure cluster is a split of
robots and cyborgs, who can actually be quite similar!).
| github_jupyter |
```
%%javascript
$('#appmode-leave').hide();
$('#copy-binder-link').hide();
$('#visit-repo-link').hide();
import ipywidgets as ipw
import json
import random
import time
import pandas as pd
import os
import webbrowser
import math
from IPython.display import display, Markdown, Math, Latex, clear_output
# set kinetic parameters
with open("rate_parameters.json") as infile:
jsdata = json.load(infile)
params = jsdata["equi4"]
```
The hydrolysis Sucrose into Glucose and Fructose is catalysed by the enzyme Invertase.
\begin{equation}
Sucrose + Invertase + \mathrm{H_2O} \to Glucose + Fructose
\end{equation}
There are however several substances that can inhibit the efficacy of the catalyst
Imagine performing a series of experiments using different initial concentration of Sucrose where you measure the rate of formation of Glucose with. The results of your experiments are affected by the presence of a contaminating substance that interferes with the catalytic reaction. Although you can somewhat control the concentration of the contaminant, you cannot completely eliminate it.
1. Determine whether the contaminating substance inhibits the catalytic reaction and the type of the inhibition mechanism, *e.g.* Competitive, Uncompetitive, Non-competitive or Mixed.
2. Determine the maximum rate achieved by the reaction, $V_{max}$ and the Michaelis constant, $K_M$, in the case you could completely eliminate the contamininat.
### Tips:
- Note that every time you restart the experiment the type of the inhibition mechanism may change.
### Instructions:
- Use the slide bar below to select temperature at which you perform the virtual experiment,
- Click `Perform measurement` to run the virtual experiment and obtain the result of the experiment,
- Click `Download CSV` to export the complete data set for all the experiments as a CSV file.
```
# define path to results.csv file
respath = os.path.join(os.getcwd(), "..", "results.csv")
# delete existing result file and setup rng
if os.path.exists(respath):
os.remove(respath)
class system:
def __init__(self, vol=0, conc=0, press=0):
self.vol = vol
self.conc = conc
self.press = press
self.inhibition = 0
self.seed = 0
self.Vm = 0
self.Km = 0
self.Ki = 0
self.Kip= 0
class data:
def __init__(self, start=-1, error=0, label='none', units='pure', value=0,
minval=-1, maxval=3, text='none'):
self.start = start
self.minval = minval
self.maxval = maxval
self.error = error
self.label = label
self.units = units
self.value = value
self.text = text
# Experiment setup (+ hidden paramters)
system = system()
def initialiseExperiment():
global n
global system
global columns_list
global scatter
scatter = 0.01
n = []
columns_list = []
n.append(len(args)) # number of input adjustable parameters
n.append(len(result)) # number of results for the experiment
for i in range(0, n[0]):
columns_list.append(f"{args[i].label} [{args[i].units}]")
for i in range(0, n[1]):
columns_list.append(f"{result[i].label} [{result[i].units}]")
# Random number seed
t = int( time.time() * 1000.0 )
system.seed = ((t & 0xff000000) >> 24) + ((t & 0x00ff0000) >> 8) +((t & 0x0000ff00) << 8) +((t & 0x000000ff) << 24)
random.seed(system.seed)
# Random inhibition type
rnd = random.random()
system.inhibition = int(5 * rnd)
if (system.inhibition > 4):
system.inhibition = 4
system.Vm = params["Vm"] * (1 + random.random()/2)
system.Km = params["Km"] * (1 + random.random()/2)
system.Ki = system.Km * random.random()
system.Kip= system.Km * random.random()
# Adjustable input parameters
def initialiseVariables():
global logScale
logScale = True
global args
args = []
args.append(
data(
label = "[S]",
minval = -3,
maxval = 1,
start = 0.001,
units = "mol/L",
value = 0.
)
)
args.append(
data(
label = "[I]",
minval = -3,
maxval = 0,
start = 0.001,
units = "mol/L",
value = 0.
)
)
# Results
def initialiseResults():
global result
result = []
result.append(
data(
label = "Reaction Rate",
start = 0.,
error = random.random() / 10.,
units = "mol/L·min"
)
)
def measure():
concS = float(args[0].text.value)
concI = float(args[0].text.value)
Vm = system.Vm
Km = system.Km
Ki = system.Ki
Kip= system.Kip
# no inhibition
a = 1
ap = 1
# competitive
if (system.inhibition == 1):
a = 1 + concI / Ki
ap = 1
adp = 1
# non-competitive
elif (system.inhibition == 4):
a = 1
ap = 1 + concI / Ki
adp = 1
# un-competitive
elif (system.inhibition == 2):
a = 1
ap = 1
adp = 1. / (1 + concI / Kip)
# mixed
elif (system.inhibition == 3):
a = 1 + concI / Ki
ap = 1
adp = 1. / (1 + concI / Kip)
res = (ap * adp) * Vm * concS / ((a * adp)*Km + concS)
return res
initialiseVariables()
out_P = ipw.Output()
out_L = ipw.Output()
out_X = ipw.Output()
with out_L:
display(Markdown("[Download CSV](../results.csv)"))
def calc(btn):
out_P.clear_output()
# Measurement result
result[0].value = measure()
# Random error
result[0].error = result[0].value * scatter * (0.5 - random.random()) * 2
# Output result
out_R[0].value = f"{result[0].value + result[0].error:.3e}"
# Read previous lines
res = pd.read_csv(respath)
var_list = []
for i in range(0, n[0]):
var_list.append(args[i].text.value)
for i in range(0, n[1]):
var_list.append(result[i].value + result[i].error)
# Append result
res.loc[len(res)] = var_list
res.to_csv(respath, index=False)
with out_P:
display(res.tail(50))
def reset(btn):
if os.path.exists(respath):
os.remove(respath)
initialiseResults()
initialiseExperiment()
res = pd.DataFrame(columns=columns_list)
res.to_csv(respath, index=False)
with out_P:
out_P.clear_output()
display(res.tail(1))
with out_X:
out_X.clear_output()
btn_reset = ipw.Button(description="Restart Laboratory", layout=ipw.Layout(width="150px"))
btn_reset.on_click(reset)
btn_calc = ipw.Button(description="Perform measurement", layout=ipw.Layout(width="150px"))
btn_calc.on_click(calc)
# ---
rows = []
reset(btn_reset)
args[0].text = ipw.Text(str(args[0].start))
rows.append(ipw.HBox([ipw.Label('Initial concentration of ' + args[0].label + ' : '),args[0].text]))
args[1].text = ipw.Text(str(args[1].start))
rows.append(ipw.HBox([ipw.Label('Initial concentration of ' + args[1].label + ' : '),args[1].text]))
out_R = []
for i in range(0, n[1]):
out_R.append(ipw.Label(value=""))
rows.append(ipw.HBox([ipw.Label(value=f"Measured {result[i].label} [{result[i].units}]:",
layout=ipw.Layout(width="250px")),
out_R[i]]))
rows.append(ipw.HBox([btn_reset, btn_calc, out_L]))
def calc2(btn):
random.seed(system.seed)
rnd = random.random()
iType = int(4 * rnd) + 1
with out_X:
out_X.clear_output()
if (iType == 1):
display(Markdown(r'Competitive inhibition'))
elif (iType == 2):
display(Markdown(r'Un-Competitive inhibition'))
elif (iType == 3):
display(Markdown(r'Mixed inhibition'))
elif (iType == 4):
display(Markdown(r'Non-Competitive inhibition'))
else:
display(Markdown(r'No inhibition'))
display(Markdown(r'$K_M$ = 'rf'{system.Km:7.5}'))
display(Markdown(r'$V_{max}$ = 'rf'{system.Ki:7.5}'))
if (iType == 1) or (iType == 3) or (iType == 4):
display(Markdown(r'$K_i$ = 'rf'{system.Ki:7.5}'))
if (iType == 2) or (iType == 3) or (iType == 4):
display(Markdown(r'$K_i^\prime$ = 'rf'{system.Kip:7.5}'))
display(out_X)
btn_calc2 = ipw.Button(description="Check Inhibition Type", layout=ipw.Layout(width="150px"))
btn_calc2.on_click(calc2)
rows.append(ipw.HBox([btn_calc2]))
rows.append(ipw.HBox([out_P]))
ipw.VBox(rows)
```
| github_jupyter |
## What is NumPy?
<ul><li>NumPy is a Python library used for working with arrays.</li>
<li>It has functions for working in domain of linear algebra, fourier transform, and matrices.</li>
<li>NumPy was created in 2005 by Travis Oliphant. It is an open source project and you can use it freely.</li></ul>
## Why Use NumPy?
<ul><li>In Python we have lists that serve the purpose of arrays, but they are slow to process.</li>
<li>NumPy aims to provide an array object that is up to 50x faster than traditional Python lists.</li>
<li>The array object in NumPy is called <b>ndarray</b>, it provides a lot of supporting functions that make working with <b>ndarray</b> very easy.</li></ul>
## Import NumPy
Once NumPy is installed, import it in your applications by adding the <b>import</b> keyword:
```
import numpy
```
## NumPy as np
<ul><li>NumPy is usually imported under the np alias.</li>
<li>Create an alias with the as keyword while importing:</li></ul>
```
import numpy as np
```
## Checking NumPy Version
The version string is stored under __ __version__ __ attribute.
```
import numpy as nk
print(nk.__version__)
```
## Create a NumPy ndarray Object
<ul><li>NumPy is used to work with arrays.</li>
<li>The array object in NumPy is called <b>ndarray</b>.</li>
<li>We can create a NumPy <b>ndarray</b> object by using the <b>array()</b> function.</li>
```
import numpy as np
arr = np.array([101, 201, 301, 401, 501])
print(arr)
print(type(arr))
```
To create an <b>ndarray</b>, we can pass a list, tuple or any array-like object into the <b>array()</b> method, and it will be converted into an <b>ndarray</b>:
```
import numpy as np
nameList = ['Angel', "Shemi", "Marvel", "Linda"]
ageTuple = (41, 32, 21, 19)
gradeDict = {"CSC102": 89, "MTH 102": 77, "CHM 102": 69, "GST 102": 99}
arr_nameList = np.array(nameList)
arr_ageTuple = np.array(ageTuple)
arr_gradeDict = np.array(gradeDict)
print(arr_nameList)
print(arr_ageTuple)
print(arr_gradeDict)
```
## Dimensions in Array
A dimension in arrays is one level of array depth (nested arrays).
### 0-Dimension
0-D arrays, or Scalars, are the elements in an array. Each value in an array is a 0-D array.
```
import numpy as np
classNum = int(input("How many students are in the CSC 102 class?"))
class_arr = np.array(classNum)
if (class_arr == 1):
print("There is only ", class_arr, "student in CSC 102 class" )
else:
print("There are", class_arr, "students in CSC 102 class" )
```
### 1-D Arrays
<ul><li>An array that has 0-D arrays as its elements is called uni-dimensional or 1-D array.</li>
<li>These are the most common and basic arrays.</li>
</ul>
```
import numpy as np
arr = np.array([1, 2, 3, 4, 5])
print(arr)
```
### 2-D Arrays
<ul><li>An array that has 1-D arrays as its elements is called a 2-D array.</li>
<li>These are often used to represent matrix or 2nd order tensors.</li></ul>
```
import numpy as np
arr = np.array([[1, 2, 3], [4, 5, 6]])
print(arr)
```
### 3-D arrays
<ul><li>An array that has 2-D arrays (matrices) as its elements is called 3-D array.</li>
<li>These are often used to represent a 3rd order tensor.</li></ul>
```
import numpy as np
arr = np.array([[[1, 2, 3], [4, 5, 6]], [[1, 2, 3], [4, 5, 6]], [[1, 2, 3], [4, 5, 6]]])
print(arr)
```
## Check Number of Dimensions?
NumPy Arrays provides the <b>ndim</b> attribute that returns an integer that tells us how many dimensions the array have
```
import numpy as np
a = np.array(42)
b = np.array([[[1, 2, 3], [4, 5, 6]], [[1, 2, 3], [4, 5, 6]], [[1, 2, 3], [4, 5, 6]]])
c = np.array([[1, 2, 3], [4, 5, 6]])
d = np.array([1, 2, 3, 4, 5])
print(a.ndim)
print(b.ndim)
print(c.ndim)
print(d.ndim)
```
## Higher Dimensional Arrays
<ul><li>An array can have any number of dimensions.</li>
<li>When the array is created, you can define the number of dimensions by using the ndmin argument.</li>
<li>In this array the innermost dimension (5th dim) has 4 elements, the 4th dim has 1 element that is the vector, the 3rd dim has 1 element that is the matrix with the vector, the 2nd dim has 1 element that is 3D array and 1st dim has 1 element that is a 4D array.</li></ul>
```
import numpy as np
arr = np.array([1, 2, 3, 4], ndmin=6)
print(arr)
print('number of dimensions :', arr.ndim)
```
## Access Array Elements
```
import numpy as np
arr = np.array([1, 2, 3, 4])
print(arr[1])
```
## Access 2-D Arrays
```
import numpy as np
arr = np.array([[1,2,3,4,5], [6,7,8,9,10]])
print('5th element on 2nd row: ', arr[1, 4])
```
## Access 3-D Arrays
```
import numpy as np
arr = np.array([[[1, 2, 3], [4, 5, 6]], [[7, 8, 9], [10, 11, 12]]])
print(arr[0, 1, 2])
```
## Negative Indexing
Use negative indexing to access an array from the end.
```
import numpy as np
arr = np.array([[1,2,3,4,5], [6,7,8,9,10]])
print('Last element from 2nd dim: ', arr[1, -1])
```
## Slicing arrays
<ul><li>Slicing in python means taking elements from one given index to another given index.</li>
<li>We pass slice instead of index like this: <b>[start:end]</b>.</li>
<li>We can also define the step, like this: <b>[start:end:step]</b>.</li>
<li>If we don't pass start its considered 0</li>
<li>If we don't pass end its considered length of array in that dimension</li>
<li>If we don't pass step its considered 1</li></ul>
```
# Slice elements from index 1 to index 5 from the following array:
import numpy as np
arr = np.array([1, 2, 3, 4, 5, 6, 7])
print(arr[1:5])
# Slice elements from index 4 to the end of the array:
import numpy as np
arr = np.array([1, 2, 3, 4, 5, 6, 7])
print(arr[4:])
# Slice elements from the beginning to index 4 (not included):
import numpy as np
arr = np.array([1, 2, 3, 4, 5, 6, 7])
print(arr[:4])
```
## Checking the Data Type of an Array
```
import numpy as np
int_arr = np.array([1, 2, 3, 4])
str_arr = np.array(['apple', 'banana', 'cherry'])
print(int_arr.dtype)
print(str_arr.dtype)
```
## NumPy Array Copy vs View
#### The Difference Between Copy and View
<ul><li>The main difference between a copy and a view of an array is that the copy is a new array, and the view is just a view of the original array.</li>
<li>The copy owns the data and any changes made to the copy will not affect original array, and any changes made to the original array will not affect the copy.</li>
<li>The view does not own the data and any changes made to the view will affect the original array, and any changes made to the original array will affect the view.</li></ul>
### Copy
```
import numpy as np
arr = np.array([1, 2, 3, 4, 5])
x = arr.copy()
arr[0] = 42
print(arr)
print(x)
```
### View
```
import numpy as np
arr = np.array([1, 2, 3, 4, 5])
x = arr.view()
arr[0] = 42
print(arr)
print(x)
```
## Check if Array Owns its Data
```
import numpy as np
arr = np.array([1, 2, 3, 4, 5])
x = arr.copy()
y = arr.view()
print(x.base)
print(y.base)
```
## Get the Shape of an Array
```
# Print the shape of a 2-D array:
import numpy as np
arr = np.array([[1, 2, 3, 4], [5, 6, 7, 8]])
print(arr.shape)
import numpy as np
arr = np.array([1, 2, 3, 4], ndmin=5)
print(arr)
print('shape of array :', arr.shape)
```
## Iterating Arrays
```
#Iterate on each scalar element of the 2-D array:
import numpy as np
arr = np.array([[1, 2, 3], [4, 5, 6]])
for x in arr:
for y in x:
print(y,x)
# Iterate on the elements of the following 3-D array:
import numpy as np
arr = np.array([[[1, 2, 3], [4, 5, 6]], [[7, 8, 9], [10, 11, 12]]])
for x in arr:
print(x[0][1])
print(x[1][0])
import numpy as np
arr = np.array([[[1, 2, 3], [4, 5, 6]], [[7, 8, 9], [10, 11, 12]]])
for x in arr:
for y in x:
for z in y:
print(z,y,x)
```
## Joining NumPy Arrays
We pass a sequence of arrays that we want to join to the concatenate() function, along with the axis. If axis is not explicitly passed, it is taken as 0.
```
# Join two arrays
import numpy as np
arr1 = np.array([1, 2, 3])
arr2 = np.array([4, 5, 6])
arr = np.concatenate((arr1, arr2))
print(arr)
```
## Splitting NumPy Arrays
<ul><li>Splitting is reverse operation of Joining.</li>
<li>Joining merges multiple arrays into one and Splitting breaks one array into multiple.</li>
<li>We use <b>array_split()</b> for splitting arrays, we pass it the array we want to split and the number of splits.</li></ul>
```
import numpy as np
arr = np.array([1, 2, 3, 4, 5, 6])
newarr = np.array_split(arr, 3)
print(newarr)
# Access splitted arrays
import numpy as np
arr = np.array([1, 2, 3, 4, 5, 6])
newarr = np.array_split(arr, 3)
print(newarr[0])
print(newarr[1])
print(newarr[2])
```
## Splitting 2-D Arrays
```
import numpy as np
arr = np.array([[1, 2], [3, 4], [5, 6], [7, 8], [9, 10], [11, 12]])
newarr = np.array_split(arr, 3)
print(newarr)
```
| github_jupyter |
__This notebook has been made to make the datasets. Please keep in mind that:__
> This is only applicable if the provided raw original datasets are used (german, compas and drug).
> Only run this once (or not), since we already included the recreated datasets. It gives insight in how the datasets are created.
```
import os
import pandas as pd
import numpy as np
import glob
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
########################### GERMAN DATASET ###########################
def recreate_german_dataset():
file_path = os.path.join(".", "resources", "german.data")
data = pd.read_csv(file_path, delim_whitespace=True, header=None)
targets = data[data.columns[-1]] # TARGET labels
data = data.drop(20, axis=1) # drop targets before rescaling
# had to change the targets since the targets were [1,2]
targets = targets.replace({1:0, 2:1})
targets = pd.DataFrame(targets).rename(columns={targets.name:"targets"})
"""
Attribute 9 (in our dataset attribute and index 8, since we start at 0, which later becomes idx 0):
Personal status and sex
A91 : male : divorced/separated
A92 : female : divorced/separated/married
A93 : male : single
A94 : male : married/widowed
A95 : female : single
"""
## Sex attribute binary
data[8] = data[8].replace({"A91": 0, "A92": 1, "A93": 0, "A94": 0, "A95":1})
group_labels = data[8]
group_labels = pd.DataFrame(group_labels).rename(columns={group_labels.name:"group_labels"})
## Sensitive feature is sex - attribute 8, make that now index 0
sensitive_feature_idx = data.pop(8)
data.insert(0, 8, sensitive_feature_idx)
data = data.rename(columns={i:j for i,j in zip(data.columns, range(13))})
# One-hot encode all categorical variables
str_columns = []
not_str = []
for i in data.columns:
if type(data[i][0]) == str:
str_columns.append(i)
else:
not_str.append(i)
# Add one-hot encoded data to the data
dummies = pd.get_dummies(data[str_columns])
data = pd.concat([data[not_str], dummies], axis=1, join='inner')
# First rescale to mean = 0 and std = 1, before adding targets to df (otherwise targets would be rescaled as well)
for i in data.columns:
data[i] = preprocessing.scale(data[i])
# Add targets and group labels to df
dataset = pd.concat([data, targets, group_labels], axis=1, join='inner')
# Thereafter reshuffle whole dataframe
dataset = dataset.sample(frac=1, random_state=2).reset_index(drop=True)
# Split dataframe in 80-20%
train, test = train_test_split(dataset, test_size=0.2, random_state=42)
# Make variable with grouplabels
group_label_train = np.array([i[0] for i in train.loc[:, "group_labels":].to_numpy()])
group_label_test = np.array([i[0] for i in test.loc[:, "group_labels":].to_numpy()])
group_label = np.concatenate((group_label_train, group_label_test))
# Drop the grouplabels from the train and test
train = train.drop(train.columns[-1], axis=1)
test = test.drop(test.columns[-1], axis=1)
# At last make x and y
X_train = train.iloc[:, :-1].to_numpy() # exclude targets
X_test = test.iloc[:, :-1].to_numpy()
y_train = train.iloc[:, -1:].to_numpy() # targets only
y_train = np.array([i[0] for i in y_train])
y_test = test.iloc[:, -1:].to_numpy() # targets only
y_test = np.array([i[0] for i in y_test])
np.savez(os.path.join("data.npz"), X_train=X_train, Y_train=y_train, X_test=X_test, Y_test=y_test)
np.savez(os.path.join("german_group_label.npz"), group_label=group_label)
######################################################################
########################### COMPAS DATASET ###########################
def recreate_compas_dataset():
data = pd.read_csv(os.path.join(".", "resources", "compas-scores-two-years.csv"))
targets = data[data.columns[-1]]
targets = pd.DataFrame(targets).rename(columns={targets.name:"targets"})
# Used columns as specified in the paper
used_cols = ["sex", "juv_fel_count", "priors_count", "race", "age_cat",
"juv_misd_count", "c_charge_degree", "juv_other_count"]
data = data[used_cols]
# Manually change the values male to 0 and female to 1
data["sex"] = data["sex"].replace({"Male":0, "Female":1})
group_labels = data["sex"]
group_labels = pd.DataFrame(group_labels).rename(columns={group_labels.name:"group_labels"})
# One-hot encode and add to data
str_columns = [i for i in data.columns if type(data[i][0]) == str]
not_str = [i for i in data.columns if type(data[i][0]) != str]
dummies = pd.get_dummies(data[str_columns])
data = pd.concat([data[not_str], dummies], axis=1, join='inner')
# First rescale to mean = 0 and std = 1, before adding targets to df (otherwise targets would be rescaled as well)
for i in data.columns:
data[i] = preprocessing.scale(data[i])
dataset = pd.concat([data, targets, group_labels], axis=1, join='inner')
# Thereafter reshuffle whole dataframe
dataset = dataset.sample(frac=1, random_state=2).reset_index(drop=True)
# Split dataframe in 80-20%
train, test = train_test_split(dataset, test_size=0.2, random_state=42)
# Make variable with grouplabels
group_label_train = np.array([i[0] for i in train.loc[:, "group_labels":].to_numpy()])
group_label_test = np.array([i[0] for i in test.loc[:, "group_labels":].to_numpy()])
group_label = np.concatenate((group_label_train, group_label_test))
# Drop the grouplabels from the train and test
train = train.drop(train.columns[-1], axis=1)
test = test.drop(test.columns[-1], axis=1)
# At last make x and y
X_train = train.iloc[:, :-1].to_numpy() # exclude targets
X_test = test.iloc[:, :-1].to_numpy()
y_train = train.iloc[:, -1:].to_numpy() # targets only
y_train = np.array([i[0] for i in y_train])
y_test = test.iloc[:, -1:].to_numpy() # targets only
y_test = np.array([i[0] for i in y_test])
np.savez(os.path.join("compas_data.npz"), X_train=X_train, Y_train=y_train, X_test=X_test, Y_test=y_test)
np.savez(os.path.join("compas_group_label.npz"), group_label=group_label)
######################################################################
########################### DRUG DATASET ###########################
def recreate_drug_dataset():
file_path = os.path.join(".", "resources", "drug_consumption.data")
data = pd.read_csv(file_path, delimiter=",", header=None)
# Targets. In the real dataset it is attribute 21 (python goes from 0, thus 20 in our case).
targets = data.iloc[:, 20]
# They only take the first 13 attributes. See below the column specifications.
data = data.iloc[:, :13]
## Sensitive feature is gender - attribute 3, make that now index 0
sensitive_feature_idx = data.pop(2)
data.insert(0, 2, sensitive_feature_idx)
data = data.rename(columns={i:j for i,j in zip(data.columns, range(13))})
"""
Our column specifications
0 = Gender, 1 = ID, 2 = Age, 3 = Education, 4 = Country, 5 = Ethinicity, 6 = NScore, 7 = EScore,
8 = OScore, 9 = AScore, 10 = CScore, 11 = Impulsiveness, 12 = SS, 20 = TARGET
"""
"""
Problem which can be solved:
* Seven class classifications for each drug separately.
* Problem can be transformed to binary classification by union of part of classes into one new class.
For example, "Never Used", "Used over a Decade Ago" form class "Non-user" and
all other classes form class "User".
"""
"""
CL0 Never Used
CL1 Used over a Decade Ago
CL2 Used in Last Decade
CL3 Used in Last Year
CL4 Used in Last Month
CL5 Used in Last Week
CL6 Used in Last Day
"""
## had to change the targets since the targets were not binary
targets = targets.replace({"CL0":0, "CL1":1, "CL2":1, "CL3":1, "CL4":1, "CL5":1, "CL6":1})
targets = pd.DataFrame(targets).rename(columns={targets.name:"targets"})
# make group labels
group_labels = {i:(1 if data[0][i] > 0 else 0) for i in data[0].index }
group_labels = pd.DataFrame.from_dict(group_labels, orient='index', columns={"group_labels"})
# First rescale to mean = 0 and std = 1, before adding targets to df (otherwise targets would be rescaled as well)
for i in data.columns:
data[i] = preprocessing.scale(data[i])
dataset = pd.concat([data, targets, group_labels], axis=1, join='inner')
# Thereafter reshuffle whole dataframe
dataset = dataset.sample(frac=1, random_state=2).reset_index(drop=True)
# Split dataframe in 80-20%
train, test = train_test_split(dataset, test_size=0.2, random_state=42)
# Make variable with grouplabels
group_label_train = np.array([i[0] for i in train.loc[:, "group_labels":].to_numpy()])
group_label_test = np.array([i[0] for i in test.loc[:, "group_labels":].to_numpy()])
group_label = np.concatenate((group_label_train, group_label_test))
# Drop the grouplabels from the train and test
train = train.drop(train.columns[-1], axis=1)
test = test.drop(test.columns[-1], axis=1)
# At last make x and y
X_train = train.iloc[:, :-1].to_numpy() # exclude targets
X_test = test.iloc[:, :-1].to_numpy()
y_train = train.iloc[:, -1:].to_numpy() # targets only
y_train = np.array([i[0] for i in y_train])
y_test = test.iloc[:, -1:].to_numpy() # targets only
y_test = np.array([i[0] for i in y_test])
# Just a check
# print(len(X_train), len(X_test), len(y_train), len(y_test), len(group_label) == len(y_train) + len(y_test))
np.savez(os.path.join("drug2_data.npz"), X_train=X_train, Y_train=y_train, X_test=X_test, Y_test=y_test)
np.savez(os.path.join("drug2_group_label.npz"), group_label=group_label)
######################################################################
# Do not run this, since this will overwrite our datasets
# recreate_german_dataset()
# recreate_compas_dataset()
# recreate_drug_dataset()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.pipeline import make_pipeline, Pipeline
from sklearn.model_selection import train_test_split
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.feature_extraction.text import HashingVectorizer
from sklearn.datasets import fetch_20newsgroups
from sklearn.neural_network import MLPClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score, StratifiedKFold
from sklearn.preprocessing import StandardScaler, PolynomialFeatures, FunctionTransformer
from sklearn.impute import SimpleImputer
data_train = fetch_20newsgroups(subset='train')
data_test = fetch_20newsgroups(subset='test')
dir(data_train), dir(data_test)
```
# Создание пайплайна с использованием стандартных функций
```
preprocessor = Pipeline(steps=[('embeddings', HashingVectorizer()),
# ('imputer', SimpleImputer()),
# ('poly', PolynomialFeatures()),
# ('log', FunctionTransformer(np.log1p)),
# ('scaler', StandardScaler())
])
pipeline = make_pipeline(preprocessor, LogisticRegression(max_iter=10000))
pipeline.fit(data_train.data, data_train.target)
pipeline.score(data_test.data, data_test.target)
```
# Создание своего класса, преобразующего данные
```
class TextTransformer(BaseEstimator, TransformerMixin):
def __init__(self):
pass
def fit(self, X, y = None):
return self
def transform(self, X , y = None):
vectorizer = HashingVectorizer()
X = vectorizer.fit_transform(X)
return X
X_train, X_test, y_train, y_test = train_test_split(data_train.data, data_train.target,
test_size=0.3,
shuffle=True,
stratify=data_train.target,
random_state=42)
for data in [X_train, y_train, X_test, y_test]:
print(len(data))
scores = []
clf = LogisticRegression()
pipeline = make_pipeline(TextTransformer(), clf)
pipeline.fit(X_train, y_train)
scores.append(pipeline.score(data_test.data, data_test.target))
scores
cv = StratifiedKFold(n_compone)
```
#### Пример разных пайплайнов для разных типов полей
```
from sklearn import set_config
set_config(display='diagram')
```
| github_jupyter |
# COVID-19 Scientific Analysis
### What is Covid-19 ?
Coronavirus disease 2019 (COVID-19) is an infectious disease caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2).
The disease was first identified in December 2019 in Wuhan, the capital of China's Hubei province, and has since spread globally, resulting in the ongoing 2019–20 coronavirus pandemic.
The first confirmed case of what was then an unknown coronavirus was traced back to November 2019 in Hubei province.Common symptoms include fever, cough, and shortness of breath.
Other symptoms may include fatigue, muscle pain, diarrhoea, sore throat, loss of smell, and abdominal pain.The time from exposure to onset of symptoms is typically around five days but may range from two to fourteen days.
While the majority of cases result in mild symptoms, some progress to viral pneumonia and multi-organ failure.
As of 23 April 2020, more than 2.62 million cases have been reported across 185 countries and territories,resulting in more than 183,000 deaths.More than 784,000 people have recovered.
<p><a href="https://commons.wikimedia.org/wiki/File:Symptoms_of_coronavirus_disease_2019_3.0.svg#/media/File:Symptoms_of_coronavirus_disease_2019_3.0.svg"><img src="https://upload.wikimedia.org/wikipedia/commons/thumb/b/b4/Symptoms_of_coronavirus_disease_2019_3.0.svg/1200px-Symptoms_of_coronavirus_disease_2019_3.0.svg.png" alt="Symptoms of coronavirus disease 2019 3.0.svg"></a>
### Covid-19 Symptoms
COVID-19 affects different people in different ways. Most infected people will develop mild to moderate symptoms.
Common symptoms:
* **Fever**
* **Tiredness**
* **Dry Cough**
* **Aches and Pains**
* **Nasal Congestion**
* **Runny Nose**
* **Sore Throat**
* **Diarrhoea**
On average it takes 5–6 days from when someone is infected with the virus for symptoms to show, however it can take up to 14 days.
People with mild symptoms who are otherwise healthy should self-isolate. Seek medical attention if you have a fever, a cough, and difficulty breathing.
### Covid-19 Preventions
<center><img class="lqwrY" src="data:image/gif;base64,R0lGODlhkACQAPIFAMzMzP//////zJmZmelCNQAAAAAAAAAAACH5BAU2AAUALAAAAACQAJAAAAP/WLrc/hACQKq9OOvN94xgKI5kGVBdqq4qEJRwLJMoa983MO+8PODAYGrQKxodgZ9wybQMXsfoLtCsVqHS7IjaGUy+4LB4TC6HlRysdo3kDASBuHxOr9vv+LsAnVGz2VwaLnmEhYZ5NX1/f4EZg4eQkYWJGH6LRo0Yj5KcnXOUF5aXUxybcWaoqaphdaAWoqMwmRems1ZMrWmxpB51t78VuRuwuxG2FbXAysIaxMUNxwTJyst0rsHPIdHT1NWfutkP277d3cyK4QzjdOXt55XpCutz7fXvoenzcvX896/Z+uLwG+gP266A0QYCK0jAWRSECiMydFgEYkSJ1sCtsXgR/+O3YRtLkVMIAM4JhROzcBzIzV5GkEdWEmS48GUzTCLZXawlEKVNdLwEjYzIM4lHOdcaBnU0lOidpBag3kgZQ2aTR1BN0tQqwApVElaZmEqq9WelpkK+astJr6bZV0970bShFkJYsXau4dFrp6vXt/+Msd2nrOiwuM3u/Kq74O5VxYnzangC+RbjAnxoob1VNHMwxJrweEO6gUiDay0Lgw61Oqro0ac8MEC9+VdRV3uFVnZLWtACuW3b3Y7sr+wcv9QKKnCVutvwPq0/Sy7nT4fn5s71EKeDnPV06m+fbE8IjLKd8XK6w/0OPjgtpu4fx85gfjMe9QTqA4YNCpROK/+m4NfQbuvVgV9RJ81EWAofWWFcgt6hpd9/rkWX3YIdNHjFPXhkNmF80nRIEojESTXVeWexF+JrFzyIYT2brKBhEwQOqCKCs+Th1IuHzfcXWrlpxmKBtVETowoz4qJikBXWqOOIPJa42JLaCUngh1Fe2BODvf1o1pNNohhmkaplmSKEG4pZgYtbIgOmm1WyRCJ0PqbZFJuZ4BiahVqSJx2aNBKI4J4E4mkiZ3OeeSgLTK5oBx96RjiXbYlGuOgKbCaCoKZv2qiicJUWeKkKbHa3qZVqejopomZaSulup45JIZZ1whjqn6OmMKisXRKAZyM4elbOkVzWitcdaCDY3a//qJJZZptp9Crfd5Gu+Wa1crYq6qvUvklIDYTsdKunuXZQLY4o4BibuuJqiyu3kqkbgADyQtIutD0Ceqwn/OYh4LDjJrlEvf1ysmgYQhCbobT7Fuwww5q4WK7C0Rq7BK0PdwIVujhQnG+5bmQsMlmE/DuYjBAvIXLGwvpqSMcBp5zWyg4LW6/JwPlJLm80e9LfIbl6LOWUPfNLCcETx2yxykX3ewHGAregtL5KNu0znJMkPDXIFVvtNddY6xz1zF5bDXbYKC9Ndtk9n402kjIHwXbRLcPs7s5Ez80yq/gObZneGbuNatpUMw14wYIPDrfach/eb+KKF1v42o5LgrPW/3eP3XjlnPRJOOR/cg6054tPvrnohoBOqNhxA4G6vQBn3joOr6cOquyMA/Fr7ToPvDUwBHOuOnx902k6EME7fnkVQhuvevKH1833LJ5RyDzvfSmIr3+au449HdIDaL00xOd+4vdxLD+tsbasiin65jfcdwG47Wc3+raOL00Bx7gvOe/Da1b/lpOv+P2vdrebkw7k0TUDDuZ1AQxb8ZSygKT4r4Coi12iFjibBh5PVwDs0wQ5eBoPno13ItwGCCxoP6ll0EhFIqEETOg74cFQf8igAQ2FsLuiBRA7OSwBC7tnrsP9MIYyGOLsHmi2G25wB0p0oAZ6+LgUlqIHUfygueqoaLnwYe6JRcii4KDXqGeZSYY8ECNnSmK5CArwilJQo5HMcK8zrkGOHYESGO+4wzzmb49swKMfnWjHUQhykNMbYTEOiUj5qfAZjGwk8pAYjkhKkgVA3F88LHnJIuJQk/GgXx876UJAhpKTpJQVHEPZQQy6EV6KZKUDUHnJTKJRlgR0ZSpXF0tcPoCWfrSlL1c4ylpScpgRyCL8PnlLZJYQg8vspTOJCU34rXKaISBZNE3WTGz+0g1s5B296tZNb35zl0EopznPiU4bqHOd7GynruA5g97tkiL0TKY8fZPPHpzAi4P0Aj6zkAAAIfkEBQYABAAsJwAaAE0AWwAAA5ZIutz+LUgBq7046827/2AojmRpnmg4AENaUe4izXED13iuk/fu/8CgcEgsGo/DADLVWzorgKenKQ0pVchrdbulcr/gsHhMLpvP6LR6ffay3/C4fE6v2zXuS+tOiPL/Tn6AHFqDbHmGiYqLjI2Oj5CRkpMMiJSXmJlqgpqdnp8Me6CjpKWXoieWpj+qqzsDhXSxrmuoNQkAIfkEBQYABQAsJgAYAE4AXwAAA/9Yutz+DggBoL04azuC/1sojpa3eBWprhgQOC8rz0raBDat6/k+SxSMADK8uAKCga/0MVmKDuijiVpGnAVkibhlYK0KQeyknELGjZ4CBz7p0Bl2O2sWwRfqrNR6V/Uvf2+CDgRPc3QPhSF9ig95O39lG32SEYeBdiOPgzN9m5s0eyueZ6AZAJuiNZlRZ5UiR1Uhql2TK0dklhuvXoS1I2INoqa2DY0wo4lpIcQMvG5+ygybxwXNGpiAjssQz8y/rNx4c6nJuuNttNbmvuiH29HSqxDXCpXV4iORzoYh90YraOXooKOStyz1hNG7UXBSwmlcetHIsSnAwRYWjgXCZ4H/IjhYfzRy+LZAXTYj+6BlULcOGwsXC1UKAaiAo0xNf3Kc1GDD5pqXG29CFDq0wMWd9IL+jLg05kqgI5vKQPpwWqAiMEugOgmCpoqsZ4Jh+PJjFyobBN+RkMMh1ocKadXaAeWWihKkcnM96EB2zRC8eRFNwfQCcN4/YrcYluvpZIXFavvsLAT5XeUBlAMvIgGAgKLK72wQ69wI9CVUPD1L1LykMzLWPgRwNA2bgewztXX4pJ0bFO/c2oDHCy58bpzixo8jL/Z0eZxrLJ0jnCWdOM/qpUBif83s9xy2G8Bu14Jz+4mqgrGTX2s+y0Xr0tdz9u7DIlT1ndSjX+2ct/jiT+CxQF9+PAw4yn51ANiagfogeAaDDrXRF2MOjgWhVpFdCEOF4Wm4RoDv8CUhiHlJMKEfJMJmYleLeBAEdgNsRcWMTVDAoWYxnoXKAO/NkQAAIfkEBQYABQAsJgAeAE4AWAAAA+JYutwLLsrpALk46807EVQoSsNonmiqrqwDtnAszzT81nh+53zv/8CgcEgsGo/IpHLJbDqf0Kh0Sq1ar9jhLosjcL/gsHhMLlMAaGPABwi43ZCwQNN+v+NkgX1fru/fZX92W19+ghYYYIJvcx1Yi3COj5AeWAOQiBxYhnuZG5uQA5VXkAGjVpx7p1Spdp6Jk5ifXKWiGWYMGrgLjRe7D7C/BcG/mcIKxL++xwWIzMNez9HMFs/D1oTW2tvCph4dJdxVa9Z4x+TP5uLr7GDZu+Ht8vP0S+9m9/X6E+r7/tvxgCQAACH5BAUGAAUALCYAGABHAF8AAAP/SLrc/jDKSau9OOvNu/9gKI5kaZ5oqq6sAwSwMLSjAN/BTH8vfu8SAGDiw+mAjB5sCCnemBCh8KR8PgbOJQSLE5iyx0XV5304y6MsVJwNoJPgEdfpar8X7TWvXec77CNjOH1ZDnN0IoJWcH6MhYl5DYo/koCQapWNbJiXTnqTMA42nBVVegSgd6ABf5EWRXqHRZmPDG0Bp1GIC7I+DaO1eK4UpAS9gwzAuwrHi8SeDbe/0o5OYRJZd7dht6zVs6/BBLdr3bTLROLkttTCmunQ7vEK5uzv2MXr8uj1FcPjlui1W+XmQjuAmsw1I2Mw4cFuBJ09C9YPYbeLuZoMq3ix9COuhqS6HfGI8YIyH1BOotxHEpbJdhCZtdz2so2OiwpUzrxxTYJOHEJ2CgV6IeLQo6EsGEU6NCMhplDBUVgadabTb1WjXt2UNetWBVS7eiwltivZslXPooWqdi3Sr2HdFtMl9+3UukjvPIiL19cEvn0pBQk89C7hnYYPt9wKmDBjxVb/Ql48YeHkIj0bWL6MDB7ngxE+x5woeh6+0uhCo05Nd/UNva1df2zrOrNG2d7Cyf6KVbTt2Kg17NbQGC2H1bz3loYNkvOHzXVDFGf6O8P0odWJE2YO4mfX5Byu62MR1CsSsMUFAMhOY4CUn+qnnJ+vIQEAIfkEBQYABQAsJwAhADwAVQAAA/9Iutz+MMpJq7046827/2AojmRpnmiqrmzrvnAsz3Rt3yUQ7AGANwKecPAj6ITCByDYaw2QSN/iiIRQm50rVDCF8hzaHVECAIwZTO9uod5JFeGvNcpoy+M8Lrv9RnvfdmsEaWpddmB8CoEBioGGbYhqUos+iwuEXg6YVQSUnY6Nhw2gnpahkKOipaB4SaltRKuirXJ1dpWBArRuT6a2bbqLwsOCv8THyDt6xsnNnq/O0YnQ0tVCfafW2tif2t7c3uHg4dsO5N7m59Zn2erN2O7W1PHJffTVzPfIgPrRe/3N/gHc126gsEkGCXZLOAwhw4YFH0qKKBGKlE0VoYzBmNFhlZGOoj6CLARnpJdlvUxey2fSnkqPCjh2bLDr4biXD14GWMagpkF2LCXypGlywkhuRDMijfRwqASZ9JxOgKpu6QSf5ThgjWYVw9ZjXTeU0aZLxYCxx3SFRXG2jNu3QDkkAAAh+QQFBgAFACwoADQAOgBCAAAD/0i63P4wykmrvTjrzbuPgBCMwJcBaOmIYxuAqEkALsnUrsrQuYe3Kt7vxWDVdKfhSKG0KYQ45KUZAAyoggVVkhrssNDhc/sIV8dUMw6tlBKMxze1OlfAf1J1S56uE+Z5TSGAfoANaXc/iTgCizVeN3OSk36RlJeYS4eZnJMOnaBklqGkLp+lqJqjqaGnrKSur6BSsqRZq7WZm7mdt0y8oLjAlDrDwb/GmZDJnCrMmc7Pl7fS08jVc9TYktrbWNfeQ9HhbeDkcX/n4nbqPwt65+/tawpX80DC50j3e1/3vmzaueFXZp4bc95AqIMEQd8EeMYOOoDIS2JBaRYhUGSVUR7hsI4UNnISwPCDyEkgOaAYaUWGhRQhGsns4rKmzQQAIfkEBQYABAAsKAA2ADoAQAAAA/9Iutz+DEhIq71LhL0F/tcwOQBnbpA0go9mesx5Ak05s43M0YSro4tfgIcTAo0bnu2HIyx/EmQAhiQ6BgOHdLglDKStm2LrMzrBEaiX3H3+skFhdNvtLtwnAZ6u+6LHVXxSezIxSGWCiSaGio18jI6RQjCAkpYylASXmyZWnJ+en5uhopaZpZuQqI5Eq5dxrpE8sZaVtI1Zt7Kauo68vYq/wIKEw0bFxlDJxMt8yM2d0FvP0lzVx4jXndTQZ9o63t8n4eIctuWz5Tvn4nfqUwt+6KraNfM57Q7ZzVb0y/1pqlkQeKHbh2SZMOxbBRADN0kJmzxMJABOExKgLjqcuE0Ko0YRKkKqsOgxAQAh+QQFBgAEACwoADcAOgA/AAADlEi63P6QjEgrBPbhBkQIwpaNwaSYUMAAaiOMDhq10SvNcJ61tKb/qRulxxHBbJaBESITAnPLy3OqiPqoT6tDu8VyMk3vCEkJ48TotHr9JLKp7ndbLibT53d4fs/v+/+AgYFcgoWGh4iJiouMjY6PkIxxi3aMZpGYmZqbnIY0hJ2hfpeihaSlqKmqQJOrrq+wFqCgGQkAIfkEBQYABQAsKAA2ADoAQAAAA6RYutz+bMBJq23AkcurhN8TOgIwdkzGCJvSXqeiou70CdU84XTv/8AghBCzBISNVy2os/WKSAgvWoBSg9OrdntRVrLcsHhMLpvP6HQarG673/B42CpXsOtJvH7PJ9P7gIGCg4SFhhR3egNNfF6Hj3KJkHiSk2sLf5aaZI59jG9FnZtilQWlo6gWnxSrqTStcgCiPqcXtVg0t1qzCgMEv8DBwsPACQAh+QQFBgABACw4ADcAKQA+AAACOYyPmaDtD6OctNqLs968+w+G4kiW5omm6sq27gvH8kzX9o3n+s73/g8MCh2DofGITCqNgqXzSXQVAAAh+QQFEgABACwrADYAKAAMAAADIBi63P4BwElrEzbrzbtXEkd8ZGkG42mFatsOLeZaspYAACH5BAUGAAMALFAAPwABAAQAAAIDhBIFACH5BAUGAAQALCgAOgA6ADwAAANvSLrc/nAFMaO9OIeQu+/bJ45NSJ6fia6Xyr4OB890bd94ru8PwK+yHyngE4o2RaNnE1SCmk6MK9qCUi2BwVWT3EaIXkw3/BCQz+jbOM1uu9/wuHxOr9th2rt+z8et2Xl9goOEhYaHiImKi19XgTkJACH5BAUGAAUALCkAOgA+ADoAAAOpWLrc/lABQqO9OLN6gddgSAxYYAZAqFpDep0muc5MW8IuTQ/Cfea6FTACMwVpQ0gxcJz1fL+m1LFkTq+FahJLqz65zSpYqh0fy+bgcpsGrdu6N3y2lM1Vyzsdp18V7X0aRYF4UYQhJ4eFX4oZJmyND0aRgpCUDQGAl0SbnZ6fnZqgG6OlpkGMpwWWqqcErbAZr6CzramnoqW3prWxvr/AGrvBxMWgrIJBCQAh+QQFBgAFACwtADsARgA4AAADyli63P4wMkKlvTg/yrX/H9eBZOkAImGuJiqy8OeOcW0Nqa1HaXUCwMFu1VMxAIGkEjAsFY0FpTTJbIaKiqnWeu0VkFop1/OMhsXjTHFwllbTF2ybClenBObzu25JVdt7fBIib2d4ghgue2eIGgCHWWGBjRqSlCVhlySWmh9ak50WmaEeW6SlbqeVqaoZU62rSbCuS7MYtbYXSrm3Aby3oL8NwcLFjsbITsnLx8zOz83QBVDPxNLXzNaJmpAr3djg4WPfywI4T+jpFAkAIfkEBQYABQAsMQBCAEUAMAAAA6JYutz+MEpGap04a2w72WDIeZ1ongVJAMDwoXDmAUFtC0OsP55g/7UBcEgsGo0ClecY8DGfUGJSSRA+d1gLDYrVaaOBLuwb3YLPT5ITChCLOla22wSomJmCuX5x2+9tbX5zgIJ6NYGFYoeJbouMXY6PWAGIkjqUlpOVmSiYnDqbn3SiMXmknaepqqsbOayvsLGyMrO1tre4ubq7vL2+v7OuvwkAIfkEBQYABQAsKQBFAE0ALwAAA3tYutz+MMrJiKU4610IAEIhfARnnhgACSXqvqKkwrRGhHVO4c+g/xMecJgRngLE3wwT8CVpt0xg+awuptbs1am1IrtZI5goHpvP6LR6zW673/C4fN7+0u9gKn7/bPH/RHocgoCFhkWHiYqLjHFcjZCRboSSlZZmlFqPfwkAIfkEBQYABQAsLgBFAEgAMAAAA5tYutz+MEoIKpg46w2C90VQEFtpYsP3AOTpvh0IEe1rZ7EYsXevXA+ZhOe7ARtCCa3IXCR3y2ZRoJuwotLT8fkQ0GrZU0BAxWHD6IX3nEZf2/DCO56e08P2uzSvb7L7RX+APUSDTIWGPl6JfoyOj5BoA5GUjFWVWpiam5wMk52geKEaYKMQAqapqnClq659R6+ysy+ttLe4GrGdCQAh+QQFBgAFACwyAEUAQwAwAAADeVi63P4wygWmvfgCkbvv1SeODyeZZAqhEqG+TDgJLvzKE25juh4Btd1oENAEhR8fRIm8BIoWYHNKoVqv1AG2qd16v+CwgynusMqjI3odJrPf8Lic5J43znZJNz/jX/Z+K4E5g4WGh4hlgIl4iY6PkJGSYI2TlpBqgQkAIfkEBRIAAQAsWgBtAAEAAQAAAgJEAQAh+QQFBgAFACxQADcADwA4AAADPli6AzArKhBCqQHIoneO3XZB4iaUJYFGg7qC73K+ZOzGeK7v+8yvoZ9wSCwaj8ikcslsOp/QaPQmfQIIWGwCACH5BAUGAAUALCkANgA2AEAAAAODWLrc/jACIMCIOOsGgv/CJo6LEDgeqWIdFABr3JwRLd+YjZPE7juXn1AUHGJ6mqIRMoBlkMtIKHpzUq/YrPYx3Y663o01rBmTMcpzJK1uu9/wuHxOr9vv+Lx+z+/7/4CBgoOEN2CFiImBZopXjI2QOIeRlJCTP2w8lSV0AASfoKGiowkAIfkEBQYABQAsOwA1ACQAQAAAA1NYutz+AIxHqxsiaA2sd0DgBN1nilRpriPrvnAsL2pMfDU8efnsV7efcEgsGo9IVi/JbDqf0Kh0Sq1ar9isdsvter/gsHhMLpvP4h167V2yP4JnAgAh+QQFBgAFACwoADMANwBDAAADbli63P4wriGrvQ2EvQH+oKJxnBee0dh0aLtCmyubMC3fjo1jQqXvuB9wSCwaGwQL5RhJVpzMxzJ6Igiplx52y+16v+CweEwum8/otHrNbrvf8Lh8Tq/b7/h8+arv+/+AgYKDhBBQhYiJiouBh0QJACH5BAUGAAUALFAAKwAPABsAAANCSLoLwFCJQMOLpOrLtOcE4GkCI46ViabLtAbdCy/yrNQNrpxv/pY7Gac2FNKEg5sxVEsyZb6eEuqowayUjLWEfSQAACH5BAUGAAUALFAAIgAQACEAAANTWLo8w3CBQAOIstaLtX7Q5FEcI4xVhKbQSqkuNLhC+C5b2Fl6VxaEmo9BwEiIRgVIUUzKnD2o4uckUJNBKVO75Ta112TYOG4Ay4vH16nWtrVZaQIAIfkEBQYABQAsUAAeABEAJQAAA01YujwALDIRaoBy2Z0V2JbQUWDVmRoqBWswdBkLrxk2M/bt6XxG9IofUAgs5nTHm6g4LBJ5LyYwOaPCllLdU8vDRG/WDiaM2yGLIjIjAQAh+QQFBgABACxVAB4ADAAQAAADFxiqNLQwgCUeFDHrqbv/gQWOZGme5AAmACH5BAUGAAEALFYAHgAKABsAAAMkSLrM8G2EOceiWAFMN/9gIIVkaQomiIZe6r7hCBJwbZcrCBAJACH5BAUSAAAALFAAIAAQAB0AAAMaCLpKDO7JSau9OOvNu+hgKI5kaZ5oqoLDlgAAIfkEBQYABQAsQwAfAB0AIgAAA0BYutyupMRHq1i36s27/xgojmRpnmh6AmrrvnAsz3Q9ACwKBDyfkYJAg5cDFR1C4+bISVacNdOkNVUNXldXtpMAACH5BAUGAAUALEIANAAPAA8AAAMmSEoj8QGsCaANk9RrJ7+S9llCYZ4oGqRsOrRwLM90IY5dbesEDiYAIfkEBQYABAAsQgAzAA4AEAAAAyJIqhAeYK32qhw1h6VzlOAihOS2kWiqrmzrrlRXxbIj1FUCACH5BAUGAAUALEIAMQAOABIAAAMqWKogEQCs+eoTs1QasHrZAhVAEE6SdyqYuVJvLMfEXNTzYO98L5eWYCUBACH5BAUGAAUALEIAKQAOABoAAAM+SLo0/ASEOQGUlNqFM2WetxGhpwxl5qBp1bRul1pyScOBEOE33Le/2c43BBaFtVDQdmQmRYNnBhDFBQACawIAIfkEBQYABQAsQgAfAA4AJAAAA09IugTAkIQ5X6TYLoAx7NkygJTGkYGwodPKuihMyiDd2Z5yxjob4BRgS+ELWIpH34CoVAh8i+eLOZVAqaillaUtdptbVJI1FmNJZbSjGEgAACH5BAUGAAUALEMAGQANACoAAANXSEogEWBJ8GqQyta4qH7T9wydCC2OyBHpt7baap6ZKdvLfKs5XvM/Vw9ImAmGn2NQoyyamrMLYRBVUGfWKkPrMSm6oi93jN2WwR/xmew1t9EadRsmjD4SACH5BAUGAAEALDgAGQARACEAAAIZjI85kO0Po5y02ouz3rz7D4biSJbmCQlXAQAh+QQFBgAFACw0ABkAGwAnAAADO1i63EwFAEdrGW3azbv/HQSOZGmeaKqubOu+cCzPdG3feK47WAoEQKCGJAg0gESL8dOzDDkC6M4lWiUAACH5BAUGAAUALDUAGQAZACQAAAM8WLrcTJC46Qhgg+pym9ze1IFkaZ5oqq5s675wLM90bd94ru98vwCCgDAwMgECjYDgdKQsS0hNBlTERTcJACH5BAUGAAEALDgAGQAWACIAAAMmGLrcOsPJqQC9OCuhu/9gKI5kaZ5oqq5s675wLI+DtZn2mItElgAAIfkEBQYAAQAsOAAZABYAHwAAAyAYutwKwMlJqxs26827/2AojmRpnmiqrmzrvjBIMAC2JQAh+QQFBgAFACw1ABkAGgAqAAADR1i63E3EyamEApFql7cHXiiGw1iAZqqubOu+cCzPdG3feK7vfAj8KJUgQCyWTMVFMeghSpykACXApFQd11yWYel5v+DCUZMAACH5BAUGAAUALDUAMwAOABAAAAMgWLqzviBICV6JMztsg1iBBSqVCF1mqq7syX5t3MLySiQAIfkEBQYABQAsNAAzAA4ADwAAAx9YqhAuYK0mSxCSVrbC7srwZWNpnuiHpWeEui3xpkICACH5BAUGAAUALDQAMQAOABIAAAMtSKoA0iuSQGsA0mqshNbLBxKDaA0EYFaougbdC0+yIs+3XdMvemO/kqxxEyQAACH5BAUGAAUALDMAKgASABoAAANUSLoz+1CBQAOIcFZ6sdpbF2lgIHhCWUEOoVYi0b4r89HB0+H5suO62+vkE76CLmAxSUPynEqJUdUi/Gii6ys7LXGZW9+FRySkwMebmecYcHiUVCABACH5BAUGAAUALDQAIQARACMAAANjWFqz/gqEGRpcks67NBVc4VFcNgKX6aGQqrGPq6UjZTmmM8HY9gSgxyDACTqGpQsxFUoRmrEnpgmQQhfWayF75UK9TXDotjC2IObYucTlRdXFdrhccD+sILtDHuKz61oFAhkJACH5BAUGAAUALDQAHwAMACEAAANAWAXRoLCIRh8cNEcWYrBLFxXiMioleK7RwL4wq64AAQs2nL/2fvor4MkVK8ZmJ+RIYJTEmCybUiF9EZvYzfGZAAAh+QQFBgABACw0ACAAAgACAAADA0hAkQAh+QQFBgABACw0AB8ABwAOAAACDIwVMMbtD6OctMLlCgAh+QQFBgABACwpAB8AFAAhAAADJxi6vARgkEbXaLPqXS//jQCOZGmeaKqubOu+cCzPdG3f2+CRogmkCQAh+QQFBgAFACwqAB8AEgAhAAADJli6vDSkSRUXmE1kTLn/YCiOZGmeaKqubOu+cCzP5QDcIBCwGpYAACH5BAUGAAUALCgAHwAUACIAAAMjWLrcRS4yIWsENuvNu/9gKI5kaZ5oqq5s674qpgxwFFQ0mAAAIfkEBQYAAwAsKAAgABUAIgAAAiWcjwjJPeuinBHSi7PevPsPhuJIluaJpuqaLkEgWFxACZ2s2V0BACH5BAUGAAAALCkAPQAHAAMAAAIGhIMByR0FACH5BAUGAAEALCkAPAALAAYAAAIMjDNzK6y4Foi02ioKACH5BAUGAAMALCgAPAALAAgAAAISnBEZYKYfHFQCTWXv0knw4HEFACH5BAUGAAIALCgAOwALAAgAAAIQlBEZYKYfHFQATXpxXpvvAgAh+QQFBgADACwoADoADAAJAAACE5wRGWCdPZaDkDHaBN68e351QgEAIfkEBQYAAgAsKAA5AAwACwAAAhWUERlgrc/Mm2a0i3OLuvsPhgIylQUAIfkEBQYABAAsKAA4AAwACwAAAxhICtEyKrZIApAUE5jjvR3FhWRpmmMpnAkAIfkEBQYABQAsKAA3AAwADQAAAx1YCtGisDTYBo23WIwBh8THeWIJPaaCikRgtmkqJAAh+QQFBgAFACwoADYADAAOAAADIkgE0QEqCkcjow6SiZ3qFQF64hh8JjoO5QiZTwsKMlh3QAIAIfkEBQYABQAsJwAvAA0AGAAAAzdIBNGgUIlGQ1yVishyix4VTSEXWlDnmeGjqJmLtemJzuVr16cLx7pe8DTM/YDHzCCptFEGzkYCACH5BAUGAAUALCYALAAOAB0AAANFSATRoBCy1kSENL+bqeVdM0RD6JBmsGHmqqQuAYMmHdodrkWpGgkzCDAVmdQkvWJvZQwxl8ikoumc9iiLayOrJZS632sCACH5BAUGAAUALCYAKwAMABUAAAMmWDoA+i9EB2WlBdenI4cQoXSPWJjgmYLD6r5wIcR03db4Ouc8NCQAIfkEBQYAAQAsKAArAAgACQAAAwwYNBMe4MlJq70YkwQAIfkEBQYAAQAsJgArAAoADAAAAxdIuszTCoQZmKRzYQrUnh3xjWRpniOQAAAh+QQFGAABACwnACwACgAOAAADEBhK2q7gPdGovDjrzbv/SwIAOw==" alt="" data-atf="1"></center>
<h1><center>STAY HOME.SAVE LIVES.</center></h1>
<h4><center>STAY home</center></h4>
<h4><center>KEEP a safe distance</center></h4>
<h4><center>WASH hands often</center></h4>
<h4><center>COVER your cough</center></h4>
<h4><center>SICK? Call the helpline</center></h4>
```
#Importing Libraries and Modules
from __future__ import print_function
from ipywidgets import interact, interactive, fixed, interact_manual
from IPython.core.display import display, HTML
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import plotly.express as px
import folium
import plotly.graph_objects as go
import seaborn as sns
import ipywidgets as widgets
```
# Importing Raw Data
```
#Collecting Data
death_df = pd.read_csv('https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_global.csv')
confirmed_df = pd.read_csv('https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv')
recovered_df = pd.read_csv('https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_recovered_global.csv')
country_df = pd.read_csv('https://raw.githubusercontent.com/CSSEGISandData/COVID-19/web-data/data/cases_country.csv')
```
# Death Cases
```
death_df.head()
```
# Confirmed Cases
```
confirmed_df.head()
```
# Recovered Cases
```
recovered_df.head()
```
# Cases According To The Countries
```
country_df.head()
#Manipulating Data
country_df.columns = map(str.lower, country_df.columns)
confirmed_df.columns = map(str.lower, confirmed_df.columns)
death_df.columns = map(str.lower, death_df.columns)
recovered_df.columns = map(str.lower, recovered_df.columns)
confirmed_df = confirmed_df.rename(columns={'province/state': 'state', 'country/region': 'country'})
recovered_df = recovered_df.rename(columns={'province/state': 'state', 'country/region': 'country'})
death_df = death_df.rename(columns={'province/state': 'state', 'country/region': 'country'})
country_df = country_df.rename(columns={'country_region': 'country'})
sorted_country_df = country_df.sort_values("confirmed",ascending=False).head(20)
```
# Countries with Highest Rates
```
def highlight_col(x):
a = 'background-color:#abbaab'
b = 'background-color:#e52d27'
c = 'background-color:#667db6'
d = 'background-color:#52c234'
e = 'background-color:#11998e'
temp_df = pd.DataFrame('',index=x.index,columns=x.columns)
temp_df.iloc[:,0]=a
temp_df.iloc[:,4]=c
temp_df.iloc[:,5]=b
temp_df.iloc[:,6]=d
temp_df.iloc[:,7]=e
return temp_df
sorted_country_df.style.apply(highlight_col ,axis=None)
fig = px.scatter(sorted_country_df.head(25),x="country" ,y="confirmed" ,size="confirmed" ,color="country" ,hover_name="country" ,size_max=70)
```
# Graphical Format
```
fig.show()
def plot_cases_for_country(country):
labels = ['confirmed', 'deaths']
colors = ['#667db6', '#e52d27']
mode_size = [6,8]
line_size = [4,5]
df_list = [confirmed_df, death_df]
fig = go.Figure()
for i, df in enumerate(df_list):
if country == 'World' or country == 'world':
x_data = np.array(list(df.iloc[:, 5:].columns))
y_data = np.sum(np.asarray(df.iloc[:, 5:]), axis=0)
else:
x_data = np.array(list(df.iloc[:, 5:].columns))
y_data = np.sum(np.asarray(df[df['country']==country].iloc[:, 5:]),axis=0)
fig.add_trace(go.Scatter(x=x_data, y=y_data, mode="lines+markers",name=labels[i],
line=dict(color=colors[i],width=line_size[i]),
connectgaps = True ,text='Total'+str(labels[i]) + ": "+ str(y_data[-1])
))
fig.show()
```
# Enter Country to View the Stats
```
interact(plot_cases_for_country, country="World");
```
# World Map
```
world_map = folium.Map(location=[11,0], tiles="cartodbpositron", zoom_start=2, max_zoom=5, min_zoom=2)
for i in range(len(confirmed_df)):
folium.Circle(
location=[confirmed_df.iloc[i]['lat'], confirmed_df.iloc[i]['long']],
fill = True,
radius = (int((np.log(confirmed_df.iloc[i,-1]+1.001))) + 0.7)*50000,
fill_color= 'blue',
color = 'red',
tooltip = "<div style='margin: 0; background-color: #2193b0; color: #200122;'>"+
"<h4 style='text-align:center;font-weight: bold'>"+confirmed_df.iloc[i]['country'] + "</h4>"
"<hr style='margin:10px;color: white;'>"+
"<ul style='color: white;;list-style-type:circle;align-item:left;padding-left:20px;padding-right:20px'>"+
"<li>Confirmed: "+str(confirmed_df.iloc[i,-1])+"</li>"+
"<li>Deaths: "+str(death_df.iloc[i,-1])+"</li>"+
"<li>Death Rate: "+ str(np.round(death_df.iloc[i,-1]/(confirmed_df.iloc[i,-1]+1.00001)*100,2))+ "</li>"+
"</ul></div>",
).add_to(world_map)
world_map
```
# Top 10 Countries with Affected Cases
```
px.bar(
sorted_country_df.head(10),
x = "country",
y = "deaths",
title= "Affected Cases",
color_discrete_sequence=["#fc466b"],
height=750,
width=1000
)
```
## Top 10 Countries with Recovered Cases
```
px.bar(
sorted_country_df.head(10),
x = "country",
y = "recovered",
title= "Recovered Cases",
color_discrete_sequence=["#56ab2f"],
height=750,
width=1000
)
```
## Top 10 Countries with Confirmed Cases
```
px.bar(
sorted_country_df.head(10),
x = "country",
y = "confirmed",
title= "Confirmed Cases",
color_discrete_sequence=["#8e2de2"],
height=750,
width=1000
)
```
## Resources
- https://en.wikipedia.org/wiki/2019%E2%80%9320_coronavirus_pandemic
- https://www.who.int/emergencies/diseases/novel-coronavirus-2019
- https://www.worldometers.info/coronavirus/
- https://github.com/CSSEGISandData/COVID-19
| github_jupyter |
#EOSC 582 Assignment V (SSMI)
```
__author__ = 'Yingkai (Kyle) Sha'
__email__ = 'yingkai@eos.ubc.ca'
import h5py
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.basemap import Basemap
% matplotlib inline
import warnings
warnings.filterwarnings('ignore')
```
Function for histogram.
```
def hist_SSMI(CWV_unfixed, CWV_both, CWV_19, CWL_unfixed, CWL_both, CWL_19):
CWV_unfixed = CWV_unfixed.flatten(); CWV_both = CWV_both.flatten(); CWV_19 = CWV_19.flatten()
CWL_unfixed = CWL_unfixed.flatten(); CWL_both = CWL_both.flatten(); CWL_19 = CWL_19.flatten()
binCWV = np.arange(0, 70+1, 1); binCWL = np.arange(0, 0.7+0.01, 0.01)
fig = plt.figure(figsize=(16, 4))
ax1=plt.subplot2grid((1, 2), (0, 0), colspan=1, rowspan=1)
ax2=plt.subplot2grid((1, 2), (0, 1), colspan=1, rowspan=1)
ax1.hist(CWV_unfixed[~np.isnan(CWV_unfixed)], binCWV, color='y', linewidth=2.5, histtype='step', label='unfixed');
ax1.hist(CWV_both[~np.isnan(CWV_both)], binCWV, color='b', linewidth=2.5, histtype='step', label='fixed: both 19, 37 GHz');
ax1.hist(CWV_19[~np.isnan(CWV_19)], binCWV, color='r', linewidth=2.5, histtype='step', label='fixed: 19 GHz only');
ax1.legend(); ax1.grid(); ax1.set_xlabel('CWV', fontsize=12)
ax1.set_title('(a) unfixed v.s. fixed CWV Histogram', fontsize=12, fontweight='bold')
ax2.hist(CWL_unfixed[~np.isnan(CWL_unfixed)], binCWL, color='y', linewidth=2.5, histtype='step', label='unfixed');
ax2.hist(CWL_both[~np.isnan(CWL_both)], binCWL, color='b', linewidth=2.5, histtype='step', label='fixed: both 19, 37 GHz');
ax2.hist(CWL_19[~np.isnan(CWL_19)], binCWL, color='r', linewidth=2.5, histtype='step', label='fixed: 19 GHz only');
ax2.legend(); ax2.grid(); ax2.set_xlabel('CWL', fontsize=12)
ax2.set_title('(b) unfixed v.s. fixed CWL Histogram', fontsize=12, fontweight='bold')
```
Function for maps
```
def single_map(ax):
proj = Basemap(projection='moll', lon_0=180, resolution='c', ax=ax)
proj.drawcoastlines()
proj.drawmeridians(np.arange(0, 360, 60));
proj.drawparallels(np.arange(-90, 90, 30));
return proj
def SSMI_map(lon, lat, CWV_unfixed, CWV_both, CWV_19, CWL_unfixed, CWL_both, CWL_19):
levCWV = np.arange(0, 80+5, 5); levCWL = np.arange(0, 0.7+0.07, 0.07)
fig = plt.figure(figsize=(12, 8))
ax1=plt.subplot2grid((3, 2), (0, 0), colspan=1, rowspan=1); ax2=plt.subplot2grid((3, 2), (1, 0), colspan=1, rowspan=1)
ax3=plt.subplot2grid((3, 2), (2, 0), colspan=1, rowspan=1); ax4=plt.subplot2grid((3, 2), (0, 1), colspan=1, rowspan=1)
ax5=plt.subplot2grid((3, 2), (1, 1), colspan=1, rowspan=1); ax6=plt.subplot2grid((3, 2), (2, 1), colspan=1, rowspan=1)
proj=single_map(ax1); x, y = proj(lon, lat)
CS = proj.contourf(x, y, CWV_unfixed, levCWV, cmap=plt.cm.RdYlGn, extend='max')
ax1.set_title('(a.1) CWV unfixed (Jan 1990)', fontsize=12, fontweight='bold', y = 1.025)
proj=single_map(ax2); x, y = proj(lon, lat)
CS = proj.contourf(x, y, CWV_both, levCWV, cmap=plt.cm.RdYlGn, extend='max')
ax2.set_title('(a.2) CWV fixed: both (Jan 1990)', fontsize=12, fontweight='bold', y = 1.025)
proj=single_map(ax3); x, y = proj(lon, lat)
CS = proj.contourf(x, y, CWV_19, levCWV, cmap=plt.cm.RdYlGn, extend='max')
ax3.set_title('(a.3) CWV fixed: 19 GHz only (Jan 1990)', fontsize=12, fontweight='bold', y = 1.025)
cax = fig.add_axes([0.175, 0.05, 0.25, 0.02])
CBar = fig.colorbar(CS, cax=cax, orientation='horizontal')
CBar.ax.tick_params(axis='x', length=12.5)
CBar.set_label('CWV $\mathrm{kg/m^2}$', fontsize=12)
proj=single_map(ax4); x, y = proj(lon, lat)
CS = proj.contourf(x, y, CWL_unfixed, levCWL, cmap=plt.cm.gist_ncar_r, extend='max')
ax4.set_title('(b.1) CWL unfixed (Jan 1990)', fontsize=12, fontweight='bold', y = 1.025)
proj=single_map(ax5); x, y = proj(lon, lat)
CS = proj.contourf(x, y, CWL_both, levCWL, cmap=plt.cm.gist_ncar_r, extend='max')
ax5.set_title('(b.2) CWL fixed: both (Jan 1990)', fontsize=12, fontweight='bold', y = 1.025)
proj=single_map(ax6); x, y = proj(lon, lat)
CS = proj.contourf(x, y, CWL_19, levCWL, cmap=plt.cm.gist_ncar_r, extend='max')
ax6.set_title('(b.3) CWL fixed: 19 GHz only (Jan 1990)', fontsize=12, fontweight='bold', y = 1.025)
cax = fig.add_axes([0.6, 0.05, 0.25, 0.02])
CBar = fig.colorbar(CS, cax=cax, orientation='horizontal')
CBar.ax.tick_params(axis='x', length=12.5)
CBar.set_label('CWL', fontsize=12)
```
# Retrieval functions
SSMI.py including functions calculates emissivity and absorption coefficient at at 19 and 37 GHz SSMI channel.
Code are Python version of <a href='http://www.aos.wisc.edu/~tristan/aos740.php'>**UW-Madison AOS-704**</a> 's FORTRAN77 code.
```
import site
site.addsitedir('_libs')
from SSMI import *
```
Approximation of windspeed and main retrieval function in Greenwald et al., 1993.
```
# windspeed
def wind_speed(sst, t19v, t22v, t37h, t37v):
"""
input: sst (K), t19v (K), t22v (K), t37h (K)
output: windspeed (m/s)
"""
speed=1.0969*(t19v)-0.4555e0*(t22v)- 1.76*(t37v)+0.786*(t37h)+ 147.9
return speed
# retrival, based on EOSC 582 Website
def SSMI_retrieval(SST, theta, T19H, T19V, T22V, T37H, T37V, iter_num=5, correction='both'):
'''
Using 4 SSMI channel brightness temperature retrive total precipitable water and liquid water path
=========================================================================
CMV, CWL = SSMI_retrieval(SST, theta, T19H, T19V, T22V, T37H, T37V, iter_num=5)
-------------------------------------------------------------------------
Input:
SST: Sea Surface Temperature (K)
theta: Incidence angle
T#H: Brightness temperature in #GHz band with horizontal polarization
T#V: Brightness temperature in #GHz band with vertical polarization
iter_num: = 0 means no correction, > 0 applies correction to CWV > 25kg/m^2
Output:
CWV: Total precipitable water
CWL: Liquid water path
==========================================================================
Author:
Yingkai (Kyle) Sha
yingkai@eos.ubc.ca
'''
M, N = np.shape(SST)
# Parameters
mu = np.cos(theta*np.pi/180.) # Incidence angle
GAMMA = -5.8E-3 # Lapse rate: -5.8 K/km = -5.8E-3 K/m
Hw = 2.2E3 # Water vapor scaling height: 2.2km
# Correction for cold bias
# (Greenwald et al., 1993)
T37H= T37H + 3.58
T37V= T37V + 3.58
# delta T
dT19 = T19H - T19V
dT37 = T37H - T37V
# Frequency bands (GHz)
freq = [19, 22, 37, 85]
# Allocate memorise
emissv = np.empty([len(freq), M, N])
emissh = np.empty([len(freq), M, N])
KL19 = np.empty([M, N])
KL37 = np.empty([M, N])
KV19 = np.empty([M, N])
KV37 = np.empty([M, N])
TOX19 = np.empty([M, N])
TOX37 = np.empty([M, N])
# Emperical windspeed
windspeed = wind_speed(SST, T19V, T22V, T37H-3.58, T37V-3.58)
# Calculate emission, absorbtion coef.
for m in range(M):
for n in range(N):
for i in range(len(freq)):
emissv[i, m, n], emissh[i, m, n] = emiss(i+1, windspeed[m, n], SST[m, n], theta)
KL19[m, n], KL37[m, n], KV19[m, n], KV37[m, n], TOX19[m, n], TOX37[m, n] = coef(SST[m, n])
# Retrieve function
R37V=(1.0 - emissv[2, :, :])
R19V=(1.0 - emissv[0, :, :])
R37H=(1.0 - emissh[2, :, :])
R19H=(1.0 - emissh[0, :, :])
# Iteration correction of F19, F37 for CWV > 25kg/m^2
# Greenwald et al., 1993) equation (4)
CWV = np.zeros(SST.shape)
#CWL = np.zeros(SST.shape)
T019 = SST
T037 = SST
for iteration in range(iter_num):
hit = CWV > 25
# transmission
Tau19V = np.exp(-1*KV19*CWV/mu)
Tau37V = np.exp(-1*KV37*CWV/mu)
f19 = np.exp(50*KV19/mu)
f37 = np.exp(50*KV37/mu)
if iteration > 0:
# in the first timestep, T019, T037 = SST
T019[hit] = SST[hit] + GAMMA*Hw*(1-f19[hit]*Tau19V[hit]**2)*TOX19[hit]
if correction == 'both':
T037[hit] = SST[hit] + GAMMA*Hw*(1-f37[hit]*Tau37V[hit]**2)*TOX37[hit]
#T037[hit] = SST[hit]
# Correction
F19 = (T19H - T019)/(T19V - T019)
F37 = (T37H - T037)/(T37V - T037)
R1 = -1*mu/2.*np.log(dT19/(SST*R19V*(1-F19)*TOX19**2.))
R2 = -1*mu/2.*np.log(dT37/(SST*R37V*(1-F37)*TOX37**2.))
# Linear algebra
M = KV19*KL37 - KL19*KV37
CWV = (R1*KL37 - R2*KL19)/M
#print('iteration step = {}'.format(iteration))
# get CWL
CWL = (R2*KV19 - R1*KV37)/M
return CWV, CWL
theta = 53.1
# boardcasting because my retrival function supports 2D array
SST = 271.75*np.ones([1, 1])
T19H = 113.57*np.ones([1, 1])
T19V = 183.24*np.ones([1, 1])
T22V = 194.80*np.ones([1, 1])
T37H = 148.13*np.ones([1, 1])
T37V = 208.11*np.ones([1, 1])
SSMI_retrieval(SST, theta, T19H, T19V, T22V, T37H, T37V, iter_num=4, correction='both')
```
# Full Retrival
## Jan
```
TB_obj = h5py.File('_data/bright_temps.h5', 'r')
lat = TB_obj['lat'][:]
lon = TB_obj['lon'][:]
SST = TB_obj['jan/sst'][:]
T19H = TB_obj['jan/t19h'][:]
T19V = TB_obj['jan/t19v'][:]
T22V = TB_obj['jan/t22v'][:]
T37H = TB_obj['jan/t37h'][:]
T37V = TB_obj['jan/t37v'][:]
TB_obj.close()
theta = 53.1
CWV1_unfixed, CWL1_unfixed = SSMI_retrieval(SST, theta, T19H, T19V, T22V, T37H, T37V, iter_num=1)
CWV1_both, CWL1_both = SSMI_retrieval(SST, theta, T19H, T19V, T22V, T37H, T37V, iter_num=5, correction='both')
CWV1_19, CWL1_19 = SSMI_retrieval(SST, theta, T19H, T19V, T22V, T37H, T37V, iter_num=5, correction='19')
hist_SSMI(CWV1_unfixed, CWV1_both, CWV1_19, CWL1_unfixed, CWL1_both, CWL1_19)
SSMI_map(lon, lat, CWV1_unfixed, CWV1_both, CWV1_19, CWL1_unfixed, CWL1_both, CWL1_19)
```
## Jul
```
TB_obj = h5py.File('_data/bright_temps.h5', 'r')
SST = TB_obj['july/sst'][:]
T19H = TB_obj['july/t19h'][:]
T19V = TB_obj['july/t19v'][:]
T22V = TB_obj['july/t22v'][:]
T37H = TB_obj['july/t37h'][:]
T37V = TB_obj['july/t37v'][:]
TB_obj.close()
CWV7_unfixed, CWL7_unfixed = SSMI_retrieval(SST, theta, T19H, T19V, T22V, T37H, T37V, iter_num=1)
CWV7_both, CWL7_both = SSMI_retrieval(SST, theta, T19H, T19V, T22V, T37H, T37V, iter_num=5, correction='both')
CWV7_19, CWL7_19 = SSMI_retrieval(SST, theta, T19H, T19V, T22V, T37H, T37V, iter_num=5, correction='19')
hist_SSMI(CWV7_unfixed, CWV7_both, CWV7_19, CWL7_unfixed, CWL7_both, CWL7_19)
SSMI_map(lon, lat, CWV7_unfixed, CWV7_both, CWV7_19, CWL7_unfixed, CWL7_both, CWL7_19)
```
## Zonal mean results
```
CWV1z_19 = np.nanmean(CWV1_19, 1); CWL1z_19 = np.nanmean(CWL1_19, 1)
CWV7z_19 = np.nanmean(CWV7_19, 1); CWL7z_19 = np.nanmean(CWL7_19, 1)
CWV1z_both = np.nanmean(CWV1_both, 1); CWL1z_both = np.nanmean(CWL1_both, 1)
CWV7z_both = np.nanmean(CWV7_both, 1); CWL7z_both = np.nanmean(CWL7_both, 1)
CWV1z_unfixed = np.nanmean(CWV1_unfixed, 1); CWL1z_unfixed = np.nanmean(CWL1_unfixed, 1)
CWV7z_unfixed = np.nanmean(CWV7_unfixed, 1); CWL7z_unfixed = np.nanmean(CWL7_unfixed, 1)
fig = plt.figure(figsize=(14, 12))
ax1=plt.subplot2grid((2, 2), (0, 0), colspan=1, rowspan=1)
ax2=plt.subplot2grid((2, 2), (0, 1), colspan=1, rowspan=1)
ax3=plt.subplot2grid((2, 2), (1, 0), colspan=1, rowspan=1)
ax4=plt.subplot2grid((2, 2), (1, 1), colspan=1, rowspan=1)
ax1.plot(lat[:, 0], CWV1z_unfixed, color=[0, 0.2, 0.4], linewidth=3.5, label='Jan unfixed');
ax1.plot(lat[:, 0], CWV1z_both, color=[0, 0.5, 0.7], linewidth=3.5, label='Jan fixed: both');
ax1.plot(lat[:, 0], CWV1z_19, color=[0, 0.8, 1], linewidth=3.5, label='Jan fixed: 19 GHz only');
ax1.grid(); ax1.legend(loc=4); ax1.set_xlim(-90, 90);
ax1.set_title('(a) Zonal mean CWV | Jan', fontsize=12, fontweight='bold')
ax2.plot(lat[:, 0], CWV7z_unfixed, color=[0.4, 0.2, 0], linewidth=3.5, label='Jul unfixed');
ax2.plot(lat[:, 0], CWV7z_both, color=[0.7, 0.5, 0], linewidth=3.5, label='Jul fixed: both');
ax2.plot(lat[:, 0], CWV7z_19, color=[1, 0.8, 0], linewidth=3.5, label='Jul fixed: 19 GHz only');
ax2.grid(); ax2.legend(loc=4); ax2.set_xlim(-90, 90);
ax2.set_title('(b) Zonal mean CWV | Jul', fontsize=12, fontweight='bold')
ax3.plot(lat[:, 0], CWL1z_unfixed, color=[0, 0.2, 0.4], linewidth=3.5, label='Jan unfixed');
ax3.plot(lat[:, 0], CWL1z_both, color=[0, 0.5, 0.7], linewidth=3.5, label='Jan fixed: both');
ax3.plot(lat[:, 0], CWL1z_19, color=[0, 0.8, 1], linewidth=3.5, label='Jan fixed: 19 GHz only');
ax3.grid(); ax3.legend(loc=4); ax3.set_xlim(-90, 90);
ax3.set_title('(c) Zonal mean CWL | Jan', fontsize=12, fontweight='bold')
ax4.plot(lat[:, 0], CWL7z_unfixed, color=[0.4, 0.2, 0], linewidth=3.5, label='Jul unfixed');
ax4.plot(lat[:, 0], CWL7z_both, color=[0.7, 0.5, 0], linewidth=3.5, label='Jul fixed: both');
ax4.plot(lat[:, 0], CWL7z_19, color=[1, 0.8, 0], linewidth=3.5, label='Jul fixed: 19 GHz only');
ax4.grid(); ax4.legend(loc=4); ax4.set_xlim(-90, 90);
ax4.set_title('(d) Zonal mean CWL | Jul', fontsize=12, fontweight='bold')
```
| github_jupyter |
# Discretization
---
In this notebook, you will deal with continuous state and action spaces by discretizing them. This will enable you to apply reinforcement learning algorithms that are only designed to work with discrete spaces.
### 1. Import the Necessary Packages
```
import sys
import gym
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# Set plotting options
%matplotlib inline
plt.style.use('ggplot')
np.set_printoptions(precision=3, linewidth=120)
```
### 2. Specify the Environment, and Explore the State and Action Spaces
We'll use [OpenAI Gym](https://gym.openai.com/) environments to test and develop our algorithms. These simulate a variety of classic as well as contemporary reinforcement learning tasks. Let's use an environment that has a continuous state space, but a discrete action space.
```
# Create an environment and set random seed
env = gym.make('MountainCar-v0')
env.seed(505);
```
Run the next code cell to watch a random agent.
```
state = env.reset()
score = 0
for t in range(200):
action = env.action_space.sample()
env.render()
state, reward, done, _ = env.step(action)
score += reward
if done:
break
print('Final score:', score)
env.close()
```
In this notebook, you will train an agent to perform much better! For now, we can explore the state and action spaces, as well as sample them.
```
# Explore state (observation) space
print("State space:", env.observation_space)
print("- low:", env.observation_space.low)
print("- high:", env.observation_space.high)
# Generate some samples from the state space
print("State space samples:")
print(np.array([env.observation_space.sample() for i in range(10)]))
# Explore the action space
print("Action space:", env.action_space)
# Generate some samples from the action space
print("Action space samples:")
print(np.array([env.action_space.sample() for i in range(10)]))
```
### 3. Discretize the State Space with a Uniform Grid
We will discretize the space using a uniformly-spaced grid. Implement the following function to create such a grid, given the lower bounds (`low`), upper bounds (`high`), and number of desired `bins` along each dimension. It should return the split points for each dimension, which will be 1 less than the number of bins.
For instance, if `low = [-1.0, -5.0]`, `high = [1.0, 5.0]`, and `bins = (10, 10)`, then your function should return the following list of 2 NumPy arrays:
```
[array([-0.8, -0.6, -0.4, -0.2, 0.0, 0.2, 0.4, 0.6, 0.8]),
array([-4.0, -3.0, -2.0, -1.0, 0.0, 1.0, 2.0, 3.0, 4.0])]
```
Note that the ends of `low` and `high` are **not** included in these split points. It is assumed that any value below the lowest split point maps to index `0` and any value above the highest split point maps to index `n-1`, where `n` is the number of bins along that dimension.
```
def create_uniform_grid(low, high, bins=(10, 10)):
"""Define a uniformly-spaced grid that can be used to discretize a space.
Parameters
----------
low : array_like
Lower bounds for each dimension of the continuous space.
high : array_like
Upper bounds for each dimension of the continuous space.
bins : tuple
Number of bins along each corresponding dimension.
Returns
-------
grid : list of array_like
A list of arrays containing split points for each dimension.
"""
# TODO: Implement this
grid = [np.linspace(low[dim], high[dim], bins[dim] + 1)[1:-1] for dim in range(len(bins))]
print("Uniform grid: [<low>, <high>] / <bins> => <splits>")
for l, h, b, splits in zip(low, high, bins, grid):
print(" [{}, {}] / {} => {}".format(l, h, b, splits))
return grid
low = [-1.0, -5.0]
high = [1.0, 5.0]
create_uniform_grid(low, high) # [test]
```
Now write a function that can convert samples from a continuous space into its equivalent discretized representation, given a grid like the one you created above. You can use the [`numpy.digitize()`](https://docs.scipy.org/doc/numpy-1.9.3/reference/generated/numpy.digitize.html) function for this purpose.
Assume the grid is a list of NumPy arrays containing the following split points:
```
[array([-0.8, -0.6, -0.4, -0.2, 0.0, 0.2, 0.4, 0.6, 0.8]),
array([-4.0, -3.0, -2.0, -1.0, 0.0, 1.0, 2.0, 3.0, 4.0])]
```
Here are some potential samples and their corresponding discretized representations:
```
[-1.0 , -5.0] => [0, 0]
[-0.81, -4.1] => [0, 0]
[-0.8 , -4.0] => [1, 1]
[-0.5 , 0.0] => [2, 5]
[ 0.2 , -1.9] => [6, 3]
[ 0.8 , 4.0] => [9, 9]
[ 0.81, 4.1] => [9, 9]
[ 1.0 , 5.0] => [9, 9]
```
**Note**: There may be one-off differences in binning due to floating-point inaccuracies when samples are close to grid boundaries, but that is alright.
```
def discretize(sample, grid):
"""Discretize a sample as per given grid.
Parameters
----------
sample : array_like
A single sample from the (original) continuous space.
grid : list of array_like
A list of arrays containing split points for each dimension.
Returns
-------
discretized_sample : array_like
A sequence of integers with the same number of dimensions as sample.
"""
# TODO: Implement this
return list(int(np.digitize(s, g)) for s, g in zip(sample, grid)) # apply along each dimension
# Test with a simple grid and some samples
grid = create_uniform_grid([-1.0, -5.0], [1.0, 5.0])
samples = np.array(
[[-1.0 , -5.0],
[-0.81, -4.1],
[-0.8 , -4.0],
[-0.5 , 0.0],
[ 0.2 , -1.9],
[ 0.8 , 4.0],
[ 0.81, 4.1],
[ 1.0 , 5.0]])
discretized_samples = np.array([discretize(sample, grid) for sample in samples])
print("\nSamples:", repr(samples), sep="\n")
print("\nDiscretized samples:", repr(discretized_samples), sep="\n")
```
### 4. Visualization
It might be helpful to visualize the original and discretized samples to get a sense of how much error you are introducing.
```
import matplotlib.collections as mc
def visualize_samples(samples, discretized_samples, grid, low=None, high=None):
"""Visualize original and discretized samples on a given 2-dimensional grid."""
fig, ax = plt.subplots(figsize=(10, 10))
# Show grid
ax.xaxis.set_major_locator(plt.FixedLocator(grid[0]))
ax.yaxis.set_major_locator(plt.FixedLocator(grid[1]))
ax.grid(True)
# If bounds (low, high) are specified, use them to set axis limits
if low is not None and high is not None:
ax.set_xlim(low[0], high[0])
ax.set_ylim(low[1], high[1])
else:
# Otherwise use first, last grid locations as low, high (for further mapping discretized samples)
low = [splits[0] for splits in grid]
high = [splits[-1] for splits in grid]
# Map each discretized sample (which is really an index) to the center of corresponding grid cell
grid_extended = np.hstack((np.array([low]).T, grid, np.array([high]).T)) # add low and high ends
grid_centers = (grid_extended[:, 1:] + grid_extended[:, :-1]) / 2 # compute center of each grid cell
locs = np.stack(grid_centers[i, discretized_samples[:, i]] for i in range(len(grid))).T # map discretized samples
ax.plot(samples[:, 0], samples[:, 1], 'o') # plot original samples
ax.plot(locs[:, 0], locs[:, 1], 's') # plot discretized samples in mapped locations
ax.add_collection(mc.LineCollection(list(zip(samples, locs)), colors='orange')) # add a line connecting each original-discretized sample
ax.legend(['original', 'discretized'])
visualize_samples(samples, discretized_samples, grid, low, high)
```
Now that we have a way to discretize a state space, let's apply it to our reinforcement learning environment.
```
# Create a grid to discretize the state space
state_grid = create_uniform_grid(env.observation_space.low, env.observation_space.high, bins=(10, 10))
state_grid
# Obtain some samples from the space, discretize them, and then visualize them
state_samples = np.array([env.observation_space.sample() for i in range(10)])
discretized_state_samples = np.array([discretize(sample, state_grid) for sample in state_samples])
visualize_samples(state_samples, discretized_state_samples, state_grid,
env.observation_space.low, env.observation_space.high)
plt.xlabel('position'); plt.ylabel('velocity'); # axis labels for MountainCar-v0 state space
```
You might notice that if you have enough bins, the discretization doesn't introduce too much error into your representation. So we may be able to now apply a reinforcement learning algorithm (like Q-Learning) that operates on discrete spaces. Give it a shot to see how well it works!
### 5. Q-Learning
Provided below is a simple Q-Learning agent. Implement the `preprocess_state()` method to convert each continuous state sample to its corresponding discretized representation.
```
class QLearningAgent:
"""Q-Learning agent that can act on a continuous state space by discretizing it."""
def __init__(self, env, state_grid, alpha=0.02, gamma=0.99,
epsilon=1.0, epsilon_decay_rate=0.9995, min_epsilon=.01, seed=505):
"""Initialize variables, create grid for discretization."""
# Environment info
self.env = env
self.state_grid = state_grid
self.state_size = tuple(len(splits) + 1 for splits in self.state_grid) # n-dimensional state space
self.action_size = self.env.action_space.n # 1-dimensional discrete action space
self.seed = np.random.seed(seed)
print("Environment:", self.env)
print("State space size:", self.state_size)
print("Action space size:", self.action_size)
# Learning parameters
self.alpha = alpha # learning rate
self.gamma = gamma # discount factor
self.epsilon = self.initial_epsilon = epsilon # initial exploration rate
self.epsilon_decay_rate = epsilon_decay_rate # how quickly should we decrease epsilon
self.min_epsilon = min_epsilon
# Create Q-table
self.q_table = np.zeros(shape=(self.state_size + (self.action_size,)))
print("Q table size:", self.q_table.shape)
def preprocess_state(self, state):
"""Map a continuous state to its discretized representation."""
# TODO: Implement this
return tuple(discretize(state, self.state_grid))
def reset_episode(self, state):
"""Reset variables for a new episode."""
# Gradually decrease exploration rate
self.epsilon *= self.epsilon_decay_rate
self.epsilon = max(self.epsilon, self.min_epsilon)
# Decide initial action
self.last_state = self.preprocess_state(state)
self.last_action = np.argmax(self.q_table[self.last_state])
return self.last_action
def reset_exploration(self, epsilon=None):
"""Reset exploration rate used when training."""
self.epsilon = epsilon if epsilon is not None else self.initial_epsilon
def act(self, state, reward=None, done=None, mode='train'):
"""Pick next action and update internal Q table (when mode != 'test')."""
state = self.preprocess_state(state)
if mode == 'test':
# Test mode: Simply produce an action
action = np.argmax(self.q_table[state])
else:
# Train mode (default): Update Q table, pick next action
# Note: We update the Q table entry for the *last* (state, action) pair with current state, reward
self.q_table[self.last_state + (self.last_action,)] += self.alpha * \
(reward + self.gamma * max(self.q_table[state]) - self.q_table[self.last_state + (self.last_action,)])
# Exploration vs. exploitation
do_exploration = np.random.uniform(0, 1) < self.epsilon
if do_exploration:
# Pick a random action
action = np.random.randint(0, self.action_size)
else:
# Pick the best action from Q table
action = np.argmax(self.q_table[state])
# Roll over current state, action for next step
self.last_state = state
self.last_action = action
return action
q_agent = QLearningAgent(env, state_grid)
```
Let's also define a convenience function to run an agent on a given environment. When calling this function, you can pass in `mode='test'` to tell the agent not to learn.
```
def run(agent, env, num_episodes=20000, mode='train'):
"""Run agent in given reinforcement learning environment and return scores."""
scores = []
max_avg_score = -np.inf
for i_episode in range(1, num_episodes+1):
# Initialize episode
state = env.reset()
action = agent.reset_episode(state)
total_reward = 0
done = False
# Roll out steps until done
while not done:
state, reward, done, info = env.step(action)
total_reward += reward
action = agent.act(state, reward, done, mode)
# Save final score
scores.append(total_reward)
# Print episode stats
if mode == 'train':
if len(scores) > 100:
avg_score = np.mean(scores[-100:])
if avg_score > max_avg_score:
max_avg_score = avg_score
if i_episode % 100 == 0:
print("\rEpisode {}/{} | Max Average Score: {}".format(i_episode, num_episodes, max_avg_score), end="")
sys.stdout.flush()
return scores
scores = run(q_agent, env)
```
The best way to analyze if your agent was learning the task is to plot the scores. It should generally increase as the agent goes through more episodes.
```
# Plot scores obtained per episode
plt.plot(scores); plt.title("Scores");
```
If the scores are noisy, it might be difficult to tell whether your agent is actually learning. To find the underlying trend, you may want to plot a rolling mean of the scores. Let's write a convenience function to plot both raw scores as well as a rolling mean.
```
def plot_scores(scores, rolling_window=100):
"""Plot scores and optional rolling mean using specified window."""
plt.plot(scores); plt.title("Scores");
rolling_mean = pd.Series(scores).rolling(rolling_window).mean()
plt.plot(rolling_mean);
return rolling_mean
rolling_mean = plot_scores(scores)
```
You should observe the mean episode scores go up over time. Next, you can freeze learning and run the agent in test mode to see how well it performs.
```
# Run in test mode and analyze scores obtained
test_scores = run(q_agent, env, num_episodes=100, mode='test')
print("[TEST] Completed {} episodes with avg. score = {}".format(len(test_scores), np.mean(test_scores)))
_ = plot_scores(test_scores)
```
It's also interesting to look at the final Q-table that is learned by the agent. Note that the Q-table is of size MxNxA, where (M, N) is the size of the state space, and A is the size of the action space. We are interested in the maximum Q-value for each state, and the corresponding (best) action associated with that value.
```
def plot_q_table(q_table):
"""Visualize max Q-value for each state and corresponding action."""
q_image = np.max(q_table, axis=2) # max Q-value for each state
q_actions = np.argmax(q_table, axis=2) # best action for each state
fig, ax = plt.subplots(figsize=(10, 10))
cax = ax.imshow(q_image, cmap='jet');
cbar = fig.colorbar(cax)
for x in range(q_image.shape[0]):
for y in range(q_image.shape[1]):
ax.text(x, y, q_actions[x, y], color='white',
horizontalalignment='center', verticalalignment='center')
ax.grid(False)
ax.set_title("Q-table, size: {}".format(q_table.shape))
ax.set_xlabel('position')
ax.set_ylabel('velocity')
plot_q_table(q_agent.q_table)
```
### 6. Modify the Grid
Now it's your turn to play with the grid definition and see what gives you optimal results. Your agent's final performance is likely to get better if you use a finer grid, with more bins per dimension, at the cost of higher model complexity (more parameters to learn).
```
# TODO: Create a new agent with a different state space grid
state_grid_new = create_uniform_grid(env.observation_space.low, env.observation_space.high, bins=(20, 20))
q_agent_new = QLearningAgent(env, state_grid_new)
q_agent_new.scores = [] # initialize a list to store scores for this agent
# Train it over a desired number of episodes and analyze scores
# Note: This cell can be run multiple times, and scores will get accumulated
q_agent_new.scores += run(q_agent_new, env, num_episodes=50000) # accumulate scores
rolling_mean_new = plot_scores(q_agent_new.scores)
# Run in test mode and analyze scores obtained
test_scores = run(q_agent_new, env, num_episodes=100, mode='test')
print("[TEST] Completed {} episodes with avg. score = {}".format(len(test_scores), np.mean(test_scores)))
_ = plot_scores(test_scores)
# Visualize the learned Q-table
plot_q_table(q_agent_new.q_table)
```
### 7. Watch a Smart Agent
```
state = env.reset()
score = 0
for t in range(200):
action = q_agent_new.act(state, mode='test')
env.render()
state, reward, done, _ = env.step(action)
score += reward
if done:
break
print('Final score:', score)
env.close()
```
| github_jupyter |
Author: Michael Gygli ([Github](https://github.com/gyglim), [Twitter](https://twitter.com/GygliMichael)), 2016-01-13
# Introduction #
This example demonstrates how to compute *C3D convolutional features* using Lasagne and Theano. C3D can be used as a general video feature and has shown strong performance. You can find more information in the paper [1] or the caffe-based reference implementation [2].
* [1]: D. Tran, L. Bourdev, R. Fergus, L. Torresani, and M. Paluri, Learning Spatiotemporal Features with 3D Convolutional Networks, ICCV 2015, http://vlg.cs.dartmouth.edu/c3d/c3d_video.pdf
* [2]: http://vlg.cs.dartmouth.edu/c3d/
# Preparation steps #
This demo uses the pretrained C3D weights, as well as the c3d module in the Lasagne Recipes modelzoo. Thus, you will need to get the Recipes from github (https://github.com/Lasagne/Recipes) first.
```
# Import models and set path
import sys
model_dir='../modelzoo/' # Path to your recipes/modelzoo
sys.path.insert(0,model_dir)
import c3d
import lasagne
import theano
# Download the weights and mean of the model
!wget -N https://s3.amazonaws.com/lasagne/recipes/pretrained/c3d/c3d_model.pkl
!wget -N https://s3.amazonaws.com/lasagne/recipes/pretrained/c3d/snipplet_mean.npy
# And the classes of Sports1m
!wget -N https://s3.amazonaws.com/lasagne/recipes/pretrained/c3d/labels.txt
# Finally, an example sniplet
!wget -N https://s3.amazonaws.com/lasagne/recipes/pretrained/c3d/example_snip.npy
# Build model
net = c3d.build_model()
# Set the weights (takes some time)
c3d.set_weights(net['prob'],'c3d_model.pkl')
# Load the video snipplet and show an example frame
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
snip=np.load('example_snip.npy')
plt.imshow(snip[0,:,:,:])
# Convert the video snipplet to the right format
# i.e. (nr in batch, channel, frameNr, y, x) and substract mean
caffe_snip=c3d.get_snips(snip,image_mean=np.load('snipplet_mean.npy'),start=0, with_mirrored=False)
# Compile prediction function
prediction = lasagne.layers.get_output(net['prob'], deterministic=True)
pred_fn = theano.function([net['input'].input_var], prediction, allow_input_downcast = True);
# Now we can get a prediction
probabilities=pred_fn(caffe_snip).mean(axis=0) # As we average over flipped and non-flipped
# Load labels
with open('labels.txt','r') as f:
class2label=dict(enumerate([name.rstrip('\n') for name in f]))
# Show the post probable ones
print('Top 10 class probabilities:')
for class_id in (-probabilities).argsort()[0:10]:
print('%20s: %.2f%%' % (class2label[class_id],100*probabilities[class_id]))
```
#### Comparison to C3D reference implementation ####
For this example, the Top 10 probabilities of the original C3D implementation are:
wiffle ball: 29.91%
knife throwing: 13.11%
croquet: 11.27%
disc golf: 5.29%
kickball: 5.18%
rounders: 4.48%
bocce: 3.53%
dodgeball: 2.27%
boomerang: 1.71%
tee ball: 1.39%
| github_jupyter |
# Plotting Target Pixel Files with Lightkurve
## Learning Goals
By the end of this tutorial, you will:
- Learn how to download and plot target pixel files from the data archive using [Lightkurve](https://docs.lightkurve.org).
- Be able to plot the target pixel file background.
- Be able to extract and plot flux from a target pixel file.
## Introduction
The [*Kepler*](https://www.nasa.gov/mission_pages/kepler/main/index.html), [*K2*](https://www.nasa.gov/mission_pages/kepler/main/index.html), and [*TESS*](https://tess.mit.edu/) telescopes observe stars for long periods of time, from just under a month to four years. By doing so they observe how the brightnesses of stars change over time.
Pixels around targeted stars are cut out and stored as *target pixel files* at each observing cadence. In this tutorial, we will learn how to use Lightkurve to download and understand the different photometric data stored in a target pixel file, and how to extract flux using basic aperture photometry.
It is useful to read the accompanying tutorial discussing how to use target pixel file products with Lightkurve before starting this tutorial. It is recommended that you also read the tutorial on using *Kepler* light curve products with Lightkurve, which will introduce you to some specifics on how *Kepler*, *K2*, and *TESS* make observations, and how these are displayed as light curves. It also introduces some important terms and concepts that are referred to in this tutorial.
*Kepler* observed a single field in the sky, although not all stars in this field were recorded. Instead, pixels were selected around certain targeted stars. These cutout images are called target pixel files, or TPFs. By combining the amount of flux in the pixels where the star appears, you can make a measurement of the amount of light from a star in that observation. The pixels chosen to include in this measurement are referred to as an *aperture*.
TPFs are typically the first port of call when studying a star with *Kepler*, *K2*, or *TESS*. They allow us to see where our data is coming from, and identify potential sources of noise or systematic trends. In this tutorial, we will use the *Kepler* mission as the main example, but these tools equally apply to *TESS* and *K2* as well.
## Imports
This tutorial requires:
- **[Lightkurve](https://docs.lightkurve.org)** to work with TPF files.
- [**Matplotlib**](https://matplotlib.org/) for plotting.
```
import lightkurve as lk
import matplotlib.pyplot as plt
%matplotlib inline
```
## 1. Downloading a TPF
A TPF contains the original imaging data from which a light curve is derived. Besides the brightness data measured by the charge-coupled device (CCD) camera, a TPF also includes post-processing information such as an estimate of the astronomical background, and a recommended pixel aperture for extracting a light curve.
First, we download a target pixel file. We will use one quarter's worth of *Kepler* data for the star named [Kepler-8](http://www.openexoplanetcatalogue.com/planet/Kepler-8%20b/), a star somewhat larger than the Sun, and the host of a [hot Jupiter planet](https://en.wikipedia.org/wiki/Hot_Jupiter).
```
search_result = lk.search_targetpixelfile("Kepler-8", mission="Kepler", quarter=4)
search_result
tpf = search_result.download()
```
This TPF contains data for every cadence in the quarter we downloaded. Let's focus on the first cadence for now, which we can select using zero-based indexing as follows:
```
first_cadence = tpf[0]
first_cadence
```
## 2. Flux and Background
At each cadence the TPF has a number of photometry data properties. These are:
- `flux_bkg`: the astronomical background of the image.
- `flux_bkg_err`: the statistical uncertainty on the background flux.
- `flux`: the stellar flux after the background is removed.
- `flux_err`: the statistical uncertainty on the stellar flux after background removal.
These properties can be accessed via a TPF object as follows:
```
first_cadence.flux.value
```
And you can plot the data as follows:
```
first_cadence.plot(column='flux');
```
Alternatively, if you are working directly with a FITS file, you can access the data in extension 1 (for example, `first_cadence.hdu[1].data['FLUX']`). Note that you can find all of the details on the structure and contents of TPF files in Section 2.3.2 of the [*Kepler* Archive Manual](http://archive.stsci.edu/files/live/sites/mast/files/home/missions-and-data/kepler/_documents/archive_manual.pdf).
When plotting data using the `plot()` function, what you are seeing in the TPF is the flux *after* the background has been removed. This background flux typically consists of [zodiacal light](https://en.wikipedia.org/wiki/Zodiacal_light) or earthshine (especially in *TESS* observations). The background is typically smooth and changes on scales much larger than a single TPF. In *Kepler*, the background is estimated for the CCD as a whole, before being extracted from each TPF in that CCD. You can learn more about background removal in Section 4.2 of the [*Kepler* Data Processing Handbook](http://archive.stsci.edu/files/live/sites/mast/files/home/missions-and-data/kepler/_documents/KSCI-19081-002-KDPH.pdf).
Now, let's compare the background to the background-subtracted flux to get a sense of scale. We can do this using the `plot()` function's `column` keyword. By default the function plots the flux, but we can change this to plot the background, as well as other data such as the error on each pixel's flux.
```
fig, axes = plt.subplots(2,2, figsize=(16,16))
first_cadence.plot(ax=axes[0,0], column='FLUX')
first_cadence.plot(ax=axes[0,1], column='FLUX_BKG')
first_cadence.plot(ax=axes[1,0], column='FLUX_ERR')
first_cadence.plot(ax=axes[1,1], column='FLUX_BKG_ERR');
```
From looking at the color scale on both plots, you may see that the background flux is very low compared to the total flux emitted by a star. This is expected — stars are bright! But these small background corrections become important when looking at the very small scale changes caused by planets or stellar oscillations. Understanding the background is an important part of astronomy with *Kepler*, *K2*, and *TESS*.
If the background is particularly bright and you want to see what the TPF looks like with it included, passing the `bkg=True` argument to the `plot()` method will show the TPF with the flux added on top of the background, representing the total flux recorded by the spacecraft.
```
first_cadence.plot(bkg=True);
```
In this case, the background is low and the star is bright, so it doesn't appear to make much of a difference.
## 3. Apertures
As part of the data processing done by the *Kepler* pipeline, each TPF includes a recommended *optimal aperture mask*. This aperture mask is optimized to ensure that the stellar signal has a high signal-to-noise ratio, with minimal contamination from the background.
The optimal aperture is stored in the TPF as the `pipeline_mask` property. We can have a look at it by calling it here:
```
first_cadence.pipeline_mask
```
As you can see, it is a Boolean array detailing which pixels are included. We can plot this aperture over the top of our TPF using the `plot()` function, and passing in the mask to the `aperture_mask` keyword. This will highlight the pixels included in the aperture mask using red hatched lines.
```
first_cadence.plot(aperture_mask=first_cadence.pipeline_mask);
```
You don't necessarily have to pass in the `pipeline_mask` to the `plot()` function; it can be any mask you create yourself, provided it is the right shape. An accompanying tutorial explains how to create such custom apertures, and goes into aperture photometry in more detail. For specifics on the selection of *Kepler*'s optimal apertures, read the [*Kepler* Data Processing Handbook](https://archive.stsci.edu/files/live/sites/mast/files/home/missions-and-data/kepler/_documents/KSCI-19081-002-KDPH.pdf), Section 7, *Finding Optimal Apertures in Kepler Data*.
## 4. Simple Aperture Photometry
Finally, let's learn how to perform simple aperture photometry (SAP) using the provided optimal aperture in `pipeline_mask` and the TPF.
Using the full TPF for all cadences in the quarter, we can perform aperture photometry using the `to_lightcurve()` method as follows:
```
lc = tpf.to_lightcurve()
```
This method returns a `LightCurve` object which details the flux and flux centroid position at each cadence:
```
lc
```
Note that this [`KeplerLightCurve`](https://docs.lightkurve.org/api/lightkurve.lightcurve.KeplerLightCurve.html) object has fewer data columns than in light curves downloaded directly from MAST. This is because we are extracting our light curve directly from the TPF using minimal processing, whereas light curves created using the official pipeline include more processing and more columns.
We can visualize the light curve as follows:
```
lc.plot();
```
This light curve is similar to the SAP light curve we previously encountered in the light curve tutorial.
### Note
The background flux can be plotted in a similar way, using the [`get_bkg_lightcurve()`](https://docs.lightkurve.org/api/lightkurve.targetpixelfile.KeplerTargetPixelFile.html#lightkurve.targetpixelfile.KeplerTargetPixelFile.get_bkg_lightcurve) method. This does not require an aperture, but instead sums the flux in the TPF's `FLUX_BKG` column at each timestamp.
```
bkg = tpf.get_bkg_lightcurve()
bkg.plot();
```
Inspecting the background in this way is useful to identify signals which appear to be present in the background rather than in the astronomical object under study.
---
## Exercises
Some stars, such as the planet-hosting star Kepler-10, have been observed both with *Kepler* and *TESS*. In this exercise, download and plot both the *TESS* and *Kepler* TPFs, along with the optimal apertures. You can do this by either selecting the TPFs from the list returned by [`search_targetpixelfile()`](https://docs.lightkurve.org/api/lightkurve.search.search_targetpixelfile.html), or by using the `mission` keyword argument when searching.
Both *Kepler* and *TESS* produce target pixel file data products, but these can look different across the two missions. *TESS* is focused on brighter stars and has larger pixels, so a star that might occupy many pixels in *Kepler* may only occupy a few in *TESS*.
How do light curves extracted from both of them compare?
```
#datalist = lk.search_targetpixelfile(...)
#soln:
datalist = lk.search_targetpixelfile("Kepler-10")
datalist
kep = datalist[6].download()
tes = datalist[15].download()
fig, axes = plt.subplots(1, 2, figsize=(14,6))
kep.plot(ax=axes[0], aperture_mask=kep.pipeline_mask, scale='log')
tes.plot(ax=axes[1], aperture_mask=tes.pipeline_mask)
fig.tight_layout();
lc_kep = kep.to_lightcurve()
lc_tes = tes.to_lightcurve()
fig, axes = plt.subplots(1, 2, figsize=(14,6), sharey=True)
lc_kep.flatten().plot(ax=axes[0], c='k', alpha=.8)
lc_tes.flatten().plot(ax=axes[1], c='k', alpha=.8);
```
If you plot the light curves for both missions side by side, you will see a stark difference. The *Kepler* data has a much smaller scatter, and repeating transits are visible. This is because *Kepler*'s pixels were smaller, and so could achieve a higher precision on fainter stars. *TESS* has larger pixels and therefore focuses on brighter stars. For stars like Kepler-10, it would be hard to detect a planet using *TESS* data alone.
## About this Notebook
**Authors:** Oliver Hall (oliver.hall@esa.int), Geert Barentsen
**Updated On**: 2020-09-15
## Citing Lightkurve and Astropy
If you use `lightkurve` or `astropy` for published research, please cite the authors. Click the buttons below to copy BibTeX entries to your clipboard.
lk.show_citation_instructions()
<img style="float: right;" src="https://raw.githubusercontent.com/spacetelescope/notebooks/master/assets/stsci_pri_combo_mark_horizonal_white_bkgd.png" alt="Space Telescope Logo" width="200px"/>
| github_jupyter |
## Imports
```
import jieba, json, math, os, re, sys, shutil, time
from datetime import datetime
import numpy as np
import tensorboardX
import torch, torchtext
import torch.nn as nn
import torch.nn.functional as F
```
## Functions
```
from fields import Field, Parms, Semantic, Vocab, _make_vocab
from utils import *
from nlp_db import nlp_db
from model_class import NLU_Classify
semantic = Semantic()
args = Parms()
vocab = Vocab(semantic)
args.manual_log = './manualog_transfromer1.log'
args.model_path = './model_stores/transformer_wiki1.pth'
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
# device = torch.device('cpu')
```
## Dialog Task
### Data Files
```
def pathFiles(rel_path):
return [
os.path.join(os.path.abspath(rel_path),
os.listdir(rel_path)[i])
for i in range(len(os.listdir(rel_path)))
]
# get train, test file names
rel_path = "../dialog_db/chinese_chatbot_corpus-master/clean_chat_corpus/"
cfiles = pathFiles(rel_path)
[print(file) for file in cfiles]
botTrainFile = cfiles[1]
botTrainFile
```
### Vocab Preprocess
```
args.vocab_path = os.path.abspath(os.path.join(rel_path, 'vocab.txt'))
def _make_chatbot_vocab(file, vocab_path, thres = 2):
word_dict = {}
with open(file, "r", encoding='utf-8') as f:
cnt = 0
for l in f.readlines():
for token in list(jieba.cut(l.strip().replace('\t',""))):
if token not in word_dict:
word_dict[token] = 0
else:
word_dict[token] += 1
if not os.path.isfile(vocab_path):
open(vocab_path,'a').close()
with open(vocab_path, 'w') as f:
for k, v in word_dict.items():
if v > thres:
print(k, file=f)
os.remove(args.vocab_path)
if not os.path.isfile(args.vocab_path):
_make_chatbot_vocab(botTrainFile, args.vocab_path)
try:
vocab.load(args.vocab_path)
except:
print("Vocab not loaded")
vocab.size, vocab.__getitem__('吃'), vocab.__getitem__(
'<pad>'), vocab.__getitem__('<unk>'), vocab.__getitem__('<sos>')
```
### Data Process => Model Parms Get
```
def get_max_sent_len(file):
with open(file, "r", encoding='utf-8') as f:
maxlen, sent_count = 0, 0
for l in f.readlines():
maxlen = max([maxlen, max([len(sent) for sent in l.split()])])
sent_count += 1
return maxlen, sent_count
args.max_sent_len, args.sent_count = get_max_sent_len(botTrainFile)
args.batch_size = 1000
dirrm(args)
```
### Sentence pad
### Define the Model
[ref: Seq2Seq - Transformer](https://pytorch.org/tutorials/beginner/transformer_tutorial.html#define-the-model)
```
# def _generate_square_subsequent_mask(sz):
# mask = (torch.triu(torch.ones(sz, sz)) == 1).transpose(0, 1)
# mask = mask.float().masked_fill(mask == 0, float('-inf')).masked_fill(mask == 1, float(0.0))
# return mask
# _generate_square_subsequent_mask(3)
class TransformerModel(nn.Module):
def __init__(self, ntoken, ninp, nhead, nhid, nlayers, dropout=0.5):
super(TransformerModel, self).__init__()
from torch.nn import TransformerEncoder, TransformerEncoderLayer
self.model_type = 'Transformer'
self.src_mask = None
self.pos_encoder = PositionalEncoding(ninp, dropout)
encoder_layers = TransformerEncoderLayer(ninp, nhead, nhid, dropout)
self.transformer_encoder = TransformerEncoder(encoder_layers, nlayers)
self.encoder = nn.Embedding(ntoken, ninp)
self.ninp = ninp
self.decoder = nn.Linear(ninp, ntoken)
self.init_weights()
def _generate_square_subsequent_mask(self, sz):
mask = (torch.triu(torch.ones(sz, sz)) == 1).transpose(0, 1)
mask = mask.float().masked_fill(mask == 0, float('-inf')).masked_fill(mask == 1, float(0.0))
return mask
def init_weights(self):
initrange = 0.1
self.encoder.weight.data.uniform_(-initrange, initrange)
self.decoder.bias.data.zero_()
self.decoder.weight.data.uniform_(-initrange, initrange)
def forward(self, src):
if self.src_mask is None or self.src_mask.size(0) != len(src):
device = src.device
mask = self._generate_square_subsequent_mask(len(src)).to(device)
self.src_mask = mask
src = self.encoder(src) * math.sqrt(self.ninp)
src = self.pos_encoder(src)
output = self.transformer_encoder(src, self.src_mask)
output = self.decoder(output)
return output
class PositionalEncoding(nn.Module):
def __init__(self, d_model, dropout=0.1, max_len=5000):
super(PositionalEncoding, self).__init__()
self.dropout = nn.Dropout(p=dropout)
pe = torch.zeros(max_len, d_model)
position = torch.arange(0, max_len, dtype=torch.float).unsqueeze(1)
div_term = torch.exp(torch.arange(0, d_model, 2).float() * (-math.log(10000.0) / d_model))
pe[:, 0::2] = torch.sin(position * div_term)
pe[:, 1::2] = torch.cos(position * div_term)
pe = pe.unsqueeze(0).transpose(0, 1)
self.register_buffer('pe', pe)
def forward(self, x):
x = x + self.pe[:x.size(0), :]
return self.dropout(x)
```
### batch data
```
from torchtext.data.utils import get_tokenizer
TEXT = torchtext.data.Field(tokenize=get_tokenizer("basic_english"),
init_token='<sos>',
eos_token='<eos>',
lower=True)
train_txt, val_txt, test_txt = torchtext.datasets.WikiText2.splits(TEXT)
TEXT.build_vocab(train_txt)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
def batchify(data, bsz):
data = TEXT.numericalize([data.examples[0].text])
# Divide the dataset into bsz parts.
nbatch = data.size(0) // bsz
# Trim off any extra elements that wouldn't cleanly fit (remainders).
data = data.narrow(0, 0, nbatch * bsz)
# Evenly divide the data across the bsz batches.
data = data.view(bsz, -1).t().contiguous()
return data.to(device)
batch_size = 20
eval_batch_size = 10
train_data = batchify(train_txt, batch_size)
val_data = batchify(val_txt, eval_batch_size)
test_data = batchify(test_txt, eval_batch_size)
bptt = 35
def get_batch(source, i):
seq_len = min(bptt, len(source) - 1 - i)
data = source[i:i+seq_len]
target = source[i+1:i+1+seq_len].view(-1)
return data, target
ntokens = len(TEXT.vocab.stoi) # the size of vocabulary
emsize = 200 # embedding dimension
nhid = 200 # the dimension of the feedforward network model in nn.TransformerEncoder
nlayers = 2 # the number of nn.TransformerEncoderLayer in nn.TransformerEncoder
nhead = 2 # the number of heads in the multiheadattention models
dropout = 0.2 # the dropout value
model = TransformerModel(ntokens, emsize, nhead, nhid, nlayers, dropout).to(device)
```
### train define
```
criterion = nn.CrossEntropyLoss()
lr = 5.0 # learning rate
optimizer = torch.optim.SGD(model.parameters(), lr=lr)
scheduler = torch.optim.lr_scheduler.StepLR(optimizer, 1.0, gamma=0.95)
import time
def train():
model.train() # Turn on the train mode
total_loss = 0.
start_time = time.time()
ntokens = len(TEXT.vocab.stoi)
for batch, i in enumerate(range(0, train_data.size(0) - 1, bptt)):
data, targets = get_batch(train_data, i)
optimizer.zero_grad()
output = model(data)
loss = criterion(output.view(-1, ntokens), targets)
loss.backward()
torch.nn.utils.clip_grad_norm_(model.parameters(), 0.5)
optimizer.step()
total_loss += loss.item()
log_interval = 200
if batch % log_interval == 0 and batch > 0:
cur_loss = total_loss / log_interval
elapsed = time.time() - start_time
print('| epoch {:3d} | {:5d}/{:5d} batches | '
'lr {:02.2f} | ms/batch {:5.2f} | '
'loss {:5.2f} | ppl {:8.2f}'.format(
epoch, batch, len(train_data) // bptt, scheduler.get_lr()[0],
elapsed * 1000 / log_interval,
cur_loss, math.exp(cur_loss)))
total_loss = 0
start_time = time.time()
def evaluate(eval_model, data_source):
eval_model.eval() # Turn on the evaluation mode
total_loss = 0.
ntokens = len(TEXT.vocab.stoi)
with torch.no_grad():
for i in range(0, data_source.size(0) - 1, bptt):
data, targets = get_batch(data_source, i)
output = eval_model(data)
output_flat = output.view(-1, ntokens)
total_loss += len(data) * criterion(output_flat, targets).item()
return total_loss / (len(data_source) - 1)
```
### trainning
```
# Train
bptt = 35
best_val_loss = float("inf")
epochs = 10 # The number of epochs
best_model = None
if os.path.isfile(args.model_path):
model.load_state_dict(torch.load(args.model_path))
for epoch in range(1, epochs + 1):
epoch_start_time = time.time()
train()
val_loss = evaluate(model, val_data)
print('-' * 89)
print('| end of epoch {:3d} | time: {:5.2f}s | valid loss {:5.2f} | '
'valid ppl {:8.2f}'.format(epoch, (time.time() - epoch_start_time),
val_loss, math.exp(val_loss)))
print('-' * 89)
if val_loss < best_val_loss:
best_val_loss = val_loss
best_model = model
torch.save(best_model.state_dict(), f = args.model_path)
scheduler.step()
test_loss = evaluate(best_model, test_data)
print('=' * 89)
print('| End of training | test loss {:5.2f} | test ppl {:8.2f}'.format(test_loss, math.exp(test_loss)))
print('=' * 89)
```
### Prediction
T,R,K; Run initialization
```
from torchtext.data.utils import get_tokenizer
TEXT = torchtext.data.Field(tokenize=get_tokenizer("basic_english"),
init_token='<sos>',
eos_token='<eos>',
lower=True)
train_txt, val_txt, test_txt = torchtext.datasets.WikiText2.splits(TEXT)
TEXT.build_vocab(train_txt)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
def batchify(data, bsz):
data = TEXT.numericalize([data.examples[0].text])
# Divide the dataset into bsz parts.
nbatch = data.size(0) // bsz
# Trim off any extra elements that wouldn't cleanly fit (remainders).
data = data.narrow(0, 0, nbatch * bsz)
# Evenly divide the data across the bsz batches.
data = data.view(bsz, -1).t().contiguous()
return data.to(device)
batch_size = 20
eval_batch_size = 10
train_data = batchify(train_txt, batch_size)
val_data = batchify(val_txt, eval_batch_size)
test_data = batchify(test_txt, eval_batch_size)
ntokens = len(TEXT.vocab.stoi) # the size of vocabulary
emsize = 200 # embedding dimension
nhid = 200 # the dimension of the feedforward network model in nn.TransformerEncoder
nlayers = 2 # the number of nn.TransformerEncoderLayer in nn.TransformerEncoder
nhead = 2 # the number of heads in the multiheadattention models
dropout = 0.2 # the dropout value
predict_model = TransformerModel(ntokens, emsize, nhead, nhid, nlayers, dropout).to(device)
predict_model.load_state_dict(torch.load(args.model_path))
# Get Results
bptt = 35
i = bptt * 2
# data, targets = get_batch(test_data, i)
data, targets = get_batch(train_data, i)
# best_model.eval()
output = predict_model(data)
def data2sent(data, func = None):
if func:
return [[TEXT.vocab.itos[func(ind).data.item()] for ind in data[:, i]]
for i in range(data.shape[1])]
else:
return [[TEXT.vocab.itos[ind.data.item()] for ind in data[:, i]]
for i in range(data.shape[1])]
# data2sent(data)
data2sent(predict_model(data[1:10,:]), func = lambda word_tensor: torch.argmax(word_tensor, dim = -1))
```
| github_jupyter |
## Import Necessary Packages
```
import numpy as np
import pandas as pd
import datetime
np.random.seed(1337) # for reproducibility
from sklearn.model_selection import train_test_split
from sklearn.metrics.classification import accuracy_score
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics.regression import r2_score, mean_squared_error, mean_absolute_error
from dbn.tensorflow import SupervisedDBNRegression
```
## Define Model Settings
```
RBM_EPOCHS = 5
DBN_EPOCHS = 150
RBM_LEARNING_RATE = 0.01
DBN_LEARNING_RATE = 0.01
HIDDEN_LAYER_STRUCT = [20, 50, 100]
ACTIVE_FUNC = 'relu'
BATCH_SIZE = 28
```
## Define Directory, Road, and Year
```
# Read the dataset
ROAD = "Vicente Cruz"
YEAR = "2015"
EXT = ".csv"
DATASET_DIVISION = "seasonWet"
DIR = "../../../datasets/Thesis Datasets/"
'''''''Training dataset'''''''
WP = False
weatherDT = "recon_weather" #orig_weather recon_weather
featureEngineering = "" #Rolling Expanding Rolling and Expanding
featuresNeeded = ['tempC', 'windspeedMiles', 'precipMM', 'visibility', #All Features
'humidity', 'pressure', 'cloudcover', 'heatIndexC',
'dewPointC', 'windChillC', 'windGustMiles', 'feelsLikeC']
# featuresNeeded = ['tempC', 'windspeedMiles', 'humidity', 'heatIndexC', #All corr > 0.1
# 'dewPointC','windChillC', 'feelsLikeC']
# featuresNeeded = ['heatIndexC', 'windspeedMiles', 'dewPointC']
ROLLING_WINDOW = 3
EXPANDING_WINDOW = 12
RECON_SHIFT = 96
RBM_EPOCHS = 5
DBN_EPOCHS = 150
RBM_LEARNING_RATE = 0.01
DBN_LEARNING_RATE = 0.01
HIDDEN_LAYER_STRUCT = [20, 50, 100]
ACTIVE_FUNC = 'relu'
BATCH_SIZE = 28
def reconstructDT(df, typeDataset='traffic', trafficFeatureNeeded=[]):
result_df = df.copy()
# Converting the index as date
result_df.index = pd.to_datetime(result_df.index, format='%d/%m/%Y %H:%M')
# result_df['year'] = result_df.index.year
result_df['month'] = result_df.index.month
# result_df['day'] = result_df.index.day
result_df['hour'] = result_df.index.hour
result_df['min'] = result_df.index.minute
result_df['dayOfWeek'] = result_df.index.dayofweek
if typeDataset == 'traffic':
for f in trafficFeatureNeeded:
result_df[f + '_' + str(RECON_SHIFT)] = result_df[f].shift(RECON_SHIFT)
result_df[f + '_' + str(RECON_SHIFT)] = result_df[f].shift(RECON_SHIFT)
result_df = result_df.iloc[96:, :]
for f in range(len(result_df.columns)):
result_df[result_df.columns[f]] = normalize(result_df[result_df.columns[f]])
return result_df
def shiftDTForReconstructed(df):
return df.iloc[shift:, :]
def getNeededFeatures(columns, arrFeaturesNeed=[], featureEngineering="Original"):
to_remove = []
if len(arrFeaturesNeed) == 0: #all features aren't needed
return []
else:
if featureEngineering == "Original":
compareTo = " "
elif featureEngineering == "Rolling" or featureEngineering == "Expanding":
compareTo = "_"
for f in arrFeaturesNeed:
for c in range(0, len(columns)):
if (len(columns[c].split(compareTo)) <= 1 and featureEngineering != "Original"):
to_remove.append(c)
if f not in columns[c].split(compareTo)[0] and columns[c].split(compareTo)[0] not in arrFeaturesNeed:
to_remove.append(c)
return to_remove
def normalize(data):
y = pd.to_numeric(data)
y = np.array(y.reshape(-1, 1))
scaler = MinMaxScaler()
y = scaler.fit_transform(y)
y = y.reshape(1, -1)[0]
return y
```
<br><br>
### Preparing Traffic Dataset
#### Importing Original Traffic (wo new features)
```
TRAFFIC_WINDOWSIZE = 3
TRAFFIC_DIR = DIR + "mmda/"
TRAFFIC_FILENAME = "mmda_" + ROAD + "_" + YEAR + "_" + DATASET_DIVISION
orig_traffic = pd.read_csv(TRAFFIC_DIR + TRAFFIC_FILENAME + EXT, skipinitialspace=True)
orig_traffic = orig_traffic.fillna(0)
#Converting index to date and time, and removing 'dt' column
orig_traffic.index = pd.to_datetime(orig_traffic.dt, format='%d/%m/%Y %H:%M')
cols_to_remove = [0]
cols_to_remove = getNeededFeatures(orig_traffic.columns, ["statusN"])
orig_traffic.drop(orig_traffic.columns[[cols_to_remove]], axis=1, inplace=True)
orig_traffic.head()
```
#### Importing Original Weather (wo new features)
```
WEATHER_DIR = DIR + "wwo/"
WEATHER_FILENAME = "wwo_" + ROAD + "_" + YEAR + "_" + DATASET_DIVISION
orig_weather = pd.read_csv(WEATHER_DIR + WEATHER_FILENAME + EXT, skipinitialspace=True)
cols_to_remove = [0]
cols_to_remove += getNeededFeatures(orig_weather.columns, arrFeaturesNeed=featuresNeeded)
orig_weather.index = pd.to_datetime(orig_weather.dt, format='%d/%m/%Y %H:%M')
orig_weather.drop(orig_weather.columns[[cols_to_remove]], axis=1, inplace=True)
orig_weather.head()
```
#### Importing Weather with Rolling features (with only needed features)
```
WEATHER_DIR = DIR + "wwo/Rolling/" + DATASET_DIVISION + "/"
WEATHER_FILENAME = "eng_win" + str(ROLLING_WINDOW) + "_wwo_" + ROAD + "_" + YEAR + "_" + DATASET_DIVISION
rolling_weather = pd.read_csv(WEATHER_DIR + WEATHER_FILENAME + EXT, skipinitialspace=True)
cols_to_remove = []
cols_to_remove += getNeededFeatures(rolling_weather.columns, orig_weather.columns, featureEngineering="Rolling")
rolling_weather.index = pd.to_datetime(rolling_weather.dt, format='%Y-%m-%d %H:%M')
rolling_weather.drop(rolling_weather.columns[[cols_to_remove]], axis=1, inplace=True)
for f in range(len(rolling_weather.columns)):
rolling_weather[rolling_weather.columns[f]] = normalize(rolling_weather[rolling_weather.columns[f]])
rolling_weather.head()
```
#### Importing Weather with Expanding features (with only needed features)
```
WEATHER_DIR = DIR + "wwo/Expanding/" + DATASET_DIVISION + "/"
WEATHER_FILENAME = "eng_win" + str(EXPANDING_WINDOW) + "_wwo_" + ROAD + "_" + YEAR + "_" + DATASET_DIVISION
expanding_weather = pd.read_csv(WEATHER_DIR + WEATHER_FILENAME + EXT, skipinitialspace=True)
cols_to_remove = []
cols_to_remove += getNeededFeatures(expanding_weather.columns, orig_weather.columns, featureEngineering="Rolling")
expanding_weather.index = pd.to_datetime(expanding_weather.dt, format='%d/%m/%Y %H:%M')
expanding_weather.drop(expanding_weather.columns[[cols_to_remove]], axis=1, inplace=True)
for f in range(len(expanding_weather.columns)):
expanding_weather[expanding_weather.columns[f]] = normalize(expanding_weather[expanding_weather.columns[f]])
expanding_weather.head()
```
#### Reconstructing Weather Input for recon_weather dataset
```
recon_weather = reconstructDT(orig_weather, 'weather', ['statusN'])
recon_weather.head()
```
### Merging datasets
```
''''''' Do not touch below '''''''
if weatherDT == "orig_weather":
print("Adding orig_weather")
arrDT = [orig_traffic, orig_weather]
elif weatherDT == "recon_weather":
print("Adding recon_weather")
arrDT = [orig_traffic.iloc[96:, :], recon_weather]
if featureEngineering == "Rolling":
print("Adding Rolling")
startIndex = np.absolute(len(arrDT[0])-len(rolling_weather))
temp = rolling_weather.iloc[startIndex:, :]
arrDT.append(temp)
elif featureEngineering == "Expanding":
print("Adding Expanding")
startIndex = np.absolute(len(arrDT[0])-len(expanding_weather))
temp = expanding_weather.iloc[startIndex:, :]
arrDT.append(temp)
elif featureEngineering == "Rolling and Expanding":
print("Adding Rolling and Expanding")
#Rolling
startIndex = np.absolute(len(arrDT[0])-len(rolling_weather))
temp = rolling_weather.iloc[startIndex:, :]
arrDT.append(temp)
#Expanding
startIndex = np.absolute(len(arrDT[0])-len(expanding_weather))
temp = expanding_weather.iloc[startIndex:, :]
arrDT.append(temp)
merged_dataset = pd.concat(arrDT, axis=1)
merged_dataset.head()
```
## Preparing Training dataset
### Merge Original (and Rolling and Expanding)
```
# To-be Predicted variable
Y = merged_dataset.statusN
Y = Y.fillna(0)
# Training Data
X = merged_dataset
X = X.drop(X.columns[[0]], axis=1)
# Splitting data
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.67, shuffle=False)
X_train = np.array(X_train)
X_test = np.array(X_test)
Y_train = np.array(Y_train)
Y_test = np.array(Y_test)
# Data scaling
# min_max_scaler = MinMaxScaler()
# X_train = min_max_scaler.fit_transform(X_train)
#Print training and testing data
pd.concat([X, Y.to_frame()], axis=1).head()
# Training
regressor = SupervisedDBNRegression(hidden_layers_structure=HIDDEN_LAYER_STRUCT,
learning_rate_rbm=RBM_LEARNING_RATE,
learning_rate=DBN_LEARNING_RATE,
n_epochs_rbm=RBM_EPOCHS,
n_iter_backprop=DBN_EPOCHS,
batch_size=BATCH_SIZE,
activation_function=ACTIVE_FUNC)
regressor.fit(X_train, Y_train)
#To check RBM Loss Errors:
rbm_error = regressor.unsupervised_dbn.rbm_layers[0].rbm_loss_error
#To check DBN Loss Errors
dbn_error = regressor.dbn_loss_error
# Test
# X_test = min_max_scaler.transform(X_test)
# Y_pred = regressor.predict(X_test)
min_max_scaler = MinMaxScaler()
X_test = min_max_scaler.fit_transform(X_test)
Y_pred = regressor.predict(X_test)
r2score = r2_score(Y_test, Y_pred)
rmse = np.sqrt(mean_squared_error(Y_test, Y_pred))
mae = mean_absolute_error(Y_test, Y_pred)
print('Done.\nR-squared: %.3f\nRMSE: %.3f \nMAE: %.3f' % (r2score, rmse, mae))
# Save the model
regressor.save('models/pm2_' + ROAD + '_' + YEAR + '.pkl')
```
### Results and Analysis below
##### Printing Predicted and Actual Results
```
startIndex = orig_weather.shape[0] - Y_pred.shape[0]
dt = orig_weather.index[startIndex:,]
temp = []
for i in range(len(Y_pred)):
temp.append(Y_pred[i][0])
d = {'Predicted': temp, 'Actual': Y_test, 'dt':dt}
df = pd.DataFrame(data=d)
df.head()
df.tail()
df.to_csv("output/pm2_" + ROAD + "_" + YEAR + EXT, encoding='utf-8')
```
#### Visualize trend of loss of RBM and DBN Training
```
import matplotlib.pyplot as plt
line1 = df.Actual
line2 = df.Predicted
plt.grid()
plt.plot(line1, c='red', alpha=0.4)
plt.plot(line2, c='blue', alpha=0.4)
plt.xlabel("Date")
plt.ylabel("Traffic Speed")
plt.yticks([0, 0.5, 1.0])
plt.show()
line1 = rbm_error
line2 = dbn_error
x = range(0, RBM_EPOCHS * len(HIDDEN_LAYER_STRUCT))
plt.plot(range(0, RBM_EPOCHS * len(HIDDEN_LAYER_STRUCT)), line1, c='red')
plt.xticks(x)
plt.xlabel("Iteration")
plt.ylabel("Error")
plt.show()
plt.plot(range(DBN_EPOCHS), line2, c='blue')
plt.xticks(x)
plt.xlabel("Iteration")
plt.ylabel("Error")
plt.show()
plt.plot(range(0, RBM_EPOCHS * len(HIDDEN_LAYER_STRUCT)), line1, c='red')
plt.plot(range(DBN_EPOCHS), line2, c='blue')
plt.xticks(x)
plt.xlabel("Iteration")
plt.ylabel("Error")
plt.show()
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import seaborn as sns
import glob
import os
import matplotlib.pyplot as plt
import shutil
from prediction_utils.util import df_dict_concat, yaml_read, yaml_write
project_dir = "/share/pi/nigam/projects/spfohl/cohorts/admissions/starr_20200523"
os.listdir(os.path.join(project_dir, 'experiments'))
experiment_name = 'baseline_tuning_fold_1_10'
baseline_files = glob.glob(
os.path.join(
project_dir,
'experiments',
experiment_name,
'**',
'result_df_training_eval.parquet'
),
recursive=True
)
baseline_df_dict = {
tuple(file_name.split('/'))[-4:-1]: pd.read_parquet(file_name)
for file_name in baseline_files
}
baseline_df = df_dict_concat(baseline_df_dict,
['task', 'config_filename', 'fold']
)
baseline_df.head()
assert (
baseline_df
.groupby(['task', 'config_filename'])
.agg(num_folds = ('fold', lambda x: len(x.unique())))
.query('num_folds != 10')
.shape[0]
) == 0
mean_performance = (
pd.DataFrame(
baseline_df
.query('metric == "loss" & phase == "val"')
.groupby(['config_filename', 'task'])
.agg(performance=('performance', 'mean'))
.reset_index()
)
)
best_model = (
mean_performance
.groupby('task')
.agg(performance=('performance','min'))
.merge(mean_performance)
)
# mean_performance
# mean_performance = (
# pd.DataFrame(
# baseline_df
# .query('metric == "loss" & phase == "val"')
# .groupby(['config_filename', 'task'])
# .agg({'performance': 'mean', 'config_filename': lambda x: x.array[-1], 'task': lambda x: x.array[-1]})
# .reset_index(drop=True)
# )
# )
# best_model = pd.DataFrame(mean_performance.groupby(['task']).performance.agg('min')).reset_index().merge(mean_performance)
# best_model
best_model_config_df = best_model[['config_filename', 'task']]
best_model_performance = baseline_df.merge(best_model_config_df)
best_model_performance[['task', 'config_filename']].drop_duplicates()
best_model_config_df
baseline_df
base_config_path = os.path.join(project_dir, 'experiments', experiment_name, 'config')
selected_config_path = os.path.join(project_dir, 'experiments', experiment_name, 'config', 'selected_models')
# Write to a new directory
for i, row in best_model_config_df.iterrows():
the_config = yaml_read(os.path.join(base_config_path, row.task, row.config_filename))
print(the_config)
the_config['label_col'] = row.task
os.makedirs(os.path.join(selected_config_path, row.task), exist_ok=True)
yaml_write(the_config, os.path.join(selected_config_path, row.task, row.config_filename))
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import sympy
import datetime as dt
import time
from math import *
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
from ipyleaflet import *
import folium
import json
import geopy.distance
from haversine import haversine
from tqdm import tqdm_notebook
sns.set()
%matplotlib inline
%config InlineBackend.figure_formats = {'png', 'retina'}
from matplotlib import font_manager, rc
plt.rcParams['axes.unicode_minus'] = False
import platform
if platform.system() == 'Darwin':
rc('font', family='AppleGothic')
elif platform.system() == 'Windows':
path = "c:/Windows/Fonts/malgun.ttf"
font_name = font_manager.FontProperties(fname=path).get_name()
rc('font', family=font_name)
import scipy as sp
import statsmodels.api as sm # statsmodel 기본 import
import statsmodels.formula.api as smf
import statsmodels.stats.api as sms
import sklearn as sk
import seaborn as sns
sns.set()
sns.set_style("whitegrid")
sns.set_color_codes()
import warnings
warnings.filterwarnings("ignore")
from patsy import dmatrix
train = pd.read_csv("../dataset/train.csv")
train.info()
train.describe()
pickup_datetime_dt = pd.to_datetime(train["pickup_datetime"])
dropoff_datetime_dt = pd.to_datetime(train["dropoff_datetime"])
train["pickup_datetime"] = pickup_datetime_dt
train["dropoff_datetime"] = dropoff_datetime_dt
train["pickup_date"] = train["pickup_datetime"].dt.date
train["dropoff_date"] = train["dropoff_datetime"].dt.date
train["pickup_month"] = train["pickup_datetime"].dt.month
train["dropoff_month"] = train["dropoff_datetime"].dt.month
train["pickup_weekday"] = train["pickup_datetime"].dt.weekday
train["dropoff_weekday"] = train["dropoff_datetime"].dt.weekday
train["pickup_hour"] = train["pickup_datetime"].dt.hour
train["dropoff_hour"] = train["dropoff_datetime"].dt.hour
train.info()
train.describe()
real = ["passenger_count", "pickup_longitude",
"pickup_latitude", "dropoff_longitude", "dropoff_latitude"]
cat = [elem for elem in train.columns if elem not in real]
cat.remove("trip_duration")
cat
len(train.columns), len(real), len(cat)
train_real = train[real]
train_real_sample = train_real.sample(10000)
train_real_sample.describe()
sns.pairplot(train_real_sample)
plt.show()
sns.boxplot(x="pickup_month", y=np.log1p(train["trip_duration"]),
data=train, palette="husl")
plt.show()
model = sm.OLS.from_formula("trip_duration ~ C(pickup_month) + 0", train)
result = model.fit()
print(result.summary())
model = sm.OLS.from_formula("trip_duration ~ C(pickup_weekday) + 0", train)
result = model.fit()
print(result.summary())
model = sm.OLS.from_formula("trip_duration ~ C(pickup_hour) + 0", train)
result = model.fit()
print(result.summary())
# pickup_hour를 카테고리값이 아닌 상수로 보고 회귀분석 시험
model = sm.OLS.from_formula("trip_duration ~ pickup_hour", train)
result = model.fit()
print(result.summary())
model = sm.OLS.from_formula("trip_duration ~ C(vendor_id) + 0", train)
result = model.fit()
print(result.summary())
model = sm.OLS.from_formula("trip_duration ~ C(store_and_fwd_flag) + 0", train)
result = model.fit()
print(result.summary())
```
| github_jupyter |
# Exercise - Hadamard (elementwise) matrix multiplication
In this exercise we are going to solidify our understanding of the process of OpenCL using a sister example of Hadamard matrix multiplication. Hadamard multiplication is elementwise multiplication. The values in matrices **D** and **E** at coordinates (i0,i1) are multiplied together to set the value at coordinates (i0,i1) in matrix **F**.
<figure style="margin-left:auto; margin-right:auto; width:80%;">
<img style="vertical-align:middle" src="../images/elementwise_multiplication.svg">
<figcaption style= "text-align:lower; margin:1em; float:bottom; vertical-align:bottom;">Elementwise multiplication of matrices D and E to get F.</figcaption>
</figure>
The source code is located in [mat_elementwise.cpp](mat_elementwise.cpp) and is similar to [mat_mult.cpp](mat_mult.cpp) in almost every aspect. Matrices **D** and **E** are read in from disk and matrix **F** is produced as the output. The code is missing some elements:
* The source code in [mat_elementwise.cpp](mat_elementwise.cpp) is missing the OpenCL machinery to write memory from **array_D** on the host to **buffer_D** on the device, and from **array_E** on the host to **buffer_E** on the device.
* In addition, the source code for the kernel in [kernels_elementwise.c](kernels_elementwise.c) is missing some code to perform the actual elementwise multiplication.
As an OpenCL developer your task is to fill in the necessary source to enable the program to work correctly.
## Constructing the inputs and solution
As before, we construct input matrices **D** and **E**, and write them to the files **array_D.dat** and **array_E.dat**.
```
import os
import sys
import numpy as np
sys.path.insert(0, os.path.abspath("../include"))
import py_helper
%matplotlib widget
# Matrices D, E, F are of size (NROWS_D, NCOLS_D)
NROWS_F = 520
NCOLS_F = 1032
# Data type
dtype = np.float32
mat_mul=py_helper.Hadamard(NROWS_F, NCOLS_F, dtype)
mat_mul.make_data()
```
## The desired answer
The source code [mat_elementwise_answer.cpp](mat_elementwise_answer.cpp) contains the full solution. By all means take a peek at the source code if you get stuck. If we run the solution and check the result we get no residual anywhere in the matrix **F**.
```
!make; ./mat_elementwise_answer.exe
```
## The code as it stands
Once the source code has been completed it will show the same results as the answer.
```
!make; ./mat_elementwise.exe
```
## Checking the output
The function **check_data** in the Python **Hadamard** class reads in **array_F.dat** and compares it to the one generated by Python.
```
mat_mul.check_data()
```
## Tasks
In these set of tasks the aim is to solidify some of the understanding developed in the walkthrough of the code. We are going to read through the documentation of a function and implement some very simple kernel code.
1. In the source file [mat_elementwise.cpp](mat_elementwise.cpp) (line 126), re-enable the memory copy from **array_D** and **array_E** on the host to **buffer_D** and **buffer_E** using the function [clEnqueueWriteBuffer](https://www.khronos.org/registry/OpenCL/sdk/3.0/docs/man/html/clEnqueueWriteBuffer.html). Read the [documentation](https://www.khronos.org/registry/OpenCL/sdk/3.0/docs/man/html/clEnqueueWriteBuffer.html) for that function and implement the copies. It may also be helpful to download the latest [OpenCL C specification](https://www.khronos.org/registry/OpenCL/specs/3.0-unified/pdf/OpenCL_C.pdf) and find that function in there.
1. Complete the kernel source code in [kernels_elementwise.c](kernels_elementwise.c) so that a new value in buffer F at coordinates (i0,i1) is constructed from the corresponding values in buffers D and E.
* Make sure you have a guard statement in place so that you don't overrun the bounds of buffer F. See the source code in [kernels_mat_mult.c](kernels_mat_mult.c) for an example.
* Use multi-dimensional indexing as shown in the <a href="../L2_Survival_C++/Lesson - Survival C++.ipynb">Survival C++</a> Lesson to index into arrays.
* If you get stuck you can just use the kernel from the answer in [kernels_elementwise_answer.c](kernels_elementwise_answer.c), just change line 124 of [mat_elementwise.cpp](mat_elementwise.cpp) to read in the kernel from the new source file.
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats
def r2(x, y):
return stats.pearsonr(x, y)[0] ** 2
%matplotlib inline
```
# Preparing Data
```
test_counts = {'yes': 256, 'no': 252, 'up': 272, 'down': 253, 'left': 267, 'right': 259, 'on': 246, 'off': 262, 'stop': 249, 'go': 251}
param_counts = {
'cnn_one_fstride4': {
'params': 220000,
'multiplies': 1430000
},
'cnn_one_fstride8': {
'params': 337000,
'multiplies': 1430000
},
'cnn_tpool2': {
'params': 1090000,
'multiplies': 103000000
},
'cnn_tpool3': {
'params': 823000,
'multiplies': 73700000
},
'cnn_trad_fpool3': {
'params': 1370000,
'multiplies': 125000000
},
'google-speech-dataset-compact': {
'params': 964000,
'multiplies': 5760000
},
'google-speech-dataset-full': {
'params': 1380000,
'multiplies': 98800000
}
}
def get_observations(fname):
observations = {'model': [], 'keyword': [], 'accuracy': [], 'time': [], 'total_energy': [], 'peak_power': [], 'params': [], 'multiplies': []}
with open(fname, 'r') as f:
for _ in range(7):
for i in range(10):
line = f.readline().rstrip()
parts = line.split(' ')
model, keyword, accuracy, time, total_energy, peak_power = parts
model = model.rstrip('\.onnx')
accuracy, time, total_energy, peak_power = list(map(float, [accuracy, time, total_energy, peak_power]))
accuracy *= 100
total_energy = 1000 * (total_energy - 1.9*time)
time *= 1000
peak_power -= 1.9
observations['model'].append(model)
observations['keyword'].append(keyword)
observations['accuracy'].append(accuracy)
observations['time'].append(time / test_counts[keyword])
observations['total_energy'].append(total_energy / test_counts[keyword])
observations['peak_power'].append(peak_power)
observations['params'].append(param_counts[model]['params'])
observations['multiplies'].append(param_counts[model]['multiplies'])
for i in range(6):
line = f.readline()
return observations
df = pd.DataFrame(get_observations('experiment_output_e2e.txt'))
df.head()
df_pre = pd.DataFrame(get_observations('experiment_output_preprocessing.txt'))
df_pre.head()
```
# Analysis
```
df_grouped = df.groupby('model')
df_grouped_means = df_grouped['accuracy', 'total_energy', 'peak_power', 'time', 'params', 'multiplies'].mean()
df_grouped_means.round(2)
df_pre_grouped = df_pre.groupby('model')
df_pre_grouped_means = df_pre_grouped['accuracy', 'total_energy', 'peak_power', 'time', 'params', 'multiplies'].mean()
df_pre_grouped_means
df_pre_grouped_means['time'].mean()
df_pre_grouped_means['total_energy'].mean()
df_pre_grouped_means['peak_power'].mean()
df_inf_only = df_grouped_means - df_pre_grouped_means
df_inf_only['peak_power'] = df_grouped_means['peak_power']
df_inf_only['params'] = df_grouped_means['params']
df_inf_only['multiplies'] = df_grouped_means['multiplies']
df_inf_only.round(2)
dims = (14, 6)
fig, ax = plt.subplots(figsize=dims)
g = sns.factorplot(x="accuracy", y="total_energy", hue="model", data=df, ax=ax)
g.set(xlim=(0, None), ylim=(0, None))
for ind, label in enumerate(ax.get_xticklabels()):
if ind % 10 == 0: # every 10th label is kept
label.set_visible(True)
else:
label.set_visible(False)
```
# Visualizations
## Energy vs. Multiplies
```
df_inf_aggregated = df_inf_only.reset_index()
ax = sns.regplot(x=df['params'], y=df['total_energy'])
ax = sns.regplot(x=df['multiplies'], y=df['total_energy'])
df.to_csv('observations.csv', index=False)
df_inf_aggregated.to_csv('observations_agg.csv', index=False)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/daemon-Lee/simplex_method_for_linear_program/blob/master/project/simplex_method/Simplex_method.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#@title Copyright 2020 Duy L.Dinh. { display-mode: "form" }
#@markdown CS1302 HE130655.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Project MAO302
This is final project of MAO302 Course make by FPT University
## Part 1: simplex method for linear program (LP)
```
import numpy as np
np.random.seed(2020)
```
### Generate input matrices of a standard linear program in matrix from
```
def gen_problem(n_var, n_contrain):
contrain = np.random.randint(low=-7, high=19, size=(n_var,n_contrain))
bacis = np.eye(n_contrain)
# A will contain the coefficients of the constraints
A = np.vstack((contrain,bacis)).T
# b will contain the amount of resources
b = np.random.randint(low=-7, high=19, size=(n_contrain,))
# c will contain coefficients of objective function Z
cz = np.random.randint(low=-7, high=19, size=(n_var,))
cb = np.zeros((n_contrain,))
c = np.concatenate([cz,cb])
return A, b, c
```
### Write a code to solve the generated LP using to phase simplex method in matrix form
```
#@title THE SIMPLEX METHOD IN MATRIX NOTATION
class Simplex_method:
#@markdown First input A, b, c, where:
#@markdown - **A** will contain the coefficients of the constraints
#@markdown - **b** will contain the amount of resources
#@markdown - **c** will contain coefficients of objective function Z
def __init__(self, A, b, c):
self.A = A
self.c = c
self.B = 0
self.n = 0
#@markdown Generate *B* and *N*
#@markdown - **B** will contain the Basic set
#@markdown - **n** will contain the nonbasic set
n_contrain = len(self.A)
n_var = len(self.c) - n_contrain
self.B = np.arange(n_var, n_var + n_contrain)[np.newaxis].T
self.n = np.arange(0, n_var)[np.newaxis].T
#@markdown - The initial values of the basic variables: xb = b
self.xb = np.transpose([b])
#@markdown - The initial values of the nonbasic dual variables: zn = -cn
self.zn = -self.c[self.n]
self.status = 'Optimal'
self.objective = 0
def solve(self, verbor=False):
self.count = 0
for i in self.n:
if True not in (self.A[:, i] > 0) and self.c[i] > 0:
print("Unbounded")
self.status = 'Unbounded'
sol = np.zeros(len(self.c))
sol[self.B] = self.xb
return {
'status': self.status,
"iter": self.count,
"objective": self.objective,
"sol": sol
}
#@markdown Find solution for problem
#@markdown - Check for Optimality. If xb ≥ 0 and zn ≥ 0, stop. The current
#@markdown solution is optimal.
if False not in (self.xb >= 0) and False not in (self.zn <= 0):
print("Optimal — the problem was trivial")
sol = np.zeros(len(self.c))
sol[self.B] = self.xb
return {
'status': self.status,
"iter": self.count,
"objective": self.objective,
"sol": sol
}
#@markdown - Since xb ≥ 0, the initial solution is **Primal feasible**, and hence
#@markdown we can apply the simplex method without needing any Phase I procedure.
elif False not in (self.xb >= 0) and False in (self.zn <= 0):
print("primal feasible")
print("run primal simplex method")
result = self.primal_simplex(verbor=verbor)
#@markdown - Since xb ≥ 0, the initial solution is **Dual feasible**
elif False in (self.xb >= 0) and False not in (self.zn <= 0):
print("run dual simplex method")
result = self.solve_two_phase(verbor=verbor)
#@markdown - Where both xb and cn have components of the wrong sign.
#@markdown In this case, we must employ a **two-phase procedure**.
else:
print("dual feasible")
print("Start convert negative components")
# self.zn = np.maximum(self.zn, -self.zn)
# self.zn = np.maximum(self.zn, 0)
print("run two phase simplex method")
result = self.solve_two_phase(verbor=verbor)
return result
def solve_two_phase(self, verbor=False):
#@markdown - In Phase I apply the dual simplex method to find an optimal solution
#@markdown of this modified problem Phase I is most likely not optimal, but it
#@markdown is feasible, and therefore the primal simplex method can be used to
#@markdown find the optimal solution to the original problem.
print("Phase one")
result = self.dual_simplex(verbor=verbor)
if result['status'] == 'Infeasible':
return result
print("Phase two")
result = self.primal_simplex(verbor=verbor)
return result
def primal_simplex(self, verbor=False):
objective = -np.inf
count = 0
Bi = self.A[:, self.B].reshape((-1, len(self.B)))
N = self.A[:, self.n].reshape((-1, len(self.n)))
if verbor:
A_hat = np.concatenate([self.B.T, self.xb.T, N.T, Bi.T]).T
print("Objective\n", np.concatenate([self.zn, self.xb]).T)
print("Dictionary\n", A_hat)
while(np.min(self.zn) < 0):
j = np.argmin(self.zn)
ej = np.zeros((1, len(self.zn))).T
ej[j] = 1
delta_xb = np.linalg.inv(Bi).dot(N).dot(ej)
t = np.max(delta_xb/self.xb)**-1
if t < 0 or t == np.inf:
self.status = 'Unbounded'
sol = np.zeros(len(self.c))
sol[self.B] = self.xb
return {
'status': self.status,
"iter": self.count,
"objective": self.objective,
"sol": sol
}
i = np.argmax(delta_xb/self.xb)
ei = np.zeros((1, len(self.xb))).T
ei[i] = 1
delta_zn = -(np.linalg.inv(Bi).dot(N)).T.dot(ei)
s = self.zn[j]/delta_zn[j]
self.xb = self.xb - t*delta_xb
self.zn = self.zn - s*delta_zn
self.xb[i] = t
self.zn[j] = s
# pivot swap
pivot = self.B[i].copy()
self.B[i] = self.n[j].copy()
self.n[j] = pivot
Bi = self.A[:, self.B].reshape((-1, len(self.B)))
N = self.A[:, self.n].reshape((-1, len(self.n)))
count += 1
self.count += 1
self.objective = self.xb.T.dot(self.c[self.B]).reshape(-1)[0]
if verbor:
A_hat = np.concatenate([self.B.T, self.xb.T, N.T, Bi.T]).T
print("iter:", count)
print("Dictionary\n", A_hat)
print("objective:", self.objective)
if self.objective > objective:
objective = self.objective
else:
self.status = 'Infeasible'
sol = np.zeros(len(self.c))
sol[self.B] = self.xb
return {
'status': self.status,
"iter": self.count,
"objective": self.objective,
"sol": sol
}
sol = np.zeros(len(self.c))
sol[self.B] = self.xb
return {
'status': self.status,
"iter": self.count,
"optimal": self.objective,
"sol": sol
}
def dual_simplex(self, verbor=False):
objective = np.inf
count = 0
Bi = self.A[:, self.B].reshape((-1, len(self.B)))
N = self.A[:, self.n].reshape((-1, len(self.n)))
if verbor:
A_hat = np.concatenate([self.B.T, self.xb.T, N.T, Bi.T]).T
print("Objective\n", np.concatenate([self.zn, self.xb]).T)
print("Dictionary\n", A_hat)
while(np.min(self.xb) < 0):
i = np.argmin(self.xb)
ei = np.zeros((1, len(self.xb))).T
ei[i] = 1
delta_zn = -(np.linalg.inv(Bi).dot(N)).T.dot(ei)
s = np.max(delta_zn/self.zn)**-1
j = np.argmax(delta_zn/self.zn)
ej = np.zeros((1, len(self.zn))).T
ej[j] = 1
delta_xb = np.linalg.inv(Bi).dot(N).dot(ej)
t = self.xb[i]/delta_xb[i]
self.xb = self.xb - t*delta_xb
self.zn = self.zn - s*delta_zn
self.xb[i] = t
self.zn[j] = s
# pivot
pivot = self.B[i].copy()
self.B[i] = self.n[j].copy()
self.n[j] = pivot
Bi = self.A[:, self.B].reshape((-1, len(self.B)))
N = self.A[:, self.n].reshape((-1, len(self.n)))
A_hat = np.concatenate([self.B.T, self.xb.T, N.T, Bi.T]).T
count += 1
self.count += 1
self.objective = self.xb.T.dot(self.c[self.B]).reshape(-1)[0]
if verbor:
A_hat = np.concatenate([self.B.T, self.xb.T, N.T, Bi.T]).T
print("iter:", count)
print("Dictionary\n", A_hat)
print("objective:", self.objective)
if self.objective < objective:
objective = self.objective
else:
self.status = 'Infeasible'
sol = np.zeros(len(self.c))
sol[self.B] = self.xb
return {
'status': self.status,
"iter": self.count,
"objective": self.objective,
"sol": sol
}
sol = np.zeros(len(self.c))
sol[self.B] = self.xb
return {
'status': self.status,
"iter": self.count,
"objective": self.objective,
"sol": sol
}
print("Exercise 2.3")
# A will contain the coefficients of the constraints
A = np.array([[-1, -1, -1, 1, 0],
[2, -1, 1, 0, 1]])
# b will contain the amount of resources
b = np.array([-2, 1])
# c will contain coefficients of objective function Z
c = np.array([2, -6, 0, 0, 0])
simplex = Simplex_method(A, b, c)
print(simplex.solve(verbor=True))
```
### Solve the genarated LP by a pulp and cplex tool
#### pulp lib
Install pulp
```
!pip install pulp
print("Exercise 2.3")
# A will contain the coefficients of the constraints
A = np.array([[-1, -1, -1, 1, 0],
[2, -1, 1, 0, 1]])
# b will contain the amount of resources
b = np.array([-2, 1])
# c will contain coefficients of objective function Z
c = np.array([2, -6, 0, 0, 0])
import pulp as p
# Generate B and N
n_contrain = len(A)
n_var = len(c) - n_contrain
B = np.arange(n_var, n_var + n_contrain)[np.newaxis].T
n = np.arange(0, n_var)[np.newaxis].T
# Create a LP Minimization problem
Lp_prob = p.LpProblem('Problem', p.LpMaximize)
# Create problem Variables
x = [p.LpVariable("x"+str(i), lowBound = 0) for i in range(1,n_var+1)]
# Objective Function
objective = 0
for i in range(n_var):
objective += c[i]*x[i]
Lp_prob += objective
# Constraints:
for i in range(n_contrain):
contrain = 0
for j in range(n_var):
contrain += A[i,j]*x[j] <= b[i]/n_var
Lp_prob += contrain
# Display the problem
print(Lp_prob)
status = Lp_prob.solve() # Solver
print(p.LpStatus[status]) # The solution status
# Printing the final solution
print(p.value(x[0]), p.value(x[1]), p.value(x[2]), p.value(Lp_prob.objective))
import pulp as p
def pulp_lib(A, b, c, verbor=False):
# Generate B and N
n_contrain = len(A)
n_var = len(c) - n_contrain
B = np.arange(n_var, n_var + n_contrain)[np.newaxis].T
n = np.arange(0, n_var)[np.newaxis].T
# Create a LP Minimization problem
Lp_prob = p.LpProblem('Problem', p.LpMaximize)
# Create problem Variables
x = [p.LpVariable("x"+str(i), lowBound = 0) for i in range(1,n_var+1)]
# Objective Function
objective = 0
for i in range(n_var):
objective += c[i]*x[i]
Lp_prob += objective
# Constraints:
for i in range(n_contrain):
contrain = 0
for j in range(n_var):
contrain += A[i,j]*x[j] <= b[i]/n_var
Lp_prob += contrain
status = Lp_prob.solve() # Solver
if verbor:
print(p.LpStatus[status]) # The solution status
# Printing the final solution
print(p.value(Lp_prob.objective))
return {
'status': p.LpStatus[status],
'objective': p.value(Lp_prob.objective)
}
```
#### cplex
```
!pip install cplex
import cplex
def cplex_lib(A, b, c):
# Input all the data and parameters here
num_constraints = len(A)
num_decision_var = len(c) - num_constraints
n = np.arange(0, num_decision_var)[np.newaxis].T
A = A[:,n.T].reshape(num_constraints, num_decision_var).tolist()
b = b.tolist()
c = c[n].T.reshape(len(n)).tolist()
# constraint_type = ["L", "L", "L"] # Less, Greater, Equal
constraint_type = ["L"]*num_constraints
# ============================================================
# Establish the Linear Programming Model
myProblem = cplex.Cplex()
# Add the decision variables and set their lower bound and upper bound (if necessary)
myProblem.variables.add(names= ["x"+str(i) for i in range(num_decision_var)])
for i in range(num_decision_var):
myProblem.variables.set_lower_bounds(i, 0.0)
# Add constraints
for i in range(num_constraints):
myProblem.linear_constraints.add(
lin_expr= [cplex.SparsePair(ind= [j for j in range(num_decision_var)], val= A[i])],
rhs= [b[i]],
names = ["c"+str(i)],
senses = [constraint_type[i]]
)
# Add objective function and set its sense
for i in range(num_decision_var):
myProblem.objective.set_linear([(i, c[i])])
myProblem.objective.set_sense(myProblem.objective.sense.maximize)
# Solve the model and print the answer
myProblem.solve()
return{
'objective': myProblem.solution.get_objective_value(),
'status': myProblem.solution.get_status_string(),
'sol': myProblem.solution.get_values()
}
print("Exercise 2.3")
# A will contain the coefficients of the constraints
A = np.array([[-1, -1, -1, 1, 0],
[2, -1, 1, 0, 1]])
# b will contain the amount of resources
b = np.array([-2, 1])
# c will contain coefficients of objective function Z
c = np.array([2, -6, 0, 0, 0])
cplex_lib(A, b, c)
```
### Repeat (1)-(3) one hundred timnes and compare the mean and standard deviation of running time of your code with those of the chosen tool.
```
n_sample = 100
np.random.seed(2020)
A_list = []
b_list = []
c_list = []
for i in range(n_sample):
n_var = np.random.randint(low=2, high=7)
n_contrain = np.random.randint(low=2, high=7)
A, b, c = gen_problem(n_var, n_contrain)
A_list.append(A)
b_list.append(b)
c_list.append(c)
from time import time
running_time_pulp = []
output_pulp = []
for i in range(n_sample):
start = time()
output_pulp.append(pulp_lib(A, b, c, verbor=False))
end = time() - start
running_time_pulp.append(end)
running_time_cplex = []
output_cplex = []
for i in range(n_sample):
start = time()
output_cplex.append(pulp_lib(A, b, c, verbor=False))
end = time() - start
running_time_cplex.append(end)
running_time_simplex_method = []
output_simplex_method= []
for i in range(n_sample):
start = time()
simplex = Simplex_method(A, b, c)
output_simplex_method.append(simplex.solve(verbor=False))
end = time() - start
running_time_simplex_method.append(end)
#@title Compare pulp and Simplex method
# Simplex method
mean_Simplex_method = np.mean(running_time_simplex_method)
std_Simplex_method = np.std(running_time_simplex_method)
# pulp
mean_pulp = np.mean(running_time_pulp)
std_pulp = np.std(running_time_pulp)
# cplex
mean_cplex = np.mean(running_time_cplex)
std_cplex = np.std(running_time_cplex)
print("mean running time of pulp - simplex_method (s):", mean_pulp - mean_Simplex_method)
print("standard deviation running time of pulp - simplex_method (s):", std_pulp - std_Simplex_method)
import seaborn as sns; sns.set()
import matplotlib.pyplot as plt
mean = np.array([mean_Simplex_method, mean_pulp, mean_cplex])[np.newaxis]
ax = sns.heatmap(mean, annot=True)
plt.title("Compare mean")
ax.set_xticklabels(['code','pulp','cplex'])
std = np.array([std_Simplex_method, std_pulp, std_cplex])[np.newaxis]
ax = sns.heatmap(std, annot=True)
plt.title("Compare standard deviation")
ax.set_xticklabels(['code','pulp','cplex'])
```
| github_jupyter |
<img src='../img/acam_logos.png' alt='Logo EU Copernicus EUMETSAT' align='right' width='65%'></img>
<br>
# 2.2 Copernicus Sentinel-5P TROPOMI Carbon monoxide (CO)
### Fire Monitoring - September 2020 - Indonesian Fires
A precursor satellite mission, Sentinel-5P aims to fill in the data gap and provide data continuity between the retirement of the Envisat satellite and NASA's Aura mission and the launch of Sentinel-5. The Copernicus Sentinel-5P mission is being used to closely monitor the changes in air quality and was launched in October 2017.
Sentinel-5P Pre-Ops data are disseminated in the `netCDF` format and can be downloaded via the [Copernicus Open Access Hub](https://scihub.copernicus.eu/).
Sentinel-5P carries the `TROPOMI` instrument, which is a spectrometer in the UV-VIS-NIR-SWIR spectral range. `TROPOMI` provides measurements on:
* `Ozone`
* `NO`<sub>`2`</sub>
* `SO`<sub>`2`</sub>
* `Formaldehyde`
* `Aerosol`
* `Carbon monoxide`
* `Methane`
* `Clouds`
This notebook provides an introduction to the Sentinel-5P TROPOMI Carbon monoxide data and how it can be used to monitor wildfires. The notebook examines the **Indonesian wildfires** which occur each year between June and September. The example shows a Sentinel-5P scene captured on 17 September 2020.
#### This module has the following outline:
* [1 - Load and browse Sentinel-5P TROPOMI Carbon Monoxide data](#load_s5p)
* [2 - Visualize Sentinel-5P TROPOMI Carbon Monoxide data](#plotting_s5p)
<hr>
#### Load required libraries
```
%matplotlib inline
import os
import xarray as xr
import numpy as np
import netCDF4 as nc
import pandas as pd
from IPython.display import HTML
import matplotlib.pyplot as plt
import matplotlib.colors
from matplotlib.cm import get_cmap
from matplotlib import animation
import cartopy.crs as ccrs
from cartopy.mpl.gridliner import LONGITUDE_FORMATTER, LATITUDE_FORMATTER
import cartopy.feature as cfeature
from matplotlib.axes import Axes
from cartopy.mpl.geoaxes import GeoAxes
GeoAxes._pcolormesh_patched = Axes.pcolormesh
import warnings
warnings.simplefilter(action = "ignore", category = RuntimeWarning)
warnings.simplefilter(action = "ignore", category = UserWarning)
```
##### Load helper functions
```
%run ../functions.ipynb
```
<hr>
## <a id="load_s5p"></a>Load and browse Sentinel-5P TROPOMI Carbon Monoxide data
A Sentinel-5P TROPOMI Carbon Monoxide (CO) Level 2 file is organised in two groups: `PRODUCT` and `METADATA`. The `PRODUCT` group stores the main data fields of the product, including `latitude`, `longitude` and the variable itself. The `METADATA` group provides additional metadata items.
Sentinel-5P TROPOMI variables have the following dimensions:
* `scanline`: the number of measurements in the granule / along-track dimension index
* `ground_pixel`: the number of spectra in a measurement / across-track dimension index
* `time`: time reference for the data
* `corner`: pixel corner index
* `layer`: number of layers
Sentinel-5P TROPOMI data is disseminated in `netCDF`. You can load a `netCDF` file with the `open_dataset()` function of the xarray library. In order to load the variable as part of a Sentinel-5P data files, you have to specify the following keyword arguments:
- `group='PRODUCT'`: to load the `PRODUCT` group
Let us load a Sentinel-5P TROPOMI data file as `xarray.Dataset` from 17 September 2020 and inspect the data structure:
```
file = xr.open_dataset('../eodata/sentinel5p/S5P_OFFL_L2__CO_____20200917T044812_20200917T062941_15178_01_010302_20200918T183114.nc', group='PRODUCT')
file
```
<br>
You see that the loaded data object contains five dimensions and five data variables:
* **Dimensions**:
* `scanline`
* `ground_pixel`
* `time`
* `corner`
* `layer`
* **Data variables**:
* `delta_time`: the offset of individual measurements within the granule, given in milliseconds
* `time_utc`: valid time stamp of the data
* `qa_value`: quality descriptor, varying between 0 (nodata) and 1 (full quality data).
* `carbonmonoxide_total_column`
* `carbonmonoxide_total_column_precision`
<br>
### <a id='data_retrieve_s5P'></a>Retrieve the variable `carbonmonoxide_total_column` as xarray.DataArray
You can specify one variable of interest by putting the name of the variable into square brackets `[]` and get more detailed information about the variable. E.g. `carbonmonoxide_total_column` is the 'Atmosphere mole content of carbon monoxide', has the unit mol m<sup>-2</sup> and has three dimensions, `time`, `scanline` and `ground_pixel` respectively.
```
co = file['carbonmonoxide_total_column']
co
```
<br>
You can do this for the available variables, but also for the dimensions latitude and longitude.
```
latitude = co['latitude']
longitude = co['longitude']
print('Latitude')
print(latitude)
print('Longitude')
print(longitude)
```
<br>
You can retrieve the array values of the variable with squared brackets: `[:,:,:]`. One single time step can be selected by specifying one value of the time dimension, e.g. `[0,:,:]`.
```
co_1709 = co[0,:,:]
co_1709
```
<br>
The attributes of the `xarray.DataArray` hold the entry `multiplication_factor_to_convert_to_molecules_percm2`, which is a conversion factor that has to be applied to convert the data from `mol per m`<sup>`2`</sup> to `molecules per cm`<sup>`2`</sup>.
```
conversion_factor = co_1709.multiplication_factor_to_convert_to_molecules_percm2
conversion_factor
```
<br>
Additionally, you can save the attribute `longname`, which you can make use of when visualizing the data.
```
longname = co_1709.long_name
longname
```
<br>
## <a id="plotting_s5p"></a>Visualize Sentinel-5P TROPOMI Carbon Monoxide data
The next step is to visualize the dataset. You can use the function [visualize_pcolormesh](./functions.ipynb#visualize_pcolormesh), which makes use of matploblib's function `pcolormesh` and the [Cartopy](https://scitools.org.uk/cartopy/docs/latest/) library.
With `?visualize_pcolormesh` you can open the function's docstring to see what keyword arguments are needed to prepare your plot.
In order to make it easier to visualize the Carbon Monoxide values, we apply the conversion factor to the `xarray.DataArray`. This converts the Carbon Monoxide values from *mol per m<sup>2</sup>* to *molecules per cm<sup>2</sup>*.
```
co_1709_converted = co_1709*conversion_factor
co_1709_converted
```
<br>
For visualization, you can use the function [visualize_pcolormesh](../functions.ipynb#visualize_pcolormesh) to visualize the data. The following keyword arguments have to be defined:
* `data_array`
* `longitude`
* `latitude`
* `projection`
* `color palette`
* `unit`
* `long_name`
* `vmin`,
* `vmax`
* `extent (lonmin, lonmax, latmin, latmax)`
* `set_global`
```
visualize_pcolormesh(data_array=co_1709_converted*1e-18,
longitude=co_1709_converted.longitude,
latitude=co_1709_converted.latitude,
projection=ccrs.Mollweide(),
color_scale='viridis',
unit='*1e-18 molecules per cm2',
long_name=longname + ' ' + str(co_1709_converted.time.data),
vmin=0,
vmax=4,
lonmin=longitude.min(),
lonmax=longitude.max(),
latmin=latitude.min(),
latmax=latitude.max(),
set_global=True)
```
<br>
You can zoom into a region by specifying a `bounding box` of interest. Let us set the extent to Borneo, with: `[-5, 8., 105., 120]`. The above plotting function [visualize_pcolormesh](./functions.ipynb#visualize_pcolormesh) allows for setting a specific bounding box. You simply have to set the `set_global` key to False. It is best to adjust the projection to `PlateCarree()`, as this will be more appropriate for a regional subset.
```
latmin = -5
latmax = 8.
lonmin = 105.
lonmax = 120.
visualize_pcolormesh(data_array=co_1709_converted*1e-18,
longitude=co_1709_converted.longitude,
latitude=co_1709_converted.latitude,
projection=ccrs.PlateCarree(),
color_scale='viridis',
unit='*1e-18 molecules per cm2',
long_name=longname + ' ' + str(co_1709_converted.time.data),
vmin=0,
vmax=2.7,
lonmin=lonmin,
lonmax=lonmax,
latmin=latmin,
latmax=latmax,
set_global=False)
```
<br>
<br>
<hr>
<img src='../img/copernicus_logo.png' alt='Logo EU Copernicus' align='right' width='20%'><br><br><br><br>
<p style="text-align:right;">This project is licensed under the <a href="./LICENSE">MIT License</a> and is developed under a Copernicus contract.
| github_jupyter |
# QMIND Workshop
In this workshop, we'll train and publish a CIFAR10 classifier to the Distributed Computer. This will allow us to submit a large number of tasks to a distributed computer to perform inference on a large number of images in parallel.
Let's begin in python.
```
%%capture
import numpy as np
import tensorflow as tf
from matplotlib import pyplot as plt
try:
import tensorflowjs as tfjs
except:
!pip install tensorflowjs tf-estimator-nightly 'ipykernel<5.0.0'
import tensorflowjs as tfjs
(_xtrain, _ytrain), (_xtest, _ytest) = tf.keras.datasets.cifar10.load_data()
_xtrain = _xtrain.astype(np.float32) / 255.
_xtest = _xtest.astype(np.float32) / 255.
_ytrain_onehot = tf.keras.utils.to_categorical(_ytrain, 10)
_ytest_onehot = tf.keras.utils.to_categorical(_ytest, 10)
print(_xtrain.min(), _xtrain.max(), _xtrain.dtype, _xtrain.shape, _ytest.shape)
class Classifier(tf.keras.Model):
def __init__(self, num_classes, name=None):
super(Classifier, self).__init__(name=name)
self.num_classes = num_classes
self.ls = [
tf.keras.layers.Conv2D(32 ,3,strides=2, padding='same', activation='relu'),
tf.keras.layers.Conv2D(64 ,3,strides=2, padding='same', activation='relu'),
tf.keras.layers.Conv2D(128,3,strides=2, padding='same', activation='relu'),
tf.keras.layers.GlobalAveragePooling2D(),
tf.keras.layers.Dense(512, activation='relu'),
tf.keras.layers.Dropout(0.5),
tf.keras.layers.Dense(self.num_classes, activation='softmax')
]
@tf.function
def call(self,x):
for l in self.ls:
x = l(x)
return x
model = Classifier(10)
model.compile(
loss = 'categorical_crossentropy',
metrics = ['accuracy']
)
_history = model.fit(_xtrain, _ytrain_onehot,
batch_size=128,
verbose=1,
epochs=10,
callbacks=[
tf.keras.callbacks.EarlyStopping(patience=3)
],
validation_data=(_xtest, _ytest_onehot))
plt.title("Loss")
plt.plot(_history.history['loss'])
plt.plot(_history.history['val_loss'])
plt.legend(['training loss', 'validation loss'])
plt.show()
plt.title("Acc")
plt.plot(_history.history['accuracy'])
plt.plot(_history.history['val_accuracy'])
plt.legend(['training accuracy', 'validation accuracy'])
plt.show()
```
# Publishing our model
Now we'll clone this helpful tensorflowjs utility and use tensorflowjs converter to convert our python model to a tensorflowjs model and then publishing this model to the Distributed Computer.
```
%%capture
!git clone https://github.com/Kings-Distributed-Systems/tfjs_util.git
!cd tfjs_util && npm i && npm run postinstall
!mkdir -p './tfjs_model'
!mkdir -p './saved_model'
model.save('./saved_model')
tfjs.converters.convert_tf_saved_model('./saved_model', './tfjs_model')
#Serialize and submit model to DCP package manager with version number X.X.XX
!node /home/mgasmallah/DCP/dcp-utils/tfjs_utils/bin/serializeModel.js -m ./tfjs_model/model.json \
-o dcp_cifar_10_ex/cifar10.js -p 0.0.6 -d
!rm -rf ./tfjs_model ./saved_model
```
# Pixiedust-node
Now we'll use a library that will enable us to run javascript in a jupyter notebook so that we can use the Distributed Compute Protocol to submit inference jobs to the cloud.
```
#!pip install pixiedust git+https://github.com/Kings-Distributed-Systems/pixiedust_node#egg=pixiedust_node
# %%capture
import pixiedust
import pixiedust_node
npm.install( ['dcp-client'] )
ID_KEY_LOC = '/home/mgasmallah/DCP/keys/id.keystore'
ACC_KEY_LOC= '/home/mgasmallah/DCP/keys/AISTEST.keystore'
SCHEDULER = 'https://demo-scheduler.distributed.computer'
node.clear();
!job-utility cancelAllJobs -I $ID_KEY_LOC --default-bank-account-file $ACC_KEY_LOC --scheduler $SCHEDULER
%%node
process.argv.push('-I', ID_KEY_LOC, '--default-bank-account-file', ACC_KEY_LOC, '--scheduler', SCHEDULER)
require('dcp-client').initSync(process.argv)
const compute = require('dcp/compute');
const wallet = require('dcp/wallet');
const dcpCli = require('dcp/dcp-cli');
var accountKeystore;
var identityKeystore;
(async function(){
identityKeystore = await dcpCli.getIdentityKeystore();
wallet.addId(identityKeystore);
accountKeystore = await dcpCli.getAccountKeystore();
console.log("Keystores loaded!");
})();
sample_cifar = _xtest[:100]
cifar_true = _ytest_onehot[:100]
sample_cifar.shape, sample_cifar[0,0,0,0], cifar_true.shape, cifar_true.dtype
%%node
var cifarResults = []
async function main(){
let x = Float32Array.from(sample_cifar.typedArray);
let y = Float32Array.from(cifar_true.typedArray);
console.log(x[0]);
let submission_array = [];
let num_elem_x = sample_cifar.shape[1]*sample_cifar.shape[2]*sample_cifar.shape[3];
let num_elem_y = cifar_true.shape[1];
for (let i = 0; i < sample_cifar.shape[0]; i++){
submission_array.push(
{
'x': Array.from(x.slice(i*num_elem_x, (i+1) * num_elem_x)),
'y': Array.from(y.slice(i*num_elem_y, (i+1) * num_elem_y))
}
);
}
let job = compute.for( submission_array, async function(data){
progress(0.);
const tf = require('tfjs');
tf.setBackend('cpu');
await tf.ready();
progress(0.15);
const model = await require('cifar10').getModel();
progress(0.45);
let inpTensor = tf.tensor(data.x, [1, 32, 32, 3], dtype='float32');
let y_true = tf.tensor(data.y, [1, 10], dtype='int32');
progress(0.5);
let y_pred = model.predict(inpTensor);
progress(0.75);
const accuracyTensor = tf.metrics.categoricalAccuracy(y_true, y_pred);
progress(0.85);
const acc = accuracyTensor.dataSync()[0];
progress(1.);
return acc;
});
job.on('accepted', ()=>{
console.log("Job Accepted...");
});
job.on('status', (status)=>{
console.log("Received a status update: ", status);
});
job.on('result', (value)=>{
cifarResults.push(value.result);
})
job.on('console', (output)=>{
console.log(output.message);
})
job.on('error', (err)=>{
console.log(err);
})
job.requires('aistensorflow/tfjs');
job.requires('dcp_cifar_10_ex/cifar10')
// job.requirements.environment.offscreenCanvas = true;
job.public.name = 'dcp-workshop';
await job.exec(compute.marketValue, accountKeystore);
console.log("Done Executing job!");
}
main();
print("Accuracy on distributed 100 set sample: ", sum(cifarResults)/len(cifarResults))
```
# Your turn!
Now try it with something like the fashion mnist dataset!
```
(_xtrain, _ytrain), (_xtest, _ytest) = tf.keras.datasets.fashion_mnist.load_data()
plt.imshow(_xtrain[np.random.randint(_xtrain.shape[0])], cmap='gray')
plt.show()
```
| github_jupyter |
# How to automate IP ranges calculations in Azure using PowerShell
> The notebook does work on Linux and [](https://mybinder.org/v2/gh/eosfor/scripting-notes/HEAD)
## Scenario
Assume we got the IP rage of `10.172.0.0/16` from the network team for planned Azure Landing Zones. We want to automate this by making a tool, which will automatically calculate IP ranges for us given some high level and simple to understand details about the future networks.
This notebook shows how to do it using the [ipmgmt](https://github.com/eosfor/ipmgmt) module. So let us go and install it first
```
Install-Module ipmgmt -Scope CurrentUser
```
And import it
```
Import-Module ipmgmt
```
The module contains only two cmdlets. The `Get-VLSMBreakdown` breaks down a range into smaller ones, meaning we can use it to break a range into VNETs and then each VNET into subnets. The `Get-IPRanges` cmdlet given the list of ranges in-use and the "root" range tries to find a free slot of the specified size. It can be used to avoid losing IP space
```
Get-Command -Module ipmgmt
```
Let us look at how we can break down our big "root" IP range into smaller ones. For that, we just need to prepare a list of smaller sub-ranges in the form of PowerShell hashtables, like so `@{type = "VNET-HUB"; size = (256-2)}`. Here we say that the name of the range is `VNET-HUB` and the size is simply `256-2`, which is the maximum number of IPs in the `/24` subnet minus 2, the first and the last.
If we need more than one, we just make an array of these hashtables
```
$subnets = @{type = "VNET-HUB"; size = (256-2)},
@{type = "VNET-A"; size = (256-2)}
```
Now everything is ready and we can try to break the "root" network
```
Get-VLSMBreakdown -Network 10.172.0.0/16 -SubnetSize $subnets | ft type, network, netmask, *usable, cidr -AutoSize
```
Here we got two ranges named `VNET-A` and `VNET-HUB`. However, by doing so we made a few unused slots in the `root` range. They are marked as `reserved`, just for our convenience. It shows what happens to the range when we break it down. The smaller sub-ranges you make, the more of such unused ranges you get in the end.
Ok, let's try to use what we've got. For that, we need to authenticate to Azure. When running locally you can just do
```
Login-AzAccount
```
In Binder however, it needs to be a bit different, like this
```
Connect-AzAccount -UseDeviceAuthentication
```
Once authenticated, we can create networks, for example, like this. Here we first filter out the `reserved` ones for simplicity.
```
$vnets = Get-VLSMBreakdown -Network 10.172.0.0/16 -SubnetSize $subnets | ? type -ne 'reserved'
$vnets | % {
New-AzVirtualNetwork -Name $_.type -ResourceGroupName 'vnet-test' `
-Location 'eastus2' -AddressPrefix "$($_.Network)/$($_.cidr)" | select name, AddressSpace, ResourceGroupName, Location
}
```
Ok, assume, at some point, we need to add a few more networks. And at the same time e may want to reuse one of those `reserved` slots, if it matches the size. This is what `Get-IPRanges` does. It takes a lit of IP ranges "in-use" and returns slots that can fit the range in question. For example, in our case, we have a "base range" of `10.10.0.0/16` and two ranges in-use `10.10.5.0/24`, `10.10.7.0/24`. We are looking for a range of the size `/22`. So the cmdlet recommends us to use the `10.172.4.0/22`, which is one of the `reserved` ranges from the previous example
```
Get-IPRanges -Networks "10.172.1.0/24", "10.172.0.0/24" -CIDR 22 -BaseNet "10.172.0.0/16" | ft -AutoSize
```
What if we need to find more than just one range at a time? Need not worry. We can do it with this simple script. And we are using Azure as a source of truth, as we can always query it for the real IP ranges, which are in use.
So what we need to do is relatively simple:
- make a list of sizes we want to create and put it into a variable - `$cidrRange`
- pull the ranges from Azure. We assume they are in use by someone - `$existingRanges`
- cast whatever we pulled from azure the `System.Net.IPNetwork` for correctness. This type is used inside the `ipmgmt` module to store information about networks and do all the calculations, comparisons, etc.
- now we run through the list of sizes, for each of them ask `Get-IPRanges` to find a proper slot, and accumulate the results
Now we just need to mark the new ranges as `free`, to see what we've got. For that, we compare what we have in Azure to what we just calculated and mark the difference accordingly
```
$cidrRange = 25,25,24,24,24,24,23,25,26,26 | sort
$existingRanges = (Get-AzVirtualNetwork -ResourceGroupName vnet-test |
select name, @{l = "AddressSpace"; e = { $_.AddressSpace.AddressPrefixes }}, ResourceGroupName, Location |
select -expand AddressSpace)
$existingNetworks = $existingRanges | % {[System.Net.IPNetwork]$_}
$nets = $existingRanges
$ret = @()
$cidrRange | % {
$ret = Get-IPRanges -Networks $nets -CIDR $_ -BaseNet "10.172.0.0/16"
$nets = ($ret | select @{l="range"; e = {"$($_.network)/$($_.cidr)"}}).range
}
$ret | % {
if ( -not ($_ -in $existingNetworks)) {$_.IsFree = $true}
}
$ret | ft -AutoSize
```
And this gives us all the necessary information to add a few more networks.
| github_jupyter |
TSG001 - Run azdata copy-logs
=============================
Steps
-----
### Common functions
Define helper functions used in this notebook.
```
# Define `run` function for transient fault handling, suggestions on error, and scrolling updates on Windows
import sys
import os
import re
import json
import platform
import shlex
import shutil
import datetime
from subprocess import Popen, PIPE
from IPython.display import Markdown
retry_hints = {}
error_hints = {}
install_hint = {}
first_run = True
rules = None
def run(cmd, return_output=False, no_output=False, retry_count=0):
"""
Run shell command, stream stdout, print stderr and optionally return output
"""
MAX_RETRIES = 5
output = ""
retry = False
global first_run
global rules
if first_run:
first_run = False
rules = load_rules()
# shlex.split is required on bash and for Windows paths with spaces
#
cmd_actual = shlex.split(cmd)
# Store this (i.e. kubectl, python etc.) to support binary context aware error_hints and retries
#
user_provided_exe_name = cmd_actual[0].lower()
# When running python, use the python in the ADS sandbox ({sys.executable})
#
if cmd.startswith("python "):
cmd_actual[0] = cmd_actual[0].replace("python", sys.executable)
# On Mac, when ADS is not launched from terminal, LC_ALL may not be set, which causes pip installs to fail
# with:
#
# UnicodeDecodeError: 'ascii' codec can't decode byte 0xc5 in position 4969: ordinal not in range(128)
#
# Setting it to a default value of "en_US.UTF-8" enables pip install to complete
#
if platform.system() == "Darwin" and "LC_ALL" not in os.environ:
os.environ["LC_ALL"] = "en_US.UTF-8"
# To aid supportabilty, determine which binary file will actually be executed on the machine
#
which_binary = None
# Special case for CURL on Windows. The version of CURL in Windows System32 does not work to
# get JWT tokens, it returns "(56) Failure when receiving data from the peer". If another instance
# of CURL exists on the machine use that one. (Unfortunately the curl.exe in System32 is almost
# always the first curl.exe in the path, and it can't be uninstalled from System32, so here we
# look for the 2nd installation of CURL in the path)
if platform.system() == "Windows" and cmd.startswith("curl "):
path = os.getenv('PATH')
for p in path.split(os.path.pathsep):
p = os.path.join(p, "curl.exe")
if os.path.exists(p) and os.access(p, os.X_OK):
if p.lower().find("system32") == -1:
cmd_actual[0] = p
which_binary = p
break
# Find the path based location (shutil.which) of the executable that will be run (and display it to aid supportability), this
# seems to be required for .msi installs of azdata.cmd/az.cmd. (otherwise Popen returns FileNotFound)
#
# NOTE: Bash needs cmd to be the list of the space separated values hence shlex.split.
#
if which_binary == None:
which_binary = shutil.which(cmd_actual[0])
if which_binary == None:
if user_provided_exe_name in install_hint and install_hint[user_provided_exe_name] is not None:
display(Markdown(f'HINT: Use [{install_hint[user_provided_exe_name][0]}]({install_hint[user_provided_exe_name][1]}) to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)")
else:
cmd_actual[0] = which_binary
start_time = datetime.datetime.now().replace(microsecond=0)
print(f"START: {cmd} @ {start_time} ({datetime.datetime.utcnow().replace(microsecond=0)} UTC)")
print(f" using: {which_binary} ({platform.system()} {platform.release()} on {platform.machine()})")
print(f" cwd: {os.getcwd()}")
# Command-line tools such as CURL and AZDATA HDFS commands output
# scrolling progress bars, which causes Jupyter to hang forever, to
# workaround this, use no_output=True
#
# Work around a infinite hang when a notebook generates a non-zero return code, break out, and do not wait
#
wait = True
try:
if no_output:
p = Popen(cmd_actual)
else:
p = Popen(cmd_actual, stdout=PIPE, stderr=PIPE, bufsize=1)
with p.stdout:
for line in iter(p.stdout.readline, b''):
line = line.decode()
if return_output:
output = output + line
else:
if cmd.startswith("azdata notebook run"): # Hyperlink the .ipynb file
regex = re.compile(' "(.*)"\: "(.*)"')
match = regex.match(line)
if match:
if match.group(1).find("HTML") != -1:
display(Markdown(f' - "{match.group(1)}": "{match.group(2)}"'))
else:
display(Markdown(f' - "{match.group(1)}": "[{match.group(2)}]({match.group(2)})"'))
wait = False
break # otherwise infinite hang, have not worked out why yet.
else:
print(line, end='')
if rules is not None:
apply_expert_rules(line)
if wait:
p.wait()
except FileNotFoundError as e:
if install_hint is not None:
display(Markdown(f'HINT: Use {install_hint} to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)") from e
exit_code_workaround = 0 # WORKAROUND: azdata hangs on exception from notebook on p.wait()
if not no_output:
for line in iter(p.stderr.readline, b''):
line_decoded = line.decode()
# azdata emits a single empty line to stderr when doing an hdfs cp, don't
# print this empty "ERR:" as it confuses.
#
if line_decoded == "":
continue
print(f"STDERR: {line_decoded}", end='')
if line_decoded.startswith("An exception has occurred") or line_decoded.startswith("ERROR: An error occurred while executing the following cell"):
exit_code_workaround = 1
if user_provided_exe_name in error_hints:
for error_hint in error_hints[user_provided_exe_name]:
if line_decoded.find(error_hint[0]) != -1:
display(Markdown(f'HINT: Use [{error_hint[1]}]({error_hint[2]}) to resolve this issue.'))
if rules is not None:
apply_expert_rules(line_decoded)
if user_provided_exe_name in retry_hints:
for retry_hint in retry_hints[user_provided_exe_name]:
if line_decoded.find(retry_hint) != -1:
if retry_count < MAX_RETRIES:
print(f"RETRY: {retry_count} (due to: {retry_hint})")
retry_count = retry_count + 1
output = run(cmd, return_output=return_output, retry_count=retry_count)
if return_output:
return output
else:
return
elapsed = datetime.datetime.now().replace(microsecond=0) - start_time
# WORKAROUND: We avoid infinite hang above in the `azdata notebook run` failure case, by inferring success (from stdout output), so
# don't wait here, if success known above
#
if wait:
if p.returncode != 0:
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(p.returncode)}.\n')
else:
if exit_code_workaround !=0 :
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(exit_code_workaround)}.\n')
print(f'\nSUCCESS: {elapsed}s elapsed.\n')
if return_output:
return output
def load_json(filename):
with open(filename, encoding="utf8") as json_file:
return json.load(json_file)
def load_rules():
try:
# Load this notebook as json to get access to the expert rules in the notebook metadata.
#
j = load_json("tsg001-copy-logs.ipynb")
except:
pass # If the user has renamed the book, we can't load ourself. NOTE: Is there a way in Jupyter, to know your own filename?
else:
if "metadata" in j and \
"azdata" in j["metadata"] and \
"expert" in j["metadata"]["azdata"] and \
"rules" in j["metadata"]["azdata"]["expert"]:
rules = j["metadata"]["azdata"]["expert"]["rules"]
rules.sort() # Sort rules, so they run in priority order (the [0] element). Lowest value first.
# print (f"EXPERT: There are {len(rules)} rules to evaluate.")
return rules
def apply_expert_rules(line):
global rules
for rule in rules:
# rules that have 9 elements are the injected (output) rules (the ones we want). Rules
# with only 8 elements are the source (input) rules, which are not expanded (i.e. TSG029,
# not ../repair/tsg029-nb-name.ipynb)
if len(rule) == 9:
notebook = rule[1]
cell_type = rule[2]
output_type = rule[3] # i.e. stream or error
output_type_name = rule[4] # i.e. ename or name
output_type_value = rule[5] # i.e. SystemExit or stdout
details_name = rule[6] # i.e. evalue or text
expression = rule[7].replace("\\*", "*") # Something escaped *, and put a \ in front of it!
# print(f"EXPERT: If rule '{expression}' satisfied', run '{notebook}'.")
if re.match(expression, line, re.DOTALL):
# print("EXPERT: MATCH: name = value: '{0}' = '{1}' matched expression '{2}', therefore HINT '{4}'".format(output_type_name, output_type_value, expression, notebook))
match_found = True
display(Markdown(f'HINT: Use [{notebook}]({notebook}) to resolve this issue.'))
print('Common functions defined successfully.')
# Hints for binary (transient fault) retry, (known) error and install guide
#
retry_hints = {'kubectl': ['A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond'], 'azdata': ['Endpoint sql-server-master does not exist', 'Endpoint livy does not exist', 'Failed to get state for cluster', 'Endpoint webhdfs does not exist', 'Adaptive Server is unavailable or does not exist', 'Error: Address already in use']}
error_hints = {'kubectl': [['no such host', 'TSG010 - Get configuration contexts', '../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb'], ['no such host', 'TSG011 - Restart sparkhistory server', '../repair/tsg011-restart-sparkhistory-server.ipynb'], ['No connection could be made because the target machine actively refused it', 'TSG056 - Kubectl fails with No connection could be made because the target machine actively refused it', '../repair/tsg056-kubectl-no-connection-could-be-made.ipynb']], 'azdata': [['azdata login', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['The token is expired', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Reason: Unauthorized', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Max retries exceeded with url: /api/v1/bdc/endpoints', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Look at the controller logs for more details', 'TSG027 - Observe cluster deployment', '../diagnose/tsg027-observe-bdc-create.ipynb'], ['provided port is already allocated', 'TSG062 - Get tail of all previous container logs for pods in BDC namespace', '../log-files/tsg062-tail-bdc-previous-container-logs.ipynb'], ['Create cluster failed since the existing namespace', 'SOP061 - Delete a big data cluster', '../install/sop061-delete-bdc.ipynb'], ['Failed to complete kube config setup', 'TSG067 - Failed to complete kube config setup', '../repair/tsg067-failed-to-complete-kube-config-setup.ipynb'], ['Error processing command: "ApiError', 'TSG110 - Azdata returns ApiError', '../repair/tsg110-azdata-returns-apierror.ipynb'], ['Error processing command: "ControllerError', 'TSG036 - Controller logs', '../log-analyzers/tsg036-get-controller-logs.ipynb'], ['ERROR: 500', 'TSG046 - Knox gateway logs', '../log-analyzers/tsg046-get-knox-logs.ipynb'], ['Data source name not found and no default driver specified', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ["Can't open lib 'ODBC Driver 17 for SQL Server", 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Control plane upgrade failed. Failed to upgrade controller.', 'TSG108 - View the controller upgrade config map', '../diagnose/tsg108-controller-failed-to-upgrade.ipynb']]}
install_hint = {'kubectl': ['SOP036 - Install kubectl command line interface', '../install/sop036-install-kubectl.ipynb'], 'azdata': ['SOP055 - Install azdata command line interface', '../install/sop055-install-azdata.ipynb']}
```
### Get the Kubernetes namespace for the big data cluster
Get the namespace of the big data cluster use the kubectl command line
interface .
NOTE: If there is more than one big data cluster in the target
Kubernetes cluster, then set \[0\] to the correct value for the big data
cluster.
```
# Place Kubernetes namespace name for BDC into 'namespace' variable
try:
namespace = run(f'kubectl get namespace --selector=MSSQL_CLUSTER -o jsonpath={{.items[0].metadata.name}}', return_output=True)
except:
from IPython.display import Markdown
print(f"ERROR: Unable to find a Kubernetes namespace with label 'MSSQL_CLUSTER'. SQL Server Big Data Cluster Kubernetes namespaces contain the label 'MSSQL_CLUSTER'.")
display(Markdown(f'HINT: Use [TSG081 - Get namespaces (Kubernetes)](../monitor-k8s/tsg081-get-kubernetes-namespaces.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [TSG010 - Get configuration contexts](../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [SOP011 - Set kubernetes configuration context](../common/sop011-set-kubernetes-context.ipynb) to resolve this issue.'))
raise
else:
print(f'The SQL Server Big Data Cluster Kubernetes namespace is: {namespace}')
```
### Run copy-logs
```
import os
import tempfile
import shutil
target_folder = os.path.join(tempfile.gettempdir(), "copy-logs", namespace)
if os.path.isdir(target_folder):
shutil.rmtree(target_folder)
```
### View the `--help` options
```
run(f'azdata bdc debug copy-logs --help')
```
### Run the `copy-logs`
NOTES:
1. The –timeout option does not work on Windows
2. Use –skip-compress on Windows if no utility available to uncompress
.tar.gz files.
```
run(f'azdata bdc debug copy-logs --namespace {namespace} --target-folder {target_folder} --exclude-dumps --skip-compress --verbose')
print(f'The logs are available at: {target_folder}')
print('Notebook execution complete.')
```
| github_jupyter |
# CrowdTruth for Multiple Choice Tasks: Relation Extraction
In this tutorial, we will apply CrowdTruth metrics to a **multiple choice** crowdsourcing task for **Relation Extraction** from sentences. The workers were asked to read a sentence with 2 highlighted terms, then pick from a multiple choice list what are the relations expressed between the 2 terms in the sentence. The task was executed on [FigureEight](https://www.figure-eight.com/). For more crowdsourcing annotation task examples, click [here](https://raw.githubusercontent.com/CrowdTruth-core/tutorial/getting_started.md).
To replicate this experiment, the code used to design and implement this crowdsourcing annotation template is available here: [template](https://raw.githubusercontent.com/CrowdTruth/CrowdTruth-core/master/tutorial/templates/Relex-Multiple-Choice/template.html), [css](https://raw.githubusercontent.com/CrowdTruth/CrowdTruth-core/master/tutorial/templates/Relex-Multiple-Choice/template.css), [javascript](https://raw.githubusercontent.com/CrowdTruth/CrowdTruth-core/master/tutorial/templates/Relex-Multiple-Choice/template.js).
This is a screenshot of the task as it appeared to workers:

A sample dataset for this task is available in [this file](https://raw.githubusercontent.com/CrowdTruth/CrowdTruth-core/master/tutorial/data/relex-multiple-choice.csv), containing raw output from the crowd on FigureEight. Download the file and place it in a folder named `data` that has the same root as this notebook. Now you can check your data:
```
import pandas as pd
test_data = pd.read_csv("../data/relex-multiple-choice.csv")
test_data.head()
```
## Declaring a pre-processing configuration
The pre-processing configuration defines how to interpret the raw crowdsourcing input. To do this, we need to define a configuration class. First, we import the default CrowdTruth configuration class:
```
import crowdtruth
from crowdtruth.configuration import DefaultConfig
```
Our test class inherits the default configuration `DefaultConfig`, while also declaring some additional attributes that are specific to the Relation Extraction task:
* **`inputColumns`:** list of input columns from the .csv file with the input data
* **`outputColumns`:** list of output columns from the .csv file with the answers from the workers
* **`annotation_separator`:** string that separates between the crowd annotations in `outputColumns`
* **`open_ended_task`:** boolean variable defining whether the task is open-ended (i.e. the possible crowd annotations are not known beforehand, like in the case of free text input); in the task that we are processing, workers pick the answers from a pre-defined list, therefore the task is not open ended, and this variable is set to `False`
* **`annotation_vector`:** list of possible crowd answers, mandatory to declare when `open_ended_task` is `False`; for our task, this is the list of relations
* **`processJudgments`:** method that defines processing of the raw crowd data; for this task, we process the crowd answers to correspond to the values in `annotation_vector`
The complete configuration class is declared below:
```
class TestConfig(DefaultConfig):
inputColumns = ["sent_id", "term1", "b1", "e1", "term2", "b2", "e2", "sentence"]
outputColumns = ["relations"]
annotation_separator = "\n"
# processing of a closed task
open_ended_task = False
annotation_vector = [
"title", "founded_org", "place_of_birth", "children", "cause_of_death",
"top_member_employee_of_org", "employee_or_member_of", "spouse",
"alternate_names", "subsidiaries", "place_of_death", "schools_attended",
"place_of_headquarters", "charges", "origin", "places_of_residence",
"none"]
def processJudgments(self, judgments):
# pre-process output to match the values in annotation_vector
for col in self.outputColumns:
# transform to lowercase
judgments[col] = judgments[col].apply(lambda x: str(x).lower())
return judgments
```
## Pre-processing the input data
After declaring the configuration of our input file, we are ready to pre-process the crowd data:
```
data, config = crowdtruth.load(
file = "../data/relex-multiple-choice.csv",
config = TestConfig()
)
data['judgments'].head()
```
## Computing the CrowdTruth metrics
The pre-processed data can then be used to calculate the CrowdTruth metrics:
```
results = crowdtruth.run(data, config)
```
`results` is a dict object that contains the quality metrics for sentences, relations and crowd workers.
The **sentence metrics** are stored in `results["units"]`:
```
results["units"].head()
```
The `uqs` column in `results["units"]` contains the **sentence quality scores**, capturing the overall workers agreement over each sentence. Here we plot its histogram:
```
import matplotlib.pyplot as plt
%matplotlib inline
plt.hist(results["units"]["uqs"])
plt.xlabel("Sentence Quality Score")
plt.ylabel("Sentences")
```
The `unit_annotation_score` column in `results["units"]` contains the **sentence-relation scores**, capturing the likelihood that a relation is expressed in a sentence. For each sentence, we store a dictionary mapping each relation to its sentence-relation score.
```
results["units"]["unit_annotation_score"].head()
```
The **worker metrics** are stored in `results["workers"]`:
```
results["workers"].head()
```
The `wqs` columns in `results["workers"]` contains the **worker quality scores**, capturing the overall agreement between one worker and all the other workers.
```
plt.hist(results["workers"]["wqs"])
plt.xlabel("Worker Quality Score")
plt.ylabel("Workers")
```
The **relation metrics** are stored in `results["annotations"]`. The `aqs` column contains the **relation quality scores**, capturing the overall worker agreement over one relation.
```
results["annotations"]
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.