text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
```
from IPython import get_ipython
IS_COLAB = False
if "google.colab" in str(get_ipython()):
# Make sure to go to Runtime > Change runtime type > Hardware Accelerator: GPU
IS_COLAB = True
!pip install fastai --upgrade -q
!pip install wandb
!pip freeze | grep fast
if IS_COLAB:
from google.colab import drive
drive.mount("/gdrive")
%ls /gdrive/MyDrive/
import json
from pathlib import Path
import pandas as pd
import wandb
from fastai.callback.wandb import WandbCallback
from fastai.learner import load_model, save_model
from fastai.metrics import accuracy
from fastai.vision.all import cnn_learner
from fastai.vision.data import ImageDataLoaders
from fastai.vision.models import xresnet
ROOT_PATH = Path("..")
TRAIN = True
if IS_COLAB:
import torch
# Make sure that this path exists
ROOT_PATH = Path("/gdrive/MyDrive/pracds_final")
print(torch.cuda.get_device_name(0))
!wandb login
# Setup y_train, y_test
df = pd.read_csv(ROOT_PATH / "data/raw/metadata.csv")
svc_ids = pd.read_json(ROOT_PATH / "data/raw/song_vs_call.json").squeeze()
svc_df = df.loc[df.id.isin(svc_ids)].copy()
with open(ROOT_PATH / "data/processed/svc_split.json") as svc_split_file:
svc_split = json.load(svc_split_file)
train_ids = svc_split["train_ids"]
test_ids = svc_split["test_ids"]
# Add response variable
type_col = svc_df.type.str.lower().str.replace(" ", "").str.split(",")
filtered_type_col = type_col.apply(lambda l: set(l) - {"call", "song"})
svc_df["label"] = type_col.apply(lambda l: "call" in l).astype(int)
y_df = svc_df.reindex(columns=["id", "pred"]).copy()
y_train, y_test = (
y_df[y_df.id.isin(train_ids)].drop(columns=["id"]).squeeze(),
y_df[y_df.id.isin(test_ids)].drop(columns=["id"]).squeeze(),
)
svc_df["name"] = svc_df.id.astype(str) + ".png"
svc_df["is_valid"] = svc_df.id.isin(test_ids)
image_df = (
svc_df.reindex(columns=["id", "name", "label", "is_valid"]).set_index("id").copy()
)
bs = 128 # Batch size
kwargs = {}
if IS_COLAB:
kwargs["num_workers"] = 0
data = (
# convert_mode is passed on intern|ally to the relevant function that will handle converting the images;
# 'L' results in one color channel
ImageDataLoaders.from_df(
image_df,
folder=ROOT_PATH / "data/raw/sonograms",
valid_col="is_valid",
bs=bs,
# num_works needs to be set to 0 for local evaluation to turn off multiprocessing
**kwargs,
)
)
learn = cnn_learner(data, xresnet.xresnet18, pretrained=True)
# Make sure this path exists on colab
fname = "sono_model.pth"
model_path = (ROOT_PATH / f"models/{fname}").resolve().absolute()
if IS_COLAB and TRAIN:
# Fine tune model
wandb.init(project="sono-model")
learn.fit_one_cycle(1, cbs=WandbCallback())
# GDrive fails when you try to use mkdir
# so we manually call `save_model`
save_path = f"/home/{fname}"
save_model(save_path, learn.model, getattr(learn, "opt", None))
%ls -al /home
from google.colab import files
files.download(save_path)
else:
load_model(model_path, learn.model, learn.opt)
learn.metrics = [accuracy]
train_metrics = learn.validate(ds_idx=0)
print(f"Train Loss: {train_metrics[0]:.4f}")
print(f"Train Accuracy: {train_metrics[1]:.4f}")
val_metrics = learn.validate()
print(f"Validation Loss: {val_metrics[0]:.4f}")
print(f"Validation Accuracy: {val_metrics[1]:.4f}")
# learn.predict(ROOT_PATH / "data/raw/sonograms/1136.png")
```
| github_jupyter |
# Using Experimental Data to Find a Soil's Weight Function
As described in Section 2.3 of the article, finding a given soil's weight functions is achieved through a set of experiments in which we measure soil degradation and "first order reversal curves" (FORCs).
This notebook shows how to process the data from these experiments to find soil's weight function for the case with two input variables: salinity, $C$, and Sodium Adsorption Ratio, SAR:
We note that this code is designed for demonstration purposes and that interested users should download the notebook *xxx* for actual use. We also encourage interested users to contact us directly (isaac.kramer@mail.huji.ac.il and yair.mau@mail.huji.ac.il)
# To use the notebook, please click Cell → Run All.
```
from IPython.display import HTML
HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide();
} else {
$('div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
<form action="javascript:code_toggle()"><input type="submit" value="View/Hide Code"></form>''')
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.gridspec import GridSpec
import scipy.interpolate as interp
from scipy.signal import argrelextrema
from palettable.colorbrewer.sequential import YlGnBu_9
from IPython.display import display_html, Markdown
class app_test(object):
def __init__(self):
# import raw data
self.urlU='https://raw.githubusercontent.com/yairmau/hysteresis-python/master/github_data_u.csv'
self.urlV='https://raw.githubusercontent.com/yairmau/hysteresis-python/master/github_data_v.csv'
self.raw_data_constant_u = pd.read_csv(self.urlU,error_bad_lines=False)
self.raw_data_constant_v = pd.read_csv(self.urlV,error_bad_lines=False)
# grid dimensions
self.hys_per_side = 101 # number of hysterons per side for weights
self.beta_grid_values = np.linspace(0, 100, self.hys_per_side) # beta grid values
self.alpha_grid_values = np.linspace(100, 0, self.hys_per_side) # alpha grid values
self.beta_grid, self.alpha_grid = np.meshgrid(self.beta_grid_values, self.alpha_grid_values)
self.beta_grids = [self.beta_grid, -np.flip(self.beta_grid, axis = 1)]
self.alpha_grids = [self.alpha_grid, -np.flip(self.alpha_grid, axis = 0)]
self.sets = [self.raw_data_constant_u, self.raw_data_constant_v]
self.weights = ["mu", "nu"]
# All of the plots, the 2 figures and the gridSpec are clustered into dictionaries
self.gs = {}
self.allPlots = {}
self.f = {}
#Side functions
def MarkDownSpace(self,n):
s = " "
for i in range(n):
s = s + " "
return s
def DisplaylTables(self,u,v,t1,t2,n1,n2,n3,n4,n5,n6):
self.fig, self.axes = plt.subplots(1, 2)
self.gs.update({"Table":(GridSpec(1, 2,
left=0.1, right=1.8,
bottom=0, top=1,
wspace=1.3, hspace=0.4))})
self.table1_axis = plt.subplot(self.gs["Table"][0, 0])
self.table2_axis = plt.subplot(self.gs["Table"][0, 1])
self.np_u = np.round(u[:5].to_numpy() , 2)
self.np_v = np.round(v[:5].to_numpy() , 2)
self.table1_axis.table(cellText = self.np_u,colLoc = 'center' ,cellLoc='center',loc='center',colWidths=[0.5,0.5,0.5]
,colLabels=[n1,n2,n3],
colColours=['gray','gray','gray'],bbox=[0, 0, 1, 1] )
self.table1_axis.axis("off")
self.table1_axis.set_title(str(t1)+"\n")
self.table2_axis.table(cellText = self.np_v, colLoc = 'center' , cellLoc='center',loc='center',colWidths=[0.5,0.5,0.5]
,colLabels=[n4,n5,n6]
,colColours=['gray','gray','gray'],bbox=[0, 0, 1, 1] )
self.table2_axis.axis("off")
self.table2_axis.set_title(str(t2)+"\n")
def draw_arrow(self,ax, start, end):
ax.annotate('', xy=end, xytext=start, xycoords='data', textcoords='data',
arrowprops=dict(headwidth=4.0, headlength=4.0, width=0.2,facecolor = "black", linewidth = 0.5)
,zorder=0)
# Variables set up functions which are necessary for both FORC graphing and interpolation
def ReversalCurveVariableSetUp1(self):
self.inputs = self.subset.input_var.to_numpy()
self.outputs = self.subset.output_var.to_numpy()
# index of local minimums (beta values from which reversal curves start)
self.minimums_index = argrelextrema(self.inputs, np.less)[0]
self.minimums_index = np.concatenate((np.array([0]), self.minimums_index)) # last value
# index of local maximum (alpha values at which reversal curves stop)
self.maximums_index = argrelextrema(self.inputs, np.greater)[0]
self.maximums_index = np.concatenate((self.maximums_index, np.array([len(self.inputs) - 1]))) # last value
# empty arrays to which we will add alpha/beta values used in experiment and output
self.betas_reversal = np.empty(0) # beta values
self.alphas_reversal = np.empty(0) # alpha values
self.F_values = np.empty(0) # output values
def ReversalCurveVariableSetUp2(self,i):
# number of points until local maximum
self.length = self.maximums_index[i] - self.minimums_index[i] + 1
# beta_0 value
self.beta_0 = np.full((self.length), self.inputs[self.minimums_index[i]]) # does not change along reversal curve
self.betas_reversal = np.concatenate((self.betas_reversal, self.beta_0)) # concentate to existing values
# alpha values
self.alphas = self.inputs[self.minimums_index[i]:self.minimums_index[i]
+ self.length] # move along reversal curve from local min to local max
self.alphas_reversal = np.concatenate((self.alphas_reversal, self.alphas))
# output values
self.reversal_curve_values = self.outputs[self.minimums_index[i]:self.minimums_index[i] + self.length]
# move along reversal curves (i.e., at beta_0, alpha)
self.F_alpha_beta_v = 0.5 * (self.reversal_curve_values - self.reversal_curve_values.min())
self.F_values = np.concatenate((self.F_values, self.F_alpha_beta_v)) # concentate to existing values
def PlotFORC(self , n1 , n2):
# Plotting
self.f.update({"FORC":plt.figure(1, figsize=(7,10))})
self.gs.update({"GsCurve1":(GridSpec(len(self.sets[0].weight_variable.unique()), 1,
left=0.1, right=0.8,
bottom=0, top=1,
wspace=0.0, hspace=0.4))})
self.gs.update({"GsCurve2":(GridSpec(len(self.sets[1].weight_variable.unique()), 1,
left=1.1, right=1.8,
bottom=0.0, top=1,
wspace=0.0, hspace=0.4))})
for jj in range(len(self.sets)):
self.constants = self.sets[jj].weight_variable.unique()
self.beta_grid = self.beta_grids[jj]
self.alpha_grid = self.alpha_grids[jj]
# Creating subplots
for n in range(len(self.constants)):
self.allPlots.update({"Data"+str(jj+1)+"Curve"+str(n):plt.subplot(self.gs["GsCurve"+str(jj+1)][n, 0])})
# Calculating Data for FORC
# The max list is created in order to determinate the y limit of the graph
for ii in range(len(self.constants)):
# data corresponding to specific u value
self.subset = self.sets[jj].loc[self.sets[jj]['weight_variable'] == self.constants[ii]]
self.ReversalCurveVariableSetUp1()
self.max_list_FORC = []
# loop for each beta_0 to isolate values associated with reversal curves
for i in range(len(self.minimums_index)):
self.ReversalCurveVariableSetUp2(i)
# plot reversal curve
self.allPlots["Data"+str(jj+1)+"Curve"+str(ii)].plot(self.alphas, self.reversal_curve_values)
self.max_list_FORC.append(self.reversal_curve_values.max())
# Graphs adjustments
self.FORC_max = max(self.max_list_FORC)
self.FORC_plot = self.allPlots["Data"+str(jj+1)+"Curve"+str(ii)]
self.FORC_plot.spines['right'].set_visible(False)
self.FORC_plot.spines['top'].set_visible(False)
self.FORC_plot.spines['left'].set_visible(False)
self.FORC_plot.spines['bottom'].set_visible(False)
if jj==0:
self.FORC_plot.set_xlim([0,106])
self.FORC_plot.set_ylim([0,self.FORC_max*1.2])
self.draw_arrow(self.FORC_plot, (0,0),(106, 0))
self.FORC_plot.annotate('', xy=(0, 0), xycoords=('data'),
xytext=(0, self.FORC_max*1.2), textcoords='data',
ha='left', va='center',
arrowprops=dict(arrowstyle='<|-', fc='black'),zorder=2)
self.FORC_plot.text(-0.02, self.FORC_max*1.3, r"$rK_s$")
self.FORC_plot.text(108, 0, "Salinity")
self.FORC_plot.set_title(n1[ii])
if jj==1:
self.FORC_plot.set_xlim([-100,6])
self.FORC_plot.set_ylim([0,self.FORC_max*1.2])
self.draw_arrow(self.FORC_plot, (-100,0),(6, 0))
self.FORC_plot.annotate('', xy=(-100, 0), xycoords=('data'),
xytext=(-100, self.FORC_max*1.2), textcoords='data',
ha='left', va='center',
arrowprops=dict(arrowstyle='<|-', fc='black'),zorder=2)
self.FORC_plot.text(-100.02, self.FORC_max*1.3, r"$rK_s$")
self.FORC_plot.text(8, 0, "SAR")
self.FORC_plot.set_title(n2[ii])
def InterpolateAndPlotWeight(self,n1,n2):
# Plotting
self.f.update({"Weight":plt.figure(2, figsize=(7,10))})
self.gs.update({"GsWeight1":(GridSpec(len(self.sets[0].weight_variable.unique()), 1,
left=0.1, right=0.8,
bottom=0, top=1,
wspace=0.0, hspace=0.4))})
self.gs.update({"GsWeight2":(GridSpec(len(self.sets[1].weight_variable.unique()), 1,
left=1.1, right=1.8,
bottom=0.0, top=1,
wspace=0.0, hspace=0.4))})
self.gs.update({"ColorbarGs":(GridSpec(1, 1,
left=1.9, right=2,
bottom=0.8, top=1,
wspace=0.0, hspace=0.4))})
self.allPlots.update({"Colorbar":plt.subplot(self.gs["ColorbarGs"][0, 0])})
for jj in range(len(self.sets)):
self.constants = self.sets[jj].weight_variable.unique()
self.beta_grid = self.beta_grids[jj]
self.alpha_grid = self.alpha_grids[jj]
for g in range(len(self.constants)):
self.allPlots.update({"Data"+str(jj+1)+"Weight"+str(g):plt.subplot(self.gs["GsWeight"+str(jj+1)][g, 0])})
self.W_plot = self.allPlots["Data"+str(jj+1)+"Weight"+str(g)]
self.W_plot.set_yticklabels([])
self.W_plot.set_xticklabels([])
self.W_plot.set_aspect('equal', adjustable='box')
self.W_plot.axis('off')
if jj==0:
self.W_plot.set_title(n1[g])
self.W_plot.set_xlim([0,115])
self.W_plot.set_ylim([0,115])
self.W_plot.annotate('', xy=(0, 0), xycoords=('data'),
xytext=(0, 110), textcoords='data',
ha='left', va='center',
arrowprops=dict(arrowstyle='<|-', fc='black'),zorder=2)
#self.W_plot.axvline(x=0,ymin=0.005,ymax=0.01,c="black",linewidth=1,zorder=10, clip_on=False)
self.draw_arrow(self.W_plot, (0,0),(110, 0))
self.W_plot.text(113, 0, "\u03B2")
self.W_plot.text(0, 113, "\u03B1")
if jj==1:
self.W_plot.set_title(n2[g])
self.W_plot.set_xlim([-100,5])
self.W_plot.set_ylim([-100,5])
self.W_plot.annotate('', xy=(-100, -100), xycoords=('data'),
xytext=(-100, 10), textcoords='data',
ha='left', va='center',
arrowprops=dict(arrowstyle='<|-', fc='black'),zorder=2)
self.draw_arrow(self.W_plot, (-100,-100),(0, -100))
self.W_plot.text(13, -100, "\u03B2")
self.W_plot.text(-100, 13, "\u03B1")
#self.allPlots.update({"Colorbar":plt.subplot(self.gs["ColorbarGs"][0, 0])})
# Setting up once again variables for interpolation
self.F_grid = np.zeros((self.hys_per_side, self.hys_per_side, len(self.constants)))# F/G values during reversal curves
self.betas_int = np.empty(0) # beta values for interpolation
self.alphas_int = np.empty(0) # alpha values for interpolation
self.u_int = np.empty(0) # output values for interpolation
self.F_int = np.empty(0) # F values for interpolation
for ii in range(len(self.constants)):
self.subset = self.sets[jj].loc[self.sets[jj]['weight_variable'] == self.constants[ii]]
self.ReversalCurveVariableSetUp1()
for i in range(len(self.minimums_index)):
self.ReversalCurveVariableSetUp2(i)
# Interpolation
self.zfun_smooth_rbf = interp.Rbf(self.betas_reversal, self.alphas_reversal, self.F_values,
function='cubic',smooth=0)
self.z_dense_smooth_rbf = self.zfun_smooth_rbf(self.beta_grid, self.alpha_grid)
self.F_grid = np.where(self.alpha_grid >= self.beta_grid, self.z_dense_smooth_rbf, np.nan)
# calculate d^2 f_{alpha,beta} and weights
self.step = self.beta_grid_values[1]
self.matrix1 = self.F_grid[:-2, 2:]
self.matrix2 = self.F_grid[2:, 2:]
self.matrix3 = self.F_grid[:-2, :-2]
self.matrix4 = self.F_grid[2:, :-2]
self.weights_grid = (self.matrix3 + self.matrix2 - self.matrix1 - self.matrix4) / (4 * self.step ** 2)
# add rows/columns containing nan to side to keep old dimensions
self.newrow = np.empty((self.hys_per_side - 2))
self.newcol = np.empty((self.hys_per_side, 1))
self.newrow[:] = np.nan
self.newcol[:] = np.nan
self.weights_grid = np.vstack([self.newrow, self.weights_grid, self.newrow])
self.weights_grid = np.hstack([self.newcol, self.weights_grid, self.newcol])
# remove values where interpolating created a negative value
with np.errstate(invalid='ignore'):
self.weights_grid = np.where(self.weights_grid < 0, 0, self.weights_grid)
# reinterpolate using 'nearest' to fill values on border
self.array = np.ma.masked_invalid(self.weights_grid)
self.beta_masked = self.beta_grid[~self.array.mask]
self.alpha_masked = self.alpha_grid[~self.array.mask]
self.newarr = self.array[~self.array.mask]
self.weights_grid = interp.griddata((self.beta_masked, self.alpha_masked), self.newarr.ravel(),
(self.beta_grid, self.alpha_grid),method='nearest')
# eliminate values outside preisach triangle
self.weights_grid = np.where(self.alpha_grid >= self.beta_grid, self.weights_grid, np.nan)
# flatten interpolated values so that they can be used later for 3-D interpolation
self.beta_flat = self.beta_grid.flatten()
self.alpha_flat = self.alpha_grid.flatten()
self.u_flat = np.full((self.hys_per_side * self.hys_per_side), self.constants[ii])
self.F_flat = self.weights_grid.flatten()
# concatenate with existing values
self.betas_int = np.concatenate((self.betas_int, self.beta_flat))
self.alphas_int = np.concatenate((self.alphas_int, self.alpha_flat))
self.u_int = np.concatenate((self.u_int, self.u_flat))
self.F_int = np.concatenate((self.F_int, self.F_flat))
# plot weight function including a colorbar
self.W_plot = self.allPlots["Data"+str(jj+1)+"Weight"+str(ii)]
self.scatter = self.W_plot.scatter(self.beta_grid, self.alpha_grid,
c=self.weights_grid, s=3,cmap=YlGnBu_9.mpl_colormap,
vmin=np.nanmin(self.weights_grid),
vmax=np.nanmax(self.weights_grid))
if jj == 0 and ii == 0:
self.cb = self.f["Weight"].colorbar(self.scatter,ax=self.allPlots["Colorbar"])
self.cb.set_ticks([self.cb.vmin,self.cb.vmax])
self.cb.set_ticklabels(["Weight = 0.0","Weight = 1.0"])
self.cb.ax.tick_params(labelsize=8)
self.allPlots["Colorbar"].axis("off")
def SaveArray(self):
self.points = (self.betas_int, self.alphas_int, self.u_int)
self.beta_cube, self.alpha_cube, self.u_cube = np.meshgrid(self.beta_grid_values, self.alpha_grid_values,
self.beta_grid_values)
self.mu = interp.griddata(self.points, self.F_int, (self.beta_cube, self.alpha_cube, self.u_cube), method='linear')
np.save('experimental_data/'+weights[jj]+'.npy', mu)
```
## 1. Import raw data
The imported data should be stored in two .csv files. The first .csv file contains data from the FORCs in which SAR was held constant. The second .csv file should contain data related to the FORCs in which salinity, $C$ was held constant. Each dataset is stored in a Pandas DataFrame, the headers of which are shown below.
```
A = app_test()
A.DisplaylTables(A.raw_data_constant_u,A.raw_data_constant_v,"Data set 1: SAR Constant","Data set 2: Salinity Constant","Salinity","Hyd. Cond.","SAR","SAR","Hyd. Cond.","Salinity")
```
## 2. Display FORCs
We use the imported data to reconstruct the FORCs.
```
A.PlotFORC([r"SAR = 0 mmol$_c^{1/2}$ L$^{-1/2}$", r"SAR = 5 mmol$_c^{1/2}$ L$^{-1/2}$", r"SAR 10 mmol$_c^{1/2}$ L$^{-1/2}$"],[r"Salinity = 0 mmol$_c$ L$^{-1}$",r"Salinity = 20 mmol$_c$ L$^{-1}$",r"Salinity = 50 mmol$_c$ L$^{-1}$"])
```
## 3. FORCs are used to calculate weight functions
We use the data from the FORCs to calculate a weight function for each of the known SAR and salinity values, respectively. This is done according to the methodology described in section 5 and by interpolating the measured data across the $alpha,beta$-plane.
```
A.InterpolateAndPlotWeight([r"SAR = 0 mmol$_c^{1/2}$ L$^{-1/2}$", r"SAR = 5 mmol$_c^{1/2}$ L$^{-1/2}$", r"SAR 10 mmol$_c^{1/2}$ L$^{-1/2}$"],[r"Salinity = 0 mmol$_c$ L$^{-1}$",r"Salinity = 20 mmol$_c$ L$^{-1}$",r"Salinity = 50 mmol$_c$ L$^{-1}$"])
```
## 4. Interpolate for all SAR and Salinity Values
The final step is to interpolate from the known SAR and salinity values, so that we have a weight function for all SAR and salinity values. This step is not performed in this notebook, but is included in *xxxx*.
| github_jupyter |
# SciPy - Library of scientific algorithms for Python
Adapted from a lecture by J.R. Johansson (jrjohansson at gmail.com)
The original version of this [IPython notebook](http://ipython.org/notebook.html) lecture is available at [http://github.com/jrjohansson/scientific-python-lectures](http://github.com/jrjohansson/scientific-python-lectures).
The other notebooks in this lecture series are indexed at [http://jrjohansson.github.io](http://jrjohansson.github.io).
```
# what is this line all about? Answer in lecture 4
%matplotlib inline
import matplotlib.pyplot as plt
from IPython.display import Image
```
## Introduction
The SciPy framework builds on top of the low-level NumPy framework for multidimensional arrays, and provides a large number of higher-level scientific algorithms. Some of the topics that SciPy covers are:
* Special functions ([scipy.special](http://docs.scipy.org/doc/scipy/reference/special.html))
* Integration ([scipy.integrate](http://docs.scipy.org/doc/scipy/reference/integrate.html))
* Optimization ([scipy.optimize](http://docs.scipy.org/doc/scipy/reference/optimize.html))
* Interpolation ([scipy.interpolate](http://docs.scipy.org/doc/scipy/reference/interpolate.html))
* Fourier Transforms ([scipy.fftpack](http://docs.scipy.org/doc/scipy/reference/fftpack.html))
* Signal Processing ([scipy.signal](http://docs.scipy.org/doc/scipy/reference/signal.html))
* Linear Algebra ([scipy.linalg](http://docs.scipy.org/doc/scipy/reference/linalg.html))
* Sparse Eigenvalue Problems ([scipy.sparse](http://docs.scipy.org/doc/scipy/reference/sparse.html))
* Statistics ([scipy.stats](http://docs.scipy.org/doc/scipy/reference/stats.html))
* Multi-dimensional image processing ([scipy.ndimage](http://docs.scipy.org/doc/scipy/reference/ndimage.html))
* File IO ([scipy.io](http://docs.scipy.org/doc/scipy/reference/io.html))
Each of these submodules provides a number of functions and classes that can be used to solve problems in their respective topics.
In this lecture we will look at how to use some of these subpackages.
To access the SciPy package in a Python program, we start by importing everything from the `scipy` module.
```
from scipy import *
```
If we only need to use part of the SciPy framework we can selectively include only those modules we are interested in. For example, to include the linear algebra package under the name `la`, we can do:
```
import scipy.linalg as la
```
## Special functions
A large number of mathematical special functions are important for many computional physics problems. SciPy provides implementations of a very extensive set of special functions. For details, see the list of functions in the reference documention at http://docs.scipy.org/doc/scipy/reference/special.html#module-scipy.special.
To demonstrate the typical usage of special functions we will look in more detail at the Bessel functions:
```
#
# The scipy.special module includes a large number of Bessel-functions
# Here we will use the functions jn and yn, which are the Bessel functions
# of the first and second kind and real-valued order. We also include the
# function jn_zeros and yn_zeros that gives the zeroes of the functions jn
# and yn.
#
from scipy.special import jn, yn, jn_zeros, yn_zeros
n = 0 # order
x = 0.0
# Bessel function of first kind
print "J_%d(%f) = %f" % (n, x, jn(n, x))
x = 1.0
# Bessel function of second kind
print "Y_%d(%f) = %f" % (n, x, yn(n, x))
x = linspace(0, 10, 100)
fig, ax = plt.subplots()
for n in range(4):
ax.plot(x, jn(n, x), label=r"$J_%d(x)$" % n)
ax.legend();
# zeros of Bessel functions
n = 0 # order
m = 4 # number of roots to compute
jn_zeros(n, m)
```
## Integration
### Numerical integration: quadrature
Numerical evaluation of a function of the type
$\displaystyle \int_a^b f(x) dx$
is called *numerical quadrature*, or simply *quadature*. SciPy provides a series of functions for different kind of quadrature, for example the `quad`, `dblquad` and `tplquad` for single, double and triple integrals, respectively.
```
from scipy.integrate import quad, dblquad, tplquad
```
The `quad` function takes a large number of optional arguments, which can be used to fine-tune the behaviour of the function (try `help(quad)` for details).
The basic usage is as follows:
```
# define a simple function for the integrand
def f(x):
return x
x_lower = 0 # the lower limit of x
x_upper = 1 # the upper limit of x
val, abserr = quad(f, x_lower, x_upper)
print "integral value =", val, ", absolute error =", abserr
```
If we need to pass extra arguments to integrand function we can use the `args` keyword argument:
```
def integrand(x, n):
"""
Bessel function of first kind and order n.
"""
return jn(n, x)
x_lower = 0 # the lower limit of x
x_upper = 10 # the upper limit of x
val, abserr = quad(integrand, x_lower, x_upper, args=(3,))
print val, abserr
```
For simple functions we can use a lambda function (name-less function) instead of explicitly defining a function for the integrand:
```
val, abserr = quad(lambda x: exp(-x ** 2), -Inf, Inf)
print "numerical =", val, abserr
analytical = sqrt(pi)
print "analytical =", analytical
```
As show in the example above, we can also use 'Inf' or '-Inf' as integral limits.
Higher-dimensional integration works in the same way:
```
def integrand(x, y):
return exp(-x**2-y**2)
x_lower = 0
x_upper = 10
y_lower = 0
y_upper = 10
val, abserr = dblquad(integrand, x_lower, x_upper, lambda x : y_lower, lambda x: y_upper)
print val, abserr
```
Note how we had to pass lambda functions for the limits for the y integration, since these in general can be functions of x.
## Ordinary differential equations (ODEs)
SciPy provides two different ways to solve ODEs: An API based on the function `odeint`, and object-oriented API based on the class `ode`. Usually `odeint` is easier to get started with, but the `ode` class offers some finer level of control.
Here we will use the `odeint` functions. For more information about the class `ode`, try `help(ode)`. It does pretty much the same thing as `odeint`, but in an object-oriented fashion.
To use `odeint`, first import it from the `scipy.integrate` module
```
from scipy.integrate import odeint, ode
```
A system of ODEs are usually formulated on standard form before it is attacked numerically. The standard form is:
$y' = f(y, t)$
where
$y = [y_1(t), y_2(t), ..., y_n(t)]$
and $f$ is some function that gives the derivatives of the function $y_i(t)$. To solve an ODE we need to know the function $f$ and an initial condition, $y(0)$.
Note that higher-order ODEs can always be written in this form by introducing new variables for the intermediate derivatives.
Once we have defined the Python function `f` and array `y_0` (that is $f$ and $y(0)$ in the mathematical formulation), we can use the `odeint` function as:
y_t = odeint(f, y_0, t)
where `t` is and array with time-coordinates for which to solve the ODE problem. `y_t` is an array with one row for each point in time in `t`, where each column corresponds to a solution `y_i(t)` at that point in time.
We will see how we can implement `f` and `y_0` in Python code in the examples below.
#### Example: double pendulum
Let's consider a physical example: The double compound pendulum, described in some detail here: http://en.wikipedia.org/wiki/Double_pendulum
```
Image(url='http://upload.wikimedia.org/wikipedia/commons/c/c9/Double-compound-pendulum-dimensioned.svg')
```
The equations of motion of the pendulum are given on the wiki page:
${\dot \theta_1} = \frac{6}{m\ell^2} \frac{ 2 p_{\theta_1} - 3 \cos(\theta_1-\theta_2) p_{\theta_2}}{16 - 9 \cos^2(\theta_1-\theta_2)}$
${\dot \theta_2} = \frac{6}{m\ell^2} \frac{ 8 p_{\theta_2} - 3 \cos(\theta_1-\theta_2) p_{\theta_1}}{16 - 9 \cos^2(\theta_1-\theta_2)}.$
${\dot p_{\theta_1}} = -\frac{1}{2} m \ell^2 \left [ {\dot \theta_1} {\dot \theta_2} \sin (\theta_1-\theta_2) + 3 \frac{g}{\ell} \sin \theta_1 \right ]$
${\dot p_{\theta_2}} = -\frac{1}{2} m \ell^2 \left [ -{\dot \theta_1} {\dot \theta_2} \sin (\theta_1-\theta_2) + \frac{g}{\ell} \sin \theta_2 \right]$
To make the Python code simpler to follow, let's introduce new variable names and the vector notation: $x = [\theta_1, \theta_2, p_{\theta_1}, p_{\theta_2}]$
${\dot x_1} = \frac{6}{m\ell^2} \frac{ 2 x_3 - 3 \cos(x_1-x_2) x_4}{16 - 9 \cos^2(x_1-x_2)}$
${\dot x_2} = \frac{6}{m\ell^2} \frac{ 8 x_4 - 3 \cos(x_1-x_2) x_3}{16 - 9 \cos^2(x_1-x_2)}$
${\dot x_3} = -\frac{1}{2} m \ell^2 \left [ {\dot x_1} {\dot x_2} \sin (x_1-x_2) + 3 \frac{g}{\ell} \sin x_1 \right ]$
${\dot x_4} = -\frac{1}{2} m \ell^2 \left [ -{\dot x_1} {\dot x_2} \sin (x_1-x_2) + \frac{g}{\ell} \sin x_2 \right]$
```
g = 9.82
L = 0.5
m = 0.1
def dx(x, t):
"""
The right-hand side of the pendulum ODE
"""
x1, x2, x3, x4 = x[0], x[1], x[2], x[3]
dx1 = 6.0/(m*L**2) * (2 * x3 - 3 * cos(x1-x2) * x4)/(16 - 9 * cos(x1-x2)**2)
dx2 = 6.0/(m*L**2) * (8 * x4 - 3 * cos(x1-x2) * x3)/(16 - 9 * cos(x1-x2)**2)
dx3 = -0.5 * m * L**2 * ( dx1 * dx2 * sin(x1-x2) + 3 * (g/L) * sin(x1))
dx4 = -0.5 * m * L**2 * (-dx1 * dx2 * sin(x1-x2) + (g/L) * sin(x2))
return [dx1, dx2, dx3, dx4]
# choose an initial state
x0 = [pi/4, pi/2, 0, 0]
# time coodinate to solve the ODE for: from 0 to 10 seconds
t = linspace(0, 10, 250)
# solve the ODE problem
x = odeint(dx, x0, t)
# plot the angles as a function of time
fig, axes = plt.subplots(1,2, figsize=(12,4))
axes[0].plot(t, x[:, 0], 'r', label="theta1")
axes[0].plot(t, x[:, 1], 'b', label="theta2")
x1 = + L * sin(x[:, 0])
y1 = - L * cos(x[:, 0])
x2 = x1 + L * sin(x[:, 1])
y2 = y1 - L * cos(x[:, 1])
axes[1].plot(x1, y1, 'r', label="pendulum1")
axes[1].plot(x2, y2, 'b', label="pendulum2")
axes[1].set_ylim([-1, 0])
axes[1].set_xlim([1, -1]);
```
Simple annimation of the pendulum motion. We will see how to make better animation in Lecture 4.
```
from IPython.display import display, clear_output
import time
fig, ax = plt.subplots(figsize=(4,4))
for t_idx, tt in enumerate(t[:200]):
x1 = + L * sin(x[t_idx, 0])
y1 = - L * cos(x[t_idx, 0])
x2 = x1 + L * sin(x[t_idx, 1])
y2 = y1 - L * cos(x[t_idx, 1])
ax.cla()
ax.plot([0, x1], [0, y1], 'r.-')
ax.plot([x1, x2], [y1, y2], 'b.-')
ax.set_ylim([-1.5, 0.5])
ax.set_xlim([1, -1])
clear_output()
display(fig)
time.sleep(0.1)
```
#### Example: Damped harmonic oscillator
ODE problems are important in computational physics, so we will look at one more example: the damped harmonic oscillation. This problem is well described on the wiki page: http://en.wikipedia.org/wiki/Damping
The equation of motion for the damped oscillator is:
$\displaystyle \frac{\mathrm{d}^2x}{\mathrm{d}t^2} + 2\zeta\omega_0\frac{\mathrm{d}x}{\mathrm{d}t} + \omega^2_0 x = 0$
where $x$ is the position of the oscillator, $\omega_0$ is the frequency, and $\zeta$ is the damping ratio. To write this second-order ODE on standard form we introduce $p = \frac{\mathrm{d}x}{\mathrm{d}t}$:
$\displaystyle \frac{\mathrm{d}p}{\mathrm{d}t} = - 2\zeta\omega_0 p - \omega^2_0 x$
$\displaystyle \frac{\mathrm{d}x}{\mathrm{d}t} = p$
In the implementation of this example we will add extra arguments to the RHS function for the ODE, rather than using global variables as we did in the previous example. As a consequence of the extra arguments to the RHS, we need to pass an keyword argument `args` to the `odeint` function:
```
def dy(y, t, zeta, w0):
"""
The right-hand side of the damped oscillator ODE
"""
x, p = y[0], y[1]
dx = p
dp = -2 * zeta * w0 * p - w0**2 * x
return [dx, dp]
# initial state:
y0 = [1.0, 0.0]
# time coodinate to solve the ODE for
t = linspace(0, 10, 1000)
w0 = 2*pi*1.0
# solve the ODE problem for three different values of the damping ratio
y1 = odeint(dy, y0, t, args=(0.0, w0)) # undamped
y2 = odeint(dy, y0, t, args=(0.2, w0)) # under damped
y3 = odeint(dy, y0, t, args=(1.0, w0)) # critial damping
y4 = odeint(dy, y0, t, args=(5.0, w0)) # over damped
fig, ax = plt.subplots()
ax.plot(t, y1[:,0], 'k', label="undamped", linewidth=0.25)
ax.plot(t, y2[:,0], 'r', label="under damped")
ax.plot(t, y3[:,0], 'b', label=r"critical damping")
ax.plot(t, y4[:,0], 'g', label="over damped")
ax.legend();
```
## Fourier transform
Fourier transforms are one of the universal tools in computational physics, which appear over and over again in different contexts. SciPy provides functions for accessing the classic [FFTPACK](http://www.netlib.org/fftpack/) library from NetLib, which is an efficient and well tested FFT library written in FORTRAN. The SciPy API has a few additional convenience functions, but overall the API is closely related to the original FORTRAN library.
To use the `fftpack` module in a python program, include it using:
```
from numpy.fft import fftfreq
from scipy.fftpack import *
```
To demonstrate how to do a fast Fourier transform with SciPy, let's look at the FFT of the solution to the damped oscillator from the previous section:
```
N = len(t)
dt = t[1]-t[0]
# calculate the fast fourier transform
# y2 is the solution to the under-damped oscillator from the previous section
F = fft(y2[:,0])
# calculate the frequencies for the components in F
w = fftfreq(N, dt)
fig, ax = plt.subplots(figsize=(9,3))
ax.plot(w, abs(F));
```
Since the signal is real, the spectrum is symmetric. We therefore only need to plot the part that corresponds to the postive frequencies. To extract that part of the `w` and `F` we can use some of the indexing tricks for NumPy arrays that we saw in Lecture 2:
```
indices = where(w > 0) # select only indices for elements that corresponds to positive frequencies
w_pos = w[indices]
F_pos = F[indices]
fig, ax = plt.subplots(figsize=(9,3))
ax.plot(w_pos, abs(F_pos))
ax.set_xlim(0, 5);
```
As expected, we now see a peak in the spectrum that is centered around 1, which is the frequency we used in the damped oscillator example.
## Linear algebra
The linear algebra module contains a lot of matrix related functions, including linear equation solving, eigenvalue solvers, matrix functions (for example matrix-exponentiation), a number of different decompositions (SVD, LU, cholesky), etc.
Detailed documetation is available at: http://docs.scipy.org/doc/scipy/reference/linalg.html
Here we will look at how to use some of these functions:
### Linear equation systems
Linear equation systems on the matrix form
$A x = b$
where $A$ is a matrix and $x,b$ are vectors can be solved like:
```
from scipy.linalg import *
A = array([[1,2,3], [4,5,6], [7,8,9]])
b = array([1,2,3])
x = solve(A, b)
x
# check
dot(A, x) - b
```
We can also do the same with
$A X = B$
where $A, B, X$ are matrices:
```
A = rand(3,3)
B = rand(3,3)
X = solve(A, B)
X
# check
norm(dot(A, X) - B)
```
### Eigenvalues and eigenvectors
The eigenvalue problem for a matrix $A$:
$\displaystyle A v_n = \lambda_n v_n$
where $v_n$ is the $n$th eigenvector and $\lambda_n$ is the $n$th eigenvalue.
To calculate eigenvalues of a matrix, use the `eigvals` and for calculating both eigenvalues and eigenvectors, use the function `eig`:
```
evals = eigvals(A)
evals
evals, evecs = eig(A)
evals
evecs
```
The eigenvectors corresponding to the $n$th eigenvalue (stored in `evals[n]`) is the $n$th *column* in `evecs`, i.e., `evecs[:,n]`. To verify this, let's try mutiplying eigenvectors with the matrix and compare to the product of the eigenvector and the eigenvalue:
```
n = 1
norm(dot(A, evecs[:,n]) - evals[n] * evecs[:,n])
```
There are also more specialized eigensolvers, like the `eigh` for Hermitian matrices.
### Matrix operations
```
# the matrix inverse
inv(A)
# determinant
det(A)
# norms of various orders
norm(A, ord=2), norm(A, ord=Inf)
```
### Sparse matrices
Sparse matrices are often useful in numerical simulations dealing with large systems, if the problem can be described in matrix form where the matrices or vectors mostly contains zeros. Scipy has a good support for sparse matrices, with basic linear algebra operations (such as equation solving, eigenvalue calculations, etc).
There are many possible strategies for storing sparse matrices in an efficient way. Some of the most common are the so-called coordinate form (COO), list of list (LIL) form, and compressed-sparse column CSC (and row, CSR). Each format has some advantanges and disadvantages. Most computational algorithms (equation solving, matrix-matrix multiplication, etc) can be efficiently implemented using CSR or CSC formats, but they are not so intuitive and not so easy to initialize. So often a sparse matrix is initially created in COO or LIL format (where we can efficiently add elements to the sparse matrix data), and then converted to CSC or CSR before used in real calcalations.
For more information about these sparse formats, see e.g. http://en.wikipedia.org/wiki/Sparse_matrix
When we create a sparse matrix we have to choose which format it should be stored in. For example,
```
from scipy.sparse import *
# dense matrix
M = array([[1,0,0,0], [0,3,0,0], [0,1,1,0], [1,0,0,1]]); M
# convert from dense to sparse
A = csr_matrix(M); A
# convert from sparse to dense
A.todense()
```
More efficient way to create sparse matrices: create an empty matrix and populate with using matrix indexing (avoids creating a potentially large dense matrix)
```
A = lil_matrix((4,4)) # empty 4x4 sparse matrix
A[0,0] = 1
A[1,1] = 3
A[2,2] = A[2,1] = 1
A[3,3] = A[3,0] = 1
A
A.todense()
```
Converting between different sparse matrix formats:
```
A
A = csr_matrix(A); A
A = csc_matrix(A); A
```
We can compute with sparse matrices like with dense matrices:
```
A.todense()
(A * A).todense()
A.todense()
A.dot(A).todense()
v = array([1,2,3,4])[:,newaxis]; v
# sparse matrix - dense vector multiplication
A * v
# same result with dense matrix - dense vector multiplcation
A.todense() * v
```
## Optimization
Optimization (finding minima or maxima of a function) is a large field in mathematics, and optimization of complicated functions or in many variables can be rather involved. Here we will only look at a few very simple cases. For a more detailed introduction to optimization with SciPy see: http://scipy-lectures.github.com/advanced/mathematical_optimization/index.html
To use the optimization module in scipy first include the `optimize` module:
```
from scipy import optimize
```
### Finding a minima
Let's first look at how to find the minima of a simple function of a single variable:
```
def f(x):
return 4*x**3 + (x-2)**2 + x**4
fig, ax = plt.subplots()
x = linspace(-5, 3, 100)
ax.plot(x, f(x));
```
We can use the `fmin_bfgs` function to find the minima of a function:
```
x_min = optimize.fmin_bfgs(f, -2)
x_min
optimize.fmin_bfgs(f, 0.5)
```
We can also use the `brent` or `fminbound` functions. They have a bit different syntax and use different algorithms.
```
optimize.brent(f)
optimize.fminbound(f, -4, 2)
```
### Finding a solution to a function
To find the root for a function of the form $f(x) = 0$ we can use the `fsolve` function. It requires an initial guess:
```
omega_c = 3.0
def f(omega):
# a transcendental equation: resonance frequencies of a low-Q SQUID terminated microwave resonator
return tan(2*pi*omega) - omega_c/omega
fig, ax = plt.subplots(figsize=(10,4))
x = linspace(0, 3, 1000)
y = f(x)
mask = where(abs(y) > 50)
x[mask] = y[mask] = NaN # get rid of vertical line when the function flip sign
ax.plot(x, y)
ax.plot([0, 3], [0, 0], 'k')
ax.set_ylim(-5,5);
optimize.fsolve(f, 0.1)
optimize.fsolve(f, 0.6)
optimize.fsolve(f, 1.1)
```
## Interpolation
Interpolation is simple and convenient in scipy: The `interp1d` function, when given arrays describing X and Y data, returns and object that behaves like a function that can be called for an arbitrary value of x (in the range covered by X), and it returns the corresponding interpolated y value:
```
from scipy.interpolate import *
def f(x):
return sin(x)
n = arange(0, 10)
x = linspace(0, 9, 100)
y_meas = f(n) + 0.1 * randn(len(n)) # simulate measurement with noise
y_real = f(x)
linear_interpolation = interp1d(n, y_meas)
y_interp1 = linear_interpolation(x)
cubic_interpolation = interp1d(n, y_meas, kind='cubic')
y_interp2 = cubic_interpolation(x)
fig, ax = plt.subplots(figsize=(10,4))
ax.plot(n, y_meas, 'bs', label='noisy data')
ax.plot(x, y_real, 'k', lw=2, label='true function')
ax.plot(x, y_interp1, 'r', label='linear interp')
ax.plot(x, y_interp2, 'g', label='cubic interp')
ax.legend(loc=3);
```
## Statistics
The `scipy.stats` module contains a large number of statistical distributions, statistical functions and tests. For a complete documentation of its features, see http://docs.scipy.org/doc/scipy/reference/stats.html.
There is also a very powerful python package for statistical modelling called statsmodels. See http://statsmodels.sourceforge.net for more details.
```
from scipy import stats
# create a (discreet) random variable with poissionian distribution
X = stats.poisson(3.5) # photon distribution for a coherent state with n=3.5 photons
n = arange(0,15)
fig, axes = plt.subplots(3,1, sharex=True)
# plot the probability mass function (PMF)
axes[0].step(n, X.pmf(n))
# plot the commulative distribution function (CDF)
axes[1].step(n, X.cdf(n))
# plot histogram of 1000 random realizations of the stochastic variable X
axes[2].hist(X.rvs(size=1000));
# create a (continous) random variable with normal distribution
Y = stats.norm()
x = linspace(-5,5,100)
fig, axes = plt.subplots(3,1, sharex=True)
# plot the probability distribution function (PDF)
axes[0].plot(x, Y.pdf(x))
# plot the commulative distributin function (CDF)
axes[1].plot(x, Y.cdf(x));
# plot histogram of 1000 random realizations of the stochastic variable Y
axes[2].hist(Y.rvs(size=1000), bins=50);
```
Statistics:
```
X.mean(), X.std(), X.var() # poission distribution
Y.mean(), Y.std(), Y.var() # normal distribution
```
### Statistical tests
Test if two sets of (independent) random data comes from the same distribution:
```
t_statistic, p_value = stats.ttest_ind(X.rvs(size=1000), X.rvs(size=1000))
print "t-statistic =", t_statistic
print "p-value =", p_value
```
Since the p value is very large we cannot reject the hypothesis that the two sets of random data have *different* means.
To test if the mean of a single sample of data has mean 0.1 (the true mean is 0.0):
```
stats.ttest_1samp(Y.rvs(size=1000), 0.1)
```
Low p-value means that we can reject the hypothesis that the mean of Y is 0.1.
```
Y.mean()
stats.ttest_1samp(Y.rvs(size=1000), Y.mean())
```
## Further reading
* http://www.scipy.org - The official web page for the SciPy project.
* http://docs.scipy.org/doc/scipy/reference/tutorial/index.html - A tutorial on how to get started using SciPy.
* https://github.com/scipy/scipy/ - The SciPy source code.
| github_jupyter |
```
from nornir import InitNornir
nr = InitNornir(config_file="config.yaml")
```
# Executing tasks
Now that you know how to initialize nornir and work with the inventory let's see how we can leverage it to run tasks on groups of hosts.
Nornir ships a bunch of tasks you can use directly without having to code them yourself. You can check them out [here](../../plugins/tasks/index.rst).
Let's start by executing the `ls -la /tmp` command on all the device in `cmh` of type `host`:
```
from nornir.plugins.tasks import commands
from nornir.plugins.functions.text import print_result
cmh_hosts = nr.filter(site="cmh", role="host")
result = cmh_hosts.run(task=commands.remote_command,
command="ls -la /tmp")
print_result(result, vars=["stdout"])
```
So what have we done here? First we have imported the `commands` and `text` modules. Then we have narrowed down nornir to the hosts we want to operate on. Once we have selected the devices we wanted to operate on we have run two tasks:
1. The task `commands.remote_command` which runs the specified `command` in the remote device.
2. The function `print_result` which just prints on screen the result of an executed task or group of tasks.
Let's try with another example:
```
from nornir.plugins.tasks import networking
cmh_spines = nr.filter(site="bma", role="spine")
result = cmh_spines.run(task=networking.napalm_get,
getters=["facts"])
print_result(result)
```
Pretty much the same pattern, just different task on different devices.
## What is a task
Let's take a look at what a task is. In it's simplest form a task is a function that takes at least a [Task](../../ref/api/task.rst#nornir.core.task.Task) object as argument. For instance:
```
def hi(task):
print(f"hi! My name is {task.host.name} and I live in {task.host['site']}")
nr.run(task=hi, num_workers=1)
```
The task object has access to `nornir`, `host` and `dry_run` attributes.
You can call other tasks from within a task:
```
def available_resources(task):
task.run(task=commands.remote_command,
name="Available disk",
command="df -h")
task.run(task=commands.remote_command,
name="Available memory",
command="free -m")
result = cmh_hosts.run(task=available_resources)
print_result(result, vars=["stdout"])
```
You probably noticed in your previous example that you can name your tasks.
Your task can also accept any extra arguments you may need:
```
def count(task, to):
print(f"{task.host.name}: {list(range(0, to))}")
cmh_hosts.run(task=count,
num_workers=1,
to=10)
cmh_hosts.run(task=count,
num_workers=1,
to=20)
```
## Tasks vs Functions
You probably noticed we introduced the concept of a `function` when we talked about `print_result`. The difference between tasks and functions is that tasks are meant to be run per host while functions are helper functions meant to be run globally.
| github_jupyter |
## Extraindo características com a VGG16, usando Transfer Learning
<b>Importando as bibliotecas</b>
```
import matplotlib.pyplot as plt
%matplotlib inline
import os
from keras.applications.vgg16 import VGG16, preprocess_input
from keras.preprocessing import image
from skimage.io import imread, imsave
import numpy as np
from keras.models import Model
import pandas as pd
import csv
from glob import glob
```
<b>Carregando os pesos da VGG16</b>
```
model = VGG16(weights='imagenet', include_top=True)
```
<b>Visualizando as camadas da VGG16</b>
```
model.summary()
```
<b>Utilizando apenas os pesos da última camada como características</b>
```
model = Model(input=model.input, output=model.get_layer('fc1').output)
```
<b>Adquirindo as imagens da base</b>
```
path_melanoma = "/home/vitoria/Área de Trabalho/ICV/SINFO/ph2/melanoma/"
path_naomelanoma = "/home/vitoria/Área de Trabalho/ICV/SINFO/ph2/naomelanoma/"
lista_melanoma = glob(path_melanoma+'*.bmp')
lista_naomelanoma = glob(path_naomelanoma+'*.bmp')
```
<b>Aplicando os pesos da VGG16 nas imagens para obter as características</b>
```
features = []
for i in range(len(lista_melanoma)):
img = image.load_img(lista_melanoma[i], target_size=(224, 224))
x = image.img_to_array(img)
# Expande o array, inserindo um novo eixo
x = np.expand_dims(x, axis=0)
# Adequa sua imagem ao formato exigido pelo modelo
x = preprocess_input(x)
# Aplica a imagem no modelo e retorna as características extraídas
feature = model.predict(x).reshape(-1)
# Transformando de np pra list
feature = list(feature)
# Adicionando label na lista
feature.append(1)
# Adicionando esse vetor de features na matriz de features
features.append(feature)
for i in range(len(lista_naomelanoma)):
img = image.load_img(lista_naomelanoma[i], target_size=(224, 224))
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
x = preprocess_input(x)
feature = model.predict(x).reshape(-1)
feature = list(feature)
feature.append(0)
features.append(feature)
print(len(features[0]))
```
<b>Salvando os dados no arquivo libsvm</b>
```
def geraSVMfile(rotulo, lista_feat, name_file, path_out, modo):
''''''
arquivo = open(path_out + name_file, modo)
featureFile = str(rotulo) + " "
arquivo.write(featureFile)
for i in range(len(lista_feat)-1):
linha = str(str(i + 1) + ":" + str(lista_feat[i]) + " ")
#print('linha ', str(i + 1), " = ", linha)
arquivo.write(linha)
arquivo.write('\n')
arquivo.close()
name = 'VGG16.libsvm'
path_file = '/home/vitoria/Área de Trabalho/ICV/SINFO/resultados/'
for i in range(len(features)):
classe = features[i][-1]
geraSVMfile(rotulo=classe, lista_feat=features[i], name_file=name, path_out=path_file, modo='a')
#print('Salvando...')
#my_df = pd.DataFrame(features)
#my_df.to_csv('/home/vitoria/Área de Trabalho/ICV/SINFO/featuresVGG16.libsvm', index=False, header=False, sep=' ')
```
| github_jupyter |
```
import numpy as np
import os,sys
sys.path.append('../../RL_lib/Agents')
sys.path.append('../../RL_lib/Policies/PPO')
sys.path.append('../../RL_lib/Policies/Common')
sys.path.append('../../RL_lib/Utils')
sys.path.append('../../Env')
sys.path.append('../../Imaging')
%load_ext autoreload
%load_ext autoreload
%autoreload 2
%matplotlib nbagg
import os
print(os.getcwd())
%%html
<style>
.output_wrapper, .output {
height:auto !important;
max-height:1000px; /* your desired max-height here */
}
.output_scroll {
box-shadow:none !important;
webkit-box-shadow:none !important;
}
</style>
```
# Optimize Policy
```
from env import Env
import env_utils as envu
from dynamics_model import Dynamics_model
from lander_model import Lander_model
from ic_gen_scene import Landing_icgen
import rl_utils
import attitude_utils as attu
from arch_policy_gtvf import Arch
from softmax_pd import Softmax_pd as PD
from policy_ppo import Policy
from value_function import Value_function
import policy_nets as policy_nets
import valfunc_nets as valfunc_nets
import cnn_nets
from agent import Agent
import torch.nn as nn
from flat_constraint import Flat_constraint
from glideslope_constraint import Glideslope_constraint
from rh_constraint import RH_constraint
from no_attitude_constraint import Attitude_constraint
from w_constraint import W_constraint
from reward_terminal_mdr import Reward
from asteroid_hfr_scene import Asteroid
from thruster_model import Thruster_model
asteroid_model = Asteroid(landing_site_override=None, omega_range=(1e-6,5e-4))
ap = attu.Quaternion_attitude()
from flash_lidar2 import Flash_lidar
import attitude_utils as attu
from triangle_ray_intersect import Triangle_ray_intersect
from isosphere import Isosphere
iso = Isosphere(recursion_level=2)
tri = Triangle_ray_intersect()
ap = attu.Quaternion_attitude()
P = 64
sensor = Flash_lidar(ap, tri, sqrt_pixels=int(np.sqrt(P)))
thruster_model = Thruster_model(pulsed=True, scale=1.0, offset=0.4)
lander_model = Lander_model(asteroid_model, thruster_model, ap, sensor, iso)
lander_model.get_state_agent = lander_model.get_state_agent_image_state_stab
logger = rl_utils.Logger()
dynamics_model = Dynamics_model(h=2)
obs_dim = 2*P
gt_dim = 13
action_dim = 12
actions_per_dim = 2
action_logit_dim = action_dim * actions_per_dim
recurrent_steps = 60
reward_object = Reward(landing_coeff=10.0, landing_rlimit=2, landing_vlimit=0.1, tracking_bias=0.01,
dalt_coeff=0.02, fuel_coeff=-0.01, use_gt=True)
glideslope_constraint = Glideslope_constraint(gs_limit=-1.0)
shape_constraint = Flat_constraint()
attitude_constraint = Attitude_constraint(ap)
w_constraint = W_constraint(w_limit=(0.1,0.1,0.1), w_margin=(0.05,0.05,0.05))
rh_constraint = RH_constraint(rh_limit=150)
wi=0.02
ic_gen = Landing_icgen(position_r=(50,500),
p_engine_fail=0.0,
p_scale=(0.01, 0.02),
engine_fail_scale=(1.0,1.0),
asteroid_axis_low=(200,200,200),
asteroid_axis_high=(300,300,300),
#position_theta=(0,np.pi/4),
lander_wll=(-wi,-wi,-wi),
lander_wul=(wi,wi,wi),
attitude_parameterization=ap,
attitude_error=(0,np.pi/16),
min_mass=450, max_mass=500,
debug=False,
inertia_uncertainty_diag=10.0,
inertia_uncertainty_offdiag=1.0)
env = Env(ic_gen, lander_model, dynamics_model, logger,
debug_done=False,
reward_object=reward_object,
glideslope_constraint=glideslope_constraint,
attitude_constraint=attitude_constraint,
w_constraint=w_constraint,
rh_constraint=rh_constraint,
tf_limit=1200.0,print_every=10,nav_period=6)
env.ic_gen.show()
arch = Arch(gt_func=lander_model.get_state_agent_gt)
cnn = cnn_nets.CNN_layer(8,2,8)
policy = Policy(policy_nets.GRU_CNN2(7, action_logit_dim, cnn, recurrent_steps=recurrent_steps),
PD(action_dim, actions_per_dim),
shuffle=False, servo_kl=False, max_grad_norm=30,
init_func=rl_utils.xn_init, scale_image_obs=True, scale_vector_obs=True)
value_function = Value_function(valfunc_nets.GRU1(gt_dim, recurrent_steps=recurrent_steps), rollout_limit=3,
shuffle=False, batch_size=9999999, max_grad_norm=30, obs_key='gt_observes')
agent = Agent(arch, policy, value_function, None, env, logger,
policy_episodes=60, policy_steps=3000, gamma1=0.95, gamma2=0.995,
recurrent_steps=recurrent_steps, monitor=env.rl_stats)
fname = "optimize-RPT2"
policy.load_params(fname)
```
# Test Policy
```
env.test_policy_batch(agent,5000,print_every=100,keys=lander_model.get_engagement_keys())
```
| github_jupyter |
# Read annotations from CoNLL
The folder `annotations-pickle` contains all documents in the corpus with its annotations as pickled Python objects (created with the script `read_annotations.py`). This notebook illustrates how to load and use these objects in Python.
Note: to be able to unpickle the files, you should make sure that your code can access the module `conll_data.py` (e.g. by putting it in the same directory).
```
import gzip
import pickle
from collections import Counter
from tqdm import tqdm
from glob import glob
```
### Single document
The following illustrates how you can read one of the documents as a Document instance and print some information about its sentences, tokens and annotations.
```
example_file = "../data/annotations-pickle/21st-Century-Wire_20170627T181355.conll.pickle.gz"
with gzip.open(example_file, "rb") as infile:
doc = pickle.load(infile)
doc.text
# print statistics
print("Number of sentences:", len(doc.sentences))
print("Number of tokens:", len(doc.tokens))
print("Number of unique words:", len(set(doc.words)))
print("Number of unique lemmas:", len(set(doc.lemmas)))
# inspect a specific sentence
sentence = doc.sentences[0]
print("Text:", sentence.text)
print("Words:", sentence.words)
print("Lemmas:", sentence.lemmas)
# inspect a specific token
token = sentence.get_token(token_id="13") # or: sentence.tokens[12]
print("Word:", token.word)
print("Lemma:", token.lemma)
print("POS:", token.pos)
print("Offset:", token.offset_start, "-", token.offset_end)
print("Full phrase:", sentence.get_full_phrase(head_id="13").text)
# print statistics on the annotations
print(f"{len(doc.events)} events annotated")
print(f"{len(doc.claims)} claims annotated")
print(f"{len(doc.attr_cues)} attribution cues annotated")
print(f"{len(doc.attr_contents)} attribution contents annotated")
print(f"{len(doc.attr_sources)} attribution sources annotated")
print(f"{len(doc.attr_relations)} attribution relations annotated")
# inspect a specific event annotation
doc.events[0].text
# inspect all multi-word events
mw_events = [event for event in doc.events if len(event.tokens) > 1]
for event in mw_events:
print(event.text)
# inspect a specific claim annotation
print(doc.claims[0].text)
# inspect a specific attribution relation;
# one AR can have multiple sources and cues
ar = doc.attr_relations[-1]
print("Content:", ar.content.text)
for source in ar.sources:
print("Source:", source.text)
for cue in ar.cues:
print("Cue:", cue.text)
```
### All documents
The following illustrates how you can read one all documents as Document instances and get some overall information on all annotations.
```
pickle_files = glob("../data/annotations-pickle/*.pickle.gz")
len(pickle_files)
# get all events
events = []
for pickle_file in tqdm(pickle_files):
with gzip.open(pickle_file, "rb") as infile:
doc = pickle.load(infile)
events.extend(doc.events)
print(len(events), "events annotated")
# most frequent events
event_texts = [event.tokens[0].lemma.lower() if len(event.tokens) == 1 else event.text.lower() for event in events]
Counter(event_texts).most_common(10)
# get all cues
cues = []
for pickle_file in tqdm(pickle_files):
with gzip.open(pickle_file, "rb") as infile:
doc = pickle.load(infile)
cues.extend(doc.attr_cues)
print(len(cues), "cues annotated")
# most frequent cues
cue_texts = [cue.tokens[0].lemma.lower() if len(cue.tokens) == 1 else cue.text.lower() for cue in cues]
Counter(cue_texts).most_common(10)
```
| github_jupyter |
[](https://colab.research.google.com/github/uwsampl/tutorial/blob/master/notebook/01_TVM_Tutorial_Intro.ipynb)
Welcome to the TVM tutorial. First we are going to get you setup so you can run the tutorial completely in the Cloud.
Google's Colab notebook's run on ephmeral nodes in the cloud. In order to preserve your build of TVM across notebooks and sessions we will be using Google Drive to cache your build of TVM.
If you have already done this step, for this notebook or another, please skip ahead to the content
after running these cells.
# Installing TVM
First we will connect to your Google drive, please follow the authentication steps below, after executing the cell.
```
from google.colab import drive
drive.mount('/content/gdrive')
```
Next we will install TVM in the Colab notebook, so we may use it for the rest of the tutorial.
```
%%bash
[[ ! -e /tools/google-cloud-sdk ]] && exit
echo "Installing Dependencies ..."
sudo apt-get update
sudo apt-get install -y -q llvm-6.0 libglfw3-dev libtinfo-dev libffi-dev zlib1g-dev clinfo
%%bash
[[ ! -e /tools/google-cloud-sdk ]] && exit
cd /content/gdrive/"My Drive"
if [[ ! -e tvm ]]; then
echo "Cloning TVM ..."
git clone --recursive https://github.com/dmlc/tvm
fi
%%bash
[[ ! -e /tools/google-cloud-sdk ]] && exit
echo "Configuring Build ..."
cd "/content/gdrive/My Drive/tvm"
mkdir -p build
cp cmake/config.cmake build
# sed -i -e 's/USE_OPONGL OFF/USE_OPONGL ON/g' build/config.cmake
sed -i -e 's/USE_CUDA OFF/USE_CUDA ON/g' build/config.cmake
sed -i -e 's/USE_CUDNN OFF/USE_CUDNN ON/g' build/config.cmake
sed -i -e 's/USE_LLVM OFF/USE_LLVM ON/g' build/config.cmake
sed -i -e 's/USE_VTA_TSIM OFF/USE_VTA_TSIM ON/g' build/config.cmake
cat build/config.cmake
%%bash
[[ ! -e /tools/google-cloud-sdk ]] && exit
echo "Running CMake ..."
cd "/content/gdrive/My Drive/tvm/build"
cmake ..
%%bash
[[ ! -e /tools/google-cloud-sdk ]] && exit
echo "Building TVM ..."
cd "/content/gdrive/My Drive/tvm/build"
make -j4
%%bash
[[ ! -e /tools/google-cloud-sdk ]] && exit
echo "Installing Python libraries ..."
cd "/content/gdrive/My Drive/tvm/"
cd python; python setup.py install; cd ..
cd topi/python; python setup.py install
```
We will set a global variable so we can later check if we are in the Cloud notebook or running locally.
```
try:
import google.colab
IN_COLAB = True
except:
IN_COLAB = False
```
Now that we have installed everything, please restart the runtime. If you have run all the above steps you are now be ready to use TVM.

```
import tvm
print(tvm)
import topi
print(topi)
```
| github_jupyter |
# Tensorboard
## Add image
```
import matplotlib.pyplot as plt
from torchvision import datasets, models, transforms
import torch.optim as optim
import torch.nn as nn
import torchvision
from torchvision.transforms import *
from torch.utils.data import DataLoader
from torch.utils.tensorboard import SummaryWriter
import torch
import numpy as np
from collections import defaultdict
import time
import copy
def matplotlib_imshow(img, one_channel=False):
if one_channel:
img = img.mean(dim=0)
# img = img / 2 + 0.5 # unnormalize
npimg = img.numpy()
if one_channel:
plt.imshow(npimg, cmap="Greys")
else:
plt.imshow(np.transpose(npimg, (1, 2, 0)))
def get_dataloaders():
def clean_up(path):
import os
from shutil import rmtree
ipynb_checkpoints = f'{path}/.ipynb_checkpoints'
if os.path.exists(ipynb_checkpoints):
rmtree(ipynb_checkpoints)
def get_dataloader(phase):
path = f'./shapes/{phase}'
clean_up(path)
if phase in ['valid']:
transform = transforms.Compose([
Resize(224),
ToTensor()])
else:
transform = transforms.Compose([
Resize(224),
RandomAffine(degrees=(30, 50), shear=5),
ToTensor()])
image_folder = datasets.ImageFolder(path, transform=transform)
# print(path, image_folder.classes)
return DataLoader(image_folder, batch_size=4, shuffle=True, num_workers=4)
return {phase: get_dataloader(phase) for phase in ['train', 'test', 'valid']}
np.random.seed(37)
torch.manual_seed(37)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
pretrained=True
num_classes = 3
writer = SummaryWriter('/root/tensorboard/shape_experiment_1')
dataloaders = get_dataloaders()
data_iter = iter(dataloaders['train'])
images, labels = data_iter.next()
img_grid = torchvision.utils.make_grid(images)
matplotlib_imshow(img_grid, one_channel=False)
writer.add_image('four_shape_images', img_grid)
writer.close()
```
## Add model
```
model = models.resnet18(pretrained=pretrained)
model.fc = nn.Linear(model.fc.in_features, num_classes)
model = model.to(device)
writer.add_graph(model, images.to(device))
writer.close()
```
## Add embedding
```
def get_data(dataloader, device):
imgs = []
lbls = []
for inputs, labels in dataloader:
inputs = inputs.to(device)
labels = labels.to(device)
imgs.append(inputs)
lbls.append(labels)
imgs = torch.cat(imgs)
lbls = torch.cat(lbls)
return imgs, lbls
def select_n_random(data, labels, n=100):
assert len(data) == len(labels)
perm = torch.randperm(len(data))
return data[perm][:n], labels[perm][:n]
images, labels = get_data(dataloaders['train'], device)
images, labels = select_n_random(images, labels)
idx_to_class = {v: k for k, v in dataloaders['train'].dataset.class_to_idx.items()}
class_labels = [idx_to_class[lab] for lab in labels.cpu().detach().numpy()]
writer.add_embedding(images.mean(dim=1).view(-1, 224 * 224), metadata=class_labels, label_img=images, global_step=1)
writer.close()
```
## Add training loss
```
import torch.nn.functional as F
def images_to_probs(net, images):
output = net(images)
_, preds_tensor = torch.max(output, 1)
preds = np.squeeze(preds_tensor.cpu().detach().numpy())
return preds, [F.softmax(el, dim=0)[i].item() for i, el in zip(preds, output)]
def plot_classes_preds(net, images, labels, classes):
preds, probs = images_to_probs(net, images)
fig = plt.figure(figsize=(12, 48))
n_images = images.shape[0]
for idx in np.arange(n_images):
ax = fig.add_subplot(1, n_images, idx+1, xticks=[], yticks=[])
matplotlib_imshow(images[idx].cpu().detach(), one_channel=False)
clazz_pred = classes[preds[idx]]
clazz_true = classes[labels[idx].cpu().detach().numpy().item()]
prob_pred = probs[idx] * 100.0
color = 'green' if preds[idx]==labels[idx].item() else 'red'
ax.set_title(f'pred={clazz_pred}, {prob_pred:.2f}\ntrue={clazz_true}', color=color)
return fig
def train(dataloaders, model, criterion, optimizer, scheduler, classes, device, writer, num_epochs=20):
def format_start_stop(start, stop):
elapsed = stop - start
return f'{elapsed//60:.0f}m {elapsed%50:.0f}s'
best_model_weights = copy.deepcopy(model.state_dict())
best_acc = -1.0
loop_start = time.time()
for epoch in range(num_epochs):
train_loss, train_acc = 0.0, 0.0
test_loss, test_acc = 0.0, 0.0
train_start, test_start = 0.0, 0.0
train_stop, test_stop = 0.0, 0.0
for i, phase in enumerate(['train', 'test']):
if phase == 'train':
optimizer.step()
scheduler.step()
model.train()
train_start = time.time()
else:
model.eval()
test_start = time.time()
running_loss = 0.0
running_corrects = 0
n = 0
dataloader = dataloaders[phase]
for inputs, labels in dataloader:
inputs = inputs.to(device)
labels = labels.to(device)
optimizer.zero_grad()
with torch.set_grad_enabled(phase == 'train'):
outputs = model(inputs)
loss = criterion(outputs, labels)
_, preds = torch.max(outputs, 1)
if phase == 'train':
loss.backward()
optimizer.step()
running_loss += loss.mean() * inputs.size(0)
running_corrects += torch.sum(preds == labels.data)
n += len(labels)
if phase == 'train':
writer.add_scalar('training loss', running_loss, epoch * len(dataloader) + i)
writer.add_figure('predictions vs. actuals',
plot_classes_preds(model, inputs, labels, classes),
global_step=epoch * len(dataloader) + i)
epoch_loss = running_loss / float(n)
epoch_acc = running_corrects.double() / float(n)
if phase == 'train':
train_stop = time.time()
train_loss, train_acc = epoch_loss, epoch_acc
else:
test_stop = time.time()
test_loss, test_acc = epoch_loss, epoch_acc
if epoch_acc > best_acc:
best_model_weights = copy.deepcopy(model.state_dict())
best_acc = epoch_acc
train_time = format_start_stop(train_start, train_stop)
test_time = format_start_stop(test_start, test_stop)
train_metrics = f'TRAIN: {train_loss:.4f}, {train_acc:.4f}, {train_time}'
test_metrics = f'TEST: {test_loss:.4f}, {test_acc:.4f}, {test_time}'
print(f'epoch {str(epoch + 1).zfill(2)}/{str(num_epochs).zfill(2)} | {train_metrics} | {test_metrics}')
loop_stop = time.time()
loop_time = format_start_stop(loop_start, loop_stop)
print(f'completed learning in {loop_time}, best accuracy {best_acc:.4f}')
model.load_state_dict(best_model_weights)
writer.close()
return model
num_epochs = 20
model = models.resnet18(pretrained=pretrained)
model.fc = nn.Linear(model.fc.in_features, num_classes)
model = model.to(device)
criterion = nn.CrossEntropyLoss()
optimizer = optim.Rprop(model.parameters(), lr=0.01)
scheduler = optim.lr_scheduler.StepLR(optimizer, step_size=1)
model = train(dataloaders, model, criterion, optimizer, scheduler, idx_to_class, device, writer, num_epochs=num_epochs)
```
## Add precision-recall curve
```
class_probs = []
class_preds = []
with torch.no_grad():
for data in dataloaders['valid']:
images, labels = data
images, labels = images.to(device), labels.to(device)
output = model(images)
class_probs_batch = [F.softmax(el, dim=0) for el in output]
_, class_preds_batch = torch.max(output, 1)
class_probs.append(class_probs_batch)
class_preds.append(class_preds_batch)
test_probs = torch.cat([torch.stack(batch) for batch in class_probs])
test_preds = torch.cat(class_preds)
test_probs
test_preds
def add_pr_curve_tensorboard(class_index, test_probs, test_preds, idx_to_class, global_step=0):
tensorboard_preds = test_preds == class_index
tensorboard_probs = test_probs[:, class_index]
writer.add_pr_curve(idx_to_class[class_index],
tensorboard_preds,
tensorboard_probs,
global_step=global_step)
writer.close()
n_classes = len(dataloaders['valid'].dataset.classes)
for i in range(n_classes):
add_pr_curve_tensorboard(i, test_probs, test_preds, idx_to_class)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/gumdropsteve/intro_to_machine_learning/blob/main/day_03/01_hyper.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import pandas as pd
df = pd.read_csv('https://github.com/gumdropsteve/datasets/raw/master/iris.csv')
df
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=1)
knn
df.columns
X = df[['sepal_length', 'sepal_width', 'petal_length', 'petal_width']]
y = df.target.astype('int')
knn.fit(X, y)
knn.predict(X)
sum(knn.predict(X) == y) / 150
from sklearn.model_selection import train_test_split
# variables that help us predict the target (y)
X = df[['sepal_length', 'sepal_width', 'petal_length', 'petal_width']]
# y is whatever you want to predict ('target', 'labels', etc...)
y = df.target.astype('int')
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.7)
X_train
len(y_test)
```
Random state...
```
random_state : int, RandomState instance or None, optional (default=None)
If int, random_state is the seed used by the random number generator;
If RandomState instance, random_state is the random number generator;
If None, the random number generator is the RandomState instance used
by np.random.
```
```
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.7, random_state=2)
X_train.tail(1)
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.7, random_state=2)
X_train.tail(1)
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.7, random_state=57)
X_train.tail(1)
model = KNeighborsClassifier(n_neighbors=1)
model.fit(X_train, y_train)
preds = model.predict(X_test)
sum(preds == y_test) / 150
```
Cross-validation,sometimes called out-of-sample testing, is any of various similar model validation techniques for assessing how the results of a statistical analysis will generalize to an independent data set.
More or less... a way to tell if your model is good by testing it with data it wasn't trained on.
```
from sklearn.model_selection import cross_val_score
model = KNeighborsClassifier(n_neighbors=3)
X = df[['sepal_length', 'sepal_width', 'petal_length', 'petal_width']]
y = df.target.astype('int')
cross_val_score(model, X, y)
df.species.value_counts()
```
```
scoring
str or callable, default=None
A str (see model evaluation documentation) or a scorer callable object / function with signature scorer(estimator, X, y) which should return only a single value.
Similar to cross_validate but only a single metric is permitted.
If None, the estimator’s default scorer (if available) is used.
```
```
cross_val_score(model, X, y, scoring='accuracy')
```
https://scikit-learn.org/stable/modules/model_evaluation.html#scoring-parameter
```
cross_val_score(model, X, y, scoring='f1_weighted')
```
## Switching to a Regression Model
```
df = pd.read_parquet('https://github.com/gumdropsteve/datasets/raw/master/nyc_taxi/yellow_tripdata_2019-12.parquet')
df.tail(3)
df = df.dropna()
X = df[['passenger_count', 'trip_distance', 'RatecodeID', 'PULocationID', 'DOLocationID']]
y = df.fare_amount
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.7)
len(X_train) + len(y_test)
from sklearn.linear_model import LinearRegression
model = LinearRegression()
# use training data to fit the model (fit means train)
model.fit(X_train, y_train)
model.predict(X_test)
preds = model.predict(X_test)
sum(preds == y_test)
from sklearn.metrics import mean_squared_error
mean_squared_error(y_true=y_test, y_pred=preds)
np.mean(y_test)
preds[-5:]
y_test[-5:]
cross_val_score(model, X, y, scoring='r2')
```
## Grid Search
```
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LinearRegression
from sklearn.pipeline import make_pipeline
def PolynomialRegression(degree=2, **kwargs):
return make_pipeline(PolynomialFeatures(degree),
LinearRegression(**kwargs))
np.arange(21)
import numpy as np
def make_data(N, err=1.0, rseed=1):
# randomly sample the data
rng = np.random.RandomState(rseed)
X = rng.rand(N, 1) ** 2
y = 10 - 1. / (X.ravel() + 0.1)
if err > 0:
y += err * rng.randn(N)
return X, y
X, y = make_data(40)
from sklearn.model_selection import GridSearchCV
param_grid = {'polynomialfeatures__degree': np.arange(21),
'linearregression__fit_intercept': [True, False],
'linearregression__normalize': [True, False]}
grid = GridSearchCV(PolynomialRegression(), param_grid, cv=7)
grid
grid.fit(X, y)
grid.best_params_
def PolynomialRegression():
return make_pipeline(PolynomialFeatures(degree=4),
LinearRegression(normalize=True, fit_intercept=False))
PolynomialRegression()
```
### Simple on Taxi
```
from sklearn.pipeline import make_pipeline
from sklearn.linear_model import LinearRegression
def LR(**kwargs):
return make_pipeline(LinearRegression(**kwargs))
param_grid = {'linearregression__fit_intercept': [True, False],
'linearregression__normalize': [True, False]}
grid = GridSearchCV(LR(), param_grid, cv=7)
df = pd.read_parquet('https://github.com/gumdropsteve/datasets/raw/master/nyc_taxi/yellow_tripdata_2019-12.parquet')
df = df.dropna()
X = df[['passenger_count', 'trip_distance', 'RatecodeID', 'PULocationID', 'DOLocationID']]
y = df.fare_amount
grid.fit(X, y)
grid.best_params_
model = LinearRegression()
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.7)
model.fit(X_train, y_train)
preds = model.predict(X_test)
from sklearn.metrics import r2_score
r2_score(y_true=y_test, y_pred=preds)
```
| github_jupyter |
Appendix plots for describing the Gaussian process model
```
from Starfish.emulator import Emulator, PCAGrid
from Starfish.utils import saveall
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1.inset_locator import zoomed_inset_axes
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
from mpl_toolkits.axes_grid1.inset_locator import mark_inset
from matplotlib.ticker import FormatStrFormatter as FSF
from matplotlib.ticker import MaxNLocator
from matplotlib.ticker import MultipleLocator
from Starfish.grid_tools import HDF5Interface
em = Emulator.open("../../libraries/PHOENIX_SPEX_M_PCA.hdf5") # All weights
pca = em.PCAGrid
temps = np.unique(pca.gparams[:,0])
#loggs = np.unique(pca.gparams[:,1])
#Zs = np.unique(pca.gparams[:,2])
#points = {"temp":temps, "logg":loggs, "Z":Zs}
# temps use just 2800 - 3200
logg = 5.0
Z = 0.0
int_temps = np.linspace(temps[0], temps[-1], num=40)
int_temps2 = np.linspace(3100, 3200, num=40)
weight_index = 4
weights = pca.w[weight_index]
pcomp = pca.pcomps[weight_index]
emw = em.WEs[weight_index] # The emulator for the first weight
nsamp = 50
mu, var = emw(np.array([3150, logg, Z]))
mu = mu[0]
var = var[0][0]
print(mu)
fig = plt.figure(figsize=(4, 3))
ax = fig.add_subplot(111)
# Load the grid-point weights
ww = []
for temp in temps:
pars = np.array([temp, logg, Z])
index = pca.get_index(pars)
ww.append(weights[index])
fparams = []
for temp in int_temps:
fparams.append([temp, logg, Z])
fparams = np.array(fparams)
wgps = []
for k in range(nsamp):
wgps.append(emw.draw_weights(fparams))
for k in range(nsamp):
ax.plot(int_temps, wgps[k] , "b", lw=0.1)
ax.plot(temps, ww, "bo")
ax.xaxis.set_major_formatter(FSF("%.0f"))
ax.xaxis.set_major_locator(MultipleLocator(100))
ax.set_xlabel(r"$T_\textrm{eff}$")
ax.set_ylabel(r"$w_5$")
# #l,b,w,h
# rect = 0.6, 0.2, 0.25, 0.3
# axins = fig.add_axes(rect)
axins = inset_axes(ax, width=1.5, height=1.5, loc=4)
# Finer temperature spacing for the inset
fparams2 = []
for temp in int_temps2:
fparams2.append([temp, logg, Z])
fparams2 = np.array(fparams2)
for k in range(nsamp):
axins.plot(int_temps2, emw.draw_weights(fparams2) , "b", lw=0.1)
axins.axvline(3150, color="k", ymin=0.2, ymax=0.8)
# sub region of the original image
x1, x2, y1, y2 = 3140, 3160, mu-0.01, mu+0.01
axins.set_xlim(x1, x2)
axins.set_ylim(y1, y2)
#ax.plot(3150, mu, "o")
axins.plot(3150, mu, "ko")
axins.xaxis.set_ticklabels([])
axins.yaxis.set_ticklabels([])
mark_inset(ax, axins, loc1=2, loc2=1, fc="none", ec="0.6")
fig.subplots_adjust(left=0.14, right=0.86, bottom=0.15)
saveall(fig, "../../plots/GP_left")
plt.show()
```
Now plot the probability distribution at 3150
```
def gauss(x):
return 1/np.sqrt(2. * np.pi * var) * np.exp(-0.5 * (x - mu)**2/var)
step = 0.1 * (y2 - y1)
xs = np.linspace(y1 + step, y2 - step, num=100)
fig = plt.figure(figsize=(3,3))
ax = fig.add_subplot(111)
ax.plot(xs, gauss(xs), "k")
ax.set_xlim(xs[0], xs[-1])
#ax.xaxis.set_major_formatter(FSF("%.0f"))
ax.xaxis.set_major_locator(MultipleLocator(0.005))
ax.set_xlabel(r"$w_5$")
ax.set_ylabel(r"$p(w_5 |\, \theta_\ast)$")
fig.subplots_adjust(left=0.18, right=0.82, bottom=0.15)
saveall(fig, "../../plots/GP_right")
plt.show()
```
| github_jupyter |
# Install basic requirements
```
pip install -U whylogs pandas
import whylogs
import pandas as pd
```
# Load example data batches
The example data is prepared from our public S3 bucket. You can use your own data if you want if you have multiple batches of data.
```
pdfs = []
for i in range(1, 8):
path = f"https://whylabs-public.s3.us-west-2.amazonaws.com/demo_batches/input_batch_{i}.csv"
print(f"Loading data from {path}")
df = pd.read_csv(path)
pdfs.append(df)
pdfs[0].describe()
```
# Configure whylogs
`whylogs`, by default, does not send statistics to WhyLabs.
There are a few small steps you need to set up. If you haven't got the access key, please onboard with WhyLabs.
**WhyLabs only requires whylogs API - your raw data never leaves your premise.**
```
from whylogs.app import Session
from whylogs.app.writers import WhyLabsWriter
import os
import datetime
import getpass
# set your org-id here
print("Enter your WhyLabs Org ID")
os.environ["WHYLABS_DEFAULT_ORG_ID"] = input()
# set your API key here
print("Enter your WhyLabs API key")
os.environ["WHYLABS_API_KEY"] = getpass.getpass()
print("Using API Key ID: ", os.environ["WHYLABS_API_KEY"][0:10])
```
## Creating session
Once the environments are set, let's create a whylogs session with a WhyLabs writer.
Note that you can add your local writer or S3 writer if you want here. Check out the API docs for more information.
```
# create WhyLabs session
writer = WhyLabsWriter("", formats=[])
session = Session(project="demo-project", pipeline="demo-pipeline", writers=[writer])
```
## Logging to WhyLabs
Ensure you have a **model ID** (also called **dataset ID**) before you start!
### Dataset Timestamp
* To avoid confusion, it's recommended that you use UTC
* If you don't set `dataset_timestamp` parameter, it'll default to `UTC` now
* WhyLabs supports real time visualization when the timestamp is **within the last 7 days**. Anything older than than will be picked up when we run our batch processing
* **If you log two profiles for the same day with different timestamps (12:00 vs 12:01), they are merged to the same batch**
### Logging Different Batches of Data
* We'll give the profiles different **dates**
* Create a new logger for each date. Note that the logger needs to be `closed` to flush out the data
```
print("Enter your model ID from WhyLabs:")
model_id = input()
for i, df in enumerate(pdfs):
# walking backwards. Each dataset has to map to a date to show up as a different batch
# in WhyLabs
dt = datetime.datetime.now(tz=datetime.timezone.utc) - datetime.timedelta(days=i)
# Create new logger for date
with session.logger(tags={"datasetId": model_id}, dataset_timestamp=dt) as ylog:
print("Log data frame for ", dt)
ylog.log_dataframe(df)
# Ensure everything is flushed
session.close()
```
## Voila
* Now check the application to see if your **statistics** are in!!
* Also, run the above cell again for the same model ID, do you see the statistics changes in WhyLabs? Especially the counters?
| github_jupyter |
```
#################
#### IMPORTS ####
#################
# Arrays
import numpy as np
import cytoolz
# Deep Learning stuff
import torch
import torchvision
import torchvision.transforms as transforms
# Images display and plots
import matplotlib.pyplot as plt
from matplotlib.colors import LinearSegmentedColormap
from matplotlib.colors import ListedColormap
import matplotlib.pylab as pl
# Fancy progress bars
import tqdm.notebook as tq
# Tensor Network Stuff
%config InlineBackend.figure_formats = ['svg']
import quimb.tensor as qtn # Tensor Network library
import quimb
import collections
import opt_einsum as oe
import itertools
import copy
import sys, os
sys.path.insert(0, '../')
# My functions
from TNutils import *
def meanpool2d_gs(npmnist, shape):
'''
Apply a meanpool convolution of an array of images (flattened)
meanpool has kernel size 2x2
'''
ds_imgs = []
for img in npmnist:
ds_img = []
for col in range(0,shape[0],2):
for row in range(0,shape[1],2):
pixel = np.mean([img.reshape(shape)[col,row], img.reshape(shape)[col,row+1],
img.reshape(shape)[col+1,row], img.reshape(shape)[col+1,row+1]])
ds_img.append(pixel)
ds_imgs.append(np.array(ds_img).reshape((shape[0]//2)*(shape[1]//2)) )
ds_imgs = np.array(ds_imgs)
return ds_imgs
def get_data_gs(train_size = 1000, test_size = 100, grayscale_threshold = .5, reduced = False):
'''
Prepare the MNIST dataset for the training algorithm:
* Choose randomly a subset from the whole dataset
* Flatten each image to mirror the mps structure
* Normalize images from [0,255] to [0,1]
* Apply a threshold for each pixels so that each value
below that threshold are set to 0, the others get set to 1.
For this algorithm we will only deal to binary states {0,1}
instead of a range from 0 to 1
'''
# Download all data
mnist = torchvision.datasets.MNIST('classifier_data', train=True, download=True,
transform = transforms.Compose([transforms.ToTensor()]) )
# Convert torch.tenor to numpy
npmnist = mnist.data.numpy()
# Check of the type of the sizes
#if ((type(train_size) != int) or (type(test_size) != int)):
# raise TypeError('train_size and test_size must be INT')
# Check if the training_size and test_size requested are bigger than
# the MNIST whole size
if ( (train_size + test_size) > npmnist.shape[0] ):
raise ValueError('Subset too big')
# Check of the positivity of sizes
if ( (train_size <= 0) or (test_size <= 0) ):
raise ValueError('Size of training set and test set cannot be negative')
# Choose just a subset of the data
# Creating a mask by randomly sampling the indexes of the full dataset
subset_indexes = np.random.choice(np.arange(npmnist.shape[0]), size=(train_size + test_size),
replace=False, p=None)
# Apply the mask
npmnist = npmnist[subset_indexes]
# Flatten every image
npmnist = np.reshape(npmnist, (npmnist.shape[0], npmnist.shape[1]*npmnist.shape[2]))
# Normalize the data from 0 - 255 to 0 - 1
if reduced:
npmnist = np.array(npmnist/npmnist.max() )
npmnist = meanpool2d_gs(npmnist, (28,28))
npmnist = np.round(3*np.array(npmnist/npmnist.max() ),0)
npmnist_bits = []
for image in npmnist:
image_bits = []
for pixel in image:
if pixel == 0:
image_bits.append(0)
image_bits.append(0)
elif pixel == 1:
image_bits.append(0)
image_bits.append(1)
elif pixel == 2:
image_bits.append(1)
image_bits.append(0)
elif pixel == 3:
image_bits.append(1)
image_bits.append(1)
npmnist_bits.append(np.array(image_bits))
# Return training set and test set
return npmnist_bits[:train_size], npmnist_bits[train_size:]
def binary_to_img(img_flat_bits, shape):
img_flat = []
for k in range(0,2*shape[0]*shape[1],2):
if img_flat_bits[k] == 0 and img_flat_bits[k+1] == 0:
img_flat.append(0)
elif img_flat_bits[k] == 0 and img_flat_bits[k+1] == 1:
img_flat.append(1)
elif img_flat_bits[k] == 1 and img_flat_bits[k+1] == 0:
img_flat.append(2)
elif img_flat_bits[k] == 1 and img_flat_bits[k+1] == 1:
img_flat.append(3)
else:
img_flat.append(-1)
img_flat = np.array(img_flat)
return img_flat
def plot_img_gs(img_flat_bits, shape, flip_color = True, savefig = ''):
'''
Display the image from the flattened form
'''
# If the image is corrupted for partial reconstruction (pixels are set to -1)
img_flat = binary_to_img(img_flat_bits, shape)
if -1 in img_flat:
img_flat = np.copy(img_flat)
img_flat[img_flat == -1] = 0
plt.figure(figsize = (2,2))
# Background white, strokes black
if flip_color:
plt.imshow(1-np.reshape(img_flat,shape), cmap='gray')
# Background black, strokes white
else:
plt.imshow(np.reshape(img_flat,shape), cmap='gray')
plt.axis('off')
if savefig != '':
# save the picture as svg in the location determined by savefig
plt.savefig(savefig, format='svg')
plt.show()
data, _ = get_data_gs(reduced = True)
data = np.array(data)[:10]
data.shape
plot_img_gs(data[0], (14,14))
mps = initialize_mps(28*14,2)
#########################
# CHACHE INITIALIZATION #
#########################
# Creating list of tensor of the images of the training set
_imgs = np.array([tens_picture(img) for img in data])
_img_cache = []
for _img in _imgs:
_left_cache = np.array([qtn.Tensor() for _ in range(len(_img))])
_right_cache = np.array([qtn.Tensor() for _ in range(len(_img))])
_img_cache.append((_left_cache,_right_cache))
_img_cache = np.array(_img_cache)
last_dirs = np.zeros(len(_img_cache),dtype =np.int32)
last_sites = np.zeros(len(_img_cache),dtype =np.int32)
last_epochs = -np.ones(len(_img_cache),dtype =np.int32)
nlls = cached_stochastic_learning_epoch(mps, data, _imgs, 20, 0.001,
_img_cache, last_dirs, last_sites, last_epochs,
batch_size = 100)
plt.plot(nlls)
bdims_imshow(mps,(28,14))
plot_img_gs(generate_sample(mps),(14,14) )
corr_img = partial_removal_img(data[6], fraction = .6, axis = 0, shape=(28,14))
recc = reconstruct(mps, corr_img)
plot_rec(binary_to_img(corr_img,(14,14)), binary_to_img(recc,(14,14)), (14,14), N=4)
plot_img(binary_to_img(corr_img,(14,14)),(14,14))
plot_img(binary_to_img(recc,(14,14)),(14,14))
```
| github_jupyter |
This notebook contains all of the necessary code to reproduce all of the figures used in the paper.
It is sectioned into a section for required auxiliary methods and variables followed by different sections for each figure.
# Import modules
```
import sqlite3
import pandas as pd
import geopandas
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import ipaddress
import csv
import json
import descartes
import cartopy
import cartopy.crs as ccrs
import collections
from collections import Counter
from matplotlib import colors
from matplotlib.lines import Line2D
from matplotlib.patches import Patch
from matplotlib.legend_handler import HandlerBase
from matplotlib.text import Text
from matplotlib.legend import Legend
from matplotlib.colors import LinearSegmentedColormap
from mpl_toolkits.axes_grid1 import make_axes_locatable
from mpl_toolkits.basemap import Basemap
from scipy import stats
from scipy.stats import gaussian_kde
```
# Auxiliary methods and variables
## Database file paths (set this before running the figure reproduction)
```
db_path = r""# file path of the Speedchecker database, e.g., r"D:/db/speedchecker_db.db"
db_path_ripe = r"" # file path of the RIPE Atlas database, e.g., r"D:/db/ripeanalysis-data.db"
```
## Global
```
continents = ["AF", "AS", "EU", "NA", "OC", "SA"]
cols = ["#c1d5e2", "#ce9c7f", "#7598b6", "#6a7465", "#914c09", "#663979", "#fbe17b"]
continent_colors = {"NA": "#7598b6", "AS": "#914c09", "EU": "#ce9c7f", "AF": "#c1d5e2",
"OC": "#663979", "SA": "#6a7465"}
lstyles = {"AF": "solid", "AS": "solid", "EU": "dashed", "NA": "dotted", "OC": "dashdot", "SA": "dotted"}
asns = [8075, 14061, 15169, 16509, 20473, 31898, 36351, 45102, 63949]
asns_dict = {
8075: "Microsoft", 14061: "DigitalOcean", 15169: "Google", 16509: "Amazon EC2", 20473: "Vultr",
31898: "Oracle", 36351: "IBM", 45102: "Alibaba", 63949: "Linode"
}
provider_abbrev = {"Alibaba": "BABA", "Amazon EC2": "AMZN", "Amazon Lightsail": "LTSL", "DigitalOcean": "DO",
"Google": "GCP", "IBM": "IBM", "Linode": "LIN", "Microsoft": "MSFT", "Oracle": "ORCL", "Vultr": "VLTR"}
plt.rcParams.update({
"text.usetex": True,
"figure.autolayout": True})
plt.rcParams["axes.axisbelow"] = True
with open("iso3.json", mode="r", encoding="utf-8") as file:
f = json.load(file)
country_codes = {value: key for key, value in f.items()} # create mapping from 2-letter to 3-letter country code
peeringdb = {}
with open("peeringdb.json", mode="r", encoding="utf-8") as file:
json_obj = json.load(file)
for elem in json_obj["data"]:
peeringdb[elem["asn"]] = {"id": elem["id"], "org_id": elem["org_id"],
"name": elem["name"], "info_type": elem["info_type"]}
def get_pdb_info(asn):
if asn in peeringdb:
pdb_dict = peeringdb[asn]
return pdb_dict["name"], pdb_dict["info_type"]
else:
return "Unknown", "Unknown"
def create_connection(db):
try:
con = sqlite3.connect(db)
return con
except sqlite3.Error as e:
print(e)
def is_private_ip(ip):
ip_obj = ipaddress.ip_address(ip)
return (ip_obj in ipaddress.ip_network("10.0.0.0/8") or ip_obj in ipaddress.ip_network("172.16.0.0/12")
or ip_obj in ipaddress.ip_network("192.168.0.0/16"))
```
## Speedchecker
```
conn = create_connection(db_path)
cursor = conn.cursor()
probe_countries = {}
probe_continents = {}
cursor.execute("select ProbeID, Country, Continent from Probes where Country != ''")
for p_id, co, cont in cursor:
probe_countries[p_id] = co
probe_continents[p_id] = cont
def get_lowest_avg_pings(ping_type, continent, filter_provider, glob, source_continent=""):
if glob:
if ping_type.lower() == "tcp":
probe_info_sql = """SELECT Ping.ProbeID, Ping1, Ping2, Ping3, Ping4, Ping5, DestinationURL,
D.Provider FROM Ping JOIN Probes P ON Ping.ProbeID = P.ProbeID JOIN Datacenters D
ON D.Url = Ping.DestinationURL WHERE P.Continent = D.Continent"""
elif ping_type.lower() == "icmp":
probe_info_sql = """WITH relevant_ids AS (
SELECT TracerouteID FROM Traceroute JOIN Datacenters D ON
D.URL = Traceroute.DestinationURL JOIN Probes P ON P.ProbeID = Traceroute.ProbeID
WHERE P.Continent = D.Continent),
ranked_hops AS (
SELECT HopNumber, TracerouteID, RTT1, RTT2, RTT3, ROW_NUMBER() OVER
(PARTITION BY TracerouteID ORDER BY HopNumber DESC) AS rn FROM Hops
WHERE TracerouteID IN relevant_ids),
tracert_id_info AS (
SELECT ProbeID, TracerouteID, DestinationURL, D.Provider FROM Traceroute T
JOIN Datacenters D on D.URL = T.DestinationURL where TracerouteID in relevant_ids)
SELECT T.ProbeID, RTT1, RTT2, RTT3, 0, 0, DestinationURL,
Provider FROM ranked_hops R JOIN tracert_id_info T
ON R.TracerouteID = T.TracerouteID WHERE rn = 1 """
else:
print("Invalid ping type")
return
cursor.execute(probe_info_sql)
else:
if ping_type.lower() == "tcp":
probe_info_sql = """SELECT Ping.ProbeID, Ping1, Ping2, Ping3, Ping4, Ping5, DestinationURL,
D.Provider FROM Ping JOIN Probes P ON Ping.ProbeID = P.ProbeID JOIN Datacenters D
ON D.Url = Ping.DestinationURL WHERE P.Continent = ? and D.Continent = ?"""
elif ping_type.lower() == "icmp":
probe_info_sql = """WITH relevant_ids AS (
SELECT TracerouteID FROM Traceroute JOIN Datacenters D ON
D.URL = Traceroute.DestinationURL JOIN Probes P ON P.ProbeID = Traceroute.ProbeID
WHERE P.Continent = ? and D.Continent = ?),
ranked_hops AS (
SELECT HopNumber, TracerouteID, RTT1, RTT2, RTT3, ROW_NUMBER() OVER
(PARTITION BY TracerouteID ORDER BY HopNumber DESC) AS rn FROM Hops
WHERE TracerouteID IN relevant_ids),
tracert_id_info AS (
SELECT ProbeID, TracerouteID, DestinationURL, D.Provider FROM Traceroute T
JOIN Datacenters D on D.URL = T.DestinationURL where TracerouteID in relevant_ids)
SELECT T.ProbeID, RTT1, RTT2, RTT3, 0, 0, DestinationURL,
Provider FROM ranked_hops R JOIN tracert_id_info T
ON R.TracerouteID = T.TracerouteID WHERE rn = 1 """
else:
print("Invalid ping type")
return
if source_continent:
cursor.execute(probe_info_sql, (source_continent, continent))
else:
cursor.execute(probe_info_sql, (continent, continent))
probe_ping_dict = {}
for probe in cursor:
probe_id = probe[0]
dc = probe[6]
if probe_id not in probe_ping_dict:
probe_ping_dict[probe_id] = {}
if filter_provider:
provider = probe[7]
if provider not in probe_ping_dict[probe_id]:
probe_ping_dict[probe_id][provider] = {}
if dc not in probe_ping_dict[probe_id][provider]:
probe_ping_dict[probe_id][provider][dc] = []
probe_ping_dict[probe_id][provider][dc] += [p for p in probe[1:6] if
p is not None and 0 < p < 1000]
else:
if dc not in probe_ping_dict[probe_id]:
probe_ping_dict[probe_id][dc] = []
probe_ping_dict[probe_id][dc] += [p for p in probe[1:6] if p is not None and 0 < p < 1000]
return probe_ping_dict
def get_lowest_avg_ping_unfiltered_geo(ping_type, continent="", country=True, glob=False,
source_continent="", minimum=False):
result_pings = {}
probe_ping_dict = get_lowest_avg_pings(ping_type, continent, False, glob, source_continent)
for probe_id in probe_ping_dict:
lowest = ()
for dc in probe_ping_dict[probe_id]:
pings = probe_ping_dict[probe_id][dc]
if len(pings) > 0:
avg = sum(pings)/len(pings)
if lowest == ():
lowest = (dc, avg)
elif avg < lowest[1]:
lowest = (dc, avg)
if lowest != ():
if country:
co = probe_countries[probe_id]
if co not in result_pings:
result_pings[co] = []
else:
co = probe_continents[probe_id]
if co not in result_pings:
result_pings[co] = []
if minimum:
result_pings[co].append(min(probe_ping_dict[probe_id][lowest[0]]))
else:
result_pings[co] += probe_ping_dict[probe_id][lowest[0]]
return result_pings
```
## RIPE Atlas
```
conn_ripe = create_connection(db_path_ripe)
cursor_ripe = conn_ripe.cursor()
probe_countries_ripe = {}
probe_continents_ripe = {}
cursor_ripe.execute("select ID, Country, Continent from Probes where Country != ''")
for p_id, co, cont in cursor_ripe:
probe_countries_ripe[p_id] = co
probe_continents_ripe[p_id] = cont
cursor_ripe.execute("select ID from Probes where home = 1")
home_ids = [x[0] for x in cursor_ripe]
def get_lowest_avg_pings_ripe(ping_type, continent, filter_provider, glob, source_continent=""):
if glob:
if ping_type.lower() == "icmp":
probe_info_sql = """SELECT Pi.probe_id, Ping1, Ping2, Ping3, Ping4, Ping5, Pi.url,
D.name FROM Ping Pi JOIN Probes P ON Pi.probe_id = P.ID JOIN Datacenter D
ON D.Url = Pi.url WHERE P.Continent = D.Continent"""
elif ping_type.lower() == "tcp":
probe_info_sql = """WITH relevant_ids AS (
SELECT TI.ID FROM TracerouteInfo TI JOIN Datacenter D ON
D.ID = TI.datacenter JOIN Probes P ON P.ID = TI.probe_id JOIN Traceroute T ON T.ID = TI.ID
WHERE P.Continent = D.Continent and protocol = 'TCP'),
ranked_hops AS (
SELECT Traceroute_ID, rtt_after, ROW_NUMBER() OVER
(PARTITION BY Traceroute_ID, attempt ORDER BY hop_number DESC) AS rn FROM Hops
WHERE Traceroute_ID IN relevant_ids),
tracert_id_info AS (
SELECT TI.probe_id, TI.ID, TI.url, D.name FROM TracerouteInfo TI JOIN Datacenter D on
TI.datacenter = D.ID where TI.ID in relevant_ids)
SELECT T.probe_id, rtt_after, url,
name FROM ranked_hops R JOIN tracert_id_info T
ON R.Traceroute_ID = T.ID WHERE rn = 1"""
cursor_ripe.execute(probe_info_sql)
else:
if ping_type.lower() == "icmp":
probe_info_sql = """SELECT Pi.probe_id, Ping1, Ping2, Ping3, Ping4, Ping5, Pi.url,
D.name FROM Ping Pi JOIN Probes P ON Pi.probe_id = P.ID JOIN Datacenter D
ON D.Url = Pi.url WHERE P.Continent = ? and D.Continent = ?"""
elif ping_type.lower() == "tcp":
probe_info_sql = """WITH relevant_ids AS (
SELECT TI.ID FROM TracerouteInfo TI JOIN Datacenter D ON
D.ID = TI.datacenter JOIN Probes P ON P.ID = TI.probe_id JOIN Traceroute T ON T.ID = TI.ID
WHERE P.Continent = ? and D.Continent = ? and protocol = 'TCP'),
ranked_hops AS (
SELECT Traceroute_ID, rtt_after, ROW_NUMBER() OVER
(PARTITION BY Traceroute_ID, attempt ORDER BY hop_number DESC) AS rn FROM Hops
WHERE Traceroute_ID IN relevant_ids),
tracert_id_info AS (
SELECT TI.probe_id, TI.ID, TI.url, D.name FROM TracerouteInfo TI JOIN Datacenter D on
TI.datacenter = D.ID where TI.ID in relevant_ids)
SELECT T.probe_id, rtt_after, url,
name FROM ranked_hops R JOIN tracert_id_info T
ON R.Traceroute_ID = T.ID WHERE rn = 1"""
if source_continent:
cursor_ripe.execute(probe_info_sql, (source_continent, continent))
else:
cursor_ripe.execute(probe_info_sql, (continent, continent))
probe_ping_dict = {}
if ping_type.lower() == "icmp":
for probe in cursor_ripe:
probe_id = probe[0]
dc = probe[6]
if probe_id not in probe_ping_dict:
probe_ping_dict[probe_id] = {}
if filter_provider:
provider = probe[7]
if provider not in probe_ping_dict[probe_id]:
probe_ping_dict[probe_id][provider] = {}
if dc not in probe_ping_dict[probe_id][provider]:
probe_ping_dict[probe_id][provider][dc] = []
probe_ping_dict[probe_id][provider][dc] += [p for p in probe[1:6] if
p is not None and 0 < p < 1000]
else:
if dc not in probe_ping_dict[probe_id]:
probe_ping_dict[probe_id][dc] = []
probe_ping_dict[probe_id][dc] += [p for p in probe[1:6] if p is not None and 0 < p < 1000]
else:
for probe in cursor_ripe:
probe_id = probe[0]
dc = probe[2]
if probe_id not in probe_ping_dict:
probe_ping_dict[probe_id] = {}
if filter_provider:
provider = probe[3]
if provider not in probe_ping_dict[probe_id]:
probe_ping_dict[probe_id][provider] = {}
if dc not in probe_ping_dict[probe_id][provider]:
probe_ping_dict[probe_id][provider][dc] = []
if probe[1] > 0:
probe_ping_dict[probe_id][provider][dc].append(probe[1])
else:
if dc not in probe_ping_dict[probe_id]:
probe_ping_dict[probe_id][dc] = []
if probe[1] > 0:
probe_ping_dict[probe_id][dc].append(probe[1])
return probe_ping_dict
def get_lowest_avg_ping_unfiltered_geo_ripe(ping_type, continent="", country=True, glob=False, source_continent="", minimum=False):
result_pings = {}
probe_ping_dict = get_lowest_avg_pings_ripe(ping_type, continent, False, glob, source_continent)
for probe_id in probe_ping_dict:
lowest = ()
for dc in probe_ping_dict[probe_id]:
pings = probe_ping_dict[probe_id][dc]
if len(pings) > 0:
avg = sum(pings)/len(pings)
if lowest == ():
lowest = (dc, avg)
elif avg < lowest[1]:
lowest = (dc, avg)
if lowest != ():
if country:
co = probe_countries_ripe[probe_id]
if co not in result_pings:
result_pings[co] = []
else:
co = probe_continents_ripe[probe_id]
if co not in result_pings:
result_pings[co] = []
if minimum:
result_pings[co].append(min(probe_ping_dict[probe_id][lowest[0]]))
else:
result_pings[co] += probe_ping_dict[probe_id][lowest[0]]
return result_pings
```
# Figure 1 - Datacenter and vantage point world maps
```
world = geopandas.read_file(geopandas.datasets.get_path('naturalearth_lowres'))
world = world[(world.name!="Antarctica")] # remove Antarctica
cursor.execute("select Country, count(*) from Datacenters group by Country")
loc_sql_dict = {country: num for country, num in cursor}
country_count = []
for name, code in zip(world["name"], world["iso_a3"]):
if name == "France":
country_count.append(loc_sql_dict[country_codes["FRA"]])
elif name == "Norway":
country_count.append(loc_sql_dict[country_codes["NOR"]])
elif code == "-99":
country_count.append(float("NaN"))
elif country_codes[code] in loc_sql_dict:
country_count.append(loc_sql_dict[country_codes[code]])
else:
country_count.append(float("NaN"))
world["count"] = country_count
fig, ax = plt.subplots(figsize=(4.66, 2), subplot_kw={'projection': ccrs.PlateCarree()})
labels = ["1", "2-5", "6-10", "11-20", "$>$20"]
cmap = colors.ListedColormap(cols[:-2])
handles = [Line2D([0], [0], marker="o", color="w", markerfacecolor=col)
for col in cols]
world.plot(facecolor ="white", edgecolor="black", ax=ax, linewidth=0.2)
world.plot(column="count", cmap=cmap, scheme='UserDefined',
classification_kwds=dict(bins=[1,5,10,20]), ax=ax)
ax.set_extent([-180, 180, -60, 90], ccrs.PlateCarree())
ax.get_xaxis().set_ticks([])
ax.get_yaxis().set_ticks([])
ax.legend(handles, labels, handlelength=1, labelspacing=0.06, columnspacing=0.5, handletextpad=0.3,
loc="lower left", fancybox=False, edgecolor="k", fontsize="small")
fig.tight_layout()
plt.savefig("Figs/Fig_1a.pdf", bbox_inches="tight", pad_inches=0)
plt.show()
world = geopandas.read_file(geopandas.datasets.get_path('naturalearth_lowres'))
world = world[(world.name!="Antarctica")]
cursor.execute("""select Country, count(*) from Probes where ProbeID in (SELECT distinct(ProbeID)
from Ping union select distinct(ProbeID) from Traceroute)
and Country is not NULL and Country != '' group by Country order by count(*)""")
probe_sql_dict = {}
for country, num in cursor:
probe_sql_dict[country] = num
# add probe numbers to the dataframe
probe_numbers = []
for name, code in zip(world["name"], world["iso_a3"]):
if name == "France":
probe_numbers.append(probe_sql_dict[country_codes["FRA"]])
elif name == "Norway":
probe_numbers.append(probe_sql_dict[country_codes["NOR"]])
elif code == "-99":
probe_numbers.append(float("NaN"))
elif country_codes[code] in probe_sql_dict:
probe_numbers.append(probe_sql_dict[country_codes[code]])
else:
probe_numbers.append(float("NaN"))
world["probe_num"] = probe_numbers
fig, ax = plt.subplots(figsize=(4.66, 2), subplot_kw={'projection': ccrs.PlateCarree()})
labels = ["$<$50", "50-200", "200-1K", "1K-5K", "$>$5K"]
cmap = colors.ListedColormap(cols[:-2])
handles = [Line2D([0], [0], marker="o", color="w", markerfacecolor=col)
for col in cols]
world.plot(facecolor ="white", edgecolor="black", ax=ax, linewidth=0.2)
world.plot(column="probe_num", cmap=cmap, scheme='UserDefined',
classification_kwds=dict(bins=[50,200,1000,5000]), ax=ax)
ax.set_extent([-180, 180, -60, 90], ccrs.PlateCarree())
ax.get_xaxis().set_ticks([])
ax.get_yaxis().set_ticks([])
ax.legend(handles, labels, handlelength=1, labelspacing=0.06, columnspacing=0.5, handletextpad=0.3,
loc="lower left", fancybox=False, edgecolor="k", fontsize="small")
ax.text(0.35, 0.03, "EU 72K\ \ \ AS 31K\ \ \ NA 5.4K\ \ \ AF 4K\ \ \ SA 2.8K\ \ \ OC 351",
transform=ax.transAxes, fontsize="x-small", verticalalignment="bottom",
bbox=dict(boxstyle="Square, pad=0.25", facecolor="none", edgecolor="black", lw=0.5))
plt.savefig("Figs/Fig_1b.pdf", bbox_inches="tight", pad_inches=0)
plt.show()
```
# Figure 2 - RIPE Atlas VP map
```
world = geopandas.read_file(geopandas.datasets.get_path('naturalearth_lowres'))
world = world[(world.name!="Antarctica")]
cursor_ripe.execute("""select Country, count(*) from Probes where ID in (SELECT distinct(probe_id)
from Ping union select distinct(probe_id) from TracerouteInfo)
and Country is not NULL and Country != '' group by Country""")
probe_sql_dict = {}
for country, num in cursor_ripe:
probe_sql_dict[country] = num
probe_numbers = []
for name, code in zip(world["name"], world["iso_a3"]):
if name == "France":
probe_numbers.append(probe_sql_dict["FRA"])
elif name == "Norway":
probe_numbers.append(probe_sql_dict["NOR"])
elif code == "-99":
probe_numbers.append(float("NaN"))
elif code in probe_sql_dict:
probe_numbers.append(probe_sql_dict[code])
else:
probe_numbers.append(float("NaN"))
world["probe_num"] = probe_numbers
fig, ax = plt.subplots(figsize=(4.66, 2), subplot_kw={'projection': ccrs.PlateCarree()})
labels = ["$<$10", "10-50", "50-200", "$>$200"]
cmap = colors.ListedColormap(cols[:-3])
handles = [Line2D([0], [0], marker="o", color="w", markerfacecolor=col)
for col in cols]
world.plot(facecolor ="white", edgecolor="black", ax=ax, linewidth=0.2)
world.plot(column="probe_num", cmap=cmap, scheme='UserDefined',
classification_kwds=dict(bins=[10,50,200]), ax=ax)
ax.set_extent([-180, 180, -60, 90], ccrs.PlateCarree())
ax.get_xaxis().set_ticks([])
ax.get_yaxis().set_ticks([])
ax.legend(handles, labels, handlelength=1, labelspacing=0.06, columnspacing=0.5, handletextpad=0.3,
loc="lower left", fancybox=False, edgecolor="k", fontsize="small")
ax.text(0.35, 0.03, "EU 5574\ \ \ AS 1083\ \ \ NA 866\ \ \ AF 261\ \ \ SA 216\ \ \ OC 289",
transform=ax.transAxes, fontsize="x-small", verticalalignment="bottom",
bbox=dict(boxstyle="Square, pad=0.25", facecolor="none", edgecolor="black", lw=0.5))
plt.savefig("Figs/Fig_2.pdf", bbox_inches="tight", pad_inches=0)
plt.show()
```
# Figure 3 - Ping latency world map
```
dc_loc_sql = "select URL, Latitude, Longitude from Datacenters"
df = pd.read_sql(dc_loc_sql, conn)
gdf = geopandas.GeoDataFrame(df, geometry=geopandas.points_from_xy(df.Longitude, df.Latitude))
world = geopandas.read_file(geopandas.datasets.get_path("naturalearth_lowres"))
world = world[(world.name!="Antarctica")]
results = get_lowest_avg_ping_unfiltered_geo("tcp", glob=True)
for co in results:
results[co] = np.median(results[co]) if len(results[co]) > 0 else 0
pings = []
for name, code in zip(world["name"], world["iso_a3"]):
if name == "France":
pings.append(results[country_codes["FRA"]])
elif name == "Norway":
pings.append(results[country_codes["NOR"]])
elif code == "-99":
pings.append(float("nan"))
elif country_codes[code] in results:
pings.append(results[country_codes[code]])
else:
pings.append(float("nan"))
world["avg_ping"] = pings
fig, ax = plt.subplots(figsize=(4.66, 2), subplot_kw={'projection': ccrs.PlateCarree()})
cmap = colors.ListedColormap(cols[:-1])
handles = [Line2D([0], [0], marker="o", color="w", markerfacecolor=col)
for col in cols]
labels = ["$<$30 ms", "30-40 ms", "40-60 ms", "60-100 ms", "100-250 ms", "$>$250 ms"]
ax.coastlines(linewidth=0.3)
world.plot(column="avg_ping", cmap=cmap, scheme='UserDefined', classification_kwds=dict(bins=[30,60,100,250]),
ax=ax, alpha=0.8)
gdf.plot(marker="D", facecolor="red", edgecolor="black", ax=ax, markersize=4, linewidth=0.5, alpha=0.5)
ax.set_extent([-180, 180, -60, 90], ccrs.PlateCarree())
ax.get_xaxis().set_ticks([])
ax.get_yaxis().set_ticks([])
ax.legend(handles, labels, handlelength=1, labelspacing=0.06, columnspacing=0.5, handletextpad=0.3,
loc="lower left", fancybox=False, edgecolor="k", fontsize="small")
fig.tight_layout()
plt.savefig("Figs/Fig_3.pdf", bbox_inches = "tight", pad_inches = 0)
plt.show()
```
# Figure 4 - Ping latency CDF
```
tcp_pings = get_lowest_avg_ping_unfiltered_geo("tcp", country=False, glob=True)
fig, ax = plt.subplots(figsize=(3.37, 2))
ax.axvline(x=20, linewidth=1, color="red", linestyle="-", alpha=0.3)
ax.axvline(x=100, linewidth=1, color="blue", linestyle="-", alpha=0.3)
ax.axvline(x=250, linewidth=1, color="green", linestyle="-", alpha=0.3)
for cont in ["EU", "NA", "OC", "SA", "AS", "AF"]:
xs = np.sort(list(filter(lambda x: x < 400, tcp_pings[cont]))) # filter out outliers above 400 ms
ys = np.arange(1, len(xs) + 1) / len(xs)
indices = []
current = xs[0]
for i, x in enumerate(xs): # only take max y value at each x value to smoothen out the graph
if x != current:
current = x
indices.append(i - 1)
indices.append(len(ys) - 1)
xs = sorted(set(xs))
ys = [ys[i] for i in indices]
ax.plot(xs, ys, label=cont, color=continent_colors[cont], linestyle=lstyles[cont])
ax.set_xlabel("Latency [ms]")
ax.set_ylabel("Percentile")
ax.set_yticks(np.arange(0, 1.25, 0.25))
ax.xaxis.get_major_formatter()._usetex = False
ax.yaxis.get_major_formatter()._usetex = False
ax.legend(loc="lower right", fontsize="small", ncol=2, edgecolor="k", handlelength=1, labelspacing=0.06,
columnspacing=0.5, handletextpad=0.3, fancybox=False)
plt.grid(True, axis='y', linestyle='-', alpha=0.7, linewidth=0.5)
ax.text(22, -0.03, "MTP", color="red", size="small")
ax.text(102, -0.03, "PL", color="blue", size="small")
ax.text(210, -0.03, "HRT", color="green", size="small")
fig.tight_layout()
plt.savefig("Figs/Fig_4.pdf", bbox_inches = "tight", pad_inches = 0)
plt.show()
```
# Figure 5 - Speedchecker vs RIPE Atlas latency difference (takes ~3 hours to calculate and plot)
```
results_sc = get_lowest_avg_ping_unfiltered_geo("tcp", country=False, glob=True)
results_ripe = get_lowest_avg_ping_unfiltered_geo_ripe("tcp", country=False, glob=True)
results_diff = {}
# calculate means of the differences between every measurement from Speedchecker and every measurement
# from RIPE Atlas towards the closest datacenter on the same continent
for cont in results_sc:
results_diff[cont] = []
for num in results_sc[cont]:
tmp = []
for num2 in results_ripe[cont]:
tmp.append(num - num2)
results_diff[cont].append(np.mean(tmp))
fig, ax = plt.subplots(figsize=(3.37,2))
ax.axvline(x=0, linewidth=0.8, color="black", linestyle="-", alpha=0.3)
for cont in sorted(results_diff.keys()):
xs = np.sort(list(filter(lambda x: x < 400, results_diff[cont])))
ys = np.arange(1, len(xs) + 1) / len(xs)
indices = []
current = xs[0]
for i, x in enumerate(xs):
if x != current:
current = x
indices.append(i - 1)
indices.append(len(ys) - 1)
xs = sorted(set(xs))
ys = [ys[i] for i in indices]
ax.plot(xs, ys, label=cont, color=continent_colors[cont], linestyle=lstyles[cont])
ax.set_xlabel("Difference in latency [ms]")
ax.set_ylabel("Percentile")
ax.set_xlim([-100,100])
ax.set_yticks([0,0.25,0.5,0.75,1])
ax.xaxis.get_major_formatter()._usetex = False
ax.yaxis.get_major_formatter()._usetex = False
plt.grid(True, axis='y', linestyle='-', alpha=0.7, linewidth=0.5)
ax.text(-95, 0.4, "Speedchecker faster", fontsize="x-small")
ax.arrow(-32, 0.35, -55, 0, head_width=0.05, head_length=5, color="black")
ax.text(55, 0.4, "Atlas faster", fontsize="x-small")
ax.arrow(55, 0.35, 32, 0, head_width=0.05, head_length=5, color="black")
ax.legend(handlelength=1, labelspacing=0.06, columnspacing=0.5, handletextpad=0.3,
loc="upper left", fancybox=False, edgecolor="k", fontsize="small", ncol=2)
fig.tight_layout()
plt.savefig("Figs/Fig_5.pdf", bbox_inches = "tight", pad_inches = 0)
plt.show()
```
# Figure 6 - Intercontinental latency for Africa and South America
```
def adjust_box(plot):
plt.setp(plot['boxes'][0], facecolor=continent_colors[target1], linewidth=1)
plt.setp(plot['medians'][0], color='yellow')
plt.setp(plot['boxes'][1], facecolor=continent_colors[target2], linewidth=1)
plt.setp(plot['medians'][1], color='yellow')
plt.setp(plot['boxes'][2], facecolor=continent_colors[cont], linewidth=1)
plt.setp(plot['medians'][2], color='yellow')
cont = "AF"
target1, target2 = "EU", "NA"
countries = ["DZ", "EG", "ET", "KE", "MA", "SN", "TN", "ZA"]
results_inter1 = get_lowest_avg_ping_unfiltered_geo("tcp", continent=target1, source_continent=cont)
results_inter2 = get_lowest_avg_ping_unfiltered_geo("tcp", continent=target2, source_continent=cont)
results_intra = get_lowest_avg_ping_unfiltered_geo("tcp", continent=cont)
fig, ax = plt.subplots(figsize=(4.66, 2))
width = 0.5
for i, co in enumerate(countries):
positions = np.arange(i * 4, i * 4 + 3)
bp = ax.boxplot([results_inter1[co], results_inter2[co], results_intra[co]], positions=positions,
widths=width, showfliers=False, patch_artist=True)
adjust_box(bp)
xspan = 4
[ax.axvspan(i * xspan - 1, i * xspan - 1 + xspan, facecolor="k", alpha=0.2)
for i in range(len(countries))
if i % 2 == 1]
ax.set_xticks(np.arange(1, 1 + 4 * len(countries), 4))
ax.set_xticklabels(countries)
ax.set_xlim([-0.5, 4 * len(countries) - 1.5])
ax.set_ylabel("Latency [ms]")
ax.set_yticks(np.arange(0, 700, 100))
ax.xaxis.get_major_formatter()._usetex = False
ax.yaxis.get_major_formatter()._usetex = False
handles = [Patch(facecolor=continent_colors[target1]), Patch(facecolor=continent_colors[target2]),
Patch(facecolor=continent_colors[cont])]
labels = [target1, target2, cont]
ax.legend(handles, labels, handlelength=1, labelspacing=0.06, columnspacing=0.5, handletextpad=0.3,
loc="upper left", fancybox=False, edgecolor="k", fontsize="small", ncol=3)
plt.grid(True, axis='y', linestyle='--')
fig.tight_layout()
plt.savefig("Figs/Fig_6a.pdf".format(cont), bbox_inches = "tight", pad_inches = 0)
plt.show()
def adjust_box(plot):
plt.setp(plot['boxes'][0], facecolor=continent_colors[target], linewidth=1)
plt.setp(plot['medians'][0], color='yellow')
plt.setp(plot['boxes'][1], facecolor=continent_colors[cont], linewidth=1)
plt.setp(plot['medians'][1], color='yellow')
cont = "SA"
target = "NA"
results_inter = get_lowest_avg_ping_unfiltered_geo("tcp", continent=target, source_continent=cont)
results_intra = get_lowest_avg_ping_unfiltered_geo("tcp", continent=cont)
countries = sorted(results_inter.keys())
fig, ax = plt.subplots(figsize=(4.66, 2))
width = 0.5
for i, co in enumerate(countries):
positions = np.arange(i * 3, i * 3 + 2)
bp = ax.boxplot([results_inter[co], results_intra[co]], positions=positions, widths=width,
showfliers=False, patch_artist=True)
adjust_box(bp)
xspan = 3
[ax.axvspan(i * xspan - 1, i * xspan - 1 + xspan, facecolor="k", alpha=0.2)
for i in range(len(countries))
if i % 2 == 1]
ax.set_xticks(np.arange(0.5, 0.5 + 3 * len(countries), 3))
ax.set_xticklabels(countries)
ax.set_xlim([-0.5, 3 * len(countries) - 1.5])
ax.set_ylabel("Latency [ms]")
ax.set_yticks(np.arange(0, 500, 100))
ax.xaxis.get_major_formatter()._usetex = False
ax.yaxis.get_major_formatter()._usetex = False
handles = [Patch(facecolor=continent_colors[target]), Patch(facecolor=continent_colors[cont])]
labels = [target, cont]
ax.legend(handles, labels, handlelength=1, labelspacing=0.06, columnspacing=0.5, handletextpad=0.3,
loc="upper left", fancybox=False, edgecolor="k", fontsize="small", ncol=3)
plt.grid(True, axis='y', linestyle='--')
fig.tight_layout()
plt.savefig("Figs/Fig_6b.pdf".format(cont), bbox_inches = "tight", pad_inches = 0)
plt.show()
```
# Figure 7, 8, 9 & 17 - Last-mile latency analysis
## Data preparation (takes ~2 hours)
```
# Speedchecker
first_hop_latency = {}
home_ids = []
other_ids = []
tracert_ids_sql = """select TracerouteID from Hops where TracerouteID in (select TracerouteID from Traceroute T
join Probes P on T.ProbeID = P.ProbeID join Datacenters D on T.DestinationURL = D.URL
where P.Continent = D.Continent) group by TracerouteID having count(*) > 2"""
tracert_latency_sql = """with ranked_hops as (
select HopNumber, TracerouteID, RTT1, RTT2, RTT3, row_number() over
(partition by TracerouteID order by HopNumber desc) as rn from Hops where TracerouteID in {})
select TracerouteID, RTT1, RTT2, RTT3 from ranked_hops where rn = 1"""
hops_sql = """with ranked_hops as (
select HopIP, HopNumber, TracerouteID, RTT1, RTT2, RTT3, row_number() over
(partition by TracerouteID order by HopNumber asc) as rn, ASN
from Hops H join NodeInfo N on H.HopIP = N.IP where TracerouteID in {})
select HopIP, TracerouteID, HopNumber, RTT1, RTT2, RTT3, rn, ASN from ranked_hops where rn in (1, 2)"""
cursor.execute(tracert_ids_sql) # get traceroute ids of paths with more than 2 identified hops
filtered_ids = tuple([r[0] for r in cursor])
cursor.execute(tracert_latency_sql.format(filtered_ids))
tracert_latencies = {}
for row in cursor:
tracert_latencies[row[0]] = [x for x in row[1:] if x is not None and 0 < x < 1000]
cursor.execute(hops_sql.format(filtered_ids))
current_id = 0
skip = False
lat = ()
home_asns = []
other_asns = []
for row in cursor:
h_ip = row[0]
t_id = row[1]
hn = row[2]
rtts = [x for x in row[3:6] if x is not None and 0 < x < 1000]
rn = row[6]
asn = row[7]
if current_id != t_id:
current_id = t_id
skip = False
if rn == 1:
if is_private_ip(h_ip):
lat = (rtts, )
else:
skip = True
if len(rtts) > 0 and hn == 1:
other_ids.append(t_id)
other_asns.append(asn)
first_hop_latency[t_id] = rtts
elif not is_private_ip(h_ip) and not skip:
lat += (rtts, )
first_hop_latency[t_id] = lat
home_ids.append(t_id)
home_asns.append(asn)
sc_tracert_loc_sql = """select TracerouteID, P.Continent from Traceroute T join Probes P on
T.ProbeID = P.ProbeID join Datacenters D on T.DestinationURL = D.URL
where D.Continent = P.Continent"""
cursor.execute(sc_tracert_loc_sql)
relevant_ids_sc = {}
for t_id, co in cursor:
if co not in relevant_ids_sc:
relevant_ids_sc[co] = []
relevant_ids_sc[co].append(t_id)
r_ids = tuple(home_ids + other_ids)
probe_tracert_sql = "select ProbeID, TracerouteID, DestinationURL from Traceroute where TracerouteID in {}"
cursor.execute(probe_tracert_sql.format(r_ids))
probe_lat_home = collections.defaultdict(lambda: collections.defaultdict(list))
tracert_id_home = collections.defaultdict(lambda: collections.defaultdict(list))
probe_lat_other = collections.defaultdict(lambda: collections.defaultdict(list))
tracert_id_other = collections.defaultdict(lambda: collections.defaultdict(list))
for p_id, t_id, url in cursor:
if t_id in home_ids:
probe_lat_home[p_id][url] += tracert_latencies[t_id]
tracert_id_home[p_id][url].append(t_id)
else:
probe_lat_other[p_id][url] += tracert_latencies[t_id]
tracert_id_other[p_id][url].append(t_id)
closest_ids_home = []
for probe_id in tracert_id_home:
lowest = ()
for dc in probe_lat_home[probe_id]:
pings = probe_lat_home[probe_id][dc]
if len(pings) > 0:
avg = sum(pings)/len(pings)
if lowest == ():
lowest = (dc, avg)
elif avg < lowest[1]:
lowest = (dc, avg)
if lowest != ():
closest_ids_home += tracert_id_home[probe_id][lowest[0]]
closest_ids_other = []
for probe_id in tracert_id_other:
lowest = ()
for dc in probe_lat_other[probe_id]:
pings = probe_lat_other[probe_id][dc]
if len(pings) > 0:
avg = sum(pings)/len(pings)
if lowest == ():
lowest = (dc, avg)
elif avg < lowest[1]:
lowest = (dc, avg)
if lowest != ():
closest_ids_other += tracert_id_other[probe_id][lowest[0]]
sc_tracert_loc_sql = """select TracerouteID, P.Continent from Traceroute T join Probes P on
T.ProbeID = P.ProbeID join Datacenters D on T.DestinationURL = D.URL
where D.Continent = P.Continent and TracerouteID in {}"""
cursor.execute(sc_tracert_loc_sql.format(tuple(closest_ids_home + closest_ids_other)))
relevant_ids_home = collections.defaultdict(list)
relevant_ids_other = collections.defaultdict(list)
for t_id, co in cursor:
if t_id in closest_ids_home:
relevant_ids_home[co].append(t_id)
else:
relevant_ids_other[co].append(t_id)
sc_per_home = collections.defaultdict(list)
sc_per_other = collections.defaultdict(list)
for co in relevant_ids_home:
for t_id in relevant_ids_home[co]:
home = first_hop_latency[t_id][0]
isp = first_hop_latency[t_id][1]
dst = tracert_latencies[t_id]
if home and isp and dst:
home = np.mean(home)
isp = np.mean(isp)
dst = np.mean(dst)
sc_per_home[co].append(100*isp / dst)
for co in relevant_ids_other:
for t_id in relevant_ids_other[co]:
isp = first_hop_latency[t_id]
dst = tracert_latencies[t_id]
if isp and dst:
isp = np.mean(isp)
dst = np.mean(dst)
sc_per_other[co].append(100*isp / dst)
# RIPE Atlas
first_hop_latency_ripe = {}
tracert_ids_sql = """select TI.ID from TracerouteInfo TI join Datacenter D on D.ID = TI.datacenter
join Probes P on P.ID = TI.probe_id join Traceroute T on T.ID = TI.ID where
P.Continent = D.Continent and protocol = 'ICMP' and home = 1"""
tracert_latency_sql = """with ranked_hops AS (
SELECT Traceroute_ID, rtt_after, attempt, ROW_NUMBER() OVER
(PARTITION BY Traceroute_ID, attempt ORDER BY hop_number DESC) AS rn FROM Hops
WHERE Traceroute_ID IN {})
select Traceroute_ID, rtt_after, attempt from ranked_hops where rn = 1"""
hops_sql = """with ranked_hops as (
select dst_ip, hop_number, Traceroute_ID, rtt_after, row_number() over
(partition by Traceroute_ID, attempt order by hop_number asc) as rn, ASN, attempt
from Hops H join NodeInfo N on H.dst_ip = N.IP where Traceroute_ID in {})
select dst_ip, Traceroute_ID, hop_number, rtt_after, rn, ASN, attempt from ranked_hops where rn = 1"""
cursor_ripe.execute(tracert_ids_sql)
filtered_ids_ripe = tuple([r[0] for r in cursor_ripe])
cursor_ripe.execute(tracert_latency_sql.format(filtered_ids_ripe))
tracert_latencies_ripe = {}
for t_id, rtt, att in cursor_ripe:
if rtt is not None and 0 < rtt < 1000:
tracert_latencies_ripe[(t_id, att)] = rtt
cursor_ripe.execute(hops_sql.format(filtered_ids_ripe))
current_id = -1
current_att = -1
skip = False
lat = ()
home_asns_ripe = []
for row in cursor_ripe:
h_ip = row[0]
t_id = row[1]
hn = row[2]
rtt = row[3]
rn = row[4]
asn = row[5]
att = row[6]
first_hop_latency_ripe[(t_id, att)] = rtt
home_asns_ripe.append(asn)
ripe_tracert_loc_sql = """select T.ID, P.Continent from TracerouteInfo T join Probes P on T.probe_id = P.ID join
Datacenter D on T.url = D.url join Traceroute TR on T.ID = TR.ID
where P.Continent = D.Continent
and protocol = 'ICMP' and home = 1"""
cursor_ripe.execute(ripe_tracert_loc_sql)
relevant_ids_ripe = {}
for t_id, co in cursor_ripe:
if co not in relevant_ids_ripe:
relevant_ids_ripe[co] = []
relevant_ids_ripe[co].append(t_id)
# Calculations
sc_per = {}
sc_abs = {}
sc_other_per = {}
sc_other_abs = {}
sc_isp_per = {}
sc_isp_abs = {}
ripe_per = {}
ripe_abs = {}
for co in relevant_ids_sc:
for t_id in relevant_ids_sc[co]:
if t_id in home_ids:
if co not in sc_abs:
sc_per[co] = []
sc_isp_per[co] = []
sc_abs[co] = []
sc_isp_abs[co] = []
home = first_hop_latency[t_id][0]
isp = first_hop_latency[t_id][1]
dst = tracert_latencies[t_id]
if home and isp and dst:
home = np.mean(home)
isp = np.mean(isp)
dst = np.mean(dst)
sc_per[co].append(100*isp / dst)
sc_abs[co].append(isp)
sc_isp_per[co].append(100*(isp - home) / dst)
sc_isp_abs[co].append(isp - home)
elif t_id in other_ids:
if co not in sc_other_abs:
sc_other_per[co] = []
sc_other_abs[co] = []
isp = first_hop_latency[t_id]
dst = tracert_latencies[t_id]
if isp and dst:
isp = np.mean(isp)
dst = np.mean(dst)
sc_other_per[co].append(100*isp / dst)
sc_other_abs[co].append(isp)
for co in relevant_ids_ripe:
for t_id in relevant_ids_ripe[co]:
if co not in ripe_abs:
ripe_per[co] = []
ripe_abs[co] = []
for i in (0,1,2):
tup = (t_id, i)
if tup in first_hop_latency_ripe and tup in tracert_latencies_ripe:
ripe_per[co].append(100*first_hop_latency_ripe[tup] / tracert_latencies_ripe[tup])
ripe_abs[co].append(first_hop_latency_ripe[tup])
```
## Plotting
### Figure 7a - Percentage share of last mile latency compared to total latency grouped by continent for Speedchecker and RIPE Atlas
```
last_mile_home = [] # global percentages
last_mile_other = []
isp_per = []
last_mile_ripe = []
# filter out percentages >= 100
for co in sc_per:
sc_per[co] = [x for x in sc_per[co] if x < 100]
sc_other_per[co] = [x for x in sc_other_per[co] if x < 100]
sc_isp_per[co] = [x for x in sc_isp_per[co] if x < 100]
last_mile_home += sc_per[co]
last_mile_other += sc_other_per[co]
isp_per += sc_isp_per[co]
for co in ripe_per:
ripe_per[co] = [x for x in ripe_per[co] if x < 100]
last_mile_ripe += ripe_per[co]
def adjust_box(plot):
for i in range(4):
plt.setp(plot['boxes'][i], facecolor=cols[i], linewidth=1)
plt.setp(plot['medians'][i], color='yellow')
fig, ax = plt.subplots(figsize=(4.66, 2))
for i, cont in enumerate(continents):
width = 0.5
positions = np.arange(i * 5, i * 5 + 4)
bp = ax.boxplot([sc_per[cont], sc_other_per[cont], sc_isp_per[cont], ripe_per[cont]],
positions=positions, showfliers=False, patch_artist=True, widths=width)
adjust_box(bp)
bp = ax.boxplot([last_mile_home, last_mile_other, isp_per, last_mile_ripe],
positions=np.arange(len(continents) * 5, len(continents) * 5 + 4), showfliers=False,
patch_artist=True, widths=width)
adjust_box(bp)
xspan = 5
[ax.axvspan(i * xspan - 1, i * xspan - 1 + xspan, facecolor="k", alpha=0.2)
for i in range(len(continents) + 1)
if i % 2 == 1]
ax.set_xticks(np.arange(1.5, 1.5 + 5 * (len(continents) + 1), 5))
ax.set_xticklabels(continents + ["Global"])
ax.set_ylabel("Last-mile / total latency [\%]")
ax.set_yticks(np.arange(0, 125, 25))
ax.set_ylim([-5,125])
ax.xaxis.get_major_formatter()._usetex = False
ax.yaxis.get_major_formatter()._usetex = False
handles = [Patch(facecolor=col) for col in cols[:4]]
labels = ["SC home (USR-ISP)", "SC cell", "SC home (RTR-ISP)", "Atlas"]
ax.legend(handles, labels, loc="upper left", fontsize="small", edgecolor="k", handlelength=1,
labelspacing=0.06, columnspacing=0.5, handletextpad=0.3, fancybox=False, ncol=4)
plt.grid(True, axis="y", linestyle="--")
fig.tight_layout()
plt.savefig("Figs/Fig_7a.pdf", bbox_inches = "tight", pad_inches = 0)
plt.show()
```
### Figure 7b - Absolute last-mile latency grouped by continent for Speedchecker and RIPE Atlas
```
last_mile_home_abs = []
last_mile_other_abs = []
isp_abs = []
last_mile_ripe_abs = []
for co in sc_abs:
sc_abs[co] = [x for x in sc_abs[co] if x < 100]
sc_other_abs[co] = [x for x in sc_other_abs[co] if x < 100]
sc_isp_abs[co] = [x for x in sc_isp_abs[co] if 0 < x < 100]
last_mile_home_abs += sc_abs[co]
last_mile_other_abs += sc_other_abs[co]
isp_abs += sc_isp_abs[co]
for co in ripe_abs:
ripe_abs[co] = [x for x in ripe_abs[co] if x < 100]
last_mile_ripe_abs += ripe_abs[co]
def adjust_box(plot):
for i in range(4):
plt.setp(plot['boxes'][i], facecolor=cols[i], linewidth=1)
plt.setp(plot['medians'][i], color='yellow')
fig, ax = plt.subplots(figsize=(4.66, 2))
for i, cont in enumerate(continents):
width = 0.5
positions = np.arange(i * 5, i * 5 + 4)
bp = ax.boxplot([sc_abs[cont], sc_other_abs[cont], sc_isp_abs[cont], ripe_abs[cont]],
positions=positions, showfliers=False, patch_artist=True, widths=width)
adjust_box(bp)
bp = ax.boxplot([last_mile_home_abs, last_mile_other_abs, isp_abs, last_mile_ripe_abs],
positions=np.arange(len(continents) * 5, len(continents) * 5 + 4), showfliers=False,
patch_artist=True, widths=width)
adjust_box(bp)
xspan = 5
[ax.axvspan(i * 5 - 1, i * 5 - 1 + xspan, facecolor="k", alpha=0.2)
for i in range(len(continents) + 1)
if i % 2 == 1]
ax.set_xticks(np.arange(1.5, 1.5 + 5 * (len(continents) + 1), 5))
ax.set_xticklabels(continents + ["Global"])
ax.set_ylabel("Last-mile latency [ms]")
ax.xaxis.get_major_formatter()._usetex = False
ax.yaxis.get_major_formatter()._usetex = False
handles = [Patch(facecolor=col) for col in cols[:4]]
labels = ["SC home (USR-ISP)", "SC cell", "SC home (RTR-ISP)", "Atlas"]
ax.legend(handles, labels, loc="upper right", fontsize="small", edgecolor="k", handlelength=1,
labelspacing=0.06, columnspacing=0.5, handletextpad=0.3, fancybox=False, ncol=4)
plt.grid(True, axis="y", linestyle="--")
fig.tight_layout()
plt.savefig("Figs/Fig_7b.pdf", bbox_inches = "tight", pad_inches = 0)
plt.show()
```
### Figure 8 - Coefficient of Variation for last-mile latency of all probes grouped by continent
```
r_ids = tuple(home_ids + other_ids)
cursor.execute("""select Pr.ProbeID, TracerouteID, Pr.Continent from Traceroute T join Probes Pr on
T.ProbeID = Pr.ProbeID where TracerouteID in {}""".format(r_ids))
probes_map = {}
probes_loc = {}
for p_id, t_id, cont in cursor:
if p_id not in probes_map:
probes_map[p_id] = []
probes_map[p_id].append(t_id)
probes_loc[p_id] = cont
probe_vars = {}
probe_env = {}
for p_id in probes_map:
lats = []
for t_id in probes_map[p_id]:
if t_id in home_ids:
lats += first_hop_latency[t_id][1]
probe_env[p_id] = 0
else:
lats += first_hop_latency[t_id]
probe_env[p_id] = 1
probe_vars[p_id] = lats
vars_home = {cont: [] for cont in continents}
vars_other = {cont: [] for cont in continents}
for p_id in probe_vars:
if len(probe_vars[p_id]) >= 5:
if probe_env[p_id] == 0:
vars_home[probes_loc[p_id]].append(stats.variation(probe_vars[p_id]))
else:
vars_other[probes_loc[p_id]].append(stats.variation(probe_vars[p_id]))
fig, ax = plt.subplots(figsize=(4.66, 2))
pos1 = np.arange(0, 3 * len(continents), 3)
pos2 = np.arange(1, 3 * len(continents) + 1, 3)
ax.boxplot([vars_home[cont] for cont in continents], positions=pos1, patch_artist=True,
showfliers=False, boxprops=dict(facecolor=cols[0], lw=1), medianprops=dict(color="yellow"))
ax.boxplot([vars_other[cont] for cont in continents], positions=pos2, patch_artist=True,
showfliers=False, boxprops=dict(facecolor=cols[1], lw=1), medianprops=dict(color="yellow"))
ax.set_xticks(np.arange(0.5, 3 * len(continents), 3))
ax.set_xticklabels(continents)
ax.set_ylabel("Coeff. of Variation ($\mathrm{C_v}$)")
ax.set_yticks(np.arange(0, 3, 0.5))
ax.set_ylim([-0.1, 2.5])
ax.set_xlim([-0.5, 3 * len(continents) - 1.5])
xspan = 3
[ax.axvspan(i * xspan - 1, i * xspan - 1 + xspan, facecolor="k", alpha=0.2)
for i in range(len(continents) + 1)
if i % 2 == 1]
handles = [Patch(facecolor=cols[0]), Patch(facecolor=cols[1])]
labels = ["SC home (USR-ISP)", "SC cell"]
ax.legend(handles, labels, loc="upper left", fontsize="small", edgecolor="k", handlelength=1,
labelspacing=0.06, columnspacing=0.5, handletextpad=0.3, fancybox=False, ncol=4)
plt.grid(True, axis="y", linestyle="--")
plt.savefig("Figs/Fig_8.pdf", bbox_inches="tight", pad_inches=0)
plt.show()
```
### Figure 9 - Coefficient of variation of all probes in two countries per continent
```
countries = ("ZA", "MA", "JP", "IR", "GB", "UA", "US", "MX", "BR", "AR")
r_ids = tuple(home_ids + other_ids)
cursor.execute("""select Pr.ProbeID, TracerouteID, Pr.Country from Traceroute T join Probes Pr on
T.ProbeID = Pr.ProbeID where TracerouteID in {} and Pr.Country in {}""".format(r_ids, countries))
probes_map = {}
probes_loc = {}
for p_id, t_id, co in cursor:
if p_id not in probes_map:
probes_map[p_id] = []
probes_map[p_id].append(t_id)
probes_loc[p_id] = co
probe_vars = {}
probe_env = {}
for p_id in probes_map:
lats = []
for t_id in probes_map[p_id]:
if t_id in home_ids:
lats += first_hop_latency[t_id][1]
probe_env[p_id] = 0
else:
lats += first_hop_latency[t_id]
probe_env[p_id] = 1
probe_vars[p_id] = lats
vars_home = collections.defaultdict(list)
vars_other = collections.defaultdict(list)
for p_id in probe_vars:
if len(probe_vars[p_id]) >= 5:
if probe_env[p_id] == 0:
vars_home[probes_loc[p_id]].append(stats.variation(probe_vars[p_id]))
else:
vars_other[probes_loc[p_id]].append(stats.variation(probe_vars[p_id]))
def adjust_box(plot, index, t):
plt.setp(plot['boxes'][index], facecolor=type_colors[t], linewidth=1)
plt.setp(plot['medians'][index], color='yellow')
type_colors = {0: cols[0], 1: cols[1]}
fig, ax = plt.subplots(figsize=(4.66, 2))
counter = 0
tick_pos = []
for co in countries:
ts = []
data = []
if len(vars_home[co]) >= 5:
ts.append(0)
data.append(vars_home[co])
if len(vars_other[co]) >= 5:
ts.append(1)
data.append(vars_other[co])
positions = np.arange(counter, counter + len(ts))
bp = ax.boxplot(data, positions=positions, widths=0.5, showfliers=False, patch_artist=True)
for i, t in enumerate(ts):
adjust_box(bp, i, t)
tick_pos.append((counter + counter + len(ts) - 1)/2)
counter += len(ts) + 1
ax.set_xticks(tick_pos)
ax.set_xticklabels(countries, fontsize="small")
ax.set_ylabel("Coeff. of Variation ($\mathrm{C_v}$)")
ax.axvspan(-1, 3, facecolor="#8dd3c7", alpha=0.6)
ax.axvspan(3, 9, facecolor="#ffffb3", alpha=0.6)
ax.axvspan(9, 15, facecolor="#bebada", alpha=0.6)
ax.axvspan(15, 21, facecolor="#fb8072", alpha=0.6)
ax.axvspan(21, 27, facecolor="#80b1d3", alpha=0.6)
ax.set_xlim([-1, 27])
legend_keys = []
legend_labels = ["SC home (USR-ISP)", "SC cell"]
for t in type_colors:
legend_keys.append(Patch(facecolor = type_colors[t]))
ax.legend(legend_keys, legend_labels, loc="upper left",
ncol=2, edgecolor="k", handlelength=1, labelspacing=0.06,
columnspacing=0.5, handletextpad=0.3, fancybox=False, fontsize="small")
plt.grid(True, axis="y", linestyle="--")
plt.savefig("Figs/Fig_9.pdf", bbox_inches = "tight", pad_inches = 0)
plt.show()
```
## Figure 17 - Coefficient of variation of measurements of all probes towards the closest datacenter
```
total_home = []
total_other = []
for co in sc_per_home:
sc_per_home[co] = [x for x in sc_per_home[co] if x < 100]
sc_per_other[co] = [x for x in sc_per_other[co] if x < 100]
total_home += sc_per_home[co]
total_other += sc_per_other[co]
def adjust_box(plot):
for i in (0, 1):
plt.setp(plot['boxes'][i], facecolor=cols[i], linewidth=1)
plt.setp(plot['medians'][i], color='yellow')
fig, ax = plt.subplots(figsize=(4.66, 2))
for i, cont in enumerate(continents):
width = 0.5
positions = np.arange(i * 3, i * 3 + 2)
bp = ax.boxplot([sc_per_home[cont], sc_per_other[cont]],
positions=positions, showfliers=False, patch_artist=True, widths=width)
adjust_box(bp)
bp = ax.boxplot([total_home, total_other],
positions=np.arange(len(continents) * 3, len(continents) * 3 + 2), showfliers=False,
patch_artist=True, widths=width)
adjust_box(bp)
xspan = 3
[ax.axvspan(i * xspan - 1, i * xspan - 1 + xspan, facecolor="k", alpha=0.2)
for i in range(len(continents) + 1)
if i % 2 == 1]
ax.set_xticks(np.arange(0.5, 0.5 + 3 * (len(continents) + 1), 3))
ax.set_xticklabels(continents + ["Global"])
ax.set_ylabel("Last-mile / total latency [\%]")
ax.set_yticks(np.arange(0, 125, 25))
ax.set_ylim([-5,125])
ax.xaxis.get_major_formatter()._usetex = False
ax.yaxis.get_major_formatter()._usetex = False
handles = [Patch(facecolor=col) for col in cols[:4]]
labels = ["SC home (USR-ISP)", "SC cell"]
ax.legend(handles, labels, loc="upper left", fontsize="small", edgecolor="k", handlelength=1,
labelspacing=0.06, columnspacing=0.5, handletextpad=0.3, fancybox=False, ncol=4)
plt.grid(True, axis="y", linestyle="--")
plt.savefig("Figs/Fig_17.pdf", bbox_inches = "tight", pad_inches = 0)
plt.show()
```
# Figure 10 - AS-level path lengths for the different cloud providers
```
def adjust_box(plot, index, cont):
plt.setp(plot["boxes"][index], facecolor=continent_colors[cont], linewidth=1)
plt.setp(plot["medians"][index], color="black")
provider_abbrev = {"Alibaba": "BABA", "Amazon EC2": "AMZN",
"DigitalOcean": "DO", "Google": "GCP", "IBM": "IBM", "Linode": "LIN",
"Microsoft": "MSFT", "Oracle": "ORCL", "Vultr": "VLTR"}
as_hops_sql = """with id_info as (select TracerouteID, Provider, D.Continent from Traceroute T join Datacenters D on
T.DestinationURL = D.URL join Probes P on T.ProbeID = P.ProbeID where P.Continent = D.Continent),
t_start as (select ASN, TracerouteID from Traceroute T join Probes P on T.ProbeID = P.ProbeID
where TracerouteID in (select TracerouteID from id_info)),
t_hops as (select N.ASN, H.TracerouteID from Hops H join
NodeInfo N on H.HopIP = N.IP where H.TracerouteID in (select TracerouteID from id_info) and N.ASN != -1),
t_complete as (select * from t_start union all select * from t_hops)
select Provider, count(distinct ASN) as cnt from t_complete T join id_info I on
T.TracerouteID = I.TracerouteID group by T.TracerouteID having cnt > 1"""
cursor.execute(as_hops_sql)
as_counts = {}
for prov, num in cursor:
if prov != "Amazon Lightsail":
if prov not in as_counts:
as_counts[prov] = []
as_counts[prov].append(num)
target_providers = sorted(provider_abbrev.keys())
results = {}
for prov in target_providers:
total = len(as_counts[prov])
zero_as = (as_counts[prov].count(2) *100) / total
one_as = (as_counts[prov].count(3) * 100) / total
more_as = 100 - zero_as - one_as
results[prov] = [zero_as, one_as, more_as]
fig, ax = plt.subplots(figsize=(4.66, 2))
labels = ["direct", "1", "2+"]
x = np.arange(len(labels))
width = 0.08
pos=np.arange(-4, 5, 1)
for i in range(len(target_providers)):
index = i//2 if i%2 == 0 else (i//2 + 2) % len(target_providers)
ax.bar(x + pos[i]*width, results[target_providers[i]], width, color=cols[index],
label=provider_abbrev[target_providers[i]],
edgecolor="white", linewidth=0.1, zorder=1, hatch="/////" if i%2 == 1 else "")
ax.bar(x + pos[i]*width, results[target_providers[i]], width, color="none",
edgecolor="k", linewidth=1, zorder=2)
ax.set_ylabel("Percentage of Paths")
ax.set_ylim(0, 100)
ax.set_xlabel("Number of intermediate AS-level Hops")
ax.set_xticks(x)
ax.set_xticklabels(labels)
ax.xaxis.get_major_formatter()._usetex = False
ax.yaxis.get_major_formatter()._usetex = False
ax.legend(loc="upper right", fontsize="small", edgecolor="k", handlelength=1, labelspacing=0.06,
columnspacing=0.5, handletextpad=0.3, fancybox=False, ncol=3)
plt.grid(True, axis="y", linestyle="--")
fig.tight_layout()
plt.savefig("Figs/Fig_10.pdf", bbox_inches = "tight", pad_inches = 0)
plt.show()
```
# Figure 11 - Pervasiveness for the different cloud providers grouped by continent
```
def adjust_box(plot, index, cont):
plt.setp(plot['boxes'][index], facecolor=continent_colors[cont], linewidth=1)
plt.setp(plot['medians'][index], color='yellow')
cloud_count_sql = "with id_prov as (select TracerouteID, Provider, Continent from Traceroute T join Datacenters D " \
"on T.DestinationURL = D.URL)" \
"select count(case when ASN in {} then 1 end) as cnt, count(*), Provider, I.Continent " \
"from Hops H join NodeInfo N on H.HopIP = N.IP join id_prov I on H.TracerouteID = I.TracerouteID " \
"where H.TracerouteID in (select TracerouteID from Traceroute T join Datacenters D " \
"on T.DestinationURL = D.URL) group by H.TracerouteID having cnt > 0"
cloud_counts_per = {}
cursor.execute(cloud_count_sql.format(tuple(asns)))
for cloud, total, prov, cont in cursor:
if prov != "Amazon Lightsail":
if prov not in cloud_counts_per:
cloud_counts_per[prov] = {}
if cont not in cloud_counts_per[prov]:
cloud_counts_per[prov][cont] = []
if total > 2:
cloud_counts_per[prov][cont].append(cloud/total)
fig, ax = plt.subplots(figsize=(4.66, 2))
counter = 0
tick_pos = []
for prov in sorted(cloud_counts_per.keys()):
conts = sorted(cloud_counts_per[prov].keys())
data = [[y for y in cloud_counts_per[prov][x] if 0 < y < 1] for x in conts]
positions = np.arange(counter, counter + len(conts))
bp = ax.boxplot(data, positions=positions, widths=0.5, showfliers=False, patch_artist=True)
for i in range(len(conts)):
adjust_box(bp, i, conts[i])
tick_pos.append((counter + counter + len(conts) - 1)/2)
counter += len(conts) + 1
ax.set_xticks(tick_pos)
ax.set_xticklabels(provider_abbrev[x] for x in sorted(cloud_counts_per.keys()))
ax.set_ylim([0, 1])
ax.set_ylabel("Pervasiveness")
ax.axvspan(4, 11, facecolor="k", alpha=0.2)
ax.axvspan(15, 21, facecolor="k", alpha=0.2)
ax.axvspan(25, 30, facecolor="k", alpha=0.2)
ax.axvspan(37, 43, facecolor="k", alpha=0.2)
plt.grid(True, axis='y', linestyle='--')
legend_elements = []
for cont in continent_colors:
legend_elements.append(Patch(color=continent_colors[cont], label=cont))
ax.legend(handles=legend_elements, handlelength=1, labelspacing=0.06, columnspacing=0.5, handletextpad=0.3,
loc="upper right", fancybox=False, edgecolor="k", fontsize="small", ncol=2)
ax.xaxis.get_major_formatter()._usetex = False
ax.yaxis.get_major_formatter()._usetex = False
plt.savefig("Figs/Fig_11.pdf", bbox_inches = "tight", pad_inches = 0)
plt.show()
```
# Figure 12, 13, 18 & 19 - Country-specific routing analysis
## Data preparation
```
def create_peering_data(country, continent):
tracert_ids_sql = "select TracerouteID from Hops where TracerouteID in (select TracerouteID from Traceroute T " \
"join Probes P on T.ProbeID = P.ProbeID join Datacenters D on T.DestinationURL = D.URL " \
"where P.Country = ? and D.Continent = ?) group by TracerouteID having count(*) > 2"
tracert_no_ixp_sql = "with t_start as (select ASN, TracerouteID, 0 as HopNumber, 0 as IsIXP from Traceroute " \
"T join Probes P on T.ProbeID = P.ProbeID where TracerouteID in {})," \
"t_hops as (select N.ASN, H.TracerouteID, HopNumber, IsIXP from Hops H join " \
"NodeInfo N on H.HopIP = N.IP where TracerouteID in {})," \
"t_complete as (select * from t_start union all select * from t_hops)," \
"complete_ids as (select TracerouteID from t_complete group by TracerouteID having " \
"count(case when IsIXP = 1 then 1 end) = 0)" \
"select * from t_complete where TracerouteID in complete_ids order by TracerouteID"
tracert_ixp_sql = "with t_start as (select ASN, TracerouteID, 0 as HopNumber, 0 as IsIXP from Traceroute " \
"T join Probes P on T.ProbeID = P.ProbeID where TracerouteID in {})," \
"t_hops as (select N.ASN, H.TracerouteID, HopNumber, IsIXP from Hops H join " \
"NodeInfo N on H.HopIP = N.IP where TracerouteID in {})" \
"select * from t_start union all select * from t_hops order by TracerouteID"
cursor.execute(tracert_ids_sql, (country, continent))
filtered_ids = tuple([r[0] for r in cursor])
paths_no_ixp = {}
cursor.execute(tracert_no_ixp_sql.format(filtered_ids, filtered_ids))
for row in cursor:
asn = row[0]
t_id = row[1]
if row[2] == 0 and asn != -1:
paths_no_ixp[t_id] = (asn, )
else:
try:
if (paths_no_ixp[t_id])[-1] != asn and asn != -1:
paths_no_ixp[t_id] += (asn, )
except KeyError:
continue
result_no_ixp = [paths_no_ixp[k] for k in paths_no_ixp]
count_no_ixp = Counter(result_no_ixp)
result_no_ixp = list(set([x for x in result_no_ixp]))
paths_ixp = {}
cursor.execute(tracert_ixp_sql.format(filtered_ids, filtered_ids))
for row in cursor:
asn = row[0]
t_id = row[1]
if row[2] == 0 and asn != -1:
paths_ixp[t_id] = (asn, )
else:
try:
if (paths_ixp[t_id])[-1] != asn and asn != -1:
paths_ixp[t_id] += (asn, )
except KeyError:
continue
tracert_ixp_ids = [k for k in paths_ixp if k not in paths_no_ixp]
result_ixp = [(paths_ixp[k], 1) if k in tracert_ixp_ids else (paths_ixp[k], 0) for k in paths_ixp]
count_ixp = Counter(result_ixp)
result_ixp = list(set([x for x in result_ixp]))
tracert_dest_sql = "select TracerouteID, Country from Traceroute T join Datacenters D on T.DestinationURL = D.URL " \
"where TracerouteID in {}"
tracert_latency_sql = "with ranked_hops as (" \
"select HopNumber, TracerouteID, RTT1, RTT2, RTT3, row_number() over " \
"(partition by TracerouteID order by HopNumber desc) as rn from Hops where TracerouteID in {})" \
"select TracerouteID, RTT1, RTT2, RTT3 from ranked_hops where rn = 1"
path_lengths_sql = "select TracerouteID, count(*) from Hops where TracerouteID in {} group by TracerouteID"
cursor.execute(tracert_dest_sql.format(filtered_ids))
tracert_dest = {}
for t_id, cn in cursor:
tracert_dest[t_id] = cn
cursor.execute(tracert_latency_sql.format(filtered_ids))
tracert_latencies = {}
for row in cursor:
tracert_latencies[row[0]] = [x for x in row[1:] if x is not None and 0 < x < 1000]
cursor.execute(path_lengths_sql.format(filtered_ids))
path_lengths = {}
for t_id, num in cursor:
path_lengths[t_id] = num
paths_filtered_ixp = {}
paths_filtered_ixp_counts = {}
for t_id in paths_ixp:
first = paths_ixp[t_id][0]
last = paths_ixp[t_id][-1]
tup = (first, last)
if get_pdb_info(first)[1] != "Content" and last in asns and None not in tup:
tup = (tup, 1) if t_id in tracert_ixp_ids else (tup, 0)
path = paths_ixp[t_id]
if tup not in paths_filtered_ixp:
paths_filtered_ixp[tup] = {}
if path not in paths_filtered_ixp[tup]:
paths_filtered_ixp[tup][path] = []
paths_filtered_ixp[tup][path].append(t_id)
if tup[0] not in paths_filtered_ixp_counts:
paths_filtered_ixp_counts[tup[0]] = 0
paths_filtered_ixp_counts[tup[0]] += 1
for tup in paths_filtered_ixp:
for path in paths_filtered_ixp[tup]:
paths_filtered_ixp[tup][path] = (round(len(paths_filtered_ixp[tup][path]) * 100 /
paths_filtered_ixp_counts[tup[0]], 3), paths_filtered_ixp[tup][path])
paths_sorted_ixp = {}
for tup in sorted(list(paths_filtered_ixp.keys()), key=lambda x: x[0]):
sorted_keys = sorted(paths_filtered_ixp[tup], key=lambda x: paths_filtered_ixp[tup][x][0], reverse=True)
paths_sorted_ixp[tup] = {}
for path in sorted_keys:
paths_sorted_ixp[tup][path] = paths_filtered_ixp[tup][path]
# Save traceroute data in csv in peering directory with more detailed information on intermediate ASs,
# destination countries, percentage shares of paths etc.
with open("peering/asn_path_map_" + country + ".csv", mode="w", encoding="utf-8") as f:
f.write("source;intermediate;destination;with_ixp;percentage;total_paths;provider;avg_path_length;" \
"avg_latency;destination_countries;ids\n")
for tup in paths_sorted_ixp:
for path in paths_sorted_ixp[tup]:
src = (str(path[0]) + ": " + get_pdb_info(path[0])[0])
dest = (str(path[-1]) + ": " + get_pdb_info(path[-1])[0])
ixp = tup[1]
per, ids = paths_sorted_ixp[tup][path]
total_num = paths_filtered_ixp_counts[tup[0]]
p_len = round(np.mean([path_lengths[x] for x in ids]), 2)
dest_counts = Counter(list(map(lambda x: tracert_dest[x], ids))).most_common()
dest_counts = list(map(str, dest_counts))
latency_dict = {}
for t_id in ids:
cn = tracert_dest[t_id]
if cn not in latency_dict:
latency_dict[cn] = []
latency_dict[cn] += tracert_latencies[t_id]
latency = [(dst, round(np.mean(latency_dict[dst]), 3)) if len(latency_dict[dst]) != 0
else (dst, -1) for dst in latency_dict]
latency = list(map(str, sorted(latency, key=lambda x: x[1])))
if len(path) == 2:
imd = " "
else:
imd = ", ".join(tuple(map(lambda x: "(" + str(x) + ": " + get_pdb_info(x)[0] + ")", \
path[1:-1])))
f.write(src + ";" + imd + ";" + dest + ";" + str(ixp) + ";" + str(per) + ";" + str(total_num) + ";"
+ asns_dict[path[-1]] + ";" + str(p_len) + ";" + ",".join(latency) + ";"
+ ",".join(dest_counts) + ";" + str(ids) + "\n")
return tracert_dest, tracert_latencies
```
## Plotting
### Figure 12 - Germany
```
class TextHandlerB(HandlerBase):
def create_artists(self, legend, text ,xdescent, ydescent,
width, height, fontsize, trans):
tx = Text(width/2.,height/2, text, fontsize=fontsize,
ha="center", va="center", fontweight="bold")
return [tx]
tracert_dest, tracert_latencies = create_peering_data("DE", "EU")
Legend.update_default_handler_map({str : TextHandlerB()})
df_ixp = pd.read_csv("peering/asn_path_map_DE.csv", delimiter=";",
converters={"intermediate": lambda x: x.split(",") if len(x) > 1 else ""})
# filter out likely erroneous paths and group them up
df_ixp = df_ixp[(df_ixp["total_paths"] > 20) & (df_ixp["percentage"] > 5)]
df_ixp["intermediate"] = df_ixp["intermediate"].apply(lambda x: len(x))
group_funcs = {"percentage": sum, "destination_countries": ",".join, "total_paths": max, "ids": ",".join}
df_ixp_grouped = df_ixp.groupby(["source", "intermediate", "destination", "with_ixp", "provider"]).agg(group_funcs).reset_index()
df_ixp_max = df_ixp_grouped.sort_values("percentage", ascending=False).drop_duplicates(["source", "destination"]).sort_index()
isps = ["3209", "3320", "6805", "6830", "8881"]
providers = list(map(str, sorted(list(asns_dict.keys()), key=lambda x: asns_dict[x])))
result = {}
path_type = {}
ixp_dict = {}
num_paths = {}
for row in df_ixp_max.itertuples(index=False):
tup = (row.source.split(":")[0], row.destination.split(":")[0])
result[tup] = row.percentage
ixp_dict[tup] = row.with_ixp
num_paths[tup] = row.total_paths
if row.intermediate >= 2:
path_type[tup] = 2
else:
path_type[tup] = row.intermediate
hmap_list = []
hmap_types = []
table_list = []
for isp in isps:
res_list = []
res_types = []
num_path_list = []
for provider in providers:
tup = (isp, provider)
if tup in result:
num_path_list.append(num_paths[tup])
res_list.append(result[tup])
if ixp_dict[tup] == 0:
res_types.append(path_type[tup])
else:
res_types.append(path_type[tup] + 10)
else:
num_path_list.append(0)
res_list.append(0)
res_types.append(-1)
hmap_list.append(res_list)
hmap_types.append(res_types)
table_list.append(num_path_list)
fig, ax1 = plt.subplots(figsize=(4.66, 3))
hmap_cols = ["red", "orange", "#377eb8", "yellowgreen", "green"]
cmap = colors.ListedColormap(hmap_cols)
heatmap = ax1.imshow(hmap_list, cmap=cmap, alpha=0, vmin=100, vmax=500)
divider = make_axes_locatable(ax1)
cax = divider.append_axes("right", size="5%", pad=0.05)
bounds = np.arange(0, 120, 20)
cb = matplotlib.colorbar.ColorbarBase(cax, cmap=cmap, boundaries=bounds, ticks=bounds, orientation="vertical")
cb.set_label("Percentage")
isp_dict = {"3209": "Vodafone", "3320": "D. Telekom", "6805": "Telefónica", "6830": "Liberty",
"8881": "1\&1"}
isp_labels = ["{} (AS {})".format(isp_dict[asn], asn) for asn in isp_dict]
provider_labels = [asns_dict[x] for x in list(map(int,providers))]
provider_labels = [provider_abbrev[x] for x in provider_labels]
ax1.set_xticks(np.arange(len(providers)))
ax1.set_yticks(np.arange(len(isps)))
ax1.set_xticklabels(provider_labels, fontsize="small")
ax1.set_yticklabels(isp_labels, fontsize="small")
plt.setp(ax1.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
for i in range(len(isps)):
for j in range(len(providers)):
if hmap_types[i][j] == -1:
ax1.text(j, i, "-", ha="center", va="center", color="black", fontsize=15)
else:
if hmap_types[i][j] == 0 or hmap_types[i][j] == 10:
fill = "full"
elif hmap_types[i][j] == 1 or hmap_types[i][j] == 11:
fill = "left"
else:
fill = "none"
ax1.plot(j, i, fillstyle=fill, markerfacecoloralt="white",
marker="o", markerfacecolor=hmap_cols[int(hmap_list[i][j]) // 20], markersize=8, color="w",
markeredgecolor=hmap_cols[int(hmap_list[i][j]) // 20])
if hmap_types[i][j] >= 10:
ax1.annotate("$\mathrm{^1}$", xy = (j + 0.2, i + 0.05))
handles = [Line2D([0], [0], marker="o", color="w", markerfacecolor="black", markersize=7,
markeredgecolor="black"),
Line2D([0], [0], fillstyle="left", marker="o", color="w", markerfacecolor="black",
markerfacecoloralt="white", markersize=7, markeredgecolor="black"),
Line2D([0], [0], marker="o", color="w", markerfacecolor="none",
markeredgecolor="black", markersize=7),
"$\mathrm{^1}$"]
labels = ["direct", "1 AS", "2+ AS", "IXP"]
fig.legend(handles=handles, labels=labels, bbox_to_anchor=(0.28, 0.73, 1., .102), loc="lower left",
ncol=5, borderaxespad=0., edgecolor="black", handletextpad=0.1, fontsize="small", fancybox=False,
columnspacing=1)
fig.tight_layout()
plt.savefig("Figs/Fig_12a.pdf", bbox_inches="tight")
plt.show()
country = "DE"
target_country = "GB"
def adjust_box(plot, index, pt):
plt.setp(plot['boxes'][index], facecolor=type_colors[pt], linewidth=1)
plt.setp(plot['medians'][index], color='yellow')
type_colors = {0: cols[0], 1: cols[1]}
latencies = {}
for row in df_ixp_max.itertuples(index=False):
src = row.source.split(":")[0]
prov = row.provider
ids = [y for x in eval(row.ids) for y in x] if type(eval(row.ids)) == tuple else eval(row.ids)
lats = [y for x in ids for y in tracert_latencies[x] if tracert_dest[x] == target_country]
if prov not in latencies:
latencies[prov] = {}
pt = 0 if row.intermediate == 0 else 1
if pt not in latencies[prov]:
latencies[prov][pt] = []
latencies[prov][pt] += lats
fig, ax = plt.subplots(figsize=(4.66, 2))
counter = 0
tick_pos = []
for prov in sorted(latencies.keys()):
pts = list(filter(lambda x: len(latencies[prov][x]) > 10, sorted(latencies[prov].keys())))
data = [latencies[prov][x] for x in pts]
positions = np.arange(counter, counter + len(pts))
if len(data) > 0:
bp = ax.boxplot(data, positions=positions, widths=0.5, showfliers=False, patch_artist=True)
for i, pt in enumerate(pts):
adjust_box(bp, i, pt)
tick_pos.append((counter + counter + len(pts) - 1)/2)
counter += len(pts) + 1
else:
del latencies[prov]
ax.set_xticks(tick_pos)
ax.set_xticklabels([provider_abbrev[x] for x in sorted(latencies.keys())], fontsize="small")
ax.set_ylabel("Latency [ms]")
ax.set_yticks(np.arange(0,140,20))
ax.xaxis.get_major_formatter()._usetex = False
ax.yaxis.get_major_formatter()._usetex = False
ax.axvspan(1, 3, facecolor="k", alpha=0.2)
ax.axvspan(6, 8, facecolor="k", alpha=0.2)
ax.axvspan(11, 14, facecolor="k", alpha=0.2)
ax.axvspan(16, 18, facecolor="k", alpha=0.2)
legend_keys = []
legend_labels = ["direct peering", "intermediate AS"]
for pt in type_colors:
legend_keys.append(Patch(facecolor = type_colors[pt]))
ax.legend(legend_keys, legend_labels, loc="upper left",
ncol=2, edgecolor="k", handlelength=1, labelspacing=0.06,
columnspacing=0.5, handletextpad=0.3, fancybox=False, fontsize="small")
plt.grid(True, axis="y", linestyle="--")
fig.tight_layout()
plt.savefig("Figs/Fig_12b.pdf", bbox_inches = "tight", pad_inches = 0)
plt.show()
```
### Figure 13 - Japan
```
class TextHandlerB(HandlerBase):
def create_artists(self, legend, text ,xdescent, ydescent,
width, height, fontsize, trans):
tx = Text(width/2.,height/2, text, fontsize=fontsize,
ha="center", va="center", fontweight="bold")
return [tx]
tracert_dest, tracert_latencies = create_peering_data("JP", "AS")
Legend.update_default_handler_map({str : TextHandlerB()})
df_ixp = pd.read_csv("peering/asn_path_map_JP.csv", delimiter=";",
converters={"intermediate": lambda x: x.split(",") if len(x) > 1 else ""})
occurence_df_ixp = df_ixp.groupby("source")[["total_paths"]].sum().reset_index()
df_ixp = df_ixp[(df_ixp["total_paths"] > 20) & (df_ixp["percentage"] > 5)]
df_ixp["intermediate"] = df_ixp["intermediate"].apply(lambda x: len(x))
group_funcs = {"percentage": sum, "destination_countries": ",".join, "total_paths": max, "ids": ",".join}
df_ixp_grouped = df_ixp.groupby(["source", "intermediate", "destination", "with_ixp", "provider"]).agg(group_funcs).reset_index()
df_ixp_max = df_ixp_grouped.sort_values("percentage", ascending=False).drop_duplicates(["source", "destination"]).sort_index()
isps = ["2516", "2518", "4713", "17511", "17676"]
providers = list(map(str, sorted(list(asns_dict.keys()), key=lambda x: asns_dict[x])))
result = {}
path_type = {}
ixp_dict = {}
num_paths = {}
for row in df_ixp_max.itertuples(index=False):
tup = (row.source.split(":")[0], row.destination.split(":")[0])
result[tup] = row.percentage
ixp_dict[tup] = row.with_ixp
num_paths[tup] = row.total_paths
if row.intermediate >= 2:
path_type[tup] = 2
else:
path_type[tup] = row.intermediate
hmap_list = []
hmap_types = []
table_list = []
for isp in isps:
res_list = []
res_types = []
num_path_list = []
for provider in providers:
tup = (isp, provider)
if tup in result:
num_path_list.append(num_paths[tup])
res_list.append(result[tup])
if ixp_dict[tup] == 0:
res_types.append(path_type[tup])
else:
res_types.append(path_type[tup] + 10)
else:
num_path_list.append(0)
res_list.append(0)
res_types.append(-1)
hmap_list.append(res_list)
hmap_types.append(res_types)
table_list.append(num_path_list)
fig, ax1 = plt.subplots(figsize=(4.66, 3))
hmap_cols = ["red", "orange", "#377eb8", "yellowgreen", "green"]
cmap = colors.ListedColormap(hmap_cols)
heatmap = ax1.imshow(hmap_list, cmap=cmap, alpha=0, vmin=100, vmax=500)
divider = make_axes_locatable(ax1)
cax = divider.append_axes("right", size="5%", pad=0.05)
bounds = np.arange(0, 120, 20)
cb = matplotlib.colorbar.ColorbarBase(cax, cmap=cmap, boundaries=bounds, ticks=bounds, orientation="vertical")
cb.set_label("Percentage")
isp_dict = {"2516": "KDDI", "2518": "BIGLOBE", "4713": "NTT", "17511": "OPTAGE",
"17676": "SoftBank"}
isp_labels = ["{} (AS {})".format(isp_dict[asn], asn) for asn in isp_dict]
provider_labels = [asns_dict[x] for x in list(map(int,providers))]
provider_labels = [provider_abbrev[x] for x in provider_labels]
ax1.set_xticks(np.arange(len(providers)))
ax1.set_yticks(np.arange(len(isps)))
ax1.set_xticklabels(provider_labels, fontsize="small")
ax1.set_yticklabels(isp_labels, fontsize="small")
plt.setp(ax1.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
for i in range(len(isps)):
for j in range(len(providers)):
if hmap_types[i][j] == -1:
ax1.text(j, i, "-", ha="center", va="center", color="black", fontsize=15)
else:
if hmap_types[i][j] == 0 or hmap_types[i][j] == 10:
fill = "full"
elif hmap_types[i][j] == 1 or hmap_types[i][j] == 11:
fill = "left"
else:
fill = "none"
ax1.plot(j, i, fillstyle=fill, markerfacecoloralt="white",
marker="o", markerfacecolor=hmap_cols[int(hmap_list[i][j]) // 20], markersize=8, color="w",
markeredgecolor=hmap_cols[int(hmap_list[i][j]) // 20])
if hmap_types[i][j] >= 10:
ax1.annotate("$\mathrm{^1}$", xy = (j + 0.2, i + 0.05))
handles = [Line2D([0], [0], marker="o", color="w", markerfacecolor="black", markersize=7,
markeredgecolor="black"),
Line2D([0], [0], fillstyle="left", marker="o", color="w", markerfacecolor="black",
markerfacecoloralt="white", markersize=7, markeredgecolor="black"),
Line2D([0], [0], marker="o", color="w", markerfacecolor="none",
markeredgecolor="black", markersize=7),
"$\mathrm{^1}$"]
labels = ["direct", "1 AS", "2+ AS", "IXP"]
fig.legend(handles=handles, labels=labels, bbox_to_anchor=(0.28, 0.73, 1., .102), loc="lower left",
ncol=5, borderaxespad=0., edgecolor="black", handletextpad=0.1, fontsize="small", fancybox=False,
columnspacing=1)
fig.tight_layout()
plt.savefig("Figs/Fig_13a.pdf", bbox_inches="tight")
plt.show()
country = "JP"
target_country = "IN"
def adjust_box(plot, index, pt):
plt.setp(plot['boxes'][index], facecolor=type_colors[pt], linewidth=1)
plt.setp(plot['medians'][index], color='yellow')
type_colors = {0: cols[0], 1: cols[1]}
latencies = {}
for row in df_ixp_max.itertuples(index=False):
src = row.source.split(":")[0]
prov = row.provider
ids = [y for x in eval(row.ids) for y in x] if type(eval(row.ids)) == tuple else eval(row.ids)
lats = [y for x in ids for y in tracert_latencies[x] if tracert_dest[x] == target_country]
if prov not in latencies:
latencies[prov] = {}
pt = 0 if row.intermediate == 0 else 1
if pt not in latencies[prov]:
latencies[prov][pt] = []
latencies[prov][pt] += lats
fig, ax = plt.subplots(figsize=(4.66, 2))
counter = 0
tick_pos = []
for prov in sorted(latencies.keys()):
pts = list(filter(lambda x: len(latencies[prov][x]) > 10, sorted(latencies[prov].keys())))
data = [latencies[prov][x] for x in pts]
positions = np.arange(counter, counter + len(pts))
if len(data) > 0:
bp = ax.boxplot(data, positions=positions, widths=0.5, showfliers=False, patch_artist=True)
for i, pt in enumerate(pts):
adjust_box(bp, i, pt)
tick_pos.append((counter + counter + len(pts) - 1)/2)
counter += len(pts) + 1
else:
del latencies[prov]
ax.set_xticks(tick_pos)
ax.set_xticklabels([provider_abbrev[x] for x in sorted(latencies.keys())], fontsize="small")
ax.set_ylabel("Latency [ms]")
ax.set_yticks(np.arange(0,500,100))
ax.set_ylim([0, 400])
ax.xaxis.get_major_formatter()._usetex = False
ax.yaxis.get_major_formatter()._usetex = False
ax.axvspan(1, 3, facecolor="k", alpha=0.2)
ax.axvspan(5, 7, facecolor="k", alpha=0.2)
ax.axvspan(9, 11, facecolor="k", alpha=0.2)
legend_keys = []
legend_labels = ["direct peering", "intermediate AS"]
for pt in type_colors:
legend_keys.append(Patch(facecolor = type_colors[pt]))
ax.legend(legend_keys, legend_labels, loc="lower left",
ncol=2, edgecolor="k", handlelength=1, labelspacing=0.06,
columnspacing=0.5, handletextpad=0.3, fancybox=False, fontsize="small")
plt.grid(True, axis="y", linestyle="--")
fig.tight_layout()
plt.savefig("Figs/Fig_13b.pdf", bbox_inches = "tight", pad_inches = 0)
plt.show()
```
### Figure 18 - Ukraine
```
class TextHandlerB(HandlerBase):
def create_artists(self, legend, text ,xdescent, ydescent,
width, height, fontsize, trans):
tx = Text(width/2.,height/2, text, fontsize=fontsize,
ha="center", va="center", fontweight="bold")
return [tx]
tracert_dest, tracert_latencies = create_peering_data("UA", "EU")
Legend.update_default_handler_map({str : TextHandlerB()})
df_ixp = pd.read_csv("peering/asn_path_map_UA.csv", delimiter=";",
converters={"intermediate": lambda x: x.split(",") if len(x) > 1 else ""})
occurence_df_ixp = df_ixp.groupby("source")[["total_paths"]].sum().reset_index()
df_ixp = df_ixp[(df_ixp["total_paths"] > 20) & (df_ixp["percentage"] > 5)]
df_ixp["intermediate"] = df_ixp["intermediate"].apply(lambda x: len(x))
group_funcs = {"percentage": sum, "destination_countries": ",".join, "total_paths": max, "ids": ",".join}
df_ixp_grouped = df_ixp.groupby(["source", "intermediate", "destination", "with_ixp", "provider"]).agg(group_funcs).reset_index()
df_ixp_max = df_ixp_grouped.sort_values("percentage", ascending=False).drop_duplicates(["source", "destination"]).sort_index()
isps = ["3255", "3326", "6849", "15895", "25229"]
providers = list(map(str, sorted(list(asns_dict.keys()), key=lambda x: asns_dict[x])))
result = {}
path_type = {}
ixp_dict = {}
num_paths = {}
for row in df_ixp_max.itertuples(index=False):
tup = (row.source.split(":")[0], row.destination.split(":")[0])
result[tup] = row.percentage
ixp_dict[tup] = row.with_ixp
num_paths[tup] = row.total_paths
if row.intermediate >= 2:
path_type[tup] = 2
else:
path_type[tup] = row.intermediate
hmap_list = []
hmap_types = []
table_list = []
for isp in isps:
res_list = []
res_types = []
num_path_list = []
for provider in providers:
tup = (isp, provider)
if tup in result:
num_path_list.append(num_paths[tup])
res_list.append(result[tup])
if ixp_dict[tup] == 0:
res_types.append(path_type[tup])
else:
res_types.append(path_type[tup] + 10)
else:
num_path_list.append(0)
res_list.append(0)
res_types.append(-1)
hmap_list.append(res_list)
hmap_types.append(res_types)
table_list.append(num_path_list)
fig, ax1 = plt.subplots(figsize=(4.66, 3))
hmap_cols = ["red", "orange", "#377eb8", "yellowgreen", "green"]
cmap = colors.ListedColormap(hmap_cols)
heatmap = ax1.imshow(hmap_list, cmap=cmap, alpha=0, vmin=100, vmax=500)
divider = make_axes_locatable(ax1)
cax = divider.append_axes("right", size="5%", pad=0.05)
bounds = np.arange(0, 120, 20)
cb = matplotlib.colorbar.ColorbarBase(cax, cmap=cmap, boundaries=bounds, ticks=bounds, orientation="vertical")
cb.set_label("Percentage")
isp_dict = {"3255": "UARnet", "3326": "Datagroup", "6849": "UKRTELNET", "15895": "Kyivstar",
"25229": "Volia"}
isp_labels = ["{} (AS {})".format(isp_dict[asn], asn) for asn in isp_dict]
provider_labels = [asns_dict[x] for x in list(map(int,providers))]
provider_labels = [provider_abbrev[x] for x in provider_labels]
ax1.set_xticks(np.arange(len(providers)))
ax1.set_yticks(np.arange(len(isps)))
ax1.set_xticklabels(provider_labels, fontsize="small")
ax1.set_yticklabels(isp_labels, fontsize="small")
plt.setp(ax1.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
for i in range(len(isps)):
for j in range(len(providers)):
if hmap_types[i][j] == -1:
ax1.text(j, i, "-", ha="center", va="center", color="black", fontsize=15)
else:
if hmap_types[i][j] == 0 or hmap_types[i][j] == 10:
fill = "full"
elif hmap_types[i][j] == 1 or hmap_types[i][j] == 11:
fill = "left"
else:
fill = "none"
ax1.plot(j, i, fillstyle=fill, markerfacecoloralt="white",
marker="o", markerfacecolor=hmap_cols[int(hmap_list[i][j]) // 20], markersize=8, color="w",
markeredgecolor=hmap_cols[int(hmap_list[i][j]) // 20])
if hmap_types[i][j] >= 10:
ax1.annotate("$\mathrm{^1}$", xy = (j + 0.2, i + 0.05))
handles = [Line2D([0], [0], marker="o", color="w", markerfacecolor="black", markersize=7,
markeredgecolor="black"),
Line2D([0], [0], fillstyle="left", marker="o", color="w", markerfacecolor="black",
markerfacecoloralt="white", markersize=7, markeredgecolor="black"),
Line2D([0], [0], marker="o", color="w", markerfacecolor="none",
markeredgecolor="black", markersize=7),
"$\mathrm{^1}$"]
labels = ["direct", "1 AS", "2+ AS", "IXP"]
fig.legend(handles=handles, labels=labels, bbox_to_anchor=(0.28, 0.73, 1., .102), loc="lower left",
ncol=5, borderaxespad=0., edgecolor="black", handletextpad=0.1, fontsize="small", fancybox=False,
columnspacing=1)
fig.tight_layout()
plt.savefig("Figs/Fig_18a.pdf", bbox_inches="tight")
plt.show()
country = "UA"
target_country = "GB"
def adjust_box(plot, index, pt):
plt.setp(plot['boxes'][index], facecolor=type_colors[pt], linewidth=1)
plt.setp(plot['medians'][index], color='yellow')
type_colors = {0: cols[0], 1: cols[1]}
latencies = {}
for row in df_ixp_max.itertuples(index=False):
src = row.source.split(":")[0]
prov = row.provider
ids = [y for x in eval(row.ids) for y in x] if type(eval(row.ids)) == tuple else eval(row.ids)
lats = [y for x in ids for y in tracert_latencies[x] if tracert_dest[x] == target_country]
if prov not in latencies:
latencies[prov] = {}
pt = 0 if row.intermediate == 0 else 1
if pt not in latencies[prov]:
latencies[prov][pt] = []
latencies[prov][pt] += lats
fig, ax = plt.subplots(figsize=(4.66, 2))
counter = 0
tick_pos = []
for prov in sorted(latencies.keys()):
pts = list(filter(lambda x: len(latencies[prov][x]) > 10, sorted(latencies[prov].keys())))
data = [latencies[prov][x] for x in pts]
positions = np.arange(counter, counter + len(pts))
if len(data) > 0:
bp = ax.boxplot(data, positions=positions, widths=0.5, showfliers=False, patch_artist=True)
for i, pt in enumerate(pts):
adjust_box(bp, i, pt)
tick_pos.append((counter + counter + len(pts) - 1)/2)
counter += len(pts) + 1
else:
del latencies[prov]
ax.set_xticks(tick_pos)
ax.set_xticklabels([provider_abbrev[x] for x in sorted(latencies.keys())], fontsize="small")
ax.set_ylabel("Latency [ms]")
ax.set_yticks(np.arange(0,250,50))
ax.xaxis.get_major_formatter()._usetex = False
ax.yaxis.get_major_formatter()._usetex = False
ax.axvspan(1, 4, facecolor="k", alpha=0.2)
ax.axvspan(7, 10, facecolor="k", alpha=0.2)
ax.axvspan(13, 16, facecolor="k", alpha=0.2)
ax.axvspan(19, 21, facecolor="k", alpha=0.2)
legend_keys = []
legend_labels = ["direct peering", "intermediate AS"]
for pt in type_colors:
legend_keys.append(Patch(facecolor = type_colors[pt]))
ax.legend(legend_keys, legend_labels, loc="upper left",
ncol=2, edgecolor="k", handlelength=1, labelspacing=0.06,
columnspacing=0.5, handletextpad=0.3, fancybox=False, fontsize="small")
plt.grid(True, axis="y", linestyle="--")
fig.tight_layout()
plt.savefig("Figs/Fig_18b.pdf", bbox_inches = "tight", pad_inches = 0)
plt.show()
```
### Figure 19 - Bahrain
```
class TextHandlerB(HandlerBase):
def create_artists(self, legend, text ,xdescent, ydescent,
width, height, fontsize, trans):
tx = Text(width/2.,height/2, text, fontsize=fontsize,
ha="center", va="center", fontweight="bold")
return [tx]
tracert_dest, tracert_latencies = create_peering_data("BH", "AS")
Legend.update_default_handler_map({str : TextHandlerB()})
df_ixp = pd.read_csv("peering/asn_path_map_BH.csv", delimiter=";",
converters={"intermediate": lambda x: x.split(",") if len(x) > 1 else ""})
occurence_df_ixp = df_ixp.groupby("source")[["total_paths"]].sum().reset_index()
df_ixp = df_ixp[(df_ixp["total_paths"] > 20) & (df_ixp["percentage"] > 5)]
df_ixp["intermediate"] = df_ixp["intermediate"].apply(lambda x: len(x))
group_funcs = {"percentage": sum, "destination_countries": ",".join, "total_paths": max, "ids": ",".join}
df_ixp_grouped = df_ixp.groupby(["source", "intermediate", "destination", "with_ixp", "provider"]).agg(group_funcs).reset_index()
df_ixp_max = df_ixp_grouped.sort_values("percentage", ascending=False).drop_duplicates(["source", "destination"]).sort_index()
isps = ["5416", "31452", "39273", "51375"]
providers = list(map(str, sorted(list(asns_dict.keys()), key=lambda x: asns_dict[x])))
result = {}
path_type = {}
ixp_dict = {}
num_paths = {}
for row in df_ixp_max.itertuples(index=False):
tup = (row.source.split(":")[0], row.destination.split(":")[0])
result[tup] = row.percentage
ixp_dict[tup] = row.with_ixp
num_paths[tup] = row.total_paths
if row.intermediate >= 2:
path_type[tup] = 2
else:
path_type[tup] = row.intermediate
hmap_list = []
hmap_types = []
table_list = []
for isp in isps:
res_list = []
res_types = []
num_path_list = []
for provider in providers:
tup = (isp, provider)
if tup in result:
num_path_list.append(num_paths[tup])
res_list.append(result[tup])
if ixp_dict[tup] == 0:
res_types.append(path_type[tup])
else:
res_types.append(path_type[tup] + 10)
else:
num_path_list.append(0)
res_list.append(0)
res_types.append(-1)
hmap_list.append(res_list)
hmap_types.append(res_types)
table_list.append(num_path_list)
fig, ax1 = plt.subplots(figsize=(4.66, 3))
hmap_cols = ["red", "orange", "#377eb8", "yellowgreen", "green"]
cmap = colors.ListedColormap(hmap_cols)
heatmap = ax1.imshow(hmap_list, cmap=cmap, alpha=0, vmin=100, vmax=500)
divider = make_axes_locatable(ax1)
cax = divider.append_axes("right", size="5%", pad=0.05)
bounds = np.arange(0, 120, 20)
cb = matplotlib.colorbar.ColorbarBase(cax, cmap=cmap, boundaries=bounds, ticks=bounds, orientation="vertical")
cb.set_label("Percentage")
isp_dict = {"5416": "Batelco", "31452": "ZAIN", "39273": "Kalaam", "51375": "stc"}
isp_labels = ["{} (AS {})".format(isp_dict[asn], asn) for asn in isp_dict]
provider_labels = [asns_dict[x] for x in list(map(int,providers))]
provider_labels = [provider_abbrev[x] for x in provider_labels]
ax1.set_xticks(np.arange(len(providers)))
ax1.set_yticks(np.arange(len(isps)))
ax1.set_xticklabels(provider_labels, fontsize="small")
ax1.set_yticklabels(isp_labels, fontsize="small")
plt.setp(ax1.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
for i in range(len(isps)):
for j in range(len(providers)):
if hmap_types[i][j] == -1:
ax1.text(j, i, "-", ha="center", va="center", color="black", fontsize=15)
else:
if hmap_types[i][j] == 0 or hmap_types[i][j] == 10:
fill = "full"
elif hmap_types[i][j] == 1 or hmap_types[i][j] == 11:
fill = "left"
else:
fill = "none"
ax1.plot(j, i, fillstyle=fill, markerfacecoloralt="white",
marker="o", markerfacecolor=hmap_cols[int(hmap_list[i][j] -0.01) // 20],
markersize=8, color="w",
markeredgecolor=hmap_cols[int(hmap_list[i][j] -0.01) // 20])
if hmap_types[i][j] >= 10:
ax1.annotate("$\mathrm{^1}$", xy = (j + 0.2, i + 0.05))
handles = [Line2D([0], [0], marker="o", color="w", markerfacecolor="black", markersize=7,
markeredgecolor="black"),
Line2D([0], [0], fillstyle="left", marker="o", color="w", markerfacecolor="black",
markerfacecoloralt="white", markersize=7, markeredgecolor="black"),
Line2D([0], [0], marker="o", color="w", markerfacecolor="none",
markeredgecolor="black", markersize=7),
"$\mathrm{^1}$"]
labels = ["direct", "1 AS", "2+ AS", "IXP"]
fig.legend(handles=handles, labels=labels, bbox_to_anchor=(0.28, 0.73, 1., .102), loc="lower left",
ncol=5, borderaxespad=0., edgecolor="black", handletextpad=0.1, fontsize="small", fancybox=False,
columnspacing=1)
fig.tight_layout()
plt.savefig("Figs/Fig_19a.pdf", bbox_inches="tight")
plt.show()
country = "BH"
target_country = "IN"
def adjust_box(plot, index, pt):
plt.setp(plot['boxes'][index], facecolor=type_colors[pt], linewidth=1)
plt.setp(plot['medians'][index], color='yellow')
type_colors = {0: cols[0], 1: cols[1]}
latencies = {}
for row in df_ixp_max.itertuples(index=False):
src = row.source.split(":")[0]
prov = row.provider
ids = [y for x in eval(row.ids) for y in x] if type(eval(row.ids)) == tuple else eval(row.ids)
lats = [y for x in ids for y in tracert_latencies[x] if tracert_dest[x] == target_country]
if prov not in latencies:
latencies[prov] = {}
pt = 0 if row.intermediate == 0 else 1
if pt not in latencies[prov]:
latencies[prov][pt] = []
latencies[prov][pt] += lats
fig, ax = plt.subplots(figsize=(4.66, 2))
counter = 0
tick_pos = []
for prov in sorted(latencies.keys()):
pts = list(filter(lambda x: len(latencies[prov][x]) > 10, sorted(latencies[prov].keys())))
data = [latencies[prov][x] for x in pts]
positions = np.arange(counter, counter + len(pts))
if len(data) > 0:
bp = ax.boxplot(data, positions=positions, widths=0.5, showfliers=False, patch_artist=True)
for i, pt in enumerate(pts):
adjust_box(bp, i, pt)
tick_pos.append((counter + counter + len(pts) - 1)/2)
counter += len(pts) + 1
else:
del latencies[prov]
ax.set_xticks(tick_pos)
ax.set_xticklabels([provider_abbrev[x] for x in sorted(latencies.keys())], fontsize="small")
ax.set_xlim([-1,13])
ax.set_ylabel("Latency [ms]")
ax.set_yticks(np.arange(0,600,100))
ax.xaxis.get_major_formatter()._usetex = False
ax.yaxis.get_major_formatter()._usetex = False
ax.axvspan(1, 3, facecolor="k", alpha=0.2)
ax.axvspan(6, 8, facecolor="k", alpha=0.2)
ax.axvspan(11, 13, facecolor="k", alpha=0.2)
legend_keys = []
legend_labels = ["direct peering", "intermediate AS"]
for pt in type_colors:
legend_keys.append(Patch(facecolor = type_colors[pt]))
ax.legend(legend_keys, legend_labels, loc="upper left",
ncol=2, edgecolor="k", handlelength=1, labelspacing=0.06,
columnspacing=0.5, handletextpad=0.3, fancybox=False, fontsize="small")
plt.grid(True, axis="y", linestyle="--")
fig.tight_layout()
plt.savefig("Figs/Fig_19b.pdf", bbox_inches = "tight", pad_inches = 0)
plt.show()
```
# Figure 14 - Vantage point density heatmap of all Speedchecker probes
```
cursor.execute("""select Latitude, Longitude from Probes where ProbeID in (SELECT distinct(ProbeID)
from Ping union select distinct(ProbeID) from Traceroute)
and Country is not NULL and Country != '' and Latitude is not NULL and Longitude is not NULL""")
lat, lon = [], []
for la, lo in cursor:
lat.append(la)
lon.append(lo)
fig, ax = plt.subplots(figsize=(4.66, 2))
m = Basemap(projection='cyl',llcrnrlat=-60,urcrnrlat=90,\
llcrnrlon=-180,urcrnrlon=180,resolution='l', ax=ax)
m.drawcoastlines()
m.drawmapboundary(fill_color='white')
x, y = m(lon, lat)
x = np.array(x)
y = np.array(y)
xy = np.vstack([x, y])
z = gaussian_kde(xy)(xy)
idx = z.argsort()
x, y, z = x[idx], y[idx], z[idx]
points=ax.scatter(x, y, c=z, s=1, marker='.', cmap='winter')
ax.text(0.35, 0.03, "EU 72K\ \ \ AS 31K\ \ \ NA 5.4K\ \ \ AF 4K\ \ \ SA 2.8K\ \ \ OC 351",
transform=ax.transAxes, fontsize="x-small", verticalalignment="bottom",
bbox=dict(boxstyle="Square, pad=0.25", facecolor="none", edgecolor="black", lw=0.5))
plt.savefig("Figs/Fig_14.pdf", bbox_inches="tight", pad_inches=0)
plt.savefig("Figs/Fig_14.png", dpi=500, bbox_inches="tight", pad_inches=0)
plt.show()
```
# Figure 15 - ICMP vs TCP latency comparison
```
tcp_pings = get_lowest_avg_ping_unfiltered_geo("tcp", country=False, glob=True)
icmp_pings = get_lowest_avg_ping_unfiltered_geo("icmp", country=False, glob=True)
fig, ax = plt.subplots(figsize=(4.66,2))
counter = 0
tick_pos = []
for cont in continents:
data = [list(filter(lambda x: x < 400, tcp_pings[cont])), list(filter(lambda x: x < 400, icmp_pings[cont]))]
positions = np.arange(counter, counter + 2)
bp = ax.boxplot(data, positions=positions, widths=0.5, showfliers=False, patch_artist=True)
for index in (0, 1):
plt.setp(bp['boxes'][index], facecolor=cols[index])
plt.setp(bp['medians'][index], color='yellow')
tick_pos.append((counter + counter + 1)/2)
counter += 3
xspan = 3
[ax.axvspan(i * xspan - 1, i * xspan - 1 + xspan, facecolor="k", alpha=0.2)
for i in range(len(continents))
if i % 2 == 1]
ax.set_xticks(tick_pos)
ax.set_xticklabels(continents)
ax.set_ylabel("Latency[ms]")
ax.set_xlim([-0.5, 3 * len(continents) - 1.5])
ax.xaxis.get_major_formatter()._usetex = False
ax.yaxis.get_major_formatter()._usetex = False
handles = [Patch(facecolor=cols[0]), Patch(facecolor=cols[1])]
labels = ["ICMP", "TCP"]
ax.legend(handles, labels, handlelength=1, labelspacing=0.06, columnspacing=0.5, handletextpad=0.3,
loc="upper right", fancybox=False, edgecolor="k", fontsize="small", ncol=3)
plt.grid(True, axis='y', linestyle='--')
plt.savefig("Figs/Fig_15.pdf", bbox_inches = "tight", pad_inches = 0)
plt.show()
```
# Figure 16 - (City, ASN, Datacenter) latency comparison between RIPE Atlas and Speedchecker
## Data preparation
```
hops_sql = """select TracerouteID, City, ASN from Hops H join NodeInfo N on H.HopIP = N.IP where
City != 'Unknown' and ASN != -1 and HopNumber = 2"""
tracert_latency_sql = """with ranked_hops as (
select TracerouteID, RTT1, RTT2, RTT3, ROW_NUMBER() OVER
(PARTITION BY TracerouteID ORDER BY HopNumber DESC) AS rn FROM Hops
WHERE TracerouteID IN {}),
tracert_id_info AS (
SELECT TracerouteID, DestinationURL FROM Traceroute T
JOIN Datacenters D on D.URL = T.DestinationURL where TracerouteID in {})
SELECT RTT1, RTT2, RTT3, DestinationURL, R.TracerouteID FROM ranked_hops R JOIN
tracert_id_info T ON R.TracerouteID = T.TracerouteID WHERE rn = 1"""
location_info = {}
cursor.execute(hops_sql)
for t_id, city, asn in cursor:
location_info[t_id] = (city, asn)
r_ids = tuple(location_info.keys())
cursor.execute(tracert_latency_sql.format(r_ids, r_ids))
latencies = collections.defaultdict(lambda: collections.defaultdict(list))
for row in cursor:
lats = [x for x in row[:3] if x and 0 < x < 1000]
latencies[row[3]][location_info[row[4]]] += lats
hops_sql = """with relevant_ids as (
select TI.ID from TracerouteInfo TI join Traceroute T on TI.ID = T.ID join Probes P on
TI.probe_id = P.ID where protocol = 'ICMP' and P.home = 1)
select Traceroute_ID, City, ASN from Hops H join NodeInfo N on H.dst_ip = N.IP
where City is not null and City != 'unknown' and City != 'Unknown' and ASN is not null and
hop_number = 2 and attempt = 0 and Traceroute_ID in relevant_ids"""
tracert_latency_sql = """with ranked_hops AS (
SELECT Traceroute_ID, rtt_after, ROW_NUMBER() OVER
(PARTITION BY Traceroute_ID, attempt ORDER BY hop_number DESC) AS rn FROM Hops
WHERE Traceroute_ID IN {}),
tracert_id_info AS (
SELECT TI.ID, TI.url FROM TracerouteInfo TI JOIN Datacenter D on
TI.datacenter = D.ID where TI.ID in {})
SELECT rtt_after, url, R.Traceroute_ID
name FROM ranked_hops R JOIN tracert_id_info T
ON R.Traceroute_ID = T.ID WHERE rn = 1"""
location_info_ripe = {}
cursor_ripe.execute(hops_sql)
for t_id, city, asn in cursor_ripe:
location_info_ripe[t_id] = (city, asn)
r_ids_ripe = tuple(location_info_ripe.keys())
cursor_ripe.execute(tracert_latency_sql.format(r_ids_ripe, r_ids_ripe))
latencies_ripe = collections.defaultdict(lambda: collections.defaultdict(list))
for row in cursor_ripe:
latencies_ripe[row[1]][location_info_ripe[row[2]]].append(row[0])
matches = {}
for url in latencies:
if url in latencies_ripe:
tuples = latencies[url].keys() & latencies_ripe[url].keys()
if tuples:
matches[url] = tuples
cities_sql = """select distinct City, Continent from NodeInfo where City is not null and City != 'Unknown'"""
cursor.execute(cities_sql)
city_map = {}
for city, cont in cursor:
city_map[city] = cont
data = collections.defaultdict(list)
for url in matches:
for tup in matches[url]:
if len(latencies[url][tup]) > 5 and len(latencies_ripe[url][tup]) > 5:
data[city_map[tup[0]]].append(np.median(latencies[url][tup]) - np.median(latencies_ripe[url][tup]))
```
## Plot
```
fig, ax = plt.subplots(figsize=(3.37,2))
ax.axvline(x=0, linewidth=0.8, color="black", linestyle="-", alpha=0.3)
for cont in data:
xs = np.sort(data[cont])
ys = np.arange(1, len(data[cont]) + 1) / len(data[cont])
ax.plot(xs, ys, label=cont, color=continent_colors[cont], linestyle=lstyles[cont])
ax.set_xlabel("Difference in latency [ms]")
ax.set_ylabel("Percentile")
ax.set_xlim([-50,50])
ax.set_xticks(range(-50, 75, 25))
ax.set_yticks([0,0.25,0.5,0.75,1])
ax.xaxis.get_major_formatter()._usetex = False
ax.yaxis.get_major_formatter()._usetex = False
ax.legend(loc="upper left", fontsize="small", ncol=3, edgecolor="k", handlelength=1, labelspacing=0.06,
columnspacing=0.5, handletextpad=0.3, fancybox=False)
plt.grid(True, axis="y", linestyle="-", alpha=0.7, linewidth=0.5)
ax.text(-48, 0.1, "Speedchecker faster", fontsize="x-small")
ax.text(28, 0.1, "Atlas faster", fontsize="x-small")
plt.savefig("Figs/Fig_16.pdf", bbox_inches = "tight", pad_inches = 0)
plt.show()
```
| github_jupyter |
# Dirichlet process mixtures for density estimation
Author: [Austin Rochford](https://github.com/AustinRochford/)
## Dirichlet processes
The [Dirichlet process](https://en.wikipedia.org/wiki/Dirichlet_process) is a flexible probability distribution over the space of distributions. Most generally, a probability distribution, $P$, on a set $\Omega$ is a [measure](https://en.wikipedia.org/wiki/Measure_(mathematics%29) that assigns measure one to the entire space ($P(\Omega) = 1$). A Dirichlet process $P \sim \textrm{DP}(\alpha, P_0)$ is a measure that has the property that, for every finite [disjoint](https://en.wikipedia.org/wiki/Disjoint_sets) partition $S_1, \ldots, S_n$ of $\Omega$,
$$(P(S_1), \ldots, P(S_n)) \sim \textrm{Dir}(\alpha P_0(S_1), \ldots, \alpha P_0(S_n)).$$
Here $P_0$ is the base probability measure on the space $\Omega$. The precision parameter $\alpha > 0$ controls how close samples from the Dirichlet process are to the base measure, $P_0$. As $\alpha \to \infty$, samples from the Dirichlet process approach the base measure $P_0$.
Dirichlet processes have several properties that make then quite suitable to [MCMC](https://en.wikipedia.org/wiki/Markov_chain_Monte_Carlo) simulation.
1. The posterior given [i.i.d.](https://en.wikipedia.org/wiki/Independent_and_identically_distributed_random_variables) observations $\omega_1, \ldots, \omega_n$ from a Dirichlet process $P \sim \textrm{DP}(\alpha, P_0)$ is also a Dirichlet process with
$$P\ |\ \omega_1, \ldots, \omega_n \sim \textrm{DP}\left(\alpha + n, \frac{\alpha}{\alpha + n} P_0 + \frac{1}{\alpha + n} \sum_{i = 1}^n \delta_{\omega_i}\right),$$
where $\delta$ is the [Dirac delta measure](https://en.wikipedia.org/wiki/Dirac_delta_function)
$$\begin{align*}
\delta_{\omega}(S)
& = \begin{cases}
1 & \textrm{if } \omega \in S \\
0 & \textrm{if } \omega \not \in S
\end{cases}
\end{align*}.$$
2. The posterior predictive distribution of a new observation is a compromise between the base measure and the observations,
$$\omega\ |\ \omega_1, \ldots, \omega_n \sim \frac{\alpha}{\alpha + n} P_0 + \frac{1}{\alpha + n} \sum_{i = 1}^n \delta_{\omega_i}.$$
We see that the prior precision $\alpha$ can naturally be interpreted as a prior sample size. The form of this posterior predictive distribution also lends itself to Gibbs sampling.
2. Samples, $P \sim \textrm{DP}(\alpha, P_0)$, from a Dirichlet process are discrete with probability one. That is, there are elements $\omega_1, \omega_2, \ldots$ in $\Omega$ and weights $w_1, w_2, \ldots$ with $\sum_{i = 1}^{\infty} w_i = 1$ such that
$$P = \sum_{i = 1}^\infty w_i \delta_{\omega_i}.$$
3. The [stick-breaking process](https://en.wikipedia.org/wiki/Dirichlet_process#The_stick-breaking_process) gives an explicit construction of the weights $w_i$ and samples $\omega_i$ above that is straightforward to sample from. If $\beta_1, \beta_2, \ldots \sim \textrm{Beta}(1, \alpha)$, then $w_i = \beta_i \prod_{j = 1}^{j - 1} (1 - \beta_j)$. The relationship between this representation and stick breaking may be illustrated as follows:
1. Start with a stick of length one.
2. Break the stick into two portions, the first of proportion $w_1 = \beta_1$ and the second of proportion $1 - w_1$.
3. Further break the second portion into two portions, the first of proportion $\beta_2$ and the second of proportion $1 - \beta_2$. The length of the first portion of this stick is $\beta_2 (1 - \beta_1)$; the length of the second portion is $(1 - \beta_1) (1 - \beta_2)$.
4. Continue breaking the second portion from the previous break in this manner forever. If $\omega_1, \omega_2, \ldots \sim P_0$, then
$$P = \sum_{i = 1}^\infty w_i \delta_{\omega_i} \sim \textrm{DP}(\alpha, P_0).$$
We can use the stick-breaking process above to easily sample from a Dirichlet process in Python. For this example, $\alpha = 2$ and the base distribution is $N(0, 1)$.
```
%matplotlib inline
from __future__ import division
from matplotlib import pyplot as plt
import numpy as np
import pymc3 as pm
import scipy as sp
import seaborn as sns
from statsmodels.datasets import get_rdataset
from theano import tensor as tt
blue, *_ = sns.color_palette()
SEED = 5132290 # from random.org
np.random.seed(SEED)
N = 20
K = 30
alpha = 2.
P0 = sp.stats.norm
```
We draw and plot samples from the stick-breaking process.
```
beta = sp.stats.beta.rvs(1, alpha, size=(N, K))
w = np.empty_like(beta)
w[:, 0] = beta[:, 0]
w[:, 1:] = beta[:, 1:] * (1 - beta[:, :-1]).cumprod(axis=1)
omega = P0.rvs(size=(N, K))
x_plot = np.linspace(-3, 3, 200)
sample_cdfs = (w[..., np.newaxis] * np.less.outer(omega, x_plot)).sum(axis=1)
fig, ax = plt.subplots(figsize=(8, 6))
ax.plot(x_plot, sample_cdfs[0], c='gray', alpha=0.75,
label='DP sample CDFs');
ax.plot(x_plot, sample_cdfs[1:].T, c='gray', alpha=0.75);
ax.plot(x_plot, P0.cdf(x_plot), c='k', label='Base CDF');
ax.set_title(r'$\alpha = {}$'.format(alpha));
ax.legend(loc=2);
```
As stated above, as $\alpha \to \infty$, samples from the Dirichlet process converge to the base distribution.
```
fig, (l_ax, r_ax) = plt.subplots(ncols=2, sharex=True, sharey=True, figsize=(16, 6))
K = 50
alpha = 10.
beta = sp.stats.beta.rvs(1, alpha, size=(N, K))
w = np.empty_like(beta)
w[:, 0] = beta[:, 0]
w[:, 1:] = beta[:, 1:] * (1 - beta[:, :-1]).cumprod(axis=1)
omega = P0.rvs(size=(N, K))
sample_cdfs = (w[..., np.newaxis] * np.less.outer(omega, x_plot)).sum(axis=1)
l_ax.plot(x_plot, sample_cdfs[0], c='gray', alpha=0.75,
label='DP sample CDFs');
l_ax.plot(x_plot, sample_cdfs[1:].T, c='gray', alpha=0.75);
l_ax.plot(x_plot, P0.cdf(x_plot), c='k', label='Base CDF');
l_ax.set_title(r'$\alpha = {}$'.format(alpha));
l_ax.legend(loc=2);
K = 200
alpha = 50.
beta = sp.stats.beta.rvs(1, alpha, size=(N, K))
w = np.empty_like(beta)
w[:, 0] = beta[:, 0]
w[:, 1:] = beta[:, 1:] * (1 - beta[:, :-1]).cumprod(axis=1)
omega = P0.rvs(size=(N, K))
sample_cdfs = (w[..., np.newaxis] * np.less.outer(omega, x_plot)).sum(axis=1)
r_ax.plot(x_plot, sample_cdfs[0], c='gray', alpha=0.75,
label='DP sample CDFs');
r_ax.plot(x_plot, sample_cdfs[1:].T, c='gray', alpha=0.75);
r_ax.plot(x_plot, P0.cdf(x_plot), c='k', label='Base CDF');
r_ax.set_title(r'$\alpha = {}$'.format(alpha));
r_ax.legend(loc=2);
```
## Dirichlet process mixtures
For the task of density estimation, the (almost sure) discreteness of samples from the Dirichlet process is a significant drawback. This problem can be solved with another level of indirection by using Dirichlet process mixtures for density estimation. A Dirichlet process mixture uses component densities from a parametric family $\mathcal{F} = \{f_{\theta}\ |\ \theta \in \Theta\}$ and represents the mixture weights as a Dirichlet process. If $P_0$ is a probability measure on the parameter space $\Theta$, a Dirichlet process mixture is the hierarchical model
$$
\begin{align*}
x_i\ |\ \theta_i
& \sim f_{\theta_i} \\
\theta_1, \ldots, \theta_n
& \sim P \\
P
& \sim \textrm{DP}(\alpha, P_0).
\end{align*}
$$
To illustrate this model, we simulate draws from a Dirichlet process mixture with $\alpha = 2$, $\theta \sim N(0, 1)$, $x\ |\ \theta \sim N(\theta, (0.3)^2)$.
```
N = 5
K = 30
alpha = 2
P0 = sp.stats.norm
f = lambda x, theta: sp.stats.norm.pdf(x, theta, 0.3)
beta = sp.stats.beta.rvs(1, alpha, size=(N, K))
w = np.empty_like(beta)
w[:, 0] = beta[:, 0]
w[:, 1:] = beta[:, 1:] * (1 - beta[:, :-1]).cumprod(axis=1)
theta = P0.rvs(size=(N, K))
dpm_pdf_components = f(x_plot[np.newaxis, np.newaxis, :], theta[..., np.newaxis])
dpm_pdfs = (w[..., np.newaxis] * dpm_pdf_components).sum(axis=1)
fig, ax = plt.subplots(figsize=(8, 6))
ax.plot(x_plot, dpm_pdfs.T, c='gray');
ax.set_yticklabels([]);
```
We now focus on a single mixture and decompose it into its individual (weighted) mixture components.
```
fig, ax = plt.subplots(figsize=(8, 6))
ix = 1
ax.plot(x_plot, dpm_pdfs[ix], c='k', label='Density');
ax.plot(x_plot, (w[..., np.newaxis] * dpm_pdf_components)[ix, 0],
'--', c='k', label='Mixture components (weighted)');
ax.plot(x_plot, (w[..., np.newaxis] * dpm_pdf_components)[ix].T,
'--', c='k');
ax.set_yticklabels([]);
ax.legend(loc=1);
```
Sampling from these stochastic processes is fun, but these ideas become truly useful when we fit them to data. The discreteness of samples and the stick-breaking representation of the Dirichlet process lend themselves nicely to Markov chain Monte Carlo simulation of posterior distributions. We will perform this sampling using `PyMC3`.
Our first example uses a Dirichlet process mixture to estimate the density of waiting times between eruptions of the [Old Faithful](https://en.wikipedia.org/wiki/Old_Faithful) geyser in [Yellowstone National Park](https://en.wikipedia.org/wiki/Yellowstone_National_Park).
```
old_faithful_df = get_rdataset('faithful', cache=True).data[['waiting']]
```
For convenience in specifying the prior, we standardize the waiting time between eruptions.
```
old_faithful_df['std_waiting'] = (old_faithful_df.waiting - old_faithful_df.waiting.mean()) / old_faithful_df.waiting.std()
old_faithful_df.head()
fig, ax = plt.subplots(figsize=(8, 6))
n_bins = 20
ax.hist(old_faithful_df.std_waiting, bins=n_bins, color=blue, lw=0, alpha=0.5);
ax.set_xlabel('Standardized waiting time between eruptions');
ax.set_ylabel('Number of eruptions');
```
Observant readers will have noted that we have not been continuing the stick-breaking process indefinitely as indicated by its definition, but rather have been truncating this process after a finite number of breaks. Obviously, when computing with Dirichlet processes, it is necessary to only store a finite number of its point masses and weights in memory. This restriction is not terribly onerous, since with a finite number of observations, it seems quite likely that the number of mixture components that contribute non-neglible mass to the mixture will grow slower than the number of samples. This intuition can be formalized to show that the (expected) number of components that contribute non-negligible mass to the mixture approaches $\alpha \log N$, where $N$ is the sample size.
There are various clever [Gibbs sampling](https://en.wikipedia.org/wiki/Gibbs_sampling) techniques for Dirichlet processes that allow the number of components stored to grow as needed. [Stochastic memoization](http://danroy.org/papers/RoyManGooTen-ICMLNPB-2008.pdf) is another powerful technique for simulating Dirichlet processes while only storing finitely many components in memory. In this introductory example, we take the much less sophistocated approach of simply truncating the Dirichlet process components that are stored after a fixed number, $K$, of components. [Ohlssen, et al.](http://fisher.osu.edu/~schroeder.9/AMIS900/Ohlssen2006.pdf) provide justification for truncation, showing that $K > 5 \alpha + 2$ is most likely sufficient to capture almost all of the mixture weight ($\sum_{i = 1}^{K} w_i > 0.99$). In practice, we can verify the suitability of our truncated approximation to the Dirichlet process by checking the number of components that contribute non-negligible mass to the mixture. If, in our simulations, all components contribute non-negligible mass to the mixture, we have truncated the Dirichlet process too early.
Our (truncated) Dirichlet process mixture model for the standardized waiting times is
$$
\begin{align*}
\alpha
& \sim \textrm{Gamma}(1, 1) \\
\beta_1, \ldots, \beta_K
& \sim \textrm{Beta}(1, \alpha) \\
w_i
& = \beta_i \prod_{j = i - 1}^i (1 - \beta_j) \\
\\
\lambda_1, \ldots, \lambda_K
& \sim U(0, 5) \\
\tau_1, \ldots, \tau_K
& \sim \textrm{Gamma}(1, 1) \\
\mu_i\ |\ \lambda_i, \tau_i
& \sim N\left(0, (\lambda_i \tau_i)^{-1}\right) \\
\\
x\ |\ w_i, \lambda_i, \tau_i, \mu_i
& \sim \sum_{i = 1}^K w_i\ N(\mu_i, (\lambda_i \tau_i)^{-1})
\end{align*}
$$
Note that instead of fixing a value of $\alpha$, as in our previous simulations, we specify a prior on $\alpha$, so that we may learn its posterior distribution from the observations.
We now construct this model using `pymc3`.
```
N = old_faithful_df.shape[0]
K = 30
def stick_breaking(beta):
portion_remaining = tt.concatenate([[1], tt.extra_ops.cumprod(1 - beta)[:-1]])
return beta * portion_remaining
with pm.Model() as model:
alpha = pm.Gamma('alpha', 1., 1.)
beta = pm.Beta('beta', 1., alpha, shape=K)
w = pm.Deterministic('w', stick_breaking(beta))
tau = pm.Gamma('tau', 1., 1., shape=K)
lambda_ = pm.Uniform('lambda', 0, 5, shape=K)
mu = pm.Normal('mu', 0, tau=lambda_ * tau, shape=K)
obs = pm.NormalMixture('obs', w, mu, tau=lambda_ * tau,
observed=old_faithful_df.std_waiting.values)
```
We sample from the model 2,000 times using NUTS initialized with ADVI.
```
with model:
trace = pm.sample(2000, n_init=50000, random_seed=SEED)
```
The posterior distribution of $\alpha$ is highly concentrated between 0.25 and 1.
```
pm.traceplot(trace, varnames=['alpha']);
```
To verify that truncation is not biasing our results, we plot the posterior expected mixture weight of each component.
```
fig, ax = plt.subplots(figsize=(8, 6))
plot_w = np.arange(K) + 1
ax.bar(plot_w - 0.5, trace['w'].mean(axis=0), width=1., lw=0);
ax.set_xlim(0.5, K);
ax.set_xlabel('Component');
ax.set_ylabel('Posterior expected mixture weight');
```
We see that only three mixture components have appreciable posterior expected weights, so we conclude that truncating the Dirichlet process to forty components has not appreciably affected our estimates.
We now compute and plot our posterior density estimate.
```
post_pdf_contribs = sp.stats.norm.pdf(np.atleast_3d(x_plot),
trace['mu'][:, np.newaxis, :],
1. / np.sqrt(trace['lambda'] * trace['tau'])[:, np.newaxis, :])
post_pdfs = (trace['w'][:, np.newaxis, :] * post_pdf_contribs).sum(axis=-1)
post_pdf_low, post_pdf_high = np.percentile(post_pdfs, [2.5, 97.5], axis=0)
fig, ax = plt.subplots(figsize=(8, 6))
n_bins = 20
ax.hist(old_faithful_df.std_waiting.values, bins=n_bins, normed=True,
color=blue, lw=0, alpha=0.5);
ax.fill_between(x_plot, post_pdf_low, post_pdf_high,
color='gray', alpha=0.45);
ax.plot(x_plot, post_pdfs[0],
c='gray', label='Posterior sample densities');
ax.plot(x_plot, post_pdfs[::100].T, c='gray');
ax.plot(x_plot, post_pdfs.mean(axis=0),
c='k', label='Posterior expected density');
ax.set_xlabel('Standardized waiting time between eruptions');
ax.set_yticklabels([]);
ax.set_ylabel('Density');
ax.legend(loc=2);
```
As above, we can decompose this density estimate into its (weighted) mixture components.
```
fig, ax = plt.subplots(figsize=(8, 6))
n_bins = 20
ax.hist(old_faithful_df.std_waiting.values, bins=n_bins, normed=True,
color=blue, lw=0, alpha=0.5);
ax.plot(x_plot, post_pdfs.mean(axis=0),
c='k', label='Posterior expected density');
ax.plot(x_plot, (trace['w'][:, np.newaxis, :] * post_pdf_contribs).mean(axis=0)[:, 0],
'--', c='k', label='Posterior expected mixture\ncomponents\n(weighted)');
ax.plot(x_plot, (trace['w'][:, np.newaxis, :] * post_pdf_contribs).mean(axis=0),
'--', c='k');
ax.set_xlabel('Standardized waiting time between eruptions');
ax.set_yticklabels([]);
ax.set_ylabel('Density');
ax.legend(loc=2);
```
The Dirichlet process mixture model is incredibly flexible in terms of the family of parametric component distributions $\{f_{\theta}\ |\ f_{\theta} \in \Theta\}$. We illustrate this flexibility below by using Poisson component distributions to estimate the density of sunspots per year.
```
sunspot_df = get_rdataset('sunspot.year', cache=True).data
sunspot_df.head()
```
For this example, the model is
$$
\begin{align*}
\alpha
& \sim \textrm{Gamma}(1, 1) \\
\beta_1, \ldots, \beta_K
& \sim \textrm{Beta}(1, \alpha) \\
w_i
& = \beta_i \prod_{j = i - 1}^i (1 - \beta_j) \\
\\
\lambda_i, \ldots, \lambda_K
& \sim U(0, 300)
\\
x\ |\ w_i, \lambda_i
& \sim \sum_{i = 1}^K w_i\ \textrm{Poisson}(\lambda_i).
\end{align*}
$$
```
K = 50
N = sunspot_df.shape[0]
with pm.Model() as model:
alpha = pm.Gamma('alpha', 1., 1.)
beta = pm.Beta('beta', 1, alpha, shape=K)
w = pm.Deterministic('w', stick_breaking(beta))
mu = pm.Uniform('mu', 0., 300., shape=K)
obs = pm.Mixture('obs', w, pm.Poisson.dist(mu), observed=sunspot_df['sunspot.year'])
with model:
step = pm.Metropolis()
trace = pm.sample(100000, step=step, tune=90000, random_seed=SEED)
```
For the sunspot model, the posterior distribution of $\alpha$ is concentrated between 0.6 and 1.2, indicating that we should expect more components to contribute non-negligible amounts to the mixture than for the Old Faithful waiting time model.
```
pm.traceplot(trace, varnames=['alpha']);
```
Indeed, we see that between ten and fifteen mixture components have appreciable posterior expected weight.
```
fig, ax = plt.subplots(figsize=(8, 6))
plot_w = np.arange(K) + 1
ax.bar(plot_w - 0.5, trace['w'].mean(axis=0), width=1., lw=0);
ax.set_xlim(0.5, K);
ax.set_xlabel('Component');
ax.set_ylabel('Posterior expected mixture weight');
```
We now calculate and plot the fitted density estimate.
```
x_plot = np.arange(250)
post_pmf_contribs = sp.stats.poisson.pmf(np.atleast_3d(x_plot),
trace['mu'][:, np.newaxis, :])
post_pmfs = (trace['w'][:, np.newaxis, :] * post_pmf_contribs).sum(axis=-1)
post_pmf_low, post_pmf_high = np.percentile(post_pmfs, [2.5, 97.5], axis=0)
fig, ax = plt.subplots(figsize=(8, 6))
ax.hist(sunspot_df['sunspot.year'].values, bins=40, normed=True, lw=0, alpha=0.75);
ax.fill_between(x_plot, post_pmf_low, post_pmf_high,
color='gray', alpha=0.45)
ax.plot(x_plot, post_pmfs[0],
c='gray', label='Posterior sample densities');
ax.plot(x_plot, post_pmfs[::200].T, c='gray');
ax.plot(x_plot, post_pmfs.mean(axis=0),
c='k', label='Posterior expected density');
ax.set_xlabel('Yearly sunspot count');
ax.set_yticklabels([]);
ax.legend(loc=1);
```
Again, we can decompose the posterior expected density into weighted mixture densities.
```
fig, ax = plt.subplots(figsize=(8, 6))
ax.hist(sunspot_df['sunspot.year'].values, bins=40, normed=True, lw=0, alpha=0.75);
ax.plot(x_plot, post_pmfs.mean(axis=0),
c='k', label='Posterior expected density');
ax.plot(x_plot, (trace['w'][:, np.newaxis, :] * post_pmf_contribs).mean(axis=0)[:, 0],
'--', c='k', label='Posterior expected\nmixture components\n(weighted)');
ax.plot(x_plot, (trace['w'][:, np.newaxis, :] * post_pmf_contribs).mean(axis=0),
'--', c='k');
ax.set_xlabel('Yearly sunspot count');
ax.set_yticklabels([]);
ax.legend(loc=1);
```
An earlier version of this example first appeared [here](http://austinrochford.com/posts/2016-02-25-density-estimation-dpm.html).
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Image/clipping.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Image/clipping.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Image/clipping.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://geemap.org). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('Installing geemap ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
import ee
import geemap
```
## Create an interactive map
The default basemap is `Google Maps`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/basemaps.py) can be added using the `Map.add_basemap()` function.
```
Map = geemap.Map(center=[40,-100], zoom=4)
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
image = ee.Image('LANDSAT/LC8_L1T_TOA/LC80440342014077LGN00')
roi = ee.Geometry.Point([-122.4481, 37.7599]).buffer(20000)
clipped = image.clip(roi)
# print(image.getInfo())
Map.setCenter(-122.1899, 37.5010, 10)
vis = {'bands': ['B5', 'B4', 'B3'], 'min': 0, 'max': 0.5, 'gamma': [0.95, 1.1, 1]}
Map.addLayer(image, vis, "Full Image", False)
Map.addLayer(clipped, vis, "Clipped Image")
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
<a href="https://colab.research.google.com/github/pachterlab/CWGFLHGCCHAP_2021/blob/master/notebooks/CellAtlasAnalysis/pseudoBulkStarvation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
!date
```
### **Download Data**
```
import requests
from tqdm import tnrange, tqdm_notebook
def download_file(doi,ext):
url = 'https://api.datacite.org/dois/'+doi+'/media'
r = requests.get(url).json()
netcdf_url = r['data'][0]['attributes']['url']
r = requests.get(netcdf_url,stream=True)
#Set file name
fname = doi.split('/')[-1]+ext
#Download file with progress bar
if r.status_code == 403:
print("File Unavailable")
if 'content-length' not in r.headers:
print("Did not get file")
else:
with open(fname, 'wb') as f:
total_length = int(r.headers.get('content-length'))
pbar = tnrange(int(total_length/1024), unit="B")
for chunk in r.iter_content(chunk_size=1024):
if chunk:
pbar.update()
f.write(chunk)
return fname
#Starvation h5ad data, all nonzero genes included, filtered for 'real cells' from de-multiplexing
download_file('10.22002/D1.1797','.gz')
#CellRanger Starvation h5ad data
download_file('10.22002/D1.1798','.gz')
#Saved DeSeq2 Results for Fed/Starved (Differentially expressed under starvation --> perturbed genes)
download_file('10.22002/D1.1810','.gz')
#Kallisto bus clustered starvation data, h5ad
download_file('10.22002/D1.1796','.gz')
#Human ortholog annotations
download_file('10.22002/D1.1819','.gz')
#Panther annotations
download_file('10.22002/D1.1820','.gz')
#GO Terms
download_file('10.22002/D1.1822','.gz')
!gunzip *.gz
#Install packages
!pip install --quiet anndata
!pip install --quiet scanpy==1.6.0
!pip3 install --quiet rpy2
```
### **Import Packages**
```
import pandas as pd
import anndata
import scanpy as sc
import numpy as np
import scipy.sparse
import warnings
warnings.filterwarnings('ignore')
from sklearn.neighbors import (KNeighborsClassifier,NeighborhoodComponentsAnalysis)
from sklearn.pipeline import Pipeline
from sklearn.manifold import TSNE
from sklearn.decomposition import PCA
from sklearn.preprocessing import scale
import random
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
%matplotlib inline
sc.set_figure_params(dpi=125)
import seaborn as sns
sns.set(style="whitegrid")
%load_ext rpy2.ipython
# See version of all installed packages, last done 11/27/2020
# !pip list -v > pkg_vers_20201127.txt
#Read in annotations
from io import StringIO
hg_ortho_df = pd.read_csv(StringIO(''.join(l.replace('|', '\t') for l in open('D1.1819'))),
sep="\t",header=None,skiprows=[0,1,2,3])
hg_ortho_df[['XLOC','TCONS']] = hg_ortho_df[13].str.split(expand=True)
hg_ortho_df[['Gene','gi']] = hg_ortho_df[3].str.split(expand=True)
hg_ortho_df['Description']= hg_ortho_df[11]
panther_df = pd.read_csv('D1.1820',
sep="\t",header=None) #skiprows=[0,1,2,3]
goTerm_df = pd.read_csv('D1.1822',
sep=" ",header=None) #skiprows=[0,1,2,3]
scDE = pd.read_csv('D1.1810')
scDE.head()
```
### **Run DeSeq2 Analysis for Starvation Pseudo-Bulk Data**
```
#Remove clusters with < 10 cells per condition
#Read in previously saved data
bus_fs_clus = anndata.read("D1.1796")
print(bus_fs_clus )
bus_fs_raw = anndata.read("D1.1797")
bus_fs_raw = bus_fs_raw[bus_fs_clus.obs_names,]
bus_fs_raw.obs['orgID'] = bus_fs_clus.obs['orgID']
bus_fs_raw.obs['fed'] = bus_fs_clus.obs['fed']
bus_fs_raw.obs['cellRanger_louvain'] = bus_fs_clus.obs['cellRanger_louvain']
sc.pp.filter_cells(bus_fs_raw, min_counts=1)
sc.pp.filter_genes(bus_fs_raw, min_counts=1)
bus_fs_raw
#clusSize
#Instantiate dataframe with gene names
def makeDF_forR(sub_raw):
counts = np.zeros((10,len(sub_raw.var_names)))
orgs = [int(i) for i in np.unique(sub_raw.obs['orgID'])]
mat = scipy.sparse.csr_matrix.toarray(sub_raw.X)
for i in np.unique(orgs):
sub_mat = mat[sub_raw.obs['orgID'] == str(i),:]
counts[i-1,:] = sub_mat.sum(axis=0)
fullDF = pd.DataFrame(counts.T, index = sub_raw.var_names.tolist(), columns= np.unique(orgs))
conds = ['True']*5 + ['False']*5
reps = [1,2,3,4,5] + [1,2,3,4,5]
sampleDF = pd.DataFrame({'org_ID': fullDF.columns}) \
.assign(condition = conds) \
.assign(replicate = reps)
sampleDF.index = sampleDF.org_ID
fullDF.to_csv('fullDF.csv')
sampleDF.to_csv('sampleDF.csv')
sampleDF.head()
makeDF_forR(bus_fs_raw)
%%R
fullDF <- read.csv(file = 'fullDF.csv')
sampleDF <- read.csv(file = 'sampleDF.csv')
head(sampleDF)
%%R
head(fullDF)
%%R
rownames(sampleDF) <- sampleDF$org_ID
rownames(fullDF) <- fullDF$X
#colnames(fullDF) <- gsub("\\.", "-", colnames(fullDF))
fullDF <- subset(fullDF, select = -c(X) )
head(fullDF)
sampleDF <- subset(sampleDF, select = -c(org_ID.1) )
# head(sampleDF)
sampleDF$condition <- factor(sampleDF$condition, labels = c("starved", "fed"))
head(sampleDF)
%%R
head(fullDF)
%%R
#Set up R environment
install.packages("BiocManager")
BiocManager::install(version = "3.10")
!sudo apt-get update
!sudo apt-get install libxml2-dev
!sudo apt-get install r-cran-xml
!sudo apt-get install libcurl4-openssl-dev
%%R
#install.packages("DESeq2",repos = "http://cran.us.r-project.org")
BiocManager::install("DESeq2")
#Make output directory
!mkdir kallistoDEAnalysis_BulkStarv
%%R
#Run DeSeq2 for each of the cell types (between control and starved cells)
#install.packages("DESeq2",repos = "http://cran.us.r-project.org")
library("DESeq2")
Genes <- c()
Condition <- c()
padj <- c()
log2FC <- c()
sampleDF$replicate <- factor(sampleDF$replicate)
dds <- DESeqDataSetFromMatrix(countData = fullDF, colData = sampleDF, design= ~replicate + condition)
#Set control condition
dds$condition <- relevel(dds$condition, ref = 'fed')
dds <- DESeq(dds)#parallel = TRUE, test="LRT", sfType="poscounts", useT=TRUE, betaPrior = FALSE, reduced=~replicate
#Starv v Fed results
res <- results(dds,alpha=0.05,name="condition_starved_vs_fed")
resLFC <- res
print(res)
resLFC <- na.omit(resLFC)
resOrdered <- resLFC[resLFC$padj < .05,]
#Keep log2 fold changes < -1 or > 1
resOrdered <- resOrdered[abs(resOrdered$log2FoldChange) > 1,]
outcomes <- resOrdered[order(resOrdered$padj),]
Genes <- c(Genes,row.names(outcomes))
Condition <- c(Condition,rep('Starved',length(row.names(outcomes))))
padj <- c(padj,outcomes$padj)
log2FC <- c(log2FC,outcomes$log2FoldChange)
deGenesDF <- data.frame(matrix(ncol = 4, nrow = length(Genes)))
names(deGenesDF) <- c("Genes", "Condition","padj","log2FC")
deGenesDF$Genes <- Genes
deGenesDF$Condition <- Condition
deGenesDF$padj <- padj
deGenesDF$log2FC <- log2FC
write.csv(deGenesDF,'./kallistoDEAnalysis_BulkStarv/deSeq2_deGenesDF_log2FCof1_singleCellReplicates_noShrinkage_subSample.csv')
head(deGenesDF)
deseq_df = pd.read_csv('./kallistoDEAnalysis_BulkStarv/deSeq2_deGenesDF_log2FCof1_singleCellReplicates_noShrinkage_subSample.csv')
deseq_df.head()
orthoGene = []
orthoDescr = []
pantherNum = []
pantherDescr = []
goTerms = []
for g in deseq_df.Genes:
sub_df = hg_ortho_df[hg_ortho_df.XLOC.isin([g])]
panth_df = panther_df[panther_df[0].isin([g])]
go_df = goTerm_df[goTerm_df[0].isin([g])]
if len(sub_df) > 0:
#Save first result for gene/description
orthoGene += [list(sub_df.Gene)[0]]
orthoDescr += [list(sub_df.Description)[0]]
else:
orthoGene += ['NA']
orthoDescr += ['NA']
if len(panth_df) > 0:
#Save first result for gene/description
pantherNum += [list(panth_df[1])]
pantherDescr += [list(panth_df[2])]
else:
pantherNum += ['NA']
pantherDescr += ['NA']
if len(go_df) > 0:
#Save first result for gene/description
goTerms += [list(go_df[1])]
else:
goTerms += ['NA']
deseq_df['orthoGene'] = orthoGene
deseq_df['orthoDescr'] = orthoDescr
deseq_df['pantherID'] = pantherNum
deseq_df['pantherDescr'] = pantherDescr
deseq_df['goTerms'] = goTerms
deseq_df.head()
deseq_df.to_csv('./kallistoDEAnalysis_BulkStarv/bulk_annos_deSeq2_deGenesDF_log2FCof1_singleCellReplicates_noShrinkage_subSample.csv',index=None)
deseq_df = pd.read_csv('./kallistoDEAnalysis_BulkStarv/bulk_annos_deSeq2_deGenesDF_log2FCof1_singleCellReplicates_noShrinkage_subSample.csv')
deseq_df.head()
scDE.head()
len(set(deseq_df.Genes).intersection(scDE.Genes))/len(deseq_df.Genes)
```
Analyze distribution and expression levels for pseudo-bulk DE genes
```
raw = bus_fs_raw.copy()
sc.pp.normalize_per_cell(raw, counts_per_cell_after=1e4)
#sc.pp.log1p(raw)
raw
allBulk = list(deseq_df.Genes)
onlyBulk = [i for i in deseq_df.Genes if i not in list(scDE.Genes)]
singleDE = raw[:,list(scDE.Genes)].X.todense().mean(axis=0).tolist()[0]
print(len(singleDE))
allBDE = raw[:,allBulk].X.todense().mean(axis=0).tolist()[0]
print(len(allBDE ))
bOnlyDE = raw[:,onlyBulk].X.todense().mean(axis=0).tolist()[0]
print(len(bOnlyDE))
def ecdf(data):
""" Compute ECDF """
x = np.sort(data)
n = len(x)
y = np.arange(n) / float(n)
return(x,y)
exprs = [singleDE,allBDE,bOnlyDE]
exprsLab = ['Single-Cell DE','All Bulk DE','Only Bulk DE']
emb = []
xs = []
ys = []
knnDF = pd.DataFrame()
for p in range(len(exprs)):
i = exprs[p]
l = exprsLab[p]
x,y = ecdf(i)
xs += list(x)
ys += list(y)
emb += [l]*len(x)
knnDF['x'] = xs
knnDF['y'] = ys
knnDF['DE'] = emb
knnDF.head()
knnDF['x'] = np.log2(knnDF['x'])
plt.figure(figsize=(8,4))
sns.scatterplot(data=knnDF, x='x', y='y', hue='DE',alpha=0.6,palette=['purple','b','g'])
plt.grid(b=None)
plt.ylabel("% of Genes")
plt.xlabel("Avg. Expression per Gene (log)")
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
plt.show()
#Plot expression histograms for scDE genes, bulk DE genes, and bulk only DE genes
plt.figure(figsize=(8,4))
plt.hist(singleDE,bins=20,color='purple',alpha=0.6,label='Single-Cell DE',density=True)
plt.hist(allBDE,bins=15, color='b',alpha=0.6,label='All Bulk DE',density=True)
plt.hist(bOnlyDE,bins=10,color = 'g',alpha=0.6,label='Only Bulk DE',density=True)
plt.legend(loc='upper right')
plt.title('DE Genes from Single-cell and Pseudo-bulk Analysis')
plt.grid(b=None)
plt.xlabel("Avg. Expression per Gene")
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
plt.show()
#Label the bulk only genes in annotations
bulkOnly = ['Bulk Only' if i in onlyBulk else 'Overlap' for i in deseq_df.Genes ]
deseq_df['BulkDE'] = bulkOnly
deseq_df.head()
deseq_df.to_csv('./kallistoDEAnalysis_BulkStarv/bulk_annos_deSeq2_deGenesDF_log2FCof1_singleCellReplicates_noShrinkage_subSample.csv',index=None)
#Expression levels for marker genes in general (is this also biased towards high expression genes)
print(bus_fs_clus)
bus_fs_raw
sc.tl.rank_genes_groups(bus_fs_clus, 'cellRanger_louvain', method='wilcoxon',n_genes=1000 )
allMarkers = []
names = np.unique(bus_fs_clus.obs['cellRanger_louvain'])
for n in names:
allMarkers += list(bus_fs_clus.uns['rank_genes_groups']['names'][str(n)])
allMarkers = np.unique(allMarkers)
len(allMarkers)
#Per cell type average expression, to compare
typeExpr = []
names = np.unique(bus_fs_clus.obs['cellRanger_louvain'])
for n in names:
markers = list(bus_fs_clus.uns['rank_genes_groups']['names'][str(n)])
typeExpr += bus_fs_raw[bus_fs_clus[bus_fs_clus.obs['cellRanger_louvain'] == n].obs_names,markers].X.todense().mean(axis=0).tolist()[0]
len(typeExpr)
expr = bus_fs_raw[:,allMarkers].X.todense().mean(axis=0).tolist()[0]
#All gene expression
allExpr = bus_fs_raw.X.todense().mean(axis=0).tolist()[0]
#Plot for expression of marker genes
plt.figure(figsize=(8,4))
plt.hist(np.log1p(expr),bins=15,color='maroon',alpha=0.6,density=True, label= 'All Cells')
plt.hist(np.log1p(typeExpr),bins=15,color='teal',alpha=0.6,density=True, label='Cell Type-Specific')
plt.hist(np.log1p(allExpr),bins=15,color='forestgreen',alpha=0.6,density=True, label='All Genes, All Cells')
plt.legend(loc='upper right')
plt.title('Average Expression of Marker Genes')
plt.grid(b=None)
plt.xlabel("Avg. Expression per Gene (log)")
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
plt.show()
```
| github_jupyter |
<div class="contentcontainer med left" style="margin-left: -50px;">
<dl class="dl-horizontal">
<dt>Title</dt> <dd> Scatter Element</dd>
<dt>Dependencies</dt> <dd>Matplotlib</dd>
<dt>Backends</dt> <dd><a href='./Scatter.ipynb'>Matplotlib</a></dd> <dd><a href='../bokeh/Scatter.ipynb'>Bokeh</a></dd>
</dl>
</div>
```
import numpy as np
import holoviews as hv
hv.extension('matplotlib')
```
The ``Scatter`` element visualizes as markers placed in a space of one independent variable, traditionally denoted as *x*, against a dependent variable, traditonally denoted as *y*. In HoloViews, the name ``'x'`` is the default dimension name used in the ``key_dimensions`` and ``'y'`` is the default dimension name used in the ``value_dimensions``. We can see this from the default axis labels when visualizing a simple ``Scatter`` element:
```
%%opts Scatter (color='k' marker='s' s=10)
np.random.seed(42)
coords = [(i, np.random.random()) for i in range(20)]
hv.Scatter(coords)
```
Here the random *y* values are considered to be the 'data' whereas the x positions express where those values are located (compare this to how [``Points``](./Points.ipynb) elements are defined). In this sense, ``Scatter`` can be thought of as a [``Curve``](./Curve.ipynb) without any lines connecting the samples and you can use slicing to view the *y* values corresponding to a chosen *x* range:
```
%%opts Scatter (color='k' marker='s' s=10)
hv.Scatter(coords)[0:12] + hv.Scatter(coords)[12:20]
```
A ``Scatter`` element must always have at least one value dimension but that doesn't mean additional value dimensions aren't supported. Here is an example with two additional quantities for each point, declared as the ``value_dimension``s *z* and α visualized as the color and size of the dots, respectively:
```
%%opts Scatter [color_index=2 size_index=3 scaling_factor=50]
np.random.seed(10)
data = np.random.rand(100,4)
scatter = hv.Scatter(data, vdims=['y', 'z', 'size'])
scatter + scatter[0.3:0.7, 0.3:0.7].hist()
```
In the right subplot, the ``hist`` method is used to show the distribution of samples along our first value dimension, (*y*).
The marker shape specified above can be any supported by [matplotlib](http://matplotlib.org/api/markers_api.html), e.g. ``s``, ``d``, or ``o``; the other options select the color and size of the marker. For convenience with the [bokeh backend](Bokeh_Backend), the matplotlib marker options are supported using a compatibility function in HoloViews.
**Note**: Although the ``Scatter`` element is superficially similar to the [``Points``](./Points.ipynb) element (they can generate plots that look identical), the two element types are semantically quite different: ``Points`` are used to visualize data where the *y* variable is *dependent*. This semantic difference also explains why the histogram generated by ``hist`` call above visualizes the distribution of a different dimension than it does for [``Points``](./Points.ipynb).
This difference means that ``Scatter`` naturally combine elements that express dependent variables in two-dimensional space such as the ``Chart`` types, such as [``Curve``](./Curve.ipynb). Similarly, ``Points`` express a independent relationship in two-dimensions and combine naturally with [``Raster``](./Raster.ipynb) types such as [``Image``](./Image.ipynb).
For full documentation and the available style and plot options, use ``hv.help(hv.Scatter).``
| github_jupyter |
<table>
<tr align=left><td><img align=left src="./images/CC-BY.png">
<td>Text provided under a Creative Commons Attribution license, CC-BY. All code is made available under the FSF-approved MIT license. (c) Kyle T. Mandli</td>
</table>
Note: The presentation below largely follows part I in "Finite Difference Methods for Ordinary and Partial Differential Equations" by LeVeque (SIAM, 2007).
```
from __future__ import print_function
%matplotlib inline
import numpy
import matplotlib.pyplot as plt
```
# Solving Boundary Value Problems
## The Problem
We want to solve an ODE (PDE) that instead of having initial conditions is contained to an interval and has values at the edges of the interval. This naturally comes about when we consider spatial problems. One of the simplest cases for this is the Poisson problem in one-dimension
$$
u_{xx} = f(x)
$$
where we will use the short-hand
$$
u_{xx} = \frac{\text{d}^2 u}{\text{d} x^2} \quad \text{or} \quad \frac{\partial^2 u}{\partial x^2}.
$$
Note that due to the order of the derivative we require two conditions to solve this. The simplest case where we are on the domain $x \in [a,b]$ is to have conditions such that we require $u(a) = u_a$ and $u(b) = u_b$ and are commonly termed boundary value problems (BVP). If these conditions are both at one end of the domain then we can actually phrase the ODE (PDE) again as an initial value problem (IVP). So what do we need to do to solve these types of problems? We will consider two approaches to this problem:
1. Rephrase the BVP to an IVP and use our standard methods for ODEs.
1. Use finite differences to represent the unknowns as a linear system and solve the resulting system.
## The Shooting Method
The shooting method takes the approach that we want to use our ability to solve IVP problems and so tries to term the problem as a root finding problem for the higher order initial condition that we are not given. This is best illustrated in an example.
Consider the problem
$$
u_{xx} = -\sin u
$$
with
$$
x \in [0, 2] \quad \text{and} \quad u(0) = 0.0, \quad u(2.0) = \frac{\pi}{2}.
$$
We can rewrite this problem as a system of two ODEs as
$$
v = \begin{bmatrix} u \\ u_x \end{bmatrix} \quad \text{and} \quad v_x = \begin{bmatrix} u_x \\ u_{xx} \end{bmatrix} = \begin{bmatrix} v_2 \\ -\sin v_1 \end{bmatrix}.
$$
We know that we want $v_1(0) = 0$ but what do we use for $v_2(0)$? Making an initial guess at $v_2(0)$ and solving the associated IVP ODE we can then find out what these initial conditions produces on the right boundary of the problem. Using a root-finding approach (or minimization routine) we can write this procedure as
$$
\min_{v_2(0)} \left | \pi / 2 - v_1(2) \right |
$$
where the parameter we vary is $v_2(0)$.
```
# Basic Shooting Method solving u_xx = -sin(u)
import scipy.integrate as integrate
# Algorithm parameters
TOLERANCE = 1e-8
MAX_ITERATIONS = 100
# Problem Statement
a = 0.0
b = 2.0
N = 100
x = numpy.linspace(a, b, N)
u_a = 0.0
u_b = numpy.pi / 2.0
# RHS function
def f(x, u):
return numpy.array([u[1], -numpy.sin(u[0])])
# Initial guess
# Slope at RHS
u_prime_rhs = 1.0
# Initial step size
du_prime = 0.5
# Plotting
fig = plt.figure()
fig.set_figwidth(fig.get_figwidth() * 2)
axes = fig.add_subplot(1, 2, 1)
# Main loop
success = False
u = numpy.empty((2, N))
convergence = numpy.zeros(MAX_ITERATIONS)
for n in range(MAX_ITERATIONS):
# Initial condition
u[0, 0] = u_a
u[1, 0] = u_prime_rhs
# Construct integrator
integrator = integrate.ode(f)
integrator.set_integrator("dopri5")
integrator.set_initial_value(u[:, 0])
# Compute solution - note that we are only producing the intermediate values
# for demonstration purposes
for (i, x_output) in enumerate(x[1:]):
integrator.integrate(x_output)
if not integrator.successful():
raise Exception("Integration Failed!")
u[:, i + 1] = integrator.y
# Stopping Criteria
convergence[n] = numpy.abs(u[0, -1] - u_b)
if numpy.abs(u[0, -1] - u_b) < TOLERANCE:
success = True
break
else:
if u[0, -1] < u_b:
u_prime_rhs += du_prime
else:
u_prime_rhs -= du_prime
du_prime *= 0.5
axes.plot(x, u[0, :], 'b')
axes.plot(b, u_b, 'ro')
axes.set_title("Shooting Method Iterations")
axes.set_xlabel("$x$")
axes.set_ylabel("$u(x)$")
axes = fig.add_subplot(1, 2, 2)
n_range = numpy.arange(n)
axes.semilogy(n_range, convergence[:n])
axes.set_title("Convergence of Shooting Method")
axes.set_xlabel("step")
axes.set_ylabel("$|u(b) - U(b)|$")
plt.show()
```
The tricky part of this procedure is coming up with the search criteria, i.e. coming up with the decision of how to change $v_2(0)$ with respect to the position of $v_2(2)$ compared to what we want $u(2)$.
In general any minimization routine can be used in a shooting method. These approaches are generally very effective at approaching non-linear BVPs where the next method we will discuss is too expensive to perform.
## Linear System Approach
### Formulation
The second approach we will consider involves the formation of a system of equations to solve based on finite difference approximations. Again let's consider an example problem where
$$
u_{xx} = f(x)
$$
with the initial conditions $u(a) = u_a$ and $u(b) = u_b$.
We know from our finite difference discussion that the second order centered difference approximation for the second derivative for a function $u(x)$ is
$$
u_{xx} \approx \frac{u(x_{i-1}) - 2 u(x_i) + u(x_{i+1})}{\Delta x^2}.
$$
If we descretize the domain of the original BVP into $N$ points (not including the boundaries) such that
$$
x_i = a + \frac{b - a}{N+1} \cdot i ~~~ \text{where} ~~~ i = 1, \ldots, N
$$
we can then write the finite difference approximation as a system of linear equations!
If for instance we take $N = 5$ then
$$\begin{aligned}
(U_{xx})_1 &\approx \frac{U_a - 2 U_1 + U_2}{\Delta x^2} \\
(U_{xx})_2 &\approx \frac{U_1 - 2 U_2 + U_3}{\Delta x^2} \\
(U_{xx})_3 &\approx \frac{U_2 - 2 U_3 + U_4}{\Delta x^2} \\
(U_{xx})_4 &\approx \frac{U_3 - 2 U_4 + U_5}{\Delta x^2} \\
(U_{xx})_5 &\approx \frac{U_4 - 2 U_5 + U_b}{\Delta x^2} \\
\end{aligned}$$
where we have used $U_a = u(a)$ and $U_b = u(b)$ as the boundary conditions.
Using these approximations to the derivatives we can then write the ODE as
$$
\frac{1}{\Delta x^2}\begin{bmatrix}
-2 & 1 & & & \\
1 & -2 & 1 & & \\
& 1 & -2 & 1 & \\
& & 1 & -2 & 1 \\
& & & 1 & -2 \\
\end{bmatrix} \begin{bmatrix}
U_1 \\ U_2 \\ U_3 \\ U_4 \\ U_5
\end{bmatrix} =
\begin{bmatrix}
f(x_1) \\ f(x_2) \\ f(x_3) \\ f(x_4) \\ f(x_5) \\
\end{bmatrix}.
$$
Note that our previous example used for the shooting method is difficult in the current context as the unknown function is in the function $f$ so that we would need to actual solve a non-linear system of equations. This is still possible in this context using an approach such as a Newton solver and has similar properties as the shooting method (although not as simple to implement).
### Boundary Conditions
This does not include the boundary conditions though. We can add these values easily for Dirichlet boundary conditions by sending the values we know to the $b$ vector:
$$\begin{aligned}
\frac{U_a - 2 U_1 + U_2}{\Delta x^2} = f(x_1) &\Rightarrow& \frac{- 2 U_1 + U_2}{\Delta x^2} = f(x_1) - \frac{U_a}{\Delta x^2} \\
\frac{U_4 - 2 U_5 + U_b}{\Delta x^2} = f(x_1) &\Rightarrow& \frac{U_4 - 2 U_5}{\Delta x^2} = f(x_5) - \frac{U_b}{\Delta x^2}
\end{aligned}$$
so that final system looks like
$$
\frac{1}{\Delta x^2} \begin{bmatrix}
-2 & 1 & & & \\
1 & -2 & 1 & & \\
& 1 & -2 & 1 & \\
& & 1 & -2 & 1 \\
& & & 1 & -2 \\
\end{bmatrix} \begin{bmatrix}
U_1 \\ U_2 \\ U_3 \\ U_4 \\ U_5
\end{bmatrix} =
\begin{bmatrix}
f(x_1) - \frac{U_a}{\Delta x^2} \\ f(x_2) \\ f(x_3) \\ f(x_4) \\ f(x_5) - \frac{U_b}{\Delta x^2} \\
\end{bmatrix}.
$$
### Example
Want to solve the BVP
$$
u_{xx} = e^x, \quad x \in [0, 1] \quad \text{with} \quad u(0) = 0.0, \text{ and } u(1) = 3
$$
via the construction of a linear system of equations.
\begin{align*}
u_{xx} &= e^x \\
u_x &= A + e^x \\
u &= Ax + B + e^x\\
u(0) &= B + 1 = 0 \Rightarrow B = -1 \\
u(1) &= A - 1 + e^{1} = 3 \Rightarrow A = 4 - e\\
~\\
u(x) &= (4 - e) x - 1 + e^x
\end{align*}
```
# Problem setup
a = 0.0
b = 1.0
u_a = 0.0
u_b = 3.0
f = lambda x: numpy.exp(x)
u_true = lambda x: (4.0 - numpy.exp(1.0)) * x - 1.0 + numpy.exp(x)
# Descretization
N = 10
x_bc = numpy.linspace(a, b, N + 2)
x = x_bc[1:-1]
delta_x = (b - a) / (N + 1)
# Construct matrix A
A = numpy.zeros((N, N))
diagonal = numpy.ones(N) / delta_x**2
A += numpy.diag(diagonal * -2.0, 0)
A += numpy.diag(diagonal[:-1], 1)
A += numpy.diag(diagonal[:-1], -1)
# Construct RHS
b = f(x)
b[0] -= u_a / delta_x**2
b[-1] -= u_b / delta_x**2
# Solve system
U = numpy.empty(N + 2)
U[0] = u_a
U[-1] = u_b
U[1:-1] = numpy.linalg.solve(A, b)
# Plot result
fig = plt.figure()
axes = fig.add_subplot(1, 1, 1)
axes.plot(x_bc, U, 'o', label="Computed")
axes.plot(x_bc, u_true(x_bc), 'k', label="True")
axes.set_title("Solution to $u_{xx} = e^x$")
axes.set_xlabel("x")
axes.set_ylabel("u(x)")
plt.show()
```
If we instead have Neumann boundary conditions it is no longer clear how to handle the boundary conditions using the above approach. Instead a **ghost cell** approach is often used. These **ghost cells** are added unknowns that represent the boundary values that we actually know.
For instance, if we had the BVP
$$
u_{xx} = e^x, \quad x \in [-1, 1] \quad \text{with} \quad u(-1) = 3, \text{ and } u_x(1) = -5
$$
then we could keep the boundary values in the vector of unknowns so that now
$$
U = \begin{bmatrix} U_0 \\ U_1 \\ \vdots \\ U_N \\ U_{N+1} \end{bmatrix}
$$
where here $U_0$ and $U_{N+1}$ are actually the boundary points.
The matrix $A$ is then modified to have the appropriate relationships. In this case the left boundary condition leads to
$$
A = \begin{bmatrix}
1 & & & & & \\
\frac{1}{\Delta x^2} & \frac{-2}{\Delta x^2} & \frac{1}{\Delta x^2} & & & \\
& \frac{1}{\Delta x^2} & \frac{-2}{\Delta x^2} & \frac{1}{\Delta x^2} & & \\
& & \ddots & \ddots & \ddots
\end{bmatrix} \quad \text{and} \quad b = \begin{bmatrix}
u(a) \\ f(x_1) \\ f(x_2) \\ \vdots
\end{bmatrix}
$$
which multiplied out simply gives
$$
U_0 = u(-1) = 3.
$$
For the right boundary condition we can use the second order backward finite difference approximation for the first derivative
$$
u_x(b) \approx \frac{3 U_{N+1} - 4 U_{N} + U_{N - 1}}{2.0 \Delta x} = -5
$$
which can be incorporated into the matrix $A$ and vector $b$ as
$$
A = \begin{bmatrix}
\ddots & \ddots & \ddots & & \\
& \frac{1}{\Delta x^2} & \frac{-2}{\Delta x^2}& \frac{1}{\Delta x^2} & \\
& & \frac{1}{\Delta x^2} & \frac{-2}{\Delta x^2} & \frac{1}{\Delta x^2} \\
& & \frac{1}{2 \Delta x} & \frac{-4}{2 \Delta x} & \frac{3}{2 \Delta x} \\
\end{bmatrix} ~~~~ \text{and} ~~~~ b = \begin{bmatrix}
\vdots \\ f(x_N) \\ u_x(b)
\end{bmatrix}.
$$
All together the new system looks like
$$
\begin{bmatrix}
1 & & & & & \\
\frac{1}{\Delta x^2} & \frac{-2}{\Delta x^2} & \frac{1}{\Delta x^2} & & & \\
& \ddots & \ddots & \ddots & \\
& & \frac{1}{\Delta x^2} & \frac{-2}{\Delta x^2} & \frac{1}{\Delta x^2} \\
& & \frac{1}{2 \Delta x} & \frac{-4}{2 \Delta x} & \frac{3}{2 \Delta x} \\
\end{bmatrix} \begin{bmatrix}
U_0 \\ U_1 \\ \vdots \\ U_N \\ U_{N+1}
\end{bmatrix} =
\begin{bmatrix}
u(a) \\ f(x_1) \\ \vdots \\ f(x_N) \\ u_x(b)
\end{bmatrix}.
$$
### Example
Want to solve the BVP
$$
u_{xx} = e^x, \quad x \in [-1, 1] \quad \text{with} \quad u(-1) = 3.0, \text{ and } u_x(1) = -5.0
$$
via the construction of a linear system of equations.
First find the true solution and then compute the solution.
\begin{align*}
u(x) &= A x + B + e^x \\
u_x(1) &= A + e^1 = -5 \Rightarrow A = -5 - e \\
u(-1) &= (5 + e) + B + e^{-1} = 3 \Rightarrow B = 3 - 5 - e - e^{-1} = -(2 + e + e^{-1}) \\
~\\
u(x) &= -(5 + e) x -(2 + e + e^{-1}) + e^{x}
\end{align*}
```
# Problem setup
a = -1.0
b = 1.0
u_a = 3.0
u_x_b = -5.0
f = lambda x: numpy.exp(x)
u_true = lambda x: -(5.0 + numpy.exp(1.0)) * x - (2.0 + numpy.exp(1.0) + numpy.exp(-1.0)) + numpy.exp(x)
# Descretization
N = 10
x_bc = numpy.linspace(a, b, N + 2)
x = x_bc[1:-1]
delta_x = (b - a) / (N + 1)
# Construct matrix A
A = numpy.zeros((N + 2, N + 2))
diagonal = numpy.ones(N + 2) / delta_x**2
A += numpy.diag(diagonal * -2.0, 0)
A += numpy.diag(diagonal[:-1], 1)
A += numpy.diag(diagonal[:-1], -1)
# Construct RHS
b = f(x_bc)
# Boundary conditions
A[0, 0] = 1.0
A[0, 1] = 0.0
A[-1, -1] = 3.0 / (2.0 * delta_x)
A[-1, -2] = -4.0 / (2.0 * delta_x)
A[-1, -3] = 1.0 / (2.0 * delta_x)
b[0] = u_a
b[-1] = u_x_b
# Solve system
U = numpy.empty(N + 2)
U = numpy.linalg.solve(A, b)
# Plot result
fig = plt.figure()
axes = fig.add_subplot(1, 1, 1)
axes.plot(x_bc, U, 'o', label="Computed")
axes.plot(x_bc, u_true(x_bc), 'k', label="True")
axes.set_title("Solution to $u_{xx} = e^x$")
axes.set_xlabel("x")
axes.set_ylabel("u(x)")
plt.show()
```
## Ways to Solve $A u = f$
We have proposed solving the linear system $A u = f$ which we have implemented naively above with the `numpy.linalg.solve` command but perhaps given the special structure of $A$ here that we can do better.
### Direct Methods (Gaussian Elimination)
We could use Gaussian elimination to solve the system (or some factorization) which leads to a solution in a finite number of steps. For large, sparse methods however these direct solvers are much more expensive in general over iterative solvers. As was discussed for eigenproblems, iterative solvers start with an initial guess and try to improve on that guess.
Consider a 3D Poisson Problem:
- Discretize using $100 \times 100 \times 100 = 10^6$ unknowns
- Gaussian elimination requires $\mathcal{O}(N^3)$ operations
- Solving this system would take $10^{18}$ floating point operations to complete
**How long?**
Today's computer is $\approx 100$ gigaflops (floating point operations per second) - $10^{11}$ flops / second. We would be waiting 115 days for the solve to finish!
**Memory?**
Matrix requires $N^2$ memory locations ($N = 10^6$ here). Single precision floating point storage (4-bytes per number) would require $4 \times 10^{12}$ bytes of 4 terabytes of memory.
The situation really is not as bad as we are making it out to be as long as we take advantage of the sparse nature of the matrices. In fact for 1 dimensional problems direct methods can be reduced to $\mathcal{O}(N)$ in the case for a tridiagonal system. The situation is not so great for higher-dimensional problems however unless most structure can be leveraged. Examples of these types of solvers include fast Fourier methods such as fast Poisson solvers.
### Iterative Methods
Iterative methods take another tact that direct methods. If we have the system $A x = b$ we form an iterative procedure that applies a function, say $L$, such that
$$
\hat{x}^{(k)} = L(\hat{x}^{(k-1)})
$$
where we want errot between the real solution $x$ and $\hat{x}^{(k)}$ goes to zero as $k \rightarrow \infty$. We will explore these methods in the next lecture.
| github_jupyter |
# Differential equation resolution
## Introduction
We present here a Perceval implementation of a Quantum Machine Learning algorithm for solving differential equations. Its aims is to approximate the solution to the differential equation considered in \[1\]:
$$
\frac{d f}{d x}+\lambda f(x)(\kappa+\tan (\lambda x))=0
$$
with boundary condition $f(0)=f_{0}$. The analytical solution is $f(x)=\exp (-\kappa \lambda x) \cos (\lambda x)+ f_0 - 1$.
### QML Loss Function Definition
In order to use QML to solve this differential equation, we first need to derive from it a loss function whose minimum is associated to its analytical solution.
Let $F\left[\left\{d^{m} f / d x^{m}\right\}_{m},f, x\right]=0$ be a general differential equation verified by $f(x)$, where $F[.]$ is an operator acting on $f(x)$, its derivatives and $x$. For the solving of a differential equation, the loss function described in \[1\] consists of two terms
$$
\mathcal{L}_{\boldsymbol{\theta}}\left[\left\{d^{m} g / d x^{m}\right\}_{m},g, x\right]:=\mathcal{L}_{\boldsymbol{\theta}}^{(\mathrm{diff})}\left[\left\{d^{m} g / d x^{m}\right\}_{m},g, x\right]+\mathcal{L}_{\boldsymbol{\theta}}^{(\text {boundary})}[g, x].
$$
The first term $\mathcal{L}_{\boldsymbol{\theta}}^{(\mathrm{diff})}$ corresponds to the differential equation which has been discretised over a fixed regular grid of $M$ points noted $x_i$:
$$
\mathcal{L}_{\boldsymbol{\theta}}^{(\mathrm{diff})}\left[\left\{d^{m} g / d x^{m}\right\}_{m},g, x\right]:=\frac{1}{M} \sum_{i=1}^{M} L\left(F\left[d_{x}^m g\left(x_{i}\right), g\left(x_{i}\right), x_{i}\right], 0\right),
$$
where $L(a,b) := (a - b)^2$ is the squared distance between two arguments. The second term $\mathcal{L}_{\boldsymbol{\theta}}^{(\text {boundary })}$ is associated to the initial conditions of our desired solution. It is defined as:
$$
\mathcal{L}_{\boldsymbol{\theta}}^{\text {(boundary) }}[g, x]:=\eta L\left(g(x_0), f_{0}\right),
$$
where $\eta$ is the weight granted to the boundary condition and $f_{0}$ is given by $f(x_0) = f_0$.
Given a function approximator $f^{(n)}(x, \boldsymbol{\theta}, \boldsymbol{\lambda})$, the loss function above will be minimised using a classical algorithm, updating the parameters $\boldsymbol{\theta}$ based on samples obtained using a quantum device.
### Quantum circuit architecture
The feature map used is presented in \[2,3,4\]. The quantum circuit architecture from \[4\] is expressed as $\mathcal{U}(x, \boldsymbol{\theta}):=\mathcal{W}^{(2)}\left(\boldsymbol{\theta}_{2}\right) \mathcal{S}(x) \mathcal{W}^{(1)}\left(\boldsymbol{\theta}_{1}\right).$ The phase-shift operator $\mathcal{S}(x)$ incorporates the $x$ dependency of the function we wish to approximate. It is sandwiched between two universal interferometers $\mathcal{W}^{(1)}(\boldsymbol{\theta_1})$ and $\mathcal{W}^{(2)}(\boldsymbol{\theta_2})$, where the beam-splitter parameters $\boldsymbol{\theta_1}$ and $\boldsymbol{\theta_2}$ of this mesh architecture are tunable to enable training of the circuit.
The output measurement operator, noted $\mathcal{M}(\boldsymbol{\lambda})$, is the projection on the Fock states obtained using photon-number resolving detectors, multiplied by some coefficients $\boldsymbol{\lambda}$ which can also be tunable. Formally, we have:
$$ \mathcal{M}(\boldsymbol{\lambda}) = \sum_{\mathbf{\left | n^{(f)}\right \rangle}}\lambda_{\mathbf{\left | n^{(f)}\right \rangle}}\mathbf{\left | n^{(f)}\right \rangle}\mathbf{\left \langle n^{(f)}\right |},
$$
where the sum is taken over all $\binom{n+m-1}{n}$ possible Fock states considering $n$ photons in $m$ modes. Let $\mathbf{\left | n^{(i)}\right \rangle} = \left |n^{(i)}_1,n^{(i)}_2,\dots,n^{(i)}_m\right \rangle$ be the input state consisting of $n$ photons where $n^{(i)}_j$ is the number of photons in input mode $j$. Given these elements, the circuit's output $f^{(n)}(x, \boldsymbol{\theta}, \boldsymbol{\lambda})$ is given by the following expectation value:
$$
f^{(n)}(x, \boldsymbol{\theta}, \boldsymbol{\lambda})=\left\langle\mathbf{n}^{(i)}\left|\mathcal{U}^{\dagger}(x, \boldsymbol{\theta}) \mathcal{M}(\boldsymbol{\lambda}) \mathcal{U}(x, \boldsymbol{\theta})\right| \mathbf{n}^{(i)}\right\rangle.
$$
This expression can be rewritten as the following Fourier series \[4\]
$$
f^{(n)}(x, \boldsymbol{\theta}, \boldsymbol{\lambda})=\sum_{\omega \in \Omega_{n}} c_{\omega}(\boldsymbol{\theta}, \boldsymbol{\lambda}) e^{i \omega x},
$$
where $\Omega_n = \{-n, -n+1, \dots, n-1, n \}$ is the frequency spectrum one can reach with $n$ incoming photons and $\{c_\omega(\boldsymbol{\theta}, \boldsymbol{\lambda})\}$ are the Fourier coefficients. The $\boldsymbol{\lambda}$ parameters are sampled randomly in the interval $[-a;a]$, with $a$ a randomly chosen integer. $f^{(n)}(x, \boldsymbol{\theta}, \boldsymbol{\lambda})$ will serve as a function approximator for this chosen differential equation. Differentiation in the loss function is discretised as $\frac{df}{dx} \simeq \frac{\Delta f}{\Delta x}$.
$n, m, \boldsymbol{\lambda}$ and $\Delta x $ are variable parameters defined below.
## Perceval Simulation
### Initialisation
```
from IPython import display
import perceval as pcvl
import perceval.lib.phys as phys
import numpy as np
from math import comb
from scipy.optimize import minimize
import time
import matplotlib.pyplot as plt
import tqdm as tqdm
```
We will run this notebook with 6 photons
```
nphotons = 6
```
### Differential equation parameters
We define here the value of the differential equation parameters and boundary condition $\lambda, \kappa, f_0$.
```
# Differential equation parameters
lambd = 8
kappa = 0.1
def F(u_prime, u, x): # DE
return (u_prime(x) + lambd * u(x) * (kappa + np.tan(lambd * x)))
# Boundary condition (f(x_0)=f_0)
x_0 = 0
f_0 = 1
# Modeling parameters
n_grid = 50 # number of grid points of the discretized differential equation
range_min = 0 # minimum of the interval on which we wish to approximate our function
range_max = 1 # maximum of the interval on which we wish to approximate our function
X = np.linspace(range_min, range_max-range_min, n_grid) # Optimisation grid
# Differential equation's exact solution - for comparison
def u(x):
return np.exp(- kappa*lambd*x)*np.cos(lambd*x)
# Parameters of the quantum machine learning procedure
N = nphotons # Number of photons
m = nphotons # Number of modes
eta = 5 # weight granted to the initial condition
a = 200 # Approximate boundaries of the interval that the image of the trial function can cover
fock_dim = comb(N + m - 1, N)
# lambda coefficients for all the possible outputs
lambda_random = 2 * a * np.random.rand(fock_dim) - a
# dx serves for the numerical differentiation of f
dx = (range_max-range_min) / 1000
# Input state with N photons and m modes
input_state = pcvl.BasicState([1]*N+[0]*(m-N))
print(input_state)
```
## Definition of the circuit
We will generate a Haar-random initial unitary using QR decomposition built in Perceval `Matrix.random_unitary`, the circuit is defined by the combination of 3 sub-circuits - the intermediate phase is a parameter.
```
"Haar unitary parameters"
# number of parameters used for the two universal interferometers (2*m**2 per interferometer)
parameters = np.random.normal(size=4*m**2)
simulator_backend = pcvl.BackendFactory().get_backend("SLOS")
px = pcvl.P("px")
c = (pcvl.Circuit(m, pcvl.Matrix.random_unitary(m, parameters[:2 * m ** 2]), name="W1")
// (0, phys.PS(px))
// pcvl.Circuit(m, pcvl.Matrix.random_unitary(m, parameters[2 * m ** 2:]), name="W2"))
pcvl.pdisplay(c)
s1 = simulator_backend(pcvl.Matrix.random_unitary(m))
s1.compile(input_state)
```
### Expectation value and loss function computation
The expectation value of the measurement operator $\mathcal{M}(\boldsymbol{\lambda})$ is obtained directly from Fock state probabilities computed by Perceval. Given this expectation value, the code snippet below computes the loss function defined in the Introduction.
Note the use of the `all_prob` simulator method giving directly access to the probabilities of all possible output states. This calculation is optimized in SLOS backend.
```
def computation(params):
global current_loss
global computation_count
"compute the loss function of a given differential equation in order for it to be optimized"
computation_count += 1
f_theta_0 = 0 # boundary condition
coefs = lambda_random # coefficients of the M observable
# initial condition with the two universal interferometers and the phase shift in the middle
U_1 = pcvl.Matrix.random_unitary(m, params[:2 * m ** 2])
U_2 = pcvl.Matrix.random_unitary(m, params[2 * m ** 2:])
px = pcvl.P("x")
c = pcvl.Circuit(m, U_2) // (0, phys.PS(px)) // pcvl.Circuit(m, U_1)
px.set_value(x_0)
U = c.compute_unitary(use_symbolic=False)
s1.U = U
f_theta_0 = np.sum(np.multiply(s1.all_prob(input_state), coefs))
# boundary condition given a weight eta
loss = eta * (f_theta_0 - f_0) ** 2 * len(X)
def ldiff(x):
i, j = 0, 0
px.set_value(x)
s1.U = c.compute_unitary()
f_theta = np.sum(np.multiply(s1.all_prob(input_state), coefs))
px.set_value(x+dx)
s1.U = c.compute_unitary()
f_theta_dx = np.sum(np.multiply(s1.all_prob(input_state), coefs))
diff_f_theta = (f_theta_dx - f_theta) / dx
return (diff_f_theta + lambd * (kappa + np.tan(lambd * x)) * f_theta) ** 2
for x in X:
loss += ldiff(x)
current_loss = loss / len(X)
return current_loss
```
### Classical optimisation
Finally the code below performs the optimisation procedure using the loss function defined in the previous section. To this end, we use a Broyden–Fletcher–Goldfarb–Shanno (BFGS) optimiser \[5\] from the SciPy library.
```
def callbackF(parameters):
"""callback function called by scipy.optimize.minimize allowing to monitor progress"""
global current_loss
global computation_count
global loss_evolution
global start_time
now = time.time()
pbar.set_description("M= %d Loss: %0.5f #computations: %d elapsed: %0.5f" %
(m, current_loss, computation_count, now-start_time))
pbar.update(1)
loss_evolution.append((current_loss, now-start_time))
computation_count = 0
start_time = now
computation_count = 0
current_loss = 0
start_time = time.time()
loss_evolution = []
pbar = tqdm.tqdm()
res = minimize(computation, parameters, callback=callbackF, method='BFGS', options={'gtol': 1E-2})
```
After the optimisation procedure has been completed, the optimal unitary parameters (in `res.x`) can be used to determine the quantum circuit beam-splitter and phase-shifter angles for an experimental realisation.
```
print("Unitary parameters", res.x)
```
### Plotting the approximation
We now plot the result of our optimisation in order to compare the QML algorithm's output and the analytical solution.
```
def plot_solution(m, N, X, optim_params, lambda_random):
Y = []
U_1 = pcvl.Matrix.random_unitary(m, optim_params[:2 * m ** 2])
U_2 = pcvl.Matrix.random_unitary(m, optim_params[2 * m ** 2:])
simulator_backend = pcvl.BackendFactory().get_backend("SLOS")
px = pcvl.P("x")
c = pcvl.Circuit(m, U_2) // (0, phys.PS(px)) // pcvl.Circuit(m, U_1)
for x in X:
px.set_value(x)
U = c.compute_unitary(use_symbolic=False)
s1.U = U
f_theta = np.sum(np.multiply(s1.all_prob(input_state), lambda_random))
Y.append(f_theta)
exact = u(X)
plt.plot(X, Y, label="Approximation with {} photons".format(N))
X = np.linspace(range_min, range_max, 200)
plot_solution(m, N, X, res.x, lambda_random)
plt.plot(X, u(X), 'r', label='Analytical solution')
plt.legend()
plt.show()
plt.plot([v[0] for v in loss_evolution])
plt.yscale("log")
plt.xlabel("Number of epochs")
plt.ylabel("Loss function value")
```
## References
\[1\] : O. Kyriienko, A. E. Paine, and V. E. Elfving, “Solving nonlinear differential equations with differentiable quantum circuits”, *Physical Review A* **103**, 052416 (2021). https://journals.aps.org/pra/abstract/10.1103/PhysRevA.103.052416
\[2\] A. Pérez-Salinas, A. Cervera-Lierta, E. Gil-Fuster, and J. I. Latorre, “Data re-uploading for a universal quantum classifier”, *Quantum* **4**, 226 (2020). https://quantum-journal.org/papers/q-2020-02-06-226/
\[3\] M. Schuld, R. Sweke, and J. J. Meyer, “Effect of data encoding on the expressive power of variational quantum-machine-learning models”, *Physical Review A* **103**, 032430 (2021). https://journals.aps.org/pra/abstract/10.1103/PhysRevA.103.032430
\[4\] B. Y. Gan, D. Leykam, D. G. Angelakis, and D. G. Angelakis, “Fock State-enhanced Expressivity of Quantum Machine Learning Models”, in *Conference on Lasers andElectro-Optics* (2021), paper JW1A.73. Optica Publishing Group, (2021). https://opg.optica.org/abstract.cfm?uri=CLEO_AT-2021-JW1A.73.
\[5\] R. Fletcher, Practical methods of optimization. John Wiley & Sons. (2013) https://onlinelibrary.wiley.com/doi/book/10.1002/9781118723203
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import rl2
from scipy.optimize import fsolve
import numba
from pyswarm import pso
%matplotlib notebook
run = rl2.run
@numba.njit(cache=True)
def solver(struct):
sin = np.sin
cos = np.cos
sqrt = np.sqrt
i = 0
Dt = struct[i].Dt
N_steps = struct[i].N_steps
N_store = struct[i].N_store
N_x = struct[i].N_x
N_y = struct[i].N_y
N_outs = 1
decimation = struct[i].decimation
eye = np.eye(N_x)
# initialization
t = 0.0
run(0.0,struct, 1)
it_store = 0
struct[i]['T'][0] = t
struct[i].X[0,:] = struct[i].x[:,0]
Y = np.zeros((N_store,N_y))
Y[0,:] = struct[i].y[:,0]
for it in range(N_steps-1):
t += Dt
perturbations(t,struct)
solver = struct[i].solvern
if solver == 1:
# forward euler solver
run(t,struct, 2)
struct[i].x[:] += Dt*struct[i].f
if solver == 2:
# bacward euler solver
x_0 = np.copy(struct[i].x[:])
for j in range(struct[i].imax):
run(t,struct, 2)
run(t,struct, 3)
run(t,struct, 10)
phi = x_0 + Dt*struct[i].f - struct[i].x
Dx = np.linalg.solve(-(Dt*struct[i].Fx - np.eye(N_x)), phi)
struct[i].x[:] += Dx[:]
if np.max(np.abs(Dx)) < struct[i].itol: break
print(struct[i].f)
if solver == 3:
# trapezoidal solver
run(t,struct, 2)
f_0 = np.copy(struct[i].f[:])
x_0 = np.copy(struct[i].x[:])
for j in range(struct[i].imax):
run(t,struct, 10)
phi = x_0 + 0.5*Dt*(f_0 + struct[i].f) - struct[i].x
Dx = np.linalg.solve(-(0.5*Dt*struct[i].Fx - np.eye(N_x)), phi)
struct[i].x[:] += Dx[:]
run(t,struct, 2)
if np.max(np.abs(Dx)) < struct[i].itol: break
if solver == 4:
#print(t)
run(t,struct, 2)
run(t,struct, 3)
x = np.copy(struct[i].x[:])
y = np.copy(struct[i].y[:])
f = np.copy(struct[i].f[:])
g = np.copy(struct[i].g[:])
for iter in range(10):
run(t,struct, 2)
run(t,struct, 3)
run(t,struct,10)
run(t,struct,11)
x_i = struct[i].x[:]
y_i = struct[i].y[:]
f_i = struct[i].f[:]
g_i = struct[i].g[:]
F_x_i = struct[i].Fx[:,:]
F_y_i = struct[i].Fy[:,:]
G_x_i = struct[i].Gx[:,:]
G_y_i = struct[i].Gy[:,:]
A_c_i = np.vstack((np.hstack((eye-0.5*Dt*F_x_i, -0.5*Dt*F_y_i)),
np.hstack((G_x_i, G_y_i))))
f_n_i = x_i - x - 0.5*Dt*(f_i+f)
Dxy_i = np.linalg.solve(-A_c_i,np.vstack((f_n_i,g_i)))
x_i = x_i + Dxy_i[0:N_x]
y_i = y_i + Dxy_i[N_x:(N_x+N_y)]
struct[i].x[:] = x_i
struct[i].y[:] = y_i
if np.max(np.abs(Dxy_i[:,0]))<1.0e-4:
break
# channels
if it >= it_store*decimation:
struct[i]['T'][it_store+1] = t
struct[i].X[it_store+1,:] = struct[i].x[:,0]
Y[it_store+1,:] = struct[i].y[:,0]
it_store += 1
return struct[i]['T'][:], struct[i].X[:], Y
@numba.njit(cache=True)
def perturbations(t,struct):
if t>0.02: struct[0].v_t_d=50.0
if t>0.02: struct[0].v_t_q=-326.6
if t>0.02: struct[0].v_s_d=0.0
if t>0.02: struct[0].v_s_q=-326.6
```
## Initialization
```
syst = rl2.rl2_class()
syst.solvern = 4
syst.decimation = 1
syst.t_end = 0.08
syst.Dt = 0.0001
syst.decimation =1
syst.update()
#syst.struct[0].H = x_pso[0]
#syst.struct[0].D = x_pso[1]
x0 = np.array([0,0,0,0,0,0]).reshape(6,1)
s = fsolve(syst.ini_problem,x0 )
syst.struct[0].x[:,0] = s[0:syst.N_x]
syst.struct[0].y[:,0] = s[syst.N_x:]
T,X,Y = solver(syst.struct)
syst.struct[0].x[:,0]
fig, axes = plt.subplots(nrows=2, ncols=1, figsize=(7, 4), sharex = True)
axes[0].plot(T[:-1], X[:-1,0])
axes[0].plot(T[:-1], X[:-1,1])
axes[1].plot(T[:-1], Y[:,2])
axes[1].plot(T[:-1], Y[:,3])
#axes[0].set_ylim([-30,100])
axes[0].set_xlim([0,0.08])
axes[0].grid(True)
syst.struct[0].x[:,0]
```
| github_jupyter |
# 9 - Advanced topics - 1 axis torque tube Shading for 1 day (Research Documentation)
## Recreating JPV 2019 / PVSC 2018 Fig. 13
Calculating and plotting shading from torque tube on 1-axis tracking for 1 day, which is figure 13 in:
Ayala Pelaez S, Deline C, Greenberg P, Stein JS, Kostuk RK. Model and validation of single-axis tracking with bifacial PV. IEEE J Photovoltaics. 2019;9(3):715–21. https://ieeexplore.ieee.org/document/8644027 and https://www.nrel.gov/docs/fy19osti/72039.pdf (pre-print, conference version)
This is what we will re-create:

Use bifacial_radiance minimum v. 0.3.1 or higher. Many things have been updated since this paper, simplifying the generation of this plot:
<ul>
<li> Sensor position is now always generated E to W on N-S tracking systems, so same sensor positions can just be added for this calculation at the end without needing to flip the sensors. </li>
<li> Torquetubes get automatically generated in makeModule. Following PVSC 2018 paper, rotation is around the modules and not around the torque tube axis (which is a new feature) </li>
<li> Simulating only 1 day on single-axis tracking easier with cumulativesky = False and gendaylit1axis(startdate='06/24', enddate='06/24' </li>
<li> Sensors get generated very close to surface, so all results are from the module surface and not the torquetube for this 1-UP case. </li>
</ul>
## Steps:
<ol>
<li> <a href='#step1'> Running the simulations for all the cases: </li>
<ol type='A'>
<li> <a href='#step1a'>Baseline Case: No Torque Tube </a></li>
<li> <a href='#step1b'> Zgap = 0.1 </a></li>
<li> <a href='#step1c'> Zgap = 0.2 </a></li>
<li> <a href='#step1d'> Zgap = 0.3 </a></li>
</ol>
<li> <a href='#step2'> Read-back the values and tabulate average values for unshaded, 10cm gap and 30cm gap </a></li>
<li> <a href='#step3'> Plot spatial loss values for 10cm and 30cm data </a></li>
<li> <a href='#step4'> Overall Shading Factor (for 1 day) </a></li>
</ol>
<a id='step1'></a>
### 1. Running the simulations for all the cases
```
import os
from pathlib import Path
testfolder = str(Path().resolve().parent.parent / 'bifacial_radiance' / 'TEMP')
# You can alternatively point to an empty directory (it will open a load GUI Visual Interface)
# or specify any other directory in your computer. I.E.:
# testfolder = r'C:\Users\sayala\Documents\RadianceScenes\Demo'
print ("Your simulation will be stored in %s" % testfolder)
# VARIABLES of the simulation:
lat = 35.1 # ABQ
lon = -106.7 # ABQ
x=1
y = 2
numpanels=1
limit_angle = 45 # tracker rotation limit angle
albedo = 'concrete' # ground albedo
hub_height = y*0.75 # H = 0.75
gcr = 0.35
pitch = y/gcr
#pitch = 1.0/gcr # Check from 1Axis_Shading_PVSC2018 file
cumulativesky = False # needed for set1axis and makeScene1axis so simulation is done hourly not with gencumsky.
limit_angle = 45 # tracker rotation limit angle
nMods=10
nRows=3
sensorsy = 200
module_type='2m_panel'
datewanted='06/24' # sunny day 6/24/1972 (index 4180 - 4195)
## Torque tube info
torquetube = False # redefined on each simulation below, since we need to compare with and without torque tube.
tubetype='round'
material = 'Metal_Grey'
diameter = 0.1
axisofrotationTorqueTube = False # Original PVSC version rotated around the modules like most other software.
# Variables that will get defined on each iteration below:
zgap = 0 # 0.2, 0.3 values tested. Re-defined on each simulation.
torquetube = False # baseline is no torque tube.
# Simulation Start.
try:
import bifacial_radiance
except ImportError:
raise RuntimeError('bifacial_radiance is required. download distribution')
import numpy as np
print(bifacial_radiance.__version__)
demo = bifacial_radiance.RadianceObj(path = testfolder)
demo.setGround(albedo)
epwfile = demo.getEPW(lat, lon)
metdata = demo.readEPW(epwfile)
trackerdict = demo.set1axis(metdata, limit_angle = limit_angle, backtrack = True, gcr = gcr, cumulativesky = False)
trackerdict = demo.gendaylit1axis(startdate=datewanted, enddate=datewanted)
sceneDict = {'pitch':pitch,'hub_height':hub_height, 'nMods': nMods, 'nRows': nRows}
```
<a id='step1a'></a>
### A. Baseline Case: No Torque Tube
When torquetube is False, zgap is the distance from axis of torque tube to module surface, but since we are rotating from the module's axis, this Zgap doesn't matter for this baseline case.
```
#CASE 0 No torque tube
# When torquetube is False, zgap is the distance from axis of torque tube to module surface, but since we are rotating from the module's axis, this Zgap doesn't matter.
# zgap = 0.1 + diameter/2.0
torquetube = False
customname = '_NoTT'
demo.makeModule(name=module_type,x=x,y=y, numpanels=numpanels, torquetube=torquetube, axisofrotationTorqueTube=axisofrotationTorqueTube)
trackerdict = demo.makeScene1axis(trackerdict,module_type,sceneDict, cumulativesky = cumulativesky)
trackerdict = demo.makeOct1axis(trackerdict)
trackerdict = demo.analysis1axis(trackerdict, sensorsy = sensorsy, customname = customname)
```
<a id='step1b'></a>
### B. ZGAP = 0.1
```
#ZGAP 0.1
zgap = 0.1
torquetube = True
customname = '_zgap0.1'
demo.makeModule(name=module_type,x=x,y=y, numpanels=numpanels,tubetype=tubetype, zgap=zgap, torquetube=torquetube, diameter=diameter, material=material, axisofrotationTorqueTube=axisofrotationTorqueTube)
trackerdict = demo.makeScene1axis(trackerdict,module_type,sceneDict, cumulativesky = cumulativesky)
trackerdict = demo.makeOct1axis(trackerdict)
trackerdict = demo.analysis1axis(trackerdict, sensorsy = sensorsy, customname = customname)
```
<a id='step1c'></a>
### C. ZGAP = 0.2
```
#ZGAP 0.2
zgap = 0.2
torquetube = True
customname = '_zgap0.2'
demo.makeModule(name=module_type,x=x,y=y, numpanels=numpanels,tubetype=tubetype, zgap=zgap, torquetube=torquetube, diameter=diameter, material=material, axisofrotationTorqueTube=axisofrotationTorqueTube)
trackerdict = demo.makeScene1axis(trackerdict,module_type,sceneDict, cumulativesky = cumulativesky)
trackerdict = demo.makeOct1axis(trackerdict)
trackerdict = demo.analysis1axis(trackerdict, sensorsy = sensorsy, customname = customname)
```
<a id='step1d'></a>
### D. ZGAP = 0.3
```
#ZGAP 0.3
zgap = 0.3
torquetube = True
customname = '_zgap0.3'
demo.makeModule(name=module_type,x=x,y=y, numpanels=numpanels,tubetype=tubetype, zgap=zgap, torquetube=torquetube, diameter=diameter, material=material, axisofrotationTorqueTube=axisofrotationTorqueTube)
trackerdict = demo.makeScene1axis(trackerdict,module_type,sceneDict, cumulativesky = cumulativesky)
trackerdict = demo.makeOct1axis(trackerdict)
trackerdict = demo.analysis1axis(trackerdict, sensorsy = sensorsy, customname = customname)
```
<a id='step2'></a>
### 2. Read-back the values and tabulate average values for unshaded, 10cm gap and 30cm gap
```
import glob
import pandas as pd
resultsfolder = os.path.join(testfolder, 'results')
print (resultsfolder)
filenames = glob.glob(os.path.join(resultsfolder,'*.csv'))
noTTlist = [k for k in filenames if 'NoTT' in k]
zgap10cmlist = [k for k in filenames if 'zgap0.1' in k]
zgap20cmlist = [k for k in filenames if 'zgap0.2' in k]
zgap30cmlist = [k for k in filenames if 'zgap0.3' in k]
# sum across all hours for each case
unsh_front = np.array([pd.read_csv(f, engine='python')['Wm2Front'] for f in noTTlist]).sum(axis = 0)
cm10_front = np.array([pd.read_csv(f, engine='python')['Wm2Front'] for f in zgap10cmlist]).sum(axis = 0)
cm20_front = np.array([pd.read_csv(f, engine='python')['Wm2Front'] for f in zgap20cmlist]).sum(axis = 0)
cm30_front = np.array([pd.read_csv(f, engine='python')['Wm2Front'] for f in zgap30cmlist]).sum(axis = 0)
unsh_back = np.array([pd.read_csv(f, engine='python')['Wm2Back'] for f in noTTlist]).sum(axis = 0)
cm10_back = np.array([pd.read_csv(f, engine='python')['Wm2Back'] for f in zgap10cmlist]).sum(axis = 0)
cm20_back = np.array([pd.read_csv(f, engine='python')['Wm2Back'] for f in zgap20cmlist]).sum(axis = 0)
cm30_back = np.array([pd.read_csv(f, engine='python')['Wm2Back'] for f in zgap30cmlist]).sum(axis = 0)
```
<a id='step3'></a>
### 3. plot spatial loss values for 10cm and 30cm data
```
import matplotlib.pyplot as plt
plt.rcParams['font.family'] = 'sans-serif'
plt.rcParams['font.sans-serif'] = ['Helvetica']
plt.rcParams['axes.linewidth'] = 0.2 #set the value globally
fig = plt.figure()
fig.set_size_inches(4, 2.5)
ax = fig.add_axes((0.15,0.15,0.78,0.75))
#plt.rc('font', family='sans-serif')
plt.rc('xtick',labelsize=8)
plt.rc('ytick',labelsize=8)
plt.rc('axes',labelsize=8)
plt.plot(np.linspace(-1,1,unsh_back.__len__()),(cm30_back - unsh_back)/unsh_back*100, label = '30cm gap',color = 'black') #steelblue
plt.plot(np.linspace(-1,1,unsh_back.__len__()),(cm20_back - unsh_back)/unsh_back*100, label = '20cm gap',color = 'steelblue', linestyle = '--') #steelblue
plt.plot(np.linspace(-1,1,unsh_back.__len__()),(cm10_back - unsh_back)/unsh_back*100, label = '10cm gap',color = 'darkorange') #steelblue
#plt.ylabel('$G_{rear}$ vs unshaded [Wm-2]')#(r'$BG_E$ [%]')
plt.ylabel('$G_{rear}$ / $G_{rear,tubeless}$ -1 [%]')
plt.xlabel('Module X position [m]')
plt.legend(fontsize = 8,frameon = False,loc='best')
#plt.ylim([0, 15])
plt.title('Torque tube shading loss',fontsize=9)
#plt.annotate('South',xy=(-10,9.5),fontsize = 8); plt.annotate('North',xy=(8,9.5),fontsize = 8)
plt.show()
```
<a id='step4'></a>
### 4. Overall Shading Loss Factor
To calculate shading loss factor, we can use the following equation:
<img src="../images_wiki/AdvancedJournals/Equation_ShadingFactor.PNG">
```
ShadingFactor = (1 - cm30_back.sum() / unsh_back.sum())*100
```
| github_jupyter |
# Named input
* **Difficulty level**: easy
* **Time need to learn**: 10 minutes or less
* **Key points**:
* Use dictionary or keyword arguments to specify labels of input
* `_input[name]` return subset of `_input `label `name`
## Named inputs
Let us first create a few temporary files as inputs of the examples
```
!touch a.txt b.txt ref.txt
```
<div class="bs-callout bs-callout-primary" role="alert">
<h4>Named inputs</h4>
<p>Keyword arguments in input to the input files and allow access to subsets of inputs with these labels</p>
</div>
in SoS, we usually specify one or more files as input of a SoS steps, and refer to them as variable `_input`:
```
input: 'a.txt', 'b.txt'
print(_input)
```
Using keyword parameters, you can assign labels to these files and access them separately:
```
input: A='a.txt', B='b.txt'
print(f'input of the substep is {_input}')
print(f'input of the substep with label A is {_input["A"]}')
print(f'input of the substep with label B is {_input["B"]}')
```
Note that although `_input['A']` and `_input['B']` are used to refer to subsets of `_input`, the variable `_input` can still be used and refers to all input files.
Named input can be used to pick a subset of input for the specification of step output. For example, in the following `print` statement, `_input["data"]`, `_input["reference"]` etc are used to obtain subsets of `_input`. These subsets of inputs are called **named inputs**. Here we use `group_by='pairlabel'` to group `step_input["data"]`. Please refer to [option `group_by`](group_by.html) for details.
```
input: data = ['a.txt', 'b.txt'], reference='ref.txt', group_by='pairlabel'
output: _input["data"].with_suffix('.res')
_output.touch()
print(f'''\
Input of step is {_input} with labels {step_input.labels}
Input data is {_input["data"]}
Reference is {_input["reference"]}
Output is {_output}
''')
```
In addition to the use of keyword arguments, you can use a dictionary directly to specify inputs with names:
```
%run
[global]
data = {
'data': ['a.txt', 'b.txt'],
'reference': 'ref.txt'
}
[10]
input: data, group_by='pairlabel'
output: _input["data"].with_suffix('.res')
_output.touch()
print(f'''\
Input of step is {_input} with labels {step_input.labels}
Input data is {_input["data"]}
Reference is {_input["reference"]}
Output is {_output}
''')
```
## Named input inherited from named output
Input created from [named output](named_output.html) will inherit their labels, unless the labels are overriden by keyword argument in the input statement.
<div class="bs-callout bs-callout-info" role="alert">
<h4>Inherit and override target labels</h4>
<p>Target lables are created and passed in SoS as follows</p>
<ul>
<li>Unnamed targets (targets specified with positional arguments) are labeled with step names</li>
<li>Labels are stored in variables <code>_input</code>, <code>_output</code>, <code>step_input</code> and <code>step_output</code>, and are passed by default to next step, or through functions <code>named_output</code> and <code>output_from</code></li>
<li>Keyword argument overrides default labels</li>
</ul>
</div>
For example, in the following workflow, step `default` gets the outputs from step `A` and `B` using function `output_from(['A', 'B'])`. Because the default labels for output from steps `A` and `B` are `A` and `B` respectively, you can differentiate the inputs using `_input['A']` and `_input['B']`.
```
%run -v0
[A]
output: 'a.txt'
_output.touch()
[B]
output: 'b.txt'
_output.touch()
[default]
input: output_from(['A', 'B'])
print(f'Input from step A is {_input["A"]}')
print(f'Input from step B is {_input["B"]}')
```
However, if you use keyword arguments in the input statement, the default or inherited labels will be overridden:
```
%run -v0
[A]
output: 'a.txt'
_output.touch()
[B]
output: 'b.txt'
_output.touch()
[default]
input: a_out=output_from('A'), b_out=output_from('B')
print(f'Input from step A is {_input["a_out"]}')
print(f'Input from step B is {_input["b_out"]}')
```
| github_jupyter |
# Generate a training set of images and annotations
We'll use the [PASCAL VOC2012](http://host.robots.ox.ac.uk/pascal/VOC/voc2012/) and [PASCAL-Part](http://www.stat.ucla.edu/~xianjie.chen/pascal_part_dataset/pascal_part.html) images and annotations as a training set for UNet.
Let's download both datasets first:
```
# Uncomment the following lines to download and extract VOC2012 and PASCAL-Part data
# !mkdir data
# !wget http://host.robots.ox.ac.uk/pascal/VOC/voc2012/VOCtrainval_11-May-2012.tar -P ./data
# !tar xf ./data/VOCtrainval_11-May-2012.tar -C ./data
# !rm -rf ./data/VOCtrainval_11-May-2012.tar
# !wget http://www.stat.ucla.edu/~xianjie.chen/pascal_part_dataset/trainval.tar.gz -P ./data
# !tar xfz ./data/trainval.tar.gz -C ./data
# !rm -rf ./data/trainval.tar.gz
import numpy as np
import os
import pickle
import warnings
from matplotlib.pyplot import imshow
from skimage.color import rgb2gray
from skimage.io import imread
from skimage.transform import resize
from anno import ImageAnnotation
warnings.filterwarnings('ignore')
%matplotlib inline
```
## Structure of input and output tensors
We'll feed the Convolutional Neural Network with 368x368 normalized grayscale frames (pixel values between 0 and 1). UNet seems to work better with a single input channel, but RGB input should be explored too. The output tensor consists of two 368x368 binary matrices. On the first one, a 1 means that the object to be segmented is being detected on that pixel or 0 otherwise. The second matrix is the opposite of the first, a 1 means that background is present in that pixel and a 0 means that the object is there.
We'll generate ground-truth data of cars to train a car-segmentation UNet model.
## Parse VOC2012 annotations
First we get a list of images that are segmented.
```
original_dir = './data/VOCdevkit/VOC2012/JPEGImages'
segmented_dir = './data/VOCdevkit/VOC2012/SegmentationClass'
with open('./data/VOCdevkit/VOC2012/ImageSets/Segmentation/trainval.txt', 'r') as f:
segmented = [line.strip() for line in f.readlines()]
```
Let's analyze a segmented car image:
```
name = '2007_001321'
original_img = imread(os.path.join(original_dir, '{}.jpg'.format(name)))
segment_img = imread(os.path.join(segmented_dir, '{}.png'.format(name)))
imshow(original_img)
imshow(segment_img)
colors = set()
for i in range(segment_img.shape[0]):
for j in range(segment_img.shape[0]):
r, g, b = segment_img[i, j, :]
colors.add((r, g, b))
print(colors)
```
Cars are segmented with gray (128, 128, 128) pixels in the VOC2012 annotations. We can then generate the correct output matrix for this segment pretty easily:
```
mask = np.all(segment_img == (128, 128, 128), axis=-1)
output_segment = np.zeros(mask.shape)
output_segment[mask] = 1.
imshow(output_segment)
```
Let's also explore a handy calculation which returns the percentage of the image covered by a car:
```
output_segment.sum() / output_segment.shape[0] / output_segment.shape[1]
```
56% of the image is covered by the car, this is useful when cars only fill a small percentage of the image and are too small to be segmented.
The last preprocess needed is converting the input to grayscale and resizing both input and output to a 368x368xChannels shape.
```
height = 368
width = 368
input_img = resize(rgb2gray(original_img), (height, width), anti_aliasing=True).reshape((height, width, 1))
imshow(input_img[:, :, 0])
input_img.shape
output_segment = resize(output_segment, (height, width), anti_aliasing=True)
output_segment[output_segment > 0.] = 1.
output_back = np.ones(output_segment.shape)
output_back[output_segment > 0.] = 0.
output = np.stack([output_segment, output_back], axis=-1)
imshow(output[:,:,0])
output.shape
```
## Generate VOC2012 train set
Now it's time to iterate through all the segmented images and add the ones with large enough cars to our train set.
Let's define the general constants to generate the train set which could be modified to generate a set of segments of any other class.
```
# Input/output sizes
height = 368
width = 368
# Class we want to detect
segment_color = (128, 128, 128) # Cars
# Percentage of the image covered by this class
threshold = 0.05
input_images = []
output_tensors = []
for name in segmented:
original_img = imread(os.path.join(original_dir, '{}.jpg'.format(name)))
segment_img = imread(os.path.join(segmented_dir, '{}.png'.format(name)))
mask = np.all(segment_img == segment_color, axis=-1)
output_segment = np.zeros(mask.shape)
output_segment[mask] = 1.
if output_segment.sum() / output_segment.shape[0] / output_segment.shape[1] > threshold:
image = resize(rgb2gray(original_img), (height, width), anti_aliasing=True).reshape((1, height, width, 1))
output_segment = resize(output_segment, (height, width), anti_aliasing=True)
output_segment[output_segment > 0.] = 1.
output_back = np.ones(output_segment.shape)
output_back[output_segment > 0.] = 0.
tensor = np.stack([output_segment, output_back], axis=-1).reshape((1, height, width, 2))
input_images.append(image)
output_tensors.append(tensor)
print("Images collected: {}".format(len(input_images)))
```
Let's check we have gathered the correct images:
```
imshow(input_images[32][0, :, :, 0])
imshow(output_tensors[32][0, :, :, 0])
```
## Parse PASCAL-Part Dataset
The 147 images from VOC2012 may be not sufficient to train a good model. There's another useful annotated dataset that we've downloaded: [PASCAL-Part](http://www.stat.ucla.edu/~xianjie.chen/pascal_part_dataset/pascal_part.html). It is based on the VOC2010 data, so we expect some of the images to be repeated. We only save those that aren't already part of the VOC2012 segmented images.
This dataset comes in `.mat` format, we use slightly modified [pascal-part-py](https://github.com/twuilliam/pascal-part-py) helpers to quickly parse these files.
```
annotations_dir = './data/Annotations_Part/'
voc2012 = set(segmented)
segmented = []
for annotation in os.listdir(annotations_dir):
name = annotation.split('.')[0]
if name in voc2012:
continue
ann = ImageAnnotation(
os.path.join(original_dir, '{}.jpg'.format(name)),
os.path.join(annotations_dir, '{}.mat'.format(name))
)
if 'car' in [ann.objects[i].class_name for i in range(ann.n_objects)]:
segmented.append(name)
print("There is an additional number of {0} images of cars in the PASCAL-Part dataset.".format(len(segmented)))
```
## Generate PASCAL-Part train set
Let's add those images where cars occupy a large enough portion of the image to the train set:
```
for name in segmented:
ann = ImageAnnotation(
os.path.join(original_dir, '{}.jpg'.format(name)),
os.path.join(annotations_dir, '{}.mat'.format(name))
)
if ann.part_mask.sum() / ann.part_mask.shape[0] / ann.part_mask.shape[1] > threshold:
image = resize(rgb2gray(ann.im), (height, width), anti_aliasing=True).reshape(1, height, width, 1)
output_segment = resize(ann.part_mask, (height, width), anti_aliasing=True)
output_segment[output_segment > 0.] = 1.
output_back = np.ones(output_segment.shape)
output_back[output_segment > 0.] = 0.
tensor = np.stack([output_segment, output_back], axis=-1).reshape((1, height, width, 2))
input_images.append(image)
output_tensors.append(tensor)
```
Again, let's check we have gathered the correct images:
```
imshow(input_images[-1][0, :, :, 0])
imshow(output_tensors[-1][0, :, :, 0])
print("Images collected: {}".format(len(input_images)))
```
## Concatenate and pickle final dataset
Let's save input and output tensors as *ndarray*s of shape (N, height, width, channels) and pickle them for training UNet networks.
```
input_images = np.concatenate(input_images, axis=0)
output_tensors = np.concatenate(output_tensors, axis=0)
print("Input tensor shape: {}".format(input_images.shape))
print("Output tensor shape: {}".format(output_tensors.shape))
with open('train_images.pkl', 'wb') as fout:
pickle.dump(input_images, fout, protocol=4)
with open('train_labels.pkl', 'wb') as fout:
pickle.dump(output_tensors, fout, protocol=4)
```
| github_jupyter |
# Dependencies
```
import os, warnings, shutil, re
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from transformers import AutoTokenizer
from sklearn.utils import shuffle
from sklearn.model_selection import StratifiedKFold
SEED = 0
warnings.filterwarnings("ignore")
pd.set_option('max_colwidth', 160)
# Preprocess data
def clean_text(text):
text = str(text)
text = re.sub(r'[0-9"]', '', text) # number
text = re.sub(r'#[\S]+\b', '', text) # hash
text = re.sub(r'@[\S]+\b', '', text) # mention
text = re.sub(r'https?\S+', '', text) # link
text = re.sub(r'\s+', ' ', text) # multiple white spaces
# text = re.sub(r'\W+', ' ', text) # non-alphanumeric
return text.strip()\
# Preprocess inputs
CLS = 0
PAD = 1
SEP = 2
def preprocess_roberta(text, tokenizer, max_seq_len):
encoded = tokenizer.encode_plus(text, return_token_type_ids=False)
# Truncate
input_ids = encoded['input_ids'][:max_seq_len]
attention_mask = encoded['attention_mask'][:max_seq_len]
# Update input_ids
input_ids[0] = CLS
input_ids[-1] = SEP
# Pad
input_ids = input_ids + [PAD] * (max_seq_len - len(input_ids))
attention_mask = attention_mask + [0] * (max_seq_len - len(attention_mask))
x = [np.asarray(input_ids, dtype=np.int32),
np.asarray(attention_mask, dtype=np.int8)]
return x
def preprocess_roberta_tail(text, tokenizer, max_seq_len):
encoded = tokenizer.encode_plus(text, return_token_type_ids=False)
# Truncate
input_ids = encoded['input_ids'][-max_seq_len:]
attention_mask = encoded['attention_mask'][-max_seq_len:]
# Update input_ids
input_ids[0] = CLS
input_ids[-1] = SEP
# Pad
input_ids = input_ids + [PAD] * (max_seq_len - len(input_ids))
attention_mask = attention_mask + [0] * (max_seq_len - len(attention_mask))
x = [np.asarray(input_ids, dtype=np.int32),
np.asarray(attention_mask, dtype=np.int8)]
# if len(encoded['input_ids']) > max_seq_len:
# return x
# else:
# return None
return x
def preprocess_roberta_tail_test(text, tokenizer, max_seq_len):
encoded = tokenizer.encode_plus(text, return_token_type_ids=False)
# Truncate
input_ids = encoded['input_ids'][-max_seq_len:]
attention_mask = encoded['attention_mask'][-max_seq_len:]
# Update input_ids
input_ids[0] = CLS
input_ids[-1] = SEP
# Pad
input_ids = input_ids + [PAD] * (max_seq_len - len(input_ids))
attention_mask = attention_mask + [0] * (max_seq_len - len(attention_mask))
x = [np.asarray(input_ids, dtype=np.int32),
np.asarray(attention_mask, dtype=np.int8)]
return x
def get_data(df, text_col, tokenizer, MAX_LEN, preprocess_fn=preprocess_roberta):
x_input_ids = []
x_attention_masks = []
y_data = []
y_data_int = []
for row in df.itertuples():
x = preprocess_fn(getattr(row, text_col), tokenizer, MAX_LEN)
if x is not None:
x_input_ids.append(x[0])
x_attention_masks.append(x[1])
y_data.append(getattr(row, 'toxic'))
y_data_int.append(getattr(row, 'toxic_int'))
x_data = [np.asarray(x_input_ids, dtype=np.int32),
np.asarray(x_attention_masks, dtype=np.int8)]
y_data = [np.asarray(y_data, dtype=np.float32),
np.asarray(y_data_int, dtype=np.int8)]
return x_data, y_data
def get_data_test(df, text_col, tokenizer, MAX_LEN, preprocess_fn=preprocess_roberta):
x_input_ids = []
x_attention_masks = []
for row in df.itertuples():
x = preprocess_fn(getattr(row, text_col), tokenizer, MAX_LEN)
x_input_ids.append(x[0])
x_attention_masks.append(x[1])
x_data = [np.asarray(x_input_ids, dtype=np.int32),
np.asarray(x_attention_masks, dtype=np.int8)]
return x_data
```
# Parameters
```
MAX_LEN = 192
tokenizer_path = 'jplu/tf-xlm-roberta-large'
sample_rate = 1
```
# Load data
```
train1 = pd.read_csv("/kaggle/input/jigsaw-multilingual-toxic-comment-classification/jigsaw-toxic-comment-train.csv")
train2 = pd.read_csv("/kaggle/input/jigsaw-multilingual-toxic-comment-classification/jigsaw-unintended-bias-train.csv")
valid_df = pd.read_csv("/kaggle/input/jigsaw-multilingual-toxic-comment-classification/validation.csv", usecols=['comment_text', 'toxic', 'lang'])
test_df = pd.read_csv("/kaggle/input/jigsaw-multilingual-toxic-comment-classification/test.csv", usecols=['content'])
n_neg_samp_df1 = len(train1.query('toxic > .5')) * sample_rate
n_neg_samp_df2 = len(train2.query('toxic > .5')) * sample_rate
train_df = pd.concat([train1[['comment_text', 'toxic']].query('toxic > .5'),
train1[['comment_text', 'toxic']].query('toxic <= .5').sample(n=n_neg_samp_df1, random_state=SEED),
train2[['comment_text', 'toxic']].query('toxic > .5'),
train2[['comment_text', 'toxic']].query('toxic <= .5').sample(n=n_neg_samp_df2, random_state=SEED)
])
print(f'Dataframe 1 samples: toxic|non-toxic | {n_neg_samp_df1 // sample_rate}|{n_neg_samp_df1}')
print(f'Dataframe 2 samples: toxic|non-toxic | {n_neg_samp_df2 // sample_rate}|{n_neg_samp_df2}')
print('Train samples %d' % len(train_df))
display(train_df.head())
display(train_df.describe())
print('Validation samples %d' % len(valid_df))
display(valid_df.head())
display(valid_df.describe())
print('Test samples %d' % len(test_df))
display(test_df.head())
display(test_df.describe())
```
# Tokenizer
```
tokenizer = AutoTokenizer.from_pretrained(tokenizer_path, lowercase=False)
```
## Preprocess
```
# Train
train_df['comment_text'] = train_df.apply(lambda x: clean_text(x['comment_text']), axis=1)
train_df['length'] = train_df['comment_text'].apply(lambda x : len(x))
train_df['word_count'] = train_df['comment_text'].apply(lambda x : len(x.split(' ')))
train_df['token_count'] = train_df['comment_text'].apply(lambda x : len(tokenizer.encode(x)))
train_df['toxic_int'] = train_df['toxic'].round().astype(int)
# Validation
valid_df['comment_text'] = valid_df.apply(lambda x: clean_text(x['comment_text']), axis=1)
valid_df['length'] = valid_df['comment_text'].apply(lambda x : len(x))
valid_df['word_count'] = valid_df['comment_text'].apply(lambda x : len(x.split(' ')))
valid_df['token_count'] = valid_df['comment_text'].apply(lambda x : len(tokenizer.encode(x)))
valid_df['toxic_int'] = valid_df['toxic'].round().astype(int)
# Test
test_df['content'] = test_df.apply(lambda x: clean_text(x['content']), axis=1)
test_df['length'] = test_df['content'].apply(lambda x : len(x))
test_df['word_count'] = test_df['content'].apply(lambda x : len(x.split(' ')))
test_df['token_count'] = test_df['content'].apply(lambda x : len(tokenizer.encode(x)))
print('Train samples %d' % len(train_df))
display(train_df.head())
display(train_df.describe())
print('Validation samples %d' % len(valid_df))
display(valid_df.head())
display(valid_df.describe())
print('Test samples %d' % len(test_df))
display(test_df.head())
display(test_df.describe())
```
## Filter
```
# train_df = train_df[train_df['word_count'] <= 250]
# train_df = train_df[train_df['token_count'] <= 1000]
train_df = shuffle(train_df, random_state=SEED).reset_index(drop=True)
print('Train samples %d' % len(train_df))
display(train_df.head())
display(train_df.describe())
```
# Data generation sanity check
```
for idx in range(5):
print('\nRow %d' % idx)
max_seq_len = 32
comment_text = train_df['comment_text'].loc[idx]
x_train, y_train = get_data(train_df[idx:idx+1], 'comment_text', tokenizer, max_seq_len, preprocess_fn=preprocess_roberta)
print('label : "%.4f"' % y_train[0])
print('label int : "%d"' % y_train[1])
print('comment_text : "%s"' % comment_text)
print('input_ids : "%s"' % x_train[0][0])
print('attention_mask: "%s"' % x_train[1][0])
x_train, _ = get_data(train_df[idx:idx+1], 'comment_text', tokenizer, max_seq_len, preprocess_fn=preprocess_roberta_tail)
if len(x_train[0]) > 0:
print('-------------------- TAIL --------------------')
print('input_ids : "%s"' % x_train[0][0])
print('attention_mask: "%s"' % x_train[1][0])
assert len(x_train[0][0]) == len(x_train[1][0]) == max_seq_len
```
# 5-Fold split
```
folds = StratifiedKFold(n_splits=5, shuffle=True, random_state=SEED)
for fold_n, (train_idx, val_idx) in enumerate(folds.split(train_df, train_df['toxic_int'])):
print('Fold: %s, Train size: %s, Validation size %s' % (fold_n+1, len(train_idx), len(val_idx)))
train_df[('fold_%s' % str(fold_n+1))] = 0
train_df[('fold_%s' % str(fold_n+1))].loc[train_idx] = 'train'
train_df[('fold_%s' % str(fold_n+1))].loc[val_idx] = 'validation'
```
# Label distribution
```
for fold_n in range(folds.n_splits):
fold_n += 1
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(18, 6))
fig.suptitle('Fold %s' % fold_n, fontsize=22)
sns.countplot(x="toxic_int", data=train_df[train_df[('fold_%s' % fold_n)] == 'train'], palette="GnBu_d", ax=ax1).set_title('Train')
sns.countplot(x="toxic_int", data=train_df[train_df[('fold_%s' % fold_n)] == 'validation'], palette="GnBu_d", ax=ax2).set_title('Validation')
sns.despine()
plt.show()
for fold_n in range(folds.n_splits):
fold_n += 1
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(18, 6))
fig.suptitle('Fold %s' % fold_n, fontsize=22)
sns.distplot(train_df[train_df[('fold_%s' % fold_n)] == 'train']['toxic'], ax=ax1).set_title('Train')
sns.distplot(train_df[train_df[('fold_%s' % fold_n)] == 'validation']['toxic'], ax=ax2).set_title('Validation')
sns.despine()
plt.show()
```
# Output 5-fold set
```
train_df.to_csv('5-fold.csv', index=False)
display(train_df.head())
for fold_n in range(folds.n_splits):
if fold_n >= 3:
fold_n += 1
base_path = 'fold_%d/' % fold_n
# Create dir
os.makedirs(base_path)
x_train, y_train = get_data(train_df[train_df[('fold_%s' % fold_n)] == 'train'], 'comment_text',
tokenizer, MAX_LEN, preprocess_fn=preprocess_roberta)
x_valid, y_valid = get_data(train_df[train_df[('fold_%s' % fold_n)] == 'validation'], 'comment_text',
tokenizer, MAX_LEN, preprocess_fn=preprocess_roberta)
y_train_float, y_train_int = y_train
y_valid_float, y_valid_int = y_valid
x_train, y_train_float, y_train_int = np.asarray(x_train), np.asarray(y_train_float), np.asarray(y_train_int)
x_valid, y_valid_float, y_valid_int = np.asarray(x_valid), np.asarray(y_valid_float), np.asarray(y_valid_int)
np.save(base_path + 'x_train', x_train)
np.save(base_path + 'y_train', y_train_float)
np.save(base_path + 'y_train_int', y_train_int)
np.save(base_path + 'x_valid', x_valid)
np.save(base_path + 'y_valid', y_valid_float)
np.save(base_path + 'y_valid_int', y_valid_int)
print('\nFOLD: %d' % (fold_n))
print('x_train shape:', x_train.shape)
print('y_train_float shape:', y_train_float.shape)
print('y_train_int shape:', y_train_int.shape)
print('x_valid shape:', x_valid.shape)
print('y_valid_float shape:', y_valid_float.shape)
print('y_valid_int shape:', y_valid_int.shape)
#################### ENCODE TAIL ONLY ####################
x_train, y_train = get_data(train_df[train_df[('fold_%s' % fold_n)] == 'train'], 'comment_text',
tokenizer, MAX_LEN, preprocess_fn=preprocess_roberta_tail)
x_valid, y_valid = get_data(train_df[train_df[('fold_%s' % fold_n)] == 'validation'], 'comment_text',
tokenizer, MAX_LEN, preprocess_fn=preprocess_roberta_tail)
y_train_float, y_train_int = y_train
y_valid_float, y_valid_int = y_valid
x_train, y_train_float, y_train_int = np.asarray(x_train), np.asarray(y_train_float), np.asarray(y_train_int)
x_valid, y_valid_float, y_valid_int = np.asarray(x_valid), np.asarray(y_valid_float), np.asarray(y_valid_int)
np.save(base_path + 'x_train_tail', x_train)
np.save(base_path + 'y_train_tail', y_train_float)
np.save(base_path + 'y_train_int_tail', y_train_int)
np.save(base_path + 'x_valid_tail', x_valid)
np.save(base_path + 'y_valid_tail', y_valid_float)
np.save(base_path + 'y_valid_int_tail', y_valid_int)
print('\nFOLD: %d [TAIL]' % (fold_n))
print('x_train shape:', x_train.shape)
print('y_train_float shape:', y_train_float.shape)
print('y_train_int shape:', y_train_int.shape)
print('x_valid shape:', x_valid.shape)
print('y_valid_float shape:', y_valid_float.shape)
print('y_valid_int shape:', y_valid_int.shape)
# Compress logs dir
# !tar -czf fold_1.tar.gz fold_1
# !tar -czf fold_2.tar.gz fold_2
# !tar -czf fold_3.tar.gz fold_3
!tar -czf fold_4.tar.gz fold_4
!tar -czf fold_5.tar.gz fold_5
# Delete logs dir
# shutil.rmtree('fold_1')
# shutil.rmtree('fold_2')
# shutil.rmtree('fold_3')
shutil.rmtree('fold_4')
shutil.rmtree('fold_5')
```
# Validation set
```
display(valid_df.head())
display(valid_df.describe())
x_valid, y_valid = get_data(valid_df, 'comment_text', tokenizer, MAX_LEN, preprocess_fn=preprocess_roberta)
y_valid_float, y_valid_int = y_valid
x_valid, y_valid_float, y_valid_int = np.asarray(x_valid), np.asarray(y_valid_float), np.asarray(y_valid_int)
np.save('x_valid', x_valid)
np.save('y_valid', y_valid_float)
np.save('y_valid_int', y_valid_int)
print('x_valid shape:', x_valid.shape)
print('y_valid_float shape:', y_valid_float.shape)
print('y_valid_int shape:', y_valid_int.shape)
#################### ENCODE TAIL ONLY ####################
x_valid, y_valid = get_data(valid_df, 'comment_text', tokenizer, MAX_LEN, preprocess_fn=preprocess_roberta_tail)
y_valid_float, y_valid_int = y_valid
x_valid, y_valid_float, y_valid_int = np.asarray(x_valid), np.asarray(y_valid_float), np.asarray(y_valid_int)
np.save('x_valid_tail', x_valid)
np.save('y_valid_tail', y_valid_float)
np.save('y_valid_int_tail', y_valid_int)
print('x_valid shape:', x_valid.shape)
print('y_valid_float shape:', y_valid_float.shape)
print('y_valid_int shape:', y_valid_int.shape)
```
# Test set
```
display(test_df.head())
display(test_df.describe())
x_test = get_data_test(test_df, 'content', tokenizer, MAX_LEN, preprocess_fn=preprocess_roberta)
x_test = np.asarray(x_test)
np.save('x_test', x_test)
print('x_test shape:', x_test.shape)
#################### ENCODE TAIL ONLY ####################
x_test = get_data_test(test_df, 'content', tokenizer, MAX_LEN, preprocess_fn=preprocess_roberta_tail_test)
x_test = np.asarray(x_test)
np.save('x_test_tail', x_test)
print('[TAIL] x_test shape:', x_test.shape)
```
# Test set EDA
## Word count distribution
```
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(20, 8.7), sharex=True)
sns.distplot(train_df['word_count'], ax=ax1).set_title("Train")
sns.distplot(test_df['word_count'], ax=ax2).set_title("Test")
sns.despine()
plt.show()
```
## Token count distribution
```
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(20, 8.7), sharex=True)
sns.distplot(train_df['token_count'], ax=ax1).set_title("Train")
sns.distplot(test_df['token_count'], ax=ax2).set_title("Test")
sns.despine()
plt.show()
```
| github_jupyter |
```
# Copyright 2021 Google LLC
# Use of this source code is governed by an MIT-style
# license that can be found in the LICENSE file or at
# https://opensource.org/licenses/MIT.
# Notebook authors: Kevin P. Murphy (murphyk@gmail.com)
# and Mahmoud Soliman (mjs@aucegypt.edu)
# This notebook reproduces figures for chapter 16 from the book
# "Probabilistic Machine Learning: An Introduction"
# by Kevin Murphy (MIT Press, 2021).
# Book pdf is available from http://probml.ai
```
<a href="https://opensource.org/licenses/MIT" target="_parent"><img src="https://img.shields.io/github/license/probml/pyprobml"/></a>
<a href="https://colab.research.google.com/github/probml/pyprobml/blob/master/book1/figures/chapter16_exemplar-based_methods_figures.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Figure 16.1:<a name='16.1'></a> <a name='knn'></a>
(a) Illustration of a $K$-nearest neighbors classifier in 2d for $K=5$. The nearest neighbors of test point $\mathbf x $ have labels $\ 1, 1, 1, 0, 0\ $, so we predict $p(y=1|\mathbf x , \mathcal D ) = 3/5$. (b) Illustration of the Voronoi tesselation induced by 1-NN. Adapted from Figure 4.13 of <a href='#Duda01'>[DHS01]</a> .
Figure(s) generated by [knn_voronoi_plot.py](https://github.com/probml/pyprobml/blob/master/scripts/knn_voronoi_plot.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
google.colab.files.view("./knn_voronoi_plot.py")
%run knn_voronoi_plot.py
```
## Figure 16.2:<a name='16.2'></a> <a name='knnThreeClass'></a>
Decision boundaries induced by a KNN classifier. (a) $K=1$. (b) $K=2$. (c) $K=5$. (d) Train and test error vs $K$.
Figure(s) generated by [knn_classify_demo.py](https://github.com/probml/pyprobml/blob/master/scripts/knn_classify_demo.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
google.colab.files.view("./knn_classify_demo.py")
%run knn_classify_demo.py
```
## Figure 16.3:<a name='16.3'></a> <a name='curse'></a>
Illustration of the curse of dimensionality. (a) We embed a small cube of side $s$ inside a larger unit cube. (b) We plot the edge length of a cube needed to cover a given volume of the unit cube as a function of the number of dimensions. Adapted from Figure 2.6 from <a href='#HastieBook'>[HTF09]</a> .
Figure(s) generated by [curse_dimensionality.py](https://github.com/probml/pyprobml/blob/master/scripts/curse_dimensionality.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
google.colab.files.view("./curse_dimensionality.py")
%run curse_dimensionality.py
```
## Figure 16.4:<a name='16.4'></a> <a name='LCA'></a>
Illustration of latent coincidence analysis (LCA) as a directed graphical model. The inputs $\mathbf x , \mathbf x ' \in \mathbb R ^D$ are mapped into Gaussian latent variables $\mathbf z , \mathbf z ' \in \mathbb R ^L$ via a linear mapping $\mathbf W $. If the two latent points coincide (within length scale $\kappa $) then we set the similarity label to $y=1$, otherwise we set it to $y=0$. From Figure 1 of <a href='#Der2012'>[ML12]</a> . Used with kind permission of Lawrence Saul.
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
show_image("/pyprobml/book1/figures/images/Figure_16.4.png")
```
## Figure 16.5:<a name='16.5'></a> <a name='tripletNet'></a>
Networks for deep metric learning. (a) Siamese network. (b) Triplet network. Adapted from Figure 5 of <a href='#Kaya2019'>[MH19]</a> .
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
show_image("/pyprobml/book1/figures/images/Figure_16.5_A.png")
show_image("/pyprobml/book1/figures/images/Figure_16.5_B.png")
```
## Figure 16.6:<a name='16.6'></a> <a name='tripletBound'></a>
Speeding up triplet loss minimization. (a) Illustration of hard vs easy negatives. Here $a$ is the anchor point, $p$ is a positive point, and $n_i$ are negative points. Adapted from Figure 4 of <a href='#Kaya2019'>[MH19]</a> . (b) Standard triplet loss would take $8 \times 3 \times 4 = 96$ calculations, whereas using a proxy loss (with one proxy per class) takes $8 \times 2 = 16$ calculations. From Figure 1 of <a href='#Do2019cvpr'>[Tha+19]</a> . Used with kind permission of Gustavo Cerneiro.
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
show_image("/pyprobml/book1/figures/images/Figure_16.6_A.png")
show_image("/pyprobml/book1/figures/images/Figure_16.6_B.png")
```
## Figure 16.7:<a name='16.7'></a> <a name='SEC'></a>
Adding spherical embedding constraint to a deep metric learning method. Used with kind permission of Dingyi Zhang.
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
show_image("/pyprobml/book1/figures/images/Figure_16.7.png")
```
## Figure 16.8:<a name='16.8'></a> <a name='smoothingKernels'></a>
A comparison of some popular normalized kernels.
Figure(s) generated by [smoothingKernelPlot.py](https://github.com/probml/pyprobml/blob/master/scripts/smoothingKernelPlot.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
google.colab.files.view("./smoothingKernelPlot.py")
%run smoothingKernelPlot.py
```
## Figure 16.9:<a name='16.9'></a> <a name='parzen'></a>
A nonparametric (Parzen) density estimator in 1d estimated from 6 data points, denoted by x. Top row: uniform kernel. Bottom row: Gaussian kernel. Left column: bandwidth parameter $h=1$. Right column: bandwidth parameter $h=2$. Adapted from http://en.wikipedia.org/wiki/Kernel_density_estimation .
Figure(s) generated by [Kernel_density_estimation](http://en.wikipedia.org/wiki/Kernel_density_estimation) [parzen_window_demo2.py](https://github.com/probml/pyprobml/blob/master/scripts/parzen_window_demo2.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
google.colab.files.view("./parzen_window_demo2.py")
%run parzen_window_demo2.py
```
## Figure 16.10:<a name='16.10'></a> <a name='kernelRegression'></a>
An example of kernel regression in 1d using a Gaussian kernel.
Figure(s) generated by [kernelRegressionDemo.py](https://github.com/probml/pyprobml/blob/master/scripts/kernelRegressionDemo.py)
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
import pyprobml_utils as pml
import colab_utils
import os
os.environ["PYPROBML"] = ".." # one above current scripts directory
import google.colab
from google.colab.patches import cv2_imshow
%reload_ext autoreload
%autoreload 2
def show_image(img_path,size=None,ratio=None):
img = colab_utils.image_resize(img_path, size)
cv2_imshow(img)
print('finished!')
google.colab.files.view("./kernelRegressionDemo.py")
%run kernelRegressionDemo.py
```
## References:
<a name='Duda01'>[DHS01]</a> R. O. Duda, P. E. Hart and D. G. Stork. "Pattern Classification". (2001).
<a name='HastieBook'>[HTF09]</a> T. Hastie, R. Tibshirani and J. Friedman. "The Elements of Statistical Learning". (2009).
<a name='Kaya2019'>[MH19]</a> K. Mahmut and B. HasanSakir. "Deep Metric Learning: A Survey". In: Symmetry (2019).
<a name='Der2012'>[ML12]</a> D. Matthew and S. LawrenceK. "Latent Coincidence Analysis: A Hidden Variable Model forDistance Metric Learning". (2012).
<a name='Do2019cvpr'>[Tha+19]</a> D. Thanh-Toan, T. Toan, R. Ian, K. Vijay, H. Tuan and C. Gustavo. "A Theoretically Sound Upper Bound on the Triplet Loss forImproving the Efficiency of Deep Distance Metric Learning". (2019).
| github_jupyter |
## Hello World with cuDF and Streamz
This notebook demonstrates use of cuDF to perform streaming word-count using a small portion of the [Streamz API](https://streamz.readthedocs.io/en/latest/).
This notebook was tested using the `rapidsai/rapidsai-dev-nightly:0.10-cuda10.0-devel-ubuntu18.04-py3.7` container from [DockerHub](https://hub.docker.com/r/rapidsai/rapidsai-nightly) and run on the NVIDIA GV100 GPU. Please be aware that your system may be different and you may need to modify the code or install packages to run the below examples.
If you think you have found a bug or an error, please file an issue here: https://github.com/rapidsai/notebooks-contrib/issues
First, make sure you have installed the [Streamz](https://github.com/python-streamz/streamz) library.
```
!conda install -c conda-forge -y streamz
```
## Getting Started
First import the required packages. We'll be programmatically generating data and process in streaming batches.
```
from streamz import Stream
import cudf, json
# create a list of static messages
messages = [
{'msg': 'Hello, World!'},
{'msg': 'hi, world!'},
{'msg': 'hey world'},
{'msg': 'Hi'}
]
```
## Define A Function to Run Per Batch
While some streaming systems deal with single events at a time, using GPUs to run a per-event process is not ideal due to the high latency of PCI-E memory transfers and kernel call overhead.
For our example, we'll focus on processing batches at a time.
```
# define function to run per batch
def process_on_gpu(messages):
# read the batch of messages into a GPU DataFrame
df = cudf.read_json('\n'.join(messages), lines=True)
# split each line into columns, one per word
tmp = df['msg'].str.split()
# combine all word columns into a single Series
words = tmp[tmp.columns[0]]
for word in tmp.columns[1:]:
words = words.append(tmp[word])
# remove punctuation, lower-case
words = words.str.fillna('').replace(',', '').replace('!', '').str.lower()
# compute and return word counts for the batch
return str(words.groupby(words).count())
```
## Setup the Stream and Emit Data
```
source = Stream()
# GPUs like to process sizable chunks of data at once
# source.partition(n) sends n events at a time to downstream functions
source.partition(10).map(process_on_gpu).sink(print)
# with 30 events partitioned by 10 events per group will give 3 "batches"
n_messages = 30
for idx in range(0, n_messages):
source.emit(json.dumps(messages[idx % len(messages)]))
```
| github_jupyter |
# WorkFlow
## Classes
## Load the data
## Test Modelling
## Modelling
**<hr>**
## Classes
```
NAME = "change the conv2d"
BATCH_SIZE = 32
import os
import cv2
import torch
import numpy as np
def load_data(img_size=112):
data = []
index = -1
labels = {}
for directory in os.listdir('./data/'):
index += 1
labels[f'./data/{directory}/'] = [index,-1]
print(len(labels))
for label in labels:
for file in os.listdir(label):
filepath = label + file
img = cv2.imread(filepath,cv2.IMREAD_GRAYSCALE)
img = cv2.resize(img,(img_size,img_size))
img = img / 255.0
data.append([
np.array(img),
labels[label][0]
])
labels[label][1] += 1
for _ in range(12):
np.random.shuffle(data)
print(len(data))
np.save('./data.npy',data)
return data
import torch
def other_loading_data_proccess(data):
X = []
y = []
print('going through the data..')
for d in data:
X.append(d[0])
y.append(d[1])
print('splitting the data')
VAL_SPLIT = 0.25
VAL_SPLIT = len(X)*VAL_SPLIT
VAL_SPLIT = int(VAL_SPLIT)
X_train = X[:-VAL_SPLIT]
y_train = y[:-VAL_SPLIT]
X_test = X[-VAL_SPLIT:]
y_test = y[-VAL_SPLIT:]
print('turning data to tensors')
X_train = torch.from_numpy(np.array(X_train))
y_train = torch.from_numpy(np.array(y_train))
X_test = torch.from_numpy(np.array(X_test))
y_test = torch.from_numpy(np.array(y_test))
return [X_train,X_test,y_train,y_test]
```
**<hr>**
## Load the data
```
REBUILD_DATA = True
if REBUILD_DATA:
data = load_data()
np.random.shuffle(data)
X_train,X_test,y_train,y_test = other_loading_data_proccess(data)
```
## Test Modelling
```
import torch
import torch.nn as nn
import torch.nn.functional as F
# class Test_Model(nn.Module):
# def __init__(self):
# super().__init__()
# self.conv1 = nn.Conv2d(1, 6, 5)
# self.pool = nn.MaxPool2d(2, 2)
# self.conv2 = nn.Conv2d(6, 16, 5)
# self.fc1 = nn.Linear(16 * 25 * 25, 120)
# self.fc2 = nn.Linear(120, 84)
# self.fc3 = nn.Linear(84, 36)
# def forward(self, x):
# x = self.pool(F.relu(self.conv1(x)))
# x = self.pool(F.relu(self.conv2(x)))
# x = x.view(-1, 16 * 25 * 25)
# x = F.relu(self.fc1(x))
# x = F.relu(self.fc2(x))
# x = self.fc3(x)
# return x
class Test_Model(nn.Module):
def __init__(self):
super().__init__()
self.pool = nn.MaxPool2d(2, 2)
self.conv1 = nn.Conv2d(1, 32, 5)
self.conv3 = nn.Conv2d(32,64,5)
self.conv2 = nn.Conv2d(64, 128, 5)
self.fc1 = nn.Linear(128 * 10 * 10, 512)
self.fc2 = nn.Linear(512, 256)
self.fc4 = nn.Linear(256,128)
self.fc3 = nn.Linear(128, 36)
def forward(self, x,shape=False):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv3(x)))
x = self.pool(F.relu(self.conv2(x)))
if shape:
print(x.shape)
x = x.view(-1, 128 * 10 * 10)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.relu(self.fc4(x))
x = self.fc3(x)
return x
device = torch.device('cuda')
model = Test_Model().to(device)
preds = model(X_test.reshape(-1,1,112,112).float().to(device),True)
preds[0]
optimizer = torch.optim.SGD(model.parameters(),lr=0.1)
criterion = nn.CrossEntropyLoss()
EPOCHS = 5
loss_logs = []
from tqdm import tqdm
PROJECT_NAME = "Sign-Language-Recognition"
def test(net,X,y):
correct = 0
total = 0
net.eval()
with torch.no_grad():
for i in range(len(X)):
real_class = torch.argmax(y[i]).to(device)
net_out = net(X[i].view(-1,1,112,112).to(device).float())
net_out = net_out[0]
predictied_class = torch.argmax(net_out)
if predictied_class == real_class:
correct += 1
total += 1
return round(correct/total,3)
import wandb
len(os.listdir('./data/'))
import random
# index = random.randint(0,29)
# print(index)
# wandb.init(project=PROJECT_NAME,name=NAME)
# for _ in tqdm(range(EPOCHS)):
# for i in range(0,len(X_train),BATCH_SIZE):
# X_batch = X_train[i:i+BATCH_SIZE].view(-1,1,112,112).to(device)
# y_batch = y_train[i:i+BATCH_SIZE].to(device)
# model.to(device)
# preds = model(X_batch.float())
# loss = criterion(preds,torch.tensor(y_batch,dtype=torch.long))
# optimizer.zero_grad()
# loss.backward()
# optimizer.step()
# wandb.log({'loss':loss.item(),'accuracy':test(model,X_train,y_train)*100,'val_accuracy':test(model,X_test,y_test)*100,'pred':torch.argmax(preds[index]),'real':torch.argmax(y_batch[index])})
# wandb.finish()
import matplotlib.pyplot as plt
import pandas as pd
df = pd.Series(loss_logs)
df.plot.line(figsize=(12,6))
test(model,X_test,y_test)
test(model,X_train,y_train)
preds
X_testing = X_train
y_testing = y_train
correct = 0
total = 0
model.eval()
with torch.no_grad():
for i in range(len(X_testing)):
real_class = torch.argmax(y_testing[i]).to(device)
net_out = model(X_testing[i].view(-1,1,112,112).to(device).float())
net_out = net_out[0]
predictied_class = torch.argmax(net_out)
# print(predictied_class)
if str(predictied_class) == str(real_class):
correct += 1
total += 1
print(round(correct/total,3))
# for real,pred in zip(y_batch,preds):
# print(real)
# print(torch.argmax(pred))
# print('\n')
```
## Modelling
```
# conv2d_output
# conv2d_1_ouput
# conv2d_2_ouput
# output_fc1
# output_fc2
# output_fc4
# max_pool2d_keranl
# max_pool2d
# num_of_linear
# activation
# best num of epochs
# best optimizer
# best loss
## best lr
class Test_Model(nn.Module):
def __init__(self,conv2d_output=128,conv2d_1_ouput=32,conv2d_2_ouput=64,output_fc1=512,output_fc2=256,output_fc4=128,output=36,activation=F.relu,max_pool2d_keranl=2):
super().__init__()
print(conv2d_output)
print(conv2d_1_ouput)
print(conv2d_2_ouput)
print(output_fc1)
print(output_fc2)
print(output_fc4)
print(activation)
self.conv2d_output = conv2d_output
self.pool = nn.MaxPool2d(max_pool2d_keranl)
self.conv1 = nn.Conv2d(1, conv2d_1_ouput, 5)
self.conv3 = nn.Conv2d(conv2d_1_ouput,conv2d_2_ouput,5)
self.conv2 = nn.Conv2d(conv2d_2_ouput, conv2d_output, 5)
self.fc1 = nn.Linear(conv2d_output * 10 * 10, output_fc1)
self.fc2 = nn.Linear(output_fc1, output_fc2)
self.fc4 = nn.Linear(output_fc2,output_fc4)
self.fc3 = nn.Linear(output_fc4, output)
self.activation = activation
def forward(self, x,shape=False):
x = self.pool(self.activation(self.conv1(x)))
x = self.pool(self.activation(self.conv3(x)))
x = self.pool(self.activation(self.conv2(x)))
if shape:
print(x.shape)
x = x.view(-1, self.conv2d_output * 10 * 10)
x = self.activation(self.fc1(x))
x = self.activation(self.fc2(x))
x = self.activation(self.fc4(x))
x = self.fc3(x)
return x
# conv2d_output
# conv2d_1_ouput
# conv2d_2_ouput
# output_fc1
# output_fc2
# output_fc4
# max_pool2d_keranl
# max_pool2d
# num_of_linear
# best num of epochs
# best loss
## best lr
# batch size
EPOCHS = 3
BATCH_SIZE = 32
# activation = nn.SELU()
# best optimizer = torch.optim.Adadelta
# best loss = nn.CrossEntropyLoss
## best lr = 0.625
# conv2d_output =
# conv2d_1_ouput =
# conv2d_2_ouput
# output_fc1
# output_fc2
# output_fc4
# max_pool2d_keranl
# max_pool2d
# num_of_linear
# best num of epochs
def get_loss(criterion,y,model,X):
preds = model(X.view(-1,1,112,112).to(device).float())
preds.to(device)
loss = criterion(preds,torch.tensor(y,dtype=torch.long).to(device))
loss.backward()
return loss.item()
conv2d_1_ouputs = [32,64,128,256,512]
for conv2d_1_ouput in conv2d_1_ouputs:
model = Test_Model(activation=nn.SELU(),conv2d_1_ouput=conv2d_1_ouput)
optimizer = torch.optim.Adadelta(model.parameters(),lr=0.625)
criterion = nn.CrossEntropyLoss()
wandb.init(project=PROJECT_NAME,name=f'conv2d_1_ouputs-{conv2d_1_ouput}')
for _ in tqdm(range(EPOCHS)):
for i in range(0,len(X_train),BATCH_SIZE):
X_batch = X_train[i:i+BATCH_SIZE].view(-1,1,112,112).to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model.to(device)
preds = model(X_batch.float())
loss = criterion(preds,torch.tensor(y_batch,dtype=torch.long))
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item(),'accuracy':test(model,X_train,y_train)*100,'val_accuracy':test(model,X_test,y_test)*100,'val_loss':get_loss(criterion,y_test,model,X_test)})
print(f'{torch.argmax(preds[1])} \n {y_batch[1]}')
print(f'{torch.argmax(preds[2])} \n {y_batch[2]}')
print(f'{torch.argmax(preds[3])} \n {y_batch[3]}')
print(f'{torch.argmax(preds[4])} \n {y_batch[4]}')
wandb.finish()
# conv2d_outputs = [32,64,128,256,512]
# for conv2d_output in conv2d_outputs:
# model = Test_Model(activation=nn.SELU(),conv2d_output=conv2d_output,conv2d_1_ouput='')
# optimizer = torch.optim.Adadelta(model.parameters(),lr=0.625)
# criterion = nn.CrossEntropyLoss()
# wandb.init(project=PROJECT_NAME,name=f'conv2d_output-{conv2d_output}')
# for _ in tqdm(range(EPOCHS)):
# for i in range(0,len(X_train),BATCH_SIZE):
# X_batch = X_train[i:i+BATCH_SIZE].view(-1,1,112,112).to(device)
# y_batch = y_train[i:i+BATCH_SIZE].to(device)
# model.to(device)
# preds = model(X_batch.float())
# loss = criterion(preds,torch.tensor(y_batch,dtype=torch.long))
# optimizer.zero_grad()
# loss.backward()
# optimizer.step()
# wandb.log({'loss':loss.item(),'accuracy':test(model,X_train,y_train)*100,'val_accuracy':test(model,X_test,y_test)*100,'val_loss':get_loss(criterion,y_test,model,X_test)})
# print(f'{torch.argmax(preds[1])} \n {y_batch[1]}')
# print(f'{torch.argmax(preds[2])} \n {y_batch[2]}')
# print(f'{torch.argmax(preds[3])} \n {y_batch[3]}')
# print(f'{torch.argmax(preds[4])} \n {y_batch[4]}')
# wandb.finish()
for real,pred in zip(y_batch,preds):
print(real)
print(torch.argmax(pred))
print('\n')
```
| github_jupyter |
1. [Value-based join of two Matrices](#JoinMatrices)
* [Filter Matrix to include only Frequent Column Values](#FilterMatrix)
* [Construct (sparse) Matrix from (rowIndex, colIndex, values) triplets](#Construct_sparse_Matrix)
* [Find and remove duplicates in columns or rows](#Find_and_remove_duplicates)
* [Set based Indexing](#Set_based_Indexing)
* [Group by Aggregate using Linear Algebra](#Multi_column_Sorting)
* [Cumulative Summation with Decay Multiplier](#CumSum_Product)
* [Invert Lower Triangular Matrix](#Invert_Lower_Triangular_Matrix)
```
from systemml import MLContext, dml, jvm_stdout
ml = MLContext(sc)
print (ml.buildTime())
```
## Value-based join of two Matrices<a id="JoinMatrices"/>
Given matrix M1 and M2, join M1 on column 2 with M2 on column 2, and return matching rows of M1.
```
prog = """
M1 = matrix ('1 1 2 3 3 3 4 4 5 3 6 4 7 1 8 2 9 1', rows = 9, cols = 2)
M2 = matrix ('1 1 2 8 3 3 4 3 5 1', rows = 5, cols = 2)
I = rowSums (outer (M1[,2], t(M2[,2]), "==")) # I : indicator matrix for M1
M12 = removeEmpty (target = M1, margin = "rows", select = I) # apply filter to retrieve join result
print ("M1 \n" + toString(M1))
print ("M2 \n" + toString(M2))
print ("M1[,2] joined with M2[,2], and return matching M1 rows\n" + toString(M12))
"""
with jvm_stdout():
ml.execute(dml(prog))
```
## Filter Matrix to include only Frequent Column Values <a id="FilterMatrix"/>
Given a matrix, filter the matrix to only include rows with column values that appear more often than MinFreq.
```
prog = """
MinFreq = 3 # minimum frequency of tokens
M = matrix ('1 1 2 3 3 3 4 4 5 3 6 4 7 1 8 2 9 1', rows = 9, cols = 2)
gM = aggregate (target = M[,2], groups = M[,2], fn = "count") # gM: group by and count (grouped matrix)
gv = cbind (seq(1,nrow(gM)), gM) # gv: add group values to counts (group values)
fg = removeEmpty (target = gv * (gv[,2] >= MinFreq), margin = "rows") # fg: filtered groups
I = rowSums (outer (M[,2] ,t(fg[,1]), "==")) # I : indicator of size M with filtered groups
fM = removeEmpty (target = M, margin = "rows", select = I) # FM: filter matrix
print (toString(M))
print (toString(fM))
"""
with jvm_stdout():
ml.execute(dml(prog))
```
## Construct (sparse) Matrix from (rowIndex, colIndex, values) triplets<a id="Construct_sparse_Matrix"></a>
Given rowIndex, colIndex, and values as column vectors, construct (sparse) matrix.
```
prog = """
I = matrix ("1 3 3 4 5", rows = 5, cols = 1)
J = matrix ("2 3 4 1 6", rows = 5, cols = 1)
V = matrix ("10 20 30 40 50", rows = 5, cols = 1)
M = table (I, J, V)
print (toString (M))
"""
ml.execute(dml(prog).output('M')).get('M').toNumPy()
```
## Find and remove duplicates in columns or rows<a id="Find_and_remove_duplicates"></a>
### Assuming values are sorted.
```
prog = """
X = matrix ("1 2 3 3 3 4 5 10", rows = 8, cols = 1)
I = rbind (matrix (1,1,1), (X[1:nrow (X)-1,] != X[2:nrow (X),])); # compare current with next value
res = removeEmpty (target = X, margin = "rows", select = I); # select where different
"""
ml.execute(dml(prog).output('res')).get('res').toNumPy()
```
### No assumptions on values.
```
prog = """
X = matrix ("3 2 1 3 3 4 5 10", rows = 8, cols = 1)
I = aggregate (target = X, groups = X[,1], fn = "count") # group and count duplicates
res = removeEmpty (target = seq (1, max (X[,1])), margin = "rows", select = (I != 0)); # select groups
"""
ml.execute(dml(prog).output('res')).get('res').toNumPy()
```
### Order the values and then remove duplicates.
```
prog = """
X = matrix ("3 2 1 3 3 4 5 10", rows = 8, cols = 1)
X = order (target = X, by = 1) # order values
I = rbind (matrix (1,1,1), (X[1:nrow (X)-1,] != X[2:nrow (X),]));
res = removeEmpty (target = X, margin = "rows", select = I);
"""
ml.execute(dml(prog).output('res')).get('res').toNumPy()
```
## Set based Indexing<a id="Set_based_Indexing"></a>
Given a matrix X, and a indicator matrix J with indices into X.
Use J to perform operation on X, e.g. add value 10 to cells in X indicated by J.
```
prog = """
X = matrix (1, rows = 1, cols = 100)
J = matrix ("10 20 25 26 28 31 50 67 79", rows = 1, cols = 9)
res = X + table (matrix (1, rows = 1, cols = ncol (J)), J, 10)
print (toString (res))
"""
ml.execute(dml(prog).output('res')).get('res').toNumPy()
```
## Group by Aggregate using Linear Algebra<a id="Multi_column_Sorting"></a>
Given a matrix PCV as (Position, Category, Value), sort PCV by category, and within each category by value in descending order. Create indicator vector for category changes, create distinct categories, and perform linear algebra operations.
```
prog = """
C = matrix ('50 40 20 10 30 20 40 20 30', rows = 9, cols = 1) # category data
V = matrix ('20 11 49 33 94 29 48 74 57', rows = 9, cols = 1) # value data
PCV = cbind (cbind (seq (1, nrow (C), 1), C), V); # PCV representation
PCV = order (target = PCV, by = 3, decreasing = TRUE, index.return = FALSE);
PCV = order (target = PCV, by = 2, decreasing = FALSE, index.return = FALSE);
# Find all rows of PCV where the category has a new value, in comparison to the previous row
is_new_C = matrix (1, rows = 1, cols = 1);
if (nrow (C) > 1) {
is_new_C = rbind (is_new_C, (PCV [1:nrow(C) - 1, 2] < PCV [2:nrow(C), 2]));
}
# Associate each category with its index
index_C = cumsum (is_new_C); # cumsum
# For each category, compute:
# - the list of distinct categories
# - the maximum value for each category
# - 0-1 aggregation matrix that adds records of the same category
distinct_C = removeEmpty (target = PCV [, 2], margin = "rows", select = is_new_C);
max_V_per_C = removeEmpty (target = PCV [, 3], margin = "rows", select = is_new_C);
C_indicator = table (index_C, PCV [, 1], max (index_C), nrow (C)); # table
sum_V_per_C = C_indicator %*% V
"""
res = ml.execute(dml(prog).output('PCV','distinct_C', 'max_V_per_C', 'C_indicator', 'sum_V_per_C'))
print (res.get('PCV').toNumPy())
print (res.get('distinct_C').toNumPy())
print (res.get('max_V_per_C').toNumPy())
print (res.get('C_indicator').toNumPy())
print (res.get('sum_V_per_C').toNumPy())
```
## Cumulative Summation with Decay Multiplier<a id="CumSum_Product"></a>
Given matrix X, compute:
Y[i] = X[i]
+ X[i-1] * C[i]
+ X[i-2] * C[i] * C[i-1]
+ X[i-3] * C[i] * C[i-1] * C[i-2]
+ ...
```
cumsum_prod_def = """
cumsum_prod = function (Matrix[double] X, Matrix[double] C, double start)
return (Matrix[double] Y)
# Computes the following recurrence in log-number of steps:
# Y [1, ] = X [1, ] + C [1, ] * start;
# Y [i+1, ] = X [i+1, ] + C [i+1, ] * Y [i, ]
{
Y = X; P = C; m = nrow(X); k = 1;
Y [1,] = Y [1,] + C [1,] * start;
while (k < m) {
Y [k + 1:m,] = Y [k + 1:m,] + Y [1:m - k,] * P [k + 1:m,];
P [k + 1:m,] = P [1:m - k,] * P [k + 1:m,];
k = 2 * k;
}
}
"""
```
In this example we use cumsum_prod for cumulative summation with "breaks", that is, multiple cumulative summations in one.
```
prog = cumsum_prod_def + """
X = matrix ("1 2 3 4 5 6 7 8 9", rows = 9, cols = 1);
#Zeros in C cause "breaks" that restart the cumulative summation from 0
C = matrix ("0 1 1 0 1 1 1 0 1", rows = 9, cols = 1);
Y = cumsum_prod (X, C, 0);
print (toString(Y))
"""
with jvm_stdout():
ml.execute(dml(prog))
```
In this example, we copy selected rows downward to all consecutive non-selected rows.
```
prog = cumsum_prod_def + """
X = matrix ("1 2 3 4 5 6 7 8 9", rows = 9, cols = 1);
# Ones in S represent selected rows to be copied, zeros represent non-selected rows
S = matrix ("1 0 0 1 0 0 0 1 0", rows = 9, cols = 1);
Y = cumsum_prod (X * S, 1 - S, 0);
print (toString(Y))
"""
with jvm_stdout():
ml.execute(dml(prog))
```
This is a naive implementation of cumulative summation with decay multiplier.
```
cumsum_prod_naive_def = """
cumsum_prod_naive = function (Matrix[double] X, Matrix[double] C, double start)
return (Matrix[double] Y)
{
Y = matrix (0, rows = nrow(X), cols = ncol(X));
Y [1,] = X [1,] + C [1,] * start;
for (i in 2:nrow(X))
{
Y [i,] = X [i,] + C [i,] * Y [i - 1,]
}
}
"""
```
There is a significant performance difference between the <b>naive</b> implementation and the <b>tricky</b> implementation.
```
prog = cumsum_prod_def + cumsum_prod_naive_def + """
X = rand (rows = 20000, cols = 10, min = 0, max = 1, pdf = "uniform", sparsity = 1.0);
C = rand (rows = 20000, cols = 10, min = 0, max = 1, pdf = "uniform", sparsity = 1.0);
Y1 = cumsum_prod_naive (X, C, 0.123);
"""
with jvm_stdout():
ml.execute(dml(prog))
prog = cumsum_prod_def + cumsum_prod_naive_def + """
X = rand (rows = 20000, cols = 10, min = 0, max = 1, pdf = "uniform", sparsity = 1.0);
C = rand (rows = 20000, cols = 10, min = 0, max = 1, pdf = "uniform", sparsity = 1.0);
Y2 = cumsum_prod (X, C, 0.123);
"""
with jvm_stdout():
ml.execute(dml(prog))
```
## Invert Lower Triangular Matrix<a id="Invert_Lower_Triangular_Matrix"></a>
In this example, we invert a lower triangular matrix using a the following divide-and-conquer approach. Given lower triangular matrix L, we compute its inverse X which is also lower triangular by splitting both matrices in the middle into 4 blocks (in a 2x2 fashion), and multiplying them together to get the identity matrix:
\begin{equation}
L \text{ %*% } X = \left(\begin{matrix} L_1 & 0 \\ L_2 & L_3 \end{matrix}\right)
\text{ %*% } \left(\begin{matrix} X_1 & 0 \\ X_2 & X_3 \end{matrix}\right)
= \left(\begin{matrix} L_1 X_1 & 0 \\ L_2 X_1 + L_3 X_2 & L_3 X_3 \end{matrix}\right)
= \left(\begin{matrix} I & 0 \\ 0 & I \end{matrix}\right)
\nonumber
\end{equation}
If we multiply blockwise, we get three equations:
$
\begin{equation}
L1 \text{ %*% } X1 = 1\\
L3 \text{ %*% } X3 = 1\\
L2 \text{ %*% } X1 + L3 \text{ %*% } X2 = 0\\
\end{equation}
$
Solving these equation gives the following formulas for X:
$
\begin{equation}
X1 = inv(L1) \\
X3 = inv(L3) \\
X2 = - X3 \text{ %*% } L2 \text{ %*% } X1 \\
\end{equation}
$
If we already recursively inverted L1 and L3, we can invert L2. This suggests an algorithm that starts at the diagonal and iterates away from the diagonal, involving bigger and bigger blocks (of size 1, 2, 4, 8, etc.) There is a logarithmic number of steps, and inside each step, the inversions can be performed in parallel using a parfor-loop.
Function "invert_lower_triangular" occurs within more general inverse operations and matrix decompositions. The divide-and-conquer idea allows to derive more efficient algorithms for other matrix decompositions.
```
invert_lower_triangular_def = """
invert_lower_triangular = function (Matrix[double] LI)
return (Matrix[double] LO)
{
n = nrow (LI);
LO = matrix (0, rows = n, cols = n);
LO = LO + diag (1 / diag (LI));
k = 1;
while (k < n)
{
LPF = matrix (0, rows = n, cols = n);
parfor (p in 0:((n - 1) / (2 * k)), check = 0)
{
i = 2 * k * p;
j = i + k;
q = min (n, j + k);
if (j + 1 <= q) {
L1 = LO [i + 1:j, i + 1:j];
L2 = LI [j + 1:q, i + 1:j];
L3 = LO [j + 1:q, j + 1:q];
LPF [j + 1:q, i + 1:j] = -L3 %*% L2 %*% L1;
}
}
LO = LO + LPF;
k = 2 * k;
}
}
"""
prog = invert_lower_triangular_def + """
n = 1000;
A = rand (rows = n, cols = n, min = -1, max = 1, pdf = "uniform", sparsity = 1.0);
Mask = cumsum (diag (matrix (1, rows = n, cols = 1)));
L = (A %*% t(A)) * Mask; # Generate L for stability of the inverse
X = invert_lower_triangular (L);
print ("Maximum difference between X %*% L and Identity = " + max (abs (X %*% L - diag (matrix (1, rows = n, cols = 1)))));
"""
with jvm_stdout():
ml.execute(dml(prog))
```
This is a naive implementation of inverting a lower triangular matrix.
```
invert_lower_triangular_naive_def = """
invert_lower_triangular_naive = function (Matrix[double] LI)
return (Matrix[double] LO)
{
n = nrow (LI);
LO = diag (matrix (1, rows = n, cols = 1));
for (i in 1:n - 1)
{
LO [i,] = LO [i,] / LI [i, i];
LO [i + 1:n,] = LO [i + 1:n,] - LI [i + 1:n, i] %*% LO [i,];
}
LO [n,] = LO [n,] / LI [n, n];
}
"""
```
The naive implementation is significantly slower than the divide-and-conquer implementation.
```
prog = invert_lower_triangular_naive_def + """
n = 1000;
A = rand (rows = n, cols = n, min = -1, max = 1, pdf = "uniform", sparsity = 1.0);
Mask = cumsum (diag (matrix (1, rows = n, cols = 1)));
L = (A %*% t(A)) * Mask; # Generate L for stability of the inverse
X = invert_lower_triangular_naive (L);
print ("Maximum difference between X %*% L and Identity = " + max (abs (X %*% L - diag (matrix (1, rows = n, cols = 1)))));
"""
with jvm_stdout():
ml.execute(dml(prog))
```
| github_jupyter |
```
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# AI Platform (Unified) AutoML text entity extractionn model
## Installation
Install the latest (preview) version of AI Platform (Unified) SDK.
```
! pip3 install -U google-cloud-aiplatform --user
```
Install the Google *cloud-storage* library as well.
```
! pip3 install google-cloud-storage
```
### Restart the Kernel
Once you've installed the AI Platform (Unified) SDK and Google *cloud-storage*, you need to restart the notebook kernel so it can find the packages.
```
import os
if not os.getenv("AUTORUN"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
```
## Before you begin
### GPU run-time
*Make sure you're running this notebook in a GPU runtime if you have that option. In Colab, select* **Runtime > Change Runtime Type > GPU**
### Set up your GCP project
**The following steps are required, regardless of your notebook environment.**
1. [Select or create a GCP project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
3. [Enable the AI Platform APIs and Compute Engine APIs.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component)
4. [Google Cloud SDK](https://cloud.google.com/sdk) is already installed in AI Platform Notebooks.
5. Enter your project ID in the cell below. Then run the cell to make sure the
Cloud SDK uses the right project for all the commands in this notebook.
**Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
#### Project ID
**If you don't know your project ID**, try to get your project ID using `gcloud` command by executing the second cell below.
```
PROJECT_ID = "[your-project-id]" #@param {type:"string"}
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
! gcloud config set project $PROJECT_ID
```
#### Region
You can also change the `REGION` variable, which is used for operations
throughout the rest of this notebook. Below are regions supported for AI Platform (Unified). We recommend when possible, to choose the region closest to you.
- Americas: `us-central1`
- Europe: `europe-west4`
- Asia Pacific: `asia-east1`
You cannot use a Multi-Regional Storage bucket for training with AI Platform. Not all regions provide support for all AI Platform services. For the latest support per region, see [Region support for AI Platform (Unified) services](https://cloud.google.com/ai-platform-unified/docs/general/locations)
```
REGION = 'us-central1' #@param {type: "string"}
```
#### Timestamp
If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append onto the name of resources which will be created in this tutorial.
```
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
```
### Authenticate your GCP account
**If you are using AI Platform Notebooks**, your environment is already
authenticated. Skip this step.
*Note: If you are on an AI Platform notebook and run the cell, the cell knows to skip executing the authentication steps.*
```
import os
import sys
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your Google Cloud account. This provides access
# to your Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# If on AI Platform, then don't execute this code
if not os.path.exists('/opt/deeplearning/metadata/env_version'):
if 'google.colab' in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this tutorial in a notebook locally, replace the string
# below with the path to your service account key and run this cell to
# authenticate your Google Cloud account.
else:
%env GOOGLE_APPLICATION_CREDENTIALS your_path_to_credentials.json
# Log in to your account on Google Cloud
! gcloud auth login
```
### Create a Cloud Storage bucket
**The following steps are required, regardless of your notebook environment.**
This tutorial is designed to use training data that is in a public Cloud Storage bucket and a local Cloud Storage bucket for your batch predictions. You may alternatively use your own training data that you have stored in a local Cloud Storage bucket.
Set the name of your Cloud Storage bucket below. It must be unique across all Cloud Storage buckets.
```
BUCKET_NAME = "[your-bucket-name]" #@param {type:"string"}
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "[your-bucket-name]":
BUCKET_NAME = PROJECT_ID + "aip-" + TIMESTAMP
```
**Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
```
! gsutil mb -l $REGION gs://$BUCKET_NAME
```
Finally, validate access to your Cloud Storage bucket by examining its contents:
```
! gsutil ls -al gs://$BUCKET_NAME
```
### Set up variables
Next, set up some variables used throughout the tutorial.
### Import libraries and define constants
#### Import AI Platform (Unified) SDK
Import the AI Platform (Unified) SDK into our Python environment.
```
import json
import os
import sys
import time
from google.cloud.aiplatform import gapic as aip
from google.protobuf import json_format
from google.protobuf.json_format import MessageToJson
from google.protobuf.struct_pb2 import Value
from google.protobuf.struct_pb2 import Struct
```
#### AI Platform (Unified) constants
Setup up the following constants for AI Platform (Unified):
- `API_ENDPOINT`: The AI Platform (Unified) API service endpoint for dataset, model, job, pipeline and endpoint services.
- `PARENT`: The AI Platform (Unified) location root path for dataset, model and endpoint resources.
```
# API Endpoint
API_ENDPOINT = "us-central1-aiplatform.googleapis.com"
# AI Platform (Unified) location root path for your dataset, model and endpoint resources
PARENT = "projects/" + PROJECT_ID + "/locations/" + REGION
```
#### AutoML constants
Next, setup constants unique to AutoML Text Entity Extraction datasets and training:
- Dataset Schemas: Tells the managed dataset service which type of dataset it is.
- Data Labeling (Annotations) Schemas: Tells the managed dataset service how the data is labeled (annotated).
- Dataset Training Schemas: Tells the managed pipelines service the task (e.g., classification) to train the model for.
```
# Text Dataset type
TEXT_SCHEMA = 'google-cloud-aiplatform/schema/dataset/metadata/text_1.0.0.yaml'
# Text Labeling type
IMPORT_SCHEMA_TEXT_EXTRACTION = "gs://google-cloud-aiplatform/schema/dataset/ioformat/text_extraction_io_format_1.0.0.yaml"
# Text Training task
TRAINING_TEXT_EXTRACTION_SCHEMA = "gs://google-cloud-aiplatform/schema/trainingjob/definition/automl_text_extraction_1.0.0.yaml"
```
## Clients
The AI Platform (Unified) SDK works as a client/server model. On your side (the Python script) you will create a client that sends requests and receives responses from the server (AI Platform).
You will use several clients in this tutorial, so set them all up upfront.
- Dataset Service for managed datasets.
- Model Service for managed models.
- Pipeline Service for training.
- Endpoint Service for deployment.
- Job Service for batch jobs and custom training.
- Prediction Service for serving. *Note*: Prediction has a different service endpoint.
```
# client options same for all services
client_options = {"api_endpoint": API_ENDPOINT}
def create_dataset_client():
client = aip.DatasetServiceClient(
client_options=client_options
)
return client
def create_model_client():
client = aip.ModelServiceClient(
client_options=client_options
)
return client
def create_pipeline_client():
client = aip.PipelineServiceClient(
client_options=client_options
)
return client
def create_endpoint_client():
client = aip.EndpointServiceClient(
client_options=client_options
)
return client
def create_prediction_client():
client = aip.PredictionServiceClient(
client_options=client_options
)
return client
def create_job_client():
client = aip.JobServiceClient(
client_options=client_options
)
return client
clients = {}
clients['dataset'] = create_dataset_client()
clients['model'] = create_model_client()
clients['pipeline'] = create_pipeline_client()
clients['endpoint'] = create_endpoint_client()
clients['prediction'] = create_prediction_client()
clients['job'] = create_job_client()
for client in clients.items():
print(client)
IMPORT_FILE = 'gs://cloud-samples-data/language/ucaip_ten_dataset.jsonl'
! gsutil cat $IMPORT_FILE | head -n 1
```
*Example output*:
```
{'text_segment_annotations': [{'endOffset': 54, 'startOffset': 27, 'displayName': 'SpecificDisease'}, {'endOffset': 173, 'startOffset': 156, 'displayName': 'SpecificDisease'}, {'endOffset': 179, 'startOffset': 176, 'displayName': 'SpecificDisease'}, {'endOffset': 246, 'startOffset': 243, 'displayName': 'Modifier'}, {'endOffset': 340, 'startOffset': 337, 'displayName': 'Modifier'}, {'endOffset': 698, 'startOffset': 695, 'displayName': 'Modifier'}], 'textContent': '1301937\tMolecular basis of hexosaminidase A deficiency and pseudodeficiency in the Berks County Pennsylvania Dutch.\tFollowing the birth of two infants with Tay-Sachs disease ( TSD ) , a non-Jewish , Pennsylvania Dutch kindred was screened for TSD carriers using the biochemical assay . A high frequency of individuals who appeared to be TSD heterozygotes was detected ( Kelly et al . , 1975 ) . Clinical and biochemical evidence suggested that the increased carrier frequency was due to at least two altered alleles for the hexosaminidase A alpha-subunit . We now report two mutant alleles in this Pennsylvania Dutch kindred , and one polymorphism . One allele , reported originally in a French TSD patient ( Akli et al . , 1991 ) , is a GT-- > AT transition at the donor splice-site of intron 9 . The second , a C-- > T transition at nucleotide 739 ( Arg247Trp ) , has been shown by Triggs-Raine et al . ( 1992 ) to be a clinically benign " pseudodeficient " allele associated with reduced enzyme activity against artificial substrate . Finally , a polymorphism [ G-- > A ( 759 ) ] , which leaves valine at codon 253 unchanged , is described .\n '}
```
## Create a dataset
### [projects.locations.datasets.create](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.datasets/create)
#### Request
```
DATA_SCHEMA = TEXT_SCHEMA
dataset = {
"display_name": "ten_" + TIMESTAMP,
"metadata_schema_uri": "gs://" + DATA_SCHEMA
}
print(MessageToJson(
aip.CreateDatasetRequest(
parent=PARENT,
dataset=dataset
).__dict__["_pb"])
)
```
*Example output*:
```
{
"parent": "projects/migration-ucaip-training/locations/us-central1",
"dataset": {
"displayName": "ten_20210301154552",
"metadataSchemaUri": "gs://google-cloud-aiplatform/schema/dataset/metadata/text_1.0.0.yaml"
}
}
```
#### Call
```
request = clients['dataset'].create_dataset(
parent=PARENT,
dataset=dataset
)
```
#### Response
```
result = request.result()
print(MessageToJson(result.__dict__["_pb"]))
```
*Example output*:
```
{
"name": "projects/116273516712/locations/us-central1/datasets/1309228077611483136",
"displayName": "ten_20210301154552",
"metadataSchemaUri": "gs://google-cloud-aiplatform/schema/dataset/metadata/text_1.0.0.yaml",
"labels": {
"aiplatform.googleapis.com/dataset_metadata_schema": "TEXT"
},
"metadata": {
"dataItemSchemaUri": "gs://google-cloud-aiplatform/schema/dataset/dataitem/text_1.0.0.yaml"
}
}
```
```
# The full unique ID for the dataset
dataset_id = result.name
# The short numeric ID for the dataset
dataset_short_id = dataset_id.split('/')[-1]
print(dataset_id)
```
### [projects.locations.datasets.import](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.datasets/import)
#### Request
```
LABEL_SCHEMA = IMPORT_SCHEMA_TEXT_EXTRACTION
import_config = {
"gcs_source": {
"uris": [IMPORT_FILE]
},
"import_schema_uri": LABEL_SCHEMA
}
print(MessageToJson(
aip.ImportDataRequest(
name=dataset_id,
import_configs=[import_config]
).__dict__["_pb"])
)
```
*Example output*:
```
{
"name": "projects/116273516712/locations/us-central1/datasets/1309228077611483136",
"importConfigs": [
{
"gcsSource": {
"uris": [
"gs://cloud-samples-data/language/ucaip_ten_dataset.jsonl"
]
},
"importSchemaUri": "gs://google-cloud-aiplatform/schema/dataset/ioformat/text_extraction_io_format_1.0.0.yaml"
}
]
}
```
#### Call
```
request = clients['dataset'].import_data(
name=dataset_id,
import_configs=[import_config]
)
```
#### Response
```
result = request.result()
print(MessageToJson(result.__dict__["_pb"]))
```
*Example output*:
```
{}
```
## Train a model
### [projects.locations.trainingPipelines.create](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.trainingPipelines/create)
#### Request
```
TRAINING_SCHEMA = TRAINING_TEXT_EXTRACTION_SCHEMA
task = Value(struct_value=Struct(
fields={
'multi_label': Value(bool_value=False),
'budget_milli_node_hours': Value(number_value=1000),
'model_type': Value(string_value="CLOUD"),
'disable_early_stopping': Value(bool_value=False)
}
))
training_pipeline = {
"display_name": "ten_" + TIMESTAMP,
"input_data_config": {
"dataset_id": dataset_short_id
},
"model_to_upload": {
"display_name": "ten_" + TIMESTAMP
},
"training_task_definition": TRAINING_SCHEMA,
"training_task_inputs": task
}
print(MessageToJson(
aip.CreateTrainingPipelineRequest(
parent=PARENT,
training_pipeline=training_pipeline,
).__dict__["_pb"])
)
```
*Example output*:
```
{
"parent": "projects/migration-ucaip-training/locations/us-central1",
"trainingPipeline": {
"displayName": "ten_20210301154552",
"inputDataConfig": {
"datasetId": "1309228077611483136"
},
"trainingTaskDefinition": "gs://google-cloud-aiplatform/schema/trainingjob/definition/automl_text_extraction_1.0.0.yaml",
"trainingTaskInputs": {
"budget_milli_node_hours": 1000.0,
"multi_label": false,
"model_type": "CLOUD",
"disable_early_stopping": false
},
"modelToUpload": {
"displayName": "ten_20210301154552"
}
}
}
```
#### Call
```
request = clients['pipeline'].create_training_pipeline(
parent=PARENT,
training_pipeline=training_pipeline
)
```
#### Response
```
print(MessageToJson(request.__dict__["_pb"]))
```
*Example output*:
```
{
"name": "projects/116273516712/locations/us-central1/trainingPipelines/4643220011912003584",
"displayName": "ten_20210301154552",
"inputDataConfig": {
"datasetId": "1309228077611483136"
},
"trainingTaskDefinition": "gs://google-cloud-aiplatform/schema/trainingjob/definition/automl_text_extraction_1.0.0.yaml",
"trainingTaskInputs": {},
"modelToUpload": {
"displayName": "ten_20210301154552"
},
"state": "PIPELINE_STATE_PENDING",
"createTime": "2021-03-01T15:55:29.352065Z",
"updateTime": "2021-03-01T15:55:29.352065Z"
}
```
```
# The full unique ID for the training pipeline
training_pipeline_id = request.name
# The short numeric ID for the training pipeline
training_pipeline_short_id = training_pipeline_id.split('/')[-1]
print(training_pipeline_id)
```
### [projects.locations.trainingPipelines.get](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.trainingPipelines/get)
#### Call
```
request = clients['pipeline'].get_training_pipeline(
name=training_pipeline_id
)
```
#### Response
```
print(MessageToJson(request.__dict__["_pb"]))
```
*Example output*:
```
{
"name": "projects/116273516712/locations/us-central1/trainingPipelines/4643220011912003584",
"displayName": "ten_20210301154552",
"inputDataConfig": {
"datasetId": "1309228077611483136"
},
"trainingTaskDefinition": "gs://google-cloud-aiplatform/schema/trainingjob/definition/automl_text_extraction_1.0.0.yaml",
"trainingTaskInputs": {},
"modelToUpload": {
"displayName": "ten_20210301154552"
},
"state": "PIPELINE_STATE_PENDING",
"createTime": "2021-03-01T15:55:29.352065Z",
"updateTime": "2021-03-01T15:55:29.352065Z"
}
```
```
while True:
response = clients["pipeline"].get_training_pipeline(name=training_pipeline_id)
if response.state != aip.PipelineState.PIPELINE_STATE_SUCCEEDED:
print("Training job has not completed:", response.state)
if response.state == aip.PipelineState.PIPELINE_STATE_FAILED:
break
else:
model_id = response.model_to_upload.name
print("Training Time:", response.end_time - response.start_time)
break
time.sleep(60)
print(model_id)
```
## Evaluate the model
### [projects.locations.models.evaluations.list](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.models.evaluations/list)
#### Call
```
request = clients["model"].list_model_evaluations(
parent=model_id
)
```
#### Response
```
model_evaluations = [
json.loads(MessageToJson(mel.__dict__["_pb"])) for mel in request
]
# The evaluation slice
evaluation_slice = request.model_evaluations[0].name
print(json.dumps(model_evaluations, indent=2))
```
*Example output*:
```
[
{
"name": "projects/116273516712/locations/us-central1/models/4400738115568795648/evaluations/7959028222912364544",
"metricsSchemaUri": "gs://google-cloud-aiplatform/schema/modelevaluation/text_extraction_metrics_1.0.0.yaml",
"metrics": {
"confusionMatrix": {
"rows": [
[
0.0,
24.0,
23.0,
1.0,
27.0
],
[
9.0,
40.0,
0.0,
0.0,
10.0
],
[
11.0,
0.0,
87.0,
0.0,
2.0
],
[
3.0,
0.0,
0.0,
5.0,
0.0
],
[
32.0,
16.0,
7.0,
1.0,
186.0
]
],
"annotationSpecs": [
{
"displayName": "NULL"
},
{
"id": "2041829376663748608",
"displayName": "DiseaseClass"
},
{
"displayName": "Modifier",
"id": "4347672385877442560"
},
{
"displayName": "CompositeMention",
"id": "6653515395091136512"
},
{
"id": "7806436899697983488",
"displayName": "SpecificDisease"
}
]
},
"confidenceMetrics": [
{
"precision": 0.74125874,
"f1Score": 0.7589499,
"recall": 0.7775061,
"confidenceThreshold": 0.04
},
{
"recall": 0.7457213,
"confidenceThreshold": 0.96,
"precision": 0.8333333,
"f1Score": 0.7870968
},
# REMOVED FOR BREVITY
{
"f1Score": 0.7596154,
"recall": 0.77261615,
"confidenceThreshold": 0.44,
"precision": 0.7470449
}
]
},
"createTime": "2021-03-01T17:59:23.638307Z",
"sliceDimensions": [
"annotationSpec"
]
}
]
```
### [projects.locations.models.evaluations.get](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.models.evaluations/get)
#### Call
```
request = clients["model"].get_model_evaluation(
name=evaluation_slice
)
```
#### Response
```
print(MessageToJson(request.__dict__["_pb"]))
```
*Example output*:
```
{
"name": "projects/116273516712/locations/us-central1/models/4400738115568795648/evaluations/7959028222912364544",
"metricsSchemaUri": "gs://google-cloud-aiplatform/schema/modelevaluation/text_extraction_metrics_1.0.0.yaml",
"metrics": {
"confusionMatrix": {
"rows": [
[
0.0,
24.0,
23.0,
1.0,
27.0
],
[
9.0,
40.0,
0.0,
0.0,
10.0
],
[
11.0,
0.0,
87.0,
0.0,
2.0
],
[
3.0,
0.0,
0.0,
5.0,
0.0
],
[
32.0,
16.0,
7.0,
1.0,
186.0
]
],
"annotationSpecs": [
{
"displayName": "NULL"
},
{
"id": "2041829376663748608",
"displayName": "DiseaseClass"
},
{
"displayName": "Modifier",
"id": "4347672385877442560"
},
{
"id": "6653515395091136512",
"displayName": "CompositeMention"
},
{
"displayName": "SpecificDisease",
"id": "7806436899697983488"
}
]
},
"confidenceMetrics": [
{
"precision": 0.74125874,
"recall": 0.7775061,
"confidenceThreshold": 0.04,
"f1Score": 0.7589499
},
{
"f1Score": 0.7870968,
"recall": 0.7457213,
"confidenceThreshold": 0.96,
"precision": 0.8333333
},
# REMOVED FOR BREVITY
{
"precision": 0.745283,
"f1Score": 0.7587035,
"recall": 0.77261615,
"confidenceThreshold": 0.43
},
{
"precision": 0.7470449,
"recall": 0.77261615,
"confidenceThreshold": 0.44,
"f1Score": 0.7596154
}
]
},
"createTime": "2021-03-01T17:59:23.638307Z",
"sliceDimensions": [
"annotationSpec"
]
}
```
## Make batch predictions
### Make a batch prediction file
```
import tensorflow as tf
import json
test_item = 'Molecular basis of hexosaminidase A deficiency and pseudodeficiency in the Berks County Pennsylvania Dutch.\\tFollowing the birth of two infants with Tay-Sachs disease ( TSD ) , a non-Jewish , Pennsylvania Dutch kindred was screened for TSD carriers using the biochemical assay . A high frequency of individuals who appeared to be TSD heterozygotes was detected ( Kelly et al . , 1975 ) . Clinical and biochemical evidence suggested that the increased carrier frequency was due to at least two altered alleles for the hexosaminidase A alpha-subunit . We now report two mutant alleles in this Pennsylvania Dutch kindred , and one polymorphism . One allele , reported originally in a French TSD patient ( Akli et al . , 1991 ) , is a GT-- > AT transition at the donor splice-site of intron 9 . The second , a C-- > T transition at nucleotide 739 ( Arg247Trp ) , has been shown by Triggs-Raine et al . ( 1992 ) to be a clinically benign " pseudodeficient " allele associated with reduced enzyme activity against artificial substrate . Finally , a polymorphism [ G-- > A ( 759 ) ] , which leaves valine at codon 253 unchanged , is described'
gcs_test_item = "gs://" + BUCKET_NAME + "/test.txt"
with tf.io.gfile.GFile(gcs_test_item, 'w') as f:
f.write(test_item + '\n')
gcs_input_uri = "gs://" + BUCKET_NAME + "/test.jsonl"
with tf.io.gfile.GFile(gcs_input_uri, 'w') as f:
f.write(json.dumps({"content": gcs_test_item, "mime_type": "text/plain"}) + '\n')
! gsutil cat $gcs_input_uri
! gsutil cat $gcs_test_item
```
*Example output*:
```
{"content": "gs://migration-ucaip-trainingaip-20210301154552/test.txt", "mime_type": "text/plain"}
Molecular basis of hexosaminidase A deficiency and pseudodeficiency in the Berks County Pennsylvania Dutch.\tFollowing the birth of two infants with Tay-Sachs disease ( TSD ) , a non-Jewish , Pennsylvania Dutch kindred was screened for TSD carriers using the biochemical assay . A high frequency of individuals who appeared to be TSD heterozygotes was detected ( Kelly et al . , 1975 ) . Clinical and biochemical evidence suggested that the increased carrier frequency was due to at least two altered alleles for the hexosaminidase A alpha-subunit . We now report two mutant alleles in this Pennsylvania Dutch kindred , and one polymorphism . One allele , reported originally in a French TSD patient ( Akli et al . , 1991 ) , is a GT-- > AT transition at the donor splice-site of intron 9 . The second , a C-- > T transition at nucleotide 739 ( Arg247Trp ) , has been shown by Triggs-Raine et al . ( 1992 ) to be a clinically benign " pseudodeficient " allele associated with reduced enzyme activity against artificial substrate . Finally , a polymorphism [ G-- > A ( 759 ) ] , which leaves valine at codon 253 unchanged , is described
```
### [projects.locations.batchPredictionJobs.create](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.batchPredictionJobs/create)
#### Request
```
batch_prediction_job = {
"display_name": "ten_" + TIMESTAMP,
"model": model_id,
"input_config": {
"instances_format": "jsonl",
"gcs_source": {
"uris": [gcs_input_uri]
}
},
"output_config": {
"predictions_format": "jsonl",
"gcs_destination": {
"output_uri_prefix": "gs://" + f"{BUCKET_NAME}/batch_output/"
}
},
"dedicated_resources": {
"machine_spec": {
"machine_type": "n1-standard-2",
"accelerator_count": 0
},
"starting_replica_count": 1,
"max_replica_count": 1
}
}
print(MessageToJson(
aip.CreateBatchPredictionJobRequest(
parent=PARENT,
batch_prediction_job=batch_prediction_job
).__dict__["_pb"])
)
```
*Example output*:
```
{
"parent": "projects/migration-ucaip-training/locations/us-central1",
"batchPredictionJob": {
"displayName": "ten_20210301154552",
"model": "projects/116273516712/locations/us-central1/models/4400738115568795648",
"inputConfig": {
"instancesFormat": "jsonl",
"gcsSource": {
"uris": [
"gs://migration-ucaip-trainingaip-20210301154552/test.jsonl"
]
}
},
"outputConfig": {
"predictionsFormat": "jsonl",
"gcsDestination": {
"outputUriPrefix": "gs://migration-ucaip-trainingaip-20210301154552/batch_output/"
}
},
"dedicatedResources": {
"machineSpec": {
"machineType": "n1-standard-2"
},
"startingReplicaCount": 1,
"maxReplicaCount": 1
}
}
}
```
#### Call
```
request = clients['job'].create_batch_prediction_job(
parent=PARENT,
batch_prediction_job=batch_prediction_job
)
```
#### Response
```
print(MessageToJson(request.__dict__["_pb"]))
```
*Example output*:
```
{
"name": "projects/116273516712/locations/us-central1/batchPredictionJobs/3588251799200464896",
"displayName": "ten_20210301154552",
"model": "projects/116273516712/locations/us-central1/models/4400738115568795648",
"inputConfig": {
"instancesFormat": "jsonl",
"gcsSource": {
"uris": [
"gs://migration-ucaip-trainingaip-20210301154552/test.jsonl"
]
}
},
"outputConfig": {
"predictionsFormat": "jsonl",
"gcsDestination": {
"outputUriPrefix": "gs://migration-ucaip-trainingaip-20210301154552/batch_output/"
}
},
"state": "JOB_STATE_PENDING",
"completionStats": {
"incompleteCount": "-1"
},
"createTime": "2021-03-01T17:59:42.777083Z",
"updateTime": "2021-03-01T17:59:42.777083Z"
}
```
```
# The fully qualified ID for the batch job
batch_job_id = request.name
# The short numeric ID for the batch job
batch_job_short_id = batch_job_id.split('/')[-1]
print(batch_job_id)
```
### [projects.locations.batchPredictionJobs.get](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.batchPredictionJobs/get)
#### Call
```
request = clients['job'].get_batch_prediction_job(
name=batch_job_id
)
```
#### Response
```
print(MessageToJson(request.__dict__["_pb"]))
```
*Example output*:
```
{
"name": "projects/116273516712/locations/us-central1/batchPredictionJobs/3588251799200464896",
"displayName": "ten_20210301154552",
"model": "projects/116273516712/locations/us-central1/models/4400738115568795648",
"inputConfig": {
"instancesFormat": "jsonl",
"gcsSource": {
"uris": [
"gs://migration-ucaip-trainingaip-20210301154552/test.jsonl"
]
}
},
"outputConfig": {
"predictionsFormat": "jsonl",
"gcsDestination": {
"outputUriPrefix": "gs://migration-ucaip-trainingaip-20210301154552/batch_output/"
}
},
"state": "JOB_STATE_PENDING",
"completionStats": {
"incompleteCount": "-1"
},
"createTime": "2021-03-01T17:59:42.777083Z",
"updateTime": "2021-03-01T17:59:42.777083Z"
}
```
```
while True:
response = clients["job"].get_batch_prediction_job(name=batch_job_id)
if response.state != aip.JobState.JOB_STATE_SUCCEEDED:
print("The job has not completed:", response.state)
if response.state == aip.JobState.JOB_STATE_FAILED:
break
else:
folder = response.output_config.gcs_destination.output_uri_prefix[:-1]
! gsutil ls $folder/prediction*/*.jsonl
! gsutil cat $folder/prediction*/*.jsonl
break
time.sleep(60)
```
*Example output*:
```
gs://migration-ucaip-trainingaip-20210301154552/batch_output/prediction-ten_20210301154552-2021-03-01T17:59:42.638222Z/predictions_00001.jsonl
{"instance":{"content":"gs://migration-ucaip-trainingaip-20210301154552/test.txt","mimeType":"text/plain"},"prediction":{"ids":["7806436899697983488","7806436899697983488","7806436899697983488","4347672385877442560","4347672385877442560","4347672385877442560"],"displayNames":["SpecificDisease","SpecificDisease","SpecificDisease","Modifier","Modifier","Modifier"],"textSegmentStartOffsets":["149","19","169","236","688","330"],"textSegmentEndOffsets":["165","45","171","238","690","332"],"confidences":[0.99957836,0.9995628,0.9995044,0.9993287,0.9993144,0.99927235]}}
```
## Make online predictions
### [projects.locations.endpoints.create](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.endpoints/create)
#### Request
```
endpoint = {
"display_name": "ten_" + TIMESTAMP
}
print(MessageToJson(
aip.CreateEndpointRequest(
parent=PARENT,
endpoint=endpoint
).__dict__["_pb"])
)
```
*Example output*:
```
{
"parent": "projects/migration-ucaip-training/locations/us-central1",
"endpoint": {
"displayName": "ten_20210301154552"
}
}
```
#### Call
```
request = clients['endpoint'].create_endpoint(
parent=PARENT,
endpoint=endpoint
)
```
#### Response
```
result =request.result()
print(MessageToJson(result.__dict__["_pb"]))
```
*Example output*:
```
{
"name": "projects/116273516712/locations/us-central1/endpoints/8916247652891361280"
}
```
```
# The fully qualified ID for the endpoint
endpoint_id = result.name
# The short numeric ID for the endpoint
endpoint_short_id = endpoint_id.split('/')[-1]
print(endpoint_id)
```
### [projects.locations.endpoints.deployModel](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.endpoints/deployModel)
#### Request
```
deployed_model = {
"model": model_id,
"display_name": "ten_" + TIMESTAMP,
"automatic_resources": {
"min_replica_count": 1,
"max_replica_count": 1
}
}
traffic_split={"0": 100}
print(MessageToJson(
aip.DeployModelRequest(
endpoint=endpoint_id,
deployed_model=deployed_model,
traffic_split=traffic_split
).__dict__["_pb"])
)
```
*Example output*:
```
{
"endpoint": "projects/116273516712/locations/us-central1/endpoints/8916247652891361280",
"deployedModel": {
"model": "projects/116273516712/locations/us-central1/models/4400738115568795648",
"displayName": "ten_20210301154552",
"automaticResources": {
"minReplicaCount": 1,
"maxReplicaCount": 1
}
},
"trafficSplit": {
"0": 100
}
}
```
#### Call
```
request = clients['endpoint'].deploy_model(
endpoint=endpoint_id,
deployed_model=deployed_model,
traffic_split=traffic_split
)
```
#### Response
```
result = request.result()
print(MessageToJson(result.__dict__["_pb"]))
```
*Example output*:
```
{
"deployedModel": {
"id": "3958065938133155840"
}
}
```
```
# The unique ID for the deployed model
deployed_model_id = result.deployed_model.id
print(deployed_model_id)
```
### [projects.locations.endpoints.predict](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.endpoints/predict)
#### Prepare data item for online prediction
```
test_item = 'Molecular basis of hexosaminidase A deficiency and pseudodeficiency in the Berks County Pennsylvania Dutch.\\tFollowing the birth of two infants with Tay-Sachs disease ( TSD ) , a non-Jewish , Pennsylvania Dutch kindred was screened for TSD carriers using the biochemical assay . A high frequency of individuals who appeared to be TSD heterozygotes was detected ( Kelly et al . , 1975 ) . Clinical and biochemical evidence suggested that the increased carrier frequency was due to at least two altered alleles for the hexosaminidase A alpha-subunit . We now report two mutant alleles in this Pennsylvania Dutch kindred , and one polymorphism . One allele , reported originally in a French TSD patient ( Akli et al . , 1991 ) , is a GT-- > AT transition at the donor splice-site of intron 9 . The second , a C-- > T transition at nucleotide 739 ( Arg247Trp ) , has been shown by Triggs-Raine et al . ( 1992 ) to be a clinically benign " pseudodeficient " allele associated with reduced enzyme activity against artificial substrate . Finally , a polymorphism [ G-- > A ( 759 ) ] , which leaves valine at codon 253 unchanged , is described'
```
#### Request
```
instances_list = [{"content": test_item}]
instances = [
json_format.ParseDict(s, Value()) for s in instances_list
]
prediction_request = aip.PredictRequest(
endpoint=endpoint_id,
)
prediction_request.instances.append(instances)
print(MessageToJson(prediction_request.__dict__["_pb"]))
```
*Example output*:
```
{
"endpoint": "projects/116273516712/locations/us-central1/endpoints/8916247652891361280",
"instances": [
[
{
"content": "Molecular basis of hexosaminidase A deficiency and pseudodeficiency in the Berks County Pennsylvania Dutch.\\tFollowing the birth of two infants with Tay-Sachs disease ( TSD ) , a non-Jewish , Pennsylvania Dutch kindred was screened for TSD carriers using the biochemical assay . A high frequency of individuals who appeared to be TSD heterozygotes was detected ( Kelly et al . , 1975 ) . Clinical and biochemical evidence suggested that the increased carrier frequency was due to at least two altered alleles for the hexosaminidase A alpha-subunit . We now report two mutant alleles in this Pennsylvania Dutch kindred , and one polymorphism . One allele , reported originally in a French TSD patient ( Akli et al . , 1991 ) , is a GT-- > AT transition at the donor splice-site of intron 9 . The second , a C-- > T transition at nucleotide 739 ( Arg247Trp ) , has been shown by Triggs-Raine et al . ( 1992 ) to be a clinically benign \" pseudodeficient \" allele associated with reduced enzyme activity against artificial substrate . Finally , a polymorphism [ G-- > A ( 759 ) ] , which leaves valine at codon 253 unchanged , is described"
}
]
]
}
```
#### Call
```
request = clients['prediction'].predict(
endpoint=endpoint_id,
instances=instances
)
```
#### Response
```
print(MessageToJson(request.__dict__["_pb"]))
```
*Example output*:
```
{
"predictions": [
{
"displayNames": [
"SpecificDisease",
"SpecificDisease",
"SpecificDisease",
"Modifier",
"Modifier",
"Modifier"
],
"confidences": [
0.9995627999305725,
0.9995783567428589,
0.9995043873786926,
0.9993286728858948,
0.999272346496582,
0.9993144273757935
],
"textSegmentStartOffsets": [
19.0,
149.0,
169.0,
236.0,
330.0,
688.0
],
"ids": [
"7806436899697983488",
"7806436899697983488",
"7806436899697983488",
"4347672385877442560",
"4347672385877442560",
"4347672385877442560"
],
"textSegmentEndOffsets": [
46.0,
166.0,
172.0,
239.0,
333.0,
691.0
]
}
],
"deployedModelId": "3958065938133155840"
}
```
### [projects.locations.endpoints.undeployModel](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.endpoints/undeployModel)
#### Call
```
request = clients['endpoint'].undeploy_model(
endpoint=endpoint_id,
deployed_model_id=deployed_model_id,
traffic_split={},
)
```
#### Response
```
result = request.result()
print(MessageToJson(result.__dict__["_pb"]))
```
*Example output*:
```
{}
```
# Cleaning up
To clean up all GCP resources used in this project, you can [delete the GCP
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
Otherwise, you can delete the individual resources you created in this tutorial.
```
delete_dataset = True
delete_model = True
delete_endpoint = True
delete_pipeline = True
delete_batchjob = True
delete_bucket = True
# Delete the dataset using the AI Platform (Unified) fully qualified identifier for the dataset
try:
if delete_dataset:
clients['dataset'].delete_dataset(name=dataset_id)
except Exception as e:
print(e)
# Delete the model using the AI Platform (Unified) fully qualified identifier for the model
try:
if delete_model:
clients['model'].delete_model(name=model_id)
except Exception as e:
print(e)
# Delete the endpoint using the AI Platform (Unified) fully qualified identifier for the endpoint
try:
if delete_endpoint:
clients['endpoint'].delete_endpoint(name=endpoint_id)
except Exception as e:
print(e)
# Delete the training pipeline using the AI Platform (Unified) fully qualified identifier for the training pipeline
try:
if delete_pipeline:
clients['pipeline'].delete_training_pipeline(name=training_pipeline_id)
except Exception as e:
print(e)
# Delete the batch job using the AI Platform (Unified) fully qualified identifier for the batch job
try:
if delete_batchjob:
clients['job'].delete_batch_prediction_job(name=batch_job_id)
except Exception as e:
print(e)
if delete_bucket and 'BUCKET_NAME' in globals():
! gsutil rm -r gs://$BUCKET_NAME
```
| github_jupyter |
# 1 - dataset_loaders.yaml
* This configuration file stores the settings of all datasets that will be utilized during the execution of an experiment, as can be seen in the example below for the **MovieLens 1M** dataset.
<!-- Which one is better? -->
<!-- * This configuration file stores the settings of all datasets required to run your experiment. Below, there is an example setting for the MovieLens 1M dataset. -->
* Template to define a dataset:
```yaml
# Dataset name
# Variables
```
* Example:
```yaml
'MovieLens 1M':
dataset_path: ./data/datasets/MovieLens 1M/
train_size: 0.8
test_consumes: 1
crono: False
random_seed: 0
```
* The standard data template for ratings consists of 4 comma-separated columns
user_id,item_id,rating,timestamp
# 2 - agents.yaml
* As its name suggests, this configuration file stores the settings of all agents utilized in the experiments.
* Template to define an agent:
```yaml
# Recommender agent name
# Agent type (irec/irec/agents.py)
# action_selection_policy:
# Action selection policy (irec/irec/action_selection_policies.py)
# Variables, if any
# value_function:
# Agent's value function (irec/irec/value_functions.py)
# Variables of the value function chosen
```
* Example
```yaml
LinearEGreedy: # Recommender agent name
SimpleAgent: # Agent type
action_selection_policy:
ASPEGreedy: # Action selection policy
# variaveis
epsilon: 0.1
value_function:
LinearEGreedy: # Agent's value function
# variáveis
item_var: 0.01
iterations: 20
num_lat: 10
stop_criteria: 0.0009
user_var: 0.01
var: 0.05
```
# 3 - agents_search.yaml
* As each agent may have a lot of variables, it is common to perform a search to find the best parameters and their combinations to achieve the best results/outcomes possible. So we created a specific file named **agents_search.yaml** to store all the combinations of parameters that a particular agent will execute.
* The agent template in this file is the same used in **agents.yaml**, however, as we will execute different combinations of the same agent, our configuration file will have multiple agents with different parameters, as shown in the following example
* In the example below, we have used the classic **EGreedy** agent with three different parameter combinations: epsilon 0.2, 0.3, and 0.5. Additionally, from the module `irec/app/generate_search_parameters.py` we can define which parameters will vary and their searching range.
```yaml
EGreedy:
- SimpleAgent:
action_selection_policy:
ASPEGreedy:
epsilon: 0.4
value_function:
EGreedy: {}
- SimpleAgent:
action_selection_policy:
ASPEGreedy:
epsilon: 0.3
value_function:
EGreedy: {}
- SimpleAgent:
action_selection_policy:
ASPEGreedy:
epsilon: 0.2
value_function:
EGreedy: {}
```
# 4 - dataset_agents.yaml
* This configuration file stores the agents and their parameters for each dataset. Usually, the agent's parameters vary according to the dataset, therefore it is convenient to store the best values found for each one. The results of a grid search on an agent will tell us which were the best parameters found for them, these data can then be stored in this file in case you want to reproduce the experiment later on.
* Template of `dataset_agents.yaml`:
```yaml
# Dataset name
# Agent 1
# Agent 2
...
# Agent n
```
* We can see in the example below the best parameters found for agents **LinUCB** and **LinearEGreedy** in the **MovieLens 1M** dataset.
```yaml
'MovieLens 1M':
LinUCB:
SimpleAgent:
action_selection_policy:
ASPGreedy: {}
value_function:
LinUCB:
alpha: 1.0
num_lat: 10
LinearEGreedy:
SimpleAgent:
action_selection_policy:
ASPEGreedy:
epsilon: 0.1
value_function:
LinearEGreedy:
item_var: 0.01
iterations: 20
num_lat: 10
stop_criteria: 0.0009
user_var: 0.01
var: 0.05
```
# 5 - evaluation_policies.yaml
* The evaluation policies are defined in this configuration file. To conduct an experiment, we need to define how the recommendation process will be executed and the user-item interactions. We specify these settings in this file according to the experiment's objectives.
* Template of `evaluation_policies.yaml`:
```yaml
# Evaluation policy name (irec/irec/evaluation_policies.py)
# Variables
```
* In the example below we can observe one of the evaluation policies implemented in the framework: **Interaction**, with its respective parameters.
```yaml
FixedInteraction: # Evaluation Policy
# Variables
num_interactions: 100 # Number of interactions for each user
interaction_size: 1 # Number of itens that will be recommended for each interaction
save_info: False # Salvar ou não informações obtidas durante a avaliação
```
# 6 - metric_evaluators.yaml
* In this file, it is defined evaluation metrics for an experiment. It specifies how to assess the interactions performed during the evaluation process..
* Template of `metric_evaluators.yaml`:
```yaml
# Evaluation metrics name (irec/irec/metric_evaluators.py)
# Variables
```
* In the example below we can see the use of an evaluation metric named **UserCumulativeInteraction** with its recpective parameters.
```yaml
UserCumulativeInteraction:
# variáveis
interaction_size: 1 # Número de itens recomendados a cada interação
interactions_to_evaluate: # Interações que serão avaliadas
- 5
- 10
- 20
- 50
- 100
num_interactions: 100 # Número de interações
relevance_evaluator_threshold: 3.999 # Rating
```
# 7 - defaults.yaml
* This configuration file defines the general settings of an experiment. We can specify not only the agents, the base, the policy, and the evaluation metric, but also some additional information.
* Example
```yaml
agent: UCB
agent_experiment: agent
data_dir: data/
dataset_experiment: dataset
dataset_loader: 'MovieLens 1M'
evaluation_experiment: evaluation
evaluation_policy: FixedInteraction
metric: Hits
metric_evaluator: UserCumulativeInteraction
pdf_dir: pdf/
tex_dir: tex/
```
| github_jupyter |
```
# default_exp models.MINIROCKET
```
# MINIROCKET
> A Very Fast (Almost) Deterministic Transform for Time Series Classification.
```
#export
from tsai.imports import *
from tsai.utils import *
from tsai.data.external import *
from tsai.models.layers import *
create_scripts
#export
from sktime.transformations.panel.rocket._minirocket import _fit as minirocket_fit
from sktime.transformations.panel.rocket._minirocket import _transform as minirocket_transform
from sktime.transformations.panel.rocket._minirocket_multivariate import _fit_multi as minirocket_fit_multi
from sktime.transformations.panel.rocket._minirocket_multivariate import _transform_multi as minirocket_transform_multi
from sktime.transformations.panel.rocket import MiniRocketMultivariate
from sklearn.linear_model import RidgeCV, RidgeClassifierCV
from sklearn.ensemble import VotingClassifier, VotingRegressor
# export
class MiniRocketClassifier(sklearn.pipeline.Pipeline):
"""Time series classification using MINIROCKET features and a linear classifier"""
def __init__(self, num_features=10_000, max_dilations_per_kernel=32, random_state=None,
alphas=np.logspace(-3, 3, 7), normalize_features=True, memory=None, verbose=False, scoring=None, class_weight=None, **kwargs):
"""
MiniRocketClassifier is recommended for up to 10k time series.
For a larger dataset, you can use MINIROCKET (in Pytorch).
scoring = None --> defaults to accuracy.
"""
self.steps = [('minirocketmultivariate', MiniRocketMultivariate(num_features=num_features,
max_dilations_per_kernel=max_dilations_per_kernel,
random_state=random_state)),
('ridgeclassifiercv', RidgeClassifierCV(alphas=alphas,
normalize=normalize_features,
scoring=scoring,
class_weight=class_weight,
**kwargs))]
store_attr()
self._validate_steps()
def __repr__(self):
return f'Pipeline(steps={self.steps.copy()})'
def save(self, fname=None, path='./models'):
fname = ifnone(fname, 'MiniRocketClassifier')
path = Path(path)
filename = path/fname
filename.parent.mkdir(parents=True, exist_ok=True)
with open(f'{filename}.pkl', 'wb') as output:
pickle.dump(self, output, pickle.HIGHEST_PROTOCOL)
MiniRocketClassifier.__doc__
#export
def load_minirocket(fname, path='./models'):
path = Path(path)
filename = path/fname
with open(f'{filename}.pkl', 'rb') as input:
output = pickle.load(input)
return output
# export
class MiniRocketRegressor(sklearn.pipeline.Pipeline):
"""Time series regression using MINIROCKET features and a linear regressor"""
def __init__(self, num_features=10000, max_dilations_per_kernel=32, random_state=None,
alphas=np.logspace(-3, 3, 7), *, normalize_features=True, memory=None, verbose=False, scoring=None, **kwargs):
"""
MiniRocketRegressor is recommended for up to 10k time series.
For a larger dataset, you can use MINIROCKET (in Pytorch).
scoring = None --> defaults to r2.
"""
self.steps = [('minirocketmultivariate', MiniRocketMultivariate(num_features=num_features,
max_dilations_per_kernel=max_dilations_per_kernel,
random_state=random_state)),
('ridgecv', RidgeCV(alphas=alphas, normalize=normalize_features, scoring=scoring, **kwargs))]
store_attr()
self._validate_steps()
def __repr__(self):
return f'Pipeline(steps={self.steps.copy()})'
def save(self, fname=None, path='./models'):
fname = ifnone(fname, 'MiniRocketRegressor')
path = Path(path)
filename = path/fname
filename.parent.mkdir(parents=True, exist_ok=True)
with open(f'{filename}.pkl', 'wb') as output:
pickle.dump(self, output, pickle.HIGHEST_PROTOCOL)
#export
def load_minirocket(fname, path='./models'):
path = Path(path)
filename = path/fname
with open(f'{filename}.pkl', 'rb') as input:
output = pickle.load(input)
return output
# export
class MiniRocketVotingClassifier(VotingClassifier):
"""Time series classification ensemble using MINIROCKET features, a linear classifier and majority voting"""
def __init__(self, n_estimators=5, weights=None, n_jobs=-1, num_features=10_000, max_dilations_per_kernel=32, random_state=None,
alphas=np.logspace(-3, 3, 7), normalize_features=True, memory=None, verbose=False, scoring=None, class_weight=None, **kwargs):
store_attr()
estimators = [(f'est_{i}', MiniRocketClassifier(num_features=num_features, max_dilations_per_kernel=max_dilations_per_kernel,
random_state=random_state, alphas=alphas, normalize_features=normalize_features, memory=memory,
verbose=verbose, scoring=scoring, class_weight=class_weight, **kwargs))
for i in range(n_estimators)]
super().__init__(estimators, voting='hard', weights=weights, n_jobs=n_jobs, verbose=verbose)
def __repr__(self):
return f'MiniRocketVotingClassifier(n_estimators={self.n_estimators}, \nsteps={self.estimators[0][1].steps})'
def save(self, fname=None, path='./models'):
fname = ifnone(fname, 'MiniRocketVotingClassifier')
path = Path(path)
filename = path/fname
filename.parent.mkdir(parents=True, exist_ok=True)
with open(f'{filename}.pkl', 'wb') as output:
pickle.dump(self, output, pickle.HIGHEST_PROTOCOL)
#export
def get_minirocket_preds(X, fname, path='./models', model=None):
if X.ndim == 1: X = X[np.newaxis][np.newaxis]
elif X.ndim == 2: X = X[np.newaxis]
if model is None:
model = load_minirocket(fname=fname, path=path)
return model.predict(X)
# export
class MiniRocketVotingRegressor(VotingRegressor):
"""Time series regression ensemble using MINIROCKET features, a linear regressor and a voting regressor"""
def __init__(self, n_estimators=5, weights=None, n_jobs=-1, num_features=10_000, max_dilations_per_kernel=32, random_state=None,
alphas=np.logspace(-3, 3, 7), normalize_features=True, memory=None, verbose=False, scoring=None, **kwargs):
store_attr()
estimators = [(f'est_{i}', MiniRocketRegressor(num_features=num_features, max_dilations_per_kernel=max_dilations_per_kernel,
random_state=random_state, alphas=alphas, normalize_features=normalize_features, memory=memory,
verbose=verbose, scoring=scoring, **kwargs))
for i in range(n_estimators)]
super().__init__(estimators, weights=weights, n_jobs=n_jobs, verbose=verbose)
def __repr__(self):
return f'MiniRocketVotingRegressor(n_estimators={self.n_estimators}, \nsteps={self.estimators[0][1].steps})'
def save(self, fname=None, path='./models'):
fname = ifnone(fname, 'MiniRocketVotingRegressor')
path = Path(path)
filename = path/fname
filename.parent.mkdir(parents=True, exist_ok=True)
with open(f'{filename}.pkl', 'wb') as output:
pickle.dump(self, output, pickle.HIGHEST_PROTOCOL)
# Univariate classification with sklearn-type API
dsid = 'OliveOil'
fname = 'MiniRocketClassifier'
X_train, y_train, X_test, y_test = get_UCR_data(dsid)
cls = MiniRocketClassifier()
cls.fit(X_train, y_train)
cls.save(fname)
pred = cls.score(X_test, y_test)
del cls
cls = load_minirocket(fname)
test_eq(cls.score(X_test, y_test), pred)
# Multivariate classification with sklearn-type API
dsid = 'NATOPS'
X_train, y_train, X_test, y_test = get_UCR_data(dsid)
cls = MiniRocketClassifier()
cls.fit(X_train, y_train)
cls.score(X_test, y_test)
# Multivariate classification with sklearn-type API
dsid = 'NATOPS'
X_train, y_train, X_test, y_test = get_UCR_data(dsid)
cls = MiniRocketVotingClassifier(5)
cls.fit(X_train, y_train)
cls.score(X_test, y_test)
# Univariate regression with sklearn-type API
from sklearn.metrics import mean_squared_error
dsid = 'Covid3Month'
fname = 'MiniRocketRegressor'
X_train, y_train, X_test, y_test = get_Monash_regression_data(dsid)
rmse_scorer = make_scorer(mean_squared_error, greater_is_better=False)
reg = MiniRocketRegressor(scoring=rmse_scorer)
reg.fit(X_train, y_train)
reg.save(fname)
del reg
reg = load_minirocket(fname)
y_pred = reg.predict(X_test)
rmse = mean_squared_error(y_test, y_pred, squared=False)
rmse
# Multivariate regression with sklearn-type API
from sklearn.metrics import mean_squared_error
dsid = 'AppliancesEnergy'
X_train, y_train, X_test, y_test = get_Monash_regression_data(dsid)
rmse_scorer = make_scorer(mean_squared_error, greater_is_better=False)
reg = MiniRocketRegressor(scoring=rmse_scorer)
reg.fit(X_train, y_train)
reg.save(fname)
del reg
reg = load_minirocket(fname)
y_pred = reg.predict(X_test)
rmse = mean_squared_error(y_test, y_pred, squared=False)
rmse
# Multivariate regression ensemble with sklearn-type API
reg = MiniRocketVotingRegressor(5, scoring=rmse_scorer)
reg.fit(X_train, y_train)
y_pred = reg.predict(X_test)
rmse = mean_squared_error(y_test, y_pred, squared=False)
rmse
#hide
out = create_scripts(); beep(out)
```
| github_jupyter |
# The Forest Fire Model
## A rapid introduction to Mesa
The [Forest Fire Model](http://en.wikipedia.org/wiki/Forest-fire_model) is one of the simplest examples of a model that exhibits self-organized criticality.
Mesa is a new, Pythonic agent-based modeling framework. A big advantage of using Python is that it a great language for interactive data analysis. Unlike some other ABM frameworks, with Mesa you can write a model, run it, and analyze it all in the same environment. (You don't have to, of course. But you can).
In this notebook, we'll go over a rapid-fire (pun intended, sorry) introduction to building and analyzing a model with Mesa.
First, some imports. We'll go over what all the Mesa ones mean just below.
```
import random
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
from mesa import Model, Agent
from mesa.time import RandomActivation
from mesa.space import Grid
from mesa.datacollection import DataCollector
from mesa.batchrunner import BatchRunner
```
## Building the model
Most models consist of basically two things: agents, and an world for the agents to be in. The Forest Fire model has only one kind of agent: a tree. A tree can either be unburned, on fire, or already burned. The environment is a grid, where each cell can either be empty or contain a tree.
First, let's define our tree agent. The agent needs to be assigned **x** and **y** coordinates on the grid, and that's about it. We could assign agents a condition to be in, but for now let's have them all start as being 'Fine'. Since the agent doesn't move, and there is only at most one tree per cell, we can use a tuple of its coordinates as a unique identifier.
Next, we define the agent's **step** method. This gets called whenever the agent needs to act in the world and takes the *model* object to which it belongs as an input. The tree's behavior is simple: If it is currently on fire, it spreads the fire to any trees above, below, to the left and the right of it that are not themselves burned out or on fire; then it burns itself out.
```
class TreeCell(Agent):
'''
A tree cell.
Attributes:
x, y: Grid coordinates
condition: Can be "Fine", "On Fire", or "Burned Out"
unique_id: (x,y) tuple.
unique_id isn't strictly necessary here, but it's good practice to give one to each
agent anyway.
'''
def __init__(self, model, pos):
'''
Create a new tree.
Args:
pos: The tree's coordinates on the grid. Used as the unique_id
'''
super().__init__(pos, model)
self.pos = pos
self.unique_id = pos
self.condition = "Fine"
def step(self):
'''
If the tree is on fire, spread it to fine trees nearby.
'''
if self.condition == "On Fire":
neighbors = self.model.grid.get_neighbors(self.pos, moore=False)
for neighbor in neighbors:
if neighbor.condition == "Fine":
neighbor.condition = "On Fire"
self.condition = "Burned Out"
```
Now we need to define the model object itself. The main thing the model needs is the grid, which the trees are placed on. But since the model is dynamic, it also needs to include time -- it needs a schedule, to manage the trees activation as they spread the fire from one to the other.
The model also needs a few parameters: how large the grid is and what the density of trees on it will be. Density will be the key parameter we'll explore below.
Finally, we'll give the model a data collector. This is a Mesa object which collects and stores data on the model as it runs for later analysis.
The constructor needs to do a few things. It instantiates all the model-level variables and objects; it randomly places trees on the grid, based on the density parameter; and it starts the fire by setting all the trees on one edge of the grid (x=0) as being On "Fire".
Next, the model needs a **step** method. Like at the agent level, this method defines what happens every step of the model. We want to activate all the trees, one at a time; then we run the data collector, to count how many trees are currently on fire, burned out, or still fine. If there are no trees left on fire, we stop the model by setting its **running** property to False.
```
class ForestFire(Model):
'''
Simple Forest Fire model.
'''
def __init__(self, height, width, density):
'''
Create a new forest fire model.
Args:
height, width: The size of the grid to model
density: What fraction of grid cells have a tree in them.
'''
# Initialize model parameters
self.height = height
self.width = width
self.density = density
# Set up model objects
self.schedule = RandomActivation(self)
self.grid = Grid(height, width, torus=False)
self.dc = DataCollector({"Fine": lambda m: self.count_type(m, "Fine"),
"On Fire": lambda m: self.count_type(m, "On Fire"),
"Burned Out": lambda m: self.count_type(m, "Burned Out")})
# Place a tree in each cell with Prob = density
for x in range(self.width):
for y in range(self.height):
if random.random() < self.density:
# Create a tree
new_tree = TreeCell(self, (x, y))
# Set all trees in the first column on fire.
if x == 0:
new_tree.condition = "On Fire"
self.grid[y][x] = new_tree
self.schedule.add(new_tree)
self.running = True
def step(self):
'''
Advance the model by one step.
'''
self.schedule.step()
self.dc.collect(self)
# Halt if no more fire
if self.count_type(self, "On Fire") == 0:
self.running = False
@staticmethod
def count_type(model, tree_condition):
'''
Helper method to count trees in a given condition in a given model.
'''
count = 0
for tree in model.schedule.agents:
if tree.condition == tree_condition:
count += 1
return count
```
## Running the model
Let's create a model with a 100 x 100 grid, and a tree density of 0.6. Remember, ForestFire takes the arguments *height*, *width*, *density*.
```
fire = ForestFire(100, 100, 0.6)
```
To run the model until it's done (that is, until it sets its **running** property to False) just use the **run_model()** method. This is implemented in the Model parent object, so we didn't need to implement it above.
```
fire.run_model()
```
That's all there is to it!
But... so what? This code doesn't include a visualization, after all.
Remember the data collector? Now we can put the data it collected into a pandas DataFrame:
```
results = fire.dc.get_model_vars_dataframe()
```
And chart it, to see the dynamics.
```
results.plot()
```
In this case, the fire burned itself out after about 90 steps, with many trees left unburned.
You can try changing the density parameter and rerunning the code above, to see how different densities yield different dynamics. For example:
```
fire = ForestFire(100, 100, 0.8)
fire.run_model()
results = fire.dc.get_model_vars_dataframe()
results.plot()
```
... But to really understand how the final outcome varies with density, we can't just tweak the parameter by hand over and over again. We need to do a batch run.
## Batch runs
Batch runs, also called parameter sweeps, allow use to systemically vary the density parameter, run the model, and check the output. Mesa provides a BatchRunner object which takes a model class, a dictionary of parameters and the range of values they can take and runs the model at each combination of these values. We can also give it reporters, which collect some data on the model at the end of each run and store it, associated with the parameters that produced it.
For ease of typing and reading, we'll first create the parameters to vary and the reporter, and then assign them to a new BatchRunner.
```
fixed_params = dict(height=50, # Height and width are constant
width=50)
# Vary density from 0.01 to 1, in 0.01 increments:
variable_params = dict(density=np.linspace(0,1,101)[1:])
# At the end of each model run, calculate the fraction of trees which are Burned Out
model_reporter = {"BurnedOut": lambda m: (ForestFire.count_type(m, "Burned Out") /
m.schedule.get_agent_count()) }
# Create the batch runner
param_run = BatchRunner(ForestFire, variable_parameters=variable_params,
fixed_parameters=fixed_params, model_reporters=model_reporter)
```
Now the BatchRunner, which we've named param_run, is ready to go. To run the model at every combination of parameters (in this case, every density value), just use the **run_all()** method.
```
param_run.run_all()
```
Like with the data collector, we can extract the data the batch runner collected into a dataframe:
```
df = param_run.get_model_vars_dataframe()
df.head()
```
As you can see, each row here is a run of the model, identified by its parameter values (and given a unique index by the Run column). To view how the BurnedOut fraction varies with density, we can easily just plot them:
```
plt.scatter(df.density, df.BurnedOut)
plt.xlim(0,1)
```
And we see the very clear emergence of a critical value around 0.5, where the model quickly shifts from almost no trees being burned, to almost all of them.
In this case we ran the model only once at each value. However, it's easy to have the BatchRunner execute multiple runs at each parameter combination, in order to generate more statistically reliable results. We do this using the *iteration* argument.
Let's run the model 5 times at each parameter point, and export and plot the results as above.
```
param_run = BatchRunner(ForestFire, variable_params, fixed_params,
iterations=5, model_reporters=model_reporter)
param_run.run_all()
df = param_run.get_model_vars_dataframe()
plt.scatter(df.density, df.BurnedOut)
plt.xlim(0,1)
```
| github_jupyter |
# Saving and Loading Models
In this notebook, I'll show you how to save and load models with PyTorch. This is important because you'll often want to load previously trained models to use in making predictions or to continue training on new data.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms
import helper
import fc_model # Library that helps implementing a model for building a fully connected classifier set.
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,))])
# Download and load the training data
trainset = datasets.FashionMNIST('~/.pytorch/F_MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
# Download and load the test data
testset = datasets.FashionMNIST('~/.pytorch/F_MNIST_data/', download=True, train=False, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=64, shuffle=True)
```
Here we can see one of the images.
```
image, label = next(iter(trainloader))
helper.imshow(image[0,:]);
```
# Train a network
To make things more concise here, I moved the model architecture and training code from the last part to a file called `fc_model`. Importing this, we can easily create a fully-connected network with `fc_model.Network`, and train the network using `fc_model.train`. I'll use this model (once it's trained) to demonstrate how we can save and load models.
```
# Create the network, define the criterion and optimizer
model = fc_model.Network(784, 10, [512, 256, 128])
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.parameters(), lr=0.001)
fc_model.train(model, trainloader, testloader, criterion, optimizer, epochs=2)
```
## Saving and loading networks
As you can imagine, it's impractical to train a network every time you need to use it. Instead, we can save trained networks then load them later to train more or use them for predictions.
The parameters for PyTorch networks are stored in a model's `state_dict`. We can see the state dict contains the weight and bias matrices for each of our layers.
```
print("Our model: \n\n", model, '\n')
print("The state dict keys: \n\n", model.state_dict().keys())
```
The simplest thing to do is simply save the state dict with `torch.save`. For example, we can save it to a file `'checkpoint.pth'`.
```
torch.save(model.state_dict(), 'checkpoint.pth')
```
Then we can load the state dict with `torch.load`.
```
state_dict = torch.load('checkpoint.pth')
print(state_dict.keys())
```
And to load the state dict in to the network, you do `model.load_state_dict(state_dict)`.
```
model.load_state_dict(state_dict)
```
Seems pretty straightforward, but as usual it's a bit more complicated. Loading the state dict works only if the model architecture is exactly the same as the checkpoint architecture. If I create a model with a different architecture, this fails.
```
# Try this
model = fc_model.Network(784, 10, [400, 200, 100])
# This will throw an error because the tensor sizes are wrong!
model.load_state_dict(state_dict)
```
This means we need to rebuild the model exactly as it was when trained. Information about the model architecture needs to be saved in the checkpoint, along with the state dict. To do this, you build a dictionary with all the information you need to compeletely rebuild the model.
```
checkpoint = {'input_size': 784,
'output_size': 10,
'hidden_layers': [each.out_features for each in model.hidden_layers],
'state_dict': model.state_dict()}
torch.save(checkpoint, 'checkpoint.pth')
```
Now the checkpoint has all the necessary information to rebuild the trained model. You can easily make that a function if you want. Similarly, we can write a function to load checkpoints.
```
def load_checkpoint(filepath):
checkpoint = torch.load(filepath)
model = fc_model.Network(checkpoint['input_size'],
checkpoint['output_size'],
checkpoint['hidden_layers'])
model.load_state_dict(checkpoint['state_dict'])
return model
model = load_checkpoint('checkpoint.pth')
print(model)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import os
import datetime
from __future__ import division
airport_data = pd.read_csv('../../Datasets/airports.csv', delimiter=',')
airlines_data = pd.read_csv('../../Datasets/airlines.csv', delimiter=',')
print "Completed Processing"
data = pd.read_csv('../../Datasets/flights.csv', delimiter=',', low_memory=False)
print "Completed Processing"
print data.shape
data['SCHEDULED_DEPARTURE'].head(100)
print data.columns.values
#Conversion of Day, Year, Month to DATE
data['FLIGHT_DATE'] = pd.to_datetime(data[['YEAR','MONTH','DAY']])
print "COMPLETED"
print data.shape
dataDict = pd.DataFrame({'DataType': data.dtypes.values, 'Missing_values': data.isnull().sum(),'Missing%': ((data.isnull().sum())/(data.shape[0])*100) }, index=data.columns.values)
dataDict
#data = data.drop(['FLIGHT_NUMBER','YEAR','MONTH','DAY','TAIL_NUMBER','TAXI_OUT','WHEELS_OFF','WHEELS_ON','TAXI_IN','WEATHER_DELAY','AIR_SYSTEM_DELAY','SECURITY_DELAY','AIRLINE_DELAY','LATE_AIRCRAFT_DELAY'], axis=1)
data = data.drop(['FLIGHT_NUMBER','DISTANCE_BINS','CANCELLATION_REASON','DIVERTED'])
print "DONE"
print data.shape
data['SCHEDULED_DEPARTURE'].head(100)
def get_time(time):
if pd.isnull(time):
np.nan
else:
h = int(time/100)
if h == 24:
h = 0
m = int(time)%100
modified_time = datetime.time(h,m)
return modified_time
data['SCHEDULED_DEPARTURE'] = data['SCHEDULED_DEPARTURE'].apply(lambda x: get_time(x))
data['DEPARTURE_TIME'] = data['DEPARTURE_TIME'].apply(lambda x: get_time(x))
data['ARRIVAL_TIME'] = data['ARRIVAL_TIME'].apply(lambda x: get_time(x))
data['SCHEDULED_ARRIVAL'] = data['SCHEDULED_ARRIVAL'].apply(lambda x: get_time(x))
print "Completed"
print data.shape
#Removing numeric values in ORIGIN and DESTINATION airports
print data.shape
data = data[data.ORIGIN_AIRPORT.str.isnumeric() == False]
print data.shape
#data = data[data.DESTINATION_AIRPORT.str.isnumeric() == False]
data.to_csv ('C:\\Users\\rashm\\PycharmProjects\\DataProgrammingProject\\ExploratoryAnalysis\\data.csv')
data.loc[:,['DEPARTURE_DELAY','ARRIVAL_DELAY']].head(30)
# Percentage of cancelled flights
(data[data.CANCELLATION_REASON.isnull() == False].DIVERTED.count())/(data.DIVERTED.count())
#airline vs arrival delay
FLIGHT_ARR_DELAY_METRIC = data.loc[data['ARRIVAL_DELAY']>15,['AIRLINE','ARRIVAL_DELAY']].groupby(['AIRLINE']).agg(['count','mean','median'])
FLIGHT_ARR_DELAY_METRIC.columns = ['TOTAL_DELAY_COUNT','MEAN_DELAY','MEDIAN_DELAY']
PERCENTAGE_FLIGHTS_DELAYED = data.loc[:,['AIRLINE','ARRIVAL_DELAY']].groupby(['AIRLINE']).count().rename(columns={"ARRIVAL_DELAY":"TOTAL_FLIGHTS"})
FLIGHT_ARR_DELAY_METRIC = pd.merge(FLIGHT_ARR_DELAY_METRIC,PERCENTAGE_FLIGHTS_DELAYED, how = 'inner', left_index = True, right_index = True )
FLIGHT_ARR_DELAY_METRIC['PERCENTAGE_DELAYED'] = (FLIGHT_ARR_DELAY_METRIC.TOTAL_DELAY_COUNT/FLIGHT_ARR_DELAY_METRIC.TOTAL_FLIGHTS)*100
FLIGHT_ARR_DELAY_METRIC
#FLIGHT_ARR_DELAY_METRIC.to_csv('C:\\Users\\rashm\\PycharmProjects\\DataProgrammingProject\\ExploratoryAnalysis\\AirlineVSArrivalDelay.csv')
arr_del_bool_mask1 = data['ARRIVAL_DELAY']>15
arr_del_bool_mask2 = data['ARRIVAL_DELAY']<=60
#airline vs arrival delay
FLIGHT_ARR_DELAY_METRIC = data.loc[arr_del_bool_mask1 & arr_del_bool_mask2,['AIRLINE','ARRIVAL_DELAY']].groupby(['AIRLINE']).agg(['count','mean','median'])
FLIGHT_ARR_DELAY_METRIC.columns = ['TOTAL_DELAY_COUNT','MEAN_DELAY','MEDIAN_DELAY']
PERCENTAGE_FLIGHTS_DELAYED = data.loc[:,['AIRLINE','ARRIVAL_DELAY']].groupby(['AIRLINE']).count().rename(columns={"ARRIVAL_DELAY":"TOTAL_FLIGHTS"})
FLIGHT_ARR_DELAY_METRIC = pd.merge(FLIGHT_ARR_DELAY_METRIC,PERCENTAGE_FLIGHTS_DELAYED, how = 'inner', left_index = True, right_index = True )
FLIGHT_ARR_DELAY_METRIC['PERCENTAGE_DELAYED'] = (FLIGHT_ARR_DELAY_METRIC.TOTAL_DELAY_COUNT/FLIGHT_ARR_DELAY_METRIC.TOTAL_FLIGHTS)*100
FLIGHT_ARR_DELAY_METRIC
#FLIGHT_ARR_DELAY_METRIC.to_csv('C:\\Users\\rashm\\PycharmProjects\\DataProgrammingProject\\ExploratoryAnalysis\\AirlineVSArrivalDelay.csv')
#airline vs departure delay
FLIGHT_DEP_DELAY_METRIC = data.loc[data['DEPARTURE_DELAY']>0,['AIRLINE','DEPARTURE_DELAY']].groupby(['AIRLINE']).agg(['count','mean'])
FLIGHT_DEP_DELAY_METRIC.columns = ['TOTAL_DELAY_COUNT','MEAN_DELAY_COUNT']
PERCENTAGE_FLIGHTS_DELAYED = data.loc[:,['AIRLINE','DEPARTURE_DELAY']].groupby(['AIRLINE']).count().rename(columns={"DEPARTURE_DELAY":"TOTAL_FLIGHTS"})
FLIGHT_DEP_DELAY_METRIC = pd.merge(FLIGHT_DEP_DELAY_METRIC,PERCENTAGE_FLIGHTS_DELAYED, how = 'inner', left_index = True, right_index = True )
FLIGHT_DEP_DELAY_METRIC['PERCENTAGE_DELAYED'] = (FLIGHT_DEP_DELAY_METRIC.TOTAL_DELAY_COUNT/FLIGHT_DEP_DELAY_METRIC.TOTAL_FLIGHTS)*100
FLIGHT_DEP_DELAY_METRIC
#FLIGHT_DEP_DELAY_METRIC.to_csv(path_or_buf='C:\\Users\\rashm\\PycharmProjects\\DataProgrammingProject\\ExploratoryAnalysis',mode = 'w',sep=',')
#Origin Airport vs Arrival Delay
AIRPORT_ARR_DELAY_METRIC = data.loc[data['ARRIVAL_DELAY']>0,['ORIGIN_AIRPORT','ARRIVAL_DELAY']].groupby(['ORIGIN_AIRPORT']).agg(['count','mean'])
AIRPORT_ARR_DELAY_METRIC.columns = ['TOTAL_DELAY_COUNT','MEAN_DELAY_COUNT']
PERCENTAGE_FLIGHTS_DELAYED = data.loc[:,['ORIGIN_AIRPORT','ARRIVAL_DELAY']].groupby(['ORIGIN_AIRPORT']).count().rename(columns={"ARRIVAL_DELAY":"TOTAL_FLIGHTS"})
AIRPORT_ARR_DELAY_METRIC = pd.merge(AIRPORT_ARR_DELAY_METRIC,PERCENTAGE_FLIGHTS_DELAYED, how = 'inner', left_index = True, right_index = True )
AIRPORT_ARR_DELAY_METRIC['PERCENTAGE_DELAYED'] = (AIRPORT_ARR_DELAY_METRIC.TOTAL_DELAY_COUNT/AIRPORT_ARR_DELAY_METRIC.TOTAL_FLIGHTS)*100
AIRPORT_ARR_DELAY_METRIC
#AIRPORT_ARR_DELAY_METRIC.to_csv(path_or_buf='C:\\Users\\rashm\\PycharmProjects\\DataProgrammingProject\\ExploratoryAnalysis',mode = 'w',sep=',')
#Origin Airport vs Departure Delay
ORIGIN_AIRPORT_DEPT_DELAY_METRIC = data.loc[data['DEPARTURE_DELAY']>0,['ORIGIN_AIRPORT','DEPARTURE_DELAY']].groupby(['ORIGIN_AIRPORT']).agg(['count','mean'])
ORIGIN_AIRPORT_DEPT_DELAY_METRIC.columns = ['TOTAL_DELAY_COUNT','MEAN_DELAY_COUNT']
PERCENTAGE_FLIGHTS_DELAYED = data.loc[:,['ORIGIN_AIRPORT','DEPARTURE_DELAY']].groupby(['ORIGIN_AIRPORT']).count().rename(columns={"DEPARTURE_DELAY":"TOTAL_FLIGHTS"})
ORIGIN_AIRPORT_DEPT_DELAY_METRIC = pd.merge(ORIGIN_AIRPORT_DEPT_DELAY_METRIC,PERCENTAGE_FLIGHTS_DELAYED, how = 'inner', left_index = True, right_index = True )
ORIGIN_AIRPORT_DEPT_DELAY_METRIC['PERCENTAGE_DELAYED'] = (ORIGIN_AIRPORT_DEPT_DELAY_METRIC.TOTAL_DELAY_COUNT/ORIGIN_AIRPORT_DEPT_DELAY_METRIC.TOTAL_FLIGHTS)*100
ORIGIN_AIRPORT_DEPT_DELAY_METRIC
#AIRPORT_ARR_DELAY_METRIC.to_csv(path_or_buf='C:\\Users\\rashm\\PycharmProjects\\DataProgrammingProject\\ExploratoryAnalysis',mode = 'w',sep=',')
#Destination Airport vs Departure Delay
DEST_AIRPORT_DEPT_DELAY_METRIC = data.loc[data['DEPARTURE_DELAY']>0,['DESTINATION_AIRPORT','DEPARTURE_DELAY']].groupby(['DESTINATION_AIRPORT']).agg(['count','mean'])
DEST_AIRPORT_DEPT_DELAY_METRIC.columns = ['TOTAL_DELAY_COUNT','MEAN_DELAY_COUNT']
PERCENTAGE_FLIGHTS_DELAYED = data.loc[:,['DESTINATION_AIRPORT','DEPARTURE_DELAY']].groupby(['DESTINATION_AIRPORT']).count().rename(columns={"DEPARTURE_DELAY":"TOTAL_FLIGHTS"})
DEST_AIRPORT_DEPT_DELAY_METRIC = pd.merge(DEST_AIRPORT_DEPT_DELAY_METRIC,PERCENTAGE_FLIGHTS_DELAYED, how = 'inner', left_index = True, right_index = True )
DEST_AIRPORT_DEPT_DELAY_METRIC['PERCENTAGE_DELAYED'] = (DEST_AIRPORT_DEPT_DELAY_METRIC.TOTAL_DELAY_COUNT/DEST_AIRPORT_DEPT_DELAY_METRIC.TOTAL_FLIGHTS)*100
DEST_AIRPORT_DEPT_DELAY_METRIC
#AIRPORT_ARR_DELAY_METRIC.to_csv(path_or_buf='C:\\Users\\rashm\\PycharmProjects\\DataProgrammingProject\\ExploratoryAnalysis',mode = 'w',sep=',')
#Destination Airport vs Arrival Delay
DEST_AIRPORT_ARR_DELAY_METRIC = data.loc[data['ARRIVAL_DELAY']>0,['DESTINATION_AIRPORT','ARRIVAL_DELAY']].groupby(['DESTINATION_AIRPORT']).agg(['count','mean'])
DEST_AIRPORT_ARR_DELAY_METRIC.columns = ['TOTAL_DELAY_COUNT','MEAN_DELAY_COUNT']
PERCENTAGE_FLIGHTS_DELAYED = data.loc[:,['DESTINATION_AIRPORT','ARRIVAL_DELAY']].groupby(['DESTINATION_AIRPORT']).count().rename(columns={"ARRIVAL_DELAY":"TOTAL_FLIGHTS"})
DEST_AIRPORT_ARR_DELAY_METRIC = pd.merge(DEST_AIRPORT_ARR_DELAY_METRIC,PERCENTAGE_FLIGHTS_DELAYED, how = 'inner', left_index = True, right_index = True )
DEST_AIRPORT_ARR_DELAY_METRIC['PERCENTAGE_DELAYED'] = (DEST_AIRPORT_ARR_DELAY_METRIC.TOTAL_DELAY_COUNT/DEST_AIRPORT_ARR_DELAY_METRIC.TOTAL_FLIGHTS)*100
DEST_AIRPORT_ARR_DELAY_METRIC
#AIRPORT_ARR_DELAY_METRIC.to_csv(path_or_buf='C:\\Users\\rashm\\PycharmProjects\\DataProgrammingProject\\ExploratoryAnalysis',mode = 'w',sep=',')
#Day Of The Week vs Arrival Delay
DAY_OF_WEEK_ARR_DELAY_METRIC = data.loc[data['ARRIVAL_DELAY']>0,['DAY_OF_WEEK','ARRIVAL_DELAY']].groupby(['DAY_OF_WEEK']).agg(['count','mean'])
DAY_OF_WEEK_ARR_DELAY_METRIC.columns = ['TOTAL_DELAY_COUNT','MEAN_DELAY_COUNT']
PERCENTAGE_FLIGHTS_DELAYED = data.loc[:,['DAY_OF_WEEK','ARRIVAL_DELAY']].groupby(['DAY_OF_WEEK']).count().rename(columns={"ARRIVAL_DELAY":"TOTAL_FLIGHTS"})
DAY_OF_WEEK_ARR_DELAY_METRIC = pd.merge(DAY_OF_WEEK_ARR_DELAY_METRIC,PERCENTAGE_FLIGHTS_DELAYED, how = 'inner', left_index = True, right_index = True )
DAY_OF_WEEK_ARR_DELAY_METRIC['PERCENTAGE_DELAYED'] = (DAY_OF_WEEK_ARR_DELAY_METRIC.TOTAL_DELAY_COUNT/DAY_OF_WEEK_ARR_DELAY_METRIC.TOTAL_FLIGHTS)*100
DAY_OF_WEEK_ARR_DELAY_METRIC
#AIRPORT_ARR_DELAY_METRIC.to_csv(path_or_buf='C:\\Users\\rashm\\PycharmProjects\\DataProgrammingProject\\ExploratoryAnalysis',mode = 'w',sep=',')
#Day Of The Week vs Departure Delay
DAY_OF_WEEK_DEPT_DELAY_METRIC = data.loc[data['DEPARTURE_DELAY']>0,['DAY_OF_WEEK','DEPARTURE_DELAY']].groupby(['DAY_OF_WEEK']).agg(['count','mean'])
DAY_OF_WEEK_DEPT_DELAY_METRIC.columns = ['TOTAL_DELAY_COUNT','MEAN_DELAY_COUNT']
PERCENTAGE_FLIGHTS_DELAYED = data.loc[:,['DAY_OF_WEEK','DEPARTURE_DELAY']].groupby(['DAY_OF_WEEK']).count().rename(columns={"DEPARTURE_DELAY":"TOTAL_FLIGHTS"})
DAY_OF_WEEK_DEPT_DELAY_METRIC = pd.merge(DAY_OF_WEEK_DEPT_DELAY_METRIC,PERCENTAGE_FLIGHTS_DELAYED, how = 'inner', left_index = True, right_index = True )
DAY_OF_WEEK_DEPT_DELAY_METRIC['PERCENTAGE_DELAYED'] = (DAY_OF_WEEK_DEPT_DELAY_METRIC.TOTAL_DELAY_COUNT/DAY_OF_WEEK_DEPT_DELAY_METRIC.TOTAL_FLIGHTS)*100
DAY_OF_WEEK_DEPT_DELAY_METRIC
#AIRPORT_ARR_DELAY_METRIC.to_csv(path_or_buf='C:\\Users\\rashm\\PycharmProjects\\DataProgrammingProject\\ExploratoryAnalysis',mode = 'w',sep=',')
# Month Vs Arrival Delay
#data.index = data['FLIGHT_DATE']
MONTHWISE_ARR_DELAY_METRICS = data.loc[data['ARRIVAL_DELAY']>0,['FLIGHT_DATE','ARRIVAL_DELAY']].groupby(pd.TimeGrouper(freq='M')).agg(['count','mean'])
MONTHWISE_ARR_DELAY_METRICS.columns = ['TOTAL_DELAY_COUNT','MEAN_DELAY_COUNT']
MONTHLY_TOTAL_FLIGHTS = data.loc[:,['ARRIVAL_DELAY']].groupby(pd.TimeGrouper(freq='M')).agg(['count'])
MONTHLY_TOTAL_FLIGHTS.columns = ['TOTAL_FLIGHTS']
MONTHWISE_ARR_DELAY_METRICS = pd.merge(MONTHWISE_ARR_DELAY_METRICS,MONTHLY_TOTAL_FLIGHTS,how = 'inner',left_index = True, right_index=True)
MONTHWISE_ARR_DELAY_METRICS['PERCENTAGE_DELAYS'] = (MONTHWISE_ARR_DELAY_METRICS.TOTAL_DELAY_COUNT/MONTHWISE_ARR_DELAY_METRICS.TOTAL_FLIGHTS)*100
MONTHWISE_ARR_DELAY_METRICS
# Month Vs Departure Delay
#data.index = data['FLIGHT_DATE']
MONTHWISE_DEPT_DELAY_METRICS = data.loc[data['DEPARTURE_DELAY']>0,['FLIGHT_DATE','DEPARTURE_DELAY']].groupby(pd.TimeGrouper(freq='M')).agg(['count','mean'])
MONTHWISE_DEPT_DELAY_METRICS.columns = ['TOTAL_DELAY_COUNT','MEAN_DELAY_COUNT']
MONTHLY_TOTAL_FLIGHTS = data.loc[:,['DEPARTURE_DELAY']].groupby(pd.TimeGrouper(freq='M')).agg(['count'])
MONTHLY_TOTAL_FLIGHTS.columns = ['TOTAL_FLIGHTS']
MONTHWISE_DEPT_DELAY_METRICS = pd.merge(MONTHWISE_DEPT_DELAY_METRICS,MONTHLY_TOTAL_FLIGHTS,how = 'inner',left_index = True, right_index=True)
MONTHWISE_DEPT_DELAY_METRICS['PERCENTAGE_DELAYS'] = (MONTHWISE_DEPT_DELAY_METRICS.TOTAL_DELAY_COUNT/MONTHWISE_DEPT_DELAY_METRICS.TOTAL_FLIGHTS)*100
MONTHWISE_DEPT_DELAY_METRICS
data
# State Vs Arrival Delay(Origin Airport)
STATE_ARR_DELAY_METRICS = pd.merge(data.loc[data['ARRIVAL_DELAY']>0,['ORIGIN_AIRPORT','ARRIVAL_DELAY']],airport_data.loc[:,['IATA_CODE','STATE']],how='outer',left_on =['ORIGIN_AIRPORT'],right_on=['IATA_CODE'])
STATE_ARR_DELAY_METRICS = STATE_ARR_DELAY_METRICS.loc[:,['STATE','ARRIVAL_DELAY']].groupby(['STATE']).agg(['count','mean'])
STATE_ARR_DELAY_METRICS.columns = ['TOTAL_DELAY_COUNT','MEAN_DELAY_COUNT']
STATE_TOTAL_FLIGHTS = pd.merge(data.loc[:,['ORIGIN_AIRPORT','ARRIVAL_DELAY']],airport_data.loc[:,['IATA_CODE','STATE']], how='outer',left_on =['ORIGIN_AIRPORT'],right_on=['IATA_CODE'])
STATE_TOTAL_FLIGHTS = STATE_TOTAL_FLIGHTS.loc[:,['STATE','ARRIVAL_DELAY']].groupby(['STATE']).agg(['count'])
STATE_TOTAL_FLIGHTS.columns = ['TOTAL_FLIGHTS']
STATE_TOTAL_FLIGHTS
STATE_ARR_DELAY_METRICS = pd.merge(STATE_ARR_DELAY_METRICS,STATE_TOTAL_FLIGHTS, how = 'inner', left_index=True, right_index=True)
STATE_ARR_DELAY_METRICS['PERCENTAGE_DELAYS'] = (STATE_ARR_DELAY_METRICS.TOTAL_DELAY_COUNT/STATE_ARR_DELAY_METRICS.TOTAL_FLIGHTS)*100
STATE_ARR_DELAY_METRICS
# State Vs Departure Delay(Origin Airport)
STATE_DEPT_DELAY_METRICS = pd.merge(data.loc[data['DEPARTURE_DELAY']>0,['ORIGIN_AIRPORT','DEPARTURE_DELAY']],airport_data.loc[:,['IATA_CODE','STATE']],how='outer',left_on =['ORIGIN_AIRPORT'],right_on=['IATA_CODE'])
STATE_DEPT_DELAY_METRICS = STATE_DEPT_DELAY_METRICS.loc[:,['STATE','DEPARTURE_DELAY']].groupby(['STATE']).agg(['count','mean'])
STATE_DEPT_DELAY_METRICS.columns = ['TOTAL_DELAY_COUNT','MEAN_DELAY_COUNT']
STATE_TOTAL_FLIGHTS = pd.merge(data.loc[:,['ORIGIN_AIRPORT','DEPARTURE_DELAY']],airport_data.loc[:,['IATA_CODE','STATE']], how='outer',left_on =['ORIGIN_AIRPORT'],right_on=['IATA_CODE'])
STATE_TOTAL_FLIGHTS = STATE_TOTAL_FLIGHTS.loc[:,['STATE','DEPARTURE_DELAY']].groupby(['STATE']).agg(['count'])
STATE_TOTAL_FLIGHTS.columns = ['TOTAL_FLIGHTS']
STATE_TOTAL_FLIGHTS
STATE_DEPT_DELAY_METRICS = pd.merge(STATE_DEPT_DELAY_METRICS,STATE_TOTAL_FLIGHTS, how = 'inner', left_index=True, right_index=True)
STATE_DEPT_DELAY_METRICS['PERCENTAGE_DELAYS'] = (STATE_DEPT_DELAY_METRICS.TOTAL_DELAY_COUNT/STATE_DEPT_DELAY_METRICS.TOTAL_FLIGHTS)*100
STATE_DEPT_DELAY_METRICS
# State Vs Departure Delay(Destination Airport)
STATE_DEPT_DELAY_METRICS = pd.merge(data.loc[data['DEPARTURE_DELAY']>0,['DESTINATION_AIRPORT','DEPARTURE_DELAY']],airport_data.loc[:,['IATA_CODE','STATE']],how='outer',left_on =['DESTINATION_AIRPORT'],right_on=['IATA_CODE'])
STATE_DEPT_DELAY_METRICS = STATE_DEPT_DELAY_METRICS.loc[:,['STATE','DEPARTURE_DELAY']].groupby(['STATE']).agg(['count','mean'])
STATE_DEPT_DELAY_METRICS.columns = ['TOTAL_DELAY_COUNT','MEAN_DELAY_COUNT']
STATE_TOTAL_FLIGHTS = pd.merge(data.loc[:,['DESTINATION_AIRPORT','DEPARTURE_DELAY']],airport_data.loc[:,['IATA_CODE','STATE']], how='outer',left_on =['DESTINATION_AIRPORT'],right_on=['IATA_CODE'])
STATE_TOTAL_FLIGHTS = STATE_TOTAL_FLIGHTS.loc[:,['STATE','DEPARTURE_DELAY']].groupby(['STATE']).agg(['count'])
STATE_TOTAL_FLIGHTS.columns = ['TOTAL_FLIGHTS']
STATE_TOTAL_FLIGHTS
STATE_DEPT_DELAY_METRICS = pd.merge(STATE_DEPT_DELAY_METRICS,STATE_TOTAL_FLIGHTS, how = 'inner', left_index=True, right_index=True)
STATE_DEPT_DELAY_METRICS['PERCENTAGE_DELAYS'] = (STATE_DEPT_DELAY_METRICS.TOTAL_DELAY_COUNT/STATE_DEPT_DELAY_METRICS.TOTAL_FLIGHTS)*100
STATE_DEPT_DELAY_METRICS
# State Vs Arrival Delay(Destination Airport)
STATE_ARR_DELAY_METRICS = pd.merge(data.loc[data['ARRIVAL_DELAY']>0,['DESTINATION_AIRPORT','ARRIVAL_DELAY']],airport_data.loc[:,['IATA_CODE','STATE']],how='outer',left_on =['DESTINATION_AIRPORT'],right_on=['IATA_CODE'])
STATE_ARR_DELAY_METRICS = STATE_ARR_DELAY_METRICS.loc[:,['STATE','ARRIVAL_DELAY']].groupby(['STATE']).agg(['count','mean'])
STATE_ARR_DELAY_METRICS.columns = ['TOTAL_DELAY_COUNT','MEAN_DELAY_COUNT']
STATE_TOTAL_FLIGHTS = pd.merge(data.loc[:,['DESTINATION_AIRPORT','ARRIVAL_DELAY']],airport_data.loc[:,['IATA_CODE','STATE']], how='outer',left_on =['DESTINATION_AIRPORT'],right_on=['IATA_CODE'])
STATE_TOTAL_FLIGHTS = STATE_TOTAL_FLIGHTS.loc[:,['STATE','ARRIVAL_DELAY']].groupby(['STATE']).agg(['count'])
STATE_TOTAL_FLIGHTS.columns = ['TOTAL_FLIGHTS']
STATE_TOTAL_FLIGHTS
STATE_ARR_DELAY_METRICS = pd.merge(STATE_ARR_DELAY_METRICS,STATE_TOTAL_FLIGHTS, how = 'inner', left_index=True, right_index=True)
STATE_ARR_DELAY_METRICS['PERCENTAGE_DELAYS'] = (STATE_ARR_DELAY_METRICS.TOTAL_DELAY_COUNT/STATE_ARR_DELAY_METRICS.TOTAL_FLIGHTS)*100
#STATE_ARR_DELAY_METRICS.to_csv ('C:\\Users\\rashm\\PycharmProjects\\DataProgrammingProject\\ExploratoryAnalysis\\STATE_ARR_DELAY_METRICS_BY_DESTINATION.csv')
print "max:%s" %(data.DISTANCE.max())
print "min:%s" %(data.DISTANCE.min())
data['DISTANCE_BINS']= pd.qcut(data.DISTANCE,11)
data.head()
test = data.loc[:,['DISTANCE','ARRIVAL_DELAY']].groupby(data['DISTANCE_BINS']).mean()
test
data.ARRIVAL_TIME.unique()
```
| github_jupyter |
<img src='https://assets.leetcode.com/uploads/2020/11/07/search1.jpg'>
```
from collections import defaultdict
class TrieNode:
def __init__(self):
self.child = defaultdict(TrieNode)
self.isWord = False
class Trie:
def __init__(self):
self.root = TrieNode()
def insert(self, word):
node = self.root
for c in word:
node = node.child[c]
node.isWord = True
class Solution:
def findWords(self, board, words):
self.res, trie = [], Trie()
node = trie.root
for w in words:
trie.insert(w)
for i, row in enumerate(board):
for j, c in enumerate(row):
self.dfs(board, node, i, j, '')
return self.res
def dfs(self, board, node, row, col, path):
if node.isWord:
self.res.append(path[:])
node.isWord = False
if row < 0 or col < 0 or row >= len(board) or col >= len(board[0]):
return
c = board[row][col]
if c not in node.child:
return
board[row][col] = '#'
for n_r, n_c in (row+1, col), (row-1, col), (row, col+1), (row, col-1):
self.dfs(board, node.child[c], n_r, n_c, path+c)
board[row][col] = c
class Solution:
def findWords(self, board, words):
pass
solution = Solution()
solution.findWords(words = ["oath","pea","eat","rain"], board =
[
['o','a','a','n'],
['e','t','a','e'],
['i','h','k','r'],
['i','f','l','v']
])
a = '2'
print(a[:])
class Node:
def __init__(self):
self.children = {}
self.is_end = False
class Trie:
def __init__(self):
self.root = Node()
def insert(self, word):
node = self.root
for char in word:
if char not in node.children:
node.children[char] = Node()
node = node.children[char]
node.is_end = True
class Solution:
def findWords(self, board, words):
self.ans = set()
self.words = words
trie = Trie()
for word in words:
trie.insert(word)
for i in range(len(board)):
for j in range(len(board[0])):
self.dfs(board, trie.root, i, j, '')
return self.ans
def dfs(self, board, node, i, j, path):
if i < 0 or j < 0 or i >= len(board) or j >= len(board[0]):
return
if node.is_end and path not in self.ans:
self.ans.add(path[:])
node.end = False
char = board[i][j]
if char not in node.children:
return
board[i][j] = '#'
for n_i, n_j in (i+1, j), (i, j+1), (i, j-1), (i-1, j):
self.dfs(board, node.children[char], n_i, n_j , path+char)
board[i][j] = char
class Node:
def __init__(self):
self.children = {}
self.is_end = False
class Trie:
def __init__(self):
self.root = Node()
def insert(self, word):
node = self.root
for char in word:
if char not in node.children:
node.children[char] = Node()
node = node.children[char]
node.is_end = True
class Solution:
def findWords(self, board, words):
self.ans = []
trie = Trie()
for word in words:
trie.insert(word)
for i in range(len(board)):
for j in range(len(board[0])):
self.dfs(board, trie.root, i, j, '')
return self.ans
def dfs(self, board, node, i, j, path):
if node.is_end:
self.ans.append(path[:])
node.end = False
if i < 0 or j < 0 or i >= len(board) or j >= len(board[0]):
return
char = board[i][j]
if char not in node.children:
return
board[i][j] = '#'
for n_i, n_j in (i+1, j), (i, j+1), (i-1, j), (i, j-1):
self.dfs(board, node.children[char], n_i, n_j , path+char)
board[i][j] = char
class Node:
def __init__(self):
self.children = {}
self.is_end = False
class Trie:
def __init__(self):
self.root = Node()
def insert(self, word):
node = self.root
for char in word:
if char not in node.children:
node.children[char] = Node()
node = node.children[char]
node.is_end = True
class Solution:
def findWords(self, board, words):
self.res, trie = [], Trie()
node = trie.root
for w in words:
trie.insert(w)
for i, row in enumerate(board):
for j, c in enumerate(row):
self.dfs(board, node, i, j, '')
return self.res
def dfs(self, board, node, row, col, path):
if node.isWord:
self.res.append(path[:])
node.isWord = False
if row < 0 or col < 0 or row >= len(board) or col >= len(board[0]):
return
c = board[row][col]
if c not in node.child:
return
board[row][col] = '#'
for n_r, n_c in (row+1, col), (row-1, col), (row, col+1), (row, col-1):
self.dfs(board, node.child[c], n_r, n_c, path+c)
board[row][col] = c
solution = Solution()
solution.findWords([["a","b"],["a","a"]], ["aba","baa","bab","aaab","aaa","aaaa","aaba"])
class Node:
def __init__(self):
self.children = {}
self.isWord = False
class Trie:
def __init__(self):
self.root = Node()
def insert(self, word):
node = self.root
for char in word:
if char not in node.children:
node.children[char] = Node()
node = node.children[char]
node.isWord = True
class Solution:
def findWords(self, board, words):
self.res, trie = [], Trie()
node = trie.root
for w in words:
trie.insert(w)
for i, row in enumerate(board):
for j, c in enumerate(row):
self.dfs(board, node, i, j, '')
return self.res
def dfs(self, board, node, row, col, path):
if node.isWord:
self.res.append(path[:])
node.isWord = False
if row < 0 or col < 0 or row >= len(board) or col >= len(board[0]):
return
c = board[row][col]
if c not in node.children:
return
board[row][col] = '#'
for n_r, n_c in (row+1, col), (row-1, col), (row, col+1), (row, col-1):
self.dfs(board, node.children[c], n_r, n_c, path+c)
board[row][col] = c
```
| github_jupyter |
# Sentiment Analysis with an RNN
In this notebook, you'll implement a recurrent neural network that performs sentiment analysis. Using an RNN rather than a feedfoward network is more accurate since we can include information about the *sequence* of words. Here we'll use a dataset of movie reviews, accompanied by labels.
The architecture for this network is shown below.
<img src="assets/network_diagram.png" width=400px>
Here, we'll pass in words to an embedding layer. We need an embedding layer because we have tens of thousands of words, so we'll need a more efficient representation for our input data than one-hot encoded vectors. You should have seen this before from the word2vec lesson. You can actually train up an embedding with word2vec and use it here. But it's good enough to just have an embedding layer and let the network learn the embedding table on it's own.
From the embedding layer, the new representations will be passed to LSTM cells. These will add recurrent connections to the network so we can include information about the sequence of words in the data. Finally, the LSTM cells will go to a sigmoid output layer here. We're using the sigmoid because we're trying to predict if this text has positive or negative sentiment. The output layer will just be a single unit then, with a sigmoid activation function.
We don't care about the sigmoid outputs except for the very last one, we can ignore the rest. We'll calculate the cost from the output of the last step and the training label.
```
import numpy as np
import tensorflow as tf
with open('../sentiment-network/reviews.txt', 'r') as f:
reviews = f.read()
with open('../sentiment-network/labels.txt', 'r') as f:
labels = f.read()
reviews[:2000]
```
## Data preprocessing
The first step when building a neural network model is getting your data into the proper form to feed into the network. Since we're using embedding layers, we'll need to encode each word with an integer. We'll also want to clean it up a bit.
You can see an example of the reviews data above. We'll want to get rid of those periods. Also, you might notice that the reviews are delimited with newlines `\n`. To deal with those, I'm going to split the text into each review using `\n` as the delimiter. Then I can combined all the reviews back together into one big string.
First, let's remove all punctuation. Then get all the text without the newlines and split it into individual words.
```
from string import punctuation
all_text = ''.join([c for c in reviews if c not in punctuation])
reviews = all_text.split('\n')
all_text = ' '.join(reviews)
words = all_text.split()
all_text[:2000]
words[:100]
```
### Encoding the words
The embedding lookup requires that we pass in integers to our network. The easiest way to do this is to create dictionaries that map the words in the vocabulary to integers. Then we can convert each of our reviews into integers so they can be passed into the network.
> **Exercise:** Now you're going to encode the words with integers. Build a dictionary that maps words to integers. Later we're going to pad our input vectors with zeros, so make sure the integers **start at 1, not 0**.
> Also, convert the reviews to integers and store the reviews in a new list called `reviews_ints`.
```
# Create your dictionary that maps vocab words to integers here
vocab_to_int = {word: i for i, word in enumerate(set(words), 1)}
# Convert the reviews to integers, same shape as reviews list, but with integers
print(reviews[1].split(" "))
reviews_ints = [[vocab_to_int[word] for word in review.split(" ") if word is not ''] for review in reviews]
print(reviews_ints[1])
```
### Encoding the labels
Our labels are "positive" or "negative". To use these labels in our network, we need to convert them to 0 and 1.
> **Exercise:** Convert labels from `positive` and `negative` to 1 and 0, respectively.
```
# Convert labels to 1s and 0s for 'positive' and 'negative'
labels_raw = labels
print(labels[:20])
labels = [1 if label == "positive" else 0 for label in labels_raw.split("\n")]
print(labels[:10])
```
If you built `labels` correctly, you should see the next output.
```
from collections import Counter
review_lens = Counter([len(x) for x in reviews_ints])
print("Zero-length reviews: {}".format(review_lens[0]))
print("Maximum review length: {}".format(max(review_lens)))
```
Okay, a couple issues here. We seem to have one review with zero length. And, the maximum review length is way too many steps for our RNN. Let's truncate to 200 steps. For reviews shorter than 200, we'll pad with 0s. For reviews longer than 200, we can truncate them to the first 200 characters.
> **Exercise:** First, remove the review with zero length from the `reviews_ints` list.
```
# Filter out that review with 0 length
print(len(reviews_ints))
reviews_ints = [r for r in reviews_ints if len(r) > 0]
print(len(reviews_ints))
```
> **Exercise:** Now, create an array `features` that contains the data we'll pass to the network. The data should come from `review_ints`, since we want to feed integers to the network. Each row should be 200 elements long. For reviews shorter than 200 words, left pad with 0s. That is, if the review is `['best', 'movie', 'ever']`, `[117, 18, 128]` as integers, the row will look like `[0, 0, 0, ..., 0, 117, 18, 128]`. For reviews longer than 200, use on the first 200 words as the feature vector.
This isn't trivial and there are a bunch of ways to do this. But, if you're going to be building your own deep learning networks, you're going to have to get used to preparing your data.
```
seq_len = 200
features = [r[:200] if (len(r) >= seq_len) else [0] * (seq_len - len(r)) + r for r in reviews_ints]
features = np.array(features)
```
If you build features correctly, it should look like that cell output below.
```
features[:10,:100]
```
## Training, Validation, Test
With our data in nice shape, we'll split it into training, validation, and test sets.
> **Exercise:** Create the training, validation, and test sets here. You'll need to create sets for the features and the labels, `train_x` and `train_y` for example. Define a split fraction, `split_frac` as the fraction of data to keep in the training set. Usually this is set to 0.8 or 0.9. The rest of the data will be split in half to create the validation and testing data.
```
features = np.array(features)
labels = np.array(labels)
split_frac = 0.8
split_len = round(split_frac * len(features))
np.random.seed(12)
indices = np.random.permutation(features.shape[0])
train_x, val_x = features[indices[:split_len]], features[indices[split_len:]]
train_y, val_y = labels[indices[:split_len]], labels[indices[split_len:]]
np.random.seed(12)
indices = np.random.permutation(val_x.shape[0])
split_frac = 0.5
split_len = round(split_frac * len(val_x))
val_x, test_x = val_x[indices[:split_len]], val_x[indices[split_len:]]
val_y, test_y = val_y[indices[:split_len]], val_y[indices[split_len:]]
print("\t\t\tFeature Shapes:")
print("Train set: \t\t{}".format(train_x.shape),
"\nValidation set: \t{}".format(val_x.shape),
"\nTest set: \t\t{}".format(test_x.shape))
```
With train, validation, and text fractions of 0.8, 0.1, 0.1, the final shapes should look like:
```
Feature Shapes:
Train set: (20000, 200)
Validation set: (2500, 200)
Test set: (2500, 200)
```
## Build the graph
Here, we'll build the graph. First up, defining the hyperparameters.
* `lstm_size`: Number of units in the hidden layers in the LSTM cells. Usually larger is better performance wise. Common values are 128, 256, 512, etc.
* `lstm_layers`: Number of LSTM layers in the network. I'd start with 1, then add more if I'm underfitting.
* `batch_size`: The number of reviews to feed the network in one training pass. Typically this should be set as high as you can go without running out of memory.
* `learning_rate`: Learning rate
```
lstm_size = 256
lstm_layers = 1
batch_size = 1000
learning_rate = 0.001
```
For the network itself, we'll be passing in our 200 element long review vectors. Each batch will be `batch_size` vectors. We'll also be using dropout on the LSTM layer, so we'll make a placeholder for the keep probability.
> **Exercise:** Create the `inputs_`, `labels_`, and drop out `keep_prob` placeholders using `tf.placeholder`. `labels_` needs to be two-dimensional to work with some functions later. Since `keep_prob` is a scalar (a 0-dimensional tensor), you shouldn't provide a size to `tf.placeholder`.
```
n_words = len(vocab_to_int)
print(n_words)
# Create the graph object
graph = tf.Graph()
# Add nodes to the graph
with graph.as_default():
inputs_ = tf.placeholder(dtype=tf.int32, shape=(batch_size, seq_len), name="inputs")
labels_ = tf.placeholder(dtype=tf.int32, shape=(batch_size, None), name="labels")
keep_prob = tf.placeholder(dtype=tf.float32, name="keep_prob")
```
### Embedding
Now we'll add an embedding layer. We need to do this because there are 74000 words in our vocabulary. It is massively inefficient to one-hot encode our classes here. You should remember dealing with this problem from the word2vec lesson. Instead of one-hot encoding, we can have an embedding layer and use that layer as a lookup table. You could train an embedding layer using word2vec, then load it here. But, it's fine to just make a new layer and let the network learn the weights.
> **Exercise:** Create the embedding lookup matrix as a `tf.Variable`. Use that embedding matrix to get the embedded vectors to pass to the LSTM cell with [`tf.nn.embedding_lookup`](https://www.tensorflow.org/api_docs/python/tf/nn/embedding_lookup). This function takes the embedding matrix and an input tensor, such as the review vectors. Then, it'll return another tensor with the embedded vectors. So, if the embedding layer has 200 units, the function will return a tensor with size [batch_size, 200].
```
# Size of the embedding vectors (number of units in the embedding layer)
embed_size = 300
with graph.as_default():
embedding = tf.Variable(tf.random_uniform((n_words+1, embed_size), -1, 1))
embed = tf.nn.embedding_lookup(embedding, inputs_)
```
### LSTM cell
<img src="assets/network_diagram.png" width=400px>
Next, we'll create our LSTM cells to use in the recurrent network ([TensorFlow documentation](https://www.tensorflow.org/api_docs/python/tf/contrib/rnn)). Here we are just defining what the cells look like. This isn't actually building the graph, just defining the type of cells we want in our graph.
To create a basic LSTM cell for the graph, you'll want to use `tf.contrib.rnn.BasicLSTMCell`. Looking at the function documentation:
```
tf.contrib.rnn.BasicLSTMCell(num_units, forget_bias=1.0, input_size=None, state_is_tuple=True, activation=<function tanh at 0x109f1ef28>)
```
you can see it takes a parameter called `num_units`, the number of units in the cell, called `lstm_size` in this code. So then, you can write something like
```
lstm = tf.contrib.rnn.BasicLSTMCell(num_units)
```
to create an LSTM cell with `num_units`. Next, you can add dropout to the cell with `tf.contrib.rnn.DropoutWrapper`. This just wraps the cell in another cell, but with dropout added to the inputs and/or outputs. It's a really convenient way to make your network better with almost no effort! So you'd do something like
```
drop = tf.contrib.rnn.DropoutWrapper(cell, output_keep_prob=keep_prob)
```
Most of the time, your network will have better performance with more layers. That's sort of the magic of deep learning, adding more layers allows the network to learn really complex relationships. Again, there is a simple way to create multiple layers of LSTM cells with `tf.contrib.rnn.MultiRNNCell`:
```
cell = tf.contrib.rnn.MultiRNNCell([drop] * lstm_layers)
```
Here, `[drop] * lstm_layers` creates a list of cells (`drop`) that is `lstm_layers` long. The `MultiRNNCell` wrapper builds this into multiple layers of RNN cells, one for each cell in the list.
So the final cell you're using in the network is actually multiple (or just one) LSTM cells with dropout. But it all works the same from an achitectural viewpoint, just a more complicated graph in the cell.
> **Exercise:** Below, use `tf.contrib.rnn.BasicLSTMCell` to create an LSTM cell. Then, add drop out to it with `tf.contrib.rnn.DropoutWrapper`. Finally, create multiple LSTM layers with `tf.contrib.rnn.MultiRNNCell`.
Here is [a tutorial on building RNNs](https://www.tensorflow.org/tutorials/recurrent) that will help you out.
```
with graph.as_default():
# Your basic LSTM cell
lstm = tf.contrib.rnn.BasicLSTMCell(lstm_size)
# Add dropout to the cell
drop = tf.contrib.rnn.DropoutWrapper(lstm, output_keep_prob=keep_prob)
# Stack up multiple LSTM layers, for deep learning
cell = tf.contrib.rnn.MultiRNNCell([drop] * lstm_layers)
# Getting an initial state of all zeros
initial_state = cell.zero_state(batch_size, tf.float32)
```
### RNN forward pass
<img src="assets/network_diagram.png" width=400px>
Now we need to actually run the data through the RNN nodes. You can use [`tf.nn.dynamic_rnn`](https://www.tensorflow.org/api_docs/python/tf/nn/dynamic_rnn) to do this. You'd pass in the RNN cell you created (our multiple layered LSTM `cell` for instance), and the inputs to the network.
```
outputs, final_state = tf.nn.dynamic_rnn(cell, inputs, initial_state=initial_state)
```
Above I created an initial state, `initial_state`, to pass to the RNN. This is the cell state that is passed between the hidden layers in successive time steps. `tf.nn.dynamic_rnn` takes care of most of the work for us. We pass in our cell and the input to the cell, then it does the unrolling and everything else for us. It returns outputs for each time step and the final_state of the hidden layer.
> **Exercise:** Use `tf.nn.dynamic_rnn` to add the forward pass through the RNN. Remember that we're actually passing in vectors from the embedding layer, `embed`.
```
with graph.as_default():
outputs, final_state = tf.nn.dynamic_rnn(cell, embed, initial_state=initial_state)
```
### Output
We only care about the final output, we'll be using that as our sentiment prediction. So we need to grab the last output with `outputs[:, -1]`, the calculate the cost from that and `labels_`.
```
with graph.as_default():
predictions = tf.contrib.layers.fully_connected(outputs[:, -1], 1, activation_fn=tf.sigmoid)
cost = tf.losses.mean_squared_error(labels_, predictions)
optimizer = tf.train.AdamOptimizer(learning_rate).minimize(cost)
```
### Validation accuracy
Here we can add a few nodes to calculate the accuracy which we'll use in the validation pass.
```
with graph.as_default():
correct_pred = tf.equal(tf.cast(tf.round(predictions), tf.int32), labels_)
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
```
### Batching
This is a simple function for returning batches from our data. First it removes data such that we only have full batches. Then it iterates through the `x` and `y` arrays and returns slices out of those arrays with size `[batch_size]`.
```
def get_batches(x, y, batch_size=100):
n_batches = len(x)//batch_size
x, y = x[:n_batches*batch_size], y[:n_batches*batch_size]
for ii in range(0, len(x), batch_size):
yield x[ii:ii+batch_size], y[ii:ii+batch_size]
```
## Training
Below is the typical training code. If you want to do this yourself, feel free to delete all this code and implement it yourself. Before you run this, make sure the `checkpoints` directory exists.
```
epochs = 10
with graph.as_default():
saver = tf.train.Saver()
with tf.Session(graph=graph) as sess:
sess.run(tf.global_variables_initializer())
iteration = 1
for e in range(epochs):
state = sess.run(initial_state)
for ii, (x, y) in enumerate(get_batches(train_x, train_y, batch_size), 1):
feed = {inputs_: x,
labels_: y[:, None],
keep_prob: 0.5,
initial_state: state}
loss, state, _ = sess.run([cost, final_state, optimizer], feed_dict=feed)
if iteration%5==0:
print("Epoch: {}/{}".format(e+1, epochs),
"Iteration: {}".format(iteration),
"Train loss: {:.3f}".format(loss))
if iteration%25==0:
val_acc = []
val_state = sess.run(cell.zero_state(batch_size, tf.float32))
for x, y in get_batches(val_x, val_y, batch_size):
feed = {inputs_: x,
labels_: y[:, None],
keep_prob: 1,
initial_state: val_state}
batch_acc, val_state = sess.run([accuracy, final_state], feed_dict=feed)
val_acc.append(batch_acc)
print("Val acc: {:.3f}".format(np.mean(val_acc)))
iteration +=1
saver.save(sess, "checkpoints/sentiment.ckpt")
```
## Testing
```
test_acc = []
with tf.Session(graph=graph) as sess:
saver.restore(sess, tf.train.latest_checkpoint('checkpoints'))
test_state = sess.run(cell.zero_state(batch_size, tf.float32))
for ii, (x, y) in enumerate(get_batches(test_x, test_y, batch_size), 1):
feed = {inputs_: x,
labels_: y[:, None],
keep_prob: 1,
initial_state: test_state}
batch_acc, test_state = sess.run([accuracy, final_state], feed_dict=feed)
test_acc.append(batch_acc)
print("Test accuracy: {:.3f}".format(np.mean(test_acc)))
```
| github_jupyter |
# Fake News Capstone Project Data Wrangling and Preprocessing
```
import s3fs
import pandas as pd
pd.set_option('display.max_columns', 100000)
pd.set_option('display.max_row', 1000000)
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import re
import tldextract
from tqdm.autonotebook import tqdm
tqdm.pandas(desc="progress-bar", leave=False)
import string
import spacy
from spacy.lang import punctuation
import unicodedata # might need to pip install unicodedate2 on aws sagemaker
import contractions
from contractions import contractions_dict
from textblob import TextBlob
import nltk
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer
from nltk import word_tokenize
from nltk.stem.wordnet import WordNetLemmatizer
import gensim
from gensim.utils import simple_preprocess
from gensim.parsing.preprocessing import preprocess_string
from gensim.parsing.preprocessing import STOPWORDS
import warnings
from afinn import Afinn
warnings.filterwarnings('ignore')
%matplotlib inline
sns.set(style='darkgrid',palette='Dark2',rc={'figure.figsize':(9,6),'figure.dpi':90})
punctuation = string.punctuation + '”' + '“' + '–' + '““' + "’’" + '”'
stopword = stopwords.words('english')
stopwords = set(STOPWORDS)
wordnet_lemmatizer = WordNetLemmatizer()
```
## Random seed
```
seed = 123
```
## Read in the dataset
```
df = pd.read_csv('FRL_Step_1_news_cleaned_2018_02_13.csv')
df.head(5)
df.info()
```
## Clean up the domain and check the unique domains
```
def extract_domain(url):
"""
Extract domain name from fld url
"""
info = tldextract.extract(url)
return info.domain
df['domain'] = df['domain'].astype(str).apply(extract_domain)
df.head(3)
```
## Cleaning up the content column by removing all the noises
```
# Portions of this are excerpts from Stack Overflow responses
def remove_special_characters(text):
"""
Removes special characters from the text document
"""
# define the pattern to keep. You can check the regex using this url https://regexr.com/
pat = r'[^a-zA-z0-9.,!?/:;\"\'\s]'
return re.sub(pat, '', text)
def remove_extra_whitespace_tabs(text):
"""
Removes extra whitespaces and remove_extra_whitespace_tabs
"""
#pattern = r'^\s+$|\s+$'
pattern = r'^\s*|\s\s*'
return re.sub(pattern, ' ', text).strip()
def remove_digits(text):
"""
Remove all digits from the text document
take string input and return a clean text without numbers.
Use regex to discard the numbers.
"""
result = ''.join(i for i in text if not i.isdigit()).lower()
return ' '.join(result.split())
def remove_newlines(text):
"""
Remove newline characters from the text document
"""
return text.replace('\\n', ' ').replace('\\r', ' ').replace('\n', ' ').replace('\r', ' ').replace('\\', ' ')
#normalize to the NFKD (Normalization Form Compatibility Decomposition) form
#that present in the Unicode standard to remain compatible with other encodings
def remove_accented_chars(text):
"""
Removes accented characters from the test
"""
new_text = unicodedata.normalize('NFKD', text).encode('ascii', 'ignore').decode('utf-8', 'ignore')
return new_text
import contractions
contractions.fix(df['content'][10])
#expands contractions found in the text
def expand_contractions(text):
#contractions_pattern = re.compile('({})'.format('|'.join(contraction_mapping.keys())), flags=re.IGNORECASE|re.DOTALL)
#def expand_match(contraction):
# match = contraction.group(0)
# first_char = match[0]
# expanded_contraction = contraction_mapping.get(match)\
# if contraction_mapping.get(match)\
# else contraction_mapping.get(match.lower())
# expanded_contraction = first_char+expanded_contraction[1:]
# return expanded_contraction
expanded_text = contractions.fix(text)
expanded_text = re.sub("'", "", expanded_text)
return expanded_text
# replace punctuation characters with spaces
def replace_punctuation(text):
filters = string.punctuation + '”' + '“' + '–'
translate_dict = dict((c, " ") for c in filters)
translate_map = str.maketrans(translate_dict)
text = text.translate(translate_map)
return text
# Remove stopwords and remove words with 2 or less characters
def stops_letters(text):
result = []
for token in gensim.utils.simple_preprocess(text):
if token not in gensim.parsing.preprocessing.STOPWORDS and len(token) > 2 and token not in stopword:
result.append(token)
return " ".join(result)
#Removes any word that starts with either http or https
def remove_urls(vTEXT):
#vTEXT = re.sub('http://\S+|https://\S+', '', vTEXT,flags=re.MULTILINE)
vTEXT = re.sub('http[s]?://\S+', '', vTEXT,flags=re.MULTILINE)
return(vTEXT)
#Remove words that starts with www
def remove_www(vTEXT):
vTEXT = re.sub('www\S+', '', vTEXT,flags=re.MULTILINE)
return(vTEXT)
```
### Covert Content and Title Fields to Lowercase
```
%%time
# Apply the functions to the dataframe
# Step 1 - convert the text to lower case
df['content']=df['content'].apply(lambda x: x.lower())
df['title']=df['title'].apply(lambda x: x.lower())
```
### Remove URLS from Content and Title Fields
```
%%time
#step 2 - Remove URLS
df['content']=df['content'].apply(remove_urls)
df['title'] = df['title'].apply(remove_urls)
```
### Remove website www from Content and Title Fields
```
%%time
#step 3 - Remove www
df['content']=df['content'].apply(remove_www)
df['title'] = df['title'].apply(remove_www)
```
### Remove special characters from Content and Title Fields
```
%%time
# Step 4 - remove special charcaters
df['content']=df['content'].apply(remove_special_characters)
df['title'] = df['title'].apply(remove_special_characters)
```
### Remove whitespace from Content and Title Fields
```
%%time
#step 5 - Remove whitespaces and tabs
df['content']=df['content'].apply(remove_extra_whitespace_tabs)
df['title'] = df['title'].apply(remove_extra_whitespace_tabs)
```
### Remove website www from Content and Title Fields
```
%%time
#step 6 - remove newlines and tabs
df['content'] = df['content'].apply(remove_newlines)
df['title'] = df['title'].apply(remove_newlines)
```
### Remove digits from Content and Title Fields
```
%%time
# step 7 - Remove digits
df['content']=df['content'].apply(remove_digits)
df['title'] = df['title'].apply(remove_digits)
```
### Remove accented characters from Content and Title Fields
```
%%time
#step 8 - remove accented characters
df['content']=df['content'].apply(remove_accented_chars)
df['title'] = df['title'].apply(remove_accented_chars)
```
### Expand Contractions within Content and Title Fields
```
%%time
#step 9 - Expand contractions
df['content']=df['content'].apply(expand_contractions)
```
### Remove punctions from Content and Title Fields, replace with single space
```
%%time
#step 10 - Replace punctuations with spaces
df['content']= df['content'].apply(replace_punctuation)
df['title'] = df['title'].apply(replace_punctuation)
```
### Remove stop letters from Content and Title Fields
```
%%time
#step 11 - Remove stopwords, tokenize and remove words with 2 or less characters
df['content']= df['content'].apply(stops_letters)
df['title'] = df['title'].apply(stops_letters)
```
## Affinity Score - Sentiment Analyis - Content Column
```
# Function to find the affinity score of a list of tweets
afinn = Afinn()
def get_affinity_scores(tweets):
scores = []
count = 0
for t in tweets:
if len(t) > 0:
scores.append(afinn.score(t) / len(t))
else:
count += 1
scores.append(0)
return scores
new_affin = get_affinity_scores(df['content'].tolist())
df['content_affin'] = new_affin
df.to_csv("./FRL_Step_2_news_cleaned_2018_02_13.csv", sep=',',index=False)
df.head()
```
## Normalization - Lemmatize the title and content columns
```
import nltk
nltk.download('punkt')
nltk.download('wordnet')
def lemmatized_word(text):
"""
lemmatize the text so as to get its root form
"""
word_tokens = nltk.word_tokenize(text)
lemmatized_word = [wordnet_lemmatizer.lemmatize(word) for word in word_tokens]
return " ".join(lemmatized_word) #combine the words into a giant string that vectorizer can accept
df['content'] = df['content'].progress_apply(lemmatized_word)
df['title'] = df['title'].progress_apply(lemmatized_word)
```
## Create additional features
```
%%time
# word counts
df['c_word_count'] = df["content"].apply(lambda x: len(str(x).split(" ")))
df['t_word_count'] = df["title"].apply(lambda x: len(str(x).split(" ")))
# Character counts
df['c_character_count'] = df["content"].apply(lambda x: sum(len(word) for word in str(x).split(" ")))
df['t_character_count'] = df["title"].apply(lambda x: sum(len(word) for word in str(x).split(" ")))
#average word length
df['c_avg_word_length'] = df['c_character_count'] / df['c_word_count']
df['t_avg_word_length'] = df['t_character_count'] / df['t_word_count']
df.to_csv("./FRL_Step_2_1_news_cleaned_2018_02_13.csv", sep=',',index=False)
```
## Identify sentiment in the title
```
# Add a new plot that shows the distribution of scores
def sentiment_check (text):
polarity_score = TextBlob(text).sentiment.polarity
df['title_sentiment_score'] = polarity_score
if polarity_score < 0:
return 'negative'
elif polarity_score == 0:
return 'neutral'
else:
return 'positive'
%%time
df['title_sentiment_label'] = df['title'].apply(sentiment_check)
df.head(3)
df.to_csv("./FRL_Step_2_full_features_news_cleaned_2018_02_13.csv", sep=',',index=False)
```
## Rename the target column
```
df = df.rename({'type': 'label'}, axis=1)
```
## Rearranged the order of the columns
```
df = df[['domain','title','content', 'content_affin','c_word_count','t_word_count','c_character_count','t_character_count','c_avg_word_length','t_avg_word_length','title_sentiment_label', 'title_sentiment_score','label']]
df.head(3)
df.info()
df.head(4)
```
## Saving the preprocessed dataframe to csv
```
# path to save the preprocessed csv file
df.to_csv("./FRL_Step_2_full_features_news_cleaned_2018_02_13.csv", sep=',',index=False)
```
| github_jupyter |
# Libraries used in this project
```
import posenet
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import cv2 as cv
import array
import os
import argparse
from sklearn.preprocessing import LabelEncoder
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import confusion_matrix, classification_report
from sklearn.metrics import accuracy_score
from sklearn.neighbors import KNeighborsClassifier
```
# Detect pose on the images
```
BODY_PARTS = { "Nose": 0, "Neck": 1, "RShoulder": 2, "RElbow": 3, "RWrist": 4,
"LShoulder": 5, "LElbow": 6, "LWrist": 7, "RHip": 8, "RKnee": 9,
"RAnkle": 10, "LHip": 11, "LKnee": 12, "LAnkle": 13, "REye": 14,
"LEye": 15, "REar": 16, "LEar": 17, "Background": 18 }
POSE_PAIRS = [ ["Neck", "RShoulder"], ["Neck", "LShoulder"], ["RShoulder", "RElbow"],
["RElbow", "RWrist"], ["LShoulder", "LElbow"], ["LElbow", "LWrist"],
["Neck", "RHip"], ["RHip", "RKnee"], ["RKnee", "RAnkle"], ["Neck", "LHip"],
["LHip", "LKnee"], ["LKnee", "LAnkle"], ["Neck", "Nose"], ["Nose", "REye"],
["REye", "REar"], ["Nose", "LEye"], ["LEye", "LEar"] ]
width = 368
height = 368
inWidth = width
inHeight = height
net = cv.dnn.readNetFromTensorflow("graph_opt.pb")
thr = 0.2
def poseDetect(frame):
frameWidth = frame.shape[1]
frameHeight = frame.shape[0]
net.setInput(cv.dnn.blobFromImage(frame, 1.0, (inWidth, inHeight), (127.5, 127.5, 127.5), swapRB=True, crop=False))
out = net.forward()
out = out[:, :19, :, :]
assert(len(BODY_PARTS) == out.shape[1])
points = []
for i in range(len(BODY_PARTS)):
heatMap = out[0, i, :, :]
_, conf, _, point = cv.minMaxLoc(heatMap)
x = (frameWidth * point[0]) / out.shape[3]
y = (frameHeight * point[1]) / out.shape[2]
points.append((int(x), int(y)) if conf > thr else None)
for pair in POSE_PAIRS:
partFrom = pair[0]
partTo = pair[1]
assert(partFrom in BODY_PARTS)
assert(partTo in BODY_PARTS)
idFrom = BODY_PARTS[partFrom]
idTo = BODY_PARTS[partTo]
if points[idFrom] and points[idTo]:
cv.line(frame, points[idFrom], points[idTo], (0, 255, 0), 3)
cv.ellipse(frame, points[idFrom], (3, 3), 0, 0, 360, (0, 0, 255), cv.FILLED)
cv.ellipse(frame, points[idTo], (3, 3), 0, 0, 360, (0, 0, 255), cv.FILLED)
t, _ = net.getPerfProfile()
return frame
```
# Here we can get the labels and features of image
```
poses = ['downdog', 'goddess', 'plank', 'tree', 'warrior2']
train_feature=[]
test_feature=[]
train_labels = []
test_labels=[]
def create_train(DIR,label,feature):
for person in poses:
path = os.path.join(DIR, person)
for img in os.listdir(path):
img_path = os.path.join(path,img)
img_array = cv.imread(img_path)
if img_array is None:
continue
else:
frame = poseDetect(img_array)
frame_new=cv.imread(img_path)
if(frame.shape[2]==frame_new.shape[2]):
frame_diff=frame-frame_new
feature.append(frame_diff)
label.append(person)
```
# For Train images
```
pose=create_train(r"C:\Users\ankit\Documents\Python Scripts\Face Recognition\Intern Project\Yoga Train Pose",train_labels,train_feature)
features_train=np.array(train_feature)
```
# For Test images
```
test_pose = create_train(r"C:\Users\ankit\Documents\Python Scripts\Face Recognition\Intern Project\Yoga Test Pose",test_labels,test_feature)
features_test=np.array(test_feature)
features_train.reshape(len(train_labels),1)
features_test.reshape(len(test_labels),1)
train_data=pd.DataFrame(features_train ,columns=['tr_feature'])
train_labels_data=pd.DataFrame(train_labels ,columns=['tr_label'])
test_data=pd.DataFrame(features_test ,columns=['te_feature'])
test_labels_data=pd.DataFrame(test_labels ,columns=['te_label'])
for i in range(len(train_data['tr_feature'])):
train_data['tr_feature'][i]=train_data['tr_feature'][i].flatten()
for i in range(len(test_data['te_feature'])):
test_data['te_feature'][i]=test_data['te_feature'][i].flatten()
```
# LabelEncoder
```
label=LabelEncoder()
labels_train=label.fit_transform(train_labels_data['tr_label'])
labels_test=label.fit_transform(test_labels_data['te_label'])
train=[]
test=[]
for i in range(len(train_data['tr_feature'])):
train.append(train_data ['tr_feature'][i].std())
for i in range(len(test_data['te_feature'])):
test.append(test_data['te_feature'][i].std())
train=np.array(train)
train=np.reshape(train,(len(labels_train),1))
test=np.array(test)
test=np.reshape(test,(len(labels_test),1))
```
# RandomForestClassifier
```
rfc=RandomForestClassifier(n_estimators=300,criterion='entropy',
max_features='sqrt',min_samples_leaf=10,random_state=100)
rfc.fit(train,labels_train)
predict=rfc.predict(test)
print(classification_report(labels_test,predict))
print(confusion_matrix(labels_test,predict))
rfc.fit(train,labels_train)
predict=rfc.predict(train)
print(classification_report(labels_train,predict))
print(confusion_matrix(labels_train,predict))
```
# KNeighborsClassifier
```
knn = KNeighborsClassifier(n_neighbors=1)
knn.fit(train,labels_train)
predict_knn=knn.predict(test)
print(classification_report(labels_test,predict_knn))
print(confusion_matrix(labels_test,predict_knn))
knn.fit(train,labels_train)
predict_rfc=knn.predict(train)
print(classification_report(labels_train,predict_rfc))
print(confusion_matrix(labels_train,predict_rfc))
```
| github_jupyter |
```
import os
from glob import glob
import pandas as pd
import numpy as np
from pathlib import Path
from scipy import stats
from matplotlib import pyplot as plt
from matplotlib.colors import Normalize
from matplotlib.backends.backend_pdf import PdfPages
from matplotlib import gridspec
import json
import torch
import gpytorch
import h5py
import collections
import scipy
import torch
import subprocess
import sys
from bnn_priors import prior
from bnn_priors.exp_utils import load_samples
from bnn_priors.notebook_utils import collect_runs, unique_cols, json_dump, json_load
%matplotlib inline
%config InlineBackend.print_figure_kwargs = {'bbox_inches':None}
```
# Load data and plot figures 3, 5 and A.10
With the repository we ship the extracted data, necessary to plot the figures. The code for *creating* the CSVs and JSONs is at the end of this notebook.
```
runs_with_eval = pd.read_csv("Plot_CIFAR10_resnet_data/sgd_runs.csv", index_col=0)
opt_dfs = json_load("Plot_CIFAR10_resnet_data/opt_dfs.json")
opt_lengthscale = json_load("Plot_CIFAR10_resnet_data/opt_lengthscale.json")
(covs, lens, conv_n_channels) = pd.read_pickle("Plot_CIFAR10_resnet_data/covs_lens.pkl.gz")
conv_keys = list(covs.keys())
conv_keys.sort(key=lambda k: (int(k.split('.')[2]), k))
fig, axes = plt.subplots(1, 2, figsize=(9, 2.3), sharex=True)
ax = axes[1]
ax.set_title("Degrees of freedom")
ax.plot([opt_dfs[k][1] for k in conv_keys])
ax.set_xticks([1, 7, 13, 18])
ax.set_xticklabels(["*L2", "*L8", "*L14", "L19"])
ax.set_yscale('log')
ax = axes[0]
ax.set_title("Lengthscale")
ax.plot([opt_lengthscale[k] for k in conv_keys])
fig.text(0.5, 0, 'Layer index (input = L0)', ha='center')
#fig.suptitle("ResNet-20, CIFAR-10 SGD: fitted parameters for T-distribution and Gaussian")
fig.tight_layout()
fig.savefig("../figures/210122_resnet_fitted.pdf", bbox_inches="tight")
# Figure 3
plt.rcParams.update({
"axes.linewidth": 0.5,
'ytick.major.width': 0.5,
'xtick.major.width': 0.5,
'ytick.minor.width': 0.5,
'xtick.minor.width': 0.5,
"figure.dpi": 300,
})
fig_width_pt = 234.8775
inches_per_pt = 1.0/72.27 # Convert pt to inches
fig_width = fig_width_pt*inches_per_pt # width in inches
fig, axes = plt.subplots(1, 1, figsize=(fig_width, 1.3), sharex=True, gridspec_kw=dict(
top=1, bottom=0.34, left=0.17, right=1))
ax = axes
ax.set_ylabel("Deg. of freedom", horizontalalignment="right", position=(0, 1))
ax.plot([opt_dfs[k][1] for k in conv_keys])
ax.set_xticks([1, 7, 13, 18])
ax.set_xticklabels(["*L2", "*L8", "*L14", "L19"])
ax.set_yscale('log')
ax.set_xlabel('Layer index (input = L0)')
fig.savefig("../figures/210126-resnet-dof.pdf")
# Figure 5
plt.rcParams.update({
"text.usetex": False,
"font.family": "sans-serif"})
fig_width_pt = 487.8225
inches_per_pt = 1.0/72.27 # Convert pt to inches
fig_width = fig_width_pt*inches_per_pt # width in inches
mean_covs = {k: covs[k]/lens[k] * conv_n_channels[k] for k in conv_keys}
plots_x = 7
plots_y = 3
margins = dict(
left=0.015,
right=0.01,
top=0.007,
bottom=0.02)
wsep = hsep = 0.002
w_cov_sep = 0.02
h_cov_sep = 0.03
height = width = (1 - w_cov_sep*(plots_x-1) - wsep*3*plots_x
- margins['left'] - margins['right'])/plots_x / 3
ttl_marg=5
fig_height_mult = (margins['bottom'] + (height*3 + hsep*2)*plots_y + h_cov_sep*plots_y + margins['top'])
# make figure rectangular and correct vertical sizes
hsep /= fig_height_mult
height /= fig_height_mult
h_cov_sep /= fig_height_mult
margins['bottom'] /= fig_height_mult
margins['top'] /= fig_height_mult
fig = plt.figure(figsize=(fig_width, fig_width *fig_height_mult))
print("fig height = ", fig_width *fig_height_mult)
cbar_height = height*3 + hsep*2
extreme = max(*(mean_covs[k].abs().max().item() for k in mean_covs.keys())) #1.68
#assert extreme < 1.7
extreme = 2
norm = Normalize(-extreme, extreme)
def plot_at(key, base_bottom, base_left, is_bottom_row=False, is_left_col=False, title="title"):
max_bottom = base_bottom
max_left = base_left
for y in range(3):
for x in range(3):
bottom = base_bottom + (height+hsep) * (2-y)
left = base_left + (width+wsep) * x
max_bottom = max(max_bottom, bottom+height+hsep)
max_left = max(max_left, left+width+wsep)
if x == 0 and is_left_col:
yticks = [1, 2, 3]
else:
yticks = []
if (y == 2 and is_bottom_row) or title=="Layer 15":
xticks = [1, 2, 3]
else:
xticks = []
ax = fig.add_axes([left, bottom, width, height], xticks=xticks, yticks=yticks)
#title=f"cov. w/ ({x + 1}, {y +1})")
extreme = 1
mappable = ax.imshow(
mean_covs[key][y*3+x, :].reshape((3, 3)) / mean_covs[key].abs().max().item() ,
cmap=plt.get_cmap('RdBu'),
extent=[0.5, 3.5, 3.5, 0.5], norm=Normalize(-extreme, extreme))
ax.plot([x+1], [y+1], marker='x', ls='none', color=('white' if title == "Layer 19" else 'white'),
ms=3, markeredgewidth=0.5)
ax.tick_params(left=False, bottom=False, labelsize="xx-small", pad=0) # remove ticks
if y==0 and x==1:
ttl = ax.set_title(title, pad=ttl_marg, size="x-small")
return max_bottom, max_left, mappable
# Iterate over the indices for axes, starting from the bottom-left of the plots
cur_bottom = margins['bottom']
for y_idx in reversed(range(0, plots_y)):
cur_left = margins['left']
for x_idx in range(min(len(conv_keys)-y_idx*plots_x, plots_x)):
key = conv_keys[y_idx*plots_x + x_idx]
if key in ['net.module.3.main.0.weight_prior.p',
'net.module.6.main.0.weight_prior.p',
'net.module.9.main.0.weight_prior.p',]:
marker = "*"
else:
marker = ""
next_bottom, cur_left, mappable = plot_at(
key, cur_bottom, cur_left,
is_bottom_row=(y_idx == plots_y-1),
is_left_col=(x_idx == 0),
title=f"{marker}Layer {conv_keys.index(key) + 1}")
cur_left += w_cov_sep
if cur_bottom == margins["bottom"]:
cbar_width = width/3
cbar_ax = fig.add_axes([cur_left, cur_bottom, cbar_width, next_bottom-cur_bottom])
fig.colorbar(mappable, cax=cbar_ax, ticks=[-extreme, -1, 0, 1, extreme])
# plot absolute variance
bottom = cur_bottom+0.02
lmarg = 2.6667*width + 2*wsep
ax = fig.add_axes([cur_left+lmarg, bottom,
1-(cur_left+lmarg + margins["right"] ), next_bottom-bottom])
ax.set_ylabel("Max. variance", size="x-small") #, horizontalalignment="right", position=(0, 1))
ax.plot([mean_covs[k].abs().max().item() for k in mean_covs.keys()])
ax.set_xticks([1, 7, 13, 18])
ax.set_xticklabels(["*L2", "*L8", "*L14", "L19"])
ax.set_ylim((0, 6))
#ax.set_yscale('log')
#ax.set_xlabel('Layer index')
ax.tick_params(labelsize="x-small")
cbar_ax.tick_params(labelsize="x-small")
cur_bottom = next_bottom + h_cov_sep
fig.savefig("../figures/210204_googleresnet_covariances_all_capped.pdf")
raise RuntimeError("Do you want to continue?")
```
# Collect SGD Runs for various data sets
Here we will only read the relevant CSV file. The cells enclosed in `if False` below were used to create it.
You need to run `jug/0_31_googleresnet_cifar10_sgd.py` to be able to run the following.
Run `eval_bnn.py` and construct the overall dataframe.
```
df = collect_runs("../logs/0_31_googleresnet_cifar10_sgd")
good_runs = df[(df["n_epochs"] == 600) & (df["status"] == "COMPLETED")]
def eval_bnn(**config):
args = [sys.executable, "eval_bnn.py", "with",
*[f"{k}={v}" for k, v in config.items()]]
print(" ".join(args))
complete = subprocess.run(args)
if complete.returncode != 0:
raise SystemError(f"Process returned with code {complete.returncode}")
#for i, (_, run) in enumerate(good_runs.iterrows()):
if False: # This would run eval_bnn.py on the relevant directory. Only needs to be run once.
print(f"run {i}/{len(good_runs)}")
config_file = str(run["the_dir"]/"config.json")
calibration_data = {
"mnist": "rotated_mnist",
"fashion_mnist": "fashion_mnist",
"cifar10": "cifar10c-gaussian_blur",
"cifar10_augmented": "cifar10c-gaussian_blur",
}[run["data"]]
eval_bnn(is_run_sgd=True, calibration_eval=True, eval_data=calibration_data,
config_file=config_file, skip_first=2, batch_size=128)
ood_data = {
"mnist": "fashion_mnist",
"fashion_mnist": "mnist",
"cifar10": "svhn",
"cifar10_augmented": "svhn",
}[run["data"]]
eval_bnn(is_run_sgd=True, ood_eval=True, eval_data=ood_data,
config_file=config_file, skip_first=2, batch_size=128)
runs_with_eval = []
for _, run in good_runs.iterrows():
corresponding = collect_runs(run["the_dir"]/"eval", metrics_must_exist=False)
new_run = [run]
for _, corr in corresponding.iterrows():
orig_keys = [k for k in corr.index if k.startswith("result.")]
if corr["calibration_eval"]:
purpose = "calibration"
assert not corr["ood_eval"]
elif corr["ood_eval"]:
purpose = "ood"
else:
raise ValueError("unknown purpose")
new_keys = [k.replace("result.", purpose+".") for k in orig_keys]
for k in new_keys:
assert k not in run.index
new_corr = corr[orig_keys]
new_corr.index = new_keys
new_run.append(new_corr)
runs_with_eval.append(pd.concat(new_run))
runs_with_eval = pd.DataFrame(runs_with_eval)
```
# Get the lengthscales and df's from each layer
```
def collect_weights(df):
samples = collections.defaultdict( lambda: [], {})
for _, row in df.iterrows():
try:
s = load_samples(row["the_dir"]/"samples.pt", idx=-1, keep_steps=False)
except pickle.UnpicklingError:
continue
assert len(samples.keys()) == 0 or set(s.keys()) == set(samples.keys())
for k in s.keys():
samples[k].append(s[k])
return {k: torch.stack(v, dim=0) for k, v in samples.items()}
samples = collect_weights(good_runs[good_runs["data"] == "cifar10_augmented"])
samples['net.module.0.weight_prior.p'].shape
for k in samples.keys():
if k.endswith(".p"):
print(k, tuple(samples[k].shape))
conv_keys = ["net.module.0.weight_prior.p", *filter(
lambda k: k.endswith(".p") and "main" in k, samples.keys())]
conv_keys.sort(key=lambda k: (int(k.split('.')[2]), k))
covs = {}
lens = {}
for k in conv_keys:
M = samples[k].view(-1, 3*3)
covs[k] = (M.t() @ M)
lens[k] = len(M)
conv_n_channels = {k: samples[k].size(-3) for k in conv_keys}
pd.to_pickle((covs, lens, conv_n_channels), "4.1_covs_lens.pkl.gz")
points = torch.from_numpy(np.mgrid[:3, :3].reshape(2, -1).T).contiguous().to(torch.float64)
import gpytorch
import math
torch.set_default_dtype(torch.float64)
kern = gpytorch.kernels.RBFKernel(batch_shape=torch.Size([1000]))
kern.lengthscale = torch.linspace(0.001**.5, 30**.5, 1000).unsqueeze(-1).pow(2)
S_inverse = kern(points).inv_matmul(torch.eye(9))
S_logdet = kern(points).logdet()
log_liks = {}
opt_lengthscale = {}
for k in covs.keys():
with torch.no_grad():
log_liks[k] = S_logdet.mul(lens[k] / -2) - 0.5 * S_inverse.mul(covs[k]).sum((-2, -1))
opt_lengthscale[k] = kern.lengthscale[torch.argmax(log_liks[k])].item()
json_dump(opt_lengthscale, "4.1_opt_lengthscale.json")
k = next(iter(log_liks.keys()))
plt.plot(kern.lengthscale.squeeze(-1).detach(), log_liks[k])
plt.ylim((-10000, 0))
plt.plot(np.arange(len(conv_keys)), [opt_lengthscale[k] for k in conv_keys])
# check that log-likelihoods aren't buggy
dist = gpytorch.distributions.MultivariateNormal(torch.zeros(9), kern[100](points))
dist.log_prob(samples[k].view(-1, 9)).sum(), log_liks[k][100] - math.log(2*math.pi) * 9 * lens[k]/2
opt_lengthscale
```
# Get max df of multivariate-T
```
class MVTFitter(torch.nn.Module):
def __init__(self, p, df, permute=None, event_dim=2):
flat_p = p.view(-1, 9)
cov = (flat_p.t() @ flat_p) / len(flat_p)
super().__init__()
self.dist = prior.MultivariateT(
p.size(), torch.zeros(9), cov.cholesky().detach().to(torch.get_default_dtype()),
df=torch.nn.Parameter(torch.tensor(df, requires_grad=True)),
event_dim=event_dim, permute=permute)
self.dist.p.requires_grad_(False)
self.dist.p[...] = p
def closure(self):
self.zero_grad()
lp = -self.dist.log_prob()
lp.backward()
return lp
opt_dfs = {}
try_df_inits = torch.linspace(math.log(2.1), math.log(1000), 300).exp()
for key in conv_keys:
max_lik = -np.inf
for permute, event_dim in [(None, 2), (None, 3), (None, 4), ((0, 2, 1, 3, 4), 3)]:
mvt = MVTFitter(samples[key], 3., permute=permute, event_dim=event_dim).cuda()
for df_init in try_df_inits:
with torch.no_grad():
mvt.dist.df[...] = df_init
lik = mvt.dist.log_prob().item()
df = mvt.dist.df.item()
if np.isnan(lik) or np.isnan(df):
print("key", key, "saw a nan with lik", lik)
if lik > max_lik:
opt_dfs[key] = (lik, df, (permute, event_dim))
max_lik = lik
json_dump(opt_dfs, "4.1_opt_dfs.json")
```
# Explore degrees of freedom of MNIST weights
You need to run `jug/0_12_mnist_no_weight_decay.py` for this.
```
#df of MVT in MNIST
mnist_weights = collections.defaultdict( lambda: [], {})
for i in range(8):
samples_file = f"../logs/sgd-no-weight-decay/mnist_classificationconvnet/{i}/samples.pt"
s = load_samples(samples_file)
for k in s.keys():
if k.endswith(".p"):
mnist_weights[k].append(s[k][-1])
mnist_weights = {k: torch.stack(v, 0) for (k, v) in mnist_weights.items()}
mnist_conv_keys = ['net.module.1.weight_prior.p', 'net.module.4.weight_prior.p', 'net.module.8.weight_prior.p']
opt_mnist_dfs = {}
try_df_inits = torch.linspace(math.log(2.1), math.log(1000), 300).exp()
for key in mnist_conv_keys:
max_lik = -np.inf
for permute, event_dim in [(None, 2), (None, 3), (None, 4), ((0, 2, 1, 3, 4), 3)]:
try:
mvt = MVTFitter(mnist_weights[key], 3., permute=permute, event_dim=event_dim).cuda()
for df_init in try_df_inits:
with torch.no_grad():
mvt.dist.df[...] = df_init
lik = mvt.dist.log_prob().item()
df = mvt.dist.df.item()
if np.isnan(lik) or np.isnan(df):
print("key", key, "saw a nan with lik", lik)
if lik > max_lik:
opt_mnist_dfs[key] = (lik, df, (permute, event_dim))
max_lik = lik
except RuntimeError as e:
dist = scipy.stats.t.fit(mnist_weights[key].numpy())
opt_mnist_dfs[key] = (None, dist[0], None)
opt_mnist_dfs
#df of MVT in MNIST
fcnn_weights = collections.defaultdict( lambda: [], {})
for i in range(10):
if i ==5 :
continue
samples_file = f"../logs/sgd-no-weight-decay/mnist_classificationdensenet/{i}/samples.pt"
s = load_samples(samples_file)
for k in s.keys():
if k.endswith("weight_prior.p"):
fcnn_weights[k].append(s[k][-1])
fcnn_weights = {k: torch.stack(v, 0) for (k, v) in fcnn_weights.items()}
{k: scipy.stats.t.fit(v)[0] for k, v in fcnn_weights.items()}
```
| github_jupyter |
```
# time series for weather
import os
import datetime
import IPython
import IPython.display
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import tensorflow as tf
mpl.rcParams['figure.figsize'] = (8, 6)
mpl.rcParams['axes.grid'] = False
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
zip_path = tf.keras.utils.get_file(
origin = 'https://storage.googleapis.com/tensorflow/tf-keras-datasets/jena_climate_2009_2016.csv.zip',
fname = 'jena_climate_2009_2016.csv.zip',
extract = True)
csv_path, _ = os.path.splitext(zip_path)
df = pd.read_csv(csv_path)
df = df[5::6]
date_time = pd.to_datetime(df.pop('Date Time'), format='%d.%m.%Y %H:%M:%S')
df.head()
plot_cols = ['T (degC)', 'p (mbar)', 'rho (g/m**3)']
plot_features = df[plot_cols]
plot_features.index = date_time
_ = plot_features.plot(subplots = True)
plot_features = df[plot_cols][:480]
plot_features.index = date_time[:480]
_ = plot_features.plot(subplots = True)
df.describe().transpose()
wv = df['wv (m/s)']
bad_wv = wv == -9999.00
wv[bad_wv] = 0.0
max_wv = df['max. wv (m/s)']
bad_max_wv = max_wv == -9999.0
max_wv[bad_max_wv] = 0.0
df['wv (m/s)'].min()
plt.hist2d(df['wd (deg)'], df['wv (m/s)'], bins=(50, 50), vmax=400)
plt.colorbar()
plt.xlabel('Wind Direction [deg]')
plt.ylabel('Wind Velocity [m/s]')
wv = df.pop('wv (m/s)')
max_wv = df.pop('max. wv (m/s)')
#Convert to radians.
wd_rad = df.pop('wd (deg)')*np.pi / 180
# Calculate the wind x and y components.
df['Wx'] = wv*np.cos(wd_rad)
df['Wy'] = wv*np.sin(wd_rad)
# Caluculate the max wind x and y components.
df['max Wx'] = max_wv*np.cos(wd_rad)
df['max Wy'] = max_wv*np.sin(wd_rad)
plt.hist2d(df['Wx'], df['Wy'], bins=(50, 50), vmax=400)
plt.colorbar()
plt.xlabel('Wind X [m/s]')
plt.ylabel('Wind Y [m/s]')
ax = plt.gca()
ax.axis('tight')
timestamp_s = date_time.map(datetime.datetime.timestamp)
day = 24*60*60
year = (365.2425) * day
df['Day sin'] = np.sin(timestamp_s * (2 * np.pi / day))
df['Day cos'] = np.cos(timestamp_s * (2 * np.pi / day))
df['Year sin'] = np.sin(timestamp_s * (2 * np.pi / day))
df['Year cos'] = np.cos(timestamp_s * (2 * np.pi / day))
plt.plot(np.array(df['Day sin'])[:25])
plt.plot(np.array(df['Day cos'])[:25])
plt.xlabel('Time [h]')
plt.title('Time of day signal')
fft = tf.signal.rfft(df['T (degC)'])
f_per_dataset = np.arange(0, len(fft))
n_samples_h = len(df['T (degC)'])
hours_per_year = 24*365.2524
years_per_dataset = n_samples_h/(hours_per_year)
f_per_year = f_per_dataset/years_per_dataset
plt.step(f_per_year, np.abs(fft))
plt.xscale('log')
plt.ylim(0, 400000)
plt.xlim([0.1, max(plt.xlim())])
plt.xticks([1, 365.2524], labels = ['1/Year', '1/day'])
_ = plt.xlabel('Frequency (log scale)')
column_indices = {name: i for i, name in enumerate(df.columns)}
n = len(df)
train_df = df[0:int(n*0.7)]
val_df = df[int(n*0.7):int(n*0.9)]
test_df = df[int(n*0.9):]
num_features = df.shape[1]
train_mean = train_df.mean()
train_std = train_df.std()
train_df = (train_df - train_mean) / train_std
val_df = (train_df - train_mean) / train_std
test_df = (train_df - train_mean) / train_std
df_std = (df - train_mean) / train_std
df_std = df_std.melt(var_name = 'Column', value_name = 'Normalized')
plt.figure(figsize = (12, 6))
ax = sns.violinplot(x='Column', y='Normalized', data=df_std)
_ = ax.set_xticklabels(df.keys(), rotation=90)
## 1. 인덱스 및 오프셋
class WindowGenerator():
def __init__(self, input_width, label_width, shift,
train_df=train_df, val_df=val_df, test_df=test_df,
label_columns=None):
# Store the raw data.
self.train_df = train_df
self.val_df = val_df
self.test_df = test_df
# Work out the label column indices.
self.label_columns = label_columns
if label_columns is not None:
self.label_columns_indices = {name: i for i, name in
enumerate(label_columns)}
self.column_indices = {name: i for i, name in
enumerate(train_df.columns)}
# Work out the window parameters.
self.input_width = input_width
self.label_width = label_width
self.shift = shift
self.total_window_size = input_width + shift
self.input_slice = slice(0, input_width)
self.input_indices = np.arange(self.total_window_size)[self.input_slice]
self.label_start = self.total_window_size - self.label_width
self.labels_slice = slice(self.label_start, None)
self.label_indices = np.arange(self.total_window_size)[self.labels_slice]
def __repr__(self):
return '\n'.join([
f'Total window size: {self.total_window_size}',
f'Input indices: {self.input_indices}',
f'Label indices: {self.label_indices}',
f'Label column name(s): {self.label_columns}'])
w1 = WindowGenerator(input_width=24, label_width=1, shift=24,
label_columns=['T (degC)'])
w1
w2 = WindowGenerator(input_width=6, label_width=1, shift=1,
label_columns=['T (degC)'])
w2
## 2. 분할
def split_window(self, features):
inputs = features[:, self.input_slice, :]
labels = features[:, self.labels_slice, :]
if self.label_columns is not None:
labels = tf.stack(
[labels[:, :, self.column_indices[name]] for name in self.label_columns],
axis=-1)
# Slicing doesn't preserve static shape information, so set the shapes
# manually. This way the `tf.data.Datasets` are easier to inspect.
inputs.set_shape([None, self.input_width, None])
labels.set_shape([None, self.label_width, None])
return inputs, labels
WindowGenerator.split_window = split_window
# Stack three slices, the length of the total window:
example_window = tf.stack([np.array(train_df[:w2.total_window_size]),
np.array(train_df[100:100+w2.total_window_size]),
np.array(train_df[200:200+w2.total_window_size])])
example_inputs, example_labels = w2.split_window(example_window)
print('All shapes are: (batch, time, features)')
print(f'Window shape: {example_window.shape}')
print(f'Inputs shape: {example_inputs.shape}')
print(f'labels shape: {example_labels.shape}')
## 3. 플롯 : 분할 창을 간단하게 시각화
w2.example = example_inputs, example_labels
def plot(self, model=None, plot_col='T (degC)', max_subplots=3):
inputs, labels = self.example
plt.figure(figsize=(12, 8))
plot_col_index = self.column_indices[plot_col]
max_n = min(max_subplots, len(inputs))
for n in range(max_n):
plt.subplot(3, 1, n+1)
plt.ylabel(f'{plot_col} [normed]')
plt.plot(self.input_indices, inputs[n, :, plot_col_index],
label='Inputs', marker='.', zorder=-10)
if self.label_columns:
label_col_index = self.label_columns_indices.get(plot_col, None)
else:
label_col_index = plot_col_index
if label_col_index is None:
continue
plt.scatter(self.label_indices, labels[n, :, label_col_index],
edgecolors='k', label='Labels', c='#2ca02c', s=64)
if model is not None:
predictions = model(inputs)
plt.scatter(self.label_indices, predictions[n, :, label_col_index],
marker='X', edgecolors='k', label='Predictions',
c='#ff7f0e', s=64)
if n == 0:
plt.legend()
plt.xlabel('Time [h]')
WindowGenerator.plot = plot
w2.plot()
## 4. tf.data.Dataset 생성
def make_dataset(self, data):
data = np.array(data, dtype=np.float32)
ds = tf.keras.preprocessing.timeseries_dataset_from_array(
data=data,
targets=None,
sequence_length=self.total_window_size,
sequence_stride=1,
shuffle=True,
batch_size=32,)
ds = ds.map(self.split_window)
return ds
WindowGenerator.make_dataset = make_dataset
@property
def train(self):
return self.make_dataset(self.train_df)
@property
def val(self):
return self.make_dataset(self.val_df)
@property
def test(self):
return self.make_dataset(self.test_df)
@property
def example(self):
"""Get and cache an example batch of 'inputs, labels' for plotting. """
result = getattr(self, '_example', None)
if result is None:
#No example batch was found, so get one from the '.train' dataset
result = next(iter(self.train))
#And cache it for net time
self._example = result
return result
WindowGenerator.train = train
WindowGenerator.val = val
WindowGenerator.test = test
WindowGenerator.example = example
# Each element is an (inputs, label) pair
w2.train.element_spec
for example_inputs, example_labels in w2.train.take(1):
print(f'Inputs shape (batch, time, features): {example_inputs.shape}')
print(f'Labels shape (batch, time, features): {example_labels.shape}')
```
# 단일 단계 모델
## 다중 출력 모델 (Multi-output models)
```
single_step_window = WindowGenerator(
# `WindowGenerator` returns all features as labels if you
# don't set the `label_columns` argument.
input_width=1, label_width=1, shift=1)
wide_window = WindowGenerator(
input_width=24, label_width=24, shift=1)
for example_inputs, example_labels in wide_window.train.take(1):
print(f'Inputs shape (batch, time, features): {example_inputs.shape}')
print(f'Labels shape (batch, time, features): {example_labels.shape}')
### Baseline
class Baseline(tf.keras.Model):
def __init__(self, label_index=None):
super().__init__()
self.label_index = label_index
def call(self, inputs):
if self.label_index is None:
return inputs
result = inputs[:, :, self.label_index]
return result[:, :, tf.newaxis]
baseline = Baseline()
baseline.compile(loss=tf.losses.MeanSquaredError(),
metrics=[tf.metrics.MeanAbsoluteError()])
val_performance = {}
performance = {}
val_performance['Baseline'] = baseline.evaluate(wide_window.val)
performance['Baseline'] = baseline.evaluate(wide_window.test, verbose=0)
wide_window.plot(baseline)
### Dense
MAX_EPOCHS = 20
def compile_and_fit(model, window, patience=2):
early_stopping = tf.keras.callbacks.EarlyStopping(monitor='val_loss',
patience=patience,
mode='min')
model.compile(loss=tf.losses.MeanSquaredError(),
optimizer=tf.optimizers.Adam(),
metrics=[tf.metrics.MeanAbsoluteError()])
history = model.fit(window.train, epochs=MAX_EPOCHS,
validation_data=window.val,
callbacks=[early_stopping])
return history
dense = tf.keras.Sequential([
tf.keras.layers.Dense(units=64, activation='relu'),
tf.keras.layers.Dense(units=64, activation='relu'),
tf.keras.layers.Dense(units=num_features)
])
history = compile_and_fit(dense, single_step_window)
IPython.display.clear_output()
val_performance['Dense'] = dense.evaluate(single_step_window.val)
performance['Dense'] = dense.evaluate(single_step_window.test, verbose=0)
wide_window.plot(dense)
### RNN
%%time
wide_window = WindowGenerator(
input_width=24, label_width=24, shift=1)
lstm_model = tf.keras.models.Sequential([
# Shape [batch, time, features] => [batch, time, lstm_units]
tf.keras.layers.LSTM(32, return_sequences=True),
# Shape => [batch, time, features]
tf.keras.layers.Dense(units=num_features)
])
history = compile_and_fit(lstm_model, wide_window)
IPython.display.clear_output()
val_performance['LSTM'] = lstm_model.evaluate( wide_window.val)
performance['LSTM'] = lstm_model.evaluate( wide_window.test, verbose=0)
print()
wide_window.plot(lstm_model)
### Advanced: Residual connections
class ResidualWrapper(tf.keras.Model):
def __init__(self, model):
super().__init__()
self.model = model
def call(self, inputs, *args, **kwargs):
delta = self.model(inputs, *args, **kwargs)
# The prediction for each timestep is the input
# from the previous time step plus the delta
# calculated by the model.
return inputs + delta
%%time
residual_lstm = ResidualWrapper(
tf.keras.Sequential([
tf.keras.layers.LSTM(32, return_sequences=True),
tf.keras.layers.Dense(
num_features,
# The predicted deltas should start small
# So initialize the output layer with zeros
kernel_initializer=tf.initializers.zeros)
]))
history = compile_and_fit(residual_lstm, wide_window)
IPython.display.clear_output()
val_performance['Residual LSTM'] = residual_lstm.evaluate(wide_window.val)
performance['Residual LSTM'] = residual_lstm.evaluate(wide_window.test, verbose=0)
print()
wide_window.plot(residual_lstm)
### Performance
x = np.arange(len(performance))
width = 0.3
metric_name = 'mean_absolute_error'
metric_index = lstm_model.metrics_names.index('mean_absolute_error')
val_mae = [v[metric_index] for v in val_performance.values()]
test_mae = [v[metric_index] for v in performance.values()]
plt.bar(x - 0.17, val_mae, width, label='Validation')
plt.bar(x + 0.17, test_mae, width, label='Test')
plt.xticks(ticks=x, labels=performance.keys(),
rotation=45)
plt.ylabel('MAE (average over all outputs)')
_ = plt.legend()
for name, value in performance.items():
print(f'{name:15s}: {value[1]:0.4f}')
```
| github_jupyter |
# The Guandu water supply system
CEDAE is the coorporation that provides drinking water and wastewater services for the Rio de Janeiro State.
They provide plenty of data regarding the quality of the water for the press and for the population, due to laws imposed by the Ministry of Health of Brazil.
Amongst the ETAs (water supply system, from Portuguese *Estação de Tratamento de Água*) managed by CEDAE is the Guandu, the largest ETA of the world, which provides drinking water for the municipalities of Nilópolis, Nova Iguaçu, Duque de Caxias, Belford Roxo, São João de Meriti, Itaguaí, Queimados and Rio de Janeiro.
First, we need to download the HTML page with the links to the PDFs.
```
import urllib.request
with urllib.request.urlopen('https://cedae.com.br/relatoriosguandu') as fp:
HTML_page = fp.read().decode() # Read from page and decode to UTF-8 string
```
Then we define an `HTMLParser` class for scrapping hyperlinks.
```
import html.parser
import urllib.parse
class HyperlinkScraper(html.parser.HTMLParser):
def __init__(self, predicate):
super().__init__()
self.link = None
self.links = dict()
self.year = None
self.in_h2 = False
self.pred = predicate
def handle_starttag(self, tag, attrs):
if tag == 'a':
for name, value in attrs:
if name == 'href':
# Heuristic: if URL has %, it must be already formatted.
# If not, format only the URL path
if '%' in value:
self.link = value
else:
urlparts = value.split('/')
filename = urllib.parse.quote(urlparts[-1])
self.link = '/'.join(urlparts[:-1] + [filename])
elif tag == 'h2':
self.in_h2 = True
def handle_data(self, data):
if self.in_h2:
# When in h2, search for a year
for word in data.split():
if word.isdigit():
number = int(word)
# CEDAE was created in 1975
# 2100 seems a good upper bound here
if 1975 <= number <= 2100:
self.year = number
break
if self.link and self.year and self.pred(data):
# If inside an <a> tag, and any link was found,
# and an year was found and the predicate is true,
# then add the links to the set of links of that year
if self.year not in self.links:
self.links[self.year] = set()
self.links[self.year].add(self.link)
def handle_endtag(self, tag):
if tag == 'a':
self.link = None
elif tag == 'h2':
self.in_h2 = False
```
Let's also make sure that the output directories are created.
```
import os
def mkdirf(directory):
try:
os.mkdir(directory)
print(f'{directory} created')
except FileExistsError:
print(f'{directory} already existed')
mkdirf('output')
mkdirf(os.path.join('output', 'guandu'))
```
# Geosmin / MIB (per ETA)
We then feed the parser with the HTML page contents and get the scrapped links. We are first interested in data relating to geosmin.
```
def geosmin_predicate(data):
return 'GEOSMINA' in data.upper()
geosmin_scraper = HyperlinkScraper(geosmin_predicate)
geosmin_scraper.feed(HTML_page)
```
Finally, we download the PDFs and convert them to pandas DataFrames.
```
import urllib.parse
import tabula
geosmin_dfs = {}
for year, links in geosmin_scraper.links.items():
assert len(links) == 1, "Assumed there is 1 report per year"
link = next(iter(links))
with urllib.request.urlopen(link) as fp:
geosmin_dfs[year] = tabula.read_pdf(fp,
pages='all',
silent=True)
```
If you were to look at the PDFs in the `data` directory, you'd see that the tables have a merged cell in the header with the text "CONCENTRAÇÃO TOTAL - GEOSMINA / MIB". Tabula simply assumes this is indeed a column. The consequence is that the subcolumns are all crammed together.
```
geosmin_dfs[2021][0][:10]
```
One important discontinuity in the data is that before 18/03/2020, the concentration of geosmin and MIB of both ETAs (new and old, VETA and NETA) were averaged. After these date, these concentrations were measured and exposed separetely, since they are independent.
A solution to this problem is to create four time series: before capture, VETA, NETA and the medium between VETA and NETA. For measurements later than 18/03/2020, calculating the weighted mean of the VETA and NETA values requires a missing data: the flow rate of each ETA.
Also, we remove the header of these tables.
```
from datetime import datetime
headerless_geosmin_dfs = {}
date_format = '%d/%m/%Y'
# Find the first row where the the value in the first
# column (date) is a valid date in the format above
# and remove any rows before it
for year, df_list in geosmin_dfs.items():
headerless_geosmin_dfs[year] = []
for df in df_list:
assert len(df.columns) == 2, "Assumed data frames have 2 columns initially"
new_df = df.copy()
new_df.columns = ['date', 'params']
for row_idx, row in new_df.iterrows():
try:
date = datetime.strptime(str(row.date), date_format)
except ValueError:
pass
else:
headerless_geosmin_dfs[year].append(new_df[row_idx:])
break
headerless_geosmin_dfs[2021][2][:10]
```
Then, we split the "params" column into "capture", "veta" and "neta" for measurements after 18/03/2020, and into "capture", "etas" before this date.
```
separated_geosmin_dfs = {}
for year, df_list in headerless_geosmin_dfs.items():
separated_geosmin_dfs[year] = []
for df_index, df in enumerate(df_list):
new_df = df.copy()
if year == 2020 and df_index == 0:
columns = ['capture_geosmin', 'etas_geosmin']
else:
columns = ['capture_geosmin', 'veta_geosmin', 'neta_geosmin']
new_df[columns] = new_df['params'].str.split(n=len(columns)-1, expand=True)
del new_df['params']
separated_geosmin_dfs[year].append(new_df)
separated_geosmin_dfs[2020][1][:10]
```
Now, we convert the strings into numbers and dates.
We see that values lower than 0.010 micrograms per liter are supressed. In order to keep that information, we'll store the upper bound. This solution is better than cramming all uncertainties to zero because throughout the time series there are different upper bounds and cramming them all to zero would mean in loss of information. Additionally, for each data column, a column will be created to indicate whether the value is uncertain or not.
```
import pandas as pd
formatted_geosmin_dfs = {}
for year, df_list in separated_geosmin_dfs.items():
formatted_geosmin_dfs[year] = []
for df_index, df in enumerate(df_list):
new_df = df.copy()
if year == 2020 and df_index == 0:
columns = ['capture_geosmin', 'etas_geosmin']
else:
columns = ['capture_geosmin', 'veta_geosmin', 'neta_geosmin']
new_df.date = pd.to_datetime(new_df.date, dayfirst=True)
for column in columns:
new_df[f'is_{column}_ub'] = df[column].str.contains('<')
new_df[column] = pd.to_numeric(df[column]
.str.replace(' ', '') # any space is removed
.str.replace('<', '') # supress uncertainties
.str.replace(',', '.')) # use . as decimal delimiter
formatted_geosmin_dfs[year].append(new_df)
formatted_geosmin_dfs[2021][0][:10]
```
We then concatenate the data frames with same columns.
```
geosmin_df1 = None
geosmin_df2 = []
for year, df_list in sorted(formatted_geosmin_dfs.items()):
for df_index, df in enumerate(df_list):
if year == 2020 and df_index == 0:
geosmin_df1 = df
else:
geosmin_df2.append(df)
geosmin_df2 = pd.concat(geosmin_df2, ignore_index=True)
```
Now that the data is nicely formatted, we can plot them with `matplotlib`
```
%matplotlib inline
import matplotlib.pyplot as plt
```
First, the data before 18/03/2020.
```
plt.plot(geosmin_df1.date, geosmin_df1.capture, label='Captação')
plt.plot(geosmin_df1.date, geosmin_df1.etas, label='VETA+NETA')
plt.xlabel('Tempo')
plt.ylabel('Concentração de Geosmina / MIB (10^-6 g/L)')
plt.title('Concentração de geosmina em 2020')
plt.legend()
plt.show()
```
And then, after this date.
```
plt.plot(geosmin_df2.date, geosmin_df2.capture, label='Captação')
plt.plot(geosmin_df2.date, geosmin_df2.veta, label='VETA')
plt.plot(geosmin_df2.date, geosmin_df2.neta, label='NETA')
plt.xlabel('Tempo')
plt.ylabel('Concentração de Geosmina / MIB (10^-6 g/L)')
plt.title('Concentração de geosmina em 2020/2021')
plt.legend()
plt.show()
```
We then export this data in the CSV format.
```
geosmin_df1.to_csv(os.path.join('output', 'guandu', 'geosmin_2020-01-26_2020-03-18.csv'), index=False)
geosmin_df2.to_csv(os.path.join('output', 'guandu', 'geosmin_2020-10-23_today.csv'), index=False)
```
# Taste and Odor (per ETA)
We are now interested in obtaining data about taste and odor (`tno`) of the water in the VETA and NETA water treatment stations. We'll be reusing the HTML data in order to extract the hyperlinks to these PDFs.
```
def tno_predicate(data):
data = data.upper()
return 'GOSTO E ODOR' in data and 'SAÍDA DE TRATAMENTO' in data
tno_scraper = HyperlinkScraper(tno_predicate)
tno_scraper.feed(HTML_page)
```
We now download the PDFs just like we did before...
```
tno_dfs = {}
for year, links in tno_scraper.links.items():
assert len(links) == 1, "Assumed there is 1 report per year"
link = next(iter(links))
with urllib.request.urlopen(link) as fp:
tno_dfs[year] = tabula.read_pdf(fp,
pages='all',
silent=True)
```
Let's take a peek at these data frames...
```
tno_dfs[2021][0][:10]
```
We see that the merged cells confound the tabula parser just like with the geosmin case. Thus, we employ a similar heuristic of removing the first rows whose values in the first column aren't dates.
```
filtered_tno_dfs = {}
for year, df_list in tno_dfs.items():
filtered_tno_dfs[year] = []
for df_idx, df in enumerate(df_list):
if len(df.columns) != 2:
print('Ignored table #%d from year #%d:' % (df_idx, year))
print(df.head())
print()
continue
new_df = df.copy()
new_df.columns = ['date', 'params']
not_dates = []
for row_idx, row in new_df.iterrows():
try:
date = datetime.strptime(str(row.date), date_format)
except ValueError:
not_dates.append(row_idx)
filtered_tno_dfs[year].append(new_df.drop(not_dates))
filtered_tno_dfs[2021][0][:10]
```
Now we split the params columns by separator.
```
columns = ['neta_taste', 'neta_odor', 'veta_taste', 'veta_odor']
split_tno_dfs = {}
for year, df_list in filtered_tno_dfs.items():
split_tno_dfs[year] = []
for df_index, df in enumerate(df_list):
new_df = df.copy()
new_df[columns] = df['params'].str.split(n=len(columns)-1, expand=True)
del new_df['params']
split_tno_dfs[year].append(new_df)
split_tno_dfs[2021][0][:10]
```
Now we format the values in a standard form.
```
import pandas as pd
formatted_tno_dfs = {}
for year, df_list in split_tno_dfs.items():
formatted_tno_dfs[year] = []
for df_index, df in enumerate(df_list):
new_df = df.copy()
new_df.date = pd.to_datetime(df.date, dayfirst=True)
for column in columns:
new_df[f'is_{column}_ub'] = df[column].str.contains('<')
new_df[column] = pd.to_numeric(df[column]
.str.replace(' ', '') # any space is removed
.str.replace('<', '')) # supress uncertainties
formatted_tno_dfs[year].append(new_df)
formatted_tno_dfs[2021][0][:10]
```
Finally, we concatenate all data frames into one. We are able to do this because the data is uniform.
```
tno_df = []
for year, df_list in sorted(formatted_tno_dfs.items()):
tno_df += df_list
tno_df = pd.concat(tno_df, ignore_index=True)
```
We now plot the time series for taste and odor separately, since the grading may not be in the same scale.
```
plt.plot(tno_df.date, tno_df.veta_taste, label='VETA')
plt.plot(tno_df.date, tno_df.neta_taste, label='NETA')
plt.xlabel('Tempo')
plt.ylabel('Gosto')
plt.title('Gosto da água das ETAs do Guandu')
plt.legend()
plt.show()
plt.plot(tno_df.date, tno_df.veta_odor, label='VETA')
plt.plot(tno_df.date, tno_df.neta_odor, label='NETA')
plt.xlabel('Tempo')
plt.ylabel('Odor')
plt.title('Odor da água das ETAs do Guandu')
plt.legend()
plt.show()
```
Like before, we export this data in the CSV format.
```
tno_df.to_csv(os.path.join('output', 'guandu', 'taste_and_odor.csv'), index=False)
```
## Taste and Odor (in the distribution network)
CEDAE also publishes data regarding taste and odor of the water in the distribution network. The data is less frequently collected than those in the ETAs, but provides more insight about the quality of the water at specific neighbourhoods in Rio.
We now search for "REDE DE DISTRIBUIÇÃO" (distribution network) instead of "SAÍDA DE TRATAMENTO" (treatment output).
```
def tno_net_predicate(data):
data = data.upper()
return 'GOSTO E ODOR' in data and 'REDE DE DISTRIBUIÇÃO' in data
tno_net_scraper = HyperlinkScraper(tno_net_predicate)
tno_net_scraper.feed(HTML_page)
```
We download the PDFs whose titles have such text. Since the PDF from 2020 is really ill-formed, we decided to manually scrape it with Tabula and store the well-formed data in a CSV. :)
```
import pandas as pd
tno_net_dfs = {}
tno_net_files = {2020: os.path.join('input', 'guandu', 'tno_net_2020.csv')}
for year, links in tno_net_scraper.links.items():
file = tno_net_files.get(year)
if file is not None:
tno_net_dfs[year] = [pd.read_csv(file)]
else:
assert len(links) == 1, "Assumed there is 1 report per year"
link = next(iter(links))
with urllib.request.urlopen(link) as fp:
tno_net_dfs[year] = tabula.read_pdf(fp,
pages='all',
lattice=True,
silent=True)
```
Let's take a peek at the data frames obtained from the PDF...
```
tno_net_dfs[2021][0][:10]
```
And the data frame obtained from the CSV...
```
tno_net_dfs[2020][0][:10]
```
We first name the columns.
```
for year, df_list in tno_net_dfs.items():
for df in df_list:
assert len(df.columns) == 8, "Assumed data frames have 8 columns"
df.columns = ['date', 'code', 'address', 'neighbourhood',
'municipality', 'collection-point', 'taste', 'odor']
```
We notice that some cells contain the carriage return character `\r` and the new line character `\n`. We decide to substitute this character with a space.
```
for year, df_list in tno_net_dfs.items():
for df in df_list:
df.replace(['\\r', '\\n'], ' ', inplace=True, regex=True)
```
Now, we remove the rows whose date is not a date. This accounts for random rows with titles and not data.
```
filtered_tno_net_dfs = {}
for year, df_list in tno_net_dfs.items():
filtered_tno_net_dfs[year] = []
for df in df_list:
not_dates = []
for row_idx, row in df.iterrows():
try:
date = datetime.strptime(str(row.date), date_format)
except ValueError:
not_dates.append(row_idx)
filtered_tno_net_dfs[year].append(df.drop(not_dates))
```
We now convert the columns to their well-behaved types.
```
converted_tno_net_dfs = {}
for year, df_list in filtered_tno_net_dfs.items():
converted_tno_net_dfs[year] = []
for df in df_list:
new_df = df.copy()
new_df.date = pd.to_datetime(new_df.date, dayfirst=True)
for column in ['taste', 'odor']:
new_df[column] = pd.to_numeric(new_df[column]
.replace(' ', ''))
converted_tno_net_dfs[year].append(new_df)
```
We then concatenate the data, which should be now uniform.
```
tno_net_df_list = []
for year, df_list in sorted(converted_tno_net_dfs.items()):
tno_net_df_list += df_list
tno_net_df = pd.concat(tno_net_df_list, ignore_index=True)
```
We can plot now the data, not considering the location of the collection point, comparing with the ETA.
```
plt.plot(tno_net_df.date, tno_net_df.taste, label='Rede de Distribuição')
plt.xlabel('Tempo')
plt.ylabel('Gosto')
plt.title('Gosto da água do Guandu nas ETAs e na rede de distribuição')
plt.legend()
plt.show()
```
And the same for odor.
```
plt.plot(tno_net_df.date, tno_net_df.odor, label='Rede de Distribuição')
plt.xlabel('Tempo')
plt.ylabel('Odor')
plt.title('Odor da água do Guandu nas ETAs e na rede de distribuição')
plt.legend()
plt.show()
```
Like before, we export this data in the CSV format.
```
tno_net_df.to_csv(os.path.join('output', 'guandu', 'taste_and_odor_net.csv'), index=False)
```
| github_jupyter |
## Midterm test and practice session
### 1. Questions.
Please, answer the following questions briefly. Two or three sentences with main idea would be enough.
Do not use external resourses in this part, please. Answer with you own words. If you forgot something, don't worry, we will discuss it later.
#### 1.0.
Please, formulate the supervised learning problem statement.
_Your answer here_
#### 1.1.
What are regression and classification problems. What’s the difference?
_Your answer here_
#### 1.2.
Write down the linear model for regression problem in matrix notation. What is Mean Squared Error (MSE) loss function? How can it be expressed?
_Your answer here_
#### 1.3.
What is the gradient of a function? How is it being used in optimization?
_Your answer here_
#### 1.4.
Write down gradient descent step for linear model and MSE for one-dimensional case.
_Your answer here_
#### 1.5.
What is validation? Cross validation?
_Your answer here_
#### 1.6.
What is regularization? How does L1 regularization differ from L2 for linear models?
_Your answer here_
#### 1.7.
What are precision and recall metrics?
_Your answer here_
#### 1.8.
What is bagging? What is the main idea beneath it?
_Your answer here_
### 2. Tackling Machine Learning problems in the wild
Now you will work with real data in classification problem. Your goal it to solve (with some quality) and make some conclusions. It's quite similar to the `assignment0_02`.
You may use external resources here.
#### 2.0 Reading the data
Today we work with the [dataset](https://archive.ics.uci.edu/ml/datasets/Statlog+%28Vehicle+Silhouettes%29), describing different cars for multiclass ($k=4$) classification problem. The data is available below.
```
# If on colab, uncomment the following lines
# !wget https://raw.githubusercontent.com/neychev/harbour_ml2020/master/assignments/assignment_Midterm/car_data.csv
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
dataset = pd.read_csv('car_data.csv', delimiter=',', header=None).values
data = dataset[:, :-1].astype(int)
target = dataset[:, -1]
print(data.shape, target.shape)
X_train, X_test, y_train, y_test = train_test_split(data, target, test_size=0.35)
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape)
```
To get some insights about the dataset, `pandas` might be used. The `train` part is transformed to `pd.DataFrame` below.
```
X_train_pd = pd.DataFrame(X_train)
# First 15 rows of our dataset.
X_train_pd.head(15)
```
Methods `describe` and `info` deliver some useful information.
```
X_train_pd.describe()
np.unique(y_train)
X_train_pd.info()
```
#### 2.1. Data preprocessing
* Make some transformations of the dataset (if necessary). Briefly explain the transformations
```
from sklearn.preprocessing import StandardScaler
### YOUR CODE HERE
```
#### 2.2. PCA: explained variance plot
* Apply the PCA to the train part of the data. Build the explaided variance plot (like in assignment 2).
```
from sklearn.decomposition import PCA
### YOUR CODE HERE
```
#### 2.3. PCA trasformation
* Select the appropriate number of components. Use `fit` and `transform` methods to transform the `train` and `test` parts.
```
### YOUR CODE HERE
```
#### 2.4. Logistic regression
* Find the optimal hyperparameters for logistic regression using cross-validation (e.g. `GridSearchCV`). You can vary only a parameter for `l2` regularization (e.g 5 different values).
* Then build a ROC curve for this classifier (`sklearn.metrics.roc_curve`). Estimate the model quality with appropriate metrics (which will you use?)
_Note: be careful with preprocessing (like scaling/PCA) to avoid data leaks._
```
from sklearn.linear_model import LogisticRegression
### YOUR CODE HERE
```
#### 2.5. Decision tree
* Now train a desicion tree on the same data. Find optimal tree depth (`max_depth`) using cross-validation (again, checking 5 variants would be fine).
* Measure the model quality using the same metrics you used above.
```
from sklearn.tree import DecisionTreeClassifier
# YOUR CODE HERE
```
#### 2.6. Random Forest
Now we will work with the Random Forest (its `sklearn` implementation).
* Vary the number of trees (from 5 to 50 with step 5) and build the plot in axes "model accuracy" - "number of trees".
* What is the optimal number of trees you've got?
```
from sklearn.ensemble import RandomForestClassifier
# YOUR CODE HERE
```
#### 2.7. Bonus part: "learning curve"
* Split the training data into 10 equal (almost) parts. Then train the models from above (Logistic regression, Desicion Tree, Random Forest) with optimal hyperparameters you have selected on 1 part, 2 parts (combined, so the train size in increased by 2 times), 3 parts and so on.
* Build a plot of accuracy and f1-score (on `test` part) varying the `train` dataset size (so the axes will be metric - dataset size.
* Analyse the final plot. Can you make any conlusions using it?
```
# YOUR CODE HERE
```
| github_jupyter |
```
# Import Libraries
import requests
from requests.adapters import HTTPAdapter
from requests.packages.urllib3.util.retry import Retry
from bs4 import BeautifulSoup
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import string
import time
import re
import os
from functools import reduce
trim = re.compile(r'[^\d.,]+')
# http://www.zipcodestogo.com/New%20York/
# http://www.zipcodestogo.com/New%20Jersey/
```
### Read in NJ Zip Codes
Source: http://www.zipcodestogo.com/New%20Jersey/
```
zipcodes = pd.read_csv("/Users/erikgregorywebb/Documents/Python/nyc-housing/Data/nj-zip-codes.csv")
zipcodes.tail()
```
### Generate Craigslist Links
```
base_links = []
for i in range(0, len(zipcodes)):
link = "https://newjersey.craigslist.org/search/apa?postal={}".format(zipcodes.iloc[i,2])
base_links.append(link)
base_links[0:5]
len(base_links)
```
### Generate Craigslist Links [2nd Method]
```
links = []
for i in range(0, 3000, 120):
link = "https://newjersey.craigslist.org/search/apa?s={}".format(i)
links.append(link)
links[0:5]
base_links = links
```
### Collect Listings Data
```
start_time = time.time()
sleep_time = 1
housing = pd.DataFrame()
length = len(base_links)
#length = 1
print("--- Expected Run Time: % seconds ---" % (length*sleep_time))
for i in range(0, length):
time.sleep(sleep_time)
s = requests.Session()
retries = Retry(total=3, backoff_factor=0.5)
s.mount('http://', HTTPAdapter(max_retries=retries))
r = s.get(base_links[i])
page = BeautifulSoup(r.content, "lxml")
dates = []
prices = []
bedrooms = []
titles = []
locations = []
links = []
rows = page.findAll('li', {'class': 'result-row'})
for row in rows:
# Date
try: date = row.find('time', {'class': 'result-date'})['datetime']
except: date = None
# Price
try: price = row.find('span', {'class': 'result-price'}).text
except: price = None
# Bedroom
try: bedroom = row.find('span', {'class': 'housing'}).text
except: bedroom = None
# Title
try: title = row.find('a', {'class': 'result-title hdrlnk'}).text
except: title = None
# Location
try: location = row.find('span', {'class': 'result-hood'}).text
except: location = None
# Link
try: link = row.find('a', href = True)['href']
except: link = None
dates.append(date)
prices.append(price)
bedrooms.append(bedroom)
titles.append(title)
locations.append(location)
links.append(link)
county = [zipcodes.iloc[i,0]] * len(dates)
city = [zipcodes.iloc[i,1]] * len(dates)
zipcode = [zipcodes.iloc[i,2]] * len(dates)
temp = pd.DataFrame(list(zip(county, city, zipcode, dates, prices, bedrooms, titles, locations, links)))
housing = pd.concat([housing, temp])
print("--- %s seconds ---" % (time.time() - start_time))
print("--- %s seconds NOT sleeping ---" % (time.time() - start_time - (sleep_time * length)))
# Clean the data
housing.columns = ["County", "City", "Zipcode", "Date", "Price", "Bedrooms", "Title", "Location", "Link"]
for i in range(0, len(housing)):
try: housing.iloc[i,4] = housing.iloc[i,4].replace('$', '')
except: housing.iloc[i,4] = housing.iloc[i,4]
try: housing.iloc[i,5] = housing.iloc[i,5].replace('\n', '')
except: housing.iloc[i,5] = housing.iloc[i,5]
try: housing.iloc[i,5] = housing.iloc[i,5].replace('-', '')
except: housing.iloc[i,5] = housing.iloc[i,5]
try: housing.iloc[i,5] = housing.iloc[i,5].strip()
except: housing.iloc[i,5] = housing.iloc[i,5]
try:
if housing.iloc[i,5].find('br') == True:
housing.iloc[i,5] = housing.iloc[i,5][0:3]
else:
housing.iloc[i,5] = None
except: None
try: housing.iloc[i,7] = housing.iloc[i,7].replace('(', '')
except: housing.iloc[i,7] = housing.iloc[i,7]
try: housing.iloc[i,7] = housing.iloc[i,7].replace(')', '')
except: housing.iloc[i,7] = housing.iloc[i,7]
# Remove Duplictates
housing = housing.drop_duplicates(subset = ['County', 'City', 'Zipcode', 'Price', 'Bedrooms', 'Location', 'Title'], keep = 'first')
housing.tail()
```
### Export the Data
```
os.chdir("/Users/erikgregorywebb/Documents/Python/nyc-housing/Data")
housing.to_csv("nj-housing-2.csv")
```
### Other Ideas
Links:
- http://www.areavibes.com/
- http://www.city-data.com/zips/10024.html
- https://www.melissadata.com/lookups/index.htm
- http://www.relocationessentials.com/aff/www/tools/community/index.aspx
Variables:
- Estimated zip code population
| github_jupyter |
# RAPA MVP Description
With RAPA, we plan to provide a robust, freely usable, and shareable tool for automated parsimony analysis.
RAPA will initially be developed on top of DataRobot’s Python API to use DataRobot as a "model-running engine." In the RAPA MVP, we will provide two primary features:
* Initial feature filtering to reduce a feature list down to a size that DataRobot can receive as input.
* Automated parsimony analysis to present to the user the trade-off between the size of Feature List and the best model performance on each Feature List, presented as a Pareto front.
Although the MVP implementation of these features will be based on basic techniques such as linear feature filters and recursive feature elimination, we plan to rapidly improve these features by integrating state-of-the-art techniques from the academic literature.
DataRobot API reference <a href="https://datarobot-public-api-client.readthedocs-hosted.com/en/v2.23.0/autodoc/api_reference.html">here</a>.
**The implementation below represents a quick first prototype that implements the core RAPA features.**
```
%matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sb
import numpy as np
import pandas as pd
import pickle
import time
from collections import defaultdict
from tqdm.notebook import tqdm
from sklearn.feature_selection import f_regression, f_classif
from sklearn.model_selection import StratifiedKFold, KFold
import datarobot as dr
pd.set_option('mode.chained_assignment', None)
local_path = "../data/"
datarobot_tokens = pickle.load(open(f'{local_path}dr-tokens.pkl', 'rb'))
dr.Client(endpoint='https://app.datarobot.com/api/v2',
token=datarobot_tokens['JOSH'])
print('DataRobot API initiated')
def create_datarobot_classification_submission(input_data_df, target_name, max_features=19990, n_splits=6,
filter_function=f_classif, random_state=None):
"""Prepares the input data for submission as a classification project on DataRobot.
Creates pre-determined k-fold cross-validation splits and filters the feature
set down to a size that DataRobot can receive as input, if necessary.
Parameters
----------
input_data_df: pandas.DataFrame
pandas DataFrame containing the feature set and prediction target.
target_name: str
Name of the prediction target column in `input_data_df`.
max_features: int, optional (default: 19990)
The number of features to reduce the feature set in `input_data_df`
down to. DataRobot's maximum feature set size is 20,000.
n_splits: int, optional (default: 6)
The number of stratified cross-validation splits to create. One of the splits
will be retained as a holdout split, so by default this function
sets up the dataset for 5-fold cross-validation with a holdout.
filter_function: callable, optional (default: sklearn.feature_selection.f_classif)
The function used to calculate the importance of each feature in
the initial filtering step that reduces the feature set down to
`max_features`.
This filter function must take a feature matrix as the first input
and the target array as the second input, then return two separate
arrays containing the feature importance of each feature and the
P-value for that correlation, in that order.
See scikit-learn's f_classif function for an example:
https://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.f_classif.html
random_state: int, optional (default: None)
The random number generator seed for RAPA. Use this parameter to make sure
that RAPA will give you the same results each time you run it on the
same input data set with that seed.
"""
input_data_df = input_data_df.copy()
only_features_df = input_data_df.drop(columns=[target_name])
kfold_type = StratifiedKFold
if len(np.unique(input_data_df[target_name].values)) == 2:
target_type = dr.enums.TARGET_TYPE.BINARY
else:
target_type = dr.enums.TARGET_TYPE.MULTICLASS
input_data_df['partition'] = 'train'
train_feature_importances = []
for fold_num, (_, fold_indices) in enumerate(
kfold_type(n_splits=n_splits, random_state=random_state, shuffle=True).split(only_features_df.values,
input_data_df[target_name].values)):
input_data_df['partition'].iloc[fold_indices] = 'CV Fold {}'.format(fold_num)
# Fold 0 is the holdout set, so don't calculate feature importances using that fold
if fold_num > 0:
feature_importances, _ = filter_function(only_features_df.iloc[fold_indices].values,
input_data_df[target_name].iloc[fold_indices].values)
train_feature_importances.append(feature_importances)
# We calculate the overall feature importance scores by averaging the feature importance scores
# across all of the training folds
avg_train_feature_importances = np.mean(train_feature_importances, axis=0)
input_data_df.loc[input_data_df['partition'] == 'CV Fold 0', 'partition'] = 'Holdout'
most_correlated_features = only_features_df.columns.values[np.argsort(avg_train_feature_importances)[::-1][:max_features]].tolist()
datarobot_upload_df = input_data_df[[target_name, 'partition'] + most_correlated_features]
return datarobot_upload_df, target_type
def create_datarobot_regression_submission(input_data_df, target_name, max_features=19900, n_splits=6,
filter_function=f_regression, random_state=None):
"""Prepares the input data for submission as a regression project on DataRobot.
Creates pre-determined k-fold cross-validation splits and filters the feature
set down to a size that DataRobot can receive as input, if necessary.
Parameters
----------
input_data_df: pandas.DataFrame
pandas DataFrame containing the feature set and prediction target.
target_name: str
Name of the prediction target column in `input_data_df`.
max_features: int, optional (default: 19990)
The number of features to reduce the feature set in `input_data_df`
down to. DataRobot's maximum feature set size is 20,000.
n_splits: int, optional (default: 6)
The number of cross-validation splits to create. One of the splits
will be retained as a holdout split, so by default this function
sets up the dataset for 5-fold cross-validation with a holdout.
filter_function: callable, optional (default: sklearn.feature_selection.f_regression)
The function used to calculate the importance of each feature in
the initial filtering step that reduces the feature set down to
`max_features`.
This filter function must take a feature matrix as the first input
and the target array as the second input, then return two separate
arrays containing the feature importance of each feature and the
P-value for that correlation, in that order.
See scikit-learn's f_classif function for an example:
https://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.f_regression.html
random_state: int, optional (default: None)
The random number generator seed for RAPA. Use this parameter to make sure
that RAPA will give you the same results each time you run it on the
same input data set with that seed.
"""
input_data_df = input_data_df.copy()
only_features_df = input_data_df.drop(columns=[target_name])
kfold_type = KFold
target_type = dr.enums.TARGET_TYPE.REGRESSION
input_data_df['partition'] = 'train'
train_feature_importances = []
for fold_num, (_, fold_indices) in enumerate(
kfold_type(n_splits=n_splits, random_state=random_state, shuffle=True).split(only_features_df.values,
input_data_df[target_name].values)):
input_data_df['partition'].iloc[fold_indices] = 'CV Fold {}'.format(fold_num)
# Fold 0 is the holdout set, so don't calculate feature importances using that fold
if fold_num > 0:
feature_importances, _ = filter_function(only_features_df.iloc[fold_indices].values,
input_data_df[target_name].iloc[fold_indices].values)
train_feature_importances.append(feature_importances)
# We calculate the overall feature importance scores by averaging the feature importance scores
# across all of the training folds
avg_train_feature_importances = np.mean(train_feature_importances, axis=0)
input_data_df.loc[input_data_df['partition'] == 'CV Fold 0', 'partition'] = 'Holdout'
most_correlated_features = only_features_df.columns.values[np.argsort(avg_train_feature_importances)[::-1][:max_features]].tolist()
datarobot_upload_df = input_data_df[[target_name, 'partition'] + most_correlated_features]
return datarobot_upload_df, target_type
def submit_datarobot_project(input_data_df, target_name, target_type, project_name,
worker_count=-1, mode=dr.AUTOPILOT_MODE.FULL_AUTO,
random_state=None):
"""Submits the input data to DataRobot as a new modeling project.
It is suggested to prepare the `input_data_df` using the
`create_datarobot_classification_submission` or
`create_datarobot_regression_submission` functions first.
Parameters
----------
input_data_df: pandas.DataFrame
pandas DataFrame containing the feature set and prediction target.
target_name: str
Name of the prediction target column in `input_data_df`.
target_type: enum
Indicator to DataRobot of whether the new modeling project should be
a binary classification, multiclass classification, or regression project.
Options:
datarobot.TARGET_TYPE.BINARY
datarobot.TARGET_TYPE.REGRESSION
datarobot.TARGET_TYPE.MULTICLASS
project_name: str
Name of the project in DataRobot.
worker_count: int, optional (default: -1)
The number of worker engines to assign to the DataRobot project.
By default, -1 tells DataRobot to use all available worker engines.
mode: enum, optional (default: datarobot.AUTOPILOT_MODE.FULL_AUTO)
The modeling mode to start the DataRobot project in.
Options:
datarobot.AUTOPILOT_MODE.FULL_AUTO
datarobot.AUTOPILOT_MODE.QUICK
datarobot.AUTOPILOT_MODE.MANUAL
datarobot.AUTOPILOT_MODE.COMPREHENSIVE: Runs all blueprints in
the repository (warning: this may be extremely slow).
random_state: int, optional (default: None)
The random number generator seed for DataRobot. Use this parameter to make sure
that DataRobot will give you the same results each time you run it on the
same input data set with that seed.
"""
project = dr.Project.create(sourcedata=input_data_df, project_name=project_name)
project.set_target(target=target_name, target_type=target_type,
worker_count=worker_count, mode=mode,
advanced_options=dr.AdvancedOptions(seed=random_state, accuracy_optimized_mb=False,
prepare_model_for_deployment=False, blend_best_models=False),
partitioning_method=dr.UserCV(user_partition_col='partition', cv_holdout_level='Holdout'))
return project
```
## Demo: RAPA with a dataset using DNA methylation to predict whether someone has been diagnosed with depression
```
depression_targets = pd.read_pickle(f'{local_path}GSE128235_GPL13534_meta_data.pkl')
print(depression_targets.shape)
print('')
print(depression_targets['diagnosis'].value_counts(dropna=False))
print('')
depression_targets.head()
depression_probes = pd.read_pickle(f'{local_path}GSE128235_published_beta_values.pkl')
print(depression_probes.shape)
print('')
print('Number probe values missing: {}'.format(depression_probes.isnull().values.sum()))
print('')
depression_probes.head()
depression_df = depression_probes.join(depression_targets)
print('Shape after join: {}'.format(depression_df.shape))
depression_df = depression_df.dropna()
print('Shape after dropna: {}'.format(depression_df.shape))
depression_df.head()
datarobot_df, target_type = create_datarobot_classification_submission(input_data_df=depression_df, target_name='diagnosis', max_features=1000, random_state=7)
print(datarobot_df.shape)
datarobot_df.head()
datarobot_project = submit_datarobot_project(input_data_df=datarobot_df, target_name='diagnosis',
target_type=target_type, project_name='RAPA_Demo_8.5.21', random_state=7)
datarobot_project
# API NOTE: Would be nice to have a progress bar here instead of print output
datarobot_project.wait_for_autopilot(verbosity=dr.VERBOSITY_LEVEL.SILENT)
datarobot_project_models = datarobot_project.get_models()
for model in datarobot_project_models:
if model.metrics['AUC']['crossValidation'] != None:
try:
model.request_feature_impact()
except dr.errors.JobAlreadyRequested:
continue
# API note: Is there a project-level wait function for all jobs, regardless of AutoPilot status?
while len(datarobot_project.get_all_jobs()) > 0:
time.sleep(10)
all_feature_importances = []
for model in datarobot_project_models:
if model.metrics['AUC']['crossValidation'] != None:
all_feature_importances.extend(model.get_feature_impact())
median_feature_importances_df = pd.DataFrame(all_feature_importances).groupby('featureName')['impactNormalized'].median().sort_values(ascending=False)
median_feature_importances_df
original_featurelist_size = median_feature_importances_df.shape[0]
for feature_pct in tqdm(np.arange(0.95, 0.04, -0.1)):
try:
desired_reduced_featurelist_size = round(original_featurelist_size * feature_pct)
reduced_features = median_feature_importances_df.head(desired_reduced_featurelist_size).index.values.tolist()
new_featurelist_name = 'RAPA Reduced to {}'.format(len(reduced_features))
reduced_featurelist = datarobot_project.create_featurelist(name=new_featurelist_name, features=reduced_features)
datarobot_project.start_autopilot(featurelist_id=reduced_featurelist.id, mode=dr.AUTOPILOT_MODE.FULL_AUTO, blend_best_models=False, prepare_model_for_deployment=False)
datarobot_project.wait_for_autopilot(verbosity=dr.VERBOSITY_LEVEL.SILENT)
datarobot_project_models = datarobot_project.get_models()
for model in datarobot_project_models:
if model.featurelist_id == reduced_featurelist.id and model.metrics['AUC']['crossValidation'] != None:
try:
model.request_feature_impact()
except dr.errors.JobAlreadyRequested:
pass
# API note: Is there a project-level wait function for all jobs, regardless of AutoPilot status?
while len(datarobot_project.get_all_jobs()) > 0:
time.sleep(10)
all_feature_importances = []
while(len(all_feature_importances) == 0):
for model in datarobot_project_models:
if model.featurelist_id == reduced_featurelist.id and model.metrics['AUC']['crossValidation'] != None:
all_feature_importances.extend(model.get_feature_impact())
time.sleep(10) #sometimes all_feature_importances is empty, so wait a little and try again?
median_feature_importances_df = pd.DataFrame(all_feature_importances).groupby('featureName')['impactNormalized'].median().sort_values(ascending=False)
except dr.errors.ClientError as e:
if 'Feature list named' in str(e) and 'already exists' in str(e):
pass
else:
raise e
# Perform a more fine-grained search at the smaller featurelist sizes
for feature_pct in tqdm(np.arange(0.04, 0.001, -0.01)):
try:
desired_reduced_featurelist_size = round(original_featurelist_size * feature_pct)
reduced_features = median_feature_importances_df.head(desired_reduced_featurelist_size).index.values.tolist()
new_featurelist_name = 'RAPA Reduced to {}'.format(len(reduced_features))
reduced_featurelist = datarobot_project.create_featurelist(name=new_featurelist_name, features=reduced_features)
datarobot_project.start_autopilot(featurelist_id=reduced_featurelist.id, mode=dr.AUTOPILOT_MODE.FULL_AUTO, blend_best_models=False, prepare_model_for_deployment=False)
datarobot_project.wait_for_autopilot(verbosity=dr.VERBOSITY_LEVEL.SILENT)
datarobot_project_models = datarobot_project.get_models()
for model in datarobot_project_models:
if model.featurelist_id == reduced_featurelist.id and model.metrics['AUC']['crossValidation'] != None:
try:
model.request_feature_impact()
except dr.errors.JobAlreadyRequested:
pass
# API note: Is there a project-level wait function for all jobs, regardless of AutoPilot status?
while len(datarobot_project.get_all_jobs()) > 0:
time.sleep(10)
while(len(all_feature_importances) == 0):
all_feature_importances = []
for model in datarobot_project_models:
if model.featurelist_id == reduced_featurelist.id and model.metrics['AUC']['crossValidation'] != None:
all_feature_importances.extend(model.get_feature_impact())
time.sleep(10)
median_feature_importances_df = pd.DataFrame(all_feature_importances).groupby('featureName')['impactNormalized'].median().sort_values(ascending=False)
except dr.errors.ClientError as e:
if 'Feature list named' in str(e) and 'already exists' in str(e):
pass
else:
raise e
```
## Plot: Distribution of model performance vs. featurelist size
```
datarobot_project_models = datarobot_project.get_models()
RAPA_model_featurelists = []
featurelist_performances = defaultdict(list)
for model in datarobot_project_models:
if 'RAPA Reduced' not in model.featurelist_name:
continue
RAPA_model_featurelists.append(model.featurelist_name)
num_features = int(model.featurelist_name.split(' ')[-1])
featurelist_performances[num_features].append(model.metrics["AUC"]["crossValidation"])
featurelist_performances_df = pd.DataFrame(featurelist_performances)[sorted(featurelist_performances.keys())]
featurelist_performances_df = featurelist_performances_df.dropna(how="all", axis=1).dropna()
with plt.style.context('tableau-colorblind10'):
sb.boxplot(data=featurelist_performances_df)
plt.ylabel('CV AUC')
plt.xlabel('Number of features')
```
## Plot: Pareto front of median model performance vs. featurelist size
```
with plt.style.context('tableau-colorblind10'):
plt.figure(figsize=(12, 10))
feature_nums = featurelist_performances_df.median().index.values
medians = featurelist_performances_df.median().values
plt.plot(feature_nums, medians, 'o-')
plt.ylabel('Median CV AUC')
plt.xlabel('Number of features')
plt.title('Pareto front trade-off')
```
## Demo take-aways
* Even when passing "only" 1,000 features (out of the original 425,000 features), substantial performance gains were achieved by continuing to reduce the featurelist size in an automated fashion (i.e., parsimony analysis).
* By default, DataRobot does not perform parsimony analysis. At best, DataRobot suggests a reduced featurelist using one round of model-based feature importance from a single model.
* Integrating parsimony analysis into DataRobot promises to improve prediction accuracy for users who work with very "wide" datasets.
| github_jupyter |
# Regression Tutorial
This guide will show how to use Tribuo’s regression models to predict wine quality based on the [UCI Wine Quality](https://archive.ics.uci.edu/ml/datasets/Wine+Quality) data set. We’ll experiment with several different regression trainers: two for training linear models (SGD and Adagrad) and one for training a tree ensemble via Tribuo’s wrapper on XGBoost (note: Tribuo's XGBoost support relies upon the Maven Central XGBoost jar which contains macOS and Linux binaries, to run this tutorial on Windows please compile DMLC's XGBoost jar from source and rebuild Tribuo). We’ll run these experiments by simply swapping in different implementations of Tribuo’s `Trainer` interface. We’ll also show how to evaluate regression models and describe some common evaluation metrics.
## Setup
First you'll need to download the winequality dataset from UCI:
`wget https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-red.csv`
then we'll load in some jars and import a few packages.
```
%jars ./tribuo-json-4.1.0-SNAPSHOT-jar-with-dependencies.jar
%jars ./tribuo-regression-sgd-4.1.0-SNAPSHOT-jar-with-dependencies.jar
%jars ./tribuo-regression-xgboost-4.1.0-SNAPSHOT-jar-with-dependencies.jar
%jars ./tribuo-regression-tree-4.1.0-SNAPSHOT-jar-with-dependencies.jar
import java.nio.file.Path;
import java.nio.file.Paths;
import org.tribuo.*;
import org.tribuo.data.csv.CSVLoader;
import org.tribuo.datasource.ListDataSource;
import org.tribuo.evaluation.TrainTestSplitter;
import org.tribuo.math.optimisers.*;
import org.tribuo.regression.*;
import org.tribuo.regression.evaluation.*;
import org.tribuo.regression.sgd.RegressionObjective;
import org.tribuo.regression.sgd.linear.LinearSGDTrainer;
import org.tribuo.regression.sgd.objectives.SquaredLoss;
import org.tribuo.regression.rtree.CARTRegressionTrainer;
import org.tribuo.regression.xgboost.XGBoostRegressionTrainer;
import org.tribuo.util.Util;
```
## Loading the data
In Tribuo, all the prediction types have an associated `OutputFactory` implementation, which can create the appropriate `Output` subclasses from an input. Here we're going to use `RegressionFactory` as we're performing regression. In Tribuo both single and multidimensional regression use the `Regressor` and `RegressionFactory` classes. We then pass the `regressionFactory` into the simple `CSVLoader` which reads all the columns into a `DataSource`. The winequality dataset uses `;` to separate the columns rather than the standard `,` so we change the default separator character. Note if your csv file isn't purely numeric or you wish to use a subset of the columns as features then you should use `CSVDataSource` which allows fine-grained control over the loading and featurisation process of your csv file. There's a columnar data tutorial which details the flexibility and power of our columnar processing infrastructure.
```
var regressionFactory = new RegressionFactory();
var csvLoader = new CSVLoader<>(';',regressionFactory);
```
We don't have a pre-defined train test split, so we take 70% as the training data, and 30% as the test data. The data is randomised using the RNG seeded by the second value. Then we feed the split data sources into the training and testing datasets. These `MutableDataset`s manage all the metadata (e.g., feature & output domains), and the mapping from feature names to feature id numbers.
```
var wineSource = csvLoader.loadDataSource(Paths.get("winequality-red.csv"),"quality");
var splitter = new TrainTestSplitter<>(wineSource, 0.7f, 0L);
Dataset<Regressor> trainData = new MutableDataset<>(splitter.getTrain());
Dataset<Regressor> evalData = new MutableDataset<>(splitter.getTest());
```
## Training the models
We're going to define a quick training function which accepts a trainer and a training dataset. It times the training and also prints the performance metrics. Evaluating on the training data is useful for debugging: if the model performs poorly in the training data, then we know something is wrong.
```
public Model<Regressor> train(String name, Trainer<Regressor> trainer, Dataset<Regressor> trainData) {
// Train the model
var startTime = System.currentTimeMillis();
Model<Regressor> model = trainer.train(trainData);
var endTime = System.currentTimeMillis();
System.out.println("Training " + name + " took " + Util.formatDuration(startTime,endTime));
// Evaluate the model on the training data (this is a useful debugging tool)
RegressionEvaluator eval = new RegressionEvaluator();
var evaluation = eval.evaluate(model,trainData);
// We create a dimension here to aid pulling out the appropriate statistics.
// You can also produce the String directly by calling "evaluation.toString()"
var dimension = new Regressor("DIM-0",Double.NaN);
System.out.printf("Evaluation (train):%n RMSE %f%n MAE %f%n R^2 %f%n",
evaluation.rmse(dimension), evaluation.mae(dimension), evaluation.r2(dimension));
return model;
}
```
Now we're going to define an equivalent testing function which accepts a model and a test dataset, printing the performance to std out.
```
public void evaluate(Model<Regressor> model, Dataset<Regressor> testData) {
// Evaluate the model on the test data
RegressionEvaluator eval = new RegressionEvaluator();
var evaluation = eval.evaluate(model,testData);
// We create a dimension here to aid pulling out the appropriate statistics.
// You can also produce the String directly by calling "evaluation.toString()"
var dimension = new Regressor("DIM-0",Double.NaN);
System.out.printf("Evaluation (test):%n RMSE %f%n MAE %f%n R^2 %f%n",
evaluation.rmse(dimension), evaluation.mae(dimension), evaluation.r2(dimension));
}
```
Now we'll define the four trainers we're going to compare.
- A linear regression trained using linear decay SGD.
- A linear regression trained using SGD and AdaGrad.
- A regression tree using the CART algorithm with a maximum depth of 6.
- An XGBoost trainer using 50 rounds of boosting.
```
var lrsgd = new LinearSGDTrainer(
new SquaredLoss(), // loss function
SGD.getLinearDecaySGD(0.01), // gradient descent algorithm
10, // number of training epochs
trainData.size()/4,// logging interval
1, // minibatch size
1L // RNG seed
);
var lrada = new LinearSGDTrainer(
new SquaredLoss(),
new AdaGrad(0.01),
10,
trainData.size()/4,
1,
1L
);
var cart = new CARTRegressionTrainer(6);
var xgb = new XGBoostRegressionTrainer(50);
```
First we'll train the linear regression with SGD:
```
var lrsgdModel = train("Linear Regression (SGD)",lrsgd,trainData);
```
## Evaluating the models
Using our evaluation function this is pretty straightforward.
```
evaluate(lrsgdModel,evalData);
```
Those numbers seem poor, but what do these evaluation metrics mean?
### RMSE
The root-mean-square error (RMSE) summarizes the magnitude of errors between our regression model's predictions and the values we observe in our data. Basically, RMSE is the standard deviation of model prediction errors on a given dataset.
$$RMSE = \sqrt{ \frac{1}{n} \sum_{i=1}^{n} (y_i - \hat{y}_i)^2 }$$
Lower is better: a perfect model for the wine data would have RMSE=0. The RMSE is sensitive to how large an error was, and is thus sensitive to outliers. This also means that RMSE can be used to compare different models on the same dataset but not across different datasets, as a "good" RMSE value on one dataset might be larger than a "good" RMSE value on a different dataset. See [Wikipedia](https://en.wikipedia.org/wiki/Root-mean-square_deviation) for more info on RMSE.
### MAE
The mean absolute error (MAE) is another summary of model error. Unlike RMSE, each error in MAE contributes proportional to its absolute value.
$$MAE = \frac{1}{n} \sum_{i=1}^{n} |y_i - \hat{y}_i|$$
### R^2
The R-squared metric (also called the "coefficient of determination") summarizes how much of the variation in observed outcomes can be explained by our model.
Let $\bar{y} = \frac{1}{n} \sum_{i=1}^{n} y_i$, i.e., the mean deviation of observed data points from the observed mean. R^2 is given by:
$$R^2 = 1 - \frac{\sum_{i=1}^{n} (y_i - \hat{y}_i)^2}{\sum_{i=1}^{n} (y_i - \bar{y})^2}$$
A value of R^2=1 means that the model accounts for all of the variation in a set of observations -- in other words, it fits a dataset perfectly. Note that R^2 can turn negative when the sum-of-squared model errors (numerator) is greater than the sum-of-squared differences between observed data points and the observed mean (denominator). In other words, when R^2 is negative, the model fits the data *worse* than simply using the observed mean to predict values.
See [Wikipedia](https://en.wikipedia.org/wiki/Coefficient_of_determination) and the [Minitab blog](https://blog.minitab.com/blog/adventures-in-statistics-2/regression-analysis-how-do-i-interpret-r-squared-and-assess-the-goodness-of-fit) for more detailed discussion of R^2.
## Improving over standard SGD with AdaGrad
It's not surprising the SGD results are bad: in linear decay SGD, the step size used for parameter updates changes over time (training iterations) but is uniform across all model parameters. This means that we use the same step size for a noisy/irrelevant feature as we would for an informative feature. There many more sophisticated approaches to gradient descent.
One of these is [AdaGrad](http://www.jmlr.org/papers/volume12/duchi11a/duchi11a.pdf), modifies the "global" learning rate for each parameter $p$ using the sum-of-squares of past gradients w.r.t. $p$, up to time $t$.
> ... the secret sauce of AdaGrad is not on necessarily accelerating gradient descent with a better step size selection, but making gradient descent more stable to not-so-good \(\eta\) choices.
> Anastasios Kyrillidis, [Note on AdaGrad](http://akyrillidis.github.io/notes/AdaGrad)
Let's try training for the same number of epochs using `AdaGrad` instead of `LinearDecaySGD`:
```
var lradaModel = train("Linear Regression (AdaGrad)",lrada,trainData);
evaluate(lradaModel,evalData);
```
Using a more robust optimizer got us a better fit in the same number of epochs. However, both the train and test R^2 scores are still substantially less than 1 and, as before, the train and test RMSE scores are very similar.
See [here](http://akyrillidis.github.io/notes/AdaGrad) and [here](http://ruder.io/optimizing-gradient-descent/index.html#adagrad) for more on AdaGrad. Also, there are many other implementations of various well-known optimizers in Tribuo, including [Adam](https://tribuo.org/learn/4.0/javadoc/org/tribuo/math/optimisers/Adam.html) and [RMSProp](https://tribuo.org/learn/4.0/javadoc/org/tribuo/math/optimisers/RMSProp.html). See the [math.optimisers](https://tribuo.org/learn/4.0/javadoc/org/tribuo/math/optimisers/package-summary.html) package.
At this point, we showed that we can improve our model by using a more robust optimizer; however, we're still using a linear model. If there are informative, non-linear relationships among wine quality features, then our current model won't be able to take advantage of them. We'll finish this tutorial by showing how to use a couple of popular non-linear models, CART and [XGBoost](https://xgboost.ai).
## Trees and ensembles
Next we'll train the CART tree:
```
var cartModel = train("CART",cart,trainData);
evaluate(cartModel,evalData);
```
Finally we'll train the XGBoost ensemble:
```
var xgbModel = train("XGBoost",xgb,trainData);
evaluate(xgbModel,evalData);
```
Using gradient boosting via XGBoost improved results by a lot. Not only are the train & test fits better, but the train and test RMSE have started to diverge, indicating that the XGBoost model isn't underfitting like the previous two linear models were. XGBoost won't always be the best model for your data, but it's often a great baseline model to try when facing a new problem or dataset.
## Conclusion
In this tutorial, we showed how to experiment with several different regression trainers (linear decay SGD, AdaGrad, CART, XGBoost). It was easy to experiment with different trainers and models by simply swapping in different implementations of the Tribuo `Trainer` interface. We also showed how to evaluate regression models and described some common evaluation metrics.
| github_jupyter |
# 横向联邦学习任务示例
这是一个使用Delta框架编写的横向联邦学习的任务示例。
数据是分布在多个节点上的[MNIST数据集](http://yann.lecun.com/exdb/mnist/),每个节点上只有其中的一部分样本。任务是训练一个卷积神经网络的模型,进行手写数字的识别。
本示例可以直接在Deltaboard中执行并查看结果。<span style="color:#FF8F8F;font-weight:bold">在点击执行之前,需要修改一下个人的Deltaboard API的地址,具体请看下面第4节的说明。</span>
## 1. 引入需要的包
我们的计算逻辑是用torch写的。所以首先引入```numpy```和```torch```,以及一些辅助的工具,然后从```delta-task```的包中,引入Delta框架的内容,包括```DeltaNode```节点,用于调用API发送任务,以及我们本示例中要执行的横向联邦学习任务```HorizontalTask```等等:
```
from typing import Dict, Iterable, List, Tuple, Any, Union
import numpy as np
import torch
from delta import DeltaNode
from delta.task import HorizontalTask
from delta.algorithm.horizontal import FedAvg
```
## 2. 定义神经网络模型
接下来我们来定义神经网络模型,这里和传统的神经网络模型定义完全一样:
```
class LeNet(torch.nn.Module):
def __init__(self):
super().__init__()
self.conv1 = torch.nn.Conv2d(1, 16, 5, padding=2)
self.pool1 = torch.nn.AvgPool2d(2, stride=2)
self.conv2 = torch.nn.Conv2d(16, 16, 5)
self.pool2 = torch.nn.AvgPool2d(2, stride=2)
self.dense1 = torch.nn.Linear(400, 100)
self.dense2 = torch.nn.Linear(100, 10)
def forward(self, x: torch.Tensor):
x = self.conv1(x)
x = torch.relu(x)
x = self.pool1(x)
x = self.conv2(x)
x = torch.relu(x)
x = self.pool2(x)
x = x.view(-1, 400)
x = self.dense1(x)
x = torch.relu(x)
x = self.dense2(x)
return x
```
## 3. 定义隐私计算任务
然后可以开始定义我们的横向联邦任务了,用横向联邦学习的方式,在多节点上训练上面定义的神经网络模型
在定义横向联邦学习任务时,有几部分内容是需要用户自己定义的:
* ***模型训练方法***:包括损失函数、优化器,以及训练步骤的定义
* ***数据预处理方法***:在执行训练步骤以前,对于加载的每个样本数据进行预处理的方法,具体的参数说明,可以参考[这篇文档](https://docs.deltampc.com/network-deployment/prepare-data)
* ***模型验证方法***:在每个节点上通过验证样本集,计算模型精确度的方法
* ***横向联邦配置***:每轮训练需要多少个节点,如何在节点上划分验证样本集合等等
```
class ExampleTask(HorizontalTask):
def __init__(self):
super().__init__(
name="example", # 任务名称,用于在Deltaboard中的展示
dataset="mnist", # 任务用到的数据集的文件名,对应于Delta Node的data文件夹下的一个文件/文件夹
max_rounds=2, # 任务训练的总轮次,每聚合更新一次权重,代表一轮
validate_interval=1, # 验证的轮次间隔,1表示每完成一轮,进行一次验证
validate_frac=0.1, # 验证集的比例,范围(0,1)
)
# 传入刚刚定义的神经网络模型
self.model = LeNet()
# 模型训练时用到的损失函数
self.loss_func = torch.nn.CrossEntropyLoss()
# 模型训练时的优化器
self.optimizer = torch.optim.SGD(
self.model.parameters(),
lr=0.1,
momentum=0.9,
weight_decay=1e-3,
nesterov=True,
)
def preprocess(self, x, y=None):
"""
数据预处理方法,会在数据加载时,对每一个样本进行预处理。
具体的参数说明,可以参考https://docs.deltampc.com/network-deployment/prepare-data
x: 原始数据集中的一个样本,类型与指定的数据集相关
y: 数据对应的标签,如果数据集中不包含标签,则为None
return: 预处理完的数据和标签(如果存在),类型需要为torch.Tensor或np.ndarray
"""
x /= 255.0
x *= 2
x -= 1
x = x.reshape((1, 28, 28))
return torch.from_numpy(x), torch.tensor(int(y), dtype=torch.long)
def train(self, dataloader: Iterable):
"""
训练步骤
dataloader: 训练数据集对应的dataloader
return: None
"""
for batch in dataloader:
x, y = batch
y_pred = self.model(x)
loss = self.loss_func(y_pred, y)
self.optimizer.zero_grad()
loss.backward()
self.optimizer.step()
def validate(self, dataloader: Iterable) -> Dict[str, float]:
"""
验证步骤,输出验证的指标值
dataloader: 验证集对应的dataloader
return: Dict[str, float],一个字典,键为指标的名称(str),值为对应的指标值(float)
"""
total_loss = 0
count = 0
ys = []
y_s = []
for batch in dataloader:
x, y = batch
y_pred = self.model(x)
loss = self.loss_func(y_pred, y)
total_loss += loss.item()
count += 1
y_ = torch.argmax(y_pred, dim=1)
y_s.extend(y_.tolist())
ys.extend(y.tolist())
avg_loss = total_loss / count
tp = len([1 for i in range(len(ys)) if ys[i] == y_s[i]])
precision = tp / len(ys)
return {"loss": avg_loss, "precision": precision}
def get_params(self) -> List[torch.Tensor]:
"""
需要训练的模型参数
在聚合更新、保存结果时,只会更新、保存get_params返回的参数
return: List[torch.Tensor], 模型参数列表
"""
return list(self.model.parameters())
def algorithm(self):
"""
聚合更新算法的配置,可选算法包含在delta.algorithm.horizontal包中
"""
return FedAvg(
merge_interval_epoch=0, # 聚合更新的间隔,merge_interval_epoch表示每多少个epoch聚合更新一次权重
merge_interval_iter=20, # 聚合更新的间隔,merge_interval_iter表示每多少个iteration聚合更新一次,merge_interval_epoch与merge_interval_iter互斥,必须有一个为0
wait_timeout=20, # 等待超时时间,用来控制一轮计算的超时时间
connection_timeout=20, # 连接超时时间,用来控制流程中每个阶段的超时时间
min_clients=2, # 算法所需的最少客户端数,至少为2
max_clients=2, # 算法所支持的最大客户端数,必须大雨等于min_clients
)
def dataloader_config(
self,
) -> Union[Dict[str, Any], Tuple[Dict[str, Any], Dict[str, Any]]]:
"""
训练集dataloader和验证集dataloader的配置,
每个配置为一个字典,对应pytorch中dataloader的配置
详情参见 https://pytorch.org/docs/stable/data.html
return: 一个或两个Dict[str, Any],返回一个时,同时配置训练集和验证集的dataloader,返回两个时,分别对应训练集和验证集
"""
train_config = {"batch_size": 64, "shuffle": True, "drop_last": True}
val_config = {"batch_size": 64, "shuffle": False, "drop_last": False}
return train_config, val_config
```
## 4. 指定执行任务用的Delta Node的API
定义好了任务,我们就可以开始准备在Delta Node上执行任务了。
Delta Task框架可以直接调用Delta Node API发送任务到Delta Node开始执行,只要在任务执行时指定Delta Node的API地址即可。
Deltaboard提供了对于Delta Node的API的封装,为每个用户提供了一个独立的API地址,支持多人同时使用同一个Delta Node,并且能够在Deltaboard中管理自己提交的任务。
在这里,我们使用Deltaboard提供的API来执行任务。如果用户自己搭建了Delta Node,也可以直接使用Delta Node的API。
在左侧导航栏中进入“个人中心”,在Deltaboard API中,复制自己的API地址,并粘贴到下面的代码中:
```
DELTA_NODE_API = "http://127.0.0.1:6704"
```
## 5. 执行隐私计算任务
接下来我们可以开始运行这个模型了:
```
task = ExampleTask()
delta_node = DeltaNode(DELTA_NODE_API)
delta_node.create_task(task)
```
## 6. 查看执行状态
点击执行后,可以从输出的日志看出,任务已经提交到了Delta Node的节点上。
接下来,可以从左侧的导航栏中,前往“任务列表”,找到刚刚提交的任务,点击进去查看具体的执行日志了。
| github_jupyter |
We will use Naive Bayes to model the "Pima Indians Diabetes" data set. This model will predict which people are likely to develop diabetes.
This dataset is originally from the National Institute of Diabetes and Digestive and Kidney Diseases. The objective of the dataset is to diagnostically predict whether or not a patient has diabetes, based on certain diagnostic measurements included in the dataset. Several constraints were placed on the selection of these instances from a larger database. In particular, all patients here are females at least 21 years old of Pima Indian heritage.
## Import Libraries
```
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import matplotlib.pyplot as plt # matplotlib.pyplot plots data
%matplotlib inline
import seaborn as sns
```
## Load and review data
```
pdata = pd.read_csv("pima-indians-diabetes.csv")
pdata.shape # Check number of columns and rows in data frame
pdata.head() # To check first 5 rows of data set
pdata.isnull().values.any() # If there are any null values in data set
columns = list(pdata)[0:-1] # Excluding Outcome column which has only
pdata[columns].hist(stacked=False, bins=100, figsize=(12,30), layout=(14,2));
# Histogram of first 8 columns
```
## Identify Correlation in data
```
pdata.corr() # It will show correlation matrix
# However we want to see correlation in graphical representation so below is function for that
def plot_corr(df, size=11):
corr = df.corr()
fig, ax = plt.subplots(figsize=(size, size))
ax.matshow(corr)
plt.xticks(range(len(corr.columns)), corr.columns)
plt.yticks(range(len(corr.columns)), corr.columns)
plot_corr(pdata)
```
In above plot yellow colour represents maximum correlation and blue colour represents minimum correlation.
We can see none of variable have correlation with any other variables.
```
sns.pairplot(pdata,diag_kind='kde')
```
## Calculate diabetes ratio of True/False from outcome variable
```
n_true = len(pdata.loc[pdata['class'] == True])
n_false = len(pdata.loc[pdata['class'] == False])
print("Number of true cases: {0} ({1:2.2f}%)".format(n_true, (n_true / (n_true + n_false)) * 100 ))
print("Number of false cases: {0} ({1:2.2f}%)".format(n_false, (n_false / (n_true + n_false)) * 100))
```
So we have 34.90% people in current data set who have diabetes and rest of 65.10% doesn't have diabetes.
Its a good distribution True/False cases of diabetes in data.
## Spliting the data
I will use 70% of data for training and 30% for testing.
```
from sklearn.model_selection import train_test_split
X = pdata.drop('class',axis=1) # Predictor feature columns (8 X m)
Y = pdata['class'] # Predicted class (1=True, 0=False) (1 X m)
x_train, x_test, y_train, y_test = train_test_split(X, Y, test_size=0.3, random_state=1)
# 1 is just any random seed number
x_train.head()
```
Lets check split of data
```
print("{0:0.2f}% data is in training set".format((len(x_train)/len(pdata.index)) * 100))
print("{0:0.2f}% data is in test set".format((len(x_test)/len(pdata.index)) * 100))
```
Now lets check diabetes True/False ratio in split data
```
print("Original Diabetes True Values : {0} ({1:0.2f}%)".format(len(pdata.loc[pdata['class'] == 1]), (len(pdata.loc[pdata['class'] == 1])/len(pdata.index)) * 100))
print("Original Diabetes False Values : {0} ({1:0.2f}%)".format(len(pdata.loc[pdata['class'] == 0]), (len(pdata.loc[pdata['class'] == 0])/len(pdata.index)) * 100))
print("")
print("Training Diabetes True Values : {0} ({1:0.2f}%)".format(len(y_train[y_train[:] == 1]), (len(y_train[y_train[:] == 1])/len(y_train)) * 100))
print("Training Diabetes False Values : {0} ({1:0.2f}%)".format(len(y_train[y_train[:] == 0]), (len(y_train[y_train[:] == 0])/len(y_train)) * 100))
print("")
print("Test Diabetes True Values : {0} ({1:0.2f}%)".format(len(y_test[y_test[:] == 1]), (len(y_test[y_test[:] == 1])/len(y_test)) * 100))
print("Test Diabetes False Values : {0} ({1:0.2f}%)".format(len(y_test[y_test[:] == 0]), (len(y_test[y_test[:] == 0])/len(y_test)) * 100))
print("")
```
# Data Preparation
### Check hidden missing values
As we checked missing values earlier but haven't got any. But there can be lots of entries with 0 values. We must need to take care of those as well.
```
x_train.head()
```
We can see lots of 0 entries above.
### Replace 0s with serial mean
```
#from sklearn.preprocessing import Imputer
#my_imputer = Imputer()
#data_with_imputed_values = my_imputer.fit_transform(original_data)
from sklearn.impute import SimpleImputer
rep_0 = SimpleImputer(missing_values=0, strategy="mean")
cols=x_train.columns
x_train = pd.DataFrame(rep_0.fit_transform(x_train))
x_test = pd.DataFrame(rep_0.fit_transform(x_test))
x_train.columns = cols
x_test.columns = cols
x_train.head()
```
# Train Naive Bayes algorithm
```
from sklearn.naive_bayes import GaussianNB # using Gaussian algorithm from Naive Bayes
# creatw the model
diab_model = GaussianNB()
diab_model.fit(x_train, y_train.ravel())
```
### Performance of our model with training data
```
diab_train_predict = diab_model.predict(x_train)
from sklearn import metrics
print("Model Accuracy: {0:.4f}".format(metrics.accuracy_score(y_train, diab_train_predict)))
print()
```
### Performance of our model with testing data
```
diab_test_predict = diab_model.predict(x_test)
from sklearn import metrics
print("Model Accuracy: {0:.4f}".format(metrics.accuracy_score(y_test, diab_test_predict)))
print()
```
### Lets check the confusion matrix and classification report
```
print("Confusion Matrix")
cm=metrics.confusion_matrix(y_test, diab_test_predict, labels=[1, 0])
df_cm = pd.DataFrame(cm, index = [i for i in ["1","0"]],
columns = [i for i in ["Predict 1","Predict 0"]])
plt.figure(figsize = (7,5))
sns.heatmap(df_cm, annot=True)
print("Classification Report")
print(metrics.classification_report(y_test, diab_test_predict, labels=[1, 0]))
```
We can see our true positive numbers with value 1 is of precision and recall is below 70%
| github_jupyter |
```
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
from kaggle_datasets import KaggleDatasets
title=[]
path=[]
title=['hpaimage-train-pad',
'hpaimage-valid-pad',
'mys0101',
'mys0102',
'mys0103',
'mys0104',
'mys0105',
'mys0106',
'mys0107',
'mys0108',
'mys0201',
'mys0202',
'mys0203',
'mys0204',
'mys0205',
'mys0206',
'mys0207',
'mys0208',
'mys0209',
'mys010102',
'mys010202',
'mys010302'
# add dataset you want to use
]
for t in title:
try:
path.append(KaggleDatasets().get_gcs_path(t))
except:
try:
path.append(KaggleDatasets().get_gcs_path(t))
except:
try:
path.append(KaggleDatasets().get_gcs_path(t))
except:
try:
path.append(KaggleDatasets().get_gcs_path(t))
except:
print('connection error!')
break
import csv
f = open('gs.csv','w',encoding='utf-8')
csv_writer = csv.writer(f)
csv_writer.writerow(["ID","URL"])
for t,p in zip(title,path):
csv_writer.writerow([t,p])
f.close()
try:
KaggleDatasets().get_gcs_path('mys020902')
except:
try:
KaggleDatasets().get_gcs_path('mys020902')
except:
KaggleDatasets().get_gcs_path('mys020902')
try:
KaggleDatasets().get_gcs_path('mys021002')
except:
try:
KaggleDatasets().get_gcs_path('mys021002')
except:
KaggleDatasets().get_gcs_path('mys021002')
try:
KaggleDatasets().get_gcs_path('mys021102')
except:
try:
KaggleDatasets().get_gcs_path('mys021102')
except:
KaggleDatasets().get_gcs_path('mys021102')
try:
KaggleDatasets().get_gcs_path('mys021202')
except:
try:
KaggleDatasets().get_gcs_path('mys021202')
except:
KaggleDatasets().get_gcs_path('mys021202')
#'gs://kds-1b4418164362b741890b32439ecc808b1b22a0d26803b8f9c7d8c837' MYS010102
#'gs://kds-ef90b2545def57e9891a01a93a3925554575b17f8195b5b475389b0c' MYS010202
#'gs://kds-017f4b39e8b0225c7dc08719cbd8d5ea561ced670e08cbf0ff05fd84' MYS010302
#'gs://kds-32c9ed20bb5228907747455eaf6da993350b0e7ac4e5a5d1c1c71a4c' MYS010402
#'gs://kds-f367bdacfcc56006d2189a1825722c9b473ab2f4fbe0b7845660ac34' MYS010502
#'gs://kds-9ef047cab1ccb4cc1ef25e142407f6c7c3c6831bde17d62c5e93bb71' MYS010602
#'gs://kds-cd39df118637847c24e8595313e36789ebabc4713a1f4f8f382a7c09' MYS010702
#'gs://kds-5d724122e8d49805639eeb2bf1c85b534cd833fd4988cf2d9191d6e9' MYS010802
'gs://kds-5f4b4abd87e9bbda926b0d0e741f07f050cbfc686d9a43ef01f12139' MYS020102
'gs://kds-6460e687552f35430943edfd530121687ae78cdd5ed59723db5d621c' MYS020202
'gs://kds-a897abf154250176fced879f98b6c7a8fc1a8648e6fa757a295e43a9' MYS020302
'gs://kds-21832b326644d6c2deeef613db6b6ff725652b480a52425438bd35bb' MYS020402
'gs://kds-5843d58caabc5225c44c6af9b79230e8924ae579bca735a4fd35a76d' MYS020502
'gs://kds-cfd75773aeb6e63b05e490701a3cf4c0baf2986172641568523226dc' MYS020602
'gs://kds-bae5a6145a74e05e64bd0dc1fbe73ab4f9ff92124ced750eada67170' MYS020702
'gs://kds-31af620f54b49fb72a88ec821e831f3682ce3f22a8e069ceb51df0ac' MYS020802
'gs://kds-bfec3db6c906ce6f35cfbd5442f047b1335be52132f3197447474f6b' MYS020902
'gs://kds-7d2926e2a3a3bc4e728e5077f51bfed267223b57faebe2018236df96' MYS021002
'gs://kds-cb3b417fd11651934b8dabf047b5feea05bbd7920f90485e43bd36ab' MYS021102
'gs://kds-7ba9627a72bcc701e4e32eae4174c565ed5de36c5ae3caa18f86d814' MYS021202
KaggleDatasets().get_gcs_path('mys020102')
KaggleDatasets().get_gcs_path('mys020202')
KaggleDatasets().get_gcs_path('mys020302')
KaggleDatasets().get_gcs_path('mys020402')
KaggleDatasets().get_gcs_path('mys020502')
KaggleDatasets().get_gcs_path('mys020602')
KaggleDatasets().get_gcs_path('mys020702')
KaggleDatasets().get_gcs_path('mys020802')
KaggleDatasets().get_gcs_path('mys020902')
KaggleDatasets().get_gcs_path('mys021002')
KaggleDatasets().get_gcs_path('mys021102')
KaggleDatasets().get_gcs_path('mys021202')
'gs://kds-03ab4a6db807c00d16132f99291dca8a454219b7dd2ad0082fd0f3ab' 0101
'gs://kds-974f38a4727e58bfaba504114c5b6685e8868c48f38ff83a14547f50' 0102
'gs://kds-4a9bb3b885a0c09270238624d571fa58e12a60e9e21794d4745e0930' 0103
'gs://kds-e2a31f5845b143a72fc6267de699db94728ee08391b3b7f7de0d9bb9' 0104
'gs://kds-e8a86031574510b98924025ba56f3d7991f8d73819c784ff050dccb4' 0105
'gs://kds-f0c74bb271550427c8f7ee031def98f1f63cd93a9286d12ff23080f0' 0106
'gs://kds-81b099e72286dd356104929768409ba2d9eb846bc9de168da14ea8a2' 0107
'gs://kds-e59c1a7f10f510a33d5534abf8f56a3d2e2cd6b5c37947f0a6707a97' 0201
'gs://kds-1b7ee7598a52aa7f0a519f2e282e29f4112ecbc3587888abfca9a067' 0202
'gs://kds-e43b0c9a8c9714bf3ae1e28fbd599476b2f4a71c4b319a687842df84' 0203
'gs://kds-f40614a201da4edbd45f23ba9991e61f84c27261a57088f6768dc800' 0204
'gs://kds-de7d5ba6ade39f4c8928690c9722793dad0d2b4900fcaa04819971e6' 0205
'gs://kds-0e4562becbdb09814309b65188723905a19bdb55b50d21c0b76cbdd1' 0206
'gs://kds-6e2468e45917d4c8f3d22de260b59e1aff13601e80a93718020e639a' 0207
'gs://kds-18336b8eeb1818f8c43da58b1d29bbf647f3bbb7d249f693085b9d9e' 0208
'gs://kds-6948c07e8d106548d3e89808264abe1cfaa1f19090dbdea1370a28b3' 0209
KaggleDatasets().get_gcs_path('mys0101')
KaggleDatasets().get_gcs_path('mys0102')
KaggleDatasets().get_gcs_path('mys0103')
KaggleDatasets().get_gcs_path('mys0104')
KaggleDatasets().get_gcs_path('mys0105')
KaggleDatasets().get_gcs_path('mys0106')
KaggleDatasets().get_gcs_path('mys0107')
KaggleDatasets().get_gcs_path('mys0108')
KaggleDatasets().get_gcs_path('mys0201')
KaggleDatasets().get_gcs_path('mys0202')
KaggleDatasets().get_gcs_path('mys0203')
KaggleDatasets().get_gcs_path('mys0204')
KaggleDatasets().get_gcs_path('mys0205')
KaggleDatasets().get_gcs_path('mys0206')
KaggleDatasets().get_gcs_path('mys0207')
KaggleDatasets().get_gcs_path('mys0208')
KaggleDatasets().get_gcs_path('mys0209')
KaggleDatasets().get_gcs_path('mys0205')
KaggleDatasets().get_gcs_path('mys0203')
KaggleDatasets().get_gcs_path('mys0204')
KaggleDatasets().get_gcs_path('mys0205')
KaggleDatasets().get_gcs_path('mys0206')
KaggleDatasets().get_gcs_path('mys0207')
KaggleDatasets().get_gcs_path('mys0208')
KaggleDatasets().get_gcs_path('mys0209')
KaggleDatasets().get_gcs_path('mys0204')
KaggleDatasets().get_gcs_path('mys0103')
KaggleDatasets().get_gcs_path('mys0104')
KaggleDatasets().get_gcs_path('mys0105')
KaggleDatasets().get_gcs_path('mys0106')
KaggleDatasets().get_gcs_path('mys0107')
KaggleDatasets().get_gcs_path('mys0108')
'gs://kds-18467a34620ba1afe1f389fd285bf108447b09f584cbe7c55a500ebe' MYS0101
'gs://kds-099b48978fad967933337af13a66cb036007b1310d5b231711934bb2' MYS0102
'gs://kds-695d786331c70c00dde5644812f3b8ea8f2ae892cab2a4f48a2f9766' MYS0103
'gs://kds-7b0babb233b7ca137d3618494467528c94939ab3ed6700474cc134e3' MYS0104
'gs://kds-1f4282d99006b8514585b07f4a361bcfb8d19454bb7872b81e7d37e7' MYS0105
'gs://kds-ac7d763f7f8eb506cb0b1868e31b45301e079bb1d509df6237aaf8db' MYS0106
'gs://kds-f2ed388129c9136f54e2645a1d8b30622c264e9dfcf1d59daab50c4f' MYS0107
'gs://kds-52c5b56223349671ebe5f640fb11981cd40a1c052da071daeca2ba8a' MYS0108
'gs://kds-81e6dfe132eb32b60d9d018d968e64a7633e530b5a212a50d6e9ded9' MYS0201
'gs://kds-da16be69b2e2a909bc95a2cffd9efede5d035fcb9793591520d96332' MYS0202
'gs://kds-a28dfde12325e031ae832c7e818f84c660a297284d409b20ed496c63' MYS0203
'gs://kds-dd992e3678f2e10af911cd0f85c5925d9bf9ed530161c790b780c23f' MYS0204
'gs://kds-df3fc43230853e82f685f4c3029c6238b8d29cfb98d20ae635a0d2d2' MYS0205
'gs://kds-30b9f186e0fe2978ad22c7f59109e4cc4e65ec9896839e5ca0b10ccb' MYS0206
'gs://kds-06683f7151402f921a76bdb3dbee0b711d13e8de449343e09a8404dd' MYS0207
'gs://kds-60c69c7f8822b869b35a0c1390c61041abf3b2932c1caced16df832d' MYS0208
'gs://kds-c9807109d89d8da8c8077075206ea4c048cd05fce211953e601a5f75' MYS0209
#'gs://kds-c90356b0c1a5565426b91f2ffea10276766a0b17367e43f88882d164' hpaexset-02tfrec
#'gs://kds-42aa49ce9fc8212676e97da0be4f4b4ddbcb0f04aa5a7481a97aac08' hpaexset-3-5tfrec
#'gs://kds-bc31e123b5089b0128a4f3e872c3dfbb6b3545a2e87468449cbda814' hpaexset-6-8tfrec
#'gs://kds-871d044987171af8883d7c7f98174dc75853a4a5eb5f28f6a30d0eb7' hpaexset-9-11
#'gs://kds-75bbca92fb91ce73f2e1c5eaa27578caa91308d0f702d9aa0b77fbce' hpaexset-12-14tfrec
#'gs://kds-cd5df74554678136acf099d07fdc0b632dd162396e04f51616040fa8' hpaexset-15-17tfrec
#'gs://kds-96c49accad963db1c4068dadc7132d6495323b4785eae6522b537115' hpaexset-18-20tfrec
#'gs://kds-c0fa84156d8e93032e87c17985a0c4051c300863de62a3f341d94acc' hpaexset-21-23tfrec
# 'gs://kds-a6c822533c6fa25a6358bb47e20d988305def64a9790104bfb4e3596' hpaexset-24-26tfrec
#'gs://kds-de5b3c0fdfbe51f9fc2e36af8665b2b76bf50daed0f7b2b705c8e0c6' hpaexset-27-30tfrec
# 'gs://kds-5566487c260e590cfab3f0b9c79fc433988134436a6ee939860b5d44' hpaexset-31-34tfrec
#'gs://kds-bc6e8d8aeb83e22f6839287353c7ee17784d3c1d852528a12679a6a8' hpaexset-35-37tfrec
#'gs://kds-004df2f4910e2bf06fc13b08ef1bdb77467be005cff1fea662d07c73' hpaexset-02tfrec
#'gs://kds-16cffa22de689b2549ff0bb18319210db5cc4566db005e35282ba293' hpaexset-3-5tfrec
#'gs://kds-5774c5e377adbfc07a2cde16dba0ef50cf79f4459280fd7ee4bb8ff3' hpaexset-6-8tfrec
#'gs://kds-20636028bcd9f63dc66fd5790cc609e05e2f836911243ef50159b008' hpaexset-9-11
#'gs://kds-c90ca8f6f770bac1662c25b73f8cd83eb94897a0632df7a20f19dbec' hpaexset-12-14tfrec
#'gs://kds-b0bf359a1d74d177f49a1f189b11142ccbaaacd7a6eb4d108f1d4fca' hpaexset-15-17tfrec
#'gs://kds-1b20087af52f7720ba8347a73cca5be42a92d20c0092489ef7b0af05' hpaexset-18-20tfrec
#'gs://kds-79a5c17813bfefd7dc4239f7ae438cd4483b713fd0610221321aa69a' hpaexset-21-23tfrec
#'gs://kds-ca83f53b04345ee5b510e901a2711465d54e134c7b7bd7d9ca52b54a' hpaexset-24-26tfrec
#'gs://kds-8ffb8be72389a3683a6925456e96ad5b7bc6516fc95457f7e2a16691' hpaexset-27-30tfrec
#'gs://kds-d33bfe4ffc561ba62405827097415975c2a0ef7babcd9f693f7aa643' hpaexset-31-34tfrec
#'gs://kds-1afb9a2e010e75655e2e686dafaf1a9d1ac2feeaf01737e8a9506d35' hpaexset-35-37tfrec
KaggleDatasets().get_gcs_path('hpatrain-green-imagetfrec')
KaggleDatasets().get_gcs_path('hpavalid-green-imagetfrec')
KaggleDatasets().get_gcs_path('hpatrain-ex08-green-imagetfrec')
KaggleDatasets().get_gcs_path('hpatrain-ex-916-green-imagetfrec')
KaggleDatasets().get_gcs_path('hpatrain-ex-1724-green-imagetfrec')
KaggleDatasets().get_gcs_path('hpatrain-ex-2532-green-imagetfrec')
KaggleDatasets().get_gcs_path('hpatrain-ex-3338-green-imagetfrec')
# 'gs://kds-db891f03f8787fad574c283c8f45d40c0072ebcdc9e44f8b821c3788' hpatrain-green-imagetfrec
#'gs://kds-f126cc8401f164a946dd39442f9eb065fe400734970516b2593a0cca' hpavalid-green-imagetfrec
#'gs://kds-3867054d379fba84653fcf956ef637375f645217ab4358a185785aaf' hpatrain-ex08-green-imagetfrec
#'gs://kds-45d6c0aafb360b188d1a74670585227dbcc2847490f9dec13e879934' hpatrain-ex-916-green-imagetfrec
#'gs://kds-47b27aa31455ce4713b8c540ab3bb52d07f01b5fe2bf4be64c673821' hpatrain-ex-1724-green-imagetfrec
#'gs://kds-833ca2c0d1e0e67d9c5b3ad7c0da7c3235f418ca65031f5e6df34fc3' hpatrain-ex-2532-green-imagetfrec
# 'gs://kds-ee5eb0076b7380bde326f8d748a8103f22ed7b4f309d43ddaadb5049' hpatrain-ex-3338-green-imagetfrec
KaggleDatasets().get_gcs_path('hpaexset-02tfrec')
KaggleDatasets().get_gcs_path('hpaexset-3-5tfrec')
KaggleDatasets().get_gcs_path('hpaexset-6-8tfrec')
KaggleDatasets().get_gcs_path('hpaexset-9-11')
KaggleDatasets().get_gcs_path('hpaexset-12-14tfrec')
KaggleDatasets().get_gcs_path('hpaexset-15-17tfrec')
KaggleDatasets().get_gcs_path('hpaexset-18-20tfrec')
KaggleDatasets().get_gcs_path('hpaexset-21-23tfrec')
KaggleDatasets().get_gcs_path('hpaexset-24-26tfrec')
KaggleDatasets().get_gcs_path('hpaexset-27-30tfrec')
KaggleDatasets().get_gcs_path('hpaexset-31-34tfrec')
KaggleDatasets().get_gcs_path('hpaexset-35-37tfrec')
KaggleDatasets().get_gcs_path('hpaexset-12-14tfrec')
KaggleDatasets().get_gcs_path('hpatrain-ex-916-green-imagetfrec')
KaggleDatasets().get_gcs_path('hpatrain-ex-1724-green-imagetfrec')
KaggleDatasets().get_gcs_path('hpatrain-ex-2532-green-imagetfrec')
KaggleDatasets().get_gcs_path('hpatrain-ex-3338-green-imagetfrec')
GCS_DS_PATH = []
GCS_DS_PATH = []
GCS_DS_PATH.append(KaggleDatasets().get_gcs_path('hpasinglecell0png'))
#'gs://kds-ee2bbb7b1b93a3aca6594b057105767bf7740a87e2c9e737df2bd14e' hpasinglecell0png
#'gs://kds-0127ffcbb8111d1467f791a3c1827f88fd682f420d074fcf3f5d5114' hpasinglecell1png
#'gs://kds-45c77f30a5cdf65369189f85ee40fd4a0f2cf13eb58e6be037942c27' hpasinglecell2png
#'gs://kds-90dcc93f1059ed7ab006ccc06e8013f83dc003e3c2000c6f7fa1e220' hpasinglecell3png
#'gs://kds-fb300e8fd046e2a7ae9baa24ab75f0e2fffdd643d56429fa3bf52edb' hpasinglecell4png
#'gs://kds-3c1b8b947b4434df5c7282f825e8c934c39d21fce642b789ffd37c34' 0
#'gs://kds-2629380fa038005cdf433ec78b56880bedc1d532134b5f5c51c728ea' ../input/hpatrain-tfrec-1
#'gs://kds-c082954115e46fdb4347e3790361e6f3caddb59cb51600319bdc0285' hpatrain-tfrec-2
#'gs://kds-63919c6ce1ace04e021733e22e37dbd216c4816667d1ba1667f8a718' hpatrain-tfrec-3
#'gs://kds-7977a755bc075ff2f47088aa134263c19820179fec133a5aa3e976a2' valid
KaggleDatasets().get_gcs_path('hpatrain-valtfrec')
KaggleDatasets().get_gcs_path('hpatrain-tfrec-1')
KaggleDatasets().get_gcs_path('hpatrain-tfrec-2')
KaggleDatasets().get_gcs_path('hpatrain-tfrec-3')
KaggleDatasets().get_gcs_path('hpavalid-tfrec')
GCS_DS_PATH=[]
GCS_DS_PATH.append(KaggleDatasets().get_gcs_path('hpatrain-valtfrec'))
GCS_DS_PATH
GCS_DS_PATH.append(KaggleDatasets().get_gcs_path('hpatrain-tfrec-1'))
GCS_DS_PATH.append(KaggleDatasets().get_gcs_path('hpatrain-tfrec-2'))
GCS_DS_PATH.append(KaggleDatasets().get_gcs_path('hpatrain-tfrec-3'))
GCS_DS_PATH.append(KaggleDatasets().get_gcs_path('hpasinglecell0png'))
GCS_DS_PATH
```
| github_jupyter |
# 重试
```
# from rtsp.rtsp_server import setup_server
#
# setup_server()
from retrying import retry
import random
import time
def retry_if_error(exception):
return exception.__class__ in {ValueError, TimeoutError}
def retry_if_not_result(result):
if result in [5, 10]:
print("result success {}".format(result))
return False
else:
print('result retry {}'.format(result))
return True
@retry(stop_max_attempt_number=50, wait_exponential_multiplier=1000, wait_exponential_max=60000,
retry_on_exception=retry_if_error)
def foo():
print(int(time.time()), end=' ')
a = random.randint(1, 15)
if a >= 2:
print('Error {}'.format(a))
raise ValueError
print('success', a)
@retry(retry_on_result=retry_if_not_result)
def foo2():
print(int(time.time()), end=' ')
a = random.randint(1, 15)
return a
# print(foo2())
foo()
```
# 超时控制
实际项目中会涉及到需要对有些函数的响应时间做一些限制,如果超时就退出函数的执行,停止等待。
可以利用python中的装饰器实现对函数执行时间的控制。
## Unix 信号机制
**限制:需要在linux系统上,并且必须在主线程中使用**
```
# 有回调函数的超时报错装饰器
import signal
import time
def set_timeout(num, callback):
def wrap(func):
def handle(
signum, frame
): # 收到信号 SIGALRM 后的回调函数,第一个参数是信号的数字,第二个参数是the interrupted stack frame.
raise RuntimeError
def to_do(*args, **kwargs):
try:
signal.signal(signal.SIGALRM, handle) # 设置信号和回调函数
signal.alarm(num) # 设置 num 秒的闹钟
print('start alarm signal.')
r = func(*args, **kwargs)
print('close alarm signal.')
signal.alarm(0) # 关闭闹钟
return r
except RuntimeError as e:
callback()
return to_do
return wrap
if __name__ == '__main__':
def after_timeout(): # 超时后的处理函数
print("函数超时")
@set_timeout(2, after_timeout) # 限时 2 秒
def connect(): # 要执行的函数
time.sleep(4) # 函数执行时间,写大于2的值,可测试超时
print('函数正常执行')
connect()
```
## func_timeout
>pip install func_timeout
```
import time
from func_timeout import func_set_timeout, FunctionTimedOut
@func_set_timeout(5)
def f():
while True:
print("1")
time.sleep(1)
if __name__ == '__main__':
try:
f()
except FunctionTimedOut:
print("timeout!")
```
## eventlet
>pip install eventlet
```
import requests
import eventlet
import time
eventlet.monkey_patch()
time_limit = 3 #set timeout time 3s
print(int(time.time()))
with eventlet.Timeout(time_limit, False):
time.sleep(5)
r = requests.get("https://me.csdn.net/dcrmg", verify=False)
print('ok')
print(int(time.time()))
print('over')
```
## Timer
- https://www.cnblogs.com/xueweihan/p/6653610.html
```
from threading import Timer
import time
def set_timeout(num):
def wrap(func):
def stop():
print(int(time.time()))
raise RuntimeError
def to_do(*args, **kwargs):
# n 秒后执行
t = Timer(num, stop)
t.start()
r = func(*args, **kwargs)
t.cancel()
return r
return to_do
return wrap
print(int(time.time()))
try:
@set_timeout(4)
def foo():
time.sleep(6)
except RuntimeError as e:
print('e', e)
foo()
```
| github_jupyter |
# Pandas fast mutate architecture
## Problem: users may need to define their own functions for SQL or pandas
In siuba, much of what users do involves expressions using `_`.
Depending on the backend they're using, these expressions are then transformed and executed.
However, sometimes no translation exists for a method.
This is not so different from pandas or SQL alchemy, where a limited number of methods are available to users.
For example, in pandas...
* you can do `some_data.cumsum()`
* you *can't* do `some_data.cumany()`
Moreover, you can use `.cummean()` on an ungrouped, but not a grouped DataFrame. And as a final cruel twist, some methods are fast when grouped, while others (e.g. `expanding().sum()`) use the slow apply route.
## What's the way out?
In pandas, it's not totally clear how you would define something like `.cumany()`, and let it run on grouped or ungrouped data, without **submitting a PR to pandas itself**.
(maybe by [registering an accessor](https://github.com/Zsailer/pandas_flavor#register-accessors), but this doesn't apply to grouped DataFrames.)
This is the tyranny of methods. The object defining the method owns the method. To add or modify a method, you need to modify the class behind the object.
Now, this isn't totally true--the class could provide a way for you to register your method (like accessors). But wouldn't it be nice if the actions we wanted to perform on data didn't have to check in with the data class itself? Why does the data class get to decide what we do with it, and why does it get priviledged methods?
### Enter singledispatch
Rather than registering functions onto your class (i.e. methods), singledispatch lets you register classes with your functions.
In singledispatch, this works by having the class of your first argument, decide which version of a function to call.
```
from functools import singledispatch
# by default dispatches on object, which everything inherits from
@singledispatch
def cool_func(x):
print("Default dispatch over:", type(x))
@cool_func.register(int)
def _cool_func_int(x):
print("Special dispatch for an integer!")
cool_func('x')
cool_func(1)
```
This concept is incredibly powerful for two reasons...
* many people can define actions over a DataFrame, without a quorum of priviledged methods.
* you can use normal importing, so don't have to worry about name conflicts
## singledispatch in siuba
siuba uses singledispatch in two places
* dispatching verbs like `mutate`, whose actions depend on the backend they're operating on (e.g. SQL vs pandas)
* creating symbolic calls
It's worth looking at symbolic calls in detail
```
from siuba.siu import symbolic_dispatch, _
import pandas as pd
@symbolic_dispatch(cls = pd.Series)
def add2(x):
return x + 2
add2(pd.Series([1,2,3]))
```
One special property of `symbolic_dispatch` is that if we pass it a symbol, then it returns a symbol.
```
sym = add2(_.astype(int))
sym
sym(pd.Series(['1', '2']))
```
Note that in this case these two bits of code work the same...
```python
ser = pd.Series(['1', '2'])
sym = add2(_.astype(int))
sym(ser)
func = lambda _: add2(_.astype(int))
func(ser)
```
siuba knows that if the function's first argument is a symbolic expression, then the function needs to return a symbolic expression.
## What should we singledispatch over
In essence, siuba needs to allow dispatching over the forms of data it can operate on, including..
* regular Series
* grouped Series
* (maybe) sqlachemy column mappings
## Are there any risks?
I'm glad you asked! There is one very big risk with singledispatch, and it's this:
singledispatch will dispatch on the "closest" matching parent class it has registered.
This means that if it has object registered, then at the very least, it will dispatch on that.
**This is a big problem since e.g. sqlalchemy column mappings and everything else is an object**.
In order to mitigate this risk, there are two compelling options...
1. Put an upper bound on dispatching classes ([related concept in type annotations](https://www.python.org/dev/peps/pep-0484/#type-variables-with-an-upper-bound))
2. Require an explicit annotation on return type
The downsides are that (1) requires a custom dispatch implementation, and (2) requires that people know about type annotations.
That said, I'm curious to explore option (2), as this has an appealing logic: an appropriate function will be a subtype of the one we typically use.
## Requiring an annotation over return type
In order to fully contextualize the process, consider the stage where something may need to be pulled from the dispatcher: call shaping via CallTreeLocal.
```
from siuba.siu import CallTreeLocal, strip_symbolic
def as_string(x):
return x.astype(str)
ctl = CallTreeLocal(local = {'as_string': as_string})
call = ctl.enter(strip_symbolic(_.as_string()))
# Call object holding function as first argument
call.__dict__
# proof it's just the function
type(call.args[0])
```
Now this setup is good and well--but how is a user going to put *their* function on CallTreeLocal?
Register it? Nah. What they need is a clear interface.
We're already "bouncing" symbolic dispatch functions when they get a symbolic expression. We can use this mechanic to make CallTreeLocal more "democratic"
Notice that when we "bounce" add2, it reports the function as a "__custom_func__".
```
@symbolic_dispatch(cls = pd.Series)
def add2(x):
return x + 2
add2(_)
```
This is because it's a special call, called a `FuncArg` (name subject to change). We can modify CallTreeLocal to perform custom behavior when it enters / exits `__custom_func__`.
```
class SpecialClass: pass
@add2.register(SpecialClass)
def _add2_special(x):
print("Wooweee!")
class CallTree2(CallTreeLocal):
# note: self.dispatch_cls already used in init for this very purpose
def enter___custom_func__(self, node):
# the function itself is the first arg
dispatcher = node.args[0]
# hardcoding for now...
return dispatcher.dispatch(self.dispatch_cls)
ctl2 = CallTree2({}, dispatch_cls = SpecialClass)
func = ctl2.enter(strip_symbolic(add2(_)))
func
type(func)
```
However, there's one major problem--CallTree2 may still dispatch the default function!
```
@symbolic_dispatch
def add3(x):
print("Calling add3 default")
call3 = ctl2.enter(strip_symbolic(add3(_)))
call3(1)
```
**THIS MEANS THAT EVERY SINGLEDISPATCH FUNCTION WILL AT LEAST USE ITS DEFAULT**
Imagine that some defined the default, but then it gets fired for SQL, and for pandas, etc etc..
What a headache.
### Keeping only when there's a compatible return type
We can check the result annotation of the function we'd dispatch, to know whether it won't. In this case, we assume it won't work if the result is not a subclass of the one our SQL tools expect: ClauseElement.
We can shut down the process early if we know the function won't return what we need.
This is because a function is a subtype of another function if it's input is contravarient (e.g. a parent), and **it's output is covariant (e.g. a subclass)**.
```
# used to get type info
import inspect
# the most basic of SQL classes
from sqlalchemy.sql.elements import ClauseElement
RESULT_CLS = ClauseElement
class CallTree3(CallTreeLocal):
# note: self.dispatch_cls already used in init for this very purpose
def enter___custom_func__(self, node):
# the function itself is the first arg
dispatcher = node.args[0]
# hardcoding for now...
f = dispatcher.dispatch(self.dispatch_cls)
sig = inspect.signature(f)
ret_type = sig.return_annotation
if issubclass(ret_type, RESULT_CLS):
return f
raise TypeError("Return type, %s, not subclass of %s" %(ret_type, RESULT_CLS))
from sqlalchemy import sql
sel = sql.select([sql.column('id'), sql.column('x'), sql.column('y')])
# this is what siuba sql expressions operate on
col_class = sel.columns.__class__
clt3 = CallTree3({}, dispatch_cls = col_class)
@symbolic_dispatch
def f_bad(x):
return x + 1
@symbolic_dispatch
def f_good(x: ClauseElement) -> ClauseElement:
return x.contains('woah')
# here is the error for the first, without that pesky stack trace
try:
clt3.enter(strip_symbolic(f_bad(_)))
except TypeError as err:
print(err)
# here is the good one going through
clt3.enter(strip_symbolic(f_good(_)))
```
### How do I get this in my life today?
Well, runtime evaluation of result types isn't the most fleshed out process in python. And there are some edge cases.
For example, what should we do if the return type is a [Union](https://www.python.org/dev/peps/pep-0484/#union-types)? [Any](https://www.python.org/dev/peps/pep-0484/#the-any-type)?
There is also a bug with the Union implementation before 3.7, where if it receives 3 classes, and 1 is the parent of the others, it just returns the parent...
```
from typing import Union
class A: pass
class B(A): pass
class C(B): pass
Union[A,B,C]
```
To be honest--I think we can be optimistic for now that anyone using a Union as their return type knows what they're doing with siuba. I think the main behaviors we want to support are...
1. Can create singledispatch, with potentially a default function
2. Don't shoot yourself in the foot when the default is fired for SQL and pandas
And even a crude result type check will ensure that. In some ways the existence of a result type is almost all the proof we need.
## To decide
* What should siuba do when dispatch function doesn't qualify? Fall back to local?
* Related: should local only look up methods? (makes sense to me)
* If so, how do we implement SQL dialects? Have ImmutableColumnCollection >= SqlColumns >= PostgresqlColumns, etc..
## Could siuba allow static type checking?
I think so. It would take a bit of work. Mostly PRs to the typing package to...
* Implement [higher-kinded types](https://www.stephanboyer.com/post/115/higher-rank-and-higher-kinded-types)
* Support static checking of singledispatch (or stubbing with @overload)
* Wait for pandas type annotations, or stub, so we can check the pipe, which uses `__rshift__` 😅
| github_jupyter |
```
import sys
sys.version
sys.version_info
import subprocess
```
## call example
```
# returns return code only
subprocess.call(["ls", "-lha"])
```
note that no exception is raised if the underlying command errors:
`bash-script-with-bad-syntax` is a shell script with bad syntax.
```
subprocess.call(["./bash-script-with-bad-syntax"])
```
## call() example using shell=True
```
subprocess.call("ls -lha", shell=True)
```
## call example capture standard output and error
```
subprocess.call(["./bash-script-with-bad-syntax"])
```
## call example, force exception if called process causes error
```
# if there's no error in the underlying process,
# this is just the same as subprocess.call
subprocess.check_call(["ls","-lha"])
# but unlike call, this throws a Called
subprocess.check_call(["./bash-script-with-bad-syntax"])
```
## call example, capture stfout and stderr
```
import subprocess
import sys
# create two files to hold the output and errors, respectively
with open('out.txt','w+') as fout:
with open('err.txt','w+') as ferr:
out=subprocess.call(["./bash-script-with-bad-syntax"],stdout=fout,stderr=ferr)
# reset file to read from it
fout.seek(0)
print('output:')
print(fout.read())
# reset file to read from it
ferr.seek(0)
print('error:')
print(ferr.read())
```
## store output in variable
```
output = subprocess.check_output(["ls","-lha"],universal_newlines=True)
print(output)
```
## run() examples
```
cp = subprocess.run(["ls","-lha"])
cp
cp = subprocess.run(["ls -lha"],shell=True)
cp
cp = subprocess.run(["ls","-lha"], universal_newlines=True,stdout=subprocess.PIPE, stderr=subprocess.PIPE)
print(cp)
cp = subprocess.run(["ls","foo bar"], stdout=subprocess.PIPE, stderr=subprocess.PIPE, universal_newlines=True)
print(cp.stdout)
print(cp.stderr)
subprocess.run(["ls","foo bar"], check=True)
try:
cp = subprocess.run(["xxxx","foo bar"], universal_newlines=True, stdout=subprocess.PIPE, stderr=subprocess.PIPE)
except FileNotFoundError as e:
print(e)
```
## popen
```
from subprocess import Popen
p = Popen(["ls","-lha"])
p.wait()
p = Popen(["ls","-lha"], stdout=subprocess.PIPE, stderr= subprocess.PIPE, universal_newlines=True)
output, errors = p.communicate()
print(output)
print(errors)
path_to_output_file = '/tmp/myoutput.txt'
myoutput = open(path_to_output_file,'w+')
p = Popen(["ls","-lha"], stdout=myoutput, stderr= subprocess.PIPE, universal_newlines=True)
output, errors = p.communicate()
print(output)
print(errors)
with open(path_to_output_file,"r") as f:
print(f.read())
path_to_output_file = '/tmp/myoutput.txt'
myoutput = open(path_to_output_file,'w+')
p = Popen(["ls","foo bar"], stdout=myoutput, stderr= myoutput, universal_newlines=True)
output, errors = p.communicate()
print(output)
print(errors)
with open(path_to_output_file,"r") as f:
print(f.read())
```
## pipe commands together
```
from subprocess import Popen,PIPE
# this is equivalent to ls -lha | grep "ipynb"
p1 = Popen(["ls","-lha"], stdout=PIPE)
p2 = Popen(["grep", "ipynb"], stdin=p1.stdout, stdout=PIPE, universal_newlines=True)
p1.stdout.close()
output = p2.communicate()[0]
print(output)
```
## async
```
import asyncio
proc = await asyncio.create_subprocess_exec(
'ls','-lha',
stdout=asyncio.subprocess.PIPE,
stderr=asyncio.subprocess.PIPE)
# if proc takes very long to complete,
# the CPUs are free to use cycles for
# other processes
stdout, stderr = await proc.communicate()
print('[return code: '+ str(proc.returncode) +']')
if stdout:
print('\n[stdout: ]\n'+str(stdout.decode()))
else:
print('stdout is empty')
if stderr:
print(f'\n[stderr]:\n'+str(stderr.decode()))
else:
print('stderr is empty')
```
| github_jupyter |
# Expected models on binned contact maps
* For intrachromosomal arm regions: P(s) by diagonal
* For interchromosomal regions: Average contact frequency by block
```
%matplotlib inline
import matplotlib.pyplot as plt
import matplotlib as mpl
import seaborn as sns
import multiprocess as mp
import numpy as np
import pandas as pd
import bioframe
import cooltools
import cooler
mm9 = bioframe.fetch_chromsizes('mm9')
chromsizes = bioframe.fetch_chromsizes('mm9')
chromosomes = list(chromsizes.index)
conditions = ['WT', 'T', 'dN']
binsize = 100000
cooler_paths = {
'WT' : f'data/UNTR.{binsize}.cool',
'T' : f'data/TAM.{binsize}.cool',
'dN' : f'data/NIPBL.{binsize}.cool',
}
long_names = {
'WT': 'Wildtype',
'T' : 'TAM',
'dN': 'NipblKO',
}
pal = sns.color_palette('colorblind')
colors = {
'WT': pal[0],
'T' : '#333333',
'dN': pal[2],
}
clrs = {
cond: cooler.Cooler(cooler_paths[cond]) for cond in conditions
}
# this cell takes a long time to run
from cooltools.expected import diagsum, blocksum_pairwise
supports = [(chrom, 0, chromsizes[chrom]) for chrom in chromosomes]
cis_exp = {}
trs_exp = {}
with mp.Pool() as pool:
for cond in conditions:
print(cond, 'cis')
tables = diagsum(
clrs[cond],
supports,
transforms={
'balanced': lambda p: p['count'] * p['weight1'] * p['weight2'],
},
chunksize=10000000,
ignore_diags=2,
map=pool.map)
cis_exp[cond] = pd.concat(
[tables[support] for support in supports],
keys=[support[0] for support in supports],
names=['chrom'])
cis_exp[cond]['balanced.avg'] = cis_exp[cond]['balanced.sum'] / cis_exp[cond]['n_valid']
cis_exp[cond].to_csv(f'data/{long_names[cond]}.{binsize//1000}kb.expected.cis.tsv', sep='\t')
print(cond, 'trans')
records = blocksum_pairwise(
clrs[cond],
supports,
transforms={
'balanced': lambda p: p['count'] * p['weight1'] * p['weight2'],
},
chunksize=10000000,
map=pool.map)
trs_exp[cond] = pd.DataFrame(
[{'chrom1': s1[0], 'chrom2': s2[0], **rec} for (s1, s2), rec in records.items()],
columns=['chrom1', 'chrom2', 'n_valid', 'count.sum', 'balanced.sum'])
trs_exp[cond].to_csv(f'data/{long_names[cond]}.{binsize//1000}kb.expected.trans.tsv', sep='\t', index=False)
cis_exp['WT'].head()
trs_exp['WT'].head()
stats = {}
for cond in conditions:
n_cis = int(cis_exp[cond]['count.sum'].sum())
n_trs = int(trs_exp[cond]['count.sum'].sum())
stats[long_names[cond]] = {
'cis': n_cis,
'trans': n_trs,
'total': n_cis + n_trs,
'cis:trans': n_cis / n_trs,
'cis:total': n_cis / (n_cis + n_trs)
}
pd.DataFrame.from_dict(stats, orient='index')
sums = {}
n_valid = {}
scalings = {}
for cond in conditions:
grouped = cis_exp[cond].groupby('diag')
n_valid[cond] = grouped['n_valid'].sum().values
sums[cond] = grouped['balanced.sum'].sum().values
scalings[cond] = (sums[cond] / n_valid[cond])
from cooltools.lib import numutils
def coarsen_geometric(sums, counts, n_bins=100):
"""Re-bin the expected sums into logarithmically growing bins.
"""
dbins = numutils.logbins(1, len(sums), N=n_bins)
spans = list(zip(dbins[:-1], dbins[1:]))
s = np.array([np.nansum(sums[lo:hi]) for lo, hi in spans])
n = np.array([np.nansum(counts[lo:hi]) for lo, hi in spans])
return dbins, s / n
gs = plt.GridSpec(nrows=2, ncols=1, height_ratios=[10, 2])
plt.figure(figsize=(8, 15))
SMOOTH = False
ax1 = plt.subplot(gs[0])
for i, cond in enumerate(conditions):
ref_point = 200000 // binsize
norm = 1 #scalings[cond][ref_point]
# cis P(s)
x = np.arange(0, len(scalings[cond]) * binsize, binsize)
y = scalings[cond] / norm
if SMOOTH:
x, y = coarsen_geometric(sums[cond], n_valid[cond], 100)
x *= binsize
x = x[:-1]
plt.plot(x, y,
color=colors[cond],
label=long_names[cond])
# average trans levels
for _, row in trs_exp[cond].iterrows():
plt.axhline(
(row['balanced.sum']/row['n_valid']) / norm,
xmin=i/len(conditions),
xmax=(i+1)/len(conditions),
c=colors[cond],
alpha=0.25)
plt.xscale('log')
plt.yscale('log')
plt.ylabel('P(s)')
plt.xlabel('separation, bp')
plt.legend()
plt.gca().set_aspect(1)
xlim = plt.xlim()
```
| github_jupyter |
# Cross-validation and hyperparameter tuning
In the previous notebooks, we saw two approaches to tune hyperparameters:
via grid-search and randomized-search.
In this notebook, we will show how to combine such hyperparameters search
with a cross-validation.
## Our predictive model
Let us reload the dataset as we did previously:
```
from sklearn import set_config
set_config(display="diagram")
import pandas as pd
adult_census = pd.read_csv("../datasets/adult-census.csv")
```
We extract the column containing the target.
```
target_name = "class"
target = adult_census[target_name]
target
```
We drop from our data the target and the `"education-num"` column which
duplicates the information from the `"education"` column.
```
data = adult_census.drop(columns=[target_name, "education-num"])
data.head()
```
Once the dataset is loaded, we split it into a training and testing sets.
```
from sklearn.model_selection import train_test_split
data_train, data_test, target_train, target_test = train_test_split(
data, target, random_state=42)
```
We will create the same predictive pipeline as seen in the grid-search
section.
```
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OrdinalEncoder
from sklearn.compose import make_column_selector as selector
categorical_columns_selector = selector(dtype_include=object)
categorical_columns = categorical_columns_selector(data)
categorical_preprocessor = OrdinalEncoder(handle_unknown="use_encoded_value",
unknown_value=-1)
preprocessor = ColumnTransformer([
('cat-preprocessor', categorical_preprocessor, categorical_columns)],
remainder='passthrough', sparse_threshold=0)
# for the moment this line is required to import HistGradientBoostingClassifier
from sklearn.experimental import enable_hist_gradient_boosting
from sklearn.ensemble import HistGradientBoostingClassifier
from sklearn.pipeline import Pipeline
model = Pipeline([
("preprocessor", preprocessor),
("classifier",
HistGradientBoostingClassifier(random_state=42, max_leaf_nodes=4))])
model
```
## Include a hyperparameter search within a cross-validation
As mentioned earlier, using a single train-test split during the grid-search
does not give any information regarding the different sources of variations:
variations in terms of test score or hyperparameters values.
To get reliable information, the hyperparameters search need to be nested
within a cross-validation.
<div class="admonition note alert alert-info">
<p class="first admonition-title" style="font-weight: bold;">Note</p>
<p class="last">To limit the computational cost, we affect <tt class="docutils literal">cv</tt> to a low integer. In
practice, the number of fold should be much higher.</p>
</div>
```
from sklearn.model_selection import cross_validate
from sklearn.model_selection import GridSearchCV
param_grid = {
'classifier__learning_rate': (0.05, 0.1),
'classifier__max_leaf_nodes': (30, 40)}
model_grid_search = GridSearchCV(model, param_grid=param_grid,
n_jobs=2, cv=2)
cv_results = cross_validate(
model_grid_search, data, target, cv=3, return_estimator=True)
```
Running the above cross-validation will give us an estimate of the
testing score.
```
scores = cv_results["test_score"]
print(f"Accuracy score by cross-validation combined with hyperparameters "
f"search:\n{scores.mean():.3f} +/- {scores.std():.3f}")
```
The hyperparameters on each fold are potentially different since we nested
the grid-search in the cross-validation. Thus, checking the variation of the
hyperparameters across folds should also be analyzed.
```
for fold_idx, estimator in enumerate(cv_results["estimator"]):
print(f"Best parameter found on fold #{fold_idx + 1}")
print(f"{estimator.best_params_}")
```
Obtaining models with unstable hyperparameters would be an issue in practice.
Indeed, it would become difficult to set them.
In this notebook, we have seen how to combine hyperparameters search with
cross-validation.
| github_jupyter |
```
import sys
import numpy as np
import pandas as pd
import scipy.sparse
import tqdm
import xswap
sys.path.insert(0, '../')
import analysis
n_perms = 1000
allow_antiparallel = False
allow_self_loops = False
directed = False
```
# 2. BioRxiv network
```
biorxiv_df = pd.read_csv('../../../data/task3/3.all_nodes/biorxiv.tsv.xz', sep='\t', compression='xz')
full_features_df = pd.DataFrame()
for network in ['train', 'test_recon', 'test_new']:
edges = list(map(tuple,
biorxiv_df
.query(f'{network} == 1')
.loc[:, ['id_a', 'id_b']]
.values
))
mat = analysis.edges_to_matrix(edges, directed=directed).tocsc()
print(network, mat.shape)
degree_matrix = np.repeat(mat.sum(axis=1), mat.shape[1], axis=1) \
+ np.repeat(mat.sum(axis=0), mat.shape[0], axis=0)
# RWR is fastest on this network with exact matrix inverse on dense array
feature_dict = {
'edge_prior': scipy.sparse.csc_matrix(mat.shape),
'rwr': analysis.rwr_approx_inv(mat, 0.25, n_iter=10),
'mean_rwr': scipy.sparse.csc_matrix(mat.shape),
'p_rwr': scipy.sparse.csc_matrix(mat.shape),
'jaccard': analysis.jaccard(mat, degree_matrix),
'mean_jaccard': np.zeros(mat.shape),
'p_jaccard': np.zeros(mat.shape),
}
perm_edges = edges.copy()
for i in tqdm.tnrange(n_perms):
perm_edges, _ = xswap.permute_edge_list(
perm_edges,
allow_self_loops=allow_self_loops,
allow_antiparallel=allow_antiparallel,
seed=i
)
perm_mat = analysis.edges_to_matrix(perm_edges, directed=directed).tocsc()
feature_dict['edge_prior'] += perm_mat
perm_rwr = analysis.rwr_approx_inv(perm_mat, 0.25, n_iter=10)
feature_dict['mean_rwr'] += perm_rwr
feature_dict['p_rwr'] += (perm_rwr >= feature_dict['rwr'])
perm_jaccard = analysis.jaccard(perm_mat, degree_matrix)
feature_dict['mean_jaccard'] += perm_jaccard
feature_dict['p_jaccard'] += (perm_jaccard >= feature_dict['jaccard'])
# Post-process features
network_features_df = None
for feature, array in feature_dict.items():
# Normalize features by the number of permutations
if feature in ['edge_prior', 'mean_rwr', 'p_rwr', 'mean_jaccard', 'p_jaccard']:
feature_dict[feature] /= n_perms
# Make features dense (for DataFrame)
if scipy.sparse.issparse(array):
array = array.toarray()
feature_df = (
pd.DataFrame(array)
.reset_index()
.melt(id_vars=['index'])
.rename(columns={'index': 'id_a', 'variable': 'id_b', 'value': feature})
.assign(
id_a=lambda df: df['id_a'].astype(int),
id_b=lambda df: df['id_b'].astype(int),
network=network,
)
.query('id_a <= id_b')
)
if network_features_df is None:
network_features_df = feature_df
else:
network_features_df = network_features_df.merge(feature_df, on=['id_a', 'id_b', 'network'])
full_features_df = pd.concat([full_features_df, network_features_df])
%%time
biorxiv_features_df = (
biorxiv_df
.melt(
id_vars=['id_a', 'id_b'],
value_vars=['train', 'test_recon', 'test_new'],
var_name='network', value_name='edge'
)
.merge(full_features_df, on=['id_a', 'id_b', 'network'], how='left')
)
biorxiv_features_df.to_csv('../../../data/task3/4.data/biorxiv.tsv.xz', sep='\t', compression='xz', index=False)
```
| github_jupyter |
Introduction to Unsupervised Learning
======
In earlier labs we learned how to use linear regression to study whether certain features are useful in predicting an observed outcome. Then we used ensemble methods to refine our predictions.
In this notebook, we shift from prediction to pattern finding.
What we'll be doing in this notebook:
-----
1. Give a general introduction to unsupervised learning.
1. Use k-means clustering as unsupervised learning technique.
1. Load and explore a dataset.
1. Find clusters with k-means algorithm.
1. Evaluate our results with the Elbow method.
Unsupervised Learning: Pattern finding in data 🔍
------
Unsupervised Learning is the process of identifying patterns in a dataset. Identifying patterns is often an early step in understanding data. Unsupervised learning methods are a set of techniques designed to _explore_ and find "hidden structure" rather than predict outcomes.
Unsupervised learning does not require labeled data, therefore works for broader range of data. In fact, most data in the world is unlabelled. However, since there are no labels / correct answers there is not always a clear feedback to validate that the results are correct.
Unsupervised Learning is also called Data Mining.
Unsupervised
------
2 Types of Unsupervised Learning
--------
1. Dimension Reduction
1. Clustering
What is Dimension Reduction?
------
Dimension reduction aims to find fewer number of features that be used to build a meaningful model. There are many reasons for reducing the number of features in a dataset, from avoiding overfitting to speeding up modeling fitting time.
One of the most common dimension reduction techniques is Principal Component Analysis (PCA).
What is Clustering?
-----
<br>
<center><img src="./images/clustering.png" width="700"/></center>
Clustering is what it sounds like: chunking your data into sub-groups (clusters) based on similar characteristics. Then these sub-groups are used for later analysis. Clustering is an intuitive to understand the various natural segments that make up the population of your data. Clustering typically makes it easier to visualizes your data.
Clustering is also called [cluster analysis](https://en.wikipedia.org/wiki/Cluster_analysis), data segmentation, or data partitioning.
We are going to focus on clustering for the rest of this notebook.
Introduction to K-means Clustering
------
<center><img src="./images/k_means.png" width="700"/></center>
K-means one of the most common clustering techniques. The goal of k-means is find a group of datapoints that close to each other (a cluster) and are far away from other datapoints (the other clusters).
How do we do k-means clustering?
-----
Initially, datapoints are <i>randomly assigned</i> to a cluster. Then the center of each cluster is calculated.
Then we alternate between two steps:
1. Assignment step: Observations are assigned to a cluster where the center is closest to them.
2. Update step: New center points of clusters are determined
The process repeats until observations shuffle are long around to different clusters anymore and the center of each cluster no longer moves.
In other words, observations are constantly being reassigned to clusters until the distance between an observation and their closest center point is minimized.
K-means Example
-----

-----
Fitting K-means to Kiva Data
------
Now we are going to fit k-means to <b>partition</b> or <b>segment</b> the Kiva data into clusters.
Let's import the relevant packages to start coding:
```
# Data loading and manipulation
import pandas as pd
import numpy as np
# K-Means clustering algorithm
from sklearn.cluster import KMeans
# Plotting
import seaborn as sns
# Places the plots in the Jupyter Notebook
%matplotlib inline
# Keep everything visible
pd.set_option('display.max_columns', 80)
pd.set_option('expand_frame_repr', True)
```
----
Load and explore the data
-----
```
# Load data
path = '../data/'
filename = 'loans.csv'
df = pd.read_csv(path+filename)
```
It is always good idea 💡 to take a peak at the raw data.
```
df.head(n=2)
print(f"There are {df.shape[1]:,} columns in the dataframe.")
print(f"There are {df.shape[0]:,} rows in the dataframe.")
df.describe()
```
Now that we have our data set up, we can begin partitioning our data into clusters based only a few features. Let's think about how to choose these…
As a potential borrower or lender, what would be interesting to explore?
In the previous notebooks, we explored a number of interesting ideas, including:
- How much a borrower should ask to borrow
- The time it takes to fund a loan
- What features can influence the loan amount
- If we partition borrowers into distinct groups based on how quickly they can fund a loan, will we be able to learn anything about these groups of borrowers?
The k-means algorithm uses continuous-valued numeric features (k-means can also be modified to work with categorical and ordinal features).
-----
Let's pick a couple of interesting continuous numeric features for analysis:
How are Funded Amount and Days to Fund related?
----
```
# Keep only the relevant columns
column_1 = 'funded_amount'
column_2 = 'repayment_term'
df = df[[column_1, column_2]]
ax = df.funded_amount.hist(grid=False);
ax.set(xlabel='Funded Amount',
ylabel='Count',
title='Histogram of Funded Amount');
```
_How can we interpret the number of loans at different funding amounts?_
<br>
<br>
<br>
```
ax = df.repayment_term.hist(grid=False);
ax.set(xlabel='Repayment Term',
ylabel='Count',
title='Histogram of Repayment Term');
```
_How can we interpret the number of loans for different amount of times?_
<br>
<br>
<br>
```
# Filter data to remove outliers
funded_small = df.funded_amount < 2_500 # Remove large loans
repayment_short = df.repayment_term < 60 # Remove long repayment terms
df = df[funded_small & repayment_short]
ax = df.funded_amount.hist(grid=False);
ax.set(xlabel='Funded Amount',
ylabel='Count',
title='Histogram of Funded Amount');
print(f"There are {df.shape[1]:,} columns in the dataframe.")
print(f"There are {df.shape[0]:,} rows in the dataframe.")
# Plot the relationship between these two variables
df.plot.scatter(x=column_1,
y=column_2);
```
_How can we interpret the relationship between the funded amount and time to fund?_
<br>
<br>
<br>
Clustering
======
----
Fitting our data with k-means using scikit-learn
----
Now we're ready to run the k-means algorithm:
Let's take quick peek at the [scikit-learn's documentation](http://scikit-learn.org/stable/modules/generated/sklearn.cluster.KMeans.html)
```
# Take KMeans class, initialize and fit it our data
kmeans = KMeans(n_clusters=2) # Number of clusters should be 2 or 3
kmeans.fit(df);
```
Now that we have clusters, the best way to understand them is to visualize them
```
# Add the cluster labels for each data point to the dataframe
df['kmeans_labels'] = kmeans.labels_
# Plot k-means
kmeans_plot = sns.lmplot(x=column_1,
y=column_2,
data=df,
fit_reg=False, # Do not fit a regression line to the data
hue="kmeans_labels", #'hue' will color code each group
legend=True);
# Plot the mean of cluster #1
kmeans_plot.ax.plot(kmeans.cluster_centers_[0][0], kmeans.cluster_centers_[0][1], color='red', marker='*');
# Plot the mean of cluster #2
kmeans_plot.ax.plot(kmeans.cluster_centers_[1][0], kmeans.cluster_centers_[1][1], color='cyan', marker='*');
# # Plot the mean of cluster #3 (if present)
# kmeans_plot.ax.plot(kmeans.cluster_centers_[2][0], kmeans.cluster_centers_[2][1], color='orange', marker='*');
```
_Why are the means where they are?_
<br>
<br>
<br>
Choosing the number of clusters
-----
The k-means algorithm is somewhat naive -- it clusters the data into k clusters, even if k is not the right number of clusters to use.
We arbitrarily set the number of clusters to be 3. But determining the appropriate number of clusters (k) is actually one of the most challenging parts of clustering.
There is no hard and fast rule for what the value of k should be because the number of clusters will depend on your data and what the goal of your analysis. The number of groups you choose to partition in your data directly influences the results you'll find. In most areas of data analysis, it's attractive to take as granular an approach as possible, but having too many clusters can be counterproductive because the grouping will not tell you much.
_Is it possible to have too many clusters? Or too few clusters?_
Think about these extreme examples:
1. A single cluster for all your data
2. A cluster for each data point
Neither of these will tell you anything new about your data!
Rather, clustering is most effective when observations in the same cluster are very similar to each other. Also, we want observations in different clusters to be as different from each other as possible.
Elbow method to explore number of clusters
------
The elbow method is a simple, intuitive way of exploring how changing the number of clusters impacts the "tightness" of the clusters.
The elbow method runs k-means clustering on the same dataset for a range of values of k (say, k is [1, 2, 3, 4, 5, 6, 7, 8, 9, 10]) and for each value of k, calculate the within-cluster sum-of-squares errors (SSE).
SSE is the distance as-the-crow* flies between each point and closest mean, squared, and summed.
SSE is a measure of internally coherent clusters are. A lower is SSE is better (an inverted score), it means the each cluster is very similar to itself. SSE is like a golf score or heart-rate, lower is better.
As k increases, the improvement in SSE () will lesson. At some point this lack of improvement will be rapid, creating the "elbow" shape.
One should choose a number of clusters so that adding another cluster doesn't give much better modeling of the data.
<sub>*The figure does not look like as-the-crow distance because each axis is on very different scales. Typically, k means data is normalized so data is on the same standard scale.</sub>

What the elbow method does is this:
1. Run the k-means algorithm over your dataset for a range of k.
2. For each value of k, calculate how the model fits.
3. If we see an "elbow" in our plotted check, then that marks a good value for k.
```
# Let's fit a different model for each value of k
k_values = range(1, 10)
# Fit a model for each value of k
k_mean_models = [KMeans(n_clusters=i) for i in k_values]
# See how the scores change
scores = [-k_mean_models[i].fit(df).score(df)
for i, model in enumerate(k_mean_models)]
# Let's plot the effect k on the clustering
ax = sns.pointplot(x=list(k_values),
y=scores);
ax.set(xlabel='k',
ylabel='Fit',
title='The Elbow Method choosing k');
```
_How can we interpret the relationship between changing k and the fit of clustering?_
_Can we see where the "bend" is that looks like an elbow in an arm?_
<br>
<br>
<br>
Summary
------
- We talked about how supervised learning finds patterns in data.
- Clustering is finding groups within a dataset.
- K-means clustering is a popular clustering technique that iteratively finds the best groups and center/means of groups.
- We fit k-means to data and evaluated the results.
Next Steps
-----
Apply the K-means clustering algorithm to a new pair of features.
Then find an useful number of clusters (k).
Further study
-----
If you want to understand k-means at a deeper level, start at the notebook found [here](https://jakevdp.github.io/PythonDataScienceHandbook/05.11-k-means.html)
If you are interested in a more theory behind k-means, a great resource is [here](https://www-users.cs.umn.edu/~kumar/dmbook/ch8.pdf).
There are many other clustering methods. Another popular method is [hierarchical clustering](https://en.wikipedia.org/wiki/Hierarchical_clustering).
<br>
<br>
<br>
----
| github_jupyter |
# Protecting against Cross-Site Scripting
This notebook provides an example to showcase the methods we use to protect against Cross-Site Scripting (XSS).
In particular, to protect against it we escape some symbols in the JSON output and we add some extra headers which further ensure that the browser won't misidentify the content.
* **JSON serialiser.** We escape any HTML symbols on the output, using their unicode sequences instead.
* **Extra headers.** On every response, we set the `X-Content-Type-Options: nosniff;` header, which ensures that the browser won't try to guess the `Content-Type` from the content.
## Setup
Before showing a couple examples on how the output is modified to protect against XSS attacks, we will setup the environment.
### Build engine image
To make sure we are running the latest version of the engine, we will build a docker image from the current code.
Note that this requires a valid JDK installation.
```
!cd ../../../engine && make build_image
```
### Create k8s cluster
Firstly, we will create a cluster using [kind](https://kind.sigs.k8s.io).
```
!kind create cluster
!export KUBECONFIG="$(kind get kubeconfig-path --name=kind)"
```
## Setup Seldon Core
Use the setup notebook to [Setup Cluster](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Setup-Cluster) with [Ambassador Ingress](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Ambassador). Instructions [also online](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html).
Next, **before installing `seldon-core`**, we load the engine image we have just built above into the cluster.
```
!kind load docker-image seldonio/engine:$(cat ../../../engine/target/version.txt)
```
We can now install `seldon-core` on the new cluster, making sure that it uses the engine image local to the nodes.
```
!helm install seldon-core \
../../../helm-charts/seldon-core-operator \
--namespace seldon-system \
--set engine.image.pullPolicy=Never \
--set usagemetrics.enabled=true \
--set ambassador.enabled=true
!kubectl rollout status statefulset.apps/seldon-operator-controller-manager -n seldon-system
```
### Dummy Model
To test how `seldon-core` processes the output to prevent XSS attacks we will use a dummy model which just replies with whatever input we send.
The code for this model can be seen below.
```
!pygmentize ./XSSModel.py
```
Firstly, we will build an appropiate image using `s2i`.
The name of this image will be `xss-model:0.1`.
```
!make build_image
```
We are now ready to spin up a service running our model.
Note that before, we need to load the image into our `kind` cluster.
```
!kind load docker-image xss-model:0.1
!pygmentize ./xss-example.json
!kubectl apply -f ./xss-example.json
```
To visualise what the model does and verify that everything is working we can make an example request using `curl`.
Note that, on the request we are passing a string field as `{"strData": "hello world"}`.
On the output, we receive the same field after being returned as-is by `XSSModel`.
```
!curl \
-X POST \
-H 'Content-Type: application/json' \
-d '{"strData": "hello world"}' \
localhost:8003/seldon/default/xss-example/api/v0.1/predictions
```
## Checking the response
### JSON serialiser
To showcase the escaping of HTML characters in the JSON output, we will submit a HTML payload in our request.
Note that the output uses the corresponding unicode value, instead of the sensitive character.
This helps to avoid undesired behaviour when the output could be mis-interpreted as HTML.
```
!curl \
-X POST \
-H 'Content-Type: application/json' \
-d '{"strData": "<div class=\"box\">This is a div</div>"}' \
localhost:8003/seldon/default/xss-example/api/v0.1/predictions
```
We can also verify that the output for anything else remains untouched.
```
!curl \
-X POST \
-H 'Content-Type: application/json' \
-d '{"strData": "Not HTML!"}' \
localhost:8003/seldon/default/xss-example/api/v0.1/predictions
!curl \
-X POST \
-H 'Content-Type: application/json' \
-d '{"data": {"ndarray": [0, 1, 2, 3, 4]}}' \
localhost:8003/seldon/default/xss-example/api/v0.1/predictions
```
### Extra headers
Similarly, we can show the response headers, to see that the `X-Content-Type-Options` header is included in the response.
This header will avoid the browser trying to infer the content type and trusting the already sent `Content-Type` header instead.
```
!curl \
-X POST \
-sD - -o /dev/null \
-H 'Content-Type: application/json' \
-d '{"strData": "<div class=\"box\">This is a div</div>"}' \
localhost:8003/seldon/default/xss-example/api/v0.1/predictions
```
### Using `SeldonClient`
To verify everything else still works as expected, we can use the `SeldonClient` to check that the responses are still interpreted correctly.
```
from seldon_core.seldon_client import SeldonClient
sc = SeldonClient(deployment_name='xss-example', namespace="default")
r = sc.predict(gateway='ambassador', str_data="<div class=\"box\">This is a div</div>")
print(r)
```
As we can see above, even though the output is now escaped, `SeldonClient` parses the `utf8`-encoded elements into their actual characters.
Therefore, using the client, the change is transparent.
| github_jupyter |
# Multi-Tissue NODDI: Including S0 responses in regular MC models
It is well known that different tissues in the brain have different T1, T2 times and proton densities. This causes the contrast that can be seen in any usual b0 image, as the bone below.
```
from dmipy.data import saved_data
scheme_hcp, data_hcp = saved_data.wu_minn_hcp_coronal_slice()
import matplotlib.pyplot as plt
import matplotlib.patches as patches
%matplotlib inline
fig, ax = plt.subplots(1)
ax.imshow(data_hcp[:, 0, :, 0].T, origin=True)
ax.set_axis_off()
ax.set_title('HCP coronal slice B0 with ROI');
```
Multi-tissue models, such as Multi-Tissue CSD *(Jeurissen et al. 2014)*, include an estimate of the S0 value of each compartment in the volume fraction estimation. In Dmipy, go further and allow any MC-model (regular MC, MC-SM or MC-SH) to have S0 tissue responses associated with the input models. Generally, this can be described as
\begin{align}
\textbf{p}^*(\textbf{x})=&\text{argmin}_{\textbf{p}}\int \left[S(\textbf{x},\textbf{A})-\hat{S}^{\text{MT}}(\textbf{A},\textbf{p})\right]^2d\textbf{A},\\
&\text{with}\,\,\hat{S}^{\text{MT}}(\textbf{A},\textbf{p})=\sum_i^NS_{0,i}f_iC_i(\textbf{A},\textbf{p}_i).
\end{align}
In this example, we demonstrate Dmipy's generalized multi-tissue modeling by implemented *Multi-Tissue NODDI*. Looking at the equation, it is clear that standard multi-compartment modeling is the same as assuming the $S_0$ of each compartment is the same and equal to the $S_0$ of the measured signal. Including the good estimate of $S_{0,i}$ will correct volume fractions in areas where compartments partial volume with different $S_0$ responses.
We will show that correcting for the differences in NODDI between CSD and white matter $S_0$ responses makes signficant different in estimated volume fractions.
# Using Dmipy to set up the Multi-Tissue NODDI-Watson model
### Recovering an estimate of the WM and CSF S0 tissue responses
First we get an estimate of the $S_0$ responses of the different tissue types, so that we can include it in the NODDI model.
```
from dmipy.tissue_response.three_tissue_response import three_tissue_response_dhollander16
[S0_wm, S0_gm, S0_csf], _, _ = three_tissue_response_dhollander16(
scheme_hcp, data_hcp, wm_algorithm='tournier07',
wm_N_candidate_voxels=150, gm_perc=0.2, csf_perc=0.4)
print 'white matter S0 response: {}'.format(S0_wm)
print 'CSF S0 response: {}'.format(S0_csf)
```
Notice that the $S_0$ response of the CSF is three times higher than that of white matter.
### Setting up Multi-Tissue NODDI
We set up the NODDI-Watson model as usual first.
```
from dmipy.signal_models import cylinder_models, gaussian_models
from dmipy.distributions.distribute_models import SD1WatsonDistributed
ball = gaussian_models.G1Ball()
stick = cylinder_models.C1Stick()
zeppelin = gaussian_models.G2Zeppelin()
watson_dispersed_bundle = SD1WatsonDistributed(models=[stick, zeppelin])
watson_dispersed_bundle.set_tortuous_parameter('G2Zeppelin_1_lambda_perp','C1Stick_1_lambda_par','partial_volume_0')
watson_dispersed_bundle.set_equal_parameter('G2Zeppelin_1_lambda_par', 'C1Stick_1_lambda_par')
watson_dispersed_bundle.set_fixed_parameter('G2Zeppelin_1_lambda_par', 1.7e-9)
```
But, when instantiating the MC-model we can now also give the `S0_tissue_responses` that we previously estimated.
```
from dmipy.core.modeling_framework import MultiCompartmentModel
NODDI_mod = MultiCompartmentModel(
models=[ball, watson_dispersed_bundle],
S0_tissue_responses=[S0_csf, S0_wm]) # notice one value for each model.
```
The last thing is to fix the diffusivity of the Ball compartment to static values.
```
NODDI_mod.set_fixed_parameter('G1Ball_1_lambda_iso', 3e-9)
```
We can visualize the model as follows, but nothing on the composition changed, only the $S_0$ values associated with the top-level models.
```
from IPython.display import Image
NODDI_mod.visualize_model_setup(view=False, cleanup=False)
Image('Model Setup.png')
```
# Fitting Multi-Tissue NODDI to Human Connectome Project data
Fitting is then the same as usual. But notice there is a second optimization happening for the multi-tissue fractions. This is because correcting for the tissue-specific $S_0$ values is independent from the estimation of the non-linear parameters of the original NODDI model.
```
NODDI_fit_hcp = NODDI_mod.fit(
scheme_hcp, data_hcp, mask=data_hcp[..., 0]>0)
```
## Compare volume fractions with and without including S0 responses
Once fitted, the fitted model has access to both the original and the multi-tissue fractions. Here, we can easily compare them and illustrate where the differences at the interfaces are significant.
```
fractions_csf = NODDI_fit_hcp.fitted_parameters['partial_volume_0']
fractions_wm = NODDI_fit_hcp.fitted_parameters['partial_volume_1']
mt_fractions_csf = NODDI_fit_hcp.fitted_multi_tissue_fractions_normalized['partial_volume_0']
mt_fractions_wm = NODDI_fit_hcp.fitted_multi_tissue_fractions_normalized['partial_volume_1']
fig, axs = plt.subplots(2, 3, figsize=[15, 8])
axs = axs.ravel()
opts = {'origin': True, 'interpolation': 'nearest'}
im = axs[0].imshow(fractions_csf.squeeze().T, vmin=0, vmax=1, **opts)
fig.colorbar(im, ax=axs[0], shrink=0.7)
axs[0].set_title('f_csf no S0')
im = axs[1].imshow(mt_fractions_csf.squeeze().T, vmin=0, vmax=1, **opts)
fig.colorbar(im, ax=axs[1], shrink=0.7)
axs[1].set_title('f_csf with S0')
im = axs[2].imshow((mt_fractions_csf - fractions_csf).squeeze().T, **opts)
fig.colorbar(im, ax=axs[2], shrink=0.7)
axs[2].set_title('(f_csf with S0) - (f_csf no S0)')
im = axs[3].imshow(fractions_wm.squeeze().T,vmin=0, vmax=1, **opts)
fig.colorbar(im, ax=axs[3], shrink=0.7)
axs[3].set_title('f_wm no S0')
im = axs[4].imshow(mt_fractions_wm.squeeze().T, vmin=0, vmax=1, **opts)
fig.colorbar(im, ax=axs[4], shrink=0.7)
axs[4].set_title('f_wm with S0')
im = axs[5].imshow((mt_fractions_wm - fractions_wm).squeeze().T, **opts)
fig.colorbar(im, ax=axs[5], shrink=0.7)
axs[5].set_title('(f_wm with S0) - (f_wm no S0)')
for ax in axs:
ax.set_axis_off()
```
## Discussion of results
As you can see, including the estimated $S_0$ responses for CSF and WM for the corresponding models makes a significant difference in the estimated volume fractions. In fact, by including an estimate of the S0 response, we are in earnest estimated a **volume** fraction giving a certain signal contribution, rather than just the **signal** fraction contribution.
As the CSF $S_0$ response is about 3x higher than that of WM, that means that the same volume of CSF tissue produces 3x more signal to the $b_0$ than the same volume of WM. By (approximately) correcting for this bias, we can see that at the interfaces between CSF and WM, the actual volume fraction can be more than 0.3 higher than estimated without the $S_0$ response. In practice, this means that the white matter bundles such as the corpus callosum grow a little more into the CSF volume, making them bigger.
In conclusion, any MC-model that models tissues with different $S_0$ responses should include an estimate of the values -- if possible. Otherwise, the estimated volume fractions will **always** be biased in the direction of the difference of the tissues' $S_0$ responses.
## References
- Zhang, Hui, et al. "NODDI: practical in vivo neurite orientation dispersion and density imaging of the human brain." Neuroimage 61.4 (2012): 1000-1016.
- Jeurissen, Ben, et al. "Multi-tissue constrained spherical deconvolution for improved analysis of multi-shell diffusion MRI data." NeuroImage 103 (2014): 411-426.
| github_jupyter |
### 008 DataMaker
This is a notebook to make a data sets for the 008 relationship plots for the tutorial and the in-class exercise. It makes:
#### two related numeric variables
* linear relationship for tutorial
* non-linear " " "
* linear for exercise
* non-lin for exercise
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
```
#### Data for tutorial
Specity whatever filenames you want:
```
fname1 = "008TutorialDataFile1.csv" # linear, linear x
fname2 = "008TutorialDataFile2.csv" # correlated, Gaussian x
fname3 = "008TutorialDataFile3.csv" # x^2 term, linear x
```
### Make some linearly related data.
Define size of data.
```
n = 100 # number of observations
nv = 2 # number of variables
xmin = 0 # minimum x value
xmax = 100
```
#### Make the numerical data (i.e. the measurements).
##### First the x axis.
It turns out that np.linspace() and np.arange() produce "vectors" **not** 1xn or nx1 arrays. **And** you cannot then concatenate or multiply it with an array; numpy won't coerce it into having a second dimension like MATLAB does. You have to explicitly add the dimension of length one. This took me over two agonizing and frustating hours to work out.
```
x = np.linspace(xmin, xmax, n)
x = x[:, np.newaxis] # makes a [n,1] array from a dimensionless vector
```
##### Now the y values
```
m = 4.2
b = 250
noysAmp = 42
noys = noysAmp*np.random.randn(n, 1)
y = b + m*x + noys
```
##### Concatenate into nx2 ndarray
```
myData = np.hstack((x,y))
np.shape(myData)
```
##### Convert the numpy array into a pandas data frame:
```
myDataFr = pd.DataFrame(myData, columns=["X", "Y"])
myDataFr.plot(x="X", y="Y", kind="scatter")
```
Save the data frame to a .csv (comma separated values) file:
```
myDataFr.to_csv(fname1, index=False)
```
### Make some correlated data
Define size of data.
```
n = 100 # number of observations
nv = 2 # number of variables
```
#### Make the numerical data (i.e. the measurements).
##### First the x axis.
```
xOff = 5
xScl = 1
x = xOff + xScl*np.random.randn(n, 1)
```
##### Now the y values
```
yOff = 5
yScl = 1
xyr = 0.42
y = yOff + yScl*np.random.randn(n, 1)
y = (1-xyr)*np.random.randn(n, 1) + (xyr)*x
```
##### Concatenate into nx2 ndarray
```
myData = np.hstack((x,y))
np.shape(myData)
```
##### Convert the numpy array into a pandas data frame:
```
myDataFr = pd.DataFrame(myData, columns=["X", "Y"])
myDataFr.plot(x="X", y="Y", kind="scatter")
```
Save the data frame to a .csv (comma separated values) file:
```
myDataFr.to_csv(fname2, index=False)
```
### Nonlinear data
Define size of data.
```
n = 200 # number of observations
nv = 2 # number of variables
xmin = 0.1 # minimum x value
xmax = 42
```
#### Make the numerical data (i.e. the measurements).
##### First the x axis.
It turns out that np.linspace() and np.arange() produce "vectors" **not** 1xn or nx1 arrays. **And** you cannot then concatenate or multiply it with an array; numpy won't coerce it into having a second dimension like MATLAB does. You have to explicitly add the dimension of length one. This took me over two agonizing and frustating hours to work out.
```
x = np.linspace(xmin, xmax, n)
x = x[:, np.newaxis] # makes a [n,1] array from a dimensionless vector
```
##### Now the y values
```
# params
a, b, c = 10, 0, 0.7
# noise
noysAmp = 100
noys = noysAmp*np.random.randn(n, 1)
# y
y = a + b*x + c*(x**2) + noys
```
##### Concatenate into nx2 ndarray
```
myData = np.hstack((x,y))
np.shape(myData)
```
##### Convert the numpy array into a pandas data frame:
```
myDataFr = pd.DataFrame(myData, columns=["X", "Y"])
myDataFr.plot(x="X", y="Y", kind="scatter")
```
Save the data frame to a .csv (comma separated values) file:
```
myDataFr.to_csv(fname3, index=False)
```
#### Data for in-class exercise
```
fname="008ExerciseFile.csv"
n = 100 # number of observations
nv = 2 # number of variables
xmin = -10 # minimum x value
xmax = 10
x = np.linspace(xmin, xmax, n)
x = x[:, np.newaxis] # makes a [n,1] array from a dimensionless vector
# poly params
a, b, c, d = 100, -50, 1.5, 0.5
# noise
noysAmp = 50
noys = noysAmp*np.random.randn(n, 1)
# y
y = a + b*x + c*(x**2) + d*(x**3) + noys
myData = np.hstack((x,y))
np.shape(myData)
myDataFr = pd.DataFrame(myData, columns=["rel_time", "m_level"])
myDataFr.plot(x="rel_time", y="m_level", kind="scatter")
myDataFr.to_csv(fname, index=False)
```
| github_jupyter |
# Batch Normalization – Lesson
1. [What is it?](#theory)
2. [What are it's benefits?](#benefits)
3. [How do we add it to a network?](#implementation_1)
4. [Let's see it work!](#demos)
5. [What are you hiding?](#implementation_2)
# What is Batch Normalization?<a id='theory'></a>
Batch normalization was introduced in Sergey Ioffe's and Christian Szegedy's 2015 paper [Batch Normalization: Accelerating Deep Network Training by Reducing Internal Covariate Shift](https://arxiv.org/pdf/1502.03167.pdf). The idea is that, instead of just normalizing the inputs to the network, we normalize the inputs to _layers within_ the network. It's called "batch" normalization because during training, we normalize each layer's inputs by using the mean and variance of the values in the current mini-batch.
Why might this help? Well, we know that normalizing the inputs to a _network_ helps the network learn. But a network is a series of layers, where the output of one layer becomes the input to another. That means we can think of any layer in a neural network as the _first_ layer of a smaller network.
For example, imagine a 3 layer network. Instead of just thinking of it as a single network with inputs, layers, and outputs, think of the output of layer 1 as the input to a two layer network. This two layer network would consist of layers 2 and 3 in our original network.
Likewise, the output of layer 2 can be thought of as the input to a single layer network, consistng only of layer 3.
When you think of it like that - as a series of neural networks feeding into each other - then it's easy to imagine how normalizing the inputs to each layer would help. It's just like normalizing the inputs to any other neural network, but you're doing it at every layer (sub-network).
Beyond the intuitive reasons, there are good mathematical reasons why it helps the network learn better, too. It helps combat what the authors call _internal covariate shift_. This discussion is best handled [in the paper](https://arxiv.org/pdf/1502.03167.pdf) and in [Deep Learning](http://www.deeplearningbook.org) a book you can read online written by Ian Goodfellow, Yoshua Bengio, and Aaron Courville. Specifically, check out the batch normalization section of [Chapter 8: Optimization for Training Deep Models](http://www.deeplearningbook.org/contents/optimization.html).
# Benefits of Batch Normalization<a id="benefits"></a>
Batch normalization optimizes network training. It has been shown to have several benefits:
1. **Networks train faster** – Each training _iteration_ will actually be slower because of the extra calculations during the forward pass and the additional hyperparameters to train during back propagation. However, it should converge much more quickly, so training should be faster overall.
2. **Allows higher learning rates** – Gradient descent usually requires small learning rates for the network to converge. And as networks get deeper, their gradients get smaller during back propagation so they require even more iterations. Using batch normalization allows us to use much higher learning rates, which further increases the speed at which networks train.
3. **Makes weights easier to initialize** – Weight initialization can be difficult, and it's even more difficult when creating deeper networks. Batch normalization seems to allow us to be much less careful about choosing our initial starting weights.
4. **Makes more activation functions viable** – Some activation functions do not work well in some situations. Sigmoids lose their gradient pretty quickly, which means they can't be used in deep networks. And ReLUs often die out during training, where they stop learning completely, so we need to be careful about the range of values fed into them. Because batch normalization regulates the values going into each activation function, non-linearlities that don't seem to work well in deep networks actually become viable again.
5. **Simplifies the creation of deeper networks** – Because of the first 4 items listed above, it is easier to build and faster to train deeper neural networks when using batch normalization. And it's been shown that deeper networks generally produce better results, so that's great.
6. **Provides a bit of regularlization** – Batch normalization adds a little noise to your network. In some cases, such as in Inception modules, batch normalization has been shown to work as well as dropout. But in general, consider batch normalization as a bit of extra regularization, possibly allowing you to reduce some of the dropout you might add to a network.
7. **May give better results overall** – Some tests seem to show batch normalization actually improves the train.ing results. However, it's really an optimization to help train faster, so you shouldn't think of it as a way to make your network better. But since it lets you train networks faster, that means you can iterate over more designs more quickly. It also lets you build deeper networks, which are usually better. So when you factor in everything, you're probably going to end up with better results if you build your networks with batch normalization.
# Batch Normalization in TensorFlow<a id="implementation_1"></a>
This section of the notebook shows you one way to add batch normalization to a neural network built in TensorFlow.
The following cell imports the packages we need in the notebook and loads the MNIST dataset to use in our experiments. However, the `tensorflow` package contains all the code you'll actually need for batch normalization.
```
# Import necessary packages
import tensorflow as tf
import tqdm
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
# Import MNIST data so we have something for our experiments
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
```
### Neural network classes for testing
The following class, `NeuralNet`, allows us to create identical neural networks with and without batch normalization. The code is heaviy documented, but there is also some additional discussion later. You do not need to read through it all before going through the rest of the notebook, but the comments within the code blocks may answer some of your questions.
*About the code:*
>This class is not meant to represent TensorFlow best practices – the design choices made here are to support the discussion related to batch normalization.
>It's also important to note that we use the well-known MNIST data for these examples, but the networks we create are not meant to be good for performing handwritten character recognition. We chose this network architecture because it is similar to the one used in the original paper, which is complex enough to demonstrate some of the benefits of batch normalization while still being fast to train.
```
class NeuralNet:
def __init__(self, initial_weights, activation_fn, use_batch_norm):
"""
Initializes this object, creating a TensorFlow graph using the given parameters.
:param initial_weights: list of NumPy arrays or Tensors
Initial values for the weights for every layer in the network. We pass these in
so we can create multiple networks with the same starting weights to eliminate
training differences caused by random initialization differences.
The number of items in the list defines the number of layers in the network,
and the shapes of the items in the list define the number of nodes in each layer.
e.g. Passing in 3 matrices of shape (784, 256), (256, 100), and (100, 10) would
create a network with 784 inputs going into a hidden layer with 256 nodes,
followed by a hidden layer with 100 nodes, followed by an output layer with 10 nodes.
:param activation_fn: Callable
The function used for the output of each hidden layer. The network will use the same
activation function on every hidden layer and no activate function on the output layer.
e.g. Pass tf.nn.relu to use ReLU activations on your hidden layers.
:param use_batch_norm: bool
Pass True to create a network that uses batch normalization; False otherwise
Note: this network will not use batch normalization on layers that do not have an
activation function.
"""
# Keep track of whether or not this network uses batch normalization.
self.use_batch_norm = use_batch_norm
self.name = "With Batch Norm" if use_batch_norm else "Without Batch Norm"
# Batch normalization needs to do different calculations during training and inference,
# so we use this placeholder to tell the graph which behavior to use.
self.is_training = tf.placeholder(tf.bool, name="is_training")
# This list is just for keeping track of data we want to plot later.
# It doesn't actually have anything to do with neural nets or batch normalization.
self.training_accuracies = []
# Create the network graph, but it will not actually have any real values until after you
# call train or test
self.build_network(initial_weights, activation_fn)
def build_network(self, initial_weights, activation_fn):
"""
Build the graph. The graph still needs to be trained via the `train` method.
:param initial_weights: list of NumPy arrays or Tensors
See __init__ for description.
:param activation_fn: Callable
See __init__ for description.
"""
self.input_layer = tf.placeholder(tf.float32, [None, initial_weights[0].shape[0]])
layer_in = self.input_layer
for weights in initial_weights[:-1]:
layer_in = self.fully_connected(layer_in, weights, activation_fn)
self.output_layer = self.fully_connected(layer_in, initial_weights[-1])
def fully_connected(self, layer_in, initial_weights, activation_fn=None):
"""
Creates a standard, fully connected layer. Its number of inputs and outputs will be
defined by the shape of `initial_weights`, and its starting weight values will be
taken directly from that same parameter. If `self.use_batch_norm` is True, this
layer will include batch normalization, otherwise it will not.
:param layer_in: Tensor
The Tensor that feeds into this layer. It's either the input to the network or the output
of a previous layer.
:param initial_weights: NumPy array or Tensor
Initial values for this layer's weights. The shape defines the number of nodes in the layer.
e.g. Passing in 3 matrix of shape (784, 256) would create a layer with 784 inputs and 256
outputs.
:param activation_fn: Callable or None (default None)
The non-linearity used for the output of the layer. If None, this layer will not include
batch normalization, regardless of the value of `self.use_batch_norm`.
e.g. Pass tf.nn.relu to use ReLU activations on your hidden layers.
"""
# Since this class supports both options, only use batch normalization when
# requested. However, do not use it on the final layer, which we identify
# by its lack of an activation function.
if self.use_batch_norm and activation_fn:
# Batch normalization uses weights as usual, but does NOT add a bias term. This is because
# its calculations include gamma and beta variables that make the bias term unnecessary.
# (See later in the notebook for more details.)
weights = tf.Variable(initial_weights)
linear_output = tf.matmul(layer_in, weights)
# Apply batch normalization to the linear combination of the inputs and weights
batch_normalized_output = tf.layers.batch_normalization(linear_output, training=self.is_training)
# Now apply the activation function, *after* the normalization.
return activation_fn(batch_normalized_output)
else:
# When not using batch normalization, create a standard layer that multiplies
# the inputs and weights, adds a bias, and optionally passes the result
# through an activation function.
weights = tf.Variable(initial_weights)
biases = tf.Variable(tf.zeros([initial_weights.shape[-1]]))
linear_output = tf.add(tf.matmul(layer_in, weights), biases)
return linear_output if not activation_fn else activation_fn(linear_output)
def train(self, session, learning_rate, training_batches, batches_per_sample, save_model_as=None):
"""
Trains the model on the MNIST training dataset.
:param session: Session
Used to run training graph operations.
:param learning_rate: float
Learning rate used during gradient descent.
:param training_batches: int
Number of batches to train.
:param batches_per_sample: int
How many batches to train before sampling the validation accuracy.
:param save_model_as: string or None (default None)
Name to use if you want to save the trained model.
"""
# This placeholder will store the target labels for each mini batch
labels = tf.placeholder(tf.float32, [None, 10])
# Define loss and optimizer
cross_entropy = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(labels=labels, logits=self.output_layer))
# Define operations for testing
correct_prediction = tf.equal(tf.argmax(self.output_layer, 1), tf.argmax(labels, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
if self.use_batch_norm:
# If we don't include the update ops as dependencies on the train step, the
# tf.layers.batch_normalization layers won't update their population statistics,
# which will cause the model to fail at inference time
with tf.control_dependencies(tf.get_collection(tf.GraphKeys.UPDATE_OPS)):
train_step = tf.train.GradientDescentOptimizer(learning_rate).minimize(cross_entropy)
else:
train_step = tf.train.GradientDescentOptimizer(learning_rate).minimize(cross_entropy)
# Train for the appropriate number of batches. (tqdm is only for a nice timing display)
for i in tqdm.tqdm(range(training_batches)):
# We use batches of 60 just because the original paper did. You can use any size batch you like.
batch_xs, batch_ys = mnist.train.next_batch(60)
session.run(train_step, feed_dict={self.input_layer: batch_xs,
labels: batch_ys,
self.is_training: True})
# Periodically test accuracy against the 5k validation images and store it for plotting later.
if i % batches_per_sample == 0:
test_accuracy = session.run(accuracy, feed_dict={self.input_layer: mnist.validation.images,
labels: mnist.validation.labels,
self.is_training: False})
self.training_accuracies.append(test_accuracy)
# After training, report accuracy against test data
test_accuracy = session.run(accuracy, feed_dict={self.input_layer: mnist.validation.images,
labels: mnist.validation.labels,
self.is_training: False})
print('{}: After training, final accuracy on validation set = {}'.format(self.name, test_accuracy))
# If you want to use this model later for inference instead of having to retrain it,
# just construct it with the same parameters and then pass this file to the 'test' function
if save_model_as:
tf.train.Saver().save(session, save_model_as)
def test(self, session, test_training_accuracy=False, include_individual_predictions=False, restore_from=None):
"""
Trains a trained model on the MNIST testing dataset.
:param session: Session
Used to run the testing graph operations.
:param test_training_accuracy: bool (default False)
If True, perform inference with batch normalization using batch mean and variance;
if False, perform inference with batch normalization using estimated population mean and variance.
Note: in real life, *always* perform inference using the population mean and variance.
This parameter exists just to support demonstrating what happens if you don't.
:param include_individual_predictions: bool (default True)
This function always performs an accuracy test against the entire test set. But if this parameter
is True, it performs an extra test, doing 200 predictions one at a time, and displays the results
and accuracy.
:param restore_from: string or None (default None)
Name of a saved model if you want to test with previously saved weights.
"""
# This placeholder will store the true labels for each mini batch
labels = tf.placeholder(tf.float32, [None, 10])
# Define operations for testing
correct_prediction = tf.equal(tf.argmax(self.output_layer, 1), tf.argmax(labels, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
# If provided, restore from a previously saved model
if restore_from:
tf.train.Saver().restore(session, restore_from)
# Test against all of the MNIST test data
test_accuracy = session.run(accuracy, feed_dict={self.input_layer: mnist.test.images,
labels: mnist.test.labels,
self.is_training: test_training_accuracy})
print('-'*75)
print('{}: Accuracy on full test set = {}'.format(self.name, test_accuracy))
# If requested, perform tests predicting individual values rather than batches
if include_individual_predictions:
predictions = []
correct = 0
# Do 200 predictions, 1 at a time
for i in range(200):
# This is a normal prediction using an individual test case. However, notice
# we pass `test_training_accuracy` to `feed_dict` as the value for `self.is_training`.
# Remember that will tell it whether it should use the batch mean & variance or
# the population estimates that were calucated while training the model.
pred, corr = session.run([tf.arg_max(self.output_layer,1), accuracy],
feed_dict={self.input_layer: [mnist.test.images[i]],
labels: [mnist.test.labels[i]],
self.is_training: test_training_accuracy})
correct += corr
predictions.append(pred[0])
print("200 Predictions:", predictions)
print("Accuracy on 200 samples:", correct/200)
```
There are quite a few comments in the code, so those should answer most of your questions. However, let's take a look at the most important lines.
We add batch normalization to layers inside the `fully_connected` function. Here are some important points about that code:
1. Layers with batch normalization do not include a bias term.
2. We use TensorFlow's [`tf.layers.batch_normalization`](https://www.tensorflow.org/api_docs/python/tf/layers/batch_normalization) function to handle the math. (We show lower-level ways to do this [later in the notebook](#implementation_2).)
3. We tell `tf.layers.batch_normalization` whether or not the network is training. This is an important step we'll talk about later.
4. We add the normalization **before** calling the activation function.
In addition to that code, the training step is wrapped in the following `with` statement:
```python
with tf.control_dependencies(tf.get_collection(tf.GraphKeys.UPDATE_OPS)):
```
This line actually works in conjunction with the `training` parameter we pass to `tf.layers.batch_normalization`. Without it, TensorFlow's batch normalization layer will not operate correctly during inference.
Finally, whenever we train the network or perform inference, we use the `feed_dict` to set `self.is_training` to `True` or `False`, respectively, like in the following line:
```python
session.run(train_step, feed_dict={self.input_layer: batch_xs,
labels: batch_ys,
self.is_training: True})
```
We'll go into more details later, but next we want to show some experiments that use this code and test networks with and without batch normalization.
# Batch Normalization Demos<a id='demos'></a>
This section of the notebook trains various networks with and without batch normalization to demonstrate some of the benefits mentioned earlier.
We'd like to thank the author of this blog post [Implementing Batch Normalization in TensorFlow](http://r2rt.com/implementing-batch-normalization-in-tensorflow.html). That post provided the idea of - and some of the code for - plotting the differences in accuracy during training, along with the idea for comparing multiple networks using the same initial weights.
## Code to support testing
The following two functions support the demos we run in the notebook.
The first function, `plot_training_accuracies`, simply plots the values found in the `training_accuracies` lists of the `NeuralNet` objects passed to it. If you look at the `train` function in `NeuralNet`, you'll see it that while it's training the network, it periodically measures validation accuracy and stores the results in that list. It does that just to support these plots.
The second function, `train_and_test`, creates two neural nets - one with and one without batch normalization. It then trains them both and tests them, calling `plot_training_accuracies` to plot how their accuracies changed over the course of training. The really imporant thing about this function is that it initializes the starting weights for the networks _outside_ of the networks and then passes them in. This lets it train both networks from the exact same starting weights, which eliminates performance differences that might result from (un)lucky initial weights.
```
def plot_training_accuracies(*args, **kwargs):
"""
Displays a plot of the accuracies calculated during training to demonstrate
how many iterations it took for the model(s) to converge.
:param args: One or more NeuralNet objects
You can supply any number of NeuralNet objects as unnamed arguments
and this will display their training accuracies. Be sure to call `train`
the NeuralNets before calling this function.
:param kwargs:
You can supply any named parameters here, but `batches_per_sample` is the only
one we look for. It should match the `batches_per_sample` value you passed
to the `train` function.
"""
fig, ax = plt.subplots()
batches_per_sample = kwargs['batches_per_sample']
for nn in args:
ax.plot(range(0,len(nn.training_accuracies)*batches_per_sample,batches_per_sample),
nn.training_accuracies, label=nn.name)
ax.set_xlabel('Training steps')
ax.set_ylabel('Accuracy')
ax.set_title('Validation Accuracy During Training')
ax.legend(loc=4)
ax.set_ylim([0,1])
plt.yticks(np.arange(0, 1.1, 0.1))
plt.grid(True)
plt.show()
def train_and_test(use_bad_weights, learning_rate, activation_fn, training_batches=50000, batches_per_sample=500):
"""
Creates two networks, one with and one without batch normalization, then trains them
with identical starting weights, layers, batches, etc. Finally tests and plots their accuracies.
:param use_bad_weights: bool
If True, initialize the weights of both networks to wildly inappropriate weights;
if False, use reasonable starting weights.
:param learning_rate: float
Learning rate used during gradient descent.
:param activation_fn: Callable
The function used for the output of each hidden layer. The network will use the same
activation function on every hidden layer and no activate function on the output layer.
e.g. Pass tf.nn.relu to use ReLU activations on your hidden layers.
:param training_batches: (default 50000)
Number of batches to train.
:param batches_per_sample: (default 500)
How many batches to train before sampling the validation accuracy.
"""
# Use identical starting weights for each network to eliminate differences in
# weight initialization as a cause for differences seen in training performance
#
# Note: The networks will use these weights to define the number of and shapes of
# its layers. The original batch normalization paper used 3 hidden layers
# with 100 nodes in each, followed by a 10 node output layer. These values
# build such a network, but feel free to experiment with different choices.
# However, the input size should always be 784 and the final output should be 10.
if use_bad_weights:
# These weights should be horrible because they have such a large standard deviation
weights = [np.random.normal(size=(784,100), scale=5.0).astype(np.float32),
np.random.normal(size=(100,100), scale=5.0).astype(np.float32),
np.random.normal(size=(100,100), scale=5.0).astype(np.float32),
np.random.normal(size=(100,10), scale=5.0).astype(np.float32)
]
else:
# These weights should be good because they have such a small standard deviation
weights = [np.random.normal(size=(784,100), scale=0.05).astype(np.float32),
np.random.normal(size=(100,100), scale=0.05).astype(np.float32),
np.random.normal(size=(100,100), scale=0.05).astype(np.float32),
np.random.normal(size=(100,10), scale=0.05).astype(np.float32)
]
# Just to make sure the TensorFlow's default graph is empty before we start another
# test, because we don't bother using different graphs or scoping and naming
# elements carefully in this sample code.
tf.reset_default_graph()
# build two versions of same network, 1 without and 1 with batch normalization
nn = NeuralNet(weights, activation_fn, False)
bn = NeuralNet(weights, activation_fn, True)
# train and test the two models
with tf.Session() as sess:
tf.global_variables_initializer().run()
nn.train(sess, learning_rate, training_batches, batches_per_sample)
bn.train(sess, learning_rate, training_batches, batches_per_sample)
nn.test(sess)
bn.test(sess)
# Display a graph of how validation accuracies changed during training
# so we can compare how the models trained and when they converged
plot_training_accuracies(nn, bn, batches_per_sample=batches_per_sample)
```
## Comparisons between identical networks, with and without batch normalization
The next series of cells train networks with various settings to show the differences with and without batch normalization. They are meant to clearly demonstrate the effects of batch normalization. We include a deeper discussion of batch normalization later in the notebook.
**The following creates two networks using a ReLU activation function, a learning rate of 0.01, and reasonable starting weights.**
```
train_and_test(False, 0.01, tf.nn.relu)
```
As expected, both networks train well and eventually reach similar test accuracies. However, notice that the model with batch normalization converges slightly faster than the other network, reaching accuracies over 90% almost immediately and nearing its max acuracy in 10 or 15 thousand iterations. The other network takes about 3 thousand iterations to reach 90% and doesn't near its best accuracy until 30 thousand or more iterations.
If you look at the raw speed, you can see that without batch normalization we were computing over 1100 batches per second, whereas with batch normalization that goes down to just over 500. However, batch normalization allows us to perform fewer iterations and converge in less time over all. (We only trained for 50 thousand batches here so we could plot the comparison.)
**The following creates two networks with the same hyperparameters used in the previous example, but only trains for 2000 iterations.**
```
train_and_test(False, 0.01, tf.nn.relu, 2000, 50)
```
As you can see, using batch normalization produces a model with over 95% accuracy in only 2000 batches, and it was above 90% at somewhere around 500 batches. Without batch normalization, the model takes 1750 iterations just to hit 80% – the network with batch normalization hits that mark after around 200 iterations! (Note: if you run the code yourself, you'll see slightly different results each time because the starting weights - while the same for each model - are different for each run.)
In the above example, you should also notice that the networks trained fewer batches per second then what you saw in the previous example. That's because much of the time we're tracking is actually spent periodically performing inference to collect data for the plots. In this example we perform that inference every 50 batches instead of every 500, so generating the plot for this example requires 10 times the overhead for the same 2000 iterations.
**The following creates two networks using a sigmoid activation function, a learning rate of 0.01, and reasonable starting weights.**
```
train_and_test(False, 0.01, tf.nn.sigmoid)
```
With the number of layers we're using and this small learning rate, using a sigmoid activation function takes a long time to start learning. It eventually starts making progress, but it took over 45 thousand batches just to get over 80% accuracy. Using batch normalization gets to 90% in around one thousand batches.
**The following creates two networks using a ReLU activation function, a learning rate of 1, and reasonable starting weights.**
```
train_and_test(False, 1, tf.nn.relu)
```
Now we're using ReLUs again, but with a larger learning rate. The plot shows how training started out pretty normally, with the network with batch normalization starting out faster than the other. But the higher learning rate bounces the accuracy around a bit more, and at some point the accuracy in the network without batch normalization just completely crashes. It's likely that too many ReLUs died off at this point because of the high learning rate.
The next cell shows the same test again. The network with batch normalization performs the same way, and the other suffers from the same problem again, but it manages to train longer before it happens.
```
train_and_test(False, 1, tf.nn.relu)
```
In both of the previous examples, the network with batch normalization manages to gets over 98% accuracy, and get near that result almost immediately. The higher learning rate allows the network to train extremely fast.
**The following creates two networks using a sigmoid activation function, a learning rate of 1, and reasonable starting weights.**
```
train_and_test(False, 1, tf.nn.sigmoid)
```
In this example, we switched to a sigmoid activation function. It appears to hande the higher learning rate well, with both networks achieving high accuracy.
The cell below shows a similar pair of networks trained for only 2000 iterations.
```
train_and_test(False, 1, tf.nn.sigmoid, 2000, 50)
```
As you can see, even though these parameters work well for both networks, the one with batch normalization gets over 90% in 400 or so batches, whereas the other takes over 1700. When training larger networks, these sorts of differences become more pronounced.
**The following creates two networks using a ReLU activation function, a learning rate of 2, and reasonable starting weights.**
```
train_and_test(False, 2, tf.nn.relu)
```
With this very large learning rate, the network with batch normalization trains fine and almost immediately manages 98% accuracy. However, the network without normalization doesn't learn at all.
**The following creates two networks using a sigmoid activation function, a learning rate of 2, and reasonable starting weights.**
```
train_and_test(False, 2, tf.nn.sigmoid)
```
Once again, using a sigmoid activation function with the larger learning rate works well both with and without batch normalization.
However, look at the plot below where we train models with the same parameters but only 2000 iterations. As usual, batch normalization lets it train faster.
```
train_and_test(False, 2, tf.nn.sigmoid, 2000, 50)
```
In the rest of the examples, we use really bad starting weights. That is, normally we would use very small values close to zero. However, in these examples we choose randome values with a standard deviation of 5. If you were really training a neural network, you would **not** want to do this. But these examples demonstrate how batch normalization makes your network much more resilient.
**The following creates two networks using a ReLU activation function, a learning rate of 0.01, and bad starting weights.**
```
train_and_test(True, 0.01, tf.nn.relu)
```
As the plot shows, without batch normalization the network never learns anything at all. But with batch normalization, it actually learns pretty well and gets to almost 80% accuracy. The starting weights obviously hurt the network, but you can see how well batch normalization does in overcoming them.
**The following creates two networks using a sigmoid activation function, a learning rate of 0.01, and bad starting weights.**
```
train_and_test(True, 0.01, tf.nn.sigmoid)
```
Using a sigmoid activation function works better than the ReLU in the previous example, but without batch normalization it would take a tremendously long time to train the network, if it ever trained at all.
**The following creates two networks using a ReLU activation function, a learning rate of 1, and bad starting weights.**<a id="successful_example_lr_1"></a>
```
train_and_test(True, 1, tf.nn.relu)
```
The higher learning rate used here allows the network with batch normalization to surpass 90% in about 30 thousand batches. The network without it never gets anywhere.
**The following creates two networks using a sigmoid activation function, a learning rate of 1, and bad starting weights.**
```
train_and_test(True, 1, tf.nn.sigmoid)
```
Using sigmoid works better than ReLUs for this higher learning rate. However, you can see that without batch normalization, the network takes a long time tro train, bounces around a lot, and spends a long time stuck at 90%. The network with batch normalization trains much more quickly, seems to be more stable, and achieves a higher accuracy.
**The following creates two networks using a ReLU activation function, a learning rate of 2, and bad starting weights.**<a id="successful_example_lr_2"></a>
```
train_and_test(True, 2, tf.nn.relu)
```
We've already seen that ReLUs do not do as well as sigmoids with higher learning rates, and here we are using an extremely high rate. As expected, without batch normalization the network doesn't learn at all. But with batch normalization, it eventually achieves 90% accuracy. Notice, though, how its accuracy bounces around wildly during training - that's because the learning rate is really much too high, so the fact that this worked at all is a bit of luck.
**The following creates two networks using a sigmoid activation function, a learning rate of 2, and bad starting weights.**
```
train_and_test(True, 2, tf.nn.sigmoid)
```
In this case, the network with batch normalization trained faster and reached a higher accuracy. Meanwhile, the high learning rate makes the network without normalization bounce around erratically and have trouble getting past 90%.
### Full Disclosure: Batch Normalization Doesn't Fix Everything
Batch normalization isn't magic and it doesn't work every time. Weights are still randomly initialized and batches are chosen at random during training, so you never know exactly how training will go. Even for these tests, where we use the same initial weights for both networks, we still get _different_ weights each time we run.
This section includes two examples that show runs when batch normalization did not help at all.
**The following creates two networks using a ReLU activation function, a learning rate of 1, and bad starting weights.**
```
train_and_test(True, 1, tf.nn.relu)
```
When we used these same parameters [earlier](#successful_example_lr_1), we saw the network with batch normalization reach 92% validation accuracy. This time we used different starting weights, initialized using the same standard deviation as before, and the network doesn't learn at all. (Remember, an accuracy around 10% is what the network gets if it just guesses the same value all the time.)
**The following creates two networks using a ReLU activation function, a learning rate of 2, and bad starting weights.**
```
train_and_test(True, 2, tf.nn.relu)
```
When we trained with these parameters and batch normalization [earlier](#successful_example_lr_2), we reached 90% validation accuracy. However, this time the network _almost_ starts to make some progress in the beginning, but it quickly breaks down and stops learning.
**Note:** Both of the above examples use *extremely* bad starting weights, along with learning rates that are too high. While we've shown batch normalization _can_ overcome bad values, we don't mean to encourage actually using them. The examples in this notebook are meant to show that batch normalization can help your networks train better. But these last two examples should remind you that you still want to try to use good network design choices and reasonable starting weights. It should also remind you that the results of each attempt to train a network are a bit random, even when using otherwise identical architectures.
# Batch Normalization: A Detailed Look<a id='implementation_2'></a>
The layer created by `tf.layers.batch_normalization` handles all the details of implementing batch normalization. Many students will be fine just using that and won't care about what's happening at the lower levels. However, some students may want to explore the details, so here is a short explanation of what's really happening, starting with the equations you're likely to come across if you ever read about batch normalization.
In order to normalize the values, we first need to find the average value for the batch. If you look at the code, you can see that this is not the average value of the batch _inputs_, but the average value coming _out_ of any particular layer before we pass it through its non-linear activation function and then feed it as an input to the _next_ layer.
We represent the average as $\mu_B$, which is simply the sum of all of the values $x_i$ divided by the number of values, $m$
$$
\mu_B \leftarrow \frac{1}{m}\sum_{i=1}^m x_i
$$
We then need to calculate the variance, or mean squared deviation, represented as $\sigma_{B}^{2}$. If you aren't familiar with statistics, that simply means for each value $x_i$, we subtract the average value (calculated earlier as $\mu_B$), which gives us what's called the "deviation" for that value. We square the result to get the squared deviation. Sum up the results of doing that for each of the values, then divide by the number of values, again $m$, to get the average, or mean, squared deviation.
$$
\sigma_{B}^{2} \leftarrow \frac{1}{m}\sum_{i=1}^m (x_i - \mu_B)^2
$$
Once we have the mean and variance, we can use them to normalize the values with the following equation. For each value, it subtracts the mean and divides by the (almost) standard deviation. (You've probably heard of standard deviation many times, but if you have not studied statistics you might not know that the standard deviation is actually the square root of the mean squared deviation.)
$$
\hat{x_i} \leftarrow \frac{x_i - \mu_B}{\sqrt{\sigma_{B}^{2} + \epsilon}}
$$
Above, we said "(almost) standard deviation". That's because the real standard deviation for the batch is calculated by $\sqrt{\sigma_{B}^{2}}$, but the above formula adds the term epsilon, $\epsilon$, before taking the square root. The epsilon can be any small, positive constant - in our code we use the value `0.001`. It is there partially to make sure we don't try to divide by zero, but it also acts to increase the variance slightly for each batch.
Why increase the variance? Statistically, this makes sense because even though we are normalizing one batch at a time, we are also trying to estimate the population distribution – the total training set, which itself an estimate of the larger population of inputs your network wants to handle. The variance of a population is higher than the variance for any sample taken from that population, so increasing the variance a little bit for each batch helps take that into account.
At this point, we have a normalized value, represented as $\hat{x_i}$. But rather than use it directly, we multiply it by a gamma value, $\gamma$, and then add a beta value, $\beta$. Both $\gamma$ and $\beta$ are learnable parameters of the network and serve to scale and shift the normalized value, respectively. Because they are learnable just like weights, they give your network some extra knobs to tweak during training to help it learn the function it is trying to approximate.
$$
y_i \leftarrow \gamma \hat{x_i} + \beta
$$
We now have the final batch-normalized output of our layer, which we would then pass to a non-linear activation function like sigmoid, tanh, ReLU, Leaky ReLU, etc. In the original batch normalization paper (linked in the beginning of this notebook), they mention that there might be cases when you'd want to perform the batch normalization _after_ the non-linearity instead of before, but it is difficult to find any uses like that in practice.
In `NeuralNet`'s implementation of `fully_connected`, all of this math is hidden inside the following line, where `linear_output` serves as the $x_i$ from the equations:
```python
batch_normalized_output = tf.layers.batch_normalization(linear_output, training=self.is_training)
```
The next section shows you how to implement the math directly.
### Batch normalization without the `tf.layers` package
Our implementation of batch normalization in `NeuralNet` uses the high-level abstraction [tf.layers.batch_normalization](https://www.tensorflow.org/api_docs/python/tf/layers/batch_normalization), found in TensorFlow's [`tf.layers`](https://www.tensorflow.org/api_docs/python/tf/layers) package.
However, if you would like to implement batch normalization at a lower level, the following code shows you how.
It uses [tf.nn.batch_normalization](https://www.tensorflow.org/api_docs/python/tf/nn/batch_normalization) from TensorFlow's [neural net (nn)](https://www.tensorflow.org/api_docs/python/tf/nn) package.
**1)** You can replace the `fully_connected` function in the `NeuralNet` class with the below code and everything in `NeuralNet` will still work like it did before.
```
def fully_connected(self, layer_in, initial_weights, activation_fn=None):
"""
Creates a standard, fully connected layer. Its number of inputs and outputs will be
defined by the shape of `initial_weights`, and its starting weight values will be
taken directly from that same parameter. If `self.use_batch_norm` is True, this
layer will include batch normalization, otherwise it will not.
:param layer_in: Tensor
The Tensor that feeds into this layer. It's either the input to the network or the output
of a previous layer.
:param initial_weights: NumPy array or Tensor
Initial values for this layer's weights. The shape defines the number of nodes in the layer.
e.g. Passing in 3 matrix of shape (784, 256) would create a layer with 784 inputs and 256
outputs.
:param activation_fn: Callable or None (default None)
The non-linearity used for the output of the layer. If None, this layer will not include
batch normalization, regardless of the value of `self.use_batch_norm`.
e.g. Pass tf.nn.relu to use ReLU activations on your hidden layers.
"""
if self.use_batch_norm and activation_fn:
# Batch normalization uses weights as usual, but does NOT add a bias term. This is because
# its calculations include gamma and beta variables that make the bias term unnecessary.
weights = tf.Variable(initial_weights)
linear_output = tf.matmul(layer_in, weights)
num_out_nodes = initial_weights.shape[-1]
# Batch normalization adds additional trainable variables:
# gamma (for scaling) and beta (for shifting).
gamma = tf.Variable(tf.ones([num_out_nodes]))
beta = tf.Variable(tf.zeros([num_out_nodes]))
# These variables will store the mean and variance for this layer over the entire training set,
# which we assume represents the general population distribution.
# By setting `trainable=False`, we tell TensorFlow not to modify these variables during
# back propagation. Instead, we will assign values to these variables ourselves.
pop_mean = tf.Variable(tf.zeros([num_out_nodes]), trainable=False)
pop_variance = tf.Variable(tf.ones([num_out_nodes]), trainable=False)
# Batch normalization requires a small constant epsilon, used to ensure we don't divide by zero.
# This is the default value TensorFlow uses.
epsilon = 1e-3
def batch_norm_training():
# Calculate the mean and variance for the data coming out of this layer's linear-combination step.
# The [0] defines an array of axes to calculate over.
batch_mean, batch_variance = tf.nn.moments(linear_output, [0])
# Calculate a moving average of the training data's mean and variance while training.
# These will be used during inference.
# Decay should be some number less than 1. tf.layers.batch_normalization uses the parameter
# "momentum" to accomplish this and defaults it to 0.99
decay = 0.99
train_mean = tf.assign(pop_mean, pop_mean * decay + batch_mean * (1 - decay))
train_variance = tf.assign(pop_variance, pop_variance * decay + batch_variance * (1 - decay))
# The 'tf.control_dependencies' context tells TensorFlow it must calculate 'train_mean'
# and 'train_variance' before it calculates the 'tf.nn.batch_normalization' layer.
# This is necessary because the those two operations are not actually in the graph
# connecting the linear_output and batch_normalization layers,
# so TensorFlow would otherwise just skip them.
with tf.control_dependencies([train_mean, train_variance]):
return tf.nn.batch_normalization(linear_output, batch_mean, batch_variance, beta, gamma, epsilon)
def batch_norm_inference():
# During inference, use the our estimated population mean and variance to normalize the layer
return tf.nn.batch_normalization(linear_output, pop_mean, pop_variance, beta, gamma, epsilon)
# Use `tf.cond` as a sort of if-check. When self.is_training is True, TensorFlow will execute
# the operation returned from `batch_norm_training`; otherwise it will execute the graph
# operation returned from `batch_norm_inference`.
batch_normalized_output = tf.cond(self.is_training, batch_norm_training, batch_norm_inference)
# Pass the batch-normalized layer output through the activation function.
# The literature states there may be cases where you want to perform the batch normalization *after*
# the activation function, but it is difficult to find any uses of that in practice.
return activation_fn(batch_normalized_output)
else:
# When not using batch normalization, create a standard layer that multiplies
# the inputs and weights, adds a bias, and optionally passes the result
# through an activation function.
weights = tf.Variable(initial_weights)
biases = tf.Variable(tf.zeros([initial_weights.shape[-1]]))
linear_output = tf.add(tf.matmul(layer_in, weights), biases)
return linear_output if not activation_fn else activation_fn(linear_output)
```
This version of `fully_connected` is much longer than the original, but once again has extensive comments to help you understand it. Here are some important points:
1. It explicitly creates variables to store gamma, beta, and the population mean and variance. These were all handled for us in the previous version of the function.
2. It initializes gamma to one and beta to zero, so they start out having no effect in this calculation: $y_i \leftarrow \gamma \hat{x_i} + \beta$. However, during training the network learns the best values for these variables using back propagation, just like networks normally do with weights.
3. Unlike gamma and beta, the variables for population mean and variance are marked as untrainable. That tells TensorFlow not to modify them during back propagation. Instead, the lines that call `tf.assign` are used to update these variables directly.
4. TensorFlow won't automatically run the `tf.assign` operations during training because it only evaluates operations that are required based on the connections it finds in the graph. To get around that, we add this line: `with tf.control_dependencies([train_mean, train_variance]):` before we run the normalization operation. This tells TensorFlow it needs to run those operations before running anything inside the `with` block.
5. The actual normalization math is still mostly hidden from us, this time using [`tf.nn.batch_normalization`](https://www.tensorflow.org/api_docs/python/tf/nn/batch_normalization).
5. `tf.nn.batch_normalization` does not have a `training` parameter like `tf.layers.batch_normalization` did. However, we still need to handle training and inference differently, so we run different code in each case using the [`tf.cond`](https://www.tensorflow.org/api_docs/python/tf/cond) operation.
6. We use the [`tf.nn.moments`](https://www.tensorflow.org/api_docs/python/tf/nn/moments) function to calculate the batch mean and variance.
**2)** The current version of the `train` function in `NeuralNet` will work fine with this new version of `fully_connected`. However, it uses these lines to ensure population statistics are updated when using batch normalization:
```python
if self.use_batch_norm:
with tf.control_dependencies(tf.get_collection(tf.GraphKeys.UPDATE_OPS)):
train_step = tf.train.GradientDescentOptimizer(learning_rate).minimize(cross_entropy)
else:
train_step = tf.train.GradientDescentOptimizer(learning_rate).minimize(cross_entropy)
```
Our new version of `fully_connected` handles updating the population statistics directly. That means you can also simplify your code by replacing the above `if`/`else` condition with just this line:
```python
train_step = tf.train.GradientDescentOptimizer(learning_rate).minimize(cross_entropy)
```
**3)** And just in case you want to implement every detail from scratch, you can replace this line in `batch_norm_training`:
```python
return tf.nn.batch_normalization(linear_output, batch_mean, batch_variance, beta, gamma, epsilon)
```
with these lines:
```python
normalized_linear_output = (linear_output - batch_mean) / tf.sqrt(batch_variance + epsilon)
return gamma * normalized_linear_output + beta
```
And replace this line in `batch_norm_inference`:
```python
return tf.nn.batch_normalization(linear_output, pop_mean, pop_variance, beta, gamma, epsilon)
```
with these lines:
```python
normalized_linear_output = (linear_output - pop_mean) / tf.sqrt(pop_variance + epsilon)
return gamma * normalized_linear_output + beta
```
As you can see in each of the above substitutions, the two lines of replacement code simply implement the following two equations directly. The first line calculates the following equation, with `linear_output` representing $x_i$ and `normalized_linear_output` representing $\hat{x_i}$:
$$
\hat{x_i} \leftarrow \frac{x_i - \mu_B}{\sqrt{\sigma_{B}^{2} + \epsilon}}
$$
And the second line is a direct translation of the following equation:
$$
y_i \leftarrow \gamma \hat{x_i} + \beta
$$
We still use the `tf.nn.moments` operation to implement the other two equations from earlier – the ones that calculate the batch mean and variance used in the normalization step. If you really wanted to do everything from scratch, you could replace that line, too, but we'll leave that to you.
## Why the difference between training and inference?
In the original function that uses `tf.layers.batch_normalization`, we tell the layer whether or not the network is training by passing a value for its `training` parameter, like so:
```python
batch_normalized_output = tf.layers.batch_normalization(linear_output, training=self.is_training)
```
And that forces us to provide a value for `self.is_training` in our `feed_dict`, like we do in this example from `NeuralNet`'s `train` function:
```python
session.run(train_step, feed_dict={self.input_layer: batch_xs,
labels: batch_ys,
self.is_training: True})
```
If you looked at the [low level implementation](#low_level_code), you probably noticed that, just like with `tf.layers.batch_normalization`, we need to do slightly different things during training and inference. But why is that?
First, let's look at what happens when we don't. The following function is similar to `train_and_test` from earlier, but this time we are only testing one network and instead of plotting its accuracy, we perform 200 predictions on test inputs, 1 input at at time. We can use the `test_training_accuracy` parameter to test the network in training or inference modes (the equivalent of passing `True` or `False` to the `feed_dict` for `is_training`).
```
def batch_norm_test(test_training_accuracy):
"""
:param test_training_accuracy: bool
If True, perform inference with batch normalization using batch mean and variance;
if False, perform inference with batch normalization using estimated population mean and variance.
"""
weights = [np.random.normal(size=(784,100), scale=0.05).astype(np.float32),
np.random.normal(size=(100,100), scale=0.05).astype(np.float32),
np.random.normal(size=(100,100), scale=0.05).astype(np.float32),
np.random.normal(size=(100,10), scale=0.05).astype(np.float32)
]
tf.reset_default_graph()
# Train the model
bn = NeuralNet(weights, tf.nn.relu, True)
# First train the network
with tf.Session() as sess:
tf.global_variables_initializer().run()
bn.train(sess, 0.01, 2000, 2000)
bn.test(sess, test_training_accuracy=test_training_accuracy, include_individual_predictions=True)
```
In the following cell, we pass `True` for `test_training_accuracy`, which performs the same batch normalization that we normally perform **during training**.
```
batch_norm_test(True)
```
As you can see, the network guessed the same value every time! But why? Because during training, a network with batch normalization adjusts the values at each layer based on the mean and variance **of that batch**. The "batches" we are using for these predictions have a single input each time, so their values _are_ the means, and their variances will always be 0. That means the network will normalize the values at any layer to zero. (Review the equations from before to see why a value that is equal to the mean would always normalize to zero.) So we end up with the same result for every input we give the network, because its the value the network produces when it applies its learned weights to zeros at every layer.
**Note:** If you re-run that cell, you might get a different value from what we showed. That's because the specific weights the network learns will be different every time. But whatever value it is, it should be the same for all 200 predictions.
To overcome this problem, the network does not just normalize the batch at each layer. It also maintains an estimate of the mean and variance for the entire population. So when we perform inference, instead of letting it "normalize" all the values using their own means and variance, it uses the estimates of the population mean and variance that it calculated while training.
So in the following example, we pass `False` for `test_training_accuracy`, which tells the network that we it want to perform inference with the population statistics it calculates during training.
```
batch_norm_test(False)
```
As you can see, now that we're using the estimated population mean and variance, we get a 97% accuracy. That means it guessed correctly on 194 of the 200 samples – not too bad for something that trained in under 4 seconds. :)
# Considerations for other network types
This notebook demonstrates batch normalization in a standard neural network with fully connected layers. You can also use batch normalization in other types of networks, but there are some special considerations.
### ConvNets
Convolution layers consist of multiple feature maps. (Remember, the depth of a convolutional layer refers to its number of feature maps.) And the weights for each feature map are shared across all the inputs that feed into the layer. Because of these differences, batch normalizaing convolutional layers requires batch/population mean and variance per feature map rather than per node in the layer.
When using `tf.layers.batch_normalization`, be sure to pay attention to the order of your convolutionlal dimensions.
Specifically, you may want to set a different value for the `axis` parameter if your layers have their channels first instead of last.
In our low-level implementations, we used the following line to calculate the batch mean and variance:
```python
batch_mean, batch_variance = tf.nn.moments(linear_output, [0])
```
If we were dealing with a convolutional layer, we would calculate the mean and variance with a line like this instead:
```python
batch_mean, batch_variance = tf.nn.moments(conv_layer, [0,1,2], keep_dims=False)
```
The second parameter, `[0,1,2]`, tells TensorFlow to calculate the batch mean and variance over each feature map. (The three axes are the batch, height, and width.) And setting `keep_dims` to `False` tells `tf.nn.moments` not to return values with the same size as the inputs. Specifically, it ensures we get one mean/variance pair per feature map.
### RNNs
Batch normalization can work with recurrent neural networks, too, as shown in the 2016 paper [Recurrent Batch Normalization](https://arxiv.org/abs/1603.09025). It's a bit more work to implement, but basically involves calculating the means and variances per time step instead of per layer. You can find an example where someone extended `tf.nn.rnn_cell.RNNCell` to include batch normalization in [this GitHub repo](https://gist.github.com/spitis/27ab7d2a30bbaf5ef431b4a02194ac60).
| github_jupyter |
# Crunch
Crunching as defined in `eodag` is a way to filter the EO products contained in a [SearchResult](../../api_reference/searchresult.rst#eodag.api.search_result.SearchResult) object. Several filters are available and further described in this document.
A [SearchResult](../../api_reference/searchresult.rst#eodag.api.search_result.SearchResult) has a [crunch()](../../api_reference/searchresult.rst#eodag.api.search_result.SearchResult.crunch) method that requires a filter instance as an argument, itself initialized with a dictionary that contains the required parameters. According to the filter used, some more kwargs may need to be passed to [crunch()](../../api_reference/searchresult.rst#eodag.api.search_result.SearchResult.crunch). The filters return a `list` of [EOProduct](../../api_reference/eoproduct.rst#eodag.api.product._product.EOProduct)s.
## Setup
Results obtained from a search of *Sentinel 2 Level-1C* products over France in March 2021 are loaded in a [SearchResult](../../api_reference/searchresult.rst#eodag.api.search_result.SearchResult).
```
from eodag import EODataAccessGateway
dag = EODataAccessGateway()
search_results = dag.deserialize("data/crunch_search_results.geojson")
print(f"This SearchResult stores {len(search_results)} products.")
# This code cell has a special metadata entry: "nbsphinx": "hidden"
# That hides it when the documentation is built with nbsphinx/sphinx.
# Uncomment these lines to regenerate the GeoJSON file used in this notebook.
#search_results, _ = dag.search(
# productType="S2_MSI_L1C",
# start="2021-03-01",
# end="2021-03-31",
# geom={"lonmin": 1, "latmin": 45, "lonmax": 5, "latmax": 47},
# items_per_page=50
#)
#dag.serialize(search_results, "data/crunch_search_results.geojson")
```
The original search geometry is used throughout the notebook as long as with its representation as a a `shapely` object which is easier to map with `folium`.
```
original_search_geometry = {"lonmin": 1, "latmin": 45, "lonmax": 5, "latmax": 47}
import shapely
search_geometry = shapely.geometry.box(
original_search_geometry["lonmin"],
original_search_geometry["latmin"],
original_search_geometry["lonmax"],
original_search_geometry["latmax"],
)
# To create interactive maps
import folium
def create_search_result_map(search_results, extent):
"""Small utility to create an interactive map with folium
that displays an extent in red and EO Producs in blue"""
fmap = folium.Map([46, 3], zoom_start=6)
folium.GeoJson(
extent,
style_function=lambda x: dict(color="red")
).add_to(fmap)
folium.GeoJson(
search_results
).add_to(fmap)
return fmap
```
## Filter by start and end date
[FilterDate](../../plugins_reference/generated/eodag.plugins.crunch.filter_date.FilterDate.rst#eodag.plugins.crunch.filter_date.FilterDate) allows to filter out products that are older than a start date (optional) or more recent than an end date (optional).
```
from eodag.plugins.crunch.filter_date import FilterDate
filtered_products = search_results.crunch(
FilterDate(dict(start="2021-03-25", end="2021-03-29"))
)
print(f"{len(search_results) - len(filtered_products)} products were filtered out by the date filter.")
```
## Filter by geometry
[FilterOverlap](../../plugins_reference/generated/eodag.plugins.crunch.filter_overlap.FilterOverlap.rst#eodag.plugins.crunch.filter_overlap.FilterOverlap) allows to filter out products that:
* whose overlap area with a geometry is less than a percentage of their area
* are not *within* a geometry
* do not *contain* a geometry
* do not *intersect* with a geometry
To execute a [FilterOverlap](../../plugins_reference/generated/eodag.plugins.crunch.filter_overlap.FilterOverlap.rst#eodag.plugins.crunch.filter_overlap.FilterOverlap), its instance must be created by passing a dictionary with either:
* `minimum_overlap` set to a number between 0 and 100. `within`, `contains` and `intersects` cannot be used in that case.
* **One** of `within`, `contains` and `intersects` (they are mutually exclusive) set to True. `minimum_overlap` cannot be used in that case.
Additionally, a geometry (shapely geometry, bounding box as a dictionary or a list) must be passed through the `geometry` parameter.
The examples below show how [FilterOverlap](../../plugins_reference/generated/eodag.plugins.crunch.filter_overlap.FilterOverlap.rst#eodag.plugins.crunch.filter_overlap.FilterOverlap) filter out products. The original products will be displayed in blue and the filtered products in green.
```
from eodag.plugins.crunch.filter_overlap import FilterOverlap
```
All the products are displayed on the next map. As it can be observed, they all intersect with the search geometry.
```
create_search_result_map(search_results, search_geometry)
```
The next two examples show how `minimum_overlap` affects the filter, with its value (i.e. percentage) set to 10 and 50%.
```
filtered_products = search_results.crunch(
FilterOverlap(dict(minimum_overlap=10)),
geometry=search_geometry
)
print(f"{len(search_results) - len(filtered_products)} products were filtered out by the geometry filter.")
fmap = create_search_result_map(search_results, search_geometry)
folium.GeoJson(
filtered_products,
style_function=lambda x: dict(color="green")
).add_to(fmap)
fmap
filtered_products = search_results.crunch(
FilterOverlap(dict(minimum_overlap=50)),
geometry=search_geometry
)
print(f"{len(search_results) - len(filtered_products)} products were filtered out by the geometry filter.")
fmap = create_search_result_map(search_results, search_geometry)
folium.GeoJson(
filtered_products,
style_function=lambda x: dict(color="green")
).add_to(fmap)
fmap
```
More and more products are filtered out when `minimum_overlap` increases. The next parameter given as an example is `within`, it is actually equivalent to setting `minimum_overlap` to 100.
```
filtered_products = search_results.crunch(
FilterOverlap(dict(within=True)),
geometry=search_geometry
)
print(f"{len(search_results) - len(filtered_products)} products were filtered out by the geometry filter.")
fmap = create_search_result_map(search_results, search_geometry)
# Create a layer that represents the search area in red
folium.GeoJson(
filtered_products,
style_function=lambda x: dict(color="green")
).add_to(fmap)
fmap
```
All the products not withing the read area are correctly filtered out by [FilterOverlap](../../plugins_reference/generated/eodag.plugins.crunch.filter_overlap.FilterOverlap.rst#eodag.plugins.crunch.filter_overlap.FilterOverlap). A new geometry is created in order to test the next parameter `intersects`.
```
from shapely.geometry import Polygon
shifted_geom = Polygon([[4, 44], [9, 44], [9, 48], [4, 48], [4, 48]])
filtered_products = search_results.crunch(
FilterOverlap(dict(intersects=True)),
geometry=shifted_geom
)
print(f"{len(search_results) - len(filtered_products)} products were filtered out by the geometry filter.")
fmap = create_search_result_map(search_results, shifted_geom)
# Create a layer that represents a smaller area in red
folium.GeoJson(
filtered_products,
style_function=lambda x: dict(color="green")
).add_to(fmap)
fmap
```
The products that do not intersect with the red area are correctly filtered out. Finally another new geometry is created to test the parameter `contains`.
```
small_geom = Polygon([[3.2, 44.4], [3.7, 44.4], [3.7, 44.9], [3.2, 44.9], [3.2, 44.4]])
filtered_products = search_results.crunch(
FilterOverlap(dict(contains=True)),
geometry=small_geom
)
print(f"{len(search_results) - len(filtered_products)} products were filtered out by the geometry filter.")
fmap = create_search_result_map(search_results, small_geom)
# Create a layer that represents a smaller area in red
folium.GeoJson(
filtered_products,
style_function=lambda x: dict(color="green")
).add_to(fmap)
fmap
```
The only product preserved is the one that contains the red area.
## Filter by property
[FilterProperty](../../plugins_reference/generated/eodag.plugins.crunch.filter_property.FilterProperty.rst#eodag.plugins.crunch.filter_property.FilterProperty) evaluates a single property of all the products against a value (e.g. cloud cover less than 10). The dictionary it requires should contain:
* A single property name from [EOProduct](../../api_reference/eoproduct.rst#eodag.api.product._product.EOProduct)`.properties` and its tested value, e.g. `dict(cloudCover=10)` or `dict(storageStatus="ONLINE")`
* One (optional) operator among `lt` (<), `le` (<=), `eq` (==), `ne` (!=), `ge` (>=), `gt` (>). `eq` by default.
```
from eodag.plugins.crunch.filter_property import FilterProperty
filtered_products = search_results.crunch(
FilterProperty(dict(cloudCover=1, operator="lt"))
)
print(f"{len(search_results) - len(filtered_products)} products were filtered out by the property filter.")
```
List comprehensions over a collection of EO products are useful to quickly extract their properties, and here to check that the filter correctly filtered the products.
```
all([p.properties["cloudCover"] < 1 for p in filtered_products])
```
## Filter the latest products intersecting a geometry
[FilterLatestIntersect](../../plugins_reference/generated/eodag.plugins.crunch.filter_latest_intersect.FilterLatestIntersect.rst#eodag.plugins.crunch.filter_latest_intersect.FilterLatestIntersect) does the following:
1. it sorts the products by date, from the newest to the oldest
2. it filters out products that do not intersect with a requested geometry (a dictionary bounding box)
3. it stops early if the requested geometry is 100% covered by the products, if not, it returns the result of 2.
This results in getting the most recent products that intersect (or completely cover) a given geometry.
```
from eodag.plugins.crunch.filter_latest_intersect import FilterLatestIntersect
filtered_products = search_results.crunch(
FilterLatestIntersect({}),
geometry=original_search_geometry
)
print(f"{len(search_results) - len(filtered_products)} products were filtered out by the property filter.")
from shapely import geometry
fmap = create_search_result_map(search_results, search_geometry)
# Create a layer that represents a smaller area in red
folium.GeoJson(
filtered_products,
style_function=lambda x: dict(color="green")
).add_to(fmap)
fmap
```
The map shows that the area is fully covered by products. The filtered products are indeed the most recent ones.
```
[p.properties["startTimeFromAscendingNode"] for p in filtered_products][::10]
```
| github_jupyter |
# Lab 01 : MNIST multi-layer -- demo
```
# For Google Colaboratory
import sys, os
if 'google.colab' in sys.modules:
# mount google drive
from google.colab import drive
drive.mount('/content/gdrive')
# find automatically the path of the folder containing "file_name" :
file_name = 'mnist_multilayer_demo.ipynb'
import subprocess
path_to_file = subprocess.check_output('find . -type f -name ' + str(file_name), shell=True).decode("utf-8")
path_to_file = path_to_file.replace(file_name,"").replace('\n',"")
# if previous search failed or too long, comment the previous line and simply write down manually the path below :
#path_to_file = '/content/gdrive/My Drive/CS5242_2021_codes/codes/labs_lecture06/lab01_mnist_multilayer'
print(path_to_file)
# change current path to the folder containing "file_name"
os.chdir(path_to_file)
!pwd
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from random import randint
import time
import utils
```
### Download the data
```
from utils import check_mnist_dataset_exists
data_path=check_mnist_dataset_exists()
train_data=torch.load(data_path+'mnist/train_data.pt')
train_label=torch.load(data_path+'mnist/train_label.pt')
test_data=torch.load(data_path+'mnist/test_data.pt')
test_label=torch.load(data_path+'mnist/test_label.pt')
```
### Make a two layer net class.
```
class two_layer_net(nn.Module):
def __init__(self, input_size, hidden_size, output_size):
super(two_layer_net , self).__init__()
self.layer1 = nn.Linear( input_size , hidden_size , bias=False )
self.layer2 = nn.Linear( hidden_size , output_size , bias=False )
def forward(self, x):
y = self.layer1(x)
y_hat = F.relu(y)
scores = self.layer2(y_hat)
return scores
```
### Build the net (recall that a one layer net had 7,840 parameters)
```
net=two_layer_net(784,50,10)
print(net)
utils.display_num_param(net)
```
### Choose the criterion, optimizer, batchsize, learning rate
```
criterion = nn.CrossEntropyLoss()
optimizer=torch.optim.SGD( net.parameters() , lr=0.01 )
bs=20
```
### Evaluate on test set
```
def eval_on_test_set():
running_error=0
num_batches=0
for i in range(0,10000,bs):
minibatch_data = test_data[i:i+bs]
minibatch_label= test_label[i:i+bs]
inputs = minibatch_data.view(bs,784)
scores=net( inputs )
error = utils.get_error( scores , minibatch_label)
running_error += error.item()
num_batches+=1
total_error = running_error/num_batches
print( 'test error = ', total_error*100 ,'percent')
```
### Training loop
```
start = time.time()
for epoch in range(200):
running_loss=0
running_error=0
num_batches=0
shuffled_indices=torch.randperm(60000)
for count in range(0,60000,bs):
# forward and backward pass
optimizer.zero_grad()
indices=shuffled_indices[count:count+bs]
minibatch_data = train_data[indices]
minibatch_label= train_label[indices]
inputs = minibatch_data.view(bs,784)
inputs.requires_grad_()
scores=net( inputs )
loss = criterion( scores , minibatch_label)
loss.backward()
optimizer.step()
# compute some stats
running_loss += loss.detach().item()
error = utils.get_error( scores.detach() , minibatch_label)
running_error += error.item()
num_batches+=1
# once the epoch is finished we divide the "running quantities"
# by the number of batches
total_loss = running_loss/num_batches
total_error = running_error/num_batches
elapsed_time = time.time() - start
# every 10 epoch we display the stats
# and compute the error rate on the test set
if epoch % 5 == 0 :
print(' ')
print('epoch=',epoch, '\t time=', elapsed_time,
'\t loss=', total_loss , '\t error=', total_error*100 ,'percent')
eval_on_test_set()
```
### Choose image at random from the test set and see how good/bad are the predictions
```
# choose a picture at random
idx=randint(0, 10000-1)
im=test_data[idx]
# diplay the picture
utils.show(im)
# feed it to the net and display the confidence scores
scores = net( im.view(1,784))
probs= F.softmax(scores, dim=1)
utils.show_prob_mnist(probs)
```
| github_jupyter |
# 15 Classes et objets
Dans cette section nous abordons la **programmation orienté objet** (POO).
## Types définis par le programmeur
Nous avons utilisés des nombreux types internes (int, float, str, bool, etc.); maintant nous allons définir un nouveau type. A titre d'example nous allons créer un type ``Point`` qui représente un point dans l'espace.
Nous allons définir une nouvelle classe. Le formalisme est similaire à une définiton de fonction:
* un mot-clé (``class``)
* un nom (``Point``)
* un deux-points (``:``)
* un bloc indenté
```
class Point:
"""Represents a 2D point in space."""
```
Ceci crée un objet **classe** dans l'espace ``__main__``
```
print(Point)
```
Une **classe** est une sorte de modèle pour créer des entitiés (instances). Créons une instance ``pt``
```
pt = Point()
pt
p0 = Point()
p1 = Point()
p0
p1
```
La création d'un nouvel objet à partir d'une classe s'appelle **instantiation**, et l'objet est une **instance** de la classe.
## Attributs
Vous pouvez attribuer des valeurs (attributs) à une instance en utilisant la **notation pointée**:
```
pt.x = 13.0
pt.y = 14.0
print(pt.x, pt.y)
pt.x + 5
```
Nous pouvons utiliser la notation pointés dans des expressions mathématiques. Par exemple:
```
import math
distance = math.sqrt(pt.x**2 + pt.y**2)
distance
```
Nous pouvons aussi utiliser un objet comme paramètre d'une fonction:
```
def print_point(p):
print('({}, {})'.format(p.x, p.y))
```
La fonction ``print_point`` prend comme argument un objet de la classe ``Point``.
```
print_point(pt)
```
**Exercice**
Ecrivez une fonction appelé ``distance_entre_points`` qui calcule la distance entre deux points.
```
p1 = Point()
p2 = Point()
p1.x = 3
p1.y = 4
p2.x = 5
p2.y = 10
def distance_entre_points(p1, p2):
dx = p2.x - p1.x
dy = p2.y - p1.y
d = math.sqrt(dx**2 + dy**2)
return d
distance_entre_points(p1,p2)
```
## Rectangles
Imaginez que vous devez concevoir une classe pour représenter des rectangles. Ils existent plusieurs manière de définir un rectangle et vous devez faire un choix approprié:
* un coin et largeur/hauteur
* le centre et largeur/hauteur
* deux points en diagonale
```
class Rectangle:
"""Represents a rectangle.
attributes: position (x, y), width (w), height (h)."""
```
To create a rectangle we write:
```
rect = Rectangle()
rect.w = 100
rect.h = 200
rect.pos = Point()
rect.pos.x = 0
rect.pos.y = 0
vars(rect)
```
## Instance comme valeur de retour
Une fonction peut prendre une instance comme argument et peux aussi retourner une instance comme résultat:
```
def find_center(rect):
p = Point()
p.x = rect.pos.x + rect.w/2
p.y = rect.pos.y + rect.h/2
return p
center = find_center(rect)
print_point(center)
```
## Les objets sont modifiables
Vous pouvez modifier des objets:
```
def resize_rect(rect, dw, dh):
"""Increase rect by delta width (dw) and delta height (dh)."""
rect.w += dw
rect.h += dh
resize_rect(rect, 99, 99)
print(rect.w, rect.h)
```
**Exercice**
Ecrivez une fonction appelé **move_rect** qui déplace un rectangle de la distance (dx, dy).
```
def move_rect(rect, dx, dy):
rect.pos.x += dx
rect.pos.y += dy
move_rect(rect, 10, 10)
print(rect.pos.x, rect.pos.y)
```
## Copier
Quand une variable pointe vers un objet, cet objet n'est pas copié. La variable est une référence (pointeur) vers l'objet en mémoire:
```
p1 = Point()
p2 = p1
```
La fonction ``is`` permet de voir si deux variable pointent sur le même objet.
```
p1 is p2
```
Nous pouvons vérifier que effectivement ``p1`` et ``p2`` pointent vers la même adresse en mémoire.
```
print(p1)
print(p2)
```
Pour faire une vraie copie, nous importons le module ``copy``.
```
import copy
p3 = copy.copy(p1)
p3 = Point()
p3.x = p1.x
p3.y = p1.y
```
Vérifions:
```
print(p1 is p3)
print(p1)
print(p3)
```
## Exercices
Ecrivez une définition pour une classe nommée **Cercle** ayant les attributs **centre** et **rayon**.
```
class Cercle:
""" Définit un carle avec centre et rayon """
c = Cercle()
c.center = Point()
c.rayon = 5
c.center.x = 4
c.center.y = 3
vars(c)
```
Instanciez un objet **Cercle** qui représente un cercle ayant son centre à (150, 100) et un rayon de 75.
```
c2 = Cercle()
c2.center = Point()
c2.rayon = 75
c2.center.x = 150
c2.center.y = 100
vars(c2)
```
Écrivez une fonction nommée **point_du_cercle** qui prend un **Cercle** et un **Point** et renvoie Vrai si le Point se trouve dans le cercle ou sur sa circonférence.
```
def point_du_cercle(c, p):
#dx = p.x - c.center.x
#dy = p.y - c.center.y
#d = maths.sqrt(dx**2 + dy**2)
d = distance_entre_points(c.center, p)
return d <= c.rayon
p0 = Point()
p0.x = -1
p0.y = 1
point_du_cercle(c, p0)
vars(p0)
```
Écrivez une fonction nommée **rect_du_cercle** qui prend un **Cercle** et un **Rectangle** et renvoie Vrai si le rectangle se trouve entièrement dans le cercle ou sur sa circonférence.
Écrivez une fonction nommée **rect_chevauche_cercle** qui prend un **Cercle** et un **Rectangle** et renvoie Vrai si l'un des coins du Rectangle se trouve à l'intérieur du cercle. Ou, version plus difficile, retourne Vrai si une partie quelconque du Rectangle se trouve à l'intérieur du cercle.
Écrivez une fonction appelée **dessine_rect** qui prend un objet **Tortue** et un **Rectangle** et utilise la Tortue pour dessiner le Rectangle. Voir le chapitre 4 pour des exemples utilisant des objets Tortue.
Écrivez une fonction appelée **dessine_cercle** qui prend une tortue et un cercle et dessine le cercle.
| github_jupyter |
```
import mido
from midiutil.MidiFile import MIDIFile
import matplotlib
import pandas as pd
import string
import os
import sys
import pandas as pd
import IPython.display as ipd
import librosa.display
import pretty_midi
from matplotlib import pyplot as plt
import numpy as np
def postprocessing(arr2):
x=arr2.shape[0]
arr3 = np.zeros((88,x,2), dtype=int)
#Array to store the start and end times of each note played in key
start=1
k=0
for j in range(arr2.shape[1]):
for i in range(arr2.shape[0]):
if arr2[i,j]==1 and start==1:
arr3[j,k,0]=i
start=0
elif arr2[i,j]==0 and start==0:
arr3[j,k,1]=i
start=1
k+=1
k=0
#Array to store the end times of each key
arr4=np.zeros((88), dtype=int)
for i in range(arr3.shape[0]):
for j in range(arr3.shape[1]):
if arr3[i,j,0]==0:
arr4[i]=j
break
return[arr3,arr4]
midi_data = pretty_midi.PrettyMIDI("/mnt/d/MAPS/test/MUS/MAPS_MUS-bor_ps6_ENSTDkCl.mid")
def plot_piano_roll(pm, start_pitch, end_pitch, sr=16000):
librosa.display.specshow(pm.get_piano_roll(sr)[start_pitch:end_pitch],
hop_length=1, sr=sr, x_axis='time', y_axis='cqt_note',
fmin=pretty_midi.note_number_to_hz(start_pitch),fmax=pretty_midi.note_number_to_hz(end_pitch))
pm = pretty_midi.PrettyMIDI('/mnt/d/MAPS/test/MUS/MAPS_MUS-bor_ps6_ENSTDkCl.mid')
plt.figure(figsize=(26, 7))
plot_piano_roll(pm,21,109,10)
from keras.models import load_model
rnn=load_model('Saved_Model.h5')
xtest1=np.load("testingCQT.npy")
xtest1=np.reshape(xtest1,(xtest1.shape[0]//100,100,252))
rnn_predictions = rnn.predict(xtest1, batch_size=1, verbose = 1)
#Now reshape Predictions back to 2D form.
rnn_predictions = np.reshape(rnn_predictions,(xtest1.shape[0]*xtest1.shape[1],88))
rnn_predictions = np.array(rnn_predictions).round()
#Convert all the values greater than 1 to 1.
rnn_predictions[rnn_predictions > 1] = 1
def arr_to_midi(source_array,duration_array):
mf = MIDIFile(1,deinterleave=False)
track = 0
time = 0
channel = 0
volume = 30
delta = 0.03125
mf.addTrackName(track, time, "Output3")
mf.addTempo(track, time, 60)
for i in range(source_array.shape[0]):
for j in range(duration_array[i]):
pitch = i + 21
time = delta*source_array[i,j,0]
duration = delta*( source_array[i,j,1] - source_array[i,j,0] )
mf.addNote(track, channel, pitch, time, duration, volume)
with open("Output3.mid", 'wb') as outf:
mf.writeFile(outf)
return mf
arr5,arr6=postprocessing(rnn_predictions)
mf=arr_to_midi(arr5,arr6)
print(arr6)
midi_data1 = pretty_midi.PrettyMIDI("Output3.mid")
plt.figure(figsize=(26, 7))
plot_piano_roll(midi_data1,21,109,10)
midi_list = []
for instrument in midi_data1.instruments:
for note in instrument.notes:
start = note.start
end = note.end
pitch = note.pitch
velocity = note.velocity
midi_list.append([start, end, pitch, velocity, instrument.name])
midi_list = sorted(midi_list, key=lambda x: (x[0], x[2]))
df = pd.DataFrame(midi_list, columns=['Start', 'End', 'Pitch', 'Velocity', 'Instrument'])
print(df)
plot_colour_graph("output2.mid", "PREDICTED")
```
| github_jupyter |
# Training metrics
*Metrics* for training fastai models are simply functions that take `input` and `target` tensors, and return some metric of interest for training. You can write your own metrics by defining a function of that type, and passing it to [`Learner`](/basic_train.html#Learner) in the [code]metrics[/code] parameter, or use one of the following pre-defined functions.
```
from fastai.gen_doc.nbdoc import *
from fastai.basics import *
```
## Predefined metrics:
```
show_doc(accuracy)
jekyll_warn("This metric is intended for classification of objects belonging to a single class.")
show_doc(accuracy_thresh)
```
Prediction are compared to `thresh` after `sigmoid` is maybe applied. Then we count the numbers that match the targets.
```
jekyll_note("This function is intended for one-hot-encoded targets (often in a multiclassification problem).")
show_doc(top_k_accuracy)
show_doc(dice)
show_doc(error_rate)
show_doc(mean_squared_error)
show_doc(mean_absolute_error)
show_doc(mean_squared_logarithmic_error)
show_doc(exp_rmspe)
show_doc(root_mean_squared_error)
show_doc(fbeta)
```
`beta` determines the value of the fbeta applied, `eps` is there for numeric stability. If `sigmoid=True`, a sigmoid is applied to the predictions before comparing them to `thresh` then to the targets. See the [F1 score wikipedia page](https://en.wikipedia.org/wiki/F1_score) for details on the fbeta score.
```
jekyll_note("This function is intended for one-hot-encoded targets (often in a multiclassification problem).")
show_doc(explained_variance)
show_doc(r2_score)
```
The following metrics are classes, don't forget to instantiate them when you pass them to a [`Learner`](/basic_train.html#Learner).
```
show_doc(RMSE, title_level=3)
show_doc(ExpRMSPE, title_level=3)
show_doc(Precision, title_level=3)
show_doc(Recall, title_level=3)
show_doc(FBeta, title_level=3)
show_doc(R2Score, title_level=3)
show_doc(ExplainedVariance, title_level=3)
show_doc(MatthewsCorreff, title_level=3)
show_doc(KappaScore, title_level=3)
show_doc(ConfusionMatrix, title_level=3)
```
## Creating your own metric
Creating a new metric can be as simple as creating a new function. If your metric is an average over the total number of elements in your dataset, just write the function that will compute it on a batch (taking `pred` and `targ` as arguments). It will then be automatically averaged over the batches (taking their different sizes into acount).
Sometimes metrics aren't simple averages however. If we take the example of precision for instance, we have to divide the number of true positives by the number of predictions we made for that class. This isn't an average over the number of elements we have in the dataset, we only consider those where we made a positive prediction for a specific thing. Computing the precision for each batch, then averaging them will yield to a result that may be close to the real value, but won't be it exactly (and it really depends on how you deal with special case of 0 positive predictions).
This why in fastai, every metric is implemented as a callback. If you pass a regular function, the library transforms it to a proper callback called `AverageCallback`. The callback metrics are only called during the validation phase, and only for the following events:
- <code>on_epoch_begin</code> (for initialization)
- <code>on_batch_begin</code> (if we need to have a look at the input/target and maybe modify them)
- <code>on_batch_end</code> (to analyze the last results and update our computation)
- <code>on_epoch_end</code>(to wrap up the final result that should be stored in `.metric`)
As an example, the following code is the exact implementation of the [`AverageMetric`](/callback.html#AverageMetric) callback that transforms a function like [`accuracy`](/metrics.html#accuracy) into a metric callback.
```
class AverageMetric(Callback):
def __init__(self, func):
self.func, self.name = func, func.__name__
def on_epoch_begin(self, **kwargs):
self.val, self.count = 0.,0
def on_batch_end(self, last_output, last_target, train, **kwargs):
self.count += last_target.size(0)
self.val += last_target.size(0) * self.func(last_output, last_target).detach().item()
def on_epoch_end(self, **kwargs):
self.metric = self.val/self.count
```
And here is another example that properly computes the precision for a given class.
```
class Precision(Callback):
def on_epoch_begin(self, **kwargs):
self.correct, self.total = 0, 0
def on_batch_end(self, last_output, last_target, **kwargs):
preds = last_output.argmax(1)
self.correct += ((preds==0) * (last_target==0)).float().sum()
self.total += (preds==0).float().sum()
def on_epoch_end(self, **kwargs):
self.metric = self.correct/self.total
```
The following custom callback class example measures peak RAM usage during each epoch:
```
import tracemalloc
class TraceMallocMetric(Callback):
def __init__(self):
super().__init__()
self.name = "peak RAM"
def on_epoch_begin(self, **kwargs):
tracemalloc.start()
def on_epoch_end(self, **kwargs):
current, peak = tracemalloc.get_traced_memory()
tracemalloc.stop()
self.metric = torch.tensor(peak)
```
To deploy it, you need to pass an instance of this custom metric in the [`metrics`](/metrics.html#metrics) argument:
```
learn = create_cnn(data, model, metrics=[accuracy, TraceMallocMetric()])
learn.fit_one_cycle(3, max_lr=1e-2)
```
And then the output changes to:
```
Total time: 00:54
epoch train_loss valid_loss accuracy peak RAM
1 0.333352 0.084342 0.973800 2395541.000000
2 0.096196 0.038386 0.988300 2342145.000000
3 0.048722 0.029234 0.990200 2342680.000000
```
As mentioner earlier, using the [`metrics`](/metrics.html#metrics) argument with a custom metrics class is limited in the number of phases of the callback system it can access, it can only return one numerical value and as you can see its output is hardcoded to have 6 points of precision in the output, even if the number is an int.
To overcome these limitations callback classes should be used instead.
For example, the following class:
* uses phases not available for the metric classes
* it reports 3 columns, instead of just one
* its column report ints, instead of floats
```
import tracemalloc
class TraceMallocMultiColMetric(LearnerCallback):
_order=-20 # Needs to run before the recorder
def __init__(self, learn):
super().__init__(learn)
self.train_max = 0
def on_train_begin(self, **kwargs):
self.learn.recorder.add_metric_names(['used', 'max_used', 'peak'])
def on_batch_end(self, train, **kwargs):
# track max memory usage during the train phase
if train:
current, peak = tracemalloc.get_traced_memory()
self.train_max = max(self.train_max, current)
def on_epoch_begin(self, **kwargs):
tracemalloc.start()
def on_epoch_end(self, **kwargs):
current, peak = tracemalloc.get_traced_memory()
tracemalloc.stop()
self.learn.recorder.add_metrics([current, self.train_max, peak])
```
Note, that it subclasses [`LearnerCallback`](/basic_train.html#LearnerCallback) and not [`Callback`](/callback.html#Callback), since the former provides extra features not available in the latter.
Also `_order=-20` is crucial - without it the custom columns will not be added - it tells the callback system to run this callback before the recorder system.
To deploy it, you need to pass the name of the class (not an instance!) of the class in the `callback_fns` argument. This is because the `learn` object doesn't exist yet, and it's required to instantiate `TraceMallocMultiColMetric`. The system will do it for us automatically as soon as the learn object has been created.
```
learn = create_cnn(data, model, metrics=[accuracy], callback_fns=TraceMallocMultiColMetric)
learn.fit_one_cycle(3, max_lr=1e-2)
```
And then the output changes to:
```
Total time: 00:53
epoch train_loss valid_loss accuracy used max_used peak
1 0.321233 0.068252 0.978600 156504 2408404 2419891
2 0.093551 0.032776 0.988500 79343 2408404 2348085
3 0.047178 0.025307 0.992100 79568 2408404 2342754
```
Another way to do the same is by using `learn.callbacks.append`, and this time we need to instantiate `TraceMallocMultiColMetric` with `learn` object which we now have, as it is called after the latter was created:
```
learn = create_cnn(data, model, metrics=[accuracy])
learn.callbacks.append(TraceMallocMultiColMetric(learn))
learn.fit_one_cycle(3, max_lr=1e-2)
```
Configuring the custom metrics in the `learn` object sets them to run in all future [`fit`](/basic_train.html#fit)-family calls. However, if you'd like to configure it for just one call, you can configure it directly inside [`fit`](/basic_train.html#fit) or [`fit_one_cycle`](/train.html#fit_one_cycle):
```
learn = create_cnn(data, model, metrics=[accuracy])
learn.fit_one_cycle(3, max_lr=1e-2, callbacks=TraceMallocMultiColMetric(learn))
```
And to stress the differences:
* the `callback_fns` argument expects a classname or a list of those
* the [`callbacks`](/callbacks.html#callbacks) argument expects an instance of a class or a list of those
* `learn.callbacks.append` expects a single instance of a class
For more examples, look inside fastai codebase and its test suite, search for classes that subclass either [`Callback`](/callback.html#Callback), [`LearnerCallback`](/basic_train.html#LearnerCallback) and subclasses of those two.
Finally, while the above examples all add to the metrics, it's not a requirement. A callback can do anything it wants and it is not required to add its outcomes to the metrics printout.
## Undocumented Methods - Methods moved below this line will intentionally be hidden
```
show_doc(FBeta.on_batch_end)
show_doc(FBeta.on_epoch_begin)
show_doc(FBeta.on_epoch_end)
show_doc(mae)
show_doc(msle)
show_doc(mse)
show_doc(rmse)
show_doc(Precision.on_epoch_end)
show_doc(FBeta.on_train_end)
show_doc(KappaScore.on_epoch_end)
show_doc(MatthewsCorreff.on_epoch_end)
show_doc(FBeta.on_train_begin)
show_doc(RMSE.on_epoch_end)
show_doc(ConfusionMatrix.on_train_begin)
show_doc(ConfusionMatrix.on_batch_end)
show_doc(ConfusionMatrix.on_epoch_end)
show_doc(Recall.on_epoch_end)
show_doc(ExplainedVariance.on_epoch_end)
show_doc(ExpRMSPE.on_epoch_end)
show_doc(ConfusionMatrix.on_epoch_begin)
show_doc(R2Score.on_epoch_end)
```
## New Methods - Please document or move to the undocumented section
| github_jupyter |
Introduction
============
Workshop description
--------------------
- This is an intermediate/advanced R course
- Appropriate for those with basic knowledge of R
- This is not a statistics course!
- Learning objectives:
- Learn the R formula interface
- Specify factor contrasts to test specific hypotheses
- Perform model comparisons
- Run and interpret variety of regression models in R
Materials and Setup
--------------------------
Lab computer users: Log in using the user name and password on the board to your left.
Laptop users:
- you should have R installed--if not, open a web browser and go to <http://cran.r-project.org> and download and install it
- also helpful to install RStudo (download from <http://rstudio.com>)
Everyone:
- Download materials from <http://tutorials.iq.harvard.edu/R/Rstatistics.zip>
- Extract materials from RStatistics.zip (on lab machines *right-click -> WinZip -> Extract to here*) and move to your desktop.
Launch RStudio<span class="tag" data-tag-name="labsetup"></span>
----------------------------------------------------------------
- Open the RStudio program from the Windows start menu
- Create a project in the Rstatistics folder you downloaded earlier:
- `File => New Project => Existing Directory => Browse` and select the Rstatistics folder.
Set working directory
---------------------
It is often helpful to start your R session by setting your working directory so you don't have to type the full path names to your data and other files
```
# set the working directory
# setwd("~/Desktop/Rstatistics")
# setwd("C:/Users/dataclass/Desktop/Rstatistics")
```
You might also start by listing the files in your working directory
```
getwd() # where am I?
list.files("dataSets") # files in the dataSets folder
```
Load the states data
--------------------
The *states.dta* data comes from <http://anawida.de/teach/SS14/anawida/4.linReg/data/states.dta.txt> and appears to have originally appeared in *Statistics with Stata* by Lawrence C. Hamilton.
```
# read the states data
states.data <- readRDS("dataSets/states.rds")
#get labels
states.info <- data.frame(attributes(states.data)[c("names", "var.labels")])
#look at last few labels
tail(states.info, 8)
```
Linear regression
=================
Examine the data before fitting models
--------------------------------------
Start by examining the data to check for problems.
```
# summary of expense and csat columns, all rows
sts.ex.sat <- subset(states.data, select = c("expense", "csat"))
summary(sts.ex.sat)
# correlation between expense and csat
cor(sts.ex.sat)
```
Plot the data before fitting models
-----------------------------------
Plot the data to look for multivariate outliers, non-linear relationships etc.
```
# scatter plot of expense vs csat
plot(sts.ex.sat)
```

Linear regression example
-------------------------
- Linear regression models can be fit with the `lm()` function
- For example, we can use `lm` to predict SAT scores based on per-pupal expenditures:
```
# Fit our regression model
sat.mod <- lm(csat ~ expense, # regression formula
data=states.data) # data set
# Summarize and print the results
summary(sat.mod) # show regression coefficients table
```
Why is the association between expense and SAT scores *negative*?
-----------------------------------------------------------------
Many people find it surprising that the per-capita expenditure on students is negatively related to SAT scores. The beauty of multiple regression is that we can try to pull these apart. What would the association between expense and SAT scores be if there were no difference among the states in the percentage of students taking the SAT?
```
summary(lm(csat ~ expense + percent, data = states.data))
```
The lm class and methods
------------------------
OK, we fit our model. Now what?
- Examine the model object:
```
class(sat.mod)
names(sat.mod)
methods(class = class(sat.mod))[1:9]
```
- Use function methods to get more information about the fit
```
confint(sat.mod)
# hist(residuals(sat.mod))
```
Linear Regression Assumptions
-----------------------------
- Ordinary least squares regression relies on several assumptions, including that the residuals are normally distributed and homoscedastic, the errors are independent and the relationships are linear.
- Investigate these assumptions visually by plotting your model:
```
par(mar = c(4, 4, 2, 2), mfrow = c(1, 2)) #optional
plot(sat.mod, which = c(1, 2)) # "which" argument optional
```
Comparing models
----------------
Do congressional voting patterns predict SAT scores over and above expense? Fit two models and compare them:
```
# fit another model, adding house and senate as predictors
sat.voting.mod <- lm(csat ~ expense + house + senate,
data = na.omit(states.data))
sat.mod <- update(sat.mod, data=na.omit(states.data))
# compare using the anova() function
anova(sat.mod, sat.voting.mod)
coef(summary(sat.voting.mod))
```
Exercise 0: least squares regression
------------------------------------
Use the *states.rds* data set. Fit a model predicting energy consumed per capita (energy) from the percentage of residents living in metropolitan areas (metro). Be sure to
1. Examine/plot the data before fitting the model
2. Print and interpret the model `summary`
3. `plot` the model to look for deviations from modeling assumptions
Select one or more additional predictors to add to your model and repeat steps 1-3. Is this model significantly better than the model with *metro* as the only predictor?
Interactions and factors
========================
Modeling interactions
---------------------
Interactions allow us assess the extent to which the association between one predictor and the outcome depends on a second predictor. For example: Does the association between expense and SAT scores depend on the median income in the state?
```
#Add the interaction to the model
sat.expense.by.percent <- lm(csat ~ expense*income,
data=states.data)
#Show the results
coef(summary(sat.expense.by.percent)) # show regression coefficients table
```
Regression with categorical predictors
--------------------------------------
Let's try to predict SAT scores from region, a categorical variable. Note that you must make sure R does not think your categorical variable is numeric.
```
# make sure R knows region is categorical
str(states.data$region)
states.data$region <- factor(states.data$region)
#Add region to the model
sat.region <- lm(csat ~ region,
data=states.data)
#Show the results
coef(summary(sat.region)) # show regression coefficients table
anova(sat.region) # show ANOVA table
```
Again, **make sure to tell R which variables are categorical by converting them to factors!**
Setting factor reference groups and contrasts
---------------------------------------------
In the previous example we use the default contrasts for region. The default in R is treatment contrasts, with the first level as the reference. We can change the reference group or use another coding scheme using the `C` function.
```
# print default contrasts
contrasts(states.data$region)
# change the reference group
coef(summary(lm(csat ~ C(region, base=4),
data=states.data)))
# change the coding scheme
coef(summary(lm(csat ~ C(region, contr.helmert),
data=states.data)))
```
See also `?contrasts`, `?contr.treatment`, and `?relevel`.
Exercise 1: interactions and factors
------------------------------------
Use the states data set.
1. Add on to the regression equation that you created in exercise 1 by generating an interaction term and testing the interaction.
2. Try adding region to the model. Are there significant differences across the four regions?
Regression with binary outcomes
===============================
Logistic regression
-------------------
This far we have used the `lm` function to fit our regression models. `lm` is great, but limited--in particular it only fits models for continuous dependent variables. For categorical dependent variables we can use the `glm()` function.
For these models we will use a different dataset, drawn from the National Health Interview Survey. From the [CDC website](http://www.cdc.gov/nchs/nhis.htm):
> The National Health Interview Survey (NHIS) has monitored the health of the nation since 1957. NHIS data on a broad range of health topics are collected through personal household interviews. For over 50 years, the U.S. Census Bureau has been the data collection agent for the National Health Interview Survey. Survey results have been instrumental in providing data to track health status, health care access, and progress toward achieving national health objectives.
Load the National Health Interview Survey data:
```
NH11 <- readRDS("dataSets/NatHealth2011.rds")
labs <- attributes(NH11)$labels
```
Logistic regression example
---------------------------
Let's predict the probability of being diagnosed with hypertension based on age, sex, sleep, and bmi
```
str(NH11$hypev) # check stucture of hypev
levels(NH11$hypev) # check levels of hypev
# collapse all missing values to NA
NH11$hypev <- factor(NH11$hypev, levels=c("2 No", "1 Yes"))
# run our regression model
hyp.out <- glm(hypev~age_p+sex+sleep+bmi,
data=NH11, family="binomial")
coef(summary(hyp.out))
```
Logistic regression coefficients
--------------------------------
Generalized linear models use link functions, so raw coefficients are difficult to interpret. For example, the age coefficient of .06 in the previous model tells us that for every one unit increase in age, the log odds of hypertension diagnosis increases by 0.06. Since most of us are not used to thinking in log odds this is not too helpful!
One solution is to transform the coefficients to make them easier to interpret
```
hyp.out.tab <- coef(summary(hyp.out))
hyp.out.tab[, "Estimate"] <- exp(coef(hyp.out))
hyp.out.tab
```
Generating predicted values
---------------------------
In addition to transforming the log-odds produced by `glm` to odds, we can use the `predict()` function to make direct statements about the predictors in our model. For example, we can ask "How much more likely is a 63 year old female to have hypertension compared to a 33 year old female?".
```
# Create a dataset with predictors set at desired levels
predDat <- with(NH11,
expand.grid(age_p = c(33, 63),
sex = "2 Female",
bmi = mean(bmi, na.rm = TRUE),
sleep = mean(sleep, na.rm = TRUE)))
# predict hypertension at those levels
cbind(predDat, predict(hyp.out, type = "response",
se.fit = TRUE, interval="confidence",
newdata = predDat))
```
This tells us that a 33 year old female has a 13% probability of having been diagnosed with hypertension, while and 63 year old female has a 48% probability of having been diagnosed.
Packages for computing and graphing predicted values
----------------------------------------------------
Instead of doing all this ourselves, we can use the effects package to compute quantities of interest for us (cf. the Zelig package).
```
library(effects)
plot(allEffects(hyp.out))
```

Exercise 2: logistic regression
-------------------------------
Use the NH11 data set that we loaded earlier.
1. Use glm to conduct a logistic regression to predict ever worked (everwrk) using age (age_p) and marital status (r_maritl).
2. Predict the probability of working for each level of marital status.
Note that the data is not perfectly clean and ready to be modeled. You will need to clean up at least some of the variables before fitting the model.
Multilevel Modeling
===================
Multilevel modeling overview
----------------------------
- Multi-level (AKA hierarchical) models are a type of mixed-effects models
- Used to model variation due to group membership where the goal is to generalize to a population of groups
- Can model different intercepts and/or slopes for each group
- Mixed-effecs models include two types of predictors: fixed-effects and random effects
- Fixed-effects -- observed levels are of direct interest (.e.g, sex, political party...)
- Random-effects -- observed levels not of direct interest: goal is to make inferences to a population represented by observed levels
- In R the lme4 package is the most popular for mixed effects models
- Use the `lmer` function for liner mixed models, `glmer` for generalized mixed models
```
library(lme4)
```
The Exam data
-------------
The Exam data set contans exam scores of 4,059 students from 65 schools in Inner London. The variable names are as follows:
| variable | Description |
|----------|----------------------------------------------------------------------------------------------------|
| school | School ID - a factor. |
| normexam | Normalized exam score. |
| schgend | School gender - a factor. Levels are 'mixed', 'boys', and 'girls'. |
| schavg | School average of intake score. |
| vr | Student level Verbal Reasoning (VR) score band at intake - 'bottom 25%', 'mid 50%', and 'top 25%'. |
| intake | Band of student's intake score - a factor. Levels are 'bottom 25%', 'mid 50%' and 'top 25%'./ |
| standLRT | Standardised LR test score. |
| sex | Sex of the student - levels are 'F' and 'M'. |
| type | School type - levels are 'Mxd' and 'Sngl'. |
| student | Student id (within school) - a factor |
```
Exam <- readRDS("dataSets/Exam.rds")
```
The null model and ICC
----------------------
As a preliminary step it is often useful to partition the variance in the dependent variable into the various levels. This can be accomplished by running a null model (i.e., a model with a random effects grouping structure, but no fixed-effects predictors).
```
# null model, grouping by school but not fixed effects.
Norm1 <-lmer(normexam ~ 1 + (1|school),
data=na.omit(Exam), REML = FALSE)
summary(Norm1)
```
The is .169/(.169 + .848) = .17: 17% of the variance is at the school level.
Adding fixed-effects predictors
-------------------------------
Predict exam scores from student's standardized tests scores
```
Norm2 <-lmer(normexam~standLRT + (1|school),
data=na.omit(Exam),
REML = FALSE)
summary(Norm2)
```
Multiple degree of freedom comparisons
--------------------------------------
As with `lm` and `glm` models, you can compare the two `lmer` models using the `anova` function.
```
anova(Norm1, Norm2)
```
Random slopes
-------------
Add a random effect of students' standardized test scores as well. Now in addition to estimating the distribution of intercepts across schools, we also estimate the distribution of the slope of exam on standardized test.
```
Norm3 <- lmer(normexam~standLRT + (standLRT|school),
data = na.omit(Exam),
REML = FALSE)
summary(Norm3)
```
Test the significance of the random slope
-----------------------------------------
To test the significance of a random slope just compare models with and without the random slope term
```
anova(Norm2, Norm3)
```
Exercise 3: multilevel modeling
-------------------------------
Use the dataset, bh1996: `data(bh1996, package="multilevel")`
From the data documentation:
> Variables are Cohesion (COHES), Leadership Climate (LEAD), Well-Being (WBEING) and Work Hours (HRS). Each of these variables has two variants - a group mean version that replicates each group mean for every individual, and a within-group version where the group mean is subtracted from each individual response. The group mean version is designated with a G. (e.g., G.HRS), and the within-group version is designated with a W. (e.g., W.HRS).
1. Create a null model predicting wellbeing ("WBEING")
2. Calculate the ICC for your null model
3. Run a second multi-level model that adds two individual-level predictors, average number of hours worked ("HRS") and leadership skills ("LEAD") to the model and interpret your output.
4. Now, add a random effect of average number of hours worked ("HRS") to the model and interpret your output. Test the significance of this random term.
Exercise solutions<span class="tag" data-tag-name="prototype"></span>
=====================================================================
Exercise 0 prototype
--------------------
Use the *states.rds* data set.
```
states <- readRDS("dataSets/states.rds")
```
Fit a model predicting energy consumed per capita (energy) from the percentage of residents living in metropolitan areas (metro). Be sure to
1. Examine/plot the data before fitting the model
```
states.en.met <- subset(states, select = c("metro", "energy"))
summary(states.en.met)
plot(states.en.met)
cor(states.en.met, use="pairwise")
```
2. Print and interpret the model `summary`
```
mod.en.met <- lm(energy ~ metro, data = states)
summary(mod.en.met)
```
3. `plot` the model to look for deviations from modeling assumptions
```
plot(mod.en.met)
```
Select one or more additional predictors to add to your model and repeat steps 1-3. Is this model significantly better than the model with *metro* as the only predictor?
```
states.en.met.pop.wst <- subset(states, select = c("energy", "metro", "pop", "waste"))
summary(states.en.met.pop.wst)
plot(states.en.met.pop.wst)
cor(states.en.met.pop.wst, use = "pairwise")
mod.en.met.pop.waste <- lm(energy ~ metro + pop + waste, data = states)
summary(mod.en.met.pop.waste)
anova(mod.en.met, mod.en.met.pop.waste)
```
Exercise 1: prototype
---------------------
Use the states data set.
1. Add on to the regression equation that you created in exercise 1 by generating an interaction term and testing the interaction.
```
mod.en.metro.by.waste <- lm(energy ~ metro * waste, data = states)
```
1. Try adding a region to the model. Are there significant differences across the four regions?
```
mod.en.region <- lm(energy ~ metro * waste + region, data = states)
anova(mod.en.region)
```
Exercise 2 prototype
--------------------
Use the NH11 data set that we loaded earlier. Note that the data is not perfectly clean and ready to be modeled. You will need to clean up at least some of the variables before fitting the model.
1. Use glm to conduct a logistic regression to predict ever worked (everwrk) using age (age_p) and marital status (r_maritl).
```
nh11.wrk.age.mar <- subset(NH11, select = c("everwrk", "age_p", "r_maritl"))
summary(nh11.wrk.age.mar)
NH11 <- transform(NH11,
everwrk = factor(everwrk,
levels = c("1 Yes", "2 No")),
r_maritl = droplevels(r_maritl))
mod.wk.age.mar <- glm(everwrk ~ age_p + r_maritl, data = NH11,
family = "binomial")
summary(mod.wk.age.mar)
```
1. Predict the probability of working for each level of marital status.
```
library(effects)
data.frame(Effect("r_maritl", mod.wk.age.mar))
```
Exercise 3 prototype
--------------------
Use the dataset, bh1996:
```
data(bh1996, package="multilevel")
```
From the data documentation:
> Variables are Cohesion (COHES), Leadership Climate (LEAD), Well-Being (WBEING) and Work Hours (HRS). Each of these variables has two variants - a group mean version that replicates each group mean for every individual, and a within-group version where the group mean is subtracted from each individual response. The group mean version is designated with a G. (e.g., G.HRS), and the within-group version is designated with a W. (e.g., W.HRS).
Note that the group identifier is named "GRP".
1. Create a null model predicting wellbeing ("WBEING")
```
library(lme4)
mod.grp0 <- lmer(WBEING ~ 1 + (1 | GRP), data = bh1996)
summary(mod.grp0)
```
3. Run a second multi-level model that adds two individual-level predictors, average number of hours worked ("HRS") and leadership skills ("LEAD") to the model and interpret your output.
```
mod.grp1 <- lmer(WBEING ~ HRS + LEAD + (1 | GRP), data = bh1996)
summary(mod.grp1)
```
3. Now, add a random effect of average number of hours worked ("HRS") to the model and interpret your output. Test the significance of this random term.
```
mod.grp2 <- lmer(WBEING ~ HRS + LEAD + (1 + HRS | GRP), data = bh1996)
anova(mod.grp1, mod.grp2)
```
Wrap-up
=======
Help us make this workshop better!
----------------------------------
- Please take a moment to fill out a very short
feedback form
- These workshops exist for you -- tell us what you need!
- <http://tinyurl.com/RstatisticsFeedback>
Additional resources
--------------------
- IQSS workshops: <http://projects.iq.harvard.edu/rtc/filter_by/workshops>
- IQSS statistical consulting: <http://dss.iq.harvard.edu>
- Zelig
- Website: <http://gking.harvard.edu/zelig>
- Documentation: <http://r.iq.harvard.edu/docs/zelig.pdf>
- Ameila
- Website: <http://gking.harvard.edu/Amelia/>
- Documetation: <http://r.iq.harvard.edu/docs/amelia/amelia.pdf>
| github_jupyter |
# Meraki Python SDK Demo: Uplink Preference Restore
*This notebook demonstrates using the Meraki Python SDK to restore Internet (WAN) and VPN traffic uplink preferences, as well as custom performance classes, from an Excel file. If you have hundreds of WAN/VPN uplink preferences, they can be a challenge to manipulate. This demo seeks to prove how using the Meraki API and Python SDK can substantially streamline such complex deployments.*
If you haven't already, please consult the corresponding **Meraki Python SDK Demo: Uplink Preference Backup**.
If an admin has backed up his Internet and VPN traffic uplink preferences and custom performance classes, this tool will restore them to the Dashboard from the Excel file backup. This is a more advanced demo, intended for intermediate to advanced Python programmers, but has been documented thoroughly with the intention that even a determined Python beginner can understand the concepts involved.
If an admin can use the appropriate template Excel file and update it with the appropriate info, e.g. subnets, ports, and WAN preference, then this tool can push those preferences to the Dashboard for the desired network's MX appliance. With the Meraki Dashboard API, its SDK and Python, we can restore hundreds of preferences without using the GUI.
---
>NB: Throughout this notebook, we will print values for demonstration purposes. In a production Python script, the coder would likely remove these print statements to clean up the console output.
In this first cell, we import the required `meraki` and `os` modules, and open the Dashboard API connection using the SDK. We also import `openpyxl` for working with Excel files, and `netaddr` for working with IP addresses.
```
# Install the relevant modules. If you are using a local editor (e.g. VS Code, rather than Colab) you can run these commands, without the preceding %, via a terminal. NB: Run `pip install meraki==` to find the latest version of the Meraki SDK.
%pip install meraki
%pip install openpyxl
# If you are using Google Colab, please ensure you have set up your environment variables as linked above, then delete the two lines of ''' to activate the following code:
'''
%pip install colab-env -qU
import colab_env
'''
# The Meraki SDK
import meraki
# The built-in OS module, to read environment variables
import os
# The openpyxl module, to manipulate Excel files
import openpyxl
# The datetime module, to generate timestamps
# We're also going to import Python's built-in JSON module, but only to make the console output pretty. In production, you wouldn't need any of the printing calls at all, nor this import!
import json
# Setting API key this way, and storing it in the env variables, lets us keep the sensitive API key out of the script itself
# The meraki.DashboardAPI() method does not require explicitly passing this value; it will check the environment for a variable
# called 'MERAKI_DASHBOARD_API_KEY' on its own. In this case, API_KEY is shown simply as an reference to where that information is
# stored.
API_KEY = os.getenv('MERAKI_DASHBOARD_API_KEY')
# Initialize the Dashboard connection.
dashboard = meraki.DashboardAPI()
# We'll also create a few reusable strings for later interactivity.
CONFIRM_STRING = 'OK, are you sure you want to do this? This script does not have an "undo" feature.'
CANCEL_STRING = 'OK. Operation canceled.'
WORKING_STRING = 'Working...'
COMPLETE_STRING = 'Operation complete.'
NETWORK_SELECTED_STRING = 'Network selected.'
# Some of the parameters we'll work with are optional. This string defines what value will be put into a cell corresponding with a parameter that is not set on that rule.
NOT_APPLICABLE_STRING = 'N/A'
# Set the filename to use for the backup workbook
WORKBOOK_FILENAME = 'downloaded_rules_workbook_2020-08-14 194610.476691 cpc and vpn.xlsx'
```
Let's make a basic pretty print formatter, `printj()`. It will make reading the JSON via Python terminal later a lot easier, but won't be necessary in production scripts, where you're not expecting to print very much to the terminal.
```
def printj(ugly_json_object):
# The json.dumps() method converts a JSON object into human-friendly formatted text
pretty_json_string = json.dumps(ugly_json_object, indent = 2, sort_keys = False)
return print(pretty_json_string)
```
## Introducing a Python class
To streamline user interaction in a re-usable way, we'll create a class called UserChoice. Think of classes like a superset of functions, where you can store related functions and variables. Later, we'll create an instance of this class to prompt the user for input, and validate that input.
It may look complex, but it will streamline our code later, and is a great example of code-reuse in Python. For more information on classes, [click here](https://docs.python.org/3/tutorial/classes.html).
```
class UserChoice:
'A re-usable CLI option prompt.'
def __init__(self, options_list=[], subject_of_choice='available options', single_option_noun='option', id_parameter='id', name_parameter='name', action_verb='choose', no_valid_options_message='no valid options'):
# options_list is a list of dictionaries containing attributes id_parameter and name_parameter
self.options_list = options_list
# subject_of_choice is a string that names the subject of the user's choice. It is typically a plural noun.
self.subject_of_choice = subject_of_choice
# single_option_noun is a string that is a singular noun corresponding to the subject_of_choice
self.single_option_noun = single_option_noun
# id_parameter is a string that represents the name of the sub-parameter that serves as the ID value for the option in options_list. It should be a unique value for usability.
self.id_parameter = id_parameter
# name_paraemter is a string that represents the name of the sub-parameter that serves as the name value for the option in options_list. It does not need to be unique.
self.name_parameter = name_parameter
# action_verb is a string that represents the verb of the user's action. For example, to "choose"
self.action_verb = action_verb
# no_valid_options_message is a string that represents an error message if options_list is empty
self.no_valid_options_message = no_valid_options_message
# Confirm there are options in the list
if len(self.options_list):
print(f'We found {len(self.options_list)} {self.subject_of_choice}:')
# Label each option and show the user their choices.
option_index = 0
for option in self.options_list:
print(f"{option_index}. {option[self.id_parameter]} with name {option[self.name_parameter]}")
option_index+=1
print(f'Which {self.single_option_noun} would you like to {self.action_verb}?')
self.active_option = int(input(f'Choose 0-{option_index-1}:'))
# Ask until the user provides valid input.
while self.active_option not in list(range(option_index)):
print(f'{self.active_option} is not a valid choice. Which {self.single_option_noun} would you like to {self.action_verb}?')
self.active_option = int(input(f'Choose 0-{option_index-1}:'))
print(f'Your {self.single_option_noun} is {self.options_list[self.active_option][self.name_parameter]}.')
self.id = self.options_list[self.active_option][self.id_parameter]
self.name = self.options_list[self.active_option][self.name_parameter]
```
## Pulling organization and network IDs
Most API calls require passing values for the organization ID and/or the network ID. In the below cell, we fetch a list of the organizations the API key can access, then pick the first org in the list, and the first network in that organization, to use for later operations. You could re-use this code presuming your API key only has access to a single organization, and that organization only contains a single network. Otherwise, you would want to review the organizations object declared and printed here to review its contents. As a side exercise, perhaps you could use the class that we defined above, `UserChoice`, to let the user decide which organization to use!
```
# Let's make it easier to call this data later
# getOrganizations will return all orgs to which the supplied API key has access
organizations = dashboard.organizations.getOrganizations()
print('Organizations:')
printj(organizations)
# This example presumes we want to use the first organization as the scope for later operations.
firstOrganizationId = organizations[0]['id']
firstOrganizationName = organizations[0]['name']
# Print a blank line for legibility before showing the firstOrganizationId
print('')
print(f'The firstOrganizationId is {firstOrganizationId}, and its name is {firstOrganizationName}.')
```
Let's see what networks are in the chosen organization.
```
networks = dashboard.organizations.getOrganizationNetworks(organizationId=firstOrganizationId)
print('Networks:')
printj(networks)
```
## Identifying networks with MX appliances
Now that we've got the organization and network values figured out, we can get to the task at hand:
> Retore a backup of the uplink selection preferences, including custom performance classes.
We can only run this on networks that have appliance devices, so we have a `for` loop that checks each entry in the `networks` list. If the network's `productTypes` value contains `appliance`, then we'll ask the user to pick one, then pull the uplink selection rules from it.
```
# Create an empty list where we can store all of the organization's networks that have appliances
networks_with_appliances = []
# Let's fill up that list
for network in networks:
# We only want to examine networks that might contain appliances
if 'appliance' in network['productTypes']:
# Add the network to networks_with_appliances
networks_with_appliances.append(network)
NO_VALID_OPTIONS_MESSAGE = 'There are no networks with appliances in this organization. Please supply an API token that has access to an organization with an appliance in one of its networks.'
```
## Prompt the user to choose a network
Now let's ask the user which network they'd like to use. Remember that `UserChoice` class we created earlier? We'll call that and supply parameters defining what the user can choose. Notice how, having defined the class earlier, we can re-use it with only a single declaration. Run this block to watch how it runs the code inside of the class.
```
# If any are found, let the user choose a network. Otherwise, let the user know that none were found. The logic for this class is defined in a cell above.
network_choice = UserChoice(
options_list=networks_with_appliances,
subject_of_choice='networks with appliances',
single_option_noun='network',
no_valid_options_message=NO_VALID_OPTIONS_MESSAGE
)
```
## Overall restore workflow
### Logical summary
The restore workflow summarized is:
1. Open a chosen Excel workbook that contains the backup information.
2. Parse each worksheet into a Python object structured according to the API documentation.
3. Restore the custom performance classes from the backup.
4. Restore the WAN (Internet) and VPN uplink preferences from the backup.
### Code summary
To structure the code, we'll break down the total functionality into discrete functions. These functions are where we define the operational logic for the restore. Most, if not all functions will return relevant information to be used by the next function in the restore procedure.
1. The first function will ingest the Excel spreadsheet with the backup information and return the data structured as a Python object, according to the API documentation.
2. Another function will restore the custom performance classes from the backup. This is a tricky operation for reasons you'll see below, but fully possible via Python methods and the Meraki SDK.
3. Another function will, if necessary, update the loaded backup object with new ID assignments, in case restoring the custom performance classes backup resulted in new class IDs.
4. Another function will restore the VPN preferences.
5. Another function will restore the WAN preferencess.
6. After definining each of those functions, we'll run them in succession to finalize the backup restoration.
Once you understand the fundamentals, consider how you might improve this script, either with additional functionality or UX improvements!
> NB: *Function* and *method* are used interchangeably. Python is an object-oriented language, and *method* is often preferred when discussing object-oriented programming.
```
# Ingest an Excel spreadsheet with the appropriate worksheets, and create an object that can be pushed as a configuration API call
def load_uplink_prefs_workbook(workbook_filename):
# Create a workbook object out of the actual workbook file
loaded_workbook = openpyxl.load_workbook(workbook_filename, read_only=True)
# Create empty rule lists to which we can add the rules defined in the workbook
loaded_custom_performance_classes = []
loaded_wan_uplink_prefs = []
loaded_vpn_uplink_prefs = []
# Open the worksheets
loaded_custom_performance_classes_worksheet = loaded_workbook['customPerformanceClasses']
loaded_wan_prefs_worksheet = loaded_workbook['wanUplinkPreferences']
loaded_vpn_prefs_worksheet = loaded_workbook['vpnUplinkPreferences']
## CUSTOM PERFORMANCE CLASSES ##
# We'll also count the number of classes to help the user know that it's working.
performance_class_count = 0
# For reference, the expected column order is
# ID [0], Name [1], Max Latency [2], Max Jitter [3], Max Loss Percentage [4]
# Append each performance class loaded_custom_performance_classes
for row in loaded_custom_performance_classes_worksheet.iter_rows(min_row=2):
# Let's start with an empty rule dictionary to which we'll add the relevant parameters
performance_class = {}
# Append the values
performance_class['customPerformanceClassId'] = row[0].value
performance_class['name'] = row[1].value
performance_class['maxLatency'] = row[2].value
performance_class['maxJitter'] = row[3].value
performance_class['maxLossPercentage'] = row[4].value
# Append the performance class to the loaded_custom_performance_classes list
loaded_custom_performance_classes.append(performance_class)
performance_class_count += 1
print(f'Loaded {performance_class_count} custom performance classes.')
## WAN PREFERENCES ##
# We'll also count the number of rules to help the user know that it's working.
rule_count = 0
# For reference, the expected column order is
# Protocol [0], Source [1], Src port [2], Destination [3], Dst port [4], Preferred uplink [5]
# Append each WAN preference to loaded_wan_uplink_prefs
for row in loaded_wan_prefs_worksheet.iter_rows(min_row=2):
# Let's start with an empty rule dictionary to which we'll add the relevant parameters
rule = {}
# We know that there will always be a preferred uplink
rule_preferred_uplink = row[5].value.lower()
# The first column is Protocol
rule_protocol = row[0].value.lower()
if rule_protocol == 'any':
# Since protocol is 'any' then src and dst ports are also 'any'
rule_src_port = 'any'
rule_dst_port = 'any'
else:
# Since protocol is not 'any', we pass these as-is
rule_src_port = row[1].value.lower()
rule_dst_port = row[4].value.lower()
# Next column is Source [1]
rule_source = row[1].value.lower()
# Last column to fill is Destination [3]
rule_destination = row[3].value.lower()
# Assemble the rule into a single Python object that uses the syntax that the corresponding API call expects
if rule_protocol == 'any':
# Protocol is any, so leave out the port numbers
rule_value = {
'protocol': rule_protocol,
'source': {
'cidr': rule_source
},
'destination': {
'cidr': rule_destination
}
}
else:
# Rule isn't any, so we need the port numbers
rule_value = {
'protocol': rule_protocol,
'source': {
'port': rule_src_port,
'cidr': rule_source
},
'destination': {
'port': rule_dst_port,
'cidr': rule_destination
}
}
# Append the trafficFilters param to the rule
rule['trafficFilters'] = [
{
'type': 'custom', # This worksheet doesn't have any Type column
'value': rule_value
}
]
# Append the preferredUplink param to the rule
rule['preferredUplink'] = rule_preferred_uplink
# Append the rule to the loaded_wan_uplink_prefs list
loaded_wan_uplink_prefs.append(rule)
rule_count += 1
print(f'Loaded {rule_count} WAN uplink preferences.')
## VPN PREFERENCES ##
# For reference, the expected column order is
# Type [0], Protocol or App ID [1], Source or App Name [2], Src port [3], Destination [4],
# Dst port [5], Preferred uplink [6], Failover criterion [7], Performance class type [8],
# Performance class name [9], Performance class ID [10]
# We'll also count the number of rules to help the user know that it's working.
rule_count = 0
# Append each WAN preference to loaded_wan_uplink_prefs
for row in loaded_vpn_prefs_worksheet.iter_rows(min_row=2):
# Since the parameters can change depending on the various options, we'll start with an empty dictionary or list depending on parameter type and then add keys along the way to correspond to the relevant values.
rule = {}
rule_traffic_filters = {}
# We know that there will always be a preferred uplink. We don't need any special logic to assign this one, so we'll keep it at the top.
rule_preferred_uplink = row[6].value
# Add it to the rule
rule['preferredUplink'] = rule_preferred_uplink
# The first column is Type, and the type will define the structure for other parameters.
rule_type = row[0].value.lower() # Always lowercase.Chimpkennuggetss
# If the rule type is application or applicationCategory then we're not concerned with destination, dst port or similar, and Protocol or App ID [1] will be 'id' not 'protocol', etc.
if 'application' in rule_type:
rule_application_id = row[1].value.lower() # Always lowercase
rule_application_name = row[2].value # Leave it capitalized
# Assign the rule value
rule_value = {
'id': rule_application_id,
'name': rule_application_name
}
else:
# Assign the rule Protocol [1]
rule_protocol = row[1].value.lower() # Always lowercase
# Regardless of protocol, we need to assign Source [2]
rule_source = row[2].value.lower() # Always lowercase
# Regardless of protocol, we need to assign Destination [4]
rule_destination = row[4].value.lower() # Always lowercase
# Assign the rule ports, if appropriate
if rule_protocol in ('any', 'icmp'):
# Since protocol is 'any' or 'icmp' then we leave out src and dst ports
rule_value = {
'protocol': rule_protocol,
'source': {
'cidr': rule_source
},
'destination': {
'cidr': rule_destination
}
}
else:
# Since protocol is not 'any', we pass these from the worksheet
rule_src_port = row[3].value.lower() # Always lowercase
rule_dst_port = row[5].value.lower() # Always lowercase
rule_value = {
'protocol': rule_protocol,
'source': {
'port': rule_src_port,
'cidr': rule_source
},
'destination': {
'port': rule_dst_port,
'cidr': rule_destination
}
}
# Assemble the rule_traffic_filters parameter
rule_traffic_filters['type'] = rule_type
rule_traffic_filters['value'] = rule_value
# Add it to the rule
rule_traffic_filters_list = [rule_traffic_filters]
rule['trafficFilters'] = rule_traffic_filters_list
# Assign the optional failOverCriterion
rule_failover_criterion = row[7].value # Leave it capitalized
if rule_failover_criterion not in (NOT_APPLICABLE_STRING, ''):
# Add it to the rule
rule['failOverCriterion'] = rule_failover_criterion
# Assign the optional performanceClass
rule_performance_class_type = row[8].value
rule_performance_class_name = row[9].value
rule_performance_class_id = row[10].value
if rule_performance_class_type not in (NOT_APPLICABLE_STRING, ''):
# Add it to the rule
rule['performanceClass'] = {}
rule['performanceClass']['type'] = rule_performance_class_type
# If the performance class type is custom, then we use customPerformanceClassId
if rule_performance_class_type == 'custom':
# Add it to the rule
rule['performanceClass']['customPerformanceClassId'] = rule_performance_class_id
# Otherwise, we use builtinPerformanceClassName
else:
# Add it to the rule
rule['performanceClass']['builtinPerformanceClassName'] = rule_performance_class_name
# Append the rule to the loaded_vpn_uplink_prefs list
loaded_vpn_uplink_prefs.append(rule)
rule_count += 1
print(f'Loaded {rule_count} VPN uplink preferences.')
return(
{
'wanPrefs': loaded_wan_uplink_prefs,
'vpnPrefs': loaded_vpn_uplink_prefs,
'customPerformanceClasses': loaded_custom_performance_classes
}
)
# We'll use the filename we specified at the top of the notebook.
# Load the workbook!
loaded_combined_uplink_prefs = load_uplink_prefs_workbook(WORKBOOK_FILENAME)
```
Let's take a look at those uplink preferences!
```
printj(loaded_combined_uplink_prefs['vpnPrefs'])
# How might we look at the other components of the loaded backup?
```
## Restoring custom performance classes
Restoring custom performance classes can be tricky. IDs are unique, but a user might have changed the name or settings of a performance class after the last backup was taken, and that does not change the performance class's ID.
Given this scenario, and many other hypotheticals, we will simplify the restore operation with a straightforward and predictable behavior. It is designed to be most in-line with common expectations about what "restoring a backup" commonly means.
First we will check if the backup's classes are a perfect match to the currently configured ones. If so, there's no need to restore anything.
Otherwise, if the backup contains a performance class with the same ID as one that exists in the current Dashboard configuration, then we will overwrite that existing class with the settings (including name) from the backup.
Otherwise, if the backup contains a performance class with a different ID but the same name as one that exists in the current Dashboard configuration, then we will overwrite that existing class with the settings from the backup, and we will return the new/old IDs
in a list of key-value pairs so that we can update the corresponding `vpnUplinkPreference` to use this new performance class ID. If that happens, then when the uplink preferences are restored in a later function `update_vpn_prefs_performance_class_ids`, they will use the same performance class settings as were backed up, but the updated performance class ID.
Finally, if the backup contains a performance class that doesn't match any of the existing classes by name or ID, then we'll create it new, and return the new/old IDs as described above for a later function `update_vpn_prefs_performance_class_ids`.
```
# A new function to compare current vs. loaded custom performance classes
# It will take as input the network ID and the list of the custom performance classes loaded from the backup
# If it has to update any VPN prefs' custom performance class IDs, it will do so, and return the new/old IDs
# in a list of key-value pairs. Otherwise, it will return None.
def restore_custom_performance_classes(*, listOfLoadedCustomPerformanceClasses, networkId):
# Let's first make a list of the custom performance classes currently configured in the dashboard
list_of_current_custom_performance_classes = dashboard.appliance.getNetworkApplianceTrafficShapingCustomPerformanceClasses(networkId=networkId)
# Let's compare the currently configured classes with those from the backup. If they're the same, there's no need to restore anything.
if list_of_current_custom_performance_classes == listOfLoadedCustomPerformanceClasses:
print('No differences found, all done!')
return(None)
# Otherwise, we will make several lists that we will use to compare the current config with the backup (loaded) config to determine what needs to be changed. We'll use the copy() method of the list object to make a new copy rather than creating a new reference to the original data.
# We'll remove from this list as we find classes that match by either ID or name
list_of_orphan_loaded_performance_classes = listOfLoadedCustomPerformanceClasses.copy()
list_of_orphan_current_performance_classes = list_of_current_custom_performance_classes.copy()
# We'll subtract from this list as we find classes that match by ID
list_of_id_unmatched_current_performance_classes = list_of_current_custom_performance_classes.copy()
# For those instances where we match by name, we'll store the new:old IDs in a new list
list_of_id_updates = []
# First let's check for current classes that match by ID. If they do, we'll update them with the loaded backup config.
for loaded_performance_class in listOfLoadedCustomPerformanceClasses:
# Let's look through each of the currently configured performance classes for that match
for current_performance_class in list_of_current_custom_performance_classes:
# Check if the IDs match up
if loaded_performance_class['customPerformanceClassId'] == current_performance_class['customPerformanceClassId']:
print(f"Matched {loaded_performance_class['customPerformanceClassId']} by ID! Restoring it from the backup.")
# Restore that class from the loaded backup configuration
dashboard.appliance.updateNetworkApplianceTrafficShapingCustomPerformanceClass(
networkId=networkId,
customPerformanceClassId=current_performance_class['customPerformanceClassId'],
maxJitter=loaded_performance_class['maxJitter'],
maxLatency=loaded_performance_class['maxLatency'],
name=loaded_performance_class['name'],
maxLossPercentage=loaded_performance_class['maxLossPercentage']
)
# Remove each from its respective orphan list
list_of_orphan_loaded_performance_classes.remove(loaded_performance_class)
list_of_orphan_current_performance_classes.remove(current_performance_class)
# Let's next check the orphan lists for classes that match by name. If they do, we'll update them with the loaded backup config.
# If we find a match, we'll also add a reference object to the name-only match list tying the new ID to the respective one from
# the loaded backup. If we find a match, we'll also remove it from both orphan lists.
for orphan_loaded_performance_class in list_of_orphan_loaded_performance_classes:
# Let's look through each of the currently configured performance classes for that match
for orphan_current_performance_class in list_of_orphan_current_performance_classes:
# Check if the names match up
if orphan_loaded_performance_class['name'] == orphan_current_performance_class['name']:
print(f"Matched custom performance class with ID {orphan_loaded_performance_class['customPerformanceClassId']} by name {orphan_loaded_performance_class['name']}! Restoring it from the backup.")
# Restore that class from the loaded backup configuration
dashboard.appliance.updateNetworkApplianceTrafficShapingCustomPerformanceClass(
networkId=networkId,
customPerformanceClassId=orphan_current_performance_class['customPerformanceClassId'],
maxJitter=orphan_loaded_performance_class['maxJitter'],
maxLatency=orphan_loaded_performance_class['maxLatency'],
name=orphan_loaded_performance_class['name'],
maxLossPercentage=orphan_loaded_performance_class['maxLossPercentage']
)
# Add it to the name-only matches list, list_of_name_matches
list_of_id_updates.append(
{
'loaded_id': orphan_loaded_performance_class['customPerformanceClassId'],
'current_id': orphan_current_performance_class['customPerformanceClassId']
}
)
# Remove each from its respective orphan list
list_of_orphan_loaded_performance_classes.remove(orphan_loaded_performance_class)
list_of_orphan_current_performance_classes.remove(orphan_current_performance_class)
# If there are any orphans left, they have not matched by ID or name. Create them new.
if len(list_of_orphan_loaded_performance_classes):
for orphan_loaded_performance_class in list_of_orphan_loaded_performance_classes:
# Re-create the loaded class from the backup and get its new ID
# We'll also add the old and new IDs to the reference object we've created for this purpose
new_performance_class = dashboard.appliance.createNetworkApplianceTrafficShapingCustomPerformanceClass(
networkId=networkId,
maxJitter=orphan_loaded_performance_class['maxJitter'],
maxLatency=orphan_loaded_performance_class['maxLatency'],
name=orphan_loaded_performance_class['name'],
maxLossPercentage=orphan_loaded_performance_class['maxLossPercentage']
)
# Add it to the name-only matches list, list_of_name_matches
list_of_id_updates.append(
{
'loaded_id': orphan_loaded_performance_class['customPerformanceClassId'],
'current_id': new_performance_class['customPerformanceClassId']
}
)
# Remove it from the list of orphans
list_of_orphan_loaded_performance_classes.remove(orphan_loaded_performance_class)
# Return the list of updated IDs in key-value pairs for later processing
return(list_of_id_updates)
```
## Restoring VPN uplink preferences
Now that the custom performance classes are restored, we can restore the VPN uplink preferences.
### Method to update the VPN prefs with updated performance class IDs
This nesting-doll of a function simply looks for `vpnUplinkPreferences` that use custom performance classes, and updates those IDs to match any corresponding new ones created, such as when a backed up performance class was deleted after the backup.
```
def update_vpn_prefs_performance_class_ids(*, loaded_vpn_prefs, performance_class_id_updates):
vpn_prefs_updates = 0
# For each update in the ID updates list
for update in performance_class_id_updates:
# For each rule in loaded_vpn_prefs
for rule in loaded_vpn_prefs:
# If the rule's performance class type is set
if 'performanceClass' in rule.keys():
# If the rule's performance class type is custom
if rule['performanceClass']['type'] == 'custom':
# And if the rule's customPerformanceClassId matches one from our ID updates list
if rule['performanceClass']['customPerformanceClassId'] == update['loaded_id']:
# Then we update it with the new ID
rule['performanceClass']['customPerformanceClassId'] = update['current_id']
vpn_prefs_updates += 1
return(vpn_prefs_updates)
```
### Method to restore the VPN preferences to Dashboard
Specify the `networkId` and provide the VPN uplink preferences as a list. This method is documented [here](https://developer.cisco.com/meraki/api-v1/#!update-network-appliance-traffic-shaping-uplink-selection).
> NB: Setting a variable `response` equal to the SDK method is a common practice, because the SDK method will return the API's HTTP response. That information is useful to confirm that the operation was successful, but it is not strictly required.
```
# A new function to push VPN preferences
def restore_vpn_prefs(vpn_prefs_list):
response = dashboard.appliance.updateNetworkApplianceTrafficShapingUplinkSelection(
networkId=network_choice.id,
vpnTrafficUplinkPreferences=vpn_prefs_list
)
return(response)
```
### Method to restore the WAN preferences to Dashboard
Specify the `networkId` and provide the WAN uplink preferences as a list. This method is documented [here](https://developer.cisco.com/meraki/api-v1/#!update-network-appliance-traffic-shaping-uplink-selection).
> NB: Notice that this relies on the same SDK method as `restore_vpn_prefs()` above. Here we've split the restore into two functions to demonstrate that you can push only specific keyword arguments when the other keyword arguments are optional. Since it's the same method, we could consolidate this function, `restore_wan_prefs()`, and `restore_vpn_prefs()`, into a single function by passing both keyword arguments `vpnTrafficUplinkPreferences` and `wanTrafficUplinkPreferences` at the same time. This would then increase the amount of work accomplished by a single API call. We recommend following a best practice of accomplishing as much as possible with as few calls as possible, when appropriate.
```
# A new function to push WAN preferences
def restore_wan_prefs(wan_prefs_list):
response = dashboard.appliance.updateNetworkApplianceTrafficShapingUplinkSelection(
networkId=network_choice.id,
wanTrafficUplinkPreferences=wan_prefs_list
)
return(response)
```
## Wrapping up!
We've now built functions to handle the discrete tasks required to restore the configuration for the three items:
* WAN uplink preferences
* VPN uplink preferences
* Custom performance classes
We had to translate the Excel workbook into a Python object that was structured according to the API specifications.
We found that some extra logic was required to properly restore custom performance classes and VPN uplink preferences that use them, and wrote custom functions to handle it.
> NB: We handled this one way of potentially many. Can you think of any other ways you might handle the problem of missing custom performance class IDs, or overlapping names or IDs?
However, we haven't actually called any of these functions, so the restore hasn't happened! To actually call those functions, we'll run them in the next cell.
### Restore the backup!
```
# Restore the custom performance classes
updated_performance_class_ids = restore_custom_performance_classes(
listOfLoadedCustomPerformanceClasses=loaded_combined_uplink_prefs['customPerformanceClasses'],
networkId=network_choice.id
)
# Update the custom perfromance class IDs
if updated_performance_class_ids:
update_vpn_prefs_performance_class_ids(
loaded_vpn_prefs=loaded_combined_uplink_prefs['vpnPrefs'],
performance_class_id_updates=updated_performance_class_ids
)
# Restore the VPN prefs
restored_vpn_prefs = restore_vpn_prefs(
loaded_combined_uplink_prefs['vpnPrefs']
)
# Restore the WAN prefs
restored_vpn_prefs = restore_wan_prefs(
loaded_combined_uplink_prefs['wanPrefs']
)
```
# Final thoughts
And we're done! Hopefully this was a useful deep dive into Python programming and interacting with the Meraki SDK and Excel workbooks. We tackled a problem that is tough to solve in the Dashboard GUI and showed how it can be done very quickly via API and the Python SDK.
Of course, writing this much Python to solve your own issues can be a daunting task. That is why it is always relevant to consider how often you plan to solve a given problem, and invest your time accordingly.
Here we used Excel workbooks, but you can imagine that there are all types of data structures that might be used instead of Excel workbooks, e.g. CSVs, plain text, YAML, XML, LibreOffice files or others, and with the right code you can use those instead of Excel.
## Further learning
Consider whether there are any "gotchas" for which we didn't prepare in this code demo. For example, what if you already have the maximum number of custom performance classes defined in the Dashboard--how might that interfere with your ability to restore the backup using this code? What might you add or change to handle that possibility?
[Meraki Interactive API Docs](https://developer.cisco.com/meraki/api-v1/#!overview): The official (and interactive!) Meraki API and SDK documentation repository on DevNet.
| github_jupyter |
##### Copyright 2019 The TensorFlow Hub Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
# Copyright 2018 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
# Semantic Search with Approximate Nearest Neighbors and Text Embeddings
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/hub/tutorials/semantic_approximate_nearest_neighbors"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/hub/blob/master/examples/colab/semantic_approximate_nearest_neighbors.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/hub/blob/master/examples/colab/semantic_approximate_nearest_neighbors.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/hub/examples/colab/semantic_approximate_nearest_neighbors.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
This tutorial illustrates how to generate embeddings from a [TensorFlow Hub](https://tfhub.dev) (TF-Hub) module given input data, and build an approximate nearest neighbours (ANN) index using the extracted embeddings. The index can then be used for real-time similarity matching and retrieval.
When dealing with a large corpus of data, it's not efficient to perform exact matching by scanning the whole repository to find the most similar items to a given query in real-time. Thus, we use an approximate similarity matching algorithm which allows us to trade off a little bit of accuracy in finding exact nearest neighbor matches for a significant boost in speed.
In this tutorial, we show an example of real-time text search over a corpus of news headlines to find the headlines that are most similar to a query. Unlike keyword search, this captures the semantic similarity encoded in the text embedding.
The steps of this tutorial are:
1. Download sample data.
2. Generate embeddings for the data using a TF-Hub module
3. Build an ANN index for the embeddings
4. Use the index for similarity matching
We use [Apache Beam](https://beam.apache.org/documentation/programming-guide/) with [TensorFlow Transform](https://www.tensorflow.org/tfx/tutorials/transform/simple) (TF-Transform) to generate the embeddings from the TF-Hub module. We also use Spotify's [ANNOY](https://github.com/spotify/annoy) library to build the approximate nearest neighbours index. You cam find benchmarking of ANN framework in this [Github repository](https://github.com/erikbern/ann-benchmarks).
This tutorial uses TensorFlow 1.0 and works only with TF1 [Hub modules](https://www.tensorflow.org/hub/tf1_hub_module) from TF-Hub. See the updated [TF2 version of this tutorial](https://github.com/tensorflow/hub/blob/master/examples/colab/tf2_semantic_approximate_nearest_neighbors.ipynb).
## Setup
Install the required libraries.
```
!pip install -q tensorflow_transform
!pip install -q apache_beam
!pip install -q sklearn
!pip install -q annoy
```
Import the required libraries
```
import os
import sys
import pathlib
import pickle
from collections import namedtuple
from datetime import datetime
import numpy as np
import apache_beam as beam
import annoy
from sklearn.random_projection import gaussian_random_matrix
import tensorflow.compat.v1 as tf
import tensorflow_transform as tft
import tensorflow_hub as hub
import tensorflow_transform.beam as tft_beam
print('TF version: {}'.format(tf.__version__))
print('TF-Hub version: {}'.format(hub.__version__))
print('TF-Transform version: {}'.format(tft.__version__))
print('Apache Beam version: {}'.format(beam.__version__))
```
## 1. Download Sample Data
[A Million News Headlines](https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/SYBGZL#) dataset contains news headlines published over a period of 15 years sourced from the reputable Australian Broadcasting Corp. (ABC). This news dataset has a summarised historical record of noteworthy events in the globe from early-2003 to end-2017 with a more granular focus on Australia.
**Format**: Tab-separated two-column data: 1) publication date and 2) headline text. We are only interested in the headline text.
```
!wget 'https://dataverse.harvard.edu/api/access/datafile/3450625?format=tab&gbrecs=true' -O raw.tsv
!wc -l raw.tsv
!head raw.tsv
```
For simplicity, we only keep the headline text and remove the publication date
```
!rm -r corpus
!mkdir corpus
with open('corpus/text.txt', 'w') as out_file:
with open('raw.tsv', 'r') as in_file:
for line in in_file:
headline = line.split('\t')[1].strip().strip('"')
out_file.write(headline+"\n")
!tail corpus/text.txt
```
## Helper function to load a TF-Hub module
```
def load_module(module_url):
embed_module = hub.Module(module_url)
placeholder = tf.placeholder(dtype=tf.string)
embed = embed_module(placeholder)
session = tf.Session()
session.run([tf.global_variables_initializer(), tf.tables_initializer()])
print('TF-Hub module is loaded.')
def _embeddings_fn(sentences):
computed_embeddings = session.run(
embed, feed_dict={placeholder: sentences})
return computed_embeddings
return _embeddings_fn
```
## 2. Generate Embeddings for the Data.
In this tutorial, we use the [Universal Sentence Encoder](https://tfhub.dev/google/universal-sentence-encoder/2) to generate emebeddings for the headline data. The sentence embeddings can then be easily used to compute sentence level meaning similarity. We run the embedding generation process using Apache Beam and TF-Transform.
### Embedding extraction method
```
encoder = None
def embed_text(text, module_url, random_projection_matrix):
# Beam will run this function in different processes that need to
# import hub and load embed_fn (if not previously loaded)
global encoder
if not encoder:
encoder = hub.Module(module_url)
embedding = encoder(text)
if random_projection_matrix is not None:
# Perform random projection for the embedding
embedding = tf.matmul(
embedding, tf.cast(random_projection_matrix, embedding.dtype))
return embedding
```
### Make TFT preprocess_fn method
```
def make_preprocess_fn(module_url, random_projection_matrix=None):
'''Makes a tft preprocess_fn'''
def _preprocess_fn(input_features):
'''tft preprocess_fn'''
text = input_features['text']
# Generate the embedding for the input text
embedding = embed_text(text, module_url, random_projection_matrix)
output_features = {
'text': text,
'embedding': embedding
}
return output_features
return _preprocess_fn
```
### Create dataset metadata
```
def create_metadata():
'''Creates metadata for the raw data'''
from tensorflow_transform.tf_metadata import dataset_metadata
from tensorflow_transform.tf_metadata import schema_utils
feature_spec = {'text': tf.FixedLenFeature([], dtype=tf.string)}
schema = schema_utils.schema_from_feature_spec(feature_spec)
metadata = dataset_metadata.DatasetMetadata(schema)
return metadata
```
### Beam pipeline
```
def run_hub2emb(args):
'''Runs the embedding generation pipeline'''
options = beam.options.pipeline_options.PipelineOptions(**args)
args = namedtuple("options", args.keys())(*args.values())
raw_metadata = create_metadata()
converter = tft.coders.CsvCoder(
column_names=['text'], schema=raw_metadata.schema)
with beam.Pipeline(args.runner, options=options) as pipeline:
with tft_beam.Context(args.temporary_dir):
# Read the sentences from the input file
sentences = (
pipeline
| 'Read sentences from files' >> beam.io.ReadFromText(
file_pattern=args.data_dir)
| 'Convert to dictionary' >> beam.Map(converter.decode)
)
sentences_dataset = (sentences, raw_metadata)
preprocess_fn = make_preprocess_fn(args.module_url, args.random_projection_matrix)
# Generate the embeddings for the sentence using the TF-Hub module
embeddings_dataset, _ = (
sentences_dataset
| 'Extract embeddings' >> tft_beam.AnalyzeAndTransformDataset(preprocess_fn)
)
embeddings, transformed_metadata = embeddings_dataset
# Write the embeddings to TFRecords files
embeddings | 'Write embeddings to TFRecords' >> beam.io.tfrecordio.WriteToTFRecord(
file_path_prefix='{}/emb'.format(args.output_dir),
file_name_suffix='.tfrecords',
coder=tft.coders.ExampleProtoCoder(transformed_metadata.schema))
```
### Generaring Random Projection Weight Matrix
[Random projection](https://en.wikipedia.org/wiki/Random_projection) is a simple, yet powerfull technique used to reduce the dimensionality of a set of points which lie in Euclidean space. For a theoretical background, see the [Johnson-Lindenstrauss lemma](https://en.wikipedia.org/wiki/Johnson%E2%80%93Lindenstrauss_lemma).
Reducing the dimensionality of the embeddings with random projection means less time needed to build and query the ANN index.
In this tutorial we use [Gaussian Random Projection](https://en.wikipedia.org/wiki/Random_projection#Gaussian_random_projection) from the [Scikit-learn](https://scikit-learn.org/stable/modules/random_projection.html#gaussian-random-projection) library.
```
def generate_random_projection_weights(original_dim, projected_dim):
random_projection_matrix = None
if projected_dim and original_dim > projected_dim:
random_projection_matrix = gaussian_random_matrix(
n_components=projected_dim, n_features=original_dim).T
print("A Gaussian random weight matrix was creates with shape of {}".format(random_projection_matrix.shape))
print('Storing random projection matrix to disk...')
with open('random_projection_matrix', 'wb') as handle:
pickle.dump(random_projection_matrix,
handle, protocol=pickle.HIGHEST_PROTOCOL)
return random_projection_matrix
```
### Set parameters
If you want to build an index using the original embedding space without random projection, set the `projected_dim` parameter to `None`. Note that this will slow down the indexing step for high-dimensional embeddings.
```
module_url = 'https://tfhub.dev/google/universal-sentence-encoder/2' #@param {type:"string"}
projected_dim = 64 #@param {type:"number"}
```
### Run pipeline
```
import tempfile
output_dir = pathlib.Path(tempfile.mkdtemp())
temporary_dir = pathlib.Path(tempfile.mkdtemp())
g = tf.Graph()
with g.as_default():
original_dim = load_module(module_url)(['']).shape[1]
random_projection_matrix = None
if projected_dim:
random_projection_matrix = generate_random_projection_weights(
original_dim, projected_dim)
args = {
'job_name': 'hub2emb-{}'.format(datetime.utcnow().strftime('%y%m%d-%H%M%S')),
'runner': 'DirectRunner',
'batch_size': 1024,
'data_dir': 'corpus/*.txt',
'output_dir': output_dir,
'temporary_dir': temporary_dir,
'module_url': module_url,
'random_projection_matrix': random_projection_matrix,
}
print("Pipeline args are set.")
args
!rm -r {output_dir}
!rm -r {temporary_dir}
print("Running pipeline...")
%time run_hub2emb(args)
print("Pipeline is done.")
!ls {output_dir}
```
Read some of the generated embeddings...
```
import itertools
embed_file = os.path.join(output_dir, 'emb-00000-of-00001.tfrecords')
sample = 5
record_iterator = tf.io.tf_record_iterator(path=embed_file)
for string_record in itertools.islice(record_iterator, sample):
example = tf.train.Example()
example.ParseFromString(string_record)
text = example.features.feature['text'].bytes_list.value
embedding = np.array(example.features.feature['embedding'].float_list.value)
print("Embedding dimensions: {}".format(embedding.shape[0]))
print("{}: {}".format(text, embedding[:10]))
```
## 3. Build the ANN Index for the Embeddings
[ANNOY](https://github.com/spotify/annoy) (Approximate Nearest Neighbors Oh Yeah) is a C++ library with Python bindings to search for points in space that are close to a given query point. It also creates large read-only file-based data structures that are mmapped into memory. It is built and used by [Spotify](https://www.spotify.com) for music recommendations.
```
def build_index(embedding_files_pattern, index_filename, vector_length,
metric='angular', num_trees=100):
'''Builds an ANNOY index'''
annoy_index = annoy.AnnoyIndex(vector_length, metric=metric)
# Mapping between the item and its identifier in the index
mapping = {}
embed_files = tf.gfile.Glob(embedding_files_pattern)
print('Found {} embedding file(s).'.format(len(embed_files)))
item_counter = 0
for f, embed_file in enumerate(embed_files):
print('Loading embeddings in file {} of {}...'.format(
f+1, len(embed_files)))
record_iterator = tf.io.tf_record_iterator(
path=embed_file)
for string_record in record_iterator:
example = tf.train.Example()
example.ParseFromString(string_record)
text = example.features.feature['text'].bytes_list.value[0].decode("utf-8")
mapping[item_counter] = text
embedding = np.array(
example.features.feature['embedding'].float_list.value)
annoy_index.add_item(item_counter, embedding)
item_counter += 1
if item_counter % 100000 == 0:
print('{} items loaded to the index'.format(item_counter))
print('A total of {} items added to the index'.format(item_counter))
print('Building the index with {} trees...'.format(num_trees))
annoy_index.build(n_trees=num_trees)
print('Index is successfully built.')
print('Saving index to disk...')
annoy_index.save(index_filename)
print('Index is saved to disk.')
print("Index file size: {} GB".format(
round(os.path.getsize(index_filename) / float(1024 ** 3), 2)))
annoy_index.unload()
print('Saving mapping to disk...')
with open(index_filename + '.mapping', 'wb') as handle:
pickle.dump(mapping, handle, protocol=pickle.HIGHEST_PROTOCOL)
print('Mapping is saved to disk.')
print("Mapping file size: {} MB".format(
round(os.path.getsize(index_filename + '.mapping') / float(1024 ** 2), 2)))
embedding_files = "{}/emb-*.tfrecords".format(output_dir)
embedding_dimension = projected_dim
index_filename = "index"
!rm {index_filename}
!rm {index_filename}.mapping
%time build_index(embedding_files, index_filename, embedding_dimension)
!ls
```
## 4. Use the Index for Similarity Matching
Now we can use the ANN index to find news headlines that are semantically close to an input query.
### Load the index and the mapping files
```
index = annoy.AnnoyIndex(embedding_dimension)
index.load(index_filename, prefault=True)
print('Annoy index is loaded.')
with open(index_filename + '.mapping', 'rb') as handle:
mapping = pickle.load(handle)
print('Mapping file is loaded.')
```
### Similarity matching method
```
def find_similar_items(embedding, num_matches=5):
'''Finds similar items to a given embedding in the ANN index'''
ids = index.get_nns_by_vector(
embedding, num_matches, search_k=-1, include_distances=False)
items = [mapping[i] for i in ids]
return items
```
### Extract embedding from a given query
```
# Load the TF-Hub module
print("Loading the TF-Hub module...")
g = tf.Graph()
with g.as_default():
embed_fn = load_module(module_url)
print("TF-Hub module is loaded.")
random_projection_matrix = None
if os.path.exists('random_projection_matrix'):
print("Loading random projection matrix...")
with open('random_projection_matrix', 'rb') as handle:
random_projection_matrix = pickle.load(handle)
print('random projection matrix is loaded.')
def extract_embeddings(query):
'''Generates the embedding for the query'''
query_embedding = embed_fn([query])[0]
if random_projection_matrix is not None:
query_embedding = query_embedding.dot(random_projection_matrix)
return query_embedding
extract_embeddings("Hello Machine Learning!")[:10]
```
### Enter a query to find the most similar items
```
#@title { run: "auto" }
query = "confronting global challenges" #@param {type:"string"}
print("Generating embedding for the query...")
%time query_embedding = extract_embeddings(query)
print("")
print("Finding relevant items in the index...")
%time items = find_similar_items(query_embedding, 10)
print("")
print("Results:")
print("=========")
for item in items:
print(item)
```
## Want to learn more?
You can learn more about TensorFlow at [tensorflow.org](https://www.tensorflow.org/) and see the TF-Hub API documentation at [tensorflow.org/hub](https://www.tensorflow.org/hub/). Find available TensorFlow Hub modules at [tfhub.dev](https://tfhub.dev/) including more text embedding modules and image feature vector modules.
Also check out the [Machine Learning Crash Course](https://developers.google.com/machine-learning/crash-course/) which is Google's fast-paced, practical introduction to machine learning.
| github_jupyter |
This is the third blog post of [Object Detection with YOLO blog series](https://fairyonice.github.io/tag/object-detection-using-yolov2-on-pascal-voc2012-series.html). This blog discusses the YOLO's model architecture. I will use PASCAL VOC2012 data. This blog assumes that the readers have read the previous two blog posts - [Part 1](https://fairyonice.github.io/Part_1_Object_Detection_with_Yolo_for_VOC_2014_data_anchor_box_clustering.html), [Part 2](https://fairyonice.github.io/Part%202_Object_Detection_with_Yolo_using_VOC_2014_data_input_and_output_encoding.html).
## Andrew Ng's YOLO lecture
- [Neural Networks - Bounding Box Predictions](https://www.youtube.com/watch?v=gKreZOUi-O0&t=0s&index=7&list=PL_IHmaMAvkVxdDOBRg2CbcJBq9SY7ZUvs)
- [C4W3L06 Intersection Over Union](https://www.youtube.com/watch?v=ANIzQ5G-XPE&t=7s)
- [C4W3L07 Nonmax Suppression](https://www.youtube.com/watch?v=VAo84c1hQX8&t=192s)
- [C4W3L08 Anchor Boxes](https://www.youtube.com/watch?v=RTlwl2bv0Tg&t=28s)
- [C4W3L09 YOLO Algorithm](https://www.youtube.com/watch?v=9s_FpMpdYW8&t=34s)
## Reference
- [You Only Look Once:Unified, Real-Time Object Detection](https://arxiv.org/pdf/1506.02640.pdf)
- [YOLO9000:Better, Faster, Stronger](https://arxiv.org/pdf/1612.08242.pdf)
- [experiencor/keras-yolo2](https://github.com/experiencor/keras-yolo2)
## Reference in my blog
- [Part 1 Object Detection using YOLOv2 on Pascal VOC2012 - anchor box clustering](https://fairyonice.github.io/Part_1_Object_Detection_with_Yolo_for_VOC_2014_data_anchor_box_clustering.html)
- [Part 2 Object Detection using YOLOv2 on Pascal VOC2012 - input and output encoding](https://fairyonice.github.io/Part%202_Object_Detection_with_Yolo_using_VOC_2014_data_input_and_output_encoding.html)
- [Part 3 Object Detection using YOLOv2 on Pascal VOC2012 - model](https://fairyonice.github.io/Part_3_Object_Detection_with_Yolo_using_VOC_2012_data_model.html)
- [Part 4 Object Detection using YOLOv2 on Pascal VOC2012 - loss](https://fairyonice.github.io/Part_4_Object_Detection_with_Yolo_using_VOC_2012_data_loss.html)
- [Part 5 Object Detection using YOLOv2 on Pascal VOC2012 - training](https://fairyonice.github.io/Part_5_Object_Detection_with_Yolo_using_VOC_2012_data_training.html)
- [Part 6 Object Detection using YOLOv2 on Pascal VOC 2012 data - inference on image](https://fairyonice.github.io/Part_6_Object_Detection_with_Yolo_using_VOC_2012_data_inference_image.html)
- [Part 7 Object Detection using YOLOv2 on Pascal VOC 2012 data - inference on video](https://fairyonice.github.io/Part_7_Object_Detection_with_Yolo_using_VOC_2012_data_inference_video.html)
## My GitHub repository
This repository contains all the ipython notebooks in this blog series and the funcitons (See backend.py).
- [FairyOnIce/ObjectDetectionYolo](https://github.com/FairyOnIce/ObjectDetectionYolo)
```
import matplotlib.pyplot as plt
import numpy as np
import os
import sys
print(sys.version)
%matplotlib inline
```
## Define anchor box
<code>ANCHORS</code> defines the number of anchor boxes and the shape of each anchor box.
The choice of the anchor box specialization is already discussed in [Part 1 Object Detection using YOLOv2 on Pascal VOC2012 - anchor box clustering](https://fairyonice.github.io/Part_1_Object_Detection_with_Yolo_for_VOC_2014_data_anchor_box_clustering.html).
Based on the K-means analysis in the previous blog post, I will select 4 anchor boxes of following width and height. The width and heights are rescaled in the grid cell scale (Assuming that the number of grid size is 13 by 13.) See [Part 2 Object Detection using YOLOv2 on Pascal VOC2012 - input and output encoding](https://fairyonice.github.io/Part%202_Object_Detection_with_Yolo_using_VOC_2014_data_input_and_output_encoding.html) to learn how I rescal the anchor box shapes into the grid cell scale.
Here I choose 4 anchor boxes. With 13 by 13 grids, every frame gets 4 x 13 x 13 = 676 bouding box predictions.
```
ANCHORS = np.array([1.07709888, 1.78171903, # anchor box 1, width , height
2.71054693, 5.12469308, # anchor box 2, width, height
10.47181473, 10.09646365, # anchor box 3, width, height
5.48531347, 8.11011331]) # anchor box 4, width, height
```
### Define Label vector containing 20 object classe names.
```
LABELS = ['aeroplane', 'bicycle', 'bird', 'boat', 'bottle',
'bus', 'car', 'cat', 'chair', 'cow',
'diningtable','dog', 'horse', 'motorbike', 'person',
'pottedplant','sheep', 'sofa', 'train', 'tvmonitor']
```
## YOLOv2 Model Architecture
While YOLO's input and output encodings are complex, and loss function of YOLO is quite complex (which will be discussed very soon), the model architecture is simple.
It repeatedly stacks Convolusion + Batch Normalization + Leaky Relu layers until the image shape reduces to the grid cell size.
Here is the model defenition, extracted from [experiencor/keras-yolo2](https://github.com/experiencor/keras-yolo2).
```
from keras.models import Sequential, Model
from keras.layers import Reshape, Activation, Conv2D, Input, MaxPooling2D, BatchNormalization, Flatten, Dense, Lambda
from keras.layers.advanced_activations import LeakyReLU
from keras.callbacks import EarlyStopping, ModelCheckpoint, TensorBoard
from keras.optimizers import SGD, Adam, RMSprop
from keras.layers.merge import concatenate
import keras.backend as K
import tensorflow as tf
# the function to implement the orgnization layer (thanks to github.com/allanzelener/YAD2K)
def space_to_depth_x2(x):
return tf.space_to_depth(x, block_size=2)
def ConvBatchLReLu(x,filters,kernel_size,index,trainable):
# when strides = None, strides = pool_size.
x = Conv2D(filters, kernel_size, strides=(1,1),
padding='same', name='conv_{}'.format(index),
use_bias=False, trainable=trainable)(x)
x = BatchNormalization(name='norm_{}'.format(index), trainable=trainable)(x)
x = LeakyReLU(alpha=0.1)(x)
return(x)
def ConvBatchLReLu_loop(x,index,convstack,trainable):
for para in convstack:
x = ConvBatchLReLu(x,para["filters"],para["kernel_size"],index,trainable)
index += 1
return(x)
def define_YOLOv2(IMAGE_H,IMAGE_W,GRID_H,GRID_W,TRUE_BOX_BUFFER,BOX,CLASS, trainable=False):
convstack3to5 = [{"filters":128, "kernel_size":(3,3)}, # 3
{"filters":64, "kernel_size":(1,1)}, # 4
{"filters":128, "kernel_size":(3,3)}] # 5
convstack6to8 = [{"filters":256, "kernel_size":(3,3)}, # 6
{"filters":128, "kernel_size":(1,1)}, # 7
{"filters":256, "kernel_size":(3,3)}] # 8
convstack9to13 = [{"filters":512, "kernel_size":(3,3)}, # 9
{"filters":256, "kernel_size":(1,1)}, # 10
{"filters":512, "kernel_size":(3,3)}, # 11
{"filters":256, "kernel_size":(1,1)}, # 12
{"filters":512, "kernel_size":(3,3)}] # 13
convstack14to20 = [{"filters":1024, "kernel_size":(3,3)}, # 14
{"filters":512, "kernel_size":(1,1)}, # 15
{"filters":1024, "kernel_size":(3,3)}, # 16
{"filters":512, "kernel_size":(1,1)}, # 17
{"filters":1024, "kernel_size":(3,3)}, # 18
{"filters":1024, "kernel_size":(3,3)}, # 19
{"filters":1024, "kernel_size":(3,3)}] # 20
input_image = Input(shape=(IMAGE_H, IMAGE_W, 3),name="input_image")
true_boxes = Input(shape=(1, 1, 1, TRUE_BOX_BUFFER , 4),name="input_hack")
# Layer 1
x = ConvBatchLReLu(input_image,filters=32,kernel_size=(3,3),index=1,trainable=trainable)
x = MaxPooling2D(pool_size=(2, 2),name="maxpool1_416to208")(x)
# Layer 2
x = ConvBatchLReLu(x,filters=64,kernel_size=(3,3),index=2,trainable=trainable)
x = MaxPooling2D(pool_size=(2, 2),name="maxpool1_208to104")(x)
# Layer 3 - 5
x = ConvBatchLReLu_loop(x,3,convstack3to5,trainable)
x = MaxPooling2D(pool_size=(2, 2),name="maxpool1_104to52")(x)
# Layer 6 - 8
x = ConvBatchLReLu_loop(x,6,convstack6to8,trainable)
x = MaxPooling2D(pool_size=(2, 2),name="maxpool1_52to26")(x)
# Layer 9 - 13
x = ConvBatchLReLu_loop(x,9,convstack9to13,trainable)
skip_connection = x
x = MaxPooling2D(pool_size=(2, 2),name="maxpool1_26to13")(x)
# Layer 14 - 20
x = ConvBatchLReLu_loop(x,14,convstack14to20,trainable)
# Layer 21
skip_connection = ConvBatchLReLu(skip_connection,filters=64,
kernel_size=(1,1),index=21,trainable=trainable)
skip_connection = Lambda(space_to_depth_x2)(skip_connection)
x = concatenate([skip_connection, x])
# Layer 22
x = ConvBatchLReLu(x,filters=1024,kernel_size=(3,3),index=22,trainable=trainable)
# Layer 23
x = Conv2D(BOX * (4 + 1 + CLASS), (1,1), strides=(1,1), padding='same', name='conv_23')(x)
output = Reshape((GRID_H, GRID_W, BOX, 4 + 1 + CLASS),name="final_output")(x)
# small hack to allow true_boxes to be registered when Keras build the model
# for more information: https://github.com/fchollet/keras/issues/2790
output = Lambda(lambda args: args[0],name="hack_layer")([output, true_boxes])
model = Model([input_image, true_boxes], output)
return(model, true_boxes)
IMAGE_H, IMAGE_W = 416, 416
GRID_H, GRID_W = 13 , 13
TRUE_BOX_BUFFER = 50
BOX = int(len(ANCHORS)/2)
CLASS = len(LABELS)
## true_boxes is the tensor that takes "b_batch"
model, true_boxes = define_YOLOv2(IMAGE_H,IMAGE_W,GRID_H,GRID_W,TRUE_BOX_BUFFER,BOX,CLASS,
trainable=False)
model.summary()
```
## Load pre-trained YOLOv2 weights
Following the instruction at [YOLO: Real-Time Object Detection](https://pjreddie.com/darknet/yolov2/), we download the pre-trained weights using wget:
<code>
wget https://pjreddie.com/media/files/yolov2.weights
</code>
The weights are saved at:
```
path_to_weight = "./yolov2.weights"
```
The following codes are extracted from [keras-yolo2/Yolo Step-by-Step.ipynb](https://github.com/experiencor/keras-yolo2/blob/master/Yolo%20Step-by-Step.ipynb)
```
class WeightReader:
# code from https://github.com/experiencor/keras-yolo2/blob/master/Yolo%20Step-by-Step.ipynb
def __init__(self, weight_file):
self.offset = 4
self.all_weights = np.fromfile(weight_file, dtype='float32')
def read_bytes(self, size):
self.offset = self.offset + size
return self.all_weights[self.offset-size:self.offset]
def reset(self):
self.offset = 4
weight_reader = WeightReader(path_to_weight)
print("all_weights.shape = {}".format(weight_reader.all_weights.shape))
```
Assign pre-trained weights to the following layers:
<code> conv_i</code>, <code>norm_i</code>, <code> i = 1, 2,..., 22</code>.
These layers do not depend on the number of object classes or the number of anchor boxes.
```
def set_pretrained_weight(model,nb_conv, path_to_weight):
weight_reader = WeightReader(path_to_weight)
weight_reader.reset()
for i in range(1, nb_conv+1):
conv_layer = model.get_layer('conv_' + str(i)) ## convolusional layer
if i < nb_conv:
norm_layer = model.get_layer('norm_' + str(i)) ## batch normalization layer
size = np.prod(norm_layer.get_weights()[0].shape)
beta = weight_reader.read_bytes(size)
gamma = weight_reader.read_bytes(size)
mean = weight_reader.read_bytes(size)
var = weight_reader.read_bytes(size)
weights = norm_layer.set_weights([gamma, beta, mean, var])
if len(conv_layer.get_weights()) > 1: ## with bias
bias = weight_reader.read_bytes(np.prod(conv_layer.get_weights()[1].shape))
kernel = weight_reader.read_bytes(np.prod(conv_layer.get_weights()[0].shape))
kernel = kernel.reshape(list(reversed(conv_layer.get_weights()[0].shape)))
kernel = kernel.transpose([2,3,1,0])
conv_layer.set_weights([kernel, bias])
else: ## without bias
kernel = weight_reader.read_bytes(np.prod(conv_layer.get_weights()[0].shape))
kernel = kernel.reshape(list(reversed(conv_layer.get_weights()[0].shape)))
kernel = kernel.transpose([2,3,1,0])
conv_layer.set_weights([kernel])
return(model)
nb_conv = 22
model = set_pretrained_weight(model,nb_conv, path_to_weight)
```
## Initialize the 23rd layer
```
def initialize_weight(layer,sd):
weights = layer.get_weights()
new_kernel = np.random.normal(size=weights[0].shape, scale=sd)
new_bias = np.random.normal(size=weights[1].shape, scale=sd)
layer.set_weights([new_kernel, new_bias])
layer = model.layers[-4] # the last convolutional layer
initialize_weight(layer,sd=GRID_H*GRID_W)
```
So that's how we define the YOLOv2 model!
The next blog will discuss the loss function of this model which will be used to train the parameters.
[FairyOnIce/ObjectDetectionYolo](https://github.com/FairyOnIce/ObjectDetectionYolo)
contains this ipython notebook and all the functions that I defined in this notebook.
| github_jupyter |
```
%matplotlib inline
import numpy as np
from numpy.fft import fft2, ifft2, fftshift, ifftshift
from scipy.stats import multivariate_normal
import matplotlib.pyplot as plt
import menpo.io as mio
from menpo.image import Image
from menpo.feature import hog, no_op
from menpo.shape import PointCloud
from menpo.visualize import visualize_images
from templatetracker.correlationfilter.correlationfilter import (
learn_mosse, learn_mccf, learn_deep_cf)
from templatetracker.correlationfilter.utils import (
build_grid, normalizenorm_vec, fast2dconv, crop)
def greyscale(i):
return i.as_greyscale('average')
def greyscale_hog(i):
return hog(greyscale(i))
def combine(i):
return Image(np.concatenate((i.pixels, greyscale(i).pixels, greyscale_hog(i).pixels)))
```
# Kernelized Correlation Filters
## Load and manipulate data
Load landmarked facial images.
```
images = []
for i in mio.import_images('/Users/joan/PhD/DataBases/faces/lfpw/trainset/*', verbose=True,
max_images=300):
i.crop_to_landmarks_proportion_inplace(0.5)
i = i.rescale_landmarks_to_diagonal_range(100)
images.append(i)
visualize_images(images)
```
Extract 31 x 31 patches around landmark number 45 (the corner of the left eye) from the previous images. Note that any other landmark could be chosen.
```
patch_shape = np.asarray((101, 101))
lm_number = 45
features = greyscale # no_op, gresycale, greyscale_hog
image_patches = []
pixel_patches = []
for i in images:
image_patches.append(i.extract_patches_around_landmarks(patch_size=patch_shape)[lm_number])
feature_patches = features(image_patches[-1])
pixel_patches.append(feature_patches.pixels)
visualize_images(image_patches)
```
Store patches as numpy array.
```
X = np.asarray(pixel_patches)
```
These are the patches that we will used in order to define and test our Kernelized Correlation Filters.
Define the desired response for each patch. Note that, because all patches are centred about the same landmark they share the same desired response, i.e. a 2 dimensional Gaussian response centred at the middle of the patch.
```
cov = 3
# define Gaussian response
mvn = multivariate_normal(mean=np.zeros(2), cov=cov)
grid = build_grid((31, 31))
y = mvn.pdf(grid)[None]
plt.title('Desired response')
plt.imshow(y[0])
```
## Learn Correlation Filter (CF)
At this point we will use the first image patch as the template from which to learn a CF. Note that we could have chosen any other image patch.
```
# img_number = 0
# x = X[img_number]
```
Apart from the kernel correlation specific parameters, we need to make some choices regarding the overall learning procedure.
```
# whether to normalize the image
normalize = True
# wheter to mask the images with a cosine mask
mask = True
# regularization parameter
l = 0.1
# type of filter
filter_type = 'deep_mosse'
# boundary padding
boundary = 'symmetric'
c1 = np.hanning(patch_shape[0])
c2 = np.hanning(patch_shape[1])
cosine_mask = c1[..., None].dot(c2[None, ...]) if mask else None
```
We are now ready to learn a CF for the first image patch.
```
X_ = np.empty_like(X)
for j, x in enumerate(X):
x_ = normalizenorm_vec(x) if normalize else x
x_ = cosine_mask * x_ if mask else x_
X_[j] = x_
if filter_type is 'mosse':
cf, _, _ = learn_mosse(X_, y, l=l, boundary=boundary)
elif filter_type is 'mccf':
cf, _, _ = learn_mccf(X_, y, l=l, boundary=boundary)
elif filter_type is 'deep_mosse':
cf, _, _ = learn_deep_cf(X_, y, learn_cf=learn_mosse, n_levels=1, l=l, boundary=boundary)
elif filter_type is 'deep_mccf':
cf, _, _ = learn_deep_cf(X_, y, learn_cf=learn_mccf, n_levels=3, l=l, boundary=boundary)
cf = cf
# only the up to the first 5 channels are shown
n_channels = np.minimum(5, cf.shape[0])
fig_size = (3*n_channels, 3*n_channels)
fig = plt.figure()
fig.set_size_inches(fig_size)
for j, c in enumerate(cf[:n_channels]):
plt.subplot(1, n_channels, j+1)
plt.title('CF in spatial domain')
plt.imshow(cf[j])
fig = plt.figure()
fig.set_size_inches(fig_size)
for j, c in enumerate(cf[:n_channels]):
plt.subplot(1, n_channels, j+1)
plt.title('CF in frequency domain')
plt.imshow(np.abs(fftshift(fft2(cf[j]))))
```
## Test KCF
In order to test the correctness of the learned KCF we will extract 61 x 61 patches centred around landmark number 31, i.e the right corner of the nose. Note that we will now expect to get responses whith peaks shifted towards the right hence, correctly localizing the eye corner for which the KCF was learned.
```
lm_test = 42
patch_shape2 =(81, 81)
image_prime_patches = []
pixel_prime_patches = []
for i in images:
image_prime_patches.append(i.extract_patches_around_landmarks(patch_size=patch_shape2)[lm_test])
feature_prime_patches = features(image_prime_patches[-1])
pixel_prime_patches.append(feature_prime_patches.pixels)
visualize_images(image_prime_patches)
```
Store patches as numpy array.
```
X_prime = np.asarray(pixel_prime_patches)
rs = []
for z in X_prime:
z_ = normalizenorm_vec(z) if normalize else z
cf_ = normalizenorm_vec(cf) if normalize else cf
# compute correlation response
r = np.sum(fast2dconv(z_, cf, boundary=boundary), axis=0)[None]
rs.append(r)
# only up to the first 5 images are shown
n_images = np.minimum(5, len(X_prime))
fig_size = (3*n_images, 3*n_images)
fig = plt.figure()
fig.set_size_inches(fig_size)
for j, r in enumerate(rs[:n_images]):
plt.subplot(1, n_images, j+1)
plt.title('response')
plt.imshow(r[0])
fig = plt.figure()
fig.set_size_inches(fig_size)
for j, (r, i) in enumerate(zip(rs[:n_images], image_prime_patches[:n_images])):
plt.subplot(1, n_images, j+1)
plt.title('original image')
peak = np.asarray(np.unravel_index(r.argmax(), r.shape))[1:]
i.landmarks['peak'] = PointCloud(peak[None, ...])
i.view_landmarks(marker_face_colour='r', figure_size=fig_size)
```
| github_jupyter |
# Value in equities
#### by Gustavo Soares
In this notebook you will apply a few things you learned in our Python lecture [FinanceHub's Python lectures](https://github.com/Finance-Hub/FinanceHubMaterials/tree/master/Python%20Lectures):
* You will use and manipulate different kinds of variables in Python such as text variables, booleans, date variables, floats, dictionaries, lists, list comprehensions, etc.;
* We will also use `Pandas.DataFrame` objects and methods which are very useful in manipulating financial time series;
* You will use if statements and loops, and;
* You will use [FinanceHub's Bloomberg tools](https://github.com/Finance-Hub/FinanceHub/tree/master/bloomberg) for fetching data from a Bloomberg terminal. If you are using this notebook within BQNT, you may want to use BQL for getting the data.
## Basic imports
You can use BQuant and BQL to download data like we did in our [notebook on K-means clustering](https://github.com/Finance-Hub/FinanceHubMaterials/blob/master/Quantitative%20Finance%20Lectures/kmeans_clusters_factors.ipynb). Here, we will use the standard terminal [FinanceHub's Python API](https://github.com/Finance-Hub/FinanceHub/tree/master/bloomberg) to grab the data but I encourage you to try it all out with the BQuant + BQL combo!
```
from scipy import stats
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
from bloomberg import BBG
bbg = BBG() # because BBG is a class, we need to create an instance of the BBG class wihtin this notebook, here deonted by bbg
```
## Value metrics: cross-sectional implementation
Value investing is an investment strategy that involves picking stocks that appear to be trading for less than their intrinsic or book value. Numerous studies document that value stocks, loosely defined as stocks with low prices relative to earnings, dividends, or book value, generate higher long-run returns than growth stocks, or stocks with high prices relative to these measures of fundamental value. The chapter on *Value Premium* of [Empirical Asset Pricing: The Cross Section of Stock Returns](https://www.amazon.com/Empirical-Asset-Pricing-Probability-Statistics/dp/1118095049) has a great summary of the academic finding regarding value strategies in equities. Here, we will focus on creating a value metric for a stock or set of stocks.
Going back to 1992 [Fama and French Three-Factor Model](https://www.investopedia.com/terms/f/famaandfrenchthreefactormodel.asp) cross-sectional value strategies are considered one of the risk factors to be added to the CAPM market risk factor. In our own [notebook on K-means clustering](https://github.com/Finance-Hub/FinanceHubMaterials/blob/master/Quantitative%20Finance%20Lectures/kmeans_clusters_factors.ipynb) we construct a value metric based on the Book-to-Price ratio for comparing stocks cross-sectionally. Here, we also discuss defining a value metric in a cross-sectional sense a bit more.
### Book-to-Price
The book-to-price or book-to-market ratio, is defined as the book value of a firm’s common equity (BE) divided by the market value of the firm’s equity (ME), or where the book value comes from the firm’s balance sheet and the market value is identical to the market capitalization of the firm. The price-to-book ratio is a close cousin of Tobin's Q, another statistic used by investors to gauge value in stocks. Tobin's Q is defined as the ratio of the market cap to the replacement cost of assets. A Q that is greater than 1 means that an investor is better off replicating the assets of a company rather than buying it. It indicates that a company is expensive. Similarly, high price-to-book ratios are a presumption of expensiveness.
The code below downloads the stocks belonging to IBOVESP, the Brazilian stock index and grabs their price to book ratio:
```
ref_date = '2019-12-04'
w_df = bbg.fetch_index_weights(index_name='IBOV Index', ref_date=pd.to_datetime(ref_date))
list_of_tickers = [x + ' Equity' for x in w_df.index]
w_df.index = list_of_tickers
sectors = bbg.fetch_contract_parameter(securities=list_of_tickers, field='GICS_SECTOR_NAME')
price_to_book = bbg.fetch_series(securities=list_of_tickers,
fields='PX_TO_BOOK_RATIO',
startdate=ref_date,
enddate=ref_date)
df = pd.concat([w_df.iloc[:,0].to_frame('weights'),
price_to_book.iloc[0].to_frame('price_to_book'),
sectors.iloc[:,0].to_frame('sector')],axis=1,sort=True)
df.sort_values(by='weights',ascending=False).head()
```
One way of computing the value metric for a value strategy is comparing it with its peers. Here, the Global Industry Classification Standard (GICS) sectors to define the peer group but you can think of [more sophisticated ways](https://www.winton.com/research/systematic-methods-for-classifying-equities) of doing that if you want.
How do we compare the value metric with the peer-group? A traditional way of doing that is using z-scores. Even when using z-scores, it's common for people to [winsorize](https://en.wikipedia.org/wiki/Winsorizing) z-scores. Here, we do that at the 0.1 limits
```
df['book_to_price_scores'] = np.nan
for s in set(df['sector']):
signals = df[df['sector']==s]['price_to_book'].dropna()**-1 # the inverse because we want B/P and not P/B
if len(signals)>=5:
df.loc[signals.index,'book_to_price_scores'] = stats.mstats.winsorize(stats.zscore(signals.values), limits=.1)
df = df.dropna()
df.head(10)
```
### Dividend yields
[Baz, Granger, Harvey, Le Roux, and Rattray (2015)](https://ssrn.com/abstract=2695101) discusses how dividend yields can also be used as a measure of value for broad equity markets. High dividend yields may indicate that stocks are cheap against bonds for example. Companies pay dividends to their shareholders but they, from time to time, also buy back stock from them. When a company buys back shares from its shareholders, it reduces the number of shares outstanding, which increases per share measures of profitability like EPS, Cash Flow Per Share, ROE. These improved metrics generally drives the share price higher over time. Hence to account for these changes, it is common to updated the yield measure to account for both dividends and buybacks. Here, to make things simple, we do not do that.
```
dvd_yield = bbg.fetch_series(securities=list(df.index),
fields='EQY_DVD_YLD_IND',
startdate=ref_date,
enddate=ref_date)
df = pd.concat([df,dvd_yield.iloc[0].to_frame('dvd_yield')],axis=1,sort=True)
for s in set(df['sector']):
signals = df[df['sector']==s]['dvd_yield'].dropna()
if len(signals)>=5:
df.loc[signals.index,'dvd_yield_scores'] = stats.mstats.winsorize(stats.zscore(signals.values), limits=.1)
df = df.dropna()
```
### Earnings yields
The earnings yield refers to the earnings per share for the most recent 12-month period divided by the current market price per share. The earnings yield (which is the inverse of the P/E ratio) shows the percentage of how much a company earned per share.
```
earnings_yield = bbg.fetch_series(securities=list(df.index),
fields='EARN_YLD',
startdate=ref_date,
enddate=ref_date)
df = pd.concat([df,earnings_yield.iloc[0].to_frame('earnings_yield')],axis=1,sort=True)
for s in set(df['sector']):
signals = df[df['sector']==s]['earnings_yield'].dropna()
if len(signals)>=5:
df.loc[signals.index,'earnings_yield_scores'] = stats.mstats.winsorize(stats.zscore(signals.values), limits=.1)
df = df.dropna()
```
### Cash flow yields
The cash flow yield is calculated by taking the free cash flow per share divided by the current share price. Free cash flow yield is similar in nature to the earnings yield metric above, which is usually meant to measure GAAP (generally accepted accounting principles) earnings per share divided by share price.
```
cf_yield = bbg.fetch_series(securities=list(df.index),
fields='FREE_CASH_FLOW_YIELD',
startdate=ref_date,
enddate=ref_date)
df = pd.concat([df,cf_yield.iloc[0].to_frame('cf_yield')],axis=1,sort=True)
for s in set(df['sector']):
signals = df[df['sector']==s]['cf_yield'].dropna()
if len(signals)>=5:
df.loc[signals.index,'cf_yield_scores'] = stats.mstats.winsorize(stats.zscore(signals.values), limits=.1)
df = df.dropna()
```
## Combining metrics
In cross-sectional as well as in time-series implementations, typically, combining different metrics has been found in empirical studies to produce better forecasts on average than methods based on the ex ante best individual metric. Even simple combinations that ignore correlations between metrics often dominate more refined combination schemes aimed at estimating the theoretically optimal combination weights. Although the reasons for the success of simple combination schemes are poorly understood (model misspecification, non-stationarities, regime switches, etc.). This [book chapter](https://www.sciencedirect.com/science/article/abs/pii/S1574070605010049) is a great reference for forecast combinations if you want to learn more about it. Here, we combine the four cross-sectional value metrics discussed above into a signle value metric using a simple average:
```
zscores_df = df[['book_to_price_scores','dvd_yield_scores','earnings_yield_scores','cf_yield_scores']].astype(float)
zscores_df['value_metric'] = zscores_df.mean(axis=1)
zscores_df['value_metric'].sort_values().plot(kind='bar',color='b',figsize=(15,10))
plt.title('Combined value metric',fontsize=18)
plt.xlabel('cheap -> expensive',fontsize=18)
plt.show()
```
Using this metric, we can think of a strategy that goes long the cheap stocks (say the 20% cheapest ones) and short the expensive stocks say the 20% most expensive ones)
```
cheap_list = zscores_df[zscores_df['value_metric']<=zscores_df['value_metric'].quantile(q=0.20)].index
print('Go long:')
print(list(cheap_list))
expensive_list = zscores_df[zscores_df['value_metric']>=zscores_df['value_metric'].quantile(q=0.80)].index
print('Go short:')
print(list(expensive_list))
```
## Value metrics: time-series implementation
[Baz, Granger, Harvey, Le Roux, and Rattray (2015)](https://ssrn.com/abstract=2695101) discusses in detail the differences between cross-sectional strategies and time series strategies for three different factors, including value. It is really important to understand the difference between the two implementations. The key difference is that now, we will look at value of a particular stock but not compare it with its peers but rather with its own historical measures. To illustrate, let's pick one particular stock, say ITUB4.
Let's start by grabbing the data we are going to need from Bloomberg:
```
ts_data = bbg.fetch_series(securities='ITUB4 BS Equity',
fields=['PX_LAST','PX_TO_BOOK_RATIO','EQY_DVD_YLD_IND','EARN_YLD','FREE_CASH_FLOW_YIELD'],
startdate='2010-01-01',
enddate=ref_date)
ts_data = ts_data.dropna().astype(float)
ts_data.tail()
```
Note that the quality of the data is not the best. This is often a serious problem when constructing quant strategies. Most of the time and effort is dedicated to cleaning data:
```
fig, axs = plt.subplots(nrows=2, ncols=2,figsize=(15,10))
axs[0, 0].plot(ts_data.iloc[:,0].dropna().index, ts_data.iloc[:,0].dropna().values)
axs[0, 0].set_title(ts_data.columns[1])
axs[0, 1].plot(ts_data.iloc[:,1].dropna().index, ts_data.iloc[:,1].dropna().values)
axs[0, 1].set_title(ts_data.columns[2])
axs[1, 0].plot(ts_data.iloc[:,2].dropna().index, ts_data.iloc[:,2].dropna().values)
axs[1, 0].set_title(ts_data.columns[3])
axs[1, 1].plot(ts_data.iloc[:,3].dropna().index, ts_data.iloc[:,3].dropna().values)
axs[1, 1].set_title(ts_data.columns[4])
plt.show()
```
Here, we are not going to directly deal with the bad data. We will just trucate at one year:
```
ts = ts_data.iloc[-252:,:]
ts.iloc[:,1] = ts.iloc[:,1]**-1 # the inverse because we want B/P and not P/B
ts = ts.rename(columns={"PX_TO_BOOK_RATIO": "BOOK_TO_PX_RATIO"})
```
Let's now calcualte time series z-scores. Note that we need to do that by looking back in the stocks past historical data and calculating z-scores with the information that was available at that point in time. The method `rolling` from `Pandas` is a great way to do that:
```
for i in range(1,5):
fig1, ax1 = plt.subplots()
ax1.set_title(ts.columns[i])
ax1.boxplot(ts.iloc[:,i])
ax1.annotate('current value', xy=(1, ts.iloc[-1,i]), xycoords='data',
xytext=(1, 0.5), textcoords='axes fraction',
arrowprops=dict(facecolor='black'),
horizontalalignment='right', verticalalignment='top',
)
plt.show()
zscores = (ts.iloc[-1,1:]-ts.iloc[:,1:].mean())/ts.iloc[:,1:].std()
zscores['mean'] = zscores.mean()
zscores.plot(kind='bar')
```
So, relative to its own history, it seems like this stock is expensive!
## Exercises
#### Beginners
Try to calculate a cross-sectional or time-series value metric that takes into account the fact that different metrics may have different volatilities, they may be correlated with each other, they may have different forecasting errors and different halflives.
#### Intermediate
Calculate a time series value strategies as in [Baz, Granger, Harvey, Le Roux, and Rattray (2015)](https://www.cmegroup.com/education/files/dissecting-investment-strategies-in-the-cross-section-and-time-series.pdf).
#### Intermediate
Create a class to calculate time-series value mterics for one stock.
#### Advanced
Create a class to calculate both cross-sectional and time-series value for a set of stock.
#### Jedi Master
Carry out the Advanced task above and make a contribution to [FinanceHub's signals class](https://github.com/Finance-Hub/FinanceHub/tree/master/signals).
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Gradient-Boosting-Machine-(GBM)" data-toc-modified-id="Gradient-Boosting-Machine-(GBM)-1"><span class="toc-item-num">1 </span>Gradient Boosting Machine (GBM)</a></span><ul class="toc-item"><li><span><a href="#Implementation" data-toc-modified-id="Implementation-1.1"><span class="toc-item-num">1.1 </span>Implementation</a></span></li><li><span><a href="#Classification" data-toc-modified-id="Classification-1.2"><span class="toc-item-num">1.2 </span>Classification</a></span><ul class="toc-item"><li><span><a href="#Softmax" data-toc-modified-id="Softmax-1.2.1"><span class="toc-item-num">1.2.1 </span>Softmax</a></span></li></ul></li><li><span><a href="#Implementation" data-toc-modified-id="Implementation-1.3"><span class="toc-item-num">1.3 </span>Implementation</a></span></li><li><span><a href="#Understanding-Model-Complexity" data-toc-modified-id="Understanding-Model-Complexity-1.4"><span class="toc-item-num">1.4 </span>Understanding Model Complexity</a></span></li></ul></li><li><span><a href="#Reference" data-toc-modified-id="Reference-2"><span class="toc-item-num">2 </span>Reference</a></span></li></ul></div>
```
# code for loading the format for the notebook
import os
# path : store the current path to convert back to it later
path = os.getcwd()
os.chdir(os.path.join('..', '..', 'notebook_format'))
from formats import load_style
load_style(css_style = 'custom2.css', plot_style = False)
os.chdir(path)
# 1. magic for inline plot
# 2. magic to print version
# 3. magic so that the notebook will reload external python modules
# 4. magic to enable retina (high resolution) plots
# https://gist.github.com/minrk/3301035
%matplotlib inline
%load_ext watermark
%load_ext autoreload
%autoreload 2
%config InlineBackend.figure_format = 'retina'
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.metrics import mean_squared_error
from sklearn.tree import DecisionTreeRegressor
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.ensemble import RandomForestRegressor, GradientBoostingRegressor
%watermark -d -t -v -p numpy,pandas,matplotlib,sklearn
```
# Gradient Boosting Machine (GBM)
Just like [Random Forest and Extra Trees](http://nbviewer.jupyter.org/github/ethen8181/machine-learning/blob/master/trees/random_forest.ipynb), Gradient Boosting Machine is also a type of Ensemble Tree method, the only difference is it is stemmed from the the boosting framework. The idea of boosting is to add a weak classifier to the ensemble at a time, and this newly added weak classifier is trained to improve upon the already trained ensemble. Meaning it will pay higher attention on examples which are misclassified or have higher errors and focus on mitigating those errors. Boosting is a general framework can be applied to any sort of weak learner, although Decision Tree models is by far the commonly used due to the fact that they have the flexibility to be weak learners by simply restricting their depth and they are quite fast to train.
Suppose we are given some dataset $(x_1, y_1), (x_2, y_2), ...,(x_n, y_n)$, and the task is to fit a model $F(x)$ to minimize square loss. After training the model, we discovered the model is good but not perfect.
There are some mistakes: $F(x_1) = 0.8$, while $y_1 = 0.9$, and $F(x_2) = 1.4$ while $y_2 = 1.3$ .... Now the question is, how can we improve this model without changing anything from $F(x)$?
How about we simply add an additional model (e.g. regression tree) $h$ to the already existing $F$, so the new prediction becomes $F(x) + h(x)$. In other words, we wish to improve upon the existing model so that $F(x_1) + h(x_1) = y_1, F(x_2) + h(x_2) = y_2 ...$ or equivalent we wish to find a new model $h$ such that $h(x_1) = y_1 - F(x_1), h(x_2) = y_2 - F(x_2) ...$. The idea is all well and good, but the bad news is probably no model $h$ (e.g. regression tree) will be able to do this perfectly. Fortunately, the good news is, some $h$ might be able to do this approximately.
The idea is, we fit the model $h$ to the data using $y_1 - F(x_1), y_2 - F(x_2)$ as the response variable. And the intuition for this is: the $y_i - F(x_i)$s are the residuals. These are the areas that the existing
model $F$ cannot do well, so now the role of $h$ is to compensate the shortcoming of existing model $F$. And if the model after adding the new model $h$, $F + h$ is still unsatisfactory, we will just add another new one.
To make sure we're actually learning the residuals, we'll employ the idea of gradient descent. Say our goal is to minimize $J$, an overall loss function additively calculated from all observations with regard to $F$, a classifier with some parameters. More formally, we're given the formula:
$$J(y, F) = \sum_i^n L\big(y_i, F(x_i)\big)$$
Where:
- $L$ is a cost/loss function comparing the response variable's value and the prediction of the model for each observation
Instead of trying to solve it directly, gradient descent is an iterative technique that allows us to approach the solution of an optimization problem. At each step of the algorithm, it will perform the following operations:
$$F_b(x_i) = F_{b-1}(x_i) - \eta \times \nabla L\big(y_i, F(x_i)\big)$$
Where:
- $F_b$ is the version of classifier at step/iteration $b$
- $\eta$ is the learning rate which controls the size of the learning process
- $\nabla$ is the gradient i.e. the first order partial derivative of the cost function with respect to the classifier
- The formula above actually refers to stochastic gradient descent as we are only computing the function for a single observation, $x_i$
For example, say we're given, sum of squares errors, a well-known quality indicator for regression model as our loss function. So now our loss function $L\big(y_i, F(x_i)\big)$ is defined as: $\frac{1}{2} \big( y_i - F(x_i) \big)^2$ (the 1/2 is simply to make the notation cleaner later). Taking the gradient of this loss function we get:
$$\frac{ \partial L\big(y_i, F(x_i)\big) }{ \partial F(x_i) } = \frac{ \partial \frac{1}{2} \big( y_i - F(x_i) \big)^2 }{ \partial F(x_i) } = F(x_i) - y_i$$
Tying this back to our original problem, we wish to update our function $F$ at iteration $b$ with a new model $h$:
\begin{align}
F_b(x_i) &= F_{b-1}(x_i) + h(x_i) \nonumber \\
&= F_{b-1}(x_i) + y_i - F_{b-1}(x_i) \nonumber \\
&= F_{b-1}(x_i) - 1 \times \frac{ \partial L\big(y_i, F_{b-1}(x_i)\big) }{ \partial F_{b-1}(x_i) }
\nonumber \\
\end{align}
As we can see, the formula above is 99% the same as as the gradient descent formula, $F_b(x_i) = F_{b-1}(x_i) - \eta \times \nabla L\big(y_i, F(x_i)\big)$. The only difference is that the learning rate $\eta$ is 1. Thus, we now have an iterative process constructing the additive model that minimizes our loss function (residuals).
In practice though, Gradient Boosting Machine is more prone to overfitting, since the week learner is tasked with optimally fitting the gradient. This means that boosting will select the optimal learner at each stage of the algorithm, although this strategy generates an optimal solution at the current stage, it has the drawbacks of not finding the optimal global model as well as overfitting the training data. A remedy for greediness is to constrain the learning process by setting the learning rate $\eta$ (also known as shrinkage). In the above algorithm, instead of directly adding the predicted value for a sample to next iteration's predicted value, so that only a fraction of the current predicted value is added to the previous iteration's predicted value. This parameter can take values between 0 and 1 and becomes another tuning parameter for the model. Small values of the learning parameter such as 0.1 tends to work better, but the value of the parameter is inversely proportional to the computation time required to find an optimal model, because more iterations is required.
To sum it all up, the process of training a GBM for regression is:
1. Initialize a predicted value for each observation (e.g. the original response or the average response or a value that minimizes the loss function). This will be our initial "residuals", $r$. It can be called the residuals because we're dealing with a regression task, but this quantity is more often referred to as the negative gradient, this terminology makes the $- \nabla \times L\big(y_i, F(x_i) \big)$ part generalizes to any loss function we might wish to employ. In short, GBM is fitting to the gradient of the loss function
2. For step = 1 to $B$ (number of iterations that we specify) do:
- Fit a regression tree $F_b$ to the training data $(X, r)$, where we use the residuals as the response variable
- Update model $F$ by adding a shrunken version of the newly fitted regression tree. Translating it to code, this means we append the new tree to the array of trees we've already stored:
$F(X) = F(X) + \eta F_{b}(X)$
- Update each observation's residual by adding the predicted value to it:
$r_{b + 1} = r_b - \eta F_b(X)$
3. In the end, our final output boosted model becomes $F(x) = \sum_{b = 1}^B \eta F_b(x)$, where we sum the values that each individual tree gives (times the learning rate)
To hit the notion home, let's conside an example using made up numbers. Suppose we have 5 observations, with responses 10, 20, 30, 40, 50. The first tree is built and gives predictions of 12, 18, 27, 39, 54 (these predictions are made up numbers). If our learning rate $\eta$ = 0.1, all trees will have their predictions scaled down by $\eta$, so the first tree will instead "predict" 1.2, 1.8, 2.7, 3.9, 5.4. The response variable passed to the next tree will then have values 8.8, 18.2, 27.3, 36.1, 44.6 (the difference between the prediction that was scaled down by the prediction and the true response). The second round then uses these response values to build another tree - and again the predictions are scaled down by the learning rate $\eta$. So tree 2 predicts say, 7, 18, 25, 40, 40, which, once scaled, become 0.7, 1.8, 2.5, 4.0, 4.0. As before, the third tree will be passed the difference between these values and the previous tree's response variable (so 8.1, 16.4, 24.8, 32.1. 40.6). And we keep iterating this process until we finished training all the trees (a parameter that we specify), in the end, the sum of the predictions from all trees will give the final prediction.
## Implementation
Here, we will use the [Wine Quality Data Set](https://archive.ics.uci.edu/ml/datasets/Wine+Quality) to test our implementation. This [link](https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-white.csv) should download the .csv file. The task is to predict the quality of the wine (a scale of 1 ~ 10) given some of its features.
```
# read in the data and shuffle the row order for model stability
np.random.seed(4321)
wine_path = os.path.join('..', 'winequality-white.csv')
wine = pd.read_csv(wine_path, sep = ';')
wine = wine.sample(frac = 1)
# train/test split the features and response column
y = wine['quality'].values
X = wine.drop('quality', axis = 1).values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 1234)
print('dimension of the dataset: ', wine.shape)
wine.head()
class GBMReg:
"""
Regression gradient boosting machine using scikit learn's
decision tree as the base tree
Parameters
----------
n_estimators: int
number of trees to train
learning_rate: float
learning rate, some calls it shrinkage,
shrinks the contribution of each tree
to prevent overfitting
max_depth: int
controls how deep to grow the tree;
this is more of a decision tree parameter,
it is tune here to make later comparison fair
all the other parameters for a decision tree like
max_features or min_sample_split also applies to GBM,
it is just not used here as that is more
related to a single decision tree
"""
def __init__(self, n_estimators, learning_rate, max_depth):
self.max_depth = max_depth
self.n_estimators = n_estimators
self.learning_rate = learning_rate
def fit(self, X, y):
self.estimators = []
# simply use the response as the original residuals
# and covert it to float type to prevent error warning
# that it's converting from int to float
residual = y.astype(np.float)
for i in range(self.n_estimators):
tree = DecisionTreeRegressor(max_depth = self.max_depth)
tree.fit(X, residual)
y_pred = tree.predict(X)
self.estimators.append(tree)
residual -= self.learning_rate * y_pred
return self
def predict(self, X):
y_pred = np.zeros(X.shape[0])
for tree in self.estimators:
y_pred += self.learning_rate * tree.predict(X)
return y_pred
# compare the results between a single decision tree,
# gradient boosting, the lower the mean square
# error, the better
tree = DecisionTreeRegressor(max_depth = 6)
tree.fit(X_train, y_train)
tree_y_pred = tree.predict(X_test)
print('tree: ', mean_squared_error(y_test, tree_y_pred))
# library to confirm result
gbm_reg = GBMReg(n_estimators = 100, learning_rate = 0.1, max_depth = 6)
gbm_reg.fit(X_train, y_train)
gbm_reg_y_pred = gbm_reg.predict(X_test)
print('gbm: ', mean_squared_error(y_test, gbm_reg_y_pred))
# gradient boosting for 100 trees and learning rate of 0.1
gbm = GradientBoostingRegressor(n_estimators = 100, learning_rate = 0.1, max_depth = 6)
gbm.fit(X_train, y_train)
gbm_y_pred = gbm.predict(X_test)
print('gbm library: ', mean_squared_error(y_test, gbm_y_pred))
```
Clearly, Gradient Boosting has some similarities to Random Forests and Extra Trees: the final prediction is based on an ensemble of models, and trees are used as the base learner, so all the tuning parameters for the tree model also controls the variability of Gradient Boosting. And for interpretability we can also access the feature importance attribute.
```
def viz_importance(model, feature_names, n_features):
"""Visualize the relative importance of predictors"""
# sort the importance in decreasing order
importances = model.feature_importances_
idx = np.argsort(importances)[-n_features:]
names = feature_names[idx]
scores = importances[idx]
y_pos = np.arange(1, n_features + 1)
plt.barh(y_pos, scores, color = 'lightskyblue', align = 'center')
plt.yticks(y_pos, names)
plt.xlabel('Importance')
plt.title('Feature Importance Plot')
# change default figure and font size
plt.rcParams['figure.figsize'] = 8, 6
plt.rcParams['font.size'] = 12
viz_importance(gbm, wine.columns[:-1], X.shape[1])
```
But the way the ensembles are constructed differs substantially between each model. In Random Forests and Extra Trees, all trees are created independently and each tree contributes equally to the final model. The trees in Gradient Boosting, however, are dependent on past trees and contribute unequally to the final model. Despite these differences, Random Forests, Extra Trees and Gradient Boosting all offer competitive predictive performance (Gradient Boosting often wins when carefully tuned). As for computation time, Gradient Boosting is often greater than for Random Forests, Extra Trees, since the two former models' procedure can be easily parallel processed given that their individual trees are created independently.
## Classification
Gradient Boosting Machine can also be extended to handle classification tasks, as we'll soon see, even in the classification context, the underlying algorithm is still a regression tree. To adapt the algorithm to a classification process, we start by defining a new loss function, cross entropy (also known as multinomial deviance), denoted as:
$$L\big(y_i, F(x_i)\big) = -\sum_k ^ K y_k(x_i) \log p_k(x_i)$$
The notation above says:
- We have a total of $K$ output class (categorical response variable) that ranges from $1, ..., K$
- $y_k(x_i)$ is a dummy indicator of the response variable that takes the value of 1 if the $i_{th}$ observation belongs to class $k$ and 0 otherwise
- $p_k(x_i)$ is the predicted probability of the $i_{th}$ observation belonging to class $k$
So the next question is how do we get $p_k(x_i)$?
### Softmax
Softmax function takes an $N$-dimensional vector of arbitrary real values and produces another $N$-dimensional vector with real values in the range (0, 1) that add up to 1. The function's formula can be written as:
$$p_i = \frac{e^{o_i}}{\sum_k^K e^{o_k}}$$
For example, in the following code chunk, we see that how the softmax function transforms a 3-element vector 1.0, 2.0, 3.0 into probabilities that sums up to 1, while still preserving the relative size of the original elements.
```
def compute_softmax(x):
"""compute the softmax of vector"""
exp_x = np.exp(x)
softmax = exp_x / np.sum(exp_x)
return softmax
# this can be interpreted as the probability
# of belonging to the three classes
compute_softmax([1, 2, 3])
```
Next, we wish to compute the derivative of this function with respect to the input $o_i$ so we can use it later when computing the derivative of the loss function. To be explicit we wish to find:
$$\frac{\partial p_i}{\partial o_j} = \frac{\partial \frac{e^{o_i}}{\sum_{k=1}^{N}e^{o_k}}}{\partial o_j}$$
For any arbitrary output $i$ and input $j$. To do so, We'll be using the quotient rule of derivatives. The rule tells us that for a function $f(x) = \frac{g(x)}{h(x)}$:
$$f'(x) = \frac{g'(x)h(x) - h'(x)g(x)}{[h(x)]^2}$$
In our case, we have:
$$
\begin{align*}
g &= e^{o_i} \nonumber \\
h &= \sum_{k=1}^{K}e^{o_k} \nonumber
\end{align*}
$$
It's important to notice that no matter which $o_j$ we compute the derivative of $h$ for the output will always be $e^{o_j}$. However, this is not the case for $g$. It's derivative will be $e^{o_j}$ only if $i = j$, because only then will it have the term $e^{o_j}$. Otherwise, the derivative is simply 0 (because it's simply taking the derivative of a constant).
So going back to using our quotient rule, we start with the $i = j$ case. In the following derivation we'll use the $\Sigma$ (Sigma) sign to represent $\sum_{k=1}^{K}e^{o_k}$ for simplicity and to prevent cluttering up the notation.
$$
\begin{align*}
\frac{\partial \frac{e^{o_i}}{\sum_{k = 1}^{N} e^{o_k}}}{\partial o_j}
&= \frac{e^{o_i}\Sigma-e^{o_j}e^{o_i}}{\Sigma^2} \nonumber \\
&= \frac{e^{o_i}}{\Sigma}\frac{\Sigma - e^{o_j}}{\Sigma} \nonumber \\
&= p_i(1 - p_j) \nonumber \\
&= p_i(1 - p_i) \nonumber
\end{align*}
$$
The reason we can perform the operation in the last line is because we're considering the scenario where $i = j$. Similarly we can do the case where $i \neq j$.
$$
\begin{align*}
\frac{\partial \frac{e^{o_i}}{\sum_{k = 1}^{N} e^{o_k}}}{\partial o_j}
&= \frac{0-e^{o_j}e^{o_i}}{\Sigma^2} \nonumber \\
&= -\frac{e^{o_j}}{\Sigma}\frac{e^{o_i}}{\Sigma} \nonumber \\
&= -p_j p_i \nonumber \\
&= -p_i p_j \nonumber
\end{align*}
$$
Just to sum it up, we now have:
$$\frac{\partial p_i}{\partial o_j} = p_i(1 - p_i),\quad i = j$$
$$\frac{\partial p_i}{\partial o_j} = -p_i p_j,\quad i \neq j$$
Now, we can tie this back to the original loss function $-\sum_k^K y_k \log p_k$ and compute its negative gradient.
$$
\begin{align}
\frac{\partial L}{\partial o_i}
&= -\sum_k y_k\frac{\partial \log p_k}{\partial o_i} \nonumber \\
&= -\sum_k y_k\frac{1}{p_k}\frac{\partial p_k}{\partial o_i} \nonumber \\
&= -y_i(1-p_i) - \sum_{k \neq i}y_k\frac{1}{p_k}(-p_kp_i) \nonumber \\
&= -y_i(1 - p_i) + \sum_{k \neq i}y_k(p_i) \nonumber \\
&= -y_i + y_i p_i + \sum_{k \neq i}y_k(p_i) \nonumber \\
&= p_i\left(\sum_ky_k\right) - y_i \nonumber \\
&= p_i - y_i \nonumber
\end{align}
$$
Remember $\sum_ky_k=1$ (as $y$ is a vector with only one non-zero element, which is $1$ when the indicating the observation belongs to the $k_{th}$ class.
After a long journey, we now see, for every class $k$, the gradient is the difference between the associated dummy variable and the predicted probability of belonging to that class. This is essentially the "residuals" from the classification gradient boosting. Given this, we can now implement the algorithm, the overall process of training a regression tree has still not changed, only now we must deal with the dummy variables, $y_k$ and fit a regression tree on the negative gradient for each dummy variable.
## Implementation
For the dataset, we'll still use the Wine Quality Data Set that was used for the regression task, except we now treat the quality of the wine (a scale of 1 ~ 10) as categorical instead of numeric.
```
from sklearn.metrics import accuracy_score
from sklearn.preprocessing import LabelEncoder
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import GradientBoostingClassifier
class GBMClass:
"""
Classification gradient boosting machine using scikit learn's
decision tree as the base tree
Parameters
----------
n_estimators: int
number of trees to train
learning_rate: float
learning rate, some calls it shrinkage,
shrinks the contribution of each tree
to prevent overfitting
max_depth: int
controls how deep to grow the tree;
this is more of a decision tree parameter,
it is tune here to make later comparison fair
all the other parameters for a decision tree like
max_features or min_sample_split also applies to GBM,
it is just not used here as that is more
related to a single decision tree
"""
def __init__(self, n_estimators, learning_rate, max_depth):
self.max_depth = max_depth
self.n_estimators = n_estimators
self.learning_rate = learning_rate
def fit(self, X, y):
# encode labels with value between 0 and n_classes - 1,
# so we can easily one-hot encode them
self.le = LabelEncoder()
labels = self.le.fit_transform(y)
Y = self._to_categorical(labels)
del labels
# the predicted probability starts out with
# a value that's uniform over all classes;
# then we compute the residuals (negative gradient),
# which is the difference between the predicted
# probability and the class label
y_proba = np.full(Y.shape, 1 / Y.shape[1])
residuals = Y - y_proba
# train a base decision tree on the residuals
# for every single class, hence we end up with
# n_estimators * n_classes base tree models
self.estimators = []
for i in range(self.n_estimators):
for j in range(self.n_classes):
tree = DecisionTreeRegressor(max_depth = self.max_depth)
tree.fit(X, residuals[:, j])
y_pred = tree.predict(X)
self.estimators.append(tree)
residuals[:, j] -= self.learning_rate * y_pred
return self
def _to_categorical(self, y):
"""one hot encode class vector y"""
self.n_classes = np.amax(y) + 1
Y = np.zeros((y.shape[0], self.n_classes))
for i in range(y.shape[0]):
Y[i, y[i]] = 1.0
return Y
def predict(self, X):
# after predicting the class remember to
# transform it back to the actual class label
y_prob = self.predict_proba(X)
y_pred = np.argmax(y_prob, axis = 1)
y_pred = self.le.inverse_transform(y_pred)
return y_pred
def predict_proba(self, X):
# add up raw score for every class and convert
# it to probability using softmax
y_raw = np.zeros((X.shape[0], self.n_classes))
# obtain the tree for each class and add up the prediction
for c in range(self.n_classes):
class_tree = self.estimators[c::self.n_classes]
for tree in class_tree:
y_raw[:, c] += self.learning_rate * tree.predict(X)
y_proba = self._compute_softmax(y_raw)
return y_proba
def _compute_softmax(self, z):
"""
compute the softmax of matrix z in a numerically stable way,
by substracting each row with the max of each row. For more
information refer to the following link:
https://nolanbconaway.github.io/blog/2017/softmax-numpy
"""
shift_z = z - np.amax(z, axis = 1, keepdims = 1)
exp_z = np.exp(shift_z)
softmax = exp_z / np.sum(exp_z, axis = 1, keepdims = 1)
return softmax
# compare the results between a single decision tree,
# gradient boosting, the higher the accuracy, the better
tree = DecisionTreeClassifier(max_depth = 6)
tree.fit(X_train, y_train)
tree_y_pred = tree.predict(X_test)
print('tree: ', accuracy_score(y_test, tree_y_pred))
# gradient boosting for 150 trees and learning rate of 0.2
# unlike random forest, gradient boosting's base tree can be shallower
# meaning that there depth can be smaller
gbm_class = GBMClass(n_estimators = 150, learning_rate = 0.2, max_depth = 3)
gbm_class.fit(X_train, y_train)
gbm_class_y_pred = gbm_class.predict(X_test)
print('gbm: ', accuracy_score(y_test, gbm_class_y_pred))
# library to confirm results are comparable
gbm = GradientBoostingClassifier(n_estimators = 150, learning_rate = 0.2, max_depth = 3)
gbm.fit(X_train, y_train)
gbm_y_pred = gbm.predict(X_test)
print('gbm library: ', accuracy_score(y_test, gbm_y_pred))
```
## Understanding Model Complexity
In the following section, we generate a Sinoide function + random gaussian noise, with 80 training samples (blue points) and 20 test samples (red points).
```
def ground_truth(x):
"""Ground truth -- function to approximate"""
return x * np.sin(x) + np.sin(2 * x)
def gen_data(low, high, n_samples):
"""generate training and testing data from the ground truth function"""
np.random.seed(15)
X = np.random.uniform(low, high, size = n_samples)
# generate the response from the ground truth function and add
# some random noise to it
y = ground_truth(X) + np.random.normal(scale = 2, size = n_samples)
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size = 0.2, random_state = 3)
return X_train, X_test, y_train, y_test
def plot_data(x_plot, X_train, X_test, y_train, y_test):
"""plot training and testing data"""
s = 20
alpha = 0.4
plt.plot(x_plot, ground_truth(x_plot), alpha = alpha, label = 'ground truth')
plt.scatter(X_train, y_train, s = s, alpha = alpha)
plt.scatter(X_test, y_test, s = s, alpha = alpha, color = 'red')
plt.xlim(( 0, 10 ))
plt.ylabel('y')
plt.xlabel('x')
plt.legend(loc = 'upper left')
plt.show()
low = 0
high = 10
x_plot = np.linspace(low, high, 500)
X_train, X_test, y_train, y_test = gen_data(low = low, high = high, n_samples = 100)
plot_data(x_plot, X_train, X_test, y_train, y_test)
```
Recall that in a single regression tree, we can use the `max_depth` parameter to control how deep to grow the tree and the deeper the tree the more variance can be explained.
```
# when using scikit-learn, the training data has to be
# a 2d-array even if it only has 1 features
tree1 = DecisionTreeRegressor(max_depth = 1)
tree1.fit(X_train[:, np.newaxis], y_train)
tree2 = DecisionTreeRegressor(max_depth = 3)
tree2.fit(X_train[:, np.newaxis], y_train)
plt.plot(x_plot, tree1.predict(x_plot[:, np.newaxis]),
label = 'RT max_depth=1', color = 'g', alpha = 0.9, linewidth = 2)
plt.plot(x_plot, tree2.predict(x_plot[:, np.newaxis]),
label = 'RT max_depth=3', color = 'g', alpha = 0.7, linewidth = 1)
plot_data(x_plot, X_train, X_test, y_train, y_test)
```
The plot above shows that the decision boundaries made by decision trees are always perpendicular to $x$ and $y$ axis (due to the fact that they consists of nested if-else statements). Let's see what happens when we use gradient boosting without tuning the parameters (by specifying a fix `max_depth`).
```
gbm = GradientBoostingRegressor(n_estimators = 300, max_depth = 6, learning_rate = 0.1)
gbm.fit(X_train[:, np.newaxis], y_train)
plt.plot(x_plot, gbm.predict(x_plot[:, np.newaxis]),
label = 'GBM max_depth=6', color = 'r', alpha = 0.9, linewidth = 2)
plot_data(x_plot, X_train, X_test, y_train, y_test)
```
Hopefully, it should be clear that compared with decision trees, gradient boosting machine is far more susceptible to overfitting the training data, hence it is common to tune parameters including `max_depth`, `max_features`, `min_samples_leaf`, `subsample` (explained below) to reduce the overfitting phenomenon from occurring.
The parameter `subsample` (technically called *stochastic gradient boosting*) borrows some idea from bagging techniques. What it does is: while iterating through each individual tree building process, it randomly select a fraction of the training data. Then the residuals and models in the remaining steps of the current iteration are based only on that sample of data. It turns out that this simple modification improved the predictive accuracy of boosting while also reducing the required computational resources (of course, this is based on the fact that you have enough observations to subsample).
The following section tunes the commonly tuned parameter and find the best one and draws the decision boundary. The resulting plot should be self-explanatory.
```
param_grid = {
'max_depth': [4, 6],
'min_samples_leaf': [3, 5, 8],
'subsample': [0.9, 1]
# 'max_features': [1.0, 0.3, 0.1] # not possible in this example (there's only 1)
}
gs_gbm = GridSearchCV(gbm, param_grid, scoring = 'neg_mean_squared_error', n_jobs = 4)
gs_gbm.fit(X_train[:, np.newaxis], y_train)
print('Best hyperparameters: %r' % gs_gbm.best_params_)
plt.plot(x_plot, gs_gbm.predict(x_plot[:, np.newaxis]),
label = 'GBM tuned', color = 'r', alpha = 0.9, linewidth = 2)
plot_data(x_plot, X_train, X_test, y_train, y_test)
```
# Reference
- [Slide: Gradent boosting](http://eric.univ-lyon2.fr/~ricco/cours/slides/en/gradient_boosting.pdf)
- [Slide: A gentle introduction to gradient boosting](http://www.ccs.neu.edu/home/vip/teach/MLcourse/4_boosting/slides/gradient_boosting.pdf)
- [Blog: The Softmax function and its derivative](http://eli.thegreenplace.net/2016/the-softmax-function-and-its-derivative/)
- [Notebook: Regression Trees and Rule-Based Models](http://nbviewer.jupyter.org/github/leig/Applied-Predictive-Modeling-with-Python/blob/master/notebooks/Chapter%208.ipynb)
- [Notebook: Gradient Boosted Regression Trees in scikit-learn](http://nbviewer.jupyter.org/github/pprett/pydata-gbrt-tutorial/blob/master/gbrt-tutorial.ipynb)
- [StackExchange: Derivative of Softmax loss function](http://math.stackexchange.com/questions/945871/derivative-of-softmax-loss-function)
- [StackExchange: How are individual trees added together in boosted regression tree?](http://stats.stackexchange.com/questions/135378/how-are-individual-trees-added-together-in-boosted-regression-tree)
- [Stackoverflow: How to access weighting of indiviual decision trees in xgboost?](https://stackoverflow.com/questions/32950607/how-to-access-weighting-of-indiviual-decision-trees-in-xgboost/34331573#34331573)
| github_jupyter |
```
from glob import glob
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from IPython.display import display, Markdown
%load_ext autoreload
%autoreload 2
from pymedphys.msq import (
mosaiq_connect, delivery_data_from_mosaiq, get_patient_fields,
get_qcls_by_date, multi_mosaiq_connect, get_incomplete_qcls_across_sites,
get_recently_completed_qcls_across_sites
)
from pymedphys.coll import calc_mu_density
from pymedphys.type import get_delivery_parameters
qa_centre = 'rccc'
treatment_centres = ['nbcc', 'sash']
centres = [qa_centre] + treatment_centres
qa_patient_ids = ['999999', '999998', '999997', '999996']
servers = {
'rccc': 'msqsql',
'nbcc': 'physics-server:31433', # nbccc-msq or physics-server:31433
'sash': 'physics-server'
}
servers_list = [
item for _, item in servers.items()
]
physics_location = {
'rccc': 'Physics_Check',
'nbcc': 'Physics',
'sash': 'Physics_Check'
}
imrt_task_names = {
'nbcc': 'Physics Check IMRT',
'rccc': 'IMRT Physics Check',
'sash': 'Physics QA '
}
comparison_field_types = [
'DMLC', 'VMAT'
]
with multi_mosaiq_connect(servers_list) as cursors:
incomplete_qcls = get_incomplete_qcls_across_sites(cursors, servers, centres, physics_location)
recently_completed_qcls = get_recently_completed_qcls_across_sites(cursors, servers, centres, physics_location)
display(incomplete_qcls)
display(recently_completed_qcls)
plans_to_verify = pd.DataFrame()
for centre in treatment_centres:
plans_to_verify = plans_to_verify.append(incomplete_qcls[(
(incomplete_qcls['centre'] == centre) &
(incomplete_qcls['task'] == imrt_task_names[centre])
)], sort=False)
plans_to_verify = plans_to_verify.append(recently_completed_qcls[(
(recently_completed_qcls['centre'] == centre) &
(recently_completed_qcls['task'] == imrt_task_names[centre])
)], sort=False)
plans_to_verify
patient_ids = plans_to_verify['patient_id'].values
patient_ids
all_qa_fields = pd.DataFrame()
with mosaiq_connect(servers[qa_centre]) as cursor:
for qa_patient_id in qa_patient_ids:
qa_fields = get_patient_fields(cursor, qa_patient_id)
qa_fields['patient_id'] = [qa_patient_id] * len(qa_fields)
all_qa_fields = all_qa_fields.append(qa_fields)
# all_qa_fields
clinical_field_ids = {}
clinical_field_tables = {}
with multi_mosaiq_connect(servers_list) as cursors:
for qcl in plans_to_verify.itertuples():
patient_id = qcl.patient_id
centre = qcl.centre
display(Markdown('### Centre: {}, Patient ID: {}'.format(centre, patient_id)))
cursor = cursors[servers[centre]]
clinical_fields = get_patient_fields(cursor, patient_id)
clinical_beam_reference = (
(clinical_fields['field_version'] == 0) &
(clinical_fields['monitor_units'] != 0) &
(clinical_fields['field_type'].isin(comparison_field_types))
)
clinical_fields = clinical_fields[clinical_beam_reference]
clinical_field_tables[(centre, patient_id)] = clinical_fields
display(clinical_fields)
clinical_field_ids[(centre, patient_id)] = clinical_fields['field_id'].values.astype(str)
clinical_field_ids
centre_patient_id_tuples = list(clinical_field_ids.keys())
centre_patient_id_tuples
qa_field_site_agreeing = {
(centre, patient_id): all_qa_fields[(
(all_qa_fields['site'] == patient_id)
)]
for centre, patient_id in centre_patient_id_tuples
}
for centre, patient_id in centre_patient_id_tuples:
display(Markdown('### Centre: {}, Patient ID: {}'.format(centre, patient_id)))
display(qa_field_site_agreeing[(centre, patient_id)])
qa_field_ids = {
key: qa_field_site_agreeing[key]['field_id'].values.astype(str)
for key in centre_patient_id_tuples
}
qa_field_ids
def determine_mu_density(cursor, field_id):
delivery_data = delivery_data_from_mosaiq(cursor, field_id)
mu, mlc, jaw = get_delivery_parameters(delivery_data)
mu_density = calc_mu_density(mu, mlc, jaw)
return mu_density
with multi_mosaiq_connect(servers_list) as cursors:
for centre, patient_id in centre_patient_id_tuples:
display(Markdown('### Centre: {}, Patient ID: {}'.format(centre, patient_id)))
display(Markdown('#### Clinical Fields'))
display(clinical_field_tables[(centre, patient_id)])
display(Markdown('#### QA Fields'))
display(qa_field_site_agreeing[(centre, patient_id)])
clinical_cursor = cursors[servers[centre]]
qa_cursor = cursors[servers[qa_centre]]
key = (centre, patient_id)
display(Markdown('#### Comparison'))
if len(clinical_field_ids[key]) == 0 or len(qa_field_ids[key]) == 0:
print('missing fields')
continue
clinical_mu_density = np.sum([
determine_mu_density(clinical_cursor, field_id)
for field_id in clinical_field_ids[key]
], axis=0)
qa_mu_density = np.sum([
determine_mu_density(qa_cursor, field_id)
for field_id in qa_field_ids[key]
], axis=0)
plt.figure()
plt.pcolormesh(clinical_mu_density)
plt.colorbar()
plt.title('Clinical MU density')
plt.xlabel('MLC direction (mm)')
plt.ylabel('Jaw direction (mm)')
plt.gca().invert_yaxis()
plt.figure()
plt.pcolormesh(qa_mu_density)
plt.colorbar()
plt.title('QA MU density')
plt.xlabel('MLC direction (mm)')
plt.ylabel('Jaw direction (mm)')
plt.gca().invert_yaxis()
plt.show()
# Verify that every pixel agrees within 0.1 MU
maximum_deviation = np.max(np.abs(clinical_mu_density - qa_mu_density))
print("Maximum MU Density deviation between fields: {}".format(maximum_deviation))
```
| github_jupyter |
In the first 2 learning units, we saw the so-called **Bag of Words** vectorization of text. Text is transformed in vectors, constituted by counts or frequencies of words. This representation has the advantage of being simple and lasted for a very long time.
A possible improvement to this idea is, instead of considering single words, to consider sequences of them. So, you can create features for 2 consecutives words, or even 3. You can even consider sequences of letters, or phonems. This approach is called **n-grams** reprensentation, and it could be very effective as well. Since we are not going to cover it in this material, you can find a very interesting explanation [here](https://www.youtube.com/watch?v=s3kKlUBa3b0&index=12&list=PL6397E4B26D00A269).
An important breakthrough in text representation has been made during recent past by using deep learning. The family of algorithms called **word embedding** - the most famous being `word2vec` - is based on training neural network to automatically learn the more informative representation for text data. The networks are trained to produce vectors that embed semantic information extracted from text. For example, words with similar meaning produces vectors that are close to one another in the vector space.
```
# Standard imports
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
import random
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer
# GenSim
import gensim
```
Train those kind of models is **very** computationally expensive. The good news is that, once trained by those guys with the big computers, we can happily use the trained models to project text to the feature space.
Thankfully, we can go [here](http://nlpserver2.inf.ufrgs.br/alexandres/vectors/lexvec.enwiki%2bnewscrawl.300d.W.pos.vectors.gz) and download a LexVec word embedding model (similar to word2vec and [GloVe](https://nlp.stanford.edu/projects/glove/)) pre-trained on Wikipedia.
Small note: the models have to be trained on text of the same language. So, one of the problems with such an approach is that most of tools are only available for English language.
Another small note: be careful, this will load a **pretty big** object in memory.
```
# Load Google's pre-trained Word2Vec model.
model = gensim.models.KeyedVectors.load_word2vec_format('./lexvec.enwiki+newscrawl.300d.W.pos.vectors')
```
As we said earlier, we have some cool properties in this vector space. We can treat words as numbers. What do you expect from `woman + king - man`?
```
model.wv.most_similar(positive=['woman', 'king'], negative=['man'])
```
Ok, that was nice. Let's see how things work with real data.
```
df = pd.read_csv('../hackathon-5-learning/02-text-in-practice/data/uci-news-aggregator.csv')
df.dtypes
```
We have already used this dataset in the last notebook but to give you some context, this dataset is made up of a different news headlines and their respective category. Let's look at some samples:
```
df = df[['TITLE', 'CATEGORY']]
df.columns = ['title', 'category']
df.head()
```
Let's plit our data into train and validation set so we can benchmark the final model!
```
_, data = train_test_split(df, test_size=0.3, random_state=42) # subsampling to help performance
train_df, validation_df = train_test_split(data, test_size=0.2, random_state=42)
```
Now that we have the dataset loaded and set up we get into the interesting part, transforming the document from a variable length sequence of words into a fixed length vector representation!
As we have talked about in previous hackathons, categorical features are generally converted not in an integer but in a onehot encoding of the category. That is, giving a small example:
If the possible values of a given feature "gender" would be:
["Male", "Female", "Unknown", "Attack Helicopter"]
The entry with the feature "Male" would be represented as:
(1, 0, 0, 0)
This procedure is important as we don't impose any information about the problem, if we would convert the values into either 1, 2, 3 or 4 we would be giving additional information as we could lead our classifier to believe that unkown and female are similar concepts since 2 and 3 are close on the line of real numbers.
Here is where word vectors come in handy! Word vectors help us find a representation in a continuous space (latent space) where similar words are close in space!
The previous example (woman + (king - queen)) aimed to show the structure of this space.
The model we loaded earlier keeps a dictionary of words and their vector representation in this space (model.wv). Looking at the word vectors of a concrete example:
```
# Load the word vector for the word "car"
king_vector = model.wv['king']
print("Word vector dimension: ", king_vector.shape)
# Load the word vector for the word "bus" and "cat"
queen_vector = model.wv['queen']
peasant_vector = model.wv['peasant']
print("Similarity between King and Queen: ", np.dot(king_vector, queen_vector))
print("Similarity between Kind and Peasant: ", np.dot(king_vector, peasant_vector))
```
In this example, we used the dot product as a means of calculating how similar two vectors are, we can see that the model assigns vectors which are similar for "king" and "queen" but separates them in space from the concept of "peasant".
Small side note: While it is curious to observe a political agenda in a word vector embedding model, this is an example of possible human bias in the input data and alerts to the dangers of giving our models biased data. In the end we are all humans, being peasant or king!
# Document vector representation
Now that we looked at our first word vectors, we are ready to move on to convert our documents into this space!
We start by splitting our sentences in words (tokens).
```
# Extract document tokens
tfidf = TfidfVectorizer(preprocessor=lambda x: x, stop_words='english')
tokenizer = tfidf.build_analyzer()
documents = train_df
tokens = [[word for word in tokenizer(document)] for document in documents['title']]
tokens[0]
```
Now that we have the sentence sepparated in tokens we can easily convert the tokens into the word vector space using the Word2Vec model's word vector dictionary.
```
# Convert each token (word) into vector representation
vectors = [np.array([model.wv[token] if token in model.wv else model.wv['the'] for token in sentence]) for sentence in tokens]
```
Looking at a single sentence, we can see that the sentence has become an array of vectors, a matrix. Why matrix you ask?
It becomes a matrix when we convert the tokens in the sentence to vecctors. Looking at the shape of a sentence we can see that it is of the dimension (9 x 300), where 9 is the number of words in the sentence and 300 the dimension of the word vector space. This means in one sentence we have 9 word vectors where each has a dimension of 300 entries. Sounds about right!
```
vectors[0].shape
```
For each document (sentence), we have now have an array of vectors. One vector for each word. Argh, that's a lot of numbers! But don't dispair, we can "summarize" each document!
There are far smarter ways to do that, but to keep things simple, we can take the mean of all the words. We end up with a fixed size vector for each document!:
```
# Average word vectors to get document representation
vectorized = np.array([document.mean(axis=0) for document in vectors])
vectorized.shape
```
Looking at a single entry we can see that the dimension of the sentence representation is 300, this is natural since we averaged the representation of all the vectors in the sentence!
Like everything in life, there is no free lunch! By reducing the array of vectors to a fixed sized representtion of 300 by averaging the word vectors we loose some information! Nevertheless this averaged representation captures the average location of words in the word vector space!
For example, if the sentence is made up of words in the general field of finance, the average of these word vectors would be somewhere in the finance region of the word vector space.
```
model.wv.similar_by_vector(vectorized[0], topn=10, restrict_vocab=None)
```
Looking at the words around the region of the sentence vector, this vector does not seem to capture a lot of information!
Let's try to fix this by weighting the average by the term frequencies so these generic words are ignored!
We start by creating a dictionary of the term frequencies:
```
tfidf = TfidfVectorizer(preprocessor=lambda x: x)
tfidf.fit(train_df['title'])
tfidf_dictionary = {w: tfidf.idf_[i] for w, i in tfidf.vocabulary_.items()}
```
Then we create a vector with each sentences token weights and average the word vectors using this weighting vector.
```
weights = [[tfidf_dictionary[token] for token in sequence] for sequence in tokens]
weighted_vectors = np.array([np.average(vectors[i], weights=weights[i], axis=0) for i in range(0, len(documents))])
model.wv.similar_by_vector(weighted_vectors[0], topn=10, restrict_vocab=None)
```
Correcting for the term frequencies we were able to get a much better document representation!
As you can see, this representation captures some semantical structure of the sentence. In some problems, this can produce very discriminative features.
This is a fairly basic approach that can be further improved in many ways, from the model used to extract the embeddings (vectors) to the strategy used to combine the word vectors into the sentence vector.
| github_jupyter |
ERROR: type should be string, got "https://github.com/d2l-ai/d2l-en/issues/1116\n\n```\nfrom d2l import torch as d2l\nimport torch\nfrom torch import nn\nimport numpy as np\nimport math\nmax_degree = 20 # Maximum degree of the polynomial\nn_train, n_test = 100, 100 # Training and test dataset sizes\ntrue_w = np.zeros(max_degree) # Allocate lots of empty space\ntrue_w[0:4] = np.array([5, 1.2, -3.4, 5.6])\n\nfeatures = np.random.normal(size=(n_train + n_test, 1))\nnp.random.shuffle(features)\npoly_features = np.power(features, np.arange(max_degree).reshape(1, -1))\nfor i in range(max_degree):\n poly_features[:, i] /= math.gamma(i + 1) # `gamma(n)` = (n-1)!\n# Shape of `labels`: (`n_train` + `n_test`,)\nlabels = np.dot(poly_features, true_w)\nlabels += np.random.normal(scale=0.1, size=labels.shape)\nfeatures[:2].T, poly_features[:2, :], labels[:2]\n# Convert from NumPy to PyTorch tensors\ntrue_w, features, poly_features, labels = [torch.from_numpy(x).type(\n torch.float32) for x in [true_w, features, poly_features, labels]]\ndef evaluate_loss(net, data_iter, loss): #@save\n \"\"\"Evaluate the loss of a model on the given dataset.\"\"\"\n metric = d2l.Accumulator(2) # Sum of losses, no. of examples\n for X, y in data_iter:\n l = loss(net(X), y)\n metric.add(l.sum(), l.numel())\n return metric[0] / metric[1]\ndef train(train_features, test_features, train_labels, test_labels,\n num_epochs=400):\n loss = nn.MSELoss()\n input_shape = train_features.shape[-1]\n # Switch off the bias since we already catered for it in the polynomial\n # features\n net = nn.Sequential(nn.Linear(input_shape, 1, bias=False))\n batch_size = min(10, train_labels.shape[0])\n train_iter = d2l.load_array((train_features, train_labels.reshape(-1,1)),\n batch_size)\n test_iter = d2l.load_array((test_features, test_labels.reshape(-1,1)),\n batch_size, is_train=False)\n trainer = torch.optim.SGD(net.parameters(), lr=0.01)\n animator = d2l.Animator(xlabel='epoch', ylabel='loss', yscale='log',\n xlim=[1, num_epochs], ylim=[1e-3, 1e2],\n legend=['train', 'test'])\n for epoch in range(num_epochs):\n d2l.train_epoch_ch3(net, train_iter, loss, trainer)\n if epoch == 0 or (epoch + 1) % 20 == 0:\n animator.add(epoch + 1, (evaluate_loss(net, train_iter, loss),\n evaluate_loss(net, test_iter, loss)))\n print('weight:', net[0].weight.data.numpy())\n# Pick the first four dimensions, i.e., 1, x, x^2/2!, x^3/3! from the\n# polynomial features\ntrain(poly_features[:n_train, :4], poly_features[n_train:, :4],\n labels[:n_train], labels[n_train:])\n# Pick from the original `features` for linear function fitting\ntrain(features[:n_train, :], features[n_train:, :], labels[:n_train],\n labels[n_train:])\n# Pick all the dimensions from the polynomial features\ntrain(poly_features[:n_train, :], poly_features[n_train:, :],\n labels[:n_train], labels[n_train:], num_epochs=1500)\n```\n\n" | github_jupyter |
# Catalyst classification tutorial
## Requirements
Download and install the latest version of catalyst and other libraries required for this tutorial.
```
!pip install -U catalyst
!pip install albumentations
!pip install pretrainedmodels
```
### Colab extras
First of all, do not forget to change the runtime type to GPU. <br/>
To do so click `Runtime` -> `Change runtime type` -> Select `"Python 3"` and `"GPU"` -> click `Save`. <br/>
After that you can click `Runtime` -> `Run` all and watch the tutorial.
To intergate visualization library `plotly` to colab, run
```
import IPython
def configure_plotly_browser_state():
display(IPython.core.display.HTML('''
<script src="/static/components/requirejs/require.js"></script>
<script>
requirejs.config({
paths: {
base: '/static/base',
plotly: 'https://cdn.plot.ly/plotly-latest.min.js?noext',
},
});
</script>
'''))
IPython.get_ipython().events.register('pre_run_cell', configure_plotly_browser_state)
```
## Setting up GPUs
PyTorch and Catalyst versions:
```
import torch, catalyst
torch.__version__, catalyst.__version__
```
You can also specify GPU/CPU usage for this turorial.
Available GPUs
```
from catalyst.utils import get_available_gpus
get_available_gpus()
import os
from typing import List, Tuple, Callable
# os.environ["CUDA_VISIBLE_DEVICES"] = "0" # "" - CPU, "0" - 1 GPU, "0,1" - MultiGPU
```
---
## Reproducibility first
Catalyst provides a special utils for research results reproducibility. <br/>
For example, `catalyst.utils.set_global_seed` fixes seed for all main DL frameworks (` PyTorch`, `Tensorflow`,` random` and `numpy`)
In case of CuDNN you can set deterministic mode and flag to use benchmark with
`catalyst.utils.prepare_cudnn`.
```
SEED = 42
from catalyst.utils import set_global_seed, prepare_cudnn
set_global_seed(SEED)
prepare_cudnn(deterministic=True)
```
## Dataset
In this tutorial we will use one of four datasets:
- [Ants / Bees dataset](https://www.dropbox.com/s/ffzfpbwzwdo9qp8/ants_bees_cleared_190806.tar.gz )
- [Stanford Dogs dataset](http://vision.stanford.edu/aditya86/ImageNetDogs/)
- [Flowers Recognition](https://www.kaggle.com/alxmamaev/flowers-recognition)
- [Best Artworks of All Time](https://www.kaggle.com/ikarus777/best-artworks-of-all-time)
If you are on MacOS and you don’t have `wget`, you can install it with:` brew install wget`.
Let's specify the dataset by `DATASET` variable:
- `ants_bees` (~ 400 pictures for 2 classes) – for CPU experiments
- `flowers` (~ 4k pictures for 5 classes) – for fast GPU experiments
- `dogs` (~ 20k pictures for 120 classes)
- `artworks` (~ 8k pictures for 50 classes) – for GPU experiments
```
DATASET = "artworks" # "ants_bees" / "flowers" / "dogs" / "artworks"
os.environ["DATASET"] = DATASET
%%bash
function gdrive_download () {
CONFIRM=$(wget --quiet --save-cookies /tmp/cookies.txt --keep-session-cookies --no-check-certificate "https://docs.google.com/uc?export=download&id=$1" -O- | sed -rn 's/.*confirm=([0-9A-Za-z_]+).*/\1\n/p')
wget --load-cookies /tmp/cookies.txt "https://docs.google.com/uc?export=download&confirm=$CONFIRM&id=$1" -O $2
rm -rf /tmp/cookies.txt
}
mkdir -p Images
rm -rf Images/
if [[ "$DATASET" == "ants_bees" ]]; then
gdrive_download 1czneYKcE2sT8dAMHz3FL12hOU7m1ZkE7 ants_bees_cleared_190806.tar.gz
tar -xf ants_bees_cleared_190806.tar.gz &>/dev/null
mv ants_bees_cleared_190806 Images
elif [[ "$DATASET" == "flowers" ]]; then
gdrive_download 1rvZGAkdLlbR_MEd4aDvXW11KnLaVRGFM flowers.tar.gz
tar -xf flowers.tar.gz &>/dev/null
mv flowers Images
elif [[ "$DATASET" == "dogs" ]]; then
# https://www.kaggle.com/jessicali9530/stanford-dogs-dataset
wget http://vision.stanford.edu/aditya86/ImageNetDogs/images.tar
tar -xf images.tar &>/dev/null
elif [[ "$DATASET" == "artworks" ]]; then
# https://www.kaggle.com/ikarus777/best-artworks-of-all-time
gdrive_download 1eAk36MEMjKPKL5j9VWLvNTVKk4ube9Ml artworks.tar.gz
tar -xf artworks.tar.gz &>/dev/null
mv artworks Images
fi
from pathlib import Path
ROOT = "Images/"
ALL_IMAGES = list(Path(ROOT).glob("**/*.jpg"))
ALL_IMAGES = list(filter(lambda x: not x.name.startswith("."), ALL_IMAGES))
print("Number of images:", len(ALL_IMAGES))
```
Let's check out the data!
```
from catalyst.utils import imread
import numpy as np
import matplotlib.pyplot as plt
def show_examples(images: List[Tuple[str, np.ndarray]]):
_indexes = [(i, j) for i in range(2) for j in range(2)]
f, ax = plt.subplots(2, 2, figsize=(16, 16))
for (i, j), (title, img) in zip(_indexes, images):
ax[i, j].imshow(img)
ax[i, j].set_title(title)
f.tight_layout()
def read_random_images(paths: List[Path]) -> List[Tuple[str, np.ndarray]]:
data = np.random.choice(paths, size=4)
result = []
for d in data:
title = f"{d.parent.name}: {d.name}"
_image = imread(d)
result.append((title, _image))
return result
```
You can restart the cell below to see more examples.
```
images = read_random_images(ALL_IMAGES)
show_examples(images)
```
## Dataset preprocessing
With Catalyst we can easily create a dataset from the following folder structure:
```
dataset/
class_1/
*.ext
...
class_2/
*.ext
...
...
class_N/
*.ext
...
```
`create_dataset` function goes through a given directory and creates a dictionary `Dict[class_name, List[image]]`
```
from catalyst.utils.dataset import create_dataset, create_dataframe, prepare_dataset_labeling
dataset = create_dataset(dirs=f"{ROOT}/*", extension="*.jpg")
```
and `create_dataframe` function creates typical `pandas.DataFrame` for further analysis
```
df = create_dataframe(dataset, columns=["class", "filepath"])
df.head()
len(df)
```
finally `prepare_dataset_labeling` creates a numerical label for each unique class name
```
tag_to_label = prepare_dataset_labeling(df, "class")
tag_to_label
```
Let's add a column with a numerical label value to the DataFrame.
It can be easily done with `map_dataframe` function.
```
from catalyst.utils.pandas import map_dataframe
df_with_labels = map_dataframe(df, tag_column="class", class_column="label", tag2class=tag_to_label, verbose=True)
df_with_labels.head()
```
additionaly let's save the `class_names` for further usage
```
class_names = [name for name, id_ in sorted(tag_to_label.items(), key=lambda x: x[1])]
class_names
```
Now let's divide our dataset into the `train` and` valid` parts.
The parameters for the split_dataframe function are the same as [sklearn.train_test_split](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html#sklearn-model-selection-train-test-split).
We also define `test_size` (it is optional) and `random_state` for reproducibility.
```
from catalyst.utils.dataset import split_dataframe
train_data, valid_data = split_dataframe(df_with_labels, test_size=0.2, random_state=SEED)
train_data, valid_data = train_data.to_dict('records'), valid_data.to_dict('records')
len(train_data), len(valid_data)
```
## Augmentations
In order to save your time during data preparation/reading/writing, Catalyst provides a special abstraction – [Reader](https://catalyst-team.github.io/catalyst/api/data.html#reader). <br/>
Reader allows you to read various structures, for example, images, strings, numerical values and perform some functions on top of them.
```
from catalyst.dl import utils
from catalyst.data.reader import ImageReader, ScalarReader, ReaderCompose, LambdaReader
from functools import partial
import torch
num_classes = len(tag_to_label)
# ReaderCompose collects different Readers into one pipeline
open_fn = ReaderCompose([
# Reads images from the `datapath` folder using the key `input_key =" filepath "` (here should be the filename)
# and writes it to the output dictionary by `output_key="features"` key
ImageReader(
input_key="filepath",
output_key="features",
datapath=ROOT
),
# Reads a number from our dataframe by the key `input_key =" label "` to np.long
# and writes it to the output dictionary by `output_key="targets"` key
ScalarReader(
input_key="label",
output_key="targets",
default_value=-1,
dtype=np.int64
),
# Same as above, but with one encoding
ScalarReader(
input_key="label",
output_key="targets_one_hot",
default_value=-1,
dtype=np.int64,
one_hot_classes=num_classes
)
])
```
For augmentation of our dataset, we will use the [albumentations library](https://github.com/albu/albumentations). <br/>
You can view the list of available augmentations on the documentation [website](https://albumentations.readthedocs.io/en/latest/api/augmentations.html).
```
from albumentations import Compose, LongestMaxSize, PadIfNeeded
from albumentations import ShiftScaleRotate, IAAPerspective, RandomBrightnessContrast, RandomGamma, \
HueSaturationValue, ToGray, CLAHE, JpegCompression
from albumentations import Normalize
from albumentations.pytorch import ToTensor
BORDER_CONSTANT = 0
BORDER_REFLECT = 2
def pre_transforms(image_size=224):
# Convert the image to a square of size image_size x image_size
# (keeping aspect ratio)
result = [
LongestMaxSize(max_size=image_size),
PadIfNeeded(image_size, image_size, border_mode=BORDER_CONSTANT)
]
return result
def hard_transforms():
result = [
# Random shifts, stretches and turns with a 50% probability
ShiftScaleRotate(
shift_limit=0.1,
scale_limit=0.1,
rotate_limit=15,
border_mode=BORDER_REFLECT,
p=0.5
),
IAAPerspective(scale=(0.02, 0.05), p=0.3),
# Random brightness / contrast with a 30% probability
RandomBrightnessContrast(
brightness_limit=0.2, contrast_limit=0.2, p=0.3
),
# Random gamma changes with a 30% probability
RandomGamma(gamma_limit=(85, 115), p=0.3),
# Randomly changes the hue, saturation, and color value of the input image
HueSaturationValue(p=0.3),
JpegCompression(quality_lower=80),
]
return result
def post_transforms():
# we use ImageNet image normalization
# and convert it to torch.Tensor
return [Normalize(), ToTensor()]
def compose(_transforms):
# combine all augmentations into one single pipeline
result = Compose([item for sublist in _transforms for item in sublist])
return result
```
Like Reader, there is a close abstraction for handling augmentations and key-value-based dataloaders – [Augmentor](https://catalyst-team.github.io/catalyst/api/data.html#augmentor).
```
from catalyst.data.augmentor import Augmentor
from torchvision import transforms
train_transforms = compose([pre_transforms(), hard_transforms(), post_transforms()])
valid_transforms = compose([pre_transforms(), post_transforms()])
show_transforms = compose([pre_transforms(), hard_transforms()])
# Takes an image from the input dictionary by the key `dict_key` and performs `train_transforms` on it.
train_data_transforms = transforms.Compose([
Augmentor(
dict_key="features",
augment_fn=lambda x: train_transforms(image=x)["image"]
)
])
# Similarly for the validation part of the dataset.
# we only perform squaring, normalization and ToTensor
valid_data_transforms = transforms.Compose([
Augmentor(
dict_key="features",
augment_fn=lambda x: valid_transforms(image=x)["image"]
)
])
```
Let's look at the augmented results. <br/>
The cell below can be restarted.
```
images = read_random_images(ALL_IMAGES)
images = [
(title, show_transforms(image=i)["image"])
for (title, i) in images
]
show_examples(images)
```
## Pytorch dataloaders
Using `catalyst.utils.get_loader`, you can immediately get loaders only from the dataset and data-converting functions.
```
import collections
bs = 64
num_workers = 4
def get_loaders(
open_fn: Callable,
train_transforms_fn: transforms.Compose,
valid_transforms_fn: transforms.Compose,
batch_size: int = 64,
num_workers: int = 4,
sampler = None
) -> collections.OrderedDict:
"""
Args:
open_fn: Reader for reading data from a dataframe
train_transforms_fn: Augmentor for train part
valid_transforms_fn: Augmentor for valid part
batch_size: batch size
num_workers: How many subprocesses to use to load data,
sampler: An object of the torch.utils.data.Sampler class
for the dataset data sampling strategy specification
"""
train_loader = utils.get_loader(
train_data,
open_fn=open_fn,
dict_transform=train_transforms_fn,
batch_size=bs,
num_workers=num_workers,
shuffle=sampler is None, # shuffle data only if Sampler is not specified (PyTorch requirement)
sampler=sampler
)
valid_loader = utils.get_loader(
valid_data,
open_fn=open_fn,
dict_transform=valid_transforms_fn,
batch_size=bs,
num_workers=num_workers,
shuffle=False,
sampler=None
)
# Catalyst expects an ordered dictionary with train/valid/infer loaders.
# The number of loaders can vary.
# For example, it can easily handle even some complex logic like:
# loaders["train_dataset1"] = train_loader_1
# loaders["train_dataset2"] = train_loader_2
# ....
# loaders["valid_1"] = valid_loader_1
# loaders["valid_2"] = valid_loader_2
# ...
# loaders["infer_1"] = infer_loader_1
# loaders["infer_2"] = infer_loader_2
# ...
loaders = collections.OrderedDict()
loaders["train"] = train_loader
loaders["valid"] = valid_loader
return loaders
loaders = get_loaders(open_fn, train_data_transforms, valid_data_transforms)
```
## Model
Let's take the classification model from [Cadene pretrain models](https://github.com/Cadene/pretrained-models.pytorch). This repository contains a huge number of pre-trained PyTorch models. <br/>
But at first, let's check them out!
```
import pretrainedmodels
pretrainedmodels.model_names
```
For this tutorial purposes, `ResNet18` is good enough, but you can try other models
```
model_name = "resnet18"
```
By `pretrained_settings` we can see what the given network expects as input and what would be the expected output.
```
pretrainedmodels.pretrained_settings[model_name]
```
The model returns logits for classification into 1000 classes from ImageNet. <br/>
Let's define a function that will replace the last fully-conected layer for our number of classes.
```
from torch import nn
def get_model(model_name: str, num_classes: int, pretrained: str = "imagenet"):
model_fn = pretrainedmodels.__dict__[model_name]
model = model_fn(num_classes=1000, pretrained=pretrained)
dim_feats = model.last_linear.in_features
model.last_linear = nn.Linear(dim_feats, num_classes)
return model
```
## Model training
```
import torch
# model creation
model = get_model(model_name, num_classes)
# as we are working on basic classification problem (no multi-class/multi-label)
# let's use standard CE loss
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters(), lr=0.0003)
scheduler = torch.optim.lr_scheduler.MultiStepLR(
optimizer, milestones=[9], gamma=0.3
)
```
To run some DL experiment, Catalyst uses a [Runner](https://catalyst-team.github.io/catalyst/api/dl.html#catalyst.dl.core.runner.Runner) abstraction. <br/>
It contains main logic about "how" you run the experiment and getting predictions.
For supervised learning case, there is an extention for Runner – [SupervisedRunner](https://catalyst-team.github.io/catalyst/api/dl.html#module-catalyst.dl.runner.supervised), which provides additional methods like `train`, `infer` and `predict_loader`.
```
from catalyst.dl.runner import SupervisedRunner
runner = SupervisedRunner()
# folder for all the experiment logs
logdir = "./logs/classification_tutorial_0"
NUM_EPOCHS = 10
```
Using [Callbacks](https://catalyst-team.github.io/catalyst/api/dl.html#catalyst.dl.core.callback.Callback), the basic functionality of the catalyst can be expanded.
A callback is a class inherited from `catalyst.dl.core.Callback` and implements one / several / all methods:
```
on_stage_start
--- on_epoch_start
------ on_loader_start
--------- on_batch_start
--------- on_batch_end
------ on_loader_end
--- on_epoch_end
on_stage_end
on_exception - if the code crashes with an error, you can catch it and reserve the parameters you need
```
You can find the list of standard callbacks [here](https://catalyst-team.github.io/catalyst/api/dl.html#module-catalyst.dl.callbacks.checkpoint).
It includes callbacks such as
- CheckpointCallback. Saves N best models in logdir
- TensorboardLogger. Logs all metrics to tensorboard
- EarlyStoppingCallback. Early training stop if metrics do not improve for the `patience` of epochs
- ConfusionMatrixCallback. Plots ConfusionMatrix per epoch in tensorboard
Many different metrics for classfication
- AccuracyCallback
- MapKCallback
- AUCCallback
- F1ScoreCallback
segmentation
- DiceCallback
- IouCallback
and many other callbacks, like LRFinder and MixupCallback
```
# as we are working on classification task
from catalyst.dl.callbacks import AccuracyCallback, AUCCallback, F1ScoreCallback
runner.train(
model=model,
logdir=logdir,
criterion=criterion,
optimizer=optimizer,
scheduler=scheduler,
loaders=loaders,
# We can specify the callbacks list for the experiment;
# For this task, we will check accuracy, AUC and F1 metrics
callbacks=[
AccuracyCallback(num_classes=num_classes),
AUCCallback(
num_classes=num_classes,
input_key="targets_one_hot",
class_names=class_names
),
F1ScoreCallback(
input_key="targets_one_hot",
activation="Softmax"
)
],
num_epochs=NUM_EPOCHS,
verbose=True
)
```
### Training analysis and model predictions
The `utils.plot_metrics` method reads tensorboard logs from the logdir and plots beautiful metrics with `plotly` package.
```
# it can take a while (colab issue)
utils.plot_metrics(
logdir=logdir,
# specify which metrics we want to plot
metrics=["loss", "accuracy01", "auc/_mean", "f1_score", "_base/lr"]
)
```
The method below will help us look at the predictions of the model for each image.
```
from torch.nn.functional import softmax
def show_prediction(
model: torch.nn.Module,
class_names: List[str],
titles: List[str],
images: List[np.ndarray],
device: torch.device
) -> None:
with torch.no_grad():
tensor_ = torch.stack([
valid_transforms(image=image)["image"]
for image in images
]).to(device)
logits = model.forward(tensor_)
probabilities = softmax(logits, dim=1)
predictions = probabilities.argmax(dim=1)
images_predicted_classes = [
(f"predicted: {class_names[x]} | correct: {title}", image)
for x, title, image in zip(predictions, titles, images)
]
show_examples(images_predicted_classes)
device = utils.get_device()
titles, images = list(zip(*read_random_images(ALL_IMAGES)))
titles = list(map(lambda x: x.rsplit(":")[0], titles))
show_prediction(model, class_names=class_names, titles=titles, images=images, device=device)
```
### Training with Focal Loss and OneCycle
In the `catalyst.contrib` there are a large number of different additional criterions, models, layers etc
For example,
[catalyst.contrib.criterion](https://catalyst-team.github.io/catalyst/api/contrib.html#module-catalyst.contrib.criterion.ce):
- HuberLoss
- CenterLoss
- FocalLossMultiClass
- DiceLoss / BCEDiceLoss
- IoULoss / BCEIoULoss
- LovaszLossBinary / LovaszLossMultiClass / LovaszLossMultiLabel
- WingLoss
Lr scheduler in [catalyst.contrib.schedulers](https://catalyst-team.github.io/catalyst/api/contrib.html#module-catalyst.contrib.schedulers.base):
- OneCycleLRWithWarmup
Moreover, in [catalyst.contrib.models](https://catalyst-team.github.io/catalyst/api/contrib.html#models) you can find various models for segmentation:
- Unet / ResnetUnet
- Linknet / ResnetLinknet
- FPNUnet / ResnetFPNUnet
- PSPnet / ResnetPSPnet
- MobileUnet
Finally, several handwritten modules in [catalyst.contrib.modules](https://catalyst-team.github.io/catalyst/api/contrib.html#module-catalyst.contrib.modules.common):
- Flatten
- TemporalAttentionPooling
- LamaPooling
- NoisyLinear
- GlobalAvgPool2d / GlobalMaxPool2d
- GlobalConcatPool2d / GlobalAttnPool2d
a bunch of others
But for now, let's take `FocalLoss` and `OneCycleLRWithWarmup` to play around.
```
from catalyst.contrib.criterion import FocalLossMultiClass
from catalyst.contrib.schedulers import OneCycleLRWithWarmup
model = get_model(model_name, num_classes)
criterion = FocalLossMultiClass()
optimizer = torch.optim.Adam(model.parameters(), lr=0.0003)
scheduler = OneCycleLRWithWarmup(
optimizer,
num_steps=NUM_EPOCHS,
lr_range=(0.001, 0.0001),
warmup_steps=1
)
# FocalLoss expects one_hot for the input
# in our Reader function we have already created the conversion of targets in one_hot
# so, all we need - respecify the target key name
runner = SupervisedRunner(input_target_key="targets_one_hot")
logdir = "./logs/classification_tutorial_1"
NUM_EPOCHS = 10
runner.train(
model=model,
logdir=logdir,
criterion=criterion,
optimizer=optimizer,
scheduler=scheduler,
loaders=loaders,
callbacks=[
AccuracyCallback(num_classes=num_classes),
AUCCallback(
num_classes=num_classes,
input_key="targets_one_hot",
class_names=class_names
),
F1ScoreCallback(
input_key="targets_one_hot",
activation="Softmax"
)
],
num_epochs=NUM_EPOCHS,
verbose=True,
)
# it can take a while (colab issue)
utils.plot_metrics(
logdir=logdir,
metrics=["loss", "accuracy01", "auc/_mean", "f1_score", "_base/lr"]
)
device = utils.get_device()
titles, images = list(zip(*read_random_images(ALL_IMAGES)))
titles = list(map(lambda x: x.rsplit(":")[0], titles))
show_prediction(model, class_names=class_names, titles=titles, images=images, device=device)
```
### Balancing classes in the dataset
There are several useful data-sampler implementations in the `catalyst.data.sampler`. For example,
- `BalanceClassSampler` allows you to create stratified sampling on an unbalanced dataset. <br/> A strategy can be either downsampling, upsampling or some prespeficied number of samples per class. <br/> Very important feature for every classification problem.
- `MiniEpochSampler` allows you to split your "very large dataset" and sample some small portion of it every epoch. <br/> This is useful for those cases where you need to check valid metrics and save checkpoints more often. <br/> For example, your 1M images dataset can be sampled in 100k per epoch with all necessary metrics.
```
from catalyst.data.sampler import BalanceClassSampler
labels = [x["label"] for x in train_data]
sampler = BalanceClassSampler(labels, mode="upsampling")
# let's re-create our loaders with BalanceClassSampler
loader = get_loaders(open_fn, train_data_transforms, valid_data_transforms, sampler=sampler)
model = get_model(model_name, num_classes)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters(), lr=0.0003)
scheduler = OneCycleLRWithWarmup(
optimizer,
num_steps=NUM_EPOCHS,
lr_range=(0.001, 0.0001),
warmup_steps=1
)
runner = SupervisedRunner()
logdir = "./logs/classification_tutorial_2"
NUM_EPOCHS = 10
runner.train(
model=model,
logdir=logdir,
criterion=criterion,
optimizer=optimizer,
scheduler=scheduler,
loaders=loaders,
callbacks=[
AccuracyCallback(num_classes=num_classes),
AUCCallback(
num_classes=num_classes,
input_key="targets_one_hot",
class_names=class_names
),
F1ScoreCallback(
input_key="targets_one_hot",
activation="Softmax"
)
],
num_epochs=NUM_EPOCHS,
verbose=True,
)
# it can take a while (colab issue)
utils.plot_metrics(
logdir=logdir,
metrics=["loss", "accuracy01", "auc/_mean", "f1_score", "_base/lr"]
)
device = utils.get_device()
titles, images = list(zip(*read_random_images(ALL_IMAGES)))
titles = list(map(lambda x: x.rsplit(":")[0], titles))
show_prediction(model, class_names=class_names, titles=titles, images=images, device=device)
```
## Model inference
With SupervisedRunner, you can easily predict entire loader with only one method call.
```
predictions = runner.predict_loader(
model, loaders["valid"],
resume=f"{logdir}/checkpoints/best.pth", verbose=True
)
```
The resulting object has shape = (number of elements in the loader, output shape from the model)
```
predictions.shape
```
Thus, we can obtain probabilities for our classes.
```
print("logits: ", predictions[0])
probabilities = torch.softmax(torch.from_numpy(predictions[0]), dim=0)
print("probabilities: ", probabilities)
label = probabilities.argmax().item()
print(f"predicted: {class_names[label]}")
```
| github_jupyter |
# Loading Image Data
So far we've been working with fairly artificial datasets that you wouldn't typically be using in real projects. Instead, you'll likely be dealing with full-sized images like you'd get from smart phone cameras. In this notebook, we'll look at how to load images and use them to train neural networks.
We'll be using a [dataset of cat and dog photos](https://www.kaggle.com/c/dogs-vs-cats) available from Kaggle. Here are a couple example images:
<img src='assets/dog_cat.png'>
We'll use this dataset to train a neural network that can differentiate between cats and dogs. These days it doesn't seem like a big accomplishment, but five years ago it was a serious challenge for computer vision systems.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import torch
from torchvision import datasets, transforms
import helper
```
The easiest way to load image data is with `datasets.ImageFolder` from `torchvision` ([documentation](http://pytorch.org/docs/master/torchvision/datasets.html#imagefolder)). In general you'll use `ImageFolder` like so:
```python
dataset = datasets.ImageFolder('path/to/data', transform=transforms)
```
where `'path/to/data'` is the file path to the data directory and `transforms` is a list of processing steps built with the [`transforms`](http://pytorch.org/docs/master/torchvision/transforms.html) module from `torchvision`. ImageFolder expects the files and directories to be constructed like so:
```
root/dog/xxx.png
root/dog/xxy.png
root/dog/xxz.png
root/cat/123.png
root/cat/nsdf3.png
root/cat/asd932_.png
```
where each class has it's own directory (`cat` and `dog`) for the images. The images are then labeled with the class taken from the directory name. So here, the image `123.png` would be loaded with the class label `cat`. You can download the dataset already structured like this [from here](https://s3.amazonaws.com/content.udacity-data.com/nd089/Cat_Dog_data.zip). I've also split it into a training set and test set.
### Transforms
When you load in the data with `ImageFolder`, you'll need to define some transforms. For example, the images are different sizes but we'll need them to all be the same size for training. You can either resize them with `transforms.Resize()` or crop with `transforms.CenterCrop()`, `transforms.RandomResizedCrop()`, etc. We'll also need to convert the images to PyTorch tensors with `transforms.ToTensor()`. Typically you'll combine these transforms into a pipeline with `transforms.Compose()`, which accepts a list of transforms and runs them in sequence. It looks something like this to scale, then crop, then convert to a tensor:
```python
transforms = transforms.Compose([transforms.Resize(255),
transforms.CenterCrop(224),
transforms.ToTensor()])
```
There are plenty of transforms available, I'll cover more in a bit and you can read through the [documentation](http://pytorch.org/docs/master/torchvision/transforms.html).
### Data Loaders
With the `ImageFolder` loaded, you have to pass it to a [`DataLoader`](http://pytorch.org/docs/master/data.html#torch.utils.data.DataLoader). The `DataLoader` takes a dataset (such as you would get from `ImageFolder`) and returns batches of images and the corresponding labels. You can set various parameters like the batch size and if the data is shuffled after each epoch.
```python
dataloader = torch.utils.data.DataLoader(dataset, batch_size=32, shuffle=True)
```
Here `dataloader` is a [generator](https://jeffknupp.com/blog/2013/04/07/improve-your-python-yield-and-generators-explained/). To get data out of it, you need to loop through it or convert it to an iterator and call `next()`.
```python
# Looping through it, get a batch on each loop
for images, labels in dataloader:
pass
# Get one batch
images, labels = next(iter(dataloader))
```
>**Exercise:** Load images from the `Cat_Dog_data/train` folder, define a few transforms, then build the dataloader.
```
data_dir = 'dogs-vs-cats/train'
trans = transforms.Compose([transforms.Resize(255),
transforms.CenterCrop(224),
transforms.ToTensor()])
dataset = datasets.ImageFolder(data_dir, transform=trans)
dataloader = torch.utils.data.DataLoader(dataset, batch_size=32, shuffle=True)
# Run this to test your data loader
images, labels = next(iter(dataloader))
helper.imshow(images[0], normalize=False)
```
If you loaded the data correctly, you should see something like this (your image will be different):
<img src='assets/cat_cropped.png', width=244>
## Data Augmentation
A common strategy for training neural networks is to introduce randomness in the input data itself. For example, you can randomly rotate, mirror, scale, and/or crop your images during training. This will help your network generalize as it's seeing the same images but in different locations, with different sizes, in different orientations, etc.
To randomly rotate, scale and crop, then flip your images you would define your transforms like this:
```python
train_transforms = transforms.Compose([transforms.RandomRotation(30),
transforms.RandomResizedCrop(100),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.5, 0.5, 0.5],
[0.5, 0.5, 0.5])])
```
You'll also typically want to normalize images with `transforms.Normalize`. You pass in a list of means and list of standard deviations, then the color channels are normalized like so
```input[channel] = (input[channel] - mean[channel]) / std[channel]```
Subtracting `mean` centers the data around zero and dividing by `std` squishes the values to be between -1 and 1. Normalizing helps keep the network work weights near zero which in turn makes backpropagation more stable. Without normalization, networks will tend to fail to learn.
You can find a list of all [the available transforms here](http://pytorch.org/docs/0.3.0/torchvision/transforms.html). When you're testing however, you'll want to use images that aren't altered (except you'll need to normalize the same way). So, for validation/test images, you'll typically just resize and crop.
>**Exercise:** Define transforms for training data and testing data below.
```
data_dir = 'dogs-vs-cats'
# TODO: Define transforms for the training data and testing data
train_transforms = transforms.Compose([transforms.RandomRotation(30),
transforms.RandomResizedCrop(100),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.5, 0.5, 0.5],
[0.5, 0.5, 0.5])])
test_transforms = transforms.Compose([transforms.Resize(255),
transforms.CenterCrop(224),
transforms.ToTensor()])
# Pass transforms in here, then run the next cell to see how the transforms look
train_data = datasets.ImageFolder(data_dir + '/train', transform=train_transforms)
test_data = datasets.ImageFolder(data_dir + '/test', transform=test_transforms)
trainloader = torch.utils.data.DataLoader(train_data, batch_size=32)
testloader = torch.utils.data.DataLoader(test_data, batch_size=32)
# change this to the trainloader or testloader
data_iter = iter(testloader)
images, labels = next(data_iter)
fig, axes = plt.subplots(figsize=(10,4), ncols=4)
for ii in range(4):
ax = axes[ii]
helper.imshow(images[ii], ax=ax, normalize=False)
```
Your transformed images should look something like this.
<center>Training examples:</center>
<img src='assets/train_examples.png' width=500px>
<center>Testing examples:</center>
<img src='assets/test_examples.png' width=500px>
At this point you should be able to load data for training and testing. Now, you should try building a network that can classify cats vs dogs. This is quite a bit more complicated than before with the MNIST and Fashion-MNIST datasets. To be honest, you probably won't get it to work with a fully-connected network, no matter how deep. These images have three color channels and at a higher resolution (so far you've seen 28x28 images which are tiny).
In the next part, I'll show you how to use a pre-trained network to build a model that can actually solve this problem.
```
# Optional TODO: Attempt to build a network to classify cats vs dogs from this dataset
import fc_model
from torch import nn, optim
model = fc_model.Network(784, 10, [512, 256, 128])
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.parameters(), lr=0.001)
fc_model.train(model, trainloader, testloader, criterion, optimizer, epochs=2)
```
| github_jupyter |
```
Copyright 2021 IBM Corporation
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
```
# Decision Tree on Credit Card Fraud Dataset
## Background
The goal of this learning task is to predict if a credit card transaction is fraudulent or genuine based on a set of anonymized features.
## Source
The raw dataset can be obtained directly from [Kaggle](https://www.kaggle.com/mlg-ulb/creditcardfraud).
In this example, we download the dataset directly from Kaggle using their API.
In order for this to work, you must login into Kaggle and folow [these instructions](https://www.kaggle.com/docs/api) to install your API token on your machine.
## Goal
The goal of this notebook is to illustrate how Snap ML can accelerate training of a decision tree model on this dataset.
## Code
```
cd ../../
CACHE_DIR='cache-dir'
import numpy as np
import time
from datasets import CreditCardFraud
from sklearn.tree import DecisionTreeClassifier
from snapml import DecisionTreeClassifier as SnapDecisionTreeClassifier
from sklearn.metrics import roc_auc_score as score
dataset = CreditCardFraud(cache_dir=CACHE_DIR)
X_train, X_test, y_train, y_test = dataset.get_train_test_split()
print("Number of examples: %d" % (X_train.shape[0]))
print("Number of features: %d" % (X_train.shape[1]))
print("Number of classes: %d" % (len(np.unique(y_train))))
# the dataset is highly imbalanced
labels, sizes = np.unique(y_train, return_counts=True)
print("%6.2f %% of the training transactions belong to class 0" % (sizes[0]*100.0/(sizes[0]+sizes[1])))
print("%6.2f %% of the training transactions belong to class 1" % (sizes[1]*100.0/(sizes[0]+sizes[1])))
from sklearn.utils.class_weight import compute_sample_weight
w_train = compute_sample_weight('balanced', y_train)
w_test = compute_sample_weight('balanced', y_test)
model = DecisionTreeClassifier(max_depth=16, random_state=42)
t0 = time.time()
model.fit(X_train, y_train, sample_weight=w_train)
t_fit_sklearn = time.time()-t0
score_sklearn = score(y_test, model.predict_proba(X_test)[:,1], sample_weight=w_test)
print("Training time (sklearn): %6.2f seconds" % (t_fit_sklearn))
print("ROC AUC score (sklearn): %.4f" % (score_sklearn))
model = SnapDecisionTreeClassifier(max_depth=16, n_jobs=4, random_state=42)
t0 = time.time()
model.fit(X_train, y_train, sample_weight=w_train)
t_fit_snapml = time.time()-t0
score_snapml = score(y_test, model.predict_proba(X_test)[:,1], sample_weight=w_test)
print("Training time (snapml): %6.2f seconds" % (t_fit_snapml))
print("ROC AUC score (snapml): %.4f" % (score_snapml))
speed_up = t_fit_sklearn/t_fit_snapml
score_diff = (score_snapml-score_sklearn)/score_sklearn
print("Speed-up: %.1f x" % (speed_up))
print("Relative diff. in score: %.4f" % (score_diff))
```
## Disclaimer
Performance results always depend on the hardware and software environment.
Information regarding the environment that was used to run this notebook are provided below:
```
import utils
environment = utils.get_environment()
for k,v in environment.items():
print("%15s: %s" % (k, v))
```
## Record Statistics
Finally, we record the enviroment and performance statistics for analysis outside of this standalone notebook.
```
import scrapbook as sb
sb.glue("result", {
'dataset': dataset.name,
'n_examples_train': X_train.shape[0],
'n_examples_test': X_test.shape[0],
'n_features': X_train.shape[1],
'n_classes': len(np.unique(y_train)),
'model': type(model).__name__,
'score': score.__name__,
't_fit_sklearn': t_fit_sklearn,
'score_sklearn': score_sklearn,
't_fit_snapml': t_fit_snapml,
'score_snapml': score_snapml,
'score_diff': score_diff,
'speed_up': speed_up,
**environment,
})
```
| github_jupyter |
<a href="https://colab.research.google.com/github/praveentn/hgwxx7/blob/master/ampligraph/Clustering_And_Classification_With_Embeddings.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
---
# Clustering and Classification using Knowledge Graph Embeddings
---
In this tutorial we will explore how to use the knowledge embeddings generated by a graph of international football matches (since the 19th century) in clustering and classification tasks. Knowledge graph embeddings are typically used for missing link prediction and knowledge discovery, but they can also be used for entity clustering, entity disambiguation, and other downstream tasks. The embeddings are a form of representation learning that allow linear algebra and machine learning to be applied to knowledge graphs, which otherwise would be difficult to do.
We will cover in this tutorial:
1. Creating the knowledge graph (i.e. triples) from a tabular dataset of football matches
2. Training the ComplEx embedding model on those triples
3. Evaluating the quality of the embeddings on a validation set
4. Clustering the embeddings, comparing to the natural clusters formed by the geographical continents
5. Applying the embeddings as features in classification task, to predict match results
6. Evaluating the predictive model on a out-of-time test set, comparing to a simple baseline
We will show that knowledge embedding clusters manage to capture implicit geographical information from the graph and that they can be a useful feature source for a downstream machine learning classification task, significantly increasing accuracy from the baseline.
---
## Requirements
A Python environment with the AmpliGraph library installed. Please follow the [install guide](http://docs.ampligraph.org/en/latest/install.html).
Some sanity check:
```
%%capture
!pip install ampligraph
import numpy as np
import pandas as pd
import ampligraph
ampligraph.__version__
```
## Dataset
We will use the [International football results from 1872 to 2019](https://www.kaggle.com/martj42/international-football-results-from-1872-to-2017) available at Kaggle (public domain). It contains over 40 thousand international football matches. Each row contains the following information:
1. Match date
2. Home team name
3. Away team name
4. Home score (goals including extra time)
5. Away score (goals including extra time)
6. Tournament (whether it was a friendly match or part of a tournament)
7. City where match took place
8. Country where match took place
9. Whether match was on neutral grounds
This dataset comes in a tabular format, therefore we will need to construct the knowledge graph ourselves.
```
import requests
url = 'https://ampligraph.s3-eu-west-1.amazonaws.com/datasets/football_graph.csv'
open('football_results.csv', 'wb').write(requests.get(url).content)
df = pd.read_csv("football_results.csv").sort_values("date")
df.isna().sum()
```
Dropping matches with unknown score:
```
df = df.dropna()
```
The training set will be from 1872 to 2014, while the test set will be from 2014 to present date. Note that a temporal test set makes any machine learning task harder compared to a random shuffle.
```
df["train"] = df.date < "2014-01-01"
df.train.value_counts()
```
## Knowledge graph creation
We are going to create a knowledge graph from scratch based on the match information. The idea is that each match is an entity that will be connected to its participating teams, geography, characteristics, and results.
The objective is to generate a new representation of the dataset where each data point is an triple in the form:
<subject, predicate, object>
First we need to create the entities (subjects and objects) that will form the graph. We make sure teams and geographical information result in different entities (e.g. the Brazilian team and the corresponding country will be different).
```
# Entities naming
df["match_id"] = df.index.values.astype(str)
df["match_id"] = "Match" + df.match_id
df["city_id"] = "City" + df.city.str.title().str.replace(" ", "")
df["country_id"] = "Country" + df.country.str.title().str.replace(" ", "")
df["home_team_id"] = "Team" + df.home_team.str.title().str.replace(" ", "")
df["away_team_id"] = "Team" + df.away_team.str.title().str.replace(" ", "")
df["tournament_id"] = "Tournament" + df.tournament.str.title().str.replace(" ", "")
df["neutral"] = df.neutral.astype(str)
```
Then, we create the actual triples based on the relationship between the entities. We do it only for the triples in the training set (before 2014).
```
triples = []
for _, row in df[df["train"]].iterrows():
# Home and away information
home_team = (row["home_team_id"], "isHomeTeamIn", row["match_id"])
away_team = (row["away_team_id"], "isAwayTeamIn", row["match_id"])
# Match results
if row["home_score"] > row["away_score"]:
score_home = (row["home_team_id"], "winnerOf", row["match_id"])
score_away = (row["away_team_id"], "loserOf", row["match_id"])
elif row["home_score"] < row["away_score"]:
score_away = (row["away_team_id"], "winnerOf", row["match_id"])
score_home = (row["home_team_id"], "loserOf", row["match_id"])
else:
score_home = (row["home_team_id"], "draws", row["match_id"])
score_away = (row["away_team_id"], "draws", row["match_id"])
home_score = (row["match_id"], "homeScores", np.clip(int(row["home_score"]), 0, 5))
away_score = (row["match_id"], "awayScores", np.clip(int(row["away_score"]), 0, 5))
# Match characteristics
tournament = (row["match_id"], "inTournament", row["tournament_id"])
city = (row["match_id"], "inCity", row["city_id"])
country = (row["match_id"], "inCountry", row["country_id"])
neutral = (row["match_id"], "isNeutral", row["neutral"])
year = (row["match_id"], "atYear", row["date"][:4])
triples.extend((home_team, away_team, score_home, score_away,
tournament, city, country, neutral, year, home_score, away_score))
```
Note that we treat some literals (year, neutral match, home score, away score) as discrete entities and they will be part of the final knowledge graph used to generate the embeddings. We limit the number of score entities by clipping the score to be at most 5.
Below we can see visualise a subset of the graph related to the infamous [Maracanazo](https://en.wikipedia.org/wiki/Uruguay_v_Brazil_(1950_FIFA_World_Cup)):

The whole graph related to this match can be summarised by the triples below:
```
triples_df = pd.DataFrame(triples, columns=["subject", "predicate", "object"])
triples_df[(triples_df.subject=="Match3129") | (triples_df.object=="Match3129")]
```
## Training knowledge graph embeddings
We split our training dataset further into training and validation, where the new training set will be used to the knowledge embedding training and the validation set will be used in its evaluation. The test set will be used to evaluate the performance of the classification algorithm built on top of the embeddings.
What differs from the standard method of randomly sampling N points to make up our validation set is that our data points are two entities linked by some relationship, and we need to take care to ensure that all entities are represented in train and validation sets by at least one triple.
To accomplish this, AmpliGraph provides the [`train_test_split_no_unseen`](https://docs.ampligraph.org/en/latest/generated/ampligraph.evaluation.train_test_split_no_unseen.html#train-test-split-no-unseen) function.
```
from ampligraph.evaluation import train_test_split_no_unseen
X_train, X_valid = train_test_split_no_unseen(np.array(triples), test_size=10000)
print('Train set size: ', X_train.shape)
print('Test set size: ', X_valid.shape)
```
AmpliGraph has implemented [several Knowledge Graph Embedding models](https://docs.ampligraph.org/en/latest/ampligraph.latent_features.html#knowledge-graph-embedding-models) (TransE, ComplEx, DistMult, HolE), but to begin with we're just going to use the [ComplEx](https://docs.ampligraph.org/en/latest/generated/ampligraph.latent_features.ComplEx.html#ampligraph.latent_features.ComplEx) model, which is known to bring state-of-the-art predictive power.
The hyper-parameter choice was based on the [best results](https://docs.ampligraph.org/en/latest/experiments.html) we have found so far for the ComplEx model applied to some benchmark datasets used in the knowledge graph embeddings community. This tutorial does not cover [hyper-parameter tuning](https://docs.ampligraph.org/en/latest/examples.html#model-selection).
```
from ampligraph.latent_features import ComplEx
model = ComplEx(batches_count=50,
epochs=300,
k=100,
eta=20,
optimizer='adam',
optimizer_params={'lr':1e-4},
loss='multiclass_nll',
regularizer='LP',
regularizer_params={'p':3, 'lambda':1e-5},
seed=0,
verbose=True)
```
Lets go through the parameters to understand what's going on:
- **`batches_count`** : the number of batches in which the training set is split during the training loop. If you are having into low memory issues than settings this to a higher number may help.
- **`epochs`** : the number of epochs to train the model for.
- **`k`**: the dimensionality of the embedding space.
- **`eta`** ($\\eta$) : the number of negative, or false triples that must be generated at training runtime for each positive, or true triple.
- **`optimizer`** : the Adam optimizer, with a learning rate of 1e-4 set via the *optimizer_params* kwarg.
- **`loss`** : pairwise loss, with a margin of 0.5 set via the *loss_params* kwarg.
- **`regularizer`** : $L_p$ regularization with $p=3$, i.e. l3 regularization. $\\lambda$ = 1e-5, set via the *regularizer_params* kwarg.
- **`seed`** : random seed, used for reproducibility.
- **`verbose`** - displays a progress bar.
Training should take around 10 minutes on a modern GPU:
```
import tensorflow as tf
tf.logging.set_verbosity(tf.logging.ERROR)
model.fit(X_train)
```
## Evaluating knowledge embeddings
AmpliGraph aims to follow scikit-learn's ease-of-use design philosophy and simplify everything down to **`fit`**, **`evaluate`**, and **`predict`** functions.
However, there are some knowledge graph specific steps we must take to ensure our model can be trained and evaluated correctly. The first of these is defining the filter that will be used to ensure that no negative statements generated by the corruption procedure are actually positives. This is simply done by concatenating our train and test sets. Now when negative triples are generated by the corruption strategy, we can check that they aren't actually true statements.
```
filter_triples = np.concatenate((X_train, X_valid))
```
For this we'll use the `evaluate_performance` function:
- **`X`** - the data to evaluate on. We're going to use our test set to evaluate.
- **`model`** - the model we previously trained.
- **`filter_triples`** - will filter out the false negatives generated by the corruption strategy.
- **`use_default_protocol`** - specifies whether to use the default corruption protocol. If True, then subj and obj are corrupted separately during evaluation.
- **`verbose`** - displays a progress bar.
```
from ampligraph.evaluation import evaluate_performance
ranks = evaluate_performance(X_valid,
model=model,
filter_triples=filter_triples,
use_default_protocol=True,
verbose=True)
```
We're going to use the mrr_score (mean reciprocal rank) and hits_at_n_score functions.
- **mrr_score**: The function computes the mean of the reciprocal of elements of a vector of rankings ranks.
- **hits_at_n_score**: The function computes how many elements of a vector of rankings ranks make it to the top n positions.
```
from ampligraph.evaluation import mr_score, mrr_score, hits_at_n_score
mr = mr_score(ranks)
mrr = mrr_score(ranks)
print("MRR: %.2f" % (mrr))
print("MR: %.2f" % (mr))
hits_10 = hits_at_n_score(ranks, n=10)
print("Hits@10: %.2f" % (hits_10))
hits_3 = hits_at_n_score(ranks, n=3)
print("Hits@3: %.2f" % (hits_3))
hits_1 = hits_at_n_score(ranks, n=1)
print("Hits@1: %.2f" % (hits_1))
```
We can interpret these results by stating that the model will rank the correct entity within the top-3 possibilities 29% of the time.
By themselves, these metrics are not enough to conclude the usefulness of the embeddings in a downstream task, but they suggest that the embeddings have learned a reasonable representation enough to consider using them in more tasks.
## Clustering and embedding visualization
To evaluate the subjective quality of the embeddings, we can visualise the embeddings on 2D space and also cluster them on the original space. We can compare the clustered embeddings with natural clusters, in this case the continent where the team is from, so that we have a ground truth to evaluate the clustering quality both qualitatively and quantitatively.
Requirements:
* seaborn
* adjustText
* incf.countryutils
For seaborn and adjustText, simply install them with `pip install seaborn adjustText`.
For incf.countryutils, do the following steps:
```bash
git clone https://github.com/wyldebeast-wunderliebe/incf.countryutils.git
cd incf.countryutils
pip install .```
Run the following code to install requirements in colab environment
```
%%capture
!pip install seaborn adjustText
!git clone https://github.com/wyldebeast-wunderliebe/incf.countryutils.git
!pip install incf.countryutils/.
from sklearn.decomposition import PCA
import matplotlib.pyplot as plt
import seaborn as sns
from adjustText import adjust_text
from incf.countryutils import transformations
%matplotlib inline
```
We create a map from the team ID (e.g. "TeamBrazil") to the team name (e.g. "Brazil") for visualization purposes.
```
id_to_name_map = {**dict(zip(df.home_team_id, df.home_team)), **dict(zip(df.away_team_id, df.away_team))}
```
We now create a dictionary with the embeddings of all teams:
```
teams = pd.concat((df.home_team_id[df["train"]], df.away_team_id[df["train"]])).unique()
team_embeddings = dict(zip(teams, model.get_embeddings(teams)))
```
We use PCA to project the embeddings from the 200 space into 2D space:
```
embeddings_2d = PCA(n_components=2).fit_transform(np.array([i for i in team_embeddings.values()]))
```
We will cluster the teams embeddings on its original 200-dimensional space using the `find_clusters` in our discovery API:
```
from ampligraph.discovery import find_clusters
from sklearn.cluster import KMeans
clustering_algorithm = KMeans(n_clusters=6, n_init=50, max_iter=500, random_state=0)
clusters = find_clusters(teams, model, clustering_algorithm, mode='entity')
```
This helper function uses the `incf.countryutils` library to translate country names to their corresponding continents.
```
def cn_to_ctn(country):
try:
return transformations.cn_to_ctn(id_to_name_map[country])
except KeyError:
return "unk"
```
This dataframe contains for each team their projected embeddings to 2D space via PCA, their continent and the KMeans cluster. This will be used alongisde Seaborn to make the visualizations.
```
plot_df = pd.DataFrame({"teams": teams,
"embedding1": embeddings_2d[:, 0],
"embedding2": embeddings_2d[:, 1],
"continent": pd.Series(teams).apply(cn_to_ctn),
"cluster": "cluster" + pd.Series(clusters).astype(str)})
```
We plot the results on a 2D scatter plot, coloring the teams by the continent or cluster and also displaying some individual team names.
We always display the names of the top 20 teams (according to [FIFA rankings](https://en.wikipedia.org/wiki/FIFA_World_Rankings)) and a random subset of the rest.
```
top20teams = ["TeamBelgium", "TeamFrance", "TeamBrazil", "TeamEngland", "TeamPortugal", "TeamCroatia", "TeamSpain",
"TeamUruguay", "TeamSwitzerland", "TeamDenmark", "TeamArgentina", "TeamGermany", "TeamColombia",
"TeamItaly", "TeamNetherlands", "TeamChile", "TeamSweden", "TeamMexico", "TeamPoland", "TeamIran"]
def plot_clusters(hue):
np.random.seed(0)
plt.figure(figsize=(12, 12))
plt.title("{} embeddings".format(hue).capitalize())
ax = sns.scatterplot(data=plot_df[plot_df.continent!="unk"], x="embedding1", y="embedding2", hue=hue)
texts = []
for i, point in plot_df.iterrows():
if point["teams"] in top20teams or np.random.random() < 0.1:
texts.append(plt.text(point['embedding1']+0.02, point['embedding2']+0.01, str(point["teams"])))
adjust_text(texts)
```
The first visualisation of the 2D embeddings shows the natural geographical clusters (continents), which can be seen as a form of the ground truth:
```
plot_clusters("continent")
```
We can see above that the embeddings learned geographical similarities even though this information was not explicit on the original dataset.
Now we plot the same 2D embeddings but with the clusters found by K-Means:
```
plot_clusters("cluster")
```
We can see that K-Means found very similar cluster to the natural geographical clusters by the continents. This shows that on the 200-dimensional embedding space, similar teams appear close together, which can be captured by a clustering algorithm.
Our evaluation of the clusters can be more objective by using a metric such as the [adjusted Rand score](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.adjusted_rand_score.html), which varies from -1 to 1, where 0 is random labelling and 1 is a perfect match:
```
from sklearn import metrics
metrics.adjusted_rand_score(plot_df.continent, plot_df.cluster)
```
## Classification
We will use the knowledge embeddings to predict future matches as a classification problem.
We can model it as a multiclass problem with three classes: home team wins, home team loses, draw.
The embeddings are used directly as features to a XGBoost classifier.
First we need to determine the target:
```
df["results"] = (df.home_score > df.away_score).astype(int) + \
(df.home_score == df.away_score).astype(int)*2 + \
(df.home_score < df.away_score).astype(int)*3 - 1
df.results.value_counts(normalize=True)
```
Now we create a function that extracts the features (knowledge embeddings for home and away teams) and the target for a particular subset of the dataset:
```
def get_features_target(mask):
def get_embeddings(team):
return team_embeddings.get(team, np.full(200, np.nan))
X = np.hstack((np.vstack(df[mask].home_team_id.apply(get_embeddings).values),
np.vstack(df[mask].away_team_id.apply(get_embeddings).values)))
y = df[mask].results.values
return X, y
clf_X_train, y_train = get_features_target((df["train"]))
clf_X_test, y_test = get_features_target((~df["train"]))
clf_X_train.shape, clf_X_test.shape
```
Note that we have 200 features by team because the ComplEx model uses imaginary and real number for its embeddings, so we have twice as many parameters as defined by `k=100` in its model definition.
We also have some missing information from the embeddings of the entities (i.e. teams) that only appear in the test set, which are unlikely to be correctly classified:
```
np.isnan(clf_X_test).sum()/clf_X_test.shape[1]
```
First install xgboost with `pip install xgboost`.
```
%%capture
!pip install xgboost
from xgboost import XGBClassifier
```
Create a multiclass model with 500 estimators:
```
clf_model = XGBClassifier(n_estimators=500, max_depth=5, objective="multi:softmax")
```
Fit the model using all of the training samples:
```
clf_model.fit(clf_X_train, y_train)
```
The baseline accuracy for this problem is 47%, as that is the frequency of the most frequent class (home team wins):
```
df[~df["train"]].results.value_counts(normalize=True)
metrics.accuracy_score(y_test, clf_model.predict(clf_X_test))
```
In conclusion, while the baseline for this classification problem was 47%, with just the knowledge embeddings alone we were able to build a classifier that achieves **54%** accuracy.
As future work, we could add more features to the model (not embeddings related) and tune the model hyper-parameters.
---
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/AssetManagement/export_TimeSeries2.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/AssetManagement/export_TimeSeries2.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=AssetManagement/export_TimeSeries2.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/AssetManagement/export_TimeSeries2.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geehydro](https://github.com/giswqs/geehydro). The **geehydro** Python package builds on the [folium](https://github.com/python-visualization/folium) package and implements several methods for displaying Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, `Map.centerObject()`, and `Map.setOptions()`.
The magic command `%%capture` can be used to hide output from a specific cell.
```
# %%capture
# !pip install earthengine-api
# !pip install geehydro
```
Import libraries
```
import ee
import folium
import geehydro
```
Authenticate and initialize Earth Engine API. You only need to authenticate the Earth Engine API once. Uncomment the line `ee.Authenticate()`
if you are running this notebook for this first time or if you are getting an authentication error.
```
# ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
This step creates an interactive map using [folium](https://github.com/python-visualization/folium). The default basemap is the OpenStreetMap. Additional basemaps can be added using the `Map.setOptions()` function.
The optional basemaps can be `ROADMAP`, `SATELLITE`, `HYBRID`, `TERRAIN`, or `ESRI`.
```
Map = folium.Map(location=[40, -100], zoom_start=4)
Map.setOptions('HYBRID')
```
## Add Earth Engine Python script
## Display Earth Engine data layers
```
Map.setControlVisibility(layerControl=True, fullscreenControl=True, latLngPopup=True)
Map
```
| github_jupyter |
## Hierarchical Regression
Hierarchical models enable sharing of features among groups. The parameters of the model are assumed to be sampled from a common distribution that models similarity between groups. The figure below shows three different scenarios that illustrate the benefit of hierarchical modeling. In the figure on the left, we have a single set of parameters $\theta$ that model the entire sequence of observations referred to as a pooled model. Here any variation in data is not modelled explicitly since we are assuming a common set of parameters that give rise to the data. On the other hand, we have an unpooled scenario where we model a different set of parameters for each observation. In the unpooled case, we are assuming that there is no sharing of parameters between groups of observations and that each parameter is independent. The hierarchical model combines the best of both worlds: it assumes that there's a common distribution from which individual parameters are sampled and therefore captures similarities between groups.
<img src="figures/hierarchical_gm.png">
In Bayesian Hierarchical Regression, we can assign priors on model parameters and use MCMC sampling to infer posterior distribution. We use the radon dataset to regress radon gas levels in houses of different counties based on the floor number (in particular if there's a basement or not). Thus our regression model will look like the following:
\begin{equation}
\alpha_c \sim N(\mu_a, \sigma_{a}^{2})\\
\beta_c \sim N(\mu_{\beta}, \sigma_{\beta}^{2})\\
\mathrm{radon}_c = \alpha_c + \beta_c \times \mathrm{floor}_{i,c} + \epsilon_c
\end{equation}
Notice that subscript $c$ indicates a county, thus we are learning an intercept and a slope for each county sampled from a shared Gaussian distribution. Thus, we are assuming a hierarchical model in which our parameters ($\alpha_c$ and $\beta_c$) are sampled from a common distribution. Having specified the model, we can implement it using PyMC3.
```
%matplotlib inline
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import pymc3 as pm
#load data
data = pd.read_csv('./radon.txt')
data.head()
county_names = data.county.unique()
county_idx = data['county_code'].values
with pm.Model() as hierarchical_model:
# Hyperpriors
mu_a = pm.Normal('mu_alpha', mu=0., sd=100**2)
sigma_a = pm.Uniform('sigma_alpha', lower=0, upper=100)
mu_b = pm.Normal('mu_beta', mu=0., sd=100**2)
sigma_b = pm.Uniform('sigma_beta', lower=0, upper=100)
# Intercept for each county, distributed around group mean mu_a
a = pm.Normal('alpha', mu=mu_a, sd=sigma_a, shape=len(data.county.unique()))
# Intercept for each county, distributed around group mean mu_a
b = pm.Normal('beta', mu=mu_b, sd=sigma_b, shape=len(data.county.unique()))
# Model error
eps = pm.Uniform('eps', lower=0, upper=100)
# Expected value
radon_est = a[county_idx] + b[county_idx] * data.floor.values
# Data likelihood
y_like = pm.Normal('y_like', mu=radon_est, sd=eps, observed=data.log_radon)
```
Having specified the graphical model, we can run inference using the state of the art No-U-Turn Sampler (NUTS) initialized with ADVI.
```
with hierarchical_model:
# Use ADVI for initialization
mu, sds, elbo = pm.variational.advi(n=100000)
step = pm.NUTS(scaling=hierarchical_model.dict_to_array(sds)**2, is_cov=True)
hierarchical_trace = pm.sample(5000, step, start=mu)
```
Let's examine the trace plots of all our latent variables in the hierarchical regression model:
```
pm.traceplot(hierarchical_trace[500:])
plt.show()
```
From the traceplots, we can see convergence in our posterior distributions for $\alpha_c$ and $\beta_c$ indicating different intercepts and slopes for different counties. In addition, we also recover the posterior distribution of the shared parameters. $\mu_{\alpha}$ tells us that the group mean of log radon levels is close to $1.5$, while $\mu_{\beta}$ tells us that the slope is negative with a mean of $-0.65$ and thus having no basement decreases radon levels.
| github_jupyter |
# Think Bayes: Chapter 5
This notebook presents code and exercises from Think Bayes, second edition.
Copyright 2016 Allen B. Downey
MIT License: https://opensource.org/licenses/MIT
```
from __future__ import print_function, division
% matplotlib inline
import warnings
warnings.filterwarnings('ignore')
import numpy as np
from thinkbayes2 import Pmf, Cdf, Suite, Beta
import thinkplot
```
## Odds
The following function converts from probabilities to odds.
```
def Odds(p):
return p / (1-p)
```
And this function converts from odds to probabilities.
```
def Probability(o):
return o / (o+1)
```
If 20% of bettors think my horse will win, that corresponds to odds of 1:4, or 0.25.
```
p = 0.2
Odds(p)
```
If the odds against my horse are 1:5, that corresponds to a probability of 1/6.
```
o = 1/5
Probability(o)
```
We can use the odds form of Bayes's theorem to solve the cookie problem:
```
prior_odds = 1
likelihood_ratio = 0.75 / 0.5
post_odds = prior_odds * likelihood_ratio
post_odds
```
And then we can compute the posterior probability, if desired.
```
post_prob = Probability(post_odds)
post_prob
```
If we draw another cookie and it's chocolate, we can do another update:
```
likelihood_ratio = 0.25 / 0.5
post_odds *= likelihood_ratio
post_odds
```
And convert back to probability.
```
post_prob = Probability(post_odds)
post_prob
```
## Oliver's blood
The likelihood ratio is also useful for talking about the strength of evidence without getting bogged down talking about priors.
As an example, we'll solve this problem from MacKay's {\it Information Theory, Inference, and Learning Algorithms}:
> Two people have left traces of their own blood at the scene of a crime. A suspect, Oliver, is tested and found to have type 'O' blood. The blood groups of the two traces are found to be of type 'O' (a common type in the local population, having frequency 60) and of type 'AB' (a rare type, with frequency 1). Do these data [the traces found at the scene] give evidence in favor of the proposition that Oliver was one of the people [who left blood at the scene]?
If Oliver is
one of the people who left blood at the crime scene, then he
accounts for the 'O' sample, so the probability of the data
is just the probability that a random member of the population
has type 'AB' blood, which is 1%.
If Oliver did not leave blood at the scene, then we have two
samples to account for. If we choose two random people from
the population, what is the chance of finding one with type 'O'
and one with type 'AB'? Well, there are two ways it might happen:
the first person we choose might have type 'O' and the second
'AB', or the other way around. So the total probability is
$2 (0.6) (0.01) = 1.2$%.
So the likelihood ratio is:
```
like1 = 0.01
like2 = 2 * 0.6 * 0.01
likelihood_ratio = like1 / like2
likelihood_ratio
```
Since the ratio is less than 1, it is evidence *against* the hypothesis that Oliver left blood at the scence.
But it is weak evidence. For example, if the prior odds were 1 (that is, 50% probability), the posterior odds would be 0.83, which corresponds to a probability of:
```
post_odds = 1 * like1 / like2
Probability(post_odds)
```
So this evidence doesn't "move the needle" very much.
**Exercise:** Suppose other evidence had made you 90% confident of Oliver's guilt. How much would this exculpatory evince change your beliefs? What if you initially thought there was only a 10% chance of his guilt?
Notice that evidence with the same strength has a different effect on probability, depending on where you started.
```
# Solution
post_odds = Odds(0.9) * like1 / like2
Probability(post_odds)
# Solution
post_odds = Odds(0.1) * like1 / like2
Probability(post_odds)
```
## Comparing distributions
Let's get back to the Kim Rhode problem from Chapter 4:
> At the 2016 Summer Olympics in the Women's Skeet event, Kim Rhode faced Wei Meng in the bronze medal match. They each hit 15 of 25 targets, sending the match into sudden death. In the first round, both hit 1 of 2 targets. In the next two rounds, they each hit 2 targets. Finally, in the fourth round, Rhode hit 2 and Wei hit 1, so Rhode won the bronze medal, making her the first Summer Olympian to win an individual medal at six consecutive summer games.
>But after all that shooting, what is the probability that Rhode is actually a better shooter than Wei? If the same match were held again, what is the probability that Rhode would win?
I'll start with a uniform distribution for `x`, the probability of hitting a target, but we should check whether the results are sensitive to that choice.
First I create a Beta distribution for each of the competitors, and update it with the results.
```
rhode = Beta(1, 1, label='Rhode')
rhode.Update((22, 11))
wei = Beta(1, 1, label='Wei')
wei.Update((21, 12))
```
Based on the data, the distribution for Rhode is slightly farther right than the distribution for Wei, but there is a lot of overlap.
```
thinkplot.Pdf(rhode.MakePmf())
thinkplot.Pdf(wei.MakePmf())
thinkplot.Config(xlabel='x', ylabel='Probability')
```
To compute the probability that Rhode actually has a higher value of `p`, there are two options:
1. Sampling: we could draw random samples from the posterior distributions and compare them.
2. Enumeration: we could enumerate all possible pairs of values and add up the "probability of superiority".
I'll start with sampling. The Beta object provides a method that draws a random value from a Beta distribution:
```
iters = 1000
count = 0
for _ in range(iters):
x1 = rhode.Random()
x2 = wei.Random()
if x1 > x2:
count += 1
count / iters
```
`Beta` also provides `Sample`, which returns a NumPy array, so we an perform the comparisons using array operations:
```
rhode_sample = rhode.Sample(iters)
wei_sample = wei.Sample(iters)
np.mean(rhode_sample > wei_sample)
```
The other option is to make `Pmf` objects that approximate the Beta distributions, and enumerate pairs of values:
```
def ProbGreater(pmf1, pmf2):
total = 0
for x1, prob1 in pmf1.Items():
for x2, prob2 in pmf2.Items():
if x1 > x2:
total += prob1 * prob2
return total
pmf1 = rhode.MakePmf(1001)
pmf2 = wei.MakePmf(1001)
ProbGreater(pmf1, pmf2)
pmf1.ProbGreater(pmf2)
pmf1.ProbLess(pmf2)
```
**Exercise:** Run this analysis again with a different prior and see how much effect it has on the results.
## Simulation
To make predictions about a rematch, we have two options again:
1. Sampling. For each simulated match, we draw a random value of `x` for each contestant, then simulate 25 shots and count hits.
2. Computing a mixture. If we knew `x` exactly, the distribution of hits, `k`, would be binomial. Since we don't know `x`, the distribution of `k` is a mixture of binomials with different values of `x`.
I'll do it by sampling first.
```
import random
def flip(p):
return random.random() < p
```
`flip` returns True with probability `p` and False with probability `1-p`
Now we can simulate 1000 rematches and count wins and losses.
```
iters = 1000
wins = 0
losses = 0
for _ in range(iters):
x1 = rhode.Random()
x2 = wei.Random()
count1 = count2 = 0
for _ in range(25):
if flip(x1):
count1 += 1
if flip(x2):
count2 += 1
if count1 > count2:
wins += 1
if count1 < count2:
losses += 1
wins/iters, losses/iters
```
Or, realizing that the distribution of `k` is binomial, we can simplify the code using NumPy:
```
rhode_rematch = np.random.binomial(25, rhode_sample)
thinkplot.Hist(Pmf(rhode_rematch))
wei_rematch = np.random.binomial(25, wei_sample)
np.mean(rhode_rematch > wei_rematch)
np.mean(rhode_rematch < wei_rematch)
```
Alternatively, we can make a mixture that represents the distribution of `k`, taking into account our uncertainty about `x`:
```
from thinkbayes2 import MakeBinomialPmf
def MakeBinomialMix(pmf, label=''):
mix = Pmf(label=label)
for x, prob in pmf.Items():
binom = MakeBinomialPmf(n=25, p=x)
for k, p in binom.Items():
mix[k] += prob * p
return mix
rhode_rematch = MakeBinomialMix(rhode.MakePmf(), label='Rhode')
wei_rematch = MakeBinomialMix(wei.MakePmf(), label='Wei')
thinkplot.Pdf(rhode_rematch)
thinkplot.Pdf(wei_rematch)
thinkplot.Config(xlabel='hits')
rhode_rematch.ProbGreater(wei_rematch), rhode_rematch.ProbLess(wei_rematch)
```
Alternatively, we could use MakeMixture:
```
from thinkbayes2 import MakeMixture
def MakeBinomialMix2(pmf):
binomials = Pmf()
for x, prob in pmf.Items():
binom = MakeBinomialPmf(n=25, p=x)
binomials[binom] = prob
return MakeMixture(binomials)
```
Here's how we use it.
```
rhode_rematch = MakeBinomialMix2(rhode.MakePmf())
wei_rematch = MakeBinomialMix2(wei.MakePmf())
rhode_rematch.ProbGreater(wei_rematch), rhode_rematch.ProbLess(wei_rematch)
```
**Exercise:** Run this analysis again with a different prior and see how much effect it has on the results.
## Distributions of sums and differences
Suppose we want to know the total number of targets the two contestants will hit in a rematch. There are two ways we might compute the distribution of this sum:
1. Sampling: We can draw samples from the distributions and add them up.
2. Enumeration: We can enumerate all possible pairs of values.
I'll start with sampling:
```
iters = 1000
pmf = Pmf()
for _ in range(iters):
k = rhode_rematch.Random() + wei_rematch.Random()
pmf[k] += 1
pmf.Normalize()
thinkplot.Hist(pmf)
```
Or we could use `Sample` and NumPy:
```
ks = rhode_rematch.Sample(iters) + wei_rematch.Sample(iters)
pmf = Pmf(ks)
thinkplot.Hist(pmf)
```
Alternatively, we could compute the distribution of the sum by enumeration:
```
def AddPmfs(pmf1, pmf2):
pmf = Pmf()
for v1, p1 in pmf1.Items():
for v2, p2 in pmf2.Items():
pmf[v1 + v2] += p1 * p2
return pmf
```
Here's how it's used:
```
pmf = AddPmfs(rhode_rematch, wei_rematch)
thinkplot.Pdf(pmf)
```
The `Pmf` class provides a `+` operator that does the same thing.
```
pmf = rhode_rematch + wei_rematch
thinkplot.Pdf(pmf)
```
**Exercise:** The Pmf class also provides the `-` operator, which computes the distribution of the difference in values from two distributions. Use the distributions from the previous section to compute the distribution of the differential between Rhode and Wei in a rematch. On average, how many clays should we expect Rhode to win by? What is the probability that Rhode wins by 10 or more?
```
# Solution
pmf = rhode_rematch - wei_rematch
thinkplot.Pdf(pmf)
# Solution
# On average, we expect Rhode to win by about 1 clay.
pmf.Mean(), pmf.Median(), pmf.Mode()
# Solution
# But there is, according to this model, a 2% chance that she could win by 10.
sum([p for (x, p) in pmf.Items() if x >= 10])
```
## Distribution of maximum
Suppose Kim Rhode continues to compete in six more Olympics. What should we expect her best result to be?
Once again, there are two ways we can compute the distribution of the maximum:
1. Sampling.
2. Analysis of the CDF.
Here's a simple version by sampling:
```
iters = 1000
pmf = Pmf()
for _ in range(iters):
ks = rhode_rematch.Sample(6)
pmf[max(ks)] += 1
pmf.Normalize()
thinkplot.Hist(pmf)
```
And here's a version using NumPy. I'll generate an array with 6 rows and 10 columns:
```
iters = 1000
ks = rhode_rematch.Sample((6, iters))
ks
```
Compute the maximum in each column:
```
maxes = np.max(ks, axis=0)
maxes[:10]
```
And then plot the distribution of maximums:
```
pmf = Pmf(maxes)
thinkplot.Hist(pmf)
```
Or we can figure it out analytically. If the maximum is less-than-or-equal-to some value `k`, all 6 random selections must be less-than-or-equal-to `k`, so:
$ CDF_{max}(x) = CDF(x)^6 $
`Pmf` provides a method that computes and returns this `Cdf`, so we can compute the distribution of the maximum like this:
```
pmf = rhode_rematch.Max(6).MakePmf()
thinkplot.Hist(pmf)
```
**Exercise:** Here's how Pmf.Max works:
def Max(self, k):
"""Computes the CDF of the maximum of k selections from this dist.
k: int
returns: new Cdf
"""
cdf = self.MakeCdf()
cdf.ps **= k
return cdf
Write a function that takes a Pmf and an integer `n` and returns a Pmf that represents the distribution of the minimum of `k` values drawn from the given Pmf. Use your function to compute the distribution of the minimum score Kim Rhode would be expected to shoot in six competitions.
```
def Min(pmf, k):
cdf = pmf.MakeCdf()
cdf.ps = 1 - (1-cdf.ps)**k
return cdf
pmf = Min(rhode_rematch, 6).MakePmf()
thinkplot.Hist(pmf)
```
## Exercises
**Exercise:** Suppose you are having a dinner party with 10 guests and 4 of them are allergic to cats. Because you have cats, you expect 50% of the allergic guests to sneeze during dinner. At the same time, you expect 10% of the non-allergic guests to sneeze. What is the distribution of the total number of guests who sneeze?
```
# Solution
n_allergic = 4
n_non = 6
p_allergic = 0.5
p_non = 0.1
pmf = MakeBinomialPmf(n_allergic, p_allergic) + MakeBinomialPmf(n_non, p_non)
thinkplot.Hist(pmf)
# Solution
pmf.Mean()
```
**Exercise** [This study from 2015](http://onlinelibrary.wiley.com/doi/10.1111/apt.13372/full) showed that many subjects diagnosed with non-celiac gluten sensitivity (NCGS) were not able to distinguish gluten flour from non-gluten flour in a blind challenge.
Here is a description of the study:
>"We studied 35 non-CD subjects (31 females) that were on a gluten-free diet (GFD), in a double-blind challenge study. Participants were randomised to receive either gluten-containing flour or gluten-free flour for 10 days, followed by a 2-week washout period and were then crossed over. The main outcome measure was their ability to identify which flour contained gluten.
>"The gluten-containing flour was correctly identified by 12 participants (34%)..."
Since 12 out of 35 participants were able to identify the gluten flour, the authors conclude "Double-blind gluten challenge induces symptom recurrence in just one-third of patients fulfilling the clinical diagnostic criteria for non-coeliac gluten sensitivity."
This conclusion seems odd to me, because if none of the patients were sensitive to gluten, we would expect some of them to identify the gluten flour by chance. So the results are consistent with the hypothesis that none of the subjects are actually gluten sensitive.
We can use a Bayesian approach to interpret the results more precisely. But first we have to make some modeling decisions.
1. Of the 35 subjects, 12 identified the gluten flour based on resumption of symptoms while they were eating it. Another 17 subjects wrongly identified the gluten-free flour based on their symptoms, and 6 subjects were unable to distinguish. So each subject gave one of three responses. To keep things simple I follow the authors of the study and lump together the second two groups; that is, I consider two groups: those who identified the gluten flour and those who did not.
2. I assume (1) people who are actually gluten sensitive have a 95% chance of correctly identifying gluten flour under the challenge conditions, and (2) subjects who are not gluten sensitive have only a 40% chance of identifying the gluten flour by chance (and a 60% chance of either choosing the other flour or failing to distinguish).
Using this model, estimate the number of study participants who are sensitive to gluten. What is the most likely number? What is the 95% credible interval?
```
# Solution
# Here's a class that models the study
class Gluten(Suite):
def Likelihood(self, data, hypo):
"""Computes the probability of the data under the hypothesis.
data: tuple of (number who identified, number who did not)
hypothesis: number of participants who are gluten sensitive
"""
# compute the number who are gluten sensitive, `gs`, and
# the number who are not, `ngs`
gs = hypo
yes, no = data
n = yes + no
ngs = n - gs
pmf1 = MakeBinomialPmf(gs, 0.95)
pmf2 = MakeBinomialPmf(ngs, 0.4)
pmf = pmf1 + pmf2
return pmf[yes]
# Solution
prior = Gluten(range(0, 35+1))
thinkplot.Pdf(prior)
# Solution
posterior = prior.Copy()
data = 12, 23
posterior.Update(data)
# Solution
thinkplot.Pdf(posterior)
thinkplot.Config(xlabel='# who are gluten sensitive',
ylabel='PMF', legend=False)
# Solution
posterior.CredibleInterval(95)
```
**Exercise** Coming soon: the space invaders problem.
```
# Solution
# Solution
# Solution
# Solution
# Solution
# Solution
# Solution
# Solution
```
| github_jupyter |
```
# Running %env without any arguments
# lists all environment variables
# The line below sets the environment
# variable CUDA_VISIBLE_DEVICES
%env CUDA_VISIBLE_DEVICES =
import numpy as np
import pandas as pd
import io
import time
from datetime import datetime
import bson # this is installed with the pymongo package
import matplotlib.pyplot as plt
from scipy.misc import imread, imsave, imshow
import tensorflow as tf
from tensorflow.python.platform import tf_logging
from tensorflow.contrib import layers
from tensorflow.contrib.training import add_gradients_summaries
from tensorflow.python.ops import math_ops
from tensorflow.python.framework import ops
from tensorflow.python.ops import array_ops
from tensorflow.contrib.layers.python.layers import layers as layers_lib
from tensorflow.python.ops import nn_ops
from tensorflow.python.ops import control_flow_ops
from tensorflow.python.training import optimizer as tf_optimizer
from tensorflow.python.ops import variables as tf_variables
import os.path
import tensorflow.contrib.slim as slim
import inception_preprocessing
import vgg_preprocessing
import logging
import resnet2
# This is a bit of magic to make matplotlib figures appear inline in the notebook
# rather than in a new window.
%matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# Some more magic so that the notebook will reload external python modules;
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
%load_ext autoreload
%autoreload 2
DATASET_PATH = '/media/rs/0E06CD1706CD0127/Kapok/kaggle/'
PRETRAINED_MODEL_PATH = DATASET_PATH + 'Resnet/logs_v2_101/model/resnet101_v2_model.ckpt-367651'
LOG_PATH = DATASET_PATH + 'Resnet/logs_v2_101/'
TRAIN_PATH = DATASET_PATH + 'Split1/Train/'
#TRAIN_PATH = '/media/rs/FC6CDC6F6CDC25E4/resample_dataset2/'
#TRAIN_PATH = '/media/rs/FC6CDC6F6CDC25E4/ResnetHardTrain/'
LR_FILE_PATH = DATASET_PATH + 'Resnet/logs_v2_101/lr_setting/resnetv2_vgg_lr_setting'
VAL_PATH = DATASET_PATH + 'Split1/Validation/'
TEST_PATH = DATASET_PATH + 'Test/'
CATEGORY_NAME_PATH = DATASET_PATH + 'category_names.csv'
CATEGORY_WEIGHT_PATH = DATASET_PATH + 'catogory_with_weight.csv'
BATCH_SIZE = 128#256
IMAGE_WIDTH = 180
IMAGE_HEIGHT = 180
NUM_CLASS = 5270
LEVEL0_CLASS = 49
LEVEL1_CLASS = 483
# validation examples num: 2319624
# train examples num: 10051704
# total step: 157057
TOTAL_EXAMPLES = 10051704
NUM_EPOCHES = 12
EPOCHES_OVER = 10
INPUT_THREADS = 12
initial_learning_rate = 0.0001
stop_learning_rate = 0.000001
moving_average_decay = 0.96# use large to be more stable?
momentum = 0.9
#Know the number steps to take before decaying the learning rate and batches per epoch
num_steps_per_epoch = TOTAL_EXAMPLES / BATCH_SIZE + 1
# get TF logger
log = logging.getLogger('tensorflow')
log.setLevel(logging.DEBUG)
# create formatter and add it to the handlers
formatter = logging.Formatter('%(asctime)s - %(name)s - %(levelname)s - %(message)s')
# create file handler which logs even debug messages
fh = logging.FileHandler(DATASET_PATH + 'tensorflow_resnet_train_vggpreprocess.log')
fh.setLevel(logging.DEBUG)
fh.setFormatter(formatter)
log.addHandler(fh)
def read_learning_rate(cur_step, num_steps_per_epoch):
def inner_lr_parser(interval_start, interval_end, lr, dict_in, default_lr, use_epoch_percent, num_steps_per_epoch):
lr = default_lr * lr
if use_epoch_percent:
interval_start = num_steps_per_epoch * interval_start
interval_end = num_steps_per_epoch * interval_end
interval_start = int(interval_start)
interval_end = int(interval_end)
if (interval_start < interval_end) and (lr > 0):
dict_in[(interval_start, interval_end)] = lr
lr_map = dict()
default_lr = initial_learning_rate
stop_lr = stop_learning_rate
line_index = -1
use_epoch_percent = True
if os.path.exists(LR_FILE_PATH):
with open(LR_FILE_PATH, 'r') as lr_setting_file:
for _, line in enumerate(lr_setting_file):
line = line.strip()
if (line != '') and (not line.startswith('#')):
line_index += 1
if line_index == 0:
default_lr = float(line.split(':')[-1].strip())
continue
if line_index == 1:
stop_lr = float(line.split(':')[-1].strip())
continue
if line_index == 2:
use_epoch_percent = ('EPOCHES_PERCENT' in (line.split(':')[-1].strip()))
continue
# this is a list desciption
if line.startswith('['):
line = [float(s.strip()) for s in line[1:-1].strip().split()]
step_interval = (line[1] - line[0])/line[-1]
lr_interval = (line[3] - line[2])/line[-1]
begin = line[0]
lr_begin = line[2]
for index in range(int(line[-1])):
inner_lr_parser(begin, begin+step_interval, lr_begin, lr_map, default_lr, use_epoch_percent, num_steps_per_epoch)
begin += step_interval
lr_begin += lr_interval
else:
interval_start, interval_end, lr = [float(s) for s in line.strip().split()]
inner_lr_parser(interval_start, interval_end, lr, lr_map, default_lr, use_epoch_percent, num_steps_per_epoch)
lr_ret = default_lr
# print(use_epoch_percent)
for (start, end), lr in lr_map.items():
if (cur_step >= start) and (cur_step <= end):
if (lr < lr_ret):
lr_ret = lr
if lr_ret < stop_lr: lr_ret = stop_lr
return lr_ret
# _ = read_learning_rate(1, num_steps_per_epoch)
# lr = []
# num_epoches_to_show = 10
# num_point = 100
# for i in [i*num_epoches_to_show*num_steps_per_epoch/num_point for i in range(num_point)]:
# lr.append(read_learning_rate(i, num_steps_per_epoch))
# plt.plot(lr)
# plt.ylabel('learning rate')
# plt.show()
def preprocess_for_inception(input_image, is_training = True):
return vgg_preprocessing.preprocess_image(input_image, 180, 180, is_training)
class LabelMapping(object):
def __init__(self, catogory_file_path):
super(LabelMapping, self).__init__()
self._category_level_csv = catogory_file_path
self._category_map, self._category_level0_map, self._category_level1_map, self._len_level0, self._len_level1 = self.cvt_csv2tfrecord()
self._catogory_weight_map = self.cvt_catogory_weight()
self._mapping_strings = tf.constant( [ str(key) for key in self._category_map.keys() ] )
self._mapping_table = tf.contrib.lookup.index_table_from_tensor(mapping=self._mapping_strings, default_value=0)
self._level0_table = tf.contrib.lookup.HashTable(tf.contrib.lookup.KeyValueTensorInitializer(list(self._category_level0_map.keys()), list(self._category_level0_map.values()), tf.int64, tf.int64), 0)
self._level1_table = tf.contrib.lookup.HashTable(tf.contrib.lookup.KeyValueTensorInitializer(list(self._category_level1_map.keys()), list(self._category_level1_map.values()), tf.int64, tf.int64), 0)
self._weight_table = tf.contrib.lookup.HashTable(tf.contrib.lookup.KeyValueTensorInitializer(list(self._catogory_weight_map.keys()), list(self._catogory_weight_map.values()), tf.int64, tf.float32), 0)
@property
def category_map(self):
return self._category_map
@property
def level0_table(self):
return self._level0_table
@property
def level1_table(self):
return self._level1_table
@property
def len_level0(self):
return self._len_level0
@property
def len_level1(self):
return self._len_level1
@property
def mapping_table(self):
return self._mapping_table
@property
def weight_table(self):
return self._weight_table
def cvt_catogory_weight(self):
category_weight_map = dict()
csv = pd.read_csv(CATEGORY_WEIGHT_PATH).values
for row in csv:
category_id, weight = row[0], row[2]
category_weight_map[int(category_id)] = weight
return category_weight_map
def cvt_csv2tfrecord(self):
level0_map, level1_map = self.create_level_map()
count = 0
category_map = dict()
category_level0_map = dict()
category_level1_map = dict()
csv = pd.read_csv(self._category_level_csv).values
for row in csv:
category_id, level0, level1 = row[0], row[1], row[2]
category_map[category_id] = count
category_level0_map[int(category_id)] = level0_map[level0]
category_level1_map[int(category_id)] = level1_map[level1]
count += 1
return category_map, category_level0_map, category_level1_map, len(level0_map), len(level1_map)
def create_level_map(self):
csv = pd.read_csv(self._category_level_csv).values
level_list = [list(), list()]
for row in csv:
for level in range(1,3):
if row[level] not in level_list[level-1]:
level_list[level-1].append(row[level])
return dict(zip(level_list[0], range(len(level_list[0])))), dict(zip(level_list[1], range(len(level_list[1]))))
class CdiscountDataset(object):
def __init__(self, data_path, file_begin_match, label_mapping, num_examples, num_classes, buffer_size, batch_size, num_epochs, is_training):
super(CdiscountDataset, self).__init__()
#self._data_file_list = [ os.path.join(data_path, x) for x in os.listdir(data_path) if lambda x: os.path.isfile(x) and x.startswith(file_begin_match) ]
self._data_file_list = data_path + file_begin_match + '*'
self._num_examples = num_examples
self._num_classes = num_classes
self._batch_size = batch_size
self._buffer_size = buffer_size
self._num_epochs = num_epochs
self._is_training = is_training
self._category_map = label_mapping.category_map
self._level0_table = label_mapping.level0_table
self._level1_table = label_mapping.level1_table
self._len_level0 = label_mapping.len_level0
self._len_level1 = label_mapping.len_level1
self._mapping_table = label_mapping.mapping_table
self._weight_table = label_mapping.weight_table
def create_dataset(self):
opts = tf.python_io.TFRecordOptions(tf.python_io.TFRecordCompressionType.ZLIB)
reader = lambda : tf.TFRecordReader(options=opts)
keys_to_features = {
'img_raw': tf.FixedLenFeature([], tf.string, default_value=''),
'product_id': tf.FixedLenFeature([], tf.int64, default_value=tf.zeros([], dtype=tf.int64)),
# notice that we don't have this feature in our TFRecord, so always default provided
'format': tf.FixedLenFeature([], tf.string, default_value='jpg'),
'category_id': tf.FixedLenFeature([], tf.int64, default_value=tf.zeros([], dtype=tf.int64))
}
items_to_handlers = {
# automated decode image from features in FixedLenFeature
'image': slim.tfexample_decoder.Image(image_key='img_raw', format_key='format'),
'label': slim.tfexample_decoder.Tensor('category_id'),
}
decoder = slim.tfexample_decoder.TFExampleDecoder(keys_to_features, items_to_handlers)
self._dataset = slim.dataset.Dataset(
data_sources = self._data_file_list,
decoder = decoder,
reader = reader,
# num_readers = 8,
num_samples = self._num_examples,
#num_classes = self._num_classes,
items_to_descriptions = None)
# notice that DatasetDataProvider can automate shuffle the examples by ParallelReader using its RandomShuffleQueue
self._data_provider = slim.dataset_data_provider.DatasetDataProvider(
self._dataset,
num_readers = INPUT_THREADS,
shuffle = True, # default is True
num_epochs = self._num_epochs,
common_queue_capacity = self._buffer_size + 4 * self._batch_size,
common_queue_min = self._buffer_size,
scope = self._is_training and 'train_files' or 'validation_files')
org_image, org_label = self._data_provider.get(['image', 'label'])
image = preprocess_for_inception(org_image, self._is_training) # final image to train
# no need for shuffle, DatasetDataProvider do this for us
batch_images, batch_labels, batch_labels_level0, batch_labels_level1, batch_weight = \
tf.train.batch([image, tf.one_hot(self._mapping_table.lookup(tf.as_string(org_label)), self._num_classes, axis=-1),\
tf.one_hot(self._level0_table.lookup(org_label), self._len_level0, axis=-1),\
tf.one_hot(self._level1_table.lookup(org_label), self._len_level1, axis=-1), self._weight_table.lookup(org_label)],\
self._batch_size,\
num_threads = INPUT_THREADS,\
capacity = self._buffer_size + 4 * self._batch_size,\
allow_smaller_final_batch = self._is_training, name = self._is_training and 'train_batch' or 'validation_batch')
return batch_images, batch_labels, batch_labels_level0, batch_labels_level1, batch_weight
def_graph = tf.Graph()
with def_graph.as_default() as graph:
def train_step(input_examples, one_hot_labels, level0_labels, level1_labels, batch_weight):
# inputs has shape [batch, 224, 224, 3]
with slim.arg_scope(resnet2.resnet_arg_scope()):
resnet2_logits, end_points = resnet2.resnet_v2_101(input_examples, None, is_training=True)
logits = tf.stop_gradient(resnet2_logits)
net = layers_lib.dropout(logits, keep_prob=0.5, is_training=True, scope='Dropout')
net = layers_lib.conv2d(
net,
NUM_CLASS, [1, 1],
activation_fn=None,
normalizer_fn=None,
scope='logits')
end_points['predictions'] = layers.softmax(net, scope='predictions')
tvars = tf.trainable_variables()
head_only_vars = [var for var in tvars if 'logits' in var.name]
variables_to_restore = slim.get_variables_to_restore(exclude = ['logits'])
end_points['logits_output_squeezed'] = tf.squeeze(net)
loss = tf.losses.softmax_cross_entropy(onehot_labels = one_hot_labels, logits = end_points['logits_output_squeezed'], weights=batch_weight, label_smoothing = 0.0)
total_loss = tf.losses.get_total_loss() # obtain the regularization losses as well
# Create the global step for monitoring the learning_rate and training.
# since supervisor will also create one global_step, so we create n advance in order to feed into exponential_decay
global_step = tf.train.get_or_create_global_step(graph = graph)
custom_learning_rate = tf.placeholder(tf.float32, shape=[])
#Now we can define the optimizer that takes on the learning rate
#optimizer = tf.train.AdamOptimizer(learning_rate = lr)
#optimizer = tf.train.RMSPropOptimizer(learning_rate = lr)
optimizer = tf.train.MomentumOptimizer(learning_rate = custom_learning_rate, momentum=momentum)
# Gather update_ops from the first clone. These contain, for example,
# the updates for the batch_norm variables created by network_fn.
# update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
moving_average_variables = slim.get_model_variables()
variable_averages = tf.train.ExponentialMovingAverage(moving_average_decay, global_step)
# Use an alternative set of update ops in addition to the default updates:
tf.add_to_collection(tf.GraphKeys.UPDATE_OPS, variable_averages.apply(moving_average_variables))
#variables_to_restore_checkpoint = (ops.get_collection(ops.GraphKeys.GLOBAL_VARIABLES) + ops.get_collection(ops.GraphKeys.SAVEABLE_OBJECTS))
#Create the train_op.
#accumulate_factor = tf.constant([1./ACCUMULATE_STEP])
#train_op, accum_ops, zero_ops = my_create_train_op(total_loss, optimizer, False, accumulate_factor)
#Create the train_op.
train_op = slim.learning.create_train_op(total_loss, optimizer, summarize_gradients=False, variables_to_train=head_only_vars)
variables_to_restore_checkpoint = slim.get_variables_to_restore()
#State the metrics that you want to predict. We get a predictions that is not one_hot_encoded.
predictions = tf.argmax(tf.squeeze(end_points['predictions']), 1)
probabilities = end_points['predictions']
accuracy, accuracy_update = tf.contrib.metrics.streaming_accuracy(predictions, tf.argmax(one_hot_labels, 1), name='train_accuracy')
metrics_op = tf.group(accuracy_update)
real_time_accuracy = tf.reduce_mean(tf.cast(tf.equal(predictions, tf.argmax(one_hot_labels, 1)), tf.float32))
#Now finally create all the summaries you need to monitor and group them into one summary op.
tf.summary.scalar('losses/Total_Loss', total_loss)
tf.summary.scalar('train/accuracy', accuracy)
tf.summary.scalar('train/real_time_accuracy', real_time_accuracy)
tf.summary.scalar('learning_rate', custom_learning_rate)
return train_op, global_step, metrics_op, variables_to_restore, variables_to_restore_checkpoint, predictions, custom_learning_rate, accuracy, real_time_accuracy, total_loss
with def_graph.as_default() as graph:
label_mapping = LabelMapping(CATEGORY_NAME_PATH)
train_dataset = CdiscountDataset(TRAIN_PATH, 'output_file', label_mapping, TOTAL_EXAMPLES, NUM_CLASS, 12000, BATCH_SIZE, NUM_EPOCHES, True)
batch_images, batch_labels, batch_level0_labels, batch_level1_labels, batch_weight = train_dataset.create_dataset()
with tf.device('/gpu:0'):
train_op, global_step, metrics_op, variables_to_restore, variables_to_restore_checkpoint, pred_op, lr, accuracy, real_time_accuracy, total_loss = train_step(batch_images, batch_labels, batch_level0_labels, batch_level1_labels, batch_weight)
summary_op = tf.summary.merge_all()
checkpoint_saver = tf.train.Saver(variables_to_restore_checkpoint)
pre_train_saver = tf.train.Saver(variables_to_restore)
#pre_train_saver = tf.train.Saver(variables_to_restore)
# Define an init function that loads the pretrained checkpoint.
# sess is the managed session passed by Supervisor
def load_pretrain(sess):
pre_train_saver.restore(sess, PRETRAINED_MODEL_PATH)
# no need for specify local_variables_initializer and tables_initializer, Supervisor will do this via default local_init_op
# init_op = tf.group(tf.global_variables_initializer(), tf.local_variables_initializer(), tf.tables_initializer())
init_op = tf.group(tf.global_variables_initializer())
#init_op = tf.group(train_iterator_initializer, val_iterator_initializer, tf.global_variables_initializer())
# Pass the init function to the supervisor.
# - The init function is called _after_ the variables have been initialized by running the init_op.
# - use default tf.Saver() for ordinary save and restore
# - save checkpoint every 1.3 hours(4800)
# - manage summary in current process by ourselves for memory saving
# - no need to specify global_step, supervisor will find this automately
# - initialize order: checkpoint -> local_init_op -> init_op -> init_func
sv = tf.train.Supervisor(logdir=LOG_PATH, init_fn = load_pretrain, init_op = init_op, summary_op = None, saver = checkpoint_saver, save_model_secs=7200, checkpoint_basename='resnet101_v2_model.ckpt')
final_loss = 0.
final_accuracy = 0.
training_state = True
cur_readed_lr = initial_learning_rate
tf_logging.info(datetime.now().strftime('%Y-%m-%d %H:%M:%S'))
config = tf.ConfigProto(log_device_placement=True, allow_soft_placement=True)
#config.gpu_options.allow_growth = True
with sv.managed_session(config=config) as sess:
#with sv.prepare_or_wait_for_session(config=tf.ConfigProto(log_device_placement=True, allow_soft_placement=True)) as sess:
#sess.run(iterator_initalizer)
# Here sess was either initialized from the pre-trained-checkpoint or
# recovered from a checkpoint saved in a previous run of this code.
for step in range(int(num_steps_per_epoch * NUM_EPOCHES)):
if sv.should_stop():
tf_logging.info('Supervisor emit finished!')
tf_logging.info('Current Loss: %s', loss)
tf_logging.info('Current Accuracy: %s', accuracy)
tf_logging.info('Saving current model to disk(maybe invalid).')
training_state = False
break
start_time = time.time()
if step % 1000 == 0:
summ, cur_global_step = sess.run([summary_op, global_step], feed_dict={lr: cur_readed_lr})
sv.summary_computed(sess, summ)
if step > EPOCHES_OVER * num_steps_per_epoch:
raise StopIteration("over epoches reached.")
cur_readed_lr = read_learning_rate(cur_global_step, num_steps_per_epoch)
with tf.device('/gpu:0'):
_, _, cur_loss, cur_acc, rt_accuracy, total_step, cur_lr = sess.run([train_op, metrics_op, total_loss, accuracy, real_time_accuracy, global_step, lr], feed_dict={lr: cur_readed_lr})
time_elapsed = time.time() - start_time
if step % 10 == 0:
final_loss = cur_loss
final_accuracy = cur_acc
tf_logging.info('Current Speed: {:5.3f}sec/batch'.format(time_elapsed))
tf_logging.info('Current Streaming Accuracy: {:5.3f}%'.format(cur_acc*100.))
tf_logging.info('Current Realtime Accuracy: {:5.3f}%'.format(rt_accuracy*100.))
tf_logging.info('Current Loss: {:5.3f}'.format(cur_loss))
tf_logging.info('Epoch %s/%s, Global Step: %s', int(total_step / num_steps_per_epoch + 1), NUM_EPOCHES, total_step)
tf_logging.info('Current Learning Rate: {}'.format(cur_lr))
if training_state:
#We log the final training loss and accuracy
tf_logging.info('Final Loss: %s', final_loss)
tf_logging.info('Final Accuracy: %s', final_accuracy)
# Once all the training has been done, save the log files and checkpoint model
tf_logging.info('Finished training! Model saved.')
sv.saver.save(sess, sv.save_path, global_step = sv.global_step)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/MIT-LCP/hack-aotearoa/blob/master/04_timeseries.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# eICU Collaborative Research Database
# Notebook 4: Timeseries for a single patient
This notebook explores timeseries data for a single patient.
## Load libraries and connect to the database
```
# Import libraries
import numpy as np
import os
import pandas as pd
import matplotlib.pyplot as plt
# Make pandas dataframes prettier
from IPython.display import display, HTML
# Access data using Google BigQuery.
from google.colab import auth
from google.cloud import bigquery
# authenticate
auth.authenticate_user()
# Set up environment variables
project_id='hack-aotearoa'
os.environ["GOOGLE_CLOUD_PROJECT"]=project_id
```
## Selecting a single patient stay
### The patient table
The patient table includes general information about the patient admissions (for example, demographics, admission and discharge details). See: http://eicu-crd.mit.edu/eicutables/patient/
```
# Display the patient table
%%bigquery
SELECT *
FROM `physionet-data.eicu_crd_demo.patient`
patient.head()
```
### The `vitalperiodic` table
The `vitalperiodic` table comprises data that is consistently interfaced from bedside vital signs monitors into eCareManager. Data are generally interfaced as 1 minute averages, and archived into the `vitalperiodic` table as 5 minute median values. For more detail, see: http://eicu-crd.mit.edu/eicutables/vitalPeriodic/
```
# Get periodic vital signs for a single patient stay
%%bigquery vitalperiodic
SELECT *
FROM `physionet-data.eicu_crd_demo.vitalperiodic`
WHERE patientunitstayid = 210014
vitalperiodic.head()
# sort the values by the observationoffset (time in minutes from ICU admission)
vitalperiodic = vitalperiodic.sort_values(by='observationoffset')
vitalperiodic.head()
# subselect the variable columns
columns = ['observationoffset','temperature','sao2','heartrate','respiration',
'cvp','etco2','systemicsystolic','systemicdiastolic','systemicmean',
'pasystolic','padiastolic','pamean','icp']
vitalperiodic = vitalperiodic[columns].set_index('observationoffset')
vitalperiodic.head()
# plot the data
plt.rcParams['figure.figsize'] = [12,8]
title = 'Vital signs (periodic) for patientunitstayid = {} \n'.format(patientunitstayid)
ax = vitalperiodic.plot(title=title, marker='o')
ax.legend(loc='center left', bbox_to_anchor=(1.0, 0.5))
ax.set_xlabel("Minutes after admission to the ICU")
ax.set_ylabel("Absolute value")
```
## Questions
- Which variables are available for this patient?
- What is the peak heart rate during the period?
### The vitalaperiodic table
The vitalAperiodic table provides invasive vital sign data that is recorded at irregular intervals. See: http://eicu-crd.mit.edu/eicutables/vitalAperiodic/
```
# Get aperiodic vital signs
%%bigquery vitalaperiodic
SELECT *
FROM `physionet-data.eicu_crd_demo.vitalaperiodic`
WHERE patientunitstayid = 210014
# display the first few rows of the dataframe
vitalaperiodic.head()
# sort the values by the observationoffset (time in minutes from ICU admission)
vitalaperiodic = vitalaperiodic.sort_values(by='observationoffset')
vitalaperiodic.head()
# subselect the variable columns
columns = ['observationoffset','noninvasivesystolic','noninvasivediastolic',
'noninvasivemean','paop','cardiacoutput','cardiacinput','svr',
'svri','pvr','pvri']
vitalaperiodic = vitalaperiodic[columns].set_index('observationoffset')
vitalaperiodic.head()
# plot the data
plt.rcParams['figure.figsize'] = [12,8]
title = 'Vital signs (aperiodic) for patientunitstayid = {} \n'.format(patientunitstayid)
ax = vitalaperiodic.plot(title=title, marker='o')
ax.legend(loc='center left', bbox_to_anchor=(1.0, 0.5))
ax.set_xlabel("Minutes after admission to the ICU")
ax.set_ylabel("Absolute value")
```
## Questions
- What do the non-invasive variables measure?
- How do you think the mean is calculated?
## 3.4. The lab table
```
# Get labs
%%bigquery lab
SELECT *
FROM `physionet-data.eicu_crd_demo.lab`
WHERE patientunitstayid = 210014
lab.head()
# sort the values by the offset time (time in minutes from ICU admission)
lab = lab.sort_values(by='labresultoffset')
lab.head()
lab = lab.set_index('labresultoffset')
columns = ['labname','labresult','labmeasurenamesystem']
lab = lab[columns]
lab.head()
# list the distinct labnames
lab['labname'].unique()
# pivot the lab table to put variables into columns
lab = lab.pivot(columns='labname', values='labresult')
lab.head()
# plot laboratory tests of interest
labs_to_plot = ['creatinine','pH','BUN', 'glucose', 'potassium']
lab[labs_to_plot].head()
# plot the data
plt.rcParams['figure.figsize'] = [12,8]
title = 'Laboratory test results for patientunitstayid = {} \n'.format(patientunitstayid)
ax = lab[labs_to_plot].plot(title=title, marker='o',ms=10, lw=0)
ax.legend(loc='center left', bbox_to_anchor=(1.0, 0.5))
ax.set_xlabel("Minutes after admission to the ICU")
ax.set_ylabel("Absolute value")
```
| github_jupyter |
<a href="https://colab.research.google.com/github/sweetpand/Algorithms/blob/master/marathone_day6.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
1. Подсчитать, сколько было выделено памяти под переменные в ранее разработанных программах в рамках первых трех уроков.
Проанализировать результат и определить программы с наиболее эффективным использованием памяти.
Для анализа возьмите любые 1-3 ваших программы. Результаты анализа вставьте в виде комментариев к коду.
P.S. Напишите в комментариях версию Python и разрядность ОС.
```
import sys
from types import ModuleType
# import platform
# from distutils import util
# print(sys.version_info)
# sys.version_info(major=3, minor=7, micro=1, releaselevel='final', serial=0)
# python -V
# Python 3.6.5
# print(platform.python_version())
# 3.7.1
# print(sys.platform)
# win32
# print(util.get_platform())
# win-amd64
# print(platform.architecture())
# ('64bit', 'WindowsPE')
def show_size(x, level=0, size=0):
print('\t' * level, f'type = {type(x)}, size = {sys.getsizeof(x)}, object = {x}')
size += sys.getsizeof(x)
if hasattr(x, '__iter__'):
if hasattr(x, 'items'):
for key, value in x.items():
size += sys.getsizeof(key)
size += sys.getsizeof(value)
show_size(key, level + 1)
show_size(value, level + 1)
elif not isinstance(x, str):
for item in x:
show_size(item, level + 1)
size += sys.getsizeof(item)
return size
def my_vars(keys):
s = 0
for v in keys:
s +=show_size(globals()[v])
print(f'summ of memory in vars: {s}\n\n')
# print('Подсчет суммы чисел ряда 1 -0.5 0.25 -0.125 ...')
n = 50
summa = 1
x = 1
for i in range(0, n - 1):
x = x / -2
summa += x
#print(f'Результат работы программы Сумма: {summa}')
my_vars(['n','summa', 'x'])
"""
type = <class 'int'>, size = 28, object = 50
type = <class 'float'>, size = 24, object = 0.6666666666666661
type = <class 'float'>, size = 24, object = -1.7763568394002505e-15
summ of memory in vars: 76
"""
"""
1. В диапазоне натуральных чисел от 2 до 99 определить, сколько из них кратны любому из чисел в диапазоне от 2 до 9.
"""
multiple_count = {}
for i in range(2, 100):
for j in range(2, 10):
if (i % j == 0):
if j not in multiple_count:
multiple_count[j] = 1
else:
multiple_count[j] += 1
#print(multiple_count)
#for i in multiple_count:
#print(f'Чисел кратных {i} - {multiple_count[i]} шт')
my_vars(['multiple_count'])
"""
type = <class 'dict'>, size = 368, object = {2: 49, 3: 33, 4: 24, 5: 19, 6: 16, 7: 14, 8: 12, 9: 11}
type = <class 'int'>, size = 28, object = 2
type = <class 'int'>, size = 28, object = 49
type = <class 'int'>, size = 28, object = 3
type = <class 'int'>, size = 28, object = 33
type = <class 'int'>, size = 28, object = 4
type = <class 'int'>, size = 28, object = 24
type = <class 'int'>, size = 28, object = 5
type = <class 'int'>, size = 28, object = 19
type = <class 'int'>, size = 28, object = 6
type = <class 'int'>, size = 28, object = 16
type = <class 'int'>, size = 28, object = 7
type = <class 'int'>, size = 28, object = 14
type = <class 'int'>, size = 28, object = 8
type = <class 'int'>, size = 28, object = 12
type = <class 'int'>, size = 28, object = 9
type = <class 'int'>, size = 28, object = 11
summ of memory in vars: 816
"""
"""
6. В одномерном массиве найти сумму элементов, находящихся между минимальным и максимальным элементами. Сами минимальный и максимальный элементы в сумму не включать.
"""
import random
random_list = random.sample(range(0,100), 10)
numers = {
'max_value': random_list[0],
'min_value': random_list[0],
'min_key': 0,
'max_key': 0,
'sum': 0
}
#print(random_list)
for k,v in enumerate(random_list):
if v > numers['max_value']:
numers['max_key'] = k
numers['max_value'] = v
elif v < numers['min_value']:
numers['min_key'] = k
numers['min_value'] = v
if numers['min_key']+1 < numers['max_key']:
for i in random_list[numers['min_key']+1:numers['max_key']]:
numers['sum'] += i
elif numers['min_key'] > numers['max_key']+1:
for i in random_list[numers['max_key']+1:numers['min_key']]:
numers['sum'] += i
else:
pass
#print('Между минимальным и максимальным элементом нет чисел')
#print(f'Сумма {numers["sum"]}')
#print(numers)
my_vars(['numers', 'random_list'])
"""
type = <class 'dict'>, size = 240, object = {'max_value': 83, 'min_value': 8, 'min_key': 6, 'max_key': 7, 'sum': 0}
type = <class 'str'>, size = 58, object = max_value
type = <class 'int'>, size = 28, object = 83
type = <class 'str'>, size = 58, object = min_value
type = <class 'int'>, size = 28, object = 8
type = <class 'str'>, size = 56, object = min_key
type = <class 'int'>, size = 28, object = 6
type = <class 'str'>, size = 56, object = max_key
type = <class 'int'>, size = 28, object = 7
type = <class 'str'>, size = 52, object = sum
type = <class 'int'>, size = 24, object = 0
type = <class 'list'>, size = 144, object = [77, 56, 64, 79, 49, 36, 8, 83, 24, 57]
type = <class 'int'>, size = 28, object = 77
type = <class 'int'>, size = 28, object = 56
type = <class 'int'>, size = 28, object = 64
type = <class 'int'>, size = 28, object = 79
type = <class 'int'>, size = 28, object = 49
type = <class 'int'>, size = 28, object = 36
type = <class 'int'>, size = 28, object = 8
type = <class 'int'>, size = 28, object = 83
type = <class 'int'>, size = 28, object = 24
type = <class 'int'>, size = 28, object = 57
summ of memory in vars: 1080
"""
```
| github_jupyter |
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
```
# Create data
```
import copy
from liegroups import SE2, SO2
params_true = {'T_1_0': SE2.identity(),
'T_2_0': SE2(SO2.identity(), -np.array([0.5, 0])),
'T_3_0': SE2(SO2.identity(), -np.array([1, 0])),
'T_4_0': SE2(SO2.from_angle(np.pi / 2),
-(SO2.from_angle(np.pi / 2).dot(np.array([1, 0.5])))),
'T_5_0': SE2(SO2.from_angle(np.pi),
-(SO2.from_angle(np.pi).dot(np.array([0.5, 0.5])))),
'T_6_0': SE2(SO2.from_angle(-np.pi / 2),
-(SO2.from_angle(-np.pi / 2).dot(np.array([0.5, 0]))))}
obs = {'T_1_0': params_true['T_1_0'],
'T_2_1': params_true['T_2_0'].dot(params_true['T_1_0'].inv()),
'T_3_2': params_true['T_3_0'].dot(params_true['T_2_0'].inv()),
'T_4_3': params_true['T_4_0'].dot(params_true['T_3_0'].inv()),
'T_5_4': params_true['T_5_0'].dot(params_true['T_4_0'].inv()),
'T_6_5': params_true['T_6_0'].dot(params_true['T_5_0'].inv()),
'T_6_2': params_true['T_6_0'].dot(params_true['T_2_0'].inv())}
params_init = copy.deepcopy(params_true)
for key in params_init.keys():
params_init[key] = SE2.exp(5 * np.random.rand(3)).dot(params_init[key])
```
# Create residual functions
```
from pyslam.residuals import PoseResidual, PoseToPoseResidual
from pyslam.utils import invsqrt
prior_stiffness = invsqrt(1e-12 * np.identity(3))
odom_stiffness = invsqrt(1e-3 * np.identity(3))
loop_stiffness = invsqrt(1e-3 * np.identity(3))
residual0 = PoseResidual(obs['T_1_0'], prior_stiffness)
residual0_params = ['T_1_0']
residual1 = PoseToPoseResidual(obs['T_2_1'], odom_stiffness)
residual1_params = ['T_1_0', 'T_2_0']
residual2 = PoseToPoseResidual(obs['T_3_2'], odom_stiffness)
residual2_params = ['T_2_0', 'T_3_0']
residual3 = PoseToPoseResidual(obs['T_4_3'], odom_stiffness)
residual3_params = ['T_3_0', 'T_4_0']
residual4 = PoseToPoseResidual(obs['T_5_4'], odom_stiffness)
residual4_params = ['T_4_0', 'T_5_0']
residual5 = PoseToPoseResidual(obs['T_6_5'], odom_stiffness)
residual5_params = ['T_5_0', 'T_6_0']
residual6 = PoseToPoseResidual(obs['T_6_2'], loop_stiffness)
residual6_params = ['T_2_0', 'T_6_0']
```
# Set up and solve the problem
```
from pyslam.problem import Problem, Options
options = Options()
options.allow_nondecreasing_steps = True
options.max_nondecreasing_steps = 3
problem = Problem(options)
problem.add_residual_block(residual0, residual0_params)
problem.add_residual_block(residual1, residual1_params)
problem.add_residual_block(residual2, residual2_params)
problem.add_residual_block(residual3, residual3_params)
problem.add_residual_block(residual4, residual4_params)
problem.add_residual_block(residual5, residual5_params)
# problem.add_residual_block(residual6, residual6_params)
problem.initialize_params(params_init)
params_final = problem.solve()
print(problem.summary(format='full'))
```
# Check results
```
print("Initial Error:")
for key in params_true.keys():
print('{}: {}'.format(key, SE2.log(params_init[key].inv().dot(params_true[key]))))
print()
print("Final Error:")
for key in params_true.keys():
print('{}: {}'.format(key, SE2.log(params_final[key].inv().dot(params_true[key]))))
```
# Optional: Compute the covariance of the final parameter estimates
```
problem.compute_covariance()
print('covariance of T_5_0:\n{}'.format( problem.get_covariance_block('T_5_0','T_5_0') ))
```
| github_jupyter |
# TensorFlow tutorial
In this tutorial we'll show how to build deep learning models in Tribuo, using Tribuo's [TensorFlow](https://tensorflow.org) interface. Tribuo uses [TensorFlow-Java](https://github.com/tensorflow/java) which is build by the TensorFlow [SIG-JVM group](https://github.com/tensorflow/community/blob/master/sigs/jvm/CHARTER.md). Tribuo's development team are active participants in SIG-JVM, and we're trying to make TensorFlow work well for everyone on the Java platform, in addition to making it work well inside Tribuo.
Note that Tribuo's TensorFlow interface is not covered by the same stability guarantee as the rest of Tribuo. SIG-JVM has not released a 1.0 version of the TensorFlow Java API, and the API is currently in flux. When TensorFlow Java has API stability we'll be able to stabilize Tribuo's TensorFlow interface to provide the same guarantees as the rest of Tribuo.
We're going to train MLPs (Multi-Layer Perceptrons) for classification and regression, along with a CNN (Convolutional Neural Network) for classifying MNIST digits. We'll discuss loading in externally trained TensorFlow models and serving them alongside Tribuo's natively trained models. Finally we'll see how to export TensorFlow models trained in Tribuo into TensorFlow's SavedModelBundle format for interop with TensorFlow Serving and the rest of the TensorFlow ecosystem.
Unfortunately TensorFlow-Java has some non-determinism in it's gradient calculations which we're working on fixing in the TensorFlow-Java project, so repeated runs of this notebook will not produce identical answers, which unfortunately breaks some of Tribuo's provenance and reproducibility guarantees. When this is fixed upstream we'll apply any necessary fixes in Tribuo.
## Setup
You'll need to get a copy of the MNIST dataset in the original IDX format. You may have this from the configuration tutorial, in which case you can skip this step.
First the training data:
`wget http://yann.lecun.com/exdb/mnist/train-images-idx3-ubyte.gz http://yann.lecun.com/exdb/mnist/train-labels-idx1-ubyte.gz`
Then the test data:
`wget http://yann.lecun.com/exdb/mnist/t10k-images-idx3-ubyte.gz http://yann.lecun.com/exdb/mnist/t10k-labels-idx1-ubyte.gz`
Tribuo's IDX loader natively reads gzipped files so you don't need to unzip them.
We'll also need to download the winequality dataset from UCI. Again, if you've followed the regression tutorial you might already have this, so you can skip this step.
`wget https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-red.csv`
Next we'll load the Tribuo TensorFlow jar and import the packages we'll need for the rest of the tutorial.
```
%jars ./tribuo-tensorflow-4.2.0-SNAPSHOT-jar-with-dependencies.jar
import java.nio.file.Path;
import java.nio.file.Paths;
import org.tribuo.*;
import org.tribuo.data.csv.CSVLoader;
import org.tribuo.datasource.IDXDataSource;
import org.tribuo.evaluation.TrainTestSplitter;
import org.tribuo.classification.*;
import org.tribuo.classification.evaluation.*;
import org.tribuo.interop.tensorflow.*;
import org.tribuo.interop.tensorflow.example.*;
import org.tribuo.regression.*;
import org.tribuo.regression.evaluation.*;
import org.tribuo.util.Util;
import org.tensorflow.*;
import org.tensorflow.framework.initializers.*;
import org.tensorflow.ndarray.Shape;
import org.tensorflow.op.*;
import org.tensorflow.op.core.*;
import org.tensorflow.types.*;
```
## Loading the data
This is the same as the configuration and regression tutorials respectively, first we instantiate a `DataSource` for the particular dataset, then feed the data sources into datasets. We'll need to split the wine quality dataset into train & test as it doesn't have a predefined train/test split.
```
// First we load winequality
var regressionFactory = new RegressionFactory();
var regEval = new RegressionEvaluator();
var csvLoader = new CSVLoader<>(';',regressionFactory);
var wineSource = csvLoader.loadDataSource(Paths.get("winequality-red.csv"),"quality");
var wineSplitter = new TrainTestSplitter<>(wineSource, 0.7f, 0L);
var wineTrain = new MutableDataset<>(wineSplitter.getTrain());
var wineTest = new MutableDataset<>(wineSplitter.getTest());
// Now we load MNIST
var labelFactory = new LabelFactory();
var labelEval = new LabelEvaluator();
var mnistTrainSource = new IDXDataSource<>(Paths.get("train-images-idx3-ubyte.gz"),Paths.get("train-labels-idx1-ubyte.gz"),labelFactory);
var mnistTestSource = new IDXDataSource<>(Paths.get("t10k-images-idx3-ubyte.gz"),Paths.get("t10k-labels-idx1-ubyte.gz"),labelFactory);
var mnistTrain = new MutableDataset<>(mnistTrainSource);
var mnistTest = new MutableDataset<>(mnistTestSource);
```
## Defining a TensorFlow graph
Tribuo's TensorFlow API operates on TensorFlow graphs. You can construct those using TensorFlow's Java API, load in ones already generated by another TensorFlow API, or use one of Tribuo's example graph generators. We're going to define a simple MLP for the wine quality regression task in the notebook, but we'll use Tribuo's example graph generators for classifying MNIST (to make this tutorial a little shorter).
TensorFlow Java is working on a higher level layer wise API (similar to [Keras](https://www.tensorflow.org/api_docs/python/tf/keras)), but at the moment we have to define the graph using the low level ops. Once the layer API is available in TensorFlow Java, we'll add entry points so that those APIs can be used with Tribuo, making the next section of this tutorial a lot shorter. For the moment it'll be rather long, but hopefully it's not too hard to follow.
Tribuo's TensorFlow trainer will add the appropriate output node, loss function and gradient optimizer, so what you need to supply is the graph which emits the output (before any softmax, sigmoid or other output function), the name of the output op, the names of the input ops and the name of the graph initialization op.
## Building a regression model using an MLP
To solve this regression task we're going to build a 3 layer neural network, where each layer is a "dense" or "MLP" layer. We'll use a sigmoid as the activation function, but any supported one in TensorFlow will work. We'll need to know the number of input features and the number of output dimensions (i.e., the number of labels or regression dimensions), which is a little unfortunate as nothing else in Tribuo requires it, but it's required to build the structure.
```
var wineGraph = new Graph();
// This object is used to write operations into the graph
var wineOps = Ops.create(wineGraph);
var wineInputName = "WINE_INPUT";
long wineNumFeatures = wineTrain.getFeatureMap().size();
var wineInitializer = new Glorot<TFloat32>(wineOps,
// Initializer distribution
VarianceScaling.Distribution.TRUNCATED_NORMAL,
// Initializer seed
Trainer.DEFAULT_SEED
);
// The input placeholder that we'll feed the features into
var wineInput = wineOps.withName(wineInputName).placeholder(TFloat32.class,
Placeholder.shape(Shape.of(-1, wineNumFeatures)));
// Fully connected layer (numFeatures -> 30)
var fc1Weights = wineOps.variable(wineInitializer.call(wineOps.array(wineNumFeatures, 30L),
TFloat32.class));
var fc1Biases = wineOps.variable(wineOps.fill(wineOps.array(30), wineOps.constant(0.1f)));
var sigmoid1 = wineOps.math.sigmoid(wineOps.math.add(wineOps.linalg.matMul(wineInput, fc1Weights),
fc1Biases));
// Fully connected layer (30 -> 20)
var fc2Weights = wineOps.variable(wineInitializer.call(wineOps.array(30L, 20L),
TFloat32.class));
var fc2Biases = wineOps.variable(wineOps.fill(wineOps.array(20), wineOps.constant(0.1f)));
var sigmoid2 = wineOps.math.sigmoid(wineOps.math.add(wineOps.linalg.matMul(sigmoid1, fc2Weights),
fc2Biases));
// Output layer (20 -> 1)
var outputWeights = wineOps.variable(wineInitializer.call(wineOps.array(20L, 1L),
TFloat32.class));
var outputBiases = wineOps.variable(wineOps.fill(wineOps.array(1), wineOps.constant(0.1f)));
var outputOp = wineOps.math.add(wineOps.linalg.matMul(sigmoid2, outputWeights), outputBiases);
// Build the Graph initialization operation
var init = wineOps.init();
// Get the operation names to pass into the trainer
var wineOutputName = outputOp.op().name();
var wineInitName = init.op().name();
```
We can query the operation names by asking the various objects for their `name()`, which Tribuo will use to supply the appropriate inputs and outputs to the graph during training and inference.
Now we have the graph, input name, output name and init name, stored in `wineGraph`, `wineInputName`, `wineOutputName` and `wineInitName` respectively. Next we'll define the gradient optimization algorithm and it's hyperparameters. These are separate from Tribuo's built in gradient optimizers as they are part of the TensorFlow native library, but it turns out that most of the same algorithms are available. We're going to use [AdaGrad](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers/Adagrad), set it's learning rate to `0.1f` and the initial accumulator value to `0.01f`.
```
var gradAlgorithm = GradientOptimiser.ADAGRAD;
var gradParams = Map.of("learningRate",0.1f,"initialAccumulatorValue",0.01f);
```
We also need to create an object to convert from Tribuo's feature representation to a TensorFlow `Tensor`, and an object that can convert to and from `Tensor` and `Regressor`. These are defined using the `ExampleTransformer` and `OutputTransformer` interfaces.
### Converting Features into Tensors with FeatureConverter
Tribuo provides two implementations of `FeatureConverter`, one for dense inputs (like those used by MLPs) called `DenseFeatureConverter` and one for image shaped inputs (like those used by CNNs) called `ImageConverter`. If you need more specialised transformations (e.g., text) then you should implement the `FeatureConverter` interface and tailor it to your task's needs.
The `FeatureConverter` needs the name of the input placeholder which the features will be fed into, so it can produce the appropriate values in the Map that is fed into the TensorFlow graph.
### Converting Outputs into Tensors (and back again) with OutputConverter
There are implementations of `OutputConverter` for `Label`, `MultiLabel` and `Regressor`, as those cover the main use cases for TensorFlow. You are free to implement these interfaces for more specialised use cases, though they should be thread-safe and idempotent. The `OutputConverter` contains the loss function and output function which is used to attach the appropriate training hooks to the graph. `LabelConverter` uses the [softmax](https://en.wikipedia.org/wiki/Softmax_function) function to produce probabilistic outputs, and the [Categorical Cross Entropy](https://www.tensorflow.org/api_docs/python/tf/keras/losses/CategoricalCrossentropy) to provide the loss for back-propagation. `RegressorConverter` uses the identity function to produce the output (as it's already producing a real value), and the [Mean-Squared Error](https://www.tensorflow.org/api_docs/python/tf/keras/losses/MeanSquaredError) as the loss function. `MultiLabelConverter` uses an independent sigmoid function for each label as the output, thresholded at 0.5, and [Binary Cross Entropy](https://www.tensorflow.org/api_docs/python/tf/keras/losses/BinaryCrossentropy) as the loss function.
```
var wineDenseConverter = new DenseFeatureConverter(wineInputName);
var wineOutputConverter = new RegressorConverter();
```
We're finally ready to build our first `TensorFlowTrainer`. We need to specify a few more parameters in the constructor, namely the training batch size, the test batch size, and the number of training epochs. We'll set the batch sizes to 16 for all experiments, and we use 100 epochs for the regression task (because it's a small dataset), 20 epochs for the MNIST MLP, and 3 for the MNIST CNN (as the CNN converges much faster than the MLP).
```
var wineTrainer = new TensorFlowTrainer<Regressor>(wineGraph,
wineOutputName,
wineInitName,
gradAlgorithm,
gradParams,
wineDenseConverter,
wineOutputConverter,
16, // training batch size
100, // number of training epochs
16, // test batch size of the trained model
-1 // disable logging of the loss value
);
// Now we close the original graph to free the associated native resources.
// The TensorFlowTrainer keeps a copy of the GraphDef protobuf to rebuild it when necessary.
wineGraph.close();
```
`TensorFlowTrainer` will accept a `Graph`, a `GraphDef` protobuf, or a path to a `GraphDef` protobuf on disk. The `Graph` should be closed after it's supplied to the trainer, to free the native resources associated with it. Tribuo manages a copy of the `Graph` inside the trainer so users don't need to worry about resource allocation. The trainer automatically adds the loss function, gradient update operations and the final output operation to the supplied graph.
We can use this trainer the way we'd use any other Tribuo trainer, we call `trainer.train()` and pass it in a dataset. In the case of TensorFlow it will throw an IllegalArgumentException if the number of features or outputs in the training dataset doesn't match what the trainer is expecting, as those parameters are coupled to the graph structure.
```
var wineStart = System.currentTimeMillis();
var wineModel = wineTrainer.train(wineTrain);
var wineEnd = System.currentTimeMillis();
System.out.println("Wine quality training took " + Util.formatDuration(wineStart,wineEnd));
```
And we can evaluate it in the same way we evaluate other Tribuo regression models:
```
var wineEvaluation = regEval.evaluate(wineModel,wineTest);
var dimension = new Regressor("DIM-0",Double.NaN);
System.out.println(String.format("Wine quality evaluation:%n RMSE %f%n MAE %f%n R^2 %f%n",
wineEvaluation.rmse(dimension),
wineEvaluation.mae(dimension),
wineEvaluation.r2(dimension)));
```
We can see the MLP did ok there, and it's managed to fit the task almost as well as the tree ensemble we showed in the regression tutorial. With further tuning of the architecture and gradient parameters we could improve on this, but let's move on to classification.
## Building a classification model using an MLP
Building classification models using the TensorFlow interface is pretty similar to building regression models, thanks to Tribuo's common API for these tasks. The differences come in the choice of `OutputConverter`.
We're going to use Tribuo's `MLPExamples` and `CNNExamples` to build the networks for MNIST, as it's a bit shorter. These classes build simple predefined TensorFlow `Graph`s which are useful for demos, Tribuo's tests and getting started with deep learning. Currently there aren't many options in those classes, but we plan to expand them over time, and we welcome community contributions to do so. If you're interested in how the graphs are constructed you can check out the source for them on [GitHub](https://github.com/oracle/tribuo). For complex tasks we recommend that users build their own `Graph`s just as we did in the regression portion of the tutorial. TensorFlow-Java exposes a wide variety of [operations](https://tensorflow.org/jvm) for building graphs, and as the high level API improves it will become easier to specify complex structures.
Tribuo's graph building functions return a `GraphDefTuple`, which is a nominal tuple for a `GraphDef` along with the strings representing the necessary operation names. As Tribuo targets Java 8 and upwards it's not a `java.lang.Record`, but it will be one day.
```
var mnistInputName = "MNIST_INPUT";
var mnistMLPTuple = MLPExamples.buildMLPGraph(
mnistInputName, // The input placeholder name
mnistTrain.getFeatureMap().size(), // The number of input features
new int[]{300,200,30}, // The hidden layer sizes
mnistTrain.getOutputs().size() // The number of output labels
);
var mnistDenseConverter = new DenseFeatureConverter(mnistInputName);
var mnistOutputConverter = new LabelConverter();
```
This built an MLP with 3 hidden layers. The first maps from the feature space to an internal dimension of size 300, then the second is also of size 200, and the third has an internal dimension of 30. Tribuo then adds an output layer mapping down from those 30 dimensions to the 10 output dimensions in MNIST, one per digit.
We'll use the same gradient optimiser as before, along with the same hyperparameters.
```
var mnistMLPTrainer = new TensorFlowTrainer<Label>(mnistMLPTuple.graphDef,
mnistMLPTuple.outputName, // the name of the logit operation
mnistMLPTuple.initName, // the name of the initialisation operation
gradAlgorithm, // the gradient descent algorithm
gradParams, // the gradient descent hyperparameters
mnistDenseConverter, // the input feature converter
mnistOutputConverter, // the output label converter
16, // training batch size
20, // number of training epochs
16, // test batch size of the trained model
-1 // disable logging of the loss value
);
```
And we train the model as before:
```
var mlpStart = System.currentTimeMillis();
var mlpModel = mnistMLPTrainer.train(mnistTrain);
var mlpEnd = System.currentTimeMillis();
System.out.println("MNIST MLP training took " + Util.formatDuration(mlpStart,mlpEnd));
```
And evaluate it in the standard way:
```
var mlpEvaluation = labelEval.evaluate(mlpModel,mnistTest);
System.out.println(mlpEvaluation.toString());
System.out.println(mlpEvaluation.getConfusionMatrix().toString());
```
An MLP works pretty well on MNIST, but when working with images it's usually better to exploit the natural structure, and for that we use a Convolutional Neural Network.
## Training a Convolutional Neural Network
This is an even smaller transition than the switch between regression and classification. All we need to do is supply a `ImageConverter` which knows the size and pixel depth of the images, and build an appropriately shaped CNN.
We'll use `CNNExamples.buildLeNetGraph` to build a version of the venerable [LeNet 5](http://yann.lecun.com/exdb/lenet/) CNN. We specify the image shape (this method assumes images are square), the pixel depth and the number of outputs. So for MNIST that's 28 pixels across, a pixel depth of 255, and 10 output classes one per digit. We'll also need the appropriate `ImageConverter` which needs the name of the input placeholder, the width and height of the image (so allowing rectangular images), and the number of colour channels. MNIST is grayscale, so there's only a single colour channel.
```
var mnistCNNTuple = CNNExamples.buildLeNetGraph(mnistInputName,28,255,mnistTrain.getOutputs().size());
var mnistImageConverter = new ImageConverter(mnistInputName,28,28,1);
```
We can build the trainer and train in the same way as before, but we will train for fewer epochs as the CNN converges faster:
```
var mnistCNNTrainer = new TensorFlowTrainer<Label>(mnistCNNTuple.graphDef,
mnistCNNTuple.outputName, // the name of the logit operation
mnistCNNTuple.initName, // the name of the initialisation operation
gradAlgorithm, // the gradient descent algorithm
gradParams, // the gradient descent hyperparameters
mnistImageConverter, // the input feature converter
mnistOutputConverter, // the output label converter
16, // training batch size
3, // number of training epochs
16, // test batch size of the trained model
-1 // disable logging of the loss value
);
// Training the model
var cnnStart = System.currentTimeMillis();
var cnnModel = mnistCNNTrainer.train(mnistTrain);
var cnnEnd = System.currentTimeMillis();
System.out.println("MNIST CNN training took " + Util.formatDuration(cnnStart,cnnEnd));
```
And evaluate it the standard way:
```
var cnnPredictions = cnnModel.predict(mnistTest);
var cnnEvaluation = labelEval.evaluate(cnnModel,cnnPredictions,mnistTest.getProvenance());
System.out.println(cnnEvaluation.toString());
System.out.println(cnnEvaluation.getConfusionMatrix().toString());
```
As we might expect, exploiting the structured nature of images lets us get better performance, with 97% accuracy after only 3 epochs. There is a wide variety of different CNN architectures, each suited for different kinds of tasks. Some are even applied to sequential data like text.
## Exporting and Importing TensorFlow models
TensorFlow's canonical model storage format is the [`SavedModelBundle`](https://www.tensorflow.org/guide/saved_model). You can export TensorFlow models trained in Tribuo in this format by calling `model.exportModel(String path)` which writes a directory at that path which contains the model as a `SavedModel`.
```
var outputPath = "./tf-cnn-mnist-model";
cnnModel.exportModel(outputPath);
```
Tribuo can also load in `SavedModel`s and serve them as an `ExternalModel`. See the external models tutorial for more details on how Tribuo works with models built in other packages. The short version is that you need to specify the mapping from Tribuo's feature names into the id numbers the model expects, and from the output indices to Tribuo's output dimensions. We'll show how to load in the CNN that we just exported, and validate that it gives the same predictions as the original.
First we'll setup the feature and output mappings. This is easy in our case as we already have the relevant information, but in most cases this requires understanding how the features were prepared when the original model was trained. We discuss this in more detail in the external models tutorial.
```
var outputMapping = new HashMap<Label,Integer>();
for (var p : cnnModel.getOutputIDInfo()) {
outputMapping.put(p.getB(),p.getA());
}
var featureIDMap = cnnModel.getFeatureIDMap();
var featureMapping = new HashMap<String,Integer>();
for (var info : featureIDMap) {
featureMapping.put(info.getName(),featureIDMap.getID(info.getName()));
}
```
Now we build the `TensorFlowSavedModelExternalModel` using it's factory, supplying the feature mapping, output mapping, the softmax output operation name, the image transformer, the label transformer and finally the path to the `SavedModel` on disk.
```
var externalModel = TensorFlowSavedModelExternalModel.createTensorflowModel(
labelFactory, // the output factory
featureMapping, // the feature mapping
outputMapping, // the output mapping
cnnModel.getOutputName(), // the name of the *softmax* output
mnistImageConverter, // the input feature converter
mnistOutputConverter, // The label converter
outputPath.toString() // path to the saved model
);
```
This model behaves like any other, so we pass it some test data and generate it's predictions.
```
var externalPredictions = externalModel.predict(mnistTest);
```
Now let's compare the output predictions. It's a little convoluted, but we're going to compare each predicted probability distribution to make sure they are the same.
```
var isEqual = true;
for (int i = 0; i < cnnPredictions.size(); i++) {
var tribuo = cnnPredictions.get(i);
var external = externalPredictions.get(i);
isEqual &= tribuo.getOutput().fullEquals(external.getOutput());
isEqual &= tribuo.distributionEquals(external);
}
System.out.println("Predictions are " + (isEqual ? "equal" : "not equal"));
```
As we can see, the models produce identical predictions, which means that we've successfully exported all our model weights and managed to load them back in as an external model.
## Conclusion
We saw how to build MLPs and CNNs in Tribuo & TensorFlow for both regression and classification, along with how to export Tribuo-trained models into TensorFlow's format, and import TensorFlow SavedModels into Tribuo.
By default Tribuo pulls in the CPU version of TensorFlow Java, but if you supply the GPU jar at runtime it will automatically run everything on a compatible Nvidia GPU. We'll look at exposing explicit GPU support from Tribuo as the relevant support matures in TensorFlow Java.
| github_jupyter |
# Numpy
___
[](http://www.youtube.com/watch?v=NVTWjd_UpzM "Numpy Playlist")
**Numpy features**
___
1. sorting data
2. mutable iterable object
3. can be indexed
4. slicing operation can be perform
**diff between list and numpy array**
____
list = different datatypes (e.g. [1,'a',3])
np.array = similar datatypes (e.g. [1, 2, 3])
```
import numpy as np
```
Numpy operation
___
* [array()](#array)
* [arange()](#arange)
* [zeros()](#zeros)
* [ones()](#ones)
* [linspace()](#linspace)
* [eye()](#eye)
* [random()](#random)
### array
```
myList = [10, 55, 16, 74, 98]
np.array(myList)
myList = [10, 55, 16, 74, 98, 26]
np.array(myList).reshape(2, -1)
myList = [10, 55, 16, 74, 98, 26]
np.array(myList).reshape(2, 3)
```
### arange
___
numpy.arange([start, ]stop, [step, ], dtype=None) -> numpy.ndarray
___
**numpy integer datatypes**
* np.int8: 8-bit signed integer (from -128 to 127)
* np.uint8: 8-bit unsigned integer (from 0 to 255)
* np.int16: 16-bit signed integer (from -32768 to 32767)
* np.uint16: 16-bit unsigned integer (from 0 to 65535)
* np.int32: 32-bit signed integer (from -2 ** 31 to 2 ** 31-1)
* np.uint32: 32-bit unsigned integer (from 0 to 2** 32-1)
* np.int64: 64-bit signed integer (from -2** 63 to 2** 63-1)
* np.uint64: 64-bit unsigned integer (from 0 to 2** 64-1)
```
np.arange(1, 10)
np.arange(10)
np.arange(10.0)
np.arange(0, 10, 2)
np.arange(10, dtype='uint8')
```
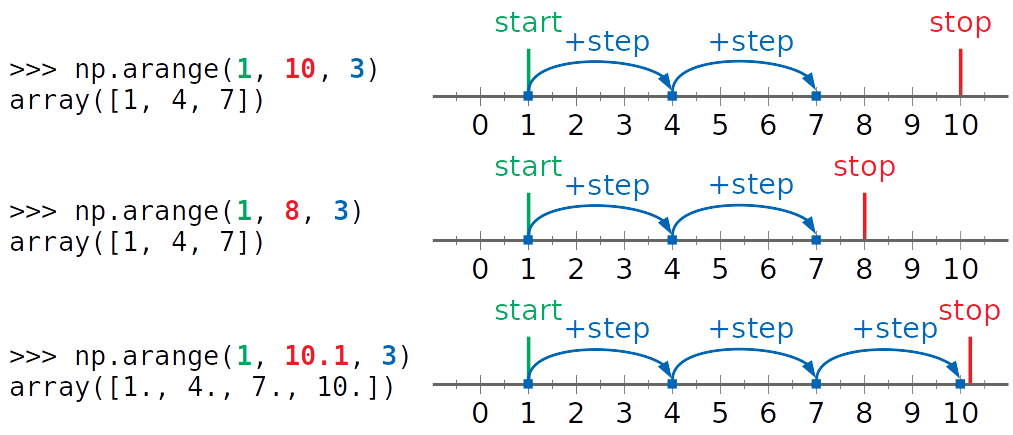
```
x = np.arange(1, 5)
x.size * x.itemsize
x**2
x = x.astype('uint8')
x.size * x.itemsize
np.arange(-1, 1.1, 0.5)
np.abs(np.arange(-1, 1.1, 0.5))
np.sin(np.arange(0, 180, 15))
%%timeit
np.arange(10)
%%timeit
[x for x in range(10)]
np.arange(8, 2)
```
### zeros
___
```python
numpy.zeros(shape, dtype = None, order = 'C')
```
___
shape : integer or sequence of integers
order : C_contiguous or F_contiguous
C-contiguous order in memory(last index varies the fastest)
C order means that operating row-rise on the array will be slightly quicker
FORTRAN-contiguous order in memory (first index varies the fastest).
F order means that column-wise operations will be faster.
dtype : [optional, float(byDeafult)] Data type of returned array.
```
np.zeros((3, 3))
np.zeros(4, dtype='int8')
```
### ones
___
```python
numpy.ones(shape, dtype = None, order = 'C')
```
___
shape : integer or sequence of integers
order : C_contiguous or F_contiguous
C-contiguous order in memory(last index varies the fastest)
C order means that operating row-rise on the array will be slightly quicker
FORTRAN-contiguous order in memory (first index varies the fastest).
F order means that column-wise operations will be faster.
dtype : [optional, float(byDeafult)] Data type of returned array.
```
np.ones(8)
np.ones(7, dtype='int')
np.ones((3, 4), dtype='uint8')
np.full((3, 4), 69)
np.empty(6)
np.empty((3, 4), dtype='int')
```
### linspace
___
**creates array filled evenly spaced values**
___
```
numpy.linspace(start,
stop,
num = 50,
endpoint = True,
retstep = False,
dtype = None)
```
___
```
-> start : [optional] start of interval range. By default start = 0
-> stop : end of interval range
-> restep : If True, return (samples, step). By deflut restep = False
-> num : [int, optional] No. of samples to generate
-> dtype : type of output array
```
```
np.linspace(1, 10)
np.linspace(1, 10, 3)
np.linspace(1, 10, 3, endpoint=False)
np.linspace(1, 100, 15)
np.linspace(1, 100, 15, retstep=True)
np.linspace(1, 100, 15, dtype='int')
np.linspace(-10, 10, 25)
%%timeit
np.linspace(-100, 100, 25000)
%%timeit
start, end, num = -100, 100, 25000
step = (abs(start) + abs(end)) / num
[start + (x*step) for x in range(num)]
m1 = np.linspace([1,2,3], [7,8,9], 3)
m1
m1.shape
m1.T
m2 = np.linspace([1,2,3], [7,8,9], 3, axis=1)
m2
import matplotlib.pyplot as plt
sin_points = np.linspace(0, 360, 50)
radian_Values = np.radians(sin_points)
sin_values = np.sin(radian_Values)
plt.plot(sin_points, sin_values)
plt.title("sin wave")
plt.ylabel("sin(x)")
plt.xlabel("x value")
plt.show()
import matplotlib.pyplot as plt
tan_points = np.linspace(0, 360, 50)
radian_Values = np.radians(tan_points)
tan_values = np.tan(radian_Values)
plt.plot(tan_points, tan_values)
plt.title("tan wave")
plt.ylabel("tan(x)")
plt.xlabel("x value")
plt.show()
import matplotlib.pyplot as plt
x_points = np.linspace(0, 360, 50)
radian_Values = np.radians(x_points)
cos_values = np.cos(radian_Values)
plt.plot(x_points, cos_values)
plt.title("cos wave")
plt.ylabel("cos(x)")
plt.xlabel("x values")
plt.show()
```
### eye
___
```
numpy.eye(R, C = None, k = 0, dtype = type <‘float’>)
```
___
```
R : Number of rows
C : [optional] Number of columns; By default M = N
k : [int, optional, 0 by default]
Diagonal we require; k>0 means diagonal above main diagonal or vice versa.
dtype : [optional, float(by Default)] Data type of returned array.
array of shape, R x C, an array where all elements
are equal to zero, except for the k-th diagonal,
whose values are equal to one.
```
___
**returns array filled with zeros except in the k-th diagonal, whose values are equal to 1**
```
np.eye(5)
np.eye(5, dtype=int)
np.eye(2, 3)
np.eye(4, k=0)
np.eye(4, k=-1)
np.eye(4, k=1)
np.identity(4)
```
### random
___
* rand()
* randn()
* ranf()
* randint()
#### rand
___
```
numpy.random.rand(d0, d1, ..., dn)
```
values will be in [0,1) interval
```
np.random.rand(5)
np.random.rand(3, 3)
np.random.rand(4, 5)
```
#### randn
___
```
The numpy.random.randn() function creates an array of specified shape and fills it with random values as per standard normal distribution.
```
```
np.random.randn(5)
np.random.randn(3, 3)
np.random.ranf()
np.random.ranf(5)
```
#### randint
___
```
Syntax : numpy.random.randint(low, high=None, size=None, dtype=’l’)
Parameters :
low : [int] Lowest (signed) integer to be drawn from the distribution.But, it works as a highest integer in the sample if high=None.
high : [int, optional] Largest (signed) integer to be drawn from the distribution.
size : [int or tuple of ints, optional] Output shape. If the given shape is, e.g., (m, n, k), then m * n * k samples are drawn. Default is None, in which case a single value is returned.
dtype : [optional] Desired output data-type.
Return : Array of random integers in the interval [low, high) or a single such random int if size not provided.
```
___
interval is [0, low) if high is not given
interval is [low, high)
```
np.random.randint(2, size=5)
np.random.randint(0, 10, size=(3, 3))
np.random.randint(0, 10, size=(3, 3, 3))
```
### Numpy attributes or property
____
```
a = np.arange(6).reshape((2, 3))
print(f'\
Shape = {a.shape}\n\
dimension = {a.ndim}\n\
total number of elements = {a.size}\n\
each element size = {a.itemsize}\n\
total size consumed by array = {a.nbytes}\n\
data type of array = {a.dtype}')
```
### Numpy Data types
___
1. **bool_** : It represents the boolean value indicating true or false. It is stored as a byte.
2. **int_** : It is the default type of integer. It is identical to long type in C that contains 64 bit or 32 bit integer.
3. **intc** : It is similar to the c integer (c int) as it represents 32 or 64-bit int.
4. **intp** : It represents the integetrs which are usedfor indexing.
5. **int8** : It is the 8-bit integer identicalto a byte. The range of the value is -128 to 127.
6. **int16** : It is the 2-byte (16-bit) integer. The range is -32768 to 32767.
7. **int32** : It is the 4-Byte(32-bit) integer. The range is -2147483648 to 2147483647.
8. **int64** : It is the 8-byte(64-bit) integer. The range is -9223372036854775808 to 9223372036854775807.
9. **uint8** : It is the 1-byte(8-bit) unsigned integer.
10. **uint16** : It is the 2-byte(16-bit) unsigned integer.
11. **uint32** : It is the 4-byte(32-bit) unsigned integer.
12. **uint64** : It is the 8-bytes(64-bit) unsigned integer.
13. **float** : It is the identical to float64.
14. **float16** : It is the half-precisionfloat. 5 bits reserved for the exponent. 10 bits are reserved for mantissa, and 1 bit is reserved for the sign.
15. **float32** : It is a single precisionfloat. 8 bits are reserved for the exponent, 23bits are reserved for mantissa, and 1 bit is reserved for the sign.
16. **float64** : It is the double precision float. 11 bits are reserved for the exponent, bits are reserved for mantissa, bit is used for the sign.
17. **complex** : It is identical to complex.
18. **complex64** : It is used to represent the complex number where real and imaginary part shares 32 bits each.
19. **complex128** : It is used to represent thecomplex number whwre real and imaginary part shares 64 bits each.
___
| github_jupyter |
```
#import libraries
import networkx as nx
import matplotlib.pyplot as plt
```
# Erdos-Renyi graph
```
# a random erdos-renyi graph with 100 nodes and a edge probability of 0.3
graph = nx.erdos_renyi_graph(100, 0.3, seed=None, directed=False)
nx.draw(graph)
plt.title("A random erdos-renyi graph with 100 nodes and a edge probability of 0.3")
#plotting the degree_centrality distribution (a histogram of the degree centrality values)
graph_deg_cent = nx.degree_centrality(graph)
plt.hist(list(graph_deg_cent.values()))
plt.title('Degree Centrality of an erdos renyi graph with p = 0.3')
plt.show()
#Changing the edge probability of the erdos-reyni graph to 0.6
graph2 = nx.erdos_renyi_graph(100, 0.6, seed=None, directed=False)
nx.draw(graph2)
plt.title("A random erdos-renyi graph with 100 nodes and a edge probability of 0.3")
#plotting the degree centrality distribution again
graph2_deg_cent = nx.degree_centrality(graph2)
plt.hist(list(graph2_deg_cent.values()))
plt.title('Degree Centrality of an erdos renyi graph with p = 0.6')
plt.show()
```
# Infering the changes in the plots
From the above histograms, it can be conculded that the difference between them is due to different edge probabilities. The frequency of the first histogram(edge probability = 0.3) looks higher than that of the second histogram (edge probability = 0.6) because the higher edge probability, the more likely it is to include graph with more edges and the less likely it is to include graph with fewer edges.
# Barabasi Albert Graph
```
# A random barabasi_albert_graph with 100 nodes and attached 3 edges to a new node in the graph (m=3)
graph3 = nx.barabasi_albert_graph(100, 3, seed=None, initial_graph=None)
nx.draw(graph3)
plt.title("A random barabasi_albert_graph with 100 nodes and attached 3 edges to a new node in the graph (m=3)")
# plotting the degree_centrality distribution (a histogram of the degree histogram)
graph3_deg_cent = nx.degree_centrality(graph3)
plt.hist(list(graph3_deg_cent.values()))
plt.title('A random barabasi_albert_graph with 100 nodes and attached 3 edges to a new node')
plt.show()
```
# Inferring the changes in the plot of two different random graphs(Erdos-Renyi graph and Barabasi Albert Graph)
The Erdos Renyi Graph has more nodes which have a higher degree centrality value and no node will have much higher degree than any other because nodes are assigned and each pair is connected with probability while for the Barabasi Albert histogram there are only very few nodes with a high degree centrality value.That is, for Barabasi Albert graph, nodes are equally assigned but they are added one at a time. When a node is added it is connected to a small number of existing nodes with probability proportional to the degree of existing nodes as a result the earlier nodes tend to have a higher degree. This explains why the histogram here has a very few nodes with high degree centrality value
| github_jupyter |
# Data Mining Challange: *Reddit Gender Text-Classification*
### Modules
```
# Numpy & matplotlib for notebooks
%pylab inline
# Pandas for data analysis and manipulation
import pandas as pd
# Sparse matrix package for numeric data.
from scipy import sparse
# Module for word embedding (word2vector)
import gensim
# Module for progress monitoring
import tqdm
# Sklearn
from sklearn.preprocessing import StandardScaler # to standardize features by removing the mean and scaling to unit variance (z=(x-u)/s)
from sklearn.neural_network import MLPClassifier # Multi-layer Perceptron classifier which optimizes the log-loss function using LBFGS or sdg.
from sklearn.svm import SVC # Support Vector Classification
from sklearn.ensemble import RandomForestClassifier # A meta-estimator that fits a number of decision tree classifiers on various sub-samples of the dataset and uses averaging to improve the predictive accuracy and control over-fitting
from sklearn.decomposition import PCA, TruncatedSVD # Principal component analysis (PCA); dimensionality reduction using truncated SVD.
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import MultinomialNB # Naive Bayes classifier for multinomial models
from sklearn.feature_extraction.text import CountVectorizer # Convert a collection of text documents to a matrix of token counts
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics import roc_auc_score as roc # Compute Area Under the Receiver Operating Characteristic Curve from prediction scores
from sklearn.metrics import roc_curve, auc # Compute ROC; Compute Area Under the Curve (AUC) using the trapezoidal rule
from sklearn.model_selection import RandomizedSearchCV, GridSearchCV # Exhaustive search over specified parameter values for a given estimator
from sklearn.model_selection import cross_val_score # Evaluate a score by cross-validation
from sklearn.model_selection import train_test_split # to split arrays or matrices into random train and test subsets
from sklearn.model_selection import KFold # K-Folds cross-validator providing train/test indices to split data in train/test sets.
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.model_selection import StratifiedKFold
import nltk
import re
from nltk.stem import WordNetLemmatizer
from bs4 import BeautifulSoup
from nltk.corpus import stopwords
from collections import defaultdict
from nltk.tokenize import word_tokenize
from nltk import pos_tag
from nltk.corpus import wordnet as wn
#XGBoost
from xgboost import XGBRegressor
# Matplotlib
import matplotlib # Data visualization
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
# Seaborn
import seaborn as sns # Statistical data visualization (based on matplotlib)
# Joblib
import joblib # To save models
```
## Data Loading and Manipulation
```
# load data
train_data = pd.read_csv("../input/dataset/train_data.csv")
target = pd.read_csv("../input/dataset/train_target.csv")
# create authors gender dictionary
author_gender = {}
for i in range(len(target)):
author_gender[target.author[i]] = target.gender[i]
# X is the aggregated comments list
X = []
# the genders
y = []
# lengths of X elements
X_len = []
for author, group in train_data.groupby("author"):
X.append(group.body.str.cat(sep = " "))
X_len.append([len(group.body)])
y.append(author_gender[author])
```
## Preprocessing
```
# preprocessing functions
def remove_number(text):
num = re.compile(r'[-+]?[.\d]*[\d]+[:,.\d]*')
return num.sub(r'NUMBER', text)
def remove_URL(text):
url = re.compile(r'https?://\S+|www\.\S+')
return url.sub(r'URL',text)
def remove_repeat_punct(text):
rep = re.compile(r'([!?.]){2,}')
return rep.sub(r'\1 REPEAT', text)
# remove words that end with one or more identical letters
def remove_elongated_words(text):
rep = re.compile(r'\b(\S*?)([a-z])\2{2,}\b')
return rep.sub(r'\1\2 ELONG', text)
def remove_allcaps(text):
caps = re.compile(r'([^a-z0-9()<>\'`\-]){2,}')
return caps.sub(r'ALLCAPS', text)
def transcription_smile(text):
eyes = "[8:=;]"
nose = "['`\-]"
smiley = re.compile(r'[8:=;][\'\-]?[)dDp]')
return smiley.sub(r'SMILE', text)
def transcription_sad(text):
eyes = "[8:=;]"
nose = "['`\-]"
smiley = re.compile(r'[8:=;][\'\-]?[(\\/]')
return smiley.sub(r'SADFACE', text)
def transcription_heart(text):
heart = re.compile(r'<3')
return heart.sub(r'HEART', text)
# tags Part of Speech (POS), because teh lemmatizer needs it
tag_map = defaultdict(lambda : wn.NOUN)
# wn does a grammatical analysis
tag_map['J'] = wn.ADJ
tag_map['V'] = wn.VERB
tag_map['R'] = wn.ADV
# create lemmatizer
word_Lemmatized = WordNetLemmatizer()
def review_to_words(raw_body):
# remove html tags
body_text = BeautifulSoup(raw_body).get_text()
#letters_only = re.sub("[^a-zA-Z]", " ", body_text)
# lowercase all text
words = body_text.lower()
# remove urls
text = remove_URL(words)
# remove numbers
text = remove_number(text)
# remove smiles
text = transcription_sad(text)
text = transcription_smile(text)
text = transcription_heart(text)
text = remove_elongated_words(text)
words = remove_repeat_punct(text)
# tokenizes and pass to lemmatizer, which lemmatizes taking tags into account (see before)
words = word_tokenize(words)
# we don't remove stop words, because doing it on combination with removing the 40 (trial & error estimated parameter) most utilized words (see below) decreases performance
#stops = set(stopwords.words("english"))
#meaningful_words = [w for w in words if not w in stops]
Final_words = []
for word, tag in pos_tag(words):
word_Final = word_Lemmatized.lemmatize(word,tag_map[tag[0]])
Final_words.append(word_Final)
# returns lemmatized texts as strings
return( " ".join(Final_words))
clean_train_comments = [review_to_words(x) for x in X]
```
## Train Countvectorizer, Optimize Input for model Training
```
#from sklearn.feature_extraction.text import TfidfVectorizer # Il CountVectorizer ha performato meglio
from sklearn.feature_extraction.text import CountVectorizer
vectorizer = CountVectorizer(analyzer = "word",
max_features = 2000)
# converts in np array
train_data_features = vectorizer.fit_transform(clean_train_comments).toarray()
print(train_data_features.shape)
# print vocabulary
vocab = vectorizer.get_feature_names()
print(vocab)
import numpy as np
# counts how many times a word appearco
dist = np.sum(train_data_features, axis=0)
for tag, count in zip(vocab, dist):
print(count, tag)
# removes the 40 most utilized words
for _ in range(40):
index = np.argmax(dist)
train_data_features = np.delete(train_data_features, index, axis = 1)
train_data_features.shape
# np array
s = np.concatenate((train_data_features,np.array(X_len)),axis = 1)
# 5000 rows (one per author), and 2000-40+1 (X_len) features
s.shape
# un np.array
y = np.array(y)
```
## Train data TruncatedSVD visualization
```
# Plot the test data along the 2 dimensions of largest variance
def plot_LSA(test_data, test_labels, savepath="PCA_demo.csv", plot=True):
lsa = TruncatedSVD(n_components=2)
lsa.fit(test_data)
lsa_scores = lsa.transform(test_data)
color_mapper = {label:idx for idx,label in enumerate(set(test_labels))}
color_column = [color_mapper[label] for label in test_labels]
colors = ['orange','blue']
if plot:
plt.scatter(lsa_scores[:,0], lsa_scores[:,1], s=8, alpha=.8, c=test_labels, cmap=matplotlib.colors.ListedColormap(colors))
orange_patch = mpatches.Patch(color='orange', label='M')
blue_patch = mpatches.Patch(color='blue', label='F')
plt.legend(handles=[orange_patch, blue_patch], prop={'size': 20})
plt.title('BOW + lenght comments only')
plt.savefig('foo.pdf')
fig = plt.figure(figsize=(8, 8))
plot_LSA(train_data_features, y)
plt.show()
```
## Model Training and Prediction
```
# splits
X_train, X_valid, y_train, y_valid = train_test_split(s, y,
train_size=0.8, test_size=0.2,
random_state=0)
# XGBoost model with parameters set with a RandomGridSearch
my_model = XGBRegressor(objective = "reg:logistic",n_estimators=10000, learning_rate=0.01, n_jobs=4,subsample = 0.9,
min_child_weight = 1,max_depth=4,gamma=1.5,colsample_bytree=0.6 )
# cross_val_score resets parameters of my_model and fits it on X_train and t_train with cross validation (we did it for consistency).
kfold = KFold(n_splits=10)
results = cross_val_score(my_model, s, y, cv=kfold, scoring='roc_auc')
print("roc = ", np.mean(results))
# fits
my_model.fit(X_train, y_train,
early_stopping_rounds=80,
#sample_weight = w,
eval_set=[(X_valid, y_valid)],
verbose=False)
# In the fit function there is the early stop, that one may set iff there is a validation set.
# The early stop interrupts the training when themodel starts overfitting.
# But, the model that will predict the test will have no validation during training, so we get here a value and heuristicallly use it also when predicting test.
print(my_model.best_iteration)
# ROC plot
y_score = my_model.predict(X_valid)
# Roc Curve for validation data
fpr, tpr, thresholds = roc_curve(y_valid, y_score)
roc_auc = auc(fpr, tpr)
plt.figure()
lw = 2
plt.plot(fpr, tpr, color='darkorange',
lw=lw, label='ROC curve (area = %0.4f)'% roc_auc )
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic example')
plt.legend(loc="lower right")
plt.show()
# Validation predictions
y_predict = my_model.predict(X_valid)
# Verification
roc(y_valid,y_predict)
# Save predictions
np.save('../working/y_predict_XGB.csv',y_predict)
# ROC evaluated on train set to evaluate overfitting
y_predict = my_model.predict(X_train)
roc(y_train,y_predict)
```
## Appendix: *GridSearch*
Thsi is the RandomGridSearch that we used to estimate the best parameters for the XGBoost
```python
from xgboost import XGBRegressor
from sklearn.metrics import roc_curve, auc
from sklearn.model_selection import RandomizedSearchCV, GridSearchCV
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.model_selection import StratifiedKFold
xgb = XGBRegressor(learning_rate=0.01, n_estimators=10000, objective='reg:logistic',
silent=True, nthread=1)
#n_esimators si assume indipendente e quindi ottimizzato a parte
params = {
'min_child_weight': [1,8],
'gamma': [0.6,0.8],
'subsample': [0.9],
'colsample_bytree': [0.6],
'max_depth': [4],
'scale_pos_weight': [1,2.70, 10, 25, 50, 75, 100, 1000]
}
folds = 5
param_comb = 18 #in realtà di più
skf = StratifiedKFold(n_splits=folds, shuffle = True, random_state = 1001)
random_search = RandomizedSearchCV(xgb, param_distributions=params, n_iter=param_comb, scoring='roc_auc', n_jobs=4, cv=skf.split(X_train,y_train), verbose=3, random_state=0)
print('\n Best score for %d-fold search with %d parameter combinations:' % (folds, param_comb))
print(random_search.best_score_)
print('\n Best hyperparameters:')
print(random_search.best_params_)
results = pd.DataFrame(random_search.cv_results_)
results.to_csv('xgb-random-grid-search-results-01.csv', index=False)
```
| github_jupyter |
# Getting Started
```
''' Required for Google Colab'''
# Mount GDrive
# from google.colab import drive
# drive.mount('/content/drive')
# # Upload Kaggle json
# !pip install -q kaggle
# !pip install -q kaggle-cli
# !mkdir -p ~/.kaggle
# !cp "/content/drive/My Drive/Kaggle/kaggle.json" ~/.kaggle/
# !cat ~/.kaggle/kaggle.json
# !chmod 600 ~/.kaggle/kaggle.json
# !kaggle competitions download -c fake-news -p dataset
# !unzip /content/dataset/train.csv.zip
# !unzip /content/dataset/test.csv.zip
''' Required for Google Colab'''
# !pip install contractions
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import rcParams
plt.rcParams['figure.figsize'] = [10,10]
import seaborn as sns
sns.set_theme(style="darkgrid")
import nltk
from nltk import sent_tokenize
''' Required for Google Colab'''
# nltk.download('punkt')
# nltk.download('averaged_perceptron_tagger')
# nltk.download('stopwords')
''' Required for Google Colab'''
from nltk.corpus import stopwords
stopwords = nltk.corpus.stopwords.words('english')
from nltk.tokenize import word_tokenize
import contractions
import string
import joblib
import re
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.naive_bayes import MultinomialNB
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier, AdaBoostClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import precision_score, recall_score, f1_score, accuracy_score
from sklearn.model_selection import GridSearchCV
import warnings
warnings.filterwarnings('ignore')
''' Required for Google Colab'''
# train_df = pd.read_csv('/content/train.csv', header=0)
# test_df = pd.read_csv('/content/test.csv', header=0)
train_df = pd.read_csv('fake-news/train.csv', header=0)
test_df = pd.read_csv('fake-news/test.csv', header=0)
train_df = train_df.fillna(' ')
test_df = test_df.fillna(' ')
train_df['text'] = train_df['text'].str.strip()
test_df['text'] = test_df['text'].str.strip()
train_df['raw_text_length'] = train_df['text'].apply(lambda x: len(x))
print(len(train_df[train_df['raw_text_length']==0]))
train_df = train_df[train_df['raw_text_length'] > 0]
train_df.shape
```
## Text Cleaning
1. Remove special characters
2. Expand contractions
3. Convert to lower-case
4. Word Tokenize
5. Remove Stopwords
```
def preprocess_text(x):
cleaned_text = re.sub(r'[^a-zA-Z\d\s\']+', '', x)
word_list = []
for each_word in cleaned_text.split(' '):
try:
word_list.append(contractions.fix(each_word).lower())
except:
print(x)
return " ".join(word_list)
```
## Got Error because of some sort of Turkish/Slavic language
ABÇin ilişkilerinde ABD ve NATOnun etkisi yazan Manlio Dinucci Uluslararası bir forumda konuşan İtalyan coğrafyacı Manlio Dinucci ABDnin tüm dünyaya egemen olabilmek için sahip olduğu silahların analizini bireşimleştirdi Suriye Rusya ve Çinin bugün elde silah herkesin açıkça kabul ettiği bu üstünlüğü dünyanın bu tek kutuplu örgütlenişi tartışılır hale getirmesinden dolayı bu makale daha da önem kazanmaktadır
Therefore I rearranged the order of preprocessing
```
text_cols = ['text', 'title', 'author']
%%time
for col in text_cols:
print("Processing column: {}".format(col))
train_df[col] = train_df[col].apply(lambda x: preprocess_text(x))
test_df[col] = test_df[col].apply(lambda x: preprocess_text(x))
%%time
for col in text_cols:
print("Processing column: {}".format(col))
train_df[col] = train_df[col].apply(word_tokenize)
test_df[col] = test_df[col].apply(word_tokenize)
%%time
for col in text_cols:
print("Processing column: {}".format(col))
train_df[col] = train_df[col].apply(
lambda x: [each_word for each_word in x if each_word not in stopwords])
test_df[col] = test_df[col].apply(
lambda x: [each_word for each_word in x if each_word not in stopwords])
train_df.head()
```
## Model Training
```
train_df['text_joined'] = train_df['text'].apply(lambda x: " ".join(x))
test_df['text_joined'] = test_df['text'].apply(lambda x: " ".join(x))
target = train_df['label'].values
count_vectorizer = CountVectorizer(ngram_range=(1, 2))
tf_idf_transformer = TfidfTransformer(smooth_idf=False)
# fit train data to count vectorizer
count_vectorizer.fit(train_df['text_joined'].values)
count_vect_train = count_vectorizer.transform(train_df['text_joined'].values)
# fit ngrams count to tfidf transformers
tf_idf_transformer.fit(count_vect_train)
tf_idf_train = tf_idf_transformer.transform(count_vect_train)
# Transform the test data as well
count_vect_test = count_vectorizer.transform(test_df['text_joined'].values)
tf_idf_test = tf_idf_transformer.transform(count_vect_test)
```
# Train Test Split
```
X_train, X_test, y_train, y_test = train_test_split(tf_idf_train, target, random_state=0)
df_perf_metrics = pd.DataFrame(columns=['Model', 'Accuracy_Training_Set', 'Accuracy_Test_Set', 'Precision', 'Recall', 'f1_score'])
```
# Machine Learning Classifier Training and Validating
```
df_perf_metrics = pd.DataFrame(columns=[
'Model', 'Accuracy_Training_Set', 'Accuracy_Test_Set', 'Precision',
'Recall', 'f1_score', 'Training Time (secs'
])
models_trained_list = []
def get_perf_metrics(model, i):
# model name
model_name = type(model).__name__
# time keeping
start_time = time.time()
print("Training {} model...".format(model_name))
# Fitting of model
model.fit(X_train, y_train)
print("Completed {} model training.".format(model_name))
elapsed_time = time.time() - start_time
# Time Elapsed
print("Time elapsed: {:.2f} s.".format(elapsed_time))
# Predictions
y_pred = model.predict(X_test)
# Add to ith row of dataframe - metrics
df_perf_metrics.loc[i] = [
model_name,
model.score(X_train, y_train),
model.score(X_test, y_test),
precision_score(y_test, y_pred),
recall_score(y_test, y_pred),
f1_score(y_test, y_pred), "{:.2f}".format(elapsed_time)
]
# keep a track of trained models
models_trained_list.append(model)
print("Completed {} model's performance assessment.".format(model_name))
models_list = [LogisticRegression(),
MultinomialNB(),
RandomForestClassifier(),
DecisionTreeClassifier(),
GradientBoostingClassifier(),
AdaBoostClassifier()]
%%time
for n, model in enumerate(models_list):
get_perf_metrics(model, n)
df_perf_metrics
```
## Adding Title and Author Information to the Text
To check if there is any improvement
```
train_df['all_info'] = train_df['text'] + train_df['title'] + train_df['author']
train_df['all_info'] = train_df['all_info'].apply(lambda x: " ".join(x))
test_df['all_info'] = test_df['text'] + test_df['title'] + test_df['author']
test_df['all_info'] = test_df['all_info'].apply(lambda x: " ".join(x))
tf_idf_transformer = TfidfTransformer(smooth_idf=False)
count_vectorizer = CountVectorizer(ngram_range=(1, 2))
count_vect_train = count_vectorizer.fit_transform(train_df['all_info'].values)
tf_idf_train = tf_idf_transformer.fit_transform(count_vect_train)
X_train, X_test, y_train, y_test = train_test_split(tf_idf_train,
target,
random_state=0)
# Transform the test data
count_vect_test = count_vectorizer.transform(test_df['all_info'].values)
tf_idf_test = tf_idf_transformer.transform(count_vect_test)
%%time
for n, model in enumerate(models_list):
get_perf_metrics(model, n)
df_perf_metrics
```
## Tuning the Logistic Regression Model
```
model = LogisticRegression()
max_iter = [100, 200, 500, 1000]
C = [0.1, 0.5, 1, 10, 50, 100]
param_grid = dict(max_iter=max_iter, C=C)
grid = GridSearchCV(estimator=model,
param_grid=param_grid,
cv=5,
scoring=['f1'],
refit='f1',
verbose=2)
%%time
grid_result = grid.fit(X_train, y_train)
grid_result.best_estimator_
grid_result.best_params_
model = grid_result.best_estimator_
y_pred = model.predict(X_test)
print('Accuracy: ', accuracy_score(y_test, y_pred))
print('Precision: ', precision_score(y_test, y_pred))
print('Recall: ', recall_score(y_test, y_pred))
print('f1-score: ', f1_score(y_test, y_pred))
```
## Retuning the Logistic Regression Classifier
### Attempt 1
In the last grid, the lowest of the max_iter list (100) and highest of the C list (100) were the best values. In this attempt, I will see if there are better fits in the values beyond these, lower than 100 for max_iter or just a bit higher, and higher than 100 for C or just a bit lower.
```
%%time
model = LogisticRegression()
max_iter = [50, 75, 100]
C = [75, 100, 125]
param_grid = dict(max_iter=max_iter, C=C)
grid = GridSearchCV(estimator=model,
param_grid=param_grid,
cv=5,
scoring=['f1'],
refit='f1',
verbose=2)
grid_result = grid.fit(X_train, y_train)
model = grid_result.best_estimator_
y_pred = model.predict(X_test)
print('Accuracy: ', accuracy_score(y_test, y_pred))
print('Precision: ', precision_score(y_test, y_pred))
print('Recall: ', recall_score(y_test, y_pred))
print('f1-score: ', f1_score(y_test, y_pred))
grid_result.best_params_
```
### Attempt 2
In the previous attempt, we saw max_iter is still at 100 while C = 125 resulted into a marginally higher precision and consequently the f1 score improved just a bit. Let's see if we can improve the results a little more.
For this I have kept C constant and I have stated a range of C from 120 to 150 which increases by a step value of 10.
```
%%time
model = LogisticRegression()
max_iter = [100]
C = [120, 130, 140, 150]
param_grid = dict(max_iter=max_iter, C=C)
grid = GridSearchCV(estimator=model,
param_grid=param_grid,
cv=5,
scoring=['f1'],
refit='f1',
verbose=2)
grid_result = grid.fit(X_train, y_train)
model = grid_result.best_estimator_
y_pred = model.predict(X_test)
print('Accuracy: ', accuracy_score(y_test, y_pred))
print('Precision: ', precision_score(y_test, y_pred))
print('Recall: ', recall_score(y_test, y_pred))
print('f1-score: ', f1_score(y_test, y_pred))
grid_result.best_params_
```
### Attempt 3
One final attempt to find the best model, keeping C = 100 and max_iter = 100, 125, 140.
```
%%time
model = LogisticRegression()
max_iter = [100]
C = [100, 125, 140]
param_grid = dict(max_iter=max_iter, C=C)
grid = GridSearchCV(estimator=model,
param_grid=param_grid,
cv=5,
scoring=['f1'],
refit='f1',
verbose=2)
grid_result = grid.fit(X_train, y_train)
model = grid_result.best_estimator_
y_pred = model.predict(X_test)
print('Accuracy: ', accuracy_score(y_test, y_pred))
print('Precision: ', precision_score(y_test, y_pred))
print('Recall: ', recall_score(y_test, y_pred))
print('f1-score: ', f1_score(y_test, y_pred))
grid_result.best_params_
file_name = 'Log_Reg_Best_Model.sav'
joblib.dump(model, file_name)
```
# Create Kaggle Submission
```
# Predicting the test dataframe and create submit.csv
# Picking up Logistic Regression since it has the best performance
submit_predictions = model.predict(tf_idf_test)
print(train_df['label'].value_counts() * 100/len(train_df))
print(np.bincount(submit_predictions)* 100 / len(submit_predictions))
## Create Submit Dataframe
submit_predictions_df = pd.DataFrame()
submit_predictions_df['id'] = test_df['id']
submit_predictions_df['label'] = submit_predictions
## Plot Genuine and Fake News
sns.countplot(submit_predictions, palette='Set3')
plt.title("Count of Genuine and Fake News of the Test Data")
plt.show()
submit_predictions_df.head()
submit_predictions_df.to_csv('kaggle_submissions/submit.csv',index=False)
```
Finally, uploaded my submission on Kaggle and checked the result.

Thank you for visiting!
| github_jupyter |
# Study of Glassdoor Data
The purpose of this study is too quickly present the Glassdoor data with the different attributes
The datasets have been cleaned with cleaning functions written in python
We will after studied missing data siginification
## Import packages and constants and helpers
```
# Packages
import pandas as pd
from autoc import DataExploration,NaImputer
from autoc.naimputer import missing_map
%pylab inline --no-import
import pylab as pl
import seaborn as sns
import matplotlib as mpl
import matplotlib.pyplot as plt
# Seaborn options
sns.set_palette("deep", desat=.6)
sns.set_context(rc={"figure.figsize": (8, 4)})
plt.style.use('ggplot') # ggplot2 style for mathplotlib
# path to cleaned datasets
path_reviews_cleaned = '~/Google Drive/Auto_clean/Datasets/Glassdoor/'
```
## Glassdoor Cleaned Reviews Data
### Quick Exploration
```
df_reviews = pd.read_csv(path_reviews_cleaned + 'glassdoor_reviews_cleaned_utf8_170415.csv')
df_reviews.head()
df_reviews[df_reviews.company_name == 'Google'].iloc[0]
df_reviews[df_reviews.company_name == 'Dataiku'] # too bad (dataset too old)
df_reviews[df_reviews.company_name == 'Uber'].iloc[0]['url']
df_reviews.columns
df_reviews.head()
exploration = DataExploration(df_reviews)
exploration.structure()
df_reviews[pd.isnull(df_reviews.stars)].company_name
df_reviews = df_reviews.drop(labels = ['benefits_below','benefits_above'],axis = 1)
df_reviews = df_reviews.dropna(subset=['stars'])
# Let's see what if Lending Club is one of the best place to work based on stars
df_sort = df_reviews[df_reviews.nb_c_reviews_detailled > 50].sort_values('stars',ascending=False).reset_index()
print('Uber is the {} happiest company'.format(str(df_sort[(df_sort.company_name == "Uber")].index[0])))
df_sort.head(5)
```
### Look at stars data
```
p = plt.hist(df_reviews.stars,bins = 30,histtype="stepfilled", color="#F08080", alpha=.5)
```
**Notes**: You can see the problem of a real life distribution (discontinuous because of small companies)
```
order = [u'1 to 5 Employees',u'6 to 15 Employees',u'16 to 50 Employees',
u'50 to 149 Employees', u'150 to 499 Employees', u'500 to 999 Employees',
u'1000 to 5000 Employees', u'5000+ Employees','Unknown']
# Violin plot
pl.figure(figsize=(20, 10))
sns.violinplot(df_reviews.stars, df_reviews['size'],order=order)
```
**Notes**: You can see the discontinuity for company with few employees
```
# Stars per size of the company
pl.figure(figsize=(20, 10))
sns.barplot("size", "stars",order = order,data = df_reviews)
big_companies = df_reviews.loc[df_reviews['size'] == "5000+ Employees"]
sns.distplot(big_companies.stars,color = '#F08080')
```
### Study on Missing data
#### Some Theory
##### Lay and Rubbin
This is a scraped and real dataset with a lot of missing data we are going to try to respond to lay and rubin theory
* **MCAR** : "Missing completely at random": the missing values are completely random and doesn/t depend from observations or any other factor.
* **MAR** : "Missing at random": missing values depends from the observations.
* **NMAR** : , "Not Missing at random": missing values depends on the unobserved original data values.
##### Purpose of the study
We are going to use statistic such as conditionnal expectation
#### Using autoc DataExploration class
```
exploration = DataExploration(df_reviews)
exploration.nacolcount()
df_test = df_reviews.copy()
df_test['is_na_interview_difficulty'] = df_test.interview_difficulty.isnull().astype(int)
def cserie(serie):
return serie[serie].index.tolist()
cserie((df_test.dtypes == int) | (df_test.dtypes == float))
def plot_hist_na(df, colname):
df_h = df.copy()
na_name = "is_na_{}".format(colname)
df_h[na_name] = df_h[colname].isnull().astype(int)
measure_col = cserie((df.dtypes == int) | (df.dtypes == float))
df_h.groupby(na_name)[measure_col].hist()
plot_hist_na(df_reviews,"revenue")
plot_hist_na(df_reviews,"interview_difficulty")
df_test.dtypes
df_test['is_na_interview_difficulty']
df_test.groupby('is_na_interview_difficulty').describe()
df_test.groupby('is_na_interview_difficulty')[['founded','ceo_rating']].hist()
# g = sns.FacetGrid(tips, col="time")
# g.map(plt.hist, "tip");
g = sns.FacetGrid(data=df_test, col='is_na_interview_difficulty')
g.map(plt.hist, "founded")
for col in ['founded','ceo_rating']:
g = sns.FacetGrid(data=df_test, col='is_na_interview_difficulty',hue="is_na_interview_difficulty")
g.map(sns.distplot, col)
g = sns.PairGrid(df_test,
y_vars=["founded", "ceo_rating", "nb_c_interviews"],
x_vars=["is_na_interview_difficulty"],
aspect=.75, size=3.5)
g.map(sns.violinplot, palette="pastel")
```
### Naimputer Examples
```
missing_map(df_reviews, nmax=1000)
na = NaImputer(df_reviews)
na.get_isna_mean(colname='ceo_rating')
na.isna_summary(colname='ceo_rating')
na.infos_na()
na.plot_corrplot_na(size=7,figsize=(20,10))
```
| github_jupyter |
# Keras for Text Classification
**Learning Objectives**
1. Learn how to tokenize and integerize a corpus of text for training in Keras
1. Learn how to do one-hot-encodings in Keras
1. Learn how to use embedding layers to represent words in Keras
1. Learn about the bag-of-word representation for sentences
1. Learn how to use DNN/CNN/RNN model to classify text in keras
## Introduction
In this notebook, we will implement text models to recognize the probable source (Github, Tech-Crunch, or The New-York Times) of the titles we have in the title dataset we constructed in the previous lab.
In a first step, we will load and pre-process the texts and labels so that they are suitable to be fed to a Keras model. For the texts of the titles we will learn how to split them into a list of tokens, and then how to map each token to an integer using the Keras Tokenizer class. What will be fed to our Keras models will be batches of padded list of integers representing the text. For the labels, we will learn how to one-hot-encode each of the 3 classes into a 3 dimensional basis vector.
Then we will explore a few possible models to do the title classification. All models will be fed padded list of integers, and all models will start with a Keras Embedding layer that transforms the integer representing the words into dense vectors.
The first model will be a simple bag-of-word DNN model that averages up the word vectors and feeds the tensor that results to further dense layers. Doing so means that we forget the word order (and hence that we consider sentences as a “bag-of-words”). In the second and in the third model we will keep the information about the word order using a simple RNN and a simple CNN allowing us to achieve the same performance as with the DNN model but in much fewer epochs.
```
# Ensure the right version of Tensorflow is installed.
!pip freeze | grep tensorflow==2.0 || pip install tensorflow==2.0
import os
import shutil
import pandas as pd
import tensorflow as tf
from tensorflow.keras.callbacks import TensorBoard, EarlyStopping
from tensorflow.keras.layers import (
Embedding,
Flatten,
GRU,
Conv1D,
Lambda,
Dense,
)
from tensorflow.keras.models import Sequential
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.utils import to_categorical
print(tf.__version__)
%matplotlib inline
```
Let's start by specifying where the information about the trained models will be saved as well as where our dataset is located:
```
LOGDIR = "./text_models"
DATA_DIR = "./data"
```
## Loading the dataset
Our dataset consists of titles of articles along with the label indicating from which source these articles have been taken from (GitHub, Tech-Crunch, or the New-York Times).
```
DATASET_NAME = "titles_full.csv"
TITLE_SAMPLE_PATH = os.path.join(DATA_DIR, DATASET_NAME)
COLUMNS = ['title', 'source']
titles_df = pd.read_csv(TITLE_SAMPLE_PATH, header=None, names=COLUMNS)
titles_df.head()
```
## Integerize the texts
The first thing we need to do is to find how many words we have in our dataset (`VOCAB_SIZE`), how many titles we have (`DATASET_SIZE`), and what the maximum length of the titles we have (`MAX_LEN`) is. Keras offers the `Tokenizer` class in its `keras.preprocessing.text` module to help us with that:
```
tokenizer = Tokenizer()
tokenizer.fit_on_texts(titles_df.title)
integerized_titles = tokenizer.texts_to_sequences(titles_df.title)
integerized_titles[:3]
VOCAB_SIZE = len(tokenizer.index_word)
VOCAB_SIZE
DATASET_SIZE = tokenizer.document_count
DATASET_SIZE
MAX_LEN = max(len(sequence) for sequence in integerized_titles)
MAX_LEN
```
Let's now implement a function `create_sequence` that will
* take as input our titles as well as the maximum sentence length and
* returns a list of the integers corresponding to our tokens padded to the sentence maximum length
Keras has the helper functions `pad_sequence` for that on the top of the tokenizer methods.
```
# TODO 1
def create_sequences(texts, max_len=MAX_LEN):
sequences = tokenizer.texts_to_sequences(texts)
padded_sequences = pad_sequences(sequences, max_len, padding='post')
return padded_sequences
sequences = create_sequences(titles_df.title[:3])
sequences
titles_df.source[:4]
```
We now need to write a function that
* takes a title source and
* returns the corresponding one-hot encoded vector
Keras `to_categorical` is handy for that.
```
CLASSES = {
'github': 0,
'nytimes': 1,
'techcrunch': 2
}
N_CLASSES = len(CLASSES)
# TODO 2
def encode_labels(sources):
classes = [CLASSES[source] for source in sources]
one_hots = to_categorical(classes)
return one_hots
encode_labels(titles_df.source[:4])
```
## Preparing the train/test splits
Let's split our data into train and test splits:
```
N_TRAIN = int(DATASET_SIZE * 0.80)
titles_train, sources_train = (
titles_df.title[:N_TRAIN], titles_df.source[:N_TRAIN])
titles_valid, sources_valid = (
titles_df.title[N_TRAIN:], titles_df.source[N_TRAIN:])
```
To be on the safe side, we verify that the train and test splits
have roughly the same number of examples per classes.
Since it is the case, accuracy will be a good metric to use to measure
the performance of our models.
```
sources_train.value_counts()
sources_valid.value_counts()
```
Using `create_sequence` and `encode_labels`, we can now prepare the
training and validation data to feed our models.
The features will be
padded list of integers and the labels will be one-hot-encoded 3D vectors.
```
X_train, Y_train = create_sequences(titles_train), encode_labels(sources_train)
X_valid, Y_valid = create_sequences(titles_valid), encode_labels(sources_valid)
X_train[:3]
Y_train[:3]
```
## Building a DNN model
The build_dnn_model function below returns a compiled Keras model that implements a simple embedding layer transforming the word integers into dense vectors, followed by a Dense softmax layer that returns the probabilities for each class.
Note that we need to put a custom Keras Lambda layer in between the Embedding layer and the Dense softmax layer to do an average of the word vectors returned by the embedding layer. This is the average that's fed to the dense softmax layer. By doing so, we create a model that is simple but that loses information about the word order, creating a model that sees sentences as "bag-of-words".
```
def build_dnn_model(embed_dim):
model = Sequential([
Embedding(VOCAB_SIZE + 1, embed_dim, input_shape=[MAX_LEN]), # TODO 3
Lambda(lambda x: tf.reduce_mean(x, axis=1)), # TODO 4
Dense(N_CLASSES, activation='softmax') # TODO 5
])
model.compile(
optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy']
)
return model
```
Below we train the model on 100 epochs but adding an `EarlyStopping` callback that will stop the training as soon as the validation loss has not improved after a number of steps specified by `PATIENCE` . Note that we also give the `model.fit` method a Tensorboard callback so that we can later compare all the models using TensorBoard.
```
%%time
tf.random.set_seed(33)
MODEL_DIR = os.path.join(LOGDIR, 'dnn')
shutil.rmtree(MODEL_DIR, ignore_errors=True)
BATCH_SIZE = 300
EPOCHS = 100
EMBED_DIM = 10
PATIENCE = 0
dnn_model = build_dnn_model(embed_dim=EMBED_DIM)
dnn_history = dnn_model.fit(
X_train, Y_train,
epochs=EPOCHS,
batch_size=BATCH_SIZE,
validation_data=(X_valid, Y_valid),
callbacks=[EarlyStopping(patience=PATIENCE), TensorBoard(MODEL_DIR)],
)
pd.DataFrame(dnn_history.history)[['loss', 'val_loss']].plot()
pd.DataFrame(dnn_history.history)[['accuracy', 'val_accuracy']].plot()
dnn_model.summary()
```
## Building a RNN model
The `build_dnn_model` function below returns a compiled Keras model that implements a simple RNN model with a single `GRU` layer, which now takes into account the word order in the sentence.
The first and last layers are the same as for the simple DNN model.
Note that we set `mask_zero=True` in the `Embedding` layer so that the padded words (represented by a zero) are ignored by this and the subsequent layers.
```
def build_rnn_model(embed_dim, units):
model = Sequential([
Embedding(VOCAB_SIZE + 1, embed_dim, input_shape=[MAX_LEN], mask_zero=True), # TODO 3
GRU(units), # TODO 5
Dense(N_CLASSES, activation='softmax')
])
model.compile(
optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy']
)
return model
```
Let's train the model with early stoping as above.
Observe that we obtain the same type of accuracy as with the DNN model, but in less epochs (~3 v.s. ~20 epochs):
```
%%time
tf.random.set_seed(33)
MODEL_DIR = os.path.join(LOGDIR, 'rnn')
shutil.rmtree(MODEL_DIR, ignore_errors=True)
EPOCHS = 100
BATCH_SIZE = 300
EMBED_DIM = 10
UNITS = 16
PATIENCE = 0
rnn_model = build_rnn_model(embed_dim=EMBED_DIM, units=UNITS)
history = rnn_model.fit(
X_train, Y_train,
epochs=EPOCHS,
batch_size=BATCH_SIZE,
validation_data=(X_valid, Y_valid),
callbacks=[EarlyStopping(patience=PATIENCE), TensorBoard(MODEL_DIR)],
)
pd.DataFrame(history.history)[['loss', 'val_loss']].plot()
pd.DataFrame(history.history)[['accuracy', 'val_accuracy']].plot()
rnn_model.summary()
```
## Build a CNN model
The `build_dnn_model` function below returns a compiled Keras model that implements a simple CNN model with a single `Conv1D` layer, which now takes into account the word order in the sentence.
The first and last layers are the same as for the simple DNN model, but we need to add a `Flatten` layer betwen the convolution and the softmax layer.
Note that we set `mask_zero=True` in the `Embedding` layer so that the padded words (represented by a zero) are ignored by this and the subsequent layers.
```
def build_cnn_model(embed_dim, filters, ksize, strides):
model = Sequential([
Embedding(VOCAB_SIZE + 1, embed_dim, input_shape=[MAX_LEN], mask_zero=True), # TODO 3
Conv1D( # TODO 5
filters=filters,
kernel_size=ksize,
strides=strides,
activation='relu',
),
Flatten(), # TODO 5
Dense(N_CLASSES, activation='softmax')
])
model.compile(
optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy']
)
return model
```
Let's train the model.
Again we observe that we get the same kind of accuracy as with the DNN model but in many fewer steps.
```
%%time
tf.random.set_seed(33)
MODEL_DIR = os.path.join(LOGDIR, 'cnn')
shutil.rmtree(MODEL_DIR, ignore_errors=True)
EPOCHS = 100
BATCH_SIZE = 300
EMBED_DIM = 5
FILTERS = 200
STRIDES = 2
KSIZE = 3
PATIENCE = 0
cnn_model = build_cnn_model(
embed_dim=EMBED_DIM,
filters=FILTERS,
strides=STRIDES,
ksize=KSIZE,
)
cnn_history = cnn_model.fit(
X_train, Y_train,
epochs=EPOCHS,
batch_size=BATCH_SIZE,
validation_data=(X_valid, Y_valid),
callbacks=[EarlyStopping(patience=PATIENCE), TensorBoard(MODEL_DIR)],
)
pd.DataFrame(cnn_history.history)[['loss', 'val_loss']].plot()
pd.DataFrame(cnn_history.history)[['accuracy', 'val_accuracy']].plot()
cnn_model.summary()
```
### Comparing the models
At last, let's compare all the models we have trained at once using TensorBoard in order
to choose the one that overfits the less for the same performance level.
Running the following command will launch TensorBoard on port 6006. This will
block the notebook execution, so you'll have to interrupt that cell first before
you can run other cells.
```
!tensorboard --logdir $LOGDIR --port 6006
```
Copyright 2019 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| github_jupyter |
```
library(ggplot2)
library(dplyr)
library(reshape2)
library(Matrix)
library(gridExtra)
# You may need to set these to your own local paths.
project_directory <- file.path(Sys.getenv("GIT_REPO_LOC"), "MicrocreditLRVB/inst/simulated_data")
raw_data_directory <- file.path(Sys.getenv("GIT_REPO_LOC"), "microcredit_vb/data")
# Read in the data that was produced by the R script.
csv_data <- read.csv(file.path(raw_data_directory, "microcredit_data_processed.csv"))
# The number of distinct groups.
n_g <- max(csv_data$site)
# Get the observations and the total number of observations.
y <- csv_data$profit
y_g <- as.integer(csv_data$site)
# The x array will indicate which rows should also get the
# treatment effect. The model accomodates generic x, but for
# this application we only need indicators.
x <- cbind(rep(1, length(y)), as.numeric(csv_data$treatment))
data_df <- data.frame(x=x, y=y, y_g=y_g)
data_df_transform <-
data_df %>%
mutate(zero_y=abs(y) < 1e-8)
# Note that 1e-8 appears to be a good filter -- we don't have any
# nearly zero entries until ~1e-1, above which they're probably actual nonzero values.
min(abs(filter(data_df_transform, !zero_y)$y))
#######################
# Look at qqplots and raw distributions with outlier trimming. Even after trimming, they do not look very normal.
trim_level <- 0.3
y_quantiles <-
filter(data_df_transform, !zero_y) %>%
group_by(y_g) %>%
summarize(qlower=quantile(y, trim_level), qupper=quantile(y, 1 - trim_level))
data_df_trim <-
filter(data_df_transform, !zero_y) %>%
inner_join(y_quantiles, by="y_g") %>%
filter(y < qupper & y > qlower) %>%
group_by(y_g) %>%
arrange(y) %>%
mutate(q=(1:length(y)) / length(y), norm=qnorm(q))
# Qqplots
ggplot(filter(data_df_trim)) +
geom_point(aes(x=norm, y=y)) +
facet_grid(y_g ~ ., scales="free")
ggplot(filter(data_df_trim)) +
geom_histogram(aes(x=y, y=..ndensity..), bins=20) +
facet_grid(~ y_g, scales="free") +
geom_vline(aes(xintercept=0))
###################
# Look at cumulative distributions to check for power law behavior.
# If it is a power law, the cumulative distribution will be a straight line
# with slope given by the power law coefficient plus one.
quantile_list <- list()
for (group in 1:max(data_df_transform$y_g)) { for (y_sign in c(-1, 1)) { for (arm in c(0, 1)) {
rows <- with(data_df_transform, (y_g == group) & (!zero_y) & (y * y_sign > 0) & (x.2 == arm))
if (sum(rows) > 0) {
quantile_df <- data.frame(y=sort(y_sign * data_df_transform[rows, ]$y),
quantile=(sum(rows):1) / sum(rows),
group=group, y_sign=y_sign, arm=arm)
quantile_list[[length(quantile_list) + 1]] <- quantile_df
}
}}}
quantile_df <- do.call(rbind, quantile_list)
ggplot(quantile_df) +
geom_point(aes(x=y_sign * log10(y), y=log10(quantile), color=factor(arm))) +
facet_grid(group ~ y_sign) +
ggtitle("Overlaid arms")
ggplot(quantile_df) +
geom_point(aes(x=log10(y), y=log10(quantile), color=factor(y_sign))) +
facet_grid(group ~ arm) +
ggtitle("Overlaid signs")
ggplot(quantile_df) +
geom_point(aes(x=log10(y), y=log10(quantile), color=paste(arm, y_sign))) +
facet_grid(group ~ .) +
ggtitle("Everything overlaid")
##########################################
# Execute box-cox transforms and save.
# A good reference for the Box-Cox transform:
# https://www.ime.usp.br/~abe/lista/pdfm9cJKUmFZp.pdf
library(MASS)
data_df_transform <-
data_df %>%
mutate(zero_y=abs(y) < 1e-8)
# Non-zero values of y_trans will be sent in the loop below.
data_df_transform$y_trans <- 0.0
data_df_transform$lambda <- NaN
for (group in 1:max(y_g)) { for (y_sign in c(-1, 1)) {
rows <- with(data_df_transform, (y_g == group) & (!zero_y) & (y * y_sign > 0))
bc_y <- y_sign * data_df_transform[rows, ]$y
if (length(bc_y) > 0) {
# The MASS boxcox function is pretty primitive. Better to do it yourself with optim.
bc <- boxcox(bc_y ~ 1, plotit=FALSE, lambda=seq(-1, 1, 0.001))
lambda <- bc$x[which.max(bc$y)]
if (abs(lambda) < 0.001) {
lambda <- 0
}
if (lambda == 0) {
y_trans <- log(bc_y)
} else {
y_trans <- ((bc_y ^ lambda) - 1) / lambda
}
qqnorm(y_trans, main=lambda)
#readline(prompt="Press [enter] to continue")
data_df_transform[rows, "y_trans"] <- y_sign * y_trans
data_df_transform[rows, "lambda"] <- lambda
}
}}
ggplot(filter(data_df_transform, !zero_y)) +
geom_histogram(aes(x=y_trans, y=..density..), bins=100) +
facet_grid(y_g ~ .)
# Look at the lambdas chosen. They are similar mostly except for group 2.
mutate(data_df_transform, y_pos=y > 0) %>%
filter(!zero_y) %>%
group_by(y_g, y_pos) %>%
summarize(lambda=unique(lambda))
```
| github_jupyter |
```
%matplotlib inline
from pathlib import Path
import requests
DATA_PATH = Path("data")
PATH = DATA_PATH / "mnist"
PATH.mkdir(parents=True, exist_ok=True)
URL = "http://deeplearning.net/data/mnist/"
FILENAME = "mnist.pkl.gz"
if not (PATH / FILENAME).exists():
content = requests.get(URL + FILENAME).content
(PATH / FILENAME).open("wb").write(content)
import pickle
import gzip
with gzip.open((PATH / FILENAME).as_posix(), "rb") as f:
((x_train, y_train), (x_valid, y_valid), _) = pickle.load(f, encoding="latin-1")
print(x_train, y_train)
print(x_valid, y_valid)
from matplotlib import pyplot
import numpy as np
pyplot.imshow(x_train[0].reshape((28, 28)), cmap="gray")
print(x_train.shape)
print(x_train)
import torch
x_train, y_train, x_valid, y_valid = map(
torch.tensor, (x_train, y_train, x_valid, y_valid)
)
n, c = x_train.shape
x_train, x_train.shape, y_train.min(), y_train.max()
print(x_train, y_train)
print(x_train.shape)
print(y_train.min(), y_train.max())
```
## Neural network from scratch
```
import math
weights = torch.randn(784, 10) / math.sqrt(784)
print(weights.shape)
weights.requires_grad_()
bias = torch.zeros(10, requires_grad=True)
def log_softmax(x):
return x - x.exp().sum(-1).log().unsqueeze(-1)
def model(xb):
return log_softmax(xb @ weights + bias)
bs = 64 # batch size
xb = x_train[0:bs] # a mini-batch from x
preds = model(xb) # predictions
preds[0], preds.shape
print(preds[0], preds.shape)
def nll(input, target):
return -input[range(target.shape[0]), target].mean()
loss_func = nll
yb = y_train[0:bs]
print(loss_func(preds, yb))
def accuracy(out, yb):
preds = torch.argmax(out, dim=1)
print(preds)
return (preds == yb).float().mean()
#print(preds)
print(yb)
print(accuracy(preds, yb))
from IPython.core.debugger import set_trace
lr = 0.5 # learning rate
epochs = 2 # how many epochs to train for
for epoch in range(epochs):
for i in range((n - 1) // bs + 1):
# set_trace()
start_i = i * bs
end_i = start_i + bs
# print(f"start: {start_i} end: {end_i}")
xb = x_train[start_i:end_i]
yb = y_train[start_i:end_i]
pred = model(xb)
loss = loss_func(pred, yb)
loss.backward()
with torch.no_grad():
weights -= weights.grad * lr
bias -= bias.grad * lr
weights.grad.zero_()
bias.grad.zero_()
print(n)
print(loss_func(model(xb), yb), accuracy(model(xb), yb))
```
## Using torch.nn.functional
```
import torch.nn.functional as F
loss_func = F.cross_entropy
def model(xb):
return xb @ weights + bias
print(loss_func(model(xb), yb), accuracy(model(xb), yb))
from torch import nn
class Mnist_Logistic(nn.Module):
def __init__(self):
super().__init__()
self.weights = nn.Parameter(torch.randn(784, 10) / math.sqrt(784))
self.bias = nn.Parameter(torch.zeros(10))
def forward(self, xb):
return xb @ self.weights + self.bias
model = Mnist_Logistic()
print(loss_func(model(xb), yb))
def fit():
for epoch in range(epochs):
for i in range((n - 1) // bs + 1):
start_i = i * bs
end_i = start_i + bs
xb = x_train[start_i:end_i]
yb = y_train[start_i:end_i]
pred = model(xb)
loss = loss_func(pred, yb)
loss.backward()
with torch.no_grad():
for p in model.parameters():
p -= p.grad * lr
model.zero_grad()
fit()
print(loss_func(model(xb), yb))
```
## Refactor using nn.Linear
```
nn.Linear?
class Mnist_Logistic(nn.Module):
def __init__(self):
super().__init__()
self.lin = nn.Linear(784, 10)
def forward(self, xb):
return self.lin(xb)
nn.Conv2d?
```
| github_jupyter |
# Introduction to Generative ML with Pyro
```
import matplotlib.pyplot as plt
import torch
import pyro
pyro.set_rng_seed(101)
```
Our goal is to understand causal modeling within the context of generative machine learning. We just examined one generative machine learning framework called Bayesian networks (BNs) and how we can use BNs as causal models.
**Bayesian Networks (BNs)** Framework that defines a probabilistic generative model of the world in terms of a directed acyclic graph.
**causal Bayesian networks:** Bayesian networks where the direction of edges in the DAG represent causality.
Bayesian networks provide a general-purpose framework for representing a causal data generating story for how the world works.
Now we will introduce probabilistic programming, a framework that is more expressive than Bayesian networks.
### What is a probabilistic programming language?
"A probabilistic programming language (PPL) is a programming language designed to describe probabilistic models and then perform inference in those models. PPLs are closely related to graphical models and Bayesian networks but are more expressive and flexible. Probabilistic programming represents an attempt to "Unify general purpose programming" with probabilistic modeling."
-Wikipedia
A PPL is a domain-specific programming language for that lets you write a data generating story as a program. As with a causal Bayesian network, you can write your program in a way that orders the steps of its execution according to cause and effect.
### How exactly do Bayesian networks and probabilistic programming differ?
**Representation of relationships between variables**. BNs restricted to representing the relationships between variables in terms of conditional probability distributions (CPDs) factored according to a DAG. Frameworks typically limit you to a small set of parametric CPDs (e.g., Gaussian, multinomial).
Just as computer programs are more expressive than flow charts, PPLs let you represent relations any way you like so long as you can represent them in code. PPL relationships can include control flow and recursion. In causal models, we will see that this allows you to be more specific about mechanism than you can with CPDs.
**DAG vs. open world models**. BNs restrict the representation of the joint distribution to a DAG. This constraint enables you to reason easily about the joint distribution through graph-theoretic operations like d-separation. PPLs need not be constrained to a DAG. For example (using an imaginary Python PPL package):
X = Bernoulli(p)
if X == 1:
Y = Gaussian(0, 1)
In a DAG, you have a fixed set of variables, i.e. a "closed world". In the above model, the variable Y is only instantiated if X==1. Y may or may not exist depending on how the generative process unfolds. For a more extreme example, consider this:
X = Poisson(λ)
Y = [Gaussian(0, 1)]
for i in range(1, X):
Y[i] = Gaussian(Y[i-1], 1))
Here you have the total number of Y variables itself being a random variable X. Further, the mean of the ith Y is a random variable given by the i-1th Y. You can't do that with a Bayes net! Unfortunately, we can't reason about this as directly as we can with a DAG. For example, recall that with the DAG, we had a convenient algorithm called `CPDAG` that converts the DAG to a partially directed acyclic graph structure called a PDAG that provides a compact representation of all the DAGs in an equivalence class. How might we define an equivalence class on this program? Certainly, enumerating all programs with an equivalent representation of the joint distribution would be very difficult even with constraints on the length of the program. In general, enumerating all programs of minimal description that provide equivalent representations of a joint distribution is an NP-hard problem.
**Inference** When you have a DAG and a constrained set of parametric CPDs, as well as constraints on the kind of inference, queries the user can make, you can implement some inference algorithms in your BN framework that will generally work in a reasonable amount of time.
PPLs are more flexible than BNs, but the trade-off s that getting inference to work is harder. PPL's develop several abstractions for inference and leave it to the user to apply them, requiring the user to become something of an expert in inference algorithms. PPL developers make design decisions to make inference easier for the user, though this often sacrifices some flexibility. One emergent pattern is to build PPLs on tensor-based frameworks like Tensorflow and PyTorch. Tensor-based PPLs allow a data scientist with experience building deep learning models to rely on that experience when doing inference.

[$$\texttt{Kevin Smith - Tutorial: Probabilistic Programming}$$](https://www.youtube.com/watch?v=9SEIYh5BCjc)
## Introduction to Pyro
Pyro is a universal probabilistic programming language (PPL) written in Python and supported by PyTorch on the backend. Pyro enables flexible and expressive deep probabilistic modeling, unifying the best of modern deep learning and Bayesian modeling.
Our purpose of this class, pyro has "do"-operator that allows intervention and counterfactual inference in these probabilistic models.
### Stochastic Functions
The basic unit of probabilistic programs is the stochastic function. A stochastic function is an arbitrary Python callable that combines two ingredients:
- deterministic Python code; and
- primitive stochastic functions that call a random number generator
For this course, we will consider these stochastic functions as **models**. Stochastic functions can be used to represent simplified or abstract descriptions of a data-generating process.
### Primitive stochastic functions
We call them distributions. We can explicitly compute the probability of the outputs given the inputs.
```
loc = 0. # mean zero
scale = 1. # unit variance
normal = torch.distributions.Normal(loc, scale) # create a normal distribution object
x = normal.rsample() # draw a sample from N(0,1)
print("sample: ", x)
```
Pyro simplifies this process of sampling from distributions. It uses `pyro.sample()`.
```
x = pyro.sample("my_sample", pyro.distributions.Normal(loc, scale))
print(x)
```
Just like a direct call to `torch.distributions.Normal().rsample()`, this returns a sample from the unit normal distribution. The crucial difference is that this sample is named. Pyro’s backend uses these names to uniquely identify sample statements and change their behavior at runtime depending on how the enclosing stochastic function is being used. This is how Pyro can implement the various manipulations that underlie inference algorithms.
Let's write a simple `weather` model.
### A simple model
```
import pyro.distributions as dist
def weather():
cloudy = pyro.sample('cloudy', dist.Bernoulli(0.3))
cloudy = 'cloudy' if cloudy.item() == 1.0 else 'sunny'
mean_temp = {'cloudy': 55.0, 'sunny': 75.0}[cloudy]
scale_temp = {'cloudy': 10.0, 'sunny': 15.0}[cloudy]
temp = pyro.sample('temp', dist.Normal(mean_temp, scale_temp))
return cloudy, temp.item()
for _ in range(3):
print(weather())
```
First two lines introduce a binary variable `cloudy`, which is given by a draw from the Bernoulli distribution with a parameter of $0.3$. The Bernoulli distribution returns either $0$ or $1$, line `2` converts that into a string. So, So according to this model, $30%$ of the time it’s cloudy and $70%$ of the time it’s sunny.
In line `4` and `5`, we initialize mean and scale of the temperature for both values. We then sample, the temperature from a Normal distribution and return that along with `cloudy` variable.
We can build complex model by modularizing and reusing the concepts into functions and use them as programmers use functions.
```
def ice_cream_sales():
cloudy, temp = weather()
expected_sales = 200. if cloudy == 'sunny' and temp > 80.0 else 50.
ice_cream = pyro.sample('ice_cream', pyro.distributions.Normal(expected_sales, 10.0))
return ice_cream
```
## Inference
As we discussed earlier, the reason we use PPLs is because they can easily go backwards and reason about cause given the observed effect. There are myriad of inference algorithms available in pyro. Let's try it on an even simpler model.
$$weight \mid guess \sim \mathcal{N}(guess, 1)$$
$$measurement \mid guess, weight \sim \mathcal{N}(weight, 0.75)$$
```
def scale(guess):
weight = pyro.sample("weight", dist.Normal(guess, 1.0))
measurement = pyro.sample("measurement", dist.Normal(weight, 0.75))
return measurement
scale(14.)
```
Suppose we observe that the `measurement` of an object was $14$ lbs. What would have we guessed if we tried to guess it's `weight` first?
This question is answered in two steps.
1. Condition the model.
```
conditioned_scale = pyro.condition(scale,{"measurement": torch.tensor(14.)})
```
2. Set the prior and infer the posterior. We will use the Hamiltonian Monte Carlo (HMC) inference algorithm with the no U-turn sampler (NUTS), like you would in a language like Stan.
```
from pyro.infer import NUTS, MCMC, EmpiricalMarginal
guess_prior = torch.tensor(13.)
nuts_kernel = NUTS(conditioned_scale, adapt_step_size=True)
posterior = MCMC(nuts_kernel,
num_samples=1000,
warmup_steps=300)
posterior.run(guess_prior)
marginal = posterior.get_samples()['weight'] # get_samples returns a dictionary keyed by the site name
plt.hist(marginal)
plt.title("P(weight | measurement = 14)")
plt.xlabel("Weight")
plt.ylabel("#")
print(posterior)
```
### Importance Sampling for inferring discrete quantities
Let us consider the following Bayesian network and the probabilities associated with each of the events as shown

Implementing the bayesian net in pyro :
#### Shapes in distribution:
We know that PyTorch tensors have single `shape` attribute, `Distribution`s have two shape attributes with special meaning.
* `.batch_shape`: Indices over `.batch_shape` denote conditionally independent random variables,
* `.event_shape`: indices over `.event_shape` denote dependent random variables (ie one draw from a distribution).
These two combine to define the total shape of a sample. Thus the total shape of `.log_prob()` of distribution is `.batch_shape`.
Also, `Distribution.sample()` also has a `sample_shape` attribute that indexes over independent and identically distributed(iid) random variables.
```
| iid | independent | dependent
------+--------------+-------------+------------
shape = sample_shape + batch_shape + event_shape
```
To learn more see [broadcasting tensors in PyTorch](https://pytorch.org/docs/master/notes/broadcasting.html).
### Examples
One way to introduce batch_shape is use `expand`.
```{python}
d = dist.MultivariateNormal(torch.zeros(3), torch.eye(3, 3)).expand([5]) # expand - 3 of these Multivariate Normal Dists
print("batch_shape: ", d.batch_shape)
print("event_shape: ", d.event_shape)
#x = d.sample(torch.Size([5]))
x = d.sample()
print("x shape: ", x.shape) # == sample_shape + batch_shape + event_shape
print("d.log_prob(x) shape:", d.log_prob(x).shape) # == batch_shape
```
The other way is using `plate` context manager.
Pyro models can use the context manager `pyro.plate` to declare that certain batch dimensions are independent. Inference algorithms can then take advantage of this independence to e.g. construct lower variance gradient estimators or to enumerate in linear space rather than exponential space.
```
def model():
burglar = pyro.sample("B", dist.Categorical(torch.tensor([0.999, 0.001])))
earthquake = pyro.sample("E", dist.Categorical(torch.tensor([0.998, 0.002])))
alarm_probs = torch.tensor([[[0.999, 0.001],[0.71,0.29]],[[0.06, 0.94],[0.05, 0.95]]])
alarm = pyro.sample("A", dist.Categorical(alarm_probs[burglar][earthquake]))
john_probs = torch.tensor([[0.95, 0.05], [0.1, 0.9]])
mary_probs = torch.tensor([[0.99, 0.01], [0.3, 0.7]])
john_calls = pyro.sample("J", dist.Categorical(john_probs[alarm]))
mary_calls = pyro.sample("M", dist.Categorical(mary_probs[alarm]))
```
Let's say we know that the alarm went off (alarm = 1 ) and Mary called. Can we use the model to infer about the possibility of an earthquake ?
```
# condition on the model with observation/evidence
conditioned_model = pyro.condition(model, data = {'M': torch.tensor(1), 'A': torch.tensor(1)})
```
Perform posterior inference using Importance sampling
```
posterior = pyro.infer.Importance(conditioned_model, num_samples=1000).run()
# get the marginal distribution of the variable we are interested in (In our case its earthquake)
marginal = pyro.infer.EmpiricalMarginal(posterior, "E")
earthquake_samples = np.array([marginal().item() for _ in range(1000)])
e_unique, e_counts = np.unique(earthquake_samples, return_counts=True)
plt.xlabel = "Earthquake"
plt.ylabel = "Count"
plt.bar(e_unique, e_counts)
plt.xticks(e_unique)
plt.title('Earthquake|Alarm=On, Mary Called')
```
What is the probability that there was an earthquake , given alarm went off
and Mary called?
```
print(earthquake_samples.mean())
```
### Gaussian Mixture Model

[$$\texttt{Blei - Build, Compute, Critique, Repeat:Data Analysis with Latent Variable Models}$$](http://www.cs.columbia.edu/~blei/papers/Blei2014b.pdf)
Here is an example of how to implement a Gaussian mixture model in Pyro.
```
from __future__ import print_function
import os
from collections import defaultdict
import numpy as np
import scipy.stats
import torch
from torch.distributions import constraints
from pyro import poutine
from pyro.contrib.autoguide import AutoDelta
from pyro.optim import Adam
from pyro.infer import SVI, TraceEnum_ELBO, config_enumerate, infer_discrete
from matplotlib import pyplot
# %matplotlib inline
pyro.enable_validation(True)
data = torch.tensor([0., 1., 10., 11., 12.])
K = 2 # Fixed number of components.
@config_enumerate
def model(data):
# Global variables.
weights = pyro.sample('weights', dist.Dirichlet(0.5 * torch.ones(K)))
scale = pyro.sample('scale', dist.LogNormal(0., 2.))
with pyro.plate('components', K):
locs = pyro.sample('locs', dist.Normal(0., 10.))
with pyro.plate('data', len(data)):
# Local variables.
assignment = pyro.sample('assignment', dist.Categorical(weights))
pyro.sample('obs', dist.Normal(locs[assignment], scale), obs=data)
```
In fact, we can also nest `plates`. The only thing we need to care about is, which dimensions are independent. Pyro automatically manages this but sometimes we need to explicitely specify the dimensions. Once we specify that, we can leverage PyTorch's CUDA enabled capabilities to run inference on GPUs.
```
with pyro.plate("x_axis", 320):
# within this context, batch dimension -1 is independent
with pyro.plate("y_axis", 200):
# within this context, batch dimensions -2 and -1 are independent
```
Note that we always count from the right by using negative indices like $-2$, $-1$.
```
with pyro.plate("x_axis", 5):
d = dist.MultivariateNormal(torch.zeros(3), torch.eye(3, 3))
x = pyro.sample("x", d)
x.shape
```
### Review of Approximate Inference
We have variables $Z$s (cluster assignments) and $X$s (data points) in our mixture model, where $X$ is observed and $Z$ is latent (unobserved). As we saw earlier, a generative model entails a joint distribution
$$p(Z,X)$$
Inference of unknown can be achieved through conditioning on the observations.
$$p(Z \mid X) = \frac{p(Z, X)}{p(X)}$$
For non-trivial interesting problems, the integral for the denominator(marginal) is not tractable.
$$p(X) = \int dZp(X \mid Z)p(Z)$$
So we have to directly approximate $p(Z \mid X)$. The most popular way of doing this is with *stochastic variational inference* because it can make use of the loss function optimization infrastructure in PyTorch.
### Variational Inference:
We can't compute $p(Z \mid X)$ directly, so let's approximate with some other distribution $q(Z; \nu)$ over Z that is tractable (for example, Gaussions or other exponential family).

[$$\texttt{David Blei - Variational Inference (NeurIPS 2016 Tutorial)}$$](https://www.youtube.com/watch?v=ogdv_6dbvVQ)
Since q is tractable, we can play with it's parameter $\nu$ such that it reaches as close to $p(Z\mid X)$ as possible. More precisely, we want to minimize the KL divergence between $q$ and $p$. With this trick, we just turned an **inference** problem to an **optimization** problem!
$$
\begin{align*}
KL(q(Z;\nu) \mid\mid p(Z\mid X)) &= -\int dZ\ q(Z) \log\frac{P(Z\mid X)}{q(Z)}\\
&= -\int dZ\ q(Z) \log
\frac{\frac{p(Z,X)}{p(X)}}{q(Z)}\\
&= -\int dZ\ q(Z) \log
\frac{p(Z,X)}{p(X)q(Z)}\\
&= -\int dZ\ q(Z)
\left[
\log
\frac{p(Z,X)}{q(Z)}
- \log p(X)
\right]\\
&= - \int dZ\ \log \frac{p(Z,X)}{q(Z)} + \underbrace{\int dZ\ q(Z)}_{\text{=1}}\log p(X)\\
&= - \int dZ\ \log \frac{p(Z,X)}{q(Z)} + \log p(X)\\
\log p(X) &= KL(q(Z;\nu)\mid\mid p(Z\mid X) + \underbrace{\int dZ\ q(Z;\nu) \log \frac{p(Z,X)}{q(Z;\nu)}}_{\mathcal{L}}\\
\end{align*}
$$
Note that we already observed $X$ and we conditioned the model to get $p(Z \mid X)$. But given $X$, $\log p(X)$ is constant! So, minimizing KL is equivalent to maximizing $\mathcal{L}$.
**How do you maximize $\mathcal{L}$**? Take $\nabla_{\nu} \mathcal{L}$.
$\mathcal{L}$ is called **variational lower bound**. It is often called ELBO.
**Stochastic Variational Inference** scales variational inference to massive data. Just like in stochastic variational inference, you subsample the data and update the posterior!
<!-- #region -->
### Stochastic Optimization
In stochastic optimization, we replace the gradient with cheaper noisy estimate which is guranteed to converge to a local optimum.
$$\nu_{t+1} = \nu_t + \rho_t \hat{\nabla}_{\nu} \mathcal{L}(\nu_t)$$
Requirements:
* Unbiased gradients, i.e.
$$\mathbb{E}[\hat{\nabla}_{\nu} \mathcal{L}(\nu_t)] = \nabla_{\nu}\mathcal{L}(\nu)$$
* Step-size sequence $\rho_t$ that follows Robbins-Monro conditions.
Stochastic variational inference takes inspiration from stochastic optimization and natural graidient. We follow the same procedure as stochastic gradient descent.
<!-- #endregion -->
### A Rough Stochastic variational inference algorithm:
* Initialize $q$ with some $\nu$
* Until Converge:
* subsample from Data:
* compute gradient $\hat{\nabla_{\nu}}\mathcal{L}_{\nu_t}$
* update global parameter $\nu_{t+1} = \nu_t + \rho_t \hat{\nabla_{\nu}}\mathcal{L}_{\nu_t}$
* Return $q(Z;\nu)$
### Training a MAP estimator
Let's start by learning model parameters `weights`, `locs`, and `scale` given priors and data. We will use `AutoDelta` guide function. Our model will learn global mixture weights, the location of each mixture component, and a shared scale that is common to both components.
During inference, `TraceEnum_ELBO` will marginalize out the assignments of datapoints to clusters.
`max_plate_nesting` lets Pyro know that we’re using the rightmost dimension plate and that Pyro can use any other dimension for parallelization.
## Some other Pyro vocabulary
* poutine - Beneath the built-in inference algorithms, Pyro has a library of composable effect handlers for creating new inference algorithms and working with probabilistic programs. Pyro’s inference algorithms are all built by applying these handlers to stochastic functions.
* poutine.block - blocks pyro premitives. By default, it blocks everything.
* param - Parameters in Pyro are basically thin wrappers around PyTorch Tensors that carry unique names. As such Parameters are the primary stateful objects in Pyro. Users typically interact with parameters via the Pyro primitive `pyro.param`. Parameters play a central role in stochastic variational inference, where they are used to represent point estimates for the parameters in parameterized families of models and guides.
* param_store - Global store for parameters in Pyro. This is basically a key-value store.
```
global_guide = AutoDelta(poutine.block(model, expose=['weights', 'locs', 'scale']))
optim = pyro.optim.Adam({'lr': 0.1, 'betas': [0.8, 0.99]})
elbo = TraceEnum_ELBO(max_plate_nesting=1)
svi = SVI(model, global_guide, optim, loss=elbo)
def initialize(seed):
pyro.set_rng_seed(seed)
pyro.clear_param_store()
# Initialize weights to uniform.
pyro.param('auto_weights', 0.5 * torch.ones(K), constraint=constraints.simplex)
# Assume half of the data variance is due to intra-component noise.
pyro.param('auto_scale', (data.var() / 2).sqrt(), constraint=constraints.positive)
# Initialize means from a subsample of data.
pyro.param('auto_locs', data[torch.multinomial(torch.ones(len(data)) / len(data), K)]);
loss = svi.loss(model, global_guide, data)
return loss
# Choose the best among 100 random initializations.
loss, seed = min((initialize(seed), seed) for seed in range(100))
initialize(seed)
print('seed = {}, initial_loss = {}'.format(seed, loss))
# Register hooks to monitor gradient norms.
gradient_norms = defaultdict(list)
for name, value in pyro.get_param_store().named_parameters():
value.register_hook(lambda g, name=name: gradient_norms[name].append(g.norm().item()))
losses = []
for i in range(200):
loss = svi.step(data)
losses.append(loss)
print('.' if i % 100 else '\n', end='')
plt.figure(figsize=(10,3), dpi=100).set_facecolor('white')
plt.plot(losses)
plt.xlabel('iters')
plt.ylabel('loss')
plt.yscale('log')
plt.title('Convergence of SVI');
map_estimates = global_guide(data)
weights = map_estimates['weights']
locs = map_estimates['locs']
scale = map_estimates['scale']
print('weights = {}'.format(weights.data.numpy()))
print('locs = {}'.format(locs.data.numpy()))
print('scale = {}'.format(scale.data.numpy()))
X = np.arange(-3,15,0.1)
Y1 = weights[0].item() * scipy.stats.norm.pdf((X - locs[0].item()) / scale.item())
Y2 = weights[1].item() * scipy.stats.norm.pdf((X - locs[1].item()) / scale.item())
plt.figure(figsize=(10, 4), dpi=100).set_facecolor('white')
plt.plot(X, Y1, 'r-')
plt.plot(X, Y2, 'b-')
plt.plot(X, Y1 + Y2, 'k--')
plt.plot(data.data.numpy(), np.zeros(len(data)), 'k*')
plt.title('Density of two-component mixture model')
plt.ylabel('probability density')
```
| github_jupyter |
## Tabular data handling
This module defines the main class to handle tabular data in the fastai library: [`TabularDataBunch`](/tabular.data.html#TabularDataBunch). As always, there is also a helper function to quickly get your data.
To allow you to easily create a [`Learner`](/basic_train.html#Learner) for your data, it provides [`tabular_learner`](/tabular.data.html#tabular_learner).
```
from fastai.gen_doc.nbdoc import *
from fastai.tabular import *
show_doc(TabularDataBunch)
```
The best way to quickly get your data in a [`DataBunch`](/basic_data.html#DataBunch) suitable for tabular data is to organize it in two (or three) dataframes. One for training, one for validation, and if you have it, one for testing. Here we are interested in a subsample of the [adult dataset](https://archive.ics.uci.edu/ml/datasets/adult).
```
path = untar_data(URLs.ADULT_SAMPLE)
df = pd.read_csv(path/'adult.csv')
valid_idx = range(len(df)-2000, len(df))
df.head()
cat_names = ['workclass', 'education', 'marital-status', 'occupation', 'relationship', 'race', 'sex', 'native-country']
dep_var = 'salary'
```
The initialization of [`TabularDataBunch`](/tabular.data.html#TabularDataBunch) is the same as [`DataBunch`](/basic_data.html#DataBunch) so you really want to use the facotry method instead.
```
show_doc(TabularDataBunch.from_df)
```
Optionally, use `test_df` for the test set. The dependent variable is `dep_var`, while the categorical and continuous variables are in the `cat_names` columns and `cont_names` columns respectively. If `cont_names` is None then we assume all variables that aren't dependent or categorical are continuous. The [`TabularProcessor`](/tabular.data.html#TabularProcessor) in `procs` are applied to the dataframes as preprocessing, then the categories are replaced by their codes+1 (leaving 0 for `nan`) and the continuous variables are normalized.
Note that the [`TabularProcessor`](/tabular.data.html#TabularProcessor) should be passed as `Callable`: the actual initialization with `cat_names` and `cont_names` is done during the preprocessing.
```
procs = [FillMissing, Categorify, Normalize]
data = TabularDataBunch.from_df(path, df, dep_var, valid_idx=valid_idx, procs=procs, cat_names=cat_names)
```
You can then easily create a [`Learner`](/basic_train.html#Learner) for this data with [`tabular_learner`](/tabular.data.html#tabular_learner).
```
show_doc(tabular_learner)
```
`emb_szs` is a `dict` mapping categorical column names to embedding sizes; you only need to pass sizes for columns where you want to override the default behaviour of the model.
```
show_doc(TabularList)
```
Basic class to create a list of inputs in `items` for tabular data. `cat_names` and `cont_names` are the names of the categorical and the continuous variables respectively. `processor` will be applied to the inputs or one will be created from the transforms in `procs`.
```
show_doc(TabularList.from_df)
show_doc(TabularList.get_emb_szs)
show_doc(TabularList.show_xys)
show_doc(TabularList.show_xyzs)
show_doc(TabularLine, doc_string=False)
```
An object that will contain the encoded `cats`, the continuous variables `conts`, the `classes` and the `names` of the columns. This is the basic input for a dataset dealing with tabular data.
```
show_doc(TabularProcessor)
```
Create a [`PreProcessor`](/data_block.html#PreProcessor) from `procs`.
## Undocumented Methods - Methods moved below this line will intentionally be hidden
```
show_doc(TabularProcessor.process_one)
show_doc(TabularList.new)
show_doc(TabularList.get)
show_doc(TabularProcessor.process)
show_doc(TabularList.reconstruct)
```
## New Methods - Please document or move to the undocumented section
| github_jupyter |
# Gaussian Mixture Model
This is a brief tutorial on training mixture models in Pyro. We'll focus on the mechanics of `config_enumerate()` and setting up mixture weights. To simplify matters, we'll train a trivial 1-D Gaussian model on a tiny 5-point dataset.
```
from __future__ import print_function
import os
from collections import defaultdict
import numpy as np
import scipy.stats
import torch
from torch.distributions import constraints
from matplotlib import pyplot
%matplotlib inline
import pyro
import pyro.distributions as dist
from pyro.optim import Adam
from pyro.infer import SVI, TraceEnum_ELBO, config_enumerate
smoke_test = ('CI' in os.environ)
pyro.enable_validation(True)
```
## Dataset
Here is our tiny dataset. It has five points.
```
data = torch.tensor([0., 1., 10., 11., 12.])
```
## Maximum likelihood approach
Let's start by optimizing model parameters `weights`, `locs`, and `scale`, rather than treating them as random variables with priors. Our model will learn global mixture weights, the location of each mixture component, and a shared scale that is common to both components. Our guide will learn soft assignment weights of each point.
Note that none of our parameters have priors. In this Maximum Likelihood approach we can embed our parameters directly in the model rather than the guide. This is equivalent to adding them in the guide as `pyro.sample(..., dist.Delta(...))` sites and using a uniform prior in the model.
```
K = 2 # Fixed number of components.
def model(data):
# Global parameters.
weights = pyro.param('weights', torch.ones(K) / K, constraint=constraints.simplex)
locs = pyro.param('locs', 10 * torch.randn(K))
scale = pyro.param('scale', torch.tensor(0.5), constraint=constraints.positive)
with pyro.iarange('data'):
# Local variables.
assignment = pyro.sample('assignment',
dist.Categorical(weights).expand_by([len(data)]))
pyro.sample('obs', dist.Normal(locs[assignment], scale), obs=data)
def guide(data):
with pyro.iarange('data'):
# Local parameters.
assignment_probs = pyro.param('assignment_probs', torch.ones(len(data), K) / K,
constraint=constraints.unit_interval)
pyro.sample('assignment', dist.Categorical(assignment_probs))
```
To run inference with this `(model,guide)` pair, we use Pyro's `config_enumerate()` function to enumerate over all assignments in each iteration. Since we've wrapped the batched Categorical assignments in a `pyro.iarange` indepencence context, this enumeration can happen in parallel: we enumerate only 2 possibilites, rather than `2**len(data) = 32`. Finally, to use the parallel version of enumeration, we inform pyro that we're only using a single `iarange` via `max_iarange_nesting=1`; this lets Pyro know that we're using the rightmost dimension `iarange` and letting use any other dimension for parallelization.
```
optim = pyro.optim.Adam({'lr': 0.2, 'betas': [0.9, 0.99]})
inference = SVI(model, config_enumerate(guide, 'parallel'), optim,
loss=TraceEnum_ELBO(max_iarange_nesting=1))
```
During training, we'll collect both losses and gradient norms to monitor convergence. We can do this using PyTorch's `.register_hook()` method.
```
pyro.set_rng_seed(1) # Set seed to make results reproducible.
pyro.clear_param_store() # Clear stale param values.
# Register hooks to monitor gradient norms.
gradient_norms = defaultdict(list)
inference.loss(model, guide, data) # Initializes param store.
for name, value in pyro.get_param_store().named_parameters():
value.register_hook(lambda g, name=name: gradient_norms[name].append(g.norm().item()))
losses = []
for i in range(500 if not smoke_test else 2):
loss = inference.step(data)
losses.append(loss)
print('.' if i % 100 else '\n', end='')
pyplot.figure(figsize=(10,3), dpi=100).set_facecolor('white')
pyplot.plot(losses)
pyplot.xlabel('iters')
pyplot.ylabel('loss')
pyplot.yscale('log')
pyplot.title('Convergence of SVI');
pyplot.figure(figsize=(10,4), dpi=100).set_facecolor('white')
for name, grad_norms in gradient_norms.items():
pyplot.plot(grad_norms, label=name)
pyplot.xlabel('iters')
pyplot.ylabel('gradient norm')
pyplot.yscale('log')
pyplot.legend(loc='best')
pyplot.title('Gradient norms during SVI');
```
Here are the learned parameters:
```
weights = pyro.param('weights')
locs = pyro.param('locs')
scale = pyro.param('scale')
print('weights = {}'.format(weights.data.numpy()))
print('locs = {}'.format(locs.data.numpy()))
print('scale = {}'.format(scale.data.numpy()))
```
The model's `weights` are as expected, with 3/5 of the data in the first component and 2/3 in the second component. We can also examine the guide's local `assignment_probs` variable.
```
assignment_probs = pyro.param('assignment_probs')
pyplot.figure(figsize=(8, 4), dpi=100).set_facecolor('white')
pyplot.plot(data.data.numpy(), assignment_probs.data.numpy()[:, 0], 'ro',
label='component with mean {:0.2g}'.format(locs[0]))
pyplot.plot(data.data.numpy(), assignment_probs.data.numpy()[:, 1], 'bo',
label='component with mean {:0.2g}'.format(locs[1]))
pyplot.title('Mixture assignment probabilities')
pyplot.xlabel('data value')
pyplot.ylabel('assignment probability')
pyplot.legend(loc='center');
```
Next let's visualize the mixture model.
```
X = np.arange(-3,15,0.1)
Y1 = weights[0].item() * scipy.stats.norm.pdf((X - locs[0].item()) / scale.item())
Y2 = weights[1].item() * scipy.stats.norm.pdf((X - locs[1].item()) / scale.item())
pyplot.figure(figsize=(10, 4), dpi=100).set_facecolor('white')
pyplot.plot(X, Y1, 'r-')
pyplot.plot(X, Y2, 'b-')
pyplot.plot(X, Y1 + Y2, 'k--')
pyplot.plot(data.data.numpy(), np.zeros(len(data)), 'k*')
pyplot.title('Densitiy of two-component mixture model')
pyplot.ylabel('probability density');
```
Finally note that optimization with mixture models is non-convex and can often get stuck in local optima. For example in this tutorial, we observed that the mixture model gets stuck in an everthing-in-one-cluster hypothesis if `scale` is initialized to be too large.
| github_jupyter |
# Custom DataLoader for Imbalanced dataset
* In this notebook we will use the higly imbalanced Protein Homology Dataset from [KDD cup 2004](https://www.kdd.org/kdd-cup/view/kdd-cup-2004/Data)
```
* The first element of each line is a BLOCK ID that denotes to which native sequence this example belongs. There is a unique BLOCK ID for each native sequence. BLOCK IDs are integers running from 1 to 303 (one for each native sequence, i.e. for each query). BLOCK IDs were assigned before the blocks were split into the train and test sets, so they do not run consecutively in either file.
* The second element of each line is an EXAMPLE ID that uniquely describes the example. You will need this EXAMPLE ID and the BLOCK ID when you submit results.
* The third element is the class of the example. Proteins that are homologous to the native sequence are denoted by 1, non-homologous proteins (i.e. decoys) by 0. Test examples have a "?" in this position.
* All following elements are feature values. There are 74 feature values in each line. The features describe the match (e.g. the score of a sequence alignment) between the native protein sequence and the sequence that is tested for homology.
```
## Initial imports
```
import numpy as np
import pandas as pd
import torch
from torch.optim import SGD, lr_scheduler
from pytorch_widedeep import Trainer
from pytorch_widedeep.preprocessing import TabPreprocessor
from pytorch_widedeep.models import TabMlp, WideDeep
from pytorch_widedeep.dataloaders import DataLoaderImbalanced, DataLoaderDefault
from torchmetrics import F1 as F1_torchmetrics
from torchmetrics import Accuracy as Accuracy_torchmetrics
from torchmetrics import Precision as Precision_torchmetrics
from torchmetrics import Recall as Recall_torchmetrics
from pytorch_widedeep.metrics import Accuracy, Recall, Precision, F1Score, R2Score
from pytorch_widedeep.initializers import XavierNormal
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
import time
import datetime
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
# increase displayed columns in jupyter notebook
pd.set_option("display.max_columns", 200)
pd.set_option("display.max_rows", 300)
header_list = ["EXAMPLE_ID", "BLOCK_ID", "target"] + [str(i) for i in range(4, 78)]
df = pd.read_csv("data/kddcup04/bio_train.dat", sep="\t", names=header_list)
df.head()
# imbalance of the classes
df["target"].value_counts()
# drop columns we won't need in this example
df.drop(columns=["EXAMPLE_ID", "BLOCK_ID"], inplace=True)
df_train, df_valid = train_test_split(
df, test_size=0.2, stratify=df["target"], random_state=1
)
df_valid, df_test = train_test_split(
df_valid, test_size=0.5, stratify=df_valid["target"], random_state=1
)
```
## Preparing the data
```
continuous_cols = df.drop(columns=["target"]).columns.values.tolist()
# deeptabular
tab_preprocessor = TabPreprocessor(continuous_cols=continuous_cols, scale=True)
X_tab_train = tab_preprocessor.fit_transform(df_train)
X_tab_valid = tab_preprocessor.transform(df_valid)
X_tab_test = tab_preprocessor.transform(df_test)
# target
y_train = df_train["target"].values
y_valid = df_valid["target"].values
y_test = df_test["target"].values
```
## Define the model
```
input_layer = len(tab_preprocessor.continuous_cols)
output_layer = 1
hidden_layers = np.linspace(
input_layer * 2, output_layer, 5, endpoint=False, dtype=int
).tolist()
deeptabular = TabMlp(
mlp_hidden_dims=hidden_layers,
column_idx=tab_preprocessor.column_idx,
continuous_cols=tab_preprocessor.continuous_cols,
)
model = WideDeep(deeptabular=deeptabular)
model
# Metrics from torchmetrics
accuracy = Accuracy_torchmetrics(average=None, num_classes=2)
precision = Precision_torchmetrics(average="micro", num_classes=2)
f1 = F1_torchmetrics(average=None, num_classes=2)
recall = Recall_torchmetrics(average=None, num_classes=2)
# # Metrics from pytorch-widedeep
# accuracy = Accuracy(top_k=2)
# precision = Precision(average=False)
# recall = Recall(average=True)
# f1 = F1Score(average=False)
# Optimizers
deep_opt = SGD(model.deeptabular.parameters(), lr=0.1)
# LR Scheduler
deep_sch = lr_scheduler.StepLR(deep_opt, step_size=3)
trainer = Trainer(
model,
objective="binary",
lr_schedulers={"deeptabular": deep_sch},
initializers={"deeptabular": XavierNormal},
optimizers={"deeptabular": deep_opt},
metrics=[accuracy, precision, recall, f1],
verbose=1,
)
start = time.time()
trainer.fit(
X_train={"X_tab": X_tab_train, "target": y_train},
X_val={"X_tab": X_tab_valid, "target": y_valid},
n_epochs=3,
batch_size=50,
custom_dataloader=DataLoaderImbalanced,
oversample_mul=5,
)
print(
"Training time[s]: {}".format(
datetime.timedelta(seconds=round(time.time() - start))
)
)
pd.DataFrame(trainer.history)
df_pred = trainer.predict(X_tab=X_tab_test)
print(classification_report(df_test["target"].to_list(), df_pred))
print("Actual predicted values:\n{}".format(np.unique(df_pred, return_counts=True)))
```
| github_jupyter |
# Cowell's formulation
For cases where we only study the gravitational forces, solving the Kepler's equation is enough to propagate the orbit forward in time. However, when we want to take perturbations that deviate from Keplerian forces into account, we need a more complex method to solve our initial value problem: one of them is **Cowell's formulation**.
In this formulation we write the two body differential equation separating the Keplerian and the perturbation accelerations:
$$\ddot{\mathbb{r}} = -\frac{\mu}{|\mathbb{r}|^3} \mathbb{r} + \mathbb{a}_d$$
<div class="alert alert-info">For an in-depth exploration of this topic, still to be integrated in poliastro, check out https://github.com/Juanlu001/pfc-uc3m</div>
## First example
Let's setup a very simple example with constant acceleration to visualize the effects on the orbit.
```
import numpy as np
from astropy import units as u
from matplotlib import ticker
from matplotlib import pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
plt.ion()
from scipy.integrate import ode
from poliastro.bodies import Earth
from poliastro.twobody import Orbit
from poliastro.examples import iss
from poliastro.twobody.propagation import func_twobody
from poliastro.util import norm
from ipywidgets.widgets import interact, fixed
def state_to_vector(ss):
r, v = ss.rv()
x, y, z = r.to(u.km).value
vx, vy, vz = v.to(u.km / u.s).value
return np.array([x, y, z, vx, vy, vz])
u0 = state_to_vector(iss)
u0
t = np.linspace(0, 10 * iss.period, 500).to(u.s).value
t[:10]
dt = t[1] - t[0]
dt
k = Earth.k.to(u.km**3 / u.s**2).value
```
To provide an acceleration depending on an extra parameter, we can use **closures** like this one:
```
def constant_accel_factory(accel):
def constant_accel(t0, u, k):
v = u[3:]
norm_v = (v[0]**2 + v[1]**2 + v[2]**2)**.5
return accel * v / norm_v
return constant_accel
constant_accel_factory(accel=1e-5)(t[0], u0, k)
help(func_twobody)
```
Now we setup the integrator manually using `scipy.integrate.ode`. We cannot provide the Jacobian since we don't know the form of the acceleration in advance.
```
res = np.zeros((t.size, 6))
res[0] = u0
ii = 1
accel = 1e-5
rr = ode(func_twobody).set_integrator('dop853') # All parameters by default
rr.set_initial_value(u0, t[0])
rr.set_f_params(k, constant_accel_factory(accel))
while rr.successful() and rr.t + dt < t[-1]:
rr.integrate(rr.t + dt)
res[ii] = rr.y
ii += 1
res[:5]
```
And we plot the results:
```
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(111, projection='3d')
ax.plot(*res[:, :3].T)
ax.view_init(14, 70)
```
## Interactivity
This is the last time we used `scipy.integrate.ode` directly. Instead, we can now import a convenient function from poliastro:
```
from poliastro.twobody.propagation import cowell
def plot_iss(thrust=0.1, mass=2000.):
r0, v0 = iss.rv()
k = iss.attractor.k
t = np.linspace(0, 10 * iss.period, 500).to(u.s).value
u0 = state_to_vector(iss)
res = np.zeros((t.size, 6))
res[0] = u0
accel = thrust / mass
# Perform the whole integration
r0 = r0.to(u.km).value
v0 = v0.to(u.km / u.s).value
k = k.to(u.km**3 / u.s**2).value
ad = constant_accel_factory(accel)
r, v = r0, v0
for ii in range(1, len(t)):
r, v = cowell(k, r, v, t[ii] - t[ii - 1], ad=ad)
x, y, z = r
vx, vy, vz = v
res[ii] = [x, y, z, vx, vy, vz]
fig = plt.figure(figsize=(8, 6))
ax = fig.add_subplot(111, projection='3d')
ax.set_xlim(-20e3, 20e3)
ax.set_ylim(-20e3, 20e3)
ax.set_zlim(-20e3, 20e3)
ax.view_init(14, 70)
return ax.plot(*res[:, :3].T)
interact(plot_iss, thrust=(0.0, 0.2, 0.001), mass=fixed(2000.))
```
## Error checking
```
rtol = 1e-13
full_periods = 2
u0 = state_to_vector(iss)
tf = ((2 * full_periods + 1) * iss.period / 2).to(u.s).value
u0, tf
iss_f_kep = iss.propagate(tf * u.s, rtol=1e-18)
r0, v0 = iss.rv()
r, v = cowell(k, r0.to(u.km).value, v0.to(u.km / u.s).value, tf, rtol=rtol)
iss_f_num = Orbit.from_vectors(Earth, r * u.km, v * u.km / u.s, iss.epoch + tf * u.s)
iss_f_num.r, iss_f_kep.r
assert np.allclose(iss_f_num.r, iss_f_kep.r, rtol=rtol, atol=1e-08 * u.km)
assert np.allclose(iss_f_num.v, iss_f_kep.v, rtol=rtol, atol=1e-08 * u.km / u.s)
#assert np.allclose(iss_f_num.a, iss_f_kep.a, rtol=rtol, atol=1e-08 * u.km)
#assert np.allclose(iss_f_num.ecc, iss_f_kep.ecc, rtol=rtol)
#assert np.allclose(iss_f_num.inc, iss_f_kep.inc, rtol=rtol, atol=1e-08 * u.rad)
#assert np.allclose(iss_f_num.raan, iss_f_kep.raan, rtol=rtol, atol=1e-08 * u.rad)
#assert np.allclose(iss_f_num.argp, iss_f_kep.argp, rtol=rtol, atol=1e-08 * u.rad)
#assert np.allclose(iss_f_num.nu, iss_f_kep.nu, rtol=rtol, atol=1e-08 * u.rad)
```
Too bad I cannot access the internal state of the solver. I will have to do it in a blackbox way.
```
u0 = state_to_vector(iss)
full_periods = 4
tof_vector = np.linspace(0, ((2 * full_periods + 1) * iss.period / 2).to(u.s).value, num=100)
rtol_vector = np.logspace(-3, -12, num=30)
res_array = np.zeros((rtol_vector.size, tof_vector.size))
for jj, tof in enumerate(tof_vector):
rf, vf = iss.propagate(tof * u.s, rtol=1e-12).rv()
for ii, rtol in enumerate(rtol_vector):
rr = ode(func_twobody).set_integrator('dop853', rtol=rtol, nsteps=1000)
rr.set_initial_value(u0, 0.0)
rr.set_f_params(k, constant_accel_factory(0.0)) # Zero acceleration
rr.integrate(rr.t + tof)
if rr.successful():
uf = rr.y
r, v = uf[:3] * u.km, uf[3:] * u.km / u.s
res = max(norm((r - rf) / rf), norm((v - vf) / vf))
else:
res = np.nan
res_array[ii, jj] = res
fig, ax = plt.subplots(figsize=(16, 6))
xx, yy = np.meshgrid(tof_vector, rtol_vector)
cs = ax.contourf(xx, yy, res_array, levels=np.logspace(-12, -1, num=12),
locator=ticker.LogLocator(), cmap=plt.cm.Spectral_r)
fig.colorbar(cs)
for nn in range(full_periods + 1):
lf = ax.axvline(nn * iss.period.to(u.s).value, color='k', ls='-')
lh = ax.axvline((2 * nn + 1) * iss.period.to(u.s).value / 2, color='k', ls='--')
ax.set_yscale('log')
ax.set_xlabel("Time of flight (s)")
ax.set_ylabel("Relative tolerance")
ax.set_title("Maximum relative difference")
ax.legend((lf, lh), ("Full period", "Half period"))
```
## Numerical validation
According to [Edelbaum, 1961], a coplanar, semimajor axis change with tangent thrust is defined by:
$$\frac{\operatorname{d}\!a}{a_0} = 2 \frac{F}{m V_0}\operatorname{d}\!t, \qquad \frac{\Delta{V}}{V_0} = \frac{1}{2} \frac{\Delta{a}}{a_0}$$
So let's create a new circular orbit and perform the necessary checks, assuming constant mass and thrust (i.e. constant acceleration):
```
ss = Orbit.circular(Earth, 500 * u.km)
tof = 20 * ss.period
ad = constant_accel_factory(1e-7)
r0, v0 = ss.rv()
r, v = cowell(k, r0.to(u.km).value, v0.to(u.km / u.s).value,
tof.to(u.s).value, ad=ad)
ss_final = Orbit.from_vectors(Earth, r * u.km, v * u.km / u.s, ss.epoch + rr.t * u.s)
da_a0 = (ss_final.a - ss.a) / ss.a
da_a0
dv_v0 = abs(norm(ss_final.v) - norm(ss.v)) / norm(ss.v)
2 * dv_v0
np.allclose(da_a0, 2 * dv_v0, rtol=1e-2)
dv = abs(norm(ss_final.v) - norm(ss.v))
dv
accel_dt = accel * u.km / u.s**2 * (t[-1] - t[0]) * u.s
accel_dt
np.allclose(dv, accel_dt, rtol=1e-2, atol=1e-8 * u.km / u.s)
```
This means **we successfully validated the model against an extremely simple orbit transfer with approximate analytical solution**. Notice that the final eccentricity, as originally noticed by Edelbaum, is nonzero:
```
ss_final.ecc
```
## References
* [Edelbaum, 1961] "Propulsion requirements for controllable satellites"
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.