text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Scale Detection
Train a model to detect the scale of an image relative to the scale of the training dataset for a model.
```
import os
import errno
import numpy as np
import deepcell
# Set up some global constants and shared filepaths
SEED = 123 # random seed for splitting data into train/test
ROOT_DIR = '/data' # TODO: Change this! Usually a mounted volume
DATA_DIR = os.path.expanduser(os.path.join('~', '.keras', 'datasets'))
MODEL_DIR = os.path.abspath(os.path.join(ROOT_DIR, 'models'))
LOG_DIR = os.path.abspath(os.path.join(ROOT_DIR, 'logs'))
MODEL_NAME = 'ScaleDetectionModel'
MODEL_PATH = os.path.join(MODEL_DIR, MODEL_NAME + '.h5')
# create directories if they do not exist
for d in (MODEL_DIR, LOG_DIR):
try:
os.makedirs(d)
except OSError as exc: # Guard against race condition
if exc.errno != errno.EEXIST:
raise
```
## Load the training data
Download data for nuclear, brightfield and fluorescent cytoplasm from `deepcell.datasets` and combine the data into a single training dataset.
```
# First, download the data from deepcell.datasets
import deepcell.datasets
# nuclear data (label type 0)
(X_train, y_train), (X_test, y_test) = deepcell.datasets.hela_s3.load_data(seed=SEED)
nuclear_train = {'X': X_train, 'y': y_train}
nuclear_test = {'X': X_test, 'y': y_test}
# brightfield phase data (label type 1)
(X_train, y_train), (X_test, y_test) = deepcell.datasets.phase.HeLa_S3.load_data(seed=SEED)
brightfield_train = {'X': X_train, 'y': y_train}
brightfield_test = {'X': X_test, 'y': y_test}
# flourescent cytoplasm data (label type 2)
(X_train, y_train), (X_test, y_test) = deepcell.datasets.cytoplasm.hela_s3.load_data(seed=SEED)
flourescent_train = {'X': X_train, 'y': y_train}
flourescent_test = {'X': X_test, 'y': y_test}
```
### Flatten All Datasets into 2D and Combine
```
# Reshape each dataset to conform to the minimum size of 216
from deepcell.utils.data_utils import reshape_matrix
RESHAPE_SIZE = 216
all_train = [nuclear_train, brightfield_train, flourescent_train]
all_test = [nuclear_test, brightfield_test, flourescent_test]
for train, test in zip(all_train, all_test):
train['X'], train['y'] = reshape_matrix(train['X'], train['y'], RESHAPE_SIZE)
test['X'], test['y'] = reshape_matrix(test['X'], test['y'], RESHAPE_SIZE)
# Stack up our data as train and test
X_train = np.vstack([
nuclear_train['X'],
brightfield_train['X'],
flourescent_train['X']
])
y_train = np.vstack([
nuclear_train['y'],
brightfield_train['y'],
flourescent_train['y']
])
X_test = np.vstack([
nuclear_test['X'],
brightfield_test['X'],
flourescent_test['X']
])
y_test = np.vstack([
nuclear_test['y'],
brightfield_test['y'],
flourescent_test['y']
])
```
## Create the Scale Detection Model
We are using the ScaleDetectionModel from `deepcell.applications`
```
from deepcell.applications import ScaleDetectionModel
# set use_pretrained_weights to False to retrain from scratch
model = ScaleDetectionModel(
backbone='mobilenetv2',
input_shape=X_train.shape[1:],
use_pretrained_weights=False)
model.summary()
from tensorflow.keras.optimizers import SGD, Adam
from deepcell.utils.train_utils import rate_scheduler
n_epoch = 20
batch_size = 64
lr = 1e-3
optimizer = Adam(lr=lr, clipnorm=.001)
lr_sched = rate_scheduler(lr=lr, decay=0.9)
model.compile(optimizer, loss='mse', metrics=['mae', 'mape'])
```
## Train the Model
Create the ImageDataGenerators and pass them to `model.fit_generator`.
```
from deepcell.image_generators import ScaleDataGenerator
# Create the image data generator for training
generator = ScaleDataGenerator(
rotation_range=180,
horizontal_flip=True,
vertical_flip=True,
zoom_range=(0.5, 2)
)
from tensorflow.python.keras import callbacks
model.fit_generator(
generator.flow({'X': X_train, 'y': y_train}, batch_size=batch_size),
steps_per_epoch=X_train.shape[0] // batch_size,
epochs=n_epoch,
validation_data=generator.flow({'X': X_test, 'y': y_test}, batch_size=batch_size),
validation_steps=X_test.shape[0] // batch_size,
callbacks=[
callbacks.LearningRateScheduler(lr_sched),
callbacks.ModelCheckpoint(
os.path.join(MODEL_DIR, MODEL_NAME + '.h5'),
verbose=1,
monitor='val_loss',
save_best_only=True)
]
)
```
## Run the model on test data
Run the model on validation data and visualize the performance.
```
model.load_weights(os.path.join(MODEL_DIR, MODEL_NAME + '.h5'))
test_data = generator.flow({'X': X_test, 'y': y_test}, batch_size=1)
true, pred = [], []
for i in range(1000):
if i % 100 == 0:
print(".", end="")
X, y = test_data.next()
true.append(y)
pred.append(model.predict(X))
true = np.array(true)
pred = np.array(pred)
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
ax.scatter(true, pred - true)
ax.set_xlabel('True Zoom')
ax.set_ylabel('Error')
```
| github_jupyter |
## What's this TensorFlow business?
You've written a lot of code in this assignment to provide a whole host of neural network functionality. Dropout, Batch Norm, and 2D convolutions are some of the workhorses of deep learning in computer vision. You've also worked hard to make your code efficient and vectorized.
For the last part of this assignment, though, we're going to leave behind your beautiful codebase and instead migrate to one of two popular deep learning frameworks: in this instance, TensorFlow (or PyTorch, if you switch over to that notebook)
#### What is it?
TensorFlow is a system for executing computational graphs over Tensor objects, with native support for performing backpropogation for its Variables. In it, we work with Tensors which are n-dimensional arrays analogous to the numpy ndarray.
#### Why?
* Our code will now run on GPUs! Much faster training. Writing your own modules to run on GPUs is beyond the scope of this class, unfortunately.
* We want you to be ready to use one of these frameworks for your project so you can experiment more efficiently than if you were writing every feature you want to use by hand.
* We want you to stand on the shoulders of giants! TensorFlow and PyTorch are both excellent frameworks that will make your lives a lot easier, and now that you understand their guts, you are free to use them :)
* We want you to be exposed to the sort of deep learning code you might run into in academia or industry.
## How will I learn TensorFlow?
TensorFlow has many excellent tutorials available, including those from [Google themselves](https://www.tensorflow.org/get_started/get_started).
Otherwise, this notebook will walk you through much of what you need to do to train models in TensorFlow. See the end of the notebook for some links to helpful tutorials if you want to learn more or need further clarification on topics that aren't fully explained here.
## Load Datasets
```
import tensorflow as tf
import numpy as np
import math
import timeit
import matplotlib.pyplot as plt
%matplotlib inline
from cs231n.data_utils import load_CIFAR10
def get_CIFAR10_data(num_training=49000, num_validation=1000, num_test=10000):
"""
Load the CIFAR-10 dataset from disk and perform preprocessing to prepare
it for the two-layer neural net classifier. These are the same steps as
we used for the SVM, but condensed to a single function.
"""
# Load the raw CIFAR-10 data
cifar10_dir = 'cs231n/datasets/cifar-10-batches-py'
X_train, y_train, X_test, y_test = load_CIFAR10(cifar10_dir)
# Subsample the data
mask = range(num_training, num_training + num_validation)
X_val = X_train[mask]
y_val = y_train[mask]
mask = range(num_training)
X_train = X_train[mask]
y_train = y_train[mask]
mask = range(num_test)
X_test = X_test[mask]
y_test = y_test[mask]
# Normalize the data: subtract the mean image
mean_image = np.mean(X_train, axis=0)
X_train -= mean_image
X_val -= mean_image
X_test -= mean_image
return X_train, y_train, X_val, y_val, X_test, y_test
# Invoke the above function to get our data.
X_train, y_train, X_val, y_val, X_test, y_test = get_CIFAR10_data()
print('Train data shape: ', X_train.shape)
print('Train labels shape: ', y_train.shape)
print('Validation data shape: ', X_val.shape)
print('Validation labels shape: ', y_val.shape)
print('Test data shape: ', X_test.shape)
print('Test labels shape: ', y_test.shape)
```
## Example Model
### Some useful utilities
. Remember that our image data is initially N x H x W x C, where:
* N is the number of datapoints
* H is the height of each image in pixels
* W is the height of each image in pixels
* C is the number of channels (usually 3: R, G, B)
This is the right way to represent the data when we are doing something like a 2D convolution, which needs spatial understanding of where the pixels are relative to each other. When we input image data into fully connected affine layers, however, we want each data example to be represented by a single vector -- it's no longer useful to segregate the different channels, rows, and columns of the data.
### The example model itself
The first step to training your own model is defining its architecture.
Here's an example of a convolutional neural network defined in TensorFlow -- try to understand what each line is doing, remembering that each layer is composed upon the previous layer. We haven't trained anything yet - that'll come next - for now, we want you to understand how everything gets set up.
In that example, you see 2D convolutional layers (Conv2d), ReLU activations, and fully-connected layers (Linear). You also see the Hinge loss function, and the Adam optimizer being used.
Make sure you understand why the parameters of the Linear layer are 5408 and 10.
### TensorFlow Details
In TensorFlow, much like in our previous notebooks, we'll first specifically initialize our variables, and then our network model.
```
# clear old variables
tf.reset_default_graph()
# setup input (e.g. the data that changes every batch)
# The first dim is None, and gets sets automatically based on batch size fed in
X = tf.placeholder(tf.float32, [None, 32, 32, 3])
y = tf.placeholder(tf.int64, [None])
is_training = tf.placeholder(tf.bool)
def simple_model(X,y):
# define our weights (e.g. init_two_layer_convnet)
# setup variables
Wconv1 = tf.get_variable("Wconv1", shape=[7, 7, 3, 32])
bconv1 = tf.get_variable("bconv1", shape=[32])
W1 = tf.get_variable("W1", shape=[5408, 10])
b1 = tf.get_variable("b1", shape=[10])
# define our graph (e.g. two_layer_convnet)
a1 = tf.nn.conv2d(X, Wconv1, strides=[1,2,2,1], padding='VALID') + bconv1
h1 = tf.nn.relu(a1)
h1_flat = tf.reshape(h1,[-1,5408])
y_out = tf.matmul(h1_flat,W1) + b1
return y_out
y_out = simple_model(X,y)
# define our loss
total_loss = tf.losses.hinge_loss(tf.one_hot(y,10),logits=y_out)
mean_loss = tf.reduce_mean(total_loss)
# define our optimizer
optimizer = tf.train.AdamOptimizer(5e-4) # select optimizer and set learning rate
train_step = optimizer.minimize(mean_loss)
```
TensorFlow supports many other layer types, loss functions, and optimizers - you will experiment with these next. Here's the official API documentation for these (if any of the parameters used above were unclear, this resource will also be helpful).
* Layers, Activations, Loss functions : https://www.tensorflow.org/api_guides/python/nn
* Optimizers: https://www.tensorflow.org/api_guides/python/train#Optimizers
* BatchNorm: https://www.tensorflow.org/api_docs/python/tf/layers/batch_normalization
### Training the model on one epoch
While we have defined a graph of operations above, in order to execute TensorFlow Graphs, by feeding them input data and computing the results, we first need to create a `tf.Session` object. A session encapsulates the control and state of the TensorFlow runtime. For more information, see the TensorFlow [Getting started](https://www.tensorflow.org/get_started/get_started) guide.
Optionally we can also specify a device context such as `/cpu:0` or `/gpu:0`. For documentation on this behavior see [this TensorFlow guide](https://www.tensorflow.org/tutorials/using_gpu)
You should see a validation loss of around 0.4 to 0.6 and an accuracy of 0.30 to 0.35 below
```
def run_model(session, predict, loss_val, Xd, yd,
epochs=1, batch_size=64, print_every=100,
training=None, plot_losses=False):
# have tensorflow compute accuracy
correct_prediction = tf.equal(tf.argmax(predict,1), y)
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
# shuffle indicies
train_indicies = np.arange(Xd.shape[0])
np.random.shuffle(train_indicies)
training_now = training is not None
# setting up variables we want to compute (and optimizing)
# if we have a training function, add that to things we compute
variables = [mean_loss,correct_prediction,accuracy]
if training_now:
variables[-1] = training
# counter
iter_cnt = 0
for e in range(epochs):
# keep track of losses and accuracy
correct = 0
losses = []
# make sure we iterate over the dataset once
for i in range(int(math.ceil(Xd.shape[0]/batch_size))):
# generate indicies for the batch
start_idx = (i*batch_size)%Xd.shape[0]
idx = train_indicies[start_idx:start_idx+batch_size]
# create a feed dictionary for this batch
feed_dict = {X: Xd[idx,:],
y: yd[idx],
is_training: training_now }
# get batch size
actual_batch_size = yd[idx].shape[0]
# have tensorflow compute loss and correct predictions
# and (if given) perform a training step
loss, corr, _ = session.run(variables,feed_dict=feed_dict)
# aggregate performance stats
losses.append(loss*actual_batch_size)
correct += np.sum(corr)
# print every now and then
if training_now and (iter_cnt % print_every) == 0:
print("Iteration {0}: with minibatch training loss = {1:.3g} and accuracy of {2:.2g}"\
.format(iter_cnt,loss,np.sum(corr)/actual_batch_size))
iter_cnt += 1
total_correct = correct/Xd.shape[0]
total_loss = np.sum(losses)/Xd.shape[0]
print("Epoch {2}, Overall loss = {0:.3g} and accuracy of {1:.3g}"\
.format(total_loss,total_correct,e+1))
if plot_losses:
plt.plot(losses)
plt.grid(True)
plt.title('Epoch {} Loss'.format(e+1))
plt.xlabel('minibatch number')
plt.ylabel('minibatch loss')
plt.show()
return total_loss,total_correct
with tf.Session() as sess:
with tf.device("/cpu:0"): #"/cpu:0" or "/gpu:0"
sess.run(tf.global_variables_initializer())
print('Training')
run_model(sess,y_out,mean_loss,X_train,y_train,1,64,100,train_step,True)
print('Validation')
run_model(sess,y_out,mean_loss,X_val,y_val,1,64)
```
## Training a specific model
In this section, we're going to specify a model for you to construct. The goal here isn't to get good performance (that'll be next), but instead to get comfortable with understanding the TensorFlow documentation and configuring your own model.
Using the code provided above as guidance, and using the following TensorFlow documentation, specify a model with the following architecture:
* 7x7 Convolutional Layer with 32 filters and stride of 1
* ReLU Activation Layer
* Spatial Batch Normalization Layer (trainable parameters, with scale and centering)
* 2x2 Max Pooling layer with a stride of 2
* Affine layer with 1024 output units
* ReLU Activation Layer
* Affine layer from 1024 input units to 10 outputs
```
# clear old variables
tf.reset_default_graph()
# define our input (e.g. the data that changes every batch)
# The first dim is None, and gets sets automatically based on batch size fed in
X = tf.placeholder(tf.float32, [None, 32, 32, 3])
y = tf.placeholder(tf.int64, [None])
is_training = tf.placeholder(tf.bool)
# define model
def complex_model(X,y,is_training):
# define our weights (e.g. init_two_layer_convnet)
# setup variables
Wconv1 = tf.get_variable("Wconv1", shape=[7, 7, 3, 32])
bconv1 = tf.get_variable("bconv1", shape=[32])
W1 = tf.get_variable("W1", shape=[5408, 1024])
b1 = tf.get_variable("b1", shape=[1024])
W2 = tf.get_variable("W2", shape=[1024, 10])
b2 = tf.get_variable("b2", shape=[10])
gamma = tf.get_variable('gamma', shape=[32])
beta = tf.get_variable('beta', shape=[32])
# define our graph (e.g. two_layer_convnet)
c1 = tf.nn.conv2d(X, Wconv1, strides=[1,1,1,1], padding='VALID') + bconv1
r1 = tf.nn.relu(c1)
mean, var = tf.nn.moments(r1, [0,1,2])
bn = tf.nn.batch_normalization(r1, mean, var, beta, gamma, 1e-6)
mp = tf.nn.max_pool(bn, [1,2,2,1], strides=[1,2,2,1], padding='VALID')
# resultant shape = (W - K + 2P)/S
# W: Original shape, K: Kernel Shape, P: Padding, S: Stride
mp_flat = tf.reshape(mp,[-1,5408])
a1 = tf.matmul(mp_flat, W1) + b1
r2 = tf.nn.relu(a1)
a2 = tf.matmul(r2, W2) + b2
return a2
y_out = complex_model(X,y,is_training)
```
To make sure you're doing the right thing, use the following tool to check the dimensionality of your output (it should be 64 x 10, since our batches have size 64 and the output of the final affine layer should be 10, corresponding to our 10 classes):
```
# Now we're going to feed a random batch into the model
# and make sure the output is the right size
x = np.random.randn(64, 32, 32,3)
with tf.Session() as sess:
with tf.device("/cpu:0"): #"/cpu:0" or "/gpu:0"
tf.global_variables_initializer().run()
ans = sess.run(y_out,feed_dict={X:x,is_training:True})
%timeit sess.run(y_out,feed_dict={X:x,is_training:True})
print(ans.shape)
print(np.array_equal(ans.shape, np.array([64, 10])))
```
You should see the following from the run above
`(64, 10)`
`True`
### GPU!
Now, we're going to try and start the model under the GPU device, the rest of the code stays unchanged and all our variables and operations will be computed using accelerated code paths. However, if there is no GPU, we get a Python exception and have to rebuild our graph. On a dual-core CPU, you might see around 50-80ms/batch running the above, while the Google Cloud GPUs (run below) should be around 2-5ms/batch.
```
try:
with tf.Session() as sess:
with tf.device("/gpu:0") as dev: #"/cpu:0" or "/gpu:0"
tf.global_variables_initializer().run()
ans = sess.run(y_out,feed_dict={X:x,is_training:True})
%timeit sess.run(y_out,feed_dict={X:x,is_training:True})
except tf.errors.InvalidArgumentError:
print("no gpu found, please use Google Cloud if you want GPU acceleration")
# rebuild the graph
# trying to start a GPU throws an exception
# and also trashes the original graph
tf.reset_default_graph()
X = tf.placeholder(tf.float32, [None, 32, 32, 3])
y = tf.placeholder(tf.int64, [None])
is_training = tf.placeholder(tf.bool)
y_out = complex_model(X,y,is_training)
```
You should observe that even a simple forward pass like this is significantly faster on the GPU. So for the rest of the assignment (and when you go train your models in assignment 3 and your project!), you should use GPU devices. However, with TensorFlow, the default device is a GPU if one is available, and a CPU otherwise, so we can skip the device specification from now on.
### Train the model.
Now that you've seen how to define a model and do a single forward pass of some data through it, let's walk through how you'd actually train one whole epoch over your training data (using the complex_model you created provided above).
Make sure you understand how each TensorFlow function used below corresponds to what you implemented in your custom neural network implementation.
First, set up an **RMSprop optimizer** (using a 1e-3 learning rate) and a **cross-entropy loss** function. See the TensorFlow documentation for more information
* Layers, Activations, Loss functions : https://www.tensorflow.org/api_guides/python/nn
* Optimizers: https://www.tensorflow.org/api_guides/python/train#Optimizers
```
# Inputs
# y_out: is what your model computes
# y: is your TensorFlow variable with label information
# Outputs
# mean_loss: a TensorFlow variable (scalar) with numerical loss
# optimizer: a TensorFlow optimizer
# This should be ~3 lines of code!
mean_loss = None
optimizer = None
# define our loss
total_loss = tf.nn.softmax_cross_entropy_with_logits_v2(labels=tf.one_hot(y,10),logits=y_out)
mean_loss = tf.reduce_mean(total_loss)
# define our optimizer
optimizer = tf.train.RMSPropOptimizer(5e-4) # select optimizer and set learning rate
# batch normalization in tensorflow requires this extra dependency
extra_update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
with tf.control_dependencies(extra_update_ops):
train_step = optimizer.minimize(mean_loss)
```
### Train the model
Below we'll create a session and train the model over one epoch. You should see a loss of 1.4 to 2.0 and an accuracy of 0.4 to 0.5. There will be some variation due to random seeds and differences in initialization
```
sess = tf.Session()
sess.run(tf.global_variables_initializer())
print('Training')
run_model(sess,y_out,mean_loss,X_train,y_train,1,64,100,train_step)
```
### Check the accuracy of the model.
Let's see the train and test code in action -- feel free to use these methods when evaluating the models you develop below. You should see a loss of 1.3 to 2.0 with an accuracy of 0.45 to 0.55.
```
print('Validation')
run_model(sess,y_out,mean_loss,X_val,y_val,1,64)
```
## Train a _great_ model on CIFAR-10!
Now it's your job to experiment with architectures, hyperparameters, loss functions, and optimizers to train a model that achieves ** >= 70% accuracy on the validation set** of CIFAR-10. You can use the `run_model` function from above.
### Things you should try:
- **Filter size**: Above we used 7x7; this makes pretty pictures but smaller filters may be more efficient
- **Number of filters**: Above we used 32 filters. Do more or fewer do better?
- **Pooling vs Strided Convolution**: Do you use max pooling or just stride convolutions?
- **Batch normalization**: Try adding spatial batch normalization after convolution layers and vanilla batch normalization after affine layers. Do your networks train faster?
- **Network architecture**: The network above has two layers of trainable parameters. Can you do better with a deep network? Good architectures to try include:
- [conv-relu-pool]xN -> [affine]xM -> [softmax or SVM]
- [conv-relu-conv-relu-pool]xN -> [affine]xM -> [softmax or SVM]
- [batchnorm-relu-conv]xN -> [affine]xM -> [softmax or SVM]
- **Use TensorFlow Scope**: Use TensorFlow scope and/or [tf.layers](https://www.tensorflow.org/api_docs/python/tf/layers) to make it easier to write deeper networks. See [this tutorial](https://www.tensorflow.org/tutorials/layers) for how to use `tf.layers`.
- **Use Learning Rate Decay**: [As the notes point out](http://cs231n.github.io/neural-networks-3/#anneal), decaying the learning rate might help the model converge. Feel free to decay every epoch, when loss doesn't change over an entire epoch, or any other heuristic you find appropriate. See the [Tensorflow documentation](https://www.tensorflow.org/versions/master/api_guides/python/train#Decaying_the_learning_rate) for learning rate decay.
- **Global Average Pooling**: Instead of flattening and then having multiple affine layers, perform convolutions until your image gets small (7x7 or so) and then perform an average pooling operation to get to a 1x1 image picture (1, 1 , Filter#), which is then reshaped into a (Filter#) vector. This is used in [Google's Inception Network](https://arxiv.org/abs/1512.00567) (See Table 1 for their architecture).
- **Regularization**: Add l2 weight regularization, or perhaps use [Dropout as in the TensorFlow MNIST tutorial](https://www.tensorflow.org/get_started/mnist/pros)
### Tips for training
For each network architecture that you try, you should tune the learning rate and regularization strength. When doing this there are a couple important things to keep in mind:
- If the parameters are working well, you should see improvement within a few hundred iterations
- Remember the coarse-to-fine approach for hyperparameter tuning: start by testing a large range of hyperparameters for just a few training iterations to find the combinations of parameters that are working at all.
- Once you have found some sets of parameters that seem to work, search more finely around these parameters. You may need to train for more epochs.
- You should use the validation set for hyperparameter search, and we'll save the test set for evaluating your architecture on the best parameters as selected by the validation set.
### Going above and beyond
If you are feeling adventurous there are many other features you can implement to try and improve your performance. You are **not required** to implement any of these; however they would be good things to try for extra credit.
- Alternative update steps: For the assignment we implemented SGD+momentum, RMSprop, and Adam; you could try alternatives like AdaGrad or AdaDelta.
- Alternative activation functions such as leaky ReLU, parametric ReLU, ELU, or MaxOut.
- Model ensembles
- Data augmentation
- New Architectures
- [ResNets](https://arxiv.org/abs/1512.03385) where the input from the previous layer is added to the output.
- [DenseNets](https://arxiv.org/abs/1608.06993) where inputs into previous layers are concatenated together.
- [This blog has an in-depth overview](https://chatbotslife.com/resnets-highwaynets-and-densenets-oh-my-9bb15918ee32)
If you do decide to implement something extra, clearly describe it in the "Extra Credit Description" cell below.
### What we expect
At the very least, you should be able to train a ConvNet that gets at **>= 70% accuracy on the validation set**. This is just a lower bound - if you are careful it should be possible to get accuracies much higher than that! Extra credit points will be awarded for particularly high-scoring models or unique approaches.
You should use the space below to experiment and train your network. The final cell in this notebook should contain the training and validation set accuracies for your final trained network.
Have fun and happy training!
```
# Feel free to play with this cell
def my_model(X, y, is_training):
conv1 = tf.layers.conv2d(
inputs=X,
filters=128,
kernel_size=(3, 3),
padding="same",
activation=tf.nn.relu)
conv2 = tf.layers.conv2d(
inputs=conv1,
filters=128,
kernel_size=(3, 3),
padding="same",
activation=tf.nn.relu)
conv3 = tf.layers.conv2d(
inputs=conv2,
filters=128,
kernel_size=(3, 3),
padding="same",
activation=tf.nn.relu)
sbn1 = tf.layers.batch_normalization(conv3, training=is_training)
pool1 = tf.layers.max_pooling2d(inputs=sbn1, pool_size=(2, 2), strides=2)
conv4 = tf.layers.conv2d(
inputs=pool1,
filters=128,
kernel_size=(3, 3),
padding="same",
activation=tf.nn.relu)
conv5 = tf.layers.conv2d(
inputs=conv4,
filters=128,
kernel_size=[3, 3],
padding="same",
activation=tf.nn.relu)
conv6 = tf.layers.conv2d(
inputs=conv5,
filters=128,
kernel_size=(3, 3),
padding="same",
activation=tf.nn.relu)
sbn2 = tf.layers.batch_normalization(conv6, training=is_training)
pool2 = tf.layers.max_pooling2d(inputs=sbn2, pool_size=(2, 2), strides=2)
# ap = tf.layers.average_pooling2d(pool2, (8, 8), (1, 1))
ap_flat = tf.reshape(pool2, [-1, 64*128])
# Note to self: Using bn here reduces accuracy
dense1 = tf.layers.dense(inputs=ap_flat, units=512, activation=tf.nn.relu)
dp1 = tf.layers.dropout(dense1, training=is_training)
dense2 = tf.layers.dense(inputs=dp1, units=512, activation=tf.nn.relu)
dp2 = tf.layers.dropout(dense2, training=is_training)
dense3 = tf.layers.dense(inputs=dp2, units=512, activation=tf.nn.relu)
dp3 = tf.layers.dropout(dense3, training=is_training)
logits = tf.layers.dense(inputs=dp3, units=10)
return logits
tf.reset_default_graph()
X = tf.placeholder(tf.float32, [None, 32, 32, 3])
y = tf.placeholder(tf.int64, [None])
is_training = tf.placeholder(tf.bool)
y_out = my_model(X,y,is_training)
total_loss = tf.nn.softmax_cross_entropy_with_logits_v2(labels=tf.one_hot(y,10), logits=y_out)
mean_loss = tf.reduce_mean(total_loss)
optimizer = tf.train.AdamOptimizer(5e-4) # select optimizer and set learning rate
train_step = optimizer.minimize(mean_loss)
# batch normalization in tensorflow requires this extra dependency
extra_update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
with tf.control_dependencies(extra_update_ops):
train_step = optimizer.minimize(mean_loss)
# Feel free to play with this cell
# This default code creates a session
# and trains your model for 10 epochs
# then prints the validation set accuracy
sess = tf.Session()
sess.run(tf.global_variables_initializer())
print('Training')
run_model(sess,y_out,mean_loss,X_train,y_train,15,64,100,train_step,True)
print('Validation')
run_model(sess,y_out,mean_loss,X_val,y_val,1,64)
# Test your model here, and make sure
# the output of this cell is the accuracy
# of your best model on the training and val sets
# We're looking for >= 70% accuracy on Validation
print('Training')
run_model(sess,y_out,mean_loss,X_train,y_train,1,64)
print('Validation')
run_model(sess,y_out,mean_loss,X_val,y_val,1,64)
```
### Describe what you did here
In this cell you should also write an explanation of what you did, any additional features that you implemented, and any visualizations or graphs that you make in the process of training and evaluating your network
The structure of the network is as follows:
\[ CONV x3 - Spatial Batchnorm - Max Pool \] x3 - Flatten - FC
I probably can train it with less epochs and switch to a deeper network but since I don't have a very good GPU I decided to stop since I got 80%+ already.
### Test Set - Do this only once
Now that we've gotten a result that we're happy with, we test our final model on the test set. This would be the score we would achieve on a competition. Think about how this compares to your validation set accuracy.
```
print('Test')
run_model(sess,y_out,mean_loss,X_test,y_test,1,64)
```
## Going further with TensorFlow
The next assignment will make heavy use of TensorFlow. You might also find it useful for your projects.
# Extra Credit Description
If you implement any additional features for extra credit, clearly describe them here with pointers to any code in this or other files if applicable.
# Extra: CNN Kernel Visualization
The following several cells display the convolution layer kernel of different layer and with different in channels.Since I have been using 3x3 kernels, there are not really much patterns to see in the visualization.
```
# First conv layer, all in channels
t = tf.get_default_graph().get_tensor_by_name('conv2d/kernel:0')
t = sess.run(t)
print(t.shape)
for j in range(t.shape[-2]):
plt.figure(j, figsize=(20,10))
for i in range(t.shape[-1]):
nt = t[:,:,j,i]
plt.subplot(8, 16, i+1)
plt.imshow((nt/np.max(np.abs(nt)) + 1)/2)
plt.axis('off')
# First conv layer, first in channel
t = tf.get_default_graph().get_tensor_by_name('conv2d_1/kernel:0')
t = sess.run(t)
print(t.shape)
for j in range(1):
plt.figure(j, figsize=(20,10))
for i in range(t.shape[-1]):
nt = t[:,:,j,i]
plt.subplot(8, 16, i+1)
plt.imshow((nt/np.max(np.abs(nt)) + 1)/2)
plt.axis('off')
# Last conv layer, first in channel
t = tf.get_default_graph().get_tensor_by_name('conv2d_5/kernel:0')
t = sess.run(t)
print(t.shape)
for j in range(1):
plt.figure(j, figsize=(20,10))
for i in range(t.shape[-1]):
nt = t[:,:,j,i]
plt.subplot(8, 16, i+1)
plt.imshow((nt/np.max(np.abs(nt)) + 1)/2)
plt.axis('off')
```
| github_jupyter |
# Evaluate
```
import logging
import regex
import unicodecsv as csv
import lemmy
logging.basicConfig(format='%(levelname)s : %(message)s', level=logging.DEBUG)
NORMS_FILE = "./data/norms.csv"
UD_TRAIN_FILE = "./data/UD_Danish/da-ud-train.conllu"
UD_DEV_FILE = "./data/UD_Danish/da-ud-dev.conllu"
```
We read the normalization rules we build in the first notebook. When evaluating, we apply these to the lemmas specified in UD. Otherwise we would risk, for example, counting "akvarie" as the incorrect lemma for "akvarier" if UD specified the other spelling, "akvarium".
```
norm_lookup = dict([row for row in csv.reader(open(NORMS_FILE, 'rb'), delimiter=",",
quotechar='"',
quoting=csv.QUOTE_MINIMAL,
encoding='utf-8',
lineterminator='\n')][1:])
```
We apply a few more normalization rules. This is due to DSN and UD not agreeing on the lemmas for certain words.
```
ud_dsn_normalization = (('PRON', 'det', 'det', 'den'),
('ADJ', 'flere', 'mange', 'flere'),
('ADJ', 'mere', 'meget', 'mere'),
('ADJ', 'meget', 'meget', 'megen'),
('ADJ', 'fleste', 'mange', 'flest'))
```
## Load The Lemmatizer
```
lemmatizer = lemmy.load('da')
def _parse_ud_line(line):
return line.split("\t")[1:4]
def _evaluate(ud_file):
correct = 0
incorrect = 0
ambiguous = 0
mistakes = {}
ambiguities = {}
pos_prev = ""
for line in open(ud_file).readlines():
if line.startswith("#") or line.strip() == "":
pos_prev = ""
continue
orth, lemma_expected, pos = _parse_ud_line(line)
if pos == "NOUN" and lemma_expected in norm_lookup:
lemma_expected = norm_lookup[lemma_expected]
else:
for pos_, orth_, expected_ud, expected_dsn in ud_dsn_normalization:
if pos != pos_ or orth.lower() != orth_ or lemma_expected != expected_ud:
continue
lemma_expected = expected_dsn
lemmas_actual = lemmatizer.lemmatize(pos, orth.lower(), pos_previous=pos_prev)
lemma_actual = lemmas_actual[0]
if len(lemmas_actual) > 1:
ambiguous += 1
ambiguities[(pos, orth)] = ambiguities.get((pos, orth), 0) + 1
elif lemma_actual.lower() == lemma_expected.lower():
correct += 1
else:
mistakes[(pos, orth, lemma_expected, lemma_actual)] = mistakes.get((pos, orth, lemma_expected, lemma_actual), 0) + 1
incorrect += 1
pos_prev = pos
print("* correct:", correct)
print("* incorrect:", incorrect)
print("* ambiguous:", ambiguous)
print("*", correct/(incorrect+ambiguous+correct))
print("*", (correct+ambiguous)/(incorrect+ambiguous+correct))
return mistakes, ambiguities
```
## Evaluate on UD Train
```
mistakes_train, ambiguities_train = _evaluate(UD_TRAIN_FILE)
```
## Evaluate on UD Dev
```
mistakes_dev, ambiguities_dev =_evaluate(UD_DEV_FILE)
```
## Mistakes
```
sorted(mistakes_train.items(), key=lambda x: (-x[1], x[0][1].lower(), x))[:10]
```
## Ambiguities
```
sorted(ambiguities_train.items(), key=lambda x: (-x[1], x[0][1].lower(), x))[:10]
```
| github_jupyter |
## Lesson 2 - Basic Data Structure
* 2.1 - List
* 2.2 - Index and Slice
* 2.3 - Common Methods of List Object
* 2.4 - List Sort
* 2.5 - Other List Operations
* 2.6 - Multiple Layers List and DeepCopy
* 2.7 - Tuple
* 2.8 - Set
In this section, we focus on discussing the basic data structure in Python: **List**, **Tuple**, and **Set**. We will discuss **Dictionary** in the later section. First of all, many people refer the Python "list" as an array in other programming language, however, You should know that the Python "list" is a much flexible and powerful data structure compares to an array in different programming language. A "tuple', on the other hand, is basically an **immutable** list. We are going to discuss some situation when we need an immutable list, tuple.
This section also discuss a relatively new Python data structure: **set**, which is an unordered collection data type that is iterable, mutable, and has no duplicate elements.
## 2.1 - List
A Python **list** is similar to an **array** in C, Java, and other programming languages, which is an ordered mutable collection of objects. A list is created by placing all the items (elements) inside a square bracket [ ], separated by commas.
Reference: https://docs.python.org/3/tutorial/datastructures.html
Here is an example of a list.
``` Python
# Create a list with 3 elements and assign it to variable x
x = [1, 2, 3]
```
It's worth to note that, unlike other programming languages, we do not need to define the type of the elements to be store in a list or the length of the list.
Python also has similar array module like in C. Here is a documentation if you are interested in it.
Reference: https://docs.python.org/3/library/array.html
```
# Create a list with 3 elements and assign it to variable x
x = [1, 2, 3]
```
Python list is different to an array in other programming lauguages: it can store different type of data, in another word, list elements can be any Python objects. Here is an example of storing different types of elements in a single list.
``` Python
# Create a list that includes integer, string, and another list of intergers.
x = [2, 'two', [1, 2, 3]]
```
```
# Create a list that includes integer, string, and another list of intergers.
x = [2, 'two', [1, 2, 3]]
```
#### Number of elements in a list?
len() is one of the most common function used for a list, it generally returns the number of items in an object. When the object is a list, the len() function returns the number of elements in the list.
e.g. Count the number of elements in list x
``` Python
x = [2, 'two', [1, 2, 3]]
len(x) # return 3 items in the list
```
Intuition: The first element in the list is an interger 2. The second element in a string "two". And the third element is a list [1, 2, 3]
Note: len() function only returns the number of elements in the first layer of the list. In the previous example, the return value should be 3 because it won't count the elements in the second layer list.
```
# Count the number of elements in list x
x = [2, 'two', [1, 2, 3]]
len(x)
# Try it yourself!
# Use the len() function to find out the number of elements in each of the list.
x = [0]
y = []
z = [[1, 3, [4, 5], 6], 7]
```
## 2.2 - Index and Slice
The method used to find or retrieve an element from a list is similar to C, which uses an **index** [n] for n position in the list. Python uses a zero-base indexing system, meaning that the first element in a sequence is located at position 0.
Here is the example,
``` Python
# Create a list with 4 string elements
x = ["first", "second", "third", "fourth"]
# Find the first element "first" from the list
x[0] # return a string "first"
# Find the third element "third" from the list
x[2] # return a string "third"
```
```
# Create a list with 4 string elements
x = ["first", "second", "third", "fourth"]
# Find the first element "first" from the list
x[0]
# Find the third element "third" from the list
x[2]
```
If we are using a negative index, the count begins from the last element in the list, which -1 is pointing at the last element in the list, -2 will be the second to the last element, etc. We can use the list in our previous example to demonstrate use of negative index.
Here is the example,
``` Python
# Assigne the last element from list x to variable a
a = x[-1]
# The string "fourth" should be assigned to variable a
a # return a string 'fourth'
# Assign the second to the last element from list x to variable b
b = x[-2]
# The string "third" should be assigned to variable b
b # return a string 'third'
```
```
# Assigne the last element from list x to variable a
a = x[-1]
# The string "fourth" should be assigned to variable a
a
# Assign the second to the last element from list x to variable b
b = x[-2]
# The string "third" should be assigned to variable b
b
# Try it yourself!
# Given a list of element, can you find the element of your interest by the index method?
x = ["Tom", 235, 10.55, "Black", "USA", "30", 2*3, "3 divided by 2", "Python"]
```
#### Slicing a List
In the previous examples, we use the index method to retrieve a single element from a list. Python also offers a slice object to specify how to slice a sequence or point to multiple elements in a list, it is sometime called **"Slicing"**. You can specify where to start the slicing and where to end by using a square bracket [ index1 : index2 ]. Note that the slicing index is not inclusive, which means the slicing will begin from index1, but do not include index2 at the end. Let's demonstrate this in the following examples.
Here is the examples:
``` Python
# Create a list with 4 string elements
x = ["first", "second", "third", "fourth"]
# Slice the list that contains the first to the third elements
x[0:3] # return a list ['first', 'second', 'third']
# Slice the list that contains the second and third elements
x[1:3] # return a list ['second', 'third']
# We can also apply the negative index,
# For instance, we can slice the list that contains second and third elements
x[-3:-1] # return a list ['second', 'third']
```
Note that if the position of index2 is in front of the position of index1, it will return an empty list.
``` Python
x[-1:2] # return an empty list [ ]
```
```
# Create a list with 4 string elements
x = ["first", "second", "third", "fourth"]
# Slice the list that contains the first to the third elements
x[0:3]
# Slice the list that contains the second and third elements
x[1:3]
# We can also apply the negative index,
# For instance, we can slice the list that contains second and third elements
x[-3:-1]
# If the position of the second index is in front of the position of the first index
# the return object will be an empty list.
x[-1:2]
```
When slicing, index1 or index2 or both can be missed in the argument. Without index1 means starts from the firt element in the list. Without index2 means ends with the last element in the list. When both are missing, it returns the original list (include all the elements from the beginning to the end).
Lets demonstrate with some examples here,
``` Python
# Slicing the first three elements from list x
x[:3] # return a list ['first', 'second', 'third']
# Slicing the last two elements from list x
x[2:] # return a list ['third', 'fourth']
# Slicing all elements from list x
x[:] # ['first', 'second', 'third', 'fourth']
```
```
# Slicing the first three elements from list x
x[:3]
# Slicing the last two elements from list x
x[2:]
# Slicing all elements from list x
x[:]
```
When we are assigning a list to a variable, we need to be careful how we assign it. Below is a very good example to demonstrate the advantage for copying a list to a new list.
``` Python
# Assign all elements to variable y
y = x[:] # y is now a list with all the elements assigned to x
# Suppose we modify the new list y
y[0] = "1 st"
# y is now changed
y # return with a list ['1 st', 'second', 'third', 'fourth']
# The original x will not be effected by any change of y
x # return the original list ['first', 'second', 'third', 'fourth']
# If we assign x to y
y = x # Now y is referencing x, which is referencing the list
# Suppose we modify y
y[0] = "1 st"
# y is now changed
y # return with a list ['1 st', 'second', 'third', 'fourth']
# x is also changed
x # return with a list ['1 st', 'second', 'third', 'fourth']
```
```
# Assign all elements to variable y
y = x[:] # y is now a list with all the elements assigned to x
# Suppose we modify the new list y
y[0] = "1 st"
# y is now changed
y # return with a list
# The original x will not be effected by any change of y
x # return the original list
# If we assign x to y
y = x # Now y is referencing x, which is referencing the list
# Suppose we modify y
y[0] = "1 st"
# y is now changed
y # return with a list
# x is also changed
x # return with a list
# Try it yourself!
# Try to use both len() and slicing to get the second half of the elements
# from a list with unknown number of elements.
# Check to see if you solution work with an example.
# Create a list
# Code your solution here!
```
#### Bonus:
When we start to program, the Python slicing index method actaully helps improving the efficiency for programming. Think about the following example,
```Python
x = [0, 1, 2, 3, 4, 5]
start = 2
length = 3
x[start:start+length] # return [2, 3, 4]
```
Since the slicing index method is not inclusive, [m:n] will have the length n-m. If the slicing index mthod is inclusive, the slicing will be x[start:start+length-1], where the length of the slice will be n-m+1. This is one of the common bug in programming when we have to -1 for one process and +1 for the other.
#### Modify element(s) in a list
Both index and slice can be used to modify the existing list. Here are some examples.
``` Python
# Suppose we want to modify the third element in a list
x = [0, 1, 2, 3, 4]
x[2] = "two" # replace by index
x
# Suppose we want to modify the second and third elements in a list
x = [0, 1, 2, 3, 4]
x[1:3] = ["one", "two"] # replace by slice
x # return [0, 1, 'two', 3, 4]
# Suppose we want to modify the second and third elements in a list
x = [0, 1, 2, 3, 4]
x[1:3] = ["one", "two"] # replace by slice
x # return [0, 'one', 'two', 3, 4]
# When using slice to modify the elements in a list a[m:n] = b,
# the length of b is not required equal to the length between n and m.
x = [0, 1, 2, 3, 4]
x[1:3] = ['one', 'two', 'three']
x # return [0, 'one', 'two', 'three', 3, 4]
x[0:4] = ['a', 'b']
x # return ['a', 'b', 3, 4]
# We can also using this method to add or insert element(s) to a list
x = [0, 1, 2, 3, 4]
# Adding 3 elements to the end of the existing list
x[len(x):] = [5, 6, 7]
x # return [0, 1, 2, 3, 4, 5, 6, 7]
# Adding 2 elements to the front of the existing list
x[:0] = [-2, -1]
x # return [-2, -1, 0, 1, 2, 3, 4, 5, 6, 7]
# Remove mutliple elements from the existing list
x[1:-1] = []
x # return [-2, 7]
```
```
# Suppose we want to modify the third element in a list
x = [0, 1, 2, 3, 4]
x[2] = "two" # replace by index
x
# Suppose we want to modify the second and third elements in a list
x = [0, 1, 2, 3, 4]
x[1:3] = ["one", "two"] # replace by slice
x
# When using slice to modify the elements in a list a[m:n] = b,
# the length of b is not required equal to the length between n and m.
x = [0, 1, 2, 3, 4]
x[1:3] = ['one', 'two', 'three']
x
x[0:4] = ['a', 'b']
x
# We can also using this method to add or insert element(s) to a list
x = [0, 1, 2, 3, 4]
# Adding 3 elements to the end of the existing list
x[len(x):] = [5, 6, 7]
x
# Adding 2 elements to the front of the existing list
x[:0] = [-2, -1]
x
# Remove mutliple elements from the existing list
x[1:-1] = []
x
```
## 2.3 - Common Methods of List Object
In this section, we discuss the common **methods** of list **object**. To programming beginners, the concept of **method** could be very vague. If we image a Python **object** is actually a physical object, **method** is bacially the action of the **object**. It's similar to when we are using a **verb** pair with a **noun** in a sentence. Python is an object oriented program. A method defines the behavior of the object and is an actiona that an object is able to perform.
Reference: https://docs.python.org/3/tutorial/datastructures.html
For instance,
| Writing | Python Program |
| :---: | :---: |
| Mom | mom |
| Mom sleeps | mom.sleep() |
Here is a list of common methods of list objects,
| List Methods | Description | Code Example |
| :---: | :---: | :---: |
| [ ] | Building an empty list | x = [ ] |
| len() | Returning the length of a list | len(x) |
| append() | Adding element at the end of a list | x.append('y') |
| extend() | Adding a list of elements at the end of a list | x.extend(['a','b']) |
| insert() | Inserting an element to an appointed position | x.insert(0, 'y') |
| del | Deleting a list of element or a slice of element | del(x[0]) |
| remove() | Removing an element from a list | x.remove('y') |
| reverse() | Reversing a list | x.reverse() |
| sort() | Sorting the elements in a list | x.sort() |
| sorted() | Returning a new list after sorting | sorted(x) |
| + | Casting two lists into a new list | x1 + x2 |
| * | Copying a list | x = ['y']*3 |
| min() | Returning the minimum value element from a list | min(x) |
| max() | Returning the maximum value element from a list | mas(x) |
| index() | Returning the index of an element from a list | x.index('y') |
| count() | Returning the frequency count of an element from a list | x.count('y') |
| sum() | Returning the sum of the elements in a list | sum(x) |
| in | Returning a Boolean True if an element exist in a list, False otherwise | 'y'in x |
Let's explore how to use these methods of list objects!
#### Adding new element to a list: Append( ) and Extend( )
Often we need to add new elements to a list object. To add new element(s) to an existing list, we can use append( ) or extend( ) methods.
e.g. for instance, if we are trying to add an element to the end of a list, we can use append( ) method.
``` Python
x = [1,2,3]
x.append("four")
x # return a list [1, 2, 3, 'four']
```
If we are using append( ) method to add a list object to an existing list, it will become multi-layer list.
``` Python
x = [1,2,3,4]
y = [5,6,7]
x.append(y)
x # return a list [1, 2, 3, 4, [5, 6, 7]]
```
Note: the return list contains 5 elements and the last element is a list object.
If we are trying to add the element from list y to the end of list x, we can use the extend( ) method.
``` Python
x = [1,2,3,4]
y = [5,6,7]
x.extend(y)
x # return a list [1, 2, 3, 4, 5, 6, 7]
```
```
# Adding new element to the end of a list
x = [1,2,3]
x.append("four")
x # return a list [1, 2, 3, 'four']
# Adding a list object to the end of a list
x = [1,2,3,4]
y = [5,6,7]
x.append(y)
x # return a list [1, 2, 3, 4, [5, 6, 7]]
# Adding the elements from list y to the end of list x
x = [1,2,3,4]
y = [5,6,7]
x.extend(y)
x # return a list [1, 2, 3, 4, 5, 6, 7]
```
#### Inserting a new element to a list: insert( )
To add an element in between two elements or at the beginning of a list, we can use insert( ) method. insert( ) method has two arguments, the first one is the index reference for the need element to be added and the second one is the element itself.
e.g. Suppose we would like to added two element to an existing list. One between two elements in the list and the other one at the beginning of the list.
``` Python
x = [1,3,4]
# Adding a string "two" between 1 and 3 in the list
x.insert(1, "two")
x # return [1, 'two', 3, 4]
# Adding a string "zero" at the beginning of the list
x.insert(0, 'zero')
x # return ['zero', 1, 'two', 3, 4]
```
We should not be surprised that insert( ) method can also reference the negative index.
e.g.
``` Python
# Using negative index with insert()
x = [1,2,4]
x.insert(-1, 'three')
x # return [1, 2, 'three', 4]
```
```
x = [1,3,4]
# Adding a string "two" between 1 and 3 in the list
x.insert(1, "two")
x # return [1, 'two', 3, 4]
# Adding a string "zero" at the beginning of the list
x.insert(0, 'zero')
x # return ['zero', 1, 'two', 3, 4]
# Using negative index with insert()
x = [1,2,4]
x.insert(-1, 'three')
x # return [1, 2, 'three', 4]
```
#### Deleting element from a list: del
Instead of slicing, we can also use "del" to delete a sequence of elements in a list. "del" is not a method of list object, but a Python keyword, therefore, it is also applied to any Python object with name. Even though "del" is not as powerful as slicing, it usually provides a more readable coding format as compare to slicing.
e.g.
``` Python
x = ['a', 2, 'c', 7, 9, 11]
# delete the second element
del x[1]
x # return ['a', 'c', 7, 9, 11]
# delete the first two elements
del x[:2]
x # return [7, 9, 11]
```
Generally speaking, del x[n] and x[n:n+1]=[ ] return the same list. Also, del x[m:n] and x[m:n]=[ ] are the same. However, just like using insert( ) method, "del" improves the readability of the programming code.
```
x = ['a', 2, 'c', 7, 9, 11]
# delete the second element
del x[1]
x # return ['a', 'c', 7, 9, 11]
# delete the first two elements
del x[:2]
x # return [7, 9, 11]
```
#### Finding the specific value from the list and remove it from the list: remove( )
remove( ) method finds the first given object from the list and removes it.
e.g.
``` Python
x = [1, 2, 3, 4, 3, 5]
# Remove the first integer 3 from the list
x.remove(3)
x # [1, 2, 4, 3, 5]
# Remove the first integer 3 from the modified list
x.remove(3)
x # [1, 2, 4, 5]
# Try to remove the first integer 3 from the modifed list
x.remove(3)
# An error will return if no such value exists in the list
```
To avoid error, we can also use 'in' Python keyword to confirm the existing of the value in the list before remove.
```
x = [1, 2, 3, 4, 3, 5]
# Remove the first integer 3 from the list
x.remove(3)
x # [1, 2, 4, 3, 5]
# Remove the first integer 3 from the modified list
x.remove(3)
x # [1, 2, 4, 5]
# Try to remove the first integer 3 from the modifed list
x.remove(3)
# An error will return if no such value exists in the list
```
#### Reversing the order of the elements in a list: reverse()
reverse( ) is a relatively special method of list object, which reverses the order of the given elements from the list efficiently and updates the list.
e.g.
``` Python
x = [1,3,5,7,10]
# Reverse the list
x.reverse()
x # [10, 7, 5, 3, 1]
```
```
# Try it yourself!
# Suppose there is a list contains 10 elements.
# How to move the last three elements to the beginning of the list and keeping their order?
# Try to code here!
```
## 2.4 - List Sort
##### Ordering a list: sort( ) and sorted( )
sort( ) method cna be used to sorts the elements of a given list in a specific order - Ascending or Descending. sort( ) method is a built-in method that modifies the list in-place, which changes the list order. There is a sorted( ) builti-in function that builds a new sorted list from an interable, which does not modify or change the orginal list order.
e.g.
``` Python
x = [2,4,1,3]
# Using sort() method
x.sort() # Note: sort() method does not return a list
x
x = [2,4,1,3]
# Using sorted() function
sorted(x) # Note: sorted() function returns a list
x = [2,4,1,3]
# To sort the list without modifying the orginal list
y = x[:]
y.sort()
y # In this case, list x will not be affected
```
```
x = [2,4,1,3]
# Using sort() method
x.sort() # Note: sort() method does not return a list
x
x = [2,4,1,3]
# Using sorted() function
sorted(x) # Note: sorted() function returns a list
x = [2,4,1,3]
# To sort the list without modifying the orginal list
y = x[:]
y.sort()
y # In this case, list x will not be affected
```
For a list of strings, sort( ) method can also be use to sort the order of the strings alphabetically, where
| Ordering Rule | Example |
| :---: | :---: |
| Alphabetically | 'a' < 'z' |
| lower case greater than upper case | 'A' < 'a' |
| Evaluate the second letter if the first letter is the same |'ab' < 'ac' |
e.g.
``` Python
x = ['a', 'e', 'w', 'k', 'q']
x.sort()
x # return ['a', 'e', 'k', 'q', 'w']
x = ['A', 'w', 'a', 'G', 'y', 'e', 'g']
x.sort()
x # return ['A', 'G', 'a', 'e', 'g', 'w', 'y']
x = ["Life", "is", "Enchanting"]
x.sort()
x # return ['Enchanting', 'Life', 'is']
```
```
x = ['a', 'e', 'w', 'k', 'q']
x.sort()
x
x = ['A', 'w', 'a', 'G', 'y', 'e', 'g']
x.sort()
x
x = ["Life", "is", "Enchanting"]
x.sort()
x
```
##### Non-Comparable Elements:
There is something we need to be careful when using the sort( ) method: the elements within a list must be comparable, which means the sort( ) method cannot be applied to a list with string mixed with numerical value.
e.g.
``` Python
x = [1, 2, 'hello', 3]
x.sort() # this will result an error
```
```
x = [1, 2, 'hello', 3]
x.sort()
```
##### Sorting a list of lists:
If a list has an element, which is also a list, the sort( ) method can be applied. The order will be determined by evaluating the first element in each list. If they are the same value, the second element will be evaluated to determine the order.
e.g.
``` Python
x = [[3,5], [2,9], [2,3], [4,1], [3,2]]
x.sort()
x # return [[2,3], [2,9], [3,2], [3,5], [4,1]]
```
```
x = [[3,5], [2,9], [2,3], [4,1], [3,2]]
x.sort()
x
```
##### Optional parameters:
sort( ) method has two optional reverse parameters, reverse and key. When reverse=True, the sorted list is reversded or sorted in a descending order. Key serves as a key for the sort comparison
e.g.
``` Python
x = [0,1,2]
# Sorting in descending order
x.sort(reverse=True)
x # return [2, 1, 0]
x = ['eric', 'Ken', 'Thomas', 'john']
# Sorting by the length of the string
x.sort(key=len)
x # return ['Ken', 'eric', 'john', 'Thomas']
```
```
x = [0,1,2]
x.sort(reverse=True)
x
x = ['eric', 'Ken', 'Thomas', 'john']
x.sort(key=len)
x
# Try it yourself!
# Suppose there is a list and each element in the list is also a list,
# [[1,2,3], [2,1,3], [4,0,1]]
# If we need to sort the order by the second element in each list,
# how should we use the key function to do it?
x = [[1,2,3], [2,1,3], [4,0,1]]
x.sort(key=lambda x:x[1])
x
```
## 2.5 - Other List Operations
##### Uisng 'in' keyword to check of an element exisits in a list
Using the Python keyword 'in' can easily check if an element exists in a list.
e.g.
``` Python
3 in [1,3,4,5] # return True
3 not in [1,3,4,5] # return False
3 in ['one', 'two', 'three'] # return False
3 not in ['one', 'two', 'three'] # return True
```
##### Using "+" to cast two lists
To cast two lists and builidng a new one, we can use the mathematical operator "+", which will not change the original lists.
e.g.
``` Python
z = [1,2,3] + [4,5,6]
z # return [1,2,3,4,5,6]
```
##### Using " * " to create a standardized list
The multiplcation operator "*" can be used as the positional expansion opeartor. We know that we can use append( ) method for adding new element(s) to a list object, but it would be more efficient to run an application by defining the length of the list (if we know the length needed in advance).
e.g.
``` Python
z = [None] * 4
z # return [None, None, None, None]
```
As you can imagine, we can also create a new list by using the list multiplication operator by copying a list and casting into a new list.
e.g.
``` Python
z = [4, 7, 9] * 2
z # return [4, 7. 9 4, 7, 9]
```
##### Using min( ) and max( ) methods to find the minimum and maximum element
Python built-in methods min( ) and max( ) methods can be used to find the minimum and the maximum element from a list. Note that if the elements in the list are not comparable, for example, a list contains numbers and strings, it will return an error message.
e.g.
``` Python
x = [3, 8, 0, -3, 11, -8]
min(x) # return -8
max(x) # return 11
x.append('two')
min(x) # TypeError Message
```
##### Using index( ) method to find the index of an element from a list
The index( ) method searches an element in the list and returns its index. In simple terms, the index( ) method finds the given element in a list and returns its position. Note that if the same element is present more than once, the method returns the index of the first occurrence of the element.
e.g.
``` Python
x = [1, 3, 'five', 7, -2]
x.index(7) # return 3
x.index(5) # return ValueError Message
```
##### Using count( ) method to find the frequency count of a given object occur in a list
Python built-in count( ) method returns count of how many times a given object occurs in a list or the number of occurances of an element in a list. count( ) method requires a signle argument, which is the element whose count is to be found in a list.
e.g.
``` Python
x = [1,2,2,3,5,2,5]
x.count(2) # return 3
x.count(5) # return 2
x.count(4) # return 0
```
```
# Try it yourself!
# What would be the return value of len([[2,1]] * 3)?
# What's the different between Python keyword "in" and index( ) method?
# Which of the following will cause an error?
min(['a', 'b', 'c'])
max([1, 2, 'three'])
[1, 2, 3].count('one')
# Suppose there is a list, we are writing a program to remove a given element from the list safely,
# which we need to check for the existence of the element before removing.
# Change the above program and only remove element that only occur in the list
# more than one time.
```
## 2.6 - Multi-Layer List Object and Deepcopy
In this section, we are getting into an intermediate topic: multi-layer list (or nested list). For beginner, you can skip this part and come back in the later time. Nested list is usually applied for matrix calculation. We can imagine a two-dimensional 3 x 3 matrix expressed below,
e.g. 3 x 3 matrix
``` Python
m = [[0, 1, 2], [10, 11, 12], [20, 21, 22]]
m[0] # return the first row of the matrix
m[0][1] # return the second element from the first row
m[2] # return the third row of the matrix
m[2][2] # return the third element from the third row
```
```
m = [[0, 1, 2], [10, 11, 12], [20, 21, 22]]
m[0] # return the first row of the matrix
m[0][1] # return the second element from the first row
m[2] # return the third row of the matrix
m[2][2] # return the third element from the third row
```
In some cases, we may encounter a problem of changing the assignment to a nested list. For example,
``` Python
nested = [0]
original = [nexted, 1]
original # return [[0], 1]
```
In this example, the original list is referencing the nested list. When we change the element in either one, the nested list will also be changed.
``` Python
nested[0] = 'zero'
original # return [['zero'], 1]
original[0][0] = 0
nested # return [0]
original # return [[0], 1]
```
To break the connection between the nested list and original list by assigning a new list object to nested list. Now, changing the value in the nested list will not have any impact to the original list.
``` Python
nested = [2]
original # return [[0], 1]
```
```
# Creating a nested list
nested = [0]
original = [nexted, 1]
original # return [[0], 1]
# Changing the element in the nested list
nested[0] = 'zero'
# Since original list is referencing the nested list,
# so the original list will be affected.
original # return [['zero'], 1]
# Changing the element in the original list
# The nested list will also be affected.
original[0][0] = 0
nested # return [0]
original # return [[0], 1]
# Assigning a new list object to nested
# Now, nested is referencing a new list, but not the value 0.
nested = [2]
original # return [[0], 1]
```
#### deepcopy( )
We have mentioned about using slice to copy a list or using '+' or '*' operators to copy a list. The methods mentioned is called "shallow copy', which should satisfy most of the application need. However, if we are trying to copy a nested list, it is possible we may need to use a deep copy. In those cases, we need to import the **copy** package and using the deepcopy( ) function.
Reference: https://docs.python.org/3/library/copy.html
e.g.
``` Python
original = [[0], 1]
shallow = original[:]
import copy
deep = copy.depcopy(original)
```
Shallow copy constructs a new compound object and then (to the extent possible) inserts **reference** into it to the objects found in the original. In this example, both original and shallow element is referencing the same second layer list [0]. Changing the second layer list in either one will affect the other.
e.g.
``` Python
shallow[1] = 2
shallow # return [[0], 2]
# The first layar is copied, so changing the element in the first layer
# will not affect the original list.
original # return [[0], 1]
shallow[0][0] = 'zero'
shallow # return [['zero'], 2]
# The second layer of both original and shallow are referencing to a
# list [0], not copied, so changing the element in the second layer
# will affect the original list.
original # return [['zero'], 1]
```
On the other hand, a deep copy constructs a new compound object and then, recursively, inserts **copies** into it of the objects found in the original. Changing the element in a deep copy will not affect the orignal list.
e.g.
``` Python
deep[0][0] = 5
deep # [[5], 1]
original # [['zero'], 1]
```
```
original = [[0], 1]
shallow = original[:]
import copy
deep = copy.depcopy(original)
shallow[1] = 2
shallow # return [[0], 2]
# The first layar is copied, so changing the element in the first layer
# will not affect the original list.
original # return [[0], 1]
shallow[0][0] = 'zero'
shallow # return [['zero'], 2]
# The second layer of both original and shallow are referencing to a
# list [0], not copied, so changing the element in the second layer
# will affect the original list.
original # return [['zero'], 1]
deep[0][0] = 5
deep # [[5], 1]
original # [['zero'], 1]
# Try it yourself!
# Suppose there is a list:
x = [[1,2,3], [4,5,6], [7,8,9]]
# Write a program to copy the list and assign to variable y.
# And changing any element in y will have no affect to the original list x.
```
## 2.7 - Tuple
**Tuple** is a similar Python data type to a **list**, however, a **tuple** can only be created and cannot be modified. Generally speaking, **list** is a mutable objects which means it can be modified after it has been created. A **tuple** is immutable objects which cannot be modified after it has been created.
The question is that if we alreayd have a mutable **list**, why Python create the **tuple** collection? The reason is that **tuple** has a very important role in data science and programming, which cannot be replaced by a list object. For instance,
1. Program execution is faster hwen manipulating a tuple that it is for the equivalent list.
2. Sometimes we don't want data to be modified. If the values in the collection are meant to remain constant for the life of the program, using a tuple instead of a list guards against accidental modification.
3. There is another Python data type called a dictionary, which requires as one of its components a value that is of an immutable type. A tuple can be used for this purpose, whereas list can't be.
Reference: https://docs.python.org/3.4/c-api/tuple.html
Create a **tuple** in Python, instead of using the square bracket [ ], it is defined by enclosing the elements in parentheses ( ).
e.g. Create a tuple with three elements
``` Python
x = ('a', 'b', 'c')
```
Once the tuple is created, the methods apply to a list also apply to a tuple since both are "sequences" type, which means each item in the sequence is accessible by index or slice.
Note: It is easy to confuse with the parentheses and the square brackets when indexing or slicing a tuple. A tuple is created by using a parentheses, but it is accessed by the square brackets [ ].
e.g.
``` Python
# Search an element by its index
x[2] # return 'c'
# Search elements by slice method
x[1:] # return ('b', 'c')
# Check the length of the tuple
len(x) # return 3
# Find the minimum or maximum value in the tuple
min(x) # return 'a'
max(x) # return 'c'
# Check if an element exist in the tuple
5 in x # return False
5 not in x # return True
```
```
# Create a tuple with 3 elements
x = ('a', 'b', 'c')
# Search an element by its index
x[2]
# Search elements by slice method
x[1:]
# Check the length of the tuple
len(x)
# Find the minimum value in the tuple
min(x)
max(x)
# Check if an element exist in the tuple
5 in x
5 not in x
```
The major different between a **list** and a **tuple** is that **tuple** is immutable. When trying to modify a **tuple**, an error message will return to indicate Python does not support assign action to a **tuple**.
``` Python
# Modify the last element in the tuple
x[2] = 'd' # TypeError Message
```
```
# Modify the last element in the tuple
x[2] = 'd'
```
Similar to a list, we can also use the mathematical operators to stack two tuples or multiply a single tuple to create a new tuple.
e.g.
``` Python
# Slice the first two elements from a tuple
x[:2] # return ('a', 'b'])
# Stack the tuple onto another one
x + x # return ('a', 'b', 'c', 'a', 'b', 'c')
# Repeat the tuple by two times
x * 2 # return ('a', 'b', 'c', 'a', 'b', 'c')
# Create a new tuple by adding two individual tuples
x + (1, 2) # return ('a', 'b', 'c', 1, 2)```
```
# Slice the first two elements from a tuple
x[:2]
# Stack the tuple onto another one
x + x
# Repeat the tuple by two times
x * 2
# Create a new tuple by adding two individual tuples
x + (1, 2)
```
##### tuple with single element needs a comma
There is a special syntax rule we need to be careful when using a tuple: because a square bracket in Python has no specific application, it is obvious when we are creating a list, [ ] means an empty list, and [8] is a list with single element. However, parentheses ( ) is widely used in mathematic operations, such that when writing a Python code, (x + y) means adding x and y before putting the combine object to a tuple or using it as a priority operator.
To resolve the issue mentioned above, Python requires the trailing comma (or comma at the end) for a single element tuple. Let's take a look of an example.
e.g.
``` Python
x = 3
y = 4
# Adding two elements
(x + y) # return 7
# Create a single element tuple with the sum of the two elements
(x+y, ) # return (7, )
# Create an empty element
() # return ( )
```
##### Packing and Unpacking of Tuples
Python has tuple assignment feature which enables users to assign more than one variable at a time. Here is an example to assign multiple elements to multiple variables by their positions.
e.g. Assigning 4 elements to 4 variables by tuple assignment
``` Python
(one, two, three, four) = (1, 2, 3, 4)
one # return 1
three # return 3
# The assignment can be re-written as the following,
one, two , three, four = 1, 2, 3, 4
# One line of code now replacing four lines of code
one = 1
two = 2
three = 3
four = 4
```
In the previous example, Python automatically **packs** the data on the right-hand side of the equal sign into a tuple (**Packing**) and assign them to the variable on the left-hand side of the equal sign,then automatically **unpack** the data to each assignment (**Unpacking**).
``` Python
one, two, three, four = 1, 2, 3, 4 # packing 1, 2, 3, 4 into a tuple (1, 2, 3, 4)
one, two, three, four = (1, 2, 3, 4) # unpacking (1, 2, 3, 4) and assign to each variable on the left
```
This feature is especially important when apply to swapping two variables. In Python, we do not need to write our code like following:
``` Python
temp = var1
var1 = var2
var2 = temp
```
Instead, we can write one line of code for the same purpose.
``` Python
var1, var2 = var2, var1
```
```
# Packing and Unpacking also applies to differet squence data structures
# Tuple
v1, v2, v3 = 1, 2, 3
# List
v1, v2, v3 = [1, 2, 3]
# String
v1, v2, v3 = "abc"
```
Packing and Unpacking not only supports multiple assignment, it can also apply to the Python control flow tools. Here is an example to demostrate how a function return a user data by its id.
``` Python
def get_user_info(id):
# using the user ID to retrieve the user data
return name, age, e-mail
# the get_user_info() function is going to return three elements
name, age, e-mail = get_user_info(id)
```
In this example, Python is packing the three elements from the return of the function into a tuple object, so the actualy return object is only one. We can also apply this function to a for loop to retrieve user infomation and assign them into three variables.
In other programming languages, such as C and Java, function can only return a single object. If we need multiple elements return from a function, we need to assign them into a matrix, list object, vector object, etc. Python Packing and Unpacking logic offers a cleaner or more elegant solution.
##### Unmatch Assignment
When assigning to multiple variables, make sure the number of elements matches with the variables. If the number of element on the right-hand side does not match with the number of variables on the left-hand side, an error message will return.
e.g.
``` Python
one, two, three = 1, 2, 3, 4 # error message return
```
To avoid such problem, Python allows to use "*" to absorb the extra elements.
e.g.
``` Python
x = (1, 2, 3, 4)
a, b, c* = x
a, b, c # return (1, 2, [3, 4])
a, b*, c = x
a, b, c # return (1, [2, 3], 4)
a*, b, c = x
a, b, c # return ([1, 2], 3, 4)
a, b, c, d, e* = x
a, b, c, d, e # return (1, 2, 3, 4, [])
```
Note: Adding * to the variable allows the variable to absorb all the extra element and assign them to a list. If no extra element to be absorbed, an empty list will return.
For some Python programmer, if only numbers of specific elements are needed to be assigned, they will usually create a "**place holder**" for assigning the unused elements. Here is an example.
e.g.
``` Python
x = (1, 2, 3, 4, 5)
# Only assign the first two elements from a tuple
a, b, *_ = x
```
In this case, the underscore "_" is just like a place holder, so the last three elements will not be assigned to anything.
You can also use tuple and list to pack or unpack data manually. Here are some examples.
e.g.
``` Python
[a, b] = [1, 2]
[c, d] = 3, 4
[e, f] = (5, 6)
(g, h) = 7, 8
i, j = [9, 10]
k, l = (11, 12)
a # return 1
[b, c, d] # return [2, 3, 4]
(e, f, g) # return (5, 6, 7)
h, i, j, k, l # return (8, 9, 10, 11, 12)
```
```
# an error message will return if the two sides are not match with number
one, two, three = 1, 2, 3, 4
# To avoid such problem, Python allows to use "*" to absorb the extra elements
x = (1, 2, 3, 4)
a, b, c* = x
a, b, c
a, b*, c = x
a, b, c
a*, b, c = x
a, b, c
a, b, c, d, e* = x
a, b, c, d, e
```
##### Switching between tuple and list
Using a list( ) function, we can switch a tuple into a list object. The list( ) function also applies to any sequence data structure and create the same sequence list object. On the other hand, the tuple( ) function can turn any seqence data structure (including list object) into a tuple. Here are some examples.
e.g.
``` Python
list((1, 2, 3, 4)) # return [1, 2, 3, 4]
tuple([1, 2, 3, 4]) # return (1, 2, 3, 4)
```
An interesting application for using the list( ) or tuple( ) functions is that it can split a string efficiently.
e.g.
``` Python
list("Hello") # return ["H", "e", "l", "l", "o"]
tuple("Hello") # return ("H", "e", "l", "l", "o")
```
# Try it yourself!
# Explain why the following operations do not apply to a tuple x = (1, 2, 3, 4)
x.append(1)
x[1] = "hello"
del x[2]
# Suppose we have a tuple x = (3, 1, 4, 2), how are we going to sort the order of this tuple?
x = (3, 1, 4, 2)
```
## 2.8 - Set
A Set is an unordered collection data type that is iterable, mutable and has no duplicate elements. Python set class represents the mathematical notion of a set. A set can be created by placing a comma-separated list of elements within braces (or curly brackets). We can use Since sets are unordered, we cannot access items using index or slice like we do in lists or tuples. Some of the operations we used on lists or tuples still apply to the sets. Here are some operation examples.
Reference: https://docs.python.org/3/library/stdtypes.html#set-type-set-frozenset
e.g.
``` Python
# Create a set with braces or curly brackets
x = {1, 2, 1, 2, 1, 2, 1, 2}
x # return {1, 2}
# We can also create a set with set( ) function
x = set([1, 2, 3, 1, 2, 3, 1, 2, 3])
x # return {1, 2, 3}
# We can use add( ) function to add element to a set
x.add(6)
x # return {1, 2, 3, 6}
# We can use remove( ) function to remove element from a set
x.remove(3)
x # return {1, 2, 6}
# We can use Python keyword "in" to check if an element exists in a set
4 in x # return False
1 in x # return True
# We can use "|" to return the collection of element from two sets (similar to OR logic operation)
y = {1, 7, 9}
x | y # return {1, 2, 6, 7, 9}
# Use "&" to return the intercept elements from two sets
x & y # return {1}
# Use "^" to find the symmeric difference collection (similar to XOR)
x ^ y # return {2, 6, 7, 9}
```
##### frozenset
As mentioned above, elements in a set must be an immutable data type. However, set itself is mutable, therefore, a set cannot be assigned to another set. To resolve this issue, Python provide a data type called "frozenset", which is an immutable set and can be assigned to anothe set. Here is an example.
e.g.
``` Python
x = set([1, 2, 3, 1, 3, 5])
z = frozenset(x)
z # return frozenset({1, 2, 3, 5})
z.add(6) # return an error message because it is immutable
# Adding an immutable set z to a mutable set x
x.add(z)
x # return {1, 2, 3, 5, fronzenset({1, 2, 3, 5})}
```
```
# Try it yourself!!
# If we are using below list to create a set, how many elements will be in the set?
[1, 2, 5, 1, 0, 2, 3, 1, 1, (1, 2, 3)]
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import plotly.express as px
data = pd.read_csv('https://raw.githubusercontent.com/PacktWorkshops/The-Data-Analysis-Workshop/master/Chapter09/Datasets/energydata_complete.csv')
data.head()
data.isnull().sum()
df1 = data.rename(columns = {
'date' : 'date_time',
'Appliances' : 'a_energy',
'lights' : 'l_energy',
'T1' : 'kitchen_temp',
'RH_1' : 'kitchen_hum',
'T2' : 'liv_temp',
'RH_2' : 'liv_hum',
'T3' : 'laun_temp',
'RH_3' : 'laun_hum',
'T4' : 'off_temp',
'RH_4' : 'off_hum',
'T5' : 'bath_temp',
'RH_5' : 'bath_hum',
'T6' : 'out_b_temp',
'RH_6' : 'out_b_hum',
'T7' : 'iron_temp',
'RH_7' : 'iron_hum',
'T8' : 'teen_temp',
'RH_8' : 'teen_hum',
'T9' : 'par_temp',
'RH_9' : 'par_hum',
'T_out' : 'out_temp',
'Press_mm_hg' : 'out_press',
'RH_out' : 'out_hum',
'Windspeed' : 'wind',
'Visibility' : 'visibility',
'Tdewpoint' : 'dew_point',
'rv1' : 'rv1',
'rv2' : 'rv2'
})
df1.head()
df1.tail()
df1.describe()
lights_box = sns.boxplot(df1.l_energy)
l = [0, 10, 20, 30, 40, 50, 60, 70]
counts = []
for i in l:
a = (df1.l_energy == i).sum()
counts.append(a)
counts
lights = sns.barplot(x = l, y = counts)
lights.set_xlabel('Energy Consumed by Lights')
lights.set_ylabel('Number of Lights')
lights.set_title('Distribution of Energy Consumed by Lights')
((df1.l_energy == 0).sum() / (df1.shape[0])) * 100
new_data = df1
new_data.drop(['l_energy'], axis = 1, inplace = True)
new_data.head()
app_box = sns.boxplot(new_data.a_energy)
out = (new_data['a_energy'] > 200).sum()
out
(out/19735) * 100
out_e = (new_data['a_energy'] > 950).sum()
out_e
(out_e/19735) * 100
energy = new_data[(new_data['a_energy'] <= 200)]
energy.describe()
new_en = energy
new_en['date_time'] = pd.to_datetime(new_en.date_time, format = '%Y-%m-%d %H:%M:%S')
new_en.head()
new_en.insert(loc = 1, column = 'month', value = new_en.date_time.dt.month)
new_en.insert(loc = 2, column = 'day', value = (new_en.date_time.dt.dayofweek)+1)
new_en.head()
import plotly.graph_objs as go
app_date = go.Scatter(x = new_en.date_time, mode = "lines", y = new_en.a_energy)
layout = go.Layout(title = 'Appliance Energy Consumed by Date', xaxis = dict(title='Date'), yaxis = dict(title='Wh'))
fig = go.Figure(data = [app_date], layout = layout)
fig.show()
app_mon = new_en.groupby(by = ['month'], as_index = False)['a_energy'].sum()
app_mon
app_mon.sort_values(by = 'a_energy', ascending = False).head()
plt.subplots(figsize = (15, 6))
am = sns.barplot(app_mon.month, app_mon.a_energy)
plt.xlabel('Month')
plt.ylabel('Energy Consumed by Appliances')
plt.title('Total Energy Consumed by Appliances per Month')
plt.show()
app_day = new_en.groupby(by = ['day'], as_index = False)['a_energy'].sum()
app_day
app_day.sort_values(by = 'a_energy', ascending = False)
plt.subplots(figsize = (15, 6))
ad = sns.barplot(app_day.day, app_day.a_energy)
plt.xlabel('Day of the Week')
plt.ylabel('Energy Consumed by Appliances')
plt.title('Total Energy Consumed by Appliances')
plt.show()
```
**Exercise 9.05: Plotting distributions of the temperature columns**
```
col_temp = ['kitchen_temp', 'liv_temp', 'laun_temp', 'off_temp',
'bath_temp', 'out_b_temp', 'iron_temp', 'teen_temp', 'par_temp']
temp = new_en[col_temp]
temp.head()
temp.hist(bins = 15, figsize = (12, 16))
```
| github_jupyter |
```
# Module import
from IPython.display import Image
import sys
import pandas as pd
# To use interact -- IPython widget
from __future__ import print_function
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
# append to path the folder that contains the analytic scanner
sys.path.append('../GaiaLab/scan/analytic_scanner')
# local imports
#from gaia_analytic_toymodel import *
from scipy import interpolate
import timeit
import frame_transformations as ft
from scanner import Scanner
from satellite import Satellite
from source import Source
import constants as const
from quaternion_implementation import Quaternion
from agis import Agis
from agis import Calc_source
from agis_functions import *
from analytic_plots import *
# Ipython magics
%load_ext autoreload
%autoreload 2
# %matplotlib notebook
# %matplotlib widget
# %matplotlib ipympl
```
# **Some functions:**
# **Initializing objects:**
```
# # create all the objects we will need:
# parameters for the notebook
t_init = 0 #1/24/60 # TODO make code work also for t_init != 0
t_end = t_init + 1/2 # 365*5
my_dt = 1/24/20 # [days]
spline_degree = 3 # actually it is the spline degree
gaia = Satellite(ti=t_init, tf=t_end, dt= my_dt, k=spline_degree)
print('Sat created')
my_times = np.linspace(t_init, t_end, num=100, endpoint=False)
real_sources = []
calc_sources = []
for t in my_times:
alpha, delta = generate_observation_wrt_attitude(gaia.func_attitude(t))
real_src_tmp = Source(str(t),np.degrees(alpha), np.degrees(delta), 0, 0, 0, 0)
calc_src_tmp = Calc_source('calc_'+str(t), [t], source=real_src_tmp)
real_sources.append(real_src_tmp)
calc_sources.append(calc_src_tmp)
print('Sources created!')
fig = plt.figure()
plt.subplot(111, projection="hammer")
alphas = []
deltas = []
#print('my_times: ', my_times)
print('len(real_sources): ', len(real_sources))
for i, s in enumerate(real_sources):
alpha, delta = s.alpha-np.pi, s.delta
#print(i, alpha, delta)
alphas.append(alpha)
deltas.append(delta)
if i==0:
plt.plot(alpha, delta, 'sk', label='first star')
else:
#plt.plot( alpha, delta,',')
pass
plt.scatter(alphas, deltas, c=my_times, marker='+', s=(72./fig.dpi)**2, cmap='jet', alpha=0.8,
label='attitude', lw=2)
plt.title("Hammer Projection of the Sky")
plt.legend(loc=9, bbox_to_anchor=(1.1, 1))
plt.grid(True)
# test if source and calc source are equal (as they should be)
np.testing.assert_array_almost_equal(np.array(real_sources[0].get_parameters()[0:5]), calc_sources[0].s_params)
# create Solver
Solver = Agis(gaia, calc_sources, real_sources, attitude_splines=[gaia.s_w, gaia.s_x, gaia.s_y, gaia.s_z], spline_degree=spline_degree,
attitude_regularisation_factor=1e-2)
# Ignore this cell if you don't want to modify the initial attitude
# Can be used to check that when recreating the splines in the solver we (almost) do not create additional errors
Solver.actualise_splines()
print('Error before Noise: ', Solver.error_function())
print('Errors after noise of attitude (not representatif):', error_between_func_attitudes(my_times, gaia.func_attitude, Solver.get_attitude))
c_noise = Solver.att_coeffs * np.random.rand(Solver.att_coeffs.shape[0], Solver.att_coeffs.shape[1]) * 1e-8
print('c_noise shape: ', c_noise.shape)
Solver.att_coeffs[:] = Solver.att_coeffs[:] + c_noise[:]
Solver.actualise_splines()
#Solver.set_splines_basis()
print('Error after Noise: ', Solver.error_function())
print('Errors after noise of attitude (not representatif):', error_between_func_attitudes(my_times, gaia.func_attitude, Solver.get_attitude))
```
# **About splines:**
```
compare_attitudes(gaia, Solver, my_times)
gaia.s_w.get_knots()
```
## test: Play with splines:
### Test compute_coeff_basis_sum:
```
# two quick tests:
np.testing.assert_array_almost_equal(t_init, my_times[0]) # could fail if we change how we define my_times
np.testing.assert_array_almost_equal(t_init, np.array(gaia.storage)[:, 0][0]) # should not fail
to_fit = gaia.s_w
plt.plot(my_times, to_fit(my_times), label='to be reconstructed')
B = Solver.att_bases
a = Solver.att_coeffs[0]
def Splines(t, a, B, my_times):
time_index = list(my_times).index(t)
left_index = get_left_index(Solver.att_knots, t, M=Solver.M)
S = 0
for i in range(len(a)):
S += a[i]*B[i, time_index]
S2 = compute_coeff_basis_sum(Solver.att_coeffs, Solver.att_bases, left_index, Solver.M, time_index)
return S, S2
S_evaluated = []
S_bis = []
for t in my_times:
S, S2 = Splines(t, a, B, my_times)
S_evaluated.append(S)
S_bis.append(S2[0])
plt.plot(my_times, S_evaluated, '+', label='manual')
plt.plot(my_times, S_bis, 'x', label='with_coeff_basis_sum!')
plt.grid(), plt.legend(), plt.show()
# Solver.iterate(1)
c, t, s = extract_coeffs_knots_from_splines([gaia.s_w, gaia.s_x, gaia.s_y, gaia.s_z], k=3)
a_ref = to_fit.get_coeffs()
plt.plot(a_ref, label='ref coeffs')
plt.plot(Solver.att_coeffs[0], label='Solver')
plt.plot(c[0], '+', label='Coeffs extracted as in solver')
plt.plot(Solver.c[0], 'rx', label='c')
plt.grid(), plt.legend(), plt.show()
```
# **Tests:**
## Test_0:
Test that the sources are aligned with the attitude
```
import astropy.units as u
eta,zeta = calculated_field_angles(calc_sources[0], gaia.func_attitude(my_times[0]), gaia, my_times[0])
print(np.array([eta,zeta])*u.rad.to(u.mas))
print('for alpha delta',calc_sources[0].s_params[:2]*u.rad.to(u.mas),'mas')
del u
```
OK
## Test_1:
Test the design and normal matrix:
line 366 in agis.py should be checked: dR_da_n = dR_da_i(dR_dq, self.att_bases[:, n_index, obs_time_index])
some times are out of the knots range...
```
left_index = get_left_index(Solver.att_knots, my_times[0], M=Solver.M)
```
## Test_2:
```
Solver.att_knots.shape[0]
print(Solver.att_knots.shape[0])
print(Solver.att_coeffs.shape[1])
N_aa = Solver.compute_attitude_LHS()
N_aa[-1:,-1:]
fig, axs = plt.subplots(1, 2, figsize=(24, 12))
plot1 = axs[0].imshow(np.abs(N_aa), vmin=None, vmax=1)
axs[0].set_title("$|N_{aa}|$")
A = N_aa.copy()
threshold = 0
A[np.where(A==threshold)] = A.max()
plot2 = axs[1].imshow(A, vmin=None, vmax=None)
axs[1].set_title("$N_{aa}$ , with zeros to max(A)")
fig.colorbar(plot1, ax=axs[0])
fig.colorbar(plot2, ax=axs[1])
fig.suptitle('Matrix visualization')
print('Is the matrix symmetric? ', helpers.check_symmetry(N_aa))
eig_vals, eig_vecs = np.linalg.eigh(N_aa)
plt.plot(eig_vals)
plt.yscale('log')
plt.title('eigenvalues'), plt.ylabel('log scale (value)'), plt.xlabel('#eigval'), plt.grid()
```
# **TEST with separated matrices:**
```
a = np.array([[1,2,3,4,5,6,7,8,9,10,0], [1,2,3,4,5,6,7,8,8,9,0]])
a[0,1:-1:2]
print(N_aa.shape)
N_aa_w = N_aa[0::4, 0::4]
N_aa_x = N_aa[1::4, 1::4]
N_aa_y = N_aa[2::4, 2::4]
N_aa_z = N_aa[3::4, 3::4]
N_aa_list = [N_aa_w, N_aa_x, N_aa_y, N_aa_z]
print('N_aa_w.shape', N_aa_w.shape)
print('N_aa_z.shape', N_aa_z.shape)
fig, axs = plt.subplots(4, 2, figsize=(24, 24))
type_list = ['w', 'x', 'y', 'z']
for i in range(4):
A = N_aa_list[i].copy()
plot1 = axs[i, 0].imshow(np.abs(A), vmin=None, vmax=1)
axs[i, 0].set_title("$|N_{aa}|$ type:"+type_list[i])
threshold = 0
A[np.where(A==threshold)] = A.max()
plot2 = axs[i, 1].imshow(A, vmin=None, vmax=None)
axs[i, 1].set_title("$N_{aa}$ , with zeros to max(A)"+type_list[i])
fig.colorbar(plot1, ax=axs[i, 0])
fig.colorbar(plot2, ax=axs[i, 1])
fig.suptitle('Matrix visualization')
for i, Naa in enumerate(N_aa_list):
eig_vals, eig_vecs = np.linalg.eigh(Naa)
plt.plot(eig_vals, label=type_list[i])
print('Is the matrix', type_list[i], 'symmetric? ', helpers.check_symmetry(Naa))
print('Condition number: ', np.linalg.cond(Naa))
print('Condition number (eig_max/eig_min): ', np.abs(eig_vals.max())/np.abs(eig_vals.min()))
print('Rank: ', np.linalg.matrix_rank(Naa, hermitian=True))
print('det(N_aa):', np.linalg.det(Naa),'\n')
plt.yscale('log'), plt.legend(), plt.grid()
plt.title('eigenvalues'), plt.ylabel('log scale (value)'), plt.xlabel('#eigval')
plt.show()
```
## **Test dR_dq = compute_dR_dq(calc_source, Solver.sat, attitude, t_L):**
**Checking that attitudes returns something similar to coeff_basis_sum:**
```
my_time_number = 0
t_L = my_times[my_time_number]
time_index = list(my_times).index(t_L)
left_index = get_left_index(Solver.att_knots, t_L, M=Solver.M)
coeff_basis_sum = compute_coeff_basis_sum(Solver.att_coeffs, Solver.att_bases, left_index, Solver.M, time_index)
calc_source = Solver.calc_sources[my_time_number]
attitude = Solver.get_attitude(t_L)
att2 = Quaternion(coeff_basis_sum[0], coeff_basis_sum[1], coeff_basis_sum[2], coeff_basis_sum[3])
dR_dq_1 = compute_dR_dq(calc_source, Solver.sat, attitude, t_L)
dR_dq_2 = compute_dR_dq(calc_source, Solver.sat, att2, t_L)
# np.testing.assert_array_almost_equal(dR_dq_1, dR_dq_2) # should not fail
print('dR_dq_1:', dR_dq_1)
print('dR_dq_2:', dR_dq_2)
```
**Checking dR_da:**
```
m_index = 3
dR_da_m = dR_da_i(dR_dq_1, Solver.att_bases[m_index, time_index])
dR_da_m
fig, axs = plt.subplots(1, 4, figsize=(24, 6))
for i, m in enumerate(range(m_index, m_index+4)):
axs[i].plot(Solver.att_bases[m, :], label='m='+str(m)), axs[i].grid(), axs[i].legend()
plt.show()
```
## Extra tests:
```
for i in range(0, 3):
# np.testing.assert_array_equal(Solver.att_bases[i], Solver.att_bases[i+1])
pass
```
### Taking a look at the splines:
```
gaia.s_w.get_knots()
basis = Solver.att_bases
f1 = plt.figure()
for b in basis:
# plt.plot(range(len(my_times)), b)
plt.plot(my_times, b)
plt.plot(Solver.att_knots, np.zeros(Solver.att_knots.shape), '+', color='orange', label='knots')
plt.grid(), plt.legend()
plt.show()
print('shape Solver.att_bases:', Solver.att_bases.shape)
print('time ranging from: ', my_times[0], ' to ', my_times[-1])
print('N=',Solver.att_coeffs.shape, ' || k=', Solver.att_knots.shape, ' || M=',Solver.M)
```
# **Taking a look to intermediate functions:**
**The N_aa[n,m] block:**
```
# set index and initialize block
m_index, n_index = (12, 14)
Naa_mn = np.zeros((4, 4))
# WARNING: here we take the knots of w since they should be the same for the 4 components
observed_times_m = get_times_in_knot_interval(Solver.all_obs_times, Solver.att_knots, m_index, Solver.M)
observed_times_n = get_times_in_knot_interval(Solver.all_obs_times, Solver.att_knots, n_index, Solver.M)
observed_times_mn = np.sort(helpers.get_lists_intersection(observed_times_m, observed_times_n))
union_times = np.sort(list(set(list(observed_times_m) + list(observed_times_n))))
print('knot m:', Solver.att_knots[m_index], 'knot m+M:', Solver.att_knots[m_index+Solver.M])
print('observed_times_mn:', observed_times_m)
print('knot n:', Solver.att_knots[n_index], 'knot n+M:', Solver.att_knots[n_index+Solver.M])
print('observed_times_mn:', observed_times_n)
print('\nobserved_times_mn:', observed_times_mn)
to_plot_n, to_plot_m, to_plot_dn, to_plot_dm = ([], [], [], [])
for t in union_times:
obs_time_index = list(Solver.all_obs_times).index(t)
to_plot_n.append(Solver.att_bases[n_index, obs_time_index])
to_plot_m.append(Solver.att_bases[m_index, obs_time_index])
for t in observed_times_mn:
obs_time_index = list(Solver.all_obs_times).index(t)
to_plot_dn.append(Solver.att_bases[n_index, obs_time_index])
to_plot_dm.append(Solver.att_bases[m_index, obs_time_index])
plt.plot(union_times, to_plot_n, '-b', label='$B_n$')
plt.plot(union_times, to_plot_m, '--r', label='$B_m$')
plt.plot(observed_times_mn, to_plot_dn, 'oc', label='for dR/d$a_n$') # label='used for $\\frac{dR}{da_n}$'
plt.plot(observed_times_mn, to_plot_dm, 'xk', label='for dR/d$a_m$')
plt.grid(), plt.legend(), plt.title('Used spline bases'), plt.show()
# compute dDL_da block
dR_da_mn = np.zeros((4, 4))
for i, t_L in enumerate(observed_times_mn):
source_index = Solver.get_source_index(t_L)
calc_source = Solver.calc_sources[source_index]
attitude = Solver.get_attitude(t_L)
left_index = get_left_index(Solver.att_knots, t_L, M=Solver.M)
obs_time_index = list(Solver.all_obs_times).index(t_L)
if i<3:
print('***** iteration #', i ,' *****')
print('t:',t_L,' // source_index:', source_index, ' // time_index:', obs_time_index, )
print('time according to time_index:', Solver.all_obs_times[obs_time_index], ' // left_index',left_index)
print('before_left_index: ',Solver.att_knots[left_index-1],
' // knot left_index:',Solver.att_knots[left_index],
' // knot after left_index:',Solver.att_knots[left_index+1])
# Ccompute the original objective function part
# # WARNING: Here we put the Across scan and the along scan together
dR_dq = compute_dR_dq(calc_source, Solver.sat, attitude, t_L)
dR_da_m = dR_da_i(dR_dq, Solver.att_bases[m_index, obs_time_index])
dR_da_n = dR_da_i(dR_dq, Solver.att_bases[n_index, obs_time_index])
if i == 0:
print('**** m:', m_index, '**** n:', n_index, '**** i:', i)
print('dR_dq: ', dR_dq)
print('dR_da_m', dR_da_m)
print('dR_da_n', dR_da_n)
print('dR_da_n @ dR_da_m.T', dR_da_n @ dR_da_m.T)
dR_da_mn += dR_da_n @ dR_da_m.T
dR_da_mn
dR_dq * Solver.att_bases[m_index, obs_time_index]
# compute regularisation block
reg_block_mn = np.zeros((4, 4))
for i, t_L in enumerate(observed_times_mn):
source_index = Solver.get_source_index(t_L)
calc_source = Solver.calc_sources[source_index]
attitude = Solver.get_attitude(t_L)
left_index = get_left_index(Solver.att_knots, t_L, M=Solver.M)
obs_time_index = list(Solver.all_obs_times).index(t_L)
# Compute the regulation part
coeff_basis_sum = compute_coeff_basis_sum(Solver.att_coeffs, Solver.att_bases,
left_index, Solver.M, obs_time_index)
# dDL_da_n = compute_DL_da_i(coeff_basis_sum, self.att_bases, obs_time_index, n_index)
# dDL_da_m = compute_DL_da_i(coeff_basis_sum, self.att_bases, obs_time_index, m_index)
dDL_da_n = compute_DL_da_i_from_attitude(attitude, Solver.att_bases, obs_time_index, n_index)
dDL_da_m = compute_DL_da_i_from_attitude(attitude, Solver.att_bases, obs_time_index, m_index)
reg_block_mn += Solver.attitude_regularisation_factor**2 * dDL_da_n @ dDL_da_m.T
reg_block_mn
Naa_mn = dR_da_mn + reg_block_mn
print('Just computed: \n',Naa_mn)
print('Solver_version: \n', Solver.compute_Naa_mn(m_index, n_index))
np.testing.assert_array_almost_equal(Naa_mn, Solver.compute_Naa_mn(m_index, n_index)) # just to check that they are the sames
```
# **Taking a look at the normal matrix:**
```
N_aa = Solver.compute_attitude_LHS()
print('N_aa computed')
h = Solver.compute_attitude_RHS()
print('shape of h: ', h.shape)
print('h/4: ', h.shape[0]/4)
print('my_times shape:', my_times.shape)
print('coeffs shape:',Solver.att_coeffs.shape)
print('Matrix dimension: ', N_aa.shape)
print('Is the matrix symmetric? ', helpers.check_symmetry(N_aa))
eig_vals, eig_vecs = np.linalg.eigh(N_aa)
print('Condition number: ', np.linalg.cond(N_aa))
print('min-max eig_vals: ',eig_vals.min(), eig_vals.max())
print('Condition number (eig_max/eig_min): ', np.abs(eig_vals.max())/np.abs(eig_vals.min()))
print('Rank: ', np.linalg.matrix_rank(N_aa, hermitian=True))
print('det(N_aa):', np.linalg.det(N_aa))
print('smallest eig_vals: ', np.sort(eig_vals)[0:4])
print('biggest eigenvalues: ', eig_vals[-5:-1])
print('eig_vals product: ', np.prod(eig_vals), ' || my_times shape:', my_times.shape,
' || coeffs shape:',Solver.att_coeffs.shape, ' || det(N_aa):', np.linalg.det(N_aa))
print('eigenvalues: ', eig_vals[0])
# print('N_aa:', N_aa)
print(Solver.compute_Naa_mn(0,1))
print(Solver.compute_Naa_mn(8,9))
print(Solver.compute_Naa_mn(9,10))
if np.linalg.det(N_aa) > 0:
L = np.linalg.cholesky(N_aa)
print(L)
else:
print('Determinant of N_aa not positive')
np.linalg.solve(N_aa, h)[0:5]
helpers.plot_sparsity_pattern(N_aa[10:18, 0:10], 1)
helpers.plot_sparsity_pattern(N_aa[0:40, 0:40], 4)
helpers.plot_sparsity_pattern(N_aa[-41:-1, -41:-1], 4)
N_aa = Solver.compute_attitude_LHS()
tol = 1
A = N_aa[0:40, 0:40]
B = A #np.zeros(A.shape)
# B[np.where( np.abs(A) > tol)] = 1
plt.matshow(B, fignum=None)
plt.colorbar()
plt.xticks(np.arange(0, B.shape[0], 4))
plt.yticks(np.arange(0, B.shape[0], 4))
plt.grid()
plt.show()
print(N_aa.min())
A = N_aa.copy()[0:44,0:44]
my_min = np.amin(A)
print('min_of_A:', my_min)
threshold = 0 #A.min()
A[np.where(A==threshold)] = A.max()
print('min of A: ',A.min())
plt.matshow(A, fignum=None)
plt.colorbar()
plt.xticks(np.arange(0, A.shape[0], 4))
plt.yticks(np.arange(0, A.shape[0], 4))
plt.grid()
plt.show()
```
## _And the righ hand side:_
```
# Plotting h
plt.matshow(h)
plt.colorbar()
plt.grid()
plt.show()
```
# **Iterating:**
```
Solver.verbose=False
# Solver.reset_iterations()
Solver.iterate(1)
plt.figure(figsize=(5,5))
c, t, s = extract_coeffs_knots_from_splines([gaia.s_w, gaia.s_x, gaia.s_y, gaia.s_z], k=3)
plt.plot(c[0], '+', label='Coeffs extracted as in solver')
plt.plot(Solver.att_coeffs[0], label='Solver')
plt.grid(), plt.legend(), plt.show()
multi_compare_attitudes(gaia, Solver, my_times)
print('MAGNITUDE:', Solver.get_attitude(0.005, unit=False).magnitude)
Solver.get_attitude(0.005, unit=False).magnitude
Solver.iterate(10)
Solver.get_attitude(0.005).magnitude
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import xarray as xr
ds = xr.open_dataset('/scratch/05488/tg847872/fluxbypass_aqua/AndKua_aqua_SPCAM3.0_sp_fbp_f4.cam2.h1.0001-01-09-00000.nc',
decode_times=False)
dsp4 = xr.open_dataset('/scratch/05488/tg847872/debug/AndKua_aqua_SPCAM3.0_sst_p4.cam2.h1.0000-01-01-00000.nc',
decode_times=False)
dsw1 = xr.open_dataset('/scratch/05488/tg847872/debug/SST_debug01.cam2.h1.0000-01-01-00000.nc',
decode_times=False)
TS = ds.TS.isel(time=0).values - 273.15
plt.imshow(TS); plt.colorbar();
plt.imshow(dsp4.TS.isel(time=0).values - 273.15); plt.colorbar();
plt.imshow(dsw1.TS.isel(time=0).values - 273.15); plt.colorbar();
plt.imshow(dsw1.TS.isel(time=0).values - ds.TS.isel(time=0).values - 273.15); plt.colorbar();
lat, lon = ds.lat.values, ds.lon.values
rlat, rlon = np.deg2rad(lat), np.deg2rad(lon)
nlat, nlon = lat.shape[0], lon.shape[0]; nlat, nlon
rlat
np.rad2deg(np.pi/3)
np.deg2rad(30.), 30/180*np.pi
def C5N(const=2.):
sst = np.empty((nlon, nlat))
for j in range(nlat):
if lat[j] < -60. or lat[j] > 60.:
zeta = 1.
elif lat[j] > 5. and lat[j] <= 60.:
zeta = (np.sin(np.pi*(lat[j]-5.)/110.))**2
elif lat[j] >= -60 and lat[j] < 5.:
zeta = (np.sin(np.pi*(lat[j]-5.)/130.))**2
for i in range(nlon):
sst[i,j] = const + 27/2. * (2 - zeta - zeta**2)
return sst
sst_c5n = C5N()
plt.imshow(sst_c5n.T); plt.colorbar(shrink=0.6);
def CTRL():
sst = np.empty((nlon, nlat))
for j in range(nlat):
for i in range(nlon):
if rlat[j] > -(np.pi/3) and rlat[j] < (np.pi/3):
sst[i,j] = 27 * (1 - np.sin(3*rlat[j]/2)**2)
else:
sst[i,j] = 0.
return sst
sst_ctrl = CTRL()
plt.imshow(sst_ctrl.T); plt.colorbar(shrink=0.6);
def s3KW1():
sst = np.empty((nlon, nlat))
lat_d, lon_0, xi = np.deg2rad(30), np.deg2rad(0), 3
shift = np.deg2rad(5)
for j in range(nlat):
for i in range(nlon):
if rlat[j] > (-lat_d+shift) and rlat[j] < (lat_d+shift):
sst[i,j] = xi * np.cos(rlon[i] - lon_0) * np.cos(np.pi/2 * (rlat[j]-shift)/lat_d)**2
else:
sst[i,j] = 0.
return sst
pert_3KW1 = s3KW1()
plt.imshow(pert_3KW1.T); plt.colorbar(shrink=0.6);
sst_3KW1 = sst_c5n + pert_3KW1
plt.imshow(sst_3KW1.T); plt.colorbar(shrink=0.6);
plt.imshow(sst_3KW1.T - sst_c5n.T); plt.colorbar(shrink=0.6);
dcrash = xr.open_dataset('/scratch/05488/tg847872/nnfullphy_fbp8_E001_3kw1/nnfullphy_fbp8_E001_3kw1.cam2.h1.0000-01-01-00000.nc',
decode_times=False)
dcrash.TS.isel(time=0).plot()
```
| github_jupyter |
This tutorial was implemented on Macbook pro (15-inch, 2018)
# Simulate scRNA-seq data
```
rm(list = ls())
library(splatter)
library(rhdf5)
i <- 1 ## set random seed
simulate <- function(nGroups=3, nGenes=2500, batchCells=1500, dropout=0) # change dropout to simulate various dropout rates
{
if (nGroups > 1) method <- 'groups'
else method <- 'single'
group.prob <- rep(1, nGroups) / nGroups
sim <- splatSimulate(group.prob=group.prob, nGenes=nGenes, batchCells=batchCells,
dropout.type="experiment", method=method,
seed=100+i, dropout.shape=-1, dropout.mid=dropout, de.facScale=0.25)
counts <- as.data.frame(t(counts(sim)))
truecounts <- as.data.frame(t(assays(sim)$TrueCounts))
dropout <- assays(sim)$Dropout
mode(dropout) <- 'integer'
cellinfo <- as.data.frame(colData(sim))
geneinfo <- as.data.frame(rowData(sim))
list(sim=sim,
counts=counts,
cellinfo=cellinfo,
geneinfo=geneinfo,
truecounts=truecounts)
}
sim <- simulate()
simulation <- sim$sim
counts <- sim$counts
geneinfo <- sim$geneinfo
cellinfo <- sim$cellinfo
truecounts <- sim$truecounts
dropout.rate <- (sum(counts==0)-sum(truecounts==0))/sum(truecounts>0)
print("Dropout rate")
print(dropout.rate)
save(counts, geneinfo, cellinfo, truecounts, file=paste("splatter_simulate_data_", i, ".RData", sep=""))
X <- t(counts) ## counts with dropout
Y <- as.integer(substring(cellinfo$Group,6))
Y <- Y-1
h5createFile(paste("splatter_simulate_data_", i, ".h5", sep=""))
h5write(X, paste("splatter_simulate_data_", i, ".h5", sep=""),"X")
h5write(Y, paste("splatter_simulate_data_", i, ".h5", sep=""),"Y")
true.X <- t(truecounts) ## counts without dropout
h5createFile(paste("splatter_simulate_truedata_", i, ".h5", sep=""))
h5write(true.X, paste("splatter_simulate_truedata_", i, ".h5", sep=""),"X")
h5write(Y, paste("splatter_simulate_truedata_", i, ".h5", sep=""),"Y")
#### make tSNE plot for the simulated data
library(Rtsne)
library(ggplot2)
library(repr)
options(repr.plot.width=3, repr.plot.height=3)
X.normalized <- apply(X, 2, function(z) z/sum(z))
X.normalized <- log(X.normalized + 1)
tsne.X <- Rtsne(t(X.normalized), max_iter = 1000)
tsne_plot.X <- data.frame(`x-tsne` = tsne.X$Y[,1], `y-tsne` = tsne.X$Y[,2],
truelabel = Y, check.names = F)
tsne_plot.X$truelabel <- factor(tsne_plot.X$truelabel, levels = c(0:max(Y)))
ggplot(tsne_plot.X) + geom_point(aes(x=`x-tsne`, y=`y-tsne`, color=truelabel), size=0.5) +
labs(color='true label') +
ggtitle("Simulated data") +
theme_bw() +
theme(panel.grid.major = element_blank(), panel.grid.minor = element_blank(),
legend.key = element_rect(fill = 'white', colour = 'white'), legend.position="none",
axis.title.y=element_blank(), axis.title.x=element_blank())
```
# Run scDeepCluster on the simulated data
```
"""
This part implements the scDeepCluster algoritm
"""
from time import time
import numpy as np
from keras.models import Model
import keras.backend as K
from keras.engine.topology import Layer, InputSpec
from keras.layers import Dense, Input, GaussianNoise, Layer, Activation
from keras.models import Model
from keras.optimizers import SGD, Adam
from keras.utils.vis_utils import plot_model
from keras.callbacks import EarlyStopping
from sklearn.cluster import KMeans
from sklearn import metrics
import h5py
import scanpy.api as sc
from layers import ConstantDispersionLayer, SliceLayer, ColWiseMultLayer
from loss import poisson_loss, NB, ZINB
from preprocess import read_dataset, normalize
import tensorflow as tf
from numpy.random import seed
seed(2211)
from tensorflow import set_random_seed
set_random_seed(2211)
MeanAct = lambda x: tf.clip_by_value(K.exp(x), 1e-5, 1e6)
DispAct = lambda x: tf.clip_by_value(tf.nn.softplus(x), 1e-4, 1e4)
def cluster_acc(y_true, y_pred):
"""
Calculate clustering accuracy. Require scikit-learn installed
# Arguments
y: true labels, numpy.array with shape `(n_samples,)`
y_pred: predicted labels, numpy.array with shape `(n_samples,)`
# Return
accuracy, in [0,1]
"""
y_true = y_true.astype(np.int64)
assert y_pred.size == y_true.size
D = max(y_pred.max(), y_true.max()) + 1
w = np.zeros((D, D), dtype=np.int64)
for i in range(y_pred.size):
w[y_pred[i], y_true[i]] += 1
from sklearn.utils.linear_assignment_ import linear_assignment
ind = linear_assignment(w.max() - w)
return sum([w[i, j] for i, j in ind]) * 1.0 / y_pred.size
def autoencoder(dims, noise_sd=0, init='glorot_uniform', act='relu'):
"""
Fully connected auto-encoder model, symmetric.
Arguments:
dims: list of number of units in each layer of encoder. dims[0] is input dim, dims[-1] is units in hidden layer.
The decoder is symmetric with encoder. So number of layers of the auto-encoder is 2*len(dims)-1
act: activation, not applied to Input, Hidden and Output layers
return:
Model of autoencoder
"""
n_stacks = len(dims) - 1
# input
sf_layer = Input(shape=(1,), name='size_factors')
x = Input(shape=(dims[0],), name='counts')
h = x
h = GaussianNoise(noise_sd, name='input_noise')(h)
# internal layers in encoder
for i in range(n_stacks-1):
h = Dense(dims[i + 1], kernel_initializer=init, name='encoder_%d' % i)(h)
h = GaussianNoise(noise_sd, name='noise_%d' % i)(h) # add Gaussian noise
h = Activation(act)(h)
# hidden layer
h = Dense(dims[-1], kernel_initializer=init, name='encoder_hidden')(h) # hidden layer, features are extracted from here
# internal layers in decoder
for i in range(n_stacks-1, 0, -1):
h = Dense(dims[i], activation=act, kernel_initializer=init, name='decoder_%d' % i)(h)
# output
pi = Dense(dims[0], activation='sigmoid', kernel_initializer=init, name='pi')(h)
disp = Dense(dims[0], activation=DispAct, kernel_initializer=init, name='dispersion')(h)
mean = Dense(dims[0], activation=MeanAct, kernel_initializer=init, name='mean')(h)
output = ColWiseMultLayer(name='output')([mean, sf_layer])
output = SliceLayer(0, name='slice')([output, disp, pi])
return Model(inputs=[x, sf_layer], outputs=output)
class ClusteringLayer(Layer):
"""
Clustering layer converts input sample (feature) to soft label, i.e. a vector that represents the probability of the
sample belonging to each cluster. The probability is calculated with student's t-distribution.
# Example
```
model.add(ClusteringLayer(n_clusters=10))
```
# Arguments
n_clusters: number of clusters.
weights: list of Numpy array with shape `(n_clusters, n_features)` witch represents the initial cluster centers.
alpha: parameter in Student's t-distribution. Default to 1.0.
# Input shape
2D tensor with shape: `(n_samples, n_features)`.
# Output shape
2D tensor with shape: `(n_samples, n_clusters)`.
"""
def __init__(self, n_clusters, weights=None, alpha=1.0, **kwargs):
if 'input_shape' not in kwargs and 'input_dim' in kwargs:
kwargs['input_shape'] = (kwargs.pop('input_dim'),)
super(ClusteringLayer, self).__init__(**kwargs)
self.n_clusters = n_clusters
self.alpha = alpha
self.initial_weights = weights
self.input_spec = InputSpec(ndim=2)
def build(self, input_shape):
assert len(input_shape) == 2
input_dim = input_shape[1]
self.input_spec = InputSpec(dtype=K.floatx(), shape=(None, input_dim))
self.clusters = self.add_weight((self.n_clusters, input_dim), initializer='glorot_uniform', name='clusters')
if self.initial_weights is not None:
self.set_weights(self.initial_weights)
del self.initial_weights
self.built = True
def call(self, inputs, **kwargs):
""" student t-distribution, as same as used in t-SNE algorithm.
q_ij = 1/(1+dist(x_i, u_j)^2), then normalize it.
Arguments:
inputs: the variable containing data, shape=(n_samples, n_features)
Return:
q: student's t-distribution, or soft labels for each sample. shape=(n_samples, n_clusters)
"""
q = 1.0 / (1.0 + (K.sum(K.square(K.expand_dims(inputs, axis=1) - self.clusters), axis=2) / self.alpha))
q **= (self.alpha + 1.0) / 2.0
q = K.transpose(K.transpose(q) / K.sum(q, axis=1))
return q
def compute_output_shape(self, input_shape):
assert input_shape and len(input_shape) == 2
return input_shape[0], self.n_clusters
def get_config(self):
config = {'n_clusters': self.n_clusters}
base_config = super(ClusteringLayer, self).get_config()
return dict(list(base_config.items()) + list(config.items()))
class SCDeepCluster(object):
def __init__(self,
dims,
n_clusters=10,
noise_sd=0,
alpha=1.0,
ridge=0,
debug=False):
super(SCDeepCluster, self).__init__()
self.dims = dims
self.input_dim = dims[0]
self.n_stacks = len(self.dims) - 1
self.n_clusters = n_clusters
self.noise_sd = noise_sd
self.alpha = alpha
self.act = 'relu'
self.ridge = ridge
self.debug = debug
self.autoencoder = autoencoder(self.dims, noise_sd=self.noise_sd, act = self.act)
# prepare clean encode model without Gaussian noise
ae_layers = [l for l in self.autoencoder.layers]
hidden = self.autoencoder.input[0]
for i in range(1, len(ae_layers)):
if "noise" in ae_layers[i].name:
next
elif "dropout" in ae_layers[i].name:
next
else:
hidden = ae_layers[i](hidden)
if "encoder_hidden" in ae_layers[i].name: # only get encoder layers
break
self.encoder = Model(inputs=self.autoencoder.input, outputs=hidden)
pi = self.autoencoder.get_layer(name='pi').output
disp = self.autoencoder.get_layer(name='dispersion').output
mean = self.autoencoder.get_layer(name='mean').output
zinb = ZINB(pi, theta=disp, ridge_lambda=self.ridge, debug=self.debug)
self.loss = zinb.loss
clustering_layer = ClusteringLayer(self.n_clusters, alpha=self.alpha, name='clustering')(hidden)
self.model = Model(inputs=[self.autoencoder.input[0], self.autoencoder.input[1]],
outputs=[clustering_layer, self.autoencoder.output])
self.pretrained = False
self.centers = []
self.y_pred = []
def pretrain(self, x, y, batch_size=256, epochs=200, optimizer='adam', ae_file='ae_weights.h5'):
print('...Pretraining autoencoder...')
self.autoencoder.compile(loss=self.loss, optimizer=optimizer)
es = EarlyStopping(monitor="loss", patience=50, verbose=1)
self.autoencoder.fit(x=x, y=y, batch_size=batch_size, epochs=epochs, callbacks=[es])
self.autoencoder.save_weights(ae_file)
print('Pretrained weights are saved to ./' + str(ae_file))
self.pretrained = True
def load_weights(self, weights_path): # load weights of scDeepCluster model
self.model.load_weights(weights_path)
def extract_feature(self, x): # extract features from before clustering layer
return self.encoder.predict(x)
def predict_clusters(self, x): # predict cluster labels using the output of clustering layer
q, _ = self.model.predict(x, verbose=0)
return q.argmax(1)
@staticmethod
def target_distribution(q): # target distribution P which enhances the discrimination of soft label Q
weight = q ** 2 / q.sum(0)
return (weight.T / weight.sum(1)).T
def fit(self, x_counts, sf, y, raw_counts, batch_size=256, maxiter=2e4, tol=1e-3, update_interval=140,
ae_weights=None, save_dir='./results/scDeepCluster', loss_weights=[1,1], optimizer='adadelta'):
self.model.compile(loss=['kld', self.loss], loss_weights=loss_weights, optimizer=optimizer)
print('Update interval', update_interval)
save_interval = int(x_counts.shape[0] / batch_size) * 5 # 5 epochs
print('Save interval', save_interval)
# Step 1: pretrain
if not self.pretrained and ae_weights is None:
print('...pretraining autoencoders using default hyper-parameters:')
print(' optimizer=\'adam\'; epochs=200')
self.pretrain(x, batch_size)
self.pretrained = True
elif ae_weights is not None:
self.autoencoder.load_weights(ae_weights)
print('ae_weights is loaded successfully.')
# Step 2: initialize cluster centers using k-means
print('Initializing cluster centers with k-means.')
kmeans = KMeans(n_clusters=self.n_clusters, n_init=20)
self.y_pred = kmeans.fit_predict(self.encoder.predict([x_counts, sf]))
y_pred_last = np.copy(self.y_pred)
self.model.get_layer(name='clustering').set_weights([kmeans.cluster_centers_])
# Step 3: deep clustering
# logging file
import csv, os
if not os.path.exists(save_dir):
os.makedirs(save_dir)
logfile = open(save_dir + '/scDeepCluster_log.csv', 'w')
logwriter = csv.DictWriter(logfile, fieldnames=['iter', 'acc', 'nmi', 'ari', 'L', 'Lc', 'Lr'])
logwriter.writeheader()
loss = [0, 0, 0]
index = 0
for ite in range(int(maxiter)):
if ite % update_interval == 0:
q, _ = self.model.predict([x_counts, sf], verbose=0)
p = self.target_distribution(q) # update the auxiliary target distribution p
# evaluate the clustering performance
self.y_pred = q.argmax(1)
if y is not None:
acc = np.round(cluster_acc(y, self.y_pred), 5)
nmi = np.round(metrics.normalized_mutual_info_score(y, self.y_pred), 5)
ari = np.round(metrics.adjusted_rand_score(y, self.y_pred), 5)
loss = np.round(loss, 5)
logwriter.writerow(dict(iter=ite, acc=acc, nmi=nmi, ari=ari, L=loss[0], Lc=loss[1], Lr=loss[2]))
print('Iter-%d: ACC= %.4f, NMI= %.4f, ARI= %.4f; L= %.5f, Lc= %.5f, Lr= %.5f'
% (ite, acc, nmi, ari, loss[0], loss[1], loss[2]))
# check stop criterion
delta_label = np.sum(self.y_pred != y_pred_last).astype(np.float32) / self.y_pred.shape[0]
y_pred_last = np.copy(self.y_pred)
if ite > 0 and delta_label < tol:
print('delta_label ', delta_label, '< tol ', tol)
print('Reached tolerance threshold. Stopping training.')
logfile.close()
break
# train on batch
if (index + 1) * batch_size > x_counts.shape[0]:
loss = self.model.train_on_batch(x=[x_counts[index * batch_size::], sf[index * batch_size:]],
y=[p[index * batch_size::], raw_counts[index * batch_size::]])
index = 0
else:
loss = self.model.train_on_batch(x=[x_counts[index * batch_size:(index + 1) * batch_size],
sf[index * batch_size:(index + 1) * batch_size]],
y=[p[index * batch_size:(index + 1) * batch_size],
raw_counts[index * batch_size:(index + 1) * batch_size]])
index += 1
# save intermediate model
if ite % save_interval == 0:
# save scDeepCluster model checkpoints
print('saving model to: ' + save_dir + '/scDeepCluster_model_' + str(ite) + '.h5')
self.model.save_weights(save_dir + '/scDeepCluster_model_' + str(ite) + '.h5')
ite += 1
# save the trained model
logfile.close()
print('saving model to: ' + save_dir + '/scDeepCluster_model_final.h5')
self.model.save_weights(save_dir + '/scDeepCluster_model_final.h5')
return self.y_pred
#### Run scDeepCluster on the simulated data
optimizer1 = Adam(amsgrad=True)
optimizer2 = 'adadelta'
data_mat = h5py.File("splatter_simulate_data_1.h5")
x = np.array(data_mat['X'])
y = np.array(data_mat['Y'])
# preprocessing scRNA-seq read counts matrix
adata = sc.AnnData(x)
adata.obs['Group'] = y
adata = read_dataset(adata,
transpose=False,
test_split=False,
copy=True)
adata = normalize(adata,
size_factors=True,
normalize_input=True,
logtrans_input=True)
input_size = adata.n_vars
print('Sample size')
print(adata.X.shape)
print(y.shape)
x_sd = adata.X.std(0)
x_sd_median = np.median(x_sd)
update_interval = int(adata.X.shape[0]/256)
# Define scDeepCluster model
scDeepCluster = SCDeepCluster(dims=[input_size, 256, 64, 32], n_clusters=3, noise_sd=2.5)
print("autocoder summary")
scDeepCluster.autoencoder.summary()
print("model summary")
scDeepCluster.model.summary()
t0 = time()
# Pretrain autoencoders before clustering
scDeepCluster.pretrain(x=[adata.X, adata.obs.size_factors], y=adata.raw.X, batch_size=256, epochs=600, optimizer=optimizer1, ae_file='ae_weights.h5')
# begin clustering, time not include pretraining part.
gamma = 1. # set hyperparameter gamma
scDeepCluster.fit(x_counts=adata.X, sf=adata.obs.size_factors, y=y, raw_counts=adata.raw.X, batch_size=256, tol=0.001, maxiter=20000,
update_interval=update_interval, ae_weights=None, save_dir='scDeepCluster', loss_weights=[gamma, 1], optimizer=optimizer2)
# Show the final results
y_pred = scDeepCluster.y_pred
acc = np.round(cluster_acc(y, scDeepCluster.y_pred), 5)
nmi = np.round(metrics.normalized_mutual_info_score(y, scDeepCluster.y_pred), 5)
ari = np.round(metrics.adjusted_rand_score(y, scDeepCluster.y_pred), 5)
print('Final: ACC= %.4f, NMI= %.4f, ARI= %.4f' % (acc, nmi, ari))
print('Clustering time: %d seconds.' % int(time() - t0))
```
Clustering result:
Final: ACC= 0.9940, NMI= 0.9640, ARI= 0.9821
# Plot the latent space
```
hidden_layer = scDeepCluster.model.get_layer(name='encoder_hidden').get_output_at(1)
hidden_output_model = Model(inputs = scDeepCluster.model.input, outputs = hidden_layer)
hidden_output = hidden_output_model.predict([adata.X, adata.obs.size_factors])
hidden_output = np.asarray(hidden_output)
np.savetxt('latent_output.csv', hidden_output, delimiter=",")
library(Rtsne)
library(ggplot2)
library(repr)
library(rhdf5)
options(repr.plot.width=3, repr.plot.height=3)
latent.dat <- read.table("latent_output.csv", header = F, sep = ",")
tsne.X <- Rtsne(latent.dat, max_iter = 1000)
Y <- h5read("splatter_simulate_data_1.h5", "Y")
tsne_plot.X <- data.frame(`x-tsne` = tsne.X$Y[,1], `y-tsne` = tsne.X$Y[,2],
truelabel = Y, check.names = F)
tsne_plot.X$truelabel <- factor(tsne_plot.X$truelabel, levels = c(0:max(Y)))
ggplot(tsne_plot.X) + geom_point(aes(x=`x-tsne`, y=`y-tsne`, color=truelabel), size=0.5) +
labs(color='true label') +
ggtitle("scDeepCluster latent space") +
theme_bw() +
theme(panel.grid.major = element_blank(), panel.grid.minor = element_blank(),
legend.key = element_rect(fill = 'white', colour = 'white'), legend.position="none",
axis.title.y=element_blank(), axis.title.x=element_blank())
```
| github_jupyter |
# Prior Sensitivity Analysis
When we don't have strong beliefs about our prior, we should choose uninformative priors. In order to show that the priors don't matter, it is better to verify that the model returns similar results using a variety of priors, that way we know that the model is dependent on the data rather than the prior.
Here we want to justify our assertion that people with bad health tend to make more doctor visits than people with good health.
```
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pymc3 as pm
import scipy.stats as stats
import statsmodels.api as sm
from scipy.special import expit
%matplotlib inline
import warnings
warnings.filterwarnings("ignore")
```
## Load the data
```
try:
badhealth_df = pd.read_csv("badhealth.csv", index=False)
except:
badhealth_df = pd.read_csv("http://vincentarelbundock.github.io/Rdatasets/csv/COUNT/badhealth.csv")
badhealth_df.to_csv("badhealth.csv")
badhealth_df.head()
```
## Model 1: Uninformative Prior
```
x_badh = badhealth_df["badh"].values
x_age = badhealth_df["age"].values
x_intx = (badhealth_df["badh"] * badhealth_df["age"]).values
y = badhealth_df["numvisit"].values
model_1 = pm.Model()
with model_1:
coeffs = pm.Normal("coeffs", mu=0, sigma=1e3, shape=4)
log_preds = (coeffs[0] +
coeffs[1] * x_badh +
coeffs[2] * x_age +
coeffs[3] * x_intx)
y_obs = pm.NegativeBinomial("y_obs", mu=pm.math.exp(log_preds), alpha=2.5, observed=y)
trace_1 = pm.sample(5000, tune=1000)
coeff_badh = trace_1.get_values("coeffs")[:, 1]
_ = pd.Series(coeff_badh).plot(kind="density")
```
Essentially all the probability mass > 0, indicating that the coefficient is positive. We have obtained this result using a relatively non-informative prior.
## Model 2: Skeptical prior
We will now use a prior that favors values of `badh` coefficient close to 0, Normal($\mu$=0, $\sigma$=0.2), so there is 99% prior probability that the coefficient < 0.6.
```
model_2 = pm.Model()
with model_2:
coeffs = pm.Normal("coeffs", mu=0, sigma=1e3, shape=2)
coeffs_2 = pm.Normal("coeffs_2", mu=0, sigma=0.2, shape=2)
log_preds = (coeffs[0] +
coeffs_2[0] * x_badh +
coeffs[1] * x_age +
coeffs_2[1] * x_intx)
y_obs = pm.NegativeBinomial("y_obs", mu=pm.math.exp(log_preds), alpha=2.5, observed=y)
trace_2 = pm.sample(5000, tune=1000)
coeff_badh = trace_2.get_values("coeffs_2")[:, 0]
_ = pd.Series(coeff_badh).plot(kind="density")
```
Under the skeptical prior, our coefficient value reduced, but it is still positive. Thus, even under the skeptical prior, bad health is associated with increased doctor visits.
## Interaction Term
Under the baseline (uninformative prior) model, coefficient of `intx` was negative, but with the skeptical prior model, it is largely positive, so this result is sensitive to choice of prior. This is probably because the skeptical prior shrinks the effect of `badh`, the interaction term tries to restore it. Thus, despite the sensitivity to the prior, our conclusion that bad health is associated with higher number of doctor visits remains unchanged.
```
coeff_intx_1 = trace_1.get_values("coeffs")[:, 3]
coeff_intx_2 = trace_2.get_values("coeffs_2")[:, 1]
intx_df = pd.DataFrame({"intx_1": coeff_intx_1, "intx_2": coeff_intx_2})
_ = intx_df.plot(kind="density")
np.mean(coeff_intx_1 > 0)
np.mean(coeff_intx_2 > 0)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/tensorflow/tpu/blob/master/tools/colab/mnist_estimator.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
##### Copyright 2018 The TensorFlow Hub Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
# Copyright 2018 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
## MNIST with the Estimator API
## Overview
This notebook trains a model to classify images based on the handwritten numbers in the MNIST dataset. After training, the model classifies incoming images into
10 categories (0 to 9) based on what it's learned from the dataset.
This notebook uses Estimator on a GPU backend. It is a reference point for converting an Estimator model to TPUEstimator and a TPU backend. This conversion is demonstrated in the [TPUEstimator](https://colab.research.google.com/github/tensorflow/tpu/blob/master/tools/colab/mnist_tpuestimator.ipynb) notebook. The conversion enables your model to take advantage of Cloud TPU to speed up training computations.
This notebook is hosted on GitHub. To view it in its original repository, after opening the notebook, select **File > View on GitHub**.
## Instructions
<h3><a href="https://cloud.google.com/gpu/"><img valign="middle" src="https://raw.githubusercontent.com/GoogleCloudPlatform/tensorflow-without-a-phd/master/tensorflow-rl-pong/images/gpu-hexagon.png" width="50"></a> Train on GPU</a></h3>
1. Create a Cloud Storage bucket for your TensorBoard logs at http://console.cloud.google.com/storage and fill in the BUCKET parameter in the "Parameters" section below.
1. On the main menu, click Runtime and select **Change runtime type**. Set "GPU" as the hardware accelerator.
1. Click Runtime again and select **Runtime > Run All** (Watch out: the "Colab-only auth" cell requires user input). You can also run the cells manually with Shift-ENTER.
<h3><a href="https://cloud.google.com/ml-engine/"><img valign="middle" src="https://raw.githubusercontent.com/GoogleCloudPlatform/tensorflow-without-a-phd/master/tensorflow-rl-pong/images/mlengine-hexagon.png" width="50"></a> Deploy to Cloud Machine Learning (ML) Engine</h3>
1. At the bottom of this notebook you can deploy your trained model to ML Engine for a serverless, autoscaled, REST API experience. You will need a GCP project name for this last part.
## Data, model, and training
### Imports
```
import os, re, math, json, shutil, pprint, datetime
import PIL.Image, PIL.ImageFont, PIL.ImageDraw
import numpy as np
import tensorflow as tf
from matplotlib import pyplot as plt
from tensorflow.python.platform import tf_logging
print("Tensorflow version " + tf.__version__)
```
### Parameters
```
BATCH_SIZE = 32 #@param {type:"integer"}
BUCKET = 'gs://' #@param {type:"string"}
assert re.search(r'gs://.+', BUCKET), 'You need a GCS bucket for your Tensorboard logs. Head to http://console.cloud.google.com/storage and create one.'
training_images_file = 'gs://mnist-public/train-images-idx3-ubyte'
training_labels_file = 'gs://mnist-public/train-labels-idx1-ubyte'
validation_images_file = 'gs://mnist-public/t10k-images-idx3-ubyte'
validation_labels_file = 'gs://mnist-public/t10k-labels-idx1-ubyte'
```
### Colab-only auth
```
# backend identification
IS_COLAB_BACKEND = 'COLAB_GPU' in os.environ # this is always set on Colab, the value is 0 or 1 depending on GPU presence
# Auth on Colab
# Little wrinkle: without auth, Colab will be extremely slow in accessing data from a GCS bucket, even public
if IS_COLAB_BACKEND:
from google.colab import auth
auth.authenticate_user()
#@title visualization utilities [RUN ME]
"""
This cell contains helper functions used for visualization
and downloads only. You can skip reading it. There is very
little useful Keras/Tensorflow code here.
"""
# Matplotlib config
plt.rc('image', cmap='gray_r')
plt.rc('grid', linewidth=0)
plt.rc('xtick', top=False, bottom=False, labelsize='large')
plt.rc('ytick', left=False, right=False, labelsize='large')
plt.rc('axes', facecolor='F8F8F8', titlesize="large", edgecolor='white')
plt.rc('text', color='a8151a')
plt.rc('figure', facecolor='F0F0F0')# Matplotlib fonts
MATPLOTLIB_FONT_DIR = os.path.join(os.path.dirname(plt.__file__), "mpl-data/fonts/ttf")
# pull a batch from the datasets. This code is not very nice, it gets much better in eager mode (TODO)
def dataset_to_numpy_util(training_dataset, validation_dataset, N):
# get one batch from each: 10000 validation digits, N training digits
unbatched_train_ds = training_dataset.unbatch()
v_images, v_labels = validation_dataset.make_one_shot_iterator().get_next()
t_images, t_labels = unbatched_train_ds.batch(N).make_one_shot_iterator().get_next()
# Run once, get one batch. Session.run returns numpy results
with tf.Session() as ses:
(validation_digits, validation_labels,
training_digits, training_labels) = ses.run([v_images, v_labels, t_images, t_labels])
# these were one-hot encoded in the dataset
validation_labels = np.argmax(validation_labels, axis=1)
training_labels = np.argmax(training_labels, axis=1)
return (training_digits, training_labels,
validation_digits, validation_labels)
# create digits from local fonts for testing
def create_digits_from_local_fonts(n):
font_labels = []
img = PIL.Image.new('LA', (28*n, 28), color = (0,255)) # format 'LA': black in channel 0, alpha in channel 1
font1 = PIL.ImageFont.truetype(os.path.join(MATPLOTLIB_FONT_DIR, 'DejaVuSansMono-Oblique.ttf'), 25)
font2 = PIL.ImageFont.truetype(os.path.join(MATPLOTLIB_FONT_DIR, 'STIXGeneral.ttf'), 25)
d = PIL.ImageDraw.Draw(img)
for i in range(n):
font_labels.append(i%10)
d.text((7+i*28,0 if i<10 else -4), str(i%10), fill=(255,255), font=font1 if i<10 else font2)
font_digits = np.array(img.getdata(), np.float32)[:,0] / 255.0 # black in channel 0, alpha in channel 1 (discarded)
font_digits = np.reshape(np.stack(np.split(np.reshape(font_digits, [28, 28*n]), n, axis=1), axis=0), [n, 28*28])
return font_digits, font_labels
# utility to display a row of digits with their predictions
def display_digits(digits, predictions, labels, title, n):
plt.figure(figsize=(13,3))
digits = np.reshape(digits, [n, 28, 28])
digits = np.swapaxes(digits, 0, 1)
digits = np.reshape(digits, [28, 28*n])
plt.yticks([])
plt.xticks([28*x+14 for x in range(n)], predictions)
for i,t in enumerate(plt.gca().xaxis.get_ticklabels()):
if predictions[i] != labels[i]: t.set_color('red') # bad predictions in red
plt.imshow(digits)
plt.grid(None)
plt.title(title)
# utility to display multiple rows of digits, sorted by unrecognized/recognized status
def display_top_unrecognized(digits, predictions, labels, n, lines):
idx = np.argsort(predictions==labels) # sort order: unrecognized first
for i in range(lines):
display_digits(digits[idx][i*n:(i+1)*n], predictions[idx][i*n:(i+1)*n], labels[idx][i*n:(i+1)*n],
"{} sample validation digits out of {} with bad predictions in red and sorted first".format(n*lines, len(digits)) if i==0 else "", n)
# utility to display training and validation curves
def display_training_curves(training, validation, title, subplot):
if subplot%10==1: # set up the subplots on the first call
plt.subplots(figsize=(10,10), facecolor='#F0F0F0')
plt.tight_layout()
ax = plt.subplot(subplot)
ax.grid(linewidth=1, color='white')
ax.plot(training)
ax.plot(validation)
ax.set_title('model '+ title)
ax.set_ylabel(title)
ax.set_xlabel('epoch')
ax.legend(['train', 'valid.'])
```
### tf.data.Dataset: parse files and prepare training and validation datasets
Please read the [best practices for building](https://www.tensorflow.org/guide/performance/datasets) input pipelines with tf.data.Dataset
```
def read_label(tf_bytestring):
label = tf.decode_raw(tf_bytestring, tf.uint8)
label = tf.reshape(label, [])
label = tf.one_hot(label, 10)
return label
def read_image(tf_bytestring):
image = tf.decode_raw(tf_bytestring, tf.uint8)
image = tf.cast(image, tf.float32)/255.0
image = tf.reshape(image, [28*28])
return image
def load_dataset(image_file, label_file):
imagedataset = tf.data.FixedLengthRecordDataset(image_file, 28*28, header_bytes=16)
imagedataset = imagedataset.map(read_image, num_parallel_calls=16)
labelsdataset = tf.data.FixedLengthRecordDataset(label_file, 1, header_bytes=8)
labelsdataset = labelsdataset.map(read_label, num_parallel_calls=16)
dataset = tf.data.Dataset.zip((imagedataset, labelsdataset))
return dataset
def get_training_dataset(image_file, label_file, batch_size):
dataset = load_dataset(image_file, label_file)
dataset = dataset.cache() # this small dataset can be entirely cached in RAM, for TPU this is important to get good performance from such a small dataset
dataset = dataset.shuffle(5000, reshuffle_each_iteration=True)
dataset = dataset.repeat() # Mandatory for Keras for now
dataset = dataset.batch(batch_size, drop_remainder=True) # drop_remainder is important on TPU, batch size must be fixed
dataset = dataset.prefetch(10) # fetch next batches while training on the current one
return dataset
def get_validation_dataset(image_file, label_file):
dataset = load_dataset(image_file, label_file)
dataset = dataset.cache() # this small dataset can be entirely cached in RAM, for TPU this is important to get good performance from such a small dataset
dataset = dataset.batch(10000, drop_remainder=True) # 10000 items in eval dataset, all in one batch
dataset = dataset.repeat() # Mandatory for Keras for now
return dataset
# instantiate the datasets
training_dataset = get_training_dataset(training_images_file, training_labels_file, BATCH_SIZE)
validation_dataset = get_validation_dataset(validation_images_file, validation_labels_file)
# In Estimator, we will need a function that returns the dataset
training_input_fn = lambda: get_training_dataset(training_images_file, training_labels_file, BATCH_SIZE)
validation_input_fn = lambda: get_validation_dataset(validation_images_file, validation_labels_file)
```
### Let's have a look at the data
```
N = 24
(training_digits, training_labels,
validation_digits, validation_labels) = dataset_to_numpy_util(training_dataset, validation_dataset, N)
display_digits(training_digits, training_labels, training_labels, "training digits and their labels", N)
display_digits(validation_digits[:N], validation_labels[:N], validation_labels[:N], "validation digits and their labels", N)
font_digits, font_labels = create_digits_from_local_fonts(N)
```
### Estimator model
If you are not sure what cross-entropy, dropout, softmax or batch-normalization mean, head here for a crash-course: [Tensorflow and deep learning without a PhD](https://github.com/GoogleCloudPlatform/tensorflow-without-a-phd/#featured-code-sample)
```
# This model trains to 99.4% sometimes 99.5% accuracy in 10 epochs
def model_fn(features, labels, mode):
x = features
is_training = (mode == tf.estimator.ModeKeys.TRAIN)
x = features
y = tf.reshape(x, [-1, 28, 28, 1])
# little wrinkle: tf.keras.layers can normally be used in an Estimator but tf.keras.layers.BatchNormalization does not work
# in an Estimator environment. Using TF layers everywhere for consistency. tf.layers and tf.ketas.layers are carbon copies of each other.
y = tf.layers.Conv2D(filters=6, kernel_size=3, padding='same', use_bias=False)(y) # no bias necessary before batch norm
y = tf.layers.BatchNormalization(scale=False, center=True)(y, training=is_training) # no batch norm scaling necessary before "relu"
y = tf.nn.relu(y) # activation after batch norm
y = tf.layers.Conv2D(filters=12, kernel_size=6, padding='same', use_bias=False, strides=2)(y)
y = tf.layers.BatchNormalization(scale=False, center=True)(y, training=is_training)
y = tf.nn.relu(y)
y = tf.layers.Conv2D(filters=24, kernel_size=6, padding='same', use_bias=False, strides=2)(y)
y = tf.layers.BatchNormalization(scale=False, center=True)(y, training=is_training)
y = tf.nn.relu(y)
y = tf.layers.Flatten()(y)
y = tf.layers.Dense(200, use_bias=False)(y)
y = tf.layers.BatchNormalization(scale=False, center=True)(y, training=is_training)
y = tf.nn.relu(y)
y = tf.layers.Dropout(0.5)(y, training=is_training)
logits = tf.layers.Dense(10)(y)
predictions = tf.nn.softmax(logits)
classes = tf.math.argmax(predictions, axis=-1)
if (mode != tf.estimator.ModeKeys.PREDICT):
loss = tf.losses.softmax_cross_entropy(labels, logits)
step = tf.train.get_or_create_global_step()
lr = 0.0001 + tf.train.exponential_decay(0.01, step, 2000, 1/math.e)
tf.summary.scalar("learn_rate", lr)
optimizer = tf.train.AdamOptimizer(lr)
# little wrinkle: batch norm uses running averages which need updating after each batch. create_train_op does it, optimizer.minimize does not.
train_op = tf.contrib.training.create_train_op(loss, optimizer)
#train_op = optimizer.minimize(loss, tf.train.get_or_create_global_step())
metrics = {'accuracy': tf.metrics.accuracy(classes, tf.math.argmax(labels, axis=-1))}
else:
loss = train_op = metrics = None # None of these can be computed in prediction mode because labels are not available
return tf.estimator.EstimatorSpec(
mode=mode,
predictions={"predictions": predictions, "classes": classes}, # name these fields as you like
loss=loss,
train_op=train_op,
eval_metric_ops=metrics
)
# Called once when the model is saved. This function produces a Tensorflow
# graph of operations that will be prepended to your model graph. When
# your model is deployed as a REST API, the API receives data in JSON format,
# parses it into Tensors, then sends the tensors to the input graph generated by
# this function. The graph can transform the data so it can be sent into your
# model input_fn. You can do anything you want here as long as you do it with
# tf.* functions that produce a graph of operations.
def serving_input_fn():
# placeholder for the data received by the API (already parsed, no JSON decoding necessary,
# but the JSON must contain one or multiple 'image' key(s) with 28x28 greyscale images as content.)
inputs = {"serving_input": tf.placeholder(tf.float32, [None, 28, 28])} # the shape of this dict should match the shape of your JSON
features = inputs['serving_input'] # no transformation needed
return tf.estimator.export.TensorServingInputReceiver(features, inputs) # features are the features needed by your model_fn
# Return a ServingInputReceiver if your features are a dictionary of Tensors, TensorServingInputReceiver if they are a straight Tensor
```
### Train and validate the model
```
EPOCHS = 8
steps_per_epoch = 60000 // BATCH_SIZE # 60,000 images in training dataset
MODEL_EXPORT_NAME = "mnist" # name for exporting saved model
tf_logging.set_verbosity(tf_logging.INFO)
now = datetime.datetime.now()
MODEL_DIR = BUCKET+"/mnistjobs/job" + "-{}-{:02d}-{:02d}-{:02d}:{:02d}:{:02d}".format(now.year, now.month, now.day, now.hour, now.minute, now.second)
training_config = tf.estimator.RunConfig(model_dir=MODEL_DIR, save_summary_steps=10, save_checkpoints_steps=steps_per_epoch, log_step_count_steps=steps_per_epoch/4)
export_latest = tf.estimator.LatestExporter(MODEL_EXPORT_NAME, serving_input_receiver_fn=serving_input_fn)
estimator = tf.estimator.Estimator(model_fn=model_fn, config=training_config)
train_spec = tf.estimator.TrainSpec(training_input_fn, max_steps=EPOCHS*steps_per_epoch)
eval_spec = tf.estimator.EvalSpec(validation_input_fn, steps=1, exporters=export_latest, throttle_secs=0) # no eval throttling: evaluates after each checkpoint
tf.estimator.train_and_evaluate(estimator, train_spec, eval_spec)
tf_logging.set_verbosity(tf_logging.WARN)
```
### Visualize predictions
```
# recognize digits from local fonts
predictions = estimator.predict(lambda: tf.data.Dataset.from_tensor_slices(font_digits).batch(N),
yield_single_examples=False) # the returned value is a generator that will yield one batch of predictions per next() call
predicted_font_classes = next(predictions)['classes']
display_digits(font_digits, predicted_font_classes, font_labels, "predictions from local fonts (bad predictions in red)", N)
# recognize validation digits
predictions = estimator.predict(validation_input_fn,
yield_single_examples=False) # the returned value is a generator that will yield one batch of predictions per next() call
predicted_labels = next(predictions)['classes']
display_top_unrecognized(validation_digits, predicted_labels, validation_labels, N, 7)
```
## Deploy the trained model to ML Engine
Push your trained model to production on ML Engine for a serverless, autoscaled, REST API experience.
You need the name of your GCS bucket and GCP project for this step. Models deployed on ML Engine autoscale to zero if not used. There will be no ML Engine charges after you are done testing.
Google Cloud Storage incurs charges. Empty the bucket after deployment if you want to avoid these. Once the model is deployed, the bucket is not useful anymore.
### Configuration
```
PROJECT = "" #@param {type:"string"}
NEW_MODEL = True #@param {type:"boolean"}
MODEL_NAME = "estimator_mnist" #@param {type:"string"}
MODEL_VERSION = "v0" #@param {type:"string"}
assert PROJECT, 'For this part, you need a GCP project. Head to http://console.cloud.google.com/ and create one.'
export_path = os.path.join(MODEL_DIR, 'export', MODEL_EXPORT_NAME)
last_export = sorted(tf.gfile.ListDirectory(export_path))[-1]
export_path = os.path.join(export_path, last_export)
print('Saved model directory found: ', export_path)
```
### Deploy the model
This uses the command-line interface. You can do the same thing through the ML Engine UI at https://console.cloud.google.com/mlengine/models
```
# Create the model
if NEW_MODEL:
!gcloud ml-engine models create {MODEL_NAME} --project={PROJECT} --regions=us-central1
# Create a version of this model (you can add --async at the end of the line to make this call non blocking)
# Additional config flags are available: https://cloud.google.com/ml-engine/reference/rest/v1/projects.models.versions
# You can also deploy a model that is stored locally by providing a --staging-bucket=... parameter
!echo "Deployment takes a couple of minutes. You can watch your deployment here: https://console.cloud.google.com/mlengine/models/{MODEL_NAME}"
!gcloud ml-engine versions create {MODEL_VERSION} --model={MODEL_NAME} --origin={export_path} --project={PROJECT} --runtime-version=1.10
```
### Test the deployed model
Your model is now available as a REST API. Let us try to call it. The cells below use the "gcloud ml-engine"
command line tool but any tool that can send a JSON payload to a REST endpoint will work.
```
# prepare digits to send to online prediction endpoint
digits = np.concatenate((font_digits, validation_digits[:100-N]))
labels = np.concatenate((font_labels, validation_labels[:100-N]))
with open("digits.json", "w") as f:
for digit in digits:
# the format for ML Engine online predictions is: one JSON object per line
data = json.dumps({"serving_input": digit.tolist()}) # "serving_input" because that is what you defined in your serving_input_fn: {"serving_input": tf.placeholder(tf.float32, [None, 28, 28])}
f.write(data+'\n')
# Request online predictions from deployed model (REST API) using the "gcloud ml-engine" command line.
predictions = !gcloud ml-engine predict --model={MODEL_NAME} --json-instances digits.json --project={PROJECT} --version {MODEL_VERSION}
predictions = np.array([int(p.split('[')[0]) for p in predictions[1:]]) # first line is the name of the input layer: drop it, parse the rest
display_top_unrecognized(digits, predictions, labels, N, 100//N)
```
## What's next
* Learn how to convert an [estimator model to a TPUEstimator model](https://colab.research.google.com/github/tensorflow/tpu/blob/master/tools/colab/mnist_tpuestimator.ipynb) to take advantage of [Cloud TPUs](https://cloud.google.com/tpu/docs).
* Explore the range of [Cloud TPU tutorials and Colabs](https://cloud.google.com/tpu/docs/tutorials) to find other examples that can be used when implementing your ML project.
On Google Cloud Platform, in addition to GPUs and TPUs available on pre-configured [deep learning VMs](https://cloud.google.com/deep-learning-vm/), you will find [AutoML](https://cloud.google.com/automl/)*(beta)* for training custom models without writing code and [Cloud ML Engine](https://cloud.google.com/ml-engine/docs/) which will allows you to run parallel trainings and hyperparameter tuning of your custom models on powerful distributed hardware.
## License
---
author: Martin Gorner<br>
twitter: @martin_gorner
---
Copyright 2018 Google LLC
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
---
This is not an official Google product but sample code provided for an educational purpose
| github_jupyter |
## These are the follow-up code samples for the following blog post: [Pandas DataFrame by Example](http://queirozf.com/entries/pandas-dataframe-by-example)
```
import pandas as pd
import numpy as np
pd.__version__
df = pd.DataFrame({
'name':['john','mary','peter','jeff','bill','lisa'],
'age':[23,78,22,19,45,33],
'state':['iowa','dc','california','texas','washington','dc'],
'num_children':[2,2,0,1,2,1],
'num_pets':[0,4,0,5,0,0]
})
# sorting columns
df=df[['name','age','state','num_children','num_pets']]
df
```
## Select rows by position
```
# select the first 2 rows
df.iloc[:2]
# select the last 2 rows
df.iloc[-2:]
```
## Select rows by index value
> compare this with `iloc` above
```
# select rows up to and including the one
# with index=2
df.loc[:2]
```
## Select rows based upon the value of columns
```
# by a simple numeric condition
df[df["age"] > 30]
# comparing the value of two columns
df[ df["num_pets"] > df[ "num_children"] ]
# using boolean AND
df[ (df["age"] > 40) & (df["num_pets"] > 0) ]
```
## select columns starting with
```
df[[colname for colname in df.columns if colname.startswith('n')]]
```
## select all columns but one
```
df.drop(['age'],axis=1)
```
## Drop a column
```
df.drop(["age","num_children"],axis=1)
```
## Apply a function to every column (as aggregates)
> Using **numpy** vectorized functions for numerical values
```
# get the mean for each of the selected columns
df[["age","num_pets","num_children"]].apply(lambda row: np.mean(row),axis=0).to_frame()
```
## Apply a function to every row (as aggregates)
> Using **numpy** vectorized functions for numerical values
```
# sum columns age, num_pets and num_children for each row
df[["age","num_pets","num_children"]].apply(lambda row: np.sum(row),axis=1).to_frame()
```
## Apply a function elementwise using apply
```
df[["age"]].apply(lambda value: value*2)
# certain numerical functions can also be used:
df[["age"]] * 2
# also works for string values
df[["name"]].apply(lambda value: value.str.upper())
```
## Apply a function elementwise using map
> use `apply` for DataFrame objects and `map` for Series objects
```
df['name'].map(lambda name: name.upper()).to_frame()
```
## Add new columns based on old ones
```
# simple sum of two columns
df["pets_and_children"] = df["num_pets"] + df["num_children"]
df
df = pd.DataFrame({
'name':['john','mary','peter','jeff','bill','lisa'],
'age':[23,78,22,19,45,33],
'state':['iowa','dc','california','texas','washington','dc'],
'num_children':[2,2,0,1,2,1],
'num_pets':[0,4,0,5,0,0]
})
# sorting columns
df=df[['name','age','state','num_children','num_pets']]
df
# you can also use custom functions we used on "elementwise application"
df["name_uppercase"] = df[["name"]].apply(lambda name: name.str.upper())
df
```
## Shuffle rows
```
df = pd.DataFrame({
'name':['john','mary','peter','jeff','bill','lisa'],
'age':[23,78,22,19,45,33],
'state':['iowa','dc','california','texas','washington','dc'],
'num_children':[2,2,0,1,2,1],
'num_pets':[0,4,0,5,0,0]
})
# sorting columns
df=df[['name','age','state','num_children','num_pets']]
df
df.reindex(np.random.permutation(df.index))
```
## Iterate over all rows
```
for index,row in df.iterrows():
print("{0} has name: {1}".format(index,row["name"]))
```
## Randomly sample rows
```
# sample 10 rows from df
random_indices = np.random.choice(df.index.values, 4, replace=False)
# iloc allows you to retrieve rows by their numeric indices
sampled_df = df.iloc[random_indices]
sampled_df
```
## Sort a dataframe
```
# sort by age, largest first
df.sort_values("age",ascending=False )
# sort by num_pets descending then sort by age ascending
df.sort_values( ["num_pets","age"], ascending=[False,True] )
```
## custom sort
```
df = pd.DataFrame({
'name':['john','mary','peter','jeff','bill','lisa'],
'age':[23,78,22,19,12,33],
'state':['N/A','dc','california','texas','N/A','dc']
})
# sorting columns
df=df[['name','age','state']]
df
def state_to_rank(state):
if state=="N/A":
return 1
else:
return 0
df['rank'] = df['state'].map(lambda x: state_to_rank(x))
df.sort_values(by=['rank','age']).drop(['rank'],axis=1).reset_index(drop=True)
```
## Perform complex selections using lambdas
```
df[df.apply(lambda row: row['name'].startswith('j'),axis=1)]
```
## Change column names
```
# use inplace=True if you want to mutate the current dataframe
df.rename(columns={"age":"age_years"} )
```
## Change column dtype
```
df['num_children'].dtype
# we don't need 64 bits for num_children
df['num_children'] = df['num_children'].astype('int32')
df['num_children'].dtype
# it looks the same but takes less space
df
```
## Veryfing that the dataframe includes specific values
This is done using the `.isin()` method, which returns a **boolean** dataframe to indicate where the passed values are.
```
df = pd.DataFrame({
'name':['john','mary','peter','jeff','bill','lisa'],
'age':[23,78,22,19,45,33],
'state':['iowa','dc','california','texas','washington','dc'],
'num_children':[2,2,0,1,2,1],
'num_pets':[0,4,0,5,0,0]
})
# sorting columns
df=df[['name','age','state','num_children','num_pets']]
# if the method is passed a simple list, it matches
# those values anywhere in the dataframe
df.isin([2,4])
# you can also pass a dict or another dataframe
# as argument
df.isin({'num_pets':[4,5]})
```
## Create an empty Dataframe and append rows one by one
```
# set column names and dtypes
new_df = pd.DataFrame(columns=['col_a','col_b']).astype({'col_a':'float32', 'col_b':'int8'})
# must reassign since the append method does not work in place
new_df = new_df.append({'col_a':5,'col_b':10}, ignore_index=True)
new_df = new_df.append({'col_a':1,'col_b':100}, ignore_index=True)
new_df
```
## Create from list of dicts
```
new_df = pd.DataFrame(columns=['id','name'])
data_dict = [
{'id':1,'name':"john"},
{'id':2,'name':"mary"},
{'id':3,'name':"peter"}
]
# must reassign since the append method does not work in place
new_df = new_df.from_records(data_dict)
new_df
```
## converting types
```
df = pd.DataFrame({
'name':['john','mary','peter'],
"date_of_birth": ['27/05/2002','10/10/1999','01/04/1985']
})
df
df['date_of_birth']=pd.to_datetime(df['date_of_birth'],format='%d/%m/%Y')
df
df.dtypes
```
| github_jupyter |
# Perceptron with Pytorch
#### Luca Laringe
In this notebook, I will show a basic implementation of a perceptron using Pytorch.
A perceptron is a single layer neural network, it is maily used for classification purposes. More info at https://en.wikipedia.org/wiki/Perceptron.
## Data
In this section I am going to generate the data we are going to fit my model on. Let's first import the libraries we need.
```
import torch # Our man library
import torch.nn as nn # To define a neural network
from torch.utils.data import Dataset, DataLoader # To put the data in a class that pytorch knows hoe to handle
from torch.distributions import multivariate_normal # To generate random data
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
plt.rc('figure', figsize=(14, 8))
plt.rc('grid', color='gray', alpha = 0.3, linestyle='solid')
from IPython.core.debugger import set_trace # Debugging tool
```
Let's now generate some data coming from two clusters (I call them $cluster_0$ and $cluser_1$). Each cluster is going to be distributed as a bivariate normal with its own mean vector and covariance matrix.
```
n_sample_0 = 100
n_sample_1 = 100
n_sample = n_sample_0 + n_sample_1
mean_0 = torch.Tensor([0.3,0.7]) # Mean vector of cluster 0
mean_1 = torch.Tensor([0.7,0.3]) # Mean vector of cluster 1
cov_0 = 0.005*torch.eye(2) # Covariance matrix of cluster 0
cov_1 = 0.005*torch.eye(2) # Covariance matrix of cluster 1
cluster_0_dist = multivariate_normal.MultivariateNormal(loc=mean_0, covariance_matrix=cov_0)
cluster_1_dist = multivariate_normal.MultivariateNormal(loc=mean_1, covariance_matrix=cov_1)
data_cluster_0 = cluster_0_dist.sample((n_sample_0,)) # Sample the features
data_cluster_0 = torch.cat((data_cluster_0, torch.zeros(n_sample_0,1)),1)# Generate the labels
data_cluster_1 = cluster_1_dist.sample((n_sample_1,)) # Sample the features
data_cluster_1 = torch.cat((data_cluster_1, torch.ones(n_sample_1,1)),1) # Generate the labels
# Aggregate the data by rows, shuffle them and then divide them into training and testing
data = torch.cat((data_cluster_0, data_cluster_1),0)
data = data[torch.randperm(n_sample).numpy(),:]
data_train = data[:int(n_sample*0.7)]
data_test = data[int(n_sample*0.7):]
# Check everything worked out as expected
print(len(data) == len(data_train) + len(data_test), len(data))
print(data[:10])
# Visualizing the data
plt.scatter(data_cluster_0[:,0], data_cluster_0[:,1]);
plt.scatter(data_cluster_1[:,0], data_cluster_1[:,1]);
plt.legend(['Cluster 0', 'Cluster 1']);
```
Now that we have our data, let's create a dataset and a dataloader so we can use batches to train the model.
First we will create the my_dataset class that inherits from the Dataset class in Pytorch. We need to do it in order to use the Dataloader later. In every dataset class you build, you shoulf always specify the __init__ method recalling that your class should call the __init__ method of the superclass, the __len__ method which returns the length of the dataset, and finally the __getitem__ method , that takes an index and returns the features and labels of the dataset at that index. Note that we are not going to use the transform function here because our data is already tidy and organized in a Pytorch tensor. Nevertheless, when your data is stored in another data structure, for example a pandas DataFrame, the transform function should be viewed as a bridge between your data structure and a Pytorch tensor.
Finally, the Dataloader function will allow us to divide the training set in batches, so that our training will be more effective. In partucular, for each epoch of training, I will divide my training set in 5 batches.
```
class my_dataset(Dataset):
def __init__(self, data, transform = None):
super().__init__()
self.data = data
self.transform = transform
def __len__(self):
return len(self.data)
def __getitem__(self, index):
features = data[index,:2]
labels = data[index, 2]
if self.transform:
features = self.transform(features)
labels = self.transform(labels)
return (features.reshape(2), labels.reshape(1))
# Define instances of my_dataset
my_data_train = my_dataset(data_train)
my_data_test = my_dataset(data_test)
# Define the data loader for the traning data
n_batches = 5
train_loader = DataLoader(batch_size=int(len(my_data_train)/n_batches),
dataset=my_data_train, shuffle=True)
# Check everything works as expected
print(my_data_train.__getitem__(0)[0])
print(my_data_train.__getitem__(0)[1])
```
## Modelling
Let's now define a perceptron.
Note that in Pytorch, every neural network you design, should inherit from the superclass nn.Module. Note also that I will built my perceptron to have a continuous output. To do so I will use the logistic function as the activation function of the output neuron. Doing so, will allow me to interpret the output of the perceptron as the probability of the data point to belong to cluster 1. If you would like a binary output, in general will have to use the unit step activation function.
```
class perceptron(nn.Module):
def __init__(self, n_features):
super().__init__()
self.n_features = n_features
self.fc1 = nn.Linear(n_features, 1)
def forward(self, x):
out = self.fc1(x)
out = torch.sigmoid(out)
return(out)
```
Now that we defined our perceptron class, we need to optimize the network. In order to do so, we will need 3 things (you will always need these regardless of the neural network model you are using):
* An *instance* of the model: in our case, we are going to call the perceptron class and initialize it to have 2 features (like our dataset).
* A loss function to optimize: I will call it criterion and I will use the MSE loss. I am not using the perceptron loss function because I have a continuous output.
* An optimizer: In this case, I will use stochastic gradient descent.
```
# Define an instance of such network
my_perceptron = perceptron(2)
# Define a loss
criterion = nn.MSELoss()
# Define an optimizer
optimizer = optimizer = torch.optim.SGD(my_perceptron.parameters(), lr=0.01)
# Check everything works as expected
print(my_perceptron(torch.Tensor([0.3,0.7])))
print(my_perceptron(my_data_train.__getitem__(0)[0]))
print(criterion(torch.zeros(1), torch.zeros(1)))
print(criterion(torch.zeros(1), torch.ones(1)))
print(my_data_train.__getitem__(0)[1].shape)
print(my_perceptron(my_data_train.__getitem__(0)[0]).shape)
my_data_train.__getitem__(0)[0]
my_data_train.__getitem__(0)[1]
```
## Training
```
n_epochs = 1000
for epoch in range(1,n_epochs+1):
loss_training = 0
for i, (x, y) in enumerate(train_loader):
my_perceptron.train() # Set the model in training mode
optimizer.zero_grad() # Set the grazient to 0
y_ = my_perceptron(x) # Compute the predicted values
loss = criterion(y_, y) # Compute the loss
loss.backward() # Computes the gradient of current tensor w.r.t. graph leaves
optimizer.step() # Updates the parameters
loss_training += loss.data.numpy()/n_batches # Averages up loss for each batch of training
if ((epoch == 1) or (epoch%(n_epochs//5) == 0)):
my_perceptron.eval() # Set the model in evaluation mode
test_features = data_test[:,:2]
test_labels = data_test[:,2].reshape(len(data_test),1)
test_predictions = my_perceptron(test_features)
loss_test = criterion(test_predictions, test_labels) # Computing (average) loss on test set
print('Epoch n. %d, Training Loss: %.4f, Test Loss: %.4f'
%(epoch, loss_training, loss_test.data.numpy()))
```
## Model Evaluation
Let's evaluate the performance of our model computing the accuracy of the predictions on the test set and visializing the decision boundary the perceptron learned.
```
# Now let's compute the predictions and plot them
y_ = my_perceptron(data_test[:,:2]) > 0.5 # Assigns one if the value is greater than 0.5
data_pred = torch.cat((data_test[:,:2], y_.float()),1)
predicted_well = (y_.reshape(len(data_test)).float() == data_test[:,2]).sum().numpy()
test_size = len(data_test)
accuracy = predicted_well/test_size
data_pred_0 = data_pred[data_pred[:,2] == 0]
data_pred_1 = data_pred[data_pred[:,2] == 1]
print('\n\nAccuracy on Test set: ', accuracy*100, '%\n')
```
Let's now try to visualize the decision boundary. To do so I will bo through some basic math.
Our perceptron takes two inputs: $x_1$ and $x_2$ and outputs a value between 0 and 1. If such value is greater than 0.5, we predict 1, else we predict 0.
The output value is given by:
$ output = sigmoid ( bias + weight_1 \times x_1 + weight_2 \times x_2)$
The boundary satisfies this condition:
$ sigmoid(bias + weight_1 \times x_1 + weight_2 \times x_2) = 0.5 $
Hence, we can solve for $x_2$ and find the equation of the line:
$ x_2 = \frac{sigmoid^{-1}(0.5)-bias}{weight_2} - \frac{weight_1}{weight_2} \times x_1 $
Also, note that the sigmoid function is:
$sigmoid(z) = \frac{1}{1+e^{-z}}$
And thus its inverse is:
$sigmoid^{-1}(x) = -\log(\frac{1-x}{x}) = \log(\frac{x}{1-x})$
Now we are ready to plot!
```
def sigmoid_inv(x):
return np.log(x/(1-x))
bias = my_perceptron.fc1.bias.detach().numpy()[0]
weight_1 = my_perceptron.fc1.weight.detach().numpy()[0][0]
weight_2 = my_perceptron.fc1.weight.detach().numpy()[0][1]
```
Let's first visualize the decision boundary and the test set.
```
xs = np.linspace(torch.min(data_test[:,0]), torch.max(data_test[:,0]),1000)
ys = (sigmoid_inv(0.5)-bias)/weight_2 - (weight_1/weight_2)*xs
data_test_0 = data_test[data_test[:,2] == 0]
data_test_1 = data_test[data_test[:,2] == 1]
plt.scatter(data_test_0[:,0], data_test_0[:,1]);
plt.scatter(data_test_1[:,0], data_test_1[:,1]);
plt.legend(['Cluster 0', 'Cluster 1']);
plt.plot(xs,ys);
```
Let's now look at the same chart for all our data.
```
plt.scatter(data_cluster_0[:,0], data_cluster_0[:,1]);
plt.scatter(data_cluster_1[:,0], data_cluster_1[:,1]);
plt.legend(['Cluster 0', 'Cluster 1']);
plt.plot(xs,ys);
```
| github_jupyter |
# Table of Contents
<p><div class="lev1 toc-item"><a href="#Setup" data-toc-modified-id="Setup-1"><span class="toc-item-num">1 </span>Setup</a></div><div class="lev1 toc-item"><a href="#Semantics" data-toc-modified-id="Semantics-2"><span class="toc-item-num">2 </span>Semantics</a></div><div class="lev2 toc-item"><a href="#Motivation" data-toc-modified-id="Motivation-21"><span class="toc-item-num">2.1 </span>Motivation</a></div><div class="lev2 toc-item"><a href="#Term-Document" data-toc-modified-id="Term-Document-22"><span class="toc-item-num">2.2 </span>Term-Document</a></div><div class="lev3 toc-item"><a href="#Bag-of-Words" data-toc-modified-id="Bag-of-Words-221"><span class="toc-item-num">2.2.1 </span>Bag-of-Words</a></div><div class="lev3 toc-item"><a href="#TF-IDF" data-toc-modified-id="TF-IDF-222"><span class="toc-item-num">2.2.2 </span>TF-IDF</a></div><div class="lev2 toc-item"><a href="#Term-Context" data-toc-modified-id="Term-Context-23"><span class="toc-item-num">2.3 </span>Term-Context</a></div><div class="lev1 toc-item"><a href="#Word2Vec" data-toc-modified-id="Word2Vec-3"><span class="toc-item-num">3 </span>Word2Vec</a></div><div class="lev2 toc-item"><a href="#Continuous-Bag-of-Words" data-toc-modified-id="Continuous-Bag-of-Words-31"><span class="toc-item-num">3.1 </span>Continuous Bag of Words</a></div><div class="lev2 toc-item"><a href="#Skip-gram" data-toc-modified-id="Skip-gram-32"><span class="toc-item-num">3.2 </span>Skip-gram</a></div><div class="lev2 toc-item"><a href="#Example" data-toc-modified-id="Example-33"><span class="toc-item-num">3.3 </span>Example</a></div><div class="lev1 toc-item"><a href="#Doc2Vec" data-toc-modified-id="Doc2Vec-4"><span class="toc-item-num">4 </span>Doc2Vec</a></div><div class="lev2 toc-item"><a href="#Doc2Vec,-the-most-powerful-extension-of-word2vec" data-toc-modified-id="Doc2Vec,-the-most-powerful-extension-of-word2vec-41"><span class="toc-item-num">4.1 </span>Doc2Vec, the most powerful extension of word2vec</a></div><div class="lev2 toc-item"><a href="#Distrubted-Memory-(DM)" data-toc-modified-id="Distrubted-Memory-(DM)-42"><span class="toc-item-num">4.2 </span>Distrubted Memory (DM)</a></div><div class="lev2 toc-item"><a href="#Distrubted-Bag-of-Words-(DBOW)" data-toc-modified-id="Distrubted-Bag-of-Words-(DBOW)-43"><span class="toc-item-num">4.3 </span>Distrubted Bag of Words (DBOW)</a></div><div class="lev1 toc-item"><a href="#Exercises" data-toc-modified-id="Exercises-5"><span class="toc-item-num">5 </span>Exercises</a></div>
# Setup
----
This notebook assumes you have done the setup required in Week 1.
In this lecture we will be using Gensim and NLTK, two widely used Python Natural Language Processing libraries.
```
reset -f -s
def pip_install(*packages):
"""
Install packages using pip
Alternatively just use command line
pip install package_name
"""
try:
import pip
for package in packages:
pip.main(["install", "--upgrade", package, "--user"])
except Exception as e:
print("Unable to install {} using pip.".format(package))
print("Exception:", e)
pip_install('gensim', 'nltk')
import nltk
nltk.download('gutenberg')
nltk.download('reuters')
import os
ROOTDIR = os.path.abspath(os.path.dirname('__file__'))
DATADIR = os.path.join(ROOTDIR, 'data')
```
# Semantics
---
## Motivation
If we want to be able to categorize text, we need to be able to generate features for articles, paragraphs, sentences and other bodies of text, based on the information they contain and what they represent. There are a number of ways to achieve this and we will go over 3 approaches.
## Term-Document
### Bag-of-Words
One of the simplest ways to extract features from text is to just count how many times a word appears in a body of text. In this model, the order of words does not matter and only the number of occurrences of each unique term for each document is taken into account.
```
import pandas as pd
#Load movie reviews dataset
df = pd.read_csv(os.path.join(DATADIR, 'movie_reviews.csv'), nrows=100000)
texts = df.text.values #pd.Series -> np.ndarray
import nltk
# Transform each review string as a list of token strings. May take a few seconds
tokenized = [nltk.word_tokenize(review) for review in texts]
n = 10 #arbitrary pick
print('Example review:\n Raw: {} \n\n Tokenized: {}'.format(texts[n], [i for i in tokenized[n]]))
def clean_text(tokenized_list):
import string
sw = nltk.corpus.stopwords.words('english')
new_list = [[token.lower() for token in tlist if token not in string.punctuation and token.lower() not in sw] for tlist in tokenized_list]
return new_list
# Remove punctuations and stopwords
cleaned = clean_text(tokenized)
from gensim import corpora
# Create a dictionary from list of documents
dictionary = corpora.Dictionary(cleaned)
# Create a Corpus based on BOW Format.
corpus = [dictionary.doc2bow(text) for text in cleaned]
print('Example review featurized in Bag of Words :\n {}'.format([(dictionary[i[0]], i[1]) for i in corpus[n]]))
```
Note that when we use this model to featurize text:
- The length of each feature vector will be the size of the vocabulary in the corpus
- Thus each body of text will have a lot of zeroes
### TF-IDF
__Term Frequency__: Number of occurrences of a word in a document
__Inverse Document Frequency__: Number of documents that contain a certain word scaled by a weight
__Term Frequency - Inverse Document Frequency__: (Number of ocurrences of word $w$ in text $T$) * $log$(Number of documents in a corpus/Number of documents containing word $w$)
Let's check out the TF-IDF scores of the previous movie review we examined.
```
from gensim import corpora, models
#Create a TFIDF Model for the corpus
tfidf = models.TfidfModel(corpus)
print('Example review featurized with TF-IDF scores : \n{}'.format([(dictionary[i[0]], round(i[1],3)) for i in tfidf[corpus[n]]]))
```
Looks much more like a feature vector that we can use for text categorization!
Note that in the TF-IDF model:
- If a term frequently occurs in the corpus(i.e. stopwords, the term $like$), it is scaled to a lower score
- Rarer terms will generally have higher scores. They tend to be more "informative" and descriptive.
- A term that occurs frequently in a small number of documents within the corpus will have the highest scores.

## Term-Context
The vast majority of NLP works regards as atomic symbols: king, queen, book, etc.
In vector space terms, this vector has one $1 $ and a lot of zeros.
$king = [1, 0, 0, 0, 0, 0, 0, 0, 0]$
$queen = [0, 1, 0, 0, 0, 0, 0, 0, 0]$
$book = [0, 0, 1, 0, 0, 0, 0, 0, 0]$
It is called a "one-hot" encoding representation. It is a common way to represent categories in models. However, it is very sparse(as we saw from the BOW model); each row is mostly 0s.
You can get more value by representing a word by its neighorbors
Instead of using entire documents, we can use small contexts to a term.
- Paragraphs
- Sentences
- A window of a sequence of consecutive terms
In this way, a word is defined over counts of context words. The assumption is that two words that appear in similar contexts are similar themselves.
But count-based models have disadvantages:
- vector sizes become huge, equal to vocabulary size
- sparsity
- curse of dimensionality
- computationally expensive
# Word2Vec
Word2Vec is an unsupervised neural network model that maximizes similarity between contextual neighbors while minimizing similarity for unseen contexts.
Initial vectors are generated randomly and converge as the models is trained on the corpus through a sliding window.
Target Vector sizes are set at the beginning of the training process, so the vectors are dense and do not need dimensionality reduction techniques.
## Continuous Bag of Words

Training objective is to maiximize the probability of observing the correct target word $w_t$ given context words $w_{c1}, w_{c2}, ... w_{cj}$
$$ C = -log p(w_t | w_{c1} ... w_{cj}) $$
The prediction vector is set as an average of all the context word vectors
## Skip-gram

Training objective is to maiximize the probability of observing the correct context words $w_{ci}$ given target word $w_{t}$
$$ C = -\sum^{j}_{i=1}log p(w_{ci} | w_{t}) $$
In this case, the prediction vector is the the target word vector.
## Example
Now let's try training our own word embeddings and looking at what we can do with them.
Word2Vec
- `size`: Number of dimensions for the word embedding model
- `window`: Number of context words to observe in each direction
- `min_count`: Minimum frequency for words included in model
- `sg` (Skip-Gram): '0' indicates CBOW model; '1' indicates Skip-Gram
- `alpha`: Learning rate (initial); prevents model from over-correcting, enables finer tuning
- `iterations`: Number of passes through dataset
- `batch_words`: Number of words to sample from data during each pass
```
from nltk.corpus import gutenberg
from gensim import models
# Training word2vec model on Gutenberg corpus. This may take a few minutes.
model = models.Word2Vec(gutenberg.sents(), size = 300, window = 5, min_count =5, sg = 0, alpha = 0.025, iter=10, batch_words = 10000)
```
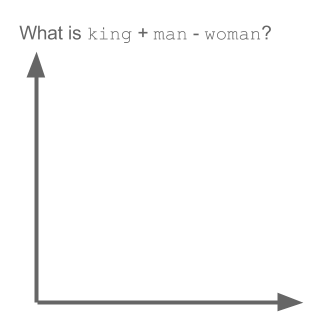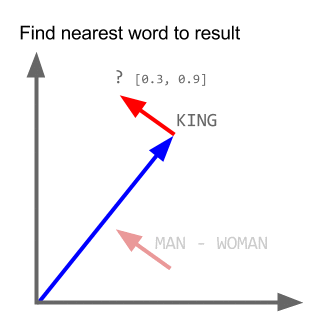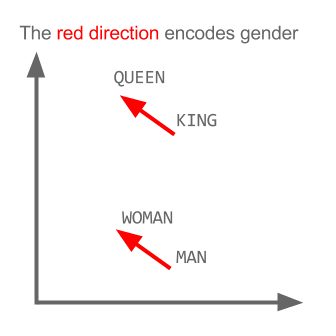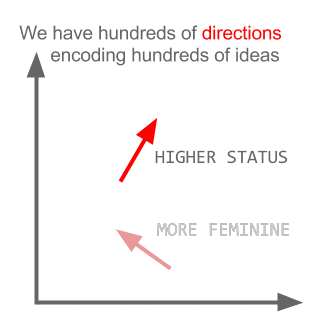
The word vectors are directions in space and can encode relationships between words.
The proximity of words to each other can be calculated through their cosine similarity.
```
model.wv.most_similar(positive=['boy'])
model.wv.most_similar(positive=['food'])
model.wv.most_similar(positive=['she','her','hers','herself'], negative=['he','him','his','himself'])
# she + her + hers + herself - he - him - his - himself
# Let's limit ourselves to top 50 words that related to food to visualize how they relate in vector space
f_tokens = [token for token,weight in model.wv.most_similar(positive=['food'], topn=50)]
from sklearn.metrics import pairwise
vectors = [model.wv[word] for word in f_tokens]
dist_matrix = pairwise.pairwise_distances(vectors, metric='cosine')
from sklearn.manifold import MDS
mds = MDS(n_components = 2, dissimilarity='precomputed')
embeddings = mds.fit_transform(dist_matrix)
import matplotlib.pyplot as plt
%matplotlib inline
_, ax = plt.subplots(figsize=(14,10))
ax.scatter(embeddings[:,0], embeddings[:,1], alpha=0)
for i in range(len(vectors)):
ax.annotate(f_tokens[i], ((embeddings[i,0], embeddings[i,1])))
```
**What kind of clusters of food-themed terms can you notice?**
# Doc2Vec
---
Doc2Vec, the most powerful extension of word2vec
---
Doc2vec (aka paragraph2vec or sentence embeddings) extrapolates the word2vec algorithm to larger blocks of text, such as sentences, paragraphs or entire documents.


Every paragraph is mapped to a unique vector, represented by a column in matrix D and every word is also mapped to a unique vector, represented by a column in matrix W .
The paragraph vector and word vectors are averaged or concatenated to predict the next word in a context.
Each additional context does not have be a fixed length (because it is vectorized and projected into the same space).
Additional parameters but the updates are sparse thus still efficent.
__2 architectures__:
1. Distrubted Memory (DM)
2. Distrubted Bag of Words (DBOW)
## Distrubted Memory (DM)
__Highlights__:
- Assign and randomly initialize paragraph vector for each doc
- Predict next word using context words and paragraph vector
- Slide context window across doc but keep paragraph vector fixed (hence: Distrubted Memory)
- Update weights via SGD and backprop
## Distrubted Bag of Words (DBOW)
__Highlights__:
- ONLY use paragraph vectors (no word vectors)
- Take a window of words in a paragraph and randomly sample which ones to predict using paragraph vector
- Simpler, more memory effecient

Let's try building our own Doc2Vec model with Gensim
Doc2Vec Parameters
- `size`: Number of dimensions for the embedding model
- `window`: Number of context words to observe in each direction within a document
- `min_count`: Minimum frequency for words included in model
- `dm` (distributed memory): '0' indicates DBOW model; '1' indicates DM
- `alpha`: Learning rate (initial); prevents model from over-correcting, enables finer tuning
- `iter`: Number of iterations through corpus
```
from gensim.models import Doc2Vec
from gensim.models.doc2vec import TaggedDocument
from nltk.corpus import reuters
# Tokenize Reuters corpus
tokenized_docs = [nltk.word_tokenize(reuters.raw(fileid)) for fileid in reuters.fileids()]
# Convert tokenized documents to TaggedDocuments
tagged_docs = [TaggedDocument(doc, tags=[idx]) for idx, doc in enumerate(tokenized_docs)]
# Create and train the doc2vec model. May take a few seconds
doc2vec = Doc2Vec(size=300, window=5, min_count=5, dm = 1, iter=10)
# Build the word2vec model from the corpus
doc2vec.build_vocab(tagged_docs)
```
You can also fortify your Doc2Vec models with pre-trained Word2Vec models.
Let's try re-training with GoogleNews-trained word vectors.
Download [here](https://drive.google.com/file/d/0B7XkCwpI5KDYNlNUTTlSS21pQmM/edit?usp=sharing)
(Size is 1.5gb)
```
#This may take a few minutes to run
w2v_loc = # your saved location of GoogleNews-vectors-negative300.bin.gz
doc2vec.intersect_word2vec_format(w2v_loc, binary=True)
doc2vec.train(tagged_docs, epochs=10, total_examples=doc2vec.corpus_count)
```
# Exercises
Using what you have learned from Week 1 and today's lecture, build a binary classifier for the movie reviews dataset in `data/movie_reviews.csv`.
| github_jupyter |
## SQLite
Embora a biblioteca Pandas seja a indica para o tratamento da maior parte das situações, por vezes torna-se necessário a utilização de bases de dados e das operações associadas às bases de dados.
Um dos motores de bases de dados disponíveis para pequenas implementações é o SQLite, cuja utilização se mostra de seguida.
Há operações que tipicamente são mais rápidas com o SQLite (select, filter e order) e outras que são mais rápidas com o Pandas (group by, load e join).
```
import sqlite3
conn = sqlite3.connect('empregados.db')
c = conn.cursor()
#Comando para criar uma tabela de dados (ex. empregado)
c.execute('CREATE TABLE empregado (nome varchar(255), departamento char(1), ano_nascimento int, salario double);')
#Criacao da tabela bonus
c.execute('CREATE TABLE bonus (nome varchar(255), bonus double);')
#Criacao de dados random para preencher a tabela empregado (populate)
#Existem bibliotecas em Python que permitem preenchimento das tabelas com dados mais parecidos com os reais ex. https://faker.readthedocs.io/en/master/#
import random
import string
def random_string(length, chars=string.ascii_letters):
return ''.join(random.choice(chars) for _ in range(length))
def gerar_empregados(n):
for _ in range(n):
yield (random_string(8), random_string(1, chars='abcdefg'), random.randint(1900, 2000), random.uniform(1e4, 1e5))
#Teste para ver se gera as linahs corretamente
for linha in gerar_empregados(3):
linha2 = "INSERT INTO empregado VALUES ('" + linha[0] + "','" + linha[1] + "'," + str(int(linha[2])) + "," + str(linha[3])+");"
print(linha2)
#Insercao de 10000 linhas utilizando o código acima
#Pode alterar o valor 10000 para o valor que desejar quanto estiver a fazer testes à BD
for lin in gerar_empregados(10000):
linha = "INSERT INTO empregado (nome, departamento, ano_nascimento, salario) VALUES ('" + lin[0] + "','" + lin[1] + "'," + str(lin[2]) + "," + str(lin[3])+");"
c.execute(linha)
for row in c.execute('SELECT nome FROM empregado'):
c.execute('INSERT INTO bonus (nome, bonus) VALUES ("' + row[0] + '",' + str(random.uniform(1e4, 1e5)) + ')')
#Guardar as alterações
conn.commit()
#Fechar a ligação à BD
conn.close()
```
De seguida vai-se testar o desempenho de execução da leitura de duas colunas à tabela empregado, uma utilizando o motor de BD SQLite e outra utilizando o Pandas (Fonte: http://bit.ly/2G93lbj).
```
#Para listar os valores das tabelas utilizaos seguintes métodos / funções:
import sqlite3
import pandas as pd
from timeit import timeit
conn = sqlite3.connect('empregados.db')
df_empregado = pd.read_sql_query('SELECT * FROM empregado', conn)
print(df_empregado.head())
def sql_select(conn):
conn.execute('SELECT nome, departamento FROM empregado')
conn.commit()
def pd_select(df_empregado):
df_empregado[["nome", "departamento"]]
%timeit sql_select(conn)
%timeit pd_select(df_empregado)
conn.close()
print("Comente os valores que obteve.")
```
## Exercício 1:
Teste as restantes situações de manipulações de dados, com as seguintes funções:
```
def sql_sort(c):
c.execute('SELECT * FROM empregado ORDER BY nome ASC;')
def pd_sort(df):
df.sort_values(by='nome')
def sql_join(c):
c.execute('SELECT empregado.nome, empregado.salario + bonus.bonus FROM empregado INNER JOIN bonus ON empregado.nome = bonus.nome')
def pd_join(df_emp, df_bonus):
joined = df_emp.merge(df_bonus, on='nome')
joined['total'] = joined['bonus'] + joined['salario']
def sql_filtrar(c):
c.execute('SELECT * FROM empregado WHERE departamento = "a";')
def pd_filtrar(df):
df[df['departamento'] == 'a']
def sql_groupby(c):
c.execute('SELECT avg(ano_nascimento), sum(salario) FROM empregado GROUP BY departamento;')
def pd_groupby(self):
df.groupby("departamento").agg({'ano_nascimento': np.mean, 'salario': np.sum})
```
## Exercício 2:
Compare a performance do SQLite vs Pandas utilizando um DataSet com 100 000 linhas.
Utilize o dataset relativo a vinhos disponível em https://www.kaggle.com/zynicide/wine-reviews/version/4
Olhe para os dados e faça medições de tempo relativas a alguns cálculos (ex. preco médio do vinho, vinhos mais baratos com a melhor classificação, etc.).
## Sintaxe Pandas vs SQL
No seguinte link http://bit.ly/2G9skev podem ver a comparação entre a sintaxe da biblioteca Pandas e a linguagem SQL.
## SQLAlchemy
Para realizar operações mais complexas consulte a seguinte biblioteca https://www.sqlalchemy.org/
## PostgreSQL
A base de dados SQLite permite guardar informação e realizar um grande conjunto de tarefas sendo uma das preferidas pelos analistas de dados.
No entanto, se desejar trabalhar com bases de dados mais robusta sugere-se a utilização das base de dados PostgreSQL.
Para poder interagir com este motor de base de dados terá primeiramente que instalar o PostGreSQL na sua máquina.
Faça import da bilioteca psycopg2 para trabalhar com o PostgreSQL.
```
import psycopg2
```
Se der erro terá de instalar o package com o comando:
```
pip install psycopg2
```
Veja um tutorial de como utilizar a biblioteca psycopg2.
| github_jupyter |
Now that we've built & trained logistic regression and decision tree models to classify the iris dataset in these previous posts:
- [LINK TO LOGISTIC REGRESSION POST]
- [LINK TO DECISION TREE POST]
We found that they were both really good in their own regard (potentially overfitting), but what if we had two models that had pros/cons of each but we wanted the best of both worlds? In machine learning such a thing exists and it's known as ensemble models where you combine multiple models together to make a single model with hopefully the strengths of each of the models that are combined. There are many methods to how we combine them together which are grouped under 2 main categories: averaging and boosting.
Averaging ensemble methods is when we build multiple models independantly and then average out their predictions. By doing this, the variance of the model is reduced and typically increases the performance of the model. Boosting ensemble methods is when we build models sequentially where each model depends on the previous and combine them in a specific strategy for the final model.
For this post, we'll use `sklearn` to train each of the models that we previously trained and combine them together to see how they fair against each other.
As always we begin by loading the data.
> Best practice would be to split the training and testing data here, but for brevity we will skip this step.
```
from sklearn.datasets import load_iris
import numpy as np
iris = load_iris()
```
Next we will train both our decision tree model and our logistic regression model.
```
from sklearn.tree import DecisionTreeClassifier
decisionTreeClassifier = DecisionTreeClassifier().fit(iris.data, iris.target)
from sklearn.linear_model import LogisticRegression
logisiticRegression = LogisticRegression().fit(iris.data, iris.target)
```
Now let's create a function that we can pass a model into that will give us a report on the score for the model, so then we can compare how well each model is performing.
```
from sklearn.metrics import classification_report
def get_model_score(model, model_name):
predictions = model.predict(iris.data)
print(f"{model_name}")
print(classification_report(iris.target, predictions))
get_model_score(logisiticRegression, 'logisiticRegression')
get_model_score(decisionTreeClassifier, 'decisionTreeClassifier')
```
As seen from the previous posts, the models are very strong in their own regard but we make sure to note that given the sample size we are potentially overfitting drastically to the dataset.
Since we will be making use of the voting classifier method later on, which takes a majority vote of the outcome of the models, we will need an odd number of models to give a worthwhile comparison, so let's train another model using k-Nearest Neighbours.
Now this model is a bit different in that we search for the best parameters for the model and then select the best model of this specific type that will go into our future ensemble. The `cv` argument stands for cross validation, which means the dataset is randomly split into a number of groups, similar to how we train/test split our dataset.
```
from sklearn.model_selection import GridSearchCV
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier()
knn_params = {'n_neighbors': np.arange(1,50)}
knn_grid_search = GridSearchCV(knn, knn_params, cv=5)
knn_grid_search.fit(iris.data, iris.target)
knn_best = knn_grid_search.best_estimator_
get_model_score(knn_best, 'KNearestNeighboursClassifier')
```
Now let's combine all of these models into a single ensemble model using the voting classifier method, this takes the majority of the models to decide on the output.
```
from sklearn.ensemble import VotingClassifier
models = [
('logisiticRegression', logisiticRegression),
('decisionTreeClassifier', decisionTreeClassifier),
('kNearestNeighboursClassifier', knn_best)
]
ensembleClassifier = VotingClassifier(models, voting='hard')
ensembleClassifier.fit(iris.data, iris.target)
get_model_score(ensembleClassifier, 'ensembleClassifier')
```
Now while our training dataset is small in this scenario, it's likely that the ensemble model will outperform the others due to it's lower variance and it being more adapted to multiple scenarios.
| github_jupyter |
# Multipe Regression
In this notebook we will learn the respective steps needed to compute Simple Linear Regression with data on house sales in King County, USA (https://www.kingcounty.gov) to predict house prices. You will:
* Upload and preprocess the data
* Write a function to compute the Multiple Regression weights
* Write a function to make predictions of the output given the input features
* Compare different models for predicting house prices
# Import all required libraries
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
#import seaborn as seabornInstance
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn import metrics
%matplotlib inline
```
# Upload, preprocess and check the data
Dataset is from house sales in King County, USA (https://www.kingcounty.gov).
```
# Import dataset
sales = pd.read_csv('housing.csv')
# Look at the table to check potential features
sales[:10]
# Check if the dataset contains NaN (Not-a-Number) values
sales.isnull().any()
# Maybe relevant for Friday/Saturday: Drop all columns / rows with NaN (Not-a-Number) values
sales_drop = sales.dropna()
# Check some statistics of the data
sales.describe()
# Understand that some variables are no good choices for linear regression,
# e.g. zipcode, lat, long, yr_renovated etc.
# Plot Scatter Matrix
plot_data_new = sales[['price', 'bedrooms', 'sqft_living', 'sqft_lot', 'sqft_above']]
from pandas.plotting import scatter_matrix
sm = scatter_matrix(plot_data_new, figsize = (15,15))
# Plot some feature relations
sales.plot(x = 'bedrooms', y = 'price', style = 'o')
plt.title('price vs. bedrooms')
plt.xlabel('bedrooms')
plt.ylabel('price')
plt.show()
# Plot some feature relations
sales.plot(x = 'sqft_living', y = 'price', style = 'o')
plt.title('price vs. sqft_living')
plt.xlabel('sqft_living')
plt.ylabel('price')
plt.show()
# Divide the data into some 'attributes' (X) and 'labels' (y) aka input and output.
X2 = sales[['bedrooms','bathrooms']]
y2 = sales['price']
```
# Split data into training and testing
```
# Split data set into 80% train and 20% test data
X_train, X_test, y_train, y_test = train_test_split(X2, y2, test_size=0.2, random_state=0)
# Look at the shape to check the split ratio
X_train.shape, X_test.shape, y_train.shape, y_test.shape
```
# Use a pre-build multiple regression function
```
# Train a Sklearn built-in function
reg = LinearRegression()
reg.fit(X_train, y_train)
X_train.columns
# See intercept and coefficients chosen by the model
print('Intercept:', reg.intercept_)
coeff_df = pd.DataFrame({'Features': X2.columns, 'Coefficients': reg.coef_}).set_index('Features')
coeff_df
# Do prediction on test data
y_pred = reg.predict(X_test)
# Check differences between actual and predicted value
df = pd.DataFrame({'Actual': y_test, 'Predicted': y_pred, 'Difference': y_pred - y_test},
columns=['Actual', 'Predicted', 'Difference']).astype(int)
df.head()
# Evaluate the performance
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred)))
# Performance of the model => r2_score = 1 - (variation unexplained / total variation)
# => How much of the variation is explained?
from sklearn.metrics import r2_score
r2_score(y_test, y_pred)
```
# Check for overfitting
```
# If r2 and RMSE in train and test differ dramatically => Overfitting!
# => Compare r2 and RSME in Test and Train!
# "Prework" needed to do the comparison
y_pred_train = reg.predict(X_train)
y_pred_test = reg.predict(X_test)
# => Compare r2 and RSME in Test and Train!
print('RSME Train:', np.sqrt(metrics.mean_squared_error(y_train, y_pred_train)))
print('RSME Test: ', np.sqrt(metrics.mean_squared_error(y_test, y_pred_test)))
print('R-2 Train:', r2_score(y_train, y_pred_train))
print('R-2 Test: ', r2_score(y_test, y_pred_test))
```
# For more information on performance evaluation see also
* https://en.wikipedia.org/wiki/Mean_absolute_error
* https://en.wikipedia.org/wiki/Mean_squared_error
* https://en.wikipedia.org/wiki/Root-mean-square_deviation
* https://en.wikipedia.org/wiki/Coefficient_of_determination
| github_jupyter |
```
import tensorflow as tf
import re
import numpy as np
import pandas as pd
from tqdm import tqdm
import collections
import itertools
from unidecode import unidecode
import malaya
import re
import json
def build_dataset(words, n_words, atleast=2):
count = [['PAD', 0], ['GO', 1], ['EOS', 2], ['UNK', 3]]
counter = collections.Counter(words).most_common(n_words - 10)
counter = [i for i in counter if i[1] >= atleast]
count.extend(counter)
dictionary = dict()
for word, _ in count:
dictionary[word] = len(dictionary)
data = list()
unk_count = 0
for word in words:
index = dictionary.get(word, 0)
if index == 0:
unk_count += 1
data.append(index)
count[0][1] = unk_count
reversed_dictionary = dict(zip(dictionary.values(), dictionary.keys()))
return data, count, dictionary, reversed_dictionary
def str_idx(corpus, dic, maxlen, UNK = 3):
X = np.zeros((len(corpus), maxlen))
for i in range(len(corpus)):
for no, k in enumerate(corpus[i][:maxlen]):
X[i, no] = dic.get(k, UNK)
return X
tokenizer = malaya.preprocessing._SocialTokenizer().tokenize
def is_number_regex(s):
if re.match("^\d+?\.\d+?$", s) is None:
return s.isdigit()
return True
def detect_money(word):
if word[:2] == 'rm' and is_number_regex(word[2:]):
return True
else:
return False
def preprocessing(string):
tokenized = tokenizer(string)
tokenized = [w.lower() for w in tokenized if len(w) > 2]
tokenized = ['<NUM>' if is_number_regex(w) else w for w in tokenized]
tokenized = ['<MONEY>' if detect_money(w) else w for w in tokenized]
return tokenized
with open('train-similarity.json') as fopen:
train = json.load(fopen)
left, right, label = train['left'], train['right'], train['label']
with open('test-similarity.json') as fopen:
test = json.load(fopen)
test_left, test_right, test_label = test['left'], test['right'], test['label']
np.unique(label, return_counts = True)
with open('similarity-dictionary.json') as fopen:
x = json.load(fopen)
dictionary = x['dictionary']
rev_dictionary = x['reverse_dictionary']
def position_encoding(inputs):
T = tf.shape(inputs)[1]
repr_dim = inputs.get_shape()[-1].value
pos = tf.reshape(tf.range(0.0, tf.to_float(T), dtype=tf.float32), [-1, 1])
i = np.arange(0, repr_dim, 2, np.float32)
denom = np.reshape(np.power(10000.0, i / repr_dim), [1, -1])
enc = tf.expand_dims(tf.concat([tf.sin(pos / denom), tf.cos(pos / denom)], 1), 0)
return tf.tile(enc, [tf.shape(inputs)[0], 1, 1])
def layer_norm(inputs, epsilon=1e-8):
mean, variance = tf.nn.moments(inputs, [-1], keep_dims=True)
normalized = (inputs - mean) / (tf.sqrt(variance + epsilon))
params_shape = inputs.get_shape()[-1:]
gamma = tf.get_variable('gamma', params_shape, tf.float32, tf.ones_initializer())
beta = tf.get_variable('beta', params_shape, tf.float32, tf.zeros_initializer())
return gamma * normalized + beta
def self_attention(inputs, is_training, num_units, num_heads = 8, activation=None):
T_q = T_k = tf.shape(inputs)[1]
Q_K_V = tf.layers.dense(inputs, 3*num_units, activation)
Q, K, V = tf.split(Q_K_V, 3, -1)
Q_ = tf.concat(tf.split(Q, num_heads, axis=2), 0)
K_ = tf.concat(tf.split(K, num_heads, axis=2), 0)
V_ = tf.concat(tf.split(V, num_heads, axis=2), 0)
align = tf.matmul(Q_, K_, transpose_b=True)
align *= tf.rsqrt(tf.to_float(K_.get_shape()[-1].value))
paddings = tf.fill(tf.shape(align), float('-inf'))
lower_tri = tf.ones([T_q, T_k])
lower_tri = tf.linalg.LinearOperatorLowerTriangular(lower_tri).to_dense()
masks = tf.tile(tf.expand_dims(lower_tri,0), [tf.shape(align)[0],1,1])
align = tf.where(tf.equal(masks, 0), paddings, align)
align = tf.nn.softmax(align)
align = tf.layers.dropout(align, 0.1, training=is_training)
x = tf.matmul(align, V_)
x = tf.concat(tf.split(x, num_heads, axis=0), 2)
x += inputs
x = layer_norm(x)
return x
def ffn(inputs, hidden_dim, activation=tf.nn.relu):
x = tf.layers.conv1d(inputs, 4* hidden_dim, 1, activation=activation)
x = tf.layers.conv1d(x, hidden_dim, 1, activation=None)
x += inputs
x = layer_norm(x)
return x
class Model:
def __init__(self, size_layer, num_layers, embedded_size,
dict_size, learning_rate, dropout, kernel_size = 5):
def cnn(x, scope):
x += position_encoding(x)
with tf.variable_scope(scope, reuse = tf.AUTO_REUSE):
for n in range(num_layers):
with tf.variable_scope('attn_%d'%n,reuse=tf.AUTO_REUSE):
x = self_attention(x, True, size_layer)
with tf.variable_scope('ffn_%d'%n, reuse=tf.AUTO_REUSE):
x = ffn(x, size_layer)
with tf.variable_scope('logits', reuse=tf.AUTO_REUSE):
return tf.layers.dense(x, size_layer)[:, -1]
self.X_left = tf.placeholder(tf.int32, [None, None])
self.X_right = tf.placeholder(tf.int32, [None, None])
self.Y = tf.placeholder(tf.float32, [None])
self.batch_size = tf.shape(self.X_left)[0]
encoder_embeddings = tf.Variable(tf.random_uniform([dict_size, embedded_size], -1, 1))
embedded_left = tf.nn.embedding_lookup(encoder_embeddings, self.X_left)
embedded_right = tf.nn.embedding_lookup(encoder_embeddings, self.X_right)
def contrastive_loss(y,d):
tmp= y * tf.square(d)
tmp2 = (1-y) * tf.square(tf.maximum((1 - d),0))
return tf.reduce_sum(tmp +tmp2)/tf.cast(self.batch_size,tf.float32)/2
self.output_left = cnn(embedded_left, 'left')
self.output_right = cnn(embedded_right, 'right')
print(self.output_left, self.output_right)
self.distance = tf.sqrt(tf.reduce_sum(tf.square(tf.subtract(self.output_left,self.output_right)),
1,keep_dims=True))
self.distance = tf.div(self.distance, tf.add(tf.sqrt(tf.reduce_sum(tf.square(self.output_left),
1,keep_dims=True)),
tf.sqrt(tf.reduce_sum(tf.square(self.output_right),
1,keep_dims=True))))
self.distance = tf.reshape(self.distance, [-1])
self.logits = tf.identity(self.distance, name = 'logits')
self.cost = contrastive_loss(self.Y,self.distance)
self.temp_sim = tf.subtract(tf.ones_like(self.distance),
tf.rint(self.distance))
correct_predictions = tf.equal(self.temp_sim, self.Y)
self.accuracy = tf.reduce_mean(tf.cast(correct_predictions, "float"))
self.optimizer = tf.train.AdamOptimizer(learning_rate = learning_rate).minimize(self.cost)
size_layer = 128
num_layers = 4
embedded_size = 128
learning_rate = 1e-4
maxlen = 50
batch_size = 128
dropout = 0.8
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model(size_layer,num_layers,embedded_size,len(dictionary),learning_rate,dropout)
sess.run(tf.global_variables_initializer())
saver = tf.train.Saver(tf.trainable_variables())
saver.save(sess, 'self-attention/model.ckpt')
strings = ','.join(
[
n.name
for n in tf.get_default_graph().as_graph_def().node
if ('Variable' in n.op
or 'Placeholder' in n.name
or 'logits' in n.name
or 'alphas' in n.name)
and 'Adam' not in n.name
and '_power' not in n.name
and 'gradient' not in n.name
and 'Initializer' not in n.name
and 'Assign' not in n.name
]
)
import time
EARLY_STOPPING, CURRENT_CHECKPOINT, CURRENT_ACC, EPOCH = 2, 0, 0, 0
while True:
lasttime = time.time()
if CURRENT_CHECKPOINT == EARLY_STOPPING:
print('break epoch:%d\n' % (EPOCH))
break
train_acc, train_loss, test_acc, test_loss = 0, 0, 0, 0
pbar = tqdm(range(0, len(left), batch_size), desc='train minibatch loop')
for i in pbar:
index = min(i+batch_size,len(left))
batch_x_left = str_idx(left[i: index], dictionary, maxlen)
batch_x_right = str_idx(right[i: index], dictionary, maxlen)
batch_y = label[i:index]
acc, loss, _ = sess.run([model.accuracy, model.cost, model.optimizer],
feed_dict = {model.X_left : batch_x_left,
model.X_right: batch_x_right,
model.Y : batch_y})
assert not np.isnan(loss)
train_loss += loss
train_acc += acc
pbar.set_postfix(cost=loss, accuracy = acc)
pbar = tqdm(range(0, len(test_left), batch_size), desc='test minibatch loop')
for i in pbar:
index = min(i+batch_size,len(test_left))
batch_x_left = str_idx(test_left[i: index], dictionary, maxlen)
batch_x_right = str_idx(test_right[i: index], dictionary, maxlen)
batch_y = test_label[i: index]
acc, loss = sess.run([model.accuracy, model.cost],
feed_dict = {model.X_left : batch_x_left,
model.X_right: batch_x_right,
model.Y : batch_y})
test_loss += loss
test_acc += acc
pbar.set_postfix(cost=loss, accuracy = acc)
train_loss /= (len(left) / batch_size)
train_acc /= (len(left) / batch_size)
test_loss /= (len(test_left) / batch_size)
test_acc /= (len(test_left) / batch_size)
if test_acc > CURRENT_ACC:
print(
'epoch: %d, pass acc: %f, current acc: %f'
% (EPOCH, CURRENT_ACC, test_acc)
)
CURRENT_ACC = test_acc
CURRENT_CHECKPOINT = 0
else:
CURRENT_CHECKPOINT += 1
print('time taken:', time.time()-lasttime)
print('epoch: %d, training loss: %f, training acc: %f, valid loss: %f, valid acc: %f\n'%(EPOCH,train_loss,
train_acc,test_loss,
test_acc))
saver.save(sess, 'self-attention/model.ckpt')
left = str_idx(['a person is outdoors, on a horse.'], dictionary, maxlen)
right = str_idx(['a person on a horse jumps over a broken down airplane.'], dictionary, maxlen)
sess.run([model.temp_sim,1-model.distance], feed_dict = {model.X_left : left,
model.X_right: right})
real_Y, predict_Y = [], []
pbar = tqdm(
range(0, len(test_left), batch_size), desc = 'validation minibatch loop'
)
for i in pbar:
index = min(i+batch_size,len(test_left))
batch_x_left = str_idx(test_left[i: index], dictionary, maxlen)
batch_x_right = str_idx(test_right[i: index], dictionary, maxlen)
batch_y = test_label[i: index]
predict_Y += sess.run(model.temp_sim, feed_dict = {model.X_left : batch_x_left,
model.X_right: batch_x_right,
model.Y : batch_y}).tolist()
real_Y += batch_y
from sklearn import metrics
print(
metrics.classification_report(
real_Y, predict_Y, target_names = ['not similar', 'similar']
)
)
strings.split(',')
def freeze_graph(model_dir, output_node_names):
if not tf.gfile.Exists(model_dir):
raise AssertionError(
"Export directory doesn't exists. Please specify an export "
'directory: %s' % model_dir
)
checkpoint = tf.train.get_checkpoint_state(model_dir)
input_checkpoint = checkpoint.model_checkpoint_path
absolute_model_dir = '/'.join(input_checkpoint.split('/')[:-1])
output_graph = absolute_model_dir + '/frozen_model.pb'
clear_devices = True
with tf.Session(graph = tf.Graph()) as sess:
saver = tf.train.import_meta_graph(
input_checkpoint + '.meta', clear_devices = clear_devices
)
saver.restore(sess, input_checkpoint)
output_graph_def = tf.graph_util.convert_variables_to_constants(
sess,
tf.get_default_graph().as_graph_def(),
output_node_names.split(','),
)
with tf.gfile.GFile(output_graph, 'wb') as f:
f.write(output_graph_def.SerializeToString())
print('%d ops in the final graph.' % len(output_graph_def.node))
freeze_graph('self-attention', strings)
def load_graph(frozen_graph_filename):
with tf.gfile.GFile(frozen_graph_filename, 'rb') as f:
graph_def = tf.GraphDef()
graph_def.ParseFromString(f.read())
with tf.Graph().as_default() as graph:
tf.import_graph_def(graph_def)
return graph
g = load_graph('self-attention/frozen_model.pb')
x1 = g.get_tensor_by_name('import/Placeholder:0')
x2 = g.get_tensor_by_name('import/Placeholder_1:0')
logits = g.get_tensor_by_name('import/logits:0')
test_sess = tf.InteractiveSession(graph = g)
test_sess.run(1-logits, feed_dict = {x1 : left, x2: right})
test_sess.run(1-logits, feed_dict = {x1 : batch_x_left, x2: batch_x_right})
```
| github_jupyter |
Центр непрерывного образования
# Программа «Python для автоматизации и анализа данных»
Неделя 1 - 1
*Татьяна Рогович, НИУ ВШЭ*
*Алла Тамбовцева, НИУ ВШЭ*
## Строки. Ввод и форматирование.
**ТЕКСТ (СТРОКИ) (STR, STRING):** любой текст внутри одинарных или двойных кавычек.
Важно: целое число в кавычках - это тоже строка, не целое число.
```
x1 = 'a'
x2 = "x"
x3 = """z z
z
z
z
zzz
z"""
x3
x = 'text'
print(type(x))
print(type('Hello, world!'))
print(type('2'))
```
Если попробовать сложить число-строку и число-число, то получим ошибку.
```
print('2' + 3)
```
Еще раз обратим внимание на текст ошибки. *TypeError: must be str, not int* - TypeError означает, что данная команда не может быть исполнена из-за типа какой-то из переменных. В данном случаем видим, что что-то должно быть "str", когда оно типа "int". Давайте попробуем сделать 3 тоже строкой. Кстати, неважно какие вы используете кавычки (только если это не текст с кавычками внутри), главное, чтобы открывающая и закрывающая кавычки были одинаковые.
```
print('2' + "3")
```
Такая операция называется контакенация (слиянием) строк.
Можно и умножить строку на число можно. Такая операция повторит нам строку заданное количество раз.
```
'2' * 3
```
Операции со строками тоже работают, если строки лежат внутри переменных.
```
word1 = 'John'
word2 = ' Brown'
print(word1 + word2)
word3 = word1 + word2 # можем результат контакенации положить в новую переменную
print(word3)
```
В прошлом блокноте мы преобразовывали строки в числа с помощью функции int().
```
print(2 + int('2512')) # нет ошибки!
print(2 + int('2512,0'))
float('12,2'.replace(',', '.'))
float('12.2')
```
С помощью функции str() мы можем превращать другие типы данных в строки.
```
print('abs' + str(123) + str(True))
```
Обратите внимание: эти функции (str, int, float и другие) не меняют тип самих данных или переменных, в которых они хранятся. Если мы не перезапишем значение, то строка станет числом только в конкретной строчке команды, а ее тип не изменится.
```
a = '2342123'
print(type(a))
print(2 + int(a))
print(type(a))
a = int(a) # перезаписываем значение переменной
print(type(a)) # теперь изменился и тип
```
# Ввод данных. input()
Познакомимся с функцией input() - ввод. Это функция позволяет нам просить
пользователя что-нибудь ввести или принять на ввод какие-то данные.
```
x = input('Как вас зовут?')
print(x)
```
'Как вас зовут?' - аргумент функции input(), который будет отображаться в строке ввода. Он необязателен, но он может быть полезен для вас (особенно, если в задаче требуется ввести больше одной строки данных и нужно не запутаться).
Обратите внимание, что функция input() все сохраняет в строковом формате данных. Но мы можем сделать из
числа-строки число-число с помощью функции int(). Введите "1" в обоих случаях и сравните результат.
```
print(type(input()))
print(type(int(input())))
```
Нужно следить за тем, какие данные вы считываете и менять их при необходимости. Сравните два примере ниже.
```
x = input('Input x value: ')
y = input('Input y value: ')
print(x + y) # произошла склейка строк, по умолчанию input() сохраняет значение типа str
x = int(input('Input x value: '))
y = int(input('Input y value: '))
print(x + y) # вот теперь произошло сложение чисел
```
## (∩`-´)⊃━☆゚.*・。゚ Задача
## Pig Latin 1
Pig Latin - вымышленный язык, который в конец каждого слова подставляет 'yay' (на самом деле оригинальные условия задачи несколько хитрее, но мы будем к ней возвращаться по ходу курса.
**Формат ввода**
Пользователь вводит слова.
**Формат вывода**
Слово на Pig Latin.
#### Примеры
Тест 1
**Ввод:**
apple
**Вывод:**
appleyay
```
# (∩`-´)⊃━☆゚.*・。゚
word = input('Введите слово: ')
print(word+'yay')
```
## (∩`-´)⊃━☆゚.*・。゚ Задача
## x потвторенное x раз в квадрате
Вводится целое число X. Повторите его X раз и возведите полученное число в квадрат. Напечатайте получившееся.
**Формат ввода**
Целое число
**Формат вывод**
Ответ на задачу
#### Примеры
Тест 1
**Ввод:**
2
**Вывод:**
484
```
x = input() # СЕЙЧАС ЭТО СТРОКА
x_2 = x * int(x)
x_2
int(x_2) ** int(x)
x = input()
print((int(x*int(x)))**2)
```
## (∩`-´)⊃━☆゚.*・。゚ Задача
## Хитрости умножения
Для умножения двузначного числа на 11 есть интересная хитрость: результат произведения можно получить если между первой и второй цифрой исходного числа вписать сумму этих цифр. Например, 15 * 11 = 1 1+5 5 = 165 или 34 * 11 = 3 3+4 4 = 374. Реализуйте программу, которая умножала бы двузначные числа на 11, используя эту хитрость. Пользоваться оператором умножения нельзя.
__Формат ввода:__
Вводится двузначное число.
__Формат вывода:__
Программа должна вывести ответ на задачу.
**Тест 1**
__Пример ввода:__
15
__Вывод:__
165
**Тест 2**
__Пример ввода:__
66
__Вывод:__
726
```
# ваше решение здесь
x = input()
n1 = int(x[0])
n2 = int(x[1])
if n1 + n2 < 10:
print(str(n1) + str(n1+n2) + str(n2))
else:
print(str(n1 + 1) + str((n1 + n2) % 10) + str(n2))
```
## Форматирование строк (string formatting)
А теперь посмотрим на то, как подставлять значения в уже имеющийся текстовый шаблон, то есть форматировать строки. Чтобы понять, о чём идет речь, можно представить, что у нас есть электронная анкета, которую заполняет пользователь, и мы должны написать программу, которая выводит на экран введенные данные, чтобы пользователь мог их проверить.
Пусть для начала пользователь вводит свое имя и возраст.
```
name = input("Введите Ваше имя: ")
age = int(input("Введите Ваш возраст: ")) # возраст будет целочисленным
```
Теперь выведем на экран сообщение вида
`Ваше имя: `имя`. Ваш возраст: `возраст`.`
Но прежде, чем это сделать, поймем, какого типа будут значения, которые мы будем подставлять в шаблон. Имя (переменная name) – это строка (string), а возраст (переменная age) – это целое число (integer).
```
result = "Ваше имя: %s. Ваш возраст: %i." % (name, age)
print(result)
```
Что за таинственные %s и %i? Все просто: оператор % в строке указывает место, на которое будет подставляться значение, а буква сразу после процента – сокращённое название типа данных (s – от string и i – от integer). Осталось только сообщить Python, что именно нужно подставлять – после кавычек поставить % и в скобках перечислить названия переменных, значения которых мы будем подставлять.
Для тех, кто работал в R: форматирование строк с помощью оператора % – аналог форматирования с помощью функции sprintf() в R.
Примечание: не теряйте часть с переменными после самой строки. Иначе получится нечто странное:
```
print("Ваше имя: %s. Ваш возраст: %i.")
```
Важно помнить, что если мы забудем указать какую-то из переменных, мы получим ошибку (точнее, исключение): Python не будет знать, откуда брать нужные значения.
```
print("Ваше имя: %s. Ваш возраст: %i." % (name))
```
Кроме того, создавая такие текстовые шаблоны, нужно обращать внимание на типы переменных, значения которых мы подставляем.
```
print("Ваше имя: %s. Ваш возраст: %s." % (name, age)) # так сработает
print("Ваше имя: %i. Ваш возраст: %s." % (name, age)) # а так нет
```
В первом случае код сработал: Python не очень строго относится к типам данных, и поэтому он легко может превратить целочисленный возраст в строку (два %s вместо %s и %i не является помехой). Во втором случае все иначе. Превратить строку, которая состоит из букв (name) в целое число никак не получится, поэтому Python справедливо ругается.
А что будет, если мы будем подставлять не целое число, а дробное, с плавающей точкой? Попробуем!
```
height = float(input("Введите Ваш рост (в метрах): "))
height
print("Ваш рост: %f м." % height) # f - от float
```
По умолчанию при подстановке значений типа float Python выводит число с шестью знаками после запятой. Но это можно исправить. Перед f нужно поставить точку и указать число знаков после запятой, которое мы хотим:
```
print("Ваш рост: %.2f м." % height) # например, два
print("Ваш рост: %.1f м. " % height) # или один
```
В случае, если указанное число знаков после запятой меньше, чем есть на самом деле (как в ячейке выше), происходит обычное арифметическое округление.
Рассмотренный выше способ форматирования строк – не единственный. Он довольно стандартный, но при этом немного устаревший. В Python 3 есть другой способ ‒ форматирование с помощью метода .format(). Кроме того, в Python 3.6 и более поздних версиях появился еще более продвинутый способ форматирования строк ‒ f-strings (formatted string literals).
F-strings очень удобны и просты в использовании: вместо % и сокращённого названия типа в фигурных скобках внутри текстового шаблона нужно указать название переменной, из которой должно подставляться значение, а перед всей строкой добавить f, чтобы Python знал, что нам нужна именно f-string.
```
print(f"Ваше имя: {name}. Ваш возраст: {age}. Рост: {height:.2f}")
```
Альтерантивой такого кода будет следующий синтаксис. Здесь переменные вставятся в порядке "упоминания" в строке.
```
print("Ваше имя: {}. Ваш возраст: {}".format(name, age))
```
Форматирование дробей добавляем так.
```
print("Ваше имя: {}. Ваш возраст: {}. Рост: {:.2f}".format(name, age, height))
```
Если указали формат переменной (.2f, например, ожидает float), то следим за порядком.
```
print("Ваше имя: {}. Ваш возраст: {}. Рост: {:.2f}".format(age, height, name))
```
Порядок заполнения можно указывать с помощью нумерации. Одну и тоже переменную можно использовать несколько раз.
```
print("Ваше имя: {2}. Ваш возраст: {0}. Рост: {1:.2f}. Все верно, {2}?".format(age, height, name))
```
Также при форматировании строк можно использовать результаты вычислений и результаты работы методах (о них немного позже). name.upper() в примере делает все символы строки name заглавными.
```
print(f"Ваше имя: {name.upper()}. Ваш возраст: {2020-1988}. Рост в футах: {height * 100 / 30.48 :.2f}")
```
## (∩`-´)⊃━☆゚.*・。゚ Задача
## Pi
Из модуля math импортируйте переменную pi. С помощью %f (первое предложение) и format (второе предложение) форматирования выведите строки из примера. Везде, где встречается число pi - это должна быть переменная pi, округленная до определенного знака после точки.
__Формат вывода:__
`Значение 22/7 (3.14) является приближением числа pi (3.1416)`
`Значение 22/7 3.142 является приближением числа pi 3.141592653589793`
```
import math
print(math.pi)
print('Значение 22/7 (%.2f) является приближением числа pi (%.4f)' % (math.pi, math.pi))
print(f'Значение 22/7 {math.pi:.3f} является приближением числа pi {math.pi}')
```
# Функция print(): аргументы и параметры
И еще немного поговорим про форматирование вывода. Это вам пригодится, если будете решать много задача на разных сайтах вроде hackerrank, где ответ нужно выводить каким-то хитрым образом. На самом деле функция print() немного сложнее, чем то, что мы уже видели. Например, мы можем печатать одной функцией больше чем один аргумент и использовать разделители вместо пробелов.
```
print(x)
a = 2
b = 4
print(a, b, sep='**', end='')
print(99)
print('2 + 3 =', 2 + 3)
z = 5
print('2 + 3 =', z)
```
У функции print есть параметры. Параметры - это такие свойства функций, которые задают значение невидимых нам аргументов внутри функции. Так существует параметр sep (separator), благодаря которому мы можем менять тип разделителя между аргументами print. В качестве разделителя может выступать любая строка.
Сравните:
```
print('1', '2', '3')
print('1', '2', '3', sep='.')
print('1', '2', '3', sep='')
print('1', '2', '3', sep='\n')
```
Параметр end задает то, что будет выведено в конце исполнения функции print. По умолчанию там стоит
невидимый символ '\n', который осуществляет переход на новую строку. Мы можем заменить его на любую строку.
Если мы хотим сохранить переход на новую строку - то обязательно прописываем наш невидимый символ внутри
выражения.
```
print('1', '2', '3', sep='.', end='!')
print('2') # строки слились
print('1', '2', '3', sep='.', end='!\n')
print('2') # вывод на новой строке
import this
n = int(input())
k = int(input())
x = n // k
print(x)
```
| github_jupyter |
<div id="image">
<img src="https://www.imt-atlantique.fr/sites/default/files/logo_mt_0_0.png" WIDTH=280 HEIGHT=280>
</div>
<div id="subject">
<CENTER>
</br>
<font size="4"></br> UE Artificial Inteligence: Project 2</font></br></div>
</CENTER>
<CENTER>
<font size="4"></br>April 2019</font></br></div>
</CENTER>
<CENTER>
<span style="color:blue">gustavo.rodrigues-dos-reis@imt-atlantique.fr</span>
</CENTER>
<CENTER>
<span style="color:blue">tales-marra@imt-atlantique.fr</span>
</CENTER>
</div>
"_Clustering is one of the most widely used techniques for exploratory data analysis. Its goal is to divide the data points into several groups such that points in the same group are similar and points in different groups are dissimilar to each other"_.
To perform a spectral clustering 3 main steps are necessary:
Create a similarity graph between our N objects to cluster
Compute the first k eigenvectors of its Laplacian matrix to define a feature vector for each object.
Run clustering algorithm (_k-means_) on these features to separate objects into k classes.
Construction of the similarity graph:
_"One can formally define an undirected graph as:_
\begin{equation*}
G=(N,E)\end{equation*}
_consisting of the set N of nodes and the set E of edges, which are unordered pairs of elements of N. The formal definition of a directed graph is similar, the only difference is that the set E contains ordered pairs of elements of N."_
There are basically two different ways of constructing a similarity graph, and they are:
**ε-neighborhood graph** - each vertex will be connected to others that fall inside the same circle of radius ε. Therefore, ε, is a parameter that needs to be adapted in order to get the local relationships.
**k-nearest neighbor graph**- each vertex will be connected to it's k-nearest neighbors.
```
import sys
import matplotlib.pyplot as plt
import networkx as nx
G = nx.grid_2d_graph(4, 4)
# write edgelist to grid.edgelist
nx.write_edgelist(G, path="grid.edgelist", delimiter=":")
# read edgelist from grid.edgelist
H = nx.read_edgelist(path="grid.edgelist", delimiter=":")
nx.draw(H)
plt.show()
```
Now that the graph is defined, it is time to deterimne it's Laplacian matrix vectors.
The adjacency matrix is a matrix that assumes the value 1 on the element _Aij_ if there is an edge between the vertex _i_ and the vertex _j_, otherwise it's 0.
And in the end we calculate the matrix D, which stands for degree (the number of edges of each vertex multiplied by the weight of each edge, if they exist).
By definition in Graph Theory, the Laplacian Matrix of a Graph is:
\begin{equation*}
L=D-A
\end{equation*}
Once we calculated L, we have to determine it's eigenvalues. An eigenvector of a matrix satisfies the condition:
\begin{equation*}
A.v=\lambda.v
\end{equation*}
Where A is the matrix,and the lambdas are the eigenvalues of A.
```
A=nx.adjacency_matrix(G)
print(A.todense())
L=nx.laplacian_matrix(G)
print(L.todense())
```
It is important to notice that, When the similarity graph is not fully connected, the multiplicity of the eigenvalue λ = 0 gives us an estimation of k.
To finish, we apply a traditional clustering algorithm,such as _k-means_ for example, into the rows of a matrix in which each column corresponds to an eigenvector.
```
from sklearn.cluster import SpectralClustering
clustering_graph = SpectralClustering()
pred=clustering_graph.fit_predict(G)
nx.draw(G,node_color=pred)
import numpy as np
xc = np.load("dataset1010-40.npz")['x']
yc = np.load("dataset1010-40.npz")['y']
xc.shape
from sklearn.cluster import SpectralClustering
clustering2= SpectralClustering(eigen_solver='arpack',affinity='nearest_neighbors')
x=xc[yc==1]
x1=x
yc_pred=clustering2.fit(x)
print(np.unique(yc_pred))
A=clustering2.affinity_matrix_
clustering3= SpectralClustering(n_clusters=5,eigen_solver='arpack',affinity='nearest_neighbors')
yc_pred_2=clustering3.fit_predict(x)
clusters=[]
for index in np.unique(yc_pred_2):
cluster=x[yc_pred_2==index]
clusters.append(cluster)
means=[]
for element in clusters:
x=element.reshape(-1, 10, 10)
mean=np.mean(x,axis=0)
means.append(mean)
import matplotlib.pyplot as plt
import seaborn as sns
for index,element in enumerate(means):
plt.figure(figsize=(12,8))
sns.heatmap(data=element,annot=True)
plt.title('Cheese Density for Cluster '+str(index))
plt.show()
x=x.reshape(-1,10,10)
mean_old=np.mean(x,axis=0)
plt.figure(figsize=(12,8))
sns.heatmap(data=mean_old,annot=True)
plt.title("Cheese Density for Python Wins Analyzing all Elements")
plt.show()
```
Now we are going to use the method of a paper published at the NIPS conference by _Lihi Zelnik-Manor_ and _Pietro Perona_ in 2005 which proposes a self tuning Spectral Clustering. As the paper proposes many different techniques to treat open issues on the spectral clustering, we've chosen to show one of the solutions proposed by them to the problem of the definition of optimal number of clusters, by doing an analysis of the eigenvalues and eigenvectors of the affinity matrix laplacian.
_param A_: Affinity matrix
_param plot_: plots the sorted eigen values for visual inspection
return A tuple containing:
- the optimal number of clusters by eigengap heuristic
- all eigen values
- all eigen vectors
This method performs the eigen decomposition on a given affinity matrix,
following the steps recommended in the paper:
- Construct the normalized affinity matrix:
\begin{equation*}
L = D^{−1/2}ÂD^{−1/2}
\end{equation*}
- Find the eigenvalues and their associated eigen vectors
- Identify the maximum gap which corresponds to the number of clusters by eigengap heuristic
_References:_
https://papers.nips.cc/paper/2619-self-tuning-spectral-clustering.pdf
http://www.kyb.mpg.de/fileadmin/user_upload/files/publications/attachments/Luxburg07_tutorial_4488%5b0%5d.pdf
```
import utils
import scipy
from scipy.sparse import csgraph
from scipy.sparse.linalg import eigsh
def eigenDecomposition(A, plot = True):
L = csgraph.laplacian(A, normed=True)
n_components = A.shape[0]
print(n_components)
# LM parameter : Eigenvalues with largest magnitude (eigs, eigsh), that is, largest eigenvalues in
# the euclidean norm of complex numbers.
eigenvalues, eigenvectors = eigsh(L, k=n_components-1, which="LM", sigma=1.0, maxiter=5000)
if plot:
plt.title('Largest eigen values of input matrix')
plt.scatter(np.arange(len(eigenvalues)), eigenvalues)
plt.grid()
# Identify the optimal number of clusters as the index corresponding
# to the larger gap between eigen values
index_largest_gap = np.argmax(np.diff(eigenvalues))
nb_clusters = index_largest_gap + 1
return nb_clusters, eigenvalues, eigenvectors
```
Here we calculate the affinity matrix based on input coordinates matrix and the numeber
of nearest neighbours.
Apply local scaling based on the k nearest neighbour
_References_:
https://papers.nips.cc/paper/2619-self-tuning-spectral-clustering.pdf
```
from scipy.spatial.distance import pdist, squareform
def getAffinityMatrix(coordinates, k = 7):
# calculate euclidian distance matrix
dists = squareform(pdist(coordinates))
# for each row, sort the distances ascendingly and take the index of the
#k-th position (nearest neighbour)
knn_distances = np.sort(dists, axis=0)[k]
knn_distances = knn_distances[np.newaxis].T
# calculate sigma_i * sigma_j
local_scale = knn_distances.dot(knn_distances.T)
affinity_matrix = dists * dists
affinity_matrix = -affinity_matrix / local_scale
# divide square distance matrix by local scale
affinity_matrix[np.where(np.isnan(affinity_matrix))] = 0.0
# apply exponential
affinity_matrix = np.exp(affinity_matrix)
np.fill_diagonal(affinity_matrix, 0)
return affinity_matrix
affinity_matrix = getAffinityMatrix(x1, k = 10)
k, _, _ = eigenDecomposition(affinity_matrix)
print(f'Optimal number of clusters {k}')
xa = np.load("dataset1010-40.npz")['x']
ya = np.load("dataset1010-40.npz")['y']
x3=x2=xa[ya!=-1]
clustering4= SpectralClustering(n_clusters=5,eigen_solver='arpack',affinity='nearest_neighbors')
ya_pred=clustering4.fit_predict(x2)
clusters1=[]
for index in np.unique(ya_pred):
cluster=x2[ya_pred==index]
clusters1.append(cluster)
means1=[]
for element in clusters:
x=element.reshape(-1, 10, 10)
mean=np.mean(x,axis=0)
means1.append(mean)
import matplotlib.pyplot as plt
import seaborn as sns
for index,element in enumerate(means1):
plt.figure(figsize=(12,8))
sns.heatmap(data=element,annot=True)
plt.title('Cheese Density for Cluster '+str(index))
plt.show()
x2=x2.reshape(-1,10,10)
mean_new=np.mean(x,axis=0)
plt.figure(figsize=(12,8))
sns.heatmap(data=mean_new,annot=True)
plt.title("Cheese Density for Python Wins and Draws Analyzing all Elements")
plt.show()
plt.figure(figsize=(12,8))
sns.heatmap(data=mean_new-mean_old,annot=True)
plt.title("Difference of the Average Cheese Density Matrices")
plt.show()
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import logging
import pandas as pd
import numpy as np
from kernel_wasserstein_flows.gradient_flow import gradient_flow
from kernel_wasserstein_flows.utils import generate_XY_mog_square
from kernel_wasserstein_flows.kernels import gaussian_kernel
from kernel_wasserstein_flows.config import LOG_LEVELS
import kernel_wasserstein_flows.gradient_flow as gf
gf.MAX_TRAJECTORY_SNAPSHOTS = 1000
LOG_LEVELS['gradient_flow'] = logging.INFO
# LOG_LEVELS['kale.gradient_descent'] = logging.INFO
# LOG_LEVELS['kale.optim.newton'] = logging.INFO
```
# Run Gradient Flows
```
results = {}
import logging
import pandas as pd
import numpy as np
from kernel_wasserstein_flows.gradient_flow import gradient_flow
from kernel_wasserstein_flows.utils import generate_XY_mog_square
from kernel_wasserstein_flows.kernels import gaussian_kernel
from kernel_wasserstein_flows.config import LOG_LEVELS
import kernel_wasserstein_flows.gradient_flow as gf
gf.MAX_TRAJECTORY_SNAPSHOTS = 1000
LOG_LEVELS['gradient_flow'] = logging.INFO
# LOG_LEVELS['kale.gradient_descent'] = logging.INFO
# LOG_LEVELS['kale.optim.newton'] = logging.INFO"
_gf_default_kwargs = dict(
max_iter=50000,
lr=0.001,
random_seed=20,
num_noisy_averages=1,
generator=generate_XY_mog_square,
generator_kwargs=dict(N=240, d=2, dist=1.8, std=1/4, random_seed=42, y_std=0.2, y_rel_dist=0.1, return_potential_functions=False),
kernel=gaussian_kernel,
kernel_kwargs={'sigma': 0.5},
)
_kale_default_kwargs={
"inner_max_iter": 200,
"inner_tol": 1e-6,
"inner_a": 0.4,
"inner_b": 0.8,
"inplace": False,
"input_check":True,
"dual_gap_tol": 1e-3
}
from kernel_wasserstein_flows.mmd import mmd, mmd_first_variation
from kernel_wasserstein_flows.kale import kale_penalized, kale_penalized_first_variation
# from kernel_wasserstein_flows.kale_torch import kale_penalized, kale_penalized_first_variation
cases = [
("mmd", mmd, mmd_first_variation, {}, lambda x:1e-10),
("kale_10000", kale_penalized, kale_penalized_first_variation, {'lambda_': 10000, **_kale_default_kwargs}, lambda x:1e-10),
("kale_01", kale_penalized, kale_penalized_first_variation, {'lambda_': 0.1, **_kale_default_kwargs}, lambda x:1e-10),
("kale_01_noise_injection", kale_penalized, kale_penalized_first_variation, {'lambda_': 0.1, **_kale_default_kwargs}, lambda x:3e-1),
("kale_0001", kale_penalized, kale_penalized_first_variation, {'lambda_': 0.001, **_kale_default_kwargs}, lambda x:1e-10)
]
results = {}
for name, loss, loss_first_var, loss_kwargs, noise_injection_callback in cases:
args, (X, Y), (trajectories, records, loss_states) = gradient_flow(
loss=loss, loss_first_variation=loss_first_var, loss_kwargs=loss_kwargs, noise_level_callback=noise_injection_callback, **_gf_default_kwargs
)
results[name] = {'args': args, 'X': X, 'Y': Y, 'trajectories': trajectories, 'records': records, 'loss_states': loss_states}
# uncomment lines below in this cell to view an interactive visualisation of the computed flows
# from kernel_wasserstein_flows.plotting import vizualize_results
# from kernel_wasserstein_flows.utils import compute_velocity_field
#
# %matplotlib ipympl
#
# _exp_name = "kale_01"
#
# args, X, Y, trajectories, records, loss_states = results[_exp_name]['args'], results[_exp_name]['X'], results[_exp_name]['Y'], results[_exp_name]['trajectories'], results[_exp_name]['records'], results[_exp_name]['loss_states']
#
# if args['generator_kwargs'].get('d') == 2:
# vs = compute_velocity_field(
# X, trajectories,
# args['kernel'], args['kernel_kwargs'], args['loss'], args['loss_kwargs'],
# args['loss_first_variation'], loss_states
# )
# else:
# vs = None
# vizualize_results(X, Y, trajectories, pd.json_normalize(records), _exp_name, metrics_subset=['loss'],
# velocities=vs)
from kernel_wasserstein_flows.extras import unadjusted_langevin_algorithm
args, (X, Y), (trajectories, records, loss_states) = unadjusted_langevin_algorithm(
max_iter=50000,
lr=0.001,
random_seed=42,
generator=generate_XY_mog_square,
generator_kwargs=dict(N=240, d=2, dist=1.8, std=1/4, random_seed=42, y_std=0.2, y_rel_dist=0.1, return_potential_functions=True),
)
results['kl'] = {'args': args, 'X': X, 'Y': Y, 'trajectories': trajectories[::50], 'records': records, 'loss_states': loss_states}
# vizualize_results(X, Y, trajectories, None, None, metrics_subset=[],velocities=None)
```
# Compare Gradient Flows Trajectories via Sinkhorn Divergence
```
import ot
import numpy as np
all_sinkhorn_distances = {}
couples_kl = (('kl', 'mmd'), ('kl', 'kale_0001'), ('kl', 'kale_01'), ('kl', 'kale_10000'))
couples_mmd = (('mmd', 'kl'), ('mmd', 'kale_0001'), ('mmd', 'kale_01'), ('mmd', 'kale_10000'))
i = 0
for reference_div, couples in zip(('kl', 'mmd'), (couples_kl, couples_mmd)):
sinkhorn_distances_reference_div = {}
for couple in couples:
print(couple)
sinkhorn_dists = []
div1, div2 = couple
for (_Y1, _Y2) in zip(
results[div1]['trajectories'][::10],
results[div2]['trajectories'][::10]
):
_X1 = results[div1]['X']
_X2 = results[div2]['X']
_M = ((_Y1[:, None, :] - _Y2[None, :, :])**2).sum(axis=2)
ret =ot.sinkhorn(
1/len(_Y1) * np.ones((len(_Y1),)), 1/len(_Y2) * np.ones((len(_Y2),)), _M, reg=0.2
)
sinkhorn_dists.append((ret * _M).sum())
sinkhorn_distances_reference_div[couple] = sinkhorn_dists
all_sinkhorn_distances[reference_div] = sinkhorn_distances_reference_div
import matplotlib.pyplot as plt
from matplotlib import rc
f, axs = plt.subplots(ncols=3, figsize=(15, 5))
for ax, _ref_div_name in zip(axs, ('mmd', 'kl')):
for exp_name in results.keys():
if exp_name in ('kl', 'mmd', 'kale_01_noise_injection'):
continue
iterations = [i * 500 for i in range(len(all_sinkhorn_distances[_ref_div_name][(_ref_div_name,exp_name)]))]
ax.plot(iterations, all_sinkhorn_distances[_ref_div_name][(_ref_div_name,exp_name)], label=exp_name)
ax.legend(loc='lower left')
ax.set_xlabel('Iteration')
ax.set_title(f'Distance to {_ref_div_name.upper()}')
axs[-1].plot([r['loss'] for r in results['kale_01']['records']], label='no noise injection')
axs[-1].plot([r['loss'] for r in results['kale_01_noise_injection']['records']], label='noise injection')
axs[-1].set_yscale('log')
axs[-1].set_xlabel('Iteration')
axs[-1].legend()
axs[-1].set_title('KALE')
```
| github_jupyter |
## Truc Huynh
- 1/11/2022
- 7:58 PM
- Workshop turn it in
# Workshop 1
This notebook will cover the following topics:
1. Basic Input/Output and formatting
2. Decision Structure and Boolean Logic
3. Basic Loop Structures
4. Data Structures
## 1.1 Basic Input/Output and formatting (Follow):
**Learning Objectives:**
1. Perform basic input and output for a user
2. Format input and output
3. Perform basic math on input and output
4. Understand different datatypes
```
# Add comments below
# This is a normal string print
print ("This is a print statement")
# This is is f tring print with number
print (f"Try printing some different things including number {5}")
# Get input from user and store in input_string
input_string = input("Enter a short string here:")
# Display the user input
print ("The string you input is: ", input_string)
# define float_number value 2
float_number = 2
# print the float number with 2 decimal place format
print("This will print a float number with 2 decimal places: ", format(float_number, '.2f'))
num1 = 5
num2 = 10
print("This will multiply {num1} times {num2} and display the result: ", num1*num2)
```
## 1.1 Basic Input/Output and formatting (Group):
1. Create a small program that will take a celcius temperature from the user and convert it to fahrenheit
2. Print the output result to 3 decimal places
3. Print a degree sign after the output
4. Try to figure out how to create the program in the fewest lines possible
Fahrenheit = (9/5)(Degrees Celcius) + 32
```
def calculate_fahrenheit(celcius_degree):
return (f"The degree in Fahrenheit is {format((float(celcius_degree)*9)/5+32,'.3f')} \N{DEGREE SIGN}F")
cel_input = input('Please enter a degree in celius: ')
print(calculate_fahrenheit(cel_input))
```
## 1.2 Decision Structure and Boolean Logic (Follow):
**Learning Objectives:**
1. Understand boolean logic and logical operators
2. Understand If/Else statements and program flow
```
#Add comments below
day = int(input('Enter a number (1-7) for the day of the week: '))
if day == 1:
print('Monday')
elif day == 2:
print('Tuesday')
elif day == 3:
print('Wednesday')
elif day == 4:
print ('Thursday')
elif day == 5:
print ('Friday')
elif day == 6:
print ('Saturday')
elif day == 7:
print ('Sunday')
else:
print ('Error: Please enter a number in the range 1-7.')
```
After running this program, think about and answer these questions:
1. What is the program doing?
2. Does every line in the code execute, explain?
3. Could you find a way to break it?
#### Answer:
1. What is the program doing: The program print out the day of the week with respective number (start with 1 for Monday and 7 for Sunday)
2. Does every line in the code execute: Yes, everyline of code execute with taking input then go through if-elif-else condition (All the syntax and else if else is in correct order and connected)
3. Could you find a way to break it: Yes, just enter a string and it will break. I include a solution below to fix this bby validation the input inside the function instead parse it in the get input
```
def day_of_week(number):
if number.isdigit() and int(number) in range(1,8):
week_days=['Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday','Saturday','Sunday']
print(week_days[int(number)-1])
else:
print('Please enter a number from 1 to 7 only!')
day = input('Enter a number (1-7) for the day of the week: ')
day_of_week(day)
#Add comments below
RED = 'red'
BLUE = 'blue'
YELLOW = 'yellow'
color1 = input('Enter the first primary color in lower case letters: ')
color2 = input('Enter the second primary color in lower case letters: ')
if color1 != RED and color1 != BLUE and color1 != YELLOW:
print('Error: The first color you entered is invalid.')
elif color2 != RED and color2 != BLUE and color2 != YELLOW:
print('Error: The second color you entered is invalid.')
elif color1 == color2:
print('Error: The two colors you entered are the same.')
else:
if color1 == RED:
if color2 == BLUE:
print('purple')
else:
print('orange')
elif color1 == BLUE:
if color2 == RED:
print('purple')
else:
print('green')
else:
if color2 == RED:
print('orange')
else:
print('green')
```
After running this program, think about and answer these questions:
1. What is the program doing?
2. Does every line in the code execute, explain?
3. Could you find a way to break it?
4. Replace the 'ands' with 'ors' and discuss how the program executes now.
### Answer:
After running this program, think about and answer these questions:
1. What is the program doing: Program take 2 color of user input (out of 3 color: red, blue green) and output the mixed color of user input's color. The prgram also validate the user input to make sure input is validate
2. Does every line in the code execute, explain: Yes, every line of code is execute. The code follow if-elif-else nested if structure and corrected indent. Therefore, every line of code is executed.
3. Could you find a way to break it? Nope, it is well structure and cant break. The only missing case is both color 1 and color is invalid, it would be nice to print it out.
4. Replace the 'ands' with 'ors' and discuss how the program executes now: Well if we add the or program will not run correctly. We are now encounter a logic error. For example in the 1st if statement, if user enter BLUE or YELLOW, the app will ignore them. it will still print out the invalid satatement eventhough the BLUE and GREEN is valid. The reason is the 'or' statement just make the print statement execute because it satisied the condition color is not 'RED'.
## 1.2 Decision Structure and Boolean Logic (Group):
Develop a program that will do the following:
1. Take input from the user for length and width for two different sets of numbers. Fours total inputs.
2. Multiply the length and width together and find the area for the two different sets of numbers.
3. Compare the areas and print which one is larger.
4. Be sure to handle non-numeric inputs.
```
def compare_area(length1, width1, length2, width2):
if length1.isdigit() and length2.isdigit() and width1.isdigit() and width2.isdigit():
area1=int(length1)*int(width1)
area2=int(length2)*int(width2)
print (f"Area of reactangle 1 is: {area1}")
print (f"Area of reactangle 2 is: {area2}")
print("Rectangle 1 have bigger area than Rectangle 2"
if area1 > area2 else "Rectangle 2 have bigger area than Rectangle 1")
else:
print("Application only take integer only")
length1 = input("Please enter the length of object 1: ")
width1 = input("Please enter the width of object 1: ")
length2 = input("Please enter the length of object 2: ")
width2 = input("Please enter the width of object 2: ")
compare_area(length1, width1, length2, width2)
```
## 1.3 Basic loop structures (Follow):
**Learning Objectives:**
1. Understanding repetative structures
2. Understanding while loops
3. Understanding for loops
4. Nested loops and order N run time
```
#Add comments below
number = 1.0
total = 0.0
# If number smaller than 0 app will stop working
while number > 0:
number = float(input('Enter a positive number (negative to quit): '))
if number > 0:
total = total + number
print (f'Total = {total:.2f}')
```
After running this program, think about and answer these questions:
1. What is the program doing?
2. How many time does each line in the code execute, explain?
3. Could you find a way to break it?
4. Try to rewrite the program using a for loop.
```
# Add comments to the program
caloriesPerMinute = 4.2
caloriesBurned = 0.0
print ('Minutes\t\tCalories Burned')
print ('-------------------------------')
# for miniute in range 10 to 30, increase by 5
for minutes in range(10, 31, 5):
caloriesBurned = caloriesPerMinute * minutes
print (minutes, "\t\t", caloriesBurned)
```
After running this program, think about and answer these questions:
1. What is the program doing?
2. How many time does each line in the code execute, explain?
3. Could you find a way to break it?
4. Try to rewrite the program using a while loop.
```
###### Add comments to the program
totalRainfall = 0.0
monthRainfall = 0.0
averageRainfall = 0.0
years = 0
totalMonths = 0
years = int(input('Enter the number of years: '))
for year in range(years):
print (f'For year {year + 1}:')
for month in range(12):
monthRainfall = float(input('Enter the rainfall amount for the month: '))
totalMonths += 1
totalRainfall += monthRainfall
averageRainfall = totalRainfall / totalMonths
print(f'For {totalMonths} months')
print(f'Total rainfall: {totalRainfall:,.2f} inches')
print(f'Average monthly rainfall: {averageRainfall:,.2f} inches')
```
After running this program, think about and answer these questions:
1. What is the program doing?
2. How many time does each line in the code execute, explain?
3. Could you find a way to break it?
4. Compare the number of executuions for the inner for loop to the outer for loop.
5. What order of magnitude larger are the inner instructions executing?
## 1.3 Basic loop structures (Group):
Develop a program that will:
1. Take input from the user asking how many organisms to start with.
2. Take input from the user asking the average daily increase for the organisms.
3. Take input from the user asking the number of days to multiply.
4. Make sure the daily increase was entered as a percentage, if not correct it.
5. Print each day and the increase in organsisms for each day.
6. Make sure to format the printing so that it is readable.
```
organism_size = int(input('please enter your organisms to start with here: '))
percentage = float(input('please enter your percentage here (format: 00.00): '))
day_count = int(input('please enter your number of days to multiply: '))
for i in range (day_count):
organism_size += organism_size*percentage
print(f"On day {i+1}, the {organism_size}" )
```
## 1.4 Data Structures (Follow):
**Learning Objectives:**
1. Understand list and tuples
2. Understand dictionaries and sets
3. Understand strings
```
# Add comments to the program
total_sales = 0.0
# define tupples
daily_sales_tuple = (7.0, 5.0, 3.2, 1.7, 6.0, 4.9, 1.1)
daily_sales_list = [7.0, 5.0, 3.2, 1.7, 6.0, 4.9, 1.1]
# define days of week list
days_of_week = ['Sunday', 'Monday', 'Tuesday',
'Wednesday', 'Thursday', 'Friday',
'Saturday']
for number in daily_sales_list:
total_sales += number
print (f'Total sales for the week: ${total_sales:,.2f}')
total_sales = 0.0
for number in daily_sales_tuple:
total_sales += number
print (f'Total sales for the week: ${total_sales:,.2f}')
print (f'Sales per day: ')
for i in range(7):
print(f"{days_of_week[i] : <10}", f"{daily_sales_list[i] : >20}")
print(f"{days_of_week[i] : <10}", f"{daily_sales_tuple[i] : >20}")
daily_sales_list[5] = 10.0
#daily_sales_tuple[5] = 10.0 #uncomment this line to see what happens
new_value = 3.2
daily_sales_list.append(new_value)
new_daily_sales_tuple = daily_sales_tuple + (new_value,)
days_of_week.append('Sunday')
print("")
print (f'Sales per day: ')
for i in range(len(days_of_week)):
print(f"{days_of_week[i] : <10}", f"{daily_sales_list[i] : >20}")
print(f"{days_of_week[i] : <10}", f"{new_daily_sales_tuple[i] : >20}")
```
After running the above code, think about and answer the following questions:
1. What is the program doing?
2. What does the append function do?
3. Which actual entry is changed when setting daily sales[5] to 10?
4. Discuss and the similarities and differences between lists and tuples.
```
# Add comments to the program
import random
capitals = {'Alabama':'Montgomery', 'Alaska':'Juneau',
'Arizona':'Phoenix', 'Arkansas':'Little Rock',
'California':'Sacramento', 'Colorado':'Denver',
'Connecticut':'Hartford', 'Delaware':'Dover',
'Florida':'Tallahassee', 'Georgia':'Atlanta',
'Hawaii':'Honolulu', 'Idaho':'Boise',
'Illinois':'Springfield', 'Indiana':'Indianapolis',
'Iowa':'Des Moines', 'Kansas':'Topeka',
'Kentucky':'Frankfort', 'Louisiana':'Baton Rouge',
'Maine':'Augusta', 'Maryland':'Annapolis',
'Massachusetts':'Boston', 'Michigan':'Lansing',
'Minnesota':'Saint Paul', 'Mississippi':'Jackson',
'Missouri':'Jefferson City', 'Montana':'Helena',
'Nebraska':'Lincoln', 'Nevada':'Carson City',
'New Hampshire':'Concord', 'New Jersey':'Trenton',
'New Mexico':'Santa Fe', 'New York':'Albany',
'North Carolina':'Raleigh', 'North Dakota':'Bismarck',
'Ohio':'Columbus', 'Oklahoma':'Oklahoma City',
'Oregon':'Salem', 'Pennsylvania':'Harrisburg',
'Rhode Island':'Providence', 'South Carolina':'Columbia',
'South Dakota':'Pierre', 'Tennessee':'Nashville',
'Texas':'Austin', 'Utah':'Salt Lake City',
'Vermont':'Montpelier', 'Virginia':'Richmond',
'Washington':'Olympia', 'West Virginia':'Charleston',
'Wisconsin':'Madison', 'Wyoming':'Cheyenne'}
correct = 0
wrong = 0
next_question = True
index = 0
user_solution = ''
while next_question:
state_iterator = iter(capitals)
index = (random.randint(1,50) - 1)
for i in range (index-1):
temp = state_iterator.__next__()
current_state = str(state_iterator.__next__())
user_solution = input(f'What is the capital of {current_state}? 'f'(or enter 0 to quit): ')
if user_solution == '0':
next_question = False
print(f'You had {correct} correct responses and 'f'{wrong} incorrect responses.')
elif user_solution == capitals[current_state]:
correct = correct + 1
print('That is correct.')
else:
wrong = wrong + 1
print('That is incorrect.')
```
After running the above code, think about and answer the following questions:
1. What is the program doing?
2. What does a key in a dictionary do? What does the value in a dictionary do?
3. Where would you be more likely to use a dictionary versus using a list or tuple?
4. How are dictionaries different from tuples and lists?
```
# add comments to the program
baseball = set(['Jodi', 'Carmen', 'Aida', 'Alicia'])
basketball = set(['Eva', 'Carmen', 'Alicia', 'Sarah'])
print('The following students are on the baseball team:')
for name in baseball:
print(name)
print()
print('The following students are on the basketball team:')
for name in basketball:
print(name)
print()
print('The following students play both baseball and basketball:')
for name in baseball.intersection(basketball):
print(name)
print()
print('The following students play either baseball or basketball:')
for name in baseball.union(basketball):
print(name)
print()
print('The following students play baseball, but not basketball:')
for name in baseball.difference(basketball):
print(name)
print()
print('The following students play basketball, but not baseball:')
for name in basketball.difference(baseball):
print(name)
print()
print('The following students play one sport, but not both:')
for name in baseball.symmetric_difference(basketball):
print(name)
```
After running the above code, think about and answer the following questions:
1. What is the program doing?
2. What does a set do and how is it used above?
3. Where would you be more likely to use a set versus using a dictionary, list, or tuple?
4. How are sets different from dictionaries, tuples, and lists?
## 1.4 Data Structures (Group):
Create a program that does the following:
1. Take the two lists provided and count the frequency of each word in each list.
2. Create a dictionary with the key equal to each unique word and count as the value.
3. Print off a list of words that are in both lists.
4. Print off a list of words that are in the first list and not the second.
5. Print off a list of words that are in the second list and not in the first.
```
list1 = ['the', 'quick', 'brown', 'fox', 'jumps', 'over', 'the', 'lazy', 'dog']
list2 = ['jack', 'be', 'nimble', 'jack', 'be', 'quick', 'jack', 'jumps', 'over', 'the', 'candlestick']
def count_character(list_):
dictlist = {}
for words in list_:
if words is not in dictlist:
dictlist[words]=1
else:
dictlist[words]+=1
dict1 = count_character(list1)
dict2 = count_character(list2)
set1 = set(list1)
set2 = set(list2)
print('The words in both list:')
for word in set1.intersection(set2):
print(word)
print('words that are in the first list:')
for word in set1.difference(set2)
print(word)
print('words that are in the second list:')
for word in set2.difference(set1)
print(word)
```
| github_jupyter |
### Dependences
```
import sys
sys.path.append("../")
import math
from tqdm import tqdm
import numpy as np
import tensorflow as tf
from PIL import Image
from tqdm import tqdm
import matplotlib.pyplot as plt
from IPython.display import clear_output
from lib.models.LinkNet import LinkNet
import lib.utils as utils
import IPython.display as ipd
```
### Loading experiment data
```
#set experiment ID
EXP_ID = "LinkNet"
utils.create_experiment_folders(EXP_ID)
utils.load_experiment_data()
```
### Model instantiation
```
model = LinkNet()
model.build((None,128,128,1))
print(model.summary())
```
### Loading Dataset
```
train_x = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/X_train.npy", mmap_mode='c')
train_y = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/y_train.npy", mmap_mode='c')
qtd_traning = train_x.shape
print("Loaded",qtd_traning, "samples")
valid_x_1 = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/X_val.npy", mmap_mode='c')
valid_y_1 = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/y_val.npy", mmap_mode='c')
qtd_traning = valid_x_1.shape
print("Loaded",qtd_traning, "samples")
```
### Dataset Normalization and Batches split
```
value = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/scale_and_shift.npy", mmap_mode='c')
print(value)
SHIFT_VALUE_X, SHIFT_VALUE_Y, SCALE_VALUE_X, SCALE_VALUE_Y = value[0], value[1], value[2], value[3]
# SHIFT_VALUE_X, SHIFT_VALUE_Y, SCALE_VALUE_X, SCALE_VALUE_Y = utils.get_shift_scale_maxmin(train_x, train_y, valid_x_1, valid_y_1)
mini_batch_size = 58
num_train_minibatches = math.floor(train_x.shape[0]/mini_batch_size)
num_val_minibatches = math.floor(valid_x_1.shape[0]/mini_batch_size)
print("train_batches:", num_train_minibatches, "valid_batches:", num_val_minibatches)
```
### Metrics
```
#default tf.keras metrics
train_loss = tf.keras.metrics.Mean(name='train_loss')
```
### Set Loss and load model weights
```
loss_object = tf.keras.losses.MeanSquaredError()
optimizer = tf.keras.optimizers.Adam(learning_rate=0.001)
#get last saved epoch index and best result in validation step
CURRENT_EPOCH, BEST_VALIDATION = utils.get_model_last_data()
if CURRENT_EPOCH > 0:
print("Loading last model state in epoch", CURRENT_EPOCH)
model.load_weights(utils.get_exp_folder_last_epoch())
print("Best validation result was PSNR=", BEST_VALIDATION)
```
### Training
```
@tf.function
def train_step(patch_x, patch_y):
with tf.GradientTape() as tape:
predictions = model(patch_x)
loss = loss_object(patch_y, predictions)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
train_loss(loss)
def valid_step(valid_x, valid_y, num_val_minibatches, mini_batch_size):
valid_mse = tf.keras.metrics.MeanSquaredError(name='train_mse')
valid_custom_metrics = utils.CustomMetric()
for i in tqdm(range(num_val_minibatches)):
data_x = valid_x[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_y = valid_y[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_x = tf.convert_to_tensor(data_x, dtype=tf.float32)
data_y = tf.convert_to_tensor(data_y, dtype=tf.float32)
data_x = ((data_x+SHIFT_VALUE_X)/SCALE_VALUE_X)+CONST_GAMA
data_y = ((data_y+SHIFT_VALUE_Y)/SCALE_VALUE_Y)+CONST_GAMA
predictions = model(data_x)
valid_mse(data_y, predictions)
predictions = predictions.numpy()
data_y = data_y.numpy()
#feed the metric evaluator
valid_custom_metrics.feed(data_y, predictions)
#get metric results
psnr, nrmse = valid_custom_metrics.result()
valid_mse_result = valid_mse.result().numpy()
valid_custom_metrics.reset_states()
valid_mse.reset_states()
return psnr, nrmse, valid_mse_result
MAX_EPOCHS = 100
EVAL_STEP = 1
CONST_GAMA = 0.001
for epoch in range(CURRENT_EPOCH, MAX_EPOCHS):
#TRAINING
print("TRAINING EPOCH", epoch)
for k in tqdm(range(0, num_train_minibatches)):
seismic_x = train_x[k * mini_batch_size : k * mini_batch_size + mini_batch_size]
seismic_y = train_y[k * mini_batch_size : k * mini_batch_size + mini_batch_size]
seismic_x = tf.convert_to_tensor(seismic_x, dtype=tf.float32)
seismic_y = tf.convert_to_tensor(seismic_y, dtype=tf.float32)
seismic_x = ((seismic_x+SHIFT_VALUE_X)/SCALE_VALUE_X)+CONST_GAMA
seismic_y = ((seismic_y+SHIFT_VALUE_Y)/SCALE_VALUE_Y)+CONST_GAMA
train_step(seismic_x, seismic_y)
#VALIDATION
if epoch%EVAL_STEP == 0:
clear_output()
print("VALIDATION EPOCH", epoch)
#saving last epoch model
model.save_weights(utils.get_exp_folder_last_epoch(), save_format='tf')
#valid with set 1
print("Validation set")
psnr_1, nrmse_1, mse_1 = valid_step(valid_x_1, valid_y_1, num_val_minibatches, mini_batch_size)
#valid with set 2
#print("Validation set 2")
#psnr_2, nrmse_2, mse_2 = valid_step(valid_x_2, valid_y_2, num_val_minibatches, mini_batch_size)
psnr_2, nrmse_2, mse_2 = 0, 0, 0
#valid with set 3
#print("Validation set 3")
#psnr_3, nrmse_3, mse_3 = valid_step(valid_x_3, valid_y_3, num_val_minibatches, mini_batch_size)
psnr_3, nrmse_3, mse_3 = 0, 0, 0
utils.update_chart_data(epoch=epoch, train_mse=train_loss.result().numpy(),
valid_mse=[mse_1,mse_2,mse_3], psnr=[psnr_1,psnr_2,psnr_3], nrmse=[nrmse_1,nrmse_2, nrmse_3])
utils.draw_chart()
#saving best validation model
if psnr_1 > BEST_VALIDATION:
BEST_VALIDATION = psnr_1
model.save_weights(utils.get_exp_folder_best_valid(), save_format='tf')
train_loss.reset_states()
utils.draw_chart()
#experimentos results
print(utils.get_experiment_results())
#load best model
model.load_weights(utils.get_exp_folder_best_valid())
CONST_GAMA = 0.001
# valid_x_1 = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/X_val.npy", mmap_mode='c')
# valid_y_1 = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/y_val.npy", mmap_mode='c')
qtd_traning = valid_x_1.shape
print("Loaded",qtd_traning, "samples")
# #normalization
# test_x = utils.shift_and_normalize(test_x, SHIFT_VALUE_X, SCALE_VALUE_X)
# test_y = utils.shift_and_normalize(test_y, SHIFT_VALUE_Y, SCALE_VALUE_Y)
#batches
num_val_minibatches = math.floor(valid_x_1.shape[0]/mini_batch_size)
# test_batches = utils.random_mini_batches(test_x, test_y, None, None, 8, seed=0)
#metrics
val_mse = tf.keras.metrics.MeanSquaredError(name='val_mse')
val_custom_metrics = utils.CustomMetric()
import json
f = open('/home/arthursrr/Documentos/Audio_Inpainting/Datasets/idx_genders_val.json', "r")
idx_gen = json.loads(f.read())
for k in idx_gen:
for i in tqdm(idx_gen[k]):
data_x = valid_x_1[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_y = valid_y_1[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_x = tf.convert_to_tensor(data_x, dtype=tf.float32)
data_y = tf.convert_to_tensor(data_y, dtype=tf.float32)
data_x = ((data_x+SHIFT_VALUE_X)/SCALE_VALUE_X)+CONST_GAMA
data_y = ((data_y+SHIFT_VALUE_Y)/SCALE_VALUE_Y)+CONST_GAMA
predictions = model(data_x)
val_mse(data_y, predictions)
predictions = predictions.numpy()
data_y = data_y.numpy()
#feed the metric evaluator
val_custom_metrics.feed(data_y, predictions)
#get metric results
psnr, nrmse = val_custom_metrics.result()
val_mse_result = val_mse.result().numpy()
val_custom_metrics.reset_states()
val_mse.reset_states()
print(k ,"\nPSNR:", psnr,"\nNRMSE:", nrmse)
# Closing file
f.close()
```
## Test
```
#load best model
model.load_weights(utils.get_exp_folder_best_valid())
CONST_GAMA = 0.001
test_x = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/X_test.npy", mmap_mode='c')
test_y = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/y_test.npy", mmap_mode='c')
qtd_traning = test_x.shape
print("Loaded",qtd_traning, "samples")
# #normalization
# test_x = utils.shift_and_normalize(test_x, SHIFT_VALUE_X, SCALE_VALUE_X)
# test_y = utils.shift_and_normalize(test_y, SHIFT_VALUE_Y, SCALE_VALUE_Y)
#batches
num_test_minibatches = math.floor(test_x.shape[0]/mini_batch_size)
# test_batches = utils.random_mini_batches(test_x, test_y, None, None, 8, seed=0)
#metrics
test_mse = tf.keras.metrics.MeanSquaredError(name='train_mse')
test_custom_metrics = utils.CustomMetric()
#test
for i in tqdm(range(num_test_minibatches)):
data_x = test_x[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_y = test_y[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_x = tf.convert_to_tensor(data_x, dtype=tf.float32)
data_y = tf.convert_to_tensor(data_y, dtype=tf.float32)
data_x = ((data_x+SHIFT_VALUE_X)/SCALE_VALUE_X)+CONST_GAMA
data_y = ((data_y+SHIFT_VALUE_Y)/SCALE_VALUE_Y)+CONST_GAMA
predictions = model(data_x)
test_mse(data_y, predictions)
predictions = predictions.numpy()
data_y = data_y.numpy()
#feed the metric evaluator
test_custom_metrics.feed(data_y, predictions)
#just show the first example of each batch until 5
# print("Spatial domain: X - Y - PREDICT - DIFF")
# plt.imshow(np.hstack((data_x[0,:,:,0], data_y[0,:,:,0], predictions[0,:,:,0], np.abs(predictions[0,:,:,0]-seismic_y[0,:,:,0]))) , cmap='Spectral', vmin=0, vmax=1)
# plt.axis('off')
# plt.pause(0.1)
#ATENÇÃO!!
#predictions = inv_shift_and_normalize(predictions, SHIFT_VALUE_Y, SCALE_VALUE_Y)
#np.save(outfile_path, predictions)
#get metric results
psnr, nrmse = test_custom_metrics.result()
test_mse_result = test_mse.result().numpy()
test_custom_metrics.reset_states()
test_mse.reset_states()
print("PSNR:", psnr,"\nNRMSE", nrmse)
#load best model
model.load_weights(utils.get_exp_folder_best_valid())
CONST_GAMA = 0.001
test_x = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/X_test.npy", mmap_mode='c')
test_y = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/y_test.npy", mmap_mode='c')
qtd_traning = test_x.shape
print("Loaded",qtd_traning, "samples")
# #normalization
# test_x = utils.shift_and_normalize(test_x, SHIFT_VALUE_X, SCALE_VALUE_X)
# test_y = utils.shift_and_normalize(test_y, SHIFT_VALUE_Y, SCALE_VALUE_Y)
#batches
num_test_minibatches = math.floor(test_x.shape[0]/mini_batch_size)
# test_batches = utils.random_mini_batches(test_x, test_y, None, None, 8, seed=0)
#metrics
test_mse = tf.keras.metrics.MeanSquaredError(name='test_mse')
test_custom_metrics = utils.CustomMetric()
import json
f = open('/home/arthursrr/Documentos/Audio_Inpainting/Datasets/idx_genders_test.json', "r")
idx_gen = json.loads(f.read())
for k in idx_gen:
for i in tqdm(idx_gen[k]):
data_x = test_x[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_y = test_y[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_x = tf.convert_to_tensor(data_x, dtype=tf.float32)
data_y = tf.convert_to_tensor(data_y, dtype=tf.float32)
data_x = ((data_x+SHIFT_VALUE_X)/SCALE_VALUE_X)+CONST_GAMA
data_y = ((data_y+SHIFT_VALUE_Y)/SCALE_VALUE_Y)+CONST_GAMA
predictions = model(data_x)
test_mse(data_y, predictions)
predictions = predictions.numpy()
data_y = data_y.numpy()
#feed the metric evaluator
test_custom_metrics.feed(data_y, predictions)
#get metric results
psnr, nrmse = test_custom_metrics.result()
test_mse_result = test_mse.result().numpy()
test_custom_metrics.reset_states()
test_mse.reset_states()
print(k ,"\nPSNR:", psnr,"\nNRMSE:", nrmse)
# Closing file
f.close()
def griffin_lim(S, frame_length=256, fft_length=255, stride=64):
'''
TensorFlow implementation of Griffin-Lim
Based on https://github.com/Kyubyong/tensorflow-exercises/blob/master/Audio_Processing.ipynb
'''
S = tf.expand_dims(S, 0)
S_complex = tf.identity(tf.cast(S, dtype=tf.complex64))
y = tf.signal.inverse_stft(S_complex, frame_length, stride, fft_length=fft_length)
for i in range(1000):
est = tf.signal.stft(y, frame_length, stride, fft_length=fft_length)
angles = est / tf.cast(tf.maximum(1e-16, tf.abs(est)), tf.complex64)
y = tf.signal.inverse_stft(S_complex * angles, frame_length, stride, fft_length=fft_length)
return tf.squeeze(y, 0)
model.load_weights(utils.get_exp_folder_best_valid())
test_x = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/X_test.npy", mmap_mode='c')
test_y = np.load("/mnt/backup/arthur/Free_Music_Archive/Spectrogramas/y_test.npy", mmap_mode='c')
qtd_traning = test_x.shape
print("Loaded",qtd_traning, "samples")
#batches
num_test_minibatches = math.floor(test_x.shape[0]/mini_batch_size)
#metrics
test_mse = tf.keras.metrics.MeanSquaredError(name='test_mse')
test_custom_metrics = utils.CustomMetric()
i=5000
CONST_GAMA = 0.001
data_x = test_x[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_y = test_y[i * mini_batch_size : i * mini_batch_size + mini_batch_size]
data_x = tf.convert_to_tensor(data_x, dtype=tf.float32)
data_norm = ((data_x+SHIFT_VALUE_X)/SCALE_VALUE_X)+CONST_GAMA
predictions = model(data_norm)
predictions = utils.inv_shift_and_normalize(predictions, SHIFT_VALUE_Y, SCALE_VALUE_Y)
predictions
audio_pred = None
for i in range(0, 58):
if i==0:
audio_pred = predictions[i,:,:,0]
else:
audio_pred = np.concatenate((audio_pred, predictions[i,:,:,0]), axis=0)
audio_pred.shape
audio_corte = None
for i in range(0, 58):
if i==0:
audio_corte = data_x[i,:,:,0]
else:
audio_corte = np.concatenate((audio_corte, data_x[i,:,:,0]), axis=0)
audio_corte.shape
audio_original = None
for i in range(0, 58):
if i==0:
audio_original = data_y[i,:,:,0]
else:
audio_original = np.concatenate((audio_original, data_y[i,:,:,0]), axis=0)
audio_original.shape
wave_original = griffin_lim(audio_original, frame_length=256, fft_length=255, stride=64)
ipd.Audio(wave_original, rate=16000)
wave_corte = griffin_lim(audio_corte, frame_length=256, fft_length=255, stride=64)
ipd.Audio(wave_corte, rate=16000)
wave_pred = griffin_lim(audio_pred, frame_length=256, fft_length=255, stride=64)
ipd.Audio(wave_pred, rate=16000)
import soundfile as sf
sf.write('x.wav', wave_corte, 16000, subtype='PCM_16')
sf.write('pred.wav', wave_pred, 16000, subtype='PCM_16')
```
| github_jupyter |
# Evaluation with JustCause
In this notebook, we examplify how to use JustCause in order to evaluate methods using reference datasets. For simplicity, we only use one dataset, but show how evaluation works with multiple methods. Both standard causal methods implemented in the framework as well as custom methods.
## Custom First
The goal of the JustCause framework is to be a modular and flexible facilitator of causal evaluation.
```
%load_ext autoreload
%autoreload 2
# Loading all required packages
import itertools
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from justcause.data import Col
from justcause.data.sets import load_ihdp
from justcause.metrics import pehe_score, mean_absolute
from justcause.evaluation import calc_scores, summarize_scores
from sklearn.linear_model import LinearRegression
```
### Setup data and methods you want to evaluate
Let's say we wanted to compare a S-Learner with propensity weighting, based on a propensity estimate of our choice. Thus, we cannot simply use the predefined SLearner from `justcause.learners`, but have to provide our own adaption, which first estimates propensities and uses these for fitting an adjusted model.
By providing a "blackbox" method like below, you can choose to do whatever you want inside. For example, you can replace your predictions available factual outcomes, estimate the propensity in different ways or even use a true propensity, in case of a generated dataset, where it is available. You can also resort to out-of-sample prediction, where no information about treatment is provided to the method.
```
from justcause.learners import SLearner
from justcause.learners.propensity import estimate_propensities
# Get the first 100 replications
replications = load_ihdp(select_rep=np.arange(100))
metrics = [pehe_score, mean_absolute]
train_size = 0.8
random_state = 42
def weighted_slearner(train, test):
"""
Custom method that takes 'train' and 'test' CausalFrames (see causal_frames.ipynb)
and returns ITE predictions for both after training on 'train'.
Implement your own method in a similar fashion to evaluate them within the framework!
"""
train_X, train_t, train_y = train.np.X, train.np.t, train.np.y
test_X, test_t, test_y = test.np.X, test.np.t, test.np.y
# Get calibrated propensity estimates
p = estimate_propensities(train_X, train_t)
# Make sure the supplied learner is able to use `sample_weights` in the fit() method
slearner = SLearner(LinearRegression())
# Weight with inverse probability of treatment (inverse propensity)
slearner.fit(train_X, train_t, train_y, weights=1/p)
return (
slearner.predict_ite(train_X, train_t, train_y),
slearner.predict_ite(test_X, test_t, test_y)
)
```
### Example Evaluation Loop
Now given a callable like `weighted_slearner` we can evaluate that method using multiple metrics on the given replications.
The result dataframe then contains two rows with the summarized scores over all replications for train and test separately.
```
results_df = list()
test_scores = list()
train_scores = list()
for rep in replications:
train, test = train_test_split(
rep, train_size=train_size, random_state=random_state
)
# REPLACE this with the function you implemented and want to evaluate
train_ite, test_ite = weighted_slearner(train, test)
# Calculate the scores and append them to a dataframe
train_scores.append(calc_scores(train[Col.ite], train_ite, metrics))
test_scores.append(calc_scores(test[Col.ite], test_ite, metrics))
# Summarize the scores and save them in a dataframe
train_result, test_result = summarize_scores(train_scores), summarize_scores(test_scores)
train_result.update({'method': 'weighted_slearner', 'train': True})
test_result.update({'method': 'weighted_slearner', 'train': False})
pd.DataFrame([train_result, test_result])
```
Now in this case, using `justcause` has hardly any advantages, because only one dataset and one method is used. You might as well just implement all the evaluation manually. However, this can simply be expanded to more methods by looping over the callables.
```
def basic_slearner(train, test):
""" """
train_X, train_t, train_y = train.np.X, train.np.t, train.np.y
test_X, test_t, test_y = test.np.X, test.np.t, test.np.y
slearner = SLearner(LinearRegression())
slearner.fit(train_X, train_t, train_y)
return (
slearner.predict_ite(train_X, train_t, train_y),
slearner.predict_ite(test_X, test_t, test_y)
)
methods = [basic_slearner, weighted_slearner]
results = list()
for method in methods:
test_scores = list()
train_scores = list()
for rep in replications:
train, test = train_test_split(
rep, train_size=train_size, random_state=random_state
)
# REPLACE this with the function you implemented and want to evaluate
train_ite, test_ite = method(train, test)
# Calculate the scores and append them to a dataframe
test_scores.append(calc_scores(test[Col.ite], test_ite, metrics))
train_scores.append(calc_scores(train[Col.ite], train_ite, metrics))
# Summarize the scores and save them in a dataframe
train_result, test_result = summarize_scores(train_scores), summarize_scores(test_scores)
train_result.update({'method': method.__name__, 'train': True})
test_result.update({'method': method.__name__, 'train': False})
results.append(train_result)
results.append(test_result)
# For visualization
pd.DataFrame(results)
```
And because in most cases, we're not changing anything within this loop for the ITE case, `justcause` provides a default implementation.
## Standard Evaluation of ITE predictions
Using the same list of method callables, we can just call `evaluate_ite` and pass all the information. The default implementation sets up a dataframe for the result following a certain convention.
First, there's two columns to define the method for which the results are as well as whether they've been calculated on train or test. Then for all supplied `metrics`, all `formats` will be listed.
Standard `metrics` like (PEHE or Mean absolute error) are implemented in `justcause.metrics`.
Standard formats used for summarizing the scores over multiple replications are `np.mean, np.median, np.std`, other possibly interesting formats could be *skewness*, *minmax*, *kurtosis*. A method provided as format must take an `axis` parameter, ensuring that it can be applied to the scores dataframe.
```
from justcause.evaluation import evaluate_ite
result = evaluate_ite(replications, methods, metrics, train_size=train_size, random_state=random_state)
pd.DataFrame(result)
```
### Adding standard causal methods to the mix
Within `justcause.learners` we've implemented a couple of standard methods that provide a `predict_ite()` method. Instead of going the tedious way like we've done in `weighted_slearner` above, we can just use these methods directly. The default implementation will use a default base learner for all the meta-learners, fit the method on train and predict the ITEs for train and test.
By doing so, we can get rid of the `basic_slearner` method above, because it just uses the default setting and procedure for fitting the model. Instead, we just use `SLearner(LinearRegression())`.
```
from justcause.learners import TLearner, XLearner, RLearner
# All in standard configuration
methods = [SLearner(LinearRegression()), weighted_slearner, TLearner(), XLearner(), RLearner(LinearRegression())]
result = evaluate_ite(replications, methods, metrics, train_size=train_size, random_state=random_state)
pd.DataFrame(result)
```
| github_jupyter |
# README
# Ref
- https://www.ieee-security.org/TC/SPW2019/DLS/doc/06-Marin.pdf
- https://www.stratosphereips.org/datasets-normal
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
%precision 4
%reload_ext autoreload
import re
import sys
import math
import random
import datetime
import numpy as np
from matplotlib import pyplot as plt
import pandas as pd
import seaborn as sns
from IPython.core.interactiveshell import InteractiveShell
import warnings
warnings.filterwarnings(action='once')
InteractiveShell.ast_node_interactivity = "all"
pd.set_option('display.max_colwidth', -1)
```
# Feature Engineering
```
from scapy.all import *
load_layer('tls')
import os
import shutil
import time
counter = 0
shape = (100, 2)
shape_size = 100 * 2
x = []
y = []
INPUTS = ['../input/botnet-capture-20110810-neris.pcap',
'../input/2013-12-17_capture1.pcap',
'../input/botnet-capture-20110815-rbot-dos.pcap']
botnet_pcap = rdpcap(INPUTS[0])
x = []
total_packets = 0
for p in botnet_pcap:
total_packets += 1
try:
tls = p[TCP][TLS]
if len(tls.payload) != 0:
k = tls.__str__().hex()
n = 2
t = [int(k[i:i+n], 16) for i in range(0, len(k), n)]
t = np.array(t)
# print("before resize", t.shape)
# print("len t", len(t))
t.resize( shape_size * math.ceil(len(t) / shape_size) )
# print("after resize", t.shape)
t = np.reshape(t, (-1, 100, 2))
# print(t)
for i in t:
x.append(i)
except IndexError as e:
if p.haslayer(TCP):
print(p[TCP].layers())
pass
total_packets
df = pd.DataFrame( np.array(x).reshape(-1, 200) )
df[df.duplicated()]
s = df.sum(axis=1)
s[s == 0]
normal_pcap = rdpcap(INPUTS[1])
n_x = []
for p in normal_pcap:
try:
tls = p[TCP][TLS]
if len(tls.payload) != 0:
k = tls.__str__().hex()
n = 2
t = [int(k[i:i+n], 16) for i in range(0, len(k), n)]
t = np.array(t)
print("before resize", t.shape)
print("len t", len(t))
t.resize( shape_size * math.ceil(len(t) / shape_size) )
print("after resize", t.shape)
t = np.reshape(t, (-1, 100, 2))
print(t)
for i in t:
n_x.append(i)
except IndexError as e:
pass
x = np.array(x)
y = np.array([1 for i in range(len(x))])
len(x), len(y)
n_x = np.array(n_x)
n_y = np.array([0 for i in range(len(n_x))])
len(n_x), len(n_y)
x = np.concatenate((x, n_x), axis=0)
y = np.concatenate((y, n_y), axis=0)
```
# Modeling
```
from __future__ import print_function
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, BatchNormalization
from keras.layers import Conv2D, MaxPooling2D, Conv1D, MaxPooling1D
from keras import backend as K
batch_size = 128
num_classes = 2
epochs = 50
# x_train, y_train = x_train, y_train
# x_test, y_test = x_train, y_train
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(
x, y, test_size=0.25, random_state=42)
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
from sklearn.utils import shuffle
x_train, y_train = shuffle(x_train, y_train, random_state=0)
x_train.shape
y_train.shape
y_train
model = Sequential()
model.add(BatchNormalization())
model.add(Conv1D(32, kernel_size=5,
activation='relu',
input_shape=(100, 2)))
# model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling1D(pool_size=(3)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(50, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(100, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.binary_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=["categorical_accuracy"])
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
model.summary()
```
### without dropout
Test loss: 0.28140440133901745
Test accuracy: 0.9353846311569214
### with dropout
Test loss: 0.27307736470149113
Test accuracy: 0.926153838634491
| github_jupyter |
This notebook walks you through on how to get a geojson shape plotted in a 2.5D view, with an overlayed satellite image on top. <br/>
For this, besides the notebook you will need this github project:<br/>
https://github.com/zhunor/threejs-dem-visualizer <br/>
It has to be cloned IN the same folder as this notebook. Afterwards install the project as follows: <br/>
From a command line: <br/>
cd threejs-dem-visualizer <br/>
yarn install <br/>
yarn build <br/>
After this is done go one with the steps in this notebook
```
import requests
from PIL import Image
import numpy as np
import matplotlib.pyplot as plt
import LMIPy
%matplotlib inline
```
To get your desired shape, go to http://geojson.io, draw a shape and copy the coordinates starting from the first square bracket '[' until the last one ']' and replace them here in the 'coordinates' attribute.
```
atts={'geojson': {'type': 'FeatureCollection',
'features': [{'type': 'Feature',
'properties': {},
'geometry': {'type': 'Polygon',
'coordinates': [
[
[
-3.5976791381835933,
40.86030420568381
],
[
-3.6038589477539062,
40.89872234775293
],
[
-3.6694335937500004,
40.886264852994955
],
[
-3.654670715332031,
40.84654093547386
],
[
-3.5976791381835933,
40.86030420568381
]
]
]
}}]}}
#g = LMIPy.Geometry(attributes=atts, server='http://localhost:9000')
g = LMIPy.Geometry(attributes=atts, server='https://staging-api.globalforestwatch.org')
#r = requests.get(f"http://localhost:9000/v1/composite-service?geostore={g.id}&get_dem=True")
r = requests.get(f"https://staging-api.globalforestwatch.org/v1/composite-service?geostore={g.id}&get_dem=True&date_range=2016-01-01, 2017-01-01")
r.json()
dem_url = r.json().get('attributes').get('dem')
surface_url = r.json().get('attributes').get('thumb_url')
dem = Image.open(requests.get(dem_url, stream=True).raw)
dem
#%cd ../..
surface = Image.open(requests.get(surface_url, stream=True).raw)
surface.save("threejs-dem-visualizer/src/textures/surface.png")
surface
dem_array = np.array(dem)
d = dem_array[:,:,0]
d.min(), d.max(), np.unique(d), np.array(d).shape
#plt.imshow(d)
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
from matplotlib._png import read_png
from pylab import *
from matplotlib.cbook import get_sample_data
fig = plt.figure(figsize=(15,15))
ax = fig.gca(projection='3d')
X = np.arange(d.shape[0])
Y = np.arange(d.shape[1])
X, Y = np.meshgrid(X, Y)
# Plot the surface.
surf = ax.plot_surface(X, Y, d, cmap=cm.coolwarm,
linewidth=0, antialiased=False)
# Customize the z axis.
# ax.set_zlim(-1.01, 1.01)
# ax.zaxis.set_major_locator(LinearLocator(10))
# ax.zaxis.set_major_formatter(FormatStrFormatter('%.02f'))
ax.zaxis.set_label_text('height above sea level (m)')
import rasterio
import rasterio.features
import rasterio.warp
import matplotlib.pyplot as plt
def open_tif(file_name):
with rasterio.open(file_name) as dataset:
# Read the dataset's valid data mask as a ndarray.
mask = dataset.dataset_mask()
# Extract feature shapes and values from the array.
for geom, val in rasterio.features.shapes(
mask, transform=dataset.transform):
# Transform shapes from the dataset's own coordinate
# reference system to CRS84 (EPSG:4326).
geom = rasterio.warp.transform_geom(
dataset.crs, 'EPSG:4326', geom, precision=6)
# Print GeoJSON shapes to stdout.
print(geom)
return dataset
#dataset = open_tif('threejs-dem-visualizer/src/textures/agri-small-dem.tif')
#open_tif('new.tif')
```
Loading the 'agri-small-dem.tif' file just to have working sample profile which is replicated
```
agri = rasterio.open('threejs-dem-visualizer/src/textures/agri-small-dem.tif')
agri_ar = agri.read(1)
# plt.imshow(agri_ar, cmap='pink')
# plt.show()
profile = agri.profile
```
Adjust the scaling factor to your liking, based on what looks right. A factor of one will leave shapes looking too flat, whereas 50 will be too much for some montaneous areas.
```
scaling_factor = 7
d2 = np.dot(d, scaling_factor)
#d2 = d + 180
with rasterio.open('threejs-dem-visualizer/src/textures/testout.tif', 'w', **agri.profile) as dst:
dst.write(d2.astype(profile['dtype']), 1)
testout = rasterio.open('threejs-dem-visualizer/src/textures/testout.tif', **profile)
testout_ar = testout.read(1)
#testout.crs = agri.crs
#testout.crs = 'EPSG:4326'
#agri_ar.crs
plt.imshow(testout_ar, cmap='pink')
plt.show()
```
## One more thing before you run the app:
You have to go in the three-js-dem-visualizer/src/js folder, open the index.js file and comment out these two lines: <br/>
//import * as terrain from "../textures/agri-small-dem.tif"; <br/>
//import * as mountainImage from "../textures/agri-small-autumn.jpg"; <br/>
### and copy the following 2 lines next
import * as terrain from "../textures/testout.tif"; <br/>
import * as mountainImage from "../textures/surface.png"; <br/>
Now the next line will run the app
```
#%cd threejs-dem-visualizer/src/
!yarn --cwd threejs-dem-visualizer/src/ dev
```
| github_jupyter |
*Donald Knuth: "Premature optimization is the root of all evil"*
**Оригинал**: https://ipython-books.github.io/chapter-5-high-performance-computing/
- Компиляция [Just-In-Time (JIT)](https://ru.wikipedia.org/wiki/JIT-%D0%BA%D0%BE%D0%BC%D0%BF%D0%B8%D0%BB%D1%8F%D1%86%D0%B8%D1%8F) кода Python.
- Использование языка более низкого уровня, такого как C, из Python.
- Распределение задач между несколькими вычислительными блоками с использованием параллельных вычислений.
### Компиляция Just-In-Time (JIT) кода Python
Благодаря JIT код Python динамически компилируется в язык более низкого уровня. Компиляция происходит во время выполнения, а не перед выполнением. Переведенный код работает быстрее, поскольку он компилируется, а не интерпретируется. JIT-компиляция является популярной техникой, поскольку она может привести к быстрым и высокоуровневым языкам, в то время как эти две характеристики раньше были взаимоисключающими.
Методы JIT-компиляции реализованы в таком пакете, как **Numba**.
PyPy ([официальный сайт](http://pypy.org) и [блог](https://morepypy.blogspot.com/)) - альтернативная реализация Python (эталонная реализация CPython) включает JIT-компилятор.
PyPy состоит из стандартного интерпретатора и транслятора.
Интерпретатор полностью реализует язык Python. Сам интерпретатор написан на ограниченном подмножестве этого же языка, называемом RPython (Restricted Python). В отличие от стандартного Python, [RPython](https://rpython.readthedocs.io/en/latest/architecture.html) является статически типизированным для более эффективной компиляции.
Транслятор является набором инструментов, который анализирует код RPython и переводит его в языки более низкого уровня, такие как C, байт-код Java или CIL. Он также поддерживает подключаемые сборщики мусора и позволяет опционально включать [Stackless](https://ru.wikipedia.org/wiki/Stackless_Python). Также он включает JIT-компилятор для трансляции кода в машинные инструкции во время исполнения программы.
### Использование языка более низкого уровня
Использование языка более низкого уровня, такого как C, является еще одним интересным методом. Популярные библиотеки включают в себя ctypes и Cython. Использование ctypes требует написания кода на C и наличия доступа к компилятору C или использования скомпилированной библиотеки C. В отличие от этого, Cython позволяет писать код в расширенном наборе Python, который переводится в C с различными результатами производительности.
### CPython and concurrent programming
Основной реализацией языка Python является CPython, написанный на C. CPython интегрирует механизм, называемый Global Interpreter Lock (GIL). Как упоминалось на http://wiki.python.org/moin/GlobalInterpreterLock:
«GIL облегчает управление памятью, предотвращая одновременное выполнение байт-кодом Python нескольких собственных потоков».
Другими словами, отключая параллельные потоки в одном процессе Python, GIL значительно упрощает систему управления памятью. Поэтому управление памятью не является поточно-ориентированным в CPython.
Важным выводом является то, что CPython делает нетривиальным использование нескольких процессоров в одном процессе Python. Это важная проблема, поскольку современные процессоры содержат все больше и больше ядер.
Какие возможные решения у нас есть, чтобы воспользоваться преимуществами многоядерных процессоров?
- Удаление GIL в CPython. Это решение было опробовано, но никогда не входило в CPython. Это приведет к слишком большой сложности при реализации CPython и ухудшит производительность однопоточных программ.
- Использование нескольких процессов вместо нескольких потоков. Это популярное решение; это можно сделать с помощью собственного многопроцессорного модуля или с помощью IPython.
- Переписать определенные части кода на Cython и заменить все переменные Python переменными C. Это позволяет временно удалить GIL в цикле, что позволяет использовать многоядерные процессоры.
- Реализация определенной части кода на языке, который предлагает лучшую поддержку для многоядерных процессоров, и вызов его из вашей программы Python.
- При создании кода используйте функции NumPy, которые получают преимущества от многоядерных процессоров, таких как numpy.dot (). NumPy необходимо скомпилировать с помощью BLAS / LAPACK / ATLAS / MKL.
Обязательную для прочтения ссылку на GIL можно найти по адресу http://www.dabeaz.com/GIL/.
### Инструкции по установке, связанные с компилятором
В Linux необходимо установить gcc. Например, в Ubuntu введите sudo apt-get install build-essential в терминале.
В macOS установите Xcode или Инструменты командной строки Xcode. Или введите gcc в терминале. Если он не установлен, macOS предоставит вам несколько вариантов его установки.
В Windows установите версию Microsoft Visual Studio, Visual C++ или Visual C++ Build Tools, соответствующую вашей версии Python. Если вы используете Python 3.6, соответствующая версия компилятора Microsoft - 2017. Все эти программы бесплатны или имеют бесплатную версию, достаточную для Python.
Вот несколько ссылок:
- Документация по установке Cython по адресу http://cython.readthedocs.io/en/latest/src/quickstart/install.html.
- Компиляторы Windows для Python, по адресу https://wiki.python.org/moin/WindowsCompilers
- Microsoft Visual Studio можно загрузить по адресу https://www.visualstudio.com/downloads/.
### Знание Python для написания более быстрого кода
Первый способ заставить код Python работать быстрее - это знать все возможности языка. Python содержит множество синтаксических функций и модулей в стандартной библиотеке, которые работают намного быстрее, чем все, что вы можете написать вручную. Более того, хотя Python может работать медленно, если вы пишете на Python, как если бы вы писали на C или Java, он часто бывает достаточно быстрым, когда вы пишете Python-код.
В этом разделе мы покажем, как плохо написанный код Python может быть значительно улучшен при использовании всех возможностей языка.
`
Помните про использование NumPy для эффективных операций с массивами
`
1. Давайте определим список нормально распределенных случайных величин, используя вместо NumPy встроенный случайный модуль.
```
import random
l = [random.normalvariate(0, 1) for i in range(100000)]
```
2. Давайте напишем функцию, которая вычисляет сумму всех чисел в этом списке. Кто-то неопытный с Python может написать на Python, как если бы это был C, который дал бы следующую функцию:
```
def sum1():
# BAD: not Pythonic and slow
res = 0
for i in range(len(l)):
res = res + l[i]
return res
sum1()
%timeit sum1()
```
5 миллисекунд для вычисления суммы «только» 100 000 чисел являются медленными, что может заставить некоторых людей довольно несправедливо сказать, что «Python медленный».
3. Теперь давайте напишем слегка улучшенную версию этого кода, учитывая тот факт, что мы можем перечислять элементы списка, используя `for x in l` вместо итерации с индексом:
```
def sum2():
# STILL BAD
res = 0
for x in l:
res = res + x
return res
sum2()
%timeit sum2()
```
Эта небольшая модификация дала нам почти двукратное улучшение скорости.
Наконец, мы помним, что в Python есть встроенная функция для вычисления суммы всех элементов в списке:
```
def sum3():
# GOOD
return sum(l)
sum3()
%timeit sum3()
```
Эта версия в 17 раз быстрее первой, и мы написали только чистый код на Python!
4. Давайте перейдем к другому примеру со строками. Мы создаем список строк, представляющих все числа в нашем предыдущем списке:
```
strings = ['%.3f' % x for x in l]
strings[:3]
```
5. Мы определяем функцию, объединяющую все строки в этом списке. Опять же, неопытный программист на Python может написать такой код:
```
def concat1():
# BAD: not Pythonic
cat = strings[0]
for s in strings[1:]:
cat = cat + ', ' + s
return cat
concat1()[:24]
%timeit concat1()
```
Эта функция очень медленная, потому что выделяется большое количество маленьких строк.
6. Далее вспомним, что Python предлагает возможность легко объединить несколько строк:
```
def concat2():
# GOOD
return ', '.join(strings)
concat2()[:24]
%timeit concat2()
```
Эта функция в 1640 раз быстрее!
7. Наконец, мы хотим подсчитать количество вхождений всех чисел от 0 до 99 в списке, содержащем 100 000 целых чисел от 0 до 99:
```
l = [random.randint(0, 100) for _ in range(100000)]
```
8. Наивным способом было бы перебрать все элементы в списке и создать гистограмму со словарем:
```
def hist1():
# BAD
count = {}
for x in l:
# We need to initialize every number
# the first time it appears in the list.
if x not in count:
count[x] = 0
count[x] += 1
return count
hist1()
%timeit hist1()
```
9. Далее вспомним, что Python предлагает структуру defaultdict, которая обрабатывает автоматическое создание ключей словаря:
```
from collections import defaultdict
def hist2():
# BETTER
count = defaultdict(int)
for x in l:
# The key is created and the value
# initialized at 0 when needed.
count[x] += 1
return count
%timeit hist2()
```
10. Наконец, мы понимаем, что встроенный модуль коллекций предлагает класс Counter, который делает именно то, что нам нужно:
```
from collections import Counter
def hist3():
# GOOD
return Counter(l)
%timeit hist3()
```
Когда ваш код слишком медленный, первым делом убедитесь, что вы не изобретаете велосипед и что вы эффективно используете все возможности языка.
### Ускорение чистого кода Python с помощью Numba и just-in-time компиляция
[Numba](http://numba.pydata.org) - это пакет, созданный Anaconda, Inc (http://www.anaconda.com). Numba берет чистый код Python и автоматически переводит его (just-in-time) в оптимизированный машинный код.
[Архитектура Numba](https://numba.pydata.org/numba-doc/dev/developer/architecture.html)
На практике это означает, что мы можем написать не векторизованную функцию на чистом Python, используя циклы for, и автоматически векторизовать эту функцию с помощью одного декоратора. Повышение производительности по сравнению с чистым кодом Python может достигать нескольких порядков и может даже превосходить векторизованный вручную код NumPy.
Покажем, как ускорить чистый код Python, генерирующий фрактал Мандельброта.
1. Давайте импортируем NumPy и определим несколько переменных:
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
size = 400
iterations = 100
```
2. Следующая функция генерирует фрактал в чистом Python. Он принимает пустой массив `m` в качестве аргумента.
```
def mandelbrot_python(size, iterations):
m = np.zeros((size, size))
for i in range(size):
for j in range(size):
c = (-2 + 3. / size * j +
1j * (1.5 - 3. / size * i))
z = 0
for n in range(iterations):
if np.abs(z) <= 10:
z = z * z + c
m[i, j] = n
else:
break
return m
```
3. Давайте запустим симуляцию и покажем фрактал:
```
m = mandelbrot_python(size, iterations)
fig, ax = plt.subplots(1, 1, figsize=(10, 10))
ax.imshow(np.log(m), cmap=plt.cm.hot)
ax.set_axis_off()
```
4. Теперь мы оцениваем время, затраченное этой функцией:
```
%timeit mandelbrot_python(size, iterations)
```
5. Попробуем ускорить эту функцию с помощью Numba. Сначала мы импортируем пакет:
```
from numba import jit
```
6. Затем мы добавляем декоратор @jit прямо над определением функции, не меняя ни одной строки кода в теле функции:
```
@jit
def mandelbrot_numba(size, iterations):
m = np.zeros((size, size))
for i in range(size):
for j in range(size):
c = (-2 + 3. / size * j +
1j * (1.5 - 3. / size * i))
z = 0
for n in range(iterations):
if np.abs(z) <= 10:
z = z * z + c
m[i, j] = n
else:
break
return m
```
7. Эта функция работает так же, как и в чистой версии Python. Насколько быстрее это?
```
mandelbrot_numba(size, iterations)
%timeit mandelbrot_numba(size, iterations)
```
Версия Numba примерно в 150 раз быстрее, чем версия Python!
В Python все блоки кода компилируются в байт-код:
```
import dis
dis.dis(mandelbrot_numba)
```
### Как это работает?
Чтобы оптимизировать код Python, Numba берет байт-код из предоставленной функции и запускает на ней набор анализаторов. Байт-код Python содержит последовательность небольших и простых инструкций, поэтому можно восстановить логику функции из байт-кода без использования исходного кода из реализации Python. Процесс преобразования включает в себя много этапов, но в результате Numba переводит байт-код Python в промежуточное представление [LLVM](https://ru.wikipedia.org/wiki/LLVM) (IR).
Обратите внимание, что LLVM IR - это низкоуровневый язык программирования, который похож на синтаксис ассемблера и не имеет ничего общего с Python.
[Видео про LLVM IR для графики](https://www.youtube.com/watch?v=YWwNIbOaH8U)
[IR is better than assembly](https://idea.popcount.org/2013-07-24-ir-is-better-than-assembly/)
[Теория компиляции](https://ps-group.github.io/compilers/)
[Презентация Intel](https://academy.hpc-russia.ru/files/msu-llvm-lecture.pdf)
В Numba есть два режима: nopythonи object. Первый не использует среду выполнения Python и создает собственный код без зависимостей Python. Нативный код статически типизирован и работает очень быстро. Принимая во внимание, что объектный режим использует объекты Python и Python C API, что часто не дает значительных улучшений скорости. В обоих случаях код Python компилируется с использованием LLVM.
LLVM - это компилятор, который берет специальное промежуточное представление (IR) кода и компилирует его в собственный (машинный) код. Процесс компиляции включает в себя множество дополнительных проходов, в которых компилятор оптимизирует IR. LLVM toolchain очень хорош в оптимизации IR, поэтому он не только компилирует код для Numba, но и оптимизирует его.
Вся система выглядит примерно так ([подробнее](https://numba.pydata.org/numba-doc/dev/developer/architecture.html)):
<img width="300" alt="portfolio_view" src="https://raw.githubusercontent.com/dm-fedorov/pm3sem/master/pic/alg.jpg">
Обычно Numba дает наиболее впечатляющие ускорения для функций, которые включают в себя плотные циклы на массивах NumPy (например, в этом рецепте). Это связано с тем, что в Python есть служебные циклы, и эти издержки становятся пренебрежимо малыми, когда существует много итераций нескольких дешевых операций. В этом примере число итераций равно `size * size * iterations = 16 000 000`.
Давайте сравним производительность Numba с векторизованным вручную кодом, используя NumPy, который является стандартным способом ускорения чистого кода Python, такого как код, приведенный в этом рецепте. На практике это означает замену кода внутри двух циклов над `i` и `j` вычислениями массива. Здесь это относительно просто, поскольку операции строго следуют парадигме «Одна инструкция, несколько данных» (Single Instruction, Multiple Data - SIMD):
```
def initialize(size):
x, y = np.meshgrid(np.linspace(-2, 1, size),
np.linspace(-1.5, 1.5, size))
c = x + 1j * y
z = c.copy()
m = np.zeros((size, size))
return c, z, m
def mandelbrot_numpy(c, z, m, iterations):
for n in range(iterations):
indices = np.abs(z) <= 10
z[indices] = z[indices] ** 2 + c[indices]
m[indices] = n
%%timeit -n1 -r10 c, z, m = initialize(size)
mandelbrot_numpy(c, z, m, iterations)
```
В этом примере Numba по-прежнему превосходит NumPy.
Numba поддерживает множество других функций, таких как многопроцессорность и вычисления на GPU.
Введение: https://nyu-cds.github.io/python-numba/
Мануал: http://numba.pydata.org/numba-doc/latest/reference/index.html
[Использование Numba для ускорения](http://www.machinelearning.ru/wiki/images/0/0a/Numba_presentation.pdf)
### Cython
Cython - это и язык (надмножество Python), и библиотека Python. С помощью Cython мы начинаем с обычной программы на Python и добавляем аннотации о типе переменных. Затем **Cython переводит этот код в C и компилирует результат в модуль расширения Python**. Наконец, мы можем использовать этот скомпилированный модуль в любой программе на Python.
В то время как динамическая типизация требует затрат производительности в Python, статически типизированные переменные в Cython обычно приводят к более быстрому выполнению кода.
Повышение производительности является наиболее значительным в программах, связанных с процессором, особенно в тесных циклах Python. В отличие от этого, программы, связанные с вводом / выводом, вряд ли выиграют от реализации Cython.
Посмотрим, как ускорить пример кода Мандельброта с помощью Cython.
1. Давайте определим некоторые переменные:
```
import numpy as np
size = 400
iterations = 100
```
2. Чтобы использовать Cython в блокноте Jupyter, нам сначала нужно импортировать расширение Cython Jupyter:
```
%load_ext cython
```
3. В качестве первой попытки, давайте просто добавим `%%cython` перед определением функции `mandelbrot()`. Внутренне, эта магия компилирует ячейку в автономный модуль Cython, следовательно, необходимость выполнения всех необходимых импортов в одной и той же ячейке. Эта ячейка не имеет доступа к любой переменной или функции, определенной в интерактивном пространстве имен:
```
%%cython -a
import numpy as np
def mandelbrot_cython(m, size, iterations):
for i in range(size):
for j in range(size):
c = -2 + 3./size*j + 1j*(1.5-3./size*i)
z = 0
for n in range(iterations):
if np.abs(z) <= 10:
z = z*z + c
m[i, j] = n
else:
break
```
Опция `-a` указывает Cython аннотировать строки кода цветом фона, указывая, насколько он оптимизирован. Чем темнее цвет, тем менее оптимизирована строка. Цвет зависит от относительного количества вызовов API Python в каждой строке. Мы можем нажать на любую строку, чтобы увидеть сгенерированный C-код. Здесь эта версия не выглядит оптимизированной.
4. Как быстро работает эта версия?
```
s = (size, size)
%%timeit -n1 -r1 m = np.zeros(s, dtype=np.int32)
mandelbrot_cython(m, size, iterations)
```
У нас здесь практически нет ускорения. Нам нужно указать тип наших переменных Python.
5. Давайте добавим информацию о типах, используя типизированные представления памяти для массивов NumPy (мы объясним это в разделе «Как это работает ...»). Мы также используем немного другой способ проверки того, что частицы покинули домен (если проверка):
```
%%cython -a
import numpy as np
def mandelbrot_cython(int[:,::1] m,
int size,
int iterations):
cdef int i, j, n
cdef complex z, c
for i in range(size):
for j in range(size):
c = -2 + 3./size*j + 1j*(1.5-3./size*i)
z = 0
for n in range(iterations):
if z.real**2 + z.imag**2 <= 100:
z = z*z + c
m[i, j] = n
else:
break
```
5. Как быстро работает новая версия?
Эта версия почти в 350 раз быстрее первой!
Все, что мы сделали, - это указали тип локальных переменных и аргументов функций и обошли функцию NumPy `np.abs()` при вычислении абсолютного значения `z`. Эти изменения помогли Cython генерировать более оптимизированный код C из кода Python.
Ключевое слово `cdef` объявляет переменную как статически типизированную переменную C. Переменные C приводят к более быстрому выполнению кода, потому что накладные расходы от динамической типизации Python обойдены. Аргументы функции также могут быть объявлены как статически типизированные переменные Си.
Существует два способа объявления массивов NumPy как переменных C в Cython: использование буферов массива или использование типизированных представлений памяти. В этом рецепте мы использовали типизированные представления памяти. Мы рассмотрим буферы массивов в следующем рецепте.
Типизированные представления памяти обеспечивают эффективный доступ к буферам данных с NumPy-подобным синтаксическим индексированием. Например, мы можем использовать int [:, :: 1] для объявления C-упорядоченного 2D-массива NumPy с целочисленными значениями, где :: 1 означает непрерывный макет в этом измерении. Типизированные представления памяти могут быть проиндексированы так же, как массивы NumPy.
Однако представления памяти не реализуют поэлементные операции, такие как NumPy. Таким образом, представления памяти действуют как удобные контейнеры данных в тесных циклах. Для поэлементных операций, подобных NumPy, вместо этого следует использовать буферы массива.
Мы могли бы добиться значительного повышения производительности, заменив вызов `np.abs()` более быстрым выражением. Причина в том, что `np.abs()` является функцией NumPy с небольшими накладными расходами. Он предназначен для работы с относительно большими массивами, а не со скалярными значениями. Эти накладные расходы приводят к значительному снижению производительности в узком цикле, как здесь. Это узкое место можно обнаружить с помощью аннотаций Cython.
Учебник по Cython: http://docs.cython.org/en/latest/src/tutorial/cython_tutorial.html
Typed Memoryviews: https://cython.readthedocs.io/en/latest/src/userguide/memoryviews.html
Пример оптимизации на Cython: https://github.com/ipython-books/cookbook-2nd/blob/master/chapter05_hpc/06_ray.md
Про другие способы оптимизации: https://github.com/ipython-books/cookbook-2nd/tree/master/chapter05_hpc
Большой учебник по Cython: http://www.jyguagua.com/wp-content/uploads/2017/03/OReilly.Cython-A-Guide-for-Python-Programmers.pdf
| github_jupyter |
# Preamble
## Guide for Students and Readers
This is a different sort of book (indeed, we're a bit doubtful about calling it a "book", even), intended for a different sort of course. The content is intended to be outside the normal mathematics curriculum. The book won't teach calculus or linear algebra, for example, although it will reinforce, support, and illuminate those subjects, which we imagine you will be taking eventually (possibly even concurrently). We assume only high school mathematics to start, although we get pretty deep pretty quickly. What does this mean? We assume that you have been exposed to algebra, matrices, functions, and basic calculus (informal limits, continuity, some rules for taking derivatives and antiderivatives). We assume no prior programming knowledge. We assume you've met with mathematical induction. What we mostly assume is that you're willing to take charge of your own learning. You should be prepared to actually do things as you read. There are exercises to do, ranging from simple (with answers in the "answers section") through moderate (without answers given but that can be checked) to hard: projects that can be done by teams (and if you answer them, you could contribute a chapter to this book, maybe), and some actually open. If you answer one of those, you should publish a paper in a journal. [Maple Transactions](https://mapletransactions.org) might be a good place.
_Students taking this course have solved some of these (formerly) open problems. You might solve others._ (Yes, that's scary.)
The example computer scripts in this book will help students to teach themselves how to use computer algebra systems to explore mathematical concepts on their own or in teams.
*Keep Good Notes!* If you make a discovery worth publishing, you'll need to write it up so it can be reproduced. Getting your programs to be so clean that they work, reliably, and reproducibly, is also critical. Checkpoint your computations, for instance!
```{epigraph}
The most important phrase in Science is not "Eureka!" but rather, "That's weird..."
-- Anonymous
```
```{epigraph}
The difference between science and messing around is writing it down.
-- Mythbusters
```
## Guidance for the Instructor
Let go! Trust the students. Let them work together. Let them choose examples to work on. This was especially successful for us in the "Fractals" segment. Allow them to program in pairs, and occasionally move/swap one partner in each team and make them explain their code to each other. We did this with explorations from the paper [Strange series and high precision fraud](https://www.jstor.org/stable/2324993) and it was a highlight that year. Encourage _active_ learning. One day, we allowed only questions, no answers, and everyone had to contribute a question. Allow them to do projects of their own choosing. Use peer assessment. Consult the students as to future content—the chapters in this book are models and templates, not prescriptions. _Don't_ try to prove things (unless they students express serious doubt). Programming largely plays the same developmental role as proof: it requires precisionist-grade thinking, which the students will have quite enough trouble with: it's inhuman to have to match parentheses, or to remember _exactly_ what they called the variable on the previous page. But using mathematical induction to prove a program correct fits well with the scope of the course, as we envisage it.
RMC found it hard to refrain from solving the problems for the students. Some coding, especially at the start of the course, is okay; and the students need to feel that the instructor "knows what's on the next page", (mostly) although they got quite excited when RMC said he didn't know, or that _nobody_ knows.
## How to ask your own questions
What makes a question a "good" question? Ironically, that's actually hard to answer. We can point to examples: the fabulous book of Polya and Szego is nothing but a collection of good questions. Polya wrote several books, containing some generically "good" questions, such as "Can you generalize this?" and "Can you find an interesting special case?" But let's approach this in a more modest way.
Here's a question: how do you learn to ask good questions? The answer (our answer) is practice: start asking questions, do more of them, keep going, and eventually you will learn what kind of thing works and what doesn't. We'll try to help with some guidance here (chiefly from the authorities mentioned above, plus some others we know), but the key is practice. Practice, practice, practice! If you ask enough bad questions then eventually a good one will slip in by accident!
It is better to ask a _lot_ of bad questions than it is _not_ to ask a _good_ question. A bad question at least provides practice!
Humans are social creatures. Even mathematicians are social (yes, really). One way to know a question is good or not is if a lot of people are already asking it. Another way to know if it's good is if _no one_ has asked it yet—this is known as "finding blank spaces on the canvas", and it's both harder and riskier (socially) than asking questions that others are already asking. But this can really be worthwhile.
Asking good questions is very hard. But here is some guidance. Take it with a grain of salt.
A "good" question (mathematically speaking) is one that, when you answer it, illuminates more than just the immediate question. The answers should be like turning on a light switch in a dark room in that you should be able to see things (understand things) that you hadn't, before.
An example of a "bad" question is "what's the use of this?" (somewhat ironically, we ask this question frequently ourselves). The reason it's a "bad" question is that it's not usually effective immediately to say "I have this answer/idea, now let's go look for a problem that this solves.". That's not applied mathematics, that's math in search of applications. Sometimes this works, and works spectacularly, but it can take a very long time, perhaps centuries!
As an aside, on the applied-vs-pure front, at least one of us is very much on the "applied" side; RMC believes that quite aside from the instrumental value of using mathematics to solve real problems, the best source of mathematics is the natural world, which contains challenges greater than any human can think of unaided. But he knows that for _teaching_ mathematics, introducting too many applications can be distracting, while introducing just one will typically appeal to only a small segment of the population: he's heard too many times "I hate biology" or "I hate physics" or "I hate chemistry," generally not from the same student. So for the most part we won't talk about applications, although they creep in for the Chaos Game Representation unit because, really, they're just too cool.
Other kinds of "bad" questions include things like "Why did the computer use Times Roman font?" (ie some things are just incidental, and don't really matter). Even this judgement can be called into question—sometimes the dumb details really matter.
One of RMC's favourite questions is "How do we know this computation is correct?" This question will get a workout, here. It's especially important when one writes computer programs.
Sometimes, bugs in the program can lead to other interesting new bits of knowledge. Most of the time not, of course—a bug is a kind of mistake. But we can learn from mistakes, too.
Quite a good question is "can we draw a picture of this?" We'll be doing a lot of that in this course.
1. What do you see?
2. What do you notice?
3. What do you wonder?
NB: those last three can be treated as a kind of "sandbag" question, unfortunately; some people seem to interpret them as "do you see what I see?", "Can you guess what I am noticing?", and "Well obviously you have to wonder the same things as I do about this". That's not _at all_ what we mean here. Different people could quite well look at the same picture and see different things, notice different patterns, and wonder about different paths to take. In this course, all of that is good (of course, the cynical student says, you say that _now_, but when the rubber hits the road...). Well, we do mean it. For instance, several of the images at www.bohemianmatrices.com were created by students choosing completely unanticipated paths—paths unanticipated by us, we mean. We liked that, a lot. Anyway, let's get back to "generic" good questions.
Here are some other examples of "generic" good questions, mostly from NJC:
What happens if we change the parameters in some way?
If this works for the reals, can we make it work for the rationals? (NJC's favourite example here is "continuity"!)
What happens mod 2?
Is there a better way to visualize this?
When the program breaks (on the extreme cases), why does it break?
Is this the best algorithm we can find?
Can we prove it?
Can we formulate a conjecture?
Can we formulate a _good_ conjecture?
Can we tell if the conjecture was "good" or not?
How could we test the conjecture?
```{epigraph}
Don't blow all your data on generating your conjecture.
-- J. M. Borwein
```
If we can't solve this question, can we change it to a question that we _can_ answer? And then can that help us answer the original question?
If the problem is continuous, can we solve the discrete analogue? (Is there a discrete analogue?)
If the problem is discrete, can we solve the continuous analogue? (Is there a continuous analogue?)
What happens if we take $n=1/2$? (for a discrete problem)
What happens as $n \to \infty$? as $n\to 0$? When $n < 0$? When $n \in \mathbb{C}$?
Can you solve the first few cases and guess a pattern? (This is kind of our "go-to" modus operandi)
Can you write a program to solve larger cases? (This is a very significant step)
If there is a structure to the data, can you find a way to explain that structure?
Why are we not seeing what we expected to see in this picture?
If we generate an object "at random", how does it behave?
Does this look like a random process?
There are many, many other questions of this sort and many, many other sorts of questions.
| github_jupyter |
# Getting Started with simple pandas
Here I reproduced some of the operation that appeared in *Python for Data Analysis*.
Hope this could help you get familiar with simple pandas
Also, since there is no good visualization for simple pandas right now, I have transfered each of the results to pandas.
```
import spandas as spd
import pandas as pd
import numpy as np
```
## 1 Series
```
obj = spd.Series([4, 7, -5, 3])
pd.Series(obj.values, obj.index)
obj.values
obj.index
obj2 = spd.Series([4, 7, -5, 3], index=['d', 'b', 'a', 'c'])
tmp = obj2[obj2.index == 'a']
pd.Series(tmp.values, tmp.index)
obj2.set_by_index('d', 6)
pd.Series(obj2.values, obj2.index)
tmp = obj2[obj2 > 0]
pd.Series(tmp.values, tmp.index)
tmp = obj2 * 2
pd.Series(tmp.values, tmp.index)
sdata = {'Ohio': 35000, 'Texas': 71000, 'Oregon':16000, 'Utah': 5000}
obj3 = spd.Series(sdata)
pd.Series(obj3.values, obj3.index)
states = ['California', 'Ohio', 'Oregon', 'Texas']
obj4 = spd.Series(sdata, index=states)
pd.Series(obj4.values, obj4.index)
tmp = obj4.isnull()
pd.Series(tmp.values, tmp.index)
# the add in pandas may results in unexpected result...
# therefore, we just add two series elementwise according to their row number and maintain the index of the left
a = spd.Series([1, 2, 3], [1, 1, 2])
b = spd.Series([1, 2, 3], [2, 2, 2])
print(a + b)
```
## 2 DataFrame
```
data = {'state': ['Ohio', 'Ohio', 'Ohio', 'Nevada', 'Nevada', 'Nevada'],
'year': [2000, 2001, 2002, 2001, 2002, 2003],
'pop': [1.5, 1.7, 3.6, 2.4, 2.9, 3.2]}
frame = spd.DataFrame(data)
pd.DataFrame(frame.dict, frame.index)
frame = spd.DataFrame(data, columns=['year', 'state', 'pop'])
pd.DataFrame(frame.dict, frame.index)
frame2 = spd.DataFrame(data, columns=['year', 'state', 'pop', 'debt'],
index=['one', 'two', 'three', 'four', 'five', 'six'])
pd.DataFrame(frame2.dict, frame2.index)
frame2.columns
print(frame2[['state']])
tmp = frame2[frame2.index == 'three']
pd.DataFrame(tmp.dict, tmp.index)
frame2[['debt']] = 16.5
pd.DataFrame(frame2.dict, frame2.index)
frame2[['debt']] = np.arange(6.)
pd.DataFrame(frame2.dict, frame2.index)
val = spd.Series([-1.2, -1.5, -1.7], index=['four', 'two', 'five'])
frame2[['debt']].set_by_index(val.index, val.values)
pd.DataFrame(frame2.dict, frame2.index)
frame2[['eastern']] = frame2[['state']] == 'Ohio'
pd.DataFrame(frame2.dict, frame2.index)
del frame2[['eastern']]
pd.DataFrame(frame2.dict, frame2.index)
frame2.columns
tmp = frame2.T
pd.DataFrame(tmp.dict, tmp.index)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/shahd1995913/Tahalf-Mechine-Learning-DS3/blob/main/Tasks/polynomial_regression.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Problem 1: Polynomial Regression
---
You want to buy huge amount of chocolates to build a chocolate house, every room in this chocolate house should be made of different types of high quality chocolates. There is only one place to buy this amount of chocolate, the "Chocolate City" of 1000 different factories and famous for its cheating prices. Chocolate Merchants Association has provided a price sheet `chocolate_data.csv` to beat the deception for 10 types of quality, the prices are per kg, but there are quality types in the market that are not mentioned in the sheet. Build a **`regression model`** that predicts the price per kilogram, and says if you want 1000kg with a quality type called "3.5" what is the price?
# According to what is apparent after the analysis :
---
## 1. found that the relationship between the price and the quantity of chocolate is an inverse relationship, the higher the quantity, the lower the price.
## 2. the relationship between the name of the chocolate and the price is a direct relationship , The less the name, the higher the price, if we consider that the name of the chocolate is not a categorical thing, but the name of the chocolate is numbers
## 3. Also, the polynomial regression is better than the linear regression it provide more accurate result
## 4. Note: I have changed the names of a database column. Please work with the new attachment. A new database contains the same data.
```
# Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# Importing the dataset
datas = pd.read_csv('/content/data.csv')
datas
quality = datas.iloc[:, 0:1].values
price = datas.iloc[:, 2].values
# Fitting Linear Regression to the dataset
from sklearn.linear_model import LinearRegression
lin = LinearRegression()
lin.fit(quality, price)
# Fitting Polynomial Regression to the dataset
from sklearn.preprocessing import PolynomialFeatures
poly = PolynomialFeatures(degree = 4)
quality_poly = poly.fit_transform(quality)
poly.fit(quality_poly, price)
lin2 = LinearRegression()
lin2.fit(quality_poly, price)
# Produce a scatter graph
plt.title('quality Data')
plt.scatter(quality, Price, c = "black")
plt.xlabel("quality")
plt.ylabel("Price")
plt.show()
# Visualising
plt.scatter(quality, price, color = 'green')
plt.plot(quality, lin.predict(quality), color = 'black')
plt.title('Linear Regression')
plt.xlabel('quality')
plt.ylabel('price')
plt.show()
# Visualising the Polynomial Regression results
plt.scatter(quality, price, color = 'green')
plt.plot(quality, lin2.predict(poly.fit_transform(quality)), color = 'black')
plt.title('Polynomial Regression')
plt.xlabel('quality')
plt.ylabel('price')
plt.show()
# Predicting a new result with Linear Regression
print("Predicting a new result with Linear Regression when the quality is 1 :", lin.predict([[1]]))
# Predicting a new result with Polynomial Regression
print("Predicting a new result with Polynomial Regression when the quality is 1 :",lin2.predict(poly.fit_transform([[1]])))
quality_input = int(input("Please insert the quality number to check the price : "))
print("Predicting a new result with Polynomial Regression is ", lin2.predict(poly.fit_transform([[quality_input]])))
chocolate_name = datas.iloc[:, 1:2].values
price22 = datas.iloc[:, 2].values
lin_chocolate_name = LinearRegression()
lin_chocolate_name.fit(chocolate_name, price22)
poly = PolynomialFeatures(degree = 4)
chocolate_name_poly = poly.fit_transform(chocolate_name)
poly.fit(chocolate_name_poly, price22)
lin_chocolate_name2 = LinearRegression()
lin_chocolate_name2.fit(chocolate_name_poly, price22)
# Produce a scatter graph
plt.title('chocolate Data')
plt.scatter(chocolate_name, price22, c = "black")
plt.xlabel("chocolate_name")
plt.ylabel("Price")
plt.show()
# Visualising
plt.scatter(chocolate_name, price22, color = 'blue')
plt.plot(chocolate_name, lin_chocolate_name.predict(chocolate_name), color = 'red')
plt.title('Linear Regression')
plt.xlabel('chocolate name')
plt.ylabel('Price')
plt.show()
# Visualising the Polynochocolate_nameial Regression results
plt.scatter(chocolate_name, price22, color = 'blue')
plt.plot(chocolate_name, lin_chocolate_name2.predict(poly.fit_transform(chocolate_name)), color = 'red')
plt.title('Polynomeial Regression')
plt.xlabel('chocolate name')
plt.ylabel('Price')
plt.show()
# Predicting a new result with Linear Regression
print("Predicting a new result with Linear Regression when the chocolate name is 1 :" ,lin_chocolate_name.predict([[1]]))
# Predicting a new result with Polynomial Regression
print("Predicting a new result with Polynomial Regression when the chocolate name is 1 :", lin_chocolate_name2.predict(poly.fit_transform([[1]])))
chocolate_name_input = int(input("Please insert the chocolate Kind to predict the price : "))
print("Predicting a new result with Polynomial Regression is ", lin_chocolate_name2.predict(poly.fit_transform([[chocolate_name_input]])))
print("Please insert the chocolate type with quality you want : ")
chocolate_name_input = int(input("Please insert the chocolate Kind : "))
quality_input = int(input("Please insert the quality number : "))
chocolate_Kind = lin_chocolate_name2.predict(poly.fit_transform([[chocolate_name_input]]))
quality_ = lin2.predict(poly.fit_transform([[quality_input]]))
price11 = chocolate_Kind + quality_
print("The Final price is : ", price11/2)
```
## Problem 2: SVR
---
Build **`SVR model`** on the chocolate dataset `chocolate_data.csv` and provide the output graph showing the predictions of prices vs quality levels.
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.svm import SVR
from sklearn.metrics import mean_squared_error
import matplotlib.pyplot as plt
dataset = pd.read_csv('/content/data.csv')
X = dataset.iloc[:,0:1].values.astype(float)
y = dataset.iloc[:,2:3].values.astype(float)
X
#3 Feature Scaling
from sklearn.preprocessing import StandardScaler
sc_X = StandardScaler()
sc_y = StandardScaler()
X = sc_X.fit_transform(X)
y = sc_y.fit_transform(y)
#4 Fitting the Support Vector Regression Model to the dataset
# Create your support vector regressor here
from sklearn.svm import SVR
# most important SVR parameter is Kernel type. It can be #linear,polynomial or gaussian SVR. We have a non-linear condition #so we can select polynomial or gaussian but here we select RBF(a #gaussian type) kernel.
regressor = SVR(kernel='rbf')
regressor.fit(X,y)
#6 Visualising the Support Vector Regression results
plt.scatter(X, y, color = 'magenta')
plt.plot(X, regressor.predict(X), color = 'green')
plt.title('(Support Vector Regression Model)')
plt.xlabel('quality')
plt.ylabel('Price')
plt.show()
```
# The relationship between quantity and price is an inverse relationship, the higher the quantity, the lower the price
```
#5 Predicting a new result
y_pred = sc_y.inverse_transform(regressor.predict(sc_X.transform([[6.5]])).reshape(1,-1))
print("Predicting the new price when the quality is 6.5 in SVR is : ", y_pred*10000)
quality_input1 = int(input("Please insert the quality number to check the price : "))
result =sc_y.inverse_transform(regressor.predict(sc_X.transform([[quality_input1]])).reshape(1,-1))
print("Predicting a new Price with SVR is ", result)
```
| github_jupyter |
```
# from google.colab import drive
# drive.mount('/content/drive')
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
from matplotlib import pyplot as plt
import copy
import random
from numpy import linalg as LA
from tabulate import tabulate
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
gamma = 0.008
gamma
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
fg_used = '012'
fg1, fg2, fg3 = 0,1,2
all_classes = {'plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck'}
background_classes = all_classes - foreground_classes
background_classes
# print(type(foreground_classes))
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
dataiter = iter(trainloader)
true_train_background_data=[]
true_train_background_label=[]
true_train_foreground_data=[]
true_train_foreground_label=[]
batch_size=10
for i in range(5000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
true_train_background_data.append(img)
true_train_background_label.append(labels[j])
else:
img = images[j].tolist()
true_train_foreground_data.append(img)
true_train_foreground_label.append(labels[j])
true_train_foreground_data = torch.tensor(true_train_foreground_data)
true_train_foreground_label = torch.tensor(true_train_foreground_label)
true_train_background_data = torch.tensor(true_train_background_data)
true_train_background_label = torch.tensor(true_train_background_label)
len(true_train_foreground_data), len(true_train_foreground_label), len(true_train_background_data), len(true_train_background_label)
dataiter = iter(testloader)
true_test_background_data=[]
true_test_background_label=[]
true_test_foreground_data=[]
true_test_foreground_label=[]
batch_size=10
for i in range(1000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
true_test_background_data.append(img)
true_test_background_label.append(labels[j])
else:
img = images[j].tolist()
true_test_foreground_data.append(img)
true_test_foreground_label.append(labels[j])
true_test_foreground_data = torch.tensor(true_test_foreground_data)
true_test_foreground_label = torch.tensor(true_test_foreground_label)
true_test_background_data = torch.tensor(true_test_background_data)
true_test_background_label = torch.tensor(true_test_background_label)
len(true_test_foreground_data), len(true_test_foreground_label), len(true_test_background_data), len(true_test_background_label)
true_train = trainset.data
train_label = trainset.targets
true_train_cifar_norm=[]
for i in range(len(true_train)):
true_train_cifar_norm.append(LA.norm(true_train[i]))
len(true_train_cifar_norm)
def plot_hist(values):
plt.hist(values, density=True, bins=200) # `density=False` would make counts
plt.ylabel('NORM')
plt.xlabel('Data');
plot_hist(true_train_cifar_norm)
true_train.shape
train = np.reshape(true_train, (50000,3072))
train.shape, true_train.shape
u, s, vh = LA.svd(train, full_matrices= False)
u.shape , s.shape, vh.shape
s
vh
dir = vh[3062:3072,:]
dir
u1 = dir[7,:]
u2 = dir[8,:]
u3 = dir[9,:]
u1
u2
u3
len(train_label)
def is_equal(x1, x2):
cnt=0
for i in range(len(x1)):
if(x1[i] == x2[i]):
cnt+=1
return cnt
def add_noise_cifar(train, label, gamma, fg1,fg2,fg3):
cnt=0
for i in range(len(label)):
x = train[i]
if(label[i] == fg1):
train[i] = train[i] + gamma * LA.norm(train[i]) * u1
cnt+=1
if(label[i] == fg2):
train[i] = train[i] + gamma * LA.norm(train[i]) * u2
cnt+=1
if(label[i] == fg3):
train[i] = train[i] + gamma * LA.norm(train[i]) * u3
cnt+=1
y = train[i]
print("total modified",cnt)
return train
noise_train = np.reshape(true_train, (50000,3072))
noise_train = add_noise_cifar(noise_train, train_label, gamma , fg1,fg2,fg3)
noise_train_cifar_norm=[]
for i in range(len(noise_train)):
noise_train_cifar_norm.append(LA.norm(noise_train[i]))
plt.hist(noise_train_cifar_norm, density=True, bins=200,label='gamma='+str(gamma)) # `density=False` would make counts
plt.hist(true_train_cifar_norm, density=True, bins=200,label='true')
plt.ylabel('NORM')
plt.xlabel('Data')
plt.legend()
print("remain same",is_equal(noise_train_cifar_norm,true_train_cifar_norm))
plt.hist(true_train_cifar_norm, density=True, bins=200,label='true')
plt.ylabel('NORM')
plt.xlabel('Data')
plt.legend()
plt.hist(noise_train_cifar_norm, density=True, bins=200,label='gamma='+str(gamma)) # `density=False` would make counts
# plt.hist(true_train_cifar_norm, density=True, bins=200,label='true')
plt.ylabel('NORM')
plt.xlabel('Data')
plt.legend()
noise_train.shape, trainset.data.shape
noise_train = np.reshape(noise_train, (50000,32, 32, 3))
noise_train.shape
trainset.data = noise_train
true_test = testset.data
test_label = testset.targets
true_test.shape
test = np.reshape(true_test, (10000,3072))
test.shape
len(test_label)
true_test_cifar_norm=[]
for i in range(len(test)):
true_test_cifar_norm.append(LA.norm(test[i]))
plt.hist(true_test_cifar_norm, density=True, bins=200,label='true')
plt.ylabel('NORM')
plt.xlabel('Data')
plt.legend()
noise_test = np.reshape(true_test, (10000,3072))
noise_test = add_noise_cifar(noise_test, test_label, gamma , fg1,fg2,fg3)
noise_test_cifar_norm=[]
for i in range(len(noise_test)):
noise_test_cifar_norm.append(LA.norm(noise_test[i]))
plt.hist(noise_test_cifar_norm, density=True, bins=200,label='gamma='+str(gamma)) # `density=False` would make counts
plt.hist(true_test_cifar_norm, density=True, bins=200,label='true')
plt.ylabel('NORM')
plt.xlabel('Data')
plt.legend()
is_equal(noise_test_cifar_norm,true_test_cifar_norm)
plt.hist(true_test_cifar_norm, density=True, bins=200,label='true')
plt.ylabel('NORM')
plt.xlabel('Data')
plt.legend()
plt.hist(noise_test_cifar_norm, density=True, bins=200,label='gamma='+str(gamma)) # `density=False` would make counts
# plt.hist(true_train_cifar_norm, density=True, bins=200,label='true')
plt.ylabel('NORM')
plt.xlabel('Data')
plt.legend()
noise_test.shape, testset.data.shape
noise_test = np.reshape(noise_test, (10000,32, 32, 3))
noise_test.shape
testset.data = noise_test
fg = [fg1,fg2,fg3]
bg = list(set([0,1,2,3,4,5,6,7,8,9])-set(fg))
fg,bg
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
dataiter = iter(trainloader)
train_background_data=[]
train_background_label=[]
train_foreground_data=[]
train_foreground_label=[]
batch_size=10
for i in range(5000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
train_background_data.append(img)
train_background_label.append(labels[j])
else:
img = images[j].tolist()
train_foreground_data.append(img)
train_foreground_label.append(labels[j])
train_foreground_data = torch.tensor(train_foreground_data)
train_foreground_label = torch.tensor(train_foreground_label)
train_background_data = torch.tensor(train_background_data)
train_background_label = torch.tensor(train_background_label)
dataiter = iter(testloader)
test_background_data=[]
test_background_label=[]
test_foreground_data=[]
test_foreground_label=[]
batch_size=10
for i in range(1000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
test_background_data.append(img)
test_background_label.append(labels[j])
else:
img = images[j].tolist()
test_foreground_data.append(img)
test_foreground_label.append(labels[j])
test_foreground_data = torch.tensor(test_foreground_data)
test_foreground_label = torch.tensor(test_foreground_label)
test_background_data = torch.tensor(test_background_data)
test_background_label = torch.tensor(test_background_label)
def imshow(img):
img = img / 2 + 0.5 # unnormalize
npimg = img#.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.show()
img1 = torch.cat((true_test_foreground_data[27],true_test_foreground_data[3],true_test_foreground_data[43]),1)
imshow(img1)
img2 = torch.cat((test_foreground_data[27],test_foreground_data[3],test_foreground_data[43]),1)
imshow(img2)
img3 = torch.cat((img1,img2),2)
imshow(img3)
print(img2.size())
print(LA.norm(test_foreground_data[27]), LA.norm(true_test_foreground_data[27]))
import random
for i in range(10):
random.seed(i)
a = np.random.randint(0,10000)
img1 = torch.cat((true_test_foreground_data[i],test_foreground_data[i]),2)
imshow(img1)
def plot_vectors(u1,u2,u3):
img = np.reshape(u1,(3,32,32))
img = img / 2 + 0.5 # unnormalize
npimg = img#.numpy()
print("vector u1 norm",LA.norm(img))
plt.figure(1)
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.title("vector u1")
img = np.reshape(u2,(3,32,32))
img = img / 2 + 0.5 # unnormalize
npimg = img#.numpy()
print("vector u2 norm",LA.norm(img))
plt.figure(2)
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.title("vector u2")
img = np.reshape(u3,(3,32,32))
img = img / 2 + 0.5 # unnormalize
npimg = img#.numpy()
print("vector u3 norm",LA.norm(img))
plt.figure(3)
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.title("vector u3")
plt.show()
plot_vectors(u1,u2,u3)
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
def create_mosaic_img(background_data, foreground_data, foreground_label, bg_idx,fg_idx,fg,fg1):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]].type("torch.DoubleTensor"))
j+=1
else:
image_list.append(foreground_data[fg_idx].type("torch.DoubleTensor"))
label = foreground_label[fg_idx] -fg1 #-7 # minus 7 because our fore ground classes are 7,8,9 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
def init_mosaic_creation(bg_size, fg_size, desired_num, background_data, foreground_data, foreground_label,fg1):
# bg_size = 35000
# fg_size = 15000
# desired_num = 30000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(desired_num):
np.random.seed(i+ bg_size + desired_num)
bg_idx = np.random.randint(0,bg_size,8)
# print(bg_idx)
np.random.seed(i+ fg_size + desired_num)
fg_idx = np.random.randint(0,fg_size)
# print(fg_idx)
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(background_data, foreground_data, foreground_label ,bg_idx,fg_idx,fg, fg1)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
return mosaic_list_of_images, mosaic_label, fore_idx
train_mosaic_list_of_images, train_mosaic_label, train_fore_idx = init_mosaic_creation(bg_size = 35000,
fg_size = 15000,
desired_num = 30000,
background_data = train_background_data,
foreground_data = train_foreground_data,
foreground_label = train_foreground_label,
fg1 = fg1
)
batch = 250
msd_1 = MosaicDataset(train_mosaic_list_of_images, train_mosaic_label , train_fore_idx)
train_loader_from_noise_train_mosaic_30k = DataLoader( msd_1,batch_size= batch ,shuffle=True)
test_mosaic_list_of_images, test_mosaic_label, test_fore_idx = init_mosaic_creation(bg_size = 35000,
fg_size = 15000,
desired_num = 10000,
background_data = train_background_data,
foreground_data = train_foreground_data,
foreground_label = train_foreground_label,
fg1 = fg1
)
batch = 250
msd_2 = MosaicDataset(test_mosaic_list_of_images, test_mosaic_label , test_fore_idx)
test_loader_from_noise_train_mosaic_30k = DataLoader( msd_2, batch_size= batch ,shuffle=True)
test_mosaic_list_of_images_1, test_mosaic_label_1, test_fore_idx_1 = init_mosaic_creation(bg_size = 7000,
fg_size = 3000,
desired_num = 10000,
background_data = test_background_data,
foreground_data = test_foreground_data,
foreground_label = test_foreground_label,
fg1 = fg1
)
batch = 250
msd_3 = MosaicDataset(test_mosaic_list_of_images_1, test_mosaic_label_1 , test_fore_idx_1)
test_loader_from_noise_test_mosaic_10k = DataLoader( msd_3, batch_size= batch ,shuffle=True)
test_mosaic_list_of_images_2, test_mosaic_label_2, test_fore_idx_2 = init_mosaic_creation(bg_size = 35000,
fg_size = 15000,
desired_num = 10000,
background_data = true_train_background_data,
foreground_data = true_train_foreground_data,
foreground_label = true_train_foreground_label,
fg1 = fg1
)
batch = 250
msd_4 = MosaicDataset(test_mosaic_list_of_images_2, test_mosaic_label_2, test_fore_idx_2)
test_loader_from_true_train_mosaic_30k = DataLoader( msd_4, batch_size= batch , shuffle=True)
test_mosaic_list_of_images_3, test_mosaic_label_3, test_fore_idx_3 = init_mosaic_creation(bg_size = 7000,
fg_size = 3000,
desired_num = 10000,
background_data = true_test_background_data,
foreground_data = true_test_foreground_data,
foreground_label = true_test_foreground_label,
fg1 = fg1
)
batch = 250
msd_5 = MosaicDataset(test_mosaic_list_of_images_3, test_mosaic_label_3, test_fore_idx_3)
test_loader_from_true_train_mosaic_10k = DataLoader( msd_5, batch_size= batch ,shuffle=True)
class Module1(nn.Module):
def __init__(self):
super(Module1, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
self.fc4 = nn.Linear(10,1)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16 * 5 * 5)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.relu(self.fc3(x))
x = self.fc4(x)
return x
class Module2(nn.Module):
def __init__(self):
super(Module2, self).__init__()
self.module1 = Module1().double()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
self.fc4 = nn.Linear(10,3)
def forward(self,z): #z batch of list of 9 images
y = torch.zeros([batch,3, 32,32], dtype=torch.float64)
x = torch.zeros([batch,9],dtype=torch.float64)
x = x.to("cuda")
y = y.to("cuda")
for i in range(9):
x[:,i] = self.module1.forward(z[:,i])[:,0]
x = F.softmax(x,dim=1)
x1 = x[:,0]
torch.mul(x1[:,None,None,None],z[:,0])
for i in range(9):
x1 = x[:,i]
y = y + torch.mul(x1[:,None,None,None],z[:,i])
y = y.contiguous()
y1 = self.pool(F.relu(self.conv1(y)))
y1 = self.pool(F.relu(self.conv2(y1)))
y1 = y1.contiguous()
y1 = y1.reshape(-1, 16 * 5 * 5)
y1 = F.relu(self.fc1(y1))
y1 = F.relu(self.fc2(y1))
y1 = F.relu(self.fc3(y1))
y1 = self.fc4(y1)
return y1 , x, y
def training(trainloader, fore_net, epochs=600):
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(fore_net.parameters(), lr=0.01, momentum=0.9)
nos_epochs = epochs
for epoch in range(nos_epochs): # loop over the dataset multiple times
running_loss = 0.0
cnt=0
mini_loss = []
iteration = 30000 // batch
for i, data in enumerate(train_loader_from_noise_train_mosaic_30k):
inputs , labels , fore_idx = data
inputs, labels, fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
optimizer.zero_grad()
outputs, alphas, avg_images = fore_net(inputs)
_, predicted = torch.max(outputs.data, 1)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
mini = 40
if cnt % mini == mini - 1: # print every 40 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, cnt + 1, running_loss / mini))
mini_loss.append(running_loss / mini)
running_loss = 0.0
cnt=cnt+1
if(np.average(mini_loss) <= 0.05):
break
print('Finished Training')
return fore_net, epoch
def testing(loader, fore_net):
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
outputs, alphas, avg_images = fore_net(inputs)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
count += 1
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
return correct, total, focus_true_pred_true, focus_false_pred_true, focus_true_pred_false, focus_false_pred_false, argmax_more_than_half
def enter_into(table, sno, correct, total, ftpt, ffpt, ftpf, ffpf, alpha_more_half , fg, bg, epoch = "NA"):
entry = []
entry = [sno,'fg = '+ str(fg),'bg = '+str(bg), epoch, total, correct,]
entry.append((100.0*correct/total))
entry.append((100 * ftpt / total))
entry.append( (100 * ffpt / total))
entry.append( ( 100 * ftpf / total))
entry.append( ( 100 * ffpf / total))
entry.append( alpha_more_half)
table.append(entry)
print(" ")
print("="*160)
print(tabulate(table, headers=['S.No.', 'fg_class','bg_class','Epoch used','total_points', 'correct','accuracy','FTPT', 'FFPT', 'FTPF', 'FFPF', 'avg_img > 0.5'] ) )
print(" ")
print("="*160)
return table
def add_average_entry(table):
entry =[]
entry = ['Avg', "","" ,"" ,"" , "",]
entry.append( np.mean(np.array(table)[:,6].astype(np.float)) )
entry.append( np.mean(np.array(table)[:,7].astype(np.float)) )
entry.append( np.mean(np.array(table)[:,8].astype(np.float)) )
entry.append( np.mean(np.array(table)[:,9].astype(np.float)) )
entry.append( np.mean(np.array(table)[:,10].astype(np.float)) )
entry.append( np.mean(np.array(table)[:,11].astype(np.float)) )
table.append(entry)
print(" ")
print("="*160)
print(tabulate(table, headers=['S.No.', 'fg_class','bg_class','Epoch used','total_points', 'correct','accuracy','FTPT', 'FFPT', 'FTPF', 'FFPF', 'avg_img > 0.5'] ) )
print(" ")
print("="*160)
return table
train_table=[]
test_table1=[]
test_table2=[]
test_table3=[]
test_table4=[]
fg = [fg1,fg2,fg3]
bg = list(set([0,1,2,3,4,5,6,7,8,9])-set(fg))
number_runs = 10
for i in range(number_runs):
fore_net = Module2().double()
fore_net = fore_net.to("cuda")
fore_net, epoch = training(train_loader_from_noise_train_mosaic_30k, fore_net)
correct, total, ftpt, ffpt, ftpf, ffpf, alpha_more_half = testing(train_loader_from_noise_train_mosaic_30k, fore_net)
train_table = enter_into(train_table, i+1, correct, total, ftpt, ffpt, ftpf, ffpf, alpha_more_half, fg, bg, str(epoch) )
correct, total, ftpt, ffpt, ftpf, ffpf, alpha_more_half = testing(test_loader_from_noise_train_mosaic_30k, fore_net)
test_table1 = enter_into(test_table1, i+1, correct, total, ftpt, ffpt, ftpf, ffpf, alpha_more_half , fg, bg )
correct, total, ftpt, ffpt, ftpf, ffpf, alpha_more_half = testing(test_loader_from_noise_test_mosaic_10k, fore_net)
test_table2 = enter_into(test_table2, i+1, correct, total, ftpt, ffpt, ftpf, ffpf, alpha_more_half, fg, bg )
correct, total, ftpt, ffpt, ftpf, ffpf, alpha_more_half = testing(test_loader_from_true_train_mosaic_30k, fore_net)
test_table3 = enter_into(test_table3, i+1, correct, total, ftpt, ffpt, ftpf, ffpf, alpha_more_half , fg, bg)
correct, total, ftpt, ffpt, ftpf, ffpf, alpha_more_half = testing(test_loader_from_true_train_mosaic_10k, fore_net)
test_table4 = enter_into(test_table4, i+1, correct, total, ftpt, ffpt, ftpf, ffpf, alpha_more_half, fg, bg )
train_table = add_average_entry(train_table)
test_table1 = add_average_entry(test_table1)
test_table2 = add_average_entry(test_table2)
test_table3 = add_average_entry(test_table3)
test_table4 = add_average_entry(test_table4)
# torch.save(fore_net.state_dict(),"/content/drive/My Drive/Research/mosaic_from_CIFAR_involving_bottop_eigen_vectors/fore_net_epoch"+str(epoch)+"_fg_used"+str(fg_used)+".pt")
```
| github_jupyter |
# <img style="float: left; padding-right: 10px; width: 45px" src="https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/iacs.png"> CS109B Data Science 2: Advanced Topics in Data Science
## Lecture 21: Adversarial Examples
**Harvard University**<br/>
**Spring 2019**<br/>
**Instructors:** Pavlos Protopapas and Mark Glickman<br/>
<hr style="height:2pt">
Code and ideas taken/borrowed/adapted from
Deep Learning From Basics to Practice, by Andrew Glassner, https://dlbasics.com, http://glassner.com, Python utilities for saving and loading files, mostly images and Keras model weights
#### Authors: Pavlos Protopapas, Amil Merchant, Alex Lin, Thomas Chang, ZiZi Zhang
```
#RUN THIS CELL
import requests
from IPython.core.display import HTML
styles = requests.get("https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/cs109.css").text
HTML(styles)
```
# Adversarial Examples
Adversarial examples for neural networks are inputs that are specifically meant to fool neural networks but be descernible to the human eye.
### Contents:
1. Datasets - MNIST
3. Cleverhans Integration
4. FGSM (non-targeted attack)
5. JSMA (targeted attack)
6. Exercise: Repeat process with CIFAR
### Datasets - MNIST / CIFAR
In this examples, we will explore some of these examples and a few attack methods on common datasets such as MNIST and CIFAR10. We first load the data in through Keras.
```
import numpy as np
from keras.datasets import mnist
from keras.datasets import cifar10
import matplotlib.pyplot as plt
import tensorflow as tf
import keras
session = tf.Session()
keras.backend.set_session(session)
(mn_x_train, mn_y_train), (mn_x_test, mn_y_test) = mnist.load_data()
```
As a refresher, MNIST is a dataset for black and white handwritten digits. This dataset is particularly simple, as we will see, neural networks can achieve very high levels of test accuracy.
Each image is 28 x 28, for a size of 784 for each input.
```
print ("Training Examples: %d" % len(mn_x_train))
print ("Test Examples: %d" % len(mn_x_test))
n_classes = 10
inds=np.array([mn_y_train==i for i in range(n_classes)])
f,ax=plt.subplots(2,5,figsize=(10,5))
ax=ax.flatten()
for i in range(n_classes):
ax[i].imshow(mn_x_train[np.argmax(inds[i])].reshape(28,28))
ax[i].set_title(str(i))
plt.show()
```
### Neural Network Training
Keras is a high level library which can be used to train neural network models. It simplies coding neural networks for the datasets, and as installed, uses tensorflow for the backend. We use Keras for its simplicity and because these models can easily be linked into the cleverhans library to generate adversarial examples.
We shall start with MNIST as the models and the results are easy to see. The second half of the notebook will repeat the results with CIFAR10. For MNIST, we will use a very simple neural network which takes in the 28 x 28 input, uses a single hidden layer of size 512, and goes uses dense connections to lead to the 10 output classes. This should be familiar and may seem too simple, we will will build up to more complex examples when we get to CIFAR10.
```
from keras import models
from keras import layers
network = models.Sequential()
network.add(layers.Dense(512, activation='relu', input_shape=(28 * 28,)))
network.add(layers.Dense(10, activation='softmax'))
network.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
```
The data must be pre-processed before analysis. We flatten the image and normalize the images so pixel values are between 0 and 1 instead of 0 and 255. The labels must also be inputted as one hot vectors, so we use built in keras functions.
```
train_images_1d = mn_x_train.reshape((60000, 28 * 28))
train_images_1d = train_images_1d.astype('float32') / 255
test_images_1d = mn_x_test.reshape((10000, 28 * 28))
test_images_1d = test_images_1d.astype('float32') / 255
from keras.utils import to_categorical #this just converts the labels to one-hot class
train_labels = to_categorical(mn_y_train)
test_labels = to_categorical(mn_y_test)
```
Training the network does not take long and can easily be done quickly on a CPU. Validation accuracy quickly rises to about 99%.
```
from keras.callbacks import ModelCheckpoint
h=network.fit(train_images_1d,
train_labels,
epochs=5,
batch_size=128,
shuffle=True,
callbacks=[ModelCheckpoint('tutorial_MNIST.h5',save_best_only=True)])
# summarize history for accuracy
plt.plot(h.history['acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train'], loc='upper left')
plt.show()
score, acc = network.evaluate(test_images_1d,
test_labels,
batch_size=128)
print ("Test Accuracy: %.5f" % acc)
network.save('tutorial_MNIST.h5')
```
### Cleverhans Integration
Cleverhans is a library written by researchers on adversarial examples, many of whom are with Google Brain. The library has wrappers that allow us to take the Keras model that we just made (or an already trained model) and create adversarial examples.
If the model has already been created and we do not want to recreate it, just run the code below to reload the model.
```
from keras.models import load_model
network = load_model('tutorial_MNIST.h5')
#%pip install -e git+https://github.com/tensorflow/cleverhans.git#egg=cleverhans
from cleverhans.utils_keras import KerasModelWrapper
wrap = KerasModelWrapper(network)
```
### FGSM
The **F**ast **G**radient **S**ign **M**ethod is a non-target attack. Using the parameters in the neural network we trained, the FGSM method calculates the $\nabla$ of the cost function for the particular input. By adding this gradient to the image times some parameter $\epsilon$, we know that the cost function will increase. If the gradient is large enough, then the predictor is likely to change.
```
x = tf.placeholder(tf.float32, shape=(None, 784))
y = tf.placeholder(tf.float32, shape=(None, 10))
```
As mentioned above, the $\epsilon \nabla$ is added to the image to create the adversarial example. $\epsilon$ is given by fgsm_rate in the code below. We chose this value since it leads low adversarial accuracy but the images are still easily discernible by eye. Too high of a value would make the images look like random noise, but too low values would leave the adversarial accuracy very high.
```
from cleverhans.attacks import FastGradientMethod
fgsm = FastGradientMethod(wrap, sess=session)
fgsm_rate = 0.08
fgsm_params = {'eps': fgsm_rate,'clip_min': 0.,'clip_max': 1.}
adv_x = fgsm.generate(x, **fgsm_params)
adv_x = tf.stop_gradient(adv_x)
adv_prob = network(adv_x)
fetches = [adv_prob]
fetches.append(adv_x)
outputs = session.run(fetches=fetches, feed_dict={x:test_images_1d})
adv_prob = outputs[0]
adv_examples = outputs[1]
adv_predicted = adv_prob.argmax(1)
adv_accuracy = np.mean(adv_predicted == mn_y_test)
print("Adversarial accuracy: %.5f" % adv_accuracy)
n_classes = 10
f,ax=plt.subplots(2,5,figsize=(10,5))
ax=ax.flatten()
for i in range(n_classes):
ax[i].imshow(adv_examples[i].reshape(28,28))
ax[i].set_title("Adv: %d, Label: %d" % (adv_predicted[i], mn_y_test[i]))
plt.show()
```
From the examples above, we can see that many of the examples are misclassified from the true labels. Since the changes are non-targetted, the adversarial labels do not seem to show significant trends but are clearly not the numbers in the picture.
### JSMA
The JSMA method is slightly more complicated than FGSM and full explanations can be found in the following paper: https://arxiv.org/abs/1511.07528. One of the main differences is that this method is that the method generates targeted examples. For example, given a 4, we could perturb the image to register as any digit we desire between 0 and 9.
```
x = tf.placeholder(tf.float32, shape=(None, 784))
y = tf.placeholder(tf.float32, shape=(None, 10))
results = np.zeros((10, 10000))
perturbations = np.zeros((10, 10000))
grid_shape = (10, 10, 28, 28, 1)
grid_data = np.zeros(grid_shape)
from cleverhans.attacks import SaliencyMapMethod
jsma = SaliencyMapMethod(wrap, sess=session)
jsma_params = {'theta': 1.,
'gamma': 0.1,
'clip_min': 0.,
'clip_max': 1.,
'y_target': None}
from cleverhans.utils import other_classes, grid_visual
for index in range(int(len(mn_x_test) / 100)):
sample = test_images_1d[index: index + 1]
current = mn_y_test[index]
target_classes = other_classes(10, current)
grid_data[current, current, :, :, :] = np.reshape(
sample, (28, 28, 1))
for target in target_classes:
one_hot_target = np.zeros((1, 10))
one_hot_target[0, target] = 1
jsma_params['y_target'] = one_hot_target
adv_x = jsma.generate_np(sample, **jsma_params)
grid_data[target, current, :, :, :] = np.reshape(
adv_x, (28, 28, 1))
if index % 10 == 0:
print(index)
print(sample.shape)
print(one_hot_target)
print(adv_x.shape)
plt.figure(figsize = (80, 80))
_ = grid_visual(grid_data)
```
Each row in the above plot shows an example for a particular starting class / digit. Using JSMA, we were able to generate examples that adversarially were designed to be predicted as $0, 1, \dots$ where each column represents the adversarial target class.
## CIFAR10
Below, you can repeat the attack mehtods described above for a neural network on CIFAR10. This library is decently larger and requires a more complex CNN to train properly.
```
import numpy as np
from keras.datasets import mnist
from keras.datasets import cifar10
import matplotlib.pyplot as plt
import tensorflow as tf
import keras
session = tf.Session()
keras.backend.set_session(session)
(c10_x_train, c10_y_train), (c10_x_test, c10_y_test) = cifar10.load_data()
```
As a refresher, CIFAR10 is a dataset for small color images which should be classifying into classes based on the object the image such as an airplane or boat.
Each image is 32 x 32 x 3, for a size of 3072 for each input.
```
print ("Training Examples: %d" % len(c10_x_train))
print ("Test Examples: %d" % len(c10_x_test))
n_classes = 10
inds=np.array([c10_y_train==i for i in range(n_classes)])
f,ax=plt.subplots(2,5,figsize=(10,5))
ax=ax.flatten()
for i in range(n_classes):
ax[i].imshow(c10_x_train[np.argmax(inds[i])].reshape(32,32,3))
ax[i].set_title(str(i))
plt.show()
```
| github_jupyter |
# SMA ROC Portfolio
1. The Security is above its 200-day moving average
2. The Security closes with sma_roc > 0, buy.
3. If the Security closes with sma_roc < 0, sell your long position.
(For a Portfolio of securities.)
```
import datetime
import matplotlib.pyplot as plt
import pandas as pd
from talib.abstract import *
import pinkfish as pf
import strategy
# Format price data.
pd.options.display.float_format = '{:0.2f}'.format
%matplotlib inline
# Set size of inline plots
'''note: rcParams can't be in same cell as import matplotlib
or %matplotlib inline
%matplotlib notebook: will lead to interactive plots embedded within
the notebook, you can zoom and resize the figure
%matplotlib inline: only draw static images in the notebook
'''
plt.rcParams["figure.figsize"] = (10, 7)
```
Yahoo finance cryptocurrencies:
https://finance.yahoo.com/cryptocurrencies/
10 largest Crypto currencies from 5 years ago:
https://coinmarketcap.com/historical/20160626/
10 largest Crypto currencies from 4 years ago:
https://coinmarketcap.com/historical/20170625/
10 largest Crypto currencies from 3 years ago:
https://coinmarketcap.com/historical/20180624/
10 largest Crypto currencies from 2 years ago:
https://coinmarketcap.com/historical/20190630/
Some global data
```
# Symbol Lists
BitCoin = ['BTC-USD']
CryptoCurrencies_2016 = ['BTC-USD', 'ETH-USD', 'XRP-USD', 'LTC-USD',
'XEM-USD', 'DASH-USD', 'MAID-USD', 'LSK-USD', 'DOGE-USD']
# 'DAO-USD' is a dead coin, so missing from above
CryptoCurrencies_2017 = ['BTC-USD', 'ETH-USD', 'XRP-USD', 'LTC-USD', 'ETC-USD',
'XEM-USD', 'MIOTA-USD', 'DASH-USD', 'BTS-USD']
# 'STRAT-USD' last trade date is 2020-11-18, so removed
CryptoCurrencies_2018 = ['BTC-USD', 'ETH-USD', 'XRP-USD', 'BCH-USD', 'EOS-USD',
'LTC-USD', 'XLM-USD', 'ADA-USD', 'TRX-USD', 'MIOTA-USD']
CryptoCurrencies_2019 = ['BTC-USD', 'ETH-USD', 'XRP-USD', 'LTC-USD', 'BCH-USD',
'EOS-USD', 'BNB-USD', 'USDT-USD', 'BSV-USD', 'CRO-USD']
Stocks_Bonds_Gold_Crypto = ['SPY', 'QQQ', 'TLT', 'GLD', 'BTC-USD']
# Set 'continuous_timeseries' : False (for mixed asset classes)
start_1900 = datetime.datetime(1900, 1, 1)
start_2016 = datetime.datetime(2016, 6, 26)
start_2017 = datetime.datetime(2017, 6, 25)
start_2018 = datetime.datetime(2018, 6, 24)
start_2019 = datetime.datetime(2019, 6, 30)
# Pick one of the above symbols and start pairs
symbols = CryptoCurrencies_2016
start = start_2016
capital = 10000
end = datetime.datetime.now()
# NOTE: Cryptocurrencies have 7 days a week timeseries. You can test them with
# their entire timeseries by setting stock_market_calendar=False. Alternatively,
# to trade with stock market calendar by setting stock_market_calendar=True.
# For mixed asset classes that include stocks or ETFs, you must set
# stock_market_calendar=True.
options = {
'use_adj' : False,
'use_cache' : True,
'use_continuous_calendar' : False,
'force_stock_market_calendar' : True,
'stop_loss_pct' : 1.0,
'margin' : 1,
'lookback' : 1,
'sma_timeperiod': 20,
'sma_pct_band': 0,
'use_regime_filter' : False,
'use_vola_weight' : True
}
```
Define Optimizations
```
# pick one
optimize_sma_timeperiod = False
optimize_sma_pct_band = True
# define SMAs ranges
if optimize_sma_timeperiod:
Xs = range(5, 40, 5)
Xs = [str(X) for X in Xs]
# define band ranges
elif optimize_sma_pct_band:
Xs = range(0, 11, 1)
Xs = [str(X) for X in Xs]
```
Run Strategy
```
strategies = pd.Series(dtype=object)
for X in Xs:
print(X, end=" ")
if optimize_sma_timeperiod:
options['sma_timeperiod'] = int(X)
elif optimize_sma_pct_band:
options['sma_pct_band'] = int(X)
strategies[X] = strategy.Strategy(symbols, capital, start, end, options)
strategies[X].run()
```
Summarize results
```
metrics = ('annual_return_rate',
'max_closed_out_drawdown',
'annualized_return_over_max_drawdown',
'best_month',
'worst_month',
'sharpe_ratio',
'sortino_ratio',
'monthly_std',
'pct_time_in_market',
'total_num_trades',
'pct_profitable_trades',
'avg_points')
df = pf.optimizer_summary(strategies, metrics)
df
```
Bar graphs
```
pf.optimizer_plot_bar_graph(df, 'annual_return_rate')
pf.optimizer_plot_bar_graph(df, 'sharpe_ratio')
pf.optimizer_plot_bar_graph(df, 'max_closed_out_drawdown')
```
Run Benchmark
```
s = strategies[Xs[0]]
benchmark = pf.Benchmark('BTC-USD', capital, s.start, s.end, use_adj=True)
benchmark.run()
```
Equity curve
```
if optimize_sma_timeperiod: Y = '20'
elif optimize_sma_pct_band: Y = '3'
pf.plot_equity_curve(strategies[Y].dbal, benchmark=benchmark.dbal)
labels = []
for strategy in strategies:
if optimize_sma_timeperiod:
label = strategy.options['sma_timeperiod']
elif optimize_sma_pct_band:
label = strategy.options['sma_pct_band']
labels.append(label)
pf.plot_equity_curves(strategies, labels)
```
| github_jupyter |
# Using GraphiPy to extract data from Tumblr
```
from graphipy.graphipy import GraphiPy
# create GraphiPy object (default to Pandas)
graphipy = GraphiPy()
```
# Creating the Tumblr Object
GraphiPy's Tumblr object needs CONSUMER_KEY, CONSUMER_SECRET, OAUTH_TOKEN, and OAUTH_SECRET in order to connect to Tumblr's API:
- To get the keys and the tokens, go to https://api.tumblr.com/console
```
# The tumblr API needs these credentials
CONSUMER_KEY = ''
CONSUMER_SECRET = ''
OAUTH_TOKEN = ''
OAUTH_SECRET = ''
tumblr_api_credentials = {
"consumer_key": CONSUMER_KEY,
"consumer_secret": CONSUMER_SECRET,
"oauth_token": OAUTH_TOKEN,
"oauth_secret": OAUTH_SECRET
}
# create the tumblr object
tumblr = graphipy.get_tumblr(tumblr_api_credentials)
```
# Find Posts by a Tag
### def fetch_posts_tagged(self, graph, tag, limit=20, before=0, filter="")
Optional parameters:
- limit: the number of results to return: 1–20, inclusive
- before: retrieve posts before the specified timestamp
Types of nodes returned:
- blog
- post
Types of edge returned:
- PUBLISHED
```
keyword = "python"
# Every function call modifies the graph that is sent as input
# tagged_posts = graphipy.create_graph()
# tumblr.fetch_posts_tagged(graph=tagged_posts, tag=keyword, limit=3)
# However, it also returns the graph modified so you can assign it to other variables like so:
tagged_posts = tumblr.fetch_posts_tagged(graph=graphipy.create_graph(), tag=keyword, limit=5)
# To get the list of available nodes
# There're posts and blogs in this case
print(tagged_posts.get_nodes().keys())
# You can get the dataframe from Pandas by specifying the node
tagged_posts_df = tagged_posts.get_df("post")
tagged_posts_df
# show the attributes of this node
tagged_posts_df.iloc[0]
# The same works with edges
pb_edges = tagged_posts.get_edges()
print(pb_edges.keys())
pb_edges["published"]
```
# Find a Blog by Blog Name
### def fetch_blog(self, graph, blog_name)
Type of node returned:
- blog
No edges returned
```
# Let's try searching for a blog
blog_to_search = tagged_posts_df.blog_name[0]
# Call the appropriate function
blog = tumblr.fetch_blog(graphipy.create_graph(), blog_name=blog_to_search)
# You can get the dataframe from Pandas by specifying the node (only 1 node in this case)
blog_df = blog.get_df("blog")
blog_df
```
# Find Blogs followed by a Given Blog
### def fetch_blogs_following(self, graph, blog_name, limit=20, offset=0)
Optional parameters:
- limit: the number of results to return: 1–20, inclusive
- offset: followed blog index to start at
Type of node returned:
- blog
Type of edge returned:
- FOLLOWING
```
# Let's use the blog name we used in the last section
blogs_following = tumblr.fetch_blogs_following(graphipy.create_graph(), blog_name=blog_to_search, limit=5)
# Grab the nodes
blogs_nodes = blogs_following.get_nodes()
print(blogs_nodes.keys())
# View the nodes
blogs_nodes["blog"]
# Grab the edges
bb_edges = blogs_following.get_edges()
print(bb_edges.keys())
# View the edges
bb_edges["following"]
```
# Find Posts Published by a Given Blog
### def fetch_published_posts(self, graph, blog_name, type=None, tag="", limit=20, offset=0)
Optional parameters:
- type: the type of post to return. Specify one of the following: text, quote, link, answer, video, audio, photo, chat
- tag: limits the response to posts with the specified tag
- limit: the number of posts to return: 1–20, inclusive
- offset: post number to start at
Types of node returned:
- blog
- post
Type of edge returned:
- PUBLISHED
```
# We can also see all the posts published by some blog we researched in previous sections
published_posts = tumblr.fetch_published_posts(graphipy.create_graph(), blog_name=blog_to_search, limit=5, type="photo")
# Grab the nodes
nodes = published_posts.get_nodes()
print(nodes.keys())
# View the posts
nodes["post"]
```
# Find Posts Liked by a Given Blog
### def fetch_liked_posts(self,graph,blog_name,limit=20,offset=0,before=0,after=0)
Optimal parameters:
- limit: the number of results to return: 1–20, inclusive
- offset: liked post number to start at
- before: retrieve posts liked before the specified timestamp
- after: retrieve posts liked after the specified timestamp
Types of nodes returned:
- blog
- post
Type of edge returned:
- LIKED
```
# We can also see all the posts published by the blog we researched in previous sections
liked_posts = tumblr.fetch_liked_posts(graphipy.create_graph(), blog_name=blog_to_search)
# Grab the nodes
nodes = liked_posts.get_nodes()
print(nodes.keys())
# View the posts
nodes["post"]
```
# Exporting Graph as CSV Files
#### For more information, see DataExportDemo.ipynb
```
# You can then export the graph into .csv files
# Just call .export_CSV_all() on the graph desired
csv_name = "tumblr"
export_path_all = liked_posts.export_all_csv(csv_name)
# You can also specify the dataframes "you want to export by using the .export_CSV() function
# Provide the label_attributes of the nodes and edges you want to export
csv_name = "specific"
nodes = {"post"}
edges = {"LIKED"}
export_path_specific = liked_posts.export_csv(csv_name, nodes, edges)
```
# Visualization with NetworkX
#### For more information, see DataExportDemo.ipynb
```
# We will visualize liked_posts graph
from matplotlib import colors as mcolors
import matplotlib.pyplot as plt
# Create the networkx exporter object
exporter = graphipy.get_nx_exporter()
nx_graph = exporter.create_from_pd(liked_posts)
# Draw the graph using graphipy
color_set = set(mcolors.CSS4_COLORS)
options = {
"node_label": "Label",
"colorful_edges": True,
"color_set": color_set
}
legend = exporter.draw_random(nx_graph, options=options, legend=plt)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import os
from IPython.core.display import display, HTML, clear_output
display(HTML("<style>.container { width:80% !important; }</style>"))
from scipy.sparse import csc_matrix
from sparsesvd import sparsesvd
```
# Import ratings data (including user data)
```
cwd = os.getcwd()
ratings = pd.read_csv(os.path.join(cwd, "..", "data", "ratings_combined_ryan.csv"))
ratings.head()
```
# Create Compressed Sparse Column matrix
```
N1 = ratings['userId'].nunique()
N2 = ratings['movieId'].nunique()
uids_raw = ratings['userId'].unique()
iids_raw = ratings['movieId'].unique()
uids_inner = np.arange(N1)
iids_inner = np.arange(N2)
uid_maptoraw = dict(zip(uids_inner, uids_raw))
uid_maptoinner = dict(zip(uids_raw, uids_inner))
iid_maptoraw = dict(zip(iids_inner, iids_raw))
iid_maptoinner = dict(zip(iids_raw, iids_inner))
ratings.rename(columns={'userId':'uid_raw', 'movieId':'iid_raw'}, inplace=True)
ratings.head()
ratings['uid_inner'] = ratings.apply(lambda x: uid_maptoinner[x['uid_raw']], axis=1)
ratings['iid_inner'] = ratings.apply(lambda x: iid_maptoinner[x['iid_raw']], axis=1)
compressed_matrix = csc_matrix((ratings['rating'], (ratings['uid_inner'], ratings['iid_inner'])), shape=(N1, N2))
```
# Perform SVD Matrix Factorization
```
ut, s, vt = sparsesvd(compressed_matrix, 50)
ut.shape, s.shape, vt.shape
```
# Make user predictions
```
user_uid_raw = ratings['uid_raw'].max(); print('user raw user id is: {}'.format(user_uid_raw))
user_uid_inner = uid_maptoinner[user_uid_raw]; print('user inner user id is: {}'.format(user_uid_inner))
```
### Dot product
```
s = np.diag(s)
preds = ut[:, user_uid_inner].dot(s).dot(vt)
preds.shape
preds = pd.DataFrame(preds, columns = ['predicted_rating'])
preds.reset_index(inplace=True)
preds.rename(inplace=True, columns={'index':'iid_inner'})
preds['iid_raw'] = preds.apply(lambda x: iid_maptoraw[x['iid_inner']], axis=1)
preds['predicted_rating'].describe()
```
# Join movie titles and genres
```
cwd = os.getcwd()
movies = pd.read_csv(os.path.join(cwd, "..", "data", "movies.csv"))
preds = pd.merge(preds, movies[['movieId', 'title', 'genres']], left_on='iid_raw', right_on='movieId')
preds = preds.sort_values('predicted_rating', ascending=False)
preds.head(20)
```
# Filter out movies that the user already rated
```
user_profile = pd.read_csv('ryan_profile.csv', index_col=0)
user_profile.head()
preds = pd.merge(preds, user_profile, on='movieId', how='left')
havent_seen_it_mask = preds['rating'].isnull()
preds[havent_seen_it_mask][['title', 'genres', 'predicted_rating']][:2000].to_csv('movie_recommendations_ryan_SVD.csv',
index=False)
preds[havent_seen_it_mask][['title', 'genres', 'predicted_rating']].head(30)
```
| github_jupyter |
## Scope
This notebook goes through how to create a training set from a set of questions and answers
## Optional: further research
* **OOB for cleaning text SpaCy** https://towardsdatascience.com/machine-learning-for-text-classification-using-spacy-in-python-b276b4051a49
* 1) BLUE SCORE FOR EVALUATION: https://machinelearningmastery.com/calculate-bleu-score-for-text-python/
* 2) Stanford SQuad (Question/Answer Scoring Datasets): https://github.com/obryanlouis/qa
* 3) https://github.com/facebookresearch/ParlAI
* 4) Seq2Seq: https://docs.chainer.org/en/stable/examples/seq2seq.html
## Initialize
```
# Data
import numpy as np
# Utility
import time
import math
import sys
import os
import glob
import pickle
# NLP
import tensorflow as tf
import re
import spacy
# Custom libraries
# from seq2seq_model import Seq2SeqModel
# from corpora_tools import *
```
## Import Data
```
def retrieve_corpora(path_to_pickle):
data = pickle.load(open(path_to_pickle,'rb'))
qa = []
for i in list(data):
qa.append((i[0]['utterance'],i[1]['utterance']))
# Split pairs into question and answer
question, answer = zip(*qa)
return question,answer
def tokenize(txt):
return str(txt).lower().split(' ')
def clean_sentences(txt):
# tokenize
tokenized_txt = tokenize(txt)
# Filter comments too long
return tokenized_txt[:20]
# Option 1: All responses are equal to the original query except for the comment by the author
# Zipped Q/A
question,answer = retrieve_corpora('../data/all_responses_equal.p')
idx = 1
print("Q:", question[idx])
print("A:", answer[idx])
```
## Create Trainable dataset
```
%%time
# Clean sentences
clean_question = [clean_sentences(s) for s in question]
clean_answer = [clean_sentences(s) for s in answer]
print(len(clean_question))
assert len(clean_question) == len(clean_answer)
idx = 2
print("Q:", clean_question[idx])
print("A:", clean_answer[idx])
def filter_sentence_length(sentences_l1, sentences_l2, min_len=0, max_len=20):
filtered_sentences_l1 = []
filtered_sentences_l2 = []
for i in range(len(sentences_l1)):
if min_len <= len(sentences_l1[i]) <= max_len and \
min_len <= len(sentences_l2[i]) <= max_len:
filtered_sentences_l1.append(sentences_l1[i])
filtered_sentences_l2.append(sentences_l2[i])
return filtered_sentences_l1, filtered_sentences_l2
import data_utils
def create_indexed_dictionary(sentences, dict_size=10000, storage_path=None):
count_words = Counter()
dict_words = {}
opt_dict_size = len(data_utils.OP_DICT_IDS)
for sen in sentences:
for word in sen:
count_words[word] += 1
dict_words[data_utils._PAD] = data_utils.PAD_ID
dict_words[data_utils._GO] = data_utils.GO_ID
dict_words[data_utils._EOS] = data_utils.EOS_ID
dict_words[data_utils._UNK] = data_utils.UNK_ID
for idx, item in enumerate(count_words.most_common(dict_size)):
dict_words[item[0]] = idx + opt_dict_size
if storage_path:
pickle.dump(dict_words, open(storage_path, "wb"))
return dict_words
def sentences_to_indexes(sentences, indexed_dictionary):
indexed_sentences = []
not_found_counter = 0
for sent in sentences:
idx_sent = []
for word in sent:
try:
idx_sent.append(indexed_dictionary[word])
except KeyError:
idx_sent.append(data_utils.UNK_ID)
not_found_counter += 1
indexed_sentences.append(idx_sent)
print('[sentences_to_indexes] Did not find {} words'.format(not_found_counter))
return indexed_sentences
def prepare_sentences(sentences_l1, sentences_l2, len_l1, len_l2):
assert len(sentences_l1) == len(sentences_l2)
data_set = []
for i in range(len(sentences_l1)):
padding_l1 = len_l1 - len(sentences_l1[i])
pad_sentence_l1 = ([data_utils.PAD_ID]*padding_l1) + sentences_l1[i]
padding_l2 = len_l2 - len(sentences_l2[i])
pad_sentence_l2 = [data_utils.GO_ID] + sentences_l2[i] + [data_utils.EOS_ID] + ([data_utils.PAD_ID] * padding_l2)
data_set.append([pad_sentence_l1, pad_sentence_l2])
return data_set
def extract_max_length(corpora):
return max([len(sentence) for sentence in corpora])
filt_clean_sen_l1, filt_clean_sen_l2 = filter_sentence_length(clean_question,
clean_answer)
print("# Filtered Corpora length (i.e. number of sentences)")
print(len(filt_clean_sen_l1))
assert len(filt_clean_sen_l1) == len(filt_clean_sen_l2)
dict_l1 = create_indexed_dictionary(filt_clean_sen_l1, dict_size=15000, storage_path="/tmp/l1_dict.p")
dict_l2 = create_indexed_dictionary(filt_clean_sen_l2, dict_size=15000, storage_path="/tmp/l2_dict.p")
idx_sentences_l1 = sentences_to_indexes(filt_clean_sen_l1, dict_l1)
idx_sentences_l2 = sentences_to_indexes(filt_clean_sen_l2, dict_l2)
print("# Same sentences as before, with their dictionary ID")
print("Q:", list(zip(filt_clean_sen_l1[0], idx_sentences_l1[0])))
print("A:", list(zip(filt_clean_sen_l2[0], idx_sentences_l2[0])))
max_length_l1 = extract_max_length(idx_sentences_l1)
max_length_l2 = extract_max_length(idx_sentences_l2)
print("# Max sentence sizes:")
print("DE:", max_length_l1)
print("EN:", max_length_l2)
```
As the final step, let's add paddings and markings to the sentences:
```
data_set = prepare_sentences(idx_sentences_l1, idx_sentences_l2, max_length_l1, max_length_l2)
print("# Prepared minibatch with paddings and extra stuff")
print("Q:", data_set[0][0])
print("A:", data_set[0][1])
print("# The sentence pass from X to Y tokens")
print("Q:", len(idx_sentences_l1[0]), "->", len(data_set[0][0]))
print("A:", len(idx_sentences_l2[0]), "->", len(data_set[0][1]))
```
After we're done with the corpora, it's now time to work on the model. This project requires again a sequence to sequence model, therefore we can use an RNN. Even more, we can reuse part of the code from the previous project: we'd just need to change how the dataset is built, and the parameters of the model. We can then copy the training script built in the previous chapter, and modify the build_dataset function, to use the Cornell dataset.
Mind that the dataset used in this chapter is bigger than the one used in the previous, therefore you may need to limit the corpora to a few dozen thousand lines. On a 4 years old laptop with 8GB RAM, we had to select only the first 30 thousand lines, otherwise, the program ran out of memory and kept swapping. As a side effect of having fewer examples, even the dictionaries are smaller, resulting in less than 10 thousands words each.
After a few iterations, you can stop the program and you'll see something similar to this output:
| github_jupyter |
Some rough fits to the sky surface brightness to use as inputs in the simulated spectra
```
import os
import h5py
import numpy as np
from scipy.optimize import curve_fit
import astropy.units as u
from feasibgs import util as UT
import desimodel.io
import desisim.simexp
import matplotlib as mpl
import matplotlib.pyplot as pl
mpl.rcParams['text.usetex'] = True
mpl.rcParams['font.family'] = 'serif'
mpl.rcParams['axes.linewidth'] = 1.5
mpl.rcParams['axes.xmargin'] = 1
mpl.rcParams['xtick.labelsize'] = 'x-large'
mpl.rcParams['xtick.major.size'] = 5
mpl.rcParams['xtick.major.width'] = 1.5
mpl.rcParams['ytick.labelsize'] = 'x-large'
mpl.rcParams['ytick.major.size'] = 5
mpl.rcParams['ytick.major.width'] = 1.5
mpl.rcParams['legend.frameon'] = False
%matplotlib inline
# ccd wavelength limit
params = desimodel.io.load_desiparams()
wavemin = params['ccd']['b']['wavemin']
wavemax = params['ccd']['z']['wavemax']
print('%f < lambda < %f' % (wavemin, wavemax))
```
Read in the bright sky spectrum produced from BOSS by Parker
```
dark_sky = np.loadtxt(UT.dat_dir()+'sky/spec-sky.dat', unpack=True, skiprows=2, usecols=[0,1])
f = UT.dat_dir()+'sky/moon_sky_spectrum.hdf5'
assert os.path.isfile(f)
f_hdf5 = h5py.File(f, 'r')
ws, ss = [], []
for i in range(4):
ws.append(f_hdf5['sky'+str(i)+'/wave'].value)
ss.append(f_hdf5['sky'+str(i)+'/sky'].value)
bright_sky_flux0= [10.*ws[2], ss[2]]
bright_sky_flux1 = [10.*ws[3], ss[3]]
```
Divide the sky flux by fiber area in $arcsec^2$. For BOSS the fiber area is $\pi\,\,arcsec^2$
```
bright_sky_sbright0 = [bright_sky_flux0[0], bright_sky_flux0[1]/np.pi]
bright_sky_sbright1 = [bright_sky_flux1[0], bright_sky_flux1[1]/np.pi]
```
Now read in the dark sky surface brightness from `spec-sky.dat`
```
waves = dark_sky[0][(dark_sky[0] > wavemin) & (dark_sky[1] < wavemax)] * u.Angstrom
config = desisim.simexp._specsim_config_for_wave((waves).to('Angstrom').value, specsim_config_file='desi')
atm_config = config.atmosphere
surface_brightness_dict = config.load_table(atm_config.sky, 'surface_brightness', as_dict=True)
fig = plt.figure(figsize=(8,4))
sub = fig.add_subplot(111)
sub.scatter(config.wavelength, surface_brightness_dict['dark'], c='k', lw=0, s=0.5, label='Dark Sky')
sub.scatter(bright_sky_sbright0[0], bright_sky_sbright0[1], c='C1', lw=0, s=1., label='Bright Sky')
sub.scatter(bright_sky_sbright1[0], bright_sky_sbright1[1], c='C1', lw=0, s=1.)
sub.set_xlabel('Wavelength', fontsize=20)
sub.set_xlim([3600., 9800.])
sub.set_ylabel('sky surface brightness [$10^{-17} erg/\AA/cm^2/s/arcsec^2$]', fontsize=20)
#sub.set_yscale("log")
sub.set_ylim([0.5, 3e1])
sub.legend(loc='upper left', markerscale=10, prop={'size':20})
def smooth(ww, sb):
''' smooth out the sufrace brightness somehow...
'''
wavebin = np.linspace(wavemin, wavemax, 10)
sb_med = np.zeros(len(wavebin)-1)
for i in range(len(wavebin)-1):
inwbin = ((wavebin[i] < ww) & (ww < wavebin[i+1]) & np.isfinite(sb))
if np.sum(inwbin) > 0.:
sb_med[i] = np.median(sb[inwbin])
return 0.5*(wavebin[1:]+wavebin[:-1]), sb_med
surface_brightness_dict['dark'].value
fig = plt.figure(figsize=(8,4))
sub = fig.add_subplot(111)
sub.scatter(config.wavelength, surface_brightness_dict['dark'], c='C0', lw=0, s=0.5, label='Dark Sky')
w_smooth_dark, sb_smooth_dark = smooth(config.wavelength.value, surface_brightness_dict['dark'].value)
sub.plot(w_smooth_dark, sb_smooth_dark, c='k', label='almost contiuum')
sub.set_xlabel('Wavelength', fontsize=20)
sub.set_xlim([3600., 9800.])
sub.set_ylabel('sky surface brightness [$10^{-17} erg/\AA/cm^2/s/arcsec^2$]', fontsize=20)
sub.set_yscale("log")
sub.set_ylim([0.5, 3e1])
sub.legend(loc='upper left', markerscale=10, prop={'size':20})
fig = plt.figure(figsize=(8,4))
sub = fig.add_subplot(111)
sub.scatter(bright_sky_sbright0[0], bright_sky_sbright0[1], c='C1', lw=0, s=1., label='Bright Sky')
w_smooth_bright0, sb_smooth_bright0 = smooth(bright_sky_sbright0[0], bright_sky_sbright0[1])
sub.plot(w_smooth_bright0, sb_smooth_bright0, c='k', label='almost contiuum')
sub.scatter(bright_sky_sbright1[0], bright_sky_sbright1[1], c='C1', lw=0, s=1.)
w_smooth_bright1, sb_smooth_bright1 = smooth(bright_sky_sbright1[0], bright_sky_sbright1[1])
sub.plot(w_smooth_bright1, sb_smooth_bright1, c='k')
sub.set_xlabel('Wavelength', fontsize=20)
sub.set_xlim([3600., 9800.])
sub.set_ylabel('sky surface brightness [$10^{-17} erg/\AA/cm^2/s/arcsec^2$]', fontsize=20)
sub.set_yscale("log")
sub.set_ylim([0.5, 3e1])
sub.legend(loc='upper left', markerscale=10, prop={'size':20})
```
combine the two smoothed bright sky surface brightnesses
```
sb_smooth_bright = np.zeros(len(w_smooth_bright0))
for i in range(len(w_smooth_bright0)):
if (sb_smooth_bright0[i] == 0) or (sb_smooth_bright1[i] == 0):
sb_smooth_bright[i] = np.max([sb_smooth_bright0[i], sb_smooth_bright1[i]])
else:
sb_smooth_bright[i] = np.mean([sb_smooth_bright0[i], sb_smooth_bright1[i]])
fig = plt.figure(figsize=(8,4))
sub = fig.add_subplot(111)
sub.scatter(w_smooth_dark, sb_smooth_dark, c='C0', lw=0, s=20.)#, label='Bright Sky')
sub.scatter(w_smooth_bright0, sb_smooth_bright, c='C1', lw=0, s=20.)
sub.set_xlabel('Wavelength', fontsize=20)
sub.set_xlim([3600., 9800.])
sub.set_ylabel('surface brightness [$10^{-17} erg/\AA/cm^2/s/arcsec^2$]', fontsize=20)
sub.set_yscale("log")
sub.set_ylim([0.5, 3e1])
sub.legend(loc='upper left', markerscale=10, prop={'size':20})
fig = plt.figure(figsize=(8,4))
sub = fig.add_subplot(111)
sub.scatter(w_smooth_dark, sb_smooth_bright - sb_smooth_dark, c='C1', lw=0, s=20.,
label='(Bright Sky) - (Dark Sky)')
sub.set_xlabel('Wavelength', fontsize=20)
sub.set_xlim([3600., 9800.])
sub.set_ylabel('surface brightness [$10^{-17} erg/\AA/cm^2/s/arcsec^2$]', fontsize=20)
sub.set_yscale("log")
sub.set_ylim([0.5, 3e1])
sub.legend(loc='upper right', prop={'size':20})
def SBright_resid(x, a, b, c):
return np.exp(b * (x-a) + c)
theta, _ = curve_fit(SBright_resid, w_smooth_dark/1000., sb_smooth_bright - sb_smooth_dark, p0=(10, -0.2, 1.))
print theta
```
best-fit of the residual surface brightness is
$$r(\lambda) = e^{-0.000488 (\lambda - 5071.) + 1.388}$$
```
fig = plt.figure(figsize=(8,4))
sub = fig.add_subplot(111)
sub.scatter(w_smooth_dark, sb_smooth_bright - sb_smooth_dark, c='C1', lw=0, s=20.,
label='(Bright Sky) - (Dark Sky)')
xarr = np.linspace(3, 12, 100)
sub.plot(1000.*xarr, SBright_resid(xarr, *theta), c='k')
sub.set_xlabel('Wavelength', fontsize=20)
sub.set_xlim([3600., 9800.])
sub.set_ylabel('surface brightness [$10^{-17} erg/\AA/cm^2/s/arcsec^2$]', fontsize=20)
sub.set_yscale("log")
sub.set_ylim([0.5, 3e1])
sub.legend(loc='upper right', prop={'size':20})
fig = plt.figure(figsize=(8,4))
sub = fig.add_subplot(111)
sub.scatter(config.wavelength, surface_brightness_dict['dark'], c='C0', lw=0, s=0.5, label='Dark Sky')
sub.scatter(bright_sky_sbright0[0], bright_sky_sbright0[1], c='C1', lw=0, s=1., label='Bright Sky')
sub.scatter(bright_sky_sbright1[0], bright_sky_sbright1[1], c='C1', lw=0, s=1.)
sub.scatter(config.wavelength,
surface_brightness_dict['dark'].value+SBright_resid(config.wavelength.value/1000., *theta), c='k', s=0.01)
#xarr = np.linspace(3, 12, 100)
#sub.plot(1000.*xarr, SBright_resid(xarr, *theta), c='k')
sub.set_xlabel('Wavelength', fontsize=20)
sub.set_xlim([3600., 9800.])
sub.set_ylabel('surface brightness [$10^{-17} erg/\AA/cm^2/s/arcsec^2$]', fontsize=20)
sub.set_yscale("log")
sub.set_ylim([0.5, 3e1])
sub.legend(loc='upper right', prop={'size':20})
```
looks pretty good.
| github_jupyter |
```
import numpy as np
from sklearn.datasets.samples_generator import make_blobs
import pandas as pd
import matplotlib.pyplot as plt
%%markdown
# k-means
samples = np.array([[1,2], [12,2], [0,1], [10,0], [9,1], \
[8,2], [0,10], [1,8], [2,9], [9,9], \
[10,8], [8,9] ], dtype = np.float)
centers = np.array([[3,2], [2,6], [9,3], [7,6]], dtype = np.float)
N = len(samples)
fig, ax = plt.subplots()
ax.scatter (samples.transpose()[0], samples.transpose()[1], marker = 'o', s = 100)
ax.scatter (centers.transpose()[0], centers.transpose()[1], marker = 's', s = 100, color='black')
plt.plot()
def distance (sample, centroids):
distances = np.zeros(len(centroids))
for i in range(0,len(centroids)):
distances[i] = np.sqrt (sum (pow (np.subtract (sample, centroids[i]), 2)))
return distances
def showcurrentstatus (samples, centers, clusters, plotnumber):
plt.subplot (620 + plotnumber)
plt.scatter (samples.transpose()[0], samples.transpose()[1], marker = 'o', s = 150, c = clusters)
plt.scatter (centers.transpose()[0], centers.transpose()[1], marker = 's', s = 100, color = 'black')
plt.plot()
def kmeans(centroids, samples, K, plotresults):
N = len(samples)
plt.figure (figsize=(20,20))
distances = np.zeros ((N,K))
new_centroids = np.zeros ((K, 2))
final_centroids = np.zeros ((K, 2))
clusters = np.zeros (N, np.int)
for i in range (0, N):
distances[i] = distance(samples[i], centroids)
clusters[i] = np.argmin(distances[i])
new_centroids[clusters[i]] += samples[i]
divisor = np.bincount(clusters).astype(np.float)
divisor.resize([K])
for j in range(0, K):
final_centroids[j] = np.nan_to_num(np.divide(new_centroids[j], divisor[j]))
if (i > 3 and plotresults == True):
showcurrentstatus(samples[:i], final_centroids, clusters[:i], i-3)
return final_centroids
finalcenters = kmeans (centers, samples, 4, True)
%%markdown
# k-Nearest Neighbors (k-NN)
data, features = make_blobs (n_samples=100, n_features = 2, centers=4, shuffle=True, cluster_std=0.8)
fig, ax = plt.subplots()
ax.scatter(data.transpose()[0], data.transpose()[1], c=features,marker = 'o', s = 100)
plt.plot()
def distance_knn (sample, data):
distances = np.zeros (len(data))
for i in range (0,len(data)):
dist = np.sqrt(sum(pow(np.subtract(sample, data[i]),2)))
distances[i] = dist
return distances
def add_sample (newsample, data, features):
distances = np.zeros ((len(data),len(data[0])))
# calculate the distance of the new sample and the current data
distances = distance_knn (newsample, data)
closestneighbors = np.argpartition (distances, 3)[:3]
closestgroups = features [closestneighbors]
return np.argmax (np.bincount (closestgroups))
def knn (newdata, data, features):
for i in newdata:
test = add_sample (i, data, features);
features = np.append (features, [test], axis = 0)
data = np.append (data, [i], axis = 0)
return (data, features)
newsamples = np.random.rand(20, 2) * 20 - 8
#newsamples
finaldata, finalfeatures = knn (newsamples, data, features)
fig, ax = plt.subplots()
ax.scatter (finaldata.transpose()[0], finaldata.transpose()[1], c = finalfeatures,marker = 'o', s = 100)
ax.scatter (newsamples.transpose()[0], newsamples.transpose()[1], c = 'none', marker = 's', s = 100)
plt.plot()
```
| github_jupyter |
# Serialization - saving, loading and checkpointing
At this point we've already covered quite a lot of ground.
We know how to manipulate data and labels.
We know how to construct flexible models capable of expressing plausible hypotheses.
We know how to fit those models to our dataset.
We know of loss functions to use for classification and for regression,
and we know how to minimize those losses with respect to our models' parameters.
We even know how to write our own neural network layers in ``gluon``.
But even with all this knowledge, we're not ready to build a real machine learning system.
That's because we haven't yet covered how to save and load models.
In reality, we often train a model on one device
and then want to run it to make predictions on many devices simultaneously.
In order for our models to persist beyond the execution of a single Python script,
we need mechanisms to save and load NDArrays, ``gluon`` Parameters, and models themselves.
```
from __future__ import print_function
import mxnet as mx
from mxnet import nd, autograd
from mxnet import gluon
ctx = mx.cpu()
# ctx = mx.gpu()
```
## Saving and loading NDArrays
To start, let's show how you can save and load a list of NDArrays for future use. Note that while it's possible to use a general Python serialization package like ``Pickle``, it's not optimized for use with NDArrays and will be unnecessarily slow. We prefer to use ``ndarray.save`` and ``ndarray.load``.
```
X = nd.ones((100, 100))
Y = nd.zeros((100, 100))
import os
dir_name = 'checkpoints'
if not os.path.exists(dir_name):
os.makedirs(dir_name)
filename = os.path.join(dir_name, "test1.params")
nd.save(filename, [X, Y])
```
It's just as easy to load a saved NDArray.
```
A, B = nd.load(filename)
print(A)
print(B)
```
We can also save a dictionary where the keys are strings and the values are NDArrays.
```
mydict = {"X": X, "Y": Y}
filename = os.path.join(dir_name, "test2.params")
nd.save(filename, mydict)
C = nd.load(filename)
print(C)
```
## Saving and loading the parameters of ``gluon`` models
Recall from [our first look at the plumbing behind ``gluon`` blocks](P03.5-C01-plumbing.ipynb])
that ``gluon`` wraps the NDArrays corresponding to model parameters in ``Parameter`` objects.
We'll often want to store and load an entire model's parameters without
having to individually extract or load the NDarrays from the Parameters via ParameterDicts in each block.
Fortunately, ``gluon`` blocks make our lives very easy by providing a ``.save_parameters()`` and ``.load_parameters()`` methods. To see them in work, let's just spin up a simple MLP.
```
num_hidden = 256
num_outputs = 1
net = gluon.nn.Sequential()
with net.name_scope():
net.add(gluon.nn.Dense(num_hidden, activation="relu"))
net.add(gluon.nn.Dense(num_hidden, activation="relu"))
net.add(gluon.nn.Dense(num_outputs))
```
Now, let's initialize the parameters by attaching an initializer and actually passing in a datapoint to induce shape inference.
```
net.collect_params().initialize(mx.init.Normal(sigma=1.), ctx=ctx)
net(nd.ones((1, 100), ctx=ctx))
```
So this randomly initialized model maps a 100-dimensional vector of all ones to the number 362.53 (that's the number on my machine--your mileage may vary).
Let's save the parameters, instantiate a new network, load them in and make sure that we get the same result.
```
filename = os.path.join(dir_name, "testnet.params")
net.save_parameters(filename)
net2 = gluon.nn.Sequential()
with net2.name_scope():
net2.add(gluon.nn.Dense(num_hidden, activation="relu"))
net2.add(gluon.nn.Dense(num_hidden, activation="relu"))
net2.add(gluon.nn.Dense(num_outputs))
net2.load_parameters(filename, ctx=ctx)
net2(nd.ones((1, 100), ctx=ctx))
```
Great! Now we're ready to save our work.
The practice of saving models is sometimes called *checkpointing*
and it's especially important for a number of reasons.
1. We can preserve and syndicate models that are trained once.
2. Some models perform best (as determined on validation data) at some epoch in the middle of training. If we checkpoint the model after each epoch, we can later select the best epoch.
3. We might want to ask questions about our trained model that we didn't think of when we first wrote the scripts for our experiments. Having the parameters lying around allows us to examine our past work without having to train from scratch.
4. Sometimes people might want to run our models who don't know how to execute training themselves or can't access a suitable dataset for training. Checkpointing gives us a way to share our work with others.
<!-- ## Serializing models themselves
[PLACEHOLDER] -->
## Next
[Convolutional neural networks from scratch](../chapter04_convolutional-neural-networks/cnn-scratch.ipynb)
For whinges or inquiries, [open an issue on GitHub.](https://github.com/zackchase/mxnet-the-straight-dope)
| github_jupyter |
**Math - Linear Algebra**
*Linear Algebra is the branch of mathematics that studies [vector spaces](https://en.wikipedia.org/wiki/Vector_space) and linear transformations between vector spaces, such as rotating a shape, scaling it up or down, translating it (ie. moving it), etc.*
*Machine Learning relies heavily on Linear Algebra, so it is essential to understand what vectors and matrices are, what operations you can perform with them, and how they can be useful.*
<table align="left">
<td>
<a href="https://colab.research.google.com/github/ageron/handson-ml2/blob/master/math_linear_algebra.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
</td>
<td>
<a target="_blank" href="https://kaggle.com/kernels/welcome?src=https://github.com/ageron/handson-ml2/blob/master/math_linear_algebra.ipynb"><img src="https://kaggle.com/static/images/open-in-kaggle.svg" /></a>
</td>
</table>
# Vectors
## Definition
A vector is a quantity defined by a magnitude and a direction. For example, a rocket's velocity is a 3-dimensional vector: its magnitude is the speed of the rocket, and its direction is (hopefully) up. A vector can be represented by an array of numbers called *scalars*. Each scalar corresponds to the magnitude of the vector with regards to each dimension.
For example, say the rocket is going up at a slight angle: it has a vertical speed of 5,000 m/s, and also a slight speed towards the East at 10 m/s, and a slight speed towards the North at 50 m/s. The rocket's velocity may be represented by the following vector:
**velocity** $= \begin{pmatrix}
10 \\
50 \\
5000 \\
\end{pmatrix}$
Note: by convention vectors are generally presented in the form of columns. Also, vector names are generally lowercase to distinguish them from matrices (which we will discuss below) and in bold (when possible) to distinguish them from simple scalar values such as ${meters\_per\_second} = 5026$.
A list of N numbers may also represent the coordinates of a point in an N-dimensional space, so it is quite frequent to represent vectors as simple points instead of arrows. A vector with 1 element may be represented as an arrow or a point on an axis, a vector with 2 elements is an arrow or a point on a plane, a vector with 3 elements is an arrow or point in space, and a vector with N elements is an arrow or a point in an N-dimensional space… which most people find hard to imagine.
## Purpose
Vectors have many purposes in Machine Learning, most notably to represent observations and predictions. For example, say we built a Machine Learning system to classify videos into 3 categories (good, spam, clickbait) based on what we know about them. For each video, we would have a vector representing what we know about it, such as:
**video** $= \begin{pmatrix}
10.5 \\
5.2 \\
3.25 \\
7.0
\end{pmatrix}$
This vector could represent a video that lasts 10.5 minutes, but only 5.2% viewers watch for more than a minute, it gets 3.25 views per day on average, and it was flagged 7 times as spam. As you can see, each axis may have a different meaning.
Based on this vector our Machine Learning system may predict that there is an 80% probability that it is a spam video, 18% that it is clickbait, and 2% that it is a good video. This could be represented as the following vector:
**class_probabilities** $= \begin{pmatrix}
0.80 \\
0.18 \\
0.02
\end{pmatrix}$
## Vectors in python
In python, a vector can be represented in many ways, the simplest being a regular python list of numbers:
```
[10.5, 5.2, 3.25, 7.0]
```
Since we plan to do quite a lot of scientific calculations, it is much better to use NumPy's `ndarray`, which provides a lot of convenient and optimized implementations of essential mathematical operations on vectors (for more details about NumPy, check out the [NumPy tutorial](tools_numpy.ipynb)). For example:
```
import numpy as np
video = np.array([10.5, 5.2, 3.25, 7.0])
video
```
The size of a vector can be obtained using the `size` attribute:
```
video.size
```
The $i^{th}$ element (also called *entry* or *item*) of a vector $\textbf{v}$ is noted $\textbf{v}_i$.
Note that indices in mathematics generally start at 1, but in programming they usually start at 0. So to access $\textbf{video}_3$ programmatically, we would write:
```
video[2] # 3rd element
```
## Plotting vectors
To plot vectors we will use matplotlib, so let's start by importing it (for details about matplotlib, check the [matplotlib tutorial](tools_matplotlib.ipynb)):
```
%matplotlib inline
import matplotlib.pyplot as plt
```
### 2D vectors
Let's create a couple very simple 2D vectors to plot:
```
u = np.array([2, 5])
v = np.array([3, 1])
```
These vectors each have 2 elements, so they can easily be represented graphically on a 2D graph, for example as points:
```
x_coords, y_coords = zip(u, v)
plt.scatter(x_coords, y_coords, color=["r","b"])
plt.axis([0, 9, 0, 6])
plt.grid()
plt.show()
```
Vectors can also be represented as arrows. Let's create a small convenience function to draw nice arrows:
```
def plot_vector2d(vector2d, origin=[0, 0], **options):
return plt.arrow(origin[0], origin[1], vector2d[0], vector2d[1],
head_width=0.2, head_length=0.3, length_includes_head=True,
**options)
```
Now let's draw the vectors **u** and **v** as arrows:
```
plot_vector2d(u, color="r")
plot_vector2d(v, color="b")
plt.axis([0, 9, 0, 6])
plt.grid()
plt.show()
```
### 3D vectors
Plotting 3D vectors is also relatively straightforward. First let's create two 3D vectors:
```
a = np.array([1, 2, 8])
b = np.array([5, 6, 3])
```
Now let's plot them using matplotlib's `Axes3D`:
```
from mpl_toolkits.mplot3d import Axes3D
subplot3d = plt.subplot(111, projection='3d')
x_coords, y_coords, z_coords = zip(a,b)
subplot3d.scatter(x_coords, y_coords, z_coords)
subplot3d.set_zlim3d([0, 9])
plt.show()
```
It is a bit hard to visualize exactly where in space these two points are, so let's add vertical lines. We'll create a small convenience function to plot a list of 3d vectors with vertical lines attached:
```
def plot_vectors3d(ax, vectors3d, z0, **options):
for v in vectors3d:
x, y, z = v
ax.plot([x,x], [y,y], [z0, z], color="gray", linestyle='dotted', marker=".")
x_coords, y_coords, z_coords = zip(*vectors3d)
ax.scatter(x_coords, y_coords, z_coords, **options)
subplot3d = plt.subplot(111, projection='3d')
subplot3d.set_zlim([0, 9])
plot_vectors3d(subplot3d, [a,b], 0, color=("r","b"))
plt.show()
```
## Norm
The norm of a vector $\textbf{u}$, noted $\left \Vert \textbf{u} \right \|$, is a measure of the length (a.k.a. the magnitude) of $\textbf{u}$. There are multiple possible norms, but the most common one (and the only one we will discuss here) is the Euclidian norm, which is defined as:
$\left \Vert \textbf{u} \right \| = \sqrt{\sum_{i}{\textbf{u}_i}^2}$
We could implement this easily in pure python, recalling that $\sqrt x = x^{\frac{1}{2}}$
```
def vector_norm(vector):
squares = [element**2 for element in vector]
return sum(squares)**0.5
print("||", u, "|| =")
vector_norm(u)
```
However, it is much more efficient to use NumPy's `norm` function, available in the `linalg` (**Lin**ear **Alg**ebra) module:
```
import numpy.linalg as LA
LA.norm(u)
```
Let's plot a little diagram to confirm that the length of vector $\textbf{v}$ is indeed $\approx5.4$:
```
radius = LA.norm(u)
plt.gca().add_artist(plt.Circle((0,0), radius, color="#DDDDDD"))
plot_vector2d(u, color="red")
plt.axis([0, 8.7, 0, 6])
plt.grid()
plt.show()
```
Looks about right!
## Addition
Vectors of same size can be added together. Addition is performed *elementwise*:
```
print(" ", u)
print("+", v)
print("-"*10)
u + v
```
Let's look at what vector addition looks like graphically:
```
plot_vector2d(u, color="r")
plot_vector2d(v, color="b")
plot_vector2d(v, origin=u, color="b", linestyle="dotted")
plot_vector2d(u, origin=v, color="r", linestyle="dotted")
plot_vector2d(u+v, color="g")
plt.axis([0, 9, 0, 7])
plt.text(0.7, 3, "u", color="r", fontsize=18)
plt.text(4, 3, "u", color="r", fontsize=18)
plt.text(1.8, 0.2, "v", color="b", fontsize=18)
plt.text(3.1, 5.6, "v", color="b", fontsize=18)
plt.text(2.4, 2.5, "u+v", color="g", fontsize=18)
plt.grid()
plt.show()
```
Vector addition is **commutative**, meaning that $\textbf{u} + \textbf{v} = \textbf{v} + \textbf{u}$. You can see it on the previous image: following $\textbf{u}$ *then* $\textbf{v}$ leads to the same point as following $\textbf{v}$ *then* $\textbf{u}$.
Vector addition is also **associative**, meaning that $\textbf{u} + (\textbf{v} + \textbf{w}) = (\textbf{u} + \textbf{v}) + \textbf{w}$.
If you have a shape defined by a number of points (vectors), and you add a vector $\textbf{v}$ to all of these points, then the whole shape gets shifted by $\textbf{v}$. This is called a [geometric translation](https://en.wikipedia.org/wiki/Translation_%28geometry%29):
```
t1 = np.array([2, 0.25])
t2 = np.array([2.5, 3.5])
t3 = np.array([1, 2])
x_coords, y_coords = zip(t1, t2, t3, t1)
plt.plot(x_coords, y_coords, "c--", x_coords, y_coords, "co")
plot_vector2d(v, t1, color="r", linestyle=":")
plot_vector2d(v, t2, color="r", linestyle=":")
plot_vector2d(v, t3, color="r", linestyle=":")
t1b = t1 + v
t2b = t2 + v
t3b = t3 + v
x_coords_b, y_coords_b = zip(t1b, t2b, t3b, t1b)
plt.plot(x_coords_b, y_coords_b, "b-", x_coords_b, y_coords_b, "bo")
plt.text(4, 4.2, "v", color="r", fontsize=18)
plt.text(3, 2.3, "v", color="r", fontsize=18)
plt.text(3.5, 0.4, "v", color="r", fontsize=18)
plt.axis([0, 6, 0, 5])
plt.grid()
plt.show()
```
Finally, subtracting a vector is like adding the opposite vector.
## Multiplication by a scalar
Vectors can be multiplied by scalars. All elements in the vector are multiplied by that number, for example:
```
print("1.5 *", u, "=")
1.5 * u
```
Graphically, scalar multiplication results in changing the scale of a figure, hence the name *scalar*. The distance from the origin (the point at coordinates equal to zero) is also multiplied by the scalar. For example, let's scale up by a factor of `k = 2.5`:
```
k = 2.5
t1c = k * t1
t2c = k * t2
t3c = k * t3
plt.plot(x_coords, y_coords, "c--", x_coords, y_coords, "co")
plot_vector2d(t1, color="r")
plot_vector2d(t2, color="r")
plot_vector2d(t3, color="r")
x_coords_c, y_coords_c = zip(t1c, t2c, t3c, t1c)
plt.plot(x_coords_c, y_coords_c, "b-", x_coords_c, y_coords_c, "bo")
plot_vector2d(k * t1, color="b", linestyle=":")
plot_vector2d(k * t2, color="b", linestyle=":")
plot_vector2d(k * t3, color="b", linestyle=":")
plt.axis([0, 9, 0, 9])
plt.grid()
plt.show()
```
As you might guess, dividing a vector by a scalar is equivalent to multiplying by its multiplicative inverse (reciprocal):
$\dfrac{\textbf{u}}{\lambda} = \dfrac{1}{\lambda} \times \textbf{u}$
Scalar multiplication is **commutative**: $\lambda \times \textbf{u} = \textbf{u} \times \lambda$.
It is also **associative**: $\lambda_1 \times (\lambda_2 \times \textbf{u}) = (\lambda_1 \times \lambda_2) \times \textbf{u}$.
Finally, it is **distributive** over addition of vectors: $\lambda \times (\textbf{u} + \textbf{v}) = \lambda \times \textbf{u} + \lambda \times \textbf{v}$.
## Zero, unit and normalized vectors
* A **zero-vector ** is a vector full of 0s.
* A **unit vector** is a vector with a norm equal to 1.
* The **normalized vector** of a non-null vector $\textbf{u}$, noted $\hat{\textbf{u}}$, is the unit vector that points in the same direction as $\textbf{u}$. It is equal to: $\hat{\textbf{u}} = \dfrac{\textbf{u}}{\left \Vert \textbf{u} \right \|}$
```
plt.gca().add_artist(plt.Circle((0,0),1,color='c'))
plt.plot(0, 0, "ko")
plot_vector2d(v / LA.norm(v), color="k")
plot_vector2d(v, color="b", linestyle=":")
plt.text(0.3, 0.3, "$\hat{u}$", color="k", fontsize=18)
plt.text(1.5, 0.7, "$u$", color="b", fontsize=18)
plt.axis([-1.5, 5.5, -1.5, 3.5])
plt.grid()
plt.show()
```
## Dot product
### Definition
The dot product (also called *scalar product* or *inner product* in the context of the Euclidian space) of two vectors $\textbf{u}$ and $\textbf{v}$ is a useful operation that comes up fairly often in linear algebra. It is noted $\textbf{u} \cdot \textbf{v}$, or sometimes $⟨\textbf{u}|\textbf{v}⟩$ or $(\textbf{u}|\textbf{v})$, and it is defined as:
$\textbf{u} \cdot \textbf{v} = \left \Vert \textbf{u} \right \| \times \left \Vert \textbf{v} \right \| \times cos(\theta)$
where $\theta$ is the angle between $\textbf{u}$ and $\textbf{v}$.
Another way to calculate the dot product is:
$\textbf{u} \cdot \textbf{v} = \sum_i{\textbf{u}_i \times \textbf{v}_i}$
### In python
The dot product is pretty simple to implement:
```
def dot_product(v1, v2):
return sum(v1i * v2i for v1i, v2i in zip(v1, v2))
dot_product(u, v)
```
But a *much* more efficient implementation is provided by NumPy with the `dot` function:
```
np.dot(u,v)
```
Equivalently, you can use the `dot` method of `ndarray`s:
```
u.dot(v)
```
**Caution**: the `*` operator will perform an *elementwise* multiplication, *NOT* a dot product:
```
print(" ",u)
print("* ",v, "(NOT a dot product)")
print("-"*10)
u * v
```
### Main properties
* The dot product is **commutative**: $\textbf{u} \cdot \textbf{v} = \textbf{v} \cdot \textbf{u}$.
* The dot product is only defined between two vectors, not between a scalar and a vector. This means that we cannot chain dot products: for example, the expression $\textbf{u} \cdot \textbf{v} \cdot \textbf{w}$ is not defined since $\textbf{u} \cdot \textbf{v}$ is a scalar and $\textbf{w}$ is a vector.
* This also means that the dot product is **NOT associative**: $(\textbf{u} \cdot \textbf{v}) \cdot \textbf{w} ≠ \textbf{u} \cdot (\textbf{v} \cdot \textbf{w})$ since neither are defined.
* However, the dot product is **associative with regards to scalar multiplication**: $\lambda \times (\textbf{u} \cdot \textbf{v}) = (\lambda \times \textbf{u}) \cdot \textbf{v} = \textbf{u} \cdot (\lambda \times \textbf{v})$
* Finally, the dot product is **distributive** over addition of vectors: $\textbf{u} \cdot (\textbf{v} + \textbf{w}) = \textbf{u} \cdot \textbf{v} + \textbf{u} \cdot \textbf{w}$.
### Calculating the angle between vectors
One of the many uses of the dot product is to calculate the angle between two non-zero vectors. Looking at the dot product definition, we can deduce the following formula:
$\theta = \arccos{\left ( \dfrac{\textbf{u} \cdot \textbf{v}}{\left \Vert \textbf{u} \right \| \times \left \Vert \textbf{v} \right \|} \right ) }$
Note that if $\textbf{u} \cdot \textbf{v} = 0$, it follows that $\theta = \dfrac{π}{2}$. In other words, if the dot product of two non-null vectors is zero, it means that they are orthogonal.
Let's use this formula to calculate the angle between $\textbf{u}$ and $\textbf{v}$ (in radians):
```
def vector_angle(u, v):
cos_theta = u.dot(v) / LA.norm(u) / LA.norm(v)
return np.arccos(np.clip(cos_theta, -1, 1))
theta = vector_angle(u, v)
print("Angle =", theta, "radians")
print(" =", theta * 180 / np.pi, "degrees")
```
Note: due to small floating point errors, `cos_theta` may be very slightly outside of the $[-1, 1]$ interval, which would make `arccos` fail. This is why we clipped the value within the range, using NumPy's `clip` function.
### Projecting a point onto an axis
The dot product is also very useful to project points onto an axis. The projection of vector $\textbf{v}$ onto $\textbf{u}$'s axis is given by this formula:
$\textbf{proj}_{\textbf{u}}{\textbf{v}} = \dfrac{\textbf{u} \cdot \textbf{v}}{\left \Vert \textbf{u} \right \| ^2} \times \textbf{u}$
Which is equivalent to:
$\textbf{proj}_{\textbf{u}}{\textbf{v}} = (\textbf{v} \cdot \hat{\textbf{u}}) \times \hat{\textbf{u}}$
```
u_normalized = u / LA.norm(u)
proj = v.dot(u_normalized) * u_normalized
plot_vector2d(u, color="r")
plot_vector2d(v, color="b")
plot_vector2d(proj, color="k", linestyle=":")
plt.plot(proj[0], proj[1], "ko")
plt.plot([proj[0], v[0]], [proj[1], v[1]], "b:")
plt.text(1, 2, "$proj_u v$", color="k", fontsize=18)
plt.text(1.8, 0.2, "$v$", color="b", fontsize=18)
plt.text(0.8, 3, "$u$", color="r", fontsize=18)
plt.axis([0, 8, 0, 5.5])
plt.grid()
plt.show()
```
# Matrices
A matrix is a rectangular array of scalars (ie. any number: integer, real or complex) arranged in rows and columns, for example:
\begin{bmatrix} 10 & 20 & 30 \\ 40 & 50 & 60 \end{bmatrix}
You can also think of a matrix as a list of vectors: the previous matrix contains either 2 horizontal 3D vectors or 3 vertical 2D vectors.
Matrices are convenient and very efficient to run operations on many vectors at a time. We will also see that they are great at representing and performing linear transformations such rotations, translations and scaling.
## Matrices in python
In python, a matrix can be represented in various ways. The simplest is just a list of python lists:
```
[
[10, 20, 30],
[40, 50, 60]
]
```
A much more efficient way is to use the NumPy library which provides optimized implementations of many matrix operations:
```
A = np.array([
[10,20,30],
[40,50,60]
])
A
```
By convention matrices generally have uppercase names, such as $A$.
In the rest of this tutorial, we will assume that we are using NumPy arrays (type `ndarray`) to represent matrices.
## Size
The size of a matrix is defined by its number of rows and number of columns. It is noted $rows \times columns$. For example, the matrix $A$ above is an example of a $2 \times 3$ matrix: 2 rows, 3 columns. Caution: a $3 \times 2$ matrix would have 3 rows and 2 columns.
To get a matrix's size in NumPy:
```
A.shape
```
**Caution**: the `size` attribute represents the number of elements in the `ndarray`, not the matrix's size:
```
A.size
```
## Element indexing
The number located in the $i^{th}$ row, and $j^{th}$ column of a matrix $X$ is sometimes noted $X_{i,j}$ or $X_{ij}$, but there is no standard notation, so people often prefer to explicitely name the elements, like this: "*let $X = (x_{i,j})_{1 ≤ i ≤ m, 1 ≤ j ≤ n}$*". This means that $X$ is equal to:
$X = \begin{bmatrix}
x_{1,1} & x_{1,2} & x_{1,3} & \cdots & x_{1,n}\\
x_{2,1} & x_{2,2} & x_{2,3} & \cdots & x_{2,n}\\
x_{3,1} & x_{3,2} & x_{3,3} & \cdots & x_{3,n}\\
\vdots & \vdots & \vdots & \ddots & \vdots \\
x_{m,1} & x_{m,2} & x_{m,3} & \cdots & x_{m,n}\\
\end{bmatrix}$
However in this notebook we will use the $X_{i,j}$ notation, as it matches fairly well NumPy's notation. Note that in math indices generally start at 1, but in programming they usually start at 0. So to access $A_{2,3}$ programmatically, we need to write this:
```
A[1,2] # 2nd row, 3rd column
```
The $i^{th}$ row vector is sometimes noted $M_i$ or $M_{i,*}$, but again there is no standard notation so people often prefer to explicitely define their own names, for example: "*let **x**$_{i}$ be the $i^{th}$ row vector of matrix $X$*". We will use the $M_{i,*}$, for the same reason as above. For example, to access $A_{2,*}$ (ie. $A$'s 2nd row vector):
```
A[1, :] # 2nd row vector (as a 1D array)
```
Similarly, the $j^{th}$ column vector is sometimes noted $M^j$ or $M_{*,j}$, but there is no standard notation. We will use $M_{*,j}$. For example, to access $A_{*,3}$ (ie. $A$'s 3rd column vector):
```
A[:, 2] # 3rd column vector (as a 1D array)
```
Note that the result is actually a one-dimensional NumPy array: there is no such thing as a *vertical* or *horizontal* one-dimensional array. If you need to actually represent a row vector as a one-row matrix (ie. a 2D NumPy array), or a column vector as a one-column matrix, then you need to use a slice instead of an integer when accessing the row or column, for example:
```
A[1:2, :] # rows 2 to 3 (excluded): this returns row 2 as a one-row matrix
A[:, 2:3] # columns 3 to 4 (excluded): this returns column 3 as a one-column matrix
```
## Square, triangular, diagonal and identity matrices
A **square matrix** is a matrix that has the same number of rows and columns, for example a $3 \times 3$ matrix:
\begin{bmatrix}
4 & 9 & 2 \\
3 & 5 & 7 \\
8 & 1 & 6
\end{bmatrix}
An **upper triangular matrix** is a special kind of square matrix where all the elements *below* the main diagonal (top-left to bottom-right) are zero, for example:
\begin{bmatrix}
4 & 9 & 2 \\
0 & 5 & 7 \\
0 & 0 & 6
\end{bmatrix}
Similarly, a **lower triangular matrix** is a square matrix where all elements *above* the main diagonal are zero, for example:
\begin{bmatrix}
4 & 0 & 0 \\
3 & 5 & 0 \\
8 & 1 & 6
\end{bmatrix}
A **triangular matrix** is one that is either lower triangular or upper triangular.
A matrix that is both upper and lower triangular is called a **diagonal matrix**, for example:
\begin{bmatrix}
4 & 0 & 0 \\
0 & 5 & 0 \\
0 & 0 & 6
\end{bmatrix}
You can construct a diagonal matrix using NumPy's `diag` function:
```
np.diag([4, 5, 6])
```
If you pass a matrix to the `diag` function, it will happily extract the diagonal values:
```
D = np.array([
[1, 2, 3],
[4, 5, 6],
[7, 8, 9],
])
np.diag(D)
```
Finally, the **identity matrix** of size $n$, noted $I_n$, is a diagonal matrix of size $n \times n$ with $1$'s in the main diagonal, for example $I_3$:
\begin{bmatrix}
1 & 0 & 0 \\
0 & 1 & 0 \\
0 & 0 & 1
\end{bmatrix}
Numpy's `eye` function returns the identity matrix of the desired size:
```
np.eye(3)
```
The identity matrix is often noted simply $I$ (instead of $I_n$) when its size is clear given the context. It is called the *identity* matrix because multiplying a matrix with it leaves the matrix unchanged as we will see below.
## Adding matrices
If two matrices $Q$ and $R$ have the same size $m \times n$, they can be added together. Addition is performed *elementwise*: the result is also a $m \times n$ matrix $S$ where each element is the sum of the elements at the corresponding position: $S_{i,j} = Q_{i,j} + R_{i,j}$
$S =
\begin{bmatrix}
Q_{11} + R_{11} & Q_{12} + R_{12} & Q_{13} + R_{13} & \cdots & Q_{1n} + R_{1n} \\
Q_{21} + R_{21} & Q_{22} + R_{22} & Q_{23} + R_{23} & \cdots & Q_{2n} + R_{2n} \\
Q_{31} + R_{31} & Q_{32} + R_{32} & Q_{33} + R_{33} & \cdots & Q_{3n} + R_{3n} \\
\vdots & \vdots & \vdots & \ddots & \vdots \\
Q_{m1} + R_{m1} & Q_{m2} + R_{m2} & Q_{m3} + R_{m3} & \cdots & Q_{mn} + R_{mn} \\
\end{bmatrix}$
For example, let's create a $2 \times 3$ matrix $B$ and compute $A + B$:
```
B = np.array([[1,2,3], [4, 5, 6]])
B
A
A + B
```
**Addition is *commutative***, meaning that $A + B = B + A$:
```
B + A
```
**It is also *associative***, meaning that $A + (B + C) = (A + B) + C$:
```
C = np.array([[100,200,300], [400, 500, 600]])
A + (B + C)
(A + B) + C
```
## Scalar multiplication
A matrix $M$ can be multiplied by a scalar $\lambda$. The result is noted $\lambda M$, and it is a matrix of the same size as $M$ with all elements multiplied by $\lambda$:
$\lambda M =
\begin{bmatrix}
\lambda \times M_{11} & \lambda \times M_{12} & \lambda \times M_{13} & \cdots & \lambda \times M_{1n} \\
\lambda \times M_{21} & \lambda \times M_{22} & \lambda \times M_{23} & \cdots & \lambda \times M_{2n} \\
\lambda \times M_{31} & \lambda \times M_{32} & \lambda \times M_{33} & \cdots & \lambda \times M_{3n} \\
\vdots & \vdots & \vdots & \ddots & \vdots \\
\lambda \times M_{m1} & \lambda \times M_{m2} & \lambda \times M_{m3} & \cdots & \lambda \times M_{mn} \\
\end{bmatrix}$
A more concise way of writing this is:
$(\lambda M)_{i,j} = \lambda (M)_{i,j}$
In NumPy, simply use the `*` operator to multiply a matrix by a scalar. For example:
```
2 * A
```
Scalar multiplication is also defined on the right hand side, and gives the same result: $M \lambda = \lambda M$. For example:
```
A * 2
```
This makes scalar multiplication **commutative**.
It is also **associative**, meaning that $\alpha (\beta M) = (\alpha \times \beta) M$, where $\alpha$ and $\beta$ are scalars. For example:
```
2 * (3 * A)
(2 * 3) * A
```
Finally, it is **distributive over addition** of matrices, meaning that $\lambda (Q + R) = \lambda Q + \lambda R$:
```
2 * (A + B)
2 * A + 2 * B
```
## Matrix multiplication
So far, matrix operations have been rather intuitive. But multiplying matrices is a bit more involved.
A matrix $Q$ of size $m \times n$ can be multiplied by a matrix $R$ of size $n \times q$. It is noted simply $QR$ without multiplication sign or dot. The result $P$ is an $m \times q$ matrix where each element is computed as a sum of products:
$P_{i,j} = \sum_{k=1}^n{Q_{i,k} \times R_{k,j}}$
The element at position $i,j$ in the resulting matrix is the sum of the products of elements in row $i$ of matrix $Q$ by the elements in column $j$ of matrix $R$.
$P =
\begin{bmatrix}
Q_{11} R_{11} + Q_{12} R_{21} + \cdots + Q_{1n} R_{n1} &
Q_{11} R_{12} + Q_{12} R_{22} + \cdots + Q_{1n} R_{n2} &
\cdots &
Q_{11} R_{1q} + Q_{12} R_{2q} + \cdots + Q_{1n} R_{nq} \\
Q_{21} R_{11} + Q_{22} R_{21} + \cdots + Q_{2n} R_{n1} &
Q_{21} R_{12} + Q_{22} R_{22} + \cdots + Q_{2n} R_{n2} &
\cdots &
Q_{21} R_{1q} + Q_{22} R_{2q} + \cdots + Q_{2n} R_{nq} \\
\vdots & \vdots & \ddots & \vdots \\
Q_{m1} R_{11} + Q_{m2} R_{21} + \cdots + Q_{mn} R_{n1} &
Q_{m1} R_{12} + Q_{m2} R_{22} + \cdots + Q_{mn} R_{n2} &
\cdots &
Q_{m1} R_{1q} + Q_{m2} R_{2q} + \cdots + Q_{mn} R_{nq}
\end{bmatrix}$
You may notice that each element $P_{i,j}$ is the dot product of the row vector $Q_{i,*}$ and the column vector $R_{*,j}$:
$P_{i,j} = Q_{i,*} \cdot R_{*,j}$
So we can rewrite $P$ more concisely as:
$P =
\begin{bmatrix}
Q_{1,*} \cdot R_{*,1} & Q_{1,*} \cdot R_{*,2} & \cdots & Q_{1,*} \cdot R_{*,q} \\
Q_{2,*} \cdot R_{*,1} & Q_{2,*} \cdot R_{*,2} & \cdots & Q_{2,*} \cdot R_{*,q} \\
\vdots & \vdots & \ddots & \vdots \\
Q_{m,*} \cdot R_{*,1} & Q_{m,*} \cdot R_{*,2} & \cdots & Q_{m,*} \cdot R_{*,q}
\end{bmatrix}$
Let's multiply two matrices in NumPy, using `ndarray`'s `dot` method:
$E = AD = \begin{bmatrix}
10 & 20 & 30 \\
40 & 50 & 60
\end{bmatrix}
\begin{bmatrix}
2 & 3 & 5 & 7 \\
11 & 13 & 17 & 19 \\
23 & 29 & 31 & 37
\end{bmatrix} =
\begin{bmatrix}
930 & 1160 & 1320 & 1560 \\
2010 & 2510 & 2910 & 3450
\end{bmatrix}$
```
D = np.array([
[ 2, 3, 5, 7],
[11, 13, 17, 19],
[23, 29, 31, 37]
])
E = A.dot(D)
E
```
Let's check this result by looking at one element, just to be sure: looking at $E_{2,3}$ for example, we need to multiply elements in $A$'s $2^{nd}$ row by elements in $D$'s $3^{rd}$ column, and sum up these products:
```
40*5 + 50*17 + 60*31
E[1,2] # row 2, column 3
```
Looks good! You can check the other elements until you get used to the algorithm.
We multiplied a $2 \times 3$ matrix by a $3 \times 4$ matrix, so the result is a $2 \times 4$ matrix. The first matrix's number of columns has to be equal to the second matrix's number of rows. If we try to multiply $D$ by $A$, we get an error because D has 4 columns while A has 2 rows:
```
try:
D.dot(A)
except ValueError as e:
print("ValueError:", e)
```
This illustrates the fact that **matrix multiplication is *NOT* commutative**: in general $QR ≠ RQ$
In fact, $QR$ and $RQ$ are only *both* defined if $Q$ has size $m \times n$ and $R$ has size $n \times m$. Let's look at an example where both *are* defined and show that they are (in general) *NOT* equal:
```
F = np.array([
[5,2],
[4,1],
[9,3]
])
A.dot(F)
F.dot(A)
```
On the other hand, **matrix multiplication *is* associative**, meaning that $Q(RS) = (QR)S$. Let's create a $4 \times 5$ matrix $G$ to illustrate this:
```
G = np.array([
[8, 7, 4, 2, 5],
[2, 5, 1, 0, 5],
[9, 11, 17, 21, 0],
[0, 1, 0, 1, 2]])
A.dot(D).dot(G) # (AB)G
A.dot(D.dot(G)) # A(BG)
```
It is also ***distributive* over addition** of matrices, meaning that $(Q + R)S = QS + RS$. For example:
```
(A + B).dot(D)
A.dot(D) + B.dot(D)
```
The product of a matrix $M$ by the identity matrix (of matching size) results in the same matrix $M$. More formally, if $M$ is an $m \times n$ matrix, then:
$M I_n = I_m M = M$
This is generally written more concisely (since the size of the identity matrices is unambiguous given the context):
$MI = IM = M$
For example:
```
A.dot(np.eye(3))
np.eye(2).dot(A)
```
**Caution**: NumPy's `*` operator performs elementwise multiplication, *NOT* a matrix multiplication:
```
A * B # NOT a matrix multiplication
```
**The @ infix operator**
Python 3.5 [introduced](https://docs.python.org/3/whatsnew/3.5.html#pep-465-a-dedicated-infix-operator-for-matrix-multiplication) the `@` infix operator for matrix multiplication, and NumPy 1.10 added support for it. If you are using Python 3.5+ and NumPy 1.10+, you can simply write `A @ D` instead of `A.dot(D)`, making your code much more readable (but less portable). This operator also works for vector dot products.
```
import sys
print("Python version: {}.{}.{}".format(*sys.version_info))
print("Numpy version:", np.version.version)
# Uncomment the following line if your Python version is ≥3.5
# and your NumPy version is ≥1.10:
#A @ D
```
Note: `Q @ R` is actually equivalent to `Q.__matmul__(R)` which is implemented by NumPy as `np.matmul(Q, R)`, not as `Q.dot(R)`. The main difference is that `matmul` does not support scalar multiplication, while `dot` does, so you can write `Q.dot(3)`, which is equivalent to `Q * 3`, but you cannot write `Q @ 3` ([more details](http://stackoverflow.com/a/34142617/38626)).
## Matrix transpose
The transpose of a matrix $M$ is a matrix noted $M^T$ such that the $i^{th}$ row in $M^T$ is equal to the $i^{th}$ column in $M$:
$ A^T =
\begin{bmatrix}
10 & 20 & 30 \\
40 & 50 & 60
\end{bmatrix}^T =
\begin{bmatrix}
10 & 40 \\
20 & 50 \\
30 & 60
\end{bmatrix}$
In other words, ($A^T)_{i,j}$ = $A_{j,i}$
Obviously, if $M$ is an $m \times n$ matrix, then $M^T$ is an $n \times m$ matrix.
Note: there are a few other notations, such as $M^t$, $M′$, or ${^t}M$.
In NumPy, a matrix's transpose can be obtained simply using the `T` attribute:
```
A
A.T
```
As you might expect, transposing a matrix twice returns the original matrix:
```
A.T.T
```
Transposition is distributive over addition of matrices, meaning that $(Q + R)^T = Q^T + R^T$. For example:
```
(A + B).T
A.T + B.T
```
Moreover, $(Q \cdot R)^T = R^T \cdot Q^T$. Note that the order is reversed. For example:
```
(A.dot(D)).T
D.T.dot(A.T)
```
A **symmetric matrix** $M$ is defined as a matrix that is equal to its transpose: $M^T = M$. This definition implies that it must be a square matrix whose elements are symmetric relative to the main diagonal, for example:
\begin{bmatrix}
17 & 22 & 27 & 49 \\
22 & 29 & 36 & 0 \\
27 & 36 & 45 & 2 \\
49 & 0 & 2 & 99
\end{bmatrix}
The product of a matrix by its transpose is always a symmetric matrix, for example:
```
D.dot(D.T)
```
## Converting 1D arrays to 2D arrays in NumPy
As we mentionned earlier, in NumPy (as opposed to Matlab, for example), 1D really means 1D: there is no such thing as a vertical 1D-array or a horizontal 1D-array. So you should not be surprised to see that transposing a 1D array does not do anything:
```
u
u.T
```
We want to convert $\textbf{u}$ into a row vector before transposing it. There are a few ways to do this:
```
u_row = np.array([u])
u_row
```
Notice the extra square brackets: this is a 2D array with just one row (ie. a 1x2 matrix). In other words it really is a **row vector**.
```
u[np.newaxis, :]
```
This is quite explicit: we are asking for a new vertical axis, keeping the existing data as the horizontal axis.
```
u[np.newaxis]
```
This is equivalent, but a little less explicit.
```
u[None]
```
This is the shortest version, but you probably want to avoid it because it is unclear. The reason it works is that `np.newaxis` is actually equal to `None`, so this is equivalent to the previous version.
Ok, now let's transpose our row vector:
```
u_row.T
```
Great! We now have a nice **column vector**.
Rather than creating a row vector then transposing it, it is also possible to convert a 1D array directly into a column vector:
```
u[:, np.newaxis]
```
## Plotting a matrix
We have already seen that vectors can be represented as points or arrows in N-dimensional space. Is there a good graphical representation of matrices? Well you can simply see a matrix as a list of vectors, so plotting a matrix results in many points or arrows. For example, let's create a $2 \times 4$ matrix `P` and plot it as points:
```
P = np.array([
[3.0, 4.0, 1.0, 4.6],
[0.2, 3.5, 2.0, 0.5]
])
x_coords_P, y_coords_P = P
plt.scatter(x_coords_P, y_coords_P)
plt.axis([0, 5, 0, 4])
plt.show()
```
Of course we could also have stored the same 4 vectors as row vectors instead of column vectors, resulting in a $4 \times 2$ matrix (the transpose of $P$, in fact). It is really an arbitrary choice.
Since the vectors are ordered, you can see the matrix as a path and represent it with connected dots:
```
plt.plot(x_coords_P, y_coords_P, "bo")
plt.plot(x_coords_P, y_coords_P, "b--")
plt.axis([0, 5, 0, 4])
plt.grid()
plt.show()
```
Or you can represent it as a polygon: matplotlib's `Polygon` class expects an $n \times 2$ NumPy array, not a $2 \times n$ array, so we just need to give it $P^T$:
```
from matplotlib.patches import Polygon
plt.gca().add_artist(Polygon(P.T))
plt.axis([0, 5, 0, 4])
plt.grid()
plt.show()
```
## Geometric applications of matrix operations
We saw earlier that vector addition results in a geometric translation, vector multiplication by a scalar results in rescaling (zooming in or out, centered on the origin), and vector dot product results in projecting a vector onto another vector, rescaling and measuring the resulting coordinate.
Similarly, matrix operations have very useful geometric applications.
### Addition = multiple geometric translations
First, adding two matrices together is equivalent to adding all their vectors together. For example, let's create a $2 \times 4$ matrix $H$ and add it to $P$, and look at the result:
```
H = np.array([
[ 0.5, -0.2, 0.2, -0.1],
[ 0.4, 0.4, 1.5, 0.6]
])
P_moved = P + H
plt.gca().add_artist(Polygon(P.T, alpha=0.2))
plt.gca().add_artist(Polygon(P_moved.T, alpha=0.3, color="r"))
for vector, origin in zip(H.T, P.T):
plot_vector2d(vector, origin=origin)
plt.text(2.2, 1.8, "$P$", color="b", fontsize=18)
plt.text(2.0, 3.2, "$P+H$", color="r", fontsize=18)
plt.text(2.5, 0.5, "$H_{*,1}$", color="k", fontsize=18)
plt.text(4.1, 3.5, "$H_{*,2}$", color="k", fontsize=18)
plt.text(0.4, 2.6, "$H_{*,3}$", color="k", fontsize=18)
plt.text(4.4, 0.2, "$H_{*,4}$", color="k", fontsize=18)
plt.axis([0, 5, 0, 4])
plt.grid()
plt.show()
```
If we add a matrix full of identical vectors, we get a simple geometric translation:
```
H2 = np.array([
[-0.5, -0.5, -0.5, -0.5],
[ 0.4, 0.4, 0.4, 0.4]
])
P_translated = P + H2
plt.gca().add_artist(Polygon(P.T, alpha=0.2))
plt.gca().add_artist(Polygon(P_translated.T, alpha=0.3, color="r"))
for vector, origin in zip(H2.T, P.T):
plot_vector2d(vector, origin=origin)
plt.axis([0, 5, 0, 4])
plt.grid()
plt.show()
```
Although matrices can only be added together if they have the same size, NumPy allows adding a row vector or a column vector to a matrix: this is called *broadcasting* and is explained in further details in the [NumPy tutorial](tools_numpy.ipynb). We could have obtained the same result as above with:
```
P + [[-0.5], [0.4]] # same as P + H2, thanks to NumPy broadcasting
```
### Scalar multiplication
Multiplying a matrix by a scalar results in all its vectors being multiplied by that scalar, so unsurprisingly, the geometric result is a rescaling of the entire figure. For example, let's rescale our polygon by a factor of 60% (zooming out, centered on the origin):
```
def plot_transformation(P_before, P_after, text_before, text_after, axis = [0, 5, 0, 4], arrows=False):
if arrows:
for vector_before, vector_after in zip(P_before.T, P_after.T):
plot_vector2d(vector_before, color="blue", linestyle="--")
plot_vector2d(vector_after, color="red", linestyle="-")
plt.gca().add_artist(Polygon(P_before.T, alpha=0.2))
plt.gca().add_artist(Polygon(P_after.T, alpha=0.3, color="r"))
plt.text(P_before[0].mean(), P_before[1].mean(), text_before, fontsize=18, color="blue")
plt.text(P_after[0].mean(), P_after[1].mean(), text_after, fontsize=18, color="red")
plt.axis(axis)
plt.grid()
P_rescaled = 0.60 * P
plot_transformation(P, P_rescaled, "$P$", "$0.6 P$", arrows=True)
plt.show()
```
### Matrix multiplication – Projection onto an axis
Matrix multiplication is more complex to visualize, but it is also the most powerful tool in the box.
Let's start simple, by defining a $1 \times 2$ matrix $U = \begin{bmatrix} 1 & 0 \end{bmatrix}$. This row vector is just the horizontal unit vector.
```
U = np.array([[1, 0]])
```
Now let's look at the dot product $U \cdot P$:
```
U.dot(P)
```
These are the horizontal coordinates of the vectors in $P$. In other words, we just projected $P$ onto the horizontal axis:
```
def plot_projection(U, P):
U_P = U.dot(P)
axis_end = 100 * U
plot_vector2d(axis_end[0], color="black")
plt.gca().add_artist(Polygon(P.T, alpha=0.2))
for vector, proj_coordinate in zip(P.T, U_P.T):
proj_point = proj_coordinate * U
plt.plot(proj_point[0][0], proj_point[0][1], "ro")
plt.plot([vector[0], proj_point[0][0]], [vector[1], proj_point[0][1]], "r--")
plt.axis([0, 5, 0, 4])
plt.grid()
plt.show()
plot_projection(U, P)
```
We can actually project on any other axis by just replacing $U$ with any other unit vector. For example, let's project on the axis that is at a 30° angle above the horizontal axis:
```
angle30 = 30 * np.pi / 180 # angle in radians
U_30 = np.array([[np.cos(angle30), np.sin(angle30)]])
plot_projection(U_30, P)
```
Good! Remember that the dot product of a unit vector and a matrix basically performs a projection on an axis and gives us the coordinates of the resulting points on that axis.
### Matrix multiplication – Rotation
Now let's create a $2 \times 2$ matrix $V$ containing two unit vectors that make 30° and 120° angles with the horizontal axis:
$V = \begin{bmatrix} \cos(30°) & \sin(30°) \\ \cos(120°) & \sin(120°) \end{bmatrix}$
```
angle120 = 120 * np.pi / 180
V = np.array([
[np.cos(angle30), np.sin(angle30)],
[np.cos(angle120), np.sin(angle120)]
])
V
```
Let's look at the product $VP$:
```
V.dot(P)
```
The first row is equal to $V_{1,*} P$, which is the coordinates of the projection of $P$ onto the 30° axis, as we have seen above. The second row is $V_{2,*} P$, which is the coordinates of the projection of $P$ onto the 120° axis. So basically we obtained the coordinates of $P$ after rotating the horizontal and vertical axes by 30° (or equivalently after rotating the polygon by -30° around the origin)! Let's plot $VP$ to see this:
```
P_rotated = V.dot(P)
plot_transformation(P, P_rotated, "$P$", "$VP$", [-2, 6, -2, 4], arrows=True)
plt.show()
```
Matrix $V$ is called a **rotation matrix**.
### Matrix multiplication – Other linear transformations
More generally, any linear transformation $f$ that maps n-dimensional vectors to m-dimensional vectors can be represented as an $m \times n$ matrix. For example, say $\textbf{u}$ is a 3-dimensional vector:
$\textbf{u} = \begin{pmatrix} x \\ y \\ z \end{pmatrix}$
and $f$ is defined as:
$f(\textbf{u}) = \begin{pmatrix}
ax + by + cz \\
dx + ey + fz
\end{pmatrix}$
This transormation $f$ maps 3-dimensional vectors to 2-dimensional vectors in a linear way (ie. the resulting coordinates only involve sums of multiples of the original coordinates). We can represent this transformation as matrix $F$:
$F = \begin{bmatrix}
a & b & c \\
d & e & f
\end{bmatrix}$
Now, to compute $f(\textbf{u})$ we can simply do a matrix multiplication:
$f(\textbf{u}) = F \textbf{u}$
If we have a matric $G = \begin{bmatrix}\textbf{u}_1 & \textbf{u}_2 & \cdots & \textbf{u}_q \end{bmatrix}$, where each $\textbf{u}_i$ is a 3-dimensional column vector, then $FG$ results in the linear transformation of all vectors $\textbf{u}_i$ as defined by the matrix $F$:
$FG = \begin{bmatrix}f(\textbf{u}_1) & f(\textbf{u}_2) & \cdots & f(\textbf{u}_q) \end{bmatrix}$
To summarize, the matrix on the left hand side of a dot product specifies what linear transormation to apply to the right hand side vectors. We have already shown that this can be used to perform projections and rotations, but any other linear transformation is possible. For example, here is a transformation known as a *shear mapping*:
```
F_shear = np.array([
[1, 1.5],
[0, 1]
])
plot_transformation(P, F_shear.dot(P), "$P$", "$F_{shear} P$",
axis=[0, 10, 0, 7])
plt.show()
```
Let's look at how this transformation affects the **unit square**:
```
Square = np.array([
[0, 0, 1, 1],
[0, 1, 1, 0]
])
plot_transformation(Square, F_shear.dot(Square), "$Square$", "$F_{shear} Square$",
axis=[0, 2.6, 0, 1.8])
plt.show()
```
Now let's look at a **squeeze mapping**:
```
F_squeeze = np.array([
[1.4, 0],
[0, 1/1.4]
])
plot_transformation(P, F_squeeze.dot(P), "$P$", "$F_{squeeze} P$",
axis=[0, 7, 0, 5])
plt.show()
```
The effect on the unit square is:
```
plot_transformation(Square, F_squeeze.dot(Square), "$Square$", "$F_{squeeze} Square$",
axis=[0, 1.8, 0, 1.2])
plt.show()
```
Let's show a last one: reflection through the horizontal axis:
```
F_reflect = np.array([
[1, 0],
[0, -1]
])
plot_transformation(P, F_reflect.dot(P), "$P$", "$F_{reflect} P$",
axis=[-2, 9, -4.5, 4.5])
plt.show()
```
## Matrix inverse
Now that we understand that a matrix can represent any linear transformation, a natural question is: can we find a transformation matrix that reverses the effect of a given transformation matrix $F$? The answer is yes… sometimes! When it exists, such a matrix is called the **inverse** of $F$, and it is noted $F^{-1}$.
For example, the rotation, the shear mapping and the squeeze mapping above all have inverse transformations. Let's demonstrate this on the shear mapping:
```
F_inv_shear = np.array([
[1, -1.5],
[0, 1]
])
P_sheared = F_shear.dot(P)
P_unsheared = F_inv_shear.dot(P_sheared)
plot_transformation(P_sheared, P_unsheared, "$P_{sheared}$", "$P_{unsheared}$",
axis=[0, 10, 0, 7])
plt.plot(P[0], P[1], "b--")
plt.show()
```
We applied a shear mapping on $P$, just like we did before, but then we applied a second transformation to the result, and *lo and behold* this had the effect of coming back to the original $P$ (we plotted the original $P$'s outline to double check). The second transformation is the inverse of the first one.
We defined the inverse matrix $F_{shear}^{-1}$ manually this time, but NumPy provides an `inv` function to compute a matrix's inverse, so we could have written instead:
```
F_inv_shear = LA.inv(F_shear)
F_inv_shear
```
Only square matrices can be inversed. This makes sense when you think about it: if you have a transformation that reduces the number of dimensions, then some information is lost and there is no way that you can get it back. For example say you use a $2 \times 3$ matrix to project a 3D object onto a plane. The result may look like this:
```
plt.plot([0, 0, 1, 1, 0, 0.1, 0.1, 0, 0.1, 1.1, 1.0, 1.1, 1.1, 1.0, 1.1, 0.1],
[0, 1, 1, 0, 0, 0.1, 1.1, 1.0, 1.1, 1.1, 1.0, 1.1, 0.1, 0, 0.1, 0.1],
"r-")
plt.axis([-0.5, 2.1, -0.5, 1.5])
plt.show()
```
Looking at this image, it is impossible to tell whether this is the projection of a cube or the projection of a narrow rectangular object. Some information has been lost in the projection.
Even square transformation matrices can lose information. For example, consider this transformation matrix:
```
F_project = np.array([
[1, 0],
[0, 0]
])
plot_transformation(P, F_project.dot(P), "$P$", "$F_{project} \cdot P$",
axis=[0, 6, -1, 4])
plt.show()
```
This transformation matrix performs a projection onto the horizontal axis. Our polygon gets entirely flattened out so some information is entirely lost and it is impossible to go back to the original polygon using a linear transformation. In other words, $F_{project}$ has no inverse. Such a square matrix that cannot be inversed is called a **singular matrix** (aka degenerate matrix). If we ask NumPy to calculate its inverse, it raises an exception:
```
try:
LA.inv(F_project)
except LA.LinAlgError as e:
print("LinAlgError:", e)
```
Here is another example of a singular matrix. This one performs a projection onto the axis at a 30° angle above the horizontal axis:
```
angle30 = 30 * np.pi / 180
F_project_30 = np.array([
[np.cos(angle30)**2, np.sin(2*angle30)/2],
[np.sin(2*angle30)/2, np.sin(angle30)**2]
])
plot_transformation(P, F_project_30.dot(P), "$P$", "$F_{project\_30} \cdot P$",
axis=[0, 6, -1, 4])
plt.show()
```
But this time, due to floating point rounding errors, NumPy manages to calculate an inverse (notice how large the elements are, though):
```
LA.inv(F_project_30)
```
As you might expect, the dot product of a matrix by its inverse results in the identity matrix:
$M \cdot M^{-1} = M^{-1} \cdot M = I$
This makes sense since doing a linear transformation followed by the inverse transformation results in no change at all.
```
F_shear.dot(LA.inv(F_shear))
```
Another way to express this is that the inverse of the inverse of a matrix $M$ is $M$ itself:
$((M)^{-1})^{-1} = M$
```
LA.inv(LA.inv(F_shear))
```
Also, the inverse of scaling by a factor of $\lambda$ is of course scaling by a factor or $\frac{1}{\lambda}$:
$ (\lambda \times M)^{-1} = \frac{1}{\lambda} \times M^{-1}$
Once you understand the geometric interpretation of matrices as linear transformations, most of these properties seem fairly intuitive.
A matrix that is its own inverse is called an **involution**. The simplest examples are reflection matrices, or a rotation by 180°, but there are also more complex involutions, for example imagine a transformation that squeezes horizontally, then reflects over the vertical axis and finally rotates by 90° clockwise. Pick up a napkin and try doing that twice: you will end up in the original position. Here is the corresponding involutory matrix:
```
F_involution = np.array([
[0, -2],
[-1/2, 0]
])
plot_transformation(P, F_involution.dot(P), "$P$", "$F_{involution} \cdot P$",
axis=[-8, 5, -4, 4])
plt.show()
```
Finally, a square matrix $H$ whose inverse is its own transpose is an **orthogonal matrix**:
$H^{-1} = H^T$
Therefore:
$H \cdot H^T = H^T \cdot H = I$
It corresponds to a transformation that preserves distances, such as rotations and reflections, and combinations of these, but not rescaling, shearing or squeezing. Let's check that $F_{reflect}$ is indeed orthogonal:
```
F_reflect.dot(F_reflect.T)
```
## Determinant
The determinant of a square matrix $M$, noted $\det(M)$ or $\det M$ or $|M|$ is a value that can be calculated from its elements $(M_{i,j})$ using various equivalent methods. One of the simplest methods is this recursive approach:
$|M| = M_{1,1}\times|M^{(1,1)}| - M_{2,1}\times|M^{(2,1)}| + M_{3,1}\times|M^{(3,1)}| - M_{4,1}\times|M^{(4,1)}| + \cdots ± M_{n,1}\times|M^{(n,1)}|$
* Where $M^{(i,j)}$ is the matrix $M$ without row $i$ and column $j$.
For example, let's calculate the determinant of the following $3 \times 3$ matrix:
$M = \begin{bmatrix}
1 & 2 & 3 \\
4 & 5 & 6 \\
7 & 8 & 0
\end{bmatrix}$
Using the method above, we get:
$|M| = 1 \times \left | \begin{bmatrix} 5 & 6 \\ 8 & 0 \end{bmatrix} \right |
- 2 \times \left | \begin{bmatrix} 4 & 6 \\ 7 & 0 \end{bmatrix} \right |
+ 3 \times \left | \begin{bmatrix} 4 & 5 \\ 7 & 8 \end{bmatrix} \right |$
Now we need to compute the determinant of each of these $2 \times 2$ matrices (these determinants are called **minors**):
$\left | \begin{bmatrix} 5 & 6 \\ 8 & 0 \end{bmatrix} \right | = 5 \times 0 - 6 \times 8 = -48$
$\left | \begin{bmatrix} 4 & 6 \\ 7 & 0 \end{bmatrix} \right | = 4 \times 0 - 6 \times 7 = -42$
$\left | \begin{bmatrix} 4 & 5 \\ 7 & 8 \end{bmatrix} \right | = 4 \times 8 - 5 \times 7 = -3$
Now we can calculate the final result:
$|M| = 1 \times (-48) - 2 \times (-42) + 3 \times (-3) = 27$
To get the determinant of a matrix, you can call NumPy's `det` function in the `numpy.linalg` module:
```
M = np.array([
[1, 2, 3],
[4, 5, 6],
[7, 8, 0]
])
LA.det(M)
```
One of the main uses of the determinant is to *determine* whether a square matrix can be inversed or not: if the determinant is equal to 0, then the matrix *cannot* be inversed (it is a singular matrix), and if the determinant is not 0, then it *can* be inversed.
For example, let's compute the determinant for the $F_{project}$, $F_{project\_30}$ and $F_{shear}$ matrices that we defined earlier:
```
LA.det(F_project)
```
That's right, $F_{project}$ is singular, as we saw earlier.
```
LA.det(F_project_30)
```
This determinant is suspiciously close to 0: it really should be 0, but it's not due to tiny floating point errors. The matrix is actually singular.
```
LA.det(F_shear)
```
Perfect! This matrix *can* be inversed as we saw earlier. Wow, math really works!
The determinant can also be used to measure how much a linear transformation affects surface areas: for example, the projection matrices $F_{project}$ and $F_{project\_30}$ completely flatten the polygon $P$, until its area is zero. This is why the determinant of these matrices is 0. The shear mapping modified the shape of the polygon, but it did not affect its surface area, which is why the determinant is 1. You can try computing the determinant of a rotation matrix, and you should also find 1. What about a scaling matrix? Let's see:
```
F_scale = np.array([
[0.5, 0],
[0, 0.5]
])
plot_transformation(P, F_scale.dot(P), "$P$", "$F_{scale} \cdot P$",
axis=[0, 6, -1, 4])
plt.show()
```
We rescaled the polygon by a factor of 1/2 on both vertical and horizontal axes so the surface area of the resulting polygon is 1/4$^{th}$ of the original polygon. Let's compute the determinant and check that:
```
LA.det(F_scale)
```
Correct!
The determinant can actually be negative, when the transformation results in a "flipped over" version of the original polygon (eg. a left hand glove becomes a right hand glove). For example, the determinant of the `F_reflect` matrix is -1 because the surface area is preserved but the polygon gets flipped over:
```
LA.det(F_reflect)
```
## Composing linear transformations
Several linear transformations can be chained simply by performing multiple dot products in a row. For example, to perform a squeeze mapping followed by a shear mapping, just write:
```
P_squeezed_then_sheared = F_shear.dot(F_squeeze.dot(P))
```
Since the dot product is associative, the following code is equivalent:
```
P_squeezed_then_sheared = (F_shear.dot(F_squeeze)).dot(P)
```
Note that the order of the transformations is the reverse of the dot product order.
If we are going to perform this composition of linear transformations more than once, we might as well save the composition matrix like this:
```
F_squeeze_then_shear = F_shear.dot(F_squeeze)
P_squeezed_then_sheared = F_squeeze_then_shear.dot(P)
```
From now on we can perform both transformations in just one dot product, which can lead to a very significant performance boost.
What if you want to perform the inverse of this double transformation? Well, if you squeezed and then you sheared, and you want to undo what you have done, it should be obvious that you should unshear first and then unsqueeze. In more mathematical terms, given two invertible (aka nonsingular) matrices $Q$ and $R$:
$(Q \cdot R)^{-1} = R^{-1} \cdot Q^{-1}$
And in NumPy:
```
LA.inv(F_shear.dot(F_squeeze)) == LA.inv(F_squeeze).dot(LA.inv(F_shear))
```
## Singular Value Decomposition
It turns out that any $m \times n$ matrix $M$ can be decomposed into the dot product of three simple matrices:
* a rotation matrix $U$ (an $m \times m$ orthogonal matrix)
* a scaling & projecting matrix $\Sigma$ (an $m \times n$ diagonal matrix)
* and another rotation matrix $V^T$ (an $n \times n$ orthogonal matrix)
$M = U \cdot \Sigma \cdot V^{T}$
For example, let's decompose the shear transformation:
```
U, S_diag, V_T = LA.svd(F_shear) # note: in python 3 you can rename S_diag to Σ_diag
U
S_diag
```
Note that this is just a 1D array containing the diagonal values of Σ. To get the actual matrix Σ, we can use NumPy's `diag` function:
```
S = np.diag(S_diag)
S
```
Now let's check that $U \cdot \Sigma \cdot V^T$ is indeed equal to `F_shear`:
```
U.dot(np.diag(S_diag)).dot(V_T)
F_shear
```
It worked like a charm. Let's apply these transformations one by one (in reverse order) on the unit square to understand what's going on. First, let's apply the first rotation $V^T$:
```
plot_transformation(Square, V_T.dot(Square), "$Square$", "$V^T \cdot Square$",
axis=[-0.5, 3.5 , -1.5, 1.5])
plt.show()
```
Now let's rescale along the vertical and horizontal axes using $\Sigma$:
```
plot_transformation(V_T.dot(Square), S.dot(V_T).dot(Square), "$V^T \cdot Square$", "$\Sigma \cdot V^T \cdot Square$",
axis=[-0.5, 3.5 , -1.5, 1.5])
plt.show()
```
Finally, we apply the second rotation $U$:
```
plot_transformation(S.dot(V_T).dot(Square), U.dot(S).dot(V_T).dot(Square),"$\Sigma \cdot V^T \cdot Square$", "$U \cdot \Sigma \cdot V^T \cdot Square$",
axis=[-0.5, 3.5 , -1.5, 1.5])
plt.show()
```
And we can see that the result is indeed a shear mapping of the original unit square.
## Eigenvectors and eigenvalues
An **eigenvector** of a square matrix $M$ (also called a **characteristic vector**) is a non-zero vector that remains on the same line after transformation by the linear transformation associated with $M$. A more formal definition is any vector $v$ such that:
$M \cdot v = \lambda \times v$
Where $\lambda$ is a scalar value called the **eigenvalue** associated to the vector $v$.
For example, any horizontal vector remains horizontal after applying the shear mapping (as you can see on the image above), so it is an eigenvector of $M$. A vertical vector ends up tilted to the right, so vertical vectors are *NOT* eigenvectors of $M$.
If we look at the squeeze mapping, we find that any horizontal or vertical vector keeps its direction (although its length changes), so all horizontal and vertical vectors are eigenvectors of $F_{squeeze}$.
However, rotation matrices have no eigenvectors at all (except if the rotation angle is 0° or 180°, in which case all non-zero vectors are eigenvectors).
NumPy's `eig` function returns the list of unit eigenvectors and their corresponding eigenvalues for any square matrix. Let's look at the eigenvectors and eigenvalues of the squeeze mapping matrix $F_{squeeze}$:
```
eigenvalues, eigenvectors = LA.eig(F_squeeze)
eigenvalues # [λ0, λ1, …]
eigenvectors # [v0, v1, …]
```
Indeed the horizontal vectors are stretched by a factor of 1.4, and the vertical vectors are shrunk by a factor of 1/1.4=0.714…, so far so good. Let's look at the shear mapping matrix $F_{shear}$:
```
eigenvalues2, eigenvectors2 = LA.eig(F_shear)
eigenvalues2 # [λ0, λ1, …]
eigenvectors2 # [v0, v1, …]
```
Wait, what!? We expected just one unit eigenvector, not two. The second vector is almost equal to $\begin{pmatrix}-1 \\ 0 \end{pmatrix}$, which is on the same line as the first vector $\begin{pmatrix}1 \\ 0 \end{pmatrix}$. This is due to floating point errors. We can safely ignore vectors that are (almost) colinear (ie. on the same line).
## Trace
The trace of a square matrix $M$, noted $tr(M)$ is the sum of the values on its main diagonal. For example:
```
D = np.array([
[100, 200, 300],
[ 10, 20, 30],
[ 1, 2, 3],
])
np.trace(D)
```
The trace does not have a simple geometric interpretation (in general), but it has a number of properties that make it useful in many areas:
* $tr(A + B) = tr(A) + tr(B)$
* $tr(A \cdot B) = tr(B \cdot A)$
* $tr(A \cdot B \cdot \cdots \cdot Y \cdot Z) = tr(Z \cdot A \cdot B \cdot \cdots \cdot Y)$
* $tr(A^T \cdot B) = tr(A \cdot B^T) = tr(B^T \cdot A) = tr(B \cdot A^T) = \sum_{i,j}X_{i,j} \times Y_{i,j}$
* …
It does, however, have a useful geometric interpretation in the case of projection matrices (such as $F_{project}$ that we discussed earlier): it corresponds to the number of dimensions after projection. For example:
```
np.trace(F_project)
```
# What next?
This concludes this introduction to Linear Algebra. Although these basics cover most of what you will need to know for Machine Learning, if you wish to go deeper into this topic there are many options available: Linear Algebra [books](http://linear.axler.net/), [Khan Academy](https://www.khanacademy.org/math/linear-algebra) lessons, or just [Wikipedia](https://en.wikipedia.org/wiki/Linear_algebra) pages.
| github_jupyter |
# Random forest classification
## RAPIDS single GPU
<img src="https://rapids.ai/assets/images/RAPIDS-logo-purple.svg" width="400">
```
import os
```
# Load data and feature engineering
Load a full month for this exercise. Note we are loading the data with RAPIDS now (`cudf.read_csv` vs. `pd.read_csv`)
```
!nvidia-smi
!nvcc --version
import cudf
import s3fs
import xgboost as xgb
s3 = s3fs.S3FileSystem(anon=True)
data = cudf.read_csv(
s3.open( 's3://kjkasjdk2934872398ojljosudfsu8fuj23/data_rev8.csv', mode='rb')
)
print(f'Num rows: {len(data)}, Size: {data.memory_usage(deep=True).sum() / 1e6} MB')
data.shape
data = data.drop(columns=['Unnamed: 0', 'Time'])
data = data.astype('float32')
features = list(data.columns[1:])
target = data.columns[0]
len(features)
```
# Train model
```
%pip install pyDOE
name = 10
n_samples = 20
tree_depth = [3,3]
sample_size = [0.676590, 0.676590]
mtry = [88, 88]
learn_rate = [0.001, 0.1]
trees = 1000
stop_iter = 20
from pyDOE import lhs
import numpy as np
np.random.seed(42)
lhd = lhs(4, samples=n_samples)
import pandas as pd
def scale_param(x, limits):
range_ = limits[1]-limits[0]
res = x*range_+min(limits)
return res
samples = pd.DataFrame({
'tree_depth': np.round(scale_param(lhd[:,0], tree_depth),0).astype(int).tolist(),
'sample_size': scale_param(lhd[:,1], sample_size).tolist(),
'mtry': np.round(scale_param(lhd[:,2], mtry),0).astype(int).tolist(),
'learn_rate': scale_param(lhd[:,3], learn_rate).tolist()
})
samples.head()
from tqdm.auto import tqdm
from cuml.metrics.regression import mean_absolute_error, mean_squared_error, r2_score
fold_train = []
fold_test = []
fold_test_np = []
n_folds = 4
folds_cumul = True
if n_folds == 4 and not folds_cumul:
for fold in tqdm(range(4), total=4):
fold_train_start = fold*40000
fold_train_end = (fold+1)*40000
fold_test_end = (fold+1)*50000
train_data_x = data[features].iloc[fold_train_start:fold_train_end]
train_data_y = data[target].iloc[fold_train_start:fold_train_end]
test_data_x = data[features].iloc[fold_train_end:fold_test_end]
test_data_y = data[target].iloc[fold_train_end:fold_test_end]
fold_train.append(xgb.DMatrix(train_data_x, label=train_data_y))
fold_test.append(xgb.DMatrix(test_data_x, label=test_data_y))
fold_test_np.append(test_data_y)
if n_folds == 4 and folds_cumul:
for fold in tqdm(range(4), total=4):
fold_train_start = 0
fold_train_end = 180000+(fold)*3000
fold_test_end = fold_train_end+2016
train_data_x = data[features].iloc[fold_train_start:fold_train_end]
train_data_y = data[target].iloc[fold_train_start:fold_train_end]
test_data_x = data[features].iloc[fold_train_end:fold_test_end]
test_data_y = data[target].iloc[fold_train_end:fold_test_end]
fold_train.append(xgb.DMatrix(train_data_x, label=train_data_y))
fold_test.append(xgb.DMatrix(test_data_x, label=test_data_y))
fold_test_np.append(test_data_y)
res = []
for sample in tqdm(list(samples.index), total=samples.shape[0]):
this_res = {}
this_res['tree_depth'] = samples.loc[sample, 'tree_depth']
this_res['sample_size'] = samples.loc[sample, 'sample_size']
this_res['mtry'] = samples.loc[sample, 'mtry']
this_res['learn_rate'] = samples.loc[sample, 'learn_rate']
this_res['res'] = {'folds': []}
for fold in tqdm(range(4), total=4, leave=False):
this_fold = {}
params = {
'verbosity': 0,
'tree_method': 'gpu_hist',
'n_gpus': 1,
'eval_metric': 'mae',
'max_depth': samples.loc[sample, 'tree_depth'],
'subsample': samples.loc[sample, 'sample_size'],
'colsample_bytree': samples.loc[sample, 'mtry']/len(features),
'eta': samples.loc[sample, 'learn_rate']
}
evallist = [(fold_train[fold], 'train'),(fold_test[fold], 'validation')]
num_round = 1000
model = xgb.train(params, fold_train[fold], num_round, evals=evallist, early_stopping_rounds=stop_iter, verbose_eval=False)
preds = model.predict(fold_test[fold])
orig = fold_test_np[fold]
this_fold['mae'] = float(mean_absolute_error(orig, preds))
this_fold['rmse'] = float(mean_squared_error(orig, preds, squared=False))
this_fold['r2'] = r2_score(orig, preds)
this_fold['best_iter'] = model.best_iteration
this_res['res']['folds'].append(this_fold)
this_res['res']['mae'] = np.mean([x['mae'] for x in this_res['res']['folds']])
this_res['res']['rmse'] = np.mean([x['rmse'] for x in this_res['res']['folds']])
this_res['res']['r2'] = np.mean([x['r2'] for x in this_res['res']['folds']])
print("tree_depth:{} sample_size:{} mtry:{} learn_rate:{} mae:{}".format(
this_res['tree_depth'],this_res['sample_size'],this_res['mtry'],this_res['learn_rate'],this_res['res']['mae']))
res.append(this_res)
rdf = pd.DataFrame({'tree_depth': [r['tree_depth'] for r in res],
'sample_size': [r['sample_size'] for r in res],
'mtry': [r['mtry'] for r in res],
'learn_rate': [r['learn_rate'] for r in res],
'mae': [r['res']['mae'] for r in res],
'rmse': [r['res']['rmse'] for r in res],
'r2': [r['res']['r2'] for r in res]})
rdf.sort_values('mae')
rdf.to_csv('xgb{}.csv'.format(name))
rdf.corr()
rdf.plot.scatter('tree_depth', 'mae');
rdf.plot.scatter('sample_size', 'mae');
rdf.plot.scatter('mtry', 'mae');
rdf.plot.scatter('learn_rate', 'mae');
rdf.sort_values('mae')
```
| github_jupyter |
TSG023 - Get all BDC objects (Kubernetes)
=========================================
Description
-----------
Get a summary of all Kubernetes resources for the system namespace and
the Big Data Cluster namespace
Steps
-----
### Common functions
Define helper functions used in this notebook.
```
# Define `run` function for transient fault handling, suggestions on error, and scrolling updates on Windows
import sys
import os
import re
import json
import platform
import shlex
import shutil
import datetime
from subprocess import Popen, PIPE
from IPython.display import Markdown
retry_hints = {} # Output in stderr known to be transient, therefore automatically retry
error_hints = {} # Output in stderr where a known SOP/TSG exists which will be HINTed for further help
install_hint = {} # The SOP to help install the executable if it cannot be found
first_run = True
rules = None
debug_logging = False
def run(cmd, return_output=False, no_output=False, retry_count=0):
"""Run shell command, stream stdout, print stderr and optionally return output
NOTES:
1. Commands that need this kind of ' quoting on Windows e.g.:
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='data-pool')].metadata.name}
Need to actually pass in as '"':
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='"'data-pool'"')].metadata.name}
The ' quote approach, although correct when pasting into Windows cmd, will hang at the line:
`iter(p.stdout.readline, b'')`
The shlex.split call does the right thing for each platform, just use the '"' pattern for a '
"""
MAX_RETRIES = 5
output = ""
retry = False
global first_run
global rules
if first_run:
first_run = False
rules = load_rules()
# When running `azdata sql query` on Windows, replace any \n in """ strings, with " ", otherwise we see:
#
# ('HY090', '[HY090] [Microsoft][ODBC Driver Manager] Invalid string or buffer length (0) (SQLExecDirectW)')
#
if platform.system() == "Windows" and cmd.startswith("azdata sql query"):
cmd = cmd.replace("\n", " ")
# shlex.split is required on bash and for Windows paths with spaces
#
cmd_actual = shlex.split(cmd)
# Store this (i.e. kubectl, python etc.) to support binary context aware error_hints and retries
#
user_provided_exe_name = cmd_actual[0].lower()
# When running python, use the python in the ADS sandbox ({sys.executable})
#
if cmd.startswith("python "):
cmd_actual[0] = cmd_actual[0].replace("python", sys.executable)
# On Mac, when ADS is not launched from terminal, LC_ALL may not be set, which causes pip installs to fail
# with:
#
# UnicodeDecodeError: 'ascii' codec can't decode byte 0xc5 in position 4969: ordinal not in range(128)
#
# Setting it to a default value of "en_US.UTF-8" enables pip install to complete
#
if platform.system() == "Darwin" and "LC_ALL" not in os.environ:
os.environ["LC_ALL"] = "en_US.UTF-8"
# When running `kubectl`, if AZDATA_OPENSHIFT is set, use `oc`
#
if cmd.startswith("kubectl ") and "AZDATA_OPENSHIFT" in os.environ:
cmd_actual[0] = cmd_actual[0].replace("kubectl", "oc")
# To aid supportabilty, determine which binary file will actually be executed on the machine
#
which_binary = None
# Special case for CURL on Windows. The version of CURL in Windows System32 does not work to
# get JWT tokens, it returns "(56) Failure when receiving data from the peer". If another instance
# of CURL exists on the machine use that one. (Unfortunately the curl.exe in System32 is almost
# always the first curl.exe in the path, and it can't be uninstalled from System32, so here we
# look for the 2nd installation of CURL in the path)
if platform.system() == "Windows" and cmd.startswith("curl "):
path = os.getenv('PATH')
for p in path.split(os.path.pathsep):
p = os.path.join(p, "curl.exe")
if os.path.exists(p) and os.access(p, os.X_OK):
if p.lower().find("system32") == -1:
cmd_actual[0] = p
which_binary = p
break
# Find the path based location (shutil.which) of the executable that will be run (and display it to aid supportability), this
# seems to be required for .msi installs of azdata.cmd/az.cmd. (otherwise Popen returns FileNotFound)
#
# NOTE: Bash needs cmd to be the list of the space separated values hence shlex.split.
#
if which_binary == None:
which_binary = shutil.which(cmd_actual[0])
if which_binary == None:
if user_provided_exe_name in install_hint and install_hint[user_provided_exe_name] is not None:
display(Markdown(f'HINT: Use [{install_hint[user_provided_exe_name][0]}]({install_hint[user_provided_exe_name][1]}) to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)")
else:
cmd_actual[0] = which_binary
start_time = datetime.datetime.now().replace(microsecond=0)
print(f"START: {cmd} @ {start_time} ({datetime.datetime.utcnow().replace(microsecond=0)} UTC)")
print(f" using: {which_binary} ({platform.system()} {platform.release()} on {platform.machine()})")
print(f" cwd: {os.getcwd()}")
# Command-line tools such as CURL and AZDATA HDFS commands output
# scrolling progress bars, which causes Jupyter to hang forever, to
# workaround this, use no_output=True
#
# Work around a infinite hang when a notebook generates a non-zero return code, break out, and do not wait
#
wait = True
try:
if no_output:
p = Popen(cmd_actual)
else:
p = Popen(cmd_actual, stdout=PIPE, stderr=PIPE, bufsize=1)
with p.stdout:
for line in iter(p.stdout.readline, b''):
line = line.decode()
if return_output:
output = output + line
else:
if cmd.startswith("azdata notebook run"): # Hyperlink the .ipynb file
regex = re.compile(' "(.*)"\: "(.*)"')
match = regex.match(line)
if match:
if match.group(1).find("HTML") != -1:
display(Markdown(f' - "{match.group(1)}": "{match.group(2)}"'))
else:
display(Markdown(f' - "{match.group(1)}": "[{match.group(2)}]({match.group(2)})"'))
wait = False
break # otherwise infinite hang, have not worked out why yet.
else:
print(line, end='')
if rules is not None:
apply_expert_rules(line)
if wait:
p.wait()
except FileNotFoundError as e:
if install_hint is not None:
display(Markdown(f'HINT: Use {install_hint} to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)") from e
exit_code_workaround = 0 # WORKAROUND: azdata hangs on exception from notebook on p.wait()
if not no_output:
for line in iter(p.stderr.readline, b''):
try:
line_decoded = line.decode()
except UnicodeDecodeError:
# NOTE: Sometimes we get characters back that cannot be decoded(), e.g.
#
# \xa0
#
# For example see this in the response from `az group create`:
#
# ERROR: Get Token request returned http error: 400 and server
# response: {"error":"invalid_grant",# "error_description":"AADSTS700082:
# The refresh token has expired due to inactivity.\xa0The token was
# issued on 2018-10-25T23:35:11.9832872Z
#
# which generates the exception:
#
# UnicodeDecodeError: 'utf-8' codec can't decode byte 0xa0 in position 179: invalid start byte
#
print("WARNING: Unable to decode stderr line, printing raw bytes:")
print(line)
line_decoded = ""
pass
else:
# azdata emits a single empty line to stderr when doing an hdfs cp, don't
# print this empty "ERR:" as it confuses.
#
if line_decoded == "":
continue
print(f"STDERR: {line_decoded}", end='')
if line_decoded.startswith("An exception has occurred") or line_decoded.startswith("ERROR: An error occurred while executing the following cell"):
exit_code_workaround = 1
# inject HINTs to next TSG/SOP based on output in stderr
#
if user_provided_exe_name in error_hints:
for error_hint in error_hints[user_provided_exe_name]:
if line_decoded.find(error_hint[0]) != -1:
display(Markdown(f'HINT: Use [{error_hint[1]}]({error_hint[2]}) to resolve this issue.'))
# apply expert rules (to run follow-on notebooks), based on output
#
if rules is not None:
apply_expert_rules(line_decoded)
# Verify if a transient error, if so automatically retry (recursive)
#
if user_provided_exe_name in retry_hints:
for retry_hint in retry_hints[user_provided_exe_name]:
if line_decoded.find(retry_hint) != -1:
if retry_count < MAX_RETRIES:
print(f"RETRY: {retry_count} (due to: {retry_hint})")
retry_count = retry_count + 1
output = run(cmd, return_output=return_output, retry_count=retry_count)
if return_output:
return output
else:
return
elapsed = datetime.datetime.now().replace(microsecond=0) - start_time
# WORKAROUND: We avoid infinite hang above in the `azdata notebook run` failure case, by inferring success (from stdout output), so
# don't wait here, if success known above
#
if wait:
if p.returncode != 0:
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(p.returncode)}.\n')
else:
if exit_code_workaround !=0 :
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(exit_code_workaround)}.\n')
print(f'\nSUCCESS: {elapsed}s elapsed.\n')
if return_output:
return output
def load_json(filename):
"""Load a json file from disk and return the contents"""
with open(filename, encoding="utf8") as json_file:
return json.load(json_file)
def load_rules():
"""Load any 'expert rules' from the metadata of this notebook (.ipynb) that should be applied to the stderr of the running executable"""
try:
# Load this notebook as json to get access to the expert rules in the notebook metadata.
#
j = load_json("tsg023-run-kubectl-get-all.ipynb")
except:
pass # If the user has renamed the book, we can't load ourself. NOTE: Is there a way in Jupyter, to know your own filename?
else:
if "metadata" in j and \
"azdata" in j["metadata"] and \
"expert" in j["metadata"]["azdata"] and \
"rules" in j["metadata"]["azdata"]["expert"]:
rules = j["metadata"]["azdata"]["expert"]["rules"]
rules.sort() # Sort rules, so they run in priority order (the [0] element). Lowest value first.
# print (f"EXPERT: There are {len(rules)} rules to evaluate.")
return rules
def apply_expert_rules(line):
"""Determine if the stderr line passed in, matches the regular expressions for any of the 'expert rules', if so
inject a 'HINT' to the follow-on SOP/TSG to run"""
global rules
for rule in rules:
# rules that have 9 elements are the injected (output) rules (the ones we want). Rules
# with only 8 elements are the source (input) rules, which are not expanded (i.e. TSG029,
# not ../repair/tsg029-nb-name.ipynb)
if len(rule) == 9:
notebook = rule[1]
cell_type = rule[2]
output_type = rule[3] # i.e. stream or error
output_type_name = rule[4] # i.e. ename or name
output_type_value = rule[5] # i.e. SystemExit or stdout
details_name = rule[6] # i.e. evalue or text
expression = rule[7].replace("\\*", "*") # Something escaped *, and put a \ in front of it!
if debug_logging:
print(f"EXPERT: If rule '{expression}' satisfied', run '{notebook}'.")
if re.match(expression, line, re.DOTALL):
if debug_logging:
print("EXPERT: MATCH: name = value: '{0}' = '{1}' matched expression '{2}', therefore HINT '{4}'".format(output_type_name, output_type_value, expression, notebook))
match_found = True
display(Markdown(f'HINT: Use [{notebook}]({notebook}) to resolve this issue.'))
print('Common functions defined successfully.')
# Hints for binary (transient fault) retry, (known) error and install guide
#
retry_hints = {'kubectl': ['A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond']}
error_hints = {'kubectl': [['no such host', 'TSG010 - Get configuration contexts', '../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb'], ['no such host', 'TSG011 - Restart sparkhistory server', '../repair/tsg011-restart-sparkhistory-server.ipynb'], ['No connection could be made because the target machine actively refused it', 'TSG056 - Kubectl fails with No connection could be made because the target machine actively refused it', '../repair/tsg056-kubectl-no-connection-could-be-made.ipynb']]}
install_hint = {'kubectl': ['SOP036 - Install kubectl command line interface', '../install/sop036-install-kubectl.ipynb']}
```
### Run kubectl get all for the system namespace
```
run("kubectl get all")
```
### Get the Kubernetes namespace for the big data cluster
Get the namespace of the Big Data Cluster use the kubectl command line
interface .
**NOTE:**
If there is more than one Big Data Cluster in the target Kubernetes
cluster, then either:
- set \[0\] to the correct value for the big data cluster.
- set the environment variable AZDATA\_NAMESPACE, before starting
Azure Data Studio.
```
# Place Kubernetes namespace name for BDC into 'namespace' variable
if "AZDATA_NAMESPACE" in os.environ:
namespace = os.environ["AZDATA_NAMESPACE"]
else:
try:
namespace = run(f'kubectl get namespace --selector=MSSQL_CLUSTER -o jsonpath={{.items[0].metadata.name}}', return_output=True)
except:
from IPython.display import Markdown
print(f"ERROR: Unable to find a Kubernetes namespace with label 'MSSQL_CLUSTER'. SQL Server Big Data Cluster Kubernetes namespaces contain the label 'MSSQL_CLUSTER'.")
display(Markdown(f'HINT: Use [TSG081 - Get namespaces (Kubernetes)](../monitor-k8s/tsg081-get-kubernetes-namespaces.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [TSG010 - Get configuration contexts](../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [SOP011 - Set kubernetes configuration context](../common/sop011-set-kubernetes-context.ipynb) to resolve this issue.'))
raise
print(f'The SQL Server Big Data Cluster Kubernetes namespace is: {namespace}')
```
### Run kubectl get all for the Big Data Cluster namespace
```
run(f"kubectl get all -n {namespace}")
print('Notebook execution complete.')
```
| github_jupyter |
<img alt="QuantRocket logo" src="https://www.quantrocket.com/assets/img/notebook-header-logo.png">
© Copyright Quantopian Inc.<br>
© Modifications Copyright QuantRocket LLC<br>
Licensed under the [Creative Commons Attribution 4.0](https://creativecommons.org/licenses/by/4.0/legalcode).
<a href="https://www.quantrocket.com/disclaimer/">Disclaimer</a>
# Risk-Constrained Portfolio Optimization
by Rene Zhang and Max Margenot
Risk management is critical for constructing portfolios and building algorithms. Its main function is to improve the quality and consistency of returns by adequately accounting for risk. Any returns obtained by *unexpected* risks, which are always lurking within our portfolio, can usually not be relied upon to produce profits over a long time. By limiting the impact of or eliminating these unexpected risks, the portfolio should ideally only have exposure to the alpha we are pursuing. In this lecture, we will focus on how to use factor models in risk management.
## Factor Models
Other lectures have discussed Factor Models and the calculation of Factor Risk Exposure, as well as how to analyze alpha factors. The notation we generally use when introducing a factor model is as follows:
$$R_i = a_i + b_{i1} F_1 + b_{i2} F_2 + \ldots + b_{iK} F_k + \epsilon_i$$
where:
$$\begin{eqnarray}
k &=& \text{the number of factors}\\
R_i &=& \text{the return for company $i$}, \\
a_i &=& \text{the intercept},\\
F_j &=& \text{the return for factor $j$, $j \in [1,k]$}, \\
b_{ij} &=& \text{the corresponding exposure to factor $j$, $j \in [1,k]$,} \\
\epsilon_i &=& \text{specific fluctuation of company $i$.}\\
\end{eqnarray}$$
To quantify unexpected risks and have acceptable risk levels in a given portfolio, we need to answer 3 questions:
1. What proportion of the variance of my portfolio comes from common risk factors?
2. How do I limit this risk?
3. Where does the return/PNL of my portfolio come from, i.e., to what do I attribute the performance?
These risk factors can be:
- Classical fundamental factors, such as those in the CAPM (market risk) or the Fama-French 3-Factor Model (price-to-book (P/B) ratio, volatility)
- Sector or industry exposure
- Macroeconomic factors, such as inflation or interest rates
- Statistical factors that are based on historical returns and derived from principal component
analysis
### Universe
The base universe of assets we use here is the TradableStocksUS.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import statsmodels.api as sm
from zipline.pipeline import Pipeline
from zipline.pipeline.data import sharadar, master, EquityPricing
from zipline.pipeline.factors import CustomFactor, Returns, AverageDollarVolume
from zipline.pipeline.filters import AllPresent, All
from zipline.research import run_pipeline
# date range for building risk model
start = "2009-01-01"
end = "2011-01-01"
```
First we pull the returns of every asset in this universe across our desired time period.
```
TradableStocksUS = (
# Market cap over $500M
(sharadar.Fundamentals.slice(dimension='ARQ', period_offset=0).MARKETCAP.latest >= 500e6)
# dollar volume over $2.5M over trailing 200 days
& (AverageDollarVolume(window_length=200) >= 2.5e6)
# price > $5
& (EquityPricing.close.latest > 5)
# no missing data for 200 days (exclude trading halts, IPOs, etc.)
& AllPresent(inputs=[EquityPricing.close], window_length=200)
& All([EquityPricing.volume.latest > 0], window_length=200)
# common stocks only
& master.SecuritiesMaster.usstock_SecurityType2.latest.eq("Common Stock")
# primary share only
& master.SecuritiesMaster.usstock_PrimaryShareSid.latest.isnull()
)
def tus_returns(start_date, end_date):
pipe = Pipeline(
columns={'Close': EquityPricing.close.latest},
screen = TradableStocksUS
)
stocks = run_pipeline(pipe, start_date=start_date, end_date=end_date, bundle='usstock-1d-bundle')
unstacked_results = stocks.unstack()
prices = (unstacked_results['Close'].fillna(method='ffill').fillna(method='bfill')
.dropna(axis=1,how='any').shift(periods=-1).dropna())
tus_returns = prices.pct_change()[1:]
return tus_returns
R = tus_returns(start, end)
print("The universe we define includes {} assets.".format(R.shape[1]))
print('The number of timestamps is {} from {} to {}.'.format(R.shape[0], start, end))
assets = R.columns
```
### Factor Returns and Exposures
We will start with the classic Fama-French factors. The Fama-French factors are the market, company size, and company price-to-book (PB) ratio. We compute each asset's exposures to these factors, computing the factors themselves using pipeline code borrowed from the Fundamental Factor Models lecture.
```
Fundamentals = sharadar.Fundamentals.slice(dimension='ARQ', period_offset=0)
def make_pipeline():
"""
Create and return our pipeline.
We break this piece of logic out into its own function to make it easier to
test and modify in isolation.
"""
# Market Cap
market_cap = Fundamentals.MARKETCAP.latest
# Book to Price ratio
book_to_price = 1/Fundamentals.PB.latest
# Build Filters representing the top and bottom 500 stocks by our combined ranking system.
biggest = market_cap.top(500, mask=TradableStocksUS)
smallest = market_cap.bottom(500, mask=TradableStocksUS)
highpb = book_to_price.top(500, mask=TradableStocksUS)
lowpb = book_to_price.bottom(500, mask=TradableStocksUS)
universe = biggest | smallest | highpb | lowpb
pipe = Pipeline(
columns = {
'returns' : Returns(window_length=2),
'market_cap' : market_cap,
'book_to_price' : book_to_price,
'biggest' : biggest,
'smallest' : smallest,
'highpb' : highpb,
'lowpb' : lowpb
},
screen=universe
)
return pipe
```
Here we run our pipeline and create the return streams for high-minus-low and small-minus-big.
```
from quantrocket.master import get_securities
from quantrocket import get_prices
pipe = make_pipeline()
# This takes a few minutes.
results = run_pipeline(pipe, start_date=start, end_date=end, bundle='usstock-1d-bundle')
R_biggest = results[results.biggest]['returns'].groupby(level=0).mean()
R_smallest = results[results.smallest]['returns'].groupby(level=0).mean()
R_highpb = results[results.highpb]['returns'].groupby(level=0).mean()
R_lowpb = results[results.lowpb]['returns'].groupby(level=0).mean()
SMB = R_smallest - R_biggest
HML = R_highpb - R_lowpb
df = pd.DataFrame({
'SMB': SMB.tz_localize(None), # company size
'HML': HML.tz_localize(None) # company PB ratio
},columns =["SMB","HML"]).shift(periods =-1).dropna()
SPY = get_securities(symbols='SPY', vendors='usstock').index[0]
MKT = get_prices(
'usstock-1d-bundle',
data_frequency='daily',
sids=SPY,
start_date=start,
end_date=end,
fields='Close').loc['Close'][SPY].pct_change()[1:]
MKT = pd.DataFrame({'MKT':MKT})
F = pd.concat([MKT,df],axis = 1).dropna()
ax = ((F + 1).cumprod() - 1).plot(subplots=True, title='Cumulative Fundamental Factors')
ax[0].set(ylabel = "daily returns")
ax[1].set(ylabel = "daily returns")
ax[2].set(ylabel = "daily returns")
plt.show()
```
### Calculating the Exposures
Running a multiple linear regression on the fundamental factors for each asset in our universe, we can obtain the corresponding factor exposure for each asset. Here we express:
$$ R_i = \alpha_i + \beta_{i, MKT} R_{i, MKT} + \beta_{i, HML} R_{i, HML} + \beta_{i, SMB} R_{i, SMB} + \epsilon_i$$
for each asset $S_i$. This shows us how much of each individual security's return is made up of these risk factors.
We calculate the risk exposures on an asset-by-asset basis in order to get a more granular view of the risk of our portfolio. This approach requires that we know the holdings of the portfolio itself, on any given day, and is computationally expensive.
```
# factor exposure
R = R.tz_localize(None)
B = pd.DataFrame(index=assets, dtype=np.float32)
epsilon = pd.DataFrame(index=R.index, dtype=np.float32)
from IPython.display import clear_output
x = sm.add_constant(F)
for i, asset in enumerate(assets):
print(f"asset {i+1} of {len(assets)}")
clear_output(wait=True)
y = R.loc[:, asset]
y_inlier = y[np.abs(y - y.mean())<=(3*y.std())]
x_inlier = x[np.abs(y - y.mean())<=(3*y.std())]
result = sm.OLS(y_inlier, x_inlier).fit()
B.loc[asset,"MKT_beta"] = result.params[1]
B.loc[asset,"SMB_beta"] = result.params[2]
B.loc[asset,"HML_beta"] = result.params[3]
epsilon[asset] = y - (x.iloc[:,0] * result.params[0] +
x.iloc[:,1] * result.params[1] +
x.iloc[:,2] * result.params[2] +
x.iloc[:,3] * result.params[3])
```
The factor exposures are shown as follows. Each individual asset in our universe will have a different exposure to the three included risk factors.
```
fig,axes = plt.subplots(3, 1)
ax1,ax2,ax3 =axes
B.iloc[0:10,0].plot.barh(ax=ax1, figsize=[15,15], title=B.columns[0])
B.iloc[0:10,1].plot.barh(ax=ax2, figsize=[15,15], title=B.columns[1])
B.iloc[0:10,2].plot.barh(ax=ax3, figsize=[15,15], title=B.columns[2])
ax1.set(xlabel='beta')
ax2.set(xlabel='beta')
ax3.set(xlabel='beta')
plt.show()
from zipline.research import sid
aapl = sid("FIBBG000B9XRY4", bundle='usstock-1d-bundle')
B.loc[aapl,:]
```
### Summary of the Setup:
1. returns of assets in universe: `R`
2. fundamental factors: `F`
3. Exposures of these fundamental factors: `B`
Currently, the `F` DataFrame contains the return streams for MKT, SMB, and HML, by date.
```
F.head(3)
```
While the `B` DataFrame contains point estimates of the beta exposures **to** MKT, SMB, and HML for every asset in our universe.
```
B.head(3)
```
Now that we have these values, we can start to crack open the variance of any portfolio that contains these assets.
### Splitting Variance into Common Factor Risks
The portfolio variance can be represented as:
$$\sigma^2 = \omega BVB^{\top}\omega^{\top} + \omega D\omega^{\top}$$
where:
$$\begin{eqnarray}
B &=& \text{the matrix of factor exposures of $n$ assets to the factors} \\
V &=& \text{the covariance matrix of factors} \\
D &=& \text{the specific variance} \\
\omega &=& \text{the vector of portfolio weights for $n$ assets}\\
\omega BVB^{\top}\omega^{\top} &=& \text{common factor variance} \\
\omega D\omega^{\top} &=& \text{specific variance} \\
\end{eqnarray}$$
#### Computing Common Factor and Specific Variance:
Here we build functions to break out the risk in our portfolio. Suppose that our portfolio consists of all stocks in the Q3000US, equally-weighted. Let's have a look at how much of the variance of the returns in this universe are due to common factor risk.
```
w = np.ones([1,R.shape[1]])/R.shape[1]
def compute_common_factor_variance(factors, factor_exposures, w):
B = np.asarray(factor_exposures)
F = np.asarray(factors)
V = np.asarray(factors.cov())
return w.dot(B.dot(V).dot(B.T)).dot(w.T)
common_factor_variance = compute_common_factor_variance(F, B, w)[0][0]
print("Common Factor Variance: {0}".format(common_factor_variance))
def compute_specific_variance(epsilon, w):
D = np.diag(np.asarray(epsilon.var())) * epsilon.shape[0] / (epsilon.shape[0]-1)
return w.dot(D).dot(w.T)
specific_variance = compute_specific_variance(epsilon, w)[0][0]
print("Specific Variance: {0}".format(specific_variance))
```
In order to actually calculate the percentage of our portfolio variance that is made up of common factor risk, we do the following:
$$\frac{\text{common factor variance}}{\text{common factor variance + specific variance}}$$
```
common_factor_pct = common_factor_variance/(common_factor_variance + specific_variance)*100.0
print("Percentage of Portfolio Variance Due to Common Factor Risk: {0:.2f}%".format(common_factor_pct))
```
So we see that if we just take every single security in the Q3000US and equally-weight them, we will end up possessing a portfolio that effectively only contains common risk.
### Risk-Constrained Optimization
Currently we are operating with an equal-weighted portfolio. However, we can reapportion those weights in such a way that we minimize the common factor risk illustrated by our common factor exposures. This is a portfolio optimization problem to find the optimal weights.
We define this problem as:
\begin{array}{ll} \mbox{$\text{minimize/maximum}$}_{w} & \text{objective function}\\
\mbox{subject to} & {\bf 1}^T \omega = 1, \quad f=B^T\omega\\
& \omega \in {\cal W}, \quad f \in {\cal F},
\end{array}
where the variable $w$ is the vector of allocations, the variable $f$ is weighted factor exposures, and the variable ${\cal F}$ provides our constraints for $f$. We set ${\cal F}$ as a vector to bound the weighted factor exposures of the porfolio. These constraints allow us to reject weightings that do not fit our criteria. For example, we can set the maximum factor exposures that our portfolios can have by changing the value of ${\cal F}$. A value of $[1,1,1]$ would indicate that we want the maximum factor exposure of the portfolio to each factor to be less than $1$, rejecting any portfolios that do not meet that condition.
We define the objective function as whichever business goal we value highest. This can be something such as maximizing the Sharpe ratio or minimizing the volatility. Ultimately, what we want to solve for in this optimization problem is the weights, $\omega$.
Let's quickly generate some random weights to see how the weighted factor exposures of the portfolio change.
```
w_0 = np.random.rand(R.shape[1])
w_0 = w_0/np.sum(w_0)
```
The variable $f$ contains the weighted factor exposures of our portfolio, with size equal to the number of factors we have. As we change $\omega$, our weights, our weighted exposures, $f$, also change.
```
f = B.T.dot(w_0)
f
```
A concrete example of this can be found [here](http://nbviewer.jupyter.org/github/cvxgrp/cvx_short_course/blob/master/applications/portfolio_optimization.ipynb), in the docs for CVXPY.
## References
* Qian, E.E., Hua, R.H. and Sorensen, E.H., 2007. *Quantitative equity portfolio management: modern techniques and applications*. CRC Press.
* Narang, R.K., 2013. *Inside the Black Box: A Simple Guide to Quantitative and High Frequency Trading*. John Wiley & Sons.
---
**Next Lecture:** [Principal Component Analysis](Lecture36-PCA.ipynb)
[Back to Introduction](Introduction.ipynb)
---
*This presentation is for informational purposes only and does not constitute an offer to sell, a solicitation to buy, or a recommendation for any security; nor does it constitute an offer to provide investment advisory or other services by Quantopian, Inc. ("Quantopian") or QuantRocket LLC ("QuantRocket"). Nothing contained herein constitutes investment advice or offers any opinion with respect to the suitability of any security, and any views expressed herein should not be taken as advice to buy, sell, or hold any security or as an endorsement of any security or company. In preparing the information contained herein, neither Quantopian nor QuantRocket has taken into account the investment needs, objectives, and financial circumstances of any particular investor. Any views expressed and data illustrated herein were prepared based upon information believed to be reliable at the time of publication. Neither Quantopian nor QuantRocket makes any guarantees as to their accuracy or completeness. All information is subject to change and may quickly become unreliable for various reasons, including changes in market conditions or economic circumstances.*
| github_jupyter |
# Sentiment analysis using IMDB dataset
```
import numpy as np
from glob import glob
import os
import matplotlib.pyplot as plt
from sklearn import svm
import zipfile
from tqdm import tqdm
from nltk.tokenize import word_tokenize
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer
import re
from scipy import sparse
import nltk
# Download any necessary nltk files for nlp
nltk.download('punkt')
```
# Get data
```
zip_file_path = './imdb_dataset.zip'
extract_dir = './'
data_dir = 'imdb_dataset'
# Extract all the files
zip_ref = zipfile.ZipFile(zip_file_path, 'r')
zip_ref.extractall(extract_dir)
zip_ref.close()
```
Let's begin by reading in all of our text files. We'll create their label according to their sentiment, either positive or negative. In addition we'll preprocess all the nexts by removing all non-alpha numeric characters.
```
# Regex to remove all Non-Alpha Numeric
SPECIAL_CHARS = re.compile(r'([^a-z\d!?.\s])', re.IGNORECASE)
def read_texts(glob_to_texts):
texts = []
labels = []
label = int("pos" in glob_to_texts)
for text_name in tqdm(glob(glob_to_texts)):
with open(text_name, 'r') as text:
# Removing all non-alphanumeric
filter_text = SPECIAL_CHARS.sub('', text.read())
texts.append(filter_text)
labels.append(label)
return texts, labels
# Get all training data
train_pos_data = read_texts(os.path.join(data_dir, "train/pos/*.txt"))
train_neg_data = read_texts(os.path.join(data_dir, "train/neg/*.txt"))
# Get all test data
test_pos_data = read_texts(os.path.join(data_dir, "test/pos/*.txt"))
test_neg_data = read_texts(os.path.join(data_dir, "test/neg/*.txt"))
train_texts = train_pos_data[0] + train_neg_data[0]
train_labels = train_pos_data[1] + train_neg_data[1]
test_texts = test_pos_data[0] + test_neg_data[0]
test_labels = test_pos_data[1] + test_neg_data[1]
```
Split the data into training and validation sets. We'll create a validation test set with 10% of the data.
```
train_texts, val_texts, train_labels, val_labels = train_test_split(train_texts, train_labels, test_size=0.1,
random_state=42)
```
## Vectorization
In order to extract information from text, we'll vectorize our word sequences. In other words, we'll transform our sentences into numerical features. There are many vectorization or embedding techniques such as Bag of Words, Pre-Trained word embeddings, but in our case we'll be using **TF-IDF**.
TF-IDF stands for "Term Frequency, Inverse Document Frequency". It's a technique that converts words into an importance score of each word in the document based on how they appear accros multiple documents. Intuitively, the TF-IDF score of a word is high when it is frequently found in a document. However, if the word appears in many documents, this word is not a unique identifier, and as such, will have a lower score. For example, common words such as "the" and "and" will have low score since they appear in many documents.
```
vec = TfidfVectorizer(ngram_range=(1, 2), tokenizer=word_tokenize,
min_df=3, max_df=0.9, strip_accents='unicode', use_idf=1,
smooth_idf=1, sublinear_tf=1)
```
We fit our vectorizer to our entire corpus of words, which includes the training, validation, and test sets. Once fitted, we'll transform each subset of the data.
```
print("Created Vectorizer %s" % vec)
print("Fitting to all docs...")
vec.fit(train_texts + val_texts + test_texts)
print("Transforming train docs...")
trn_term_doc = vec.transform(train_texts)
print("Transforming val docs...")
val_term_doc = vec.transform(val_texts)
print("Transforming test docs...")
test_term_doc = vec.transform(test_texts)
```
# Model
If you're unfamiliar or want a refresher on SVM's you should check out our [CV tutorial](https://github.com/abhmul/DataScienceTrack/blob/master/CV/Tutorial.ipynb)!
```
from sklearn.base import BaseEstimator, ClassifierMixin
from sklearn.utils.validation import check_X_y, check_is_fitted
from sklearn.svm import LinearSVC
class NbSvmClassifier(BaseEstimator, ClassifierMixin):
def __init__(self, C=1.0, dual='auto', verbose=0):
self.C = C
self.dual = dual
self.verbose = verbose
self._clf = None
print("Creating model with C=%s" % C)
def predict(self, x):
# Verify that model has been fit
check_is_fitted(self, ['_r', '_clf'])
return self._clf.predict(x.multiply(self._r))
def score(self, x, y):
check_is_fitted(self, ['_r', '_clf'])
return self._clf.score(x.multiply(self._r), y)
def fit(self, x, y):
# Check that X and y have correct shape
x, y = check_X_y(x, y, accept_sparse=True)
def pr(x, y_i, y):
p = x[y == y_i].sum(0)
return (p + 1) / ((y == y_i).sum() + 1)
self._r = sparse.csr_matrix(np.log(pr(x, 1, y) / pr(x, 0, y)))
x_nb = x.multiply(self._r)
if self.dual == 'auto':
self.dual = x_nb.shape[0] <= x_nb.shape[1]
self._clf = LinearSVC(C=self.C, dual=self.dual, verbose=self.verbose)
self._clf.fit(x_nb, y)
return self
```
## Finding optimal parameters
We'll perform a grid search across the C parameter to find the optimal parameter for our dataset.
```
# Search for the appropriate C
Cs = [1e-2, 1e-1, 1e0, 1e1, 1e2]
best_model = None
best_val = -float("inf")
best_C = None
for C in Cs:
print("Fitting with C={}".format(C))
model = NbSvmClassifier(C=C, verbose=0).fit(trn_term_doc, train_labels)
# Evaluate the model
val_preds = model.predict(val_term_doc)
score = np.mean(val_labels == val_preds)
print("Model had val score of %s" % score)
if score > best_val:
print("New maximum score improved from {} to {}".format(best_val, score))
best_model = model
best_val = score
best_C = C
score = best_val
print("Best score with C={} is {}".format(best_C, score))
```
## Test score
```
best_model.score(test_term_doc, test_labels)
```
## Takeaways
From this tutorial, we learned how to work with text data and use a basic embedding. In addition, we realize that deep learning isn't always the way to go! We trained a fast and powerful linear model that achieved ~**91**%!
## Sample Texts
```
train_pos_sample_ind = np.random.randint(len(train_pos_data[0]))
train_neg_sample_ind = np.random.randint(len(train_neg_data[0]))
print("Positive Sentiment example")
print(train_pos_data[0][train_pos_sample_ind])
print("---------------------------")
print("Negative Sentiment example")
print(train_neg_data[0][train_neg_sample_ind])
from collections import defaultdict
word_counts = defaultdict(int)
# Compute the frequency of each unique
for text in tqdm(train_texts + val_texts + test_texts):
# Splits sentences
for word in word_tokenize(text):
word_counts[word] += 1
vocab = ['<PAD>'] + sorted(word_counts, key=lambda word: word_counts[word], reverse=True)
word2id = {word: i for i, word in enumerate(vocab)}
# Examine the most common words
print("Number of unique words", len(vocab))
print("Most frequent word: ", vocab[1], "occurs", word_counts[vocab[1]], "times")
print(vocab[:100])
np.savez('glove_embeddings.npz', embeddings=embeddings)
glove_embeddings = np.load('glove_embeddings.npz')['embeddings']
def map_texts(texts, word2id):
return [[word2id[word] for word in word_tokenize(text)] for text in tqdm(texts)]
train_map_text = map_texts(train_texts, word2id)
val_map_text = map_texts(val_texts, word2id)
test_map_text = map_texts(test_texts, word2id)
x_train = keras.preprocessing.sequence.pad_sequences(train_map_text)
x_val = keras.preprocessing.sequence.pad_sequences(val_map_text)
x_test = keras.preprocessing.sequence.pad_sequences(test_map_text)
import keras
from keras import layers
from keras import models
def get_LSTM_model(embedding_matrix):
inp = layers.Input(shape=(None,))
x = layers.Embedding(*(embedding_matrix.shape),
weights=[embedding_matrix],
trainable=False)(inp)
x = layers.Bidirectional(layers.LSTM(50, return_sequences=True, dropout=0.1, recurrent_dropout=0.1))(x)
x = layers.GlobalMaxPool1D()(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(50, activation="relu")(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(1)(x)
model = models.Model(inputs=inp, outputs=x)
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
return model
model = get_LSTM_model(glove_embeddings)
model.summary()
model.fit(x_train,
train_labels,
validation_data=(x_val, val_labels),
batch_size=128,
epochs=20,
shuffle=True)
```
| github_jupyter |
```
#default_exp callback.training
```
# Training Callbacks
> Callbacks to help during training, including `fit_one_cycle`, the LR Finder, and hyper-parameter scheduling
```
#export
# Contains code used/modified by fastai_minima author from fastai
# Copyright 2019 the fast.ai team.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language
#hide
from nbdev.showdoc import *
from fastcore.test import *
#export
from fastcore.basics import store_attr, patch
from fastcore.foundation import L
from fastcore.meta import delegates
from fastcore.xtras import is_listy
from fastai_minima.optimizer import convert_params
from fastai_minima.callback.core import Callback, CancelValidException, CancelFitException
from fastai_minima.learner import Learner, Recorder
from fastai_minima.utils import tensor, defaults, params
import functools, math, collections
import os
import matplotlib.pyplot as plt
import numpy as np
import torch
```
## Annealing
```
#export
class _Annealer:
def __init__(self, f, start, end): store_attr('f,start,end')
def __call__(self, pos): return self.f(self.start, self.end, pos)
#export
def annealer(f):
"Decorator to make `f` return itself partially applied."
@functools.wraps(f)
def _inner(start, end): return _Annealer(f, start, end)
return _inner
```
This is the decorator we will use for all of our scheduling functions, as it transforms a function taking `(start, end, pos)` to something taking `(start, end)` and return a function depending of `pos`.
```
#export
#TODO Jeremy, make this pickle
#@annealer
#def SchedLin(start, end, pos): return start + pos*(end-start)
#@annealer
#def SchedCos(start, end, pos): return start + (1 + math.cos(math.pi*(1-pos))) * (end-start) / 2
#@annealer
#def SchedNo (start, end, pos): return start
#@annealer
#def SchedExp(start, end, pos): return start * (end/start) ** pos
#
#SchedLin.__doc__ = "Linear schedule function from `start` to `end`"
#SchedCos.__doc__ = "Cosine schedule function from `start` to `end`"
#SchedNo .__doc__ = "Constant schedule function with `start` value"
#SchedExp.__doc__ = "Exponential schedule function from `start` to `end`"
#export
def sched_lin(start, end, pos): return start + pos*(end-start)
def sched_cos(start, end, pos): return start + (1 + math.cos(math.pi*(1-pos))) * (end-start) / 2
def sched_no (start, end, pos): return start
def sched_exp(start, end, pos): return start * (end/start) ** pos
def SchedLin(start, end):
"Linear schedule function from `start` to `end`"
return _Annealer(sched_lin, start, end)
def SchedCos(start, end):
"Cosine schedule function from `start` to `end`"
return _Annealer(sched_cos, start, end)
def SchedNo (start, end):
"Constant schedule function with `start` value"
return _Annealer(sched_no, start, end)
def SchedExp(start, end):
"Exponential schedule function from `start` to `end`"
return _Annealer(sched_exp, start, end)
#hide
import pickle
tst = pickle.dumps(SchedCos(0, 5))
annealings = "NO LINEAR COS EXP".split()
p = torch.linspace(0.,1,100)
fns = [SchedNo, SchedLin, SchedCos, SchedExp]
#export
def SchedPoly(start, end, power):
"Polynomial schedule (of `power`) function from `start` to `end`"
def _inner(pos): return start + (end - start) * pos ** power
return _inner
for fn, t in zip(fns, annealings):
plt.plot(p, [fn(2, 1e-2)(o) for o in p], label=t)
f = SchedPoly(2,1e-2,0.5)
plt.plot(p, [f(o) for o in p], label="POLY(0.5)")
plt.legend();
show_doc(SchedLin)
sched = SchedLin(0, 2)
test_eq(L(map(sched, [0., 0.25, 0.5, 0.75, 1.])), [0., 0.5, 1., 1.5, 2.])
show_doc(SchedCos)
sched = SchedCos(0, 2)
test_close(L(map(sched, [0., 0.25, 0.5, 0.75, 1.])), [0., 0.29289, 1., 1.70711, 2.])
show_doc(SchedNo)
sched = SchedNo(0, 2)
test_close(L(map(sched, [0., 0.25, 0.5, 0.75, 1.])), [0., 0., 0., 0., 0.])
show_doc(SchedExp)
sched = SchedExp(1, 2)
test_close(L(map(sched, [0., 0.25, 0.5, 0.75, 1.])), [1., 1.18921, 1.41421, 1.68179, 2.])
show_doc(SchedPoly)
sched = SchedPoly(0, 2, 2)
test_close(L(map(sched, [0., 0.25, 0.5, 0.75, 1.])), [0., 0.125, 0.5, 1.125, 2.])
p = torch.linspace(0.,1,100)
pows = [0.5,1.,2.]
for e in pows:
f = SchedPoly(2, 0, e)
plt.plot(p, [f(o) for o in p], label=f'power {e}')
plt.legend();
#export
def combine_scheds(pcts, scheds):
"Combine `scheds` according to `pcts` in one function"
assert sum(pcts) == 1.
pcts = tensor([0] + L(pcts))
assert torch.all(pcts >= 0)
pcts = torch.cumsum(pcts, 0)
def _inner(pos):
if int(pos) == 1: return scheds[-1](1.)
idx = (pos >= pcts).nonzero().max()
actual_pos = (pos-pcts[idx]) / (pcts[idx+1]-pcts[idx])
return scheds[idx](actual_pos.item())
return _inner
```
`pcts` must be a list of positive numbers that add up to 1 and is the same length as `scheds`. The generated function will use `scheds[0]` from 0 to `pcts[0]` then `scheds[1]` from `pcts[0]` to `pcts[0]+pcts[1]` and so forth.
```
p = torch.linspace(0.,1,100)
f = combine_scheds([0.3,0.7], [SchedCos(0.3,0.6), SchedCos(0.6,0.2)])
plt.plot(p, [f(o) for o in p]);
p = torch.linspace(0.,1,100)
f = combine_scheds([0.3,0.2,0.5], [SchedLin(0.,1.), SchedNo(1.,1.), SchedCos(1., 0.)])
plt.plot(p, [f(o) for o in p]);
#hide
test_close([f(0.), f(0.15), f(0.3), f(0.4), f(0.5), f(0.7), f(1.)],
[0., 0.5, 1., 1., 1., 0.65451, 0.])
#export
def combined_cos(pct, start, middle, end):
"Return a scheduler with cosine annealing from `start`→`middle` & `middle`→`end`"
return combine_scheds([pct,1-pct], [SchedCos(start, middle), SchedCos(middle, end)])
```
This is a useful helper function for the [1cycle policy](https://sgugger.github.io/the-1cycle-policy.html). `pct` is used for the `start` to `middle` part, `1-pct` for the `middle` to `end`. Handles floats or collection of floats. For example:
```
f = combined_cos(0.25,0.5,1.,0.)
plt.plot(p, [f(o) for o in p]);
#hide
test_close([f(0.), f(0.1), f(0.25), f(0.5), f(1.)], [0.5, 0.67275, 1., 0.75, 0.])
f = combined_cos(0.25, np.array([0.25,0.5]), np.array([0.5,1.]), np.array([0.,0.]))
for a,b in zip([f(0.), f(0.1), f(0.25), f(0.5), f(1.)],
[[0.25,0.5], [0.33638,0.67275], [0.5,1.], [0.375,0.75], [0.,0.]]):
test_close(a,b)
```
## ParamScheduler -
```
#export
class ParamScheduler(Callback):
"Schedule hyper-parameters according to `scheds`"
order,run_valid = 60,False
def __init__(self, scheds): self.scheds = scheds
def before_fit(self):
"Initialize container for hyper-parameters"
self.hps = {p:[] for p in self.scheds.keys()}
def before_batch(self):
"Set the proper hyper-parameters in the optimizer"
self._update_val(self.pct_train)
def _update_val(self, pct):
for n,f in self.scheds.items(): self.opt.set_hyper(n, f(pct))
def after_batch(self):
"Record hyper-parameters of this batch"
for p in self.scheds.keys(): self.hps[p].append(self.opt.hypers[-1][p])
def after_fit(self):
"Save the hyper-parameters in the recorder if there is one"
if hasattr(self.learn, 'recorder') and hasattr(self, 'hps'): self.recorder.hps = self.hps
```
`scheds` is a dictionary with one key for each hyper-parameter you want to schedule, with either a scheduler or a list of schedulers as values (in the second case, the list must have the same length as the the number of parameters groups of the optimizer).
```
#hide
import torch
from torch.utils.data import TensorDataset, DataLoader
from fastai_minima.learner import DataLoaders
from torch import nn
def synth_dbunch(a=2, b=3, bs=16, n_train=10, n_valid=2):
"A simple dataset where `x` is random and `y = a*x + b` plus some noise."
def get_data(n):
x = torch.randn(int(bs*n))
return TensorDataset(x, a*x + b + 0.1*torch.randn(int(bs*n)))
train_ds = get_data(n_train)
valid_ds = get_data(n_valid)
train_dl = DataLoader(train_ds, batch_size=bs, shuffle=True, num_workers=0)
valid_dl = DataLoader(valid_ds, batch_size=bs, num_workers=0)
return DataLoaders(train_dl, valid_dl)
def synth_learner(n_train=10, n_valid=2, lr=defaults.lr, **kwargs):
data = synth_dbunch(n_train=n_train,n_valid=n_valid)
return Learner(data, RegModel(), loss_func=nn.MSELoss(), lr=lr, **kwargs)
class RegModel(nn.Module):
"A r"
def __init__(self):
super().__init__()
self.a,self.b = nn.Parameter(torch.randn(1)),nn.Parameter(torch.randn(1))
def forward(self, x): return x*self.a + self.b
learn = synth_learner()
sched = {'lr': SchedLin(1e-3, 1e-2)}
learn.fit(1, cbs=ParamScheduler(sched))
n = len(learn.dls.train)
test_close(learn.recorder.hps['lr'], [1e-3 + (1e-2-1e-3) * i/n for i in range(n)])
#hide
#test discriminative lrs
from torch import optim
from functools import partial
def _splitter(m): return convert_params([[m.a], [m.b]])
learn = synth_learner(splitter=_splitter)
sched = {'lr': combined_cos(0.5, np.array([1e-4,1e-3]), np.array([1e-3,1e-2]), np.array([1e-5,1e-4]))}
learn.fit(1, cbs=ParamScheduler(sched))
show_doc(ParamScheduler.before_fit)
show_doc(ParamScheduler.before_batch)
show_doc(ParamScheduler.after_batch)
show_doc(ParamScheduler.after_fit)
#export
@patch
def fit_one_cycle(self:Learner, n_epoch, lr_max=None, div=25., div_final=1e5, pct_start=0.25, wd=None,
moms=None, cbs=None, reset_opt=False):
"Fit `self.model` for `n_epoch` using the 1cycle policy."
if self.opt is None: self.create_opt()
self.opt.set_hyper('lr', self.lr if lr_max is None else lr_max)
lr_max = np.array([h['lr'] for h in self.opt.hypers])
scheds = {'lr': combined_cos(pct_start, lr_max/div, lr_max, lr_max/div_final),
'mom': combined_cos(pct_start, *(self.moms if moms is None else moms))}
self.fit(n_epoch, cbs=ParamScheduler(scheds)+L(cbs), reset_opt=reset_opt, wd=wd)
```
The 1cycle policy was introduced by Leslie N. Smith et al. in [Super-Convergence: Very Fast Training of Neural Networks Using Large Learning Rates](https://arxiv.org/abs/1708.07120). It schedules the learning rate with a cosine annealing from `lr_max/div` to `lr_max` then `lr_max/div_final` (pass an array to `lr_max` if you want to use differential learning rates) and the momentum with cosine annealing according to the values in `moms`. The first phase takes `pct_start` of the training. You can optionally pass additional `cbs` and `reset_opt`.
```
#Integration test: training a few epochs should make the model better
learn = synth_learner(lr=1e-2)
xb,yb = learn.dls.one_batch()
init_loss = learn.loss_func(learn.model(xb), yb)
learn.fit_one_cycle(2)
xb,yb = learn.dls.one_batch()
final_loss = learn.loss_func(learn.model(xb), yb)
assert final_loss < init_loss
#Scheduler test
lrs,moms = learn.recorder.hps['lr'],learn.recorder.hps['mom']
test_close(lrs, [combined_cos(0.25,1e-2/25,1e-2,1e-7)(i/20) for i in range(20)])
test_close(moms, [combined_cos(0.25,0.95,0.85,0.95)(i/20) for i in range(20)])
#export
@patch
def plot_sched(self:Recorder, keys=None, figsize=None):
keys = self.hps.keys() if keys is None else L(keys)
rows,cols = (len(keys)+1)//2, min(2, len(keys))
figsize = figsize or (6*cols,4*rows)
_, axs = plt.subplots(rows, cols, figsize=figsize)
axs = axs.flatten() if len(keys) > 1 else L(axs)
for p,ax in zip(keys, axs):
ax.plot(self.hps[p])
ax.set_ylabel(p)
#hide
#test discriminative lrs
def _splitter(m): return convert_params([[m.a], [m.b]])
learn = synth_learner(splitter=_splitter)
learn.fit_one_cycle(1, lr_max=slice(1e-3,1e-2))
#n = len(learn.dls.train)
#est_close(learn.recorder.hps['lr'], [1e-3 + (1e-2-1e-3) * i/n for i in range(n)])
learn = synth_learner()
learn.fit_one_cycle(2)
learn.recorder.plot_sched()
#export
@patch
def fit_flat_cos(self:Learner, n_epoch, lr=None, div_final=1e5, pct_start=0.75, wd=None,
cbs=None, reset_opt=False):
"Fit `self.model` for `n_epoch` at flat `lr` before a cosine annealing."
if self.opt is None: self.create_opt()
self.opt.set_hyper('lr', self.lr if lr is None else lr)
lr = np.array([h['lr'] for h in self.opt.hypers])
scheds = {'lr': combined_cos(pct_start, lr, lr, lr/div_final)}
self.fit(n_epoch, cbs=ParamScheduler(scheds)+L(cbs), reset_opt=reset_opt, wd=wd)
learn = synth_learner()
learn.fit_flat_cos(2)
learn.recorder.plot_sched()
#export
@patch
def fit_sgdr(self:Learner, n_cycles, cycle_len, lr_max=None, cycle_mult=2, cbs=None, reset_opt=False, wd=None):
"Fit `self.model` for `n_cycles` of `cycle_len` using SGDR."
if self.opt is None: self.create_opt()
self.opt.set_hyper('lr', self.lr if lr_max is None else lr_max)
lr_max = np.array([h['lr'] for h in self.opt.hypers])
n_epoch = cycle_len * (cycle_mult**n_cycles-1)//(cycle_mult-1)
pcts = [cycle_len * cycle_mult**i / n_epoch for i in range(n_cycles)]
scheds = [SchedCos(lr_max, 0) for _ in range(n_cycles)]
scheds = {'lr': combine_scheds(pcts, scheds)}
self.fit(n_epoch, cbs=ParamScheduler(scheds)+L(cbs), reset_opt=reset_opt, wd=wd)
```
This schedule was introduced by Ilya Loshchilov et al. in [SGDR: Stochastic Gradient Descent with Warm Restarts](https://arxiv.org/abs/1608.03983). It consists of `n_cycles` that are cosine annealings from `lr_max` (defaults to the `Learner` lr) to 0, with a length of `cycle_len * cycle_mult**i` for the `i`-th cycle (first one is `cycle_len`-long, then we multiply the length by `cycle_mult` at each epoch). You can optionally pass additional `cbs` and `reset_opt`.
```
#slow
learn = synth_learner()
with learn.no_logging(): learn.fit_sgdr(3, 1)
test_eq(learn.n_epoch, 7)
iters = [k * len(learn.dls.train) for k in [0,1,3,7]]
for i in range(3):
n = iters[i+1]-iters[i]
#The start of a cycle can be mixed with the 0 of the previous cycle with rounding errors, so we test at +1
test_close(learn.recorder.lrs[iters[i]+1:iters[i+1]], [SchedCos(learn.lr, 0)(k/n) for k in range(1,n)])
learn.recorder.plot_sched()
#export
@patch
@delegates(Learner.fit_one_cycle)
def fine_tune(self:Learner, epochs, base_lr=2e-3, freeze_epochs=1, lr_mult=100,
pct_start=0.3, div=5.0, **kwargs):
"Fine tune with `freeze` for `freeze_epochs` then with `unfreeze` from `epochs` using discriminative LR"
self.freeze()
self.fit_one_cycle(freeze_epochs, slice(base_lr), pct_start=0.99, **kwargs)
base_lr /= 2
self.unfreeze()
self.fit_one_cycle(epochs, slice(base_lr/lr_mult, base_lr), pct_start=pct_start, div=div, **kwargs)
learn.fine_tune(1)
```
## LRFind -
```
#export
class LRFinder(ParamScheduler):
"Training with exponentially growing learning rate"
def __init__(self, start_lr=1e-7, end_lr=10, num_it=100, stop_div=True):
if is_listy(start_lr):
self.scheds = {'lr': [SchedExp(s, e) for (s,e) in zip(start_lr,end_lr)]}
else: self.scheds = {'lr': SchedExp(start_lr, end_lr)}
self.num_it,self.stop_div = num_it,stop_div
def before_fit(self):
"Initialize container for hyper-parameters and save the model"
super().before_fit()
self.learn.save('_tmp')
self.best_loss = float('inf')
def before_batch(self):
"Record hyper-parameters of this batch and potentially stop training"
self._update_val(self.train_iter/self.num_it)
def after_batch(self):
"Set the proper hyper-parameters in the optimizer"
super().after_batch()
if self.smooth_loss < self.best_loss: self.best_loss = self.smooth_loss
if self.smooth_loss > 4*self.best_loss and self.stop_div: raise CancelFitException()
if self.train_iter >= self.num_it: raise CancelFitException()
def before_validate(self):
"Skip the validation part of training"
raise CancelValidException()
def after_fit(self):
"Save the hyper-parameters in the recorder if there is one and load the original model"
self.learn.opt.zero_grad() #Need to zero the gradients of the model before detaching the optimizer for future fits
tmp_f = self.path/self.model_dir/'_tmp.pth'
if tmp_f.exists():
self.learn.load('_tmp', with_opt=True)
os.remove(tmp_f)
#slow
import tempfile
from fastcore.basics import range_of
from fastcore.xtras import Path
with tempfile.TemporaryDirectory() as d:
learn = synth_learner(path=Path(d))
init_a,init_b = learn.model.a,learn.model.b
with learn.no_logging(): learn.fit(20, cbs=LRFinder(num_it=100))
assert len(learn.recorder.lrs) <= 100
test_eq(len(learn.recorder.lrs), len(learn.recorder.losses))
#Check stop if diverge
if len(learn.recorder.lrs) < 100: assert learn.recorder.losses[-1] > 4 * min(learn.recorder.losses)
#Test schedule
test_eq(learn.recorder.lrs, [SchedExp(1e-7, 10)(i/100) for i in range_of(learn.recorder.lrs)])
#No validation data
test_eq([len(v) for v in learn.recorder.values], [1 for _ in range_of(learn.recorder.values)])
#Model loaded back properly
test_eq(learn.model.a, init_a)
test_eq(learn.model.b, init_b)
test_eq(learn.opt.state_dict()['state'], {})
show_doc(LRFinder.before_fit)
show_doc(LRFinder.before_batch)
show_doc(LRFinder.after_batch)
show_doc(LRFinder.before_validate)
#export
@patch
def plot_lr_find(self:Recorder, skip_end=5):
"Plot the result of an LR Finder test (won't work if you didn't do `learn.lr_find()` before)"
lrs = self.lrs if skip_end==0 else self.lrs [:-skip_end]
losses = self.losses if skip_end==0 else self.losses[:-skip_end]
fig, ax = plt.subplots(1,1)
ax.plot(lrs, losses)
ax.set_ylabel("Loss")
ax.set_xlabel("Learning Rate")
ax.set_xscale('log')
#export
SuggestedLRs = collections.namedtuple('SuggestedLRs', ['lr_min', 'lr_steep'])
#export
@patch
def lr_find(self:Learner, start_lr=1e-7, end_lr=10, num_it=100, stop_div=True, show_plot=True, suggestions=True):
"Launch a mock training to find a good learning rate, return lr_min, lr_steep if `suggestions` is True"
n_epoch = num_it//len(self.dls.train) + 1
cb=LRFinder(start_lr=start_lr, end_lr=end_lr, num_it=num_it, stop_div=stop_div)
with self.no_logging(): self.fit(n_epoch, cbs=cb)
if show_plot: self.recorder.plot_lr_find()
if suggestions:
lrs,losses = tensor(self.recorder.lrs[num_it//10:-5]),tensor(self.recorder.losses[num_it//10:-5])
if len(losses) == 0: return
lr_min = lrs[losses.argmin()].item()
grads = (losses[1:]-losses[:-1]) / (lrs[1:].log()-lrs[:-1].log())
lr_steep = lrs[grads.argmin()].item()
return SuggestedLRs(lr_min/10.,lr_steep)
```
First introduced by Leslie N. Smith in [Cyclical Learning Rates for Training Neural Networks](https://arxiv.org/pdf/1506.01186.pdf), the LR Finder trains the model with exponentially growing learning rates from `start_lr` to `end_lr` for `num_it` and stops in case of divergence (unless `stop_div=False`) then plots the losses vs the learning rates with a log scale.
A good value for the learning rates is then either:
- one tenth of the minimum before the divergence
- when the slope is the steepest
Those two values are returned by default by the Learning Rate Finder.
```
#slow
with tempfile.TemporaryDirectory() as d:
learn = synth_learner(path=Path(d))
weights_pre_lr_find = L(learn.model.parameters())
lr_min,lr_steep = learn.lr_find()
weights_post_lr_find = L(learn.model.parameters())
test_eq(weights_pre_lr_find, weights_post_lr_find)
print(f"Minimum/10: {lr_min:.2e}, steepest point: {lr_steep:.2e}")
```
| github_jupyter |
# <img src="https://img.icons8.com/bubbles/100/000000/3d-glasses.png" style="height:50px;display:inline"> EE 046746 - Technion - Computer Vision
#### Elias Nehme
## Tutorial 14 - Deep Computational Imaging
---
<img src="./assets/tut_14_teaser.gif" style="width:800px">
* <a href="https://www.nature.com/articles/s41377-020-00403-7">Image source</a>
## <img src="https://img.icons8.com/bubbles/50/000000/checklist.png" style="height:50px;display:inline"> Agenda
---
* [What is Computational Imaging?](#-What-is-Computational-Imaging?)
* [Compressive Imaging](#-Compressive-Imaging)
* [Depth Encoding PSF](#-Depth-Encoding-PSF)
* [Deep "Optics"](#-Deep-Optics)
* [Computer Vision Pipelines](#-Computer-Vision-Pipelines)
* [Differentiable Optics](#-Differentiable-Optics)
* [Applications](#-Applications)
* [Recommended Videos](#-Recommended-Videos)
* [Credits](#-Credits)
### <img src="https://img.icons8.com/color/64/000000/fantasy.png" style="height:50px;display:inline"> What is Computational Imaging?
---
<img src="./assets/tut14_ci_1.jpg" width="800">
### <img src="https://img.icons8.com/color/64/000000/fantasy.png" style="height:50px;display:inline"> What is Computational Imaging?
---
<img src="./assets/tut14_ci_2.jpg" width="800">
### <img src="https://img.icons8.com/wired/64/000000/zip.png" style="height:50px;display:inline"> Compressive Imaging
---
* Depth encoding PSF
#### <img src="https://img.icons8.com/wired/64/000000/zip.png" style="height:50px;display:inline"> Depth Encoding PSF
---
* Measurement is a 2D image:
<img src="./assets/tut14_cs_0.jpg" width="400">
* <a href="https://www.osapublishing.org/optica/fulltext.cfm?uri=optica-5-1-1&id=380297">Image source - Optica 2018</a>
#### <img src="https://img.icons8.com/wired/64/000000/zip.png" style="height:50px;display:inline"> Depth Encoding PSF
---
* Recovery is a 3D volume:
<img src="./assets/tut14_cs_1.gif" width="400">
* What?? How?!
* <a href="https://www.osapublishing.org/optica/fulltext.cfm?uri=optica-5-1-1&id=380297">Image source - Optica 2018</a>
#### <img src="https://img.icons8.com/wired/64/000000/zip.png" style="height:50px;display:inline"> Depth Encoding PSF
---
* Depth Encoding Impulse Response/Point Spread Function (PSF)
* Main idea is to encode depth in the shape generated on the 2D sensor
<img src="./assets/tut14_cs_2.gif" width="600">
* <a href="https://www.osapublishing.org/optica/fulltext.cfm?uri=optica-5-1-1&id=380297">Image source - Optica 2018</a>
#### <img src="https://img.icons8.com/wired/64/000000/zip.png" style="height:50px;display:inline"> Depth Encoding PSF
---
* Writing down the problem in matrix formulation
<img src="./assets/tut14_cs_3.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/zip.png" style="height:50px;display:inline"> Depth Encoding PSF
---
* Solution given by "MAP" estimator under certain conditions
<img src="./assets/tut14_cs_4.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/zip.png" style="height:50px;display:inline"> Depth Encoding PSF
---
* Overall framework:
<img src="./assets/tut14_cs_5.jpg" width="800">
* <a href="https://www.osapublishing.org/optica/fulltext.cfm?uri=optica-5-1-1&id=380297">Image source - Optica 2018</a>
### <img src="https://img.icons8.com/wired/64/000000/switch-camera.png" style="height:50px;display:inline"> Deep Optics
---
* Computer Vision Pipelines
* Differentiable Optics
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
<img src="./assets/tut14_cv_pipe_1.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
<img src="./assets/tut14_cv_pipe_2.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
<img src="./assets/tut14_cv_pipe_3.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
<img src="./assets/tut14_cv_pipe_4.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
<img src="./assets/tut14_cv_pipe_5.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
<img src="./assets/tut14_cv_pipe_6.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
<img src="./assets/tut14_cv_pipe_7.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
<img src="./assets/tut14_cv_pipe_8.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
<img src="./assets/tut14_cv_pipe_9.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
* Animal vision is adapted to the surrounding and the day-to-day "task"
<img src="./assets/tut14_cv_pipe_10.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
* "Standard" deep image processing
<img src="./assets/tut14_cv_pipe_11.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
* Deep computational imaging
<img src="./assets/tut14_cv_pipe_12.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
* Image classification with specialized "optics"
<img src="./assets/tut14_cv_pipe_13.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
* Main idea: optimize the optics and the algorithm jointly to excel in the final task
<img src="./assets/tut14_cv_pipe_14.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-identification.png" style="height:50px;display:inline"> Computer Vision Pipelines
---
<img src="./assets/tut14_cv_pipe_15.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
<img src="./assets/tut14_diff_opt_1.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
<img src="./assets/tut14_diff_opt_2.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
<img src="./assets/tut14_diff_opt_3.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
<img src="./assets/tut14_diff_opt_4.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
<img src="./assets/tut14_diff_opt_5.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
<img src="./assets/tut14_diff_opt_6.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
<img src="./assets/tut14_diff_opt_7.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
<img src="./assets/tut14_diff_opt_8.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
<img src="./assets/tut14_diff_opt_9.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
<img src="./assets/tut14_diff_opt_10.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
<img src="./assets/tut14_diff_opt_11.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
<img src="./assets/tut14_diff_opt_12.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
<img src="./assets/tut14_diff_opt_13.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
<img src="./assets/tut14_diff_opt_14.jpg" width="800">
#### <img src="https://img.icons8.com/wired/64/000000/camera-addon.png" style="height:50px;display:inline"> Differentiable Optics
---
<img src="./assets/tut14_diff_opt_15.jpg" width="800">
### <img src="https://img.icons8.com/wired/64/000000/untested.png" style="height:50px;display:inline"> Applications
---
* Extended Depth of Field
* Monocular Depth Estimation / Depth from Defocus
* High Dynamic Range Imaging
* Video Compressive Sensing
* Computational Microscopy (Will not show examples)
#### <img src="https://img.icons8.com/wired/64/000000/aperture.png" style="height:50px;display:inline"> Application 1: Extended Depth of Field (EDOF)
---
<img src="./assets/tut14_app_edof_0.jpg" width="800">
* <a href="https://dl.acm.org/doi/10.1145/3197517.3201333">Image source - ACM SIGGRAPH 2018</a>
#### <img src="https://img.icons8.com/wired/64/000000/aperture.png" style="height:50px;display:inline"> Application 1: Extended Depth of Field (EDOF)
---
<img src="./assets/tut14_app_edof_1.jpg" width="800">
* <a href="https://dl.acm.org/doi/10.1145/3197517.3201333">Image source - ACM SIGGRAPH 2018</a>
#### <img src="https://img.icons8.com/wired/64/000000/aperture.png" style="height:50px;display:inline"> Application 1: Extended Depth of Field (EDOF)
---
<img src="./assets/tut14_app_edof_2.jpg" width="800">
* <a href="https://dl.acm.org/doi/10.1145/3197517.3201333">Image source - ACM SIGGRAPH 2018</a>
#### <img src="https://img.icons8.com/ios-glyphs/64/000000/abscissa.png" style="height:50px;display:inline"> Application 2: Monocular Depth Estimation
---
<img src="./assets/tut14_app_dfd_0.jpg" width="800">
* <a href="https://openaccess.thecvf.com/content_ICCV_2019/papers/Chang_Deep_Optics_for_Monocular_Depth_Estimation_and_3D_Object_Detection_ICCV_2019_paper.pdf">Image source - ICCV 2019</a>
#### <img src="https://img.icons8.com/ios-glyphs/64/000000/abscissa.png" style="height:50px;display:inline"> Application 2: Monocular Depth Estimation
---
<img src="./assets/tut14_app_dfd_1.jpg" width="800">
* <a href="https://openaccess.thecvf.com/content_ICCV_2019/papers/Chang_Deep_Optics_for_Monocular_Depth_Estimation_and_3D_Object_Detection_ICCV_2019_paper.pdf">Image source - ICCV 2019</a>
```
from IPython.display import Video
Video("./assets/input_vid_dfd.mp4")
Video("./assets/output_vid_dfd.mp4")
Video("./assets/bb_3d.mp4")
```
#### <img src="https://img.icons8.com/officel/60/000000/width.png" style="height:50px;display:inline"> Application 3: High Dynamic Range Imaging
---
<img src="./assets/tut14_app_hdr_0.jpg" width="800">
* <a href="https://openaccess.thecvf.com/content_CVPR_2020/papers/Metzler_Deep_Optics_for_Single-Shot_High-Dynamic-Range_Imaging_CVPR_2020_paper.pdf">Image source - CVPR 2020</a>
#### <img src="https://img.icons8.com/officel/60/000000/width.png" style="height:50px;display:inline"> Application 3: High Dynamic Range Imaging
---
<img src="./assets/tut14_app_hdr_1.jpg" width="800">
* <a href="https://openaccess.thecvf.com/content_CVPR_2020/papers/Metzler_Deep_Optics_for_Single-Shot_High-Dynamic-Range_Imaging_CVPR_2020_paper.pdf">Image source - CVPR 2020</a>
#### <img src="https://img.icons8.com/officel/60/000000/width.png" style="height:50px;display:inline"> Application 3: High Dynamic Range Imaging
---
<img src="./assets/tut14_app_hdr_2.jpg" width="800">
* <a href="https://openaccess.thecvf.com/content_CVPR_2020/papers/Metzler_Deep_Optics_for_Single-Shot_High-Dynamic-Range_Imaging_CVPR_2020_paper.pdf">Image source - CVPR 2020</a>
#### <img src="https://img.icons8.com/officel/60/000000/width.png" style="height:50px;display:inline"> Application 3: High Dynamic Range Imaging
---
<img src="./assets/tut14_app_hdr_3.jpg" width="800">
* <a href="https://openaccess.thecvf.com/content_CVPR_2020/papers/Metzler_Deep_Optics_for_Single-Shot_High-Dynamic-Range_Imaging_CVPR_2020_paper.pdf">Image source - CVPR 2020</a>
#### <img src="https://img.icons8.com/material-outlined/48/000000/video-trimming.png" style="height:50px;display:inline"> Application 4: Video Compressive Sensing
---
* Challenge is to learn the binary masks $\Phi$ to recover video $x$ from snapshot $y$
<img src="./assets/tut14_app_vid_cs_0.jpg" width="800">
* <a href="https://www.sciencedirect.com/science/article/pii/S1051200419301459">Image source - DSP Magazine 2020</a>
#### <img src="https://img.icons8.com/material-outlined/48/000000/video-trimming.png" style="height:50px;display:inline"> Application 4: Video Compressive Sensing
---
<img src="./assets/tut14_app_vid_cs_1.jpg" width="800">
* <a href="https://ieeexplore.ieee.org/document/9064896">Image source - ICCP 2020</a>
#### <img src="https://img.icons8.com/material-outlined/48/000000/video-trimming.png" style="height:50px;display:inline"> Application 4: Video Compressive Sensing
---
<img src="./assets/tut14_app_vid_cs_2.jpg" width="400">
* <a href="https://ieeexplore.ieee.org/document/9064896">Image source - ICCP 2020</a>
#### <img src="https://img.icons8.com/material-outlined/48/000000/video-trimming.png" style="height:50px;display:inline"> Application 4: Video Compressive Sensing
---
* 64 frames recovered from 4 measurements (16 frames/capture)
<img src="./assets/tut14_app_vid_cs_3.gif" width="400">
* <a href="https://ieeexplore.ieee.org/document/9064896">Image source - ICCP 2020</a>
### <img src="https://img.icons8.com/color/96/000000/code.png" style="height:50px;display:inline"> Available resources online
---
* https://github.com/computational-imaging/opticalCNN
* https://github.com/vsitzmann/deepoptics
* https://github.com/computational-imaging/DeepOpticsHDR
* https://github.com/EliasNehme/DeepSTORM3D
* https://github.com/computational-imaging/DepthFromDefocusWithLearnedOptics
* etc.
### <img src="https://img.icons8.com/bubbles/50/000000/video-playlist.png" style="height:50px;display:inline"> Recommended Videos
---
#### <img src="https://img.icons8.com/cute-clipart/64/000000/warning-shield.png" style="height:30px;display:inline"> Warning!
* These videos do not replace the lectures and tutorials.
* Please use these to get a better understanding of the material, and not as an alternative to the written material.
#### Video By Subject
* End-to-end Optimization of Optics and Image Processing - <a href="https://www.youtube.com/watch?v=iJdsxXOfqvw&t=266s&ab_channel=DingzeyuLi">Vincent Sitzmann</a>
* Neural Sensors - <a href="https://www.youtube.com/watch?v=tTYHoxA2RVg&ab_channel=StanfordComputationalImagingLab">J.N.P Martel</a>
* High Dynamic Range Imaging - <a href="https://www.youtube.com/watch?v=Pla8p9Nqlb8&ab_channel=StanfordComputationalImagingLab">Christopher Metzler</a>
* 3D Single Molecule Localization Microscopy - <a href="https://app.quantitativebioimaging.com/video/17">Elias Nehme</a>
* Towards Neural Imaging & Signal Processing - <a href="https://www.youtube.com/watch?v=vTio0tuizHw&t=3984s&ab_channel=IEEESignalProcessingSociety">Gordon Wetzstein</a>
## <img src="https://img.icons8.com/dusk/64/000000/prize.png" style="height:50px;display:inline"> Credits
----
* <a href="https://vsitzmann.github.io/deepoptics/">End-to-end optimization of optics and image processing</a> - Vincent Sitzmann
* <a href="https://dl.acm.org/doi/abs/10.1145/3388769.3407486">ACM SIGGRAPH 2020 Courses</a> - Yifan (Evan) Peng, Ashok Veeraraghavan, Wolfgang Heidrich, Gordon Wetzstein
* <a href="http://stanford.edu/class/ee367/">Stanford EE367/CS448I </a> (Computational Imaging and Display) - Gordon Wetzstein
* <a href="https://sites.google.com/view/sps-space/home?authuser=0">IEEE SPACE Webinar</a> - IEEE Computational Imaging TC
* Research papers:
* <a href="https://dl.acm.org/doi/10.1145/3197517.3201333">End-to-end optimization of optics and image processing for achromatic extended depth of field and super-resolution imaging</a>
* <a href="https://openaccess.thecvf.com/content_ICCV_2019/papers/Chang_Deep_Optics_for_Monocular_Depth_Estimation_and_3D_Object_Detection_ICCV_2019_paper.pdf">Deep Optics for Monocular Depth Estimation and 3D Object Detection</a>
* <a href="https://openaccess.thecvf.com/content_CVPR_2020/papers/Metzler_Deep_Optics_for_Single-Shot_High-Dynamic-Range_Imaging_CVPR_2020_paper.pdf">Deep Optics for Single-shot High-dynamic-range Imaging</a>
* <a href="https://ieeexplore.ieee.org/document/9064896">Neural Sensors: Learning Pixel Exposures for HDR Imaging and Video Compressive Sensing With Programmable Sensors</a>
* <a href="https://arxiv.org/abs/1709.07223">Convolutional neural networks that teach microscopes how to image</a>
* <a href="https://www.nature.com/articles/s41592-020-0853-5">DeepSTORM3D: dense 3D localization microscopy and PSF design by deep learning</a>
* <a href="http://www.computationalimaging.org/publications/deepopticsdfd/">Depth From Defocus With Learned Optics</a>
* etc.
* Icons from <a href="https://icons8.com/">Icon8.com</a> - https://icons8.com
| github_jupyter |
# Transfer Learning
In this notebook, you'll learn how to use pre-trained networks to solved challenging problems in computer vision. Specifically, you'll use networks trained on [ImageNet](http://www.image-net.org/) [available from torchvision](http://pytorch.org/docs/0.3.0/torchvision/models.html).
ImageNet is a massive dataset with over 1 million labeled images in 1000 categories. It's used to train deep neural networks using an architecture called convolutional layers. I'm not going to get into the details of convolutional networks here, but if you want to learn more about them, please [watch this](https://www.youtube.com/watch?v=2-Ol7ZB0MmU).
Once trained, these models work astonishingly well as feature detectors for images they weren't trained on. Using a pre-trained network on images not in the training set is called transfer learning. Here we'll use transfer learning to train a network that can classify our cat and dog photos with near perfect accuracy.
With `torchvision.models` you can download these pre-trained networks and use them in your applications. We'll include `models` in our imports now.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import torch
import numpy as np
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms, models
```
Most of the pretrained models require the input to be 224x224 images. Also, we'll need to match the normalization used when the models were trained. Each color channel was normalized separately, the means are `[0.485, 0.456, 0.406]` and the standard deviations are `[0.229, 0.224, 0.225]`.
```
data_dir = 'Cat_Dog_data'
# TODO: Define transforms for the training data and testing data
train_transforms = transforms.Compose([transforms.RandomRotation(30),
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
test_transforms = transforms.Compose([transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
# Pass transforms in here, then run the next cell to see how the transforms look
train_data = datasets.ImageFolder(data_dir + '/train', transform=train_transforms)
test_data = datasets.ImageFolder(data_dir + '/test', transform=test_transforms)
trainloader = torch.utils.data.DataLoader(train_data, batch_size=64, shuffle=True)
testloader = torch.utils.data.DataLoader(test_data, batch_size=32)
```
We can load in a model such as [DenseNet](http://pytorch.org/docs/0.3.0/torchvision/models.html#id5). Let's print out the model architecture so we can see what's going on.
```
model = models.densenet121(pretrained=True)
model
```
This model is built out of two main parts, the features and the classifier. The features part is a stack of convolutional layers and overall works as a feature detector that can be fed into a classifier. The classifier part is a single fully-connected layer `(classifier): Linear(in_features=1024, out_features=1000)`. This layer was trained on the ImageNet dataset, so it won't work for our specific problem. That means we need to replace the classifier, but the features will work perfectly on their own. In general, I think about pre-trained networks as amazingly good feature detectors that can be used as the input for simple feed-forward classifiers.
```
# Freeze parameters so we don't backprop through them
for param in model.parameters():
param.requires_grad = False
from collections import OrderedDict
classifier = nn.Sequential(OrderedDict([
('fc1', nn.Linear(1024, 500)),
('relu', nn.ReLU()),
('fc2', nn.Linear(500, 2)),
('output', nn.LogSoftmax(dim=1))
]))
model.classifier = classifier
```
With our model built, we need to train the classifier. However, now we're using a **really deep** neural network. If you try to train this on a CPU like normal, it will take a long, long time. Instead, we're going to use the GPU to do the calculations. The linear algebra computations are done in parallel on the GPU leading to 100x increased training speeds. It's also possible to train on multiple GPUs, further decreasing training time.
PyTorch, along with pretty much every other deep learning framework, uses [CUDA](https://developer.nvidia.com/cuda-zone) to efficiently compute the forward and backwards passes on the GPU. In PyTorch, you move your model parameters and other tensors to the GPU memory using `model.to('cuda')`. You can move them back from the GPU with `model.to('cpu')` which you'll commonly do when you need to operate on the network output outside of PyTorch. As a demonstration of the increased speed, I'll compare how long it takes to perform a forward and backward pass with and without a GPU.
```
import time
for device in ['cpu', 'cuda']:
criterion = nn.NLLLoss()
# Only train the classifier parameters, feature parameters are frozen
optimizer = optim.Adam(model.classifier.parameters(), lr=0.001)
model.to(device)
for ii, (inputs, labels) in enumerate(trainloader):
# Move input and label tensors to the GPU
inputs, labels = inputs.to(device), labels.to(device)
start = time.time()
outputs = model.forward(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
if ii==3:
break
print(f"DEVICE = {device}; Time per batch: {(time.time() - start)/3:.3f} seconds")
```
You can write device agnostic code which will automatically use CUDA if it's enabled like so:
```python
# at beginning of the script
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
...
# then whenever you get a new Tensor or Module
# this won't copy if they are already on the desired device
input = data.to(device)
model = MyModule(...).to(device)
```
From here, I'll let you finish training the model. The process is the same as before except now your model is much more powerful. You should get better than 95% accuracy easily.
>**Exercise:** Train a pretrained models to classify the cat and dog images. Continue with the DenseNet model, or try ResNet, it's also a good model to try out first. Make sure you are only training the classifier and the parameters for the features part are frozen.
```
# TODO: Train a model with a pre-trained network
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.classifier.parameters(), lr=0.001)
epochs = 3
print_every = 40
steps = 0
# change to cuda
model.to('cuda')
for e in range(epochs):
running_loss = 0
for ii, (inputs, labels) in enumerate(trainloader):
steps += 1
inputs, labels = inputs.to('cuda'), labels.to('cuda')
optimizer.zero_grad()
# Forward and backward passes
outputs = model.forward(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
if steps % print_every == 0:
print("Epoch: {}/{}... ".format(e+1, epochs),
"Loss: {:.4f}".format(running_loss/print_every))
running_loss = 0
correct = 0
total = 0
with torch.no_grad():
for data in testloader:
images, labels = data
outputs = model(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (100 * correct / total))
# Putting the above into functions, so they can be used later
def do_deep_learning(model, trainloader, epochs, print_every, criterion, optimizer, device='cpu'):
epochs = epochs
print_every = print_every
steps = 0
# change to cuda
model.to('cuda')
for e in range(epochs):
running_loss = 0
for ii, (inputs, labels) in enumerate(trainloader):
steps += 1
inputs, labels = inputs.to('cuda'), labels.to('cuda')
optimizer.zero_grad()
# Forward and backward passes
outputs = model.forward(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
if steps % print_every == 0:
print("Epoch: {}/{}... ".format(e+1, epochs),
"Loss: {:.4f}".format(running_loss/print_every))
running_loss = 0
def check_accuracy_on_test(testloader):
correct = 0
total = 0
with torch.no_grad():
for data in testloader:
images, labels = data
outputs = model(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (100 * correct / total))
do_deep_learning(model, trainloader, 3, 40, criterion, optimizer, 'gpu')
check_accuracy_on_test(testloader)
```
| github_jupyter |
[](http://rpi.analyticsdojo.com)
<center><h1>Introduction to Python - Kaggle Baseline</h1></center>
<center><h3><a href = 'http://rpi.analyticsdojo.com'>rpi.analyticsdojo.com</a></h3></center>
# Kaggle Baseline
## Running Code using Kaggle Notebooks
- Kaggle utilizes Docker to create a fully functional environment for hosting competitions in data science.
- You could download/run this locally or view the [published version](https://www.kaggle.com/analyticsdojo/titanic-baseline-models-analyticsdojo-python/editnb) and `fork` it.
- Kaggle has created an incredible resource for learning analytics. You can view a number of *toy* examples that can be used to understand data science and also compete in real problems faced by top companies.
```
!wget https://raw.githubusercontent.com/rpi-techfundamentals/spring2019-materials/master/input/train.csv
!wget https://raw.githubusercontent.com/rpi-techfundamentals/spring2019-materials/master/input/test.csv
import numpy as np
import pandas as pd
# Input data files are available in the "../input/" directory.
# Let's input them into a Pandas DataFrame
train = pd.read_csv("train.csv")
test = pd.read_csv("test.csv")
```
## `train` and `test` set on Kaggle
- The `train` file contains a wide variety of information that might be useful in understanding whether they survived or not. It also includes a record as to whether they survived or not.
- The `test` file contains all of the columns of the first file except whether they survived. Our goal is to predict whether the individuals survived.
```
train.head()
test.head()
```
## Baseline Models: No Survivors
- The Titanic problem is one of classification, and often the simplest baseline of all 0/1 is an appropriate baseline.
- Think of the baseline as the simplest model you can think of that can be used to lend intuition on how your model is working.
- Even if you aren't familiar with the history of the tragedy, by checking out the [Wikipedia Page](https://en.wikipedia.org/wiki/RMS_Titanic) we can quickly see that the majority of people (68%) died.
- As a result, our baseline model will be for no survivors.
```
test["Survived"] = 0
submission = test.loc[:,["PassengerId", "Survived"]]
submission.head()
```
## Write to CSV
The code below will write your dataframe to a CSV.
```
submission.to_csv('everyone_dies.csv', index=False)
```
## Download from Colab
Working on colab requires you to download a file via a google specific package.
```
from google.colab import files
files.download('everyone_dies.csv')
```
## The First Rule of Shipwrecks
- You may have seen it in a movie or read it in a novel, but [women and children first](https://en.wikipedia.org/wiki/Women_and_children_first) has at it's roots something that could provide our first model.
- Now let's recode the `Survived` column based on whether was a man or a woman.
- We are using conditionals to *select* rows of interest (for example, where test['Sex'] == 'male') and recoding appropriate columns.
```
#Here we can code it as Survived, but if we do so we will overwrite our other prediction.
#Instead, let's code it as PredGender
test.loc[test['Sex'] == 'male', 'PredGender'] = 0
test.loc[test['Sex'] == 'female', 'PredGender'] = 1
#test.PredGender.astype(int)
test
submission = test.loc[:,['PassengerId', 'PredGender']]
# But we have to change the column name.
# Option 1: submission.columns = ['PassengerId', 'Survived']
# Option 2: Rename command.
submission.rename(columns={'PredGender': 'Survived'}, inplace=True)
```
## Writeout and then Download your File
Try your first submission to Kaggle!
```
submission.to_csv('women_survive.csv', index=False)
from google.colab import files
files.download('women_survive.csv')
```
| github_jupyter |
### Set Data Path
```
from pathlib import Path
base_dir = Path("data")
train_dir = base_dir/Path("train")
validation_dir = base_dir/Path("validation")
test_dir = base_dir/Path("test")
```
### Image Transform Function
```
from torchvision import transforms
transform = transforms.Compose([
transforms.Resize((224, 224)),
transforms.ToTensor(),
transforms.Normalize(mean=(.5, .5, .5), std=(.5, .5, .5))
])
```
### Load Training Data (x: features, y: labels)
```
from PIL import Image
x, y = [], []
for file_name in train_dir.glob("*.jpg"):
bounding_box_file = file_name.with_suffix('.txt')
with open(bounding_box_file) as file:
lines = file.readlines()
if(len(lines) > 1):
continue
else:
line = lines[0].strip('\n')
(classes, cen_x, cen_y, box_w, box_h) = list(map(float, line.split(' ')))
torch_data = torch.FloatTensor([cen_x, cen_y, box_w, box_h])
y.append(torch_data)
img = Image.open(str(file_name)).convert('RGB')
img = transform(img)
x.append(img)
```
### Put Training Data into Torch Loader
```
import torch.utils.data as Data
tensor_x = torch.stack(x)
tensor_y = torch.stack(y)
torch_dataset = Data.TensorDataset(tensor_x, tensor_y)
loader = Data.DataLoader(dataset=torch_dataset, batch_size=32, shuffle=True, num_workers=2)
```
### Load Pretrained RestNet18 Model
```
import torchvision
model = torchvision.models.resnet18(pretrained=True)
fc_in_size = model.fc.in_features
model.fc = nn.Linear(fc_in_size, 4)
model = model.cuda()
```
### Parameters
```
EPOCH = 10
LR = 1e-3
```
### Loss Function & Optimizer
```
loss_func = nn.SmoothL1Loss().cuda()
opt = torch.optim.Adam(model.parameters(), lr=LR)
```
### Training
```
for epoch in range(EPOCH):
for step, (batch_x, batch_y) in enumerate(loader):
batch_x = batch_x.cuda()
batch_y = batch_y.cuda()
output = model(batch_x)
loss = loss_func(output, batch_y)
opt.zero_grad()
loss.backward()
opt.step()
if(step % 5 == 0):
print("Epoch {} | Step {} | Loss {}".format(epoch, step, loss))
```
### Show some of the Prediction
```
%matplotlib inline
import cv2
from matplotlib import pyplot as plt
import numpy as np
model = model.cpu()
for batch_x, batch_y in loader:
predict = model(batch_x)
for x, pred, y in zip(batch_x, predict, batch_y):
(pos_x, pos_y, box_w, box_h) = pred
pos_x *= 224
pos_y *= 224
box_w *= 224
box_h *= 224
image = transforms.ToPILImage()(x)
img = cv2.cvtColor(np.asarray(image), cv2.COLOR_RGB2BGR)
img = cv2.rectangle(img, (pos_x - box_w/2, pos_y - box_h/2), (pos_x + box_w/2, pos_y + box_h/2), (255, 0, 0), 3)
plt.imshow(img)
plt.show()
break
```
| github_jupyter |
GONG PFSS extrapolation
=======================
Calculating PFSS solution for a GONG synoptic magnetic field map.
First, import required modules
```
import astropy.constants as const
import astropy.units as u
from astropy.coordinates import SkyCoord
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import numpy as np
import sunpy.map
import pfsspy
from pfsspy import coords
from pfsspy import tracing
#from gong_helpers import get_gong_map
%matplotlib inline
import warnings
warnings.filterwarnings("ignore")
```
Load a GONG magnetic field map. If 'gong.fits' is present in the current
directory, just use that, otherwise download a sample GONG map.
```
def get_gong_map():
"""
Automatically download and unzip a sample GONG synoptic map.
"""
if not os.path.exists('190310t0014gong.fits') and not os.path.exists('190310t0014gong.fits.gz'):
import urllib.request
urllib.request.urlretrieve(
'https://gong2.nso.edu/oQR/zqs/201903/mrzqs190310/mrzqs190310t0014c2215_333.fits.gz',
'190310t0014gong.fits.gz')
if not os.path.exists('190310t0014gong.fits'):
import gzip
with gzip.open('190310t0014gong.fits.gz', 'rb') as f:
with open('190310t0014gong.fits', 'wb') as g:
g.write(f.read())
return '190310t0014gong.fits'
gong_fname = get_gong_map()
```
We can now use SunPy to load the GONG fits file, and extract the magnetic
field data.
The mean is subtracted to enforce div(B) = 0 on the solar surface: n.b. it is
not obvious this is the correct way to do this, so use the following lines
at your own risk!
```
gong_map = sunpy.map.Map(gong_fname)
# Remove the mean
gong_map = sunpy.map.Map(gong_map.data - np.mean(gong_map.data), gong_map.meta)
```
The PFSS solution is calculated on a regular 3D grid in (phi, s, rho), where
rho = ln(r), and r is the standard spherical radial coordinate. We need to
define the number of rho grid points, and the source surface radius.
```
nrho = 35
rss = 2.5
```
From the boundary condition, number of radial grid points, and source
surface, we now construct an Input object that stores this information
```
input = pfsspy.Input(gong_map, nrho, rss)
def set_axes_lims(ax):
ax.set_xlim(0, 360)
ax.set_ylim(0, 180)
```
Using the Input object, plot the input field
```
m = input.map
fig = plt.figure()
ax = plt.subplot(projection=m)
m.plot()
plt.colorbar()
ax.set_title('Input field')
set_axes_lims(ax)
```
Now calculate the PFSS solution, and plot the polarity inversion line.
```
output = pfsspy.pfss(input)
# output.plot_pil(ax)
```
Using the Output object we can plot the source surface field, and the
polarity inversion line.
```
ss_br = output.source_surface_br
# Create the figure and axes
fig = plt.figure()
ax = plt.subplot(projection=ss_br)
# Plot the source surface map
ss_br.plot()
# Plot the polarity inversion line
ax.plot_coord(output.source_surface_pils[0])
# Plot formatting
plt.colorbar()
ax.set_title('Source surface magnetic field')
set_axes_lims(ax)
```
It is also easy to plot the magnetic field at an arbitrary height within
the PFSS solution.
```
# Get the radial magnetic field at a given height
ridx = 15
br = output.bc[0][:, :, ridx]
# Create a sunpy Map object using output WCS
br = sunpy.map.Map(br.T, output.source_surface_br.wcs)
# Get the radial coordinate
r = np.exp(output.grid.rc[ridx])
# Create the figure and axes
fig = plt.figure()
ax = plt.subplot(projection=br)
# Plot the source surface map
br.plot(cmap='RdBu')
# Plot formatting
plt.colorbar()
ax.set_title('$B_{r}$ ' + f'at r={r:.2f}' + '$r_{\\odot}$')
set_axes_lims(ax)
```
Finally, using the 3D magnetic field solution we can trace some field lines.
In this case 64 points equally gridded in theta and phi are chosen and
traced from the source surface outwards.
```
%matplotlib inline
fig = plt.figure(figsize=(15, 8))
ax = fig.add_subplot(111, projection='3d' )
tracer = tracing.PythonTracer()
r = 1.2 * const.R_sun
lat = np.linspace(-np.pi / 2, np.pi / 2, 8, endpoint=False)
lon = np.linspace(0, 2 * np.pi, 8, endpoint=False)
lat, lon = np.meshgrid(lat, lon, indexing='ij')
lat, lon = lat.ravel() * u.rad, lon.ravel() * u.rad
seeds = SkyCoord(lon, lat, r, frame=output.coordinate_frame)
field_lines = tracer.trace(seeds, output)
for field_line in field_lines:
color = {0: 'black', -1: 'tab:blue', 1: 'tab:red'}.get(field_line.polarity)
coords = field_line.coords
coords.representation_type = 'cartesian'
ax.plot(coords.x / const.R_sun,
coords.y / const.R_sun,
coords.z / const.R_sun,
color=color, linewidth=1)
ax.set_title('PFSS solution')
plt.show()
# sphinx_gallery_thumbnail_number = 4
```
| github_jupyter |
# Synthesis with a configuration file
Perhaps the simplest approach to Hazel is to use configuration files. In this notebook we show how to use a configuration file to run Hazel in different situations.
## Single pixel synthesis
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as pl
import hazel
import h5py
print(hazel.__version__)
label = ['I', 'Q', 'U', 'V']
```
Configuration files can be used very simply by just instantiating the `Model` with a configuration file. First, let's print the configuration file. The architecture of the file is explained in the documentation.
```
%cat conf_single.ini
```
We use this file and do the synthesis.
```
mod = hazel.Model('../configuration/conf_single.ini', working_mode='synthesis', verbose=3)
mod.synthesize()
```
And now do some plots.
```
f, ax = pl.subplots(nrows=2, ncols=2, figsize=(10,10))
ax = ax.flatten()
for i in range(4):
ax[i].plot(mod.spectrum['spec1'].wavelength_axis - 10830, mod.spectrum['spec1'].stokes[i,:])
for i in range(4):
ax[i].set_xlabel('Wavelength - 10830[$\AA$]')
ax[i].set_ylabel('{0}/Ic'.format(label[i]))
ax[i].set_xlim([-4,3])
pl.tight_layout()
```
It is possible to save the output of a single pixel to a file. In this case, just open the file, do the synthesis, write to the file and close it.
```
mod = hazel.Model('../configuration/conf_single.ini', working_mode='synthesis')
mod.open_output()
mod.synthesize()
mod.write_output()
mod.close_output()
```
The output file contains a dataset for the currently synthesized spectral region. Here you can see the datasets and their content:
```
f = h5py.File('output.h5', 'r')
print(list(f.keys()))
print(list(f['spec1'].keys()))
f['spec1']['stokes'].shape
```
And then we do some plots:
```
fig, ax = pl.subplots(nrows=2, ncols=2, figsize=(10,10))
ax = ax.flatten()
for i in range(4):
ax[i].plot(f['spec1']['wavelength'][:] - 10830, f['spec1']['stokes'][0,0,0,i,:])
for i in range(4):
ax[i].set_xlabel('Wavelength - 10830[$\AA$]')
ax[i].set_ylabel('{0}/Ic'.format(label[i]))
ax[i].set_xlim([-4,3])
pl.tight_layout()
f.close()
```
## Many pixels synthesis
Synthesizing many pixels can be exhausting and time consuming if you do it one by one. For this reason, Hazel can admit HDF5 files with different models for the synthesis.
### Without MPI
The simplest option is to iterate over all pixels with a single CPU. To this end, we make use of the `iterator`. You first instantiate the iterator and then pass the model to the iterator, which will take care of iterating through all pixels.
```
iterator = hazel.Iterator(use_mpi=False)
rank = iterator.get_rank()
mod = hazel.Model('../configuration/conf_nonmpi_syn1d.ini', working_mode='synthesis', verbose=2)
iterator.use_model(model=mod)
iterator.run_all_pixels()
f = h5py.File('output.h5', 'r')
print('(npix,nrandom,ncycle,nstokes,nlambda) -> {0}'.format(f['spec1']['stokes'].shape))
fig, ax = pl.subplots(nrows=2, ncols=2, figsize=(10,10))
ax = ax.flatten()
for j in range(2):
for i in range(4):
ax[i].plot(f['spec1']['wavelength'][:] - 10830, f['spec1']['stokes'][j,0,0,i,:])
for i in range(4):
ax[i].set_xlabel('Wavelength - 10830[$\AA$]')
ax[i].set_ylabel('{0}/Ic'.format(label[i]))
ax[i].set_xlim([-4,1])
pl.tight_layout()
f.close()
```
### With MPI
In case you want to synthesize large maps in a supercomputer, you can use `mpi4py` and run many pixels in parallel. To this end, you pass the `use_mpi=True` keyword to the iterator. Then, this piece of code should be called with `mpiexec` to run it using MPI:
`mpiexec -n 10 python code.py`
```
iterator = hazel.iterator(use_mpi=True)
rank = iterator.get_rank()
mod = hazel.Model('conf_mpi_synh5.ini', working_mode='synthesis', rank=rank)
iterator.use_model(model=mod)
iterator.run_all_pixels()
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
#argument 'thousands=','' avoids that comma is kept in numeric values
dataset_train = pd.read_csv('Google_Price_Train_2012-2016.csv', thousands=',')
#select just the Close column
dataset_train_close = dataset_train['Close']
#change datatype of Close column from object to float
dataset_train_close = pd.to_numeric(dataset_train_close, errors='coerce')
#extract the 'Open' column
dataset_train_open = dataset_train['Open']
#transform series into arrays
dataset_train_open_array = dataset_train_open.values
dataset_train_close_array = dataset_train_close.values
#transpose the close array
#add mean value of close array to beginning of close array
#and remove last value of close array
dataset_train_close_array.mean()
dataset_train_close_array.shape
dataset_train_close_array_insert = np.insert(dataset_train_close_array, 0, dataset_train_close_array.mean())
dataset_train_close_array_insert.shape
#remove last value of array
dataset_train_close_array_transp = dataset_train_close_array_insert[:-1]
dataset_train_close_array_transp.shape
#reshape to vertical structure and make 2D array
open = dataset_train_open_array.reshape(-1, 1)
close = dataset_train_close_array_transp.reshape(-1, 1)
#concatenate the 2 arrays
training_set = np.concatenate((open, close), axis = 1)
training_set.shape
#feature scaling with normalisation
from sklearn.preprocessing import MinMaxScaler
#'sc' is an object of the MinMaxScaler class
#the scaled stock prices will be between 0 and 1
sc = MinMaxScaler(feature_range = (0, 1))
#apply the scaler on the data
training_set_scaled = sc.fit_transform(training_set)
#create data structure with 60 timesteps and 1 output
#timesteps means: for one output in time t, it will check the values
#of the 60 moments before
X_train = []
y_train = []
#60 is the starting point (we need the days before to create first training value)
#1258 is the index of the last day
for i in range(60, 1258):
#get previous 60 values
#'1' specifies the columns (we have two columns)
X_train.append(training_set_scaled[i-60:i, 0:2])
y_train.append(training_set_scaled[i, 0])
#make arrays from the lists
X_train, y_train = np.array(X_train), np.array(y_train)
#reshape data and add more dimensions
#define number of indicators
#currently we have two indicators ('open' and 'close' stock price)
#we for example can add stock prices of another company
#'X_train.shape[0]' is the number of lines
#'X_train.shape[1]' is the number of columns
#'2' is the number of indicators (here we take 2 indicators)
X_train = np.reshape(X_train, (X_train.shape[0], X_train.shape[1], 2))
X_train.shape
#import keras libraries & packages
#that helps us to set up a sequence of layers
from keras.models import Sequential
#'Dense' produces the output layer
from keras.layers import Dense
#getting the LSTM layers
from keras.layers import LSTM
#use dropout to avoid overfitting
from keras.layers import Dropout
from keras import optimizers
#initialize the RNN
#'regressor' is used to predict a continuous value
#'regressor' is an object of the 'Sequential' class
regressor = Sequential()
#add first LSTM layer and dropout regularization
#LSTM object has 3 arguments:
#1) number of units (neurons; LSTM cells)
#2) return sequences: has to be true, cause we build a stacked LSTM
#3) input shape (relates to shape of X_train; 2 last dimensions are enough)
regressor.add(LSTM(units = 50, return_sequences = True, input_shape = (X_train.shape[1], 2)))
#add dropout regularization
#Dropout rate has one argument:
#--> dropout rate (number of neurons, that should be switched off
#during each iteration of the training)
regressor.add(Dropout(0.2))
#2nd layer
#we don't need to specifiy input shape anymore
regressor.add(LSTM(units = 50, return_sequences = True))
regressor.add(Dropout(0.2))
#3nd layer
regressor.add(LSTM(units = 50, return_sequences = True))
regressor.add(Dropout(0.2))
#4th layer
#we remove 'return_sequences = True' and keep the default value --> 'false'
regressor.add(LSTM(units = 50))
regressor.add(Dropout(0.2))
#add output layer
regressor.add(Dense(units = 1))
#compile RNN with right optimizer
#and right loss function
#use 'compile' method of the 'sequential' class
#different optimizers can be found on keras documentation
regressor.compile(optimizer = 'Adam', loss = 'mean_squared_error')
#fit RNN to training set
#'batch_size' means: in every epoch, the model takes 32 observations
#and updates the weights accordingly
# 1258 total observations / 32 observations per batch =^ 38
regressor.fit(X_train, y_train, epochs = 100, batch_size = 32)
#save the model
from keras.models import load_model
regressor.save('reg_ind2_open_close_transp_2012-2016.h5')
#making predictions and visualize results
#get real stock price of January 2017
dataset_test = pd.read_csv('Google_Stock_Price_Test.csv')
#only consider 'Open' and 'Close' columns
dataset_test = dataset_test[['Open', 'Close']]
#real_stock_price = dataset_test.iloc[:, 1:3].values
real_stock_price = dataset_test.values
#transpose the close array
#add mean value of close array to beginning of close array
#and remove last value of close
real_stock_price[:, 1].mean()
#insert the mean value of the close column at the first position of the close column
real_stock_price_close_insert = np.insert(real_stock_price[:, 1], 0, real_stock_price[:, 1].mean())
real_stock_price_close_insert.shape
#remove last value of array
real_stock_price_close_transp = real_stock_price_close_insert[:-1]
real_stock_price_close_transp.shape
#reshape to vertical structure and make 2D array
close_test = real_stock_price_close_transp.reshape(-1, 1)
#remove previous close column and
#concatenate close_test to real_stock_price_transp
real_stock_price_transp = real_stock_price[:,0].reshape(-1, 1)
real_stock_price_transp_final = np.concatenate((real_stock_price_transp, close_test), axis = 1)
#'real_stock_price_transp_final' contains 20 observations
real_stock_price_transp_final.shape
#get predicted stock price of 2017
#in order to predict stockprice of one day in Jan 17,
#we need 60 previous days
#for these 60 days, we need training and test set
#we will concatenate the initial dataframes
#then we will scale the values
#we have to scale the input values
#cause the rnn was trained on the scaled values
#for vertical concatenation, we take: axis = 0
#for horizontal concatenation, we take: axis = 1
dataset_total = pd.concat((dataset_train[['Open', 'Close']], dataset_test[['Open', 'Close']]), axis = 0)
#get inputs of 60 previous days
#first financial day that we want to predict is Jan 3
#we get the index with this expression: [len(dataset_total) - len(dataset_test)]
#--> length of dataset_test is 20
#we get the lower bound with this expression: len(dataset_total) - len(dataset_test) - 60
#the upper bound is the last index of the whole dataset
#with 'values' we get a numpy array
#'inputs' gives us all information to predict values of Jan 2017
inputs = dataset_total[len(dataset_total) - len(dataset_test) - 60:].values
#reshape to get right numpy shape
#now we have the observations in lines and in 2 columns
inputs = inputs.reshape(-1, 2)
inputs.shape
#transpose close values for
#inputs
#transpose the close array
#add mean value of close array to beginning of close array
#and remove last value of close
inputs[:, 1].mean()
#insert the mean value of the close column at the first position of the close column
inputs_close_insert = np.insert(inputs[:, 1], 0, inputs[:, 1].mean())
#remove last value of array
inputs_close_transp = inputs_close_insert[:-1]
#reshape to vertical structure and make 2D array
inputs_close = inputs_close_transp.reshape(-1, 1)
#remove previous close column and
#concatenate close_test to real_stock_price_transp
inputs_transp = inputs[:,0].reshape(-1, 1)
inputs_transp_final = np.concatenate((inputs_transp, inputs_close), axis = 1)
#'inputs_transp_final' contains 80 observations
inputs_transp_final.shape
#scale the inputs (but not the test values)
#here we don't take the 'fit_transform' method, because
#'sc' was already prepared
inputs_transp_final = sc.transform(inputs_transp_final)
#make 3d structure for test set
X_test = []
#upper bound is 80: 60 + 20 (we have 20 financial days in the test set)
for i in range(60, 80):
#get previous 60 values for each of the stock prices in Jan 2017
#':' specifies the columns (we take all columns)
X_test.append(inputs_transp_final[i-60:i, :])
#make arrays from the lists
X_test = np.array(X_test)
#reshape data and add more dimensions
#define number of indicators
#currently we have one indicator ('open' stock price)
#we for example can add stock prices of another company
#'X_train.shape[0]' is the number of lines
#'X_train.shape[1]' is the number of columns
#'2' is the number of indicators (here we only take google stock price)
X_test = np.reshape(X_test, (X_test.shape[0], X_test.shape[1], 2))
```
## Get predictions based on training set from 2012 to 2016
```
from keras.models import load_model
regressor = load_model('reg_ind2_open_close_transp_2012-2016.h5')
#make predictions on the values of X_test (Jan 2017)
predicted_stock_price = regressor.predict(X_test)
#make 2D structure of array
predicted_stock_price_2D = np.reshape(predicted_stock_price, (-1, 1))
#change training_set_scaled[:,1] to 2D array and only take last 20 values
#B = np.reshape(A, (-1, 2))
close_price_2D = np.reshape(training_set_scaled[:20,1], (-1, 1))
#add close_price_2D to array 'predicted_stock_price_2D'
predicted_stock_close_price_2D = np.concatenate((predicted_stock_price_2D, close_price_2D), axis = 1)
#inverse scaling of predictions
predicted_stock_close_price_2D = sc.inverse_transform(predicted_stock_close_price_2D)
#visualize the results
plt.plot(real_stock_price[:, 0], color = 'red', label = 'Real Google Stock Price for Jan 2017')
plt.plot(predicted_stock_close_price_2D[:, 0], color = 'blue', label = 'Predicted Google Stock Price for Jan 2017')
plt.title('Google Stock Price Prediction (based on training between 2012 & 2016),\n considering indicators Open Price and transposed Close Price')
plt.xlabel('Time')
plt.ylabel('Stock Price')
plt.legend()
plt.show()
import math
from sklearn.metrics import mean_squared_error
#divide rmse by 840 (range of stock price of Jan 2017) --> so we get a relative stock price
rmse_04_16 = (math.sqrt(mean_squared_error(real_stock_price[:, 0], predicted_stock_close_price_2D[:, 0]))) / 840
rmse_04_16
#reshape arrays to vertical structure
real_stock_vert = np.reshape(real_stock_price[:, 0], (-1, 1))
pred_stock_vert = np.reshape(predicted_stock_close_price_2D[:, 0], (-1, 1))
#concat arrays horizontally
arrays_concat = np.concatenate((real_stock_vert, pred_stock_vert), axis = 1)
real_val_list = arrays_concat[:,0].tolist()
pred_val_list = arrays_concat[:,1].tolist()
#loop through columns of array and check directions
#if second value bigger than first value: give a 2
#if second value smaller than first value: give a 1
#get directions of real values
direction_list_realval = []
n = 0
for x in real_val_list:
if real_val_list[n] > real_val_list[n-1]:
direction_list_realval.append(2)
else:
direction_list_realval.append(1)
n = n + 1
#get directions of predicted values
direction_list_predval = []
n = 0
for x in pred_val_list:
if pred_val_list[n] > pred_val_list[n-1]:
direction_list_predval.append(2)
else:
direction_list_predval.append(1)
n = n + 1
#change lists to arrays
real_val_array = np.array(real_val_list)
direction_array_realval = np.array(direction_list_realval)
pred_val_array = np.array(pred_val_list)
direction_array_predval = np.array(direction_list_predval)
#reshape to 2D array
real_val_2d = np.reshape(real_val_array, (-1, 1))
direction_2d_realval = np.reshape(direction_array_realval, (-1, 1))
pred_val_2d = np.reshape(pred_val_array, (-1, 1))
direction_2d_predval = np.reshape(direction_array_predval, (-1, 1))
val_dir = np.concatenate((real_val_2d, direction_2d_realval, pred_val_2d, direction_2d_predval), axis = 1)
#select direction columns from 2D array
list_ind = [1, 3]
val_bin = val_dir[:,list_ind]
correct_list = []
for x in val_bin:
if x[0] == x[1]:
correct_list.append(1)
else:
correct_list.append(0)
correct_list_array = np.reshape(correct_list, (-1, 1))
val_bin_correct = np.concatenate((val_bin, correct_list_array), axis = 1)
#calculate direction accuracy (in percent)
dir_acc = sum(val_bin_correct[:,2]) / len(val_bin_correct)
dir_acc
```
| github_jupyter |
## Example for class imbalance
The dataset used in this notebook is of '[IEEE-CIS Fraud Detection](https://www.kaggle.com/c/ieee-fraud-detection/data)'. This notebook will introduce you to class imbalance problem.
Data set link: [Fraud Dataset](https://drive.google.com/file/d/1q8SYcjOJULdSkETv5S_gd7xNq1GrBHAO/view)
The dataset can be directly accessed with the link: https://raw.githubusercontent.com/dphi-official/Imbalanced_classes/master/fraud_data.csv
#### Imbalanced Problem
Imbalanced classes are a common problem in machine learning classification where there are a disproportionate ratio of observations in each class. Class imbalance can be found in many different areas including medical diagnosis, spam filtering, and fraud detection.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# importing the fraud dataset
fraud_data = pd.read_csv("https://raw.githubusercontent.com/dphi-official/Imbalanced_classes/master/fraud_data.csv")
# take a look at the data
fraud_data.shape
fraud_data.head()
# Taking a look at the target variable
# isFraud = 0 --> normal transaction
# isFraud = 1 --> fraudulent transaction
fraud_data.isFraud.value_counts()
fraud_data.isFraud.value_counts(normalize=True) * 100
# visualize the target variable column
sns.countplot(fraud_data.isFraud)
# interpreting the results - 3.40% of transactions are fraudulent and 99.60% of transactions are normal
# Missing values - To get percentage of missing data in each column
fraud_data.isnull().sum() / len(fraud_data) * 100
# getting all the numerical columns
num_cols = fraud_data.select_dtypes(include=np.number).columns
# filling missing values of numerical columns with mean value
fraud_data[num_cols] = fraud_data[num_cols].fillna(fraud_data[num_cols].mean()) # fills the missing values with mean
# getting all the categorical columns
cat_cols = fraud_data.select_dtypes(include = 'object').columns
# fills the missing values with maximum occuring element in the column
fraud_data[cat_cols] = fraud_data[cat_cols].fillna(fraud_data[cat_cols].mode().iloc[0])
# Let's have a look if there still exist any missing values
fraud_data.isnull().sum() / len(fraud_data) * 100
```
## One hot encoding
Machine learning models require all input and output variables to be numeric. Run the model with data as-is and then iterate for feature engineering. This means that if your data contains categorical data, you must encode it to numbers before you can fit and evaluate a model.
Ordinal encoding - The integer values have a natural ordered relationship between each other and machine learning algorithms may be able to understand and harness this relationship.
For categorical variables where no ordinal relationship exists, the integer encoding may not be enough, at best, or misleading to the model at worst. In this case, a one-hot encoding can be applied to the ordinal representation.
The one-hot encoding creates one binary variable for each category. The problem is that this representation includes redundancy. For example, if we know that [1, 0, 0] represents “blue” and [0, 1, 0] represents “green” we don’t need another binary variable to represent “red“, instead we could use 0 values for both “blue” and “green” alone, e.g.. [0, 0].
This is called a dummy variable encoding, and always represents C categories with C-1 binary variables. In addition to being slightly less redundant, a dummy variable representation is required for some models.
For example, in the case of a linear regression model (and other regression models that have a bias term), a one hot encoding will case the matrix of input data to become singular, meaning it cannot be inverted and the linear regression coefficients cannot be calculated using linear algebra. For these types of models a dummy variable encoding must be used instead.
```
# earlier we have collected all the categorical columns in cat_cols
fraud_data.shape
fraud_data[cat_cols] = fraud_data[cat_cols].fillna(fraud_data[cat_cols].mode().iloc[0])
fraud_data = pd.get_dummies(fraud_data, columns=cat_cols)
fraud_data.shape
fraud_data.head()
new_df = fraud_data.drop(['DeviceInfo_rv:54.0'], axis=1)
# new_df['DeviceInfo_rv:54.0']
```
## Feature transformation
In most cases, the numerical features of the dataset do not have a certain range and they differ from each other. In real life, it is nonsense to expect age and income columns to have the same range. But from the machine learning point of view, how these two columns can be compared?
Scaling solves this problem.
The continuous features become identical in terms of the range, after a scaling process. This process is not mandatory for many algorithms, but it might be still nice to apply. However, the algorithms based on distance calculations such as k-NN or k-Means need to have scaled continuous features as model input.
*Normalization (or min-max normalization)* scale all values in a fixed range between 0 and 1. This transformation does not change the distribution of the feature and due to the decreased standard deviations, the effects of the outliers increases. Therefore, before normalization, it is recommended to handle the outliers.
*Standardization (or z-score normalization)* scales the values while taking into account standard deviation. If the standard deviation of features is different, their range also would differ from each other. This reduces the effect of the outliers in the features.
```
# Separate input features and output feature
# input features
X = fraud_data.drop(columns = ['isFraud'])
# output feature
Y = fraud_data.isFraud
from sklearn.preprocessing import StandardScaler
scaled_features = StandardScaler().fit_transform(X)
scaled_features = pd.DataFrame(data=scaled_features)
scaled_features.columns= X.columns
# Let's see how the data looks after scaling
scaled_features.head()
```
## Splitting the data
##### Split into train and validation set
Train – what we use to train the model
Validation – what we use to evaluate the model
Test – data that is unexposed to the model
```
fraud_data.isFraud.value_counts()
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.3, random_state = 42, stratify=Y)
# X_train: independent feature data for training the model
# Y_train: dependent feature data for training the model
# X_test: independent feature data for testing the model; will be used to predict the target values
# Y_test: original target values of X_test; We will compare this values with our predicted values.
# test_size = 0.3: 30% of the data will go for test set and 70% of the data will go for train set
# random_state = 42: this will fix the split i.e. there will be same split for each time you run the code
```
## Dealing with imbalanced data
An imbalanced classification problem is an example of a classification problem where the distribution of examples across the known classes is biased or skewed ( let us look only at binary classification)
Imbalance can occur due to:
- Biased sampling – E.g., Sampling only from a single geographic location
- Nature of the problem statement – E.g., any fraudulent transactions like credit card frauds, etc.
The imbalance could be
- Slight Imbalance (gender distribution – 60% male; 40% female)
- Severe Imbalance (claims prediction in insurance)
- Terms (Minority class - that has few examples; Majority class - that has many examples)
### Over sampling minority class
Oversampling can be defined as adding more copies of the minority class. In other words, we are creating artificial/synthetic data of the minority class (or group). Oversampling could be a good choice when you don’t have a lot of data to work with.
```
# 'resample' is located under sklearn.utils
from sklearn.utils import resample
# concatenate training data back together
train_data = pd.concat([X_train, Y_train], axis = 1)
# separate minority and majority class
not_fraud = train_data[train_data.isFraud==0]
fraud = train_data[train_data.isFraud==1]
# Unsample minority; we are oversampling the minority class to match the number of majority classs
fraud_upsampled = resample(fraud,
replace = True, # Sample with replacement
n_samples = len(not_fraud), # Match number in majority class
random_state=27)
# combine majority and upsampled minority
upsampled = pd.concat([not_fraud, fraud_upsampled])
# Now let's check the classes count
upsampled.isFraud.value_counts()
# We can notice here after resampling we have an equal ratio of data points for each class!
```
### Under sampling majority class
Undersampling can be defined as removing some observations of the majority class. Undersampling can be a good choice when you have a ton of data -think millions of rows. But a drawback is that we are removing information that may be valuable. This could lead to underfitting and poor generalization to the test set.
```
# we are still using our separated class i.e. fraud and not_fraud from above
# Again we are removing the observations of the majority class to mathch the number of minority class
# downsample majority
not_fraud_downsampled = resample(not_fraud,
replace = False, # sample without replacement
n_samples = len(fraud), # match minority n
random_state = 27)
# combine minority and downsampled majority
downsampled = pd.concat([not_fraud_downsampled, fraud]) # Concatenation
# let's check the classes counts
downsampled.isFraud.value_counts()
# we have an equal ratio of fraud to not fraud data points, but in this case
# a much smaller quantity of data to train the model on.
```
### SMOTE - Synthetic Minority Oversampling Technique
Here we will use imblearn’s SMOTE or Synthetic Minority Oversampling Technique. SMOTE uses a nearest neighbors' algorithm to generate new and synthetic data we can use for training our model.
```
# !pip install delayed
```
Having trouble with SMOTE installation
-> Download using `conda install -c conda-forge imbalanced-learn`
```
# import SMOTE
from imblearn import under_sampling
from imblearn import over_sampling
from imblearn.over_sampling import SMOTE
# random_state = 25, sampling_strategy = 1.0
sm = SMOTE() # again we are eqalizing both the classes
# fit the sampling
X_train, Y_train = sm.fit_resample(X_train, Y_train)
X_train.head()
Y_train.head()
# distribution of target class after sythetic sampling
Y_train.value_counts()
```
# Modelling
```
from sklearn.linear_model import LogisticRegression # Logistic regression model
from sklearn.feature_selection import SelectFromModel
from sklearn.metrics import accuracy_score, f1_score
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
from sklearn.metrics import
model = LogisticRegression() # The maximum number of iterations will be 1000. This will help you prevent from convergence warning.
# model.fit(X_train,Y_train)
f1_score(Y_test, model.predict(X_test))
forest = RandomForestClassifier(random_state=1, n_estimators=1000, max_depth=5)
forest.fit(X_train, Y_train)
#Model Evaluation
f1_score(Y_test, forest.predict(X_test))
```
# End
| github_jupyter |
# Detecting and mitigating age bias on credit decisions
The goal of this tutorial is to introduce the basic functionality of AI Fairness 360.
### Biases and Machine Learning
A machine learning model makes predictions of an outcome for a particular instance. (Given an instance of a loan application, predict if the applicant will repay the loan.) The model makes these predictions based on a training dataset, where many other instances (other loan applications) and actual outcomes (whether they repaid) are provided. Thus, a machine learning algorithm will attempt to find patterns, or generalizations, in the training dataset to use when a prediction for a new instance is needed. (For example, one pattern it might discover is "if a person has salary > USD 40K and has outstanding debt < USD 5, they will repay the loan".) In many domains this technique, called supervised machine learning, has worked very well.
However, sometimes the patterns that are found may not be desirable or may even be illegal. For example, a loan repay model may determine that age plays a significant role in the prediction of repayment because the training dataset happened to have better repayment for one age group than for another. This raises two problems: 1) the training dataset may not be representative of the true population of people of all age groups, and 2) even if it is representative, it is illegal to base any decision on a applicant's age, regardless of whether this is a good prediction based on historical data.
AI Fairness 360 is designed to help address this problem with _fairness metrics_ and _bias mitigators_. Fairness metrics can be used to check for bias in machine learning workflows. Bias mitigators can be used to overcome bias in the workflow to produce a more fair outcome.
The loan scenario describes an intuitive example of illegal bias. However, not all undesirable bias in machine learning is illegal it may also exist in more subtle ways. For example, a loan company may want a diverse portfolio of customers across all income levels, and thus, will deem it undesirable if they are making more loans to high income levels over low income levels. Although this is not illegal or unethical, it is undesirable for the company's strategy.
As these two examples illustrate, a bias detection and/or mitigation toolkit needs to be tailored to the particular bias of interest. More specifically, it needs to know the attribute or attributes, called _protected attributes_, that are of interest: race is one example of a _protected attribute_ and age is a second.
### The Machine Learning Workflow
To understand how bias can enter a machine learning model, we first review the basics of how a model is created in a supervised machine learning process.

First, the process starts with a _training dataset_, which contains a sequence of instances, where each instance has two components: the features and the correct prediction for those features. Next, a machine learning algorithm is trained on this training dataset to produce a machine learning model. This generated model can be used to make a prediction when given a new instance. A second dataset with features and correct predictions, called a _test dataset_, is used to assess the accuracy of the model.
Since this test dataset is the same format as the training dataset, a set of instances of features and prediction pairs, often these two datasets derive from the same initial dataset. A random partitioning algorithm is used to split the initial dataset into training and test datasets.
Bias can enter the system in any of the three steps above. The training data set may be biased in that its outcomes may be biased towards particular kinds of instances. The algorithm that creates the model may be biased in that it may generate models that are weighted towards particular features in the input. The test data set may be biased in that it has expectations on correct answers that may be biased. These three points in the machine learning process represent points for testing and mitigating bias. In AI Fairness 360 codebase, we call these points _pre-processing_, _in-processing_, and _post-processing_.
## AI Fairness 360
We are now ready to utilize AI Fairness 360 (`aif360`) to detect and mitigate bias. We will use the German credit dataset, splitting it into a training and test dataset. We will look for bias in the creation of a machine learning model to predict if an applicant should be given credit based on various features from a typical credit application. The protected attribute will be "Age", with "1" (older than or equal to 25) and "0" (younger than 25) being the values for the privileged and unprivileged groups, respectively.
For this first tutorial, we will check for bias in the initial training data, mitigate the bias, and recheck. More sophisticated machine learning workflows are given in the author tutorials and demo notebooks in the codebase.
Here are the steps involved
#### Step 1: Write import statements
#### Step 2: Set bias detection options, load dataset, and split between train and test
#### Step 3: Compute fairness metric on original training dataset
#### Step 4: Mitigate bias by transforming the original dataset
#### Step 5: Compute fairness metric on transformed training dataset
## Step 1 Import Statements
As with any python program, the first step will be to import the necessary packages. Below we import several components from the `aif360` package. We import the GermanDataset, metrics to check for bias, and classes related to the algorithm we will use to mitigate bias.
```
# Load all necessary packages
import sys
sys.path.insert(1, "../")
import numpy as np
np.random.seed(0)
from aif360.datasets import GermanDataset
from aif360.metrics import BinaryLabelDatasetMetric, DatasetMetric
from aif360.algorithms.preprocessing import Reweighing
from aif360.explainers import MetricTextExplainer, MetricJSONExplainer
from IPython.display import Markdown, display
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
import json
from collections import OrderedDict
```
## Step 2 Load dataset, specifying protected attribute, and split dataset into train and test
In Step 2 we load the initial dataset, setting the protected attribute to be age. We then split the original dataset into training and testing datasets. Although we will use only the training dataset in this tutorial, a normal workflow would also use a test dataset for assessing the efficacy (accuracy, fairness, etc.) during the development of a machine learning model. Finally, we set two variables (to be used in Step 3) for the privileged (1) and unprivileged (0) values for the age attribute. These are key inputs for detecting and mitigating bias, which will be Step 3 and Step 4.
#### What is the German Credit Risk dataset?
The original dataset contains 1000 entries with 20 categorial/symbolic attributes prepared by Prof. Hofmann. In this dataset, each entry represents a person who takes a credit by a bank. Each person is classified as good or bad credit risks according to the set of attributes. The original dataset can be found at https://archive.ics.uci.edu/ml/datasets/Statlog+%28German+Credit+Data%29
#### Loading dataset
<b>protected_attribute</b> means the attribute on which the bias can occur, basically the attribute you want to test bias for.
<b>privileged_classes</b> means a subset of protected attribute values which are considered privileged from a fairness perspective.
In the german dataset: Old (age >= 25) are the privileged class and Young (age < 25) are the unprivileged class.
Here we have binary membership in a protected group (age) and this is a binary classification problem.
Here, age -> sensitive attribute and Old (age >= 25) is the protected group -> historically systematic advantage group.
The dataset is already encoded as the algorithms need the dataset to have numerical values and not categorical.
```
dataset_orig = GermanDataset(protected_attribute_names=['age'], # this dataset also contains protected
# attribute for "sex" which we do not
# consider in this evaluation
privileged_classes=[lambda x: x >= 25], # age >=25 is considered privileged
features_to_drop=['personal_status', 'sex']) # ignore sex-related attributes
dataset_orig_train, dataset_orig_test = dataset_orig.split([0.7], shuffle=True)
privileged_groups = [{'age': 1}]
unprivileged_groups = [{'age': 0}]
print("Original one hot encoded german dataset shape: ",dataset_orig.features.shape)
print("Train dataset shape: ", dataset_orig_train.features.shape)
print("Test dataset shape: ", dataset_orig_test.features.shape)
```
The object dataset_orig is an aif360 dataset, which has some useful methods and attributes taht you can explore:
<b>instance_weights: </b> Weighting for each instance. All equal (ones) by default.
<b>metadata</b> : returns a dict which contains details about the creation of the dataset.
<b>convert_to_dataframe</b> : converts a structured dataset to a pandas dataframe.
<b>de_dummy_code = True</b> : converts dummy-coded columns to categories.
<b>set_category = True</b> : sets the de-dummy coded features to categorical type.
More documentation is available at https://aif360.readthedocs.io/en/latest/modules/datasets.html.
For now, we'll just transform it into a pandas dataframe.
```
df, dict_df = dataset_orig.convert_to_dataframe()
print("Shape: ", df.shape)
print(df.columns)
df.head(5)
```
Let's take a look at our primary variables of interest.
```
print("Key: ", dataset_orig.metadata['protected_attribute_maps'][1])
df['age'].value_counts().plot(kind='bar')
plt.xlabel("Age (0 = under 25, 1 = over 25)")
plt.ylabel("Frequency")
print("Key: ", dataset_orig.metadata['label_maps'])
df['credit'].value_counts().plot(kind='bar')
plt.xlabel("Credit (1 = Good Credit, 2 = Bad Credit)")
plt.ylabel("Frequency")
```
Take a minute to explore the relationship between these two variables. Do credit scores vary with age?
```
# Your code here
```
## Step 3 Compute fairness metric on original training dataset
Now that we've identified the protected attribute 'age' and defined privileged and unprivileged values, we can use aif360 to detect bias in the dataset.
##### Mean Difference (same as statistical parity)
Compare the percentage of favorable results for the privileged and unprivileged groups, subtracting the former percentage from the latter.
The ideal value of this metric is 0.
A value < 0 indicates less favorable outcomes for the unprivileged groups. This is implemented in the method called mean_difference on the BinaryLabelDatasetMetric class. The code below performs this check and displays the output, showing that the difference is -0.169905.
```
metric_orig_train = BinaryLabelDatasetMetric(dataset_orig_train,
unprivileged_groups=unprivileged_groups,
privileged_groups=privileged_groups)
print("Original training dataset")
print("Difference in mean outcomes between unprivileged and privileged groups = %f" % metric_orig_train.mean_difference())
```
##### Disparate Impact
Computed as the ratio of rate of favorable outcome for the unprivileged group to that of the privileged group. The ideal value of this metric is 1.0.
A value < 1 implies higher benefit for the privileged group and a value >1 implies a higher benefit for the unprivileged group.
```
print("Original training dataset")
print("Disparate Impact = %f" % metric_orig_train.disparate_impact())
```
#### Explainers
###### Text Explanations
```
text_expl = MetricTextExplainer(metric_orig_train)
json_expl = MetricJSONExplainer(metric_orig_train)
#print(text_expl.mean_difference())
#print(text_expl.disparate_impact())
```
##### JSON Explanations
```
def format_json(json_str):
return json.dumps(json.loads(json_str, object_pairs_hook=OrderedDict), indent=2)
#print(format_json(json_expl.mean_difference()))
#print(format_json(json_expl.disparate_impact()))
```
## Step 4 Mitigate bias by transforming the original dataset
The previous step showed that the privileged group was getting 17% more positive outcomes in the training dataset. Since this is not desirable, we are going to try to mitigate this bias in the training dataset. As stated above, this is called _pre-processing_ mitigation because it happens before the creation of the model.
AI Fairness 360 implements several pre-processing mitigation algorithms. We will choose the Reweighing algorithm [1], which is implemented in the `Reweighing` class in the `aif360.algorithms.preprocessing` package. This algorithm will transform the dataset to have more equity in positive outcomes on the protected attribute for the privileged and unprivileged groups.
We then call the fit and transform methods to perform the transformation, producing a newly transformed training dataset (dataset_transf_train).
`[1] F. Kamiran and T. Calders, "Data Preprocessing Techniques for Classification without Discrimination," Knowledge and Information Systems, 2012.`
<b>Reweighing:</b> Reweighing is a data preprocessing technique that recommends generating weights for the training examples in each (group, label) combination differently to ensure fairness before classification. The idea is to apply appropriate weights to different tuples in the training dataset to make the training dataset discrimination free with respect to the sensitive attributes. Instead of reweighing, one could also apply techniques (non-discrimination constraints) such as suppression (remove sensitive attributes) or massaging the dataset — modify the labels (change the labels appropriately to remove discrimination from the training data). However, the reweighing technique is more effective than the other two mentioned earlier.
```
RW = Reweighing(unprivileged_groups=unprivileged_groups,
privileged_groups=privileged_groups)
dataset_transf_train = RW.fit_transform(dataset_orig_train)
dataset_transf_train.instance_weights
len(dataset_transf_train.instance_weights)
```
## Step 5 Compute fairness metric on transformed dataset
Now that we have a transformed dataset, we can check how effective it was in removing bias by using the same metric we used for the original training dataset in Step 3. Once again, we use the function mean_difference in the BinaryLabelDatasetMetric class. We see the mitigation step was very effective, the difference in mean outcomes is now 0.0. So we went from a 17% advantage for the privileged group to equality in terms of mean outcome.
```
metric_transf_train = BinaryLabelDatasetMetric(dataset_transf_train,
unprivileged_groups=unprivileged_groups,
privileged_groups=privileged_groups)
print("Transformed training dataset")
print("Difference in mean outcomes between unprivileged and privileged groups = %f" % metric_transf_train.mean_difference())
print("Transformed training dataset")
print("Disparate Impact = %f" % metric_transf_train.disparate_impact())
```
## Step 6: Try another algorithm!
There are numerous other pre-processing mitigation algorithms implemented in aif360, described at the following link:
https://aif360.readthedocs.io/en/latest/modules/preprocessing.html#
Take a minute to read about these options, then repeat Steps 4 and 5 above using a different algorithm. How do the fairness metrics compare? What could explain your similar/different results?
```
# Set up the pre-processing mitigation algorithm
# Fit and transform the data
# Compute fairness metrics using your transformed data
```
### Summary
The purpose of this tutorial is to give a new user to bias detection and mitigation a gentle introduction to some of the functionality of AI Fairness 360. A more complete use case would take the next step and see how the transformed dataset impacts the accuracy and fairness of a trained model. This is implemented in the demo notebook in the examples directory of toolkit, called demo_reweighing_preproc.ipynb. I highly encourage readers to view that notebook as it is generalization and extension of this simple tutorial.
There are many metrics one can use to detect the presence of bias. AI Fairness 360 provides many of them for your use. Since it is not clear which of these metrics to use, we also provide some guidance. Likewise, there are many different bias mitigation algorithms one can employ, many of which are in AI Fairness 360. Other tutorials will demonstrate the use of some of these metrics and mitigations algorithms.
As mentioned earlier, both fairness metrics and mitigation algorithms can be performed at various stages of the machine learning pipeline. We recommend checking for bias as often as possible, using as many metrics are relevant for the application domain. We also recommend incorporating bias detection in an automated continouus integration pipeline to ensure bias awareness as a software project evolves.
| github_jupyter |
```
import pandas as pd
# Read .csv file of total schools and grades from Florida Department of Education
file = "./SchoolGrades19_clean.csv"
school_grades = pd.read_csv(file)
# Read file with school zip code info
file2 = "./hills_schools_zip.csv"
school_zip = pd.read_csv(file2)
pd.set_option('display.max_columns', None)
# Drop all but Hillsborough County schools
index_names1 = school_grades[(school_grades['District Name'] != "HILLSBOROUGH")].index
school_grades_hill = school_grades.drop(index_names1, inplace = False)
school_grades_hill
school_grades_hill.count()
# Create stripped-down dataframe that only contains grades as far back as 2015
# Also removes: collocated rule columns because they were empty
# and miscellaneous other columns:
school_grades_basic = school_grades_hill.loc[:, ["District Number", "District Name", "School Number", "School Name",
"Total Points Earned", "Total Components", "Percent of Total Possible Points",
"Percent Tested", "Grade 2019", "Grade 2018", "Grade 2017", "Grade 2016",
"Informational Baseline Grade 2015", "Charter School", "Title I",
"Alternative/ESE Center School", "School Type", "Percent of Minority Students",
"Percent of Economically Disadvantaged Students"]]
# Drop schools with missing grade data from 2015
indexNames2 = school_grades_basic[school_grades_basic['Informational Baseline Grade 2015'] == '0'].index
# Delete these row indices from dataFrame
school_grades_basic = school_grades_basic.drop(indexNames2)
school_grades_basic.head()
# Create dataframe with only elementary schools
indexNames3 = school_grades_basic[school_grades_basic['School Type'] != 1].index
school_grades_elem = school_grades_basic.drop(indexNames3)
school_grades_elem.head()
# Create dataframe without charter schools
indexNames4 = school_grades_elem[school_grades_elem['Charter School'] == 'YES'].index
# Delete these row indices from dataFrame
school_grades_elem_xc = school_grades_elem.drop(indexNames4)
# Add column for zip code
school_grades_elem_xc['ZIP']= ""
school_grades_elem_xc
# Convert numbers in school_zip so they match number formats in school_grades_elem_xc
school_zip.astype({'School Number': 'int64'}, {'School ZIP': 'int64'}).dtypes
for i, row in school_zip.iterrows():
school_grades_elem_xc.loc[school_grades_elem_xc['School Number'] == row['School Number'], "ZIP"] = row['School ZIP']
# school_grades_elem_xc.astype({'ZIP': 'int64'}).dtypes
school_grades_elem_xc.head()
# school_grades_final = school_grades_elem_xc.sort_values(by=['ZIP'])
indexNames5 = school_grades_elem_xc[school_grades_elem_xc['ZIP'] == ''].index
# Delete these row indices from dataFrame
school_grades_final = school_grades_elem_xc.drop(indexNames5)
# school_grades_elem_xc.astype({'ZIP': 'int64'}).dtypes
school_grades_final.count()
school_grades_final.to_csv('school_grades_final.csv')
school_grades_final.count()
```
| github_jupyter |
# Lab 05 : Final code -- demo
```
# For Google Colaboratory
import sys, os
if 'google.colab' in sys.modules:
# mount google drive
from google.colab import drive
drive.mount('/content/gdrive')
# find automatically the path of the folder containing "file_name" :
file_name = 'final_demo.ipynb'
import subprocess
path_to_file = subprocess.check_output('find . -type f -name ' + str(file_name), shell=True).decode("utf-8")
path_to_file = path_to_file.replace(file_name,"").replace('\n',"")
# if previous search failed or too long, comment the previous line and simply write down manually the path below :
#path_to_file = '/content/gdrive/My Drive/CS5242_2021_codes/codes/labs_lecture05/lab05_final'
print(path_to_file)
# change current path to the folder containing "file_name"
os.chdir(path_to_file)
!pwd
import torch
import torch.nn as nn
import torch.optim as optim
from random import randint
import time
import utils
```
### Download the data
```
from utils import check_mnist_dataset_exists
data_path=check_mnist_dataset_exists()
train_data=torch.load(data_path+'mnist/train_data.pt')
train_label=torch.load(data_path+'mnist/train_label.pt')
test_data=torch.load(data_path+'mnist/test_data.pt')
test_label=torch.load(data_path+'mnist/test_label.pt')
```
### Make a one layer net class.
```
class one_layer_net(nn.Module):
def __init__(self, input_size, output_size):
super(one_layer_net , self).__init__()
self.linear_layer = nn.Linear( input_size, output_size , bias=False)
def forward(self, x):
scores = self.linear_layer(x)
return scores
```
### Build the net
```
net=one_layer_net(784,10)
print(net)
utils.display_num_param(net)
```
### Choose the criterion, batchsize
```
criterion = nn.CrossEntropyLoss()
bs=200
```
### Evaluate on test set
```
def eval_on_test_set():
running_error=0
num_batches=0
for i in range(0,10000,bs):
minibatch_data = test_data[i:i+bs]
minibatch_label= test_label[i:i+bs]
inputs = minibatch_data.view(bs,784)
scores=net( inputs )
error = utils.get_error( scores , minibatch_label)
running_error += error.item()
num_batches+=1
total_error = running_error/num_batches
print( 'test error = ', total_error*100 ,'percent')
```
### Training loop
```
start = time.time()
lr = 0.05 # initial learning rate
for epoch in range(200):
# learning rate strategy : divide the learning rate by 1.5 every 10 epochs
if epoch%10==0 and epoch>10:
lr = lr / 1.5
# create a new optimizer at the beginning of each epoch: give the current learning rate.
optimizer=torch.optim.SGD( net.parameters() , lr=lr )
running_loss=0
running_error=0
num_batches=0
shuffled_indices=torch.randperm(60000)
for count in range(0,60000,bs):
# forward and backward pass
optimizer.zero_grad()
indices=shuffled_indices[count:count+bs]
minibatch_data = train_data[indices]
minibatch_label= train_label[indices]
inputs = minibatch_data.view(bs,784)
inputs.requires_grad_()
scores=net( inputs )
loss = criterion( scores , minibatch_label)
loss.backward()
optimizer.step()
# compute some stats
running_loss += loss.detach().item()
error = utils.get_error( scores.detach() , minibatch_label)
running_error += error.item()
num_batches+=1
# once the epoch is finished we divide the "running quantities"
# by the number of batches
total_loss = running_loss/num_batches
total_error = running_error/num_batches
elapsed_time = time.time() - start
# every 10 epoch we display the stats
# and compute the error rate on the test set
if epoch % 10 == 0 :
print(' ')
print('epoch=',epoch, ' time=', elapsed_time,
' loss=', total_loss , ' error=', total_error*100 ,'percent lr=', lr)
eval_on_test_set()
```
### Choose image at random from the test set and see how good/bad are the predictions
```
# choose a picture at random
idx=randint(0, 10000-1)
im=test_data[idx]
# diplay the picture
utils.show(im)
# feed it to the net and display the confidence scores
scores = net( im.view(1,784))
probs= torch.softmax(scores, dim=1)
utils.show_prob_mnist(probs)
```
| github_jupyter |
# SVI Part III: ELBO Gradient Estimators
## Setup
We've defined a Pyro model with observations ${\bf x}$ and latents ${\bf z}$ of the form $p_{\theta}({\bf x}, {\bf z}) = p_{\theta}({\bf x}|{\bf z}) p_{\theta}({\bf z})$. We've also defined a Pyro guide (i.e. a variational distribution) of the form $q_{\phi}({\bf z})$. Here ${\theta}$ and $\phi$ are variational parameters for the model and guide, respectively. (In particular these are _not_ random variables that call for a Bayesian treatment).
We'd like to maximize the log evidence $\log p_{\theta}({\bf x})$ by maximizing the ELBO (the evidence lower bound) given by
$${\rm ELBO} \equiv \mathbb{E}_{q_{\phi}({\bf z})} \left [
\log p_{\theta}({\bf x}, {\bf z}) - \log q_{\phi}({\bf z})
\right]$$
To do this we're going to take (stochastic) gradient steps on the ELBO in the parameter space $\{ \theta, \phi \}$ (see references [1,2] for early work on this approach). So we need to be able to compute unbiased estimates of
$$\nabla_{\theta,\phi} {\rm ELBO} = \nabla_{\theta,\phi}\mathbb{E}_{q_{\phi}({\bf z})} \left [
\log p_{\theta}({\bf x}, {\bf z}) - \log q_{\phi}({\bf z})
\right]$$
How do we do this for general stochastic functions `model()` and `guide()`? To simplify notation let's generalize our discussion a bit and ask how we can compute gradients of expectations of an arbitrary cost function $f({\bf z})$. Let's also drop any distinction between $\theta$ and $\phi$. So we want to compute
$$\nabla_{\phi}\mathbb{E}_{q_{\phi}({\bf z})} \left [
f_{\phi}({\bf z}) \right]$$
Let's start with the easiest case.
## Easy Case: Reparameterizable Random Variables
Suppose that we can reparameterize things such that
$$\mathbb{E}_{q_{\phi}({\bf z})} \left [f_{\phi}({\bf z}) \right]
=\mathbb{E}_{q({\bf \epsilon})} \left [f_{\phi}(g_{\phi}({\bf \epsilon})) \right]$$
Crucially we've moved all the $\phi$ dependence inside of the expectation; $q({\bf \epsilon})$ is a fixed distribution with no dependence on $\phi$. This kind of reparameterization can be done for many distributions (e.g. the normal distribution); see reference [3] for a discussion. In this case we can pass the gradient straight through the expectation to get
$$\nabla_{\phi}\mathbb{E}_{q({\bf \epsilon})} \left [f_{\phi}(g_{\phi}({\bf \epsilon})) \right]=
\mathbb{E}_{q({\bf \epsilon})} \left [\nabla_{\phi}f_{\phi}(g_{\phi}({\bf \epsilon})) \right]$$
Assuming $f(\cdot)$ and $g(\cdot)$ are sufficiently smooth, we can now get unbiased estimates of the gradient of interest by taking a Monte Carlo estimate of this expectation.
## Tricky Case: Non-reparameterizable Random Variables
What if we can't do the above reparameterization? Unfortunately this is the case for many distributions of interest, for example all discrete distributions. In this case our estimator takes a bit more complicated form.
We begin by expanding the gradient of interest as
$$\nabla_{\phi}\mathbb{E}_{q_{\phi}({\bf z})} \left [
f_{\phi}({\bf z}) \right]=
\nabla_{\phi} \int d{\bf z} \; q_{\phi}({\bf z}) f_{\phi}({\bf z})$$
and use the chain rule to write this as
$$ \int d{\bf z} \; \left \{ (\nabla_{\phi} q_{\phi}({\bf z})) f_{\phi}({\bf z}) + q_{\phi}({\bf z})(\nabla_{\phi} f_{\phi}({\bf z}))\right \} $$
At this point we run into a problem. We know how to generate samples from $q(\cdot)$—we just run the guide forward—but $\nabla_{\phi} q_{\phi}({\bf z})$ isn't even a valid probability density. So we need to massage this formula so that it's in the form of an expectation w.r.t. $q(\cdot)$. This is easily done using the identity
$$ \nabla_{\phi} q_{\phi}({\bf z}) =
q_{\phi}({\bf z})\nabla_{\phi} \log q_{\phi}({\bf z})$$
which allows us to rewrite the gradient of interest as
$$\mathbb{E}_{q_{\phi}({\bf z})} \left [
(\nabla_{\phi} \log q_{\phi}({\bf z})) f_{\phi}({\bf z}) + \nabla_{\phi} f_{\phi}({\bf z})\right]$$
This form of the gradient estimator—variously known as the REINFORCE estimator or the score function estimator or the likelihood ratio estimator—is amenable to simple Monte Carlo estimation.
Note that one way to package this result (which is convenient for implementation) is to introduce a surrogate objective function
$${\rm surrogate \;objective} \equiv
\log q_{\phi}({\bf z}) \overline{f_{\phi}({\bf z})} + f_{\phi}({\bf z})$$
Here the bar indicates that the term is held constant (i.e. it is not to be differentiated w.r.t. $\phi$). To get a (single-sample) Monte Carlo gradient estimate, we sample the latent random variables, compute the surrogate objective, and differentiate. The result is an unbiased estimate of $\nabla_{\phi}\mathbb{E}_{q_{\phi}({\bf z})} \left [
f_{\phi}({\bf z}) \right]$. In equations:
$$\nabla_{\phi} {\rm ELBO} = \mathbb{E}_{q_{\phi}({\bf z})} \left [
\nabla_{\phi} ({\rm surrogate \; objective}) \right]$$
## Variance or Why I Wish I Was Doing MLE Deep Learning
We now have a general recipe for an unbiased gradient estimator of expectations of cost functions. Unfortunately, in the more general case where our $q(\cdot)$ includes non-reparameterizable random variables, this estimator tends to have high variance. Indeed in many cases of interest the variance is so high that the estimator is effectively unusable. So we need strategies to reduce variance (for a discussion see reference [4]). We're going to pursue two strategies. The first strategy takes advantage of the particular structure of the cost function $f(\cdot)$. The second strategy effectively introduces a way to reduce variance by using information from previous estimates of
$\mathbb{E}_{q_{\phi}({\bf z})} [ f_{\phi}({\bf z})]$. As such it is somewhat analogous to using momentum in stochastic gradient descent.
### Reducing Variance via Dependency Structure
In the above discussion we stuck to a general cost function $f_{\phi}({\bf z})$. We could continue in this vein (the approach we're about to discuss is applicable in the general case) but for concreteness let's zoom back in. In the case of stochastic variational inference, we're interested in a particular cost function of the form <br/><br/>
$$\log p_{\theta}({\bf x} | {\rm Pa}_p ({\bf x})) +
\sum_i \log p_{\theta}({\bf z}_i | {\rm Pa}_p ({\bf z}_i))
- \sum_i \log q_{\phi}({\bf z}_i | {\rm Pa}_q ({\bf z}_i))$$
where we've broken the log ratio $\log p_{\theta}({\bf x}, {\bf z})/q_{\phi}({\bf z})$ into an observation log likelihood piece and a sum over the different latent random variables $\{{\bf z}_i \}$. We've also introduced the notation
${\rm Pa}_p (\cdot)$ and ${\rm Pa}_q (\cdot)$ to denote the parents of a given random variable in the model and in the guide, respectively. (The reader might worry what the appropriate notion of dependency would be in the case of general stochastic functions; here we simply mean regular ol' dependency within a single execution trace). The point is that different terms in the cost function have different dependencies on the random variables $\{ {\bf z}_i \}$ and this is something we can leverage.
To make a long story short, for any non-reparameterizable latent random variable ${\bf z}_i$ the surrogate objective is going to have a term
$$\log q_{\phi}({\bf z}_i) \overline{f_{\phi}({\bf z})} $$
It turns out that we can remove some of the terms in $\overline{f_{\phi}({\bf z})}$ and still get an unbiased gradient estimator; furthermore, doing so will generally decrease the variance. In particular (see reference [4] for details) we can remove any terms in $\overline{f_{\phi}({\bf z})}$ that are not downstream of the latent variable ${\bf z}_i$ (downstream w.r.t. to the dependency structure of the guide). Note that this general trick—where certain random variables are dealt with analytically to reduce variance—often goes under the name of Rao-Blackwellization.
In Pyro, all of this logic is taken care of automatically by the `SVI` class. In particular as long as we use a `TraceGraph_ELBO` loss, Pyro will keep track of the dependency structure within the execution traces of the model and guide and construct a surrogate objective that has all the unnecessary terms removed:
```python
svi = SVI(model, guide, optimizer, TraceGraph_ELBO())
```
Note that leveraging this dependency information takes extra computations, so `TraceGraph_ELBO` should only be used in the case where your model has non-reparameterizable random variables; in most applications `Trace_ELBO` suffices.
### An Example with Rao-Blackwellization:
Suppose we have a gaussian mixture model with $K$ components. For each data point we: (i) first sample the component distribution $k \in [1,...,K]$; and (ii) observe the data point using the $k^{\rm th}$ component distribution. The simplest way to write down a model of this sort is as follows:
```python
ks = pyro.sample("k", dist.Categorical(probs)
.to_event(1))
pyro.sample("obs", dist.Normal(locs[ks], scale)
.to_event(1),
obs=data)
```
Since the user hasn't taken care to mark any of the conditional independencies in the model, the gradient estimator constructed by Pyro's `SVI` class is unable to take advantage of Rao-Blackwellization, with the result that the gradient estimator will tend to suffer from high variance. To address this problem the user needs to explicitly mark the conditional independence. Happily, this is not much work:
```python
# mark conditional independence
# (assumed to be along the rightmost tensor dimension)
with pyro.plate("foo", data.size(-1)):
ks = pyro.sample("k", dist.Categorical(probs))
pyro.sample("obs", dist.Normal(locs[ks], scale),
obs=data)
```
That's all there is to it.
### Aside: Dependency tracking in Pyro
Finally, a word about dependency tracking. Tracking dependency within a stochastic function that includes arbitrary Python code is a bit tricky. The approach currently implemented in Pyro is analogous to the one used in WebPPL (cf. reference [5]). Briefly, a conservative notion of dependency is used that relies on sequential ordering. If random variable ${\bf z}_2$ follows ${\bf z}_1$ in a given stochastic function then ${\bf z}_2$ _may be_ dependent on ${\bf z}_1$ and therefore _is_ assumed to be dependent. To mitigate the overly coarse conclusions that can be drawn by this kind of dependency tracking, Pyro includes constructs for declaring things as independent, namely `plate` and `markov` ([see the previous tutorial](svi_part_ii.ipynb)). For use cases with non-reparameterizable variables, it is therefore important for the user to make use of these constructs (when applicable) to take full advantage of the variance reduction provided by `SVI`. In some cases it may also pay to consider reordering random variables within a stochastic function (if possible).
### Reducing Variance with Data-Dependent Baselines
The second strategy for reducing variance in our ELBO gradient estimator goes under the name of baselines (see e.g. reference [6]). It actually makes use of the same bit of math that underlies the variance reduction strategy discussed above, except now instead of removing terms we're going to add terms. Basically, instead of removing terms with zero expectation that tend to _contribute_ to the variance, we're going to add specially chosen terms with zero expectation that work to _reduce_ the variance. As such, this is a control variate strategy.
In more detail, the idea is to take advantage of the fact that for any constant $b$, the following identity holds
$$\mathbb{E}_{q_{\phi}({\bf z})} \left [\nabla_{\phi}
(\log q_{\phi}({\bf z}) \times b) \right]=0$$
This follows since $q(\cdot)$ is normalized:
$$\mathbb{E}_{q_{\phi}({\bf z})} \left [\nabla_{\phi}
\log q_{\phi}({\bf z}) \right]=
\int \!d{\bf z} \; q_{\phi}({\bf z}) \nabla_{\phi}
\log q_{\phi}({\bf z})=
\int \! d{\bf z} \; \nabla_{\phi} q_{\phi}({\bf z})=
\nabla_{\phi} \int \! d{\bf z} \; q_{\phi}({\bf z})=\nabla_{\phi} 1 = 0$$
What this means is that we can replace any term
$$\log q_{\phi}({\bf z}_i) \overline{f_{\phi}({\bf z})} $$
in our surrogate objective with
$$\log q_{\phi}({\bf z}_i) \left(\overline{f_{\phi}({\bf z})}-b\right) $$
Doing so doesn't affect the mean of our gradient estimator but it does affect the variance. If we choose $b$ wisely, we can hope to reduce the variance. In fact, $b$ need not be a constant: it can depend on any of the random choices upstream (or sidestream) of ${\bf z}_i$.
#### Baselines in Pyro
There are several ways the user can instruct Pyro to use baselines in the context of stochastic variational inference. Since baselines can be attached to any non-reparameterizable random variable, the current baseline interface is at the level of the `pyro.sample` statement. In particular the baseline interface makes use of an argument `baseline`, which is a dictionary that specifies baseline options. Note that it only makes sense to specify baselines for sample statements within the guide (and not in the model).
##### Decaying Average Baseline
The simplest baseline is constructed from a running average of recent samples of $\overline{f_{\phi}({\bf z})}$. In Pyro this kind of baseline can be invoked as follows
```python
z = pyro.sample("z", dist.Bernoulli(...),
infer=dict(baseline={'use_decaying_avg_baseline': True,
'baseline_beta': 0.95}))
```
The optional argument `baseline_beta` specifies the decay rate of the decaying average (default value: `0.90`).
#### Neural Baselines
In some cases a decaying average baseline works well. In others using a baseline that depends on upstream randomness is crucial for getting good variance reduction. A powerful approach for constructing such a baseline is to use a neural network that can be adapted during the course of learning. Pyro provides two ways to specify such a baseline (for an extended example see the [AIR tutorial](air.ipynb)).
First the user needs to decide what inputs the baseline is going to consume (e.g. the current datapoint under consideration or the previously sampled random variable). Then the user needs to construct a `nn.Module` that encapsulates the baseline computation. This might look something like
```python
class BaselineNN(nn.Module):
def __init__(self, dim_input, dim_hidden):
super().__init__()
self.linear = nn.Linear(dim_input, dim_hidden)
# ... finish initialization ...
def forward(self, x):
hidden = self.linear(x)
# ... do more computations ...
return baseline
```
Then, assuming the BaselineNN object `baseline_module` has been initialized somewhere else, in the guide we'll have something like
```python
def guide(x): # here x is the current mini-batch of data
pyro.module("my_baseline", baseline_module)
# ... other computations ...
z = pyro.sample("z", dist.Bernoulli(...),
infer=dict(baseline={'nn_baseline': baseline_module,
'nn_baseline_input': x}))
```
Here the argument `nn_baseline` tells Pyro which `nn.Module` to use to construct the baseline. On the backend the argument `nn_baseline_input` is fed into the forward method of the module to compute the baseline $b$. Note that the baseline module needs to be registered with Pyro with a `pyro.module` call so that Pyro is aware of the trainable parameters within the module.
Under the hood Pyro constructs a loss of the form
$${\rm baseline\; loss} \equiv\left(\overline{f_{\phi}({\bf z})} - b \right)^2$$
which is used to adapt the parameters of the neural network. There's no theorem that suggests this is the optimal loss function to use in this context (it's not), but in practice it can work pretty well. Just as for the decaying average baseline, the idea is that a baseline that can track the mean $\overline{f_{\phi}({\bf z})}$ will help reduce the variance. Under the hood `SVI` takes one step on the baseline loss in conjunction with a step on the ELBO.
Note that in practice it can be important to use a different set of learning hyperparameters (e.g. a higher learning rate) for baseline parameters. In Pyro this can be done as follows:
```python
def per_param_args(module_name, param_name):
if 'baseline' in param_name or 'baseline' in module_name:
return {"lr": 0.010}
else:
return {"lr": 0.001}
optimizer = optim.Adam(per_param_args)
```
Note that in order for the overall procedure to be correct the baseline parameters should only be optimized through the baseline loss. Similarly the model and guide parameters should only be optimized through the ELBO. To ensure that this is the case under the hood `SVI` detaches the baseline $b$ that enters the ELBO from the autograd graph. Also, since the inputs to the neural baseline may depend on the parameters of the model and guide, the inputs are also detached from the autograd graph before they are fed into the neural network.
Finally, there is an alternate way for the user to specify a neural baseline. Simply use the argument `baseline_value`:
```python
b = # do baseline computation
z = pyro.sample("z", dist.Bernoulli(...),
infer=dict(baseline={'baseline_value': b}))
```
This works as above, except in this case it's the user's responsibility to make sure that any autograd tape connecting $b$ to the parameters of the model and guide has been cut. Or to say the same thing in language more familiar to PyTorch users, any inputs to $b$ that depend on $\theta$ or $\phi$ need to be detached from the autograd graph with `detach()` statements.
#### A complete example with baselines
Recall that in the [first SVI tutorial](svi_part_i.ipynb) we considered a bernoulli-beta model for coin flips. Because the beta random variable is non-reparameterizable (or rather not easily reparameterizable), the corresponding ELBO gradients can be quite noisy. In that context we dealt with this problem by using a Beta distribution that provides (approximate) reparameterized gradients. Here we showcase how a simple decaying average baseline can reduce the variance in the case where the Beta distribution is treated as non-reparameterized (so that the ELBO gradient estimator is of the score function type). While we're at it, we also use `plate` to write our model in a fully vectorized manner.
Instead of directly comparing gradient variances, we're going to see how many steps it takes for SVI to converge. Recall that for this particular model (because of conjugacy) we can compute the exact posterior. So to assess the utility of baselines in this context, we setup the following simple experiment. We initialize the guide at a specified set of variational parameters. We then do SVI until the variational parameters have gotten to within a fixed tolerance of the parameters of the exact posterior. We do this both with and without the decaying average baseline. We then compare the number of gradient steps we needed in the two cases. Here's the complete code:
(_Since apart from the use of_ `plate` _and_ `use_decaying_avg_baseline`, _this code is very similar to the code in parts I and II of the SVI tutorial, we're not going to go through the code line by line._)
```
import os
import torch
import torch.distributions.constraints as constraints
import pyro
import pyro.distributions as dist
# Pyro also has a reparameterized Beta distribution so we import
# the non-reparameterized version to make our point
from pyro.distributions.testing.fakes import NonreparameterizedBeta
import pyro.optim as optim
from pyro.infer import SVI, TraceGraph_ELBO
import sys
# enable validation (e.g. validate parameters of distributions)
assert pyro.__version__.startswith('1.4.0')
pyro.enable_validation(True)
# this is for running the notebook in our testing framework
smoke_test = ('CI' in os.environ)
max_steps = 2 if smoke_test else 10000
def param_abs_error(name, target):
return torch.sum(torch.abs(target - pyro.param(name))).item()
class BernoulliBetaExample:
def __init__(self, max_steps):
# the maximum number of inference steps we do
self.max_steps = max_steps
# the two hyperparameters for the beta prior
self.alpha0 = 10.0
self.beta0 = 10.0
# the dataset consists of six 1s and four 0s
self.data = torch.zeros(10)
self.data[0:6] = torch.ones(6)
self.n_data = self.data.size(0)
# compute the alpha parameter of the exact beta posterior
self.alpha_n = self.data.sum() + self.alpha0
# compute the beta parameter of the exact beta posterior
self.beta_n = - self.data.sum() + torch.tensor(self.beta0 + self.n_data)
# initial values of the two variational parameters
self.alpha_q_0 = 15.0
self.beta_q_0 = 15.0
def model(self, use_decaying_avg_baseline):
# sample `latent_fairness` from the beta prior
f = pyro.sample("latent_fairness", dist.Beta(self.alpha0, self.beta0))
# use plate to indicate that the observations are
# conditionally independent given f and get vectorization
with pyro.plate("data_plate"):
# observe all ten datapoints using the bernoulli likelihood
pyro.sample("obs", dist.Bernoulli(f), obs=self.data)
def guide(self, use_decaying_avg_baseline):
# register the two variational parameters with pyro
alpha_q = pyro.param("alpha_q", torch.tensor(self.alpha_q_0),
constraint=constraints.positive)
beta_q = pyro.param("beta_q", torch.tensor(self.beta_q_0),
constraint=constraints.positive)
# sample f from the beta variational distribution
baseline_dict = {'use_decaying_avg_baseline': use_decaying_avg_baseline,
'baseline_beta': 0.90}
# note that the baseline_dict specifies whether we're using
# decaying average baselines or not
pyro.sample("latent_fairness", NonreparameterizedBeta(alpha_q, beta_q),
infer=dict(baseline=baseline_dict))
def do_inference(self, use_decaying_avg_baseline, tolerance=0.80):
# clear the param store in case we're in a REPL
pyro.clear_param_store()
# setup the optimizer and the inference algorithm
optimizer = optim.Adam({"lr": .0005, "betas": (0.93, 0.999)})
svi = SVI(self.model, self.guide, optimizer, loss=TraceGraph_ELBO())
print("Doing inference with use_decaying_avg_baseline=%s" % use_decaying_avg_baseline)
# do up to this many steps of inference
for k in range(self.max_steps):
svi.step(use_decaying_avg_baseline)
if k % 100 == 0:
print('.', end='')
sys.stdout.flush()
# compute the distance to the parameters of the true posterior
alpha_error = param_abs_error("alpha_q", self.alpha_n)
beta_error = param_abs_error("beta_q", self.beta_n)
# stop inference early if we're close to the true posterior
if alpha_error < tolerance and beta_error < tolerance:
break
print("\nDid %d steps of inference." % k)
print(("Final absolute errors for the two variational parameters " +
"were %.4f & %.4f") % (alpha_error, beta_error))
# do the experiment
bbe = BernoulliBetaExample(max_steps=max_steps)
bbe.do_inference(use_decaying_avg_baseline=True)
bbe.do_inference(use_decaying_avg_baseline=False)
```
**Sample output:**
```
Doing inference with use_decaying_avg_baseline=True
....................
Did 1932 steps of inference.
Final absolute errors for the two variational parameters were 0.7997 & 0.0800
Doing inference with use_decaying_avg_baseline=False
..................................................
Did 4908 steps of inference.
Final absolute errors for the two variational parameters were 0.7991 & 0.2532
```
For this particular run we can see that baselines roughly halved the number of steps of SVI we needed to do. The results are stochastic and will vary from run to run, but this is an encouraging result. This is a pretty contrived example, but for certain model and guide pairs, baselines can provide a substantial win.
## References
[1] `Automated Variational Inference in Probabilistic Programming`,
<br/>
David Wingate, Theo Weber
[2] `Black Box Variational Inference`,<br/>
Rajesh Ranganath, Sean Gerrish, David M. Blei
[3] `Auto-Encoding Variational Bayes`,<br/>
Diederik P Kingma, Max Welling
[4] `Gradient Estimation Using Stochastic Computation Graphs`,
<br/>
John Schulman, Nicolas Heess, Theophane Weber, Pieter Abbeel
[5] `Deep Amortized Inference for Probabilistic Programs`
<br/>
Daniel Ritchie, Paul Horsfall, Noah D. Goodman
[6] `Neural Variational Inference and Learning in Belief Networks`
<br/>
Andriy Mnih, Karol Gregor
| github_jupyter |
# Scientific Computing with Python (Second Edition)
# Chapter 12
This notebook file requires Jupyter notebook version >= 5 as it uses cell tagging to avoid that execution stops when intentionally exceptions are raises in a a cell.
*We start by importing all from Numpy. As explained in Chapter 01
the examples are written assuming this import is initially done.*
```
from numpy import *
def f(x):
return 1/x
f(2.5)
f(0)
a = arange(8.0)
a
a[3] = 'string'
```
### 12.1.1 Basic principles
```
raise Exception("Something went wrong")
print("The algorithm did not converge.")
raise Exception("The algorithm did not converge.")
def factorial(n):
if not (isinstance(n, (int, int32, int64))):
raise TypeError("An integer is expected")
if not (n >=0):
raise ValueError("A positive number is expected")
...
print(factorial(5.2))
print(factorial(-5))
try:
f = open('data.txt', 'r')
data = f.readline()
value = float(data)
except FileNotFoundError as FnF:
print(f' {FnF.strerror}: {FnF.filename}')
except ValueError:
print("Could not convert data to float.")
```
### 12.1.2 User-defined exceptions
```
class MyError(Exception):
def __init__(self, expr):
self.expr = expr
def __str__(self):
return str(self.expr)
try:
x = random.rand()
if x < 0.5:
raise MyError(x)
except MyError as e:
print("Random number too small", e.expr)
else:
print(x)
```
### 12.1.3 Context managers – the with statement
```
with open('data.txt', 'w') as f:
process_file_data(f)
f = open('data.txt', 'w')
try:
# some function that does something with the file
process_file_data(f)
except:
...
finally:
f.close()
import numpy as np # note, sqrt in NumPy and SciPy
# behave differently in that example
with errstate(invalid='ignore'):
print(np.sqrt(-1)) # prints 'nan'
with errstate(invalid='warn'):
print(np.sqrt(-1)) # prints 'nan' and
# 'RuntimeWarning: invalid value encountered in sqrt'
with errstate(invalid='raise'):
print(np.sqrt(-1)) # prints nothing and raises FloatingPointError
```
## 12.2 Finding errors: debugging
### 12.2.1 Bugs
no code
### 12.2.2 The stack
```
def f():
g()
def g():
h()
def h():
1//0
f()
def f(a):
g(a)
def g(a):
h(a)
def h(a):
raise Exception(f'An exception just to provoke a strack trace and a value a={a}')
f(23)
```
### 12.2.3 The Python debugger
We do not present code here as the section requires interactive user actions.
### 12.2.4 Overview – debug commands
No code in this section.
### 12.2.5 Debugging in IPython
We do not present code here as the section requires interactive user actions.
| github_jupyter |
# Rough draft of getting _vcf_ files
### Overview
Different tertiary analysis software often works with _vcf_ or _gene expression_ files. Depending on the API they use, you may want to _point to the actual file_ **or** accept _an http download link_. We are going to show both approaches here, but adding a download step is trivial<sup>1</sup>.
There are **two** approaches we will show to get **vcf download links**. This may be something you would like to do locally or as a _push-button_ from within a tertiary analysis GUI.
* Specify a **project** and grab **all** vcf files within that project (a.k.a. the _lazy way_)
* Specify both a **task** and **project** and look for _outputs_ of that task which are vcf format.
Obvious extensions would be to keep track of previously imported file names, or only take tasks more recent than a certain date.
### Prerequisites
1. You need your _authentication token_ and the API needs to know about it. See <a href="Setup_API_environment.ipynb">**Setup_API_environment.ipynb**</a> for details.
2. You understand how to <a href="../../Recipes/SBPLAT/projects_listAll.ipynb" target="_blank">list</a> **projects** you are a member of (we will just use that call directly and pick one here).
3. You understand how to <a href="../../Recipes/SBPLAT/files_listAll.ipynb" target="_blank">list</a> **files** within one of your projects.
4. You understand how to <a href="../../Recipes/SBPLAT/tasks_monitorAndGetResults.ipynb" target="_blank">deal with</a> **tasks** within one of your projects.
<sup>1</sup> the **.download()** method for a _file object_ will do this directly. You are welcome to use your favorite flavor of downloader.
## Imports
We import the _Api_ class from the official sevenbridges-python bindings below.
```
import sevenbridges as sbg
```
## Initialize the object
The _Api_ object needs to know your **auth\_token** and the correct path. Here we assume you are using the .sbgrc file in your home directory. For other options see <a href="Setup_API_environment.ipynb">Setup_API_environment.ipynb</a>
```
# [USER INPUT] specify platform {cgc, sbg}
prof = 'cgc'
config_file = sbg.Config(profile=prof)
api = sbg.Api(config=config_file)
```
# Approach 1: Get all files from a project
This is a more _brute-force_ approach to get all files in a project. This does not discriminate between _reference_ or _output_ vcf files. **Approach 1** consists of three steps:
* Find my project
* Find all files in my project
* Get download links
## Find my project
A **list**-call for projects returns the following *attributes*:
* **id** _Unique_ identifier for the project, generated based on Project Name
* **name** Name of project specified by the user, maybe _non-unique_
* **href** Address<sup>2</sup> of the project.\n"
All list API calls will feature pagination, by _default_ 50 items will be returned. We will also show how to specify a different limit and page forward and backwards. If you want to deal with pagination, look here: <a href="../../Recipes/SBPLAT/files_listAll.ipynb" target="_blank">list</a>. Otherwise use the **.all()** method to avoid it.
<sup>2</sup> This is the address where, by using API you can get this resource
```
# [USER INPUT] Set project name:
project_name = 'WES project'
# List all projects
my_project = [p for p in api.projects.query(limit=100).all()
if p.name == project_name][0]
if not my_project: # exploit fact that empty list is False, {list, tuple, etc} is True
print('Project {} was not found, please check spelling (especially trailing spaces)'
.format(project_name))
raise KeyboardInterrupt
```
## Find all files in my project
A **list**-call for files returns the following *attributes*:
* **id** _Unique_ identifier for each file
* **name** Name of file, maybe _non-unique_
* **href** Address<sup>3</sup> of the file.
<sup>3</sup> This *address* is for the API, but will not work in a browser.
```
# [USER INPUT] Set the input file extension we are looking for
input_ext = 'vcf'
# LIST all files in the source and target project
my_files = [f for f in api.files.query(limit = 100, project = my_project.id)
if f.name[-len(input_ext):] == input_ext]
print('Project {} has {} matching files:'
.format(my_project.name, len(my_files)))
for f in my_files:
print(' ' + f.name)
```
## Get download links
Files objects have two methods for downloading:
* **.download_info()**
* **.download()**
Here we use the first method to build and print a list of download links to pass to a tertiary analysis provider.
```
# Make download list
list_of_urls = []
for f in my_files:
list_of_urls.append(f.download_info())
for url in list_of_urls:
print(url)
```
# Approach 2: Get all files from a particular task in a project
This is a _targeted_ approach to get _specific_ output files from a _particular_ task. **Approach 2** consists of four steps:
* Find my project
* Find a **particular** task
* Find all files in my project
* Get download links
## Find my project
Same code as in Approach 1 for finding a particular project.
```
# [USER INPUT] Set project name:
project_name = 'WES project'
# List all projects
my_project = [p for p in api.projects.query(limit=100).all()
if p.name == project_name][0]
if not my_project: # exploit fact that empty list is False, {list, tuple, etc} is True
print('Project {} was not found, please check spelling (especially trailing spaces)'
.format(project_name))
raise KeyboardInterrupt
```
## Find a particular task
Here we will first return all tasks within the project selected above, then search for a particular one.
#### NOTE
It would be **much cleaner** to work with tasks list (my_tasks) and a cutoff date.
```
# [USER INPUT] Set the task we are looking for
task_name = 'Whole Exome Sequencing GATK 2.3.9.-lite run - 03-25-16 18:08:25'
# LIST all tasks
my_tasks = api.tasks.query(limit = 100, project = my_project.id)
if not my_tasks: # exploit fact that empty list is False, {list, tuple, etc} is True
print("There are no tasks in project {}, cannot continue".format(my_project.name))
raise KeyboardInterrupt
else:
print("The project {} has {} tasks."
.format(my_project.name, my_tasks.total))
# this is NOT robust to users naming the task the same, but will take the first one
single_task = [t for t in my_tasks.all() if t.name == task_name]
if len(single_task) == 0:
print('No task exists with name {}, please check spelling'
.format(task_name))
elif len(single_task) > 1:
print('WARNING!! Multiple tasks have the same name, using the first one')
single_task = single_task[0]
else:
print('Task {} found.'.format(task_name))
```
## Find specific output files in the task
First use the **.outputs** attribute of the task to find particular files you want to download. Then query all files and check for matching ids.
```
# [USER INPUT] Specify which task outputs you like
outputs = ['raw_vcf',
'annotated']
# get the file ids assigned to those outputs
ids_to_get = []
for out in outputs:
ids_to_get.append(single_task[0].outputs[out].id)
# LIST files matching those ids
my_files = [f for f in api.files.query(limit = 100, project = my_project.id).all()
if f.id in ids_to_get]
my_file_names = [f.name for f in my_files]
print('There are {} files for your selected outputs:'
.format(len(my_file_names)))
for f in my_file_names:
print(' ' + f)
```
## Get download links
Files objects have two methods for downloading:
* **.download_info()**
* **.download()**
Here we also show the second method and download the first file to pass to a tertiary analysis provider.
```
# Make download list
list_of_urls = []
for f in my_files:
list_of_urls.append(f.download_info())
for url in list_of_urls:
print(url)
# Download first file to local directory
f = my_files[0]
f.download(path=f.name)
```
# Additional Information
Detailed documentation of this REST architectural style API is available [here](http://docs.cancergenomicscloud.org/docs/the-cgc-api). Details of particular API calls are in the linked _recipes_. The sevenbridges-python bindings are available on [github](https://github.com/sbg/sevenbridges-python) along with binding documentation [here](http://sevenbridges-python.readthedocs.io/en/latest/quickstart).
| github_jupyter |
####################################
# Our early approaches of producing test heat-events-dataset
####################################
```
from matplotlib import pyplot as plt
import matplotlib.dates as mdates
from matplotlib.patches import Rectangle
from datetime import datetime, timedelta
import numpy as np
import pandas as pd
import xarray as xr
import zarr
import fsspec
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
plt.rcParams['figure.figsize'] = 12,8
################################
# Read the dataset (only reads the metadata) - HAWAII
################################
# Ref:
# https://daac.ornl.gov/DAYMET/guides/Daymet_Daily_V4.html#datasetoverview
import pystac
import fsspec
import xarray as xr
account_name = "daymeteuwest"
container_name = "daymet-zarr"
collection = pystac.Collection.from_file(
"https://planetarycomputer.microsoft.com/api/stac/v1/collections/daymet-daily-hi"
)
asset = collection.assets["zarr-https"]
store = fsspec.get_mapper(asset.href)
ds = xr.open_zarr(store, **asset.extra_fields["xarray:open_kwargs"])
################################
# core algorithm to flag heat extremes
################################
# input: 1D tmax data
# output: same-size 1D flags where heat event == True
# algorithm coefs:
TEMP_DIFF_THRESHOLD = 1 # Celcius (or K)
PERSISTED_FOR_MIN = 3 # days
def flag_heat_events(arr_tmax1d: np.array, timestamps: np.array) -> np.array:
"""
# Same logic as in Notebook2, but this time it does not slice the
# xarray Dataset every time. Instead, it operates on the numpy array - faster.
"""
df = pd.DataFrame({'tmax':arr_tmax1d})
df['mov_avg'] = df.rolling(15, center=True).mean()
df['diff'] = df['tmax'] - df['mov_avg']
df['time'] = timestamps
df['hot'] = df['diff'] > TEMP_DIFF_THRESHOLD
df['label'] = df['hot'].diff().ne(False).cumsum()
df = df.reset_index().reset_index()
# filter
summer_months = [5,6,7,8,9]
df['isSummer'] = df['time'].dt.month.isin(summer_months)
dff = df[df['isSummer'] & df['hot']].dropna(subset=['diff'])
dfg = dff.groupby('label').agg({
'index':[np.min,np.max,len],
})
dfg.columns = ['i1','i2','count']
dfg = dfg[dfg['count'] >= PERSISTED_FOR_MIN]
dfg = dfg.drop('count', axis=1)
dfg = dfg.reset_index(drop=True)
arr = np.zeros(len(df), dtype=int)
for _, (i, j) in dfg.iterrows():
arr[i:j+1] = 1
return arr
################################
# Utils
################################
def get_xy_meshgrid(arr_tmax3d:np.ndarray) -> np.ndarray:
"""create grid for all x,y coordinate pairs [0,1],[0,2],..[283,583]"""
shape_yx = arr_tmax3d.shape[1:] # np.shape order -> zyx
arr_y = np.arange(shape_yx[0])
arr_x = np.arange(shape_yx[1])
ac = np.array(np.meshgrid(arr_x, arr_y)).T.reshape(-1, 2)
return ac
def print_stats(arr:np.ndarray) -> None:
size = round(arr_tmax3d.nbytes/1e9,2)
shp = arr_tmax3d.shape
print(f"""processing.. year={year}, shape z,y,x={shp}, in-memory={size} GB""")
%%time
import time
years = np.unique(ds['time'].dt.year).astype(str)
for year in years:
t1 = time.time()
# Read the entire tmax data into memory as np.ndarray
arr_tmax3d = ds['tmax'].sel(time=year).values
# Create same size empty 3d array to populate with heat event flags
arr_heat3d = np.zeros(arr_tmax3d.shape).astype(int)
print_stats(arr_heat3d)
# loop through all iX,iY pairs
meshgrid = get_xy_meshgrid(arr_tmax3d)
for i, j in meshgrid:
arr_tmax1d = arr_tmax3d[:,j,i]
no_data = np.isnan(arr_tmax1d).all()
if no_data:
arr_heat1d = np.zeros(arr_tmax1d.shape, dtype=int)
else:
timestamps = ds['tmax'].sel(time=year)['time'].values
arr_heat1d = flag_heat_events(arr_tmax1d, timestamps)
arr_heat3d[:,j,i] = arr_heat1d
np.save(f'./arr_heat3d/arr_heat3d-{year}.npy', arr_heat3d)
print(f'{round((time.time() - t1)/60, 2)}min')
```
| github_jupyter |
# Inferring prompt types
This notebook demos two transformers, which broadly aim at producing abstract representations of an utterance in terms of its phrasing and its rhetorical intent:
* The `PhrasingMotifs` transformer extracts representations of utterances in terms of how they are phrased;
* The `PromptTypes` transformer computes latent representations of utterances in terms of their rhetorical intention -- the _responses_ they aim at prompting -- and assigns utterances to different (automatically-inferred) types of intentions.
It also demos some additional transformers used in preprocessing steps.
Together, these transformers implement the methodology detailed in the [paper](http://www.cs.cornell.edu/~cristian/Asking_too_much.html),
```
Asking Too Much? The Rhetorical Role of Questions in Political Discourse
Justine Zhang, Arthur Spirling, Cristian Danescu-Niculescu-Mizil
Proceedings of EMNLP 2017
```
ConvoKit also includes an end-to-end implementation, `PromptTypesWrapper`, that runs the transformers one after another, and handles the particular pre-processing steps found in the paper. See [this notebook](https://github.com/CornellNLP/Cornell-Conversational-Analysis-Toolkit/blob/master/examples/prompt-types/prompt-type-wrapper-demo.ipynb) for a demonstration of this end-to-end transformer.
This is a really clear example of a method which reflects both good (we think) ideas and somewhat ad-hoc implementation decisions. As such, there are lots of options and potential variations to consider (beyond the deeper question of what phrasings and intentions even are) -- I'll detail these as I go along.
Note that due to small methodological tweaks and changes in the random seed, the particular output of the transformers as presently implemented may not totally match the output from the paper, but the broad types of questions returned are comparable.
## Preliminaries
First we load the corpus. We will examine a dataset of questions from question periods that take place in the British House of Commons (also detailed in the paper).
```
import convokit
from convokit import download
```
We'll load the corpus, plus some pre-computed dependency parses (see [this notebook](https://github.com/CornellNLP/Cornell-Conversational-Analysis-Toolkit/blob/master/examples/text-processing/text_preprocessing_demo.ipynb) for a demonstration of how to get these parses on your own; for this dataset they should be included with our release).
```
# OPTION 1: DOWNLOAD CORPUS
# UNCOMMENT THESE LINES TO DOWNLOAD CORPUS
# DATA_DIR = '<YOUR DIRECTORY>'
# ROOT_DIR = download('parliament-corpus', data_dir=DATA_DIR)
# OPTION 2: READ PREVIOUSLY-DOWNLOADED CORPUS FROM DISK
# UNCOMMENT THIS LINE AND REPLACE WITH THE DIRECTORY WHERE THE PARLIAMENT-CORPUS IS LOCATED
# ROOT_DIR = '<YOUR DIRECTORY>'
corpus = convokit.Corpus(ROOT_DIR)
corpus.load_info('utterance',['parsed'])
import warnings
warnings.filterwarnings('ignore')
VERBOSITY = 10000
```
Our specific goal, which we'll use ConvoKit to accomplish, is to produce an abstract representation of questions asked by members of parliament, in terms of:
* how they are phrased: what phrasing, or lexico-syntatic "motif", does a question have?
* their rhetorical intention: what's the intent of the asker -- which we take to mean the response the asker aims to prompt?
In other words, what are the different types of questions people ask in parliament?
Here's an example of an utterance:
```
test_utt_id = '1997-01-27a.4.0'
utt = corpus.get_utterance(test_utt_id)
utt.text
```
To state our goals more precisely:
* For each _sentence_ that has a question (all but the last), we want to come up with a representation of the sentence's phrasing. Intuitively, for instance, the first two sentences sound like they could both be thought of as a "Does X agree that Y?" -- whether Y is asking about a yacht or a harbour.
* For each utterance, we want to come up with a representation of the utterance's rhetorical intent. Intuitively, all the questions could be construed as asking if the answerer is in agreement with the asker -- whether they "agree" with the opinion or "share" the opinion. We might think of this as being an example of an "agreeing" type of question.
Intuitively, if we want to get at this higher level of abstraction, we have to look beyond the particular n-grams: it doesn't seem plausible that there is a meaningful type of question about yachts (unless our specific context is the parliamentary subcommittee on yachts).
## Preprocessing step: Arcs
One place to start is to look at the structural "skeleton" of sentences -- i.e., its dependency parse. Thus, we are first going to provide a representation of questions in terms of their dependency parse by extracting all the parent-to-child token edges, or "arcs". We will use the `TextToArcs` class to do this:
```
from convokit.text_processing import TextToArcs
```
`get_arcs` is a transformer (actually a `TextProcessor`) that will read the dependency parse of an utterance and write the resultant arcs to a field called `'arcs'`:
(demo continues after long output)
```
get_arcs = TextToArcs('arcs', verbosity=VERBOSITY)
corpus = get_arcs.transform(corpus)
```
(demo continued)
`'arcs'` is a list where each element corresponds to a sentence in the utterance. Each sentence is represented in terms of its arcs, in a space-separated string.
Each arc, in turn, can be read as follows:
* `x_y` means that `x` is the parent and `y` is the child token (e.g., `agree_does` = `agree --> does`)
* `x_*` means that `x` is a token with at least one descendant, which we do not resolve (this is roughly like bigrams backing off to unigrams)
* `x>y` means that `x` and `y` are the first two tokens in the sentence (the decision here was that how the sentence starts is a signal of "phrasing structure" on par with the dependency tree structure)
* `x>*` means that `x` is the first token in the sentence.
```
utt.retrieve_meta('arcs')
```
### Further preprocessing: cleaned-up arcs
At this point, while we've got the methodology to start making sense of the dependency tree, we arguably haven't progressed beyond producing fancy bigram representations of sentences. One problem is perhaps that the default arc extraction is a bit too permissive -- it gives us _all_ of the arcs. We might not want this for a few reasons:
* We only want to learn about question phrasings; we don't actually care about non-question sentences.
* The structure of a question might be best encapsulated by the arcs that go out of the _root_ of the tree; as you get further down we might end up with less structural and more content-specific representations.
* Likewise, the particular _nouns_ used (e.g., `yacht`) might not be good descriptions of the more abstract phrasing pattern.
All of these points are debatable, and the resultant modules I'll show below hopefully allow you to play around with them. Taking these point as is for now, though, we'll do the following.
```
from convokit.phrasing_motifs import CensorNouns, QuestionSentences
from convokit.convokitPipeline import ConvokitPipeline
```
We will actually create a pipeline to extract the arcs we want. This pipeline has the following components, in order:
* `CensorNouns`: a transformer that removes all the nouns and pronouns from a dependency parse. This transformer also collapses constructions like `What time [is it]` into `What [is it]`.
* `TextToArcs`: calling the arc extractor from above with an extra parameter: `root_only=True` which will only extract arcs attached to the root (in addition to the first two tokens, though this is also tunable by passing in parameter `use_start=True`).
* `QuestionSentences`: a transformer that, given utterance fields consisting of a list of sentences, removes all the sentences which contain question marks. Here, we want to focus on questions that are asked as part of the procedure of questions period, not questions that a minister who is playing the role of answer raises in their response. Thus, we pass an extra parameter `input_filter=question_filter`, telling it to ignore utterances which aren't labeled as (official) questions in the corpus.
* (you may wonder how this transformer can tell whether a sentence has a question mark in it, given that the output of `TextToArcs` doesn't have any punctuation. Under the hood, `QuestionSentences` looks at the dependency parse of the sentence and checks whether the last token is a question.)
* `QuestionSentences` also omits any sentences which don't begin in capital letters. To turn this off, pass parameter `use_caps=False`.
```
def question_filter(utt):
return utt.retrieve_meta('is_question')
q_arc_pipe = ConvokitPipeline([
('censor_nouns', CensorNouns('parsed_censored', verbosity=VERBOSITY)),
('shallow_arcs', TextToArcs('arcs_censored', input_field='parsed_censored',
root_only=True, verbosity=VERBOSITY)),
('question_sentence_filter', QuestionSentences('question_arcs', input_field='arcs_censored',
input_filter=question_filter, verbosity=VERBOSITY))
])
```
The pipeline should accordingly annotate each utterance with arcs for questions only, in a field called `question_arcs`.
(demo continues after long output)
```
corpus = q_arc_pipe.transform(corpus)
```
(demo continues)
This pipeline results in a more minimalistic representation of utterances, in terms of just the arcs at the root of dependency trees, just the questions, and no nouns:
```
utt.retrieve_meta('question_arcs')
```
Here's another example:
```
test_utt_id_1 = '2015-06-09c.1041.5'
utt1 = corpus.get_utterance(test_utt_id_1)
utt1.text
utt1.retrieve_meta('question_arcs')
```
## Phrasing motifs
Finally, to arrive at our representation of phrasings, we can go one further level of abstraction. In short, some of these arcs feel less fully-specified than others. While `agree_does` sounds like it hints at a coherent question, `doing_is` seems like it's not meaningful until you consider that it occurs in the same sentence as `doing_ensure` (i.e., "_what is the Government doing to ensure...?_")
Our intuition is to think of phrasings as frequently-cooccurring sets of multiple arcs. To extract these frequent arc-sets (which may remind you of the data mining idea of extracting frequent itemsets) we will use the `PhrasingMotifs` class.
```
from convokit.phrasing_motifs import PhrasingMotifs
pm_model = PhrasingMotifs('motifs','question_arcs',min_support=100,fit_filter=question_filter,
verbosity=VERBOSITY)
```
Here, `pm_model` will:
* extract all sets of arcs, as read from the `question_arcs` field, which occur at least 50 times in a corpus. These frequently-occurring arc sets will constitute the set, or "vocabulary", of phrasings.
* write the resultant output -- the phrasings that an utterance contains -- to a field called `question_motifs`.
On the latter point, `pm_model` will only transform (i.e., label phrasings for) utterances which are questions, i.e., `question_filter(utterance) = True`. That is, in both the train and transform steps, we totally ignore non-questions.
Note that the phrasings learned by `pm_model` are therefore _corpus-specific_ -- different corpora may have different frequently-occurring sets, resulting in different vocabularies of phrasings. For instance, you wouldn't expect people in the British House of Commons to ask questions that sound like questions asked to tennis players. In this respect, think of `PhrasingMotifs` like models from scikit learn (e.g., `LogisticRegression`) -- it is fit to a particular dataset:
(demo continues after long output)
```
pm_model.fit(corpus)
```
(demo continues)
Here are the most common phrasings and how often they occur in the data (in # of sentences). Note that `('*',)` denotes the null phrasing -- i.e., it encapsulates sentences with _any_ root word.
```
pm_model.print_top_phrasings(25)
```
Having "trained", or fitted our model, we can then use it to annotate each (question) utterance in the corpus with the phrasings this utterance contains, in a field called `motifs`.
(demo continues after long output)
```
corpus = pm_model.transform(corpus)
```
(demo continues)
One thing to note here is that each sentence can and probably will have multiple phrasings it embodies. For instance, two sentences with phrasing `agree_do` and `agree_will` will also have phrasing `agree_*`. Intuitively, more finely-specified phrasings (i.e., `agree_does`) more closely specify the phrasing embodied by a sentence (we could imagine "Do you agree..." and "Will you agree..." being very different, but perhaps also more similar to each other than "Can you explain..").
We want to keep track of both the complete set of phrasings and the most finely-specified phrasing you can have for each utterance. Therefore, `PhrasingMotifs` actually annotates utterances with _two_ fields.
`motifs` lists all the phrasings (arcs in a phrasing motif are separated by two underscores, `'__'`):
```
utt.retrieve_meta('motifs')
```
and `motifs__sink` lists the most finely specified _sink phrasings_ (they are "sinks" in the sense that if you think of phrasings as a directed graph where A-->B when B is a more finely-specified version of A, these sinks have no child phrasings which are contained in the utterance)
```
utt.retrieve_meta('motifs__sink')
```
### model persistence
We can save `pm_model` to disk and later reload it, thus caching the trained model (i.e., the motifs in a corpus and the internal representation of these motifs). Here, we save the model to a `pm_model` subfolder in the corpus directory via `dump_model()`:
```
import os
pm_model.dump_model(os.path.join(ROOT_DIR, 'pm_model'))
```
This subfolder then stores the motifs, as well as relations between the motifs that facilitate transforming new utterances.
```
pm_model_dir = os.path.join(ROOT_DIR, 'pm_model')
!ls $pm_model_dir
```
Suppose we later initialize a new `PhrasingMotifs` model, `new_pm_model`.
```
new_pm_model = PhrasingMotifs('motifs_new','question_arcs',min_support=100,fit_filter=question_filter,
verbosity=VERBOSITY)
```
Calling `load_model()` then reloads the stored model from our earlier run into this new model:
```
new_pm_model.load_model(os.path.join(ROOT_DIR, 'pm_model'))
```
Just to check that we've loaded the same thing that we previously saved, we'll get the motifs in our test utterance using `new_pm_model`:
```
utt = new_pm_model.transform_utterance(utt)
```
This is the output from the original run:
```
utt.retrieve_meta('motifs__sink')
```
And we see the new output matches.
```
utt.retrieve_meta('motifs_new__sink')
```
### example variation: not removing the nouns
**note** this takes a while to run, and is somewhat of an extension -- you can safely skip these cells.
There are other ways to use `PhrasingMotifs` that might be more or less suited to your own application. For instance, you may wonder what happens if we do not remove the nouns (as we did with `CensorNouns` above). To try this out, we can create an alternate pipeline that uses `TextToArcs` to generate root arcs (setting argument `root_only=True`) on the original parses, not the noun-censored ones.
```
q_arc_pipe_full = ConvokitPipeline([
('shallow_arcs_full', TextToArcs('root_arcs', input_field='parsed',
root_only=True, verbosity=VERBOSITY)),
('question_sentence_filter', QuestionSentences('question_arcs_full', input_field='root_arcs',
input_filter=question_filter, verbosity=VERBOSITY)),
])
corpus = q_arc_pipe_full.transform(corpus)
```
We can then train a new `PhrasingMotifs` model that finds phrasings with the nouns still included.
(demo continues after long output)
```
noun_pm_model = PhrasingMotifs('motifs_full','question_arcs_full',min_support=100,
fit_filter=question_filter,
verbosity=VERBOSITY)
noun_pm_model.fit(corpus)
```
(demo continues)
The most common phrasings, of course, won't be very topic-specific (unless people talk about yachts very very frequently in parliament). However, we do see that phrasings now reflect the pronoun used (which may be troublesome if we believe that "Does _he_ agree" and "Does _she_ agree" are getting at similar things).
```
noun_pm_model.print_top_phrasings(25)
```
Here are the sink phrasings for our example utterance from earlier, comparing against the noun-less run:
```
utt = noun_pm_model.transform_utterance(utt)
utt.retrieve_meta('motifs__sink')
utt.retrieve_meta('motifs_full__sink')
```
We see that we get this extra "hon" -- which actually stands for "honourable [member]" -- an artefact of parliamentary etiquette.
For our particular dataset, removing nouns has the benefit of removing most of these etiquette-related words. However, you may also imagine cases where nouns actually carry a lot of useful information about rhetorical intent (including in this domain -- one could argue that asking about a person versus asking about a department is a strong signal of trying to get at different things, for instance). As such, noun-removal is something that you may want to play around with, and/or try to improve upon.
## PromptTypes
As we intuited above, "do you agree" and "do you share my opinion" are both getting at similar intentions. However, extracting these phrasings alone won't allow us to make this association. Rather, our strategy will be to produce vector representations of them which encode this similarity. Clustering these representations then gives us different "types of question".
Our key intuition here is that questions with similar intentions will tend to be answered in similiar ways. Thus, "do you agree" and "do you share" may both often be answered with "yes, I agree"; if tomorrow I asked a new question of this ilk ("do you agree that we should invest in planes, instead of yachts"), I might be expecting a similar sort of answer.
For a full explanation of this idea, and how we operationalized it, you can read our paper. In ConvoKit, we implement this methodology of producing vector representations and clusterings via the `PromptTypes` transformer:
```
from convokit.prompt_types import PromptTypes
```
`PromptTypes` will train a model -- a low-dimensional embedding, along with a k-means clustering -- by using question-answer pairs as input.
```
def question_filter(utt):
return utt.retrieve_meta('is_question')
def response_filter(utt):
return (not utt.retrieve_meta('is_question')) and (utt.reply_to is not None)
```
We initialize `pt` with the following arguments:
* `n_types=8`: we want to infer 8 types of questions.
* `prompt_field='motifs'`: we want to encode questions in terms of the phrasing motifs we extracted above. thus, `pt` will produce representations of these motifs (rather than, e.g., the raw tokens in a question)
* `reference_field='arcs_censored'`: we will encode responses in terms of the noun-less arcs we extracted above (in practice, this appears to work better than using phrasings of responses as well, perhaps because responses are noisier)
* `prompt_transform_field='motifs__sink'`: while we want to come up with a representation of _all_ phrasing motifs, when we produce a vector representation of a _particular_ utterance we want to use the most finely-specified phrasing.
There are some other arguments you can set, which are listed in the docstring.
```
pt = PromptTypes(n_types=8, prompt_field='motifs', reference_field='arcs_censored',
prompt_transform_field='motifs__sink',
output_field='prompt_types',
random_state=1000, verbosity=1)
```
We can fit `pt` to the corpus -- that is, learn the associations between question phrasings and response dependency arcs that allow us to produce our vector representations, as well as a clsutering of these representations that gives us our different question types. To focus on questions, we will use the following filters to select a subset of the corpus as training data:
* `prompt_selector=question_filter` and `ref_selector=response_filter`: To tell the transformer what counts as a question and an answer, we will pass the constructor the above filters (i.e., boolean functions). Note that in a less questions-heavy dataset, we could omit these filters and hence infer types of "prompts" beyond questions.
```
pt.fit(corpus, prompt_selector=question_filter, reference_selector=response_filter)
```
calling `summarize()` as below will print the question phrasings, response arcs, and prototypical questions and responses that are associated with each inferred type of question. We will examine some of these types more closely by way of examples, below.
(long output follows)
```
pt.summarize(corpus=corpus, k=15)
```
(demo continues)
When this trained model is used to transform a corpus, it will output several representations or features associated with each utterance.
Detail: The fields corresponding to representations and features computed with the model are each titled `prompt_types__<feature name>`. The prefix, `prompt_types` can be modified by changing the `output_field` argument of the constructor.
```
utt = pt.transform_utterance(utt)
```
First, a vector representation encapsulating the utterance's rhetorical intent (in short, an embedding of the utterance based on the responses associated with questions containing its constituent phrasings):
```
utt.retrieve_meta('prompt_types__prompt_repr')
```
Second, the distance between the vector of that utterance and the centroid of each cluster, or type of question.
```
utt.retrieve_meta('prompt_types__prompt_dists.8')
```
The particular type of question this utterance is, i.e., the centroid that its vector representation is closest to, as well its distance to the centroid (roughly, how well it fits that question type):
```
utt.retrieve_meta('prompt_types__prompt_type_dist.8')
```
Here, we see that our running example is of a question type exemplified by phrasings like `does [the Minister] agree...` -- we may characterize the entire cluster as encapsulating "agreeing" questions which are perhaps asked helpfully to bolster the answerer's reputation.
(long output)
```
pt.summarize(corpus,type_ids=utt.retrieve_meta('prompt_types__prompt_type.8'), k=15)
```
(demo continues)
We can transform the other utterances in the corpus as such:
```
corpus = pt.transform(corpus)
```
Having transformed the corpus, we can first see what the prompt types of other utterances are.
This utterance is of a type that's perhaps more information-seeking and querying for an update ("what steps is the Government taking, what are they doing to ensure", etc)
```
utt1.text
utt1.retrieve_meta('motifs__sink')
utt1.retrieve_meta('prompt_types__prompt_type.8')
```
(long output)
```
pt.summarize(corpus,type_ids=utt1.retrieve_meta('prompt_types__prompt_type.8'), k=15)
```
(demo continues)
This utterance, on the other hand is a lot more aggressive -- perhaps _accusatory_ to the ends of putting the answerer on the spot ("will the secretary admit that the policy is a failure?")
```
utt2 = corpus.get_utterance('1987-03-04a.857.5')
utt2.text
utt2.retrieve_meta('motifs__sink')
utt2.retrieve_meta('prompt_types__prompt_type.8')
```
(long output)
```
pt.summarize(corpus,type_ids=utt2.retrieve_meta('prompt_types__prompt_type.8'), k=15)
```
(demo continues)
The transform step also annotates _responses_ to questions in question period with the prompt type that the utterance is the most appropriate response to, according to our model.
For instance, consider the following response: (which actually responds to the utterance in the first example)
```
response_utt = corpus.get_utterance('1997-01-27a.4.1')
corpus.get_utterance(response_utt.reply_to).text
response_utt.text
```
The response has the following type:
```
response_utt.retrieve_meta('prompt_types__reference_type.8')
```
corresponding to questions which, like the first example, are relatively friendly and agreeable. In other words, out of all the prompt types, the response would be the most appropriate reply to these agreeable questions.
(detail: the term `reference_type` refers to the fact that the set of question responses are used as "references" to structure the space of questions, per the methodology detailed in the paper. We keep the terminology deliberately generic, as opposed to calling it a "response type", to suggest that other data could serve as references -- for instance, reversing the role of questions and answers in the method. This possibility is something we'll explore in future work.)
### vector representations
As mentioned above, `PromptTypes` produces a few vector representations of utterances. For efficiency, rather than storing these representations attached to the utterance (as values in `utterance.meta`), we store them in corpus-wide matrices. (See the following demo [here](https://github.com/CornellNLP/Cornell-Conversational-Analysis-Toolkit/blob/master/examples/vectors/vector_demo.ipynb) for more details on using vectors in ConvoKit.)
To view (a subset of) these matrices, we'll call `corpus.get_vectors(matrix name, [utterance ids])` which allows us to access the vectors corresponding to all utterances in the list of [utterance ids]).
Each row of the matrix `prompt_types__prompt_repr` corresponds to the vector representation of the utterance's latent intent. The first row should be exactly the latent representation we printed above, for the first example question:
(here I've returned the matrix as a dataframe to highlight that each row corresponds to an utterance indicated by the ID in the index)
```
corpus.get_vectors('prompt_types__prompt_repr', ids=[utt.id, utt1.id, utt2.id],
as_dataframe=True).head()
```
Each row of the matrix `prompt_types__prompt_dists.8` lists the distances between each question (as its vector representation) and each type centroid. The first row should be exactly the distances corresponding to the first example question, as we printed above.
```
corpus.get_vectors('prompt_types__prompt_dists.8', ids=[utt.id, utt1.id, utt2.id],
as_dataframe=True)
```
Note that these vectors could be used as features in a prediction task: [this notebook](https://github.com/CornellNLP/Cornell-Conversational-Analysis-Toolkit/blob/master/examples/conversations-gone-awry/Conversations_Gone_Awry_Prediction.ipynb) has an example for predicting derailed conversations, using the distance between utterances and type centroids (`prompt_dists.#`); using the latent representations (`prompt_repr`) is another good option.
To save all of these representations to disk, we can call the following:
```
corpus.dump_vectors('prompt_types__prompt_repr')
corpus.dump_vectors('prompt_types__prompt_dists.8')
```
These vector representations can later be re-loaded:
```
new_corpus = convokit.Corpus(ROOT_DIR, preload_vectors=['prompt_types__prompt_repr',
'prompt_types__prompt_dists.8'])
new_corpus.get_vectors('prompt_types__prompt_repr', ids=[utt.id, utt1.id, utt2.id],
as_dataframe=True)
new_corpus.get_vectors('prompt_types__prompt_dists.8', ids=[utt.id, utt1.id, utt2.id],
as_dataframe=True)
```
### a few caveats and potential modifications
One thorn in our sides might be that the model occasionally gets caught up on very generic motifs e.g., `'is>*'`, and as such, will fit many questions to the type containing `'is>*'` instead of going with a better signal; various optional parameters detailed in the documentation may provide incomplete solutions to this. Another caveat is that while this model allows us to associate together lexically-diverging phrasings (e.g., "will the Minister admit" and "does the Minister not realise" both serve to be accusatory towards the Minister), we are ultimately relying on the fact that our domain has a sufficient amount of lexical regularity (e.g., the institutional norms of how people talk in parliament) -- we might need to be cleverer when dealing with noisier settings where this regularity isn't guaranteed (like social media data).
Finally, as a data-specific note, one of the types may be a result of the parser assuming that "Will the learned Gentleman please answer my question?" has "learned" as the root verb -- an artefact of parliamentary discourse we haven't handled. You may wish to play around with this by modifying how the data is preprocessed.
### model persistence
We can save our trained `pt_model` to disk for later use:
```
import os
pt.dump_model(os.path.join(ROOT_DIR, 'pt_model'))
```
In broad strokes, what's loaded to disk is:
* TfIdf models that store the distribution of phrasings and arcs in the training data;
* SVD models that allow us to map raw phrasing/arc counts to vector representations;
* a KMeans model to cluster vector representations.
```
pt_model_dir = os.path.join(ROOT_DIR, 'pt_model')
!ls $pt_model_dir
```
Initializing a new `PromptTypes` model and loading our saved model then allows us to use it again:
```
new_pt = PromptTypes(prompt_field='motifs', reference_field='arcs_censored',
prompt_transform_field='motifs__sink',
output_field='prompt_types_new', prompt__tfidf_min_df=100,
reference__tfidf_min_df=100,
random_state=1000, verbosity=1)
new_pt.load_model(pt_model_dir)
utt = new_pt.transform_utterance(utt)
utt.retrieve_meta('prompt_types_new__prompt_type.8')
```
## examples of potential variations
### trying other numbers of prompt types:
Calling `refit_types(n)` will retrain the clustering component of the `PromptType` model to infer a different number of types. Suppose we only wanted 4 types of questions:
```
pt.refit_types(4)
```
(long output)
```
pt.summarize(corpus, type_key=4, k=15)
```
(demo continues)
### trying other input formats
We may also experiment with different representations of the input text -- for instance, in lieu of using phrasing motifs we may instead pass questions into the model as just the raw arcs, similar to the responses. This may help with datasets where the phrasing motifs are relatively sparse (due to size or noise/linguistic variability). This can be modified by changing the `prompt_field` argument:
```
pt_arcs = PromptTypes(prompt_field='arcs_censored', reference_field='arcs_censored',
prompt_transform_field='arcs_censored',
output_field='prompt_types_arcs', prompt__tfidf_min_df=100,
reference__tfidf_min_df=100, n_types=8,
random_state=1000, verbosity=1)
pt_arcs.fit(corpus)
```
(long output)
```
pt_arcs.summarize(corpus, k=10)
```
(demo continues)
### going beyond root arcs
If we initialize the `TextToArcs` transformer with `root_only=False`, we will use arcs beyond those attached to the root of the dependency parse. This may produce neater output, especially in domains where utterances are less well-structured (see [this notebook](https://github.com/CornellNLP/Cornell-Conversational-Analysis-Toolkit/blob/master/examples/conversations-gone-awry/Conversations_Gone_Awry_Prediction.ipynb) for a demo of this on Wikipedia talk page data)
| github_jupyter |
```
from IPython.display import YouTubeVideo
YouTubeVideo('W-ZsWqcl1_c')
```
# 如何使用和开发微信聊天机器人的系列教程
# A workshop to develop & use an intelligent and interactive chat-bot in WeChat
### WeChat is a popular social media app, which has more than 800 million monthly active users.
<img src='http://www.kudosdata.com/wp-content/uploads/2016/11/cropped-KudosLogo1.png' width=30% style="float: right;">
<img src='reference/WeChat_SamGu_QR.png' width=10% style="float: right;">
### http://www.KudosData.com
by: Sam.Gu@KudosData.com
April 2017 ========== Scan the QR code to become trainer's friend in WeChat ========>>
### 第一课:使用微信问答机制
### Lesson 1: Basic usage of WeChat Python API
* 使用和开发微信个人号聊天机器人:一种Python编程接口 (Use WeChat Python API)
* 用微信App扫QR码图片来自动登录 (Log-in, contact scan, and processing of text, image, file, video, etc)
* 查找指定联系人或群组 (Scan ccontact list)
* 发送信息(文字、图片、文件、音频、视频等) (Send message: text, image, file, voice, video, etc)
* 接收信息 (Receive message, and keep 'listening')
* 自动回复 (Receive message and then automaticaly reply)
* 自定义复杂消息处理,例如:信息存档、回复群组中被@的消息 (Advanced message processing and reply)
### 导入需要用到的一些功能程序库:
```
# from __future__ import unicode_literals, division
import time, datetime, requests, itchat
from itchat.content import *
```
### * 用微信App扫QR码图片来自动登录
```
itchat.auto_login(hotReload=True) # hotReload=True: 退出程序后暂存登陆状态。即使程序关闭,一定时间内重新开启也可以不用重新扫码。
# itchat.auto_login(enableCmdQR=-2) # enableCmdQR=-2: 命令行显示QR图片
```
### * 查找指定联系人或群组
使用search_friends方法可以搜索用户,有几种搜索方式:
1.仅获取自己的用户信息
2.获取昵称'NickName'、微信号'Alias'、备注名'RemarkName'中的任何一项等于name键值的用户
3.获取分别对应相应键值的用户
```
# 获取自己的用户信息,返回自己的属性字典
friend = itchat.search_friends()
print(friend)
print('NickName : %s' % friend['NickName'])
print('Alias A-ID: %s' % friend['Alias'])
print('RemarkName: %s' % friend['RemarkName'])
print('UserName : %s' % friend['UserName'])
# 获取任何一项等于name键值的用户。
# 'NickName' 昵称, set by that friend, changeable
# 'Alias' ID微信号 = wechatAccount, one time set by that friend, cannot change
# 'RemarkName' 备注名, set by current login account owner, changeable by login account owner
# 注意:返回可能包含多个朋友。为什么呢?
friend = itchat.search_friends(name=u'Sam Gu')
# friend = itchat.search_friends(name=u'Mr. R')
# friend = itchat.search_friends(name=u'Ms. S')
for i in range(0, len(friend)):
print('NickName : %s' % friend[i]['NickName'])
print('Alias A-ID: %s' % friend[i]['Alias'])
print('RemarkName: %s' % friend[i]['RemarkName'])
print('UserName : %s' % friend[i]['UserName'])
# 获取分别对应相应键值的用户。
# friend = itchat.search_friends(nickName=u'Sam Gu')
# friend = itchat.search_friends(wechatAccount=u'Sam Gu')
friend = itchat.search_friends(remarkName=u'Sam Gu')
# friend = itchat.search_friends(userName=u'Sam Gu')
for i in range(0, len(friend)):
print('NickName : %s' % friend[i]['NickName'])
print('Alias A-ID: %s' % friend[i]['Alias'])
print('RemarkName: %s' % friend[i]['RemarkName'])
print('UserName : %s' % friend[i]['UserName'])
# 查找群组
# group = itchat.search_chatrooms(name=u'Data Science')
group = itchat.search_chatrooms(name=u'陪聊妹UAT')
for i in range(0, len(group)):
print('NickName : %s' % group[i]['NickName'])
print('Alias A-ID: %s' % group[i]['Alias'])
print('RemarkName: %s' % group[i]['RemarkName'])
print('UserName : %s' % group[i]['UserName'])
print('Is Owner? : %s ( 0 for No | 1 for Yes )' % group[0]['IsOwner'])
print('Is Admin? : %s' % group[i]['IsAdmin'])
print('')
```
### * 发送信息(文字、图片、文件、音频、视频等)
```
# 文字
reply = itchat.send(u'别来无恙啊!\n发送时间:\n{:%Y-%b-%d %H:%M:%S}'.format(datetime.datetime.now()), friend[0]['UserName'])
print(reply['BaseResponse']['ErrMsg'])
# 图片
reply = itchat.send_image('./reference/WeChat_SamGu_QR.png', friend[0]['UserName'])
print(reply['BaseResponse']['ErrMsg'])
# 文件
reply = itchat.send_file('./reference/logo.pdf', friend[0]['UserName'])
print(reply['BaseResponse']['ErrMsg'])
# 音频(语音可以先转成MP3)
reply = itchat.send_file('./reference/audio.mp3', friend[0]['UserName'])
print(reply['BaseResponse']['ErrMsg'])
# 视频
reply = itchat.send_video('./reference/video.mp4', friend[0]['UserName'])
print(reply['BaseResponse']['ErrMsg'])
# 发送信息去群组: group[0]['UserName']
# 文字
reply = itchat.send(u'别来无恙啊!\n发送时间:\n{:%Y-%b-%d %H:%M:%S}'.format(datetime.datetime.now()), group[0]['UserName'])
print(reply['BaseResponse']['ErrMsg'])
```
### * 接收信息
显示发给自己的文本消息:
```
# itchat.auto_login(hotReload=True) # hotReload=True: 退出程序后暂存登陆状态。即使程序关闭,一定时间内重新开启也可以不用重新扫码。
@itchat.msg_register(itchat.content.TEXT)
def text_reply(msg):
print(msg['Text'])
# 长期有效地运行(术语叫做:开始监听)
itchat.run()
```
回复发给自己的文本消息:
```
# interupt, then re-login
itchat.auto_login(hotReload=True)
@itchat.msg_register(itchat.content.TEXT)
def text_reply(msg):
print(msg['Text'])
return u'谢谢亲[嘴唇]我收到 I received:\n' + msg['Text']
itchat.run()
```
### * 自定义复杂消息处理,例如:信息存档、回复群组中被@的消息
```
# interupt, then re-login
itchat.auto_login(hotReload=True)
# 如果收到[TEXT, MAP, CARD, NOTE, SHARING]类的信息,会自动回复:
@itchat.msg_register([TEXT, MAP, CARD, NOTE, SHARING]) # 文字、位置、名片、通知、分享
def text_reply(msg):
itchat.send('%s: %s' % (msg['Type'], msg['Text']), msg['FromUserName'])
# 如果收到[PICTURE, RECORDING, ATTACHMENT, VIDEO]类的信息,会自动保存:
@itchat.msg_register([PICTURE, RECORDING, ATTACHMENT, VIDEO]) # 图片、语音、文件、视频
def download_files(msg):
msg['Text'](msg['FileName'])
return '@%s@%s' % ({'Picture': 'img', 'Video': 'vid'}.get(msg['Type'], 'fil'), msg['FileName'])
# 如果收到新朋友的请求,会自动通过验证添加加好友,并主动打个招呼:幸会幸会!Nice to meet you!
@itchat.msg_register(FRIENDS)
def add_friend(msg):
itchat.add_friend(**msg['Text']) # 该操作会自动将新好友的消息录入,不需要重载通讯录
itchat.send_msg(u'幸会幸会!Nice to meet you!', msg['RecommendInfo']['UserName'])
# 在群里,如果收到@自己的文字信息,会自动回复:
@itchat.msg_register(TEXT, isGroupChat=True)
def text_reply(msg):
if msg['isAt']:
itchat.send(u'@%s\u2005I received: %s' % (msg['ActualNickName'], msg['Content']), msg['FromUserName'])
itchat.run()
# interupt, then logout
itchat.logout() # 安全退出
```
### 恭喜您!已经能够使用微信问答机制了。
* 使用和开发微信个人号聊天机器人:一种Python编程接口 (Use WeChat Python API)
* 用微信App扫QR码图片来自动登录 (Log-in, contact scan, and processing of text, image, file, video, etc)
* 查找指定联系人或群组 (Scan ccontact list)
* 发送信息(文字、图片、文件、音频、视频等) (Send message: text, image, file, voice, video, etc)
* 接收信息 (Receive message, and keep 'listening')
* 自动回复 (Receive message and then automaticaly reply)
* 自定义复杂消息处理,例如:信息存档、回复群组中被@的消息 (Advanced message processing and reply)
### 下一课是第二课:图像识别和处理
### Lesson 2: Image Recognition & Processing
* 识别图片消息中的物体名字 (Recognize objects in image)
* 识别图片消息中的文字 (OCR: Extract text from image)
* 识别人脸 (Recognize human face)
* 基于人脸的表情来识别喜怒哀乐等情绪 (Identify semtiment and emotion from human face)
<img src='http://www.kudosdata.com/wp-content/uploads/2016/11/cropped-KudosLogo1.png' width=30% style="float: right;">
<img src='reference/WeChat_SamGu_QR.png' width=10% style="float: left;">
| github_jupyter |
<span style="color:#8735fb; font-size:22pt"> **Demo Overview** </span>
Automated Model Tuning (AMT) also known as Hyper-Parameter Optimization (HPO) helps to find the best version of a model by exploring the space of possible configurations. While generally desirable, this search is computationally expensive and can feel prohibitive.
In the notebook demo below, we show how [SageMaker Studio](https://docs.aws.amazon.com/sagemaker/latest/dg/studio.html) and RAPIDS working together can tackle model tuning by accelerating compute parallelism within a node's GPUs; and simultaneously accelerating the search by leveraging sets of cloud nodes running parallel experiments.
For more check out our [AWS blog](https://aws.amazon.com/blogs/machine-learning/rapids-and-amazon-sagemaker-scale-up-and-scale-out-to-tackle-ml-challenges/).
<span style="color:#8735fb; font-size:22pt"> **0. Preamble** </span>
This notebook was tested in an Amazon SageMaker Studio notebook, on a ml.t3.medium instance with Python 3 (Data Science) kernel.
To get things rolling let's make sure we can query our AWS SageMaker execution role and session as well as our account ID and AWS region.
```
import sagemaker
from helper_functions import *
execution_role = sagemaker.get_execution_role()
session = sagemaker.Session()
account=!(aws sts get-caller-identity --query Account --output text)
region = [session.boto_region_name]
account, region
```
<span style="color:#8735fb; font-size:22pt"> **1. Key Choices** </span>
<span style="color:#8735fb; font-size:18pt"> 1.1 - HPO Configurations </span>
Let's go ahead and choose the configuration options for our HPO run. We will be using the default algorithm configurations in [code/Dockerfile](code/Dockerfile) (three-fold cross validation with XGBoost on a single GPU), which are explained in detail in our [extended notebook example](rapids_sagemaker_hpo_extended.ipynb). If you are using your own workflow and training scripts, be sure to write your Dockerfile accordingly.
```
# please choose dataset S3 bucket and directory
data_bucket = 'sagemaker-rapids-hpo-' + region[0]
dataset_directory = '3_year' # '1_year', '3_year', '10_year', 'NYC_taxi'
# please choose output bucket for trained model(s)
model_output_bucket = session.default_bucket()
s3_data_input = f"s3://{data_bucket}/{dataset_directory}"
s3_model_output = f"s3://{model_output_bucket}/trained-models"
# please choose HPO search ranges
hyperparameter_ranges = {
'max_depth' : sagemaker.parameter.IntegerParameter ( 5, 15 ),
'num_boost_round' : sagemaker.parameter.IntegerParameter ( 100, 500 ),
'max_features' : sagemaker.parameter.ContinuousParameter ( 0.1, 1.0 ),
}
```
<span style="color:#8735fb; font-size:18pt"> 1.2 - Experiment Scale </span>
We also need to decide how may total experiments to run, and how many should run in parallel. Below we have a very conservative number of maximum jobs to run so that you don't accidently spawn large computations when starting out, however for meaningful HPO searches this number should be much higher (e.g., in our experiments we often run 100 max_jobs). Note that you may need to request a [quota limit increase](https://docs.aws.amazon.com/general/latest/gr/sagemaker.html) for additional `max_parallel_jobs` parallel workers.
```
# please choose total number of HPO experiments[ we have set this number very low to allow for automated CI testing ]
max_jobs = 2
# please choose number of experiments that can run in parallel
max_parallel_jobs = 2
```
Let's also set the max duration for an individual job to 24 hours so we don't have run-away compute jobs taking too long.
```
max_duration_of_experiment_seconds = 60 * 60 * 24
```
<span style="color:#8735fb; font-size:18pt"> 1.3 - Compute Platform </span>
Depending on the workflow you have chosen, your instance should reflect the specifications needed. For example, for the singleGPU workflow, you should choose an instance with a GPU, such as the p3.2xlarge instance. You can [read about Amazon EC2 Instance Types here](https://aws.amazon.com/ec2/instance-types/).
> e.g., For the 10_year dataset option, we suggest ml.p3.8xlarge instances (4 GPUs) and ml.m5.24xlarge CPU instances ( we will need upwards of 200GB CPU RAM during model training).
```
# we will recommend a compute instance type, feel free to modify
instance_type = 'ml.p3.2xlarge' # recommend_instance_type(ml_workflow_choice, dataset_directory)
```
In addition to choosing our instance type, we can also enable significant savings by leveraging [AWS EC2 Spot Instances](https://aws.amazon.com/ec2/spot/).
We **highly recommend** that you set this flag to `True` as it typically leads to 60-70% cost savings. Note, however that you may need to request a [quota limit increase](https://docs.aws.amazon.com/general/latest/gr/sagemaker.html) to enable Spot instances in SageMaker.
```
# please choose whether spot instances should be used
use_spot_instances_flag = True
```
<span style="color:#8735fb; font-size:22pt"> **2. Build Estimator** </span>
To build a RAPIDS enabled SageMaker HPO we first need to build a [SageMaker Estimator](https://sagemaker.readthedocs.io/en/stable/api/training/estimators.html). An Estimator is a container image that captures all the software needed to run an HPO experiment. The container is augmented with entrypoint code that will be trggered at runtime by each worker. The entrypoint code enables us to write custom models and hook them up to data.
In order to work with SageMaker HPO, the entrypoint logic should parse hyperparameters (supplied by AWS SageMaker), load and split data, build and train a model, score/evaluate the trained model, and emit an output representing the final score for the given hyperparameter setting. We've already built multiple variations of this code.
If you would like to make changes by adding your custom model logic feel free to modify the **train.py** and/or the specific workflow files in the `code/workflows` directory.
First, let's switch our working directory to the location of the Estimator entrypoint and library code.
```
%cd code
```
<span style="color:#8735fb; font-size:20pt"> 2.1 - Containerize and Push to ECR </span>
Now let's turn to building our container so that it can integrate with the AWS SageMaker HPO API.
Let's first decide on the full name of our container.
```
rapids_base_container = 'rapidsai/rapidsai-cloud-ml:latest'
image_base = 'cloud-ml-sagemaker'
image_tag = rapids_base_container.split(':')[1]
ecr_fullname = f"{account[0]}.dkr.ecr.{region[0]}.amazonaws.com/{image_base}:{image_tag}"
ecr_fullname
```
Lastly, let's ensure that our Dockerfile correctly captured our base image selection.
```
validate_dockerfile(rapids_base_container)
!cat Dockerfile
```
<span style="color:#8735fb; font-size:18pt"> 2.1.1 Build and Publish to ECR</span>
In order to build and push to the ECR from SageMaker Studio, we must first install sm-docker:
```
!pip install sagemaker-studio-image-build
```
Now we’re ready to start taking advantage of the new CLI to easily build our custom bring-your-own Docker image from Amazon SageMaker Studio without worrying about the underlying setup and configuration of build services. Once in ECR, AWS SageMaker will be able to leverage our image to build Estimators and run experiments. We are able to build and publish to the ECR with:
```
%%time
!sm-docker build . --repository cloud-ml-sagemaker:latest
```
<span style="color:#8735fb; font-size:20pt"> 2.2 - Create Estimator </span>
Having built our container [ +custom logic] and pushed it to ECR, we can finally compile all of efforts into an Estimator instance.
```
# 'volume_size' - EBS volume size in GB, default = 30
estimator_params = {
'image_uri': ecr_fullname,
'role': execution_role,
'instance_type': instance_type,
'instance_count': 1,
'input_mode': 'File',
'output_path': s3_model_output,
'use_spot_instances': use_spot_instances_flag,
'max_run': max_duration_of_experiment_seconds, # 24 hours
'sagemaker_session': session,
}
if use_spot_instances_flag == True:
estimator_params.update({'max_wait' : max_duration_of_experiment_seconds + 1})
estimator = sagemaker.estimator.Estimator(**estimator_params)
```
<span style="color:#8735fb; font-size:20pt"> 2.3 - Test Estimator </span>
Now we are ready to test by asking SageMaker to run the BYOContainer logic inside our Estimator. This is a useful step if you've made changes to your custom logic and are interested in making sure everything works before launching a large HPO search.
> Note: This verification step will use the default hyperparameter values declared in our custom train code, as SageMaker HPO will not be orchestrating a search for this single run.
```
estimator.fit(inputs = s3_data_input)
```
<span style="display: block; text-align: center; color:#8735fb; font-size:30pt"> **3. Run HPO** </span>
With a working SageMaker Estimator in hand, the hardest part is behind us. In the key choices section we <a href='#strategy-and-param-ranges'>already defined our search strategy and hyperparameter ranges</a>, so all that remains is to choose a metric to evaluate performance on. For more documentation check out the AWS SageMaker [Hyperparameter Tuner documentation](https://sagemaker.readthedocs.io/en/stable/tuner.html).
<span style="color:#8735fb; font-size:20pt"> 3.1 - Define Metric </span>
We only focus on a single metric, which we call 'final-score', that captures the accuracy of our model on the test data unseen during training. You are of course welcome to add aditional metrics, see [AWS SageMaker documentation on Metrics](https://docs.aws.amazon.com/sagemaker/latest/dg/automatic-model-tuning-define-metrics.html). When defining a metric we provide a regular expression (i.e., string parsing rule) to extract the key metric from the output of each Estimator/worker.
```
metric_definitions = [{'Name': 'final-score', 'Regex': 'final-score: (.*);'}]
objective_metric_name = 'final-score'
```
<span style="color:#8735fb; font-size:20pt"> 3.2 - Define Tuner </span>
Finally we put all of the elements we've been building up together into a HyperparameterTuner declaration.
```
hpo = sagemaker.tuner.HyperparameterTuner(estimator=estimator,
metric_definitions=metric_definitions,
objective_metric_name=objective_metric_name,
objective_type='Maximize',
hyperparameter_ranges=hyperparameter_ranges,
strategy='Random',
max_jobs=max_jobs,
max_parallel_jobs=max_parallel_jobs)
```
<span style="color:#8735fb; font-size:20pt"> 3.3 - Run HPO </span>
Let's be sure we take a moment to confirm before launching all of our HPO experiments. Depending on your configuration options running this cell can kick off a massive amount of computation!
> Once this process begins, we recommend that you use the SageMaker UI to keep track of the <a href='../img/gpu_hpo_100x10.png'>health of the HPO process and the individual workers</a>.
```
import random
import string
tuning_job_name = 'unified-hpo-' + ''.join(random.choices(string.digits, k = 5))
hpo.fit( inputs=s3_data_input,
job_name=tuning_job_name,
wait=True,
logs='All')
hpo.wait() # block until the .fit call above is completed
```
<span style="color:#8735fb; font-size:20pt"> 3.4 - Results and Summary </span>
Once your job is complete there are multiple ways to analyze the results. Below we display the performance of the best job, as well printing each HPO trial/job as a row of a dataframe.
```
hpo_results = summarize_hpo_results(tuning_job_name)
sagemaker.HyperparameterTuningJobAnalytics(tuning_job_name).dataframe()
```
<span style="color:#8735fb; font-size:30pt">RAPIDS References</span>
> [cloud-ml-examples](http://github.com/rapidsai/cloud-ml-examples)
> [RAPIDS HPO](https://rapids.ai/hpo)
> [cuML Documentation](https://docs.rapids.ai/api/cuml/stable/)
<span style="color:#8735fb; font-size:30pt">SageMaker References</span>
> [SageMaker Training Toolkit](https://github.com/aws/sagemaker-training-toolkit)
> [Getting Started with SageMaker Studio](https://sagemaker-examples.readthedocs.io/en/latest/aws_sagemaker_studio/index.html)
> [Using the Amazon SageMaker Studio Image Build CLI to build container images from your Studio notebooks](https://aws.amazon.com/blogs/machine-learning/using-the-amazon-sagemaker-studio-image-build-cli-to-build-container-images-from-your-studio-notebooks/)
> [Docker containers with SageMaker](https://docs.aws.amazon.com/sagemaker/latest/dg/docker-basics.html)
> [Estimator Parameters](https://sagemaker.readthedocs.io/en/stable/api/training/estimators.html)
> Spot Instances [docs](https://docs.aws.amazon.com/sagemaker/latest/dg/model-managed-spot-training.html), and [blog]()
| github_jupyter |
# Experimental Features
Here are some examples of experimental features we're planning to add to our API. These wrappers abstract away some parts of the protobuf structs, and provide easier access to geometries and `datetime`.
## First Code Example
This is an alternative to the original version [here](./README.md#first-code-example). This simplifies the setting of the `observed` field and hides the use of more verbose protobuf classes for both time filter (`observed`) and spatial filter (`intersects`).
```
import tempfile
from datetime import date
from IPython.display import Image, display
from epl.geometry import Point
from nsl.stac import enum, utils
from nsl.stac.experimental import NSLClientEx, StacRequestWrap
# the client package stubs out a little bit of the gRPC connection code
# get a client interface to the gRPC channel. This client singleton is threadsafe
client = NSLClientEx()
# create our request. this interface allows us to set fields in our protobuf object
request = StacRequestWrap()
# our area of interest will be the coordinates of the UT Stadium in Austin Texas
# the order of coordinates here is longitude then latitude (x, y). The results of our query
# will be returned only if they intersect this point geometry we've defined (other geometry
# types besides points are supported)
#
# This string format, POINT(float, float) is the well-known-text geometry format:
# https://en.wikipedia.org/wiki/Well-known_text_representation_of_geometry
#
# the epsg # defines the WGS-84 elispsoid (`epsg=4326`) spatial reference
# (the latitude longitude spatial reference most commonly used)
#
# the epl.geometry Point class is an extension of shapely's Point class that supports
# the protobuf definitions we use with STAC. To extract a shapely geometry from it use
# the shapely_dump property
request.intersects = Point.import_wkt(wkt="POINT(-97.7323317 30.2830764)", epsg=4326)
# The `set_observed` method allows for making sql-like queries on the observed field and the
# LTE is an enum that means less than or equal to the value in the query field
#
# This Query is for data from August 25, 2019 UTC or earlier
request.set_observed(rel_type=enum.FilterRelationship.LTE, value=date(2019, 8, 25))
# search_one_ex method requests only one item be returned that meets the query filters in the StacRequestWrap
# the item returned is a wrapper of the protobuf message; StacItemWrap. search_one_ex, will only return the most
# recently observed results that matches the time filter and spatial filter
stac_item = client.search_one_ex(request)
# get the thumbnail asset from the assets map. The other option would be a Geotiff,
# with asset key 'GEOTIFF_RGB'
asset_wrap = stac_item.get_asset(asset_type=enum.AssetType.THUMBNAIL)
print(asset_wrap)
# uncomment to display image
# with tempfile.TemporaryDirectory() as d:
# filename = utils.download_asset(asset=asset, save_directory=d)
# display(Image(filename=filename))
```
## Simple Query and the Makeup of a StacItem
The easiest query to construct is a `StacRequestWrap` constructor with no variables, and the next simplest, is the case where we know the STAC item `id` that we want to search. If we already know the STAC `id` of an item, we can construct the `StacRequestWrap` as follows:
```
from nsl.stac.experimental import NSLClientEx, StacRequestWrap
# get a client interface to the gRPC channel
client_ex = NSLClientEx()
# create a request for a specific STAC item
request = StacRequestWrap(id='20190822T183518Z_746_POM1_ST2_P')
# for this request we might as well use the search one, as STAC ids ought to be unique
stac_item = client.search_one_ex(request)
print(stac_item)
```
## Spatial Queries
### Query by Bounds
You can query for STAC items intersecting a bounding box of minx, miny, maxx, and maxy. An [epsg](https://en.wikipedia.org/wiki/EPSG_Geodetic_Parameter_Dataset) code is required (4326 is the most common epsg code used for longitude and latitude). Remember, this finds STAC items that intersect the bounds, **not** STAC items contained by the bounds.
```
from nsl.stac.experimental import StacRequestWrap, NSLClientEx
# get a client interface to the gRPC channel
client_ex = NSLClientEx()
# request wrapper
request = StacRequestWrap()
# define our area of interest bounds using the xmin, ymin, xmax, ymax coordinates of an area on
# the WGS-84 ellipsoid
neighborhood_box = (-97.7352547645, 30.27526474757116, -97.7195692, 30.28532)
# setting the bounds tests for intersection (not contains)
request.set_bounds(neighborhood_box, epsg=4326)
request.limit = 3
# Search for data that intersects the bounding box
epsg_4326_ids = []
for stac_item in client_ex.search_ex(request):
print("STAC item id: {}".format(stac_item.id))
print("bounds:")
print(stac_item.geometry.bounds)
print("bbox (EnvelopeData protobuf):")
print(stac_item.bbox)
print("geometry:")
print(stac_item.geometry)
epsg_4326_ids.append(stac_item.id)
```
### Query by Bounds; Projection Support
Querying using geometries defined in a different projection requires defining the epsg number for the spatial reference of the data. In this example we use an epsg code for a [UTM projection](https://epsg.io/3744).
Notice that the results below are the same as the cell above (look at the `TEST-->` section of the printout)
```
from nsl.stac.experimental import StacRequestWrap, NSLClientEx
# get a client interface to the gRPC channel
client_ex = NSLClientEx()
# request wrapper
request = StacRequestWrap()
# define our area of interest bounds using the xmin, ymin, xmax, ymax coordinates of in UTM 14N in NAD83 (epsg 3744)
neighborhood_box = (621636.1875228449, 3349964.520449501, 623157.4212553708, 3351095.8075163467)
# setting the bounds tests for intersection (not contains)
request.set_bounds(neighborhood_box, epsg=3744)
request.limit = 3
# Search for data that intersects the bounding box
for stac_item in client_ex.search_ex(request):
print("STAC item id: {}".format(stac_item.id))
print("bounds:")
print(stac_item.geometry.bounds)
print("bbox (EnvelopeData protobuf):")
print(stac_item.bbox)
print("geometry:")
print(stac_item.geometry)
print("TEST RESULT '{1}': stac_item id {0} from 3744 bounds is in the set of the 4326 bounds search results.".format(stac_item.id, stac_item.id in epsg_4326_ids))
print()
```
### Query by GeoJSON
Use a GeoJSON geometry to define the `intersects` property.
```
import json
import requests
from epl.geometry import shape as epl_shape
from nsl.stac.experimental import StacRequestWrap, NSLClientEx
# get a client interface to the gRPC channel
client_ex = NSLClientEx()
request = StacRequestWrap()
# retrieve a coarse geojson foot print of Travis County, Texas
r = requests.get("http://raw.githubusercontent.com/johan/world.geo.json/master/countries/USA/TX/Travis.geo.json")
travis_shape = epl_shape(r.json()['features'][0]['geometry'], epsg=4326)
# search for any data that intersects the travis county geometry
request.intersects = travis_shape
# limit results to 2 (instead of default of 10)
request.limit = 2
geojson_ids = []
# get a client interface to the gRPC channel
for stac_item in client_ex.search_ex(request):
print("STAC item id: {}".format(stac_item.id))
print("Stac item observed: {}".format(stac_item.observed))
geojson_ids.append(stac_item.id)
```
### Query by WKT
Use a [WKT](https://en.wikipedia.org/wiki/Well-known_text_representation_of_geometry) geometry to define the `intersects` property.
```
from epl.geometry import Polygon
# Same geometry as above, but a wkt geometry instead of a geojson
travis_wkt = "POLYGON((-97.9736 30.6251, -97.9188 30.6032, -97.9243 30.5703, -97.8695 30.5484, \
-97.8476 30.4717, -97.7764 30.4279, -97.5793 30.4991, -97.3711 30.4170, \
-97.4916 30.2089, -97.6505 30.0719, -97.6669 30.0665, -97.7107 30.0226, \
-98.1708 30.3567, -98.1270 30.4279, -98.0503 30.6251, -97.9736 30.6251))"
request.intersects = Polygon.import_wkt(wkt=travis_wkt, epsg=4326)
request.limit = 2
for stac_item in client_ex.search_ex(request):
print("STAC item id: {0} from wkt filter intersects result from geojson filter: {1}"
.format(stac_item.id, stac_item.id in geojson_ids))
```
## Temporal Queries
When it comes to Temporal queries there are a few things to note.
- we assume all dates are UTC unless specified
- datetime and date are treated differently in different situations
- date for EQ and NEQ is a 24 hour period
- datetime for EQ and NEQ is almost useless (it is a timestamp accurate down to the nanosecond)
- date for GTE or LT is defined as the date from the first nanosecond of that date
- date for LTE or GT is defined as the date at the final nanosecond of that date
- date for start and end with BETWEEN has a start with minimum time and an end with the max time. same for NOT_BETWEEN
- datetime for GTE, GT, LTE, LT, BETWEEN and NOT_BETWEEN is interpreted strictly according to the nanosecond of that datetime definition(s) provided
When creating a time query filter, we want to use the >, >=, <, <=, ==, != operations and inclusive and exclusive range requests. We do this by using the `set_observed` method and the `value` set to a date/datetime combined with `rel_typ` set to `GTE`,`GT`, `LTE`, `LT`, `EQ`, or `NEQ`. If we use `rel_type` set to `BETWEEN` OR `NOT_BETWEEN` then we must set the `start` **and** the `end` variables.
### Everything After A Secific Date
```
from datetime import date, datetime, timezone
from nsl.stac.experimental import NSLClientEx, StacRequestWrap
from nsl.stac import utils, enum
# get a client interface to the gRPC channel
client_ex = NSLClientEx()
request = StacRequestWrap()
# make a filter that selects all data on or after August 21st, 2019
request.set_observed(rel_type=enum.FilterRelationship.GTE, value=date(2019, 8, 21))
request.limit = 2
demonstration_datetime = datetime.combine(date(2019, 8, 21), datetime.min.time())
for stac_item in client_ex.search_ex(request):
print("STAC item date, {0}, is after {1}: {2}".format(
stac_item.observed,
demonstration_datetime,
datetime.timestamp(stac_item.observed) > datetime.timestamp(demonstration_datetime)))
```
#### Everything Between Two Dates
Now we're going to do a range request and select data between two dates using the `start` and `end` parameters instead of the `value` parameter:
```
from datetime import datetime, timezone, timedelta
from nsl.stac.client import NSLClient
from nsl.stac import utils, enum, StacRequest
# Query data from August 1, 2019
start = datetime(2019, 8, 1, 0, 0, 0, tzinfo=timezone.utc)
# ... up until August 10, 2019
end = start + timedelta(days=9)
request.set_observed(rel_type=enum.FilterRelationship.BETWEEN, start=start, end=end)
request.limit = 2
# get a client interface to the gRPC channel
client_ex = NSLClientEx()
for stac_item in client_ex.search_ex(request):
print("STAC item date, {0}, is between {1} and {2}".format(
stac_item.observed,
datetime.combine(start, datetime.min.time()),
datetime.combine(end, datetime.min.time())))
```
In the above print out we are returned STAC items that are between the dates of Aug 1 2019 and Aug 10 2019. Also, notice there's no warnings as we defined our utc timezone on the datetime objects.
#### Select Data for One Day
Now we'll search for everything on a specific day using a python `datetime.date` for the `value` and `rel_type` set to use equals (`FilterRelationship.EQ`). Python's `datetime.datetime` is a specific value and if you use it combined with `EQ` the query would insist that the time relationship match down to the second. But since `datetime.date` is only specific down to the day, the filter is created for the entire day. This will check for everything from the start until the end of the 8th of August, specifically in the Austin, Texas timezone (UTC -6).
```
from datetime import datetime, timezone, timedelta, date
from nsl.stac.experimental import NSLClientEx, StacItemWrap
from nsl.stac import utils, enum
request = StacRequestWrap()
texas_utc_offset = timezone(timedelta(hours=-6))
value = date(2019, 8, 6)
# Query all data for the entire day of August 6, 2019
request.set_observed(rel_type=enum.FilterRelationship.EQ, value=value, tzinfo=texas_utc_offset)
request.limit = 2
# get a client interface to the gRPC channel
client_ex = NSLClientEx()
for stac_item in client.search_ex(request):
print(datetime.fromtimestamp(stac_item.observed.timestamp(), tz=timezone.utc))
```
## Downloading from AssetWrap
```
import os
import tempfile
from IPython.display import Image, display
from epl.geometry import LineString
from nsl.stac.experimental import NSLClientEx, StacRequestWrap
from nsl.stac import utils, enum
request = StacRequestWrap()
request.intersects = LineString.import_wkt(wkt='LINESTRING(622301.8284206488 3350344.236542711, 622973.3950196661 3350466.792693002)',
epsg=3744)
request.set_observed(value=date(2019, 8, 25), rel_type=enum.FilterRelationship.LTE)
request.limit = 3
client_ex = NSLClientEx()
for stac_item in client_ex.search_ex(request):
# get the thumbnail asset from the assets map
asset_wrap = stac_item.get_asset(asset_type=enum.AssetType.THUMBNAIL)
print(asset_wrap)
print()
# (side-note delete=False in NamedTemporaryFile is only required for windows.)
with tempfile.NamedTemporaryFile(suffix=".png", delete=False) as file_obj:
asset_wrap.download(file_obj=file_obj)
print("downloaded file {}".format(os.path.basename(asset_wrap.object_path)))
print()
# uncomment to display
# display(Image(filename=file_obj.name))
```
## View
For our ground sampling distance query we're using another query filter; this time it's the [FloatFilter](https://geo-grpc.github.io/api/#epl.protobuf.v1.FloatFilter). It behaves just as the TimestampFilter, but with floats for `value` or for `start` + `end`.
In order to make our off nadir query we need to insert it inside of an [ViewRequest](https://geo-grpc.github.io/api/#epl.protobuf.v1.ViewRequest) container and set that to the `view` field of the `StacRequest`.
```
from datetime import datetime, timezone
from epl.geometry import Point
from nsl.stac.experimental import NSLClientEx, StacRequestWrap
from nsl.stac.enum import FilterRelationship, Mission
request = StacRequestWrap()
# create our off_nadir query to only return data captured with an angle of less than or
# equal to 10 degrees
request.set_off_nadir(rel_type=FilterRelationship.GTE, value=30.0)
request.set_gsd(rel_type=FilterRelationship.LT, value=1.0)
# define ourselves a point in Texas
request.intersects = Point.import_wkt("POINT(621920.1090935947 3350833.389847579)", epsg=26914)
# the above could also be defined using longitude and latitude as follows:
# request.intersects = Point.import_wkt("POINT(-97.7323317 30.2830764)", epsg=4326)
# create a StacRequest with geometry, gsd, and off nadir and a limit of 4
request.limit = 4
# get a client interface to the gRPC channel
client_ex = NSLClientEx()
for stac_item in client_ex.search_ex(request):
print("{0} STAC item '{1}' from {2}\nhas a off_nadir\t{3:.2f}, which should be greater than or "
"equal to requested off_nadir\t{4:.3f} (confirmed {5})".format(
stac_item.mission.name,
stac_item.id,
stac_item.observed,
stac_item.off_nadir,
request.stac_request.view.off_nadir.value,
request.stac_request.view.off_nadir.value < stac_item.off_nadir))
print("has a gsd\t{0:.3f}, which should be less than "
"the requested\t\t gsd\t\t{1:.3f} (confirmed {2})".format(
stac_item.gsd,
request.stac_request.gsd.value,
request.stac_request.gsd.value < stac_item.gsd))
```
## Shapely Geometry
In order to extract a shapely geometry from the STAC item geometry use the `shapely_dump` method.
```
from epl.geometry import LineString
from nsl.stac.experimental import NSLClientEx, StacRequestWrap
from nsl.stac import utils, enum
request = StacRequestWrap()
request.intersects = LineString.import_wkt(wkt='LINESTRING(-97.72842049283962 30.278624772098176,-97.72142529172878 30.2796624743974)',
epsg=4326)
request.set_observed(value=date(2019, 8, 25), rel_type=enum.FilterRelationship.LTE)
request.limit = 10
client_ex = NSLClientEx()
unioned = None
for stac_item in client_ex.search_ex(request):
if unioned is None:
unioned = stac_item.geometry.shapely_dump
else:
# execute shapely union
unioned = unioned.union(stac_item.geometry.shapely_dump)
print(unioned)
```
## Enum Classes
There are a number of different enum classes used for both STAC requests and STAC items.
```
from nsl.stac import enum
import inspect
[m for m in inspect.getmembers(enum) if not m[0].startswith('_') and m[0][0].isupper() and m[0] != 'IntFlag']
# Specific to Queries
print("for defining the sort direction of a query")
for s in enum.SortDirection:
print(s.name)
print("for defining the query relationship with a value (EQ, LTE, GTE, LT, GT, NEQ), a start-end range \n(BETWEEN, NOT_BETWEEN), a set (IN, NOT_IN) or a string (LIKE, NOT_LIKE). To be noted, EQ is the default relationship type")
for f in enum.FilterRelationship:
print(f.name)
# Specific to Assets
print("these can be useful when getting a specific Asset from a STAC item by the type of Asset")
for a in enum.AssetType:
print(a.name)
# STAC Item details
print("Mission, Platform and Instrument are aspects of data that can be used in queries.\nBut as NSL currently only has one platform and one instrument, these may not be useful")
print("missions:")
for a in enum.Mission:
print(a.name)
print("\nplatforms:")
for a in enum.Platform:
print(a.name)
print("\ninstruments:")
for a in enum.Instrument:
print(a.name)
```
| github_jupyter |
## Dataset: Labeled Faces in the Wild ##
## Experiment: (experiment_1) Image based gender classification ##
```
import torch
import torchvision
import torch.nn as nn
from torch.utils.data import Dataset, DataLoader
import torch.optim as optim
from torchvision import transforms
import torch.nn.functional as F
from torch.utils.tensorboard import SummaryWriter
import pandas as pd
import os
import numpy as np
# import cv2
from PIL import Image
import matplotlib.pyplot as plt
from tqdm import tqdm_notebook
%matplotlib inline
```
### Parameters ###
```
epochs = 40
batch_size = 64
learning_rate = 0.001
# momentum = 0.5
num_workers = 2
data_path = '../data/lfw/data/'
# default `log_dir` is "runs" - we'll be more specific here
!rm -rf ./runs/experiment_1
writer = SummaryWriter('runs/experiment_1')
```
### Dataset class used to instanciate the data loader ###
```
class LFWDataset(Dataset):
"""Face Landmarks dataset."""
def __init__(self, data_path, attributes_df, transform=None):
self.attributes_df = attributes_df
self.data_path = data_path
self.transform = transform
def __len__(self):
return len(self.attributes_df)
def __getitem__(self, idx):
img_path = os.path.join(self.data_path, "lfw_home/lfw_funneled", self.attributes_df.iloc[idx]['person'].replace(' ', '_'),"{}_{:04d}.jpg".format(self.attributes_df.iloc[idx]['person'].replace(' ', '_'),self.attributes_df.iloc[idx]['imagenum']))
# img = torch.from_numpy(cv2.imread(img_path))
img = Image.open(img_path, mode='r')
label = self.attributes_df.iloc[idx]['Male']>0
if self.transform:
img = self.transform(img)
return img, torch.tensor(label, dtype=torch.float)
```
### Creating datasets ###
```
# read file
attributes_df = pd.read_csv(os.path.join(data_path,'lfw_attributes.txt'))
# split data into training val and test
all_names = attributes_df.person.unique()
tt_msk = np.random.rand(len(all_names)) < 0.8
temp_train_names = all_names[tt_msk]
tv_msk = np.random.rand(len(temp_train_names)) < 0.8
train_names = temp_train_names[tv_msk]
val_names = temp_train_names[~tv_msk]
test_names = all_names[~tt_msk]
del all_names, tt_msk, temp_train_names, tv_msk
# create train val and test dataframes
train_df = attributes_df.loc[attributes_df['person'].isin(train_names)]
val_df = attributes_df.loc[attributes_df['person'].isin(val_names)]
test_df = attributes_df.loc[attributes_df['person'].isin(test_names)]
del attributes_df, train_names, val_names, test_names
# create datasets
train_dataset = LFWDataset(data_path, train_df, transform=transforms.Compose([
# transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor()
]))
val_dataset = LFWDataset(data_path, val_df, transform=transforms.Compose([
# transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor()]))
test_dataset = LFWDataset(data_path, test_df, transform=transforms.Compose([
# transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor()]))
del train_df, val_df, test_df
print(len(train_dataset), len(val_dataset), len(test_dataset))
```
### Declaring dataloaders ###
```
train_dataloader = DataLoader(train_dataset, batch_size=batch_size, shuffle=True, num_workers=num_workers)
val_dataloader = DataLoader(val_dataset, batch_size=batch_size, shuffle=True, num_workers=num_workers)
test_dataloader = DataLoader(test_dataset, batch_size=batch_size, shuffle=True, num_workers=num_workers)
print(len(train_dataloader), len(val_dataloader), len(test_dataloader))
```
### Sanity check ###
```
# for i , (img, label) in enumerate(train_dataloader):
# for im, lab in zip(img, label):
# plt.imshow(np.moveaxis(np.asarray(im), 0, -1))
# plt.show()
# print(lab)
# if i>2:
# break
```
### Define Model ###
```
class ResNet(nn.Module):
def __init__(self):
super(ResNet, self).__init__()
self.conv1 = nn.Conv2d(3, 8, 5)
self.pool1 = nn.MaxPool2d(4,4)
self.conv2 = nn.Conv2d(8, 16, 5)
self.pool2 = nn.MaxPool2d(4,4)
self.conv3 = nn.Conv2d(16, 32, 5)
self.pool3 = nn.MaxPool2d(2,2)
# self.conv4 = nn.Conv2d(32, 32, 5)
self.fc1 = nn.Linear(800, 512)
self.fc2 = nn.Linear(512, 64)
self.fc3 = nn.Linear(64, 1)
self.dropout_layer1 = nn.Dropout(p=0.6)
self.dropout_layer2 = nn.Dropout(p=0.5)
# self.dropout_layer3 = nn.Dropout(p=0.2)
def forward(self, x):
x = self.pool1(F.relu(self.conv1(x)))
x = self.pool2(F.relu(self.conv2(x)))
x = self.pool3(F.relu(self.conv3(x)))
# x = F.relu(self.conv4(x))
x = x.view(x.shape[0],-1)
x = self.dropout_layer1(x)
x = F.relu(self.fc1(x))
x = self.dropout_layer2(x)
# x = self.dropout_layer3(x)
x = F.relu(self.fc2(x))
x = F.sigmoid(self.fc3(x))
return x
model = ResNet().cuda()
# model = ResNet()
print(model)
```
### All layers except the last are frozen ###
```
# print(model)
# def unfreeze_layer4():
# for p in model.rnet.layer4.parameters():
# p.requires_grad = True
# def unfreeze_layer3():
# for p in model.rnet.layer3.parameters():
# p.requires_grad = True
# def unfreeze_layer2():
# for p in model.rnet.layer2.parameters():
# p.requires_grad = True
# def unfreeze_layer1():
# for p in model.rnet.layer1.parameters():
# p.requires_grad = True
# for p in model.parameters():
# p.requires_grad = False
# for p in model.rnet.fc.parameters():
# p.requires_grad = True
# for p in model.parameters():
# print(p.requires_grad)
# unfreeze_layer4()
```
### Define training and validation functions ###
```
optimizer = optim.Adam(model.parameters(), lr=learning_rate)
def perform_evaluation(val_model, dataloader):
with torch.no_grad():
epoch_loss = 0
epoch_accuracy = 0
for batch_idx, (data, target) in tqdm_notebook(enumerate(dataloader), total=len(dataloader)):
# move data batch to GPU
data = data.cuda()
target = target.cuda()
# forward pass
output = val_model(data)
loss = F.binary_cross_entropy(output, target.unsqueeze(1))
# compute average loss an accuracy
output = output.to('cpu')
target = target.to('cpu')
current_acc = torch.tensor(((output>0.5)== torch.tensor(target.unsqueeze(1), dtype=torch.bool)).sum(), dtype=torch.float)/torch.tensor(len(target), dtype=torch.float)
epoch_loss = ((epoch_loss*batch_idx) + loss.item())/(batch_idx+1)
epoch_accuracy = ((epoch_accuracy*batch_idx) + current_acc.item())/(batch_idx+1)
print("testing loss: {} and testing accuracy: {}".format(epoch_loss, epoch_accuracy))
return epoch_loss, epoch_accuracy
def perform_validation(val_model, dataloader):
with torch.no_grad():
epoch_loss = 0
epoch_accuracy = 0
for batch_idx, (data, target) in tqdm_notebook(enumerate(dataloader), total=len(dataloader)):
# move data batch to GPU
data = data.cuda()
target = target.cuda()
# forward pass
output = val_model(data)
loss = F.binary_cross_entropy(output, target.unsqueeze(1))
# compute average loss an accuracy
output = output.to('cpu')
target = target.to('cpu')
current_acc = torch.tensor(((output>0.5)== torch.tensor(target.unsqueeze(1), dtype=torch.bool)).sum(), dtype=torch.float)/torch.tensor(len(target), dtype=torch.float)
epoch_loss = ((epoch_loss*batch_idx) + loss.item())/(batch_idx+1)
epoch_accuracy = ((epoch_accuracy*batch_idx) + current_acc.item())/(batch_idx+1)
print("val loss: {} and val accuracy: {}".format(epoch_loss, epoch_accuracy))
return epoch_loss, epoch_accuracy
def perform_training(val_model, dataloader):
epoch_loss = 0
epoch_accuracy = 0
for batch_idx, (data, target) in tqdm_notebook(enumerate(dataloader), total=len(dataloader)):
# move data batch to GPU
data = data.cuda()
target = target.cuda()
# zero the parameter gradients
optimizer.zero_grad()
# forward pass
output = val_model(data)
# print(output)
loss = F.binary_cross_entropy(output, target.unsqueeze(1))
# backward pass
loss.backward()
optimizer.step()
# compute average loss an accuracy
output = output.to('cpu')
target = target.to('cpu')
current_acc = torch.tensor(((output>0.5)== torch.tensor(target.unsqueeze(1), dtype=torch.bool)).sum(), dtype=torch.float)/torch.tensor(len(target), dtype=torch.float)
epoch_loss = ((epoch_loss*batch_idx) + loss.item())/(batch_idx+1)
epoch_accuracy = ((epoch_accuracy*batch_idx) + current_acc.item())/(batch_idx+1)
print("train loss: {} and train accuracy: {}".format(epoch_loss, epoch_accuracy))
return epoch_loss, epoch_accuracy
```
### Initiate training ###
```
train_losses=[]
train_accuracies=[]
val_losses=[]
val_accuracies=[]
max_val_acc = 0
for epoch in range(epochs):
# run train and val epochs
print("Epoch: {}".format(epoch))
model.train()
train_loss, train_acc = perform_training(model, train_dataloader)
train_losses.append(train_loss)
train_accuracies.append(train_acc)
writer.add_scalar('training loss', train_loss, epoch)
writer.add_scalar('training accuracy', train_acc, epoch)
model.eval()
val_loss, val_acc = perform_validation(model, val_dataloader)
val_losses.append(val_loss)
val_accuracies.append(val_acc)
writer.add_scalar('validation loss', val_loss, epoch)
writer.add_scalar('validation accuracy', val_acc, epoch)
if val_acc > max_val_acc:
print("saving model on epoch {}".format(epoch))
torch.save(model.state_dict(), "models/best_model.pt")
writer.close()
```
### Evaluate the model ###
```
# evaluate final model
model.eval()
test_loss, test_acc = perform_evaluation(model, test_dataloader)
# evaluate best model
eval_model = ResNet().cuda()
eval_model.load_state_dict(torch.load("models/best_model.pt"))
eval_model.eval()
best_test_loss, best_test_acc = perform_evaluation(eval_model, test_dataloader)
print("final model loss: {}, final model accuracy: {}, best model loss: {}, best model accuracy: {}".format(test_loss, test_acc, best_test_loss, best_test_acc))
torch.save(model.state_dict(), "final_model.pt")
```
| github_jupyter |
Deep Learning Models -- A collection of various deep learning architectures, models, and tips for TensorFlow and PyTorch in Jupyter Notebooks.
- Author: Sebastian Raschka
- GitHub Repository: https://github.com/rasbt/deeplearning-models
# Model Zoo -- Simple RNN
Demo of a simple RNN for sentiment classification (here: a binary classification problem with two labels, positive and negative). Note that a simple RNN usually doesn't work very well due to vanishing and exploding gradient problems. Also, this implementation uses padding for dealing with variable size inputs. Hence, the shorter the sentence, the more `<pad>` placeholders will be added to match the length of the longest sentence in a batch.
Note that this RNN trains about 4 times slower than the equivalent with packed sequences, [./rnn-simple-packed-imdb.ipynb](./rnn-simple-packed-imdb.ipynb).
```
%load_ext watermark
%watermark -a 'Sebastian Raschka' -v -p torch
import torch
import torch.nn.functional as F
from torchtext import data
from torchtext import datasets
import time
import random
torch.backends.cudnn.deterministic = True
```
## General Settings
```
RANDOM_SEED = 123
torch.manual_seed(RANDOM_SEED)
VOCABULARY_SIZE = 20000
LEARNING_RATE = 1e-4
BATCH_SIZE = 128
NUM_EPOCHS = 15
DEVICE = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
EMBEDDING_DIM = 128
HIDDEN_DIM = 256
OUTPUT_DIM = 1
```
## Dataset
Load the IMDB Movie Review dataset:
```
TEXT = data.Field(tokenize = 'spacy')
LABEL = data.LabelField(dtype = torch.float)
train_data, test_data = datasets.IMDB.splits(TEXT, LABEL)
train_data, valid_data = train_data.split(random_state=random.seed(RANDOM_SEED),
split_ratio=0.8)
print(f'Num Train: {len(train_data)}')
print(f'Num Valid: {len(valid_data)}')
print(f'Num Test: {len(test_data)}')
```
Build the vocabulary based on the top "VOCABULARY_SIZE" words:
```
TEXT.build_vocab(train_data, max_size=VOCABULARY_SIZE)
LABEL.build_vocab(train_data)
print(f'Vocabulary size: {len(TEXT.vocab)}')
print(f'Number of classes: {len(LABEL.vocab)}')
```
The TEXT.vocab dictionary will contain the word counts and indices. The reason why the number of words is VOCABULARY_SIZE + 2 is that it contains to special tokens for padding and unknown words: `<unk>` and `<pad>`.
Make dataset iterators:
```
train_loader, valid_loader, test_loader = data.BucketIterator.splits(
(train_data, valid_data, test_data),
batch_size=BATCH_SIZE,
device=DEVICE)
```
Testing the iterators (note that the number of rows depends on the longest document in the respective batch):
```
print('Train')
for batch in train_loader:
print(f'Text matrix size: {batch.text.size()}')
print(f'Target vector size: {batch.label.size()}')
break
print('\nValid:')
for batch in valid_loader:
print(f'Text matrix size: {batch.text.size()}')
print(f'Target vector size: {batch.label.size()}')
break
print('\nTest:')
for batch in test_loader:
print(f'Text matrix size: {batch.text.size()}')
print(f'Target vector size: {batch.label.size()}')
break
```
## Model
```
import torch.nn as nn
class RNN(nn.Module):
def __init__(self, input_dim, embedding_dim, hidden_dim, output_dim):
super().__init__()
self.embedding = nn.Embedding(input_dim, embedding_dim)
self.rnn = nn.RNN(embedding_dim, hidden_dim)
self.fc = nn.Linear(hidden_dim, output_dim)
def forward(self, text):
#[sentence len, batch size] => [sentence len, batch size, embedding size]
embedded = self.embedding(text)
#[sentence len, batch size, embedding size] =>
# output: [sentence len, batch size, hidden size]
# hidden: [1, batch size, hidden size]
output, hidden = self.rnn(embedded)
return self.fc(hidden.squeeze(0)).view(-1)
INPUT_DIM = len(TEXT.vocab)
EMBEDDING_DIM = 64
HIDDEN_DIM = 128
OUTPUT_DIM = 1
torch.manual_seed(RANDOM_SEED)
model = RNN(INPUT_DIM, EMBEDDING_DIM, HIDDEN_DIM, OUTPUT_DIM)
model = model.to(DEVICE)
optimizer = torch.optim.Adam(model.parameters(), lr=LEARNING_RATE)
```
## Training
```
def compute_binary_accuracy(model, data_loader, device):
model.eval()
correct_pred, num_examples = 0, 0
with torch.no_grad():
for batch_idx, batch_data in enumerate(data_loader):
logits = model(batch_data.text)
predicted_labels = (torch.sigmoid(logits) > 0.5).long()
num_examples += batch_data.label.size(0)
correct_pred += (predicted_labels == batch_data.label.long()).sum()
return correct_pred.float()/num_examples * 100
start_time = time.time()
for epoch in range(NUM_EPOCHS):
model.train()
for batch_idx, batch_data in enumerate(train_loader):
### FORWARD AND BACK PROP
logits = model(batch_data.text)
cost = F.binary_cross_entropy_with_logits(logits, batch_data.label)
optimizer.zero_grad()
cost.backward()
### UPDATE MODEL PARAMETERS
optimizer.step()
### LOGGING
if not batch_idx % 50:
print (f'Epoch: {epoch+1:03d}/{NUM_EPOCHS:03d} | '
f'Batch {batch_idx:03d}/{len(train_loader):03d} | '
f'Cost: {cost:.4f}')
with torch.set_grad_enabled(False):
print(f'training accuracy: '
f'{compute_binary_accuracy(model, train_loader, DEVICE):.2f}%'
f'\nvalid accuracy: '
f'{compute_binary_accuracy(model, valid_loader, DEVICE):.2f}%')
print(f'Time elapsed: {(time.time() - start_time)/60:.2f} min')
print(f'Total Training Time: {(time.time() - start_time)/60:.2f} min')
print(f'Test accuracy: {compute_binary_accuracy(model, test_loader, DEVICE):.2f}%')
import spacy
nlp = spacy.load('en')
def predict_sentiment(model, sentence):
# based on:
# https://github.com/bentrevett/pytorch-sentiment-analysis/blob/
# master/2%20-%20Upgraded%20Sentiment%20Analysis.ipynb
model.eval()
tokenized = [tok.text for tok in nlp.tokenizer(sentence)]
indexed = [TEXT.vocab.stoi[t] for t in tokenized]
tensor = torch.LongTensor(indexed).to(DEVICE)
tensor = tensor.unsqueeze(1)
prediction = torch.sigmoid(model(tensor))
return prediction.item()
print('Probability positive:')
predict_sentiment(model, "I really love this movie. This movie is so great!")
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Build a linear model with Estimators
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r1/tutorials/estimators/linear.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r1/tutorials/estimators/linear.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
> Note: This is an archived TF1 notebook. These are configured
to run in TF2's
[compatbility mode](https://www.tensorflow.org/guide/migrate)
but will run in TF1 as well. To use TF1 in Colab, use the
[%tensorflow_version 1.x](https://colab.research.google.com/notebooks/tensorflow_version.ipynb)
magic.
This tutorial uses the `tf.estimator` API in TensorFlow to solve a benchmark binary classification problem. Estimators are TensorFlow's most scalable and production-oriented model type. For more information see the [Estimator guide](https://www.tensorflow.org/r1/guide/estimators).
## Overview
Using census data which contains data about a person's age, education, marital status, and occupation (the *features*), we will try to predict whether or not the person earns more than 50,000 dollars a year (the target *label*). We will train a *logistic regression* model that, given an individual's information, outputs a number between 0 and 1—this can be interpreted as the probability that the individual has an annual income of over 50,000 dollars.
Key Point: As a modeler and developer, think about how this data is used and the potential benefits and harm a model's predictions can cause. A model like this could reinforce societal biases and disparities. Is each feature relevant to the problem you want to solve or will it introduce bias? For more information, read about [ML fairness](https://developers.google.com/machine-learning/fairness-overview/).
## Setup
Import TensorFlow, feature column support, and supporting modules:
```
import tensorflow.compat.v1 as tf
import tensorflow.feature_column as fc
import os
import sys
import matplotlib.pyplot as plt
from IPython.display import clear_output
```
And let's enable [eager execution](https://www.tensorflow.org/r1/guide/eager) to inspect this program as we run it:
## Download the official implementation
We'll use the [wide and deep model](https://github.com/tensorflow/models/tree/master/official/r1/wide_deep/) available in TensorFlow's [model repository](https://github.com/tensorflow/models/). Download the code, add the root directory to your Python path, and jump to the `wide_deep` directory:
```
! pip install requests
! git clone --depth 1 https://github.com/tensorflow/models
```
Add the root directory of the repository to your Python path:
```
models_path = os.path.join(os.getcwd(), 'models')
sys.path.append(models_path)
```
Download the dataset:
```
from official.r1.wide_deep import census_dataset
from official.r1.wide_deep import census_main
census_dataset.download("/tmp/census_data/")
```
### Command line usage
The repo includes a complete program for experimenting with this type of model.
To execute the tutorial code from the command line first add the path to tensorflow/models to your `PYTHONPATH`.
```
#export PYTHONPATH=${PYTHONPATH}:"$(pwd)/models"
#running from python you need to set the `os.environ` or the subprocess will not see the directory.
if "PYTHONPATH" in os.environ:
os.environ['PYTHONPATH'] += os.pathsep + models_path
else:
os.environ['PYTHONPATH'] = models_path
```
Use `--help` to see what command line options are available:
```
!python -m official.r1.wide_deep.census_main --help
```
Now run the model:
```
!python -m official.r1.wide_deep.census_main --model_type=wide --train_epochs=2
```
## Read the U.S. Census data
This example uses the [U.S Census Income Dataset](https://archive.ics.uci.edu/ml/datasets/Census+Income) from 1994 and 1995. We have provided the [census_dataset.py](https://github.com/tensorflow/models/tree/master/official/r1/wide_deep/census_dataset.py) script to download the data and perform a little cleanup.
Since the task is a *binary classification problem*, we'll construct a label column named "label" whose value is 1 if the income is over 50K, and 0 otherwise. For reference, see the `input_fn` in [census_main.py](https://github.com/tensorflow/models/tree/master/official/r1/wide_deep/census_main.py).
Let's look at the data to see which columns we can use to predict the target label:
```
!ls /tmp/census_data/
train_file = "/tmp/census_data/adult.data"
test_file = "/tmp/census_data/adult.test"
```
[pandas](https://pandas.pydata.org/) provides some convenient utilities for data analysis. Here's a list of columns available in the Census Income dataset:
```
import pandas
train_df = pandas.read_csv(train_file, header = None, names = census_dataset._CSV_COLUMNS)
test_df = pandas.read_csv(test_file, header = None, names = census_dataset._CSV_COLUMNS)
train_df.head()
```
The columns are grouped into two types: *categorical* and *continuous* columns:
* A column is called *categorical* if its value can only be one of the categories in a finite set. For example, the relationship status of a person (wife, husband, unmarried, etc.) or the education level (high school, college, etc.) are categorical columns.
* A column is called *continuous* if its value can be any numerical value in a continuous range. For example, the capital gain of a person (e.g. $14,084) is a continuous column.
## Converting Data into Tensors
When building a `tf.estimator` model, the input data is specified by using an *input function* (or `input_fn`). This builder function returns a `tf.data.Dataset` of batches of `(features-dict, label)` pairs. It is not called until it is passed to `tf.estimator.Estimator` methods such as `train` and `evaluate`.
The input builder function returns the following pair:
1. `features`: A dict from feature names to `Tensors` or `SparseTensors` containing batches of features.
2. `labels`: A `Tensor` containing batches of labels.
The keys of the `features` are used to configure the model's input layer.
Note: The input function is called while constructing the TensorFlow graph, *not* while running the graph. It is returning a representation of the input data as a sequence of TensorFlow graph operations.
For small problems like this, it's easy to make a `tf.data.Dataset` by slicing the `pandas.DataFrame`:
```
def easy_input_function(df, label_key, num_epochs, shuffle, batch_size):
label = df[label_key]
ds = tf.data.Dataset.from_tensor_slices((dict(df),label))
if shuffle:
ds = ds.shuffle(10000)
ds = ds.batch(batch_size).repeat(num_epochs)
return ds
```
Since we have eager execution enabled, it's easy to inspect the resulting dataset:
```
ds = easy_input_function(train_df, label_key='income_bracket', num_epochs=5, shuffle=True, batch_size=10)
for feature_batch, label_batch in ds.take(1):
print('Some feature keys:', list(feature_batch.keys())[:5])
print()
print('A batch of Ages :', feature_batch['age'])
print()
print('A batch of Labels:', label_batch )
```
But this approach has severly-limited scalability. Larger datasets should be streamed from disk. The `census_dataset.input_fn` provides an example of how to do this using `tf.decode_csv` and `tf.data.TextLineDataset`:
<!-- TODO(markdaoust): This `input_fn` should use `tf.contrib.data.make_csv_dataset` -->
```
import inspect
print(inspect.getsource(census_dataset.input_fn))
```
This `input_fn` returns equivalent output:
```
ds = census_dataset.input_fn(train_file, num_epochs=5, shuffle=True, batch_size=10)
for feature_batch, label_batch in ds.take(1):
print('Feature keys:', list(feature_batch.keys())[:5])
print()
print('Age batch :', feature_batch['age'])
print()
print('Label batch :', label_batch )
```
Because `Estimators` expect an `input_fn` that takes no arguments, we typically wrap configurable input function into an object with the expected signature. For this notebook configure the `train_inpf` to iterate over the data twice:
```
import functools
train_inpf = functools.partial(census_dataset.input_fn, train_file, num_epochs=2, shuffle=True, batch_size=64)
test_inpf = functools.partial(census_dataset.input_fn, test_file, num_epochs=1, shuffle=False, batch_size=64)
```
## Selecting and Engineering Features for the Model
Estimators use a system called [feature columns](https://www.tensorflow.org/r1/guide/feature_columns) to describe how the model should interpret each of the raw input features. An Estimator expects a vector of numeric inputs, and feature columns describe how the model should convert each feature.
Selecting and crafting the right set of feature columns is key to learning an effective model. A *feature column* can be either one of the raw inputs in the original features `dict` (a *base feature column*), or any new columns created using transformations defined over one or multiple base columns (a *derived feature columns*).
A feature column is an abstract concept of any raw or derived variable that can be used to predict the target label.
### Base Feature Columns
#### Numeric columns
The simplest `feature_column` is `numeric_column`. This indicates that a feature is a numeric value that should be input to the model directly. For example:
```
age = fc.numeric_column('age')
```
The model will use the `feature_column` definitions to build the model input. You can inspect the resulting output using the `input_layer` function:
```
fc.input_layer(feature_batch, [age]).numpy()
```
The following will train and evaluate a model using only the `age` feature:
```
classifier = tf.estimator.LinearClassifier(feature_columns=[age])
classifier.train(train_inpf)
result = classifier.evaluate(test_inpf)
clear_output() # used for display in notebook
print(result)
```
Similarly, we can define a `NumericColumn` for each continuous feature column
that we want to use in the model:
```
education_num = tf.feature_column.numeric_column('education_num')
capital_gain = tf.feature_column.numeric_column('capital_gain')
capital_loss = tf.feature_column.numeric_column('capital_loss')
hours_per_week = tf.feature_column.numeric_column('hours_per_week')
my_numeric_columns = [age,education_num, capital_gain, capital_loss, hours_per_week]
fc.input_layer(feature_batch, my_numeric_columns).numpy()
```
You could retrain a model on these features by changing the `feature_columns` argument to the constructor:
```
classifier = tf.estimator.LinearClassifier(feature_columns=my_numeric_columns)
classifier.train(train_inpf)
result = classifier.evaluate(test_inpf)
clear_output()
for key,value in sorted(result.items()):
print('%s: %s' % (key, value))
```
#### Categorical columns
To define a feature column for a categorical feature, create a `CategoricalColumn` using one of the `tf.feature_column.categorical_column*` functions.
If you know the set of all possible feature values of a column—and there are only a few of them—use `categorical_column_with_vocabulary_list`. Each key in the list is assigned an auto-incremented ID starting from 0. For example, for the `relationship` column we can assign the feature string `Husband` to an integer ID of 0 and "Not-in-family" to 1, etc.
```
relationship = fc.categorical_column_with_vocabulary_list(
'relationship',
['Husband', 'Not-in-family', 'Wife', 'Own-child', 'Unmarried', 'Other-relative'])
```
This creates a sparse one-hot vector from the raw input feature.
The `input_layer` function we're using is designed for DNN models and expects dense inputs. To demonstrate the categorical column we must wrap it in a `tf.feature_column.indicator_column` to create the dense one-hot output (Linear `Estimators` can often skip this dense-step).
Note: the other sparse-to-dense option is `tf.feature_column.embedding_column`.
Run the input layer, configured with both the `age` and `relationship` columns:
```
fc.input_layer(feature_batch, [age, fc.indicator_column(relationship)])
```
If we don't know the set of possible values in advance, use the `categorical_column_with_hash_bucket` instead:
```
occupation = tf.feature_column.categorical_column_with_hash_bucket(
'occupation', hash_bucket_size=1000)
```
Here, each possible value in the feature column `occupation` is hashed to an integer ID as we encounter them in training. The example batch has a few different occupations:
```
for item in feature_batch['occupation'].numpy():
print(item.decode())
```
If we run `input_layer` with the hashed column, we see that the output shape is `(batch_size, hash_bucket_size)`:
```
occupation_result = fc.input_layer(feature_batch, [fc.indicator_column(occupation)])
occupation_result.numpy().shape
```
It's easier to see the actual results if we take the `tf.argmax` over the `hash_bucket_size` dimension. Notice how any duplicate occupations are mapped to the same pseudo-random index:
```
tf.argmax(occupation_result, axis=1).numpy()
```
Note: Hash collisions are unavoidable, but often have minimal impact on model quality. The effect may be noticable if the hash buckets are being used to compress the input space.
No matter how we choose to define a `SparseColumn`, each feature string is mapped into an integer ID by looking up a fixed mapping or by hashing. Under the hood, the `LinearModel` class is responsible for managing the mapping and creating `tf.Variable` to store the model parameters (model *weights*) for each feature ID. The model parameters are learned through the model training process described later.
Let's do the similar trick to define the other categorical features:
```
education = tf.feature_column.categorical_column_with_vocabulary_list(
'education', [
'Bachelors', 'HS-grad', '11th', 'Masters', '9th', 'Some-college',
'Assoc-acdm', 'Assoc-voc', '7th-8th', 'Doctorate', 'Prof-school',
'5th-6th', '10th', '1st-4th', 'Preschool', '12th'])
marital_status = tf.feature_column.categorical_column_with_vocabulary_list(
'marital_status', [
'Married-civ-spouse', 'Divorced', 'Married-spouse-absent',
'Never-married', 'Separated', 'Married-AF-spouse', 'Widowed'])
workclass = tf.feature_column.categorical_column_with_vocabulary_list(
'workclass', [
'Self-emp-not-inc', 'Private', 'State-gov', 'Federal-gov',
'Local-gov', '?', 'Self-emp-inc', 'Without-pay', 'Never-worked'])
my_categorical_columns = [relationship, occupation, education, marital_status, workclass]
```
It's easy to use both sets of columns to configure a model that uses all these features:
```
classifier = tf.estimator.LinearClassifier(feature_columns=my_numeric_columns+my_categorical_columns)
classifier.train(train_inpf)
result = classifier.evaluate(test_inpf)
clear_output()
for key,value in sorted(result.items()):
print('%s: %s' % (key, value))
```
### Derived feature columns
#### Make Continuous Features Categorical through Bucketization
Sometimes the relationship between a continuous feature and the label is not linear. For example, *age* and *income*—a person's income may grow in the early stage of their career, then the growth may slow at some point, and finally, the income decreases after retirement. In this scenario, using the raw `age` as a real-valued feature column might not be a good choice because the model can only learn one of the three cases:
1. Income always increases at some rate as age grows (positive correlation),
2. Income always decreases at some rate as age grows (negative correlation), or
3. Income stays the same no matter at what age (no correlation).
If we want to learn the fine-grained correlation between income and each age group separately, we can leverage *bucketization*. Bucketization is a process of dividing the entire range of a continuous feature into a set of consecutive buckets, and then converting the original numerical feature into a bucket ID (as a categorical feature) depending on which bucket that value falls into. So, we can define a `bucketized_column` over `age` as:
```
age_buckets = tf.feature_column.bucketized_column(
age, boundaries=[18, 25, 30, 35, 40, 45, 50, 55, 60, 65])
```
`boundaries` is a list of bucket boundaries. In this case, there are 10 boundaries, resulting in 11 age group buckets (from age 17 and below, 18-24, 25-29, ..., to 65 and over).
With bucketing, the model sees each bucket as a one-hot feature:
```
fc.input_layer(feature_batch, [age, age_buckets]).numpy()
```
#### Learn complex relationships with crossed column
Using each base feature column separately may not be enough to explain the data. For example, the correlation between education and the label (earning > 50,000 dollars) may be different for different occupations. Therefore, if we only learn a single model weight for `education="Bachelors"` and `education="Masters"`, we won't capture every education-occupation combination (e.g. distinguishing between `education="Bachelors"` AND `occupation="Exec-managerial"` AND `education="Bachelors" AND occupation="Craft-repair"`).
To learn the differences between different feature combinations, we can add *crossed feature columns* to the model:
```
education_x_occupation = tf.feature_column.crossed_column(
['education', 'occupation'], hash_bucket_size=1000)
```
We can also create a `crossed_column` over more than two columns. Each constituent column can be either a base feature column that is categorical (`SparseColumn`), a bucketized real-valued feature column, or even another `CrossColumn`. For example:
```
age_buckets_x_education_x_occupation = tf.feature_column.crossed_column(
[age_buckets, 'education', 'occupation'], hash_bucket_size=1000)
```
These crossed columns always use hash buckets to avoid the exponential explosion in the number of categories, and put the control over number of model weights in the hands of the user.
## Define the logistic regression model
After processing the input data and defining all the feature columns, we can put them together and build a *logistic regression* model. The previous section showed several types of base and derived feature columns, including:
* `CategoricalColumn`
* `NumericColumn`
* `BucketizedColumn`
* `CrossedColumn`
All of these are subclasses of the abstract `FeatureColumn` class and can be added to the `feature_columns` field of a model:
```
import tempfile
base_columns = [
education, marital_status, relationship, workclass, occupation,
age_buckets,
]
crossed_columns = [
tf.feature_column.crossed_column(
['education', 'occupation'], hash_bucket_size=1000),
tf.feature_column.crossed_column(
[age_buckets, 'education', 'occupation'], hash_bucket_size=1000),
]
model = tf.estimator.LinearClassifier(
model_dir=tempfile.mkdtemp(),
feature_columns=base_columns + crossed_columns,
optimizer=tf.train.FtrlOptimizer(learning_rate=0.1))
```
The model automatically learns a bias term, which controls the prediction made without observing any features. The learned model files are stored in `model_dir`.
## Train and evaluate the model
After adding all the features to the model, let's train the model. Training a model is just a single command using the `tf.estimator` API:
```
train_inpf = functools.partial(census_dataset.input_fn, train_file,
num_epochs=40, shuffle=True, batch_size=64)
model.train(train_inpf)
clear_output() # used for notebook display
```
After the model is trained, evaluate the accuracy of the model by predicting the labels of the holdout data:
```
results = model.evaluate(test_inpf)
clear_output()
for key,value in sorted(results.items()):
print('%s: %0.2f' % (key, value))
```
The first line of the output should display something like: `accuracy: 0.84`, which means the accuracy is 84%. You can try using more features and transformations to see if you can do better!
After the model is evaluated, we can use it to predict whether an individual has an annual income of over 50,000 dollars given an individual's information input.
Let's look in more detail how the model performed:
```
import numpy as np
predict_df = test_df[:20].copy()
pred_iter = model.predict(
lambda:easy_input_function(predict_df, label_key='income_bracket',
num_epochs=1, shuffle=False, batch_size=10))
classes = np.array(['<=50K', '>50K'])
pred_class_id = []
for pred_dict in pred_iter:
pred_class_id.append(pred_dict['class_ids'])
predict_df['predicted_class'] = classes[np.array(pred_class_id)]
predict_df['correct'] = predict_df['predicted_class'] == predict_df['income_bracket']
clear_output()
predict_df[['income_bracket','predicted_class', 'correct']]
```
For a working end-to-end example, download our [example code](https://github.com/tensorflow/models/tree/master/official/r1/wide_deep/census_main.py) and set the `model_type` flag to `wide`.
## Adding Regularization to Prevent Overfitting
Regularization is a technique used to avoid overfitting. Overfitting happens when a model performs well on the data it is trained on, but worse on test data that the model has not seen before. Overfitting can occur when a model is excessively complex, such as having too many parameters relative to the number of observed training data. Regularization allows you to control the model's complexity and make the model more generalizable to unseen data.
You can add L1 and L2 regularizations to the model with the following code:
```
model_l1 = tf.estimator.LinearClassifier(
feature_columns=base_columns + crossed_columns,
optimizer=tf.train.FtrlOptimizer(
learning_rate=0.1,
l1_regularization_strength=10.0,
l2_regularization_strength=0.0))
model_l1.train(train_inpf)
results = model_l1.evaluate(test_inpf)
clear_output()
for key in sorted(results):
print('%s: %0.2f' % (key, results[key]))
model_l2 = tf.estimator.LinearClassifier(
feature_columns=base_columns + crossed_columns,
optimizer=tf.train.FtrlOptimizer(
learning_rate=0.1,
l1_regularization_strength=0.0,
l2_regularization_strength=10.0))
model_l2.train(train_inpf)
results = model_l2.evaluate(test_inpf)
clear_output()
for key in sorted(results):
print('%s: %0.2f' % (key, results[key]))
```
These regularized models don't perform much better than the base model. Let's look at the model's weight distributions to better see the effect of the regularization:
```
def get_flat_weights(model):
weight_names = [
name for name in model.get_variable_names()
if "linear_model" in name and "Ftrl" not in name]
weight_values = [model.get_variable_value(name) for name in weight_names]
weights_flat = np.concatenate([item.flatten() for item in weight_values], axis=0)
return weights_flat
weights_flat = get_flat_weights(model)
weights_flat_l1 = get_flat_weights(model_l1)
weights_flat_l2 = get_flat_weights(model_l2)
```
The models have many zero-valued weights caused by unused hash bins (there are many more hash bins than categories in some columns). We can mask these weights when viewing the weight distributions:
```
weight_mask = weights_flat != 0
weights_base = weights_flat[weight_mask]
weights_l1 = weights_flat_l1[weight_mask]
weights_l2 = weights_flat_l2[weight_mask]
```
Now plot the distributions:
```
plt.figure()
_ = plt.hist(weights_base, bins=np.linspace(-3,3,30))
plt.title('Base Model')
plt.ylim([0,500])
plt.figure()
_ = plt.hist(weights_l1, bins=np.linspace(-3,3,30))
plt.title('L1 - Regularization')
plt.ylim([0,500])
plt.figure()
_ = plt.hist(weights_l2, bins=np.linspace(-3,3,30))
plt.title('L2 - Regularization')
_=plt.ylim([0,500])
```
Both types of regularization squeeze the distribution of weights towards zero. L2 regularization has a greater effect in the tails of the distribution eliminating extreme weights. L1 regularization produces more exactly-zero values, in this case it sets ~200 to zero.
| github_jupyter |
# Gradient Checking
Welcome to the final assignment for this week! In this assignment you will learn to implement and use gradient checking.
You are part of a team working to make mobile payments available globally, and are asked to build a deep learning model to detect fraud--whenever someone makes a payment, you want to see if the payment might be fraudulent, such as if the user's account has been taken over by a hacker.
But backpropagation is quite challenging to implement, and sometimes has bugs. Because this is a mission-critical application, your company's CEO wants to be really certain that your implementation of backpropagation is correct. Your CEO says, "Give me a proof that your backpropagation is actually working!" To give this reassurance, you are going to use "gradient checking".
Let's do it!
```
# Packages
import numpy as np
from testCases import *
from gc_utils import sigmoid, relu, dictionary_to_vector, vector_to_dictionary, gradients_to_vector
```
## 1) How does gradient checking work?
Backpropagation computes the gradients $\frac{\partial J}{\partial \theta}$, where $\theta$ denotes the parameters of the model. $J$ is computed using forward propagation and your loss function.
Because forward propagation is relatively easy to implement, you're confident you got that right, and so you're almost 100% sure that you're computing the cost $J$ correctly. Thus, you can use your code for computing $J$ to verify the code for computing $\frac{\partial J}{\partial \theta}$.
Let's look back at the definition of a derivative (or gradient):
$$ \frac{\partial J}{\partial \theta} = \lim_{\varepsilon \to 0} \frac{J(\theta + \varepsilon) - J(\theta - \varepsilon)}{2 \varepsilon} \tag{1}$$
If you're not familiar with the "$\displaystyle \lim_{\varepsilon \to 0}$" notation, it's just a way of saying "when $\varepsilon$ is really really small."
We know the following:
- $\frac{\partial J}{\partial \theta}$ is what you want to make sure you're computing correctly.
- You can compute $J(\theta + \varepsilon)$ and $J(\theta - \varepsilon)$ (in the case that $\theta$ is a real number), since you're confident your implementation for $J$ is correct.
Lets use equation (1) and a small value for $\varepsilon$ to convince your CEO that your code for computing $\frac{\partial J}{\partial \theta}$ is correct!
## 2) 1-dimensional gradient checking
Consider a 1D linear function $J(\theta) = \theta x$. The model contains only a single real-valued parameter $\theta$, and takes $x$ as input.
You will implement code to compute $J(.)$ and its derivative $\frac{\partial J}{\partial \theta}$. You will then use gradient checking to make sure your derivative computation for $J$ is correct.
<img src="images/1Dgrad_kiank.png" style="width:600px;height:250px;">
<caption><center> <u> **Figure 1** </u>: **1D linear model**<br> </center></caption>
The diagram above shows the key computation steps: First start with $x$, then evaluate the function $J(x)$ ("forward propagation"). Then compute the derivative $\frac{\partial J}{\partial \theta}$ ("backward propagation").
**Exercise**: implement "forward propagation" and "backward propagation" for this simple function. I.e., compute both $J(.)$ ("forward propagation") and its derivative with respect to $\theta$ ("backward propagation"), in two separate functions.
```
# GRADED FUNCTION: forward_propagation
def forward_propagation(x, theta):
"""
Implement the linear forward propagation (compute J) presented in Figure 1 (J(theta) = theta * x)
Arguments:
x -- a real-valued input
theta -- our parameter, a real number as well
Returns:
J -- the value of function J, computed using the formula J(theta) = theta * x
"""
### START CODE HERE ### (approx. 1 line)
J = theta * x
### END CODE HERE ###
return J
x, theta = 2, 4
J = forward_propagation(x, theta)
print ("J = " + str(J))
```
**Expected Output**:
<table style=>
<tr>
<td> ** J ** </td>
<td> 8</td>
</tr>
</table>
**Exercise**: Now, implement the backward propagation step (derivative computation) of Figure 1. That is, compute the derivative of $J(\theta) = \theta x$ with respect to $\theta$. To save you from doing the calculus, you should get $dtheta = \frac { \partial J }{ \partial \theta} = x$.
```
# GRADED FUNCTION: backward_propagation
def backward_propagation(x, theta):
"""
Computes the derivative of J with respect to theta (see Figure 1).
Arguments:
x -- a real-valued input
theta -- our parameter, a real number as well
Returns:
dtheta -- the gradient of the cost with respect to theta
"""
### START CODE HERE ### (approx. 1 line)
dtheta = x
### END CODE HERE ###
return dtheta
x, theta = 2, 4
dtheta = backward_propagation(x, theta)
print ("dtheta = " + str(dtheta))
```
**Expected Output**:
<table>
<tr>
<td> ** dtheta ** </td>
<td> 2 </td>
</tr>
</table>
**Exercise**: To show that the `backward_propagation()` function is correctly computing the gradient $\frac{\partial J}{\partial \theta}$, let's implement gradient checking.
**Instructions**:
- First compute "gradapprox" using the formula above (1) and a small value of $\varepsilon$. Here are the Steps to follow:
1. $\theta^{+} = \theta + \varepsilon$
2. $\theta^{-} = \theta - \varepsilon$
3. $J^{+} = J(\theta^{+})$
4. $J^{-} = J(\theta^{-})$
5. $gradapprox = \frac{J^{+} - J^{-}}{2 \varepsilon}$
- Then compute the gradient using backward propagation, and store the result in a variable "grad"
- Finally, compute the relative difference between "gradapprox" and the "grad" using the following formula:
$$ difference = \frac {\mid\mid grad - gradapprox \mid\mid_2}{\mid\mid grad \mid\mid_2 + \mid\mid gradapprox \mid\mid_2} \tag{2}$$
You will need 3 Steps to compute this formula:
- 1'. compute the numerator using np.linalg.norm(...)
- 2'. compute the denominator. You will need to call np.linalg.norm(...) twice.
- 3'. divide them.
- If this difference is small (say less than $10^{-7}$), you can be quite confident that you have computed your gradient correctly. Otherwise, there may be a mistake in the gradient computation.
```
# GRADED FUNCTION: gradient_check
def gradient_check(x, theta, epsilon = 1e-7):
"""
Implement the backward propagation presented in Figure 1.
Arguments:
x -- a real-valued input
theta -- our parameter, a real number as well
epsilon -- tiny shift to the input to compute approximated gradient with formula(1)
Returns:
difference -- difference (2) between the approximated gradient and the backward propagation gradient
"""
# Compute gradapprox using left side of formula (1). epsilon is small enough, you don't need to worry about the limit.
### START CODE HERE ### (approx. 5 lines)
thetaplus = theta + epsilon # Step 1
thetaminus = theta - epsilon # Step 2
J_plus = forward_propagation(x, thetaplus) # Step 3
J_minus = forward_propagation(x, thetaminus) # Step 4
gradapprox = (J_plus - J_minus)/(2*epsilon) # Step 5
### END CODE HERE ###
# Check if gradapprox is close enough to the output of backward_propagation()
### START CODE HERE ### (approx. 1 line)
grad = backward_propagation(x, theta)
### END CODE HERE ###
### START CODE HERE ### (approx. 1 line)
numerator = np.linalg.norm(grad - gradapprox) # Step 1'
denominator = np.linalg.norm(grad) + np.linalg.norm(gradapprox) # Step 2'
difference = numerator/denominator # Step 3'
### END CODE HERE ###
if difference < 1e-7:
print ("The gradient is correct!")
else:
print ("The gradient is wrong!")
return difference
x, theta = 2, 4
difference = gradient_check(x, theta)
print("difference = " + str(difference))
```
**Expected Output**:
The gradient is correct!
<table>
<tr>
<td> ** difference ** </td>
<td> 2.9193358103083e-10 </td>
</tr>
</table>
Congrats, the difference is smaller than the $10^{-7}$ threshold. So you can have high confidence that you've correctly computed the gradient in `backward_propagation()`.
Now, in the more general case, your cost function $J$ has more than a single 1D input. When you are training a neural network, $\theta$ actually consists of multiple matrices $W^{[l]}$ and biases $b^{[l]}$! It is important to know how to do a gradient check with higher-dimensional inputs. Let's do it!
## 3) N-dimensional gradient checking
The following figure describes the forward and backward propagation of your fraud detection model.
<img src="images/NDgrad_kiank.png" style="width:600px;height:400px;">
<caption><center> <u> **Figure 2** </u>: **deep neural network**<br>*LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SIGMOID*</center></caption>
Let's look at your implementations for forward propagation and backward propagation.
```
def forward_propagation_n(X, Y, parameters):
"""
Implements the forward propagation (and computes the cost) presented in Figure 3.
Arguments:
X -- training set for m examples
Y -- labels for m examples
parameters -- python dictionary containing your parameters "W1", "b1", "W2", "b2", "W3", "b3":
W1 -- weight matrix of shape (5, 4)
b1 -- bias vector of shape (5, 1)
W2 -- weight matrix of shape (3, 5)
b2 -- bias vector of shape (3, 1)
W3 -- weight matrix of shape (1, 3)
b3 -- bias vector of shape (1, 1)
Returns:
cost -- the cost function (logistic cost for one example)
"""
# retrieve parameters
m = X.shape[1]
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
W3 = parameters["W3"]
b3 = parameters["b3"]
# LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SIGMOID
Z1 = np.dot(W1, X) + b1
A1 = relu(Z1)
Z2 = np.dot(W2, A1) + b2
A2 = relu(Z2)
Z3 = np.dot(W3, A2) + b3
A3 = sigmoid(Z3)
# Cost
logprobs = np.multiply(-np.log(A3),Y) + np.multiply(-np.log(1 - A3), 1 - Y)
cost = 1./m * np.sum(logprobs)
cache = (Z1, A1, W1, b1, Z2, A2, W2, b2, Z3, A3, W3, b3)
return cost, cache
```
Now, run backward propagation.
```
def backward_propagation_n(X, Y, cache):
"""
Implement the backward propagation presented in figure 2.
Arguments:
X -- input datapoint, of shape (input size, 1)
Y -- true "label"
cache -- cache output from forward_propagation_n()
Returns:
gradients -- A dictionary with the gradients of the cost with respect to each parameter, activation and pre-activation variables.
"""
m = X.shape[1]
(Z1, A1, W1, b1, Z2, A2, W2, b2, Z3, A3, W3, b3) = cache
dZ3 = A3 - Y
dW3 = 1./m * np.dot(dZ3, A2.T)
db3 = 1./m * np.sum(dZ3, axis=1, keepdims = True)
dA2 = np.dot(W3.T, dZ3)
dZ2 = np.multiply(dA2, np.int64(A2 > 0))
dW2 = 1./m * np.dot(dZ2, A1.T) * 2
db2 = 1./m * np.sum(dZ2, axis=1, keepdims = True)
dA1 = np.dot(W2.T, dZ2)
dZ1 = np.multiply(dA1, np.int64(A1 > 0))
dW1 = 1./m * np.dot(dZ1, X.T)
db1 = 4./m * np.sum(dZ1, axis=1, keepdims = True)
gradients = {"dZ3": dZ3, "dW3": dW3, "db3": db3,
"dA2": dA2, "dZ2": dZ2, "dW2": dW2, "db2": db2,
"dA1": dA1, "dZ1": dZ1, "dW1": dW1, "db1": db1}
return gradients
```
You obtained some results on the fraud detection test set but you are not 100% sure of your model. Nobody's perfect! Let's implement gradient checking to verify if your gradients are correct.
**How does gradient checking work?**.
As in 1) and 2), you want to compare "gradapprox" to the gradient computed by backpropagation. The formula is still:
$$ \frac{\partial J}{\partial \theta} = \lim_{\varepsilon \to 0} \frac{J(\theta + \varepsilon) - J(\theta - \varepsilon)}{2 \varepsilon} \tag{1}$$
However, $\theta$ is not a scalar anymore. It is a dictionary called "parameters". We implemented a function "`dictionary_to_vector()`" for you. It converts the "parameters" dictionary into a vector called "values", obtained by reshaping all parameters (W1, b1, W2, b2, W3, b3) into vectors and concatenating them.
The inverse function is "`vector_to_dictionary`" which outputs back the "parameters" dictionary.
<img src="images/dictionary_to_vector.png" style="width:600px;height:400px;">
<caption><center> <u> **Figure 2** </u>: **dictionary_to_vector() and vector_to_dictionary()**<br> You will need these functions in gradient_check_n()</center></caption>
We have also converted the "gradients" dictionary into a vector "grad" using gradients_to_vector(). You don't need to worry about that.
**Exercise**: Implement gradient_check_n().
**Instructions**: Here is pseudo-code that will help you implement the gradient check.
For each i in num_parameters:
- To compute `J_plus[i]`:
1. Set $\theta^{+}$ to `np.copy(parameters_values)`
2. Set $\theta^{+}_i$ to $\theta^{+}_i + \varepsilon$
3. Calculate $J^{+}_i$ using to `forward_propagation_n(x, y, vector_to_dictionary(`$\theta^{+}$ `))`.
- To compute `J_minus[i]`: do the same thing with $\theta^{-}$
- Compute $gradapprox[i] = \frac{J^{+}_i - J^{-}_i}{2 \varepsilon}$
Thus, you get a vector gradapprox, where gradapprox[i] is an approximation of the gradient with respect to `parameter_values[i]`. You can now compare this gradapprox vector to the gradients vector from backpropagation. Just like for the 1D case (Steps 1', 2', 3'), compute:
$$ difference = \frac {\| grad - gradapprox \|_2}{\| grad \|_2 + \| gradapprox \|_2 } \tag{3}$$
```
# GRADED FUNCTION: gradient_check_n
def gradient_check_n(parameters, gradients, X, Y, epsilon = 1e-7):
"""
Checks if backward_propagation_n computes correctly the gradient of the cost output by forward_propagation_n
Arguments:
parameters -- python dictionary containing your parameters "W1", "b1", "W2", "b2", "W3", "b3":
grad -- output of backward_propagation_n, contains gradients of the cost with respect to the parameters.
x -- input datapoint, of shape (input size, 1)
y -- true "label"
epsilon -- tiny shift to the input to compute approximated gradient with formula(1)
Returns:
difference -- difference (2) between the approximated gradient and the backward propagation gradient
"""
# Set-up variables
parameters_values, _ = dictionary_to_vector(parameters)
grad = gradients_to_vector(gradients)
num_parameters = parameters_values.shape[0]
J_plus = np.zeros((num_parameters, 1))
J_minus = np.zeros((num_parameters, 1))
gradapprox = np.zeros((num_parameters, 1))
# Compute gradapprox
for i in range(num_parameters):
# Compute J_plus[i]. Inputs: "parameters_values, epsilon". Output = "J_plus[i]".
# "_" is used because the function you have to outputs two parameters but we only care about the first one
### START CODE HERE ### (approx. 3 lines)
thetaplus = np.copy(parameters_values) # Step 1
thetaplus[i][0] += epsilon # Step 2
J_plus[i], _ = forward_propagation_n(X, Y, vector_to_dictionary(thetaplus)) # Step 3
### END CODE HERE ###
# Compute J_minus[i]. Inputs: "parameters_values, epsilon". Output = "J_minus[i]".
### START CODE HERE ### (approx. 3 lines)
thetaminus = np.copy(parameters_values) # Step 1
thetaminus[i][0] -= epsilon # Step 2
J_minus[i], _ = forward_propagation_n(X, Y, vector_to_dictionary(thetaminus)) # Step 3
### END CODE HERE ###
# Compute gradapprox[i]
### START CODE HERE ### (approx. 1 line)
gradapprox[i] = (J_plus[i] - J_minus[i])/(2*epsilon)
### END CODE HERE ###
# Compare gradapprox to backward propagation gradients by computing difference.
### START CODE HERE ### (approx. 1 line)
numerator = np.linalg.norm(grad - gradapprox) # Step 1'
denominator = np.linalg.norm(grad) + np.linalg.norm(gradapprox) # Step 2'
difference = numerator/denominator # Step 3'
### END CODE HERE ###
if difference > 2e-7:
print ("\033[93m" + "There is a mistake in the backward propagation! difference = " + str(difference) + "\033[0m")
else:
print ("\033[92m" + "Your backward propagation works perfectly fine! difference = " + str(difference) + "\033[0m")
return difference
X, Y, parameters = gradient_check_n_test_case()
cost, cache = forward_propagation_n(X, Y, parameters)
gradients = backward_propagation_n(X, Y, cache)
difference = gradient_check_n(parameters, gradients, X, Y)
```
**Expected output**:
<table>
<tr>
<td> ** There is a mistake in the backward propagation!** </td>
<td> difference = 0.285093156781 </td>
</tr>
</table>
It seems that there were errors in the `backward_propagation_n` code we gave you! Good that you've implemented the gradient check. Go back to `backward_propagation` and try to find/correct the errors *(Hint: check dW2 and db1)*. Rerun the gradient check when you think you've fixed it. Remember you'll need to re-execute the cell defining `backward_propagation_n()` if you modify the code.
Can you get gradient check to declare your derivative computation correct? Even though this part of the assignment isn't graded, we strongly urge you to try to find the bug and re-run gradient check until you're convinced backprop is now correctly implemented.
**Note**
- Gradient Checking is slow! Approximating the gradient with $\frac{\partial J}{\partial \theta} \approx \frac{J(\theta + \varepsilon) - J(\theta - \varepsilon)}{2 \varepsilon}$ is computationally costly. For this reason, we don't run gradient checking at every iteration during training. Just a few times to check if the gradient is correct.
- Gradient Checking, at least as we've presented it, doesn't work with dropout. You would usually run the gradient check algorithm without dropout to make sure your backprop is correct, then add dropout.
Congrats, you can be confident that your deep learning model for fraud detection is working correctly! You can even use this to convince your CEO. :)
<font color='blue'>
**What you should remember from this notebook**:
- Gradient checking verifies closeness between the gradients from backpropagation and the numerical approximation of the gradient (computed using forward propagation).
- Gradient checking is slow, so we don't run it in every iteration of training. You would usually run it only to make sure your code is correct, then turn it off and use backprop for the actual learning process.
| github_jupyter |
```
# Imports needed for this exercice
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import hcipy as hc # get at https://docs.hcipy.org/0.4.0/
# You might need those later on in the class
# import os
# import exoscene.image
# import exoscene.star
# import exoscene.planet
# from exoscene.planet import Planet
# import astropy.units as u
```
For the sake of this exercise let's define the 2D Fourier transfrom as transforming a function f(x,y) into F(u,v)
\begin{equation}
F(\xi,\eta) = \frac{1}{\sqrt{2\pi}} \int_{-\infty}^{+\infty} f(x,y) e^{\left[i(x \xi +y \eta )\right]} dx dy
\end{equation}
When we use the FT for optics purposes it is often more convenient to use the following definition:
\begin{equation}
F(u,v) = \frac{1}{\sqrt{2\pi}} \int_{-\infty}^{+\infty} f(x,y) e^{\left[i 2 \pi (x u +y u )\right]} dx dy
\end{equation}
where we have simply replace the variables $ (\xi,\eta) = 2 \pi (u,v)$. We also assume for now that the spatial extent of $f(x,y)$ is finite on a unit square, that is $x \in [-1/2,1/2]$. When the bounding area (or volume for higher dimensional FTs) is not unitary (say $\tilde{x} \in [-D/2,D/2]$) then we can also use the definition above after the simple change of variables $x = \tilde{x}/D$.
We call "units of angular resolution" each unit increment in fourier space (eg in u and v)
The "physics" reasons for doing this will be detailed in Lecture 3. For now let's just roll with these definitions.
Note: Whether the forward transform has a $+i$ and the reverse transform a $-i$ or vice versa does not make a difference, but you have to use the same convention in the entire calculation/theory. Anand uses the opposite of this, $-i$ for forward transforms.
# How to make a circular aperture in hcipy?
These are defined as abstract objects with a center and a radius. We need to:
1) Decide on the number of points (numerical sampling) we call it Npup
2) Define the abstract object (we use aperture.circular_aperture)
3) Define the grid (we use field.make_pupil_grid)
4) Evaluate the object on the grid
5) Display the array a figure (we use imshow_field). Note that all HCIPy fields are collapsed oin one dimesion. We could also reshape them into a 2D array and use the native imshow of matplotlib.
```
Npup = 100
print('number of pixels in aperture array = ' + str(Npup))
Diam = 1
print('diameter of the physical array = ' + str(Diam))
circular_aperture_object = hc.aperture.circular_aperture(1., center=None)
pupil_grid = hc.field.make_pupil_grid(Npup, diameter=1.)
circular_aperture_field = circular_aperture_object(pupil_grid)
fig = plt.figure(figsize = [10,10])
hc.imshow_field(circular_aperture_field)
circular_aperture_field_square_array = np.reshape(circular_aperture_field,[Npup, Npup])
fig = plt.figure(figsize = [10,10])
plt.imshow(circular_aperture_field_square_array)
```
# Take the fourier transform of this aperture with MFT
Fourier transforms are defined as abstract methods. We first define their parameters and the use them on our freshly define array. Here is out to do this:
1) define q, nunber of pixels per units of angular resolution
2) define number of elements of angular resolution num_airy
3) define focal plane grid (hint use field.make_focal_grid and ignore all arguments after num_airy). Note that this function is defined in $(\xi,\eta)$ so we will need to multiply coordinates $2 \pi$
4) define the matrix fourier transform operator (we use fourier.MatrixFourierTransform)
5) take the fourier transform of the aperture (we use the .forward method)
6) calculate the amplitude of the image of aperture
7) calculate the intensity of the image of aperture (amplitude square of complex number, numpy can do that for you)
```
q = 2
print('number of pixels per unit of angular resolution = ' + str(q))
num_airy = 10
num_airy_two_pi = 2*np.pi*num_airy
print('physical size of the image in units of angular resolution = ' + str(num_airy))
print('physical size of the image in units of spatial frequency = ' + str(num_airy_two_pi))
focal_grid = hc.field.make_focal_grid(q, num_airy_two_pi)
print('predicted size of the array containing the FT = ' + str(2*num_airy*q))
MFT = hc.fourier.MatrixFourierTransform(pupil_grid,focal_grid, precompute_matrices=None, allocate_intermediate=None)
fourier_transform_with_MFT = MFT.forward(circular_aperture_field)
intensity = np.abs(fourier_transform_with_MFT)**2
print('actual size of the array containing the FT = ' + str(np.sqrt(intensity.shape[0])))
fig = plt.figure(figsize = [10,10])
hc.imshow_field(np.sqrt(intensity))
plt.colorbar()
fig = plt.figure(figsize = [10,10])
hc.imshow_field(np.log10(intensity))
```
7) compare the summed intensities of the input aperture and of the image. What do you notice? Is energy conserved?
```
np.sum(intensity)
np.sum(circular_aperture_field**2)
```
8) The energy is not conserved. Change the parameters (q,Npup and num_airy) and see how the ratio of the intensities changes. What is happenning is that by "summing" over the array we forgot to mutiply by $dx$ and $du$.
### Q1 Try to fix code so the sum of the intensities is equal. You can either derive $dx$ and $du$ based on (q,Npup and num_airy) or read directly in focal_grid.coords.delta and pupil_grid.coords.delta
# Explore the fourier transform of this aperture with MFT
1) Right now everything is the fourier plane is in units of angular resolution. If interrested play with the , spatial_resolution, pupil_diameter, focal_length, f_number, reference_wavelength to see how images scale. This will make a lot more sense after lecture 3.
```
q = 20
print('number of pixels per unit of angular resolution = ' + str(q))
num_airy = 10
num_airy_two_pi = 2*np.pi*num_airy
print('physical size of the image in units of angular resolution = ' + str(num_airy))
print('physical size of the image in units of spatial frequency = ' + str(num_airy_two_pi))
focal_grid = hc.field.make_focal_grid(q, num_airy_two_pi)
print('predicted size of the array containing the FT = ' + str(2*num_airy*q))
MFT = hc.fourier.MatrixFourierTransform(pupil_grid,focal_grid, precompute_matrices=None, allocate_intermediate=None)
fourier_transform_with_MFT = MFT.forward(circular_aperture_field)
intensity = np.abs(fourier_transform_with_MFT)**2
print('actual size of the array containing the FT = ' + str(np.sqrt(intensity.shape[0])))
fig = plt.figure(figsize = [10,10])
hc.imshow_field(np.sqrt(intensity))
plt.colorbar()
fig = plt.figure(figsize = [10,10])
hc.imshow_field(np.log10(intensity))
plt.colorbar()
```
# Take the fourier transform of this aperture with FFT
Similar steps as before:
1) define q, nunber of pixels per units of angular resolution (now this is done via zero padding, we will discuss that in class if you are not familiar with it )
2) define the matrix fourier transform operator (we use fourier.MatrixFourierTransform)
3) take the fourier transform of the aperture (we use the .forward method)
4) calculate the amplitude of the image of aperture
5) calculate the intensity of the image of aperture (amplitude square of complex number, numpy can do that for you)
6) compare the summed intensities of the input aperture and of the image. What do you notice? Is energy conserved?
7) fix the summed intensities to make sure that energy is conserved (hint use focal_grid.coords.delta). Note that there is a factor of 2 pi missing in in front of the DFT so you will need to fix this.
```
q0 = 4.
print('number of pixels per unit of angular resolution = ' + str(q0))
FFT = hc.fourier.FastFourierTransform(pupil_grid, q=q0, fov=1, shift=0, emulate_fftshifts=None)
fourier_transform_with_FFT = FFT.forward(circular_aperture_field)
print('predicted size of the array containing the FT = ' + str(q0*np.sqrt(pupil_grid.x.shape[0])))
intensity = np.abs(fourier_transform_with_FFT)**2
print('actual size of the array containing the FT = ' + str(np.sqrt(intensity.shape[0])))
fig = plt.figure(figsize = [10,10])
hc.imshow_field(np.log10(intensity))
plt.colorbar()
```
### Q2: Same exercise as before: fix the energy ratio so it is as close to one as possible. Except that this time the samplings are different
# A few example to explore now that you know the basics.
### Q3: Answer one as part of your homework (you can have a crack at all of them if you want)
1) Try the FFT with the pathological q = 2. What is going on here?
2) Figure out the value of num_airy so that the result from the MFT is the same as the result from the FFT (or at least close enough). Try to explain what is happenning.
3) Try to apply the forward and backward methods to see what happens. What are differences between FFT and MFT? Try to explain what is happenning.
4) Explore what zero padding actuattly does with both MFT and FFT by defining the aperture as circular_aperture_object = hc.aperture.circular_aperture(0.5, center=None) [or whatever nuber <1 you d like]. Try to explain what is happenning.
| github_jupyter |
# SQL Queries 01
For more SQL examples in the SQLite3 dialect, seee [SQLite3 tutorial](https://www.techonthenet.com/sqlite/index.php).
For a deep dive, see [SQL Queries for Mere Mortals](https://www.amazon.com/SQL-Queries-Mere-Mortals-Hands/dp/0134858336/ref=dp_ob_title_bk).
## Data
```
%load_ext sql
%sql sqlite:///data/faculty.db
%%sql
SELECT * FROM sqlite_master WHERE type='table';
```
Note: You can save results as a variable
```
%%sql master <<
SELECT * FROM sqlite_master WHERE type='table'
master.DataFrame()
```
## Basic Structure
```SQL
SELECT DISTINCT value_expression AS alias
FROM tables AS alias
WHERE predicate
ORDER BY value_expression
```
### Types
- Character (Fixed width, variable width)
- National Character (Fixed width, variable width)
- Binary
- Numeric (Exact, Arpproximate)
- Boolean
- DateTime
- Interval
**CHAR** and **NCHAR** are vendor-dependent. Sometimes they mean the same thing, and sometimes CHAR means bytes and NCHAR means Unicode.
The SQL standard specifies that character strings and datetime literals are enclosed by single quotes. Two single quotes wihtin a string is intepreted as a literal single quote.
```sql
'Gilligan''s island'
```
#### The CAST function
```sql
CAST(X as CHARACTER(10))
```
### Value expression
- Literal
- Column reference
- Function
- CASES
- (Value expression)
which may be prefixed with unary operators `-` and `+` and combined with binary operators appropriate for the data type.
Literal
```
%sql SELECT 23
```
Column reference
```
%sql SELECT first, last FROM person LIMIT 3
```
Function
```
%sql SELECT count(*) FROM person
```
Cases
```
%%sql
SELECT first, last, age,
CASE
WHEN age < 50 THEN 'Whippernapper'
WHEN age < 70 THEN 'Old codger'
ELSE 'Dinosaur'
END comment
FROM person
LIMIT 4
```
Value expression
```
%%sql
SELECT first || ' ' || last AS name, age, age - 10 AS fake_age
FROM person
LIMIT 3
```
### Bineary operators
#### Concatenation
```SQL
A || B
```
#### Mathematical
```SQL
A + B
A - B
A * B
A / B
```
#### Data and time arithmetic
```SQL
'2018-08-29' + 3
'11:59' + '00:01'
```
```
%%sql
SELECT DISTINCT language_name
FROM language
LIMIT 5;
```
### Sorting
```SQL
SELECT DISTINCT value_expression AS alias
FROM tables AS alias
ORDER BY value_expression
```
```
%%sql
SELECT DISTINCT language_name
FROM language
ORDER BY language_name ASC
LIMIT 5;
%%sql
SELECT DISTINCT language_name
FROM language
ORDER BY random()
LIMIT 5;
```
### Filtering
For efficiency, place the most stringent filters first.
```SQL
SELECT DISTINCT value_expression AS alias
FROM tables AS alias
WHERE predicate
ORDER BY value_expression
```
#### Predicates for filtering rows
- Comparison operators (=, <>, <, >, <=, >=)
- BETWEEN start AND end
- IN(A, B, C)
- LIKE
- IS NULL
- REGEX
Use NOT prefix for negation
#### Combining predicates
```sql
AND
OR
```
USe parenthesis to indicate order of evaluation for compound statements.
```
%%sql
SELECT first, last, age
FROM person
WHERE age BETWEEN 16 AND 17
LIMIT 5;
```
### Joins
Joins combine data from 1 or more tables to form a new result set.
Note: To join on multiple columns just use `AND` in the `ON` expression
#### Natural join
Uses all common columns in Tables 1 and 2 for JOIN
```SQL
FROM Table1
NATURAL INNER JOIN Table 2
```
#### Inner join
General form of INNER JOIN uisng ON
```SQL
FROM Table1
INNER JOIN Table2
ON Table1.Column = Table2.Column
```
**Note**: This is equivalent to an EQUIJOIN but more flexible in that additional JOIN conditions can be specified.
```SQL
SELECT *
FROM Table1, Table2
WHERE Table1.Column = Table2.Column
```
If there is a common column in both tables
```SQL
FROM Table1
INNER JOIN Table2
USING Column
```
Joining more than two tables
```SQL
From (Table1
INNER JOIN Table2
ON Table1.column1 = Table2.Column1)
INNER JOIN Table3
ON Table3.column2 = Table2.Column2
```
#### Outer join
General form of OUTER JOIN uisng ON
```SQL
FROM Table1
RIGHT OUTER JOIN Table2
ON Table1.Column = Table2.Column
```
```SQL
FROM Table1
LEFT OUTER JOIN Table2
ON Table1.Column = Table2.Column
```
```SQL
FROM Table1
FULL OUTER JOIN Table2
ON Table1.Column = Table2.Column
```
```
%%sql
SELECT first, last, language_name
FROM person
INNER JOIN person_language
ON person.person_id = person_language.person_id
INNER JOIN language
ON language.language_id = person_language.language_id
LIMIT 10;
```
### Set operations
```SQL
SELECT a, b
FROM table1
SetOp
SELECT a, b
FROM table2
```
wehre SetOp is `INTERSECT`, `EXCEPT`, `UNION` or `UNION ALL`.
#### Intersection
```sql
INTERSECT
```
Alternative using `INNER JOIN`
#### Union
```SQL
UNION
UNION ALL (does not eliminate duplicate rows)
```
#### Difference
```SQL
EXCEPT
```
Alternative using `OUTER JOIN` with test for `NULL`
```
%%sql
DROP VIEW IF EXISTS language_view;
CREATE VIEW language_view AS
SELECT first, last, language_name
FROM person
INNER JOIN person_language
ON person.person_id = person_language.person_id
INNER JOIN language
ON language.language_id = person_language.language_id
;
%%sql
SELECt *
FROM language_view
LIMIT 10;
%%sql
SELECt *
FROM language_view
WHERE language_name = 'Python'
UNION
SELECt *
FROM language_view
WHERE language_name = 'Haskell'
LIMIT 10;
%%sql
SELECt *
FROM language_view
WHERE language_name IN ('Python', 'Haskell')
ORDER BY first
LIMIT 10;
```
### Aggregate functions
```SQL
COUNT
MIN
MAX
AVG
SUM
```
```
%%sql
SELECT count(language_name)
FROM language_view;
```
### Grouping
```SQL
SELECT a, MIN(b) AS min_b, MAX(b) AS max_b, AVG(b) AS mean_b
FROM table
GROUP BY a
HAVING mean_b > 5
```
The `HAVING` is analagous to the `WHERE` clause, but filters on aggregate conditions. Note that the `WHERE` statement filters rows BEFORE the grouping is done.
Note: Any variable in the SELECT part that is not an aggregte function needs to be in the GROUP BY part.
```SQL
SELECT a, b, c, COUNT(d)
FROM table
GROUP BY a, b, c
```
```
%%sql
SELECT language_name, count(*) AS n
FROM language_view
GROUP BY language_name
HAVING n > 45;
```
### The CASE switch
#### Simple CASE
```SQL
SELECT name,
(CASE sex
WHEN 'M' THEN 1.5*dose
WHEN 'F' THEN dose
END) as adjusted_dose
FROM table
```
#### Searched CASE
```SQL
SELECT name,
(CASE
WHEN sex = 'M' THEN 1.5*dose
WHEN sex = 'F' THEN dose
END) as adjusted_dose
FROM table
```
```
%%sql
SELECT first, last, language_name,
(CASE
WHEN language_name LIKE 'H%' THEN 'Hire'
ELSE 'FIRE'
END
) AS outcome
FROM language_view
LIMIT 10;
```
## User defined functions (UDF)
```
import sqlite3
import random
import statistics
con = sqlite3.connect(":memory:")
```
#### Row functions
```
con.create_function("rnorm", 2, random.normalvariate)
cr = con.cursor()
cr.execute('CREATE TABLE foo(num REAL);')
cr.execute("""
INSERT INTO foo(num)
VALUES
(rnorm(0,1)),
(rnorm(0,1)),
(rnorm(0,1)),
(rnorm(0,1)),
(rnorm(0,1)),
(rnorm(0,1)),
(rnorm(0,1)),
(rnorm(0,1))
""")
cr.execute('SELECT * from foo')
cr.fetchall()
```
#### Aggregate functions
```
class Var:
def __init__(self):
self.acc = []
def step(self, value):
self.acc.append(value)
def finalize(self):
if len(self.acc) < 2:
return 0
else:
return statistics.variance(self.acc)
con.create_aggregate("Var", 1, Var)
cr.execute('SELECT Var(num) FROM foo')
cr.fetchall()
con.close()
```
| github_jupyter |
# Reminder
<a href="#/slide-1-0" class="navigate-right" style="background-color:blue;color:white;padding:10px;margin:2px;font-weight:bold;">Continue with the lesson</a>
<font size="+1">
By continuing with this lesson you are granting your permission to take part in this research study for the Hour of Cyberinfrastructure: Developing Cyber Literacy for GIScience project. In this study, you will be learning about cyberinfrastructure and related concepts using a web-based platform that will take approximately one hour per lesson. Participation in this study is voluntary.
Participants in this research must be 18 years or older. If you are under the age of 18 then please exit this webpage or navigate to another website such as the Hour of Code at https://hourofcode.com, which is designed for K-12 students.
If you are not interested in participating please exit the browser or navigate to this website: http://www.umn.edu. Your participation is voluntary and you are free to stop the lesson at any time.
For the full description please navigate to this website: <a href="gateway-1.ipynb">Gateway Lesson Research Study Permission</a>.
</font>
```
# This code cell starts the necessary setup for Hour of CI lesson notebooks.
# First, it enables users to hide and unhide code by producing a 'Toggle raw code' button below.
# Second, it imports the hourofci package, which is necessary for lessons and interactive Jupyter Widgets.
# Third, it helps hide/control other aspects of Jupyter Notebooks to improve the user experience
# This is an initialization cell
# It is not displayed because the Slide Type is 'Skip'
from IPython.display import HTML, IFrame, Javascript, display
from ipywidgets import interactive
import ipywidgets as widgets
from ipywidgets import Layout
import getpass # This library allows us to get the username (User agent string)
# import package for hourofci project
import sys
sys.path.append('../../supplementary') # relative path (may change depending on the location of the lesson notebook)
import hourofci
# Retreive the user agent string, it will be passed to the hourofci submit button
agent_js = """
IPython.notebook.kernel.execute("user_agent = " + "'" + navigator.userAgent + "'");
"""
Javascript(agent_js)
# load javascript to initialize/hide cells, get user agent string, and hide output indicator
# hide code by introducing a toggle button "Toggle raw code"
HTML('''
<script type="text/javascript" src=\"../../supplementary/js/custom.js\"></script>
<input id="toggle_code" type="button" value="Toggle raw code">
''')
```
# Types of Computational Systems in Cyberinfrastructure
In this section we will look "under the hood" and cover different types of computational systems that are commonly used in cyberinfrastructure.
## GPUs
<img src="supplementary/gpu.png" width="400"/>
<small>CC BY 4.0 https://commons.wikimedia.org/wiki/File:NvidiaTesla.jpg</small>
- GPUs – Graphical Processing Units – are a very powerful type of processor (currently being used to display this text on your computer screen)
- GPUs make up a very large part of the computational power of many computing systems
- GPUs were originally developed for rendering graphical images, but it turns out that they are very fast for some (but not all) kinds of mathematically-oriented calculations.
## Quantum Computers
- Quantum computers are the newest new thing
- Normal computers work on a very simple principle: you do the same calculation over and over, you get the same results
- Quantum computers are different
- Rather than operating with a string of things called “bits” each of which is a 0 or a 1 like a current digital computer, Quantum computers operate on things called Qbits that are either 0 or 1 with a certain probability.
- So when you run a program with a quantum computer, you don’t get an answer. You get a probability distribution of answers
- Quantum computers are very important for some kinds of challenges but it will be a long time before they matter much to people using GIS applications!
### High Throughput Computing (HTC) Systems
- HTC systems have been around for a long time
- Sometimes certain data analysis problems involved doing lots of analysis (or lots of computations) that can happen pretty much independently
- So a lot of work is done, and then the results are collected
- This is different than the kind of jobs you usually run in parallel on a supercomputer
- A bit of a good way to think about high performance parallel computing as opposed to high throughput computing is this:
- If you care about how long one job takes, you’re probably doing high performance parallel computing
- If you care about how many thousand jobs you run per month, you’re probably doing high throughput computing
## Let's take a closer look at High Throughput Computing Systems in GIS (at Clemson University)
<img src="supplementary/htcs.png" width="400"/>
- HTC systems used to analyze GIS data
- An example problem: calculate the AADT (Annual Average Daily Traffic) through Greenville South Carolina
- **Problem:** calculate all possible intersects between analyzed traffic routes (1.9 million observations) and all the traffic data collection sites that are spread throughout the city of Greenville
- The Clemson University HTC system used a very famous and important piece of HTC software called Condor
- Read more about the HTC system at Clemson at - https://www.clemsongis.org/high-throughput-computing-for-gis
### How High Throughput Computing was used
<img src="supplementary/htc.png" width="400"/>
And you can see how this is cyberinfrastructure: lots of data, lots of data storage, broken up and sent across a network to lots of different computers organized into something called a “Condor Pool.” A “Condor Pool” is what a group of computational systems is called within the Condor High Throughput Computing software system.
These images also from https://www.clemsongis.org/high-throughput-computing-for-gis
### Another example: The Large Hadron Collider (LHC)
<img src="supplementary/hadron.png" width="400"/>
- The LHC is the single biggest physics experiment in the world
- It produces lots of small-ish blocks of data
- The data tell what happened when subatomic particles are smashed together
- Most of the time nothing new happens
- So the data analysis task is to look at a whole bunch of data and determine if anything novel has happened
- PERFECT for HTC – which is a very important kind of cyberinfrastructure
From: https://home.cern/science/accelerators/large-hadron-collider
<img src="supplementary/congratulations.png" width="400"/>
## You just learned a lot more detail about what computational systems can be part of cyberinfrastructure systems
Really, anything that can connect to a digital network and can either produce data or do calculations can be considered cyberinfrastructure if it is put to work as part of “infrastructure for knowledge”
<a href="cyberinfrastructure-5.ipynb">Click here to move on to the next segment where you will learn more about the importance of CI in scientific discovery</a>
| github_jupyter |
<img src='https://assets.leetcode-cn.com/aliyun-lc-upload/uploads/2020/10/11/p1.png'>
```
# 初始状态mask里面的1代表这次选的所有node,然后BFS尝试把所有node连在一起,每连一个node,就把1设置成0
from collections import defaultdict, deque
class Solution:
def countSubgraphsForEachDiameter(self, n: int, edges):
graph = defaultdict(list)
for s, e in edges:
graph[s-1].append(e-1)
graph[e-1].append(s-1)
print(graph)
def get_max_dist(mask):
# 1、检查mask是否可以构成一颗树
# 2、如果可以构成一颗树,返回这个树的最大 distance
# if str(bin(mask)[2:]).count('1') <= 1:
# return 0
# 对于只选择一颗树的情况,最大距离就是 1
if mask & (mask - 1) == 0:
return 0
ans = 0
for i in range(n):
if mask & (1 << i):
queue = deque([i])
temp_mask = mask
temp_mask ^= 1 << i # 求异或
d = 0
while queue:
que_len = len(queue)
for _ in range(que_len):
node = queue.popleft()
for nei in graph[node]:
if temp_mask & (1 << nei):
queue.append(nei)
temp_mask ^= 1 << nei
d += 1
if temp_mask:
return 0
ans = max(ans, d)
return ans - 1
res = [0] * (n-1)
for mask in range(1<<n): # 遍历每一个树选或者不选
dist = get_max_dist(mask)
if dist:
res[dist-1] += 1
return res
import collections, itertools
class Solution:
def countSubgraphsForEachDiameter(self, n: int, edges):
vertices = [i for i in range(1, n+1)]
ans = [0]*(n-1)
graph = collections.defaultdict(set)
for a, b in edges:
graph[a].add(b)
graph[b].add(a)
def checkValid(tree):
tree = set(tree)
start = tree.pop() # 随机弹出一个 节点
tree.add(start)
trav = set()
# 随机从一个subtree的一个节点出发,查看是否能够遍历sub_tree的所有节点
def helper(curr):
nonlocal trav
trav.add(curr)
for nei in graph[curr]:
if (nei not in trav) and (nei in tree):
helper(nei)
helper(start)
return trav == tree
def MaxDis(tree):
# should be the diameter of tree
tree = set(tree)
start = tree.pop()
tree.add(start)
diameter = 0
def DFS(curr, trav):
nonlocal diameter
neighbors_dep = []
for nei in graph[curr]:
if nei not in trav and nei in tree:
neighbors_dep.append(DFS(nei, trav | {nei}))
if not neighbors_dep:
dia = 0
diameter = max(diameter, dia)
return 0
elif len(neighbors_dep)==1:
dia = 1+neighbors_dep[0]
diameter = max(diameter, dia)
return dia
else:
dia = sum(sorted(neighbors_dep)[-2:])+2
diameter = max(diameter, dia)
return max(neighbors_dep)+1
DFS(start, {start})
return diameter
for l in range(1, n+1):
# 从vertices中,返回长度为 l 的组合可能
subtrees = list(itertools.combinations(vertices, l))
for subtree in subtrees:
# 验证每一个subtree
if checkValid(subtree): # 首先验证 subtree 是不是有效的tree
max_dis = MaxDis(subtree) # 如果是有效的节点,计算节点之间的最大距离
if max_dis > 0:
ans[max_dis - 1] += 1
return ans
solution = Solution()
solution.countSubgraphsForEachDiameter(n = 4, edges = [[1,2],[2,3],[2,4]])
a = 1
b = 2
c = a ^ b
c
mask = 3
n_mask = mask - 1
print(bin(mask), bin(n_mask))
mask = 0
if mask & (mask - 1) == 0:
print(mask)
a = [1, 2]
for i in range(len(a)):
a.append(i)
print(i, a)
a = {1}
b = {2}
c = a and b
c
```
| github_jupyter |
```
import pandas as pd
from sklearn.preprocessing import LabelEncoder
from unidecode import unidecode
import re
def cleaning(string):
string = unidecode(string)
string = re.sub(r'\w+:\/{2}[\d\w-]+(\.[\d\w-]+)*(?:(?:\/[^\s/]*))*', '', string)
string = re.sub(r'[ ]+', ' ', string).strip().split()
string = [w for w in string if w[0] != '@']
return ' '.join(string)
df = pd.read_csv('news-sentiment/sentiment-data-v2.csv')
Y = LabelEncoder().fit_transform(df.label)
with open('polarity/polarity-negative-translated.txt','r') as fopen:
texts = fopen.read().split('\n')
labels = [0] * len(texts)
with open('polarity/polarity-positive-translated.txt','r') as fopen:
positive_texts = fopen.read().split('\n')
labels += [1] * len(positive_texts)
texts += positive_texts
texts += df.iloc[:,1].tolist()
labels += Y.tolist()
assert len(labels) == len(texts)
import json
with open('multidomain-sentiment/bm-amazon.json') as fopen:
amazon = json.load(fopen)
with open('multidomain-sentiment/bm-imdb.json') as fopen:
imdb = json.load(fopen)
with open('multidomain-sentiment/bm-yelp.json') as fopen:
yelp = json.load(fopen)
texts += amazon['negative']
labels += [0] * len(amazon['negative'])
texts += amazon['positive']
labels += [1] * len(amazon['positive'])
texts += imdb['negative']
labels += [0] * len(imdb['negative'])
texts += imdb['positive']
labels += [1] * len(imdb['positive'])
texts += yelp['negative']
labels += [0] * len(yelp['negative'])
texts += yelp['positive']
labels += [1] * len(yelp['positive'])
import os
for i in [i for i in os.listdir('twitter-sentiment/negative') if 'Store' not in i]:
with open('twitter-sentiment/negative/'+i) as fopen:
a = json.load(fopen)
texts += a
labels += [0] * len(a)
for i in [i for i in os.listdir('twitter-sentiment/positive') if 'Store' not in i]:
with open('twitter-sentiment/positive/'+i) as fopen:
a = json.load(fopen)
texts += a
labels += [1] * len(a)
with open('emotion/happy-twitter-malaysia.json') as fopen:
a = json.load(fopen)
texts += a
labels += [1] * len(a)
with open('emotion/anger-twitter-malaysia.json') as fopen:
a = json.load(fopen)
texts += a
labels += [0] * len(a)
len(texts), len(labels)
rules_normalizer = {
'ty': 'terima kasih',
'january': 'januari',
'february': 'februari',
'march': 'mac',
'may': 'mei',
'june': 'jun',
'july': 'julai',
'august': 'ogos',
'october': 'oktober',
'december': 'disember',
'dec': 'dis',
'oct': 'okt',
'monday': 'isnin',
'mon': 'isn',
'tuesday': 'selasa',
'tues': 'sel',
'wednesday': 'rabu',
'wed': 'rab',
'thursday': 'khamis',
'thurs': 'kha',
'friday': 'jumaat',
'fri': 'jum',
'saturday': 'sabtu',
'sat': 'sab',
'sunday': 'ahad',
'sun': 'ahd',
'gemen': 'kerajaan',
'camtu': 'macam itu',
'experience': 'pengalaman',
'kpd': 'kepada',
'bengng': 'bengang',
'mntak': 'minta',
'bagasi': 'bagasi',
'kg': 'kampung',
'kilo': 'kilogram',
'g': 'pergi',
'grm': 'gram',
'k': 'okay',
'abgkat': 'abang dekat',
'abis': 'habis',
'ade': 'ada',
'adoi': 'aduh',
'adoii': 'aduh',
'aerodarat': 'kapal darat',
'agkt': 'angkat',
'ahh': 'ah',
'ailior': 'air liur',
'airasia': 'air asia x',
'airasiax': 'penerbangan',
'airline': 'penerbangan',
'airlines': 'penerbangan',
'airport': 'lapangan terbang',
'airpot': 'lapangan terbang',
'aje': 'sahaja',
'ajelah': 'sahajalah',
'ajer': 'sahaja',
'ak': 'aku',
'aq': 'aku',
'all': 'semua',
'ambik': 'ambil',
'amek': 'ambil',
'amer': 'amir',
'amik': 'ambil',
'ana': 'saya',
'angkt': 'angkat',
'anual': 'tahunan',
'apapun': 'apa pun',
'ape': 'apa',
'arab': 'arab',
'area': 'kawasan',
'aritu': 'hari itu',
'ask': 'tanya',
'astro': 'astro',
'at': 'pada',
'attitude': 'sikap',
'babi': 'khinzir',
'back': 'belakang',
'bag': 'beg',
'bang': 'abang',
'bangla': 'bangladesh',
'banyk': 'banyak',
'bard': 'pujangga',
'bargasi': 'bagasi',
'bawak': 'bawa',
'bawanges': 'bawang',
'be': 'jadi',
'behave': 'berkelakuan baik',
'belagak': 'berlagak',
'berdisiplin': 'berdisplin',
'berenti': 'berhenti',
'beskal': 'basikal',
'bff': 'rakan karib',
'bg': 'bagi',
'bgi': 'bagi',
'biase': 'biasa',
'big': 'besar',
'bike': 'basikal',
'bile': 'bila',
'binawe': 'binatang',
'bini': 'isteri',
'bkn': 'bukan',
'bla': 'bila',
'blom': 'belum',
'bnyak': 'banyak',
'body': 'tubuh',
'bole': 'boleh',
'boss': 'bos',
'bowling': 'boling',
'bpe': 'berapa',
'brand': 'jenama',
'brg': 'barang',
'briefing': 'taklimat',
'brng': 'barang',
'bro': 'abang',
'bru': 'baru',
'bruntung': 'beruntung',
'bsikal': 'basikal',
'btnggjwb': 'bertanggungjawab',
'btul': 'betul',
'buatlh': 'buatlah',
'buh': 'letak',
'buka': 'buka',
'but': 'tetapi',
'bwk': 'bawa',
'by': 'dengan',
'byr': 'bayar',
'bz': 'sibuk',
'camera': 'kamera',
'camni': 'macam ini',
'cane': 'macam mana',
'cant': 'tak boleh',
'carakerja': 'cara kerja',
'care': 'jaga',
'cargo': 'kargo',
'cctv': 'kamera litar tertutup',
'celako': 'celaka',
'cer': 'cerita',
'cheap': 'murah',
'check': 'semak',
'ciput': 'sedikit',
'cite': 'cerita',
'citer': 'cerita',
'ckit': 'sikit',
'cikit': 'sikit',
'ckp': 'cakap',
'class': 'kelas',
'cm': 'macam',
'cmni': 'macam ini',
'cmpak': 'campak',
'committed': 'komited',
'company': 'syarikat',
'complain': 'aduan',
'corn': 'jagung',
'couldnt': 'tak boleh',
'cr': 'cari',
'crew': 'krew',
'cube': 'cuba',
'cuma': 'cuma',
'curinyaa': 'curinya',
'cust': 'pelanggan',
'customer': 'pelanggan',
'd': 'di',
'da': 'dah',
'dn': 'dan',
'dahh': 'dah',
'damaged': 'rosak',
'dapek': 'dapat',
'day': 'hari',
'dazrin': 'dazrin',
'dbalingnya': 'dibalingnya',
'de': 'ada',
'deep': 'dalam',
'deliberately': 'sengaja',
'depa': 'mereka',
'dessa': 'desa',
'dgn': 'dengan',
'dh': 'dah',
'didunia': 'di dunia',
'diorang': 'mereka',
'diorng': 'mereka',
'direct': 'secara terus',
'diving': 'junam',
'dkt': 'dekat',
'dlempar': 'dilempar',
'dlm': 'dalam',
'dlt': 'padam',
'dlu': 'dulu',
'done': 'siap',
'dont': 'jangan',
'dorg': 'mereka',
'dpermudhkn': 'dipermudahkan',
'dpt': 'dapat',
'dri': 'dari',
'dsb': 'dan sebagainya',
'dy': 'dia',
'educate': 'mendidik',
'ensure': 'memastikan',
'everything': 'semua',
'ewahh': 'wah',
'expect': 'sangka',
'fb': 'facebook',
'fired': 'pecat',
'first': 'pertama',
'fkr': 'fikir',
'flight': 'kapal terbang',
'for': 'untuk',
'free': 'percuma',
'friend': 'kawan',
'fyi': 'untuk pengetahuan anda',
'gantila': 'gantilah',
'gantirugi': 'ganti rugi',
'gentlemen': 'lelaki budiman',
'gerenti': 'jaminan',
'gile': 'gila',
'gk': 'juga',
'gnti': 'ganti',
'go': 'pergi',
'gomen': 'kerajaan',
'goment': 'kerajaan',
'good': 'baik',
'ground': 'tanah',
'guarno': 'macam mana',
'hampa': 'mereka',
'hampeh': 'teruk',
'hanat': 'jahanam',
'handle': 'kawal',
'handling': 'kawalan',
'hanta': 'hantar',
'haritu': 'hari itu',
'harini': 'hari ini',
'hate': 'benci',
'have': 'ada',
'hawau': 'celaka',
'henpon': 'telefon',
'heran': 'hairan',
'him': 'dia',
'his': 'dia',
'hmpa': 'mereka',
'hntr': 'hantar',
'hotak': 'otak',
'hr': 'hari',
'i': 'saya',
'hrga': 'harga',
'hrp': 'harap',
'hu': 'sedih',
'humble': 'merendah diri',
'ibon': 'ikon',
'ichi': 'inci',
'idung': 'hidung',
'if': 'jika',
'ig': 'instagram',
'iklas': 'ikhlas',
'improve': 'menambah baik',
'in': 'masuk',
'isn t': 'tidak',
'isyaallah': 'insyallah',
'ja': 'sahaja',
'japan': 'jepun',
'jd': 'jadi',
'saja': 'sahaja',
'saje': 'sahaja',
'je': 'sahaja',
'jee': 'sahaja',
'jek': 'sahaja',
'jepun': 'jepun',
'jer': 'sahaja',
'jerr': 'sahaja',
'jez': 'sahaja',
'jg': 'juga',
'jgk': 'juga',
'jgn': 'jangan',
'jgnla': 'janganlah',
'jibake': 'celaka',
'jjur': 'jujur',
'job': 'kerja',
'jobscope': 'skop kerja',
'jogja': 'jogjakarta',
'jpam': 'jpam',
'jth': 'jatuh',
'jugak': 'juga',
'ka': 'ke',
'kalo': 'kalau',
'kalu': 'kalau',
'kang': 'nanti',
'kantoi': 'temberang',
'kasi': 'beri',
'kat': 'dekat',
'kbye': 'ok bye',
'kearah': 'ke arah',
'kecik': 'kecil',
'keja': 'kerja',
'keje': 'kerja',
'kejo': 'kerja',
'keksongan': 'kekosongan',
'kemana': 'ke mana',
'kene': 'kena',
'kenekan': 'kenakan',
'kesah': 'kisah',
'ketempat': 'ke tempat',
'kije': 'kerja',
'kijo': 'kerja',
'kiss': 'cium',
'kite': 'kita',
'kito': 'kita',
'kje': 'kerja',
'kjr': 'kerja',
'kk': 'okay',
'kmi': 'kami',
'kt': 'kat',
'tlg': 'tolong',
'kl': 'kuala lumpur',
'klai': 'kalau',
'klau': 'kalau',
'klia': 'klia',
'klo': 'kalau',
'klu': 'kalau',
'kn': 'kan',
'knapa': 'kenapa',
'kne': 'kena',
'ko': 'kau',
'kompom': 'sah',
'korang': 'kamu semua',
'korea': 'korea',
'korg': 'kamu semua',
'kot': 'mungkin',
'krja': 'kerja',
'ksalahan': 'kesalahan',
'kta': 'kita',
'kuar': 'keluar',
'kut': 'mungkin',
'la': 'lah',
'laa': 'lah',
'lahabau': 'celaka',
'lahanat': 'celaka',
'lainda': 'lain dah',
'lak': 'pula',
'last': 'akhir',
'le': 'lah',
'leader': 'ketua',
'leave': 'pergi',
'ler': 'lah',
'less': 'kurang',
'letter': 'surat',
'lg': 'lagi',
'lgi': 'lagi',
'lngsong': 'langsung',
'lol': 'hehe',
'lorr': 'lah',
'low': 'rendah',
'lps': 'lepas',
'luggage': 'bagasi',
'lumbe': 'lumba',
'lyak': 'layak',
'maap': 'maaf',
'maapkan': 'maafkan',
'mahai': 'mahal',
'mampos': 'mampus',
'mart': 'kedai',
'mau': 'mahu',
'mcm': 'macam',
'mcmtu': 'macam itu',
'memerlukn': 'memerlukan',
'mengembirakan': 'menggembirakan',
'mengmbilnyer': 'mengambilnya',
'mengtasi': 'mengatasi',
'mg': 'memang',
'mihak': 'memihak',
'min': 'admin',
'mingu': 'minggu',
'mintak': 'minta',
'mjtuhkn': 'menjatuhkan',
'mkyong': 'mak yong',
'mlibatkn': 'melibatkan',
'mmg': 'memang',
'mmnjang': 'memanjang',
'mmpos': 'mampus',
'mn': 'mana',
'mna': 'mana',
'mntak': 'minta',
'mntk': 'minta',
'mnyusun': 'menyusun',
'mood': 'suasana',
'most': 'paling',
'mr': 'tuan',
'msa': 'masa',
'msia': 'malaysia',
'mst': 'mesti',
'mu': 'awak',
'much': 'banyak',
'muko': 'muka',
'mum': 'emak',
'n': 'dan',
'nah': 'nah',
'nanny': 'nenek',
'napo': 'kenapa',
'nati': 'nanti',
'ngan': 'dengan',
'ngn': 'dengan',
'ni': 'ini',
'nie': 'ini',
'nii': 'ini',
'nk': 'nak',
'nmpk': 'nampak',
'nye': 'nya',
'ofis': 'pejabat',
'ohh': 'oh',
'oii': 'hoi',
'one': 'satu',
'online': 'dalam talian',
'or': 'atau',
'org': 'orang',
'orng': 'orang',
'otek': 'otak',
'p': 'pergi',
'paid': 'dah bayar',
'palabana': 'kepala otak',
'pasni': 'lepas ini',
'passengers': 'penumpang',
'passengger': 'penumpang',
'pastu': 'lepas itu',
'pd': 'pada',
'pegi': 'pergi',
'pekerje': 'pekerja',
'pekrja': 'pekerja',
'perabih': 'perabis',
'perkerja': 'pekerja',
'pg': 'pergi',
'phuii': 'puih',
'pikir': 'fikir',
'pilot': 'juruterbang',
'pk': 'fikir',
'pkerja': 'pekerja',
'pkerjaan': 'pekerjaan',
'pki': 'pakai',
'please': 'tolong',
'pls': 'tolong',
'pn': 'pun',
'pnh': 'pernah',
'pnt': 'penat',
'pnya': 'punya',
'pon': 'pun',
'priority': 'keutamaan',
'properties': 'harta benda',
'ptugas': 'petugas',
'pub': 'kelab malam',
'pulak': 'pula',
'puye': 'punya',
'pwrcuma': 'percuma',
'pyahnya': 'payahnya',
'quality': 'kualiti',
'quit': 'keluar',
'ramly': 'ramly',
'rege': 'harga',
'reger': 'harga',
'report': 'laporan',
'resigned': 'meletakkan jawatan',
'respect': 'hormat',
'rizal': 'rizal',
'rosak': 'rosak',
'rosok': 'rosak',
'rse': 'rasa',
'sacked': 'buang',
'sado': 'tegap',
'salute': 'sanjung',
'sam': 'sama',
'same': 'sama',
'samp': 'sampah',
'sbb': 'sebab',
'sbgai': 'sebagai',
'sblm': 'sebelum',
'sblum': 'sebelum',
'sbnarnya': 'sebenarnya',
'sbum': 'sebelum',
'sdg': 'sedang',
'sebb': 'sebab',
'sebijik': 'sebiji',
'see': 'lihat',
'seen': 'dilihat',
'selangor': 'selangor',
'selfie': 'swafoto',
'sempoi': 'cantik',
'senaraihitam': 'senarai hitam',
'seorg': 'seorang',
'service': 'perkhidmatan',
'sgt': 'sangat',
'shared': 'kongsi',
'shirt': 'kemeja',
'shut': 'tutup',
'sib': 'nasib',
'skali': 'sekali',
'sket': 'sikit',
'sma': 'sama',
'smoga': 'semoga',
'smpoi': 'cantik',
'sndiri': 'sendiri',
'sndr': 'sendiri',
'sndri': 'sendiri',
'sne': 'sana',
'so': 'jadi',
'sop': 'tatacara pengendalian piawai',
'sorang': 'seorang',
'spoting': 'pembintikan',
'sronok': 'seronok',
'ssh': 'susah',
'staff': 'staf',
'standing': 'berdiri',
'start': 'mula',
'steady': 'mantap',
'stiap': 'setiap',
'stress': 'stres',
'student': 'pelajar',
'study': 'belajar',
'studycase': 'kajian kes',
'sure': 'pasti',
'sykt': 'syarikat',
'tah': 'entah',
'taik': 'tahi',
'takan': 'tak akan',
'takat': 'setakat',
'takde': 'tak ada',
'takkan': 'tak akan',
'taknak': 'tak nak',
'tang': 'tentang',
'tanggungjawab': 'bertanggungjawab',
'taraa': 'sementara',
'tau': 'tahu',
'tbabit': 'terbabit',
'team': 'pasukan',
'terbaekk': 'terbaik',
'teruknye': 'teruknya',
'tgk': 'tengok',
'that': 'itu',
'thinking': 'fikir',
'those': 'itu',
'time': 'masa',
'tk': 'tak',
'tnggongjwb': 'tanggungjawab',
'tngok': 'tengok',
'tngu': 'tunggu',
'to': 'kepada',
'tosak': 'rosak',
'tp': 'tapi',
'tpi': 'tapi',
'tpon': 'telefon',
'transfer': 'pindah',
'trgelak': 'tergelak',
'ts': 'tan sri',
'tstony': 'tan sri tony',
'tu': 'itu',
'tuh': 'itu',
'tula': 'itulah',
'umeno': 'umno',
'unfortunately': 'malangnya',
'unhappy': 'tidak gembira',
'up': 'naik',
'upkan': 'naikkan',
'ur': 'awak',
'utk': 'untuk',
'very': 'sangat',
'viral': 'tular',
'vote': 'undi',
'warning': 'amaran',
'warranty': 'waranti',
'wassap': 'whatsapp',
'wat': 'apa',
'weii': 'wei',
'well': 'maklumlah',
'win': 'menang',
'with': 'dengan',
'wt': 'buat',
'x': 'tak',
'tw': 'tahu',
'ye': 'ya',
'yee': 'ya',
'yg': 'yang',
'yng': 'yang',
'you': 'awak',
'your': 'awak',
'sakai': 'selekeh',
'rmb': 'billion ringgit',
'rmj': 'juta ringgit',
'rmk': 'ribu ringgit',
'rm': 'ringgit',
':*': '<kiss>',
':-*': '<kiss>',
':x': '<kiss>',
':-)': '<happy>',
':-))': '<happy>',
':-)))': '<happy>',
':-))))': '<happy>',
':-)))))': '<happy>',
':-))))))': '<happy>',
':)': '<happy>',
':))': '<happy>',
':)))': '<happy>',
':))))': '<happy>',
':)))))': '<happy>',
':))))))': '<happy>',
':)))))))': '<happy>',
':o)': '<happy>',
':]': '<happy>',
':3': '<happy>',
':c)': '<happy>',
':>': '<happy>',
'=]': '<happy>',
'8)': '<happy>',
'=)': '<happy>',
':}': '<happy>',
':^)': '<happy>',
'|;-)': '<happy>',
":'-)": '<happy>',
":')": '<happy>',
'\o/': '<happy>',
'*\\0/*': '<happy>',
':-D': '<laugh>',
':D': '<laugh>',
'8-D': '<laugh>',
'8D': '<laugh>',
'x-D': '<laugh>',
'xD': '<laugh>',
'X-D': '<laugh>',
'XD': '<laugh>',
'=-D': '<laugh>',
'=D': '<laugh>',
'=-3': '<laugh>',
'=3': '<laugh>',
'B^D': '<laugh>',
'>:[': '<sad>',
':-(': '<sad>',
':-((': '<sad>',
':-(((': '<sad>',
':-((((': '<sad>',
':-(((((': '<sad>',
':-((((((': '<sad>',
':-(((((((': '<sad>',
':(': '<sad>',
':((': '<sad>',
':(((': '<sad>',
':((((': '<sad>',
':(((((': '<sad>',
':((((((': '<sad>',
':(((((((': '<sad>',
':((((((((': '<sad>',
':-c': '<sad>',
':c': '<sad>',
':-<': '<sad>',
':<': '<sad>',
':-[': '<sad>',
':[': '<sad>',
':{': '<sad>',
':-||': '<sad>',
':@': '<sad>',
":'-(": '<sad>',
":'(": '<sad>',
'D:<': '<sad>',
'D:': '<sad>',
'D8': '<sad>',
'D;': '<sad>',
'D=': '<sad>',
'DX': '<sad>',
'v.v': '<sad>',
"D-':": '<sad>',
'(>_<)': '<sad>',
':|': '<sad>',
'>:O': '<surprise>',
':-O': '<surprise>',
':-o': '<surprise>',
':O': '<surprise>',
'°o°': '<surprise>',
'o_O': '<surprise>',
'o_0': '<surprise>',
'o.O': '<surprise>',
'o-o': '<surprise>',
'8-0': '<surprise>',
'|-O': '<surprise>',
';-)': '<wink>',
';)': '<wink>',
'*-)': '<wink>',
'*)': '<wink>',
';-]': '<wink>',
';]': '<wink>',
';D': '<wink>',
';^)': '<wink>',
':-,': '<wink>',
'>:P': '<tong>',
':-P': '<tong>',
':P': '<tong>',
'X-P': '<tong>',
'x-p': '<tong>',
'xp': '<tong>',
'XP': '<tong>',
':-p': '<tong>',
':p': '<tong>',
'=p': '<tong>',
':-Þ': '<tong>',
':Þ': '<tong>',
':-b': '<tong>',
':b': '<tong>',
':-&': '<tong>',
'>:\\': '<annoyed>',
'>:/': '<annoyed>',
':-/': '<annoyed>',
':-.': '<annoyed>',
':/': '<annoyed>',
':\\': '<annoyed>',
'=/': '<annoyed>',
'=\\': '<annoyed>',
':L': '<annoyed>',
'=L': '<annoyed>',
':S': '<annoyed>',
'>.<': '<annoyed>',
':-|': '<annoyed>',
'<:-|': '<annoyed>',
':-X': '<seallips>',
':X': '<seallips>',
':-#': '<seallips>',
':#': '<seallips>',
'O:-)': '<angel>',
'0:-3': '<angel>',
'0:3': '<angel>',
'0:-)': '<angel>',
'0:)': '<angel>',
'0;^)': '<angel>',
'>:)': '<devil>',
'>:D': '<devil>',
'>:-D': '<devil>',
'>;)': '<devil>',
'>:-)': '<devil>',
'}:-)': '<devil>',
'}:)': '<devil>',
'3:-)': '<devil>',
'3:)': '<devil>',
'o/\o': '<highfive>',
'^5': '<highfive>',
'>_>^': '<highfive>',
'^<_<': '<highfive>',
'<3': '<heart>',
}
from unidecode import unidecode
permulaan = [
'bel',
'se',
'ter',
'men',
'meng',
'mem',
'memper',
'di',
'pe',
'me',
'ke',
'ber',
'pen',
'per',
]
hujung = ['kan', 'kah', 'lah', 'tah', 'nya', 'an', 'wan', 'wati', 'ita']
def naive_stemmer(word):
assert isinstance(word, str), 'input must be a string'
hujung_result = [e for e in hujung if word.endswith(e)]
if len(hujung_result):
hujung_result = max(hujung_result, key = len)
if len(hujung_result):
word = word[: -len(hujung_result)]
permulaan_result = [e for e in permulaan if word.startswith(e)]
if len(permulaan_result):
permulaan_result = max(permulaan_result, key = len)
if len(permulaan_result):
word = word[len(permulaan_result) :]
return word
def cleaning(string):
string = re.sub(
'http\S+|www.\S+',
'',
' '.join(
[i for i in string.split() if i.find('#') < 0 and i.find('@') < 0]
),
)
string = unidecode(string).replace('.', ' . ').replace(',', ' , ')
string = re.sub('[^A-Za-z ]+', ' ', string)
string = re.sub(r'[ ]+', ' ', string.lower()).strip()
string = [rules_normalizer.get(w, w) for w in string.split()]
string = [naive_stemmer(word) for word in string]
return ' '.join([word for word in string if len(word) > 1])
from tqdm import tqdm
for i in tqdm(range(len(texts))):
texts[i] = cleaning(texts[i])
import numpy as np
np.unique(labels, return_counts = True)
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.naive_bayes import ComplementNB
tfidf = TfidfVectorizer(ngram_range=(1, 3),min_df=3, max_features=800000).fit(texts)
vectors = tfidf.transform(texts)
vectors.shape
from sklearn.model_selection import train_test_split
from sklearn import metrics
train_X, test_X, train_Y, test_Y = train_test_split(vectors, labels, test_size = 0.2)
multinomial = ComplementNB().fit(train_X, train_Y)
print(
metrics.classification_report(
train_Y,
multinomial.predict(train_X),
target_names = ['negative', 'positive'],
)
)
print(
metrics.classification_report(
test_Y,
multinomial.predict(test_X),
target_names = ['negative', 'positive'],
digits = 5
)
)
text = 'kerajaan sebenarnya sangat bencikan rakyatnya, minyak naik dan segalanya'
multinomial.predict_proba(tfidf.transform([cleaning(text)]))
delattr(tfidf, 'stop_words_')
import pickle
with open('multinomial-sentiment.pkl','wb') as fopen:
pickle.dump(multinomial,fopen)
with open('tfidf-multinomial-sentiment.pkl','wb') as fopen:
pickle.dump(tfidf,fopen)
import boto3
bucketName = 'huseinhouse-storage'
Key = 'multinomial-sentiment.pkl'
outPutname = "v30/sentiment/multinomial-sentiment.pkl"
s3 = boto3.client('s3',
aws_access_key_id='',
aws_secret_access_key='')
s3.upload_file(Key,bucketName,outPutname)
bucketName = 'huseinhouse-storage'
Key = 'tfidf-multinomial-sentiment.pkl'
outPutname = "v30/emotion/multinomial-sentiment-tfidf.pkl"
s3 = boto3.client('s3',
aws_access_key_id='',
aws_secret_access_key='')
s3.upload_file(Key,bucketName,outPutname)
```
| github_jupyter |
```
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
# Setup
%matplotlib inline
```
# Load some house value vs. crime rate data
Dataset is from Philadelphia, PA and includes average house sales price in a number of neighborhoods. The attributes of each neighborhood we have include the crime rate ('CrimeRate'), miles from Center City ('MilesPhila'), town name ('Name'), and county name ('County').
```
sales = pd.read_csv('https://courses.cs.washington.edu/courses/cse416/18sp/notebooks/Philadelphia_Crime_Rate_noNA.csv')
sales.head()
```
# Exploring the data
The house price in a town is correlated with the crime rate of that town. Low crime towns tend to be associated with higher house prices and vice versa.
```
sales.plot.scatter(x="CrimeRate", y="HousePrice")
```
# Fit the regression model using crime as the feature
```
features = sales["CrimeRate"].to_numpy().reshape(-1, 1)
target = sales["HousePrice"].to_numpy().reshape(-1, 1)
crime_model = LinearRegression()
regression = crime_model.fit(features, target)
print(f"coeficients: {regression.coef_}")
print(f"intercept: {regression.intercept_}")
crime_model.predict(features[0].reshape(-1, 1))
```
# Let's see what our fit looks like
Matplotlib is a Python plotting library that is also useful for plotting. You can install it with:
'pip install matplotlib'
```
plt.plot(sales['CrimeRate'],sales['HousePrice'],'.',
sales['CrimeRate'], crime_model.predict(features),'-')
```
Above: blue dots are original data, green line is the fit from the simple regression.
# Remove Center City and redo the analysis
Center City is the one observation with an extremely high crime rate, yet house prices are not very low. This point does not follow the trend of the rest of the data very well. A question is how much including Center City is influencing our fit on the other datapoints. Let's remove this datapoint and see what happens.
```
sales_noCC = sales[sales['MilesPhila'] != 0.0]
sales_noCC.plot.scatter(x="CrimeRate", y="HousePrice")
```
### Refit our simple regression model on this modified dataset:
```
features_noCC = sales_noCC["CrimeRate"].to_numpy().reshape(-1, 1)
target_noCC = sales_noCC["HousePrice"].to_numpy().reshape(-1, 1)
crime_model_noCC = LinearRegression()
regression_noCC = crime_model_noCC.fit(features_noCC, target_noCC)
print(f"coeficients: {regression_noCC.coef_}")
print(f"intercept: {regression_noCC.intercept_}")
```
### Look at the fit:
```
plt.plot(sales_noCC['CrimeRate'], sales_noCC['HousePrice'], '.',
sales_noCC['CrimeRate'], crime_model_noCC.predict(features_noCC), '-')
```
# Compare coefficients for full-data fit versus no-Center-City fit
Visually, the fit seems different, but let's quantify this by examining the estimated coefficients of our original fit and that of the modified dataset with Center City removed.
```
print(f"coeficients: {regression.coef_}")
print(f"intercept: {regression.intercept_}")
print(f"coeficients: {regression_noCC.coef_}")
print(f"intercept: {regression_noCC.intercept_}")
```
Above: We see that for the "no Center City" version, per unit increase in crime, the predicted decrease in house prices is 2,288. In contrast, for the original dataset, the drop is only 576 per unit increase in crime. This is significantly different!
### High leverage points:
Center City is said to be a "high leverage" point because it is at an extreme x value where there are not other observations. As a result, recalling the closed-form solution for simple regression, this point has the *potential* to dramatically change the least squares line since the center of x mass is heavily influenced by this one point and the least squares line will try to fit close to that outlying (in x) point. If a high leverage point follows the trend of the other data, this might not have much effect. On the other hand, if this point somehow differs, it can be strongly influential in the resulting fit.
### Influential observations:
An influential observation is one where the removal of the point significantly changes the fit. As discussed above, high leverage points are good candidates for being influential observations, but need not be. Other observations that are *not* leverage points can also be influential observations (e.g., strongly outlying in y even if x is a typical value).
### Plotting the two models
Confirm the above calculations by looking at the plots. The orange line is the model trained removing Center City, and the green line is the model trained on all the data. Notice how much steeper the green line is, since the drop in value is much higher according to this model.
```
plt.plot(sales_noCC['CrimeRate'], sales_noCC['HousePrice'], '.',
sales_noCC['CrimeRate'], crime_model.predict(features_noCC), '-',
sales_noCC['CrimeRate'], crime_model_noCC.predict(features_noCC), '-')
```
# Remove high-value outlier neighborhoods and redo analysis
Based on the discussion above, a question is whether the outlying high-value towns are strongly influencing the fit. Let's remove them and see what happens.
```
# sales_nohighend = sales_noCC[sales_noCC['HousePrice'] < 350000]
# crime_model_nohighend = turicreate.linear_regression.create(
# sales_nohighend,
# target='HousePrice',
# features=['CrimeRate'],
# validation_set=None,
# verbose=False
# )
sales_noHE = sales_noCC[sales_noCC['HousePrice'] < 350000]
features_noHE = sales_noHE["CrimeRate"].to_numpy().reshape(-1, 1)
target_noHE = sales_noHE["HousePrice"].to_numpy().reshape(-1, 1)
crime_model_noHE = LinearRegression()
regression_noHE= crime_model_noHE.fit(features_noHE, target_noHE)
```
### Do the coefficients change much?
```
print(f"coeficients: {regression_noCC.coef_}")
print(f"intercept: {regression_noCC.intercept_}")
print(f"coeficients: {regression_noHE.coef_}")
print(f"intercept: {regression_noHE.intercept_}")
```
Above: We see that removing the outlying high-value neighborhoods has *some* effect on the fit, but not nearly as much as our high-leverage Center City datapoint.
### Compare the two models
Confirm the above calculations by looking at the plots. The orange line is the no high-end model, and the green line is the no-city-center model.
```
plt.plot(sales_noHE['CrimeRate'], sales_noHE['HousePrice'], '.',
sales_noHE['CrimeRate'], crime_model_noHE.predict(features_noHE), '-',
sales_noHE['CrimeRate'], crime_model_noCC.predict(features_noHE), '-')
```
| github_jupyter |
# Comparing Russell Westbrook and Oscar Robertson's Triple Double Seasons
### Author: Rohan Patel
NBA player Russell Westbrook who plays for the Oklahoma City Thunder just finished an historic NBA basketball season as he became the second basketball player in NBA history to average a triple double for an entire season. A triple double entails having atleast 3 of the stat totals of points, assists, rebounds, steals, and blocks to be in double figures. A triple double is most commonly obtained through points, rebounds, and assists. During the 2017 NBA regular season, Westbrook averaged 31.6 points per game, 10.4 assists per game, and 10.7 rebounds per game.
Former NBA basketball player Oscar Robertson who used to play for the Cincinatti Royals is the only other player to average a triple double for an entire regular season as he did so 55 years ago. During the 1962 NBA regular season, Oscar Robertson averaged 30.8 points per game, 11.4 assists per game, and 12.5 rebounds per game. Many thought no one would ever average a triple double for an entire season ever again.
My project is going to compare the 2 seasons. Since it has been 55 years in between the 2 seasons, much has changed about the NBA and how basketball is played. I want to compare the differences in the way the game is played by examining their respective seasons in order to obtain a better understanding of who had the more impressive season.
```
# importing packages
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# all data is obtained through basketball-reference.com
# http://www.basketball-reference.com/teams/OKC/2017.html
# http://www.basketball-reference.com/teams/CIN/1962.html
# http://www.basketball-reference.com/leagues/NBA_stats.html
# all 2017 okc thunder player per game stats
okc = pd.read_csv('/Users/rohanpatel/Downloads/Per_Game_OKC_2017.csv')
okc.head()
# all 1962 cincinatti royals player per game stats
cin = pd.read_csv('/Users/rohanpatel/Downloads/Per_Game_CincRoy_1962.csv')
cin.head()
# only russell westbrook's points, rebounds, assists, and minutes per game
RW = okc.loc[:0]
RW = RW[['PTS/G', 'TRB', 'AST', 'MP']]
RW = RW.rename(columns={'PTS/G': 'PTS'})
RW
# only oscar robertson's points, rebounds, assists, and minutes per game
OR = cin.loc[:0]
OR = OR[['PTS', 'TRB', 'AST', 'MP']]
OR
# robertson played a considerable amount of more minutes than westbrook
# adjusting per game stats by 36 minutes played
rw_min_factor = 36/RW['MP']
or_min_factor = 36/OR['MP']
RW[['PTS', 'TRB', 'AST']] = RW[['PTS', 'TRB', 'AST']].apply(lambda x: x*rw_min_factor)
RW_36 = RW[['PTS', 'TRB', 'AST']]
print(RW_36)
OR[['PTS', 'TRB', 'AST']] = OR[['PTS', 'TRB', 'AST']].apply(lambda x: x*or_min_factor)
OR_36 = OR[['PTS', 'TRB', 'AST']]
print(OR_36)
# difference between Westbrook and Robertson's per 36 minute stats
RW_36 - OR_36
```
### After adjusting Westbrook and Robertson's per game stats to a per minute basis, Westbrook has the edge. He averages about 8 more points, 1 more rebound, and 1.5 more assists per 36 minutes than Robertson did during the 1962 season. Now I will look into their respective team seasons to see if there are any other adjustments that should be made when comparing the two seasons
```
# 2017 NBA stats
df_2017 = pd.read_csv('/Users/rohanpatel/Downloads/2017_NBA_Stats.csv')
df_2017
# 2017 okc thunder stats
okc_2017 = df_2017.loc[9]
okc_2017
# 1962 NBA stats
df_1962 = pd.read_csv('/Users/rohanpatel/Downloads/1962_NBA_Stats.csv')
df_1962
# 1962 cincinatti royal stats
cin_1962 = df_1962.loc[4]
cin_1962
```
# PACE
There is a noticable difference in the 'pace' stat between the 2017 Thunder and 1962 Royals. The pace stat measures how many possessions per game that a team plays per 48 minutes. The higher the pace total, the more possessions per game that the team plays. The 1962 Cincinatti Royals played about 125 possessions per game while the 2017 Oklahoma City Thunder played about 98 possessions per game. The number of possessions in a game would seem to have an impact on the stat totals of players. It would be estimated that the more possessions a team plays with, the more totals of stats such as points, rebounds, and assists would accumulate. I am going to see how the pace of teams has changed over time and how well that correlates with the number of points, rebounds, and assists that have been accumulated over time to see if Westbrook and Robertson's stats should be adjusted for the number of possessions played.
```
# nba averages per game for every season
nba_avgs = pd.read_csv('/Users/rohanpatel/Downloads/NBA_Averages_Over_Time.csv')
nba_avgs = nba_avgs[['Pace', 'PTS', 'AST', 'TRB', 'FGA']]
# pace values after the 44th row are missing
nba_avgs = nba_avgs.iloc[:44]
print(nba_avgs)
# scatterplots of stats against number of possessions
fig, ax = plt.subplots(nrows = 4, ncols = 1, sharex = True, figsize=(10, 20))
ax[0].scatter(nba_avgs['Pace'], nba_avgs['PTS'], color = 'green')
ax[1].scatter(nba_avgs['Pace'], nba_avgs['TRB'], color = 'blue')
ax[2].scatter(nba_avgs['Pace'], nba_avgs['AST'], color = 'red')
ax[3].scatter(nba_avgs['Pace'], nba_avgs['FGA'], color = 'orange')
ax[0].set_ylabel('POINTS', fontsize = 18)
ax[1].set_ylabel('REBOUNDS', fontsize = 18)
ax[2].set_ylabel('ASSISTS', fontsize = 18)
ax[3].set_ylabel('SHOT ATTEMPTS', fontsize = 18)
ax[3].set_xlabel('NUMBER OF POSSESSIONS', fontsize = 18)
plt.suptitle('STAT TOTALS VS NUMBER OF POSSESSIONS (PER GAME)', fontsize = 22)
plt.show()
```
### It seems pretty clear that the more possessions that a team plays with, the more stat totals they will accumulate. Pace seems to predict the number of shot attempts and rebounds very well just by looking at the scatterplots. Assists and points also increase as pace increases, but it seems to dip off towards the higher paces. Robertson played with a very high pace. I will perform a linear regression for assists and points.
```
import statsmodels.api as sm
from pandas.tools.plotting import scatter_matrix
y = np.matrix(nba_avgs['PTS']).transpose()
x1 = np.matrix(nba_avgs['Pace']).transpose()
X = sm.add_constant(x1)
model = sm.OLS(y,X)
f = model.fit()
print(f.summary())
```
### P-Value shows that pace is a statistically significant predictor of points and R-squared shows that about 74% of the variation in points comes from the variation in possessions which is pretty significant. It makes sense to adjust Robertson and Westbrook's points for pace
```
y = np.matrix(nba_avgs['AST']).transpose()
x1 = np.matrix(nba_avgs['Pace']).transpose()
X = sm.add_constant(x1)
model = sm.OLS(y,X)
f = model.fit()
print(f.summary())
```
### Possessions also significant in predicting the number of assists
```
# adjusting both player's per 36 minute points, rebounds, and assists per 100 team possessions
rw_pace_factor = 100/okc_2017['Pace']
or_pace_factor = 100/cin_1962['Pace']
RW_36_100 = RW_36.apply(lambda x: x*rw_pace_factor)
print(RW_36_100)
OR_36_100 = OR_36.apply(lambda x: x*or_pace_factor)
print(OR_36_100)
print(RW_36_100 - OR_36_100)
# westbrook's per 36 minute stats adjusted for 1962 Cincinatti Royals pace
RW_36_1962 = RW_36 * (cin_1962['Pace']/okc_2017['Pace'])
print(RW_36_1962)
# robertson's per 36 minute stats adjusted for 2017 OKC Thunder Pace
OR_36_2017 = OR_36 * (okc_2017['Pace']/cin_1962['Pace'])
print(OR_36_2017)
# difference between the two if westbrook played at 1962 robertson's pace per 36 minutes
print(RW_36_1962 - OR_36)
# difference between the two if robertson played at 2017 westbrook's pace per 36 minutes
print(RW_36 - OR_36_2017)
# huge advantages for westbrook after adjusting for possessions
```
# CONCLUSION
PACE MATTERS. Pace is something that almost never gets mentioned in any basketball debate when comparing across eras. Per-game statistics are what is mainly used when comparing players to see who is better. But as we saw, the number of possessions a player plays with varies largely between teams and plays a major factor in the amount of total statistics he is able to accumulate. Pace has steadily slowed down as the NBA has gotten older other than the slight recent surge the past few years. In Robertson's time, a team playing with 130 possessions per game was not unusual, but 130 possessions per game today is unheard of. It now does not seem like a coincidence that 1962 was also the the season that Wilt Chamberlain mythically averaged 50 points per game and scored a 100 points in a single game
This goes to show how impressive Westbrook's 2017 NBA season actually was. Many recognize the greatness in that it's only the 2nd time ever someone has averaged a triple double for an entire season. But people do not realize how much better it was than Robertson's season. As most people do, just glancing at the per game statistics makes the seasons look similar and might even give Robertson an edge. But as we broke down the statistics and adjusted for pace and minutes played, Westbrook averaged about 13.5 more points, 3.25 more rebounds, and 3.65 more assists than Robertson did per 36 minutes and 100 possessions. Robertson's averages of about 20 points, 8 rebounds, and 7.5 assists per game after adjusting for pace and minutes is about what you'd expect from a regular all-star player today. The numbers are still very good, but not what they seem by just looking at the box score. The typical NBA box-score is limited because it does not take into account many factors that lead to the accumulation of statistics as per-game numbers can be very misleading.
| github_jupyter |
## Loading dataset
```
from beta_rec.datasets.movielens import Movielens_100k
from beta_rec.data import BaseData
dataset = Movielens_100k()
split_dataset = dataset.load_leave_one_out(n_test=1)
data = BaseData(split_dataset)
```
### Model config
```
config = {
"config_file":"../configs/mf_default.json"
}
# the 'config_file' key is required, that is used load a default config. Other keys can be specified to replace the default settings.
```
### Model intialization and training
```
from beta_rec.recommenders import MatrixFactorization
model = MatrixFactorization(config)
model.train(data)
# @To be discussed
# model.train(train_df) # Case 1, without validation, stop training by loss or max_epoch
# model.train(train_df,valid_df[0]) # Case 2, with validation, stop training by performance on validation set
# model.train(train_df,valid_df[0],test_df[0]) # Case 3, same as Case 2, but also evaluate performance for each epoch on test set.
## Note that the best model will be save automatically, and record the model-save-dir.
```
### Model trainng
```
model.test(data.test[0])
```
### Load a pre-trained Model, and Predict for new dataset
```
from beta_rec.recommenders import MatrixFactorization
config = {
"config_file":"../configs/mf_default.json",
}
# model_dir = model.config["system"]["model_save_dir"] # default saving dir for current model
model_dir = "/home/zm324/workspace/beta_rec/checkpoints/MF_default_20200912_173445_gccwwj/mf.model" # a specfic model
model = MatrixFactorization(config)
model.init_engine(data)
# this is necessary, since we cannot intialize a model before we get the the numbers of users and items
model.load(model_dir = model_dir)
scores = model.predict(data.test[0])
scores
scores.shape
```
## Model tuning
```
## Before using tune, you need to install beta_rec to your local python lib
## E.g. go to the project folder, run:
## python setup.py install --record files.txt
## if something wrong, you can uninstall by: xargs rm -rf < files.txt
from beta_rec.datasets.movielens import Movielens_100k
from beta_rec.data import BaseData
dataset = Movielens_100k()
split_dataset = dataset.load_leave_one_out(n_test=1)
data = BaseData(split_dataset)
config = {
"config_file":"../configs/mf_default.json",
"tune":True,
"max_epoch":2
}
from beta_rec.recommenders import MatrixFactorization
model = MatrixFactorization(config)
tune_result = model.train(data)
tune_result
```
### Note that ray version should be 0.8.5.
```
import ray
ray.__version__
ray.init()
```
| github_jupyter |
First, run the command to get the embeddings of the upcoming weeks. We also note this model (to get the embeddings as well as one first model to learn the embeddings). This model will then be used to genereate embeddings for all upcoming weeks.
```
day = 20160701
! export CUDA_VISIBLE_DEVICES=3 && python main.py --data real-t --semi_supervised 0 --batch_size 128 --sampling hybrid --subsamplings bATE/DATE --weights 0/1 --mode scratch --train_from 20160101 --test_from $day --test_length 350 --valid_length 30 --initial_inspection_rate 20 --final_inspection_rate 5 --epoch 5 --closs bce --rloss full --save 0 --numweeks 1 --inspection_plan direct_decay
```
This will store the embeddings in the directory './intermediary/embeddings/embd_0.pickle'
These embeddings will be domain-shift calculated between weeks
Calculate Domain shifts between weeks
```
import pandas as pd
import numpy as np
import torch
import random
import pickle
import ot
with open("./intermediary/embeddings/embd_0.pickle","rb") as f :
processed_data = pickle.load(f)
```
Matching the datapoints together
```
datestart = '16-07-01'
from datetime import datetime, timedelta
date = datetime.strptime(datestart, "%y-%m-%d")
datelist0 = []
for i in range(51):
delta = i * 7
new_date = date + timedelta(days=delta)
datelist0.append(new_date.strftime('%y-%m-%d'))
enddate = datelist0[-1]
data = pd.read_csv('tdata.csv')
data = data[(data['sgd.date'] >= datestart) & (data['sgd.date'] < enddate)]
data.reset_index(drop=True, inplace=True)
```
Concat weekly data together
```
full_data = []
for i in range(len(datelist0) - 1):
data0 = data[(data['sgd.date'] >= datelist0[i]) & (data['sgd.date'] < datelist0[i+1])]
start = data0.index[0]
end = data0.index[-1]
xd = []
for j in range(start, end + 1):
xd.append(processed_data[datelist[j]].reshape(1, -1))
full_data.append(torch.cat(xd, axis=0))
def domain_shift(xs, xt):
M = torch.cdist(xs, xt)
lol = torch.max(M)
M = M/lol
M = M.data.cpu().numpy()
a = [1/xs.shape[0]] * xs.shape[0]
b = [1/xt.shape[0]] * xt.shape[0]
prep = ot.emd2(a, b, M)
return prep * lol
arraycom = []
for i in range(81, len(full_data) - 1):
stack = []
for j in range(60):
indices = torch.tensor(random.sample(range(full_data[i].shape[0]), 500))
indices2 = torch.tensor(random.sample(range(full_data[i+1].shape[0]), 500))
xs = full_data[i][indices]
xt = full_data[i+1][indices2]
print(domain_shift(xs, xt))
stack.append(domain_shift(xs, xt))
xd = np.mean(stack)
print(xd)
arraycom.append(xd)
```
arraycom will contain domainshifts information through 100 weeks
This part is about extracting the best probability of resampling through each week
```
def extraction(lol):
for i in lol:
if 'Pr@5.0' in i:
xd = i.split(', ')[0]
value = float(xd.split(':')[1])
return value
probs = []
for i in range(0, len(datelist0) - 2):
highest = 0
day = datelist0[i]
for j in range(1, 10):
ratio = j * 0.1
bratio = 1 - ratio
lol = ! export CUDA_VISIBLE_DEVICES=3 && python main.py --data real-t --semi_supervised 0 --batch_size 128 --sampling hybrid --subsamplings xgb/random --weights $ratio/$bratio --mode scratch --train_from 20160101 --test_from $day --test_length 7 --valid_length 30 --initial_inspection_rate 20 --final_inspection_rate 5 --epoch 5 --closs bce --rloss full --save 0 --numweeks 2 --inspection_plan direct_decay
value = extract(lol)
if value > highest:
smt = bratio
probs.append(smt)
```
probs should contain the best resampling ratio. Now we come to find the coefficients. First transform the probs result
```
news = []
for i in probs:
news.append(np.log(i/(1-i)))
newcom = np.reshape(arraycom, (-1, 1))
from sklearn.linear_model import LinearRegression
reg = LinearRegression()
reg.fit(newcom, news)
reg.coef_
reg.intercept_
```
| github_jupyter |
# Part III: Tradeoffs
In this notebook, we'll explore the pros and cons of a few variations of the Babble Labble framework.
1. Data Programming or Majority Vote
2. Explanations or Traditional Labels
3. Including LFs as features
As with all machine learning tools, no one tool fits all situations; there are always tradeoffs.
Also, note that the relative performance of each of these variants can vary widely across applications and different sets of labeling functions, so take the results of any single run with a grain of salt.
## 0. Setup
Once again, we need to first load the data (candidates and labels) from the pickle. This time, we'll also load our label matrices and training set predictions from Tutorial 1.
```
%load_ext autoreload
%autoreload 2
import pickle
DATA_FILE = 'data/tutorial_data.pkl'
with open(DATA_FILE, 'rb') as f:
Cs, Ys = pickle.load(f)
with open("Ls.pkl", 'rb') as f:
Ls = pickle.load(f)
with open("Y_p.pkl", 'rb') as f:
Y_p = pickle.load(f)
```
## 1. Data Programming or Majority Vote
When it comes to label aggregation, there are a variety of ways to reweight and combine the outputs of the labeling functions. Perhaps the simplest approach is to use the majority vote on a per-candidate basis (effectively making the naive assumption that all labeling functions have equal accuracy). While simple, this is often an effective baseline.
As is described in the [VLDB paper](https://arxiv.org/abs/1711.10160) on Snorkel, in the regimes of very low label density and very high label density, the expected benefits of learning the accuracies of the functions with data programming decreases. When label density is low (i.e. few LFs and/or low coverage LFs), there are few conflicts to resolve, and minimal overlaps from which to learn. When label density is very high, it can be shown that under certain conditions, majority vote converges to an optimal solution, so long as the average labeling function accuracy is better than random.
Because many applications of interest occur in the middle regime, and because the data programming label model can effectively reduce to majority vote with sufficient regularization, we tend to use data programming for label aggregation.
```
# TEMP
from metal import MajorityLabelVoter
print("MajorityVoter")
mv = MajorityLabelVoter(seed=123, verbose=False)
_ = mv.score(Ls[1], Ys[1], metric='f1')
```
And indeed, for our sample set of labeling functions, we see that data programming does indeed outperform majority vote (69.7% vs. 50.8%).
## 2. Explanations or Traditional Labels
We can also consider when it's worthwhile to just use traditional (manually generated) training labels vs weakly supervised (programatically generated) ones. Since the weakly supervised training set will almost by definition not have perfect labels, if you have a large number of ground truth labels to train on, then use that! Where weak supervision makes more sense is situations where labeled data is sparse and/or hard to collect, or when you have the ability to create a much larger training set out of unlabeled data (e.g., 100 "perfect" training labels may not perform as well as 100k "good enough" labels that were automatically generated from labeling functions).
Other aspects to consider:
* static vs dynamic: If the data distribution shifts over time or the task requirements change even slightly, a hand-labeled training set can quickly depreciate in value, as it no longer accurately reflects what you want your model to learn. If your training data is automatically generated, however, you can modify or add a small number of labeling functions to "reshape" your dataset in the appropriate way; no tedious relabeling required.
* label provenance: While we often treat training data creation as a black box process, in actuality, there are often bugs even in manual label collection (e.g., crowdsource workers of varying quality, systematic biases, etc.). We've written about how to debug training data in these blog posts [1](https://dawn.cs.stanford.edu/2018/08/30/debugging2/), [2](https://dawn.cs.stanford.edu/2018/06/21/debugging/).
Finally, most training labels are of approximately equal value. That is, we'd expect a classifier trained with 500 randomly selected labels to achieve approximately the same performance. But it's worth asking:
**What is the value of a labeling function?**
It depends. A labeling function may:
* Label one example: `Label 1 if the candidate ID is 8675309`
* Label one distant supervision pair: `Label 1 if X is "Barack" and Y is "Michelle"`
* Label a whole database-worth: `Label 1 if the tuple of X and Y is in my known_spouses dictionary`
* Label based on a feature (1 or 1000s): `Label 2 if the last word of X is different than the last word of Y`
And it isn't just quantity (coverage) that matters; a labeling function that contributes to many labels may be "worth less" in our application than one with lower coverage but higher accuracy. And one that captures a new type of signal not reflected in our current set of LFs will also have relatively higher value than one re-using the same type of signal (e.g., the same keyword list) over and over. The upshot of this is that we can't simply say "An explanation/labeling function is worth this many labels."
Here we train our same discriminative model using 1000 traditional labels (100x the number of explanations in our sample set).
```
from metal.contrib.featurizers.ngram_featurizer import RelationNgramFeaturizer
NUM_LABELS = 1000
featurizer = RelationNgramFeaturizer()
Xs_ts = []
Xs_ts.append(featurizer.fit_transform(Cs[0][:NUM_LABELS]))
Xs_ts.append(featurizer.transform(Cs[1]))
Xs_ts.append(featurizer.transform(Cs[2]))
X_train = Xs_ts[0]
Y_train = Ys[0][:NUM_LABELS]
from metal.tuners import RandomSearchTuner
from metal.metrics import metric_score
from babble.disc_model import LogisticRegressionWrapper
search_space = {
'C': {'range': [0.0001, 1000], 'scale': 'log'},
'penalty': ['l1', 'l2'],
}
tuner = RandomSearchTuner(LogisticRegressionWrapper, seed=123)
disc_model = tuner.search(
search_space,
train_args=[X_train, Y_train],
X_dev=Xs_ts[1], Y_dev=Ys[1],
max_search=20, verbose=False, metric='f1')
scores = disc_model.score(Xs_ts[1], Ys[1], metric=['precision', 'recall', 'f1'])
```
For this particular set of LFs and this particular set of manual labels, the 10 explanations resulted in a better classifier than 1000 labels (69.3% vs 63.2%). The exact multiplicative factor for any particular LF set will vary (and is not linear, as both collecting more manual labels and collecting more labeling functions experience diminishing returns after a point).
## Including LFs as Features
Once we have collected user explanations, there are a number of ways this extra information can be used. In our paper, we described using these explanations as functions for generating training data. Another option is to use them as essentially hand-crafted features, treating the label matrix as a feature matrix instead. Not surprisingly, these features tend to be highly relevant for their respective tasks. However, as we described in Tutorial 1, there may still be good reasons for not including them. For example:
* We may want to make sure our classifier generalizes beyond the signals described by the explanations.
* We may want to capitalize on representation learning, using the larger training set generated by using them as functions.
* We may be in a cross-modal setting, where the features we have at training time are different than the features that our classifier will have access to at deployment time.
Regardless, we find that even in situations where we do want to include the labeling function outputs as features, we can usually achieve additional quality by using them as labeling functions as well, thanks to the larger training set and the access to additional features relevant to the task at hand.
### LF as features only
First, we consider using the labeling function outputs as our only features.
```
import numpy as np
from data.sample_explanations import explanations
candidate_ids = [exp.candidate for exp in explanations]
indices = []
for c1 in candidate_ids:
for i, c2 in enumerate(Cs[0]):
if c1 == c2.mention_id:
indices.append(i)
break
X_train = Ls[0][indices, :]
Y_train = np.array([exp.label for exp in explanations])
from metal.tuners import RandomSearchTuner
from metal.metrics import metric_score
from babble.disc_model import LogisticRegressionWrapper
search_space = {
'C': {'range': [0.0001, 1000], 'scale': 'log'},
'penalty': ['l1', 'l2'],
}
tuner = RandomSearchTuner(LogisticRegressionWrapper, seed=123)
disc_model = tuner.search(
search_space,
train_args=[X_train, Y_train],
X_dev=Ls[1], Y_dev=Ys[1],
max_search=20, verbose=False, metric='f1')
scores = disc_model.score(Ls[1], Ys[1], metric=['precision', 'recall', 'f1'])
```
As expected, these hand-engineered (or shall we say "natural-language-engineered"?) features get us pretty far. But in situations where we do want to give our discriminative model access to the labeling function outputs as features, this approach can nearly always be trumped by combining the two uses for labeling functions--using them to make the larger training set, and then also providing them as features.
### LFs as features and labelers
```
from metal.contrib.featurizers.ngram_featurizer import RelationNgramFeaturizer
featurizer = RelationNgramFeaturizer(min_df=3)
featurizer.fit(Cs[0])
Xs = [featurizer.transform(C) for C in Cs]
from scipy.sparse import hstack, csr_matrix
Xs_new = []
for i in [0,1,2]:
X_new = csr_matrix(hstack([Ls[i], Xs[i]]))
Xs_new.append(X_new)
from metal.tuners import RandomSearchTuner
from metal.metrics import metric_score
from babble.disc_model import LogisticRegressionWrapper
search_space = {
'C': {'range': [0.0001, 1000], 'scale': 'log'},
'penalty': ['l1', 'l2'],
}
tuner = RandomSearchTuner(LogisticRegressionWrapper, seed=123)
disc_model = tuner.search(
search_space,
train_args=[Xs_new[0], Y_p],
X_dev=Xs_new[1], Y_dev=Ys[1],
max_search=20, verbose=False, metric='f1')
scores = disc_model.score(Xs_new[1], Ys[1], metric=['precision', 'recall', 'f1'])
```
## Conclusions
This concludes the tutorial!
If you'd like to stay up-to-date on the latest tools we're working on in the weak supervision Snorkel ecosystem, we post regular updates to the landing page at [snorkel.stanford.edu](https://hazyresearch.github.io/snorkel/)
| github_jupyter |
<img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAAgAAAADhCAYAAAC+/w30AAAABmJLR0QA/wD/AP+gvaeTAAAACXBIWXMAAC4jAAAuIwF4pT92AAAAB3RJTUUH4wEeDgYF/Qy0kwAAIABJREFUeNrsnXl4G9W5/z8zWrxNHDt29j0kBEgQMYQdwtqW0CKgtHS5XShtb1vfbrSltOpduro/StfbW3VvaaG0QEtApQ1Q9n1JIlDCEkL23fES2+NFsjTz++MckUHYia0ZW5J9vs+jR7JkHc3MOXPe77uDgoKCgoKCgoKCgoKCgoKCgkIBYSbCA75WUFBQUFBQGAfCX5EABQUFBQWvoalLUNQkwAD+AuwGvmCEYt3qqigoKCgoeAFdXYKixqnA24ELgXnqcigoKCgoKAIwPjBZPvcDvepyKCgoKCgoAjA+UCuf04Ay/ysoKCgoKAIwTlDjIAA96nIoKCgoKCgCMD5QrQiAgoKCgoIiAOPYAmCEYpliOjBVo0BBQUFBEQCF0bEAFJXwN0IxzETYr0iAgoKCgiIACt6jUj73FpOQlcL/XKDfTIR/aoRiGKGYmi0FBQUFRQAUPLYApLKCt4jwHfm8TE2TgoKCgiIAYwJFZM6uks9dxXRdzET4ZOAMIAncUWTXTEFBQUFhCPCrSwBmIqwBc4H5wFojFOvM+rkLjAr5nCyG6+S4Hl+Tz/uA3+R8pqCgoKCgLAAlgzLgv4E7gXcXifAHEQNgA31FpP0vBc6Sb99ihGJdSvtXUFBQUASgVGEBBxA+9w8BM4rkuCrksRVTGeBPICoU9gPfU0tHQUFBQRGAkoURiqWA24AMsAI4tkgOrUw+20VwjTAT4fnA2XLd/N4IxQ5mP1NQUFBQUASgVLEN4QIAuMpMhCuKwLTtkxaAziK5Rhc7yNF31JJRUFBQUARgLFgBWoG75J8fAKYUgWZbXegDcPj+a4G3AkHgLiMU26F8/woKCgqKAJQ0HILsSeA5+foTOZ8V4nhs+ShYGWAHCToROFe+/nERBUoqKCgoKCgC4E7IGaHYZuBh+XajmQhXFkLI5fymTYEbAZmJcDnwNoRF4hkgroS/goKCgiIAY80KsArYDUwEriqUFaDIUA98UL7+DUVSmEhBQUFBQREAL60ATwEJ+fZXBtDIxxsx0hG+/2nAZuAZIxSz1IpRUFBQUARgLOL3CLP7DDMRvnwcC38Q1SI/Ld9aDbzi+ExBQUFBQRGAMWUNuB1R5tYHfHa8Cn9p+TgeaADagQeMUKxfBQAqKCgoKAIwVrVegP+TzyEzET5lHGu9X5XPLwCPSoKkFoqCgoKCIgBjTvvPvvwVItitFvjweBN8svLfVOCdiLK/DxuhWJsy/SsoKCgoAjCmrQBGKNaNiHjXgLPMRHjBeLECOM7x0/L8W4Bb1MpQUFBQUARgvFgBvi+flwArx4sVwHGOH0HUIXjWCMU2jZfzV1BQUFAEQGE/8BdEMOBbzUR48njR/s1E+N1AHZBGWEIKbv1Q7gcFBQUFRQBGCxngp/L1hYiI+DEt6Bwa/oeBcqDZCMXuLqT2n70eMi6hykyEa8xEWFPLU0FBQUERgJEUhi8DjwGVwEpZFne0hHB2jiaMsvZ/nIPs/LxYtG8zEa4D7kCkJB6jVqiCgoKCIgAjiQ7gJvn6Q8DUUfztdIG0/0sRlf8Afl1I7d+h+ZcD30RUJQSYpJamgoKCQokTgFztslh8vTIbwAKeAF4FpkgrgDZKx9iDiMIPjuI5TwDOlr95txGKNRdyPsxEOLsePgV8Ur79LeBpFROgoKCgUMIEIFtVzkyE/WYiPNnxdzFdp1cQZXABvjTKv60BZaNIwpYDy+TrHxRS+3dUHFwJfFeu1xuB7xuhWEbdvgoKCgolTACksA8CX0CU3/13MxHWiiHdzNEgyALuQ2QFHAVcNErHd1ASgMBonStwGjAd2A6sLQJiOB+4TZKgx4GvG6FYZyGJiYKCgoIiAN7BB8yUx/NL4OJslHcRWQLuB9bL15FR+k1bEoCKURK6MxHZDgA3A30FFv7lwEOAIQnJV41QbLsy/SsoKCiMAQIgN/te4CfAs/LtvwMrHBaCYjjGlDyuHkRlwKWjQFCyFoDRigGYB5wnX99phGL9BRT+GnAXMBdRkvk7Rij2uGpEpKCgoDCGLAByU9+CCPJaL4XePWYifGExkACHwPkTsFe+/lrOZyOBlHyuHOnrbybCAeACee0fAXYVUPgDfA94i/zo/4xQ7NdK81dQUFAYQwQgK0Dl5h8HPgFsRRSh+YuZCIeLgQTI42sF/oEwzV9kJsKzRtgK0CGfR8MFUAG8X75eDTQXQvhLXI2I+teA3xuhWGQAMqagoKCgUOoWAOemboRiTwHvQwTc1QE/KxYSIPF9wEL4pf9jhH8rawGYMArX/zhgMdAKPGeEYtZoXmsHETwLkeZXBfwLERyKMv0rKCgojEECkCvcjVDsGYQv2gRmAVEzEb64kBqgIyNgJxAD/IiaADUjeExx+dw5kpq3xAfl8zpgQyGusZkIT0dUHpwBbAY+aYRiB5XwV1BQUBjDBGAAEvAycDoiAGwmcIuZCJ+bI7RGW0BlX35HPs8HLh/BY/oGsBR4z0gTGwcBWGOEYs2jSbYcMQi/kedrAh+TcSFK+CsoKCiMdQLg3Oyl1rcBOAvYDUxEBAaei/ANjzoRyBIUIxRbi8iRr0akLAZGyD1hG6HYi0Yo1j2S52omwm9BuBn2AY+O1rXN+Y1vAxcD/Yh0v4cLSfYUFBQUFAEoMBEwQrEEcCWwCVEM5kHgnYWqGOjQRr8rn0+Xj5H8rZHWgrPa/25EsZ0R17pzzPqfBL6MCK78DfCLAf5HQUFBQWE8EACn9meEYk8iGvEkpPZ/O/A5MxH2FYIEyN+7F1GcZiZwoZkI+0tYWCWANcDNRihmjvT1dAp2MxF+N/C/8qP7gK8ZoVi6lIR/Q2PTqP/eaP9mqWE0r4+aC4VSRdH3VM8RFkuA3wGnIEzFPwb+xwjFekdTYEgB6UOkq/0UuB5oMkKxZJ4asJfadD7f9wELgG1GKNY/ktfSQeowE+HzgT9KEvUyosTyjmIX/g2NTcSjkYHeDwJzEEGMUxEdC2sR7pUqROBopeO+s4EkkEEUmOqVz13y0SEfrYiiUC3xaKQv3+Mby8J+sPNtaGyaLdfXFGCyYz4q5cPPGwtt9SG6cKbkXHQjYlJMRDBum5yTNqA1Ho2Yaj4UFAEYHUsAZiI8DdEQ5m1y47wF+IwRinWMtuAwE+EKRKW6ZqBtsN8+3HGZiXA1ItMhd4MK5sxPSgqFTikQ9gG7jFCszQ0pyP2/kb6GDtfNUkS54RPkeZ1ihGKvFKvwH2gTb2hsWoLonbAckUZZj6inUI5wVwURfRwCHNnaZsv1nJECKC1Jbr+c++xzUs7/HmmB2glsk4/N8WjEGg+CaJD5mIGIGco2tJqeMx9lci788nEkZOcikzMf2bnIPjoQhbN2ysd2OR9b4tFIz+GsBuOZGJT6emxobNLi0YhdytdRK5WLnUMCKoCfAR+RHz8IvMcIxVoKKUBy4xJyj8NMhE9AxAucJDeoBVJI6HIuss/aYYRE9mHJ5xSwA2HGX4cw5T9thGLpkbI4eHCdpgJ/kCQO4DSZ+ll0fn/nzdXQ2OQHLkJkZbwVUQvCL61BvlE+NGuAhy3JwUZpUdkAvAisj0cjXWNhE8493obGptPlfKwEZst7yDdEAT/S82FJK8Im4CX52ABsiEcj+wotvIAocJkkNJq0QCU9/BlbzkW1fI1jn/t5PBr5dqmSgOxxNzQ2XYWIXzLkR2mp0OgeX8cq3tgUTkdUpW2MRyPPjWkLwCBCJIBIk/uqfGuz3JS3OKL1R1XwO/7OCoRZwNuBS4FzRnlT2oDwqf9dEoNeIG2EYnYhSIGDHBmINsP/Lj96H3ArIuOh2AR+thPjYuDzwHsZ4bLMI4iDkiTG5WMtsEVqt1lhlZGbWtFsyk5NWb72IdwsH0FUi6wv0flIIcqerwOel8T9ZSmMrey8xKMReyTmo6GxSZeKw8wCXoPaeDRysIQtABWIJnFnFPAwPhyPRv44rghAjqb9BeC/EWmC3cA7gQeMUCwzmCY+goK/DuH/fYcUbMcXySXrAe6WgnYt0GKEYt0FmK8goofCf8uP/gv4f7nWiiLRLOsRDamuQZiVxyo2SELwPPCYG21ihMlYLcLN8jng3WN4Pl4DXpBz8jjwbDwa6R2Ba6sj3BSzC3iuH41HI78rVStUQ2PTmQg35rwCHs6H4tHITfl+2V+Kd0iOhv9DMxHeh8gjnw/cA3zVTIR/nu0bP1KariM48VjJAi+XGn+xoRKRSnklIl7hDjMR/hew1gjFto+kRSBnzC86hP8vgR8VQ8R/jqCpkQTuUwVm9qOFpfLxQXn+1YO5Cwqk9ZchrGcflet3rGOhfFwh/74IkXE0FnEFIqi7pOAgKw0FtqC4hl6qB57ja78F+ACii50G/D/gF2YivMBrC4AzRc5MhE8yE+HvSc36N0Uq/HMxBZF3fxuiuuI3zUT45BxC47nwNxPhrwHZfKk7EOl+3UUm/LNZCTeOE+GfizSHfJkFnQ8p/JciUkTvGCfCfyCUjeFzO6WhsamsRGMAAogA5oAiAAUmAVKIPClJwE/lx+8D7syWD3Yr2HKCEGeaifAvgL8B11I8pv7hwCeF3H8Bt5qJ8K1mIrzCKyKQI/y/AmTv8oeBTxihWGuRCf/vIsx5lzD6AX3FAhsRK1Bw7aqhsenfgL8iYkWqxul8dCNid8YqqoALS/TY55Tovj92CECusDJCsV3AV4CPI4Jojgf+aSbC1+ZaDYYjyHKEWQQRuPMxRArgWMB8qWHFzER4tZkIh/K9XgMI/88C/4lwQ8SBy7LZGsWgaTY0Nk1qaGx6EOGemM44x0j4m4c6H47X35VWmMXjfDq8jsovNgQR5b9LppiS4zjnSgtASUMbqytLFpn5nUNI3we8ywjFunI1+iGOtwxh5j9pnGw+NyECLFtySNZQr5cGfBj4FcJMthFYnq00WCjNP0frXyS1zBAKAN3xaMQYzaCsnPnQEe60d6mpAERtgXfHo5GnR+C6F0MQIMCaeDRycilNirx2n0EUois0XAUB6mPtjnFo7A8C5yLSNPoRKYIbzUT4wmwJ4SONIV9Xm4nwfwBPjSPhDyIobB/wBTMRnuC0CBxJe5fC/63A76XwfwFYUWjhLzXc7E18ghL+b5465zUa5U3VAP6lhP8b0IdIFxzLmNLQ2HRyKVkBEBln546Fiz/mCIARijmzBLYZodhbgO8jiiZMR2QJfMNMhGsGEkRZASXHOBbRn/7/ENXExht8iJz9+81E+Pxs18MhuAZ0YBHCp/wE8M7RbjE8mLYpn48Ffq2E/5vQXghrjEzx+ytwvpqCcUcAJjNCzdRGmACsUASgyImAw8wfQdTtf0AKta8NNIE5vutLEZHy71f7EKcAdwLfkqWYDyvIjVAsgwio+xBwlRGKbcle30IKfylsJktSd7Ka1jehpwDzYQA/4VBVSIVD6EfEMo1lVAAnQmmURZbFwY5F9PlQBKCESMA9CLP2f0pt49XDCP8vIvz9S9Ue9DomANcBq7LZAodzBxih2EEjFLvZCMVeyw2kLAQcm8uvEHnlCm9G5ygLfx1RaEmR7PFrAQA4pqGxaUGJHKuP0kj3VgQgV1M1QrG9wA2ICP5Xc4W/mQjrZiL8beBblG6J0ZHGacBdZiL81SG6A153qRRS+5fP1yNqnysMjFGpDukgY2+VpNKnLv2AyDaFGutYLB9FHQfgOLYxs4fo4+VOclgCUkYo1mGEYlZObn8Zoprg1xBmKYXBUYNwB9yRjQsodFrfELTNcxCRuwqDo3cU52Ua8D3Gb47/UJBEuAHGw36yJIccFh3ksS2ixKv/OeEfL3fSEaL+yxF54F8t1PHZaNi2Lp812TpLG+C/AM1GE/+Jrllo2NlPRhM+ROnjx8xE+L1GKLYtS7SKpZufQ/iXISrKVRx5HsCv2ZQFLGxbo7dfz39KD2lwGcejFRFsZyJ87lkfb0AKQ00eZ6V8rpbXOtslUj/Ms+6S1JujNC8+4NOMfiGV3PlII9weLdL60cGhrnXZGzDbha1azkcVh6olDnVONBfHO9ZjALI4raGxqS4ejbQWsyKB6DUzZjBuCMBAFgGpuQaALyHM/iMKDRtNswGNZKaMZKacpBWkN1PJwVQtHakazLRBb6ZSfubHtg/tHQE9Q7kvSbmvB8PfTXWgg9qydir9XQT1FGV6knJfH7qWfp1MjAJOBe42E+FPGKHYE6PdiXEIjB1EtcZjDve/GVvDr9vMmJDkjDkdfODEPaTSPq66fQk9QycBFqLXQgui095aRMe3V4Gtg/WGH+IGpCHiMGrko1Y+6uRjsnxMRQQoleU8yh3Pg5ncR1TbdGyixyIsbaOBduAAsBNRwOsF4BVgSzwaaXd5PlWSGEx0zMck+ZiCcCPWyzmpQhS+GWg+gkcgLOOCAMj1W5QEwLGXjF8CMNDGXmz92/M4j+tGWvhrmoVPy9CXruJAcjKtyTqa+6ZyoG8Kbck6uvoNNKnJa1K7ZxDx/brubx96LvelqC1ro66slcllB5hU1kJtsI3asnaCepKM7cOyR9TbswS42UyEP2+EYncVw7pwaP/HIMpCBwdU0TMauqZx3BSTU2Z38LajWzhuWie2BXu7yqnw23T3H1GFywDPAI8CDwJPxaMR8whCcFjnEY9GbKmtdiLauB7pe35JDOod5CD7mCKFVY1DeNUh+tWPxib601FYAusRtTseQHQ53OvlfMjz6ZaWg71D/O5Eef3rcuZiinzPSe4mIwoBjeVSwE7MBBY0NDa9UqxugIbGpjpkxsJYwbBVRDMRPgrRoepZIxR7rtROeIAa9U2MUEVEDYugL01bsp5NnYvY2TOblr7JHEzVkLE1fJrlEPj5w0kILFunwpd8nRDMqNjNPGMrdWUHyNg+MvaIxlu1AF80QrE/ymJAdoFJgIbodfCNXMGfyujowPKZXVy48AAnzupk8WQTXbfY1V7F6o2TeWTrJF5qrjzSzzyKyBh5KB6N7MpXsIyAlj1UgVTrEDrr49FIywgTskuAkVwUryIyPe6LRyPr3Qj7AsxHpWMuJgHN8Whk4wgdV7FUAnTih0AkHo0UZfnjhsamj8p7vZjgqhJgPgTgdkS71GapidwP3GKEYptKwSrgMP1/VC646pH4nYDeT1d/NevaTmFL13w6UhNJWsHXffYjCRuwpcZf5ktR5e9mWsU+jq9Zz/wJr5G2RpQIdALfMEKxHxaKBDiEzSJE/YLjsp+lMsJte9acDt55/D6OmdzFlAlJyvwWbd3l3PL8DB7aUsuujjKSaR2fPuhcWYigwjvi0ci+Qgv+QgisPMd/gZErwPQb4IZ4NPJqqcyHs/XxaM5JkRKAVxA1R7qKac4c+8m9iMyVcU0APoso3JFFv3y8hKi9f5MRipnFSAYcwv8c4A+MQDOfbEDehoPLeLblFMz+qkIE6L3JQqBj4dfT1JW1cebkJ5hrbMYauTiBHuALRij2y0LNv9zgrgJ+C5C2RJDlyqNbee8Ju1lQ103AZ1HmtzjYW8ZN62Zy10uT6e7XSVvaka7KDkQqUCIejWTGguAfpQ30EkStf6+zbNLAZ4GfqzkoaQIAsCgejbxWhNcriCiNXjuWCMCwHMNyM/9fRITy1cDjUuEsB5YDUaDDTIRXmYnwuTK1Ts8do1CQwn8OwiQ8Ip38DqYmcdv2f+PePW+lq98ouPB3kpJ+K8C+3qn8dccV3LHjStpT9SN1fJXA92R2wKjOe/Z3LJuJwFWaBkbQ4n2hZlZ/ZA3ffNvLLJ7Shd9n0WKW8eNHj+LtN57EH9ZNozPpI3Nk4b8euDAejcRxRGgrwTM4HNfmE3hfUrsfuCwejfzcqVErjCq+DZyNN7Ed4WKaR8dxXOTB2n0GkTl1DaNUdMtTAuCI8O41QrHfG6HY2VKQXgc8B+yXm+JlwEMIf1zETISPMRPh2uwY2Y16tMmATPf7HB5XgtOw6beCxNtO5vebP8ru7un4teIN3tWw2dy1gD9svoo1LafTk64aCbdENXCDmQif51w7IyX0c7s7njP/4DGL6nrP/szpu7jzQ2v5ynmvUl/VR0dvgE0tBj94ZCHv+lMDv183nVQG9KHxoI3AVfFoZJPS+Ie3gTY0NoUQaX9eMs408K54NPIPR7Ckuuijj1XxaORx4HrgZZdjFVWUvWM9XcTg2RpDQTvw3Xg0cieiTHpPMZzfsNMAczU6IxTbh2i2830zEV4uGdwKRNrVHESE/beAO81EeJUkChuNUMxyWBVGXCuUvxFGtLj1jkFpNi19U3j8wAo2dR6FX0ujlUCTZZ+WwQYe2ncOr5kLOX3y48yt2oZPS3uZMTAL+KGZCP+bEYq95PU8Oxs3yb/L5Lo7qaV766fqKvsxKpL0JYXQ33jA4MHN9Ty0pYZkWqPcL1wAQ0Qb8JV4NLJOaZl5baBvB6Z5PPwX49FITJGxgiPgEHIPItI888WZDY1N1fFopLPIzvE03FWsbAMelq9nUCQp+HkfhHMzd2zEa4A1ZiJcBZwlzULnAWdIq8BlwIvAg2Yi/C/gXiMUS40kEXD4/echuvp5rEkv4vHmsznQV1/UWv/Axw9lviR7e6bwj12XsKz2eU6sew7D3+VlkOAySQLeY4RiHV7M8wCCfxaik9wKYIWu2Ytm1/TQl/bx2ObJPLezhjW7q1m/z8AGArpFRWBYFg8b+KVk77mCTeHIVoBKuQcEPRz2j8Av1FwUFdnraWhsehb4D5dDvR34cxGt39MQqZr5wkbEC3XIv61iOTdPWIjTKiA35m7gXuBeKXiPkaadKxE540uA9wKvmInwXcAfjFCsJceygJfHhkgNmuyd5m+xvr2BJ5rPpDtdiU8r3YJdPs2i3/KzpvUkDiQnc87Uh6gra8byjgS8TVqJPu6BFce53sLApcBJwEKgSgM6+wLWfZum6I9vq2FrWwXN3UEsGwK+vO+7V4Cvy81ACZyhb5zZaxXCkYnhAbYAP4pHIyl1lYsOLwOvyfsxX7wX+HOh7zXH75+JqNOQLzKIbKSig6fVYXKtAvK9bbIT3zVSG/wPeQNPlhaCbwPrzET4Z2YiPNvLxjHZYzAT4U8Bb/GMNWlp1rWeygN7z6cnU4GuWSV/12ZjALaZc7lr5zvZ1zvL67iAj5mJ8MeGGwuQSwjNRLhBrpUtUgu8GjgBUWntqcqq1BUfv2Pp3dGnZ/HMzokc6A6gazZ+3dW5fDEejaSU8B+2Rph9uQTwstvbH+PRyPPZTVqh6AiA24JS5xeDZcfx+yfiLgDQAu4Y8wRgMKuA/LsbkXYSlexwJfC0vLCzgUZgk5kI/8lMhBfkCoB8j8FMhOuA//FOW06zpvUMHtp3DjbaiOf0F4IIHExVc9u297KrZ67X5/dzMxFedCQS4PxM/u8CMxH+tpkIb0aUc/0UMB9Rxa4d+AFwdNXxa87Qjrrnjl0d5eck0zq6ZnsRj7E2Ho2sVvt63lpUlbTQeClg7hpgk1YoDtJnAs+Dq42joqGx6a1Fsn4XAUe7HObJwSqDjlkCMJBVwPHaNkKxe4xQ7HSgAfgLohGHD9EbfLOZCN9uJsINiJQyhkMIcv7nO3jU2lfXLJ5vP4WH9q0oKq3ftm0qggFqqrzJsNKwSdsat217D5vNxV6SAD9wm5kIawORABnNHwAmmonwMjMR/rqZCL8CbEbUjl+AiJ5tk4z67UYoNskIxb5khGKbNG0PDY1NJ0pi4BW+roSNK0wGTvZwvGey2r9C0eIJREaYG7l0SZFYeI7BnTsDKd+KEqPeDjgneAsjFHveCMXeh3AP/C+wAZHb+y6p7d1iJsIXmYlwvUMjHNJvmYnwKYiqha6d2bpm8WrncTy67+yiE/66rnHBCQu46KRFnmUgaICuZfj7zkvY2LlEtj3zhAgcD3xzkLmsAD4pNYi4tNwsBg4igkdvl5/PNUKxK4xQ7J/ZdZSTr+sV9gP3KjOzK9TLe9sLtAOrikQwKAyOpxhif4TDbD9F4QYAliLKM7vBHcU6UXohf9wpAGSswBeBixF1Be5H+E4uBVYDN5qJ8FVmIlx/pOIyclwNUQnOde9mXbPY3TOHR/evoN8urgaKmqaxfOFMjpszhQXTJjFjUjWW7Z3Z3sbm/r0X8rIkAR7AB1xtJsKnDvBZLaJYzDxEV7CHJSlsBC42QrErjVDsJulOGtDKhIexHoh83dIP8CgQZLW5xXiX8tQM3KOsMUU958SjkS5E10U3987khsamhgKfSz2iNLEbPBePRg4UK2HVC30AA8QK7DRCsR8BH0ZkDayS//p24JfAHWYi/DkzEa4YiAg4Xp8stX+XAtams7+Gx/avoCNVXVQ+f9uG+VNrOWvJXPw+MZUXLjsKy/LuGDUgZQV4bP8KtpgLvbJ+zAA+bSbCRo4VoAVRp6EReDfwQSMU+5wRiv3ZCMV2HI70yVKzPilwvMJdQEYJm7wRRORPewEL0WUxpbT/ksDdQJ8b0QBcUGBrzxQP1u+fipmwFo06OwAR2AP8zUyEH0T0Hvgiwi90NiKo6CNmIvxTRAphOkf7D0jC4LrOddoKsKblFHb3zEQvslS/iUYZF598NBXBwOvvzayr5oR503l+214CPm/4nYaN2V/FI/vPozrYQV3wgBclhN8F3Ab83VFhMgXcl/uPA6UADoJleFdqdiewXQl/VwggSoR7gYwUKgpFDMf9cjeQJCeGaxioAE4vlPCUpGMh7otX3VnM86UX2wFlYwQcRKAd0XL1MrkgHpeL6gREbv81AwwzC9ED3r0U6J7LurYTi074a8Blpx7HxMo3y7tzT5hHRcCP7aErQNcsWpM1PLzvApKWJzK2HJEaWOcgbm8Q+nnUhFgGlHl0yk9TJPW6SxFyAw0ggny9gI2oLaLM/6VBBJJyr3aDBQ2NTQsKdArlwIUux1gLHFAEwIVFQL62AcsIxZ6W/QcuQjRlaQd25QgNXZptFrk9hmSmnHv3rCw64W/ZNueGFjCrfuBOxhMqyrhg2VGkM96inH6aAAAgAElEQVS6r32axeau+cRbT/aqiVAYOD4r7HMzRoYq+B0mwiV4V21uLWCi4EYTnOYhIXstHo2YyvxfUrjd5fdnS1JfCDeAFwTg74iAdkUAvCIEUlDca4RiIeAoIxT7c45fuAwRSOZa4310//mY6cqi8vtnLIuF0+s4/ZjZhzl2jWNnT2bxrHoylrfHHtRTPN58Jnt7Z3sVD/B5t0LboRF6GXD2SjwaSSuB484Q4OFYDyvtv+SwCneBgHUIS28h5n0K7noaAPwrHo0oAjCCloGOAT6bi8tufz4tzfbuBWw4uKSo6vtbto1RHuTik45s3KgsC3DGsXOZWFnmaVYAgE9Pc+/ut5HMVHhBji4F5rmt/tjQ2FSGR/UeEClMzWr/do3jPRzrKXU5S4j5iWwAE3jM5VAnyGj80cYlLr+/AYd1WhGAESYFDivAB9yMpWGTtoI8tn9F0Zn+MxmLFUvnUz3Egj9zJk/kpIUzX88Q8AqiWuBEnjhwNgGfJwT3k4DbdsEzEZHDXmAXIg1RaZzu4GX9/7XqcpYOHPfNH91yCWQw92hY4xy/8W6XQz2MyGpSBGCUcbWbL/v1fjYcPIG25KTiEv6Wxfxpkzh65vDI8KmLZzG7fqL3C0ez2NhxNLu653rRCOmjWSLnAtMR/QC8sgC0qm3cNTxLyYxHIy+ry1mSuMvl9+cAR40WGZfpxDXAqS6HeiYejfQoAjC6VoAGKQjy1mx7M1W82HEcGbvYLo3G0rlTmVAxPHe536dz8fKjCfp9nh9RX6acta0no2uu3QDVZiK80uUYUz0kAAcQ1QcV3GlRR3k05M7R0gAVPF8HB3HvBjhLtpQeLbzd5fdfQ3QQLfo1OyYIgMN07Mps49MyvNZ1NAdTtUV1frZtU1NVxuTq/O6BWqOCC5ct9DQtEERe1t7eaWzvnu+FFeADLr8/mfxzjnPREo9GMii40aIMPCjBLbFptDRABW/XASII8G8uhzoXmDCKh+7W/L8BeLUU1uyYiQGQcOVE7reCbOuaR0+6wpPjylgWi6bXMaOu2tU4lp1h3baHWbf1ybzHWLZgGgtn1HmaFaABnalqNncu9CItMK8a/g6GPQlRPMQLXtOqNE7XmOPhWNvU5SxZ2MjyzS5wAlA/ivej206EL8Sjkc5S2D/GjAvATIQn4cLnqGsWzX1T2dc33bNmP5Zls2B6LeFTjqEimH922v7OLTy5eTXfi/2MF7ZtyHuct5+8GKM86HGvANjdM5O2ZL3bjICqQfoDDEXLANFHwIviBL2IboNK4yweArBTXc6StgK0As+6HOrC0ZBXsg2xm0pn+yihjJWxFAR4nitBZms0902lPVnrSd6/ZdvUGhXMmFTNpAkVnLVkHv3p4ROL3n6TNdtWY9tpdrc3c0MsSm+yNz9LSXmQ809Y4GmvAEGcptDcN8Wt+PUhTH35wisTYS+iNbWCO0z1cKy96nKWNDo9sAJcJgX0SAn+7MvLXQ61D3iyVBSIsUQAznBzPn2ZCnb3zMLyqOedZdksmlHHlBoRl3bSwhksmF47rOp8Pj1AfPt9tHbvxaf7qQiW8/jGNdz2ZP7lpY+eWcdR0yd5WiUwZQXY3TObVMZVPR8fcCYMPx2wobEpiHcpgElFADzBNA/H2q8uZ0lbAVKI0tpucC4wYaSEqmPcc8hflbGA52U3RGUBGA3kdP/L+3x6MpXs7p2FjvvYL9u2mTShgiVzp+LTdakpa1x04iL8Pn1I9gVd87GjdT2vNa/Drx9q9lMeKOPHq39Dd193XsdWEQxw0lEzCAzxOIYkubUMu3pm05dx5YLXgKNySwIP9bTwLgMgBXSj4BaTPRxLpWSWKBya9VYg4XK4t4zwsZ7kct32IlrXl0z8UMkTAJn+58dF+h+A2V9NW1+tFylt6LrOwul1zJn8xvz7SRMqWbFkHv3pI5EMDctO8/SWGFoOF9U0DbOvl98+cHPexzezvpq5U2o9cwXomsX+3sl0Z1wr4Qb5xXGU4V0XwBTQg4JbeFlIo11dzpLV/rMvtwNrXA737pEQro7xzsedJbEP2bGyVOKHxooLYCEuIsAtW2d370w0j4L/aqvKWbF03ps1ZV3j+PlTmVVXfVjh69f9rNt+Lz2pLgayRvl9fv78ZP5FcyZUlLFgWi0+XfNsAmw09nbPxHZXP6GS/Jo4BT0kAGnJ5BXcocbDsVRXxhK3AsSjkV5ENUc3Wsc7RkK4OsY7y+U+8lw8GukppeyhsUIA5rqZOMvW2d87zZPof13XWLF0HpVlgQE/n1hZzmmLZxPwD2yC1zUfbd172NwcP/yO2NfNA4lH8j7OOVNqqDUq8CohQNcs9vZNdxtDUQnMz9MCUOHRWupXFoCiIwAHVUrm2DAIIFwBee8PDY1NK0aIpCwA5rkc5i+lpP2PJQIwGxdtRy10DvRNQcMdAUhnLI6bPYXj5kw57P8dPaueRTPqB+TCNhYbdj9GMn14JdS2be594eG8j3VKdRUTKio8XEgWzX3T3FoAyiWZG24goB/Re94LZBBuAAV38LJym6ZSMktY6h+auzjuazq8K2tV8Mo6IXGilCNu9o07S21uxgoBmO5GAKStAAdTE135/23bpraqnLcsO3L106Dfx6mLZ2FUBN/AAXy6n/0dW9nb8RrWEciIbdts2Pkq2PmRFps0O1rXk7a86VapaxZtyVoy7paUjoweH2YgYADv+s6nEZkACu7gVVpmBoqoH7eCGyLQB6yT91i+WOmllu0YZxmilki+eCoejXSUmqVqrBCAKbjoKd/ZX4Nlu/OHJ9MZ3nriIirKhsZDZtZVc8L8aeiOKL/+TJLtresxkx1oQzClm33dbNy9Oa/jPdjdwdptD9Bm7vZsEtKWTne/68ZD1WYiPNx16fNqLds2mfiOmiERAJfdC8c6gh6N068uZenDIRjvx11Mx9SGxqbjPT62yUDI5TA3leK8+Et5UTlSxvKuAqdh09Ffg+ZC+89YFotn1jN/2vAI5LlL5/PKzgO0mb1omk67uYvtrS8NSfgD9Kb6eGXPayyeNfy4udaudsxkN89t+weXnPBpbA+ULE2zOZiqpq5sv5vSwFXARIYX+X1EC4Bti0BFW5g/sAd4T9OgptzW9vzXg/WTv/eOqmRar3SsLV3+TlISji1GKLYpz7TF8QCv0jK7lAVgTGj/TgLQQf5ZImWIsuHrZXChK1Iivz8XUW7YDf7qpWVCEYAhwLHxuvI3dvcbrqr/WRactnjW8DvuaaI87x8feh7LSrGr/RU6+9oI+IamPPWkenlp96tcyvAb6bWZB+lLJWnu2sPm5rUsnLocy3YXA6Fh05NxbfmtRKTitA+gbesDrAGrKpjx96VFwQXtdTIiXtvYVPqhrjJNndHH5Mo0kyr7mVLVx6SKDHXyuaain+ryJMBpaUvbkUwf2aBgJsIhIxRb77GW5JOPQgm9fo82Ma8sAFYh95iGxiYd75oa5YN0PBoZEwRICtxMQ2PTE4iAOy3PdXUhcIPbder4/gLcla5+EhmoOooEwJL7oav7w8/YgKvNpjdTnjcByFgWC6bVMqUmP8E3e/JETlwwnYdffFEW/Rn6lPRn0mzZvwPbstD04VnA28x2elK9+PUgz21bzZy6JQT85a46Bop2yhWg2ZC/S+VNEf1S+M9G5OkGHcJxYs/6S7Rt7S8e+8Le6lqfbjExaFFdnmZieYpJFRlqK1MEA9KKbMujfN0CoJGxNPotjbSl0ZX0o2mky/2ZToTv2ZIP23GjZa0BQbx3ofmBq4F/QwQijubGr0nSdTVgejSeJwKwwBaA98r5CIzy7+oIU/k3gefHmBXgb8D7XBCreQ2NTbPj0YjrHhGyzfCZHmj/9ggK/wyiOFmPfLTINbEOeGhcEwDpL3bF0FNW0NV2tWhGHRMq8ucg55+wgH/EV9Pe3UxZYOiR+Zqmsb/jAJv3b2Ph9AXD+s1Ws52eZA+appFM97B2+z2cufBdpG0XLlcNUpZfat55YyBzfgD4OPBfb6LBtsb8Sd0cO7UT5w/3p/30pXUO9vnoM4Mk0xq9/T5SGZ3etE6y30dfWqc75cdM+ulM6nQlAwR99itfOW/TV3v6fb22TR/C5N+PKPKhyUcl0GWEYq967AIIAFcAKwp4S/1YajRu4VVdhu4CE4BLgIsL+Pt/GysEwIF/SkGWr7lwMqL0+60eaN3VUrFwg3s9XqPtQLN87Ae2AC8DLwGvxKORzgEsK+4JQLH7M7NBV14fo4U/r/mzbZvqynKmT3Jn9g76fZx4VB2r1qYpG4aeoaGxp30/m/ZtHT4B6DpIX3+KoD+IpulsbVnPgskNTJu4gIyVf5CuB3UFBkrpywCPIRqKZCerF7B1jb7dHeWT4nsmntmVDFT2pXV6+zV6Un560z56+3V6Un56+jXMVIDefh0zpdPbH6CnXyNjZd0FNpoGts3er37qx3cPdT2OwP1SyAwEL7Vtr7IyNA+tCfnOh13AYyioC8RrSGGVamhsegRZ2CcP1AKnALe6jQFAZB0tdXFKa4H9wzwOnUMVB01gsxTyWxEVE3fIx854NHLgMNcx17KSPwHIbmZmIlyHSIko45BvvYxDptdyxwatyb/9g2gAQxFnNiLQZ6AbLPt+ErjXCMW2FVPktW1D/YRK6ia4S3nWNI0LQ+fyx0f/xu72Zvy6b8jfa+/uYOPu11jZcMGQf68/3U+r2f6Gnb4/k2TDrkeYPGG2XJ8FU7retOEboZhlJsIPSE0ou6b6ASqmdyb/869nnr69vWJpf0avTEuTfnZpafB6OeVsXICGDZpF+cD2ryFbk1Tw3xHvawV1Hd8Ah7C6yQUBAFjS0Ng0KR6NtLkYQ8dlF1lE7f/husxagC9JId8sNf42oC0ejViHE/YDXEfX2lbuZvY7DrVl1QbYlHM36IH+Hi5ztwZZ7JaDTPzUTIS/aoRinldp08jkRfBtYGJVOUaF+3inWXUz+Nj57+c/b/0e/uDw3AAbdr5Cc8cBpkwcWh+Lrt4u2rra35CCCLC/aztbDrzA0VNPdlEfwPv9SpJTCxiQDYc+9TZT17LC3iboU7JHQaHIcZfL7y+Wj6fyMYFL7V8H3GqUT8SjkeFa7fYBv4xHI+kjafdeCvvBGNBAmnu1fEyQDwOR1lMlrQIVjke5tBCUSe2sB2Ge7UH474by6OVQ/fUyxyP7GxqiJ3jvAFqYhcvKbUG9L6/vBXw6kyZ4U01P13RWHHsapx61jHRm6CZ4n+5j/Y6X2dM29I6pXX3dtJrtaDnTn0z3sOXA83T1tQ05FTFX9pf70m7dABlyCoUcSdP2abamoaCgUEKWgCTCd54v5gHH5Csg5XcqHcpuPljPMCsbSuFuZYX/QIWDRjOV8E0uAOBSRFpEtiVqtjuaLk0dabkpF0TNGsTv6spvWulLYtvasI0AAb/OhHLvgoPnTJ7NxSdewIZdr2INUYrqms6egwdIbH+R4+cei28I7oPOnk5aOlvRcsSmrvnYc3ATu9pfYdHUk/OQ/xoVvl5cukv78yB0liQOCsWDbkQ9B7dQ3G7s4s/A21x8/+SGxqbb4tFIvu27h/rblpQxSbk3HQReAG5nmL0NRsqU75oASP8/RijWh4g2dCOQR0X4O/7ucDNuVcAcduEa27bpTraRzEzCZSfiN2Dlsgv4Z/xBnt/+Ero2tCyzskCQe55/iEtPWcnEyuoj/n9Hj0lzZ+ubXABCiNu8svdpptcsxCgbXmEjG41Kf5fbS5Bk+N348iENgxoU1L7sCdIejVOpLuWYxX0uv386UO9QVoejhQO8Z5B/6UL45duBVkSA3nr5SMSjkdZBxis5vCEMKh9BPpqBULm/5fi7WQqBYavjNhq1wXZhARgGMnaaJ167iwUz3sryhcd5do5Taiazctl5vLJnM6n00Pzwfp+fZzY/z/bmHYTmHTmg9UDHATp7TarKqwawKPg4YO5kS3Oc0OzzhqWA2bZGTaDTTRVApPA3hz2N3gUf6A2NTfpAwTgKBSEAPmUFGHuQpu8ORB57voF4y4AZiMj5YWnhDY1N2ZTbpPz+TvnYIQX+FmBLPBrZfQQSQSk3qhorhYD2SA0wL3v8hEAHfn3o+72maexue5nNzRv46zMZTpizlOPnLfHsZN5zxmX8+Yk72dG6d2jHg0bGsrntibuOSACS/Um2Nu9AO4yrwO8Lsn73o8yrD1FdUT/k4w7oGS8sAN0Mv1Z4Gu9qxvsQGS99jE94JWx7PRqnAoUxBymEU4gOem4i8U9vaGx6brCAusPpk4jaItlI/GageaBxsn760QrMG02MlWZAO3ARB+DT+qkNtmMNsZWtZVms2X4vfj3Aq3u28MnffpXP/OYr/P251bR0HHC/45VV8I4TL3yTj/5wKA8E+duae2jtOnxWTF9/ks3NO/AdpnKghkZ/Jsmabavx6UPjVJatUxM8iE9z7YrvMkKx4ZrzR4IAjFd4lWXT5dE45coCMGZJQBp4CnfWu5VAWR5d+Dri0chv4tHIffFo5Pl4NLJnsMC8eDTCWG1HPVYIwBY3GpuuWUyt2Ic1hMvh0/28tOcxzGQ7mqZjI+rqP/DSU0Ru/R4Xfvt9XHHD1Ty0/lFXJ/TBFe8ecgxA1iqRSqf59b8O35SqN9XHq3s3HzFY0Kf72dn+MvsOvjak47DRqC87gKa5spzbiGyP4cLrGIAyxi+8IlLdHo0TVARgTGMv8LSL718IVAxHQGcj8QcS9mNJux9PBGCjG83Fh8XMyt1HtABomkZfyiSx62F07Y0C1LZt0pkMyXSKDbs38e+//gpfv/V6unrzU4RqjBqWz1+CZQ1doAb9AW5+/A7Mw/xmV08Xm/ZuHVK2gKbprNm++k3nOpgFYGpFs6umSpLEbc/jeym8q6AXRAWeeYFOD8dSBGDs4gDw+BAUg2w5blOShtXA/0gLwLD2/qyAX3N5mHg0wprLw6y5PBxcc3m4fM3lYW3N5eOnzfdY6AWAEYqlzUR4O3BUPpuFpllMq9iD/wjma58W4Pmdfz9ikRxd09B8fn73yO34dB9fuvTTVASHXxr9bSecx2Mb1xIcRqOftGXxw7//gv++8to330W2TWL7i2TsoYXpaWi0mnvY0bqB2XVLDlsi2EZjavledHcWgB5EWczhIol3pms/yu/sBdo8HGsiIvVKYQxBauLJhsam53I+6kW4kLokkdyCqAIaB+LxaGTPIGMN6/eXr4qx5vKwBpwMfAiYAvwMeJRxUsmy5AmAIxPgCURRh7zSuKp8JtMr97KnZxq+AYSYrvlo697NttYNQ1ZZKssq+e1Dt3LcrMVccfolwz6m0xadSMbKMJzYRk3TiK27n/eddTmLZizM0dItntq0loBv6OPZ2Ly453FmTToWBmnzY6MxIdDNxGC72+nsRjS9yMdy4FXQWRDvetnng7FilWv1cKxa8rMMeQGVhTDy2AD8Rd53O4BNCKvuxng0svUw5OFNWv1wIDX9ZcDfpfAHOBuYxTipK+IfQ+fyAJC386bS38Ocqh3s6p45IAFAg5f3Pkmyf+iKpgYEA2VcH4ty/NxjOTpHIB8J86bOzas9b0+qj5/deyM//si332gdyKR59KWnCfiHN+1tPfvZ3b6RWZOOHdAKYNk60yv2UuZzbYXvMEKx1/L4Xi/e+pwLRQCsYQpOq4hJwwEPx6orEUtGxkEaFI4Ah9B+DfgcIjAvOZCwzxXybv30Uvj7gE86hH+WbI4bjBkCYIRij5qJcC95tpj06/3MqNiN4e+mzyp7gy9b13wc6NzO3o7NWFjDKpPr03Xaujv40k3f4LZrfkX5sOr857evW7bFYxuf465nV3PpKStff//+Fx6muauNiuDwLNypdA9bWl5gZu3igXc928fMyl2U+1xlzlmIQhvDLi4Vj0Z6GxqbTI+WUjneVLDLBylEy+NfcGQTZIZD9Q9mAz8E5hfRLbnfw7GmFfA8vgf8aQgky1mN0kJUiTtKifnDQ2ry/Yg0vCMRBa+hA+/Nee9xxlEjqzFBABwC41/AO/MZw7Y1plXsZVrFPraa89A02yFQM+xoe4mO3pYhBcTlIuDz8+q+bXzyV1/mxk//dMjf292ye1ipgIcsDxpmXzc/ved3zJ8yh9C8JVhWhutjUcoC5XlcG2gxd9Fi7qTemI1lH7KQ2GhUB7qYUr4Pn5YecirlIALtITfLwKPlVIbog1EIjcjmUCvQ4WyiuxElSYuJAOwpdQIghdNuYHce3908jgiA7mLNF+SApf+/YYB7PTaeCMCY8Dc6tMWb8x1D+LE7mGtsI6D3v0GYHuzZz862l/NrkOMY59nNCRp/fR396aFlrN365F34fflxNF3T2dm2l0/8+lqa7vgJ4es/THNn24Dlf49sidDo6DnA3o4tgxCnfUwubx52NcUcpCWBG5b270jjafdoOb1uAcgjt7hQ0Ci+vvG7PRxrRoEImZuvj6VKkrbDwpFGBOY9DtwAvAtRF79k4Ijyv3yAj+9TFoDSxZ0If3BeUdxp28fR1Rt56eASWpJ18i622N+5jRZzNwFfmcu7yOb+DY/z6d9+la+98/PMmTx70P99eedGfv/IbQR8+U+Rrum0d3fx24f+gl/3oeu6q2Pfe/A15tefQFVwIra8R/x6mrlV26kOdNJvuVpO24xQbHse5v/sy1ZEMGC5yzVUhvQ5j6d84BHANg/HmqUu56gixaEOrb2IKPx1wBpgTTwa2TKItaQkTm75qthgBOAFoNXxuSIApQIpOGwzEb4R+FReQs7WmVTWwoIJW2hNTsJGI9nfw6b9z+HTvblUPt3HvYnH2N6ym09e+CFOO/okptVOff3zTCbNui0JvnJLE/2W5Tr8WNM0gn73HQs1TWdf51Y6epqpCk583WpSV9bGouqNpG3XxqSbXX6/RW5W5R5M0yQlA1xrz60eWlAWqCs6IsjeK/skgW5B5Ni/jGgI92I8GhkwK2c0e9aPkBVgOpAb1PQg3rkSFQEYTTi0xl/kSwAAUpkgJ9Y9x8bOYziYmkCLuZMDXTvw+7wrDlcRLOe1/Tv48p++w+lHn8iJ849nWs0UMlaGV/du4d4XHqbV7Dhsud7RhoZGKt3H7vaNTKmei08P4NMyHFO9kdpgKynLdfXcm3LmcbhoRmQCeBHFW9fQ2FQej0bGaz8AV3AIhx3AHA+GXFRqWmYJwAa+LwXeVkSa5bZ4NNIzFgX+ABgoVuyJ5ativeNpEYwpF4CZCAO8iosOUzYaVX6Tc6Y+xKodl/PK3qfQNO+zegI+PzbwxKtreXzjc1QFK8jYFn39Kfw+X1EJ/9cXix5g84HnWTrrHHx6gEnBgyybtIZ+y7WF4WkjFNvpsrX0XrxLBZwC1EjNSGH42n/25SaPCMDEhsamysGEk0Jec2QjMhyOKOzHiMDPav5ZF8AVOR/tQBYhc/zPmIc+Bs8pCfzEzQBpy88C4zXOqLuLzS2bPTP/v1mrFkQg6A+SsjJYtk3QHxhWD4BRtQJoOt2pDvYc3ETG9nHetAcJ6Cm37X8B/ncQMvem10cgAF41oJmOcgN4gY0ejnWcupzeY7zVwpfR/1XACTkfPY8MXC208L9u5bUDvlYE4AgwQjGMUMwG1uKuwQQ2cPLkV1la7yM9CvG8pVJqzKf7eXnfOk6sizO7aisZ27V1pAO4Ozt/WYFvhGKYibAv9/3ByEA8GunwkADMBOoH2yCLEMUatbzBw7GWK3E9IpaA8XjaF/DmQPH48lWxA4U8qKywv371DVy38tqp16281pCvFQEYJhHYBdzoXtjBV86oQtdsbFttFiCK/kwr28n50+7zQviDqL3d6xTsUvhPAHaYiXCfmQj/0UyEV0qtvMJMhN/Al77642uyL7fiTfpVHTL3vEQ2SJ3h1IsePTLhJQE4o4QImUJx4yLe2PK7HUjAG1IECyX4z75u5bVbEe7HfdetvPYd16++QRGAocIhSO5F9Jp2hdBUPx9fViFKro1zEmDZUKbDNadWuin440QSuMUIxdID+P4nyxuzDPgg8E+Ej+4XwBVmIrzUTIQnAXz38z/KfuclvGsLvKShsSlYIlPjZQtjL/1dXroAzhrHGquCtziVN5Zr3o6sZVBI8/91K689DVgFzJNvVQEjykhKjgAMZAJ2moazgsQIxbYBtyByw/NG0AcfOL6cixYEPVePSgm2LdwU/95QTmiqZzLiZgZo8iJdAFuAf0e0/LwLkVdeg+jadTtwD/BzMxH+vJkIn2vb8yu2t1c8m8roScubSVpOnmWlCzE9Hi5NT/ogSE09iajz7gVmNzQ2TS+lW0bJ2eLDmsvDJ0vlwokty1fFhtyF9LqV1w5ols/HVH/dymuzmn8d8C3e3PdiRNdRyRCAXAFvJsJnmYnw18xEeHmu9uggCb8DnnOr9daWa3zh1ApOm+kfl1aAbBmwS48O8v6l5fi9WTUdwN+MUMwcbL6NUOxJIxT7phT67wE+Kud0B8JPfyXwI+Dmvg1L71rz2SeuumBBe6AyAF1JH6mM5ma+ziohAlB0kJp6P6KAjFd4u4NcFDuCahUUleDPvjwNGd8j0Qc8k/M/R4QU2vXXrbz2E9etvDZy3cpr5+bjr5ff0eQed37Oxylg17glAANp9mYi/B4zEX5AaoHfloLhDelj0oeMEYr1AP8pNZG8kbZgqqHzjRVVHDNJxxpnJMC24ezZAT5/SiVGUPOKBN0HPDKQVUcGczrnvtMIxZ41QrHfAZ9HtH2+COEO2AfMzNjaW6ZUpa6+7rzXKn9zRYLrL3qN8xd0ENChO+UjbQ07zLIaaGhobFKtYPNHyi0Bd8AHXFZC514xXiZ5iFk6BT0Oh2n/pJy56czuQ0Mx/zs09ulAFPgxwkp5z3Urrw3m6a9fDHxhAHnchchOGDH4i3lROYR+FfAJ4DOIFK2sv7MNePRw48gugb8GPu3meDIWTKnSia6cwIfu6mKXaaGPE9GwuM7Hd86pZGK55pNQZN4AACAASURBVBX5aQH+IgnaoLn/zvez68EIxbqALjMR3oao9/Bl4HjgKuDK2orUxEmVKebU9HD+wgN0p/ys3VXDPa9O4cntE7AAXbOHOnfvBu5uaGzqH0e+Z8/obTwaSTc0NnlFADREXMaMeDSypwSuY0nHVw1Wk8NMhGs41EBnvxGKJXOzd0ZT2OcqfgAdz15anmz31VlpUcDFX24n6y+4Y/8Dc8PT7H4WaX6caVfty1fFnslDY79E7g9ZHGOjLUZ2NB0KpLWgHPgiA5e77gKeHfMEYAAtXwP8ZiI8VV6cj3PIN9mPiPb+AfBrIxRLDbTwchbGZ2QUuavuXJYNtRU6t19RzYf/3sWrbZkxTwLmVOv8cuUEaso1Mt5ZPp42QrE7hvOFAebXNhPhlBGKpYAn5ePfb7z5U9dUBjI/PGt+G5XBfiaWp3jL0fu56Jh9dKeCPPhaHf/aNJnEvirSFmQs7XAS7z1AYzwaOTiOtPZOj8drBnYiWha7Rb3ceH9ZAtex5NZMzj7qMxPhIBBCFM25AFiWS2zMRPggoonXz6QyZo8UGehcdynVJ97lJFg+MxE+GngbwmV3AjYL0Gwqp6ffdG52Bjt5QLNaHgTzVRtsrKwCOcziP/XA+52MuUrr54MVL+373jCEvyQTpwAfG+Tftl+/+ob9IznnRcFSHSZfzUyEpyB8ITfLjePzQCWiv/g9wLuNUGyBEYr9bDDhP4hZ6DJcBgSCMIeXBzRuvGQC584OoGuMOZeALcnOMZN83HhJNbUVngr/VuA7RzLZDZcUZMe66qen/PK/71+Qvuh3y/nyP5bwj1emsaXV4IBZRsCX5vKle/jFO1/gzg+t40srtnP6nE5mVCcxghlsWxOEwH7D/XE1jKv0M68p7QFEExlPtgrgHVm3jEoJ9F4BMxPhgJkILwQ+C7yIqKdyLXDiIPKiRmrCDwN/MhPh2pE4vo5nhPDf/9g7ta4XwvMRFuEEGhs0nR9oOpfbGRZkkpA2Id0F/R3iOW1CpgfsDFr5dNtXs1xD87++1d0NQ4/+l1r7AuCc1xVDNI72H7Q6reBHhyn8K4HfDvJvaYSbdETrAPgLuehyNP5lwNnAe4HTX7+2IojoUeDPRii2Jvf7R2KZjniADWYi/FngV16QgMqAxv+7wOBX63q57ZUknUnbq+C4giJLZs6dE+AbKyqF5u9tIaQ/GaHY015rCEYoJkuYfqSn4qSmNTac9sSOCTy0dSK15RkaZnRy0qwOjq7vZnZNHzOre7ly2U6uXLaLHa0Ga3dXs25PNdvaK9hxsJzWngAaNj7dvvak/2iKrv1ZpK/Ia9EXJQ2NRyMt0g1wuUdDngy8Jbs5Kni6Dy9GBNZ+CuFqHS7eB/SYiXCjtMy51xYeuRwjtEqYVJ68dLa/Kn0ZGp9FY6HmE4I92Qz9bRrJA5Bqg4wJVj9YKdADoPlALwf/BPBVQvdmG1sYCHqHs46kINYR7kaHELXtuXqHnkGvGebp/Rew8DD38z9AuB3GBAFwMk0pmMsQkb0rEYFd2YvRLU/+HuARmRI2LME/iKZ4I7BUslvXgrLCD43LK1hc5+MX63rZ1J4h6Ctdn0DGEmmPVx5bxidOrKC6zFPNH0Re+IhJUIdwvkmD04I+m6AvQzIDT2yfyMNba5lcmWZhfTeL6rtZMqWb46d1saDeZE6dyWVL9rK9vYpNLZVsbDF4cZ/B+n0Tpm3rKPsy8M1pVZminTpcBrqOBByE6QVETQcvtMOpwJUNjU2Px6ORHtUgyL3wlwrYB4FrpCI2EJKIPittUm7MYWC3Thj4M/CAW5Lf8dylTDx5Vfb15XrA/iwa52o+sNPQ8aKG+RL07oTUAVGsTdMdNizNQYtFoqxt26K1i2zv8szyVbG+oZr/pdZejYjYlzeexjxflxXUMj5bVKAdqvZ/srzeg6H5+tU3vDDSa2BUCYCDadYhUrrCiAjIeoe58JeIoi+vGKFYuxvBP8Dv95uJ8LcQhRZch65mbFEt8G1HBTmm3s+NL/Tyl5eTBH0apcYD0hZMLINrTqnk4oVByv3aSLg2rjZCse5RCBa6DeGTBEDXQPfZBHwZzJTG2t0TWLd7AhPLM9RXpZhT08fpsw9y+tyDzKs3mVdnctb8Ng50B2npDvJy88TPfKn/x7euvuHzG4tU4NiSBBQrEogiTWd6NN4H5Bzfp4S/u/3YTIQrgSZEnFXlIKT914gYmxaEG1VDBK+9DxGE6/zeZKlkPZDvPX5IUbyLPXdeETTm9H9D89kf1zTqNB+kWjWa7xOafKYHNA20AGhija0DtshjPSiPrQ4RJ7AiZ1u+PY/Du8x5vpats8DXriFi0/4+FBIh8SPeXLzLdtCXe52EYaQw4kbrnKYuc81E+OeISMlvyg2hHtE17KPAEuBbRij2VFb4DxTt6dL60CKZlyfpFbYtrAFzJ+p8+YxKfv/2Ccys0ulNl05gQL8FR0/y8dt3VHPZ4jLKfCMi/L+GrMw4ksJf+oVbsjdQLjQN/LqNT7fpSulsbSvn0a01/PjJOVx1+1KuvrWBv8Rn09UXYGZ1H0undbJy8b76E2aa/5u1MhSh77ko6WZWOMejkV14Ww+gDPhBQ2NTDQp574VS+P8R+NwAwr8HYSk9A/iJ3JM3GaHYTiMU22GEYq8aodg3GLja43QzEc67NLXDQlxXvaD/Ft3PdZpGneaHzhc1tv0Kul60sXpB09mCxrWIAO+zgE8iYoyiiDiyXwExRPpfLv4GQ0//k/jC68IfjTq915qoJXXgr8tXxfqGMsZ1K6+9Jud4OhGB7c77+I4cwlD8BCA3qMthYjrDTITvRlRz+ySHUvkeBN5ihGJHG6HY74xQ7AA5pVy9EhY5qSpbgA/jYZEFy4Yyn8YpMwP89V3VNDZUUObTir4cmA18/IRybn9nNYtqfa+/5zEeAqKyUdOoCB2GECmuSUIAkEzrHOzz8/zeKq5/ZC4X37icT9xxArEXZ/C39TPY1ek/r6Gx6ds5v6EwNEIGwp3X7OHQS4H/a2hs0nN+R+EIe7Sj0dZfeHNbXCRZW2yEYj9FmPzTh9nnWwbYMrR8jy37bCbCcxDZBVcAGjrsX62x62abTLcNsAuNDwDHLF8V+/7yVbEtiJS+biC1fFUss3xVzEb0D/kXb65wuWH5qljLUIr/OMz2RyNSjl8nAAv8HVpQswC+OxTt/7qV1x4FfERaULLb7XZkK2IHVo/GetC9WlQ5wSQBmcL3fjMRfgF4AuHrTyHatv4BONYIxS4wQrH7Byr4M8JmL4xQLIFI8/I8zaLCr/HF0yu49fIJXDg3QE25hkZxZAtka8YGfRoNU/3cdlk1Xzit0tNasjnYA1xjhGIHR6tgiBTQjyLbew5bndbAsm2e22XwP/cv4CdPziboswLANQ2NTR9TUeh5WQH+KbUcL/FvwHcaGpv8WcuMmpMjC3+JXyBSKt+gxwA3G6HYSUYotmsgt2vOPl8DLBpA4O8zQrF+F8e2RMqMBjTI9GrsvEmj9REbzUe3lB9Llq+K/Wn5qlh/VojnavJrLg9XI4r8THOj/TvwyTdq/31M101Nx75/+arY+sHIRE4U/yecJAIRG3MzhwLfAf51/eobrKFG/+f+32CligeC34sF5VgQExF9uy9E5DbOcZzkSwiz7G+NUGzPSGj5wyQBmhGKPWkmwlfICZjnpZDtS8PcGh//t3ICa/b089eXU6w/kGZnZ4bufhu/ro1qDYGMDZYlyhofW+/jkkVlvGNREF2DVGbEmEk3cJ0Rir1QgP2uC+G7/Hq+A/ikq8CBSkTlL39DY9Pv4tFISgWhDc0KIK/RnxHpZF52LvwKUNHQ2PTNeDTSlvN7YwFeNbdy7tOfl/uz0+dsAT8yQrEvDSCQBxqjGuHHzi1gsxPZin2osT45pOJ8hG9+Ehok92nsjUHvDpv/3955x8lR1o//vVdzZC+VkNAJhBLKQuBEqkQFZQQOgmIQpPhTkW8URf2Oo4nYkMAwwFdUIirNQhPxZFEGQZp0CASPXkInvecu5dr+/ng+wz333Ozu7N7utczn9ZrXttmZZ57P83x6SVSyDLiwoSn9a2HwNDSlezDx4Lv5MxpHy3X2yXLbO6POm2aG/7DwTyUZdq1cQ32ijQxcmCuQMPi/Y9lHoOJXdLhbrCi6hSKS+d/oIjhGcLHS9b3FUQWAoiwAIYV7dmhpbjwL+A0qr/JnwvzfQeU5ngtYyVT6wmQqvShXX/d+FAIysvAeFUvAgpIz3S5obcuQ2qaKiz8xkss+OZLzP1LHZ3arYdLICjZ1ZNjUkaGjqzzP2JmBzZ0ZNnZk2Km+gpP3qmHO4VtxxdFJPrtXLZkMZbu3EBQPuLEQYlBCzbNNNvmyEl96JCrA8NJps+buEjCaWPPMbwUQgWxJCS8dMK9vAX+YNmvuR4z7DQcoiZqgmdcP1oRi/drzRJgyGXLYtY4G/oRyo1YZAtkfKCDWx7jXmcL8xpGA1tcTfHAbbHwnQ6KCRcCXTeZvaPwB8x8pgvoxWW67BHg1qvlfXo9BgtUzwKTKVnauXAtkbgaezmdJkJx/M71yDaqM8JeM0/8VZVyu7wXMf5rQpCbgV45l7xg1dqAoC4CGsH1RKRFHAfvTHdX4rGjVDwHPB+agUkTyl8ESQDKVfqqlufEM4EpUxauSQnsXtHdlmDy2kinjqjh+YxfvrOtk4eouFixp57mlHby6spPODFRVQGWFyiIoxEKQyYiWn1ECRQbYPlnBR7ar5pDtqtlzfCW7jq2kvqaC9q5MfwQp/g64WBO0BkLjfBEVLf6NEt+iQpjO4dNmzb0W+O2CebMzuiDQz0xo0EecCk42TJs19wrRHEsNxwP7TZs19wbg8gXzZq8fYGvAoMKJ0LoK4EfA6BCG87NkKt1h7lVdyUNF0h+PaqcbVifgesDV6GohzP/bIpiMSlTCuhcSLLkzQ8c6SFSyDvhqQ1P6rhzm/oD5V6NSjc/IcWsfI64hApyKNHiqS3SyX9UKRtC5opPEbxqa0hvzCRAijJxm/PwTVHrlEdp3LwGLc0X/6785lm2JohVYOnZA1TaIVO+m2GCNKlSE6AWojmlBb+V/AL8WAWBVMpXuZAhBS3PjdsCvgJPLLdIHFQRb2zO0tmVYuznDSys6eGF5J6+t6mDhqk6Wb8yoFBc0YUA+B9kHGXk/uibBDqMqmDK2kqlbV5LapprtR1UwsjpBska5GzKZfqNKtwJnSNrlgAp802bNPUq0lR3LdIsNwMvANcB1YnmgnMzHvO60WXPHi6DziT5eugM4YMG82S+Wa8wStPd2GfERdFD7E3DVgnmzl+eauzKuu3+XSJn43IJ5s2/vq/YvTPkkWad6y9l3gJnJVHc9fKnPsisq0LJBGNSOqKp/2Tpk/hC4PJlKbypkz0sw4g+El9QkqmD9Kwk+uCVD12bJ61ea828lqC8rzJ/RmEC1EJ9Hbuv251FR+5HIoWPZI1FpkKkMCQ6v+YAdK9bRRcIFZjc0pbtyuQAcy64FXqGnm/kVYBqqv41eQfgK4Puu77VHGNfpqJL4Ew2rwlmu70VCQLECwAhUasUxIsH8EbgE1aa1M4j27q/AvlKYxzRJtBL4pSy6fk2vymSU3TwoRbuxI8OidZ0s25Bh1cYuWtqUHz8D1NfC+LoKJo5MsG19JXXVKtAwiGwfwB4FtwKnRtUC+sEKwLRZc69GBd+UE7pQcQf/RPke710wb3ZrAUy8IMY0bdbcrYRI7yIWuG/Rd9962QQAQwg4lvJHOXehcrMfFWvkXQvmzV5awJopFB9VwM4o1+dBommXop10nwUAjc5dg0q31mEV8ALKtTUa2AYVoR6Qk4o8zPQllBX42cDaF4Xmy3mVwIUiAJCohA3vJXj3mgyZbtXxL8A5DU3ptTkYf/D20yifei5YDxzW0JR+Ieq8OZZ9LHBDJ4mJ06qXM7VyBV0kngE+29CUfieX9i8m+osR94oGlut7dzuW/QI94xQ+DdyTS/sXfJwryqqJm7+5vvfZQpTRYhZSAhW1+DHgd8lUepXJSIciGILAOcBlDHBP+MACELzq9sWMZgEYJHALKjK7azAIfhrDmYxKBdqtn4fwtljDXkKl+byHyjpZLZYDcy9WipY1Wl7HowqrTJIjqL62K+XpN19WASDAiRCt2+n/9r5BX4KXUPnr7wo+Vgo+zIiYKlTg5xhUFcMAHxNQJvAdRACbjOpTUA44bcG82TeXQPvfGxWPc0Bf9RSxsrwFeNKiO7LJP6ALYmW4MhDMExWwaVmCd6+Bzo0fErTFwFkNTel7c/n85f3+qEp8ldopjwmudte++zdwRkNTOnIsimPZP8+Q+MEeVWsqDqpaTAeJtRk49+Cm9C3ZNH8jddCslXAXKu5sG3qm/60ADnN97/UQph9cb4RYW+aECLx3u753nH7/fFBsDECG7g5sQ0bTj/BcwbMkkqn071qaGx8Xk9lBxsLqV6tAJth2gxc6UGlF302m0l0a0RnQdRGkhi2YN/utabPmXiaWnep+HMIucpw8hLZBotw4AbqmzZprowqBTejHZ5uAKjtuDSF8lGrn7wbsVeR/O8VSsBxVxO3PyVT6H2GKU0TlaiIqYHwGkKGCxOaVCT64BTo3ZPQV+J+GpvS9EO7z197vjWpGVGlYJs6ld3D348JoI8GXP31BHWzaf9fKtRUHVC2hkwRLupKvn3DHTbeEjSuE+V5v/NwG/J/rey2OZX/P+O0xUQ5Cr+VY9iQRmj5v/G8T8HvX975ZCPOHEtUBGEyBfSV6liBw7XmUX9Wj9NHkwwXWiUT63bDujEFFr5bmxnpdWOxPIUBerwZuitGVF+r66T4LAQdlpo+hjPRMYCe6i8/kgjahdS8JU70RZab/MnBUMpU+NWD+hZj7Neb/EeAOguZQCRKdrbQsuTOzdvPSHsx/bbBfs0XrS9Df3sDfxUqjW95OQ+XbJwxF5b8NTemOfBkAwe+TKjdO2btq1W4HVC+lii6WdCU7H2vb/h8Af2j8Sj7LwXn0zO9H5vNxeW+mBD6pCwAG8z8QFeVvMv9lgF0M8y/aArAlbZxkKt0K/KClufFeVBnI4+LZ+RDekjn5e5jPX76bikp1qW9pbvxxMpWeLy6kTH8JjZo/95so8/mRMeqyQnU/4SMzbdbcW1Dd/f4nnvbygZjbdwyxLNyICkbrRNXsaEH5yFej3CJLk6n0YpOZF6r0acz/dBEmJn9obqpg2ZpnuLH1dU5NVPTITliMijPLFfG/OyrNXDfxrwS+3dCU/u/8GY2zDQHgVVTZ+bzMv6EpTeYseGTtW6fWJDr3GEEHi7qSPN0+iQ2ZqlsBzkpfk1X7dyx7L1Rcjn7/D4DrXd9rldS9yYbg9YLre50hzP94VHDg7sbt3gC+7freP4ph/luUAJDNTJXPfKWlLt7f0ty4AJUGcynh1aW2JLgdVeRnYZ55nAh8CuVD3beluXF2MpXu19oAmitg3bRZc89GpcnsRgxhkOlHfGycNmvuD1ENwT4RT31ZhbrRIXi+KplKP1EI/Sxkv2quwCqUFfWr9Cx4s7miljOX+UxMVPXojJcBHhImn63Qz7Yoc/ghBhO9oKEp/ff5MxorUK5bnQG/JkfWiH39fs+sa7yyLtH+tWq6KpdkRvJ0+yQ2Zirfv/Lui1/JovEHzL8SPuxPEEAX0OT63sPy+Qx6FmJ6FdXEyGT+X5K5G2/crhkV7f9cscwf+qEZ0GDT6FuaGz/a0tz4Wktz45UtzY3JfAva8GWvFql5KqqD1pYI7WIOPCMC8yeZSj8o57eizJC/aWlunGPObT8ynTdRAThLiGHAQar3fQFlco6hfJCIQv+z7cdCGb/G/HcBnhZtWGf+G0gwPT2FexNVH7aB15nlfWHXFuY/SpiiGcvxi4am9G/k/eH0dAt0Ac/natijCRdj5s9o/A/wzUq6ahd1JXmybVtaM9UkJMsgrNKexoBPorep/j2xfgRwQoh14i1DoJgDXBfC/B8DPt5X5r9FCABaBayKlubGY1F+rd1l8YwuRHgIFlIylV6TTKXnoNJ+mii8qMRQhftRPRyuS6a6i19EsKA0oYqHLEZlVfyspbnx/1qaG+v6uTpg8PoMyv+2OOYLA8b4gQ/dActQOfML45kpm1UnjEaN6gujzyU4CPM/T7TaA0KY3V7J/dJP7HYySXqXE84g3SO1KP/gtQ5VQOd04z+3NTSlHc23/zF6ZmasAJ4yGb4eCyDM/zhUbYQjE8DirnqebN+WlsyHnrF7DGaPLhA4lr0L8L/GvdsB1/W9ZXLOUSFM/SXX94K04QrHsq8Dfh4y1X9zfe9w1/dW6dkBxcKwFQAMKXQEKiLURwXCvI2qDldwsxjDx/1uMpU+GZWHfQ/K/zTcoA1V6OYr0rxpYSGlnLXYgBdRhUUek3V3PjBPii/RX+Whg4YxC+bNvg+Vv/xqzB8GVhAQfCxBFZ15Mp6Vsuzh1SEWgam5tP5CaK1mYa0BDm5pbnwIlXVjBuHdBnwkmUq/J6y+NoQZZhqa0gtDNPNaVODot+npqvp3Q1P687rAIApHrXbOMlMACPoIzJ/RmJg/o3Hn+TMaf48qZjcK2LQyU/f8o+3bbxbNP4BHzefXTP+1qHiWQ4xTHnF97zfa5+PpmV6+HMlWcCx7G1Sq4pdCcDg3yPHvi9Y/7AUAI990gpiLrgqQAZySTKXv6asJWrvPY8lU+tMild4sUu9Qhw5UzvTPgcOSqfS1+mYvRFPQhIBVouldJ1Lx2cC1Lc2N++aqPV5GzfPfKPPzv2MeMSiEgDWoWIDrUalNMZQApBT7+yECwKdKofUHzdWkz8AlqE6cHzNOX4LKFjotmUqv174Pai2YzO5DDV2Y9AjR/H+sjR/gYeBzhpVgN7ob0QUWhYUNTenl+nnyfhKqMdJ98grKMnjR3Zt2+U57pqK2olvWaHV9b2mORjufA8zUvhWongmBsDACOIye8XfLgKckMPBe4OPGNZYDX3d9b04pmf+wFAB0BtXS3LiXMJugFvyNKN/1fLOFcZEby9wI/0qm0qeJIPBD4AFUhO1QY/z/Eg19hjRwWtPXnH5NCNgkUvJPUXEBxwI3tTQ3flrDW38KAQtQKUNzZTwxDKwQsAFVHOY7KFNsDCVQhkQAMGvWH9zS3NhQiAVOP0ejf0eLtn+baOe1xt/uBk5PptJu0G8gRBjRoV3X0oX5X0rvanpPAf+voSm91kjrS9HTrdCGcl/q1xw5f0bjl1Alo39Hd8De/cDZDU3pn1cluqYkesbEbjQ1f8eyE6L9fwpVgtiEWa7vvacJDYcawgmoglT7oQKrU8ZvbwBnu753TamZf9jED6eF3wD8FtV+FJEcr0ym0mvLVaDGFChamhu3B6agXATHAQcnKjLU1naR6KeZ7+pM0NZWEaVa4CpUydQ7geZkKr0s7JlKNUdScvlU2TSjgEXABYVUFisFGDXqpwuROWYL5RcdwMcWzJv9+EANwCjHux/Kn3rmFszDv7Bg3uxbSrDv9kDl1R9kaMaPAmcmU+m3otI1+W5rVKDbCcK8tg/7K6pi3V+SKVV5z7zW/BmNE1CpfCcYAsAODU3pZfNnNI5FNY860+BXLwNfbGhKP2vedP6Mxh/Tsw34OmBaQ1P6zfkzGmtQlscvCm8IAhM3ARcD1zY0pT/4nmVXJxQt+Jlx+ZNd32syBIHPiBBhzsFcVEnoTi2qfw49fftdqNiXEfRO1XwOFenfXA7mP+wEAI25HI6qIb2d/HQ68NewQjXlkrgNQaAGqK+szIxd+Fry9EcfHP+TFcuqSSTKm3GVySSYstdGPnnsUkbUhRoiMsLw/yimtPV6cF+55srwGR6Oqp8/GtXI4qJkKn3ZQAgB8n4Mykd3SRaiNtwFgOkL5s1+dCAHYeCjDlUrwAMO3gIFgD6VAjb23S9Q0fgmDVgoytINyVR6RY7/7ydC8vGo7q+jyF446lZUg5+3dK3f3M/zZzQmUQ1tzjHG9F9UHYCjRWvWedX7wMkNTemn5Rp6sOBYVNtpvR7+6yiXxDdQZvqdjHE/i2rK82RDUzrIw69EWSp/ZTzXarFGXCVj+i4wC2kVrMHfgK+5vrdCixEYL8LOiaaeRm9r/L3ATNf3VpeL+Q8rAUBj/p9GBXJUoapJzQTuGYi2tOb9vv8ZuyaTYY5Ihf0CmzdVcNasD9h9z3X6AvaBv8oiawkRWMo+T4YQsDMqOHA7YUI/Bi4Na0/aj0yoBmWK/iGqZveWAG3A5AXzZi8a6IHobZXlfaVYZi6m7/XshwqsA2YsmDf7/hLtuX1RWUtTsigDCZQ/+g2hEx0imO+IKmldEYFvvIgquPVgMpXuikJP5s9o/Dqqi2y2MemwCji6oSm9wGT+8nlfET72DrmWOfY2ocVeQ1O6S7sGDU1pHMs+jJCgP3rXyjDH+BTwxZCa/gfK2KbkQdVvXd87V/tfWZj/sBAANMafENPOdfLTm6jI9QcGy1gdy64TCfAL/XXPzo4Ehxy5+r6jj1t2U1V15j/JVPqNfOa9AcLfWFQwXuCy+QmqvWhLf48xpFPf50UYOFBMdSOG2DYJ0sDatdd2VCOcl4RovwQ8smDe7OX91TK3D/g5TDSvwzQtdKjRsgAXOl5Wo7JSXpbjPwvmzX6rxPvsbNG4x5VwbbWi/NgXJ1PpP0elLVqA30dRwdOT89xrMWBJlb/QYj7zZzQehio1vHWO8W4UJfG8hibl6gy7nmPZE1A9Tgrp5fEccKbre8+bjNux7E/K2EZmI9fAD13fu6QUKX7DWgAwIlDrUL7CwF8zHzgvmUo/MZgaFQ2EAACwaVPFd698wL1isDD9PELAH1EmRoBfAD9LptKrB2LMIYLATkIMPika0QQhNJWDYBq7UB3t9CMo77oYlfr6lhwLF8yb/XaUZx5kjN/Ex1igEVXTY0+6O/XVDJIhbxQc6PhoRUXEv6PhZKHgpLOcOGlpnjQL7AAAIABJREFUbkyIJfQ0lG9+ah94QAsqjfq/wI3JVDrdF/oyf0bjj0Qbz7aXHgNOb2hKv52N+ct1PipMdmIWAeJR4PKGJlUBMUI3v4NQDeHyWZ3aUJkEs1zfeztMa3cs+xgZW5jbZBHwA9f3/tifC3RICgAG8x8rjD+I9L8bOD+ZSr862LoUigBwHSr4rV9vDVzh+l7HEMDteJS/92xZnzejGg0tHkB3QC8CPG3W3B1REbv7oCKId0C5MMrBhDajarS3yLFOPq9DxU2sRaUbrUClDC2TY+mCebNbcz0XMKi1/ajjFmFgGqrWxBSUn3dbVMnuremdatYX6BA8BALWWsHHWsHJapSpeoXgYblYW5YumDd77QDTzUAI2AUVWHe4CE87kjsrbJ0wqbdR5XQXAA8H1UD7wPgDK0AdKvPji/TsWvgWymx+SRDtn435y/W2QbkTTtG+XoBKTWxqaEo/FJFWf6iBO5adQmVFWfQuAd+BMvn/FbjK9b22bCZ7x7J3Q2WifdQQ3O8CLnN97yFd+IgFgGjM4tcoP39CmOucZCq9ZDC2KJYc0Hn0LvJQbvgKqglF1xAR6upRLoDvyM//Bs4NihANJE6zaWPTZs0NrAFj5RhFdw/5UcKAtpJ1OlJj7B2iPWzWNMRWjdlvkiNUu8/F4IeCVl9GPI0RQWw8qiRsveBjHMqvnUSlq5mCWkbmt0M0+E0GPjYKrjbI+w0m3rJp8oPN4ibvt0GZ3reR+RlDd456Rp55jRzLgUVBhlCpLIpGEN8BIlCPl7l/EXgmF9MPud6uqFz6USKwvNzQlH4l7H4FCAFboVyAU2W+KlGF314F/uv63tv5mLdj2QkRUoMqtGtR7ZWfdH1vaX8z/yEpAGjm4npU3mSQsjUXFTi2drCZuLUFUCHCSn+2pe0CDnZ975khZtmpRjXUuEh+fgL4UjKVfmWwCHdRGOu0WXOrhKBW0m3erNRwk9FeO1B+wM4F82ZnSj2WLYTx55yHabPmVhr4CKOBnYKPTsFN54J5s7uGEz5yNUfT5ySZSmcK+X8phIBsvwMFnTN/RmOioal7/FEZf5ggYPjya2WO2nSlKg/z1xv8BDShI7DKlsLnf9GtDwIwZ+b0vOcF5wwpAUBj/luhKvpNk58uQAWMbRyszF9bCNViptyD8nde60T56Ba6vpcZaniW9+egUpQAnkEVFHl1sOO5P4WMGPLPIZTG1TFc8RG1K2p/QzFMu1hGX6ggEIXxl/L/OuOOIAhsj3JLbo9y72yPCjadN2fm9A+GlAXAYAhbiza4G8p0ehEqUIyhwBQcy65HdYvaTwSAcjHmhGgvDwH3u77XNoStPV9EVexCBJrPBu6AwWAJiCGGGGLoC9OWcxNCtyuyvCZQbprtsxwBw89Wm2ETcN6cmdOvGTICgMH890AFTOyG8k1dkkylLxpICbVA5l+JSie7qp9vfaTre48MZc2kpbnxDFT8RBLlN5uZTKVfjoWAGGKIYTAzdjm/EhVvUgNUa++Dz3WomJWJqEDDSagg1m3kdVtCujeGwGY52kKOVwFnzszpC4eEAGAw/4OBG1BBGOtRZWOvHCrMXwSAOpQ5+4x+vvUs4Heu73UO5U3X0tx4JnAFKkDodeALyVT6mVgIiCGGGAZIawcV1JtEBfkmtc8j5ahHBQiPE9qlHxNEq88HHcL31qMCT9fTMzh1PSrrJMg+WSnHcmDVnJnTV4Y9X9VgRoTG/A9D5WJORUVOzkqm0jcNJeavQfUA3HMUQ7zmgzQK+mNLc+MmEaJ2B25uaW78cjKVfrgUzZ1iiCGGmKkH511064NB5khwjBEmPkY+jxbamtRe6+UYJb9Habi3Rpj1Knm/SvtuNSoFcx3dKad6KvD6OTOnbyx2HgYtU9BMvwej6vrvLA99SjKVvmcoMv+BKgSEampx+VCoA5BvPcj741G1tqtROcnnJFPph2JLQAwxxMw9wvmB5r0N3XU7JoqWvg3ddSNqtWMEylxfJ++jKM9rUfUflmmvwfugXsdmlG8+9HXOzOllTd0elBYAjfnvi6pbP04m6zSkreMQ1vYGIhe/hfJnHJQV9CDPZCr9D+n5cD8qm+L3Lc2NZyVT6cfL3Uo4hhhiGHwa+0W3PlgrTDzwlwc+9ImowLiJcmwlWnkFKg1Uf62KqLGvRFUVXCTHEnldjKqOuES08x6ppMb7rjkzp2dKPReFwqCyABha3lRUSd+tRGL6UjKV9ofyopb8z28Bl/UTQw7wOz2oMjUcQBMQjxIhoAIVGPj5oE5AbAWIIYbBy9xD+E/Y+0rRxrcT5r2Dxti3k/fBa5Ry3BnjVX/fJkpmwMQXGe8XAR/MmTl9bdRnLBfTHpYCgMH8DwQeRPlSFqPq+t8+HEy8jmWPQ5nkp4o02FUGYSCQcBNiQbne9b2Nw4mIaELAscBtKP9bM3BiMpV+OyazMcTQf1q49p+gwJJeAMt8DSLegyj3SfI6UXu/Lcr0ng+6hHkHRbQ6tKMTZUoPSjEvltcl8rpImP6iOTOntw8nxj6kBACD+R8hBH2SIOqbyVT6tmHC/MMaRGyLCmgrFXQB813f25Tv3sNIGPgCKrVyLHBTMpU+PSbfMcRQMqaeoLsLZp0w5jrjuxGowDc92j14P47ussz1EW+7ie5Sy8H7Tcb3QaS7GfX+4XdRzezDkbkPKQuAxvxvQOX5LwW+mkyl7xzOwV2OZT9Bz+YQpYBfuL737eG+eA3B8WxUw46bkqn0pTGpjyGG3oxcZ4gX3frgSGHa9SgLWtAjoV77Xn8/KuR9EAkfBTpR0e3rtdcgwt18HzRWMhstrZszc3pLuYWeWADof2K+D6qj0l5ilpmZTKUfHObMv1qk1foSX/pp1/cO3hIWsOYKqEK16H0vmUpvjrd2DDGEMsNPoIqRjTa0d/39VmTvWR8GgfYd5KCvpjsnfZX8FjRS2mho9EFTpU1RzfAxYy8dDKYsgAnC/FsC5h/8MIwDuqpFKi41VGwpC1jLDugA3jAtAzHEEEMP+AYwI8J5esraMmHoi1Em9uXyfg3Z/e8f+uHLlcoWM//hJQA8DhwtC+7FLYGQu763IegCVWLYsCUtYnONxMw/hhiywu+Eaa+hOyguCIwLguP0viGZbK8F5t7HDDsWAMJBGP1m4D7ju2E78VpQ3u3AyYSnpkQFM4XmhnhpxxBDDCFM+G7g7v5m1jHzH5yQiKdgQIWAhOt7GceyR2iMP6hEFdVslhHJfS3K9N/m+l5ncO14lmOIIYYYYhi0FoAtGDIAesqeY9knAA49zXC5BLi1wGzX9/5lWBdi5h9DDDHEEEMsAAxGyJKXvwiVzx41M+B14N0I140hhhhiiCGGD6EinoJBB2lUo5uoFoSrXd97eUuesDIFUhZ174EcS75x6mMbrOMcKuOLIYaBWqNh9433Sx8RNRgmMBiDY9mVjmVf5Vh2JsfR4Vj214baAihknMU+U3/MRS7GP5hwETLOmlKPs5zPK/0zhvz6NgWwGIYuLwnZU9X9tBdKvu8SwxVJgRncseygy1MtkAJ2RdUcqEVVlnof+C/K9N4JdA6kCT1Aqut7OJa9D6p50NGo2thdwEKxEvzC9b0V+vmDmSDK8wS4SAL7AZNRVcRqUdW+WlABjW+h0pHaxcrRJXjJGNceAdwFfFz7eqbre3/pp2f7M6CXHf4pMNf1vbZBhoPTUKWSx8hXy4C9XN9b3dc9JvitFhxOQfW4GC33Wo9KSV2OclO9g6rN3qUf+toVYvptwNVud5vre58fQvQnBfwPypXXBdzn+t61w4CuBiWBs8UXdbi+1xG17LhcL1e9/07X99oH6VychCpcFzQhWgxMdn1vc5nvWyX7qoZuC/7Bru89Xcz1hl0MgMH8xwHHA18Bjszz10Wo9LmbHct+WSLp+52xamPH9b0XgXOiPOsQwMV44FhUIZJDIl7ifeAVVKe/u4F7jN8rgH2N7z4G/KWfHvFA4/PeqCpqbYMMFRM05o8IYAcAD/SR+dcB04BvoorL1ET4+1rgZcHrA6i+H3qjqkoRdnX4+BAjQ/8LnKF9nupY9i2u77UOcfLaADwl61tnzAEz+iXwnQJo0pGopm8ZYw0ggsathoA9mGAsPTsQbiv7f0GZ75uUudFhT8eyn3F9r+CCS8MuBkBjOIcBdwB/iMD8QbWVnA08BHzdsexqTcMZyOfIal4e7Mxfe3+AMOU/F8D8QbX/PFo0wm9l0xKMz+/142N2hDC3zkGIDrPjZKeMtS/MfzxwIfAoMDMi80esA4cAZwPfQ3WA0yETomGuHGJkKGE8Q9swobU7aQx/pHZUCzM8xbHsPQugDefJXFUY1xsp1xs/iOcizAqyZIDuu44iO8oOKwuAtrAOBm5E1YYvFMYBV4pE94PBItAMBaYfNnbHsvcTISzVx8sti3jemgF85NZBKgCEEZH1fdT8LwfO6uM4AhdBlPOGElTR071aQ/S6HoMZ1kUQ7o4AXs1lnZQ1lAA+E0FwHUqwvJ+ES0IUj1gAEKgDrs7C/O9FmR3fQZmwxglj+qxoIhltgr/vWPaLru/9eTi30i2XICabvB6wszD/N1H++zdRzUMCLWAEylw9TnAStBF9d5Bqevk251DSYKIKo18DTstCiO4CmlFNYDYLTmsEp+MFr9ui2n1/EIGpDEW4W+jPSJSV6M5hYP6PwpDrRfm6NgK9PBLlLhtO0B8Cy4ZS7OVhKQAI0zkb5ZfUoV2kzSeEKLVrQWm1wMUoc+bZxv+uEN9dR8zWi4KPinClQyfwM7GyBL7ETmGegTmwAmUC1I/B2OGvZUtDqGPZk4HPo8y+OtyFiu9YquE0I7hMhOC0Ss7ZNAyn6WZUoG4Fmn97mCoSrfTsHLiXY9nbub63KJdyAJwaYl0YQXRX0mCEsisAru+1ldItPRwtAE7Id590fe9hcwNK0MRG4H3Hsr8nGsoJ2v/GowIIrzYC2kI3cpQNXkCELEZ0dJgmRrYsgFzZAcWOoUBBbATKf29K+Ve5vvezkPFlNAGh3y0Whc6hJliW7H755jxX/Ec/MpeDgY8Y3/0XOMn1vfaQcXSWE2/55qqY9V/oXBqZLkgWSFs23EW5X641Wcgz6dkaRazvKHAcKjg3YNx7oDJBFuWxIp1k7KMfAP8POKgv+M9FJ8tJ8/oy1gHcy8MuBmBfYGfj6z8CT+UjFq7vLXcs+1rgGLqjLCtEUr3aEBwCBhf49hLABskcCNLcRmjz2ymCxvoguyAfcZJ7VIpZrVbutRWwybHsNtGcWoOUsxDhJrjeCNHWuujuFbBZrB8jUS6TGnmGLiFcLa7vbcxGOCLCWGC68d164PwSEJ0+bUB59gBH1TIPnY5lbxRitFHmoCgToGPZW2nzWilz2ylWjPXZcGaMsUrDe7WspSp53+5YdofgqrUEuIo6h1Wo7IEqg8FfmIX5lx2fWipZjTZXFYLbhOC0Q/bLetf3uvKNUcPBSNlzespVF92tbtu0/aTThmrBf2AS7nR9b2M+IUXWTXC/WqDGsewWuc9G1/da9MDkEJpUKfcNNNG2IC1Nu0e10JQa1/dKEbT2KvCcCIagXDtT0Rq7hTz3AaieJwG8hnLL1hSCe20uAjxVa/uu1bHszYL3Ftf3OqMIjRFxv1nuU5BSpOE4CJwcAVQJjtuFh7T2x15GY3DDCayQ725xfW9zxMl8XhazDlNkMZjwQ1Sg4dXyeoxj2QeiIlv/jvJtvyfHm6iMhK+KeSxUKzAW4K4oX+vtssneRaVOvS3HXYDjWPa+jmVXhGUsyOdzgT8B81DBeOc7lr0zKk3pennm9+T676PaMl/iWPbBfcRFvTALHW6X5kelxHlnVIIh76eIpvFn0Vzfl/l9A+WTflpweopj2Vvn094MmIRyeVwO3C9ETZ/b+4DZjmXvpWuMpmDpWPZ0VObDr4E7ZU1+INd7Q17fB/4DeI5lH+pYdmU/ZK2MQ9Vv0KHF9b3b+1uok2fdCxWIeBFwE/Awqk7G+zJPr8v7l1ApZWc7lj0hG061QlwjHMs+HvgN8Ixc410Njy8LLv8A/MSx7KTx7KfInrsauBaYExQ0ymI1GOlY9ieBK2T/fSA04xW537PAbx3LPtGx7PHZ6AewO3CNjOsPwI+1+yVlT9uAL2u+FDBClKweViLHskfnsGCdZJjLX5KjosC9PEH2229Q7t0PBO8vyR55GZVm+j+y78m2R/S4JceyPy308ukQ3L8o8/c/hViHHMuucyz7WOAX2lgXCh4CujPPsewTHMuu768MtOEmABxhfF4szDKSdCaLxjy/ht655ogmezwqT/UEIfr/kk18lCEhjkDlp/8GuMGx7Km5zD6OZX8GVWTiKuATqOhac0yHoHzpdwNnOZZdFbJoxgCNwInAmahc7QtQ/skbZPNsY1x7N1Re9x2OZX+mmIUo50+id5GP+0vMKDJEiyIPxvU5VDri7wVnk0JO21lw+hfgesey9y5gDj4nxPBcVOCj7idPAPsLUU4LkQmDycA/gUtRMSmH0zsvPrjeVODrQpDO6getYbQwGR2eK1BIKiX8VoTY74gpeq8smtk44NPCjK91LHtc2DwJnmtl7u8QIXnHLNat/WU/fT+EPnxB9t3pqHiJE0P2cHC/rUWAuVsE/ikh99sBFXT5d+DXjmXvkQXPO6FSMmfI8QPHsrd3LPtEYWj3yL0OBkaZQkmRUCNMVofDQuiKvjaP1XhPuwhZK4jgQ9do5H6C+78KnnbNgqdjgF8JPZthFLAymf8Ewf3dQi93DrnmeKG9+xVgpRolfOFO4KtZxrqz3DMtON62P4SAYSEAaJNkapxvUEAKkZiJzDzyWsIzCszgpb1RbXzzwTHAhYEkH6K1f1II27SIw95eNMXjpQWwySRMP/VIoqXkTQKucyx7bBFMpUKYkwnNZUB/JuImnFHgvCIC3q2OZW8dcTNGJai7A5cGgqBx3cDVUyhj/gkq6LKcRGMk3bngPXA6QMFtY4r4zwkiOGerr3GhWPEKoY3mHjODhjdgxAQEmj+qeuS3Clg7p4rVZ2LI2gmzht0igs8ZhhCSoWfwXtE8xPW9ZcCThhKxS5b9OMVgrMtQtVeqC6D324ul47gCxrk38DsRPsLcJxWC+3NLaaUSuE6uW0W0iP0zRVGsyBYHFgsA4RM9wfhpCb0rTOUTIt4z/lNF72Il+YjBvwXpf0GlQ5lwomj2PQJ0HMueKJaFHYzzfdFg9gY+JeZFHbYCfi6aTlR4WyTdwFz4eMg5E1FBkMWsqzDp+Y2BWBtStOanIfPzKjBXNK/zhKiYBXL2FQ2iEPhA5vZ6sQi8FHJOSrQ1k3muQLmU7pL//lLGeAHwXdF2Lxbrlg47IjEXZWTGW9G7CtnCAdz6N4vV7Tax6lwu++B7MldOiHYKyhVXHcIEpsr/TOGyHWXyv1mY6Y0o19y9YklrDrHO5BRUJXbhKGCWcd5yYA4qgPZosRiZ+eWNYlmIgusjCC+ok3F9b20JcfFn4/PHs/SdOJqenU6XuL73RBQBQIqi1ch+PThEIfuV/PZlEfLeMs7ZGvieY9mTQsZ1HL0zlkC5ES6WNTVXhIi7yZPzr5n+v25cN4GyNM8SK9JBqMqR7xiXOFr2uukuKmnc3rBxAYgpzXyeFiKWZdU20gpDu68SU1I+eFG0i9GolMMgV3p7IeYZ45rTg9gC7d4Hidapw2Wu730GuFe6/v0b5fc0Ccc+wKFCWAJoC3n+tcAZru9NFkLyFTk+LgzJlFA/X4T0mQgzAZLFXB/12kW6IkCZZHczfv4LKpr9AmH880RKP5zecSDHO5a9WwRi+yCwp9yrEVXG+cuu7+2DinI2UxkPF21Gh0Uo0/9JQsi+C/xIiNAvUOmTc4R4mLnlu0vthXJZ2cL2waK+XrcPmo0ne+502Q+OWEKukLm6TDRf8wY1aNHm2v2/adCQhKyT7VHxRWfKGjlb1tRxwFeDAEgNWnNpenJujSFsADwia+cSlLvsftFK96J3lbkTHMversD52iwC09wQa2lf4e/GZwsYERKw+DHN8tAh2n82oSmbEGpWBX1XLA7ny16+QQSAKUCTKZgAqZBxfcKw4HaKsLcfKt7rCtmHX5c1d0FEfjLb+OkJlBX4atf3ml3fe9b1vcuBkw1FoQIVs2JadUeWEmnDKQtgTMgi2kjhaUjrQ0x6UQSl3wL/DILcNM2+07HsU2QD68T5QNFIW4Uo1AVWAQ2ed33PlsWakYUVBNFdjQpmazA0A13Y6AohRItd3wuk9XZtcXU4ln2PmAy/YJjOitEqTZ/nujxaeo0wNVNgqZZN/47re+8UaRk6ip5m9XdRkevrTZ+g63svOpZ9qWz+wJ9ci/KpXpbnls8B7+kNQbTrXiLrQO8fsAfKv/+BFtmcCTEh92IgkrXygCEw1hdiSi0QElmIT0uOcVai3B1mA5mE7NeVwIvF1DCXOe3Mtb9lnjY7lv2kCPZbG9Ytc52cZFxiPvA91/eWF7DWyEdzZL2NC9nvZ+lrUnuGVZKmrAfbHSJrJ58A1orysf8B1VRpvTE/pVofq4DHUP5/RMAY7/reOu1+O9AzxmEzKtaiEGveoagGYjqc5vre0rBUbceyZwpN1+ORjhPBY7Ocu6PQHtN6fI7re11ZUvjy1iWRoL9xhvJ1qet7r4fEIDzrWPaNYvGp0SwWR6CCW8sCwykIsCaL2a3QKkltFFfRKShw8uFi0YSADcDfjPOnGkwyCRxqnPN/YQRG3idE2tXhoxHGuTLb5nd9b40Qix5rJCyiNwKzqMhlAg2Bj6IajTxnHE/LZv1jkRrmtqg+Dz00Btf3XjBxZRD+VwxBOYp/vcIUQg0/7a3G+TuQJW4k1320sS4K0Y76W6jPVQxplGi1CwycLkBV5Py7aLdlAW2e1tM7Fsg0yW8Xwlj+VajQWQAcbXx+EliZLZ3X9b0/GYJFPb1dnmFwuut7R7m+d10gXJSpg2hbiBWg0VjLB9AzNmCV63v/KfA+ZmOo94HnwuZNvmsPETIOMfbJtiKM63BHNuZfgOXx48Z9npf9kA3+aVh6RhIeSxVbAEIgTGsqhpHXlFIw0hbP3fSsnT5amL5OvPc2/r4g20aVxflf4+sd85jSMoHFIUcWwlK6awYEzHwChTWPCdNiKyOY9vIRmGJggiFodQbzmo3YClFZbgg0uxRLNLX/PB4iMIzVz9FznOVzlWjL9bJekmKZqKSwgMZSQNh+qouwFnIJD0WXTw2po5DU9lWS7piFBvLHx+wSsnfeLoOmHIDpw14IrM1zn3dRWSIfMi4JFMs1h805LBWlFLY6HMt+yPh6JnClds9phtDyzyJuZQqMT6PiGXLtuycRV6bA7qj8++Cc8Sg3jw73lmC/72/w2BWBNSkL3XmNnm7SGsIzlWIBIATCTMy1RTDzMC2qFDWenw/5bry2CKvpHdU8x7Hst7Iw9QS900mimH8TeRbtBjmS2vmjinheU2BI6u6REChXSdikIVx0IcFBOZqVbHQs21xPdRI41pf+5GG51+PEz5cxTJiHofyouwnzGqUJAUExkZEhQnBZ6pHL+MLq2Y/uw2W7yOPuyMH8gzkbK8T9QJRZf7TMUz3dBWJGRRBAxxu0Yi2qR0W5mKaZTtkAXClFvrIJUhND9ns+33l/0vhFwmwDS+ShkkW0WlwepsD6x3x0KQS2CRGK8rl5zeDjsajsBf1zFHpdKJj42t+x7Muz7NGMrNNJxpxUlxNhw0YAcH1vSYjZdIwIAYU04tianpHOnZSm5ntYS9OgWlm2lJzPFXiPUnTCC/OrFlqfOyOWBJNo1+s+yBBrx+9E+BiJ8rlvXYLn2Yqe/r9MQNjzaJVhFow6iiz/KxCWERK0Pu3QKkxegIrvKFT6L9Z91Zf1lWuMG1DBWPvI+j81RGgtuH664CjjWPbRqLiMKRQeHJXIQwvbKW//CVPY34PeZuh8sJrB1X1yGSoQVndFNqLiD3aiZ/zLCtf3Hi9i7W4dsubzuReXZsO/BE2PziLMFA0S/2KuqclIZH9E6MhFq0oBw60QkElgJxCxZKMmPOxgaIwdRG9FW+hcJ/rAZLMt9EwZ5jVZhGb3dsj3k3PM/TpUFLaNStl7v0RjD2Mw1YNovwRNcwJz/9dQkcOTchCF5SjTrkkc6iJoun2BDSHC9JQc529GBaue7/reHJQLpM8CilYF8BqUmTUb89+EMqu+GLIv6geYVpViLS0fTM2FpMT1fENIDgKKd6NnavBfDbrbXuTayBRJA/LhorMMdKdQaM+iOMYCQBYwfeI7RNUMtI20YwgRe7sEYwvTZlvp9l+tzbIAgo55+Q5QpU/LAYUu5i56BtEFkDI2/YdzL8dm6bzYXkJBZoOhySXIkdaprYOaEMbb1770YfUkWjRiExSGMdefi/IZj3B9r9r1vW1c39sflaJm4qmcHcla6d2Wed9cc+n6Xrvre5tKRFR1OD9krz6KSqfaFahyfa/O9b09Uel77+RZ06ZlpzZQBMpUWGlTyJ5pj3h0yLoZjC2GX6ZnOtvHJTX1Y8Z5Nxr7rVgr1OgIfGzrbIKDZN1sjPifQoShMHdcIThuF/wuLieyhls3wMdQBXMC2F00qdeiNIGQGvmmlroReKEEYwuLdl6hp+HJotQJ01GGmWxIgMzlUpk73QJzFL0LhpQbWg1iWxHgOEcHtboQq8fmoKFIH2CfPGvAzAwB+Jzre/8w1+oACfBrRcjUI5MPyjK2soGU693PeP5ngM8YaWfBeGojWH1WGgS7HgkcLNMzmSbmO13fO6mIuRhsLYZfFYvL/hqP+SJwpMHEnyhy7MtChOqKPNcyrVSrDGF0VRZ6vaiPc2EKeb7re8cPJhwPNwvAPSHfnZqvUYo2uSl613h+2/W9KH6YTDbkBZKw8dNyQ4LfTO8yxLuVUQMpBIrRxluQaHsNZjiWXddfjS4CBmtYVyoDRpyjleiO9Aw2ygQaZDHj1v4zPURoueXpAAALxElEQVS4XJODUG0AHsrRBjjTz+tgDb2rGialzDL9uFa3oXdgZ7Pre+uyzFUn+YMN3wr5bpegsFYZnutl85mkEU3BwvZgAWFUHSg3wGbN0hL0xgjgtj6sXbPy5AHZ+Jg2l4eGXKNT+30NveO8Di/Bfn/TFColaLWg65YTx8NNAHiC3iUav4pWNlPPgzWKMYxHVRMzI95vioiwihyWhTpU+V+TAKw1mMGLxjnn6OOOsODKRYA3FvGfVahcbx3GI8V0cnQ0CxhbSZib63vv09uMtptj2fsH9w8ObaMdQM/KgR2iYebbjJkwvGgtYk0N7x16pxvqUI3h0zcq55luiq5yCgWSAfEsPV0qlcAPgy6X5h4zcFyqsSVCPlfn2Bdd9I6N6DKebQm93XCfEmEwa+vuPuy5x4zPhwJTw+avn/d6KeARuvPZE8L89fm/HegqkrGZdQN2AfbP0eCnjp5W4cBa1KHdf6kwax1OMmmvMd9tERj2c4alYSqqBkFOmt6fOB5OpYCDib/YIDZVqO5KFzmWPUnPsdaY0FGoOt/HGpddgqrpH0UKmyzSb4+FILnJ19C7o9tTBvFfF7K4j3Qs+/JAa44iIYacVwqiW3CqlgQE3RdiXvuKY9nXOpadq8JgBSUIitQ2z+OGOW5b4EdB200j735PVG3uemMOm6JYSqTok7m+dkL1BdgzRAjUfervhggA0038yrraKtBSNBhTLsKhXe8RVO61DtOAW6SLZY/5NHA8gtLEKKwy8JkApgb4DLnvASH7b3zIsz0Q8lyeY9m75Np3RVq0nqF3lsF1jmUfZs5frr0+mIQAbT6eRvXDCINlwEviey+GTj0SMm9XO5Y9OWRuEqgKraZb7V8GA38/xCKzn2PZPwrBc9Kx7ONQbo188A/juSYAPw1arUfBcch+aSslzoZTGmDw9gZUwR29tOM4VHT52Y5lLxJzX7sQzN2FGIQVCvma63ttEX0wX5Fc13eEmSdRJt2P0LszVgvwiDDJYPwdjmU/Qu+SpeehTOdBb/A2bWGMkWM7eYajgLe0sVZS3qjwfMLYw6jqd3rv7BpULfUTHMteLExvjTzPaLqLcmxbQiH2OhnDZG3uTgQecSw7jQpYHC+4OobeucYPSB+GfHCmCDabtbFNRAWjTjDG2gbcL9UXAwiLNbnWseyUjLFeW1N70Ltp1BHA1xzL/oXrey3l2F+u7y12LPs6YY4jtfk8ElVjfTkqaHaZaNnVMp+TCC+2k9eSEjKWFseyzSyRA4C/OZZ9C8p1sh0qQLEBlYJm7oNvOJb9qOt7z2j75Qp6lsEG1cjlKMeyg57wG+SZRss9xgEHB2nIcq2KCMLUem1dBrAPqlX0m4LvFs2qM1LuuTUqmv5mVDnrDYOJDmtz8B9Zp6Zl5l7CA55rIs7bSlT3U72Pwr7Ao1KIKMhC2F32sulWexZ4VhdApELi02IpTmjC92zHsk8V4aBDrIKTBBf1EfbMi45lP0XPNvUNwN1CzxdqtA9UvNRouccE4HHX987uqzK2RQgA2uJb7Vj22aia+Nsa2tR2chyURbrSYY7re+kCbl+Hav6Tz1wJquezb5qJRVP9MyrCWR/3ZDkyea6/Ez19mXVETIMsE7PocCz7AlRcxREG054gR0p7rkK1w4DB5hvHSsey56AKj1RpwlEQ86EHXyZChLUzDeKWDcaJ2TgKPBxYlzQtrlUI+xeMa/4kzxj1+bhQiIhtENdEifYXru9d71j2gcK8dMY6Wo4p9K79XwhOo6Sd3i6m3XoNn0cDn4w4V7sDtzmWfXoQaOv63nzHsm9FujRq150oR0OWZ9oZ1dVOx1kiz7psdyz7l8J0xhuWifHGvcL2++6ytzcMUnL8D+AbIQLAfXqgpgaj8ln9ZN7aHMu+GtVwbaph1Ztp4M7EwSbAc33v3RCB5XZU4yhdcayVe+wVcR1nQq57Dj3jZoIspEMRd0AOHIdp++NLiaRhFQOgEfznZIE8nWUSs6VLdYpE9nXX9+YWUTM7EXLosBnV4eu7umVBbxwkBPyOPOPOdv11IVJ1jfH/0RGeoaJI4h3GMFYKLu4ge0GlKOlr7Vn+Vx8iiIWN42axpqwPYU4VIWPoRAVlHuL63rIszL+YudkkZszTXN/boK8BSZebS++YhbAxttFd4z4Tco9cc5Iodn9pe+I8VOOStYSn9yUi4LUzZOxhlScTIWO5RUy57XnmKiNMcm2WNWRqVGejitl0FfBM5nybZa2rw9al63uvoDpxLo+41/X7FtPorK9QXQANfhRV/nyh0OHXBF8PG0JvACOJ2BLY9b03gC+Jxawrx5zpjHk58ANZN2Z8Dq7vvS3r+f0I9GkT4SmYo829IpbDzxIeR5ULxxnCq9uOLiVCh1saoI7Y5xzLPkI29OfErDxaNmegEXUIYlrEtPQk8EtZYIVGXwaRpHVy/UrZoJuFSC9D+ZG9sCYT2oJZJVHVtiycicLkamXMnUL8Nwlha5F7Pwa8FtJb/hmxegQEY36e51iBit4fpZkf1/WFYUj1v5Mcy24UfOxu4KJS7tNuPFur3Ps9IR5hxPufcr02lI/5zSzjSLi+d7Vj2Y+iCu3sL2MICE9GcNUic3AvcLHre2tzaP6viEm2UvBTq5kyq+mOPt8oa2AJcLvre7/KZlFwfe8Fx7JPRNUDCNICawQXG4WRLUH5qx+QPXyuWLW2kvGYWSsvokqbVmvPuLYvOJX3F4nGfL5oNGNEe9fT7oKc5k0y/lY5lqLy9peHCHqPo7rKBQLjM1ksEac4ln0xqj3reMFlhayFYE+/Jha3gBAfLxp6veBkg3HdTY5lHyP77yRt/wVld9uN53id3mVjH6HbjZcRRrU+yzP83bHs94Dviyl7rNCQKrpLJW/W7rlOmNS1hvsooEELNMGnhtJUBw1glcxni6z5ijBGqK2PU/K4CXR4W/Ac0LcuwktnB/9/UipB/i8qu2Ybbe0FeGqVMT+P6knwZNj9Nfpwh2PZ76Da/u4juKiVudwkc/kBqunRGlRb7g1yvy5zT2n072+OZR+uXXe00KoAxwHdC2jeepmPa0Ie/115nhEypjpgdbGZAgmGKYQ0CtlJTJPBhq6QCV8pG+pVrWBJXnOvY9krDHPMr1Ed/8YJgmtl464WpC0ImnZEqUkg72tR5TN3kjFXCdLXy8JeAnwQlvusvW4l/28NxpOrnr0EzgQaWBeq+MzyvuSihuBiojDtiXKvWiF0G4TArRSBaZHrexv1sZnBQxJdv6M8W3sObd2c221RzZe2E8bRJZv6PVRKWWuE56oTN0Y93fEYARHaStvUy1HtjN/Itb6M8SVQfvZAWGqX67whmqP535SYorcH7nF9702DIO8s62cd0CmCZp9SjELW6j5y/zEypxlNSF0l41/i+t6qPOujQvbqZlnvq/V4mZB7byMC0CR5xvVCqF8MuVfQknY7wc8/jTWmX3eECIo7aZamVlmfi1Gtn9eZGqVGb7rkHq2u77WGPKf5nymoQNHxsn465H5rRShdJFkt2eYhcJ8EFryKUuDZoCsTBCdVAZMNCegr9tr1spc/ZIQikOWbtyTKnbej7JWE/H8JKuBwUdj/wmhf8CyOZe8hpv8xso5XiYL1urY/xwmOKoE2iU3JxzemyHXHC/MOcLxG9seiYLwhYwrmKYj7Wi2C4bpiW2oPeygkSjZX+k2YAOBYdkY7vlHqsfT1OaOkmUS9ZymijYu9RinmotQ4KOezRD0nSrpYrt8HI05LuWajPqv5W6FzEyVtq5RrKWx8fZmfgdoLpaBJBdDrsu/PUuO41Ptgi7IA9INgYVoAvg382kwFjCGGGGKIIYbBCBXxFMQQQwwxxBBDLADEEEMMMcQQQwyxABCDCZqvpTqejRhiiCGGGGIBYAsBLcIzb65yDDHEEEMMMcQCwPADM02sjf7vzhZDDDHEEEMMsQDQH6C5AD4GzEA1EDoa+Cv9X5krhhhiiCGGGGIYACEghhhiiCGGGGKIIYYYYoghhhhiiCGGGGKIIYYYBin8f6QOdVsixkBdAAAAAElFTkSuQmCC" align="left">
# Simulación simple: hielo sobre una inclinación
En el laboratorio, investigamos cómo se desplazó nuestro "glacier goo" sobre varias inclinaciones. En este cuaderno, vamos a estudiar la misma cosa con una simulación de computadora.
Primero, configuramos las herramientas que usaremos. No necesitarás modificar estos.
```
# import oggm-edu helper package
import oggm_edu as edu
# import modules, constants and set plotting defaults
%matplotlib inline
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = (9, 6) # Default plot size
# Scientific packages
import numpy as np
# Constants
from oggm import cfg
cfg.initialize()
# OGGM models
from oggm.core.massbalance import LinearMassBalance
from oggm.core.flowline import FluxBasedModel, RectangularBedFlowline
# There are several numerical implementations in OGGM core.
# We use the "FluxBasedModel" with some default options
from functools import partial
FlowlineModel = partial(FluxBasedModel, min_dt=0, cfl_number=0.01)
```
Salga una caja roja? No te preocupes, esa caja presenta información sobre la configuración del modelo, `oggm.cfg`--no es un error.
## Lo esencial
Ahora configuramos nuestra primera inclinación y le decimos al modelo que queremos estudiar un glaciar allí.
### Base del glaciar
```
# define horizontal resolution of the model:
# nx: number of grid points
# map_dx: grid point spacing in meters
nx = 200
map_dx = 100
# define glacier slope and how high up it is
slope = 0.1
top = 3400 # m, the peak elevation
# create a linear bedrock profile from top to bottom
bottom = top - nx * map_dx * slope # m, elevation of the bottom of the incline based on the slope we defined
bed_h, surface_h = edu.define_linear_bed(top, bottom, nx)
# ask the model to calculate the distance from the top to the bottom of the glacier in km
distance_along_glacier = edu.distance_along_glacier(nx, map_dx)
# plot the glacier bedrock profile and the initial glacier surface
plt.plot(distance_along_glacier, surface_h, label='Initial surface')
edu.plot_xz_bed(x=distance_along_glacier, bed=bed_h);
```
Tenemos que decidir qué tan ancho es nuestro glaciar. Por ahora, utilizamos en la celda abajo la forma `RectangularBedFlowline` con un ancho `initial_width` de 300 m. La forma transversal "rectangular" significa que las paredes del "valle" glacial son rectas y el ancho *w* es el mismo por todo:

En este imagen, las lineas negras significan las paredes del "valle". La linea azul significa la superficie del glaciar.
**Qué significa *h* en el imagen--qué opinas?**
Si estás interesado/a, puedes leer más sobre formas de la base en la [documentación de OGGM](http://docs.oggm.org/en/latest/ice-dynamics.html#rectangular) (en inglés).
```
initial_width = 300 # width in meters
# Now describe the widths in "grid points" for the model, based on grid point spacing map_dx
widths = np.zeros(nx) + initial_width/map_dx
# Define our bed
init_flowline = RectangularBedFlowline(surface_h=surface_h, bed_h=bed_h, widths=widths, map_dx=map_dx)
```
Definimos un objeto llamado `init_flowline`, que almacena toda la información sobre la inclinación que usa el modelo. También nos puede dar información sobre el glaciar que tenemos allí. Por ejemplo, `volume_km3` puede decirnos cuánto hielo está almacenado actualmente en el glaciar.
**¿Cuánto volumen de hielo crees que hay ahora?**
```
print('Glacier length:', init_flowline.length_m)
print('Glacier area:', init_flowline.area_km2)
print('Glacier volume:', init_flowline.volume_km3)
```
Hasta ahora hemos configurado un valle. Hemos definido su inclinación, su ancho, su forma transversal--la geometría del hábitat en que va a crecer nuestro glaciar.
### Añadimos el hielo
Ahora queremos agregar hielo a nuestra inclinación y hacer un glaciar. Nosotros glaciólogos describimos la cantidad de hielo que se agrega o elimina sobre toda la superficie del glaciar como el "balance de masa" (en inglés: "mass balance"). Aquí abajo ves una ilustración del sitio web de OGGM:

Un balance de masa lineal se define por la altitud de línea de equilibrio (ELA, por el inglés "Equilibrium Line Altitude") y un gradiente de altitud (en [mm m$^{-1}$]). En arriba de la ELA, agregamos hielo (de nieve) y abajo de la linea removemos hielo (derritiéndolo). Aprenderemos más sobre esto en el [cuaderno 2](2_balance_de_masa.ipynb).
```
# ELA at 3000m a.s.l., gradient 4 mm m-1
ELA = 3000 # equilibrium line altitude in meters above sea level
altgrad = 4 # altitude gradient in mm/m
mb_model = LinearMassBalance(ELA, grad=altgrad)
```
El modelo de balance de masa, "mb_model", nos brinda el balance de masa para cualquier altitud que deseamos, en unidades [m s$^{-1}$]. Calculemos el balance de masa *anual* a lo largo del perfil del glaciar:
```
annual_mb = mb_model.get_annual_mb(surface_h) * cfg.SEC_IN_YEAR
# Plot it
plt.plot(annual_mb, bed_h, color='C2', label='Mass balance')
plt.xlabel('Annual mass balance (m yr-1)')
plt.ylabel('Altitude (m)')
plt.legend(loc='best')
# Display equilibrium line altitude, where annual mass balance = 0
plt.axvline(x=0, color='k', linestyle='--', linewidth=0.8)
plt.axhline(y=mb_model.ela_h, color='k', linestyle='--', linewidth=0.8);
```
**Qué notaste en el gráfico?** Que son los ejes? En cuales altitudes se agrega hielo al glaciar (balance de masa positivo)? En cuales altitudes se derrite el hielo?
### Ejecución del modelo
Cuando hicimos nuestros experimentos en el laboratorio, tuvimos una inclinación y algo de material para verter nuestro "glaciar". También decidimos cuándo comenzaría nuestro experimento, es decir, cuándo verteríamos el material. Configurar nuestro modelo de glaciar en la computadora es muy similar. Hemos recopilado nuestra inclinación (`init_flowline`), el balance de masa (`mb_model`), y ahora le diremos al modelo que comience en el año 0 (`y0 = 0`, para "year 0" en ingles).
```
# The model requires the initial glacier bed, a mass balance model, and an initial time (the year y0)
model = FlowlineModel(init_flowline, mb_model=mb_model, y0=0.)
```
Primero ejecutemos el modelo por un año:
```
runtime = 1
model.run_until(runtime)
edu.glacier_plot(x=distance_along_glacier, bed=bed_h, model=model, mb_model=mb_model, init_flowline=init_flowline)
print('Year:', model.yr)
print('Glacier length (m):', model.length_m)
print('Glacier area (km2):', model.area_km2)
print('Glacier volume (km3):', model.volume_km3)
```
El "glaciar" modelado ya llena toda el fundamento y su longitud sube hasta la ELA (línea discontinua), pero es extremadamente delgada.
Ahora podemos ejecutar el modelo durante 150 años.
**En el siguiente código, ¿puedes identificar dónde le decimos al modelo cuántos años queremos estudiar?**
```
runtime = 150
model.run_until(runtime)
edu.glacier_plot(x=distance_along_glacier, bed=bed_h, model=model, mb_model=mb_model, init_flowline=init_flowline)
```
Veamos cómo ha cambiado nuestro glaciar.
```
print('Current model year:', model.yr)
print('Glacier length (m):', model.length_m)
print('Glacier area (km2):', model.area_km2)
print('Glacier volume (km3):', model.volume_km3)
```
**Compara este "glaciar numérico" con el "goo glaciar" que hicimos en el laboratorio:**
- **cuanto tiempo necesita el goo glaciar para "crecer" y bajar en la inclinación? Cuanto tiempo necesita el glaciar numérico?**
- **bajo de la linea ELA (3000 m) ocurre algo para el glaciar numérico (que ocurre para glaciares reales también) que no ocurro en el goo glaciar: que es?**
Si queremos calcular más tiempo, tenemos que establecer la fecha deseada.
**Intenta editar la celda siguiente para pedirle al modelo que se ejecute durante 500 años en lugar de 150.**
```
runtime = 150
model.run_until(runtime)
edu.glacier_plot(x=distance_along_glacier, bed=bed_h, model=model, mb_model=mb_model, init_flowline=init_flowline)
print('Current model year:', model.yr)
print('Glacier length (m):', model.length_m)
print('Glacier area (km2):', model.area_km2)
print('Glacier volume (km3):', model.volume_km3)
```
Basado en esta información, ¿crees que modificaste la celda correctamente?
Es importante tener en cuenta que el modelo no se calculará atrás en el tiempo.
Una vez calculado para 500 años, el modelo no volverá a la fecha 450 años y se mantiene en año 500. Intenta ejecutar la celda de abajo. ¿La salida coincide con lo que esperabas?
```
model.run_until(450)
print('Current model year:', model.yr)
print('Glacier length (m):', model.length_m)
```
Para estudiar varios puntos en el tiempo, será útil almacenar algunos pasos intermedios de la evolución del glaciar. Hacemos un bucle "for" para que el modelo nos informe varias veces.
```
# Reinitialize the model
model = FlowlineModel(init_flowline, mb_model=mb_model, y0=0.)
# Year 0 to 600 in 5 years step
yrs = np.arange(0, 601, 5)
# Array to fill with data
nsteps = len(yrs)
length = np.zeros(nsteps)
vol = np.zeros(nsteps)
# Loop
for i, yr in enumerate(yrs):
model.run_until(yr)
length[i] = model.length_m
vol[i] = model.volume_km3
# I store the final results for later use
simple_glacier_h = model.fls[-1].surface_h
```
Ahora podemos trazar la evolución de la longitud y el volumen del glaciar en tiempo:
```
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(14, 5))
ax1.plot(yrs, length);
ax1.set_xlabel('Years')
ax1.set_ylabel('Length (m)');
ax2.plot(yrs, vol);
ax2.set_xlabel('Years')
ax2.set_ylabel('Volume (km3)');
```
**Qué paso en este imagen?**
La longitud del glaciar es una función escalonada en el primer año de simulación porque, por encima de la altitud de la línea de equilibrio (ELA), solo está pasando la acumulación.
Después de eso, al principio la longitud del glaciar permanece constante. El hielo no es muy espeso, por lo que la porción que llega por debajo del ELA no sobrevive a la fusión que se produce allí. Pero no importa qué tan espeso sea el glaciar, la parte sobre el ELA siempre está ganando masa. Eventualmente, hay suficiente hielo para persistir por debajo de la ELA. Aprenderemos más sobre el ELA en el [siguiente cuaderno](2_balance_de_masa.ipynb).
```
print('Glacier length from the top to the equilibrium line altitude ('+str(mb_model.ela_h)+' m) is: '
+ str(length[1])+'m')
```
Después de varios siglos, el glaciar se equilibra con su clima. Su longitud y volumen no cambiarán más si el clima se mantiene aproximadamente constante.
## Un primer experimento
Ok, hemos visto lo básico. Ahora, ¿qué tal un glaciar en una inclinación diferente? A veces los glaciares son muy escarpados, como este en Nepal.

Ajustemos la inclinación en el modelo para observar un glaciar más inclinado:
**En el siguiente código, ¿puedes identificar dónde le decimos al modelo la nueva inclinación que vamos a usar?**
**¿Que inclinación usamos para el primero experimento? Busca esa información al principio del cuaderno.**
```
# define new slope
new_slope = 0.2
top = 3400 #m, the peak elevation
# create a linear bedrock profile from top to bottom
bottom_1 = top - nx*map_dx*new_slope #m, elevation of the bottom of the incline based on the slope we defined
bed_1, surface_1 = edu.define_linear_bed(top, bottom_1, nx)
# Define new flowline
flowline_1 = RectangularBedFlowline(surface_h=surface_1, bed_h=bed_1, widths=widths, map_dx=map_dx)
```
Ahora ejecutaremos nuestro modelo con las nuevas condiciones iniciales (nuevamente durante 600 años), y almacenaremos la salida en una nueva variable para comparación:
```
# Reinitialize the model with the new input
model_1 = FlowlineModel(flowline_1, mb_model=mb_model, y0=0.)
# Array to fill with data
nsteps = len(yrs)
length_w = np.zeros(nsteps)
vol_w = np.zeros(nsteps)
# Loop
for i, yr in enumerate(yrs):
model_1.run_until(yr)
length_w[i] = model_1.length_m
vol_w[i] = model_1.volume_km3
# I store the final results for later use
new_glacier_h = model_1.fls[-1].surface_h
```
### Comparando nuestros glaciares
Comparemos ahora el glaciar que estudiamos primero con nuestro nuevo glaciar en una pendiente diferente. **¿Crees que el nuevo glaciar se verá más espeso o más delgado que el primero?**
```
# Plot the final results - first our previous glacier:
plt.plot(distance_along_glacier, simple_glacier_h, label='First glacier')
edu.plot_xz_bed(x=distance_along_glacier, bed=bed_h);
plt.ylim([1200, 3700]);
# and now our new glacier:
plt.plot(distance_along_glacier, new_glacier_h, label='New glacier')
edu.plot_xz_bed(x=distance_along_glacier, bed=bed_1)
plt.ylim([1200, 3700]); # same range as above for comparison
```
Probablemente ya puedas adivinar cómo se compara el volumen del nuevo glaciar con el anterior. Grafiquemos cómo el volumen de ambos glaciares crece a tiempo para ver:
```
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(14, 5))
ax1.plot(yrs, length, label='First glacier');
ax1.plot(yrs, length_w, label='New glacier');
ax1.legend(loc='best')
ax1.set_xlabel('Years')
ax1.set_ylabel('Length (m)');
ax2.plot(yrs, vol, label='First glacier');
ax2.plot(yrs, vol_w, label='New glacier');
ax2.legend(loc='best')
ax2.set_xlabel('Years')
ax2.set_ylabel('Volume (km3)');
```
### Actividad
**Ahora, trata de cambiar la inclinación a otro valor, y ejecutar el modelo otra vez. Qué ves?**
## Qué sigue?
Hacia adelante al [cuaderno 2](2_balance_de_masa.ipynb)!
| github_jupyter |
##### Copyright 2020 The EvoFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 & Creative Common licence 4.0
# EvoFlow and its tutorials are released under the Apache 2.0 licence
# its documentaton is licensed under the Creative Common licence 4.0
```
# Visualization callback setup
As seen above visualizing the route on the map is useful to quickly assess how good the current best route is so its a good idea to do the same while our model evolve its population.
To do this we are going to use a callback that will redraw the best route at the end of each generation. Callbacks, similarly to [keras.callbacks](https://keras.io/api/callbacks/) allows you to add your own custom codes at key points of the algorithms so it is easy to customize the evolution loop.
```
class MapCallback(Callback):
def __init__(self, chart_data, idx2city):
# init graph
self.chart_data = chart_data
self.idx2city = idx2city
self.fig = go.Figure(go.Scattergeo(
lat = chart_data['lat'],
lon = chart_data['lon'],
text = chart_data['name'],
marker_color= chart_data['population']))
self.fig.update_layout(title = 'Intial map', showlegend=False,
geo = go.layout.Geo(scope='europe', showframe = False,
projection_type = 'mercator',
lonaxis_range=lon_axis,
lataxis_range=lat_axis))
self.fig.show()
def on_generation_end(self, generation, metrics, fitness_scores, populations):
# get the best model
# the model only have one population hence the [0]
route = populations[0][0] # population is sorted by fitness score
best_distance = fitness_scores[0][0] # best fitness score = shortest distance
frames = []
initial_city = self.idx2city[int(route[0])]
for idx in range(len(route) - 1):
start_city = self.idx2city[int(route[idx])]
stop_city = self.idx2city[int(route[idx + 1])]
distance = distances[start_city['idx']][stop_city['idx']]
frames.append(go.Scattergeo(
lon = [start_city['lon'], stop_city['lon']],
lat = [start_city['lat'], stop_city['lat']],
mode = 'lines',
line = dict(width = 1,color = 'red')))
# last leg
frames.append(go.Scattergeo(
lon = [stop_city['lon'], initial_city['lon']],
lat = [stop_city['lat'], initial_city['lat']],
mode = 'lines',
line = dict(width = 1,color = 'red')))
self.fig.update_layout(title = 'Generation %d Total distance %d kms' % (generation, best_distance))
```
| github_jupyter |
<h1><font size=12>
Weather Derivatites </h1>
<h1> Rainfall Simulator -- Full modeling <br></h1>
Developed by [Jesus Solano](mailto:ja.solano588@uniandes.edu.co) <br>
16 September 2018
```
# Import needed libraries.
import numpy as np
import pandas as pd
import random as rand
import matplotlib.pyplot as plt
from scipy.stats import bernoulli
from scipy.stats import gamma
import pickle
import time
import datetime
```
# Generate artificial Data
```
### ENSO probabilistic forecast.
# Open saved data.
ensoForecast = pickle.load(open('../datasets/ensoForecastProb/ensoForecastProbabilities.pickle','rb'))
# Print an example .. ( Format needed)
ensoForecast['2005-01']
### Create total dataframe.
def createTotalDataFrame(daysNumber, startDate , initialState , initialPrep , ensoForecast ):
# Set variables names.
totalDataframeColumns = ['state','Prep','Month','probNina','probNino', 'nextState']
# Create dataframe.
allDataDataframe = pd.DataFrame(columns=totalDataframeColumns)
# Number of simulation days(i.e 30, 60)
daysNumber = daysNumber
# Simulation start date ('1995-04-22')
startDate = startDate
# State of rainfall last day before start date --> Remember 0 means dry and 1 means wet.
initialState = initialState
initialPrep = initialPrep # Only fill when initialState == 1
dates = pd.date_range(startDate, periods = daysNumber + 2 , freq='D')
for date in dates:
# Fill precipitation amount.
allDataDataframe.loc[date.strftime('%Y-%m-%d'),'Prep'] = np.nan
# Fill month of date
allDataDataframe.loc[date.strftime('%Y-%m-%d'),'Month'] = date.month
# Fill El Nino ENSO forecast probability.
allDataDataframe.loc[date.strftime('%Y-%m-%d'),'probNino'] = float(ensoForecast[date.strftime('%Y-%m')].loc[0,'El Niño'].strip('%').strip('~'))/100
# Fill La Nina ENSO forecast probability.
allDataDataframe.loc[date.strftime('%Y-%m-%d'),'probNina'] = float(ensoForecast[date.strftime('%Y-%m')].loc[0,'La Niña'].strip('%').strip('~'))/100
# Fill State.
allDataDataframe.loc[date.strftime('%Y-%m-%d'),'state'] = np.nan
simulationDataFrame = allDataDataframe[:-1]
# Fill initial conditions.
simulationDataFrame['state'][0] = initialState
if initialState == 1:
simulationDataFrame['Prep'][0] = initialPrep
else:
simulationDataFrame['Prep'][0] = 0.0
return simulationDataFrame
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-08-18', initialState = 1 , initialPrep = 0.4, ensoForecast = ensoForecast)
simulationDataFrame.head()
### Load transitions and amount parameters.
# Transitions probabilites.
transitionsParametersDry = pd.read_csv('../results/visibleMarkov/transitionsParametersDry.csv', sep = ' ', header=None, names = ['variable', 'value'])
transitionsParametersDry.index += 1
transitionsParametersDry
transitionsParametersWet = pd.read_csv('../results/visibleMarkov/transitionsParametersWet.csv', sep = ' ', header=None, names = ['variable', 'value'])
transitionsParametersWet.index += 1
transitionsParametersWet
amountParametersGamma = pd.read_csv('../results/visibleMarkov/amountGammaPro.csv', sep = ' ', header=None, names = ['variable', 'mu', 'shape'])
amountParametersGamma.index += 1
'''
# !!!!!! Delete !!!!!!!!!!!!!11.
amountParametersGamma = pd.read_csv('../results/visibleMarkov/fittedGamma.csv', index_col=0)
'''
print(transitionsParametersDry)
print('\n * Intercept means firts month (January) ')
```
## Simulation Function Core
```
### Build the simulation core.
# Updates the state of the day based on yesterday state.
def updateState(yesterdayIndex, simulationDataFrame, transitionsParametersDry, transitionsParametersWet):
# Additional data of day.
yesterdayState = simulationDataFrame['state'][yesterdayIndex]
yesterdayPrep = simulationDataFrame['Prep'][yesterdayIndex]
yesterdayProbNino = simulationDataFrame['probNino'][yesterdayIndex]
yesterdayProbNina = simulationDataFrame['probNina'][yesterdayIndex]
yesterdayMonth = simulationDataFrame['Month'][yesterdayIndex]
# Calculate transition probability.
if yesterdayState == 0:
# Includes month factor + probNino value + probNino value.
successProbabilityLogit = transitionsParametersDry['value'][1]+transitionsParametersDry['value'][yesterdayMonth] + yesterdayProbNino*transitionsParametersDry['value'][13] + yesterdayProbNina*transitionsParametersDry['value'][14]
if yesterdayMonth==1:
# Includes month factor + probNino value + probNino value.
successProbabilityLogit = transitionsParametersDry['value'][yesterdayMonth] + yesterdayProbNino*transitionsParametersDry['value'][13] + yesterdayProbNina*transitionsParametersDry['value'][14]
successProbability = (np.exp(successProbabilityLogit))/(1+np.exp(successProbabilityLogit))
elif yesterdayState == 1:
# Includes month factor + probNino value + probNino value + prep value .
successProbabilityLogit = transitionsParametersDry['value'][1]+ transitionsParametersDry['value'][yesterdayMonth] + yesterdayProbNino*transitionsParametersWet['value'][14] + yesterdayProbNina*transitionsParametersWet['value'][15] + yesterdayPrep*transitionsParametersWet['value'][13]
if yesterdayMonth==1:
# Includes month factor + probNino value + probNino value + prep value .
successProbabilityLogit = transitionsParametersDry['value'][yesterdayMonth] + yesterdayProbNino*transitionsParametersWet['value'][14] + yesterdayProbNina*transitionsParametersWet['value'][15] + yesterdayPrep*transitionsParametersWet['value'][13]
successProbability = (np.exp(successProbabilityLogit))/(1+np.exp(successProbabilityLogit))
else:
print('State of date: ', simulationDataFrame.index[yesterdayIndex],' not found.')
#print(successProbability)
#successProbability = monthTransitions['p'+str(yesterdayState)+'1'][yesterdayMonth]
todayState = bernoulli.rvs(successProbability)
return todayState
# Simulates one run of simulation.
def oneRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma):
# Define the total rainfall amount over the simulation.
rainfall = 0
# Total rainfall days.
wetDays = 0
# Loop over days in simulation to calculate rainfall ammount.
for day in range(1,len(simulationDataFrame)):
# Get today date.
dateOfDay = datetime.datetime.strptime(simulationDataFrame.index[day],'%Y-%m-%d')
# Update today state based on the yesterday state.
todayState = updateState(day-1, simulationDataFrame, transitionsParametersDry, transitionsParametersWet)
# Write new day information.
simulationDataFrame['state'][day] = todayState
simulationDataFrame['nextState'][day-1] = todayState
# Computes total accumulated rainfall.
if todayState == 1:
# Sum wet day.
wetDays+=1
# Additional data of day.
todayProbNino = simulationDataFrame['probNino'][day]
todayProbNina = simulationDataFrame['probNina'][day]
todayMonth = simulationDataFrame['Month'][day]
# Calculates gamma log(mu).
gammaLogMu = amountParametersGamma['mu'][1] + amountParametersGamma['mu'][todayMonth]+ todayProbNino*amountParametersGamma['mu'][13]+todayProbNino*amountParametersGamma['mu'][13]
#print(gammaMu)
# Calculates gamma scale
gammaLogShape = amountParametersGamma['shape'][1] + amountParametersGamma['shape'][todayMonth]+ todayProbNino*amountParametersGamma['shape'][13]+todayProbNino*amountParametersGamma['shape'][13]
#print(gammaShape)
if todayMonth==1:
# Calculates gamma log(mu).
gammaLogMu = amountParametersGamma['mu'][todayMonth]+ todayProbNino*amountParametersGamma['mu'][13]+todayProbNino*amountParametersGamma['mu'][13]
#print(gammaMu)
# Calculates gamma scale
gammaLogShape = amountParametersGamma['shape'][todayMonth]+ todayProbNino*amountParametersGamma['shape'][13]+todayProbNino*amountParametersGamma['shape'][13]
#print(gammaShape)
# Update mu
gammaMu = np.exp(gammaLogMu)
# Update shape
gammaShape = np.exp(gammaLogShape)
# Calculate gamma scale.
gammaScale = gammaMu / gammaShape
# Generate random rainfall.
todayRainfall = gamma.rvs(a = gammaShape, scale = gammaScale)
'''
# !!!!!! Delete !!!!!!!!!!!!!11.
todayRainfall = gamma.rvs(amountParametersGamma['Shape'][0],amountParametersGamma['Loc'][0],amountParametersGamma['Scale'][0])
'''
# Write new day information.
simulationDataFrame['Prep'][day] = todayRainfall
# Updates rainfall amount.
rainfall += todayRainfall
else:
# Write new day information.
simulationDataFrame['Prep'][day] = 0
yesterdayState = todayState
return rainfall,wetDays
updateState(0, simulationDataFrame, transitionsParametersDry, transitionsParametersWet)
# Run only one iteration(Print structure of results)
# Simulations iterations.
iterations = 10000
oneRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma)
```
## Complete Simulation
```
# Run total iterations.
def totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations):
# Initialize time
startTime = time.time()
# Array to store all precipitations.
rainfallPerIteration = [None]*iterations
wetDaysPerIteration = [None]*iterations
# Loop over each iteration(simulation)
for i in range(iterations):
simulationDataFrameC = simulationDataFrame.copy()
iterationRainfall,wetDays = oneRun(simulationDataFrameC, transitionsParametersDry, transitionsParametersWet, amountParametersGamma)
rainfallPerIteration[i] = iterationRainfall
wetDaysPerIteration[i] = wetDays
# Calculate time
currentTime = time.time() - startTime
# Print mean of wet days.
print('The mean of wet days is: ', np.mean(wetDaysPerIteration))
# Logging time.
print('The elapsed time over simulation is: ', currentTime, ' seconds.')
return rainfallPerIteration
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-12-18', initialState = 1 , initialPrep = 0.4, ensoForecast = ensoForecast)
simulationDataFrame.head()
```
## Final Results
```
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='steelblue',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
```
### Enero
```
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-01-01', initialState = 0 , initialPrep = 0.4, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='lightgreen',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-01-01', initialState = 1 , initialPrep = 0.4, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='skyblue',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-01-01', initialState = 1 , initialPrep = 5.0, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='steelblue',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
```
### Abril
```
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-04-01', initialState = 0 , initialPrep = 0.4, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='lightgreen',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-04-01', initialState = 1 , initialPrep = 0.4, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='skyblue',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-04-01', initialState = 1 , initialPrep = 5.0, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='steelblue',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
```
### Octubre
```
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-10-01', initialState = 0 , initialPrep = 0.4, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='lightgreen',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-10-01', initialState = 1 , initialPrep = 0.4, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='skyblue',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-10-01', initialState = 1 , initialPrep = 5.0, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='steelblue',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
```
# Financial Analysis
```
def calculatePrice(strikePrice, interestRate, finalSimulationData):
presentValueArray = [0]*len(finalSimulationData)
for i in range(len(finalSimulationData)):
tempDiff = finalSimulationData[i]-strikePrice
realDiff = max(0,tempDiff)
presentValue = realDiff*np.exp(-interestRate/12)
presentValueArray[i] = presentValue
print('The option price should be: \n ' , np.mean(presentValueArray))
calculatePrice(50,0.20,finalSimulation)
```
## Compare Data
```
import pickle
path = '../datasets'
allDataDataframe = pickle.load(open(path+'/fullDataset/completeDailyDataset.pickle','rb'))
allDataDataframe.tail()
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2011-08-01', initialState = 1 , initialPrep = 10, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='steelblue',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
allDataDataframe['2011-08-01':'2011-08-30']['state'].sum()
allDataDataframe['2011-08-01':'2011-08-30']['Prep'].sum()
allDataDataframe.ix['2012-04-01']
allDataDataframe['2012-01-01':'2012-01-30']
# 2012-04-26
manoMuGamma = np.exp(0.532197085346794 + 0.02*(-0.641890886355037) + 0.1*(-0.0188551459242916))
print(manoGamma)
manoShapeGamma = np.exp((-0.0667884480565946)+(0.00326485317705059)*0.02+(0.152284551058496)*0.1)
print(manoShapeGamma)
todayRainfall = gamma.rvs(a = manoShapeGamma, scale = manoGamma/manoShapeGamma)
todayRainfall
values = []
for i in range(1,100):
values.append(gamma.rvs(a = manoShapeGamma, scale = manoGamma/manoShapeGamma))
np.mean(values)
# 2012-01-26
manoMuGamma = np.exp(0.532197085346794 + 0.02*(-0.641890886355037) + 0.1*(-0.0188551459242916))
print(manoGamma)
manoShapeGamma = np.exp((-0.0667884480565946)+(0.00326485317705059)*0.02+(0.152284551058496)*0.1)
print(manoShapeGamma)
todayRainfall = gamma.rvs(a = manoShapeGamma, scale = manoGamma/manoShapeGamma)
todayRainfall
```
| github_jupyter |
# *CoNNear*: A convolutional neural-network model of human cochlear mechanics and filter tuning for real-time applications
Python notebook for reproducing the evaluation results of the proposed CoNNear model.
## Prerequisites
- First, let us compile the cochlea_utils.c file that is used for solving the transmission line (TL) model of the cochlea. This requires some C++ compiler which should be installed beforehand. Then go the connear folder from the terminal and run:
```
gcc -shared -fpic -O3 -ffast-math -o tridiag.so cochlea_utils.c
```
- Install numpy, scipy, keras and tensorflow
- If running on google colab: add the following as a code block and run it to compile cochlea_utils.c in the runtime machine.
```
!gcc -shared -fpic -O3 -ffast-math -o tridiag.so cochlea_utils.c
```
## Import required python packages and functions
Import required python packages and load the connear model.
```
import os
#os.add_dll_directory(os.getcwd())
import numpy as np
from scipy import signal
import scipy.signal as sp_sig
import matplotlib.pyplot as plt
import keras
from keras.models import model_from_json
from keras.utils import CustomObjectScope
from keras.initializers import glorot_uniform
from tlmodel.get_tl_vbm_and_oae import tl_vbm_and_oae
from helper_ops import *
json_file = open("connear/Gmodel.json", "r")
loaded_model_json = json_file.read()
json_file.close()
with CustomObjectScope({'GlorotUniform': glorot_uniform()}):
connear = model_from_json(loaded_model_json)
with CustomObjectScope({'GlorotUniform': glorot_uniform()}):
connear.load_weights("connear/Gmodel.h5")
connear.summary()
```
Define parameters
```
# Define model specific variables
down_rate = 2
fs = 20e3
fs_tl = 100e3
factor_fs = int(fs_tl / fs)
p0 = 2e-5
right_context = 256
left_context = 256
# load CFs
CF = np.loadtxt('tlmodel/cf.txt')
channels = CF.size
```
## Click response
Compare the responses of the models to a click stimulus.
**Notice that for all the simulations, TL model operates at 100kHz and the CoNNear model operates at 20kHz.**
```
#Define the click stimulus
dur = 128.0e-3 # for 2560 samples (2048 window length, 2x256 context)
stim = np.zeros((1, int(dur * fs)))
L = 70.0
samples = dur * fs
click_duration = 2 # 100 us click
click_duration_tl = factor_fs * click_duration
silence = 60 #samples in silence
samples = int(samples - right_context - left_context)
################# TL ##########################################
stim = np.zeros((1, (samples + right_context + left_context)*factor_fs))
stim[0, (factor_fs * (right_context+silence)) : (factor_fs * (right_context+silence)) + click_duration_tl] = 2 * np.sqrt(2) * p0 * 10**(L/20)
output = tl_vbm_and_oae(stim , L)
CF_tl_full = output[0]['cf']
channels_tl = CF_tl_full.size
CF_tl = CF_tl_full[::down_rate]
# basilar membrane motion for click response, the context samples (first and last 256 samples) are removed. Also downsample it to 20kHz
bmm_click_out_full = np.array(output[0]['v'])
stimrange = range(right_context*factor_fs, (right_context*factor_fs) + (factor_fs*samples))
bmm_click_tl = sp_sig.resample_poly(output[0]['v'][stimrange,::down_rate], fs, fs_tl)
bmm_click_tl = bmm_click_tl.T
################ CoNNear ####################################
stim = np.zeros((1, int(dur * fs)))
stim[0, right_context + silence : right_context + silence + click_duration] = 2 * np.sqrt(2) * p0 * 10**(L/20)
stim = np.expand_dims(stim, axis=2)
connear_pred_click = connear.predict(stim.T, verbose=1)
bmm_click_connear = connear_pred_click[0,:,:].T * 1e-6
```
Plotting the results.
```
################ Plots ######################################
#Plot input stimulus
plt.plot(stim[0,256:-256]), plt.xlim(0,2000)
plt.show()
# Plot the TL response
plt.imshow(bmm_click_tl, aspect='auto', cmap='jet')
plt.xlim(0,2000), plt.clim(-4e-7,5e-7)
plt.colorbar()
plt.show()
# Plot the CoNNear response
plt.imshow(bmm_click_connear, aspect='auto', cmap='jet')
plt.xlim(0,2000), plt.clim(-4e-7,5e-7)
plt.colorbar()
plt.show()
```
## Cochlear Excitation Patterns
Here, we plot the simulated RMS levels of basilar memberane (BM) displacement across CF for tone stimuli presented at SPLs between 0 and 90 dB SPL.
**You can change the `f_tone` variable to have tone stimuli of different frequencies, say 500Hz, 1kHz, 2kHz, etc..**
```
f_tone = 1e3 # You can change this tone frequency to see how the excitation pattern changes with stimulus frequency
dur = 102.4e-3 # for 2048 samples
window_len = int(fs * dur)
L = np.arange(0., 91.0, 10.) # SPLs from 0 to 90dB
total_length = window_len + right_context + left_context #total length = 2560
################# TL ##########################################
t = np.arange(0., dur, 1./fs_tl)
hanlength = int(10e-3 * fs_tl)
stim_sin = np.sin(2 * np.pi * f_tone * t)
han = signal.hanning(hanlength)
stim_sin[:int(hanlength/2)] = stim_sin[:int(hanlength/2)] * han[:int(hanlength/2)]
stim_sin[-int(hanlength/2):] = stim_sin[-int(hanlength/2):] * han[int(hanlength/2):]
total_length_tl = factor_fs * total_length
stim = np.zeros((len(L), total_length_tl))
for j in range(len(L)):
stim[j, factor_fs*right_context:factor_fs*(window_len+right_context)] = p0 * np.sqrt(2) * 10**(L[j]/20) * stim_sin
output = tl_vbm_and_oae(stim, L)
tl_target_tone_full = np.zeros((len(L), total_length_tl,channels_tl))
tl_target_tone = np.zeros((len(L), window_len,channels))
stimrange = range(right_context*factor_fs, (right_context*factor_fs) + (factor_fs*samples))
for i in range(len(L)):
tl_target_tone_full[i,:,:] = np.array(output[i]['v'])
tl_target_tone[i,:,:] = sp_sig.resample_poly(output[i]['v'][stimrange,::down_rate], fs, fs_tl)
tl_target_tone[i,:,:] = tl_target_tone[i,:,:] * 1e6
tl_target_tone_rms = np.vstack([rms(tl_target_tone[i]) for i in range(len(L))])
################ CoNNear ####################################
t = np.arange(0., dur, 1./fs)
hanlength = int(10e-3 * fs) # 10ms length hanning window
stim_sin = np.sin(2 * np.pi * f_tone * t)
han = signal.windows.hann(hanlength)
stim_sin[:int(hanlength/2)] = stim_sin[:int(hanlength/2)] * han[:int(hanlength/2)]
stim_sin[-int(hanlength/2):] = stim_sin[-int(hanlength/2):] * han[int(hanlength/2):]
stim = np.zeros((len(L), total_length))
for j in range(len(L)):
stim[j,right_context:window_len+right_context] = p0 * np.sqrt(2) * 10**(L[j]/20) * stim_sin
# prepare for feeding to the DNN
stim = np.expand_dims(stim, axis=2)
connear_pred_tone = connear.predict(stim, verbose=1)
bmm_tone_connear = connear_pred_tone
# Compute rms for each level
cochlear_pred_tone_rms = np.vstack([rms(bmm_tone_connear[i]) for i in range(len(L))])
################ Plots ######################################
# Plot the RMS for the TL
cftile=np.tile(CF_tl, (len(L),1))
plt.semilogx((cftile.T)/10e2, 20.*np.log10(tl_target_tone_rms.T))
plt.xlim(0.25,8.), plt.grid(which='both'),
plt.xticks(ticks=(0.25, 0.5, 1., 2., 4., 8.) , labels=(0.25, 0.5, 1., 2., 4., 8.))
plt.ylim(-80, 20)
plt.xlabel('CF (kHz)')
plt.ylabel('RMS of y_bm (dB)')
plt.title('TL Target')
plt.show()
# Plot the RMS for CoNNear
cftile=np.tile(CF, (len(L),1))
plt.semilogx((cftile.T), 20.*np.log10(cochlear_pred_tone_rms.T))
plt.xlim(0.25,8.), plt.grid(which='both'),
plt.xticks(ticks=(0.25, 0.5, 1., 2., 4., 8.) , labels=(0.25, 0.5, 1., 2., 4., 8.))
plt.ylim(-80, 20)
plt.xlabel('CF (kHz)')
plt.ylabel('RMS of y_bm (dB)')
plt.title('CoNNear Predicted')
plt.show()
```
## QERB Plots
Next, the level dependent tuning properties of the cochlea based on QERB are shown. For this, we choose click stimuli at three levels 0dB, 40dB and 70dB, compute their cochlear BM response and compute QERB from it.
```
L = [0., 40., 70.] # We will plot it for three SPLs
################# TL ##########################################
stim_tl = np.zeros(((len(L),int(dur * fs_tl))))
QERB_tl = np.zeros(((len(L),channels_tl)))
tl_predicted = np.zeros((len(L),(window_len)*factor_fs,channels_tl))
for i in range (len(L)):
stim_tl[i, (right_context + silence)*factor_fs : (right_context + silence + click_duration)*factor_fs] = 2 * np.sqrt(2) * p0 * 10**(L[i]/20)
#Get TL outputs
output = tl_vbm_and_oae(stim_tl, L)
for i in range(len(L)):
tl_predicted[i,:,:] = np.array(output[i]['v'])
QERB_tl[i,:] = QERB_calculation(tl_predicted[i, :, :].T, CF_tl_full*1e3, fs_tl)
################ CoNNear ####################################
stim_con = np.zeros(((len(L),int(dur * fs)+right_context+left_context,1)))
QERB_connear = np.zeros(((len(L),channels)))
for i in range (len(L)):
stim_con[i, right_context + silence : right_context + silence + click_duration, 0] = 2 * np.sqrt(2) * p0 * 10**(L[i]/20)
#Get CoNNear outputs
con_predicted = connear.predict(stim_con)
for i in range (len(L)):
QERB_connear[i,:] = QERB_calculation(con_predicted[i, :, :].T, CF*1e3, fs)
################ Plots ######################################
# Plot QERB of tl model
plt.semilogx(CF_tl_full[0::10]/10e2, QERB_tl[0,0::10]/10e2,':gs', label='0dB')
plt.semilogx(CF_tl_full[0::10]/10e2, QERB_tl[1,0::10]/10e2,'r', label='40dB')
plt.semilogx(CF_tl_full[0::10]/10e2, QERB_tl[2,0::10]/10e2,':rs', label='70dB')
plt.xlim(0.25,8.), plt.grid(which='both'),
plt.xticks(ticks=(0.25, 0.5, 1., 2., 4., 8.) , labels=(0.25, 0.5, 1., 2., 4., 8.))
plt.yticks(ticks=(5, 10, 15, 20) , labels=(5, 10, 15, 20))
plt.xlabel('CF (kHz)')
plt.ylim(2,20)
plt.ylabel('QERB')
plt.title('TL Target')
plt.legend()
plt.show()
# Plot QERB of CoNNear model
plt.semilogx(CF[0::5], (QERB_connear[0,0::5]),':gs', label='0dB')
plt.semilogx(CF[0::5], (QERB_connear[1,0::5]),'r', label='40dB')
plt.semilogx(CF[0::5], (QERB_connear[2,0::5]),':rs', label='70dB')
plt.xlim(0.25,8.), plt.grid(which='both'),
plt.xticks(ticks=(0.25, 0.5, 1., 2., 4., 8.) , labels=(0.25, 0.5, 1., 2., 4., 8.))
plt.yticks(ticks=(5, 10, 15, 20) , labels=(5, 10, 15, 20))
plt.xlabel('CF (kHz)')
plt.ylim(2,20)
plt.ylabel('QERB')
plt.title('CoNNear Predicted')
plt.legend()
plt.show()
```
## Speech Input
Here, a sentence from the Dutch speech matrix (unseen during training) will be input to both the TL and the CoNNear model. By adapting fragment_length parameter, various input lengths can be compared and visualised.
**Notice that this part is computationally more expensive with a higher fragment_duration, both in terms of memory and time.**
```
#load in speechfile
wavfile = 'dutch_sentence.wav'
signal_wav, fs_signal = wavfile_read(wavfile)
signalr = sp_sig.resample_poly(signal_wav, fs_tl, fs_signal)
L = np.array([70]) #sound-level of 70 dB SPL
stim_full = np.zeros((len(L), signalr.size))
for j in range(len(L)):
stim_full[j, :] = p0 * 10**(L[j]/20) * signalr/rms(signalr)
fragment_length = 12345 #define fragment length (max 40000 for the included wav-file)
stim_length_init = factor_fs*(fragment_length+right_context+left_context)
stim_length = stim_length_init
#adapt fragment duration if no multiple of 16 (due to the CNN character of CoNNear, we need multiples of 16)
zero_pad = fragment_length%16
zeros = 0
if zero_pad != 0:
zeros = 16-zero_pad
stim_length = factor_fs*(fragment_length+right_context+left_context+zeros)
################# TL ##########################################
stim = np.zeros((len(L), int(stim_length)))
stimrange = range(0, stim_length_init)
stim[:,stimrange] = stim_full[:,0:stim_length_init]
output = tl_vbm_and_oae(stim, L)
tl_target = []
tl_target = sp_sig.resample_poly(output[0]['v'][0:,::down_rate], fs, fs_tl)
tl_target = tl_target.T * 1e6
################ CoNNear ####################################
stim=sp_sig.resample_poly(stim, fs, fs_tl, axis=1)
stim=np.expand_dims(stim, axis=2)
tl_pred = connear.predict(stim)
tl_pred = tl_pred[0, :, :].T
################ Plots ######################################
fig, axarr = plt.subplots(4, sharex=True)
axarr[0].set_ylim(-0.35, 0.35)
axarr[0].plot(stim[0,(left_context):-(right_context),0])
axarr[0].set_title('Segment of Audio Input')
cax1 = axarr[1].imshow(tl_target[:,left_context:-right_context], cmap='bwr',aspect='auto', vmin=-0.5, vmax=0.5)
axarr[1].set_title('Output of TL-model')
axarr[1].set(ylabel='Center Frequency')
cax2 = axarr[2].imshow(tl_pred, cmap='bwr',aspect='auto', vmin=-0.5, vmax=0.5)
axarr[2].set_title('Output of CoNNear')
axarr[2].set(ylabel='Center Frequency')
cax3 = axarr[3].imshow(tl_pred-tl_target[:,left_context:-right_context], cmap='bwr',aspect='auto', vmin=-0.5, vmax=0.5)
axarr[3].set_title('Difference TL output and CoNNear')
axarr[3].set(ylabel='Center Frequency')
plt.show()
```
## DPOAE Plots
The frequency response of the 12-kHz CF channel is evaluated as a proxy for the otoacoustic emissions recorded in the ear-canal. Frequency responses of model simulations are shown in response to two pure tones of $f_{1,2}$ of 2.0 and 2.4 kHz.
The most pronounced distortion product in humans occurs at $2f_1 - f_2$ (1.6 kHz).
**Notice that this part is computationally expensive, both in terms of memory and time.**
```
# Create the stimulus
L = [70.0]
f1 = 2000.
f2 = 1.2 * f1
L2 = 50.
L1 = 39 + 0.4 * L2 # scissors paradigm
print ("The tone frequencies are " + str(f1) + " and " + str(f2))
print ("with levels " + str(L1) + " and " + str(L2))
trailing_silence = 0.
# Here we will pick a stimulus longer than 2048 samples to get a better FFT
# We will prepare it with fs_tl sampling frequency
dur_sin_samples = np.lcm(int(f1), int(f2))
min_duration = 0.25 # in seconds
if dur_sin_samples < (min_duration * fs_tl):
dur_sin = (((min_duration * fs_tl) // dur_sin_samples) + 1) * dur_sin_samples
else:
dur_sin = dur_sin_samples
dur_sin = (dur_sin / fs_tl)
t = np.arange(0., dur_sin, 1./fs_tl)
hanlength = int(10e-3 * fs_tl) # 10ms length hanning window
#f1
stim_sin1 = np.sin(2 * np.pi * f1 * t)
han = signal.windows.hann(hanlength)
stim_sin1 = p0 * np.sqrt(2) * 10**(L1/20) * stim_sin1
#f2
stim_sin2 = np.sin(2 * np.pi * f2 * t)
stim_sin2 = p0 * np.sqrt(2) * 10**(L2/20) * stim_sin2
stim_sin = stim_sin1 + stim_sin2
total_length = int(trailing_silence * fs_tl) + len(stim_sin)
stim = np.zeros((1, int(total_length)))
stimrange = range(int(trailing_silence * fs_tl), int(trailing_silence * fs_tl) + len(stim_sin))
stim[0, stimrange] = stim_sin
################# TL ##########################################
output = tl_vbm_and_oae(stim , L)
tl_out_full = []
downrate = 2 # downsampling across CF axis
tl_out_full = sp_sig.resample_poly(output[0]['v'][stimrange,::downrate], fs, fs_tl)
dpoae = sp_sig.resample_poly(output[0]['e'][stimrange,], fs, fs_tl)
tl_out_full = np.expand_dims(tl_out_full, axis=0)
################# CoNNear ####################################
# prepare for feeding to the CoNNear model
# first resample it to fs
shift_stim = 1
stim = stim[:, :]
stim = signal.decimate(stim, factor_fs, axis=1)
stim_1 = np.array(stim[0,:])
# window the signal into chunks of 2560 samples to be fed to the CoNNer model
stim = slice_1dsignal(stim_1, 2048, shift_stim, 256, left_context=256, right_context=256)
connear_out_chunks = connear.predict(stim,verbose=1)
# undo the windowing to get back the full response
connear_out_full = undo_window (connear_out_chunks, 2048, shift_stim, ignore_first_set=0)
connear_out_full = connear_out_full[:,:stim_1.shape[0],:] * 1e-6
##############################################################
f_cf = 12000.
tone_index, tone_cf = min(enumerate(CF*1000), key=lambda x: abs( x [1]- f_cf))
print("CF nearest to " + str(f_cf) + " is " + str(CF[tone_index]*1000))
scale_val = (p0* np.sqrt(2))
################ Plots ######################################
# Plot the DPOAE TL
tl_dpoae, nfft_tl = get_dpoae(tl_out_full, cf_location=tone_index)
freq_bins_tl = np.linspace(0, fs, num = nfft_tl)
plt.semilogx(freq_bins_tl[:int(nfft_tl/2)]/1000, 20 * np.log10(tl_dpoae/scale_val))
plt.title("DPOAEs - TL")
plt.xlabel('Frequency [kHz]'), plt.ylabel('Magnitudes dbSPL'), plt.xlim((0.25, 8)), plt.grid()
plt.show()
# Plot the DPOAE CoNNear
connear_dpoae, nfft_connear = get_dpoae(connear_out_full, cf_location=tone_index)
freq_bins_connear = np.linspace(0, fs, num = nfft_connear)
plt.semilogx(freq_bins_connear[:int(nfft_connear/2)]/1000, 20 * np.log10(connear_dpoae/scale_val))
plt.title("DPOAEs - CoNNear")
plt.xlabel('Frequency [kHz]'), plt.ylabel('Magnitudes dbSPL'), plt.xlim((0.25, 8)), plt.grid()
plt.show()
```
## RMS error between the excitation patterns
Here we will generate tones at different frequencies and levels, pass them through the TL and CoNNear model, compute the excitation patterns and compute the L2 loss between the excitation patterns. Then they are plotted as a scatter plot CFs vs RMS error for different levels.
```
L = np.arange(0., 91.0, 10.)
CFs = [250., 500., 1000., 2000., 4000., 8000., 10000., 12000.]
dur = 102.4e-3 # 2048 samples
window_len = int(fs * dur)
CF_TL = np.loadtxt('tlmodel/cf.txt')
################# TL ##########################################
t = np.arange(0., dur, 1./fs_tl)
hanlength = int(10e-3 * fs_tl)
# initialize an empty array for storing all the excitation patterns
y_bm_tl_all = np.zeros((len(L)*len(CFs), len(CF_TL)))
tl_pred_all= np.zeros((len(L)*len(CFs), window_len, len(CF_TL))) # to store all prediction ouputs
for idx, f_tone in enumerate(CFs):
stim_sin = np.sin(2 * np.pi * f_tone * t)
han = signal.hanning(hanlength)
stim_sin[:int(hanlength/2)] = stim_sin[:int(hanlength/2)] * han[:int(hanlength/2)]
stim_sin[-int(hanlength/2):] = stim_sin[-int(hanlength/2):] * han[int(hanlength/2):]
total_length_tl = factor_fs * (window_len + right_context + left_context)
stim = np.zeros((len(L), total_length_tl))
for j in range(len(L)):
stim[j, factor_fs*right_context:factor_fs*(window_len+right_context)] = p0 * np.sqrt(2) * 10**(L[j]/20) * stim_sin
output = tl_vbm_and_oae(stim, L)
tl_target_tone_full = np.zeros((len(L), total_length_tl,channels_tl))
tl_target_tone = np.zeros((len(L), window_len,channels))
stimrange = range(right_context*factor_fs, (right_context*factor_fs) + (factor_fs*samples))
for i in range(len(L)):
tl_target_tone_full[i,:,:] = np.array(output[i]['v'])
tl_target_tone[i,:,:] = sp_sig.resample_poly(output[i]['v'][stimrange,::down_rate], fs, fs_tl)
tl_target_tone[i,:,:] = tl_target_tone[i,:,:] * 1e6
tl_target_tone_rms = np.vstack([rms(tl_target_tone[i]) for i in range(len(L))])
y_bm_tl_all[ idx*len(L):(idx+1)*len(L) , :] = tl_target_tone_rms
tl_pred_all[idx*len(L):(idx+1)*len(L) , :, :] = tl_target_tone
################# CoNNear ####################################
t = np.arange(0., dur, 1./fs)
hanlength = int(10e-3 * fs)
# initialize an empty array for storing all the excitation patterns
y_bm_connear_all = np.zeros((len(L)*len(CFs), len(CF_TL)))
connear_pred_all= np.zeros((len(L)*len(CFs), window_len, len(CF_TL))) # to store all prediction ouputs
for idx, f_tone in enumerate(CFs):
stim_sin = np.sin(2 * np.pi * f_tone * t)
han = signal.windows.hann(hanlength)
stim_sin[:int(hanlength/2)] = stim_sin[:int(hanlength/2)] * han[:int(hanlength/2)]
stim_sin[-int(hanlength/2):] = stim_sin[-int(hanlength/2):] * han[int(hanlength/2):]
stim = np.zeros((len(L), int(len(stim_sin))))
total_length = window_len + right_context + left_context
stim = np.zeros((len(L), total_length))
for j in range(len(L)):
stim[j,right_context:window_len+right_context] = p0 * np.sqrt(2) * 10**(L[j]/20) * stim_sin
# prepare for feeding to the DNN
stim = np.expand_dims(stim, axis=2)
connear_pred_tone = connear.predict(stim, verbose=1)
bmm_tone_connear = connear_pred_tone
bmm_tone_connear.shape
connear_pred_tone_rms = np.vstack([rms(bmm_tone_connear[i]) for i in range(len(L))])
y_bm_connear_all[ idx*len(L):(idx+1)*len(L) , :] = connear_pred_tone_rms
connear_pred_all[idx*len(L):(idx+1)*len(L) , :, :] = bmm_tone_connear
################ Plots ######################################
L2Loss = [[] for _ in L]
cf_index = -1
for row in range(len(CFs) * len(L)):
level_index = row % len(L)
if level_index == 0:
cf_index += 1
l2_loss = np.sqrt(np.mean(np.square(y_bm_connear_all[row,:] - y_bm_tl_all[row,:])))
cf = CFs[cf_index]
L2Loss[level_index].append([cf, l2_loss])
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
colors=['red', 'blue', 'green', 'black', 'orange', 'cyan','gray','yellow','magenta']
for data, color, level in zip(L2Loss, colors, L):
x = []
y = []
for x_, y_ in data:
x.append(x_)
y.append(y_)
ax.scatter(x, y, alpha=0.8, color=color, edgecolors='none', s=30, label=level)
ax.set_xscale('log')
plt.title('Scatter plot of RMS Error')
plt.legend(loc="upper left")
plt.show()
```
| github_jupyter |
```
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
import torch.optim as optim
from matplotlib import pyplot as plt
import copy
import pickle
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print(device)
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
background_classes = {'cat', 'deer', 'dog', 'frog', 'horse','ship', 'truck'}
# print(type(foreground_classes))
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(5000): #5000*batch_size = 50000 data points
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
def imshow(img):
img = img / 2 + 0.5 # unnormalize
npimg = img#.numpy()
plt.imshow(np.transpose(npimg, axes = (1, 2, 0)))
plt.show()
# img1 = torch.cat((background_data[0],background_data[1],background_data[2]),1)
# imshow(img1)
# img2 = torch.cat((background_data[27],background_data[3],background_data[43]),1)
# imshow(img2)
# img3 = torch.cat((img1,img2),2)
# imshow(img3)
# print(img2.size())
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]].type("torch.DoubleTensor"))
j+=1
else:
image_list.append(foreground_data[fg_idx].type("torch.DoubleTensor"))
label = foreground_label[fg_idx] #-7 # minus 7 because our fore ground classes are 7,8,9 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
desired_num = 30000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(desired_num):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
qw=10010
print(fore_idx[qw])
imshow(mosaic_list_of_images[qw][fore_idx[qw]])
# print(mosaic_list_of_images[0])
print(classes[mosaic_label[qw]]) # add 7 as we had subtracted 7 while saving
# imshow(mosaic_list_of_images[13][2])
# print(type(mosaic_list_of_images[1][0]))
# print(mosaic_label)
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
batch = 250
msd = MosaicDataset(mosaic_list_of_images, mosaic_label , fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=True)
class Wherenet(nn.Module):
def __init__(self):
super(Wherenet, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
self.fc4 = nn.Linear(10,1)
def forward(self, z):
x = torch.zeros([batch,9],dtype=torch.float64)
y = torch.zeros([batch,3, 32,32], dtype=torch.float64)
x,y = x.to("cuda"),y.to("cuda")
for i in range(9):
x[:,i] = self.helper(z[:,i])[:,0]
x = F.softmax(x,dim=1) # alphas
x1 = x[:,0]
torch.mul(x1[:,None,None,None],z[:,0])
for i in range(9):
x1 = x[:,i]
y = y + torch.mul(x1[:,None,None,None],z[:,i])
return y , x
def helper(self,x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16 * 5 * 5)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.relu(self.fc3(x))
x = self.fc4(x)
return x
class Whatnet(nn.Module):
def __init__(self):
super(Whatnet, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
self.fc4 = nn.Linear(10,3)
def forward(self,y): #z batch of list of 9 images
y1 = self.pool(F.relu(self.conv1(y)))
y1 = self.pool(F.relu(self.conv2(y1)))
y1 = y1.view(-1, 16 * 5 * 5)
y1 = F.relu(self.fc1(y1))
y1 = F.relu(self.fc2(y1))
y1 = F.relu(self.fc3(y1))
y1 = self.fc4(y1)
return y1
test_images =[] #list of mosaic images, each mosaic image is saved as laist of 9 images
fore_idx_test =[] #list of indexes at which foreground image is present in a mosaic image
test_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(10000):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx_test.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
test_images.append(image_list)
test_label.append(label)
test_data = MosaicDataset(test_images,test_label,fore_idx_test)
test_loader = DataLoader( test_data,batch_size= batch ,shuffle=False)
where = Wherenet().double()
where = where.to(device)
# out_where,alphas = where(input1)
# out_where.shape,alphas.shape
what = Whatnet().double()
what =what.to(device)
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
cifar_acc = []
col1=[]
col2=[]
col3=[]
col4=[]
col5=[]
col6=[]
col7=[]
col8=[]
col9=[]
col10=[]
col11=[]
col12=[]
col13=[]
criterion = nn.CrossEntropyLoss()
optimizer_where = optim.SGD(where.parameters(), lr=0.01, momentum=0.9)
optimizer_what = optim.SGD(what.parameters(), lr=0.01, momentum=0.9)
nos_epochs = 150
train_loss=[]
test_loss =[]
train_acc = []
test_acc = []
ig = np.random.randint(0,250)
for epoch in range(nos_epochs): # loop over the dataset multiple times
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
running_loss = 0.0
cnt=0
c = 0
iteration = desired_num // batch
#training data set
for i, data in enumerate(train_loader):
inputs , labels , fore_idx = data
inputs,labels,fore_idx = inputs.to(device),labels.to(device),fore_idx.to(device)
# zero the parameter gradients
optimizer_what.zero_grad()
optimizer_where.zero_grad()
avg_inp,alphas = where(inputs)
outputs = what(avg_inp)
_, predicted = torch.max(outputs.data, 1)
# display plots
# if(c==0):
# disp_plot(inputs[ig,:],avg_inp[ig,:],ig,labels[ig].item(),predicted[ig].item(), alphas[ig,:], fore_idx[ig].item())
# c+=1
loss = criterion(outputs, labels)
loss.backward()
optimizer_what.step()
optimizer_where.step()
running_loss += loss.item()
if cnt % 40 == 39: # print every 6 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, cnt + 1, running_loss / 40))
running_loss = 0.0
cnt=cnt+1
if epoch % 5 == 4:
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
if epoch % 5 == 4:
col1.append(epoch)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
#************************************************************************
#testing data set
with torch.no_grad():
full_batch_true = 0
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for data in trainloader:
inputs,labels = data
inputs,labels = inputs.to(device), labels.to(device)
outputs = what(inputs.double())
_,predicted = torch.max(outputs.data,1)
#print(predicted.cpu().numpy(),"labels",labels.cpu().numpy())
full_batch_true+=sum(predicted.cpu().numpy()== labels.cpu().numpy())
cifar_acc.append(full_batch_true)
print("focibly_true_accuracy: ",full_batch_true)
for data in test_loader:
inputs, labels , fore_idx = data
inputs,labels,fore_idx = inputs.to(device),labels.to(device),fore_idx.to(device)
# print(inputs.shtorch.save(where.state_dict(),"model_epoch"+str(epoch)+".pt")ape,labels.shape)
avg_inp,alphas = where(inputs)
outputs = what(avg_inp)
_, predicted = torch.max(outputs.data, 1)
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
torch.save(where.state_dict(),"weights_forcibly_true/where_model_epoch"+str(epoch)+".pt")
torch.save(what.state_dict(),"weights_forcibly_true/what_model_epoch"+str(epoch)+".pt")
print('Finished Training')
torch.save(where.state_dict(),"weights_forcibly_true/where_model_epoch"+str(nos_epochs)+".pt")
torch.save(what.state_dict(),"weights_forcibly_true/what_model_epoch"+str(epoch)+".pt")
full_train_acc = 0
for data in train_loader:
inputs,labels,fore_idx = data
inputs,labels,fore_idx = inputs.to(device), labels.to(device),fore_idx.to(device)
avg_inp,alphas = where(inputs)
outputs = what(avg_inp)
_,predicted = torch.max(outputs.data,1)
#print(predicted.cpu().numpy(),"labels",labels.cpu().numpy())
full_train_acc+=sum(predicted.cpu().numpy()== labels.cpu().numpy())
print("mosaic_data_training_accuracy :",full_train_acc/30000)
full_test_acc = 0
for data in test_loader:
inputs,labels,fore_idx = data
inputs,labels,fore_idx = inputs.to(device), labels.to(device),fore_idx.to(device)
avg_inp,alphas = where(inputs)
outputs = what(avg_inp)
_,predicted = torch.max(outputs.data,1)
#print(predicted.cpu().numpy(),"labels",labels.cpu().numpy())
full_test_acc+=sum(predicted.cpu().numpy()== labels.cpu().numpy())
print("mosaic_data_test_accuracy :",full_test_acc/10000)
columns = ["epochs", "argmax > 0.5" ,"argmax < 0.5", "focus_true_pred_true", "focus_false_pred_true", "focus_true_pred_false", "focus_false_pred_false" ]
df_train = pd.DataFrame()
df_test = pd.DataFrame()
df_train[columns[0]] = col1
df_train[columns[1]] = col2
df_train[columns[2]] = col3
df_train[columns[3]] = col4
df_train[columns[4]] = col5
df_train[columns[5]] = col6
df_train[columns[6]] = col7
df_test[columns[0]] = col1
df_test[columns[1]] = col8
df_test[columns[2]] = col9
df_test[columns[3]] = col10
df_test[columns[4]] = col11
df_test[columns[5]] = col12
df_test[columns[6]] = col13
df_train
plt.plot(col1,col2, label='argmax > 0.5')
plt.plot(col1,col3, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.title("On Training set")
plt.show()
plt.plot(col1,col4, label ="focus_true_pred_true ")
plt.plot(col1,col5, label ="focus_false_pred_true ")
plt.plot(col1,col6, label ="focus_true_pred_false ")
plt.plot(col1,col7, label ="focus_false_pred_false ")
plt.title("On Training set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.show()
df_test
plt.plot(col1,col8, label='argmax > 0.5')
plt.plot(col1,col9, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.title("On Testing set")
plt.show()
plt.plot(col1,col10, label ="focus_true_pred_true ")
plt.plot(col1,col11, label ="focus_false_pred_true ")
plt.plot(col1,col12, label ="focus_true_pred_false ")
plt.plot(col1,col13, label ="focus_false_pred_false ")
plt.title("On Testing set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.show()
plt.plot(cifar_acc,"forcibly true accuracy")
plt.xlabel("every 5th epoch")
plt.ylabel("accuracy")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.title("forcibly true accuracy")
```
| github_jupyter |
# Lung Segmentation - Montgomery Dataset
```
%reload_ext autoreload
%autoreload 2
import os
import tempfile
import tensorflow as tf
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import cv2
from tensorflow.keras.models import load_model
import fastestimator as fe
from fastestimator.architecture import UNet
from fastestimator.dataset import montgomery
from fastestimator.trace import Dice, ModelSaver
from fastestimator.op.numpyop import ImageReader, Reshape, Resize
from fastestimator.op.tensorop import Augmentation2D, BinaryCrossentropy, Minmax, ModelOp
from fastestimator.op import NumpyOp
from fastestimator import RecordWriter
```
## Download and prepare the montgomery dataset
The montgomery dataset API will generate a summary CSV file for the data. The path of the csv file is returned as csv_path. The dataset path is returned as path. Inside the CSV file, the file paths are all relative to path.
```
train_csv_path, path = montgomery.load_data()
df = pd.read_csv(train_csv_path)
df.head()
```
## Create TF Records from input image and mask
RecordWriter api shall convert the training images and masks into 'TF Records'. Writing of training dataset into TF-Records is preceded by the following processing of the feature 'image', 'mask_left' and 'mask_left'. <br>
Processing( transformation ) are carried out using 'Operations'.<br>
Operation can be seen as processing step over feature(s). Operation can generate newer feature(s) or can update existing feature(s).
Preprocessing step for feature 'image' : <br>
'image' = ImageReader('image') <br>
'image' = Resize('image') <br>
'image' = Reshape('image') <br>
Preprocessing step for feature 'mask_left' and 'mask_right': <br>
'mask' = ImageReader('mask_left') + ImageReader('mask_right') <br>
'CombinedLeftRightMask' does the addition between left mask and right mask <br>
'mask' = Resize('mask') <br>
'mask' = Reshape('mask') <br>
```
class CombineLeftRightMask(NumpyOp):
def forward(self, data, state):
mask_left, mask_right = data
data = mask_left + mask_right
return data
writer = RecordWriter(
save_dir=os.path.join(path, "tfrecords"),
train_data=train_csv_path,
validation_data=0.2,
ops=[
ImageReader(grey_scale=True, inputs="image", parent_path=path, outputs="image"),
ImageReader(grey_scale=True, inputs="mask_left", parent_path=path, outputs="mask_left"),
ImageReader(grey_scale=True, inputs="mask_right", parent_path=path, outputs="mask_right"),
CombineLeftRightMask(inputs=("mask_left", "mask_right")),
Resize(target_size=(512, 512)),
Reshape(shape=(512, 512, 1), outputs="mask"),
Resize(inputs="image", target_size=(512, 512)),
Reshape(shape=(512, 512, 1), outputs="image"),
],
write_feature=["image", "mask"])
```
## Create data pipeline
Pipeline api generates sample batches one at a time, from TF Records. Preprocessing steps on feature 'image' and 'mask' are as follow:<br>
'image' = Augmentation('image') - rotation and flip being the type of augmentation <br>
'mask' = Augmentation('mask') - rotation and flip <br>
'image' = Minmax('image') <br>
'mask' = Minmax('mask') <br>
```
batch_size=4
epochs = 25
steps_per_epoch = None
validation_steps = None
pipeline = fe.Pipeline(
batch_size=batch_size,
data=writer,
ops=[
Augmentation2D(inputs=["image", "mask"], outputs=["image", "mask"], mode="train", rotation_range=10, flip_left_right=False),
Minmax(inputs="image", outputs="image"),
Minmax(inputs="mask", outputs="mask")
])
```
## Check the results of preprocessing
```
idx = np.random.randint(low=0,high=batch_size)
fig, ax = plt.subplots(nrows=1, ncols=2, sharex=True, sharey=True, figsize = (6,6), squeeze=True)
sample_data = pipeline.show_results(mode='train')
sample_img = sample_data[0]['image'][idx]
sample_mask = sample_data[0]['mask'][idx]
ax[0].imshow(np.squeeze(sample_img), cmap='gray')
ax[1].imshow(np.squeeze(sample_mask), cmap='gray')
plt.tight_layout()
plt.show()
```
## Creation of the network
FEModel api enables to create a model for training. It accepts a function as argument which return tf.keras.model,
it also accept optimizer type to perform optimization on network weights.
```
model_dir=tempfile.mkdtemp()
model = fe.build(model_def=lambda: UNet(input_size=(512, 512, 1)),
model_name="lung_segmentation",
optimizer=tf.keras.optimizers.Adam(learning_rate=0.0001),
loss_name="loss")
```
Network api enables user to constitute a network as a sequence of operations.
The model defined in the previous step is seen as an operation.
Operation can be seen as processing step over feature(s). Operation can generate newer feature(s) or can update existing feature(s).
Along with model defined in previous step, BinaryCrossEntropy is another operation added to the network. By default this operation generates a 'loss' and in this example, from feature 'pred_segment' which is prediction result and 'mask' which is ground truth.
```
network = fe.Network(ops=[
ModelOp(inputs="image", model=model, outputs="pred_segment"),
BinaryCrossentropy(y_true="mask", y_pred="pred_segment", outputs="loss")
])
```
Trace defined here are 'Dice' and 'ModelSaver'. Trace can be seen as tf.keras callbacks.
```
traces = [
Dice(true_key="mask", pred_key="pred_segment"),
ModelSaver(model_name="lung_segmentation", save_dir=model_dir, save_best=True)
]
```
Estimator api accepts network object, pipeline object , number of epochs and trace object and returns an estimator object which can orchestrate the training loop on the training data along with evaluaton on validation data.
```
estimator = fe.Estimator(network=network,
pipeline=pipeline,
epochs=epochs,
log_steps=20,
traces=traces,
steps_per_epoch=steps_per_epoch,
validation_steps=validation_steps)
estimator.fit()
```
## Let's do Inference
Inference in done here on validation batch set.
```
pipeline = fe.Pipeline(
batch_size=batch_size,
data=writer,
ops=[
Augmentation2D(inputs=["image", "mask"], outputs=["image", "mask"], mode="train", rotation_range=10, flip_left_right=False),
Minmax(inputs="image", outputs="image"),
Minmax(inputs="mask", outputs="mask")
])
sample_data = pipeline.show_results(mode='eval')
predict_model = load_model(os.path.join(model_dir, 'lung_segmentation_best_loss.h5'), compile=False)
predict_batch = predict_model.predict(sample_data[0]['image'])
```
Display a image, predicted mask and overlay version. Image sample is randomly selected from the validation batch.
```
idx = np.random.randint(low=0,high=batch_size)
img = sample_data[0]['image'][idx]
img = img.numpy()
img = img.reshape(512,512)
img_rgb = cv2.cvtColor(img, cv2.COLOR_GRAY2RGB)
img_rgb = (img_rgb*255).astype(np.uint8)
mask_gt = sample_data[0]['mask'][idx]
mask_gt = mask_gt.numpy()
mask_gt = mask_gt.reshape(512, 512)
mask_gt_rgb = cv2.cvtColor(mask_gt, cv2.COLOR_GRAY2RGB)
mask_gt_rgb = (mask_gt_rgb).astype(np.uint8)
mask = predict_batch[idx]
mask = mask.reshape(512,512)
mask_rgb = cv2.cvtColor(mask, cv2.COLOR_GRAY2RGB)
mask_rgb = (mask_rgb*255).astype(np.uint8)
ret, mask_thres = cv2.threshold(mask, 0.5,1, cv2.THRESH_BINARY)
mask_overlay = mask_rgb * np.expand_dims(mask_thres,axis=-1)
mask_overlay = np.where( mask_overlay != [0,0,0], [255,0,0] ,[0,0,0])
mask_overlay = mask_overlay.astype(np.uint8)
img_with_mask = cv2.addWeighted(img_rgb, 0.7, mask_overlay, 0.3,0 )
maskgt_with_maskpred = cv2.addWeighted(mask_rgb, 0.7, mask_overlay, 0.3, 0)
fig, ax = plt.subplots(nrows=1, ncols=3,figsize=(18,8))
ax[0].imshow(img_rgb)
ax[0].set_title('original lung')
ax[1].imshow(maskgt_with_maskpred)
ax[1].set_title('mask gt - predict mask')
ax[2].imshow(img_with_mask)
ax[2].set_title('img - predict mask ')
plt.show()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from itertools import combinations
from catboost import CatBoostClassifier, CatBoostRegressor
from sklearn.model_selection import train_test_split, KFold
from sklearn.metrics import mean_squared_error, accuracy_score, recall_score, precision_score, f1_score, roc_auc_score
import warnings
warnings.filterwarnings("ignore")
np.random.seed(42)
df = pd.read_csv('./train_merc.csv')
y = df.y
df.drop(['ID', 'y'], axis = 1, inplace=True)
cat_features_ids = np.where(df.apply(pd.Series.nunique) < 30000)[0].tolist()
pred = [10,10,10,10,10,10,10,10,10,10]
y_real = [10,10,10,10,10,10,10,10,10,100]
print(np.sqrt(mean_squared_error(pred, y_real)))
pred = [25,25,25,25,25,25,25,25,25,25]
y_real = [10,10,10,10,10,10,10,10,10,100]
print(np.sqrt(mean_squared_error(pred, y_real)))
train, test, y_train, y_test = train_test_split(df, y, test_size = 0.1)
clf = CatBoostRegressor(learning_rate=0.1, iterations=100, random_seed=42, logging_level='Silent')
clf.fit(train, y_train, cat_features=cat_features_ids)
prediction = clf.predict(test)
print('RMSE score:', np.sqrt(mean_squared_error(y_test, prediction)))
print('RMSLE score:', np.sqrt(mean_squared_error(np.log1p(y_test), np.log1p(prediction))))
df = pd.read_csv('./train_sample.csv.zip')
y = df.is_attributed
df.drop(['click_time', 'attributed_time', 'is_attributed'], axis = 1, inplace=True)
cat_features_ids = np.where(df.apply(pd.Series.nunique) < 30000)[0].tolist()
y_positive = np.ones_like(y)
y_negative = np.zeros_like(y)
print('\t\t $a(x)$ = 1 \t\t\n')
print('Accuracy all positive:', accuracy_score(y, y_positive))
print('Recall all positive:', recall_score(y, y_positive))
print('Precision all positive:', precision_score(y, y_positive))
print('F1 score all positive:', f1_score(y, y_positive))
print('Roc auc score all positive:', roc_auc_score(y, y_positive))
print('\n\n')
print('\t\t $a(x)$ = 0 \t\t\n')
print('Accuracy all negative:', accuracy_score(y, y_negative))
print('Recall all negative:', recall_score(y, y_negative))
print('Precision all negative:', precision_score(y, y_negative))
print('F1 score all negative:', f1_score(y, y_negative))
print('Roc auc score all positive:', roc_auc_score(y, y_negative))
print('\n\n')
print('\t\t Catboost \t\t\n')
train, test, y_train, y_test = train_test_split(df, y, test_size = 0.1)
clf = CatBoostClassifier(learning_rate=0.1, iterations=100, random_seed=42,
eval_metric='AUC', logging_level='Silent', l2_leaf_reg=3,
model_size_reg = 3)
clf.fit(train, y_train, cat_features=cat_features_ids)
prediction = clf.predict_proba(test)
print('Accuracy using Catboost:', accuracy_score(y_test, prediction[:, 1] > 0.5))
print('Recall using Catboost:', recall_score(y_test, prediction[:, 1] > 0.5))
print('Precision using Catboost:', precision_score(y_test, prediction[:, 1] > 0.5))
print('F1 score using Catboost:', f1_score(y_test, prediction[:, 1] > 0.5))
print('Roc auc score using Catboost:', roc_auc_score(y_test, prediction[:, 1]))
```
| github_jupyter |
# MARATONA BEHIND THE CODE 2021
## DESAFIO 2: QUANAM
##### Autor: Rodrigo Oliveira
##### LinkedIn: https://www.linkedin.com/in/rodrigolima82/
- `"ID":` número identificador da amostra
- `"ILLUM":` iluminação
- `"HUMID":` humidade
- `"CO2":` CO2
- `"SOUND":` som
- `"TEMP":` temperatura
- `"RYTHM":` ritmo cardíaco
# Parte 01. Importando as bibliotecas
```
# Pacotes padrao
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pylab as plt
%matplotlib inline
# Desativando warnings no jupyter
import warnings
warnings.filterwarnings('ignore')
```
# Parte 02. Carregando o Dataset
```
# Carregando o dataset IOT
iotDF = pd.read_csv('../../data/IOT.csv')
# Visualizando os primeiros registros
print(iotDF.shape)
iotDF.head()
# Carregando o dataset ANSWERS
answers = pd.read_csv('../../data/answers.csv')
# Visualizando os primeiros registros
print(answers.shape)
answers.head()
iotDF.describe()
answers.describe()
```
# Parte 03. EDA
```
sns.set(rc={'figure.figsize':(12, 12)})
corr = iotDF.corr()
plt.figure()
ax = sns.heatmap(corr, linewidths=.5, annot=True, cmap="YlGnBu", fmt='.1g')
plt.show()
```
# Parte 04. Tratamento dos dados
```
df = iotDF.copy()
# features do modelo
features = ['CO2','SOUND','TEMP','ILLUM','HUMID']
# coluna a ser prevista
target = ['RYTHM']
df.info()
```
# Parte 05. Modelagem Preditiva
```
# Models
from sklearn.linear_model import ElasticNet
# Misc
from sklearn.model_selection import train_test_split, cross_val_predict
from sklearn.metrics import r2_score
# Split features and labels
X = df[features]
y = df[target]
# Verificando o shape apos o split entre feature e target
X.shape, y.shape
# Separando o dataset em dados de treino e teste (considerando a proporção 70/30)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
# Elastic Net Regression
eNet = ElasticNet(alpha=0.0005, random_state=42)
# Fit do modelo com os dados de treino
model_enet = eNet.fit(np.array(X_train), np.array(y_train))
# Fazendo as previsoes nos dados de teste
y_pred = cross_val_predict(model_enet, X_test, y_test, cv=5)
# Previsão
score = r2_score(y_test, y_pred)
print("R2 nos dados de Teste: {}".format(score))
# Visualizar
fig, ax = plt.subplots()
ax.scatter(y_test, y_pred, edgecolors=(0, 0, 0))
ax.plot([y_test.min(), y_test.max()], [y_test.min(), y_test.max()], 'k--', lw=4)
ax.set_xlabel('Measured')
ax.set_ylabel('Predicted')
plt.show()
```
# Parte 06. Realizando Previsoes
```
# Deixando somente as features do modelo no dataset
X_test = answers[features]
# Realizando as previsoes
answers['RYTHM'] = model_enet.predict(np.array(X_test))
sns.set(rc={'figure.figsize':(8, 4)})
plt.figure()
plt.hist(answers['RYTHM'])
plt.show()
```
# Parte 07. Salvar resultado
```
answers.to_csv('../../submissao/ANSWERS.csv', index=False)
```
| github_jupyter |
```
from pyspark.mllib.linalg import SparseVector
from pyspark.mllib.linalg.distributed import RowMatrix
import numpy as np
from sklearn.metrics.pairwise import cosine_similarity
import time
from collections import defaultdict
from pyspark.sql import functions as sfunc
from pyspark.sql import types as stypes
import math
import sys
from pyspark.ml.linalg import SparseVector
from pyspark.mllib.linalg.distributed import RowMatrix
from operator import itemgetter
import operator
import random
schema = stypes.StructType().add("fv", stypes.StringType()).add("sku", stypes.StringType()).add("score", stypes.FloatType())
train_df = spark.read.csv('gs://lbanor/pyspark/train_query*.gz', header=True, schema=schema)
train_df.createOrReplaceTempView('test1')
train_df.head(3)
print(train_df.rdd.filter(lambda x: x.sku == 'FI911SHF89UBM-50').take(3))
# query = """
# SELECT
# sku,
# ROW_NUMBER() OVER (ORDER BY SUM(1)) -1 idx
# FROM test1
# GROUP BY 1
# """
# skus_rdd = spark.sql(query).rdd
query_statistics = """
SELECT
sku,
SQRT(10 * LOG(COUNT(sku) OVER()) / {threshold}) / SQRT(SUM(score * score)) p,
IF(SQRT(10 * LOG(COUNT(sku) OVER()) / {threshold}) > SQRT(SUM(score * score)), SQRT(SUM(score * score)), SQRT(10 * LOG(COUNT(sku) OVER()) / {threshold})) q --- implements the min(gamma, ||c||)
FROM test1
GROUP BY 1
"""
skus_stats = spark.sql(query_statistics.format(threshold=0.1))
print(skus_stats.rdd.filter(lambda x: x.sku == 'FI911SHF89UBM-50').take(3))
sku_stats.take(3)
print(skus_stats.rdd.filter(lambda x: x.sku == 'FI911SHF89UBM-50').take(3))
# query_statistics = """
# SELECT
# sku,
# {gamma} / SQRT(SUM(score * score)) p,
# IF({gamma} > SQRT(SUM(score * score)), SQRT(SUM(score * score)), {gamma}) q
# FROM test1
# GROUP BY 1
# """
# def get_gamma(threshold, numCols):
# return math.sqrt(10 * math.log(numCols) / threshold) if threshold > 10e-6 else math.inf
# gamma_b = sc.broadcast(get_gamma(10e-2))
# print(gamma_b.value)
# skus_stats = spark.sql(query_statistics.format(gamma=gamma_b.value))
# skus_stats.head(2)
pq_b = sc.broadcast({row.sku: [row.p, row.q] for row in skus_stats.collect()})
pq_b.value['FI911SHF89UBM-50']
#skus_idx_b = sc.broadcast({sku: idx for idx, sku in enumerate(pq_b.value.keys())})
#idx_skus_b = sc.broadcast({value: key for key, value in skus_idx_b.value.items()})
# d = {row.sku: row.idx for row in skus_rdd.collect()}
# db = sc.broadcast(d)
# id_ = {value: key for key, value in d.items()}
# id_b = sc.broadcast(id_)
#numCols = sc.broadcast(len(idx_skus_b.value))
# p = [0] * numCols.value
# for row in skus_stats
#p = {row.sku: gamma_b.value / row.norm for row in skus_stats.collect()} # if 0 happens as the ``norm`` we expected an Exception to be raised.
#p_b = sc.broadcast(p)
#q = {row.sku: gamma_b.value / row.norm for row in skus_stats.collect()}
#numCols.value
#skus_s['NI531SRM74IHX']
query_users_items = """
SELECT
data
FROM(
SELECT
fv,
COLLECT_LIST(STRUCT(sku, score)) data
FROM test1
GROUP BY 1
)
WHERE SIZE(data) BETWEEN 2 AND 200
"""
t0 = time.time()
users = spark.sql(query_users_items)
users_rdd = users.rdd
users.head(2)
def map_cosines(row):
for i in range(len(row)):
value_i = row[i].score / pq_b.value[row[i].sku][1]
if random.random() < pq_b.value[row[i].sku][0]:
for j in range(i + 1, len(row)):
value_j = row[j].score / pq_b.value[row[j].sku][1]
if random.random() < pq_b.value[row[i].sku][0]:
yield ((row[i].sku, row[j].sku), value_i * value_j)
users2 = users.rdd.flatMap(lambda row: map_cosines(row.data))
users2.take(2)
final = users2.reduceByKey(operator.add)
t0 = time.time()
print(final.take(3))
print(time.time() - t0)
import numpy as np
a = np.random.randn(12288, 150) # a.shape = (12288, 150)
b = np.random.randn(150, 45) # b.shape = (150, 45)
c = np.dot(a,b)
c.shape
b = np.random.randn(4, 1)
b
b[3]
a = np.random.randn(3, 3)
b = np.random.randn(3, 1)
c = a*b
a
b
c
```
| github_jupyter |
[View in Colaboratory](https://colab.research.google.com/github/findingfoot/ML_practice-codes/blob/master/Lasso_and_Ridge_regression.ipynb)
```
import warnings
warnings.filterwarnings('ignore')
import sys
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from sklearn import datasets
from tensorflow.python.framework import ops
regression_type = 'lasso'
ops.reset_default_graph()
sess = tf.Session()
iris = datasets.load_iris()
x_vals = np.array([x[3] for x in iris.data])
y_vals = np.array([y[0] for y in iris.data])
#declare the batch size
batch_size = 25
#define the placeholders
x_data = tf.placeholder(shape = [None,1], dtype=tf.float32)
y_target = tf.placeholder(shape = [None,1], dtype=tf.float32)
#set the seed
seed = 23
np.random.seed(seed)
tf.set_random_seed(seed)
#set the variables for the regression
A = tf.Variable(tf.random_normal(shape = [1,1]))
b = tf.Variable(tf.random_normal(shape = [1,1]))
#model output
model_output = tf.add(tf.matmul(x_data, A), b)
#Loss function
if regression_type == 'lasso':
# Declare Lasso loss function
# Lasso Loss = L2_Loss + heavyside_step,
# Where heavyside_step ~ 0 if A < constant, otherwise ~ 99
lasso_param = tf.constant(0.9)
heavyside_step = tf.truediv(1., tf.add(1., tf.exp(tf.multiply(-50., tf.subtract(A, lasso_param)))))
regularization_param = tf.multiply(heavyside_step, 99.)
loss = tf.add(tf.reduce_mean(tf.square(y_target - model_output)), regularization_param)
elif regression_type == 'ridge':
#Declare the Ridge loss function
# Ridge loss = L2_loss + L2 norm of slope
ridge_param = tf.constant(1.)
ridge_loss = tf.reduce_mean(tf.square(A))
loss = tf.expand_dims(tf.add(tf.reduce_mean(tf.square(y_target - model_output)),
tf.multiply(ridge_param, ridge_loss)), 0)
else:
print('Invalid regression_type parameter value',file=sys.stderr)
#optimizer
my_opt = tf.train.GradientDescentOptimizer(0.001)
train_step = my_opt.minimize(loss)
#initialize the variable
init = tf.global_variables_initializer()
sess.run(init)
#calculate the regression
loss_vec = []
for i in range(1500):
rand_index = np.random.choice(len(x_vals), size = batch_size)
rand_x = np.transpose([x_vals[rand_index]])
rand_y = np.transpose([y_vals[rand_index]])
sess.run(train_step, feed_dict={x_data: rand_x, y_target: rand_y})
temp_loss = sess.run(loss, feed_dict={x_data: rand_x, y_target: rand_y})
#print(temp_loss.shape)
loss_vec.append(temp_loss[0])
if (i+1)%300==0:
print('Step #' + str(i+1) + ' A = ' + str(sess.run(A)) + ' b = ' + str(sess.run(b)))
print('Loss = ' + str(temp_loss))
print('\n')
#get the regression coeeficients
[slope] = sess.run(A)
[intercept] = sess.run(b)
best_fit = []
for i in x_vals:
best_fit.append(slope*i+ intercept)
print(len(loss_vec))
import jupyterthemes as jt
from jupyterthemes import jtplot
jtplot.style(context='talk', fscale=1.4, spines=False, gridlines='--')
plt.plot(x_vals, y_vals, 'go', label = 'Data')
plt.plot(x_vals, best_fit, 'b-', label = 'Best_fit')
plt.legend(loc ='best')
plt.show()
#plot loss over time
plt.plot(loss_vec, 'r-')
plt.title(regression_type + ' Loss per Generation')
plt.xlabel('Generation')
plt.ylabel('Loss')
plt.show()
```
| github_jupyter |
# Datashading LandSat8 raster satellite imagery
Datashader is fundamentally a rasterizing library, turning data into rasters (image-like arrays), but it is also useful for already-rasterized data like satellite imagery. For raster data, datashader uses the separate [xarray](http://xarray.pydata.org/) library to re-render the data to whatever new bounding box and resolution the user requests, and the rest of the datashader pipeline can then be used to visualize and analyze the data. This demo shows how to work with a set of raster satellite data, generating images as needed and overlaying them on geographic coordinates using [HoloViews](http://holoviews.org), [GeoViews](http://geo.holoviews.org), and [Bokeh](http://bokeh.pydata.org).
This notebook currently relies on HoloViews 1.9 or above. Run `conda install -c ioam/label/dev holoviews` to install it.
```
import numpy as np
import xarray as xr
import holoviews as hv
import geoviews as gv
import datashader as ds
import cartopy.crs as ccrs
from holoviews.operation.datashader import regrid, shade
from bokeh.tile_providers import STAMEN_TONER
hv.extension('bokeh', width=80)
```
### Load LandSat Data
LandSat data is measured in different frequency bands, revealing different types of information:
```
import pandas as pd
band_info = pd.DataFrame([
(1, "Aerosol", " 0.43 - 0.45", 0.440, "30", "Coastal aerosol"),
(2, "Blue", " 0.45 - 0.51", 0.480, "30", "Blue"),
(3, "Green", " 0.53 - 0.59", 0.560, "30", "Green"),
(4, "Red", " 0.64 - 0.67", 0.655, "30", "Red"),
(5, "NIR", " 0.85 - 0.88", 0.865, "30", "Near Infrared (NIR)"),
(6, "SWIR1", " 1.57 - 1.65", 1.610, "30", "Shortwave Infrared (SWIR) 1"),
(7, "SWIR2", " 2.11 - 2.29", 2.200, "30", "Shortwave Infrared (SWIR) 2"),
(8, "Panc", " 0.50 - 0.68", 0.590, "15", "Panchromatic"),
(9, "Cirrus", " 1.36 - 1.38", 1.370, "30", "Cirrus"),
(10, "TIRS1", "10.60 - 11.19", 10.895, "100 * (30)", "Thermal Infrared (TIRS) 1"),
(11, "TIRS2", "11.50 - 12.51", 12.005, "100 * (30)", "Thermal Infrared (TIRS) 2")],
columns=['Band', 'Name', 'Wavelength Range (µm)', 'Nominal Wavelength (µm)', 'Resolution (m)', 'Description']).set_index(["Band"])
band_info
file_path = '../data/MERCATOR_LC80210392016114LGN00_B%s.TIF'
bands = list(range(1, 12)) + ['QA']
bands = [xr.open_rasterio(file_path%band).load()[0] for band in bands]
```
## Rendering LandSat data as images
The bands measured by LandSat include wavelengths covering the visible spectrum, but also other ranges, and so it's possible to visualize this data in many different ways, in both true color (using the visible spectrum directly) or false color (usually showing other bands). Some examples are shown in the sections below.
### Just the Blue Band
Using datashader's default histogram-equalized colormapping, the full range of data is visible in the plot:
```
nodata= 1
def one_band(b):
xs, ys = b['x'], b['y']
b = ds.utils.orient_array(b)
a = (np.where(np.logical_or(np.isnan(b),b<=nodata),0,255)).astype(np.uint8)
col, rows = b.shape
return hv.RGB((xs, ys[::-1], b, b, b, a), vdims=list('RGBA'))
%opts RGB [width=600 height=600]
tiles = gv.WMTS(STAMEN_TONER)
tiles * shade(regrid(one_band(bands[1])), cmap=['black', 'white']).redim(x='Longitude', y='Latitude')
```
You will usually want to zoom in, which will re-rasterize the image if you are in a live notebook, and then re-equalize the colormap to show all the detail available. If you are on a static copy of the notebook, only the original resolution at which the image was rendered will be available, but zooming will still update the map tiles to whatever resolution is requested.
The plots below use a different type of colormap processing, implemented as a custom transfer function:
```
from datashader.utils import ngjit
nodata= 1
@ngjit
def normalize_data(agg):
out = np.zeros_like(agg)
min_val = 0
max_val = 2**16 - 1
range_val = max_val - min_val
col, rows = agg.shape
c = 40
th = .125
for x in range(col):
for y in range(rows):
val = agg[x, y]
norm = (val - min_val) / range_val
norm = 1 / (1 + np.exp(c * (th - norm))) # bonus
out[x, y] = norm * 255.0
return out
def combine_bands(r, g, b):
xs, ys = r['x'], r['y']
r, g, b = [ds.utils.orient_array(img) for img in (r, g, b)]
a = (np.where(np.logical_or(np.isnan(r),r<=nodata),0,255)).astype(np.uint8)
r = (normalize_data(r)).astype(np.uint8)
g = (normalize_data(g)).astype(np.uint8)
b = (normalize_data(b)).astype(np.uint8)
return hv.RGB((xs, ys[::-1], r, g, b, a), vdims=list('RGBA'))
```
### True Color
Mapping the Red, Green, and Blue bands to the R, G, and B channels of an image reconstructs the image as it would appear to an ordinary camera from that viewpoint:
```
true_color = combine_bands(bands[3], bands[2], bands[1]).relabel("True Color (R=Red, G=Green, B=Blue)")
tiles * regrid(true_color)
```
Again, the raster data will only refresh to a new resolution if you are running a live notebook, because that data is not actually present in the web page; it's held in a separate Python server.
### False Color
[Other combinations](https://blogs.esri.com/esri/arcgis/2013/07/24/band-combinations-for-landsat-8/) highlight particular features of interest based on the different spectral properties of reflectances from various objects and surfaces, with full data redrawing on zooming if you have a live Python process:
```
%%opts RGB Curve [width=350 height=350 xaxis=None yaxis=None] {+framewise}
combos = pd.DataFrame([
(4,3,2,"True color",""),
(7,6,4,"Urban","False color"),
(5,4,3,"Vegetation","Color Infrared"),
(6,5,2,"Agriculture",""),
(7,6,5,"Penetration","Atmospheric Penetration"),
(5,6,2,"Healthy Vegetation",""),
(5,6,4,"Land vs. Water",""),
(7,5,3,"Atmosphere Removal","Natural With Atmospheric Removal"),
(7,5,4,"Shortwave Infrared",""),
(6,5,4,"Vegetation Analysis","")],
columns=['R', 'G', 'B', 'Name', 'Description']).set_index(["Name"])
combos
def combo(name):
c=combos.loc[name]
return regrid(combine_bands(bands[c.R-1],bands[c.G-1],bands[c.B-1])).relabel(name)
(combo("Urban") + combo("Vegetation") + combo("Agriculture") + combo("Land vs. Water")).cols(2)
```
All the various ways of combining aggregates supported by [xarray](http://xarray.pydata.org) are available for these channels, making it simple to make your own custom visualizations highlighting any combination of bands that reveal something of interest.
### Revealing the spectrum
The above plots all map some of the measured data into the R,G,B channels of an image, showing all spatial locations but only a restricted set of wavelengths. Alternatively, you could sample across all the measured wavelength bands to show the full spectrum at any given location:
```
%%opts Curve [width=800 height=300 logx=True]
band_map = hv.HoloMap({i: hv.Image(band) for i, band in enumerate(bands)})
def spectrum(x, y):
try:
spectrum_vals = band_map.sample(x=x, y=y)['z'][:-1]
point = gv.Points([(x, y)], crs=ccrs.GOOGLE_MERCATOR)
point = gv.operation.project_points(point, projection=ccrs.PlateCarree())
label = label = 'Lon: %.3f, Lat: %.3f' % tuple(point.array()[0])
except:
spectrum_vals = np.zeros(11)
label = 'Lon: -, Lat: -'
return hv.Curve((band_info['Nominal Wavelength (µm)'].values, spectrum_vals), label=label,
kdims=['Wavelength (µm)'], vdims=['Luminance']).sort()
spectrum(x=-9880000, y=3570000) # Location in Web Mercator coordinates
```
We can now combine these two approaches to let you explore the full hyperspectral information at any location in the true-color image, updating the curve whenever you hover over an area of the image:
```
%%opts Curve RGB [width=450 height=450] Curve [logx=True]
tap = hv.streams.PointerXY(source=true_color)
spectrum_curve = hv.DynamicMap(spectrum, streams=[tap]).redim.range(Luminance=(0, 30000))
tiles * regrid(true_color) + spectrum_curve
```
(Of course, just as for the raster data resolution, the plot on the right will update only in a live notebook session, because it needs to run Python code for each mouse pointer position.)
As you can see, even though datashader is not a GIS system, it can be a flexible, high-performance way to explore GIS data when combined with HoloViews, GeoViews, and Bokeh.
| github_jupyter |
## Linear regression using PyTorch built-ins
```
import torch.nn as nn
import numpy as np
import torch
# Input (temp, rainfall, humidity)
inputs = np.array([[73, 67, 43], [91, 88, 64], [87, 134, 58],
[102, 43, 37], [69, 96, 70], [73, 67, 43],
[91, 88, 64], [87, 134, 58], [102, 43, 37],
[69, 96, 70], [73, 67, 43], [91, 88, 64],
[87, 134, 58], [102, 43, 37], [69, 96, 70]],
dtype='float32')
# Targets (apples, oranges)
targets = np.array([[56, 70], [81, 101], [119, 133],
[22, 37], [103, 119], [56, 70],
[81, 101], [119, 133], [22, 37],
[103, 119], [56, 70], [81, 101],
[119, 133], [22, 37], [103, 119]],
dtype='float32')
inputs = torch.from_numpy(inputs)
targets = torch.from_numpy(targets)
inputs
```
## Dataset and DataLoader
```
from torch.utils.data import TensorDataset
# Define dataset
train_ds = TensorDataset(inputs, targets)
train_ds[0:3]
from torch.utils.data import DataLoader
# Define data loader
batch_size = 5
train_dl = DataLoader(train_ds, batch_size, shuffle=True)
for xb, yb in train_dl:
print(xb)
print(yb)
break
```
## nn.Linear
Instead of initializing the weights & biases manually, we can define the model using the `nn.Linear` class from PyTorch, which does it automatically.
```
# Define model
model = nn.Linear(3, 2)
print(model.weight)
print(model.bias)
# Parameters
list(model.parameters())
# Generate predictions
preds = model(inputs)
preds
```
## Loss Function
```
# Import nn.functional
import torch.nn.functional as F
# Define loss function
loss_fn = F.mse_loss
loss = loss_fn(model(inputs), targets)
print(loss)
```
## Optimizer
```
# Define optimizer
opt = torch.optim.SGD(model.parameters(), lr=1e-5)
```
## Train the model
1. Generate predictions
2. Calculate the loss
3. Compute gradients w.r.t the weights and biases
4. Adjust the weights by subtracting a small quantity proportional to the gradient
5. Reset the gradients to zero
```
# Utility function to train the model
def fit(num_epochs, model, loss_fn, opt, train_dl):
# Repeat for given number of epochs
for epoch in range(num_epochs):
# Train with batches of data
for xb,yb in train_dl:
# 1. Generate predictions
pred = model(xb)
# 2. Calculate loss
loss = loss_fn(pred, yb)
# 3. Compute gradients
loss.backward()
# 4. Update parameters using gradients
opt.step()
# 5. Reset the gradients to zero
opt.zero_grad()
# Print the progress
if (epoch+1) % 10 == 0:
print('Epoch [{}/{}], Loss: {:.4f}'.format(epoch+1, num_epochs, loss.item()))
fit(500, model, loss_fn, opt,train_dl)
# Generate predictions
preds = model(inputs)
preds
# Compare with targets
targets
```
| github_jupyter |
```
from PIL import Image, ImageDraw, ImageFont, ImageFilter, ImageChops
import pandas as pd
import numpy as np
import pathlib
import os
from fontTools.ttLib import TTFont
from fontTools.unicode import Unicode
characters = ['1','2','3','4','5','6','7','8','9']
IMG_WIDTH = 100
IMG_HEIGHT = 100
count = 0
def drawCharacter(path, character, font_path):
background_color = min(int(200 + np.random.normal() * 20),255)
foreground_color = int(20 + np.random.normal() * 20)
image = Image.new("L", (IMG_WIDTH, IMG_HEIGHT), (background_color,))
draw = ImageDraw.Draw(image)
font = ImageFont.truetype(font_path, 50)
w, h = draw.textsize(character, font=font)
draw.text(((IMG_WIDTH - w) / 2, (IMG_HEIGHT - h) / 2), character, (foreground_color), font=font)
blurred = image.filter(filter=ImageFilter.GaussianBlur(1))
left = IMG_WIDTH
top = IMG_HEIGHT
right = 0
bottom = 0
image_pixels = image.load()
for x in range(image.size[0]):
for y in range(image.size[1]): # For every row
if image_pixels[x,y] != background_color:
left = min(left, x)
right = max(right, x)
top = min(top, y)
bottom = max(bottom, y)
width = right - left + 4
height = bottom - top + 4
x = (left + right)/2
y = (top + bottom)/2
character_image = blurred.crop((x - width/2, y-height/2, x + width/2, y+height/2))
pixels = character_image.load() # create the pixel map
for x in range(character_image.size[0]):
for y in range(character_image.size[1]): # For every row
new_value = pixels[x,y] + np.random.normal() * 2
pixels[x,y] = (int(new_value),)
character_image.save(f'{path}/{count}.png')
required_ordinals = [ord(glyph) for glyph in ['1','2','3','4','5','6','7','8','9']]
def supportsDigits(fontPath):
font = TTFont(fontPath)
for table in font['cmap'].tables:
for o in required_ordinals:
if not o in table.cmap.keys():
return False
return True
image_lookup = [];
font_blacklist = [
'#',
"italic",
"Italic",
"VAZTEK",
"antique",
"aztek",
"RosewoodStd-Regular",
"ShishoniBrush",
"VAVOI",
"seguili",
'UVNThayGiaoNang_I',
'VAVOBI',
'VNI-Nhatban',
'VNI-Script',
'VNI-Trung Kien',
'segoeuii.ttf',
'VNI-Viettay',
'Brush',
'VREDROCK',
'UVNHaiBaTrung',
'UVNButLong',
'AmaticSC',
'UVNMucCham',
'UVNThayGiao_BI.TTF',
'VKUN',
'segoeuiz.ttf',
'seguibli.ttf',
'MyriadPro',
'Lobster-Regular',
'Bangers-Regular',
'VNI',
'VUSALI.TTF',
'VPEINOT.TTF',
'UVNSangSong_R',
'VDURHAM.TTF',
'Vnhltfap.ttf',
'UVNVienDu',
'UVNBucThu',
'UVNSangSong',
'VSCRIPT',
'VAUCHON',
'Vnthfap3',
'VCAMPAI',
'BungeeOutline',
'VBROADW',
'BungeeHairline-Regular',
'UVNMinhMap',
'scripti',
'UVN',
'brushsbi',
'Montserrat',
'VHELVCI.TTF',
'VFREE',
'BRUSH'
'seguis'
'VSLOGAN.TTF']
def check_blacklist(font):
for bl in font_blacklist:
if bl in font:
return False
return True
with open('fonts/fonts.list', 'r') as fonts:
for font in fonts:
font = font.strip()
can_use = supportsDigits(font);
if can_use and check_blacklist(font):
for character in characters:
path = f'testing_data/{character}' if (count % 10) == 0 else f'training_data/{character}'
image_lookup.append(f'{count}:{font}')
pathlib.Path(path).mkdir(parents=True, exist_ok=True)
drawCharacter(path, character, font)
count = count+1
else:
print(f'Skipping {font}')
with open("image_to_font_map.txt", "w") as map_file:
map_file.write("\n".join(image_lookup))
```
| github_jupyter |
```
import urllib
import pandas as pd
import numpy as np
import dask.dataframe as dd
import dask.bag as db
import dask.diagnostics as dg
# Determine which stations we need
# {column name:extents of the fixed-width fields}
columns = {"ID": (0,11), "LATITUDE": (12, 20), "LONGITUDE": (21, 30), "ELEVATION": (31, 37),"STATE": (38, 40),
"NAME": (41, 71), "GSN FLAG": (72, 75), "HCN/CRN FLAG": (76, 79),"WMO ID": (80, 85)}
df = pd.read_fwf("http://noaa-ghcn-pds.s3.amazonaws.com/ghcnd-stations.txt",
colspecs=list(columns.values()), names=list(columns.keys()),
dtype={'ID': str, 'LATITUDE':float, 'LONGITUDE':float,
'ELEVATION':float, 'STATE':str, 'NAME':str,
'GSN FLAG': str, 'HCN/CRN FLAG': str, 'WMO ID':str})
df[df['ID'].str.startswith('US')]['STATE'].unique()
# So lets get all the US country codes where the STATE is not in ['PR', 'VI'] 'cause it's easier to map
# Also we need a WMO ID 'cause thats how we cross reference against mos
# this is just a check that our filter is correct
mask = (df['ID'].str.startswith('US') & ~ df['STATE'].isin(['PR', 'VI']) & ~df['WMO ID'].isnull())
df[mask]['STATE'].unique()
df[mask]
# we're getting the list of US unique stations
us_stations = df[mask]
len(us_stations['ID'])
us_stations.head()
#Lets sample that!
#Do we really need all our stations?
%conda install geopandas
#
import geopandas as gpd
gdf = gpd.GeoDataFrame(us_stations, geometry=gpd.points_from_xy(us_stations['LONGITUDE'], us_stations['LATITUDE']))
gdf.plot(figsize=(10,5))
gdf.sort_values('LONGITUDE', ascending=False).head()
# let's drop them:
gdf_w = gdf[gdf['LONGITUDE']<0]
gdf_w.plot()
#We really don't need that dense of a network of plots, let's sample by 10 percent and see what happens
gdf_w.sample(frac=.10).plot(figsize=(10,5))
#1% which is about 6 stations
gdf_w.sample(frac=.10).plot(figsize=(10,5))
# try your own
#frac = ? I will try 10%
gdf_w.sample(frac=.10).plot(figsize=(10,5))
# lets use the 60 stations from our sample to filter down our dataset
us_stations = gdf_w.sample(frac=.1)['ID']
#Filter MOS down to our sample set of stations
#OUR MOS codes aren't matching up w/ our WMO above so we're
#gonna find the nearest neighbors to the above stations
#This is a spreadsheet of where the mos stations are
mdf = pd.read_csv("https://www.weather.gov/source/mdl/tables/MOS_stationtable_20190702.csv")
# us stations only have 4 values
mosdf = mdf[mdf['Station'].str.len()==4]
mgdf = gpd.GeoDataFrame(mosdf,geometry=gpd.points_from_xy(mosdf['Longitude'], mosdf['Latitude']))
len(mgdf['Station'].unique())
#Let's find our nearest neighbor
# compute distance between each station and all the others
def distance(row):
return mgdf['geometry'].distance(row['geometry'])
distances = gdf_w.apply(distance, axis=1)
distances.shape # rows in gdf_w x rows in mgdf
# get the id of the closet station by finding the index of the station with the minimum distance
# and then create a new column in our gdf_w with the closest MOS station
stationloc = np.argmin(distances.values,axis=1)
stationloc.shape, mgdf.shape, gdf_w.shape
# from mgdf pull the station closest in distance to the gdf station for that row
gdf_w['neighbor'] = mgdf.iloc[stationloc]['Station'].values
gdf_w['distance'] = distances.min(axis=1)
# this is our lookup comparison
gdf_w[['ID', 'neighbor']].head()
# this is our lookup comparison
gdf_w[['ID', 'neighbor']].head()
#lets merge in the mgdf since we're going to end up wanting the center lat/lon for plotting
gdmo = gdf_w.merge(mgdf, left_on = 'neighbor', right_on='Station', suffixes=('_ghcn', '_mos'))
gdmo.head()
#let's save out what we need:
gdmo[['ID', 'LATITUDE', 'LONGITUDE', 'Station', 'Latitude', 'Longitude']].to_csv("ghcn_mos_lookup.csv")
# the above lines up w/ mos, let's filter GHCN down to the us_stations
%conda install s3fs
# the above lines up w/ mos, let's filter GHCN down to the us_stations
YEAR = 2019
names = ['ID', 'DATE', 'ELEMENT', 'DATA_VALUE', 'M-FLAG', 'Q-FLAG', 'S-FLAG', 'OBS-TIME']
ds = dd.read_csv(f's3://noaa-ghcn-pds/csv/{YEAR}.csv', storage_options={'anon':True},
names=names, dtype={'DATA_VALUE':'object'})
# Here we're filtering to just the rows where these conditions are met
dsus = ds[ds['ID'].isin(gdmo['ID']) & ds['ELEMENT'].isin(['PRCP', 'TMAX', 'TMIN'])]
dsus.head()
# Lets only get the columns we want
dsus_subset = dsus[['ID', 'DATE', 'ELEMENT', 'DATA_VALUE', 'M-FLAG', 'Q-FLAG']]
dsus_subset.head()
# let's save out to a single largeish CSV for the GHCN data,
sample = 100
dsus.compute().to_csv(f"GHCN_{YEAR}.csv")
#Filter MOS down to just stations in our lookup
columns = ['station', 'short_model', 'model', 'runtime', 'ftime', 'N/X', 'X/N',
'TMP', 'DPT', 'WDR', 'WSP', 'CIG', 'VIS', 'P06', 'P12', 'POS', 'POZ',
'SNW', 'CLD', 'OBV', 'TYP', 'Q06', 'Q12', 'T06', 'T12']
mos = dd.read_csv(f'mos/modelrun/mav{YEAR}*.csv', names=columns)
mos.head()
# row filters
#N/X = nighttime minimum/daytime maximum surface temperatures.
#TMP = surface temperature valid at that hour.
#Q06 = quantitative precipitation forecast (QPF) category for liquid equivalent precipitation amount during a 6-h period ending at that time.
#Q12 = QPF category for liquid equivalent precipitation amount during a 12-h period ending at the indicated time.
mosus = mos[mos['station'].isin(gdmo['Station'])]
# column filter
mosv = mosus[['station', 'runtime', 'ftime', 'N/X', 'X/N','Q06', 'Q12']]
# save to csv , compute turns it to a dataframe so we only have 1 file to worry about
mosv.compute().to_csv(f"MOS_{YEAR}.csv")
import seaborn as sns
import matplotlib.pyplot as plt
sns.set(style="ticks", color_codes=True)
#Not using seaborns
#titanic = sns.load_dataset("titanic")
#sns.catplot(x="deck", kind="count", palette="ch:.25", data=titanic);
```
| github_jupyter |
# Arvo to PostgreSQL
## Generating a fully-functional CSV. (Err... repairing)
When my database processing script ran and saved as CSV something happened and corrupted the CSV.
As a result it only contained 44,000 rows.
Luckily I also saved a version as a .avro file.
Here's the steps I took to make this work.
* I pulled the .avro file into pandas.
* Traverse the array cell by cell and ensure that it is endcoding correctly.
* Then I processed the file into a CSV.
* Used my terminal & psql to send the csv to our AWS RDS PostgreSQL instance.
Here's the terminal commands I used to upload the CSV:
``` sh
foo@bar:~$ brew install postgres
foo@bar:~$ sudo mkdir -p /etc/paths.d && echo /Applications/Postgres.app/Contents/Versions/latest/bin | sudo tee /etc/paths.d/postgresapp
foo@bar:~$ psql "host=awsinstancename.awslocation.rds.amazonaws.com port=5432 dbname=lambdaRPG user=lambdaRPG"
foo@bar:~$ lambdaRPG=> \copy commentor_data (commentor, commentor_sentiment, commentor_total_happyness, commentor_total_saltiness, commentor_upvotes_mean, commentor_upvotes_total, qty_non_salty_comments, qty_salty_comments, salty_comments, sweet_comments, total_comments) from 'commentor_data.csv' CSV HEADER;
COPY 183926
```
I will likely always export my important df outputs to more than one filetype from now on as a precaution (in my personal projects).
### Import and install packages
```
#!pip install pandavro
import pandas as pd
import sqlite3
import psycopg2
import pandavro as pdx
import sqlalchemy
from sqlalchemy import create_engine
from sqlite3 import dbapi2 as sqlite
from tqdm import tqdm, tqdm_pandas
import pandavro as pdx
```
### Import and check dataframe shape
```
df = pdx.read_avro('data/hn_commentors_db.avro')
df.shape
df.tail()
```
### Make sure it isn't an encoding issue. Check each cell individually.
```
# This little section of code makes sure everything is encoding/decoding correctly.
for column in df.columns:
for idx in df[column].index:
x = df.get_value(idx,column)
try:
x = x if type(x) == str else str(x).encode('utf-8','ignore').decode('utf-8','ignore')
df.set_value(idx,column,x)
except Exception:
print('encoding error: {0} {1}'.format(idx,column))
df.set_value(idx,column,'')
continue
```
### Export the clean, intact data back to csv.
```
df.to_csv('data/commentor_data_repaired.csv',index=False)
```
### Reimport the data from CSV to DataFrame and inspect.
```
df2 = pd.read_csv("data/commentor_data_repaired.csv")
df2.tail()
df2.shape
```
# AND IT WORKED!
| github_jupyter |
# Ejemplos de programas en Python
Veamos aquí algunos ejemplos de Python.
## Números de Fibonacci
Los numéros de Fibonacci siguen una secuencia de números enteros y se caracterizan porque cada número en la secuencia (después de los primeros dos números) son la suma de los dos numeros precedentes ($x_n = x_{n-1} + x_{n-2}$).
Estos números son (los primeros 10): 0, 1, 1, 2, 3, 5, 8, 13, 21, 34, 55, ...
```
from IPython.display import Image
Image(filename = './flores.jpg')
# http://www.geohikers.es/matematicas-en-la-naturaleza-ii-y-sin-embargo-el-universo-tambien-es-euclidiano/
# programa para calculas la secuencia de Fibonacci
def recu_fibo(n):
"""Funcion recursiva para imprimir la secuencia de Fibonacci"""
if n <= 1:
return n
else:
return(recu_fibo(n-1) + recu_fibo(n-2))
nterminos = 10
# si queremos que el usuario determine el numero de terminos
# nterminos = int(input("¿Cuantos terminos quiere?"))
# hacemos una revision para ver si el numero es valido
if nterminos <= 0:
print('Por favor ingrese un entero positivo')
else:
print('Secuencia de Fibonacci')
for i in range(nterminos):
print(recu_fibo(i))
```
## Triangulo de Pascal
Cuando se eleva un binomio al cuadrado se tiene lo siguiente $(x + y)^2 = x^2 + 2 x y + y^2$; al cubo $(x + y)^3 = x^3 + 3 x^2 y + 3 x y^3 + y^3$ y así sucesivamente. Estos coeficientes que acompañan a la(s) variables pueden obtenerse el triangulo de Pascal.
```
from IPython.display import Image
Image(filename = './pascals3.jpg')
# http://jwilson.coe.uga.edu/EMAT6680Su12/Berryman/6690/BerrymanK-Pascals/BerrymanK-Pascals.html
def pascal_triangulo(n):
trow = [1]
y = [0]
for x in range(max(n,0)):
print(trow)
trow = [l + r for l,r in zip(trow + y, y + trow)]
return n >= 1
pascal_triangulo(8)
```
## Adivine el número
```
import random
adivina_real = 0
nombre = input("¡Hola! ¿Cual es tu nombre?")
numero = random.randint(1, 20)
print('Bueno, {0}, estoy pensando en un numero entre el 1 y el 20.'.format(nombre))
while adivina_real < 6:
adivina = int(input("¿Que numero estoy pensando?: "))
adivina_real += 1
if adivina < numero:
print('Estas muy abajo')
if adivina > numero:
print('Estas muy arriba')
if adivina == numero:
break
if adivina == numero:
print('¡Excelente, {0}! Adivinaste mi numero en {1} intento(s)'.format(nombre,adivina_real))
else:
print("No, el numero que estaba pensando era {0}".format(numero))
```
| github_jupyter |
```
%matplotlib inline
```
# Overview
**This code is for analyzing patterns of the different weight files which should have already been computed**
**Basic imports here**
```
import numpy as np
import glob
from scipy import stats
from matplotlib import pyplot as plt
```
**Just some details for making prettier plots**
```
import matplotlib
matplotlib.rcParams['xtick.labelsize'] = 10
matplotlib.rcParams['ytick.labelsize'] = 10
matplotlib.rcParams['axes.labelsize'] = 12
matplotlib.rcParams['axes.titlesize'] = 12
matplotlib.rcParams['axes.grid'] = True
matplotlib.rcParams['grid.color'] = '0.5'
matplotlib.rcParams['grid.linewidth'] = '0.5'
matplotlib.rcParams['axes.edgecolor'] = '0.25'
matplotlib.rcParams['xtick.color'] = '0'
matplotlib.rcParams['ytick.color'] = '0'
matplotlib.rcParams['xtick.major.width'] = 1
matplotlib.rcParams['ytick.major.width'] = 1
matplotlib.rcParams['ytick.major.size'] = 5
matplotlib.rcParams['xtick.major.size'] = 5
matplotlib.rcParams['axes.spines.right'] = True
matplotlib.rcParams['axes.spines.left'] = True
matplotlib.rcParams['axes.spines.top'] = True
matplotlib.rcParams['axes.spines.bottom'] = True
matplotlib.rcParams['font.family'] = 'sans-serif'
matplotlib.rcParams['font.sans-serif'] = 'helvetica'
matplotlib.rcParams['font.weight']='normal'
matplotlib.rcParams['axes.axisbelow'] = True
```
**And I use this for setting up the creation of new results folders depending on the month/year. This is where files/images will be saved so feel free to alter accordingly**
```
import datetime
year = datetime.date.today().year
month = datetime.date.today().month
import os
figs_dir = '../Results/Figures/{}_{:02}'.format(year, month)
if not os.path.exists(figs_dir):
os.makedirs(figs_dir)
```
**Functions for plotting the lorenz curve and calculating the GINI coefficient. Largely stolen from online sources and triple checked to make sure they make sense/work as intended**
```
def get_gini(arr):
count = arr.size
coefficient = 2. / count
indexes = np.arange(1, count + 1)
weighted_sum = (indexes * arr).sum()
total = arr.sum()
constant = (count + 1) / count
return coefficient * weighted_sum / total - constant
def get_lorenz(arr):
X_lorenz = arr.cumsum() / arr.sum()
X_lorenz = np.insert(X_lorenz, 0, 0)
return (np.linspace(0.0, 1.0, X_lorenz.size), X_lorenz)
```
# Looking at an example protein
```
example_prot_name = '1aoeA'
methods_of_interest = ['uniform', 'simple_0.8', 'HH_meanScale', 'GSC_meanScale', 'ACL_meanScale']
weights_dict = {}
for method in methods_of_interest:
weights_dict[method] = []
for weight_file in sorted(glob.glob('../Data/weights/{}*.weights'.format(example_prot_name)))[:]:
prot_name = weight_file.split('/')[-1].split('_')[0]
method = weight_file.split('/')[-1].replace(prot_name+'_', '').replace('.weights', '')
if method in methods_of_interest:
weights = np.genfromtxt(weight_file)
weights_dict[method] = weights
```
** These are most certainly not the most efficiently drawn plots but look nice! **
```
fig, ax = plt.subplots(figsize=(5,3))
ginis = []
for method_of_interest in methods_of_interest[:]:
print(method_of_interest)
X = np.array(sorted(weights_dict[method_of_interest]))
ginis.append(get_gini(X))
x,y = get_lorenz(X)
ax.plot(x, y, marker='')
ax.lines[0].set_label('Uniform ({:.2f})'.format(ginis[0]))
ax.lines[1].set_label('0.8 sequence\n identity ({:.2f})'.format(ginis[1]))
ax.lines[2].set_label('HH mean\n scaled ({:.2f})'.format(ginis[2]))
ax.lines[3].set_label('GSC mean\n scaled ({:.2f})'.format(ginis[3]))
ax.lines[4].set_label('ACL mean\n scaled ({:.2f})'.format(ginis[4]))
box = ax.get_position()
ax.set_position([box.x0, box.y0, box.width * 0.8, box.height])
# Put a legend to the right of the current axis
leg = ax.legend(loc='center left', bbox_to_anchor=(1, 0.5), fontsize=12)
leg.set_title('Method (GINI)',prop={'size':12})
ax.set_xlabel('Cumulative fraction of sequences')
ax.set_ylabel('Cumulative fraction\nof weights')
# plt.savefig('{}/gini_example.pdf'.format(figs_dir), bbox_inches='tight')
```
# Repeating for lots of proteins
```
methods_of_interest = ['uniform', 'simple_0.8', 'HH_meanScale', 'GSC_meanScale', 'ACL_meanScale']
weights_dict = {}
for method in methods_of_interest:
weights_dict[method] = []
for weight_file in sorted(glob.glob('../Data/weights/*.weights')):
prot_name = weight_file.split('/')[-1].split('_')[0]
method = weight_file.split('/')[-1].replace(prot_name+'_', '').replace('.weights', '')
if method in methods_of_interest:
weights = np.genfromtxt(weight_file)
weights_dict[method].append(weights)
gini_dict = {}
for method in methods_of_interest:
for i in weights_dict[method]:
arr = np.sort(i)
try:
gini_dict[method].append(get_gini(arr))
except:
gini_dict[method] = [get_gini(arr)]
print(methods_of_interest)
```
**Draw out some more complicated figures that look good**
```
boxes = []
for method in methods_of_interest:
boxes.append(gini_dict[method])
labels = ['Uniform', '0.8 sequence identity','HH mean scaled', 'GSC mean scaled', 'ACL mean scaled']
fig, ax = plt.subplots(figsize=(4,5))
bplot = ax.boxplot(boxes[::-1], labels=labels[::-1], widths=0.6, vert=False, patch_artist=True);
for patch in bplot['boxes']:
patch.set_facecolor('white')
patch.set_linewidth(2)
for median in bplot['medians']:
median.set_linewidth(2.5)
for whisker in bplot['whiskers']:
whisker.set_linewidth(2)
for cap in bplot['caps']:
cap.set_linewidth(2)
ax.set_xlabel('GINI coefficient', fontsize=16)
ax.tick_params(axis='x', labelsize=14)
ax.tick_params(axis='y', labelsize=16)
# plt.savefig('{}/gini_all.pdf'.format(figs_dir), bbox_inches='tight')
```
**Test the correlation between different methods**
```
methods_of_interest = ['uniform', 'simple_0.8', 'HH_meanScale',\
'GSC_meanScale', 'ACL_meanScale']
weights_dict = {}
for method in methods_of_interest:
weights_dict[method] = []
for weight_file in sorted(glob.glob('../Data/*.weights'))[:]:
prot_name = weight_file.split('/')[-1].split('_')[0]
method = weight_file.split('/')[-1].replace(prot_name+'_', '').replace('.weights', '')
if method in methods_of_interest:
weights = np.genfromtxt(weight_file)
weights_dict[method].append(weights)
matrix = []
for i, method_a in enumerate(methods_of_interest):
new_line = []
if method_a == 'uniform':
continue
print('##############################')
for method_b in methods_of_interest:
if method_b == 'uniform':
continue
print(method_a, method_b)
zipped = zip(weights_dict[method_a], weights_dict[method_b])
correlations = []
for weights_a,weights_b in zipped:
correlations.append(stats.spearmanr(weights_a,weights_b)[0])
print(np.median(correlations))
new_line.append(np.median(correlations))
matrix.append(new_line)
print(methods_of_interest)
```
**And plot it out**
```
fig, ax = plt.subplots(figsize=(3,3))
matrix = np.array(matrix)
cax = ax.matshow(matrix, vmin=0, vmax=1.0)
ax.grid(False)
labels = ['0.8 sequence identity', 'HH mean scaled', 'GSC mean scaled', 'ACL mean scaled']
ax.set_xticklabels(['']+labels, rotation=30, ha='left')
labels = ['0.8 sequence\n identity', 'HH mean\n scaled', 'GSC mean\n scaled', 'ACL mean\n scaled']
ax.set_yticklabels(['']+labels)
plt.colorbar(cax, label=r"Spearman's $\rho$ (median)", fraction=0.047, pad=0.10)
for y in range(matrix.shape[0]):
for x in range(matrix.shape[1]):
if x == y:
color='black'
tformat = '%.1f'
else:
color='white'
tformat = '%.3f'
plt.text(x , y, tformat % matrix[y, x],
horizontalalignment='center',
verticalalignment='center', color=color
)
# plt.savefig('{}/weights_corr.pdf'.format(figs_dir), bbox_inches='tight')
```
# Comparing regular and RelTime weights
```
methods_of_interest = ['GSC_meanScale', 'ACL_meanScale', 'GSC_meanScale.RelTime', 'ACL_meanScale.RelTime']
weights_dict = {}
for method in methods_of_interest:
weights_dict[method] = []
for weight_file in sorted(glob.glob('../Data/weights/*.weights')):
prot_name = weight_file.split('/')[-1].split('_')[0]
method = weight_file.split('/')[-1].replace(prot_name+'_', '').replace('.weights', '')
if method in methods_of_interest:
weights = np.genfromtxt(weight_file)
weights_dict[method].append(weights)
gini_dict = {}
for method in methods_of_interest:
for i in weights_dict[method]:
arr = np.sort(i)
try:
gini_dict[method].append(get_gini(arr))
except:
gini_dict[method] = [get_gini(arr)]
print(np.median(gini_dict['ACL_meanScale']), np.median(gini_dict['ACL_meanScale.RelTime']))
print(stats.wilcoxon(gini_dict['ACL_meanScale'], gini_dict['ACL_meanScale.RelTime']))
print(np.median(gini_dict['GSC_meanScale']), np.median(gini_dict['GSC_meanScale.RelTime']))
print(stats.wilcoxon(gini_dict['GSC_meanScale'], gini_dict['GSC_meanScale.RelTime']))
```
| github_jupyter |
# 1D Kalman Filter
Now, you're ready to implement a 1D Kalman Filter by putting all these steps together. Let's take the case of a robot that moves through the world. As a robot moves through the world it locates itself by performing a cycle of:
1. sensing and performing a measurement update and
2. moving and performing a motion update
You've programmed each of these steps individually, so now let's combine them in a cycle!
After implementing this filter, you should see that you can go from a very uncertain location Gaussian to a more and more certain Gaussian, as pictured below. The code in this notebooks is really just a simplified version of the Kalman filter that runs in the Google self-driving car that is used to track surrounding vehicles and other objects.
<img src='images/gaussian_updates.png' height=70% width=70% />
---
Below is our usual Gaussian equation and imports.
```
# import math functions
from math import *
import matplotlib.pyplot as plt
import numpy as np
# gaussian function
def f(mu, sigma2, x):
''' f takes in a mean and squared variance, and an input x
and returns the gaussian value.'''
coefficient = 1.0 / sqrt(2.0 * pi *sigma2)
exponential = exp(-0.5 * (x-mu) ** 2 / sigma2)
return coefficient * exponential
```
You've also been given the complete `update` code that performs a parameter update when an initial belief and new measurement information are merged. And the complete `predict` code that performs an update to a Gasuuain after a motion is incorporated.
```
# the update function
def update(mean1, var1, mean2, var2):
''' This function takes in two means and two squared variance terms,
and returns updated gaussian parameters.'''
# Calculate the new parameters
new_mean = (var2*mean1 + var1*mean2)/(var2+var1)
new_var = 1/(1/var2 + 1/var1)
return [new_mean, new_var]
# the motion update/predict function
def predict(mean1, var1, mean2, var2):
''' This function takes in two means and two squared variance terms,
and returns updated gaussian parameters, after motion.'''
# Calculate the new parameters
new_mean = mean1 + mean2
new_var = var1 + var2
return [new_mean, new_var]
```
### QUIZ: For the given measurements and motions, write complete 1D Kalman filter code that loops through all of these in order.
Your complete code should look at sensor measurements then motions in that sequence until all updates are done!
### Initial Uncertainty
You'll see that you are given initial parameters below, and this includes and nitial location estimation, `mu` and squared variance, `sig`. Note that the initial estimate is set to the location 0, and the variance is extremely large; this is a state of high confusion much like the *uniform* distribution we used in the histogram filter. There are also values given for the squared variance associated with the sensor measurements and the motion, since neither of those readings are perfect, either.
You should see that even though the initial estimate for location (the initial `mu`) is far from the first measurement, it should catch up fairly quickly as you cycle through measurements and motions.
```
# measurements for mu and motions, U
measurements = [5., 6., 7., 9., 10.]
motions = [1., 1., 2., 1., 1.]
# initial parameters
measurement_sig = 4.
motion_sig = 2.
mu = 0.
sig = 10000.
## TODO: Loop through all measurements/motions
## Print out and display the resulting Gaussian
# your code here
for i in range(len(measurements)):
mu, sig = update(mu,sig,measurements[i],measurement_sig)
print('update: ',mu,sig)
mu, sig = predict(mu,sig,motions[i],motion_sig)
print('predict: ',mu,sig)
```
### Plot a Gaussian
Plot a Gaussian by looping through a range of x values and creating a resulting list of Gaussian values, `g`, as shown below. You're encouraged to see what happens if you change the values of `mu` and `sigma2`.
```
# display the *initial* gaussian over a range of x values
# define the parameters
mu = 0
sigma2 = 10000
# define a range of x values
x_axis = np.arange(-20, 20, 0.1)
# create a corresponding list of gaussian values
g = []
for x in x_axis:
g.append(f(mu, sigma2, x))
# plot the result
plt.plot(x_axis, g)
```
| github_jupyter |
# Principal Component Analysis with Intel® Data Analytics Acceleration Library in Amazon SageMaker
## Introduction
Intel® Data Analytics Acceleration Library (Intel® DAAL) is the library of Intel® architecture optimized building blocks covering all stages of data analytics: data acquisition from a data source, preprocessing, transformation, data mining, modeling, validation, and decision making. One of its algorithms is PCA.
Principal Component Analysis (PCA) is a method for exploratory data analysis. PCA transforms a set of observations of possibly correlated variables to a new set of uncorrelated variables, called principal components. Principal components are the directions of the largest variance, that is, the directions where the data is mostly spread out.
Because all principal components are orthogonal to each other, there is no redundant information. This is a way of replacing a group of variables with a smaller set of new variables. PCA is one of powerful techniques for dimension reduction.
Intel® DAAL developer guide: https://software.intel.com/en-us/daal-programming-guide
Intel® DAAL documentation for PCA: https://software.intel.com/en-us/daal-programming-guide-principal-component-analysis
## PCA Usage with SageMaker Estimator
Firstly, you need to import SageMaker package, get execution role and create session.
```
import sagemaker
role = sagemaker.get_execution_role()
sess = sagemaker.Session()
```
Secondly, you can specify parameters of PCA.
#### Hyperparameters
All hyperparameters of PCA algorithm are optional.
<table style="border: 1px solid black;">
<tr>
<td><strong>Parameter name</strong></td>
<td><strong>Type</strong></td>
<td><strong>Default value</strong></td>
<td><strong>Description</strong></td>
</tr>
<tr>
<td>fptype</td>
<td>str</td>
<td>"double"</td>
<td>The floating-point type that the algorithm uses for intermediate computations. Can be "float" or "double"</td>
</tr>
<tr>
<td>method</td>
<td>str</td>
<td>"correlationDense"</td>
<td>Available methods for PCA computation: "correlationDense" ("defaultDense") or "svdDense"</td>
</tr>
<tr>
<td>resultsToCompute</td>
<td>str</td>
<td>"none"</td>
<td>Provide one of the following values to request a single characteristic or use bitwise OR to request a combination of the characteristics: mean, variance, eigenvalue. For example, combination of all is "mean|variance|eigenvalue"</td>
</tr>
<tr>
<td>nComponents</td>
<td>int</td>
<td>0</td>
<td>Number of principal components.<br/> If it is zero, the algorithm will compute the result for number of principal components = number of features.<br/>Remember that number of components must be equal or less than number of features for PCA algorithm</td>
</tr>
<tr>
<td>isDeterministic</td>
<td>bool</td>
<td>False</td>
<td>If True, the algorithm applies the "sign flip" technique to the results</td>
</tr>
<tr>
<td>transformOnTrain</td>
<td>bool</td>
<td>False</td>
<td>If True, training data will be transformed and saved in model package on training stage</td>
</tr>
</table>
Example of hyperparameters dictionary:
```
pca_params = {
"fptype": "float",
"method": "svdDense",
"resultsToCompute": "mean|eigenvalue",
"nComponents": 4,
"isDeterministic": True
}
```
Then, you need to create SageMaker Estimator instance with following parameters:
<table style="border: 1px solid black;">
<tr>
<td><strong>Parameter name</strong></td>
<td><strong>Description</strong></td>
</tr>
<tr>
<td>image_name</td>
<td>The container image to use for training</td>
</tr>
<tr>
<td>role</td>
<td>An AWS IAM role. The SageMaker training jobs and APIs that create SageMaker endpoints use this role to access training data and models</td>
</tr>
<tr>
<td>train_instance_count</td>
<td>Number of Amazon EC2 instances to use for training. Should be 1, because it is not distributed version of algorithm</td>
</tr>
<tr>
<td>train_instance_type</td>
<td>Type of EC2 instance to use for training. See available types on Amazon Marketplace page of algorithm</td>
</tr>
<tr>
<td>input_mode</td>
<td>The input mode that the algorithm supports. May be "File" or "Pipe"</td>
</tr>
<tr>
<td>output_path</td>
<td>S3 location for saving the trainig result (model artifacts and output files)</td>
</tr>
<tr>
<td>sagemaker_session</td>
<td>Session object which manages interactions with Amazon SageMaker APIs and any other AWS services needed</td>
</tr>
<tr>
<td>hyperparameters</td>
<td>Dictionary containing the hyperparameters to initialize this estimator with</td>
</tr>
</table>
Full SageMaker Estimator documentation: https://sagemaker.readthedocs.io/en/latest/estimators.html
```
daal_pca_arn = "<algorithm-arn>" # you can find it on algorithm page in your subscriptions
daal_pca = sagemaker.algorithm.AlgorithmEstimator(
algorithm_arn=daal_pca_arn,
role=role,
base_job_name="<base-job-name>",
train_instance_count=1,
train_instance_type='ml.m4.xlarge',
input_mode="File",
output_path="s3://<bucket-name>/<output-path>",
sagemaker_session=sess,
hyperparameters=pca_params
)
```
### Training stage
On training stage, PCA algorithm consume input data from S3 location and computes eigen vectors and other results (if they are specified in "resultsToCompute" parameter).
This container supports only .csv ("comma-separated values") files.
```
daal_pca.fit({"training": "s3://<bucket-name>/<training-data-path>"})
```
### Real-time prediction
On prediction stage, PCA algorithm transforms input data using previously computed results.
Firstly, you need to deploy SageMaker endpoint that consumes data.
```
predictor = daal_pca.deploy(1, "ml.m4.xlarge", serializer=sagemaker.predictor.csv_serializer)
```
Secondly, you should pass data as numpy array to predictor instance and get transformed data as space-separated values.
In this example we are passing random data, but you can use any numpy 2D array with one specific condition for PCA: training data and data to transform must have same numbers of features.
```
import numpy as np
predict_data = np.random.random(size=(10,10))
print(predictor.predict(predict_data).decode("utf-8"))
```
Don't forget to delete endpoint if you don't need it anymore.
```
sess.delete_endpoint(predictor.endpoint)
```
### Batch transform job
If you don't need real-time prediction, you can use transform job. It uses saved model, compute transformed data one time and saves it in specified or auto-generated output path.
More about transform jobs: https://docs.aws.amazon.com/sagemaker/latest/dg/how-it-works-batch.html
Transformer API: https://sagemaker.readthedocs.io/en/latest/transformer.html
```
transformer = daal_pca.transformer(1, 'ml.m4.xlarge')
transformer.transform("s3://<bucket-name>/<prediction-data-path>", content_type='text/csv')
transformer.wait()
print(transformer.output_path)
```
| github_jupyter |
# Тест. Доверительные интервалы для долей
```
import numpy as np
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
```
Большая часть млекопитающих неспособны во взрослом возрасте переваривать лактозу, содержащуюся в молоке. У людей за расщепление лактозы отвечает фермент лактаза, кодируемый геном LCT. У людей с вариантом 13910T этого гена лактаза продолжает функционировать на протяжении всей жизни. Распределение этого варианта гена сильно варьируется в различных генетических популяциях.
Из 50 исследованных представителей народа майя вариант 13910T был обнаружен у одного. Постройте нормальный 95% доверительный интервал для доли носителей варианта 13910T в популяции майя. Чему равна его нижняя граница? Округлите ответ до 4 знаков после десятичной точки.
```
size = 50
data_gen = np.zeros(size)
data_gen[0] = 1
data_gen
from statsmodels.stats.proportion import proportion_confint
normal_interval = proportion_confint(sum(data_gen), len(data_gen), method = 'normal')
print('Normal interval [%.4f, %.4f] with width %f' % (normal_interval[0],
normal_interval[1],
normal_interval[1] - normal_interval[0]))
```
В условиях предыдущей задачи постройте 95% доверительный интервал Уилсона для доли носителей варианта 13910T в популяции майя. Чему равна его нижняя граница? Округлите ответ до 4 знаков после десятичной точки.
```
wilson_interval = proportion_confint(sum(data_gen), len(data_gen), method = 'wilson')
print('Wilson interval [%.4f, %.4f] with width %f' % (wilson_interval[0],
wilson_interval[1],
wilson_interval[1] - wilson_interval[0]))
```
Пусть в популяции майя действительно 2% носителей варианта 13910T, как в выборке, которую мы исследовали. Какой объём выборки нужен, чтобы с помощью нормального интервала оценить долю носителей гена 13910T с точностью ±0.01 на уровне доверия 95%?
```
from statsmodels.stats.proportion import samplesize_confint_proportion
n_samples = int(np.ceil(samplesize_confint_proportion(data_gen.mean(), 0.01)))
n_samples
```
Постройте график зависимости объёма выборки, необходимой для оценки для доли носителей гена 13910T с точностью ±0.01 на уровне доверия 95%, от неизвестного параметра p. Посмотрите, при каком значении p нужно больше всего испытуемых. Как вы думаете, насколько вероятно, что выборка, которую мы анализируем, взята из случайной величины с этим значением параметра?
Как бы вы не ответили на последний вопрос, рассмотреть объём выборки, необходимый при таком p, всё равно полезно — это даёт максимально пессимистичную оценку необходимого объёма выборки.
Какой объём выборки нужен в худшем случае, чтобы с помощью нормального интервала оценить долю носителей гена 13910T с точностью ±0.01 на уровне доверия 95%?
```
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
proportion = np.linspace(0, 1, 101)
proportion
n_samples = np.empty(proportion.shape)
for i, p in enumerate(proportion):
n_samples[i] = int(np.ceil(samplesize_confint_proportion(p, 0.01)))
n_samples
plt.plot(proportion, n_samples);
n_samples[np.where(proportion == 0.5)]
```
| github_jupyter |
<a href="https://colab.research.google.com/github/parekhakhil/pyImageSearch/blob/main/1403_multi_template_matching.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>

# Multi-template matching with OpenCV
### by [PyImageSearch.com](http://www.pyimagesearch.com)
## Welcome to **[PyImageSearch Plus](http://pyimg.co/plus)** Jupyter Notebooks!
This notebook is associated with the [Multi-template matching with OpenCV](https://www.pyimagesearch.com/2021/03/29/multi-template-matching-with-opencv/) blog post published on 2021-03-29.
Only the code for the blog post is here. Most codeblocks have a 1:1 relationship with what you find in the blog post with two exceptions: (1) Python classes are not separate files as they are typically organized with PyImageSearch projects, and (2) Command Line Argument parsing is replaced with an `args` dictionary that you can manipulate as needed.
We recommend that you execute (press ▶️) the code block-by-block, as-is, before adjusting parameters and `args` inputs. Once you've verified that the code is working, you are welcome to hack with it and learn from manipulating inputs, settings, and parameters. For more information on using Jupyter and Colab, please refer to these resources:
* [Jupyter Notebook User Interface](https://jupyter-notebook.readthedocs.io/en/stable/notebook.html#notebook-user-interface)
* [Overview of Google Colaboratory Features](https://colab.research.google.com/notebooks/basic_features_overview.ipynb)
As a reminder, these PyImageSearch Plus Jupyter Notebooks are not for sharing; please refer to the **Copyright** directly below and **Code License Agreement** in the last cell of this notebook.
Happy hacking!
*Adrian*
<hr>
***Copyright:*** *The contents of this Jupyter Notebook, unless otherwise indicated, are Copyright 2021 Adrian Rosebrock, PyimageSearch.com. All rights reserved. Content like this is made possible by the time invested by the authors. If you received this Jupyter Notebook and did not purchase it, please consider making future content possible by joining PyImageSearch Plus at http://pyimg.co/plus/ today.*
### Download the code zip file
```
!wget https://pyimagesearch-code-downloads.s3-us-west-2.amazonaws.com/multi-template-matching/multi-template-matching.zip
!unzip -qq multi-template-matching.zip
%cd multi-template-matching
```
## Blog Post Code
### Import Packages
```
# import the necessary packages
from imutils.object_detection import non_max_suppression
from matplotlib import pyplot as plt
import numpy as np
import argparse
import cv2
```
### Function to display images in Jupyter Notebooks and Google Colab
```
def plt_imshow(title, image):
# convert the image frame BGR to RGB color space and display it
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
plt.imshow(image)
plt.title(title)
plt.grid(False)
plt.show()
```
### Implementing multi-template matching with OpenCV
```
# construct the argument parser and parse the arguments
#ap = argparse.ArgumentParser()
#ap.add_argument("-i", "--image", type=str, required=True,
# help="path to input image where we'll apply template matching")
#ap.add_argument("-t", "--template", type=str, required=True,
# help="path to template image")
#ap.add_argument("-b", "--threshold", type=float, default=0.8,
# help="threshold for multi-template matching")
#args = vars(ap.parse_args())
# since we are using Jupyter Notebooks we can replace our argument
# parsing code with *hard coded* arguments and values
args = {
"image": "images/8_diamonds.png",
"template": "images/diamonds_template.png",
"threshold": 0.8
}
# load the input image and template image from disk, then grab the
# template image spatial dimensions
print("[INFO] loading images...")
image = cv2.imread(args["image"])
template = cv2.imread(args["template"])
(tH, tW) = template.shape[:2]
# display the image and template to our screen
plt_imshow("Image", image)
plt_imshow("Template", template)
# convert both the image and template to grayscale
imageGray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
templateGray = cv2.cvtColor(template, cv2.COLOR_BGR2GRAY)
# perform template matching
print("[INFO] performing template matching...")
result = cv2.matchTemplate(imageGray, templateGray,
cv2.TM_CCOEFF_NORMED)
# find all locations in the result map where the matched value is
# greater than the threshold, then clone our original image so we
# can draw on it
(yCoords, xCoords) = np.where(result >= args["threshold"])
clone = image.copy()
print("[INFO] {} matched locations *before* NMS".format(len(yCoords)))
# loop over our starting (x, y)-coordinates
for (x, y) in zip(xCoords, yCoords):
# draw the bounding box on the image
cv2.rectangle(clone, (x, y), (x + tW, y + tH),
(255, 0, 0), 3)
# show our output image *before* applying non-maxima suppression
plt_imshow("Before NMS", clone)
# initialize our list of rectangles
rects = []
# loop over the starting (x, y)-coordinates again
for (x, y) in zip(xCoords, yCoords):
# update our list of rectangles
rects.append((x, y, x + tW, y + tH))
# apply non-maxima suppression to the rectangles
pick = non_max_suppression(np.array(rects))
print("[INFO] {} matched locations *after* NMS".format(len(pick)))
# loop over the final bounding boxes
for (startX, startY, endX, endY) in pick:
# draw the bounding box on the image
cv2.rectangle(image, (startX, startY), (endX, endY),
(255, 0, 0), 3)
# show the output image
plt_imshow("After NMS", image)
```
For a detailed walkthrough of the concepts and code, be sure to refer to the full tutorial, [*Multi-template matching with OpenCV*](https://www.pyimagesearch.com/2021/03/29/multi-template-matching-with-opencv/) published on 2021-03-29.
# Code License Agreement
```
Copyright (c) 2021 PyImageSearch.com
SIMPLE VERSION
Feel free to use this code for your own projects, whether they are
purely educational, for fun, or for profit. THE EXCEPTION BEING if
you are developing a course, book, or other educational product.
Under *NO CIRCUMSTANCE* may you use this code for your own paid
educational or self-promotional ventures without written consent
from Adrian Rosebrock and PyImageSearch.com.
LONGER, FORMAL VERSION
Permission is hereby granted, free of charge, to any person obtaining
a copy of this software and associated documentation files
(the "Software"), to deal in the Software without restriction,
including without limitation the rights to use, copy, modify, merge,
publish, distribute, sublicense, and/or sell copies of the Software,
and to permit persons to whom the Software is furnished to do so,
subject to the following conditions:
The above copyright notice and this permission notice shall be
included in all copies or substantial portions of the Software.
Notwithstanding the foregoing, you may not use, copy, modify, merge,
publish, distribute, sublicense, create a derivative work, and/or
sell copies of the Software in any work that is designed, intended,
or marketed for pedagogical or instructional purposes related to
programming, coding, application development, or information
technology. Permission for such use, copying, modification, and
merger, publication, distribution, sub-licensing, creation of
derivative works, or sale is expressly withheld.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND,
EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES
OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND
NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS
BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN
ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN
CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
SOFTWARE.
```
| github_jupyter |
This is a jupyter notebook to get the graphs from geo maps.
```
import random
import os
import numpy as np
import networkx as nx
import geopandas as gpd
import pandas as pd
import matplotlib
from matplotlib import pyplot as plt
from matplotlib.collections import PatchCollection
# Load the box module from shapely to create box objects
from shapely.geometry import box, Point, Polygon, MultiPoint, shape, LineString
from shapely.ops import triangulate
from shapely.affinity import translate
from earthpy import clip as cl
import pickle #to save the resulting graphs
# to display images inline
get_ipython().magic(u'matplotlib inline')
matplotlib.use('Agg')# not sure what I used it for
```
# RELATIVE NEIGHBORHOOD GRAPH
This notebook demonstrated how RNG graphs can be calculated from OSM data in the form of shapefiles.
Recent OSM data can be downloded from the project website, here we use a French region called Lorraine.
#### Pre-processing:
We processed data in QGIS to merge the buildings which touch, and to extract certain separate categories from the buildings.
#### Categories
A pre-selected set of objects is defined. The nodes of our graphs are formed by these objects,each category is cooded as a unique integer. Some of categories are missing from OSM data, but we keep the codes to be consistent.
aerodroms: 0
houses: 1
public build: 2
railway stat: 3
sport build: 4
arcs, towers: 5
churches: 6
castles: 7
forts: 8
monuments: 9
cemetries: 10
sport build: 11
roads: 12
water: 13
railroads: 14
```
# define the region folder - folder where OSM data are unpacked
global_path = "./lorraine"
```
Load all shapefiles as geopandas dataframes. We also convert the data to Lambert 93 on the way.
## ROADS
```
# load all the shapely files related to ROADS
fp_road = global_path + "/gis_osm_roads_free_1.shp"
all_roads = gpd.read_file(fp_road)
all_roads = all_roads.to_crs("EPSG:2154")
all_roads.head() # small demo of the roads and the info we have
```
Data type - geographical projection of he data used.
## Houses
```
# Load all the data from the BUILDINGS caegory
fp_bati = global_path + "/buildings_merged.shp"
# Read file using gpd.read_file()
all_buildings = gpd.read_file(fp_bati)
all_buildings = all_buildings.to_crs("EPSG:2154")
```
## Churches
```
#places of worship
churches = global_path + "/gis_osm_pofw_free_1.shp"
churches = gpd.read_file(churches)
churches=churches.to_crs("EPSG:2154")
```
## Towers, castles, forts, monuments
```
fp_poi = global_path + "/gis_osm_pois_free_1.shp"
# Read file using gpd.read_file()
poi = gpd.read_file(fp_poi)
poi=poi.to_crs("EPSG:2154")
poi.fclass.unique()
towers = poi.loc[poi['fclass'] == 'tower']
castels = poi.loc[poi['fclass'] == 'castle']
forts = poi.loc[poi['fclass']=='fort']
monuments = poi.loc[poi['fclass']=='monument']
print("statisics over all POI objects. There %d churches, %d towers, %d monuments, %d forts, %d castles and %d ordinary buildings"
%(len(churches),len(towers),len(monuments), len(forts), len(castels), len(all_buildings)))
```
## WATER
```
fp_water = global_path + "/gis_osm_waterways_free_1.shp"
all_water = gpd.read_file(fp_water)
all_water = all_water.to_crs("EPSG:2154")
```
## SPORT TERRITORIES
```
fp_sport = global_path + "/gis_osm_pois_free_1.shp"
data_sport = gpd.read_file(fp_sport)
data_sport=data_sport.to_crs("EPSG:2154")
data_sport = data_sport.loc[data_sport['fclass'] == 'stadium']
```
## CEMETRIES
```
fp_cemetries = global_path + "/gis_osm_pois_a_free_1.shp"
data_cemetries = gpd.read_file(fp_cemetries)
data_cemetries=data_cemetries.to_crs("EPSG:2154")
#pick the graveyards
data_cemetries = data_cemetries.loc[data_cemetries['fclass'] =='graveyard']
```
## RAILROADS
```
#railroads
fp_rail = global_path + "/gis_osm_railways_free_1.shp"
data_rail = gpd.read_file(fp_rail)
data_rail = data_rail.to_crs("EPSG:2154")
print("statisics over all line objects. There %d road segments, %d water segements, %d railroad segments"
%(len(all_roads),len(all_water),len(data_rail)))
```
## BOUNDING BOXES FROM THE IMAGES
We use the pre-save bounding boxes, just change the paths to the right one. The bounding box just limits the zone to create different graphs.
```
image_polygons = gpd.read_file('image_polygons_57.shp') # image polygons for POI in Moselle region
image_polygons
```
## graph creation routine
Below are the functions defined for each step of RNG graph creation procedure.
```
# parameters of the nodes
def calculate_eccentricity(geometry):
''' function calculates eccentricity measure (w/l of min envelop rectangle) '''
envelop_rect = geometry.minimum_rotated_rectangle
minx, miny, maxx, maxy = envelop_rect.bounds
width = maxx - minx
height = maxy - miny
if height ==0 or width ==0:
return 0
if width < height:
return width/height
else:
return height/width
def calculate_perimeter(geometry):
''' function calculates the perimeter of the shapely geometry '''
return geometry.length
# unit tests
# calculate_perimeter(buildings3.geometry[1])
def get_node_attributes(shapely_geometry, poly_bound, nature, within_poly = True):
""" function returns attributes of road/house node:
nature, normed_lenght and eccentricity """
attributes = {}
obj_type = nature
if within_poly:
obj_length = calculate_perimeter(shapely_geometry)
frame_perimeter = poly_bound.length
obj_normed_length = obj_length/frame_perimeter
obj_points = calculate_eccentricity(shapely_geometry)
else:
obj_normed_length = 0
obj_points = 0
attributes = {'nature': obj_type, 'normed_length':obj_normed_length,
'eccentricity':obj_points}
return attributes
def calculate_distance(obj1, obj2):
''' this fucntion calculates the minimum distance between the objects:
if an object is a polygon, its center is considered as a central point
if an object is a line, the distance is the lenght of the normal from another object,
if both objects are lines, the distance is 0'''
d = None
if obj1.type == 'Polygon' and obj2.type == 'Polygon':
#print("polyg polyg")
proj_dist = obj1.centroid.distance(obj2.centroid) #euclidean distance
d = proj_dist
elif obj1.type == 'LineString' and obj2.type == 'Polygon':
#print("linestr polyg")
proj_dist= obj2.centroid.distance(obj1)
# print(proj_dist)
# print(obj1.project(obj2.centroid))
d = proj_dist
elif obj2.type=='LineString' and obj1.type == 'Polygon':
#print(" polyg linestr")
proj_dist= obj1.centroid.distance(obj2)
d = proj_dist
elif obj1.type == 'LineString' and obj2.type=='LineString':
#print(" linestr linestr")
if obj1.intersects( obj2):
d = 0
else:
d = None
elif obj1.type == 'shapely.geometry.LineString' and obj2.type=='shapely.geometry.LineString':
#print(" linestr linestr")
if obj1.intersects( obj2):
d = 0
else:
d = None
else:
print(" unkn unkn")
try:
print(type(obj1),type(obj2))
d = obj2.distance(obj1.centroid)
except:
print('Distance calc didnt work')
d = None
#print(f"d is {d} for objects {type(obj1)} {type(obj2)}")
return d
def determine_if_connected_rng(geodf, idx1, idx2, distance_matrix):
''' check if the nodes are connected in the graph (the algorithm specified in
here https://en.wikipedia.org/wiki/Relative_neighborhood_graph is used, it is not optimized
for big graphs (polynomial time version is implemented))'''
connected = False
dist = distance_matrix[idx1][idx2]
dmax =[]
N = distance_matrix.shape[0]
if N>2:
for i in range(N):
if i!=idx1 and i!=idx2:
dmax.append(max(distance_matrix[idx1][i], distance_matrix[idx2][i]))
connected = dist <= min(dmax)
else:
connected = True
return connected,dist
def calculate_distance_matrix(geodf):
''' returns distance matrix between all the nodes'''
dist_matrix = np.zeros((len(geodf),len(geodf)))
for shp1 in range(0, len(geodf)):
for shp2 in range(0, len(geodf)):
dist_matrix[shp1][shp2] = geodf.centroid[shp1].distance(geodf.centroid[shp2])
np.fill_diagonal(dist_matrix, float('inf'))
return dist_matrix
def calculate_centroids(geodf):
''' calculates geometrical centers of geometries'''
geodf = geodf.assign(centroid="")
for i in range(len(geodf)):
if type(geodf.loc[i, ('geometry')]) == 'Point':
geodf.loc[i, ('centroid')] = geodf.loc[i, ('geometry')]
else:
geodf.centroid[i] = geodf.loc[i, ('geometry')].centroid
return geodf
def create_rng_graph(gp_frame, poly = None):
''' function takes the pandas frame and creates the graph, where the nodes are rivers and roads,
and buildings, and sportive objects etc.
Nodes have the following attributes:
a) type of object (road/water/house/church etc)
b) eccentricity
c) length (divided by HxW of the polygon to scale)
Edges have attributes based on the normalized distance between the points:
they are calculated by the algorithm proposed
here:
https://en.wikipedia.org/wiki/Relative_neighborhood_graph
'''
pos = {} #dictionary for node coord
net = nx.Graph() # empty graph
attr = {}
dist_matrix = calculate_distance_matrix(gp_frame)
for shp1 in range(0, len(gp_frame)-1): # for each object
# the geometry property here may be specific to my shapefile
object1 = gp_frame['geometry'].iloc[shp1] #get the line
pos[shp1] = [gp_frame['geometry'].iloc[shp1].centroid.x, gp_frame['geometry'].iloc[shp1].centroid.y]
# get all line attributes
attributes = get_node_attributes(object1, poly, gp_frame['nature'].iloc[shp1],gp_frame['withinPoly'].iloc[shp1])
net.add_node(shp1) # add node
attr[shp1]= attributes # nested dict
for shp2 in range(shp1+1, len(gp_frame)):
object2 = gp_frame['geometry'].iloc[shp2] #get the second object
connected, dist = determine_if_connected_rng(gp_frame, shp1, shp2, dist_matrix)
if connected: # if intersects
net.add_edge(shp1, shp2, distance = dist) # edge with an attribute
# add the last element - coz first loop is not for all values, and last node needs attributes
attributes = get_node_attributes(object2, poly, gp_frame['nature'].iloc[shp2],gp_frame['withinPoly'].iloc[shp2])
attr[len(gp_frame)-1]= attributes # nested dict
net.add_node(shp2) # add node
nx.set_node_attributes(net, attr)
pos[shp2] = [gp_frame['geometry'].iloc[shp2].centroid.x, gp_frame['geometry'].iloc[shp2].centroid.y]
return net, pos
from shapely.ops import split
def divide_road_into_segments(final_data, nature, geoobject, inside_polygon =1, max_line_seg_len = 10 ):
''' this function divides the road into a set of segments '''
number_of_segments = int(geoobject.length// max_line_seg_len)
if number_of_segments>1:
for i in range(1, number_of_segments):
#cutting_point=geoobject.interpolate(i/number_of_segments)
#dist= geoobject.project(cutting_point)
segment = cut(geoobject, max_line_seg_len)
segment, geoobject = segment[0], segment[1]
final_data.append([nature, 1, segment])
else:
final_data.append([nature, 1, geoobject])
return final_data
def cut(line, distance):
# Cuts a line in two at a distance from its starting point
if distance <= 0.0 or distance >= line.length:
return [LineString(line)]
coords = list(line.coords)
for i, p in enumerate(coords):
pd = line.project(Point(p))
if pd == distance:
return [
LineString(coords[:i+1]),
LineString(coords[i:])]
if pd > distance:
cp = line.interpolate(distance)
return [
LineString(coords[:i] + [(cp.x, cp.y)]),
LineString([(cp.x, cp.y)] + coords[i:])]
## version with splitting!!!!!
def clean_and_append(final_data, data_segment, nature, polygon_bbox, max_line_seg_len):
''' function copies data from a data frame to a new data list of a following structure:
nature
within the image (bool) if the object is entirely inside the polygon or not
geometry
returns list of lists with objects like [[nat, within, geom], [], []...] '''
# if it is line object
if data_segment.empty == True:
pass
else:
# check if object is completely within the polygon
for index, row in data_segment.iterrows(): # for each element
if row['geometry'].geom_type == 'MultiPolygon' or row['geometry'].geom_type == 'MultiLineString':
for single_obj in row['geometry']:
if nature in [12,13,14]:
final_data = divide_road_into_segments(final_data, nature, single_obj, 1, 15)
else:
final_data.append([nature, 1, single_obj])
print('a multistring detected and transformed')
elif row['geometry'].is_empty or row['geometry']==[]:
pass
else:
if nature in [12,13,14]:
final_data = divide_road_into_segments(final_data, nature, row['geometry'], 1, max_line_seg_len)
else:
final_data.append([nature, 1, row['geometry']]) # polygons are classified depending on their status always within
return final_data
def clip_data(pd_obj, pd_polyg):
try:
sg = cl.clip_shp(pd_obj, pd_polyg) #all_roads[all_roads.geometry.intersects(polygon_bbox)] #extract segments of roads
except:
sg = pd.DataFrame()
return sg
#some variables to define
max_line_seg_len = 10 # if the roads should be cut, define this one, if not -> put a big value
name = './range_graphs/' #folder name where the graphs will be save as pickle files
for i in range(1,2) : # len(image_polygons)
# randomly shift the polygon box
polygon_bbox = image_polygons.iloc[i-1:i]
data = []
# lines
sg_roads = clip_data(all_roads, polygon_bbox) #extract segments of roads
data = clean_and_append(data, sg_roads, 12, polygon_bbox, max_line_seg_len)
sg_water = clip_data(all_water, polygon_bbox) #extract segments of water
data = clean_and_append(data, sg_water, 13, polygon_bbox, max_line_seg_len)
sg_data_rail = clip_data(data_rail, polygon_bbox)
data = clean_and_append(data, sg_data_rail, 14, polygon_bbox,max_line_seg_len)
# objects
sg_houses = clip_data(all_buildings, polygon_bbox) #extract segments of buildings
data = clean_and_append(data, sg_houses, 1, polygon_bbox,max_line_seg_len)
sg_towers = clip_data(towers, polygon_bbox)#towers[towers.intersects(polygon_bbox)]
data = clean_and_append(data, sg_towers, 5, polygon_bbox, max_line_seg_len)
sg_churches = clip_data(churches, polygon_bbox) # churches[churches.intersects(polygon_bbox)]
data = clean_and_append(data, sg_churches, 6, polygon_bbox, max_line_seg_len)
sg_castels = clip_data(castels, polygon_bbox)#castels[castels.intersects(polygon_bbox)]
data = clean_and_append(data, sg_castels, 7, polygon_bbox, max_line_seg_len)
sg_forts = clip_data(forts, polygon_bbox)#forts[forts.intersects(polygon_bbox)]
data = clean_and_append(data, sg_forts, 8, polygon_bbox,max_line_seg_len)
sg_monuments = clip_data(monuments, polygon_bbox)#monuments[monuments.intersects(polygon_bbox)]
data = clean_and_append(data, sg_monuments, 9, polygon_bbox,max_line_seg_len)
sg_data_cemetries = clip_data(data_cemetries, polygon_bbox)#data_cemetries[data_cemetries.geometry.intersects(polygon_bbox)]
data = clean_and_append(data, sg_data_cemetries, 10, polygon_bbox,max_line_seg_len)
sg_sport = clip_data(data_sport, polygon_bbox)#data_sport[data_sport.geometry.intersects(polygon_bbox)] #extract segments of sport things
data = clean_and_append(data, sg_sport, 11, polygon_bbox,max_line_seg_len)
combined_pd = pd.DataFrame(data, columns = ['nature','withinPoly', 'geometry'])
sg_houses_rem = pd.concat([sg_sport, sg_towers,sg_churches, sg_data_cemetries, sg_castels,sg_forts,sg_monuments],ignore_index=True)
combined_pd = calculate_centroids(combined_pd)
print(name +'graph_main'+ str(i).zfill(5) +'.pickle')
print(len(combined_pd))
if combined_pd.empty:
G = nx.empty_graph()
else:
G, pos = create_rng_graph(gp_frame=combined_pd, poly=polygon_bbox)
nx.write_gpickle(G, name + str(i).zfill(5) +'_graph' +'.pickle', protocol=4)
with open(name + str(i).zfill(5) +'_position.pickle', 'wb') as handle:
pickle.dump(pos, handle)
```
## Resulting graphs demo
```
combined_pd # all objetcs in a graph (nodes)
sg_roads = sg_roads [~sg_roads.is_empty] #sometimes, we get empty geometries which should be cleaned
sg_houses = sg_houses [~sg_houses.is_empty]
sg_water = sg_water [~sg_water.is_empty]
fig, ax = plt.subplots(figsize=(8, 8))
for p in combined_pd.centroid.tolist():
ax.scatter(p.x,p.y)
nx.draw_networkx_edges(G=G, pos=pos,ax=ax)
sg_roads.plot(linewidth=10.0, edgecolor='#FFA500', color='#FFA500', alpha = 0.3, ax=ax)
sg_houses.plot(color='#FF0000', alpha = 0.5, ax=ax)
#sg_houses_rem.plot(color='#FFFF00', ax=ax)
sg_water.plot(linewidth=10.0, color='#0000FF', alpha = 0.5, ax=ax)
#sg_data_rail.plot(linewidth=10.0, color='#FF00FF',alpha = 0.5, ax=ax)
nx.draw_networkx(G, pos =pos)
```
That is it! The graphs will be saved along with their positions in pickle files in a specified folder.
| github_jupyter |
```
%matplotlib inline
```
# Post pruning decision trees with cost complexity pruning
.. currentmodule:: sklearn.tree
The :class:`DecisionTreeClassifier` provides parameters such as
``min_samples_leaf`` and ``max_depth`` to prevent a tree from overfiting. Cost
complexity pruning provides another option to control the size of a tree. In
:class:`DecisionTreeClassifier`, this pruning technique is parameterized by the
cost complexity parameter, ``ccp_alpha``. Greater values of ``ccp_alpha``
increase the number of nodes pruned. Here we only show the effect of
``ccp_alpha`` on regularizing the trees and how to choose a ``ccp_alpha``
based on validation scores.
See also `minimal_cost_complexity_pruning` for details on pruning.
```
print(__doc__)
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_breast_cancer
from sklearn.tree import DecisionTreeClassifier
```
Total impurity of leaves vs effective alphas of pruned tree
---------------------------------------------------------------
Minimal cost complexity pruning recursively finds the node with the "weakest
link". The weakest link is characterized by an effective alpha, where the
nodes with the smallest effective alpha are pruned first. To get an idea of
what values of ``ccp_alpha`` could be appropriate, scikit-learn provides
:func:`DecisionTreeClassifier.cost_complexity_pruning_path` that returns the
effective alphas and the corresponding total leaf impurities at each step of
the pruning process. As alpha increases, more of the tree is pruned, which
increases the total impurity of its leaves.
```
X, y = load_breast_cancer(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
clf = DecisionTreeClassifier(random_state=0)
path = clf.cost_complexity_pruning_path(X_train, y_train)
ccp_alphas, impurities = path.ccp_alphas, path.impurities
```
In the following plot, the maximum effective alpha value is removed, because
it is the trivial tree with only one node.
```
fig, ax = plt.subplots()
ax.plot(ccp_alphas[:-1], impurities[:-1], marker='o', drawstyle="steps-post")
ax.set_xlabel("effective alpha")
ax.set_ylabel("total impurity of leaves")
ax.set_title("Total Impurity vs effective alpha for training set")
```
Next, we train a decision tree using the effective alphas. The last value
in ``ccp_alphas`` is the alpha value that prunes the whole tree,
leaving the tree, ``clfs[-1]``, with one node.
```
clfs = []
for ccp_alpha in ccp_alphas:
clf = DecisionTreeClassifier(random_state=0, ccp_alpha=ccp_alpha)
clf.fit(X_train, y_train)
clfs.append(clf)
print("Number of nodes in the last tree is: {} with ccp_alpha: {}".format(
clfs[-1].tree_.node_count, ccp_alphas[-1]))
```
For the remainder of this example, we remove the last element in
``clfs`` and ``ccp_alphas``, because it is the trivial tree with only one
node. Here we show that the number of nodes and tree depth decreases as alpha
increases.
```
clfs = clfs[:-1]
ccp_alphas = ccp_alphas[:-1]
node_counts = [clf.tree_.node_count for clf in clfs]
depth = [clf.tree_.max_depth for clf in clfs]
fig, ax = plt.subplots(2, 1)
ax[0].plot(ccp_alphas, node_counts, marker='o', drawstyle="steps-post")
ax[0].set_xlabel("alpha")
ax[0].set_ylabel("number of nodes")
ax[0].set_title("Number of nodes vs alpha")
ax[1].plot(ccp_alphas, depth, marker='o', drawstyle="steps-post")
ax[1].set_xlabel("alpha")
ax[1].set_ylabel("depth of tree")
ax[1].set_title("Depth vs alpha")
fig.tight_layout()
```
Accuracy vs alpha for training and testing sets
----------------------------------------------------
When ``ccp_alpha`` is set to zero and keeping the other default parameters
of :class:`DecisionTreeClassifier`, the tree overfits, leading to
a 100% training accuracy and 88% testing accuracy. As alpha increases, more
of the tree is pruned, thus creating a decision tree that generalizes better.
In this example, setting ``ccp_alpha=0.015`` maximizes the testing accuracy.
```
train_scores = [clf.score(X_train, y_train) for clf in clfs]
test_scores = [clf.score(X_test, y_test) for clf in clfs]
fig, ax = plt.subplots()
ax.set_xlabel("alpha")
ax.set_ylabel("accuracy")
ax.set_title("Accuracy vs alpha for training and testing sets")
ax.plot(ccp_alphas, train_scores, marker='o', label="train",
drawstyle="steps-post")
ax.plot(ccp_alphas, test_scores, marker='o', label="test",
drawstyle="steps-post")
ax.legend()
plt.show()
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.