text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Plotly's Python API User Guide
## Section 7: Plotly's Streaming API
Welcome to Plotly's Python API User Guide.
> Links to the other sections are on the User Guide's [homepage](https://plotly.com/python/userguide)
Section 7 is divided, into separate notebooks, as follows:
* [7.0 Streaming API introduction](https://plotly.com/python/intro_streaming)
* [7.1 A first Plotly streaming plot](https://plotly.com/python/streaming_part1)
* [7.2
Quickstart (initialize Plotly figure object and send 1 data point through a stream):
>>> import plotly.plotly as py
>>> from plotly.graph_objs import *
>>> # auto sign-in with credentials or use py.sign_in()
>>> trace1 = Scatter(
x=[],
y=[],
stream=dict(token='my_stream_id')
)
>>> data = Data([trace1])
>>> py.plot(data)
>>> s = py.Stream('my_stream_id')
>>> s.open()
>>> s.write(dict(x=1, y=2))
>>> s.close()
<hr>
Check which version is installed on your machine and please upgrade if needed.
```
# (*) Import plotly package
import plotly
# Check plolty version (if not latest, please upgrade)
plotly.__version__
```
<hr>
Plotly's Streaming API enables your Plotly plots to update in real-time, without refreshing your browser. In other words, users can *continuously* send data to Plotly's servers and visualize this data in *real-time*.
For example, imagine that you have a thermometer (hooked to an Arduino for example) in your attic and you would like to monitor the temperature readings from your laptop. Plotly together with its streaming API makes this project easy to achieve.
With Ploly's Streaming API:
> Everyone looking at a Plotly streaming plot sees the same data updating at the same time. Like all Plotly plots, Plotly streaming plots are immediately shareable by shortlink, embedded in website, or in an IPython notebook. Owners of the Plotly plot can edit the plot with the Plotly web GUI and all viewers will see these changes live.
And for the skeptical among us, *it's fast*: plots update up to 20 times per second.
In this section, we present examples of how to make Plotly streaming plots.
Readers looking for info about the nuts and bolts of Plotly's streaming API should refer to <a href="https://plotly.com/streaming/" target="_blank">this link</a>.
So, we first import a few modules and sign in to Plotly using our credentials file:
```
# (*) To communicate with Plotly's server, sign in with credentials file
import plotly.plotly as py
# (*) Useful Python/Plotly tools
import plotly.tools as tls
# (*) Graph objects
from plotly.graph_objs import *
import numpy as np # (*) numpy for math functions and arrays
```
##### What do Plotly streaming plots look like?
```
# Embed an existing Plotly streaming plot
tls.embed('streaming-demos','6')
# Note that the time point correspond to internal clock of the servers,
# that is UTC time.
```
Data is sent in real-time. <br>
Plotly draws the data in real-time. <br>
Plotly's interactibility happens in real-time.
##### Get your stream tokens
Making Plotly streaming plots requires no modifications to the sign in process; however, users must generate stream *tokens* or *ids*. To do so, first sign-in to <a href="https://plotly.com" target="_blank">plot.ly</a>. Once that is done, click on the *Settings* button in the upper-right corner of the page:
<img src="http://i.imgur.com/RNExysW.png" style="margin-top:1em; margin-bottom:1em" />
<p style="margin-top:1.5em;,margin-bottom:1.5em">Under the <b>Stream Tokens</b> tab, click on the <b>Generate Token</b> button:</p>
<img src="http://i.imgur.com/o5Uguye.png" />
And there you go, you have generated a stream token. Please note that:
> You must generate one stream token per **trace** in your Plotly streaming plot.
If you are looking to run this notebook with you own account, please generate 4 unique stream tokens and add them to your credentials file by entering:
>>> tls.set_credentials_file(stream_ids=[
"ab4kf5nfdn",
"kdf5bn4dbn",
"o72o2p08y5",
"81dygs4lcy"
])
where the `stream_ids` keyword argument is filled in with your own stream ids.
Note that, in the above, `tls.set_credentials()` overwrites the existing stream tokens (if any) but does not clear the other keys in your credentials file such as `username` and `api_key`.
Once your credentials file is updated with your stream tokens (or stream ids, a synonym), retrieve them as a list:
```
stream_ids = tls.get_credentials_file()['stream_ids']
```
We are now ready to start making Plotly streaming plots!
The content of this section has been divided into separate IPython notebooks as loading multiple streaming plots at once can cause performance slow downs on some internet connections.
Here are the links to the subsections' notebooks:
* [7.0 Streaming API introduction](https://plotly.com/python/intro_streaming)
* [7.1 A first Plotly streaming plot](https://plotly.com/python/streaming_part1)
In addition, here is a notebook of another Plotly streaming plot:
* <a href="http://nbviewer.ipython.org/gist/empet/a03885a54c256a21c514" target="_blank">Streaming the Poisson Pareto Burst Process</a> by
<a href="https://github.com/empet" target="_blank">Emilia Petrisor</a>
<div style="float:right; \">
<img src="http://i.imgur.com/4vwuxdJ.png"
align=right style="float:right; margin-left: 5px; margin-top: -10px" />
</div>
<h4>Got Questions or Feedback? </h4>
Reach us here at: <a href="https://community.plot.ly" target="_blank">Plotly Community</a>
<h4> What's going on at Plotly? </h4>
Check out our twitter:
<a href="https://twitter.com/plotlygraphs" target="_blank">@plotlygraphs</a>
```
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install publisher --upgrade
import publisher
publisher.publish(
's7_streaming.ipynb', 'python/intro_streaming//', 'Getting Started with Plotly Streaming',
'Getting Started with Plotly Streaming',
title = 'Getting Started with Plotly Streaming',
thumbnail='', language='python',
layout='user-guide', has_thumbnail='false')
```
| github_jupyter |
> Texto fornecido sob a Creative Commons Attribution license, CC-BY. Todo o código está disponível sob a FSF-approved BSD-3 license.<br>
> (c) Original por Lorena A. Barba, Gilbert F. Forsyth em 2017, traduzido por Felipe N. Schuch em 2020.<br>
> [@LorenaABarba](https://twitter.com/LorenaABarba) - [@fschuch](https://twitter.com/fschuch)
12 passos para Navier–Stokes
======
***
Olá! Bem-vindo aos **12 passos para Navier-Stokes**. Esse é um módulo prático usado inicialmente como um curso interativo de Dinâmica dos Fluidos Computacional (CFD, do Inglês *Computational Fluid Dynamics*), ministrado pela [Prof. Lorena Barba](http://lorenabarba.com) desde a primavera de 2009 na Universidade de Boston. O curso assume que o leitor tenha conhecimentos básicos sobre programação (em qualquer linguagem) e alguma familiaridade com equações diferenciais e mecânica dos fluidos. Os "passos" foram inspirados pelas ideias do Dr. Rio Yokota, que era um pós-doc no laboratório da Prof. Barba. As lições foram refinadas pela Professora e seus estudantes ao longo de vários semestres. O curso é ensinado inteiramente em Python, e os alunos que não conhecem Python tem a oportunidade de aprender conforme progredimos pelas lições. Adaptado e traduzido para português por [Felipe N. Schuch](https://fschuch.github.io/).
Esse [Jupyter notebook](https://jupyter-notebook.readthedocs.io/en/stable/) vai lhe guiar através do primeiro passo para programar sua própria ferramenta de resolução de Navier-Stokes em Python. Colocamos a mão na massa agora mesmo. Não se preocupe se inicialmente não entender tudo que está acontecendo, cobriremos cada detalhe à medida que avançamos, além disso, material complementar para o aprendizado está disponível em vídeo (em inglês) em [Prof. Barba's lectures on YouTube](http://www.youtube.com/playlist?list=PL30F4C5ABCE62CB61).
Para obter melhores resultados, uma vez concluido esse notebook, escreva seu próprio código para o Passo 1, seja em um script Python ou em um novo Jupyter notebook.
Passo 1: Convecção Linear Unidimensional
-----
***
O equação de Convecção Linear 1D é a versão mais simplificada, o modelo mais básico que se pode empregar para aprender algo sobre CFD. É supreendente o quanto essa pequena equação é capaz de nos ensinar! Aqui está ela:
$$\frac{\partial u(x,t)}{\partial t} + c \frac{\partial u(x,t)}{\partial x} = 0$$
Para uma dada condição inicial (denominada como *onda*), a equação descreve a propagação dessa *onda* inicial com uma velocidade $c$, sem mudar seu formato. Considerando a condição inicial como $u(x,0)=u_0(x)$, então a solução exata da equação é dada por $u(x,t)=u_0(x-ct)$.
Usamos o métodos das diferenças finitas para discretizar a equação no tempo e no espaço, usando um esquema de diferença para frente para a derivada temporal e um esquema de diferença para trás para a derivada espacial. Assim, se discretiza a coordenada espacial $x$ em pontos com índice $i$, sendo $0 \le i \le N$, e discretizamos o tempo em $n$ passos discretos de tamanho $\Delta t$.
À partir da definição de uma derivada (e simplesmente removendo o limite), sabemos que:
$$\frac{\partial u}{\partial x}\approx \frac{u(x+\Delta x)-u(x)}{\Delta x}$$
Nossa equação discreta é, então, dada por:
$$\frac{u_i^{n+1}-u_i^n}{\Delta t} + c \frac{u_i^n - u_{i-1}^n}{\Delta x} = 0 $$
Onde $n$ e $n+1$ são dois passos consecutivos no tempo, enquanto $i-1$ e $i$ são dois pontos vizinhos na discretização da coordenada $x$. Se a condição inicial é conhecida, então a única incógnita na discretização é $u_i^{n+1}$. Podemos isolar nossa incógnita para obter uma equação que nos permita avançar no tempo, que é escrita como:
$$u_i^{n+1} = u_i^n - c \frac{\Delta t}{\Delta x}(u_i^n-u_{i-1}^n)$$
Agora, é hora de implementá-la em Python.
Vamos começar importando algumas bibliotecas que nos serão úteis:
* `numpy` é uma biblioteca que fornece uma série de ferramentas para manipulação matricial, similar ao MATLAB;
* `matplotlib` é uma biblioteca para produção gráfica 2D, usaremos para mostrar os resultados;
* `time` e `sys` proporcionam utilidades básicas de tempo e sistema, que usaremos para reduzir a velocidade da visualização de animações.
```
#Lembrete: comentários em Python são indicados pelo jogo da velha
import numpy #Aqui carregamos numpy
from matplotlib import pyplot #Aqui carregamos matplotlib
import time, sys #E carregamos algumas utilidades
#isso faz os gráficos matplotlib aparecerem no notebook (em vez de em uma nova janela)
%matplotlib inline
```
Agora vamos definir alguns parâmetros. Queremos criar um domínio espacial com pontos igualmente espaçados e duas unidades de comprimento, isso é, $x_i\in(0,2)$. Para tanto, a função [numpy.linspace](https://docs.scipy.org/doc/numpy/reference/generated/numpy.linspace.html) é exatamente o que precisamos.
```
#Tente mudar esse número de 41 para 81
#então execute tudo (Run All).
#O que acontece?
x = numpy.linspace(0., 2., num = 41)
nt = 25 #Número de passos de tempo que queremos calcular
dt = .025 #Tamanho de cada passo de tempo
c = 1 #Velocidade de propagação da onda
```
É sempre possível acessar a documentação de uma dada função se você quiser saber mais detalhes. Tente executar uma célula com o comando:
```python
help(numpy.linspace)
```
À partir de `x`, extraímos o parâmetro `nx` como o número de pontos na malha e `dx` será o espaçamento entre eles.
```
nx = x.size
dx = x[1] - x[0]
```
Também precisamos definir a condição inicial (CI). Digamos que a velocidade inicial $u_0$ é dada por $u_0 = 2$ no intervalo $0,5 \leq x \leq 1$, senão $u_0 = 1$ (isso é, uma função chapéu).
Aqui, utilizamos a função [numpy.ones_like](https://docs.scipy.org/doc/numpy/reference/generated/numpy.ones_like.html), onde `u` terá o mesmo formato de `x`, porém o valor em todos os elementos será 1.
```
u = numpy.ones_like(x) #Função ones_like do numpy
u[(0.5<=x) & (x<=1)] = 2 #Define u = 2 entre 0,5 e 1,
#de acordo com nossa CI
print(u) #Verificamos o resultado na tela
```
**Nota:** perceba que nesse contexto, tanto `numpy.ones_like(x)` quanto `numpy.ones(nx)` produzem o mesmo resultado. Experimente!
Vamos visualizar essa condição inicial representando-a graficamente com [Matplotlib](https://matplotlib.org/). Nós já importamos o pacote gráfico `pyplot` da biblioteca `matplotlib`, agora vamos usar a função chamada `plot`, ao invocar `pyplot.plot`. Para saber mais sobre toda a capacidade de Matplotlib, explore a [Galeria de exemplos](http://matplotlib.org/gallery.html).
Aqui, nós usamos a sintaxe simples para gráficos bidimensionais `plot(x,y)`:
```
pyplot.plot(x, u);
```
Por que a função chapéu (*hat function*) não apresenta linhas perfeitamente retas? Pense sobre isso.
Agora é o momento de implementar a equação de convecção discretizada pelos esquemas de diferenças finitas.
Para cada elemento do nosso arranjo `u`, nós precisamos realizar a operação $u_i^{n+1} = u_i^n - c \frac{\Delta t}{\Delta x}(u_i^n-u_{i-1}^n)$.
Vamos armazenar o resultado em um novo (temporário) arranjo `un`, que será a solução $u$ para o passo de tempo seguinte. Repetiremos essa operação por tantos passos de tempo quando especificamos anteriormente, e então poderemos ver o quão longe a onda se moveu por convecção.
Começamos inicializando o arranjo auxiliar `un` para mantermos os valores que calculamos para o passo de tempo $n+1$, novamente utilizando a função NumPy `ones_like()`.
Finalmente, podemos pensar que temos duas operações iterativas: uma no tempo e uma no espaço (veremos uma abordagem diferente mais tarde). Então, começamos pelo aninhamento um laço dentro de outro. Note que usamos a elegante função `range()`. Quando escrevemos `for i in range(1,nx)`, nós vamos interar através do arranjo `u`, mas vamos pular o primeiro elemento (0 ou elemento zero). *Por quê?*
```
un = numpy.ones_like(u) #Inicializar o arranjo temporário
for n in range(nt): #Laço para os valores de 0 a nt-1, então será executado nt vezes
un = u.copy() ##Cópia dos valores de u para un
for i in range(1, nx): ##Você pode tentar comentar essa linha...
#for i in range(nx): ## ... e descomentar essa linha, para ver o que acontece!
u[i] = un[i] - c * dt / dx * (un[i] - un[i-1])
```
**Nota:** Vamos aprender mais tarde que o código escrito acima é bastante ineficiente, e que existem melhores maneiras para escreve-lo ao estilo Python. Porém, vamos continuar.
Agora, vamos graficar nosso arranjo `u` após o avanço no tempo.
```
pyplot.plot(x, u);
```
Certo! Então nossa função chapéu definitivamente se moveu para a direita, entretanto, ela não se parece mais com um chapéu. **O que aconteceu?**
Material Complementar
-----
***
Para uma explicação mais detalhada sobre o método de diferenças finitas, incluindo assuntos como erro de truncamento, ordem de convergência e outros detalhes, assista (EN) **Video Lessons 2 and 3** por Prof. Barba no YouTube.
```
from IPython.display import YouTubeVideo
YouTubeVideo('iz22_37mMkk')
YouTubeVideo('xq9YTcv-fQg')
```
Para uma explicação passo à passo sobre a discretização da equaçaõ de convecção linear com diferenças finitas (e também os passos seguintes, até o Passo 4), assista **Video Lesson 4** por Prof. Barba no YouTube.
```
YouTubeVideo('y2WaK7_iMRI')
```
## Por último, mas não menos importante
**Lembre-se** de reescrever o Passo 1 como um novo script Python, ou seu próprio Jupyter notebook e então experimente mudar os parâmetros de discretização. Uma vez feito isso, você pode prosseguir para o [Passo 2](./02_Passo_2.ipynb).
***
```
from IPython.core.display import HTML
def css_styling():
styles = open("../styles/custom.css", "r").read()
return HTML(styles)
css_styling()
```
> A célula acima executa o estilo para esse notebook. Nós modificamos o estilo encontrado no GitHub de [CamDavidsonPilon](https://github.com/CamDavidsonPilon), [@Cmrn_DP](https://twitter.com/cmrn_dp).
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Using the SavedModel format
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/guide/saved_model"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/saved_model.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/saved_model.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/saved_model.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
A SavedModel contains a complete TensorFlow program, including weights and computation. It does not require the original model building code to run, which makes it useful for sharing or deploying (with [TFLite](https://tensorflow.org/lite), [TensorFlow.js](https://js.tensorflow.org/), [TensorFlow Serving](https://www.tensorflow.org/tfx/serving/tutorials/Serving_REST_simple), or [TensorFlow Hub](https://tensorflow.org/hub)).
This document dives into some of the details of how to use the low-level `tf.saved_model` api:
- If you are using a `tf.keras.Model` the `keras.Model.save(output_path)` method may be all you need: See the [Keras save and serialize](keras/save_and_serialize.ipynb)
- If you just want to save/load weights during training see the [guide to training checkpoints](./checkpoint.ipynb).
## Creating a SavedModel from Keras
For a quick introduction, this section exports a pre-trained Keras model and serves image classification requests with it. The rest of the guide will fill in details and discuss other ways to create SavedModels.
```
import os
import tempfile
from matplotlib import pyplot as plt
import numpy as np
import tensorflow as tf
tmpdir = tempfile.mkdtemp()
physical_devices = tf.config.experimental.list_physical_devices('GPU')
if physical_devices:
tf.config.experimental.set_memory_growth(physical_devices[0], True)
file = tf.keras.utils.get_file(
"grace_hopper.jpg",
"https://storage.googleapis.com/download.tensorflow.org/example_images/grace_hopper.jpg")
img = tf.keras.preprocessing.image.load_img(file, target_size=[224, 224])
plt.imshow(img)
plt.axis('off')
x = tf.keras.preprocessing.image.img_to_array(img)
x = tf.keras.applications.mobilenet.preprocess_input(
x[tf.newaxis,...])
```
We'll use an image of Grace Hopper as a running example, and a Keras pre-trained image classification model since it's easy to use. Custom models work too, and are covered in detail later.
```
labels_path = tf.keras.utils.get_file(
'ImageNetLabels.txt',
'https://storage.googleapis.com/download.tensorflow.org/data/ImageNetLabels.txt')
imagenet_labels = np.array(open(labels_path).read().splitlines())
pretrained_model = tf.keras.applications.MobileNet()
result_before_save = pretrained_model(x)
decoded = imagenet_labels[np.argsort(result_before_save)[0,::-1][:5]+1]
print("Result before saving:\n", decoded)
```
The top prediction for this image is "military uniform".
```
mobilenet_save_path = os.path.join(tmpdir, "mobilenet/1/")
tf.saved_model.save(pretrained_model, mobilenet_save_path)
```
The save-path follows a convention used by TensorFlow Serving where the last path component (`1/` here) is a version number for your model - it allows tools like Tensorflow Serving to reason about the relative freshness.
We can load the SavedModel back into Python with `tf.saved_model.load` and see how Admiral Hopper's image is classified.
```
loaded = tf.saved_model.load(mobilenet_save_path)
print(list(loaded.signatures.keys())) # ["serving_default"]
```
Imported signatures always return dictionaries. To customize signature names and output dictionary keys, see [Specifying signatures during export](#specifying_signatures_during_export).
```
infer = loaded.signatures["serving_default"]
print(infer.structured_outputs)
```
Running inference from the SavedModel gives the same result as the original model.
```
labeling = infer(tf.constant(x))[pretrained_model.output_names[0]]
decoded = imagenet_labels[np.argsort(labeling)[0,::-1][:5]+1]
print("Result after saving and loading:\n", decoded)
```
## Running a SavedModel in TensorFlow Serving
SavedModels are usable from Python (more on that below), but production environments typically use a dedicated service for inference without running Python code. This is easy to set up from a SavedModel using TensorFlow Serving.
See the [TensorFlow Serving REST tutorial](https://www.tensorflow.org/tfx/tutorials/serving/rest_simple) for more details about serving, including instructions for installing `tensorflow_model_server` in a notebook or on your local machine. As a quick sketch, to serve the `mobilenet` model exported above just point the model server at the SavedModel directory:
```bash
nohup tensorflow_model_server \
--rest_api_port=8501 \
--model_name=mobilenet \
--model_base_path="/tmp/mobilenet" >server.log 2>&1
```
Then send a request.
```python
!pip install requests
import json
import numpy
import requests
data = json.dumps({"signature_name": "serving_default",
"instances": x.tolist()})
headers = {"content-type": "application/json"}
json_response = requests.post('http://localhost:8501/v1/models/mobilenet:predict',
data=data, headers=headers)
predictions = numpy.array(json.loads(json_response.text)["predictions"])
```
The resulting `predictions` are identical to the results from Python.
## The SavedModel format on disk
A SavedModel is a directory containing serialized signatures and the state needed to run them, including variable values and vocabularies.
```
!ls {mobilenet_save_path}
```
The `saved_model.pb` file stores the actual TensorFlow program, or model, and a set of named signatures, each identifying a function that accepts tensor inputs and produces tensor outputs.
SavedModels may contain multiple variants of the model (multiple `v1.MetaGraphDefs`, identified with the `--tag_set` flag to `saved_model_cli`), but this is rare. APIs which create multiple variants of a model include [`tf.Estimator.experimental_export_all_saved_models`](https://www.tensorflow.org/api_docs/python/tf/estimator/Estimator#experimental_export_all_saved_models) and in TensorFlow 1.x `tf.saved_model.Builder`.
```
!saved_model_cli show --dir {mobilenet_save_path} --tag_set serve
```
The `variables` directory contains a standard training checkpoint (see the [guide to training checkpoints](./checkpoint.ipynb)).
```
!ls {mobilenet_save_path}/variables
```
The `assets` directory contains files used by the TensorFlow graph, for example text files used to initialize vocabulary tables. It is unused in this example.
SavedModels may have an `assets.extra` directory for any files not used by the TensorFlow graph, for example information for consumers about what to do with the SavedModel. TensorFlow itself does not use this directory.
## Saving a custom model
`tf.saved_model.save` supports saving `tf.Module` objects and its subclasses, like `tf.keras.Layer` and `tf.keras.Model`.
Let's look at an example of saving and restoring a `tf.Module`.
```
class CustomModule(tf.Module):
def __init__(self):
super(CustomModule, self).__init__()
self.v = tf.Variable(1.)
@tf.function
def __call__(self, x):
print('Tracing with', x)
return x * self.v
@tf.function(input_signature=[tf.TensorSpec([], tf.float32)])
def mutate(self, new_v):
self.v.assign(new_v)
module = CustomModule()
```
When you save a `tf.Module`, any `tf.Variable` attributes, `tf.function`-decorated methods, and `tf.Module`s found via recursive traversal are saved. (See the [Checkpoint tutorial](./checkpoint.ipynb) for more about this recursive traversal.) However, any Python attributes, functions, and data are lost. This means that when a `tf.function` is saved, no Python code is saved.
If no Python code is saved, how does SavedModel know how to restore the function?
Briefly, `tf.function` works by tracing the Python code to generate a ConcreteFunction (a callable wrapper around `tf.Graph`). When saving a `tf.function`, you're really saving the `tf.function`'s cache of ConcreteFunctions.
To learn more about the relationship between `tf.function` and ConcreteFunctions, see the [tf.function guide](../../guide/function).
```
module_no_signatures_path = os.path.join(tmpdir, 'module_no_signatures')
module(tf.constant(0.))
print('Saving model...')
tf.saved_model.save(module, module_no_signatures_path)
```
## Loading and using a custom model
When you load a SavedModel in Python, all `tf.Variable` attributes, `tf.function`-decorated methods, and `tf.Module`s are restored in the same object structure as the original saved `tf.Module`.
```
imported = tf.saved_model.load(module_no_signatures_path)
assert imported(tf.constant(3.)).numpy() == 3
imported.mutate(tf.constant(2.))
assert imported(tf.constant(3.)).numpy() == 6
```
Because no Python code is saved, calling a `tf.function` with a new input signature will fail:
```python
imported(tf.constant([3.]))
```
<pre>
ValueError: Could not find matching function to call for canonicalized inputs ((<tf.Tensor 'args_0:0' shape=(1,) dtype=float32>,), {}). Only existing signatures are [((TensorSpec(shape=(), dtype=tf.float32, name=u'x'),), {})].
</pre>
### Basic fine-tuning
Variable objects are available, and we can backprop through imported functions. That is enough to fine-tune (i.e. retrain) a SavedModel in simple cases.
```
optimizer = tf.optimizers.SGD(0.05)
def train_step():
with tf.GradientTape() as tape:
loss = (10. - imported(tf.constant(2.))) ** 2
variables = tape.watched_variables()
grads = tape.gradient(loss, variables)
optimizer.apply_gradients(zip(grads, variables))
return loss
for _ in range(10):
# "v" approaches 5, "loss" approaches 0
print("loss={:.2f} v={:.2f}".format(train_step(), imported.v.numpy()))
```
### General fine-tuning
A SavedModel from Keras provides [more details](https://github.com/tensorflow/community/blob/master/rfcs/20190509-keras-saved-model.md#serialization-details) than a plain `__call__` to address more advanced cases of fine-tuning. TensorFlow Hub recommends to provide the following of those, if applicable, in SavedModels shared for the purpose of fine-tuning:
* If the model uses dropout or another technique in which the forward pass differs between training and inference (like batch normalization), the `__call__` method takes an optional, Python-valued `training=` argument that defaults to `False` but can be set to `True`.
* Next to the `__call__` attribute, there are `.variable` and `.trainable_variable` attributes with the corresponding lists of variables. A variable that was originally trainable but is meant to be frozen during fine-tuning is omitted from `.trainable_variables`.
* For the sake of frameworks like Keras that represent weight regularizers as attributes of layers or sub-models, there can also be a `.regularization_losses` attribute. It holds a list of zero-argument functions whose values are meant for addition to the total loss.
Going back to the initial MobileNet example, we can see some of those in action:
```
loaded = tf.saved_model.load(mobilenet_save_path)
print("MobileNet has {} trainable variables: {}, ...".format(
len(loaded.trainable_variables),
", ".join([v.name for v in loaded.trainable_variables[:5]])))
trainable_variable_ids = {id(v) for v in loaded.trainable_variables}
non_trainable_variables = [v for v in loaded.variables
if id(v) not in trainable_variable_ids]
print("MobileNet also has {} non-trainable variables: {}, ...".format(
len(non_trainable_variables),
", ".join([v.name for v in non_trainable_variables[:3]])))
```
## Specifying signatures during export
Tools like TensorFlow Serving and `saved_model_cli` can interact with SavedModels. To help these tools determine which ConcreteFunctions to use, we need to specify serving signatures. `tf.keras.Model`s automatically specify serving signatures, but we'll have to explicitly declare a serving signature for our custom modules.
By default, no signatures are declared in a custom `tf.Module`.
```
assert len(imported.signatures) == 0
```
To declare a serving signature, specify a ConcreteFunction using the `signatures` kwarg. When specifying a single signature, its signature key will be `'serving_default'`, which is saved as the constant `tf.saved_model.DEFAULT_SERVING_SIGNATURE_DEF_KEY`.
```
module_with_signature_path = os.path.join(tmpdir, 'module_with_signature')
call = module.__call__.get_concrete_function(tf.TensorSpec(None, tf.float32))
tf.saved_model.save(module, module_with_signature_path, signatures=call)
imported_with_signatures = tf.saved_model.load(module_with_signature_path)
list(imported_with_signatures.signatures.keys())
```
To export multiple signatures, pass a dictionary of signature keys to ConcreteFunctions. Each signature key corresponds to one ConcreteFunction.
```
module_multiple_signatures_path = os.path.join(tmpdir, 'module_with_multiple_signatures')
signatures = {"serving_default": call,
"array_input": module.__call__.get_concrete_function(tf.TensorSpec([None], tf.float32))}
tf.saved_model.save(module, module_multiple_signatures_path, signatures=signatures)
imported_with_multiple_signatures = tf.saved_model.load(module_multiple_signatures_path)
list(imported_with_multiple_signatures.signatures.keys())
```
By default, the output tensor names are fairly generic, like `output_0`. To control the names of outputs, modify your `tf.function` to return a dictionary that maps output names to outputs. The names of inputs are derived from the Python function arg names.
```
class CustomModuleWithOutputName(tf.Module):
def __init__(self):
super(CustomModuleWithOutputName, self).__init__()
self.v = tf.Variable(1.)
@tf.function(input_signature=[tf.TensorSpec([], tf.float32)])
def __call__(self, x):
return {'custom_output_name': x * self.v}
module_output = CustomModuleWithOutputName()
call_output = module_output.__call__.get_concrete_function(tf.TensorSpec(None, tf.float32))
module_output_path = os.path.join(tmpdir, 'module_with_output_name')
tf.saved_model.save(module_output, module_output_path,
signatures={'serving_default': call_output})
imported_with_output_name = tf.saved_model.load(module_output_path)
imported_with_output_name.signatures['serving_default'].structured_outputs
```
## SavedModels from Estimators
Estimators export SavedModels through [`tf.Estimator.export_saved_model`](https://www.tensorflow.org/api_docs/python/tf/estimator/Estimator#export_saved_model). See the [guide to Estimator](https://www.tensorflow.org/guide/estimator) for details.
```
input_column = tf.feature_column.numeric_column("x")
estimator = tf.estimator.LinearClassifier(feature_columns=[input_column])
def input_fn():
return tf.data.Dataset.from_tensor_slices(
({"x": [1., 2., 3., 4.]}, [1, 1, 0, 0])).repeat(200).shuffle(64).batch(16)
estimator.train(input_fn)
serving_input_fn = tf.estimator.export.build_parsing_serving_input_receiver_fn(
tf.feature_column.make_parse_example_spec([input_column]))
estimator_base_path = os.path.join(tmpdir, 'from_estimator')
estimator_path = estimator.export_saved_model(estimator_base_path, serving_input_fn)
```
This SavedModel accepts serialized `tf.Example` protocol buffers, which are useful for serving. But we can also load it with `tf.saved_model.load` and run it from Python.
```
imported = tf.saved_model.load(estimator_path)
def predict(x):
example = tf.train.Example()
example.features.feature["x"].float_list.value.extend([x])
return imported.signatures["predict"](
examples=tf.constant([example.SerializeToString()]))
print(predict(1.5))
print(predict(3.5))
```
`tf.estimator.export.build_raw_serving_input_receiver_fn` allows you to create input functions which take raw tensors rather than `tf.train.Example`s.
## Load a SavedModel in C++
The C++ version of the SavedModel [loader](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/cc/saved_model/loader.h) provides an API to load a SavedModel from a path, while allowing SessionOptions and RunOptions. You have to specify the tags associated with the graph to be loaded. The loaded version of SavedModel is referred to as SavedModelBundle and contains the MetaGraphDef and the session within which it is loaded.
```C++
const string export_dir = ...
SavedModelBundle bundle;
...
LoadSavedModel(session_options, run_options, export_dir, {kSavedModelTagTrain},
&bundle);
```
<a id=saved_model_cli/>
## Details of the SavedModel command line interface
You can use the SavedModel Command Line Interface (CLI) to inspect and
execute a SavedModel.
For example, you can use the CLI to inspect the model's `SignatureDef`s.
The CLI enables you to quickly confirm that the input
Tensor dtype and shape match the model. Moreover, if you
want to test your model, you can use the CLI to do a sanity check by
passing in sample inputs in various formats (for example, Python
expressions) and then fetching the output.
### Install the SavedModel CLI
Broadly speaking, you can install TensorFlow in either of the following
two ways:
* By installing a pre-built TensorFlow binary.
* By building TensorFlow from source code.
If you installed TensorFlow through a pre-built TensorFlow binary,
then the SavedModel CLI is already installed on your system
at pathname `bin/saved_model_cli`.
If you built TensorFlow from source code, you must run the following
additional command to build `saved_model_cli`:
```
$ bazel build tensorflow/python/tools:saved_model_cli
```
### Overview of commands
The SavedModel CLI supports the following two commands on a SavedModel:
* `show`, which shows the computations available from a SavedModel.
* `run`, which runs a computation from a SavedModel.
### `show` command
A SavedModel contains one or more model variants (technically, `v1.MetaGraphDef`s), identified by their tag-sets. To serve a model, you might wonder what kind of `SignatureDef`s are in each model variant, and what are their inputs and outputs. The `show` command let you examine the contents of the SavedModel in hierarchical order. Here's the syntax:
```
usage: saved_model_cli show [-h] --dir DIR [--all]
[--tag_set TAG_SET] [--signature_def SIGNATURE_DEF_KEY]
```
For example, the following command shows all available tag-sets in the SavedModel:
```
$ saved_model_cli show --dir /tmp/saved_model_dir
The given SavedModel contains the following tag-sets:
serve
serve, gpu
```
The following command shows all available `SignatureDef` keys for a tag set:
```
$ saved_model_cli show --dir /tmp/saved_model_dir --tag_set serve
The given SavedModel `MetaGraphDef` contains `SignatureDefs` with the
following keys:
SignatureDef key: "classify_x2_to_y3"
SignatureDef key: "classify_x_to_y"
SignatureDef key: "regress_x2_to_y3"
SignatureDef key: "regress_x_to_y"
SignatureDef key: "regress_x_to_y2"
SignatureDef key: "serving_default"
```
If there are *multiple* tags in the tag-set, you must specify
all tags, each tag separated by a comma. For example:
<pre>
$ saved_model_cli show --dir /tmp/saved_model_dir --tag_set serve,gpu
</pre>
To show all inputs and outputs TensorInfo for a specific `SignatureDef`, pass in
the `SignatureDef` key to `signature_def` option. This is very useful when you
want to know the tensor key value, dtype and shape of the input tensors for
executing the computation graph later. For example:
```
$ saved_model_cli show --dir \
/tmp/saved_model_dir --tag_set serve --signature_def serving_default
The given SavedModel SignatureDef contains the following input(s):
inputs['x'] tensor_info:
dtype: DT_FLOAT
shape: (-1, 1)
name: x:0
The given SavedModel SignatureDef contains the following output(s):
outputs['y'] tensor_info:
dtype: DT_FLOAT
shape: (-1, 1)
name: y:0
Method name is: tensorflow/serving/predict
```
To show all available information in the SavedModel, use the `--all` option.
For example:
<pre>
$ saved_model_cli show --dir /tmp/saved_model_dir --all
MetaGraphDef with tag-set: 'serve' contains the following SignatureDefs:
signature_def['classify_x2_to_y3']:
The given SavedModel SignatureDef contains the following input(s):
inputs['inputs'] tensor_info:
dtype: DT_FLOAT
shape: (-1, 1)
name: x2:0
The given SavedModel SignatureDef contains the following output(s):
outputs['scores'] tensor_info:
dtype: DT_FLOAT
shape: (-1, 1)
name: y3:0
Method name is: tensorflow/serving/classify
...
signature_def['serving_default']:
The given SavedModel SignatureDef contains the following input(s):
inputs['x'] tensor_info:
dtype: DT_FLOAT
shape: (-1, 1)
name: x:0
The given SavedModel SignatureDef contains the following output(s):
outputs['y'] tensor_info:
dtype: DT_FLOAT
shape: (-1, 1)
name: y:0
Method name is: tensorflow/serving/predict
</pre>
### `run` command
Invoke the `run` command to run a graph computation, passing
inputs and then displaying (and optionally saving) the outputs.
Here's the syntax:
```
usage: saved_model_cli run [-h] --dir DIR --tag_set TAG_SET --signature_def
SIGNATURE_DEF_KEY [--inputs INPUTS]
[--input_exprs INPUT_EXPRS]
[--input_examples INPUT_EXAMPLES] [--outdir OUTDIR]
[--overwrite] [--tf_debug]
```
The `run` command provides the following three ways to pass inputs to the model:
* `--inputs` option enables you to pass numpy ndarray in files.
* `--input_exprs` option enables you to pass Python expressions.
* `--input_examples` option enables you to pass `tf.train.Example`.
#### `--inputs`
To pass input data in files, specify the `--inputs` option, which takes the
following general format:
```bsh
--inputs <INPUTS>
```
where *INPUTS* is either of the following formats:
* `<input_key>=<filename>`
* `<input_key>=<filename>[<variable_name>]`
You may pass multiple *INPUTS*. If you do pass multiple inputs, use a semicolon
to separate each of the *INPUTS*.
`saved_model_cli` uses `numpy.load` to load the *filename*.
The *filename* may be in any of the following formats:
* `.npy`
* `.npz`
* pickle format
A `.npy` file always contains a numpy ndarray. Therefore, when loading from
a `.npy` file, the content will be directly assigned to the specified input
tensor. If you specify a *variable_name* with that `.npy` file, the
*variable_name* will be ignored and a warning will be issued.
When loading from a `.npz` (zip) file, you may optionally specify a
*variable_name* to identify the variable within the zip file to load for
the input tensor key. If you don't specify a *variable_name*, the SavedModel
CLI will check that only one file is included in the zip file and load it
for the specified input tensor key.
When loading from a pickle file, if no `variable_name` is specified in the
square brackets, whatever that is inside the pickle file will be passed to the
specified input tensor key. Otherwise, the SavedModel CLI will assume a
dictionary is stored in the pickle file and the value corresponding to
the *variable_name* will be used.
#### `--input_exprs`
To pass inputs through Python expressions, specify the `--input_exprs` option.
This can be useful for when you don't have data
files lying around, but still want to sanity check the model with some simple
inputs that match the dtype and shape of the model's `SignatureDef`s.
For example:
```bsh
`<input_key>=[[1],[2],[3]]`
```
In addition to Python expressions, you may also pass numpy functions. For
example:
```bsh
`<input_key>=np.ones((32,32,3))`
```
(Note that the `numpy` module is already available to you as `np`.)
#### `--input_examples`
To pass `tf.train.Example` as inputs, specify the `--input_examples` option.
For each input key, it takes a list of dictionary, where each dictionary is an
instance of `tf.train.Example`. The dictionary keys are the features and the
values are the value lists for each feature.
For example:
```bsh
`<input_key>=[{"age":[22,24],"education":["BS","MS"]}]`
```
#### Save output
By default, the SavedModel CLI writes output to stdout. If a directory is
passed to `--outdir` option, the outputs will be saved as `.npy` files named after
output tensor keys under the given directory.
Use `--overwrite` to overwrite existing output files.
| github_jupyter |
<div align="right" style="text-align: right"><i>Peter Norvig<br>July 2021</i></div>
# Olympic Climbing Wall
From the 538 Riddler on [23 July 2021](https://fivethirtyeight.com/features/can-you-hop-across-the-chessboard/) (rephrased):
>Today marks the beginning of the Summer Olympics! One of the brand-new events this year is [sport climbing](https://olympics.com/tokyo-2020/en/sports/sport-climbing/).
>
>Suppose the organizers place climbing holds uniformly at randomly on a 10-by-10 meter climbing wall until there is a **path**: a series of moves from the bottom of the wall to a hold, and then to successive holds, and finally to the top of the wall, where each move is no more than 1 meter distance. There are two climbing events:
> - For the first event, all the holds are placed on a single vertical line, at random heights.
> - For the second event, holds are placed anywhere on the wall, at random.
>
> On average, how many holds (not including the bottom and top of the wall) have to be placed to make a path in each event?
# First Event
A hold can be represented by a single number, the vertical height off the ground. I'll define `place_holds` to randomly place holds until a path is formed (as detected by `is_path`). Internally to the function, the bottom and top of the wall are considered to be holds, but these are excluded from the output of the function.
```
import random
from typing import List, Tuple, Iterable
from statistics import mean
def place_holds(top=10) -> List[float]:
"""Randomly place holds on wall until there is a path from bottom to top."""
holds = [0, top]
while not is_path(holds):
holds.append(random.uniform(0, top))
holds.sort()
return holds[1:-1] # (not including the bottom and top of the wall)
def is_path(holds) -> bool:
"""Do the sorted holds form a path where each move has distance <= 1?"""
return all(holds[i + 1] - holds[i] <= 1
for i in range(len(holds) - 1))
```
For example, here are random holds that form a path on a 3 meter tall wall:
```
place_holds(3)
```
I can use a [Monte Carlo algorithm](https://en.wikipedia.org/wiki/Monte_Carlo_algorithm) to estimate the expected number of holds by averaging the `len` of repetitions of `place_holds`:
```
def monte_carlo(fn, *args, repeat=50_000, key=len) -> float:
"""Mean value of `repeat` repetitions of key(fn(*args))."""
return mean(key(fn(*args)) for _ in range(repeat))
monte_carlo(place_holds)
```
**Answer: The expected number of holds is about 43** (which I found surprisingly large).
# Second Event
For this event a hold is represented by a point in 2-D space: an `(x, y)` tuple of two numbers:
```
Hold = Point = Tuple[float, float]
def X_(point): return point[0]
def Y_(point): return point[1]
def distance(A: Point, B: Point) -> float:
"""Distance between two 2-D points."""
return abs(complex(*A) - complex(*B))
```
To make it easier to determine when there is a path from bottom to top, I'll keep track, for every hold, of the highest hold that can be reached from that hold (in any number of moves). The data structure `Wall` will be a mapping of `{hold: highest_reachable_hold}`. A `Wall` also has an attribute, `wall.paths`, that is a dict whose entries are `{hold_near_bottom: hold_near_top}` pairs denoting paths from bottom to top.
When a new `hold` is added to the wall, update the wall as follows:
- Find all holds that are within 1 meter of the new `hold` (including the `hold` itself).
- For each of those holds, look up the highest hold they can reach. That set of holds is called `reachable_holds`.
- The highest of the reachable holds is called `highest_hold`.
- Any hold that can reach one of `reachable_holds` can reach all of them (via `hold`), and thus can reach `highest_hold`.
- So update each such hold to say that it can reach `highest_hold`.
- Also, if `highest_hold` is within a meter of the top, and a hold `h` that can reach it is within a meter of the bottom, update the `paths` attribute to include the path `{h: highest_hold}`.
```
class Wall(dict):
"""A Wall is a mapping of {hold: highest_reachable_hold}.
Also keep track of `wall.paths`: a map of {start_hold: end_hold}
where there is a path from start to end, and start is within 1
of the bottom, and end is within 1 of the top."""
def __init__(self, top=10):
self.top = top
self.paths = {} # Paths of the form {hold_near_bottom: hold_near_top}
def add(self, hold: Point):
"""Add hold to this Wall, and merge groups of holds."""
self[hold] = hold # A hold can at least reach itself
self.merge({self[h] for h in self if distance(hold, h) <= 1})
def merge(self, reachable_holds):
"""If you can reach one of these holds, you can reach the highest of them."""
if len(reachable_holds) > 1:
highest_hold = max(reachable_holds, key=Y_)
for h in self:
if self[h] in reachable_holds:
self[h] = highest_hold
if Y_(h) <= 1 and self.top - Y_(highest_hold) <= 1:
self.paths[h] = highest_hold
```
*Note: This could be made more efficient with an [integer lattice](https://en.wikipedia.org/wiki/Fixed-radius_near_neighbors) to quickly find holds within 1 meter, and a [union-find forest](https://en.wikipedia.org/wiki/Disjoint-set_data_structure) to quickly merge groups of holds. But since the expected number of points is small, I opted for simplicity, not efficiency.*
Now `place_holds_2d` is analagous to `place_holds`, but places holds in two dimensions:
```
def place_holds_2d(top=10) -> Wall:
"""Randomly place holds on a square wall until there is a path from bottom to top."""
wall = Wall(top)
while not wall.paths:
wall.add((random.uniform(0, top), random.uniform(0, top)))
return wall
```
Finally, we can estimate the expected number of holds:
```
monte_carlo(place_holds_2d, repeat=5000)
```
**Answer: The expected number of holds is about 143** (which I found surprisingly small).
# Visualization
To get an idea what random climbing walls look like, and to gain confidence in this program, I'll plot some climbing walls, with green dots indicating the random climbing holds, and yellow lines indicating possible paths from bottom to top.
```
import matplotlib.pyplot as plt
def plot_wall(wall):
"""Plot the holds on the wall, and the paths from bottom to top."""
plt.gca().set_aspect('equal', adjustable='box')
plt.xlim(0, wall.top); plt.ylim(0, wall.top)
ends = set(wall.paths.values())
for h in wall:
if wall[h] in ends:
if Y_(h) <= 1: # Plot vertical move from bottom
plot_points([h, (X_(h), 0)], 'y-')
if wall.top - Y_(h) <= 1: # Plot vertical move to top
plot_points([h, (X_(h), wall.top)], 'y-')
for h2 in wall:
if distance(h, h2) <= 1:
plot_points([h, h2], 'y-') # Plot move between holds
plot_points(wall, 'g.') # Plot all holds
plt.title(f'holds: {len(wall)} starts: {len(wall.paths)}')
def plot_points(points, fmt):
"""Plot (x, y) points with given format."""
plt.plot([X_(p) for p in points], [Y_(p) for p in points], fmt)
for i in range(10):
plot_wall(place_holds_2d(10))
plt.show()
```
To get a feel for the internals of a `Wall`, let's look at a smaller one:
```
wall = place_holds_2d(2)
plot_wall(wall)
wall
wall.paths
```
We can also show a wall from the first event:
```
height = 4
wall1 = Wall(height)
for hold in place_holds(height):
wall1.add((height / 2, hold))
plot_wall(wall1)
wall1
```
# Different Size Walls
What if the wall had a size other than 10 meters?
My guess would be that the expected number of required holds goes up roughly linearly on the 1-D wall, and roughly quadratically on the 2-D wall. I can plot expected number of holds for different wall heights, and fit a quadratic polynomial to the data (using `np.polyfit` and `np.poly1d`):
```
import numpy as np
def fit(X, fn, key=len, repeat=1000, degree=2) -> np.array:
"""Fit key(fn(x)) to a polynomial; plot; return polynomial coefficients."""
Y = [monte_carlo(fn, x, key=key, repeat=repeat) for x in X]
coefs = np.polyfit(X, Y, 2)
poly = np.poly1d(coefs)
plt.plot(X, Y, 'o-', label=fn.__name__);
plt.plot(X, [poly(x) for x in X], '.:', label=poly_name(coefs))
plt.legend()
return coefs
def poly_name(coefs, ndigits=2) -> str:
"""A str representing a polynomial."""
degree = len(coefs) - 1
return ' + '.join(term(round(coef, ndigits), degree - i)
for i, coef in enumerate(coefs))
def term(coef, d) -> str:
"""A str representing a term in a polynomial."""
return f'{coef}' + ('' if d == 0 else 'x' if d == 1 else f'x^{d}')
```
First 1-D walls—we see the best-fit quadratic is almost a straight line, but has a slight upward bend:
```
fit(range(2, 41), place_holds);
```
Now 2-D walls—we see a prominent quadratic shape:
```
fit(range(2, 26), place_holds_2d, repeat=100);
```
# Do the Math
The Monte Carlo approach can only give an approximation. To get an exact result requires a level of math that is above my ability. Fortunately, a real mathematician, [George Hauser](https://www.math.rutgers.edu/component/comprofiler/userprofile/gdh43?Itemid=753), provided the following analysis of the first event:
- If you choose uniformly randomly *n* numbers between 0 and 1 and put them in order (including 0 and 1 in the list) and look at the *n*+1 gaps between them, the probability that any given *k* of the gaps are greater than *x* is (1-*kx*)<sup>*n*</sup> if *kx* ≤ 1 and 0 otherwise. So by inclusion-exclusion, the probability that the largest gap is greater than *x* is the sum of the probabilities that each individual gap is greater than *x*, minus the sum of the probabilities that each pair of gaps are simultaneously greater than *x*, plus the sum of all triples, etc.
- So as a formula it is Pr(*X*<sub><i>n</i></sub> > *x*) = ∑<sub><i>k</i> ≤ 1/<i>x</i></sub> (-1)<sup><i>k</i>-1</sup> (*n*+1 choose *k*) (1-*kx*)<sup><i>n</i></sup>.
- Here *X*<sub><i>n</i></sub> is the largest gap that appears in a sample of *n* random points between 0 and 1.
- What we are interested in is *N*, the first step at which *X*<sub><i>n</i></sub> < *x*, and E(*N*) the expectation of *N*.
- This expectation is ∑<sub><i>n</i> ≥ 1</sub> *n* Pr(*X*<sub><i>n</i></sub> < *x* and *X*<sub>*n-1*</sub> > *x*).
- But the sequence *X*<sub><i>n</i></sub> is decreasing since the biggest gap can only get smaller when you add a new hold.
- So this series just telescopes into ∑<sub>*n* ≥ 1</sub> Pr(*X*<sub><i>n</i></sub> > *x*).
- So combining the two formulas we need to evaluate ∑<sub><i>n</i> ≥ 1</sub>∑<sub><i>k</i> ≤ 1/<i>x</i></sub> (-1)<sup><i>k</i>-1</sup> (*n*+1 choose *k*) (1-*kx*)<sup><i>n</i></sup>.
- If you sum first over n, this gives ∑<sub><i>k</i> ≤ 1/<i>x</i></sub> (-1)<sup><i>k</i>-1</sup> (*kx*)<sup>-2</sup> (1/(*kx*)-1)<sup><i>k</i> - 1</sup>.
- I couldn't really simplify this further, but it is easy enough to plug in *x* = 1/10 (i.e. 1 out of 10 meters) and get the answer.
We can use Hauser's formula to do the computation with exact rationals and with floating point numbers:
```
from fractions import Fraction
def hauser(x):
"""George Hauser's formula for the expected number of holds in the first Event."""
return sum((-1) ** (k - 1) * (k * x) ** -2 * (1 / (k * x) - 1) ** (k - 1)
for k in range(1, int(1/x) + 1))
print(hauser(Fraction(1, 10)), '≅', hauser(1 / 10))
```
This agrees well with my Monte Carlo estimate.
| github_jupyter |
# Tables
```
cd ..
import NotebookImport
from DX_screen import *
dx_rna.sort('p').to_csv(FIG_DIR + 'f_up_genes.csv')
dx_mir.sort('p').to_csv(FIG_DIR + 'f_up_miR.csv')
dx_meth.sort('p').to_csv(FIG_DIR + 'f_up_meth.csv')
from metaPCNA import *
dp = -1*meta_pcna_all.unstack()[['01','11']].dropna().T.diff().ix['11']
dp = dp[dp > 0]
dp.name = 'proliferation change'
dx = matched_tn
dx = dx.xs('01',1,1) - dx.xs('11',1,1)
pcna_corr_genes = dx.T.corrwith(dp)
pearson_pandas(pcna_corr_genes, dx_rna.frac)
dx = matched_mir
dx = dx.xs('01',1,1) - dx.xs('11',1,1)
pts = dx.columns.intersection(dp.index)
dx = dx.ix[:, pts]
pcna_corr_mir = dx.T.corrwith(dp.ix[pts])
len(pts)
dx_mir_match = binomial_test_screen(matched_mir.ix[:, pts], 1.)
dx_mir_match = dx_mir_match[dx_mir_match.num_dx > 300]
pearson_pandas(pcna_corr_mir, dx_mir_match.frac)
series_scatter(pcna_corr_mir, dx_mir.frac)
%%time
dx = matched_meth
dx = dx.xs('01',1,1) - dx.xs('11',1,1)
pts = dx.columns.intersection(dp.index)
dx = dx.ix[:, pts]
pcna_corr_meth = dx.T.corrwith(dp.ix[pts])
len(pts)
dx_meth_match = binomial_test_screen(matched_meth.ix[:, pts], 1.)
dx_meth_match = dx_meth_match[dx_meth_match.num_dx > 300]
pearson_pandas(pcna_corr_meth, dx_meth_match.frac)
m = pd.rolling_mean(dx_rna.frac.ix[pcna_corr_genes.order().index].dropna(),
window=50, center=True).dropna()
f_win_genes = (dx_rna.frac - m).dropna()
f_win_genes.name = 'fraction overexpressed (detrended)'
m = pd.rolling_mean(dx_mir_match.frac.ix[pcna_corr_mir.order().index].dropna(),
window=50, center=True).dropna()
f_win_mir = (dx_mir_match.frac - m).dropna()
f_win_mir.name = 'fraction overexpressed (detrended)'
m = pd.rolling_mean(dx_meth_match.frac.ix[pcna_corr_meth.order().index].dropna(),
window=50, center=True).dropna()
f_win_meth = (dx_meth_match.frac - m).dropna()
f_win_meth.name = 'fraction overexpressed (detrended)'
df = pd.concat({'fraction upregulated': dx_rna.frac,
'proliferation score': pcna_corr_genes,
'detrended f_up': f_win_genes}, 1)[[1,2,0]]
df = df.ix[dx_rna.index].sort('detrended f_up', ascending=False)
df.to_csv(FIG_DIR + 'proliferation_score_genes.csv')
df = pd.concat({'fraction upregulated': dx_mir.frac,
'fraction upregulated (matched patients)': dx_mir_match.frac,
'proliferation score': pcna_corr_mir,
'detrended f_up': f_win_mir}, 1)[[1,2,3, 0]]
df = df.ix[dx_mir.index].sort('detrended f_up', ascending=False)
df.to_csv(FIG_DIR + 'proliferation_score_mir.csv')
df = pd.concat({'fraction upregulated': dx_meth.frac,
'fraction upregulated (matched patients)': dx_meth_match.frac,
'proliferation score': pcna_corr_meth,
'detrended f_up': f_win_meth}, 1)[[1,2,3, 0]]
df = df.ix[dx_meth.index].sort('detrended f_up', ascending=False)
df.to_csv(FIG_DIR + 'proliferation_score_meth.csv')
```
| github_jupyter |
# VII - Parallel and Distributed Execution
In this notebook, we will execute training across multiple nodes (or in parallel across a single node over multiple GPUs). We will train an image classification model with Resnet20 on the CIFAR-10 data set across multiple nodes in this notebook.
Azure Batch and Batch Shipyard have the ability to perform "gang scheduling" or scheduling multiple nodes for a single task. This is most commonly used for Message Passing Interface (MPI) jobs.
* [Setup](#section1)
* [Configure and Submit MPI Job and Submit](#section2)
* [Delete Multi-Instance Job](#section3)
<a id='section1'></a>
## Setup
Create a simple alias for Batch Shipyard
```
%alias shipyard SHIPYARD_CONFIGDIR=config python $HOME/batch-shipyard/shipyard.py %l
```
Check that everything is working
```
shipyard
```
Read in the account information we saved earlier
```
import json
import os
def read_json(filename):
with open(filename, 'r') as infile:
return json.load(infile)
def write_json_to_file(json_dict, filename):
""" Simple function to write JSON dictionaries to files
"""
with open(filename, 'w') as outfile:
json.dump(json_dict, outfile)
account_info = read_json('account_information.json')
storage_account_key = account_info['storage_account_key']
storage_account_name = account_info['storage_account_name']
STORAGE_ALIAS = account_info['STORAGE_ALIAS']
```
We will need to delete the pool from earlier as we need a different Docker image on the pool. Due to limited core quota on a default Batch account, we'll need to wait for this pool to delete first before proceeding.
```
shipyard pool del -y --wait
IMAGE_NAME = 'alfpark/cntk:2.1-gpu-1bitsgd-py35-cuda8-cudnn6-refdata'
```
This CNTK image contains the 1-bit SGD version of CNTK for use with GPUs. Additionally it already comes preloaded with the CIFAR-10 reference data, so there is no need to download and convert the training data as it is already baked into the image.
Note that if we were using Infiniband/RDMA enabled instances, we would opt to use the `intelmpi` versions of the CNTK images instead.
Now we will create the config structure:
```
config = {
"batch_shipyard": {
"storage_account_settings": STORAGE_ALIAS
},
"global_resources": {
"docker_images": [
IMAGE_NAME
]
}
}
```
Now we'll create the pool specification with a few modifications for this particular execution:
- `inter_node_communication_enabled` will ensure nodes are allocated such that they can communicate with each other (e.g., send and receive network packets)
**Note:** Most often it is better to scale up the execution first, prior to scale out. Due to the default Batch core quota of just 20 cores, we are using 3 `STANDARD_NC6` nodes. In real production runs, we'd most likely scale up to multiple GPUs within a single node (parallel execution) such as `STANDARD_NC12` or `STANDARD_NC24` prior to scaling out to multiple NC nodes (parallel and distributed execution). We can further improve performance by opting to utilize the `STANDARD_NC24r` instances which are Infiniband/RDMA-capable with GPUs. To use this VM, we would also change the `IMAGE` to use the `intelmpi` versions of the Docker images along with `OpenLogic CentOS-HPC 7.3` as the `platform_image`.
```
POOL_ID = 'gpupool-multi-instance'
pool = {
"pool_specification": {
"id": POOL_ID,
"vm_configuration": {
"platform_image": {
"publisher": "Canonical",
"offer": "UbuntuServer",
"sku": "16.04-LTS"
},
},
"vm_size": "STANDARD_NC6",
"vm_count": {
"dedicated": 3
},
"ssh": {
"username": "docker"
},
"inter_node_communication_enabled": True
}
}
!mkdir config
write_json_to_file(config, os.path.join('config', 'config.json'))
write_json_to_file(pool, os.path.join('config', 'pool.json'))
print(json.dumps(config, indent=4, sort_keys=True))
print(json.dumps(pool, indent=4, sort_keys=True))
```
Create the pool, please be patient while the compute nodes are allocated.
```
shipyard pool add -y
```
Ensure that all compute nodes are `idle` and ready to accept tasks:
```
shipyard pool listnodes
```
<a id='section2'></a>
## Configure MPI Job and Submit
MPI jobs in Batch require execution as a multi-instance task. Essentially this allows multiple compute nodes to be used for a single task.
A few things to note in this jobs configuration:
- The `COMMAND` executes the `run_cntk.sh` script which is embedded into the Docker image. This helper script removes complexities needed in order to execute a distributed MPI CNTK job.
- `auto_complete` is being set to `True` which forces the job to move from `active` to `completed` state once all tasks complete. Note that once a job has moved to `completed` state, no new tasks can be added to it.
- `multi_instance` property is populated which enables multiple nodes, e.g., `num_instances` to participate in the execution of this task. The `coordination_command` is the command that is run on all nodes prior to the `command`. Here, we are simply executing the Docker image to run the SSH server for the MPI daemon (e.g., orted, hydra, etc.) to initialize all of the nodes prior to running the application command.
```
JOB_ID = 'cntk-mpi-job'
# reduce the nubmer of epochs to 20 for purposes of this notebook
COMMAND = '/cntk/run_cntk.sh -s /cntk/Examples/Image/Classification/ResNet/Python/TrainResNet_CIFAR10_Distributed.py -- --network resnet20 -q 1 -a 0 --datadir /cntk/Examples/Image/DataSets/CIFAR-10 --outputdir $AZ_BATCH_TASK_WORKING_DIR/output'
jobs = {
"job_specifications": [
{
"id": JOB_ID,
"auto_complete": True,
"tasks": [
{
"image": IMAGE_NAME,
"command": COMMAND,
"gpu": True,
"multi_instance": {
"num_instances": "pool_current_dedicated",
}
}
]
}
]
}
write_json_to_file(jobs, os.path.join('config', 'jobs.json'))
print(json.dumps(jobs, indent=4, sort_keys=True))
```
Submit the job and tail `stdout.txt`:
```
shipyard jobs add --tail stdout.txt
```
Using the command below we can check the status of our jobs. Once all jobs have an exit code we can continue. You can also view the **heatmap** of this pool on [Azure Portal](https://portal.azure.com) to monitor the progress of this job on the compute nodes under your Batch account.
<a id='section3'></a>
## Delete Multi-instance Job
Deleting multi-instance jobs running as Docker containers requires a little more care. We will need to first ensure that the job has entered `completed` state. In the above `jobs` configuration, we set `auto_complete` to `True` enabling the Batch service to automatically complete the job when all tasks finish. This also allows automatic cleanup of the running Docker containers used for executing the MPI job.
Special logic is required to cleanup MPI jobs since the `coordination_command` that runs actually detaches an SSH server. The job auto completion logic Batch Shipyard injects ensures that these containers are killed.
```
shipyard jobs listtasks
```
Once we are sure that the job is completed, then we issue the standard delete command:
```
shipyard jobs del -y --termtasks --wait
shipyard pool del -y --wait
```
[Next notebook: Advanced - Keras Single GPU Training with Tensorflow](08_Keras_Single_GPU_Training_With_Tensorflow.ipynb)
| github_jupyter |
# 2-x-filter Overlay - Demostration Notebook
通过HLS高层次综合工具,可以很方便的通过C/C++语言将算法综合为可在Vivado中直接例化的硬件IP,利用FPGA并行计算的优势,帮助我们实现算法加速,提高系统响应速度。在本示例中通过HLS工具实现了一个阶数与系数均可实时修改的FIR滤波器IP。
2-x-filter Overlay实现了对该滤波器的系统集成,里面包含了2个FIR滤波器,Block Design如下图所示,ARM处理器可通过AXI总线和DMA访问该IP。
<img src="./images/2-x-order_filter.PNG"/>
*注:Overlay可以理解为具体的FPGA比特流 + 相应的Python API驱动*
Address Map如下图所示:
<img src="./images/AddressMap.PNG"/>
而在PYNQ框架下,通过Python API我们可以很方便的对Overlay中的IP进行调用。而基于Python的生态,导入数据分析库如numpy和图形库matplotlib,通过简单的几行代码即可对FIR滤波器进行分析和验证。在本notebook中我们展示了通过numpy库产生的多个频率的叠加信号作为FIR滤波器的输入,并对经过FIR滤波器滤波前后的信号在时域和频频进行了分析。
下表为HLS工具自动为IP产生的驱动头文件,在notebook中需要对照该头文件来对IP进行调用。
==============================================================
File generated on Mon Oct 07 01:59:23 +0800 2019
Vivado(TM) HLS - High-Level Synthesis from C, C++ and SystemC v2018.3 (64-bit)
SW Build 2405991 on Thu Dec 6 23:38:27 MST 2018
IP Build 2404404 on Fri Dec 7 01:43:56 MST 2018
Copyright 1986-2018 Xilinx, Inc. All Rights Reserved.
==============================================================
AXILiteS
0x00 : Control signals
bit 0 - ap_start (Read/Write/COH)
bit 1 - ap_done (Read/COR)
bit 2 - ap_idle (Read)
bit 3 - ap_ready (Read)
bit 7 - auto_restart (Read/Write)
others - reserved
0x04 : Global Interrupt Enable Register
bit 0 - Global Interrupt Enable (Read/Write)
others - reserved
0x08 : IP Interrupt Enable Register (Read/Write)
bit 0 - Channel 0 (ap_done)
bit 1 - Channel 1 (ap_ready)
others - reserved
0x0c : IP Interrupt Status Register (Read/TOW)
bit 0 - Channel 0 (ap_done)
bit 1 - Channel 1 (ap_ready)
others - reserved
0x10 : Data signal of coe
bit 31~0 - coe[31:0] (Read/Write)
0x14 : reserved
0x18 : Data signal of ctrl
bit 31~0 - ctrl[31:0] (Read/Write)
0x1c : reserved
(SC = Self Clear, COR = Clear on Read, TOW = Toggle on Write, COH = Clear on Handshake)
为了帮助我们在notebook上对算法进行验证,我们通过matlab工具设计了2个滤波器,预设信号频率分量最高为750Hz,根据采样定理知采样频率要大于信号频率2倍,在设计的2个滤波器中,均设置扫描频率为1800Hz。
下图为在matlab中设计的的FIR低通滤波器幅频曲线,示例中设计了1个截至频率为500Hz的10阶FIR低通滤波器。
<img src="./images/MagnitudeResponse.PNG" width="70%" height="70%"/>
导出系数:[107,280,-1193,-1212,9334,18136,9334,-1212,-1193,280,107]
修改滤波器设置,重新设计1个截至频率为500Hz的15阶FIR高通滤波器.
<img src="./images/MagnitudeResponse_500Hz_HP.png" width="70%" height="70%"/>
导出系数:[-97,-66,435,0,-1730,1101,5506,-13305,13305,-5506,-1101,1730,0,-435,66,97]
# 步骤1 - 导入Python库,实例化用于控制FIR滤波器的DMA设备。
### 注:我们可以通过“Shift + Enter”组合键来逐一执行notebook中每一个cell内的python脚本。cell左边的"*"号表示脚本正在执行,执行完毕后会变为数字。
```
#导入必要的python库
import pynq.lib.dma #导入访问FPGA内侧DMA的库
import numpy as np #numpy为pyrhon的数值分析库
from pynq import Xlnk #Xlnk()可实现连续内存分配,访问FPGA侧的DMA需要该库
from scipy.fftpack import fft,ifft #python的FFT库
import matplotlib.pyplot as plt #python图表库
import scipy as scipy
#加载FPGA比特流
firn = pynq.Overlay("2-x-order_filter.bit")
#实例化Overlay内的x_order_fir_0和DMA0模块
dma_0 = firn.axi_dma_0
fir_filter_0 = firn.x_order_fir_0
#实例化Overlay内的x_order_fir_1和DMA1模块
dma_1 = firn.axi_dma_1
fir_filter_1 = firn.x_order_fir_1
#对Overlay内的DMA进行配置,每次传输1800个数据点。
xlnk = Xlnk()
in_buffer_0 = xlnk.cma_array(shape=(1800,), dtype=np.int32)
out_buffer_0 = xlnk.cma_array(shape=(1800,), dtype=np.int32)
in_buffer_1 = xlnk.cma_array(shape=(1800,), dtype=np.int32)
out_buffer_1 = xlnk.cma_array(shape=(1800,), dtype=np.int32)
#coe_buffer = xlnk.cma_array(shape=(11,), dtype=np.int32)
coe_buffer_0 = xlnk.cma_array(shape=(16,), dtype=np.int32)
ctrl_buffer_0 = xlnk.cma_array(shape=(2,), dtype=np.int32)
coe_buffer_1 = xlnk.cma_array(shape=(16,), dtype=np.int32)
ctrl_buffer_1 = xlnk.cma_array(shape=(2,), dtype=np.int32)
#coe = [107,280,-1193,-1212,9334,18136,9334,-1212,-1193,280,107]
coe_0 = [-97,-66,435,0,-1730,1101,5506,-13305,13305,-5506,-1101,1730,0,-435,66,97]
for i in range (16):
coe_buffer_0[i] = coe[i]
coe_1 = [-97,-66,435,0,-1730,1101,5506,-13305,13305,-5506,-1101,1730,0,-435,66,97]
for i in range (16):
coe_buffer_1[i] = coe[i]
ctrl_buffer_0[0] = 1
#ctrl_buffer[1] = 10
ctrl_buffer_0[1] = 16
ctrl_buffer_1[0] = 1
#ctrl_buffer[1] = 10
ctrl_buffer_1[1] = 16
coe_buffer_0.physical_address
fir_filter_0.write(0x10,coe_buffer_0.physical_address)
fir_filter_0.write(0x18,ctrl_buffer_0.physical_address)
fir_filter_1.write(0x10,coe_buffer_1.physical_address)
fir_filter_1.write(0x18,ctrl_buffer_1.physical_address)
fir_filter_0.write(0x00,0x81)
fir_filter_1.write(0x00,0x81)
```
# 步骤2 - 叠加多个不同频率和幅值的信号,作为滤波器的输入信号。
```
#采样频率为1800Hz,即1秒内有1800个采样点,我们将采样点个数选择1800个。
x=np.linspace(0,1,1800)
#产生滤波器输入信号
f1 = 600 #设置第1个信号分量频率设置为600Hz
a1 = 100 #设置第1个信号分量幅值设置为100
f2 = 450 #设置第2个信号分量频率设置为450Hz
a2 = 100 #设置第2个信号分量幅值设置为100
f3 = 200 #设置第3个信号分量频率设置为200Hz
a3 = 100 #设置第3个信号分量幅值设置为100
f4 = 650 #设置第4个信号分量频率设置为650Hz
a4 = 100 #设置第5个信号分量幅值设置为100
#产生2个不同频率分量的叠加信号,将其作为滤波器的输入信号,我们还可以叠加更多信号。
#y=np.int32(a1*np.sin(2*np.pi*f1*x) + a2*np.sin(2*np.pi*f2*x))
y_0=np.int32(a1*np.sin(2*np.pi*f1*x) + a2*np.sin(2*np.pi*f2*x) + a3*np.sin(2*np.pi*f3*x) + a4*np.sin(2*np.pi*f4*x))
#绘制滤波器输入信号波形图
fig_0_1 = plt.figure()
ax_0_1 = fig_0_1.gca()
plt.plot(y_0[0:50]) #为便于观察,这里仅显示前50个点的波形,如需要显示更多的点,请将50改为其它数值
plt.title('input signal',fontsize=10,color='b')
y_1=np.int32(a1*np.sin(2*np.pi*f1*x) + a2*np.sin(2*np.pi*f2*x) + a3*np.sin(2*np.pi*f3*x))
#绘制滤波器输入信号波形图
fig_1_1 = plt.figure()
ax_1_1 = fig_1_1.gca()
plt.plot(y_1[0:50]) #为便于观察,这里仅显示前50个点的波形,如需要显示更多的点,请将50改为其它数值
plt.title('input signal',fontsize=10,color='b')
#通过DMA将数据发送in_buffer内的数值到FIR滤波器的输入端
for i in range(1800):
in_buffer_0[i] = y_0[i]
dma_0.sendchannel.transfer(in_buffer_0)
#通过DMA将数据发送in_buffer内的数值到FIR滤波器的输入端
for i in range(1800):
in_buffer_1[i] = y_1[i]
dma_1.sendchannel.transfer(in_buffer_1)
#获取滤波器的输出信号数据存储在out_buffer中
dma_0.recvchannel.transfer(out_buffer_0)
#获取滤波器的输出信号数据存储在out_buffer中
dma_1.recvchannel.transfer(out_buffer_1)
#绘制滤波器输出信号图
fig_0_2 = plt.figure()
ax_0_2 = fig2.gca()
plt.plot(out_buffer_0[0:50]/32768) #除于32768的原因是滤波器系数为16位有符号定点小数,运算过程中被当作整数计算。
plt.title('output signal',fontsize=10,color='b')
#绘制滤波器输出信号图
fig_1_2 = plt.figure()
ax_1_2 = fig2.gca()
plt.plot(out_buffer_1[0:50]/32768) #除于32768的原因是滤波器系数为16位有符号定点小数,运算过程中被当作整数计算。
plt.title('output signal',fontsize=10,color='b')
```
# 步骤3 - 对滤波器输入和输出信号做频域分析
```
#FFT变换函数体
def fft(signal_buffer,points):
yy = scipy.fftpack.fft(signal_buffer)
yreal = yy.real # 获取实部
yimag = yy.imag # 获取虚部
yf1 = abs(yy)/((len(points)/2)) #归一化处理
yf2 = yf1[range(int(len(points)/2))] #由于对称性,只取一半区间
xf1 = np.arange(len(signal_buffer)) # 频率
xf2 = xf1[range(int(len(points)/2))] #取一半区间
#混合波的FFT(双边频率范围)
#plt.subplot(222)
plt.plot(xf2,yf2,'r') #显示原始信号的FFT模值,本例只显示其中的750个点,如需要显示更多请调整750为其它数值
plt.title('FFT of Mixed wave',fontsize=10,color='r') #注意这里的颜色可以查询颜色代码
return
#对输入信号做FFT变换
fft(in_buffer_0,x)
#对输入信号做FFT变换
fft(in_buffer_1,x)
#对输出信号做FFT变换
fft(out_buffer_0/32768,x)#除于32768的原因是滤波器系数为16位有符号定点小数,运算过程中被当作整数计算。
#对输出信号做FFT变换
fft(out_buffer_1/32768,x)#除于32768的原因是滤波器系数为16位有符号定点小数,运算过程中被当作整数计算。
#dma.sendchannel.wait()
#dma.recvchannel.wait()
in_buffer_0.close()
out_buffer_0.close()
in_buffer_1.close()
out_buffer_1.close()
xlnk.xlnk_reset()
```
| github_jupyter |
# Process data
Here we import data from all conditions (for one experiment at a time) and do the necessary processing. This results in a large `.csv` file (eg `EXPERIMENT1DATA.csv`) which is ready for the next stage, parameter estimation.
```
from glob import glob
import os
import numpy as np
import pandas as pd
```
# Experiment 1
```
def import_files(files, paradigm, reward_mag_level):
"""Import raw discounting data from a list of filenames.
The user can adapt this function and the related functions in to come up
with the appropriately structured dataframe.
inputs:
"""
data = []
for i,fname in enumerate(files):
df = pd.read_csv(fname)
df = _new_col_of_value(df, 'paradigm', paradigm)
df = _new_col_of_value(df, 'reward_mag', reward_mag_level)
df.drop(columns=['block_order', 'group', 'index', 'trial'], inplace=True)
df.rename(columns={'A':'RA',
'B': 'RB'}, inplace=True)
data.append(df)
return(pd.concat(data))
def _new_col_of_value(df, colname, value):
df[colname] = pd.Series(value, index=df.index)
return df
def _generate_trial_col(df):
df = df.reset_index()
df['trial'] = df.index
return df
expt = 1
reward_levels = ['low', 'high']
paradigms = ['deferred', 'online']
data = []
for reward_level in reward_levels:
for paradigm in paradigms:
file_location = f'data/raw_data_expt{expt}/{paradigm}_{reward_level}'
files = glob(file_location + '/*.csv')
print(f'{len(files)} files found in {file_location}')
data.append(import_files(files, paradigm, reward_level))
data = pd.concat(data)
```
Need to create a new `id` column from 0 - total number of participants
```
# new column `id` which is the factors of `Participant`
factors, keys = data.Participant.factorize()
data['id'] = pd.Series(factors, index=data.index)
data.head()
```
Recode the condition values into numerical values. We are going to factorize `paradigm` and `reward_mag_level` so we end up with conditions = 1, 2, 3, 4.
```
# multi-column factorize
tuples = data[['paradigm', 'reward_mag']].apply(tuple, axis=1)
# work out factoring and print the condition key. Important for decoding the results by condition!
print('*** CONDITION KEY ***')
factors, keys = pd.factorize( tuples )
for i in np.unique(factors):
print(f'{i} = {keys[i]}')
data['condition'] = pd.factorize( tuples )[0]
data.reset_index()
```
Change `trial` column to equate to each unique trial in the whole dataset rather than counting actual trial number in each experiment.
```
data['trial'] = np.arange(data.shape[0])
data.to_csv('data/processed/EXPERIMENT1DATA.csv')
```
# Experiment 2
```
def import_files(files, paradigm, domain):
"""Import raw discounting data from a list of filenames.
The user can adapt this function and the related functions in to come up
with the appropriately structured dataframe.
inputs:
"""
data = []
for i,fname in enumerate(files):
df = pd.read_csv(fname)
df = _new_col_of_value(df, 'paradigm', paradigm)
df = _new_col_of_value(df, 'domain', domain)
df.drop(columns=['block_order', 'index', 'trial'], inplace=True)
df.rename(columns={'A': 'RA',
'B': 'RB'}, inplace=True)
data.append(df)
return(pd.concat(data))
def _new_col_of_value(df, colname, value):
df[colname] = pd.Series(value, index=df.index)
return df
def _generate_trial_col(df):
df = df.reset_index()
df['trial'] = df.index
return df
expt = 2
domains = ['gain', 'loss']
paradigms = ['deferred', 'online']
data = []
for domain in domains:
for paradigm in paradigms:
file_location = f'data/raw_data_expt{expt}/{paradigm}_{domain}'
files = glob(file_location + '/*.csv')
print(f'{len(files)} files found in {file_location}')
data.append(import_files(files, paradigm, domain))
data = pd.concat(data)
# new column `id` which is the factors of `Participant`
factors, keys = data.Participant.factorize()
data['id'] = pd.Series(factors, index=data.index)
data.head()
# multi-column factorize
tuples = data[['paradigm', 'domain']].apply(tuple, axis=1)
# work out factoring and print the condition key. Important for decoding the results by condition!
print('*** CONDITION KEY ***')
factors, keys = pd.factorize( tuples )
for i in np.unique(factors):
print(f'{i} = {keys[i]}')
data['condition'] = pd.factorize( tuples )[0]
data.reset_index()
data['trial'] = np.arange(data.shape[0])
data.to_csv('data/processed/EXPERIMENT2DATA.csv')
```
| github_jupyter |
```
import time
from tqdm import tqdm
import pandas as pd
import timeit #Для проверки времени работы программы
from multiprocessing.dummy import Pool
from PIL import Image
import matplotlib.pyplot as plt
import collections
import os
import ast
from pandas.core.common import flatten
from more_itertools import sliced
import numpy as np
import urllib.request
from google.colab import drive
drive.mount('/content/drive/')
```
# All posts labeling
After we manually marked up users images merges from different parts of their profile, we created `labeled_merges_upd.csv` file with the user_ids and labels.
File `posts_for_merge.csv` is a main file, where all posts with images urls are presented (it was prepared in `selected_merges_creation.ipynb` notebook).
This notebook contains code with the automatic posts labeling, based on the manually retrieved category labels for selected user image merges.
```
merges_with_labeling = pd.read_csv('labeled_merges_upd.csv', sep=';')
posts_for_labeling = pd.read_csv('posts_for_merge.csv')
# Add user id column to the dataframe
merges_with_labeling['user_id'] = [int(x.split('_')[0]) for x in merges_with_labeling['merge_name'].values]
merges_with_labeling['merge_id'] = [int(x.split('merge')[1].split('.')[0]) - 1 for x in merges_with_labeling['merge_name'].values]
```
In the `labeled_merges_upd` dataset, there are three labels for each user's photo merges:
* for the first merge from the downloaded period
* for the central merge from the period
* for the last merge.
This is done in order to track whether the quality of content in the user's profile has changed during the selected download period.
If two consecutive labels for one user are the same, we will assume that the quality of this user's content did not change during this period, and all other photo merges between them will also be assigned this label.
```
# Labels assignment for the other photo merges according to the hypothesis above
def get_user_df(user_id):
return posts_for_labeling[posts_for_labeling['id'] == user_id].reset_index(drop=True)
def get_labels_for_user(user_id):
return merges_with_labeling[merges_with_labeling['user_id'] == user_id]['lbl'].values, merges_with_labeling[merges_with_labeling['user_id'] == user_id]['merge_id'].values
def create_merges(user_id, lbls_lst, merge_id_start=0, merge_id_end=2, only_borders=False):
user_names = []
merge_names = []
images_to_merge = []
merge_labels = []
user_df = get_user_df(user_id)
labeled_merges_ids = [0, int((len(user_df) - 9) // 2), len(user_df) - 9]
post_id_start = labeled_merges_ids[merge_id_start] # post_id_start included
post_id_end = labeled_merges_ids[merge_id_end] + 1 # post_id_end included
if only_borders:
for lbl_id in range(3):
start_id = labeled_merges_ids[lbl_id]
user_imgs_to_merge = []
for i in range(9):
user_imgs_to_merge.append(user_df.im_url[start_id + i])
merge_labels.append(lbls_lst[lbl_id])
images_to_merge.append(user_imgs_to_merge)
merge_names.append(str(user_id) + '_merge' + str(start_id + 1))
user_names.append(user_id)
return merge_labels, merge_names, images_to_merge, user_names
for start_id in range(post_id_start, post_id_end):
# Shift on one photo
user_imgs_to_merge = []
# Create photo merge
for i in range(9):
user_imgs_to_merge.append(user_df.im_url[start_id + i])
# Append created merge (as a set of 9 links to the images in this merge)
# to the all merges list for given user
images_to_merge.append(user_imgs_to_merge)
# Append merge name for created merge
merge_names.append(str(user_id) + '_merge' + str(start_id + 1))
user_names.append(user_id)
merge_labels.append(lbls_lst[0])
return merge_labels, merge_names, images_to_merge, user_names
def create_labeled_merges(user_ids):
all_users_ids = []
all_merges_imgs = []
all_merges_names = []
all_merges_labels = []
for user_id in user_ids:
merge_names = []
merges_imgs = []
users_ids = []
merge_labels = []
lbl_vals, merges_ids = get_labels_for_user(user_id)
if (len(lbl_vals) == 2) and (merges_ids[1] - merges_ids[0] == 1) and (lbl_vals[0] == lbl_vals[1]):
merge_labels, merge_names, merges_imgs, users_ids = create_merges(user_id, [lbl_vals[0]], merges_ids[0], merges_ids[1])
if len(lbl_vals) == 3:
if (lbl_vals[0] == lbl_vals[1]) and (lbl_vals[1] == lbl_vals[2]):
merge_labels, merge_names, merges_imgs, users_ids = create_merges(user_id, [lbl_vals[0]], 0, 2)
elif lbl_vals[0] == lbl_vals[1]:
merge_labels, merge_names, merges_imgs, users_ids = create_merges(user_id, [lbl_vals[0]], 0, 1)
elif lbl_vals[1] == lbl_vals[2]:
merge_labels, merge_names, merges_imgs, users_ids = create_merges(user_id, [lbl_vals[1]], 1, 2)
else:
merge_labels, merge_names, merges_imgs, users_ids = create_merges(user_id, [lbl_vals[0], lbl_vals[1], lbl_vals[2]], 0, 2, only_borders=True)
all_users_ids.extend(users_ids)
all_merges_imgs.extend(merges_imgs)
all_merges_names.extend(merge_names)
all_merges_labels.extend(merge_labels)
return all_merges_labels, all_merges_names, all_merges_imgs, all_users_ids
all_merges_labels, all_merges_names, all_merges_imgs, all_users_ids = create_labeled_merges(list(set(merges_with_labeling['user_id'].values)))
all_merges_dict = {'merge_name':all_merges_names, 'merge_lbl':all_merges_labels, 'user_id':all_users_ids, 'images':all_merges_imgs}
all_merges_df = pd.DataFrame.from_dict(all_merges_dict)
# save labeled lists for images uploading to csv file
all_merges_df.to_csv('all_labeled_merges.csv')
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
from keras.applications import VGG16
import numpy as np
```
# Adding VGG16 on base_model
```
# VGG16 was designed to work on 224 x 224 pixel input images sizes
img_rows = 224
img_cols = 224
#include_top=False removes the output layer of the model
base_model = VGG16(weights = 'imagenet',
include_top = False,
input_shape = (img_rows, img_cols, 3))
#To get only the name
base_model.layers[0].__class__.__name__
#To see the input/output of a particular layer
base_model.layers[0].input
#Freezing all the layers by making their trainable=False
for layer in base_model.layers:
layer.trainable=False
#Checking this
base_model.layers[12].trainable
# Let's print our layers
for (i,layer) in enumerate(base_model.layers):
print(str(i) + " "+ layer.__class__.__name__, layer.trainable)
#Ouput of the current model
base_model.output
#Now we add our dense layers for a new prediction on top of the base model
from keras.layers import Dense, Flatten
from keras.models import Sequential
top_model = base_model.output
top_model = Flatten()(top_model)
top_model = Dense(512, activation='relu')(top_model) #First added FCL dense layer
top_model = Dense(512, activation='relu')(top_model) #Second added FCL dense layer
top_model = Dense(256, activation='relu')(top_model) #Third added FCL dense layer
top_model = Dense(7, activation='softmax')(top_model) #Output layer with 2 class labels
#Now let's see the top_model output
top_model
base_model.input
#IMP: Mounting the base_model with the top_model and forming newmodel
from keras.models import Model
newmodel = Model(inputs=base_model.input, outputs=top_model)
newmodel.output
newmodel.layers
newmodel.summary()
#Importing our images for recognition
from keras.preprocessing.image import ImageDataGenerator
train_data="/content/drive/My Drive/MLOPS AND DEVOPS/train"
test_data="/content/drive/My Drive/MLOPS AND DEVOPS/validate"
#Data image augmentation
train_datagen = ImageDataGenerator(
rescale=1./255,
rotation_range=20,
width_shift_range=0.2,
height_shift_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
test_datagen = ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory(
train_data,
target_size=(img_rows, img_cols),
class_mode='categorical')
'''#Converting to 4D
for img_train in train_generator:
img_train=np.expand_dims(img_train, axis=0)'''
test_generator = test_datagen.flow_from_directory(
test_data,
target_size=(img_rows, img_cols),
class_mode='categorical',
shuffle=False)
'''#Converting to 4D
for img_test in test_generator:
img_test=np.expand_dims(img_test, axis=0)'''
#Now let's compile our model
from keras.optimizers import RMSprop
newmodel.compile(optimizer = RMSprop(lr=0.0001),
loss = 'categorical_crossentropy',
metrics =['accuracy']
)
history = newmodel.fit_generator(train_generator, epochs=5,steps_per_epoch=100, validation_data=test_generator)
newmodel.save('FaceRecog_VGG16.h5')
from keras.models import load_model
classifier = load_model('FaceRecog_VGG16.h5')
#Testing the Model
%matplotlib inline
#The line above is necesary to show Matplotlib's plots inside a Jupyter Notebook
import cv2
from matplotlib import pyplot as plt
import os
import cv2
from os import listdir
from os.path import isfile, join
five_celeb_dict = {"[0]": "ben_afflek",
"[1]": "elton_john",
"[2]": "jerry_seinfeld",
"[3]": "madonna",
"[4]": "mindy_kaling",
"[5]": "naman_ghumare",
"[6]": "rohit_ghumare",
}
five_celeb_dict_n = {"ben_afflek": "ben_afflek",
"elton_john": "elton_john",
"jerry_seinfeld": "jerry_seinfeld",
"madonna": "madonna",
"mindy_kaling": "mindy_kaling",
"naman_ghumare": "Naman",
"rohit_ghumare": "Rohit"
}
def draw_test(name, pred, im):
celeb =five_celeb_dict[str(pred)]
BLACK = [0,0,0]
expanded_image = cv2.copyMakeBorder(im, 80, 0, 0, 100 ,cv2.BORDER_CONSTANT,value=BLACK)
cv2.putText(expanded_image, celeb, (20, 60) , cv2.FONT_HERSHEY_SIMPLEX,1, (0,0,255), 2)
cv2.imshow(name, expanded_image)
def getRandomImage(path):
"""function loads a random images from a random folder in our test path """
folders = list(filter(lambda x: os.path.isdir(os.path.join(path, x)), os.listdir(path)))
random_directory = np.random.randint(0,len(folders))
path_class = folders[random_directory]
print("Class - " + five_celeb_dict_n[str(path_class)])
file_path = path + path_class
file_names = [f for f in listdir(file_path) if isfile(join(file_path, f))]
random_file_index = np.random.randint(0,len(file_names))
image_name = file_names[random_file_index]
return cv2.imread(file_path+"/"+image_name)
for i in range(0,10):
input_im = getRandomImage("/content/drive/My Drive/MLOPS AND DEVOPS/validate/")
input_original = input_im.copy()
input_original = cv2.resize(input_original, None, fx=0.5, fy=0.5, interpolation = cv2.INTER_LINEAR)
input_im = cv2.resize(input_im, (224, 224), interpolation = cv2.INTER_LINEAR)
input_im = input_im / 255.
input_im = input_im.reshape(1,224,224,3)
# Get Prediction
res = np.argmax(classifier.predict(input_im, 1, verbose = 0), axis=1)
# Show image with predicted class
draw_test("Prediction", res, input_original)
cv2.waitKey(0)
cv2.destroyAllWindows()
```
| github_jupyter |
# 📝 Exercise M6.02
The aim of this exercise it to explore some attributes available in
scikit-learn random forest.
First, we will fit the penguins regression dataset.
```
import pandas as pd
from sklearn.model_selection import train_test_split
penguins = pd.read_csv("../datasets/penguins_regression.csv")
feature_names = ["Flipper Length (mm)"]
target_name = "Body Mass (g)"
data, target = penguins[feature_names], penguins[target_name]
data_train, data_test, target_train, target_test = train_test_split(
data, target, random_state=0)
```
<div class="admonition note alert alert-info">
<p class="first admonition-title" style="font-weight: bold;">Note</p>
<p class="last">If you want a deeper overview regarding this dataset, you can refer to the
Appendix - Datasets description section at the end of this MOOC.</p>
</div>
Create a random forest containing three trees. Train the forest and
check the statistical performance on the testing set in terms of mean
absolute error.
```
# Write your code here.
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_absolute_error
model = RandomForestRegressor()
model.get_params()
model = RandomForestRegressor(n_estimators = 3)
model.fit(data_train, target_train)
target_pred = model.predict(data_test)
mae1 = mean_absolute_error(target_pred, target_test)
mae2 = mean_absolute_error(target_test, target_pred)
print(mae1, mae2)
```
The next steps of this exercise are to:
- create a new dataset containing the penguins with a flipper length
between 170 mm and 230 mm;
- plot the training data using a scatter plot;
- plot the decision of each individual tree by predicting on the newly
created dataset;
- plot the decision of the random forest using this newly created dataset.
<div class="admonition tip alert alert-warning">
<p class="first admonition-title" style="font-weight: bold;">Tip</p>
<p class="last">The trees contained in the forest that you created can be accessed
with the attribute <tt class="docutils literal">estimators_</tt>.</p>
</div>
```
# Write your code here.
import seaborn as sns
#data[data["Flipper Length (mm)"] >= 170 and data["Flipper Length (mm)"] <= 230]
#data[data["Flipper Length (mm)"] >= 170 and data["Flipper Length (mm)"] <= 230]
#data[data["Flipper Length (mm)"] >= 170 and data["Flipper Length (mm)"] <= 230]
penguins2 = penguins[(penguins["Flipper Length (mm)"] >= 170) & (penguins["Flipper Length (mm)"] <= 230)].sort_values("Flipper Length (mm)")
print(penguins2.head())
feature_name = "Flipper Length (mm)"
target_name = "Body Mass (g)"
data2 = penguins2[[feature_name]]
target2 = penguins2[target_name] # Avec les double crochets, on a un DF
target2
import numpy as np
from matplotlib import pyplot as plt
data_ranges = pd.DataFrame(np.linspace(170, 235, num=300),
columns=data.columns)
for i, tree in enumerate(model.estimators_):
print(i)
predictions = tree.predict(data_ranges)
plt.plot(data_ranges, predictions, label = i)
predictions_rf = model.predict(data_ranges)
plt.plot(data_ranges, predictions, label = "RF", color="tab:red")
sns.scatterplot(data=penguins, x=feature_names[0], y=target_name,
color="black", alpha=0.5)
_ = plt.legend()
# Pour le scatterplot, on peut aussi utiliser sans utiliser argument data
sns.scatterplot(x=penguins[feature_name], y=penguins[target_name],
color="black", alpha=0.5)
```
| github_jupyter |
# Resampling an Image onto Another's Physical Space
The purpose of this Notebook is to demonstrate how the physical space described by the meta-data is used when resampling onto a reference image.
```
%matplotlib inline
import matplotlib.pyplot as plt
import SimpleITK as sitk
# If the environment variable SIMPLE_ITK_MEMORY_CONSTRAINED_ENVIRONMENT is set, this will override the ReadImage
# function so that it also resamples the image to a smaller size (testing environment is memory constrained).
%run setup_for_testing
print(sitk.Version())
from myshow import myshow
# Download data to work on
%run update_path_to_download_script
from downloaddata import fetch_data as fdata
OUTPUT_DIR = "Output"
```
Load the RGB cryosectioning of the Visible Human Male dataset. The data is about 1GB so this may take several seconds, or a bit longer if this is the first time the data is downloaded from the midas repository.
```
fixed = sitk.ReadImage(fdata("vm_head_rgb.mha"))
moving = sitk.ReadImage(fdata("vm_head_mri.mha"))
print(fixed.GetSize())
print(fixed.GetOrigin())
print(fixed.GetSpacing())
print(fixed.GetDirection())
print(moving.GetSize())
print(moving.GetOrigin())
print(moving.GetSpacing())
print(moving.GetDirection())
import sys
resample = sitk.ResampleImageFilter()
resample.SetReferenceImage(fixed)
resample.SetInterpolator(sitk.sitkBSpline)
resample.AddCommand(
sitk.sitkProgressEvent,
lambda: print(f"\rProgress: {100*resample.GetProgress():03.1f}%...", end=""),
)
resample.AddCommand(sitk.sitkProgressEvent, lambda: sys.stdout.flush())
out = resample.Execute(moving)
```
Because we are resampling the moving image using the physical location of the fixed image without any transformation (identity), most of the resulting volume is empty. The image content appears in slice 57 and below.
```
myshow(out)
# combine the two images using a checkerboard pattern:
# because the moving image is single channel with a high dynamic range we rescale it to [0,255] and repeat
# the channel 3 times
vis = sitk.CheckerBoard(
fixed,
sitk.Compose([sitk.Cast(sitk.RescaleIntensity(out), sitk.sitkUInt8)] * 3),
checkerPattern=[15, 10, 1],
)
myshow(vis)
```
Write the image to the Output directory: (1) original as a single image volume and (2) as a series of smaller JPEG images which can be constructed into an animated GIF.
```
import os
sitk.WriteImage(vis, os.path.join(OUTPUT_DIR, "example_resample_vis.mha"))
temp = sitk.Shrink(vis, [3, 3, 2])
sitk.WriteImage(
temp, [os.path.join(OUTPUT_DIR, f"r{i:03d}.jpg") for i in range(temp.GetSize()[2])]
)
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#H2O-API-Walkthrough" data-toc-modified-id="H2O-API-Walkthrough-1"><span class="toc-item-num">1 </span>H2O API Walkthrough</a></span><ul class="toc-item"><li><span><a href="#General-Setup" data-toc-modified-id="General-Setup-1.1"><span class="toc-item-num">1.1 </span>General Setup</a></span></li><li><span><a href="#Modeling" data-toc-modified-id="Modeling-1.2"><span class="toc-item-num">1.2 </span>Modeling</a></span></li><li><span><a href="#Hyperparameter-Tuning" data-toc-modified-id="Hyperparameter-Tuning-1.3"><span class="toc-item-num">1.3 </span>Hyperparameter Tuning</a></span></li><li><span><a href="#Model-Interpretation" data-toc-modified-id="Model-Interpretation-1.4"><span class="toc-item-num">1.4 </span>Model Interpretation</a></span></li></ul></li><li><span><a href="#Reference" data-toc-modified-id="Reference-2"><span class="toc-item-num">2 </span>Reference</a></span></li></ul></div>
```
# code for loading the format for the notebook
import os
# path : store the current path to convert back to it later
path = os.getcwd()
os.chdir(os.path.join('..', '..', 'notebook_format'))
from formats import load_style
load_style(plot_style = False)
os.chdir(path)
# 1. magic for inline plot
# 2. magic to print version
# 3. magic so that the notebook will reload external python modules
# 4. magic to enable retina (high resolution) plots
# https://gist.github.com/minrk/3301035
%matplotlib inline
%load_ext watermark
%load_ext autoreload
%autoreload 2
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
%watermark -a 'Ethen' -d -t -v -p h2o,matplotlib
```
# H2O API Walkthrough
## General Setup
The first question you might be asking is why H2O instead of scikit-learn or Spark MLlib.
- People would prefer H2O over scikit-learn because it is much straightforward to integrate ML models into an existing non-Python system, i.e., Java-based product.
- Performance wise, H2O is extremely fast and can outperform scikit-learn by a significant amount when the data size we're dealing with large datset. As for Spark, while it is a decent tool for ETL on raw data (which often is indeed "big"), its ML libraries are often times outperformed (in training time, memory footprint and even accuracy) by much better tools by orders of magnitude. Please refer to the benchmark at the following link for more detailed number-based proofs. [Github: A minimal benchmark for scalability, speed and accuracy of commonly used open source implementations](https://github.com/szilard/benchm-ml)
```bash
# for installing h2o in python
pip install h2o
```
```
# Load the H2O library and start up the H2O cluter locally on your machine
import h2o
# Number of threads, nthreads = -1, means use all cores on your machine
# max_mem_size is the maximum memory (in GB) to allocate to H2O
h2o.init(nthreads = -1, max_mem_size = 8)
```
In this example, we will be working with a cleaned up version of the Lending Club Bad Loans dataset. The purpose here is to predict whether a loan will be bad (i.e. not repaid to the lender). The response column, `bad_loan`, is 1 if the loan was bad, and 0 otherwise.
```
filepath = 'https://raw.githubusercontent.com/h2oai/app-consumer-loan/master/data/loan.csv'
data = h2o.import_file(filepath)
print('dimension:', data.shape)
data.head(6)
```
Since the task we're dealing at hand is a binary classification problem, we must ensure that our response variable is encoded as a `factor` type. If the response is represented as numerical values of 0/1, H2O will assume we want to train a regression model.
```
# encode the binary repsonse as a factor
label_col = 'bad_loan'
data[label_col] = data[label_col].asfactor()
# this is an optional step that checks the factor level
data[label_col].levels()
# if we check types of each column, we can see which columns
# are treated as categorical type (listed as 'enum')
data.types
```
Next, we perform a three-way split:
- 70% for training
- 15% for validation
- 15% for final testing
We will train a data set on one set and use the others to test the validity of the model by ensuring that it can predict accurately on data the model has not been shown. i.e. to ensure our model is generalizable.
```
# 1. for the splitting percentage, we can leave off
# the last proportion, and h2o will generate the
# number for the last subset for us
# 2. setting a seed will guarantee reproducibility
random_split_seed = 1234
train, valid, test = data.split_frame([0.7, 0.15], seed = random_split_seed)
print(train.nrow)
print(valid.nrow)
print(test.nrow)
```
Here, we extract the column name that will serve as our response and predictors. These informations will be used during the model training phase.
```
# .names, .col_names, .columns are
# all equivalent way of accessing the list
# of column names for the h2o dataframe
input_cols = data.columns
# remove the response and the interest rate
# column since it's correlated with our response
input_cols.remove(label_col)
input_cols.remove('int_rate')
input_cols
```
## Modeling
We'll now jump into the model training part, here Gradient Boosted Machine (GBM) is used as an example. We will set the number of trees to be 500. Increasing the number of trees in a ensemble tree-based model like GBM is one way to increase performance of the model, however, we have to be careful not to overfit our model to the training data by using too many trees. To automatically find the optimal number of trees, we can leverage H2O's early stopping functionality.
There are several parameters that could be used to control early stopping. The three that are generic to all the algorithms are: `stopping_rounds`, `stopping_metric` and `stopping_tolerance`.
- `stopping metric` is the metric by which we'd like to measure performance, and we will choose `AUC` here.
- `score_tree_interval` is a parameter specific to tree-based models. e.g. setting score_tree_interval=5 will score the model after every five trees.
The parameters we have specify below is saying that our model will stop training after there have been three scoring intervals where the AUC has not increased more than 0.0005. Since we have specified a validation frame, the stopping tolerance will be computed on validation AUC rather than training AUC.
```
from h2o.estimators.gbm import H2OGradientBoostingEstimator
# we specify an id for the model so we can refer to it more
# easily later
gbm = H2OGradientBoostingEstimator(
seed = 1,
ntrees = 500,
model_id = 'gbm1',
stopping_rounds = 3,
stopping_metric = 'auc',
score_tree_interval = 5,
stopping_tolerance = 0.0005)
# note that it is .train not .fit to train the model
# just in case you're coming from scikit-learn
gbm.train(
y = label_col,
x = input_cols,
training_frame = train,
validation_frame = valid)
# evaluating the performance, printing the whole
# model performance object will give us a whole bunch
# of information, we'll only be accessing the auc metric here
gbm_test_performance = gbm.model_performance(test)
gbm_test_performance.auc()
```
To examine the scoring history, use the `scoring_history` method on a trained model. When early stopping is used, we see that training stopped at before the full 500 trees. Since we also used a validation set in our model, both training and validation performance metrics are stored in the scoring history object. We can take a look at the validation AUC to observe that the correct stopping tolerance was enforced.
```
gbm_history = gbm.scoring_history()
gbm_history
# change default style figure and font size
plt.rcParams['figure.figsize'] = 8, 6
plt.rcParams['font.size'] = 12
plt.plot(gbm_history['training_auc'], label = 'training_auc')
plt.plot(gbm_history['validation_auc'], label = 'validation_auc')
plt.xticks(range(gbm_history.shape[0]), gbm_history['number_of_trees'].apply(int))
plt.title('GBM training history')
plt.legend()
plt.show()
```
## Hyperparameter Tuning
When training machine learning algorithm, often times we wish to perform hyperparameter search. Thus rather than training our model with different parameters manually one-by-one, we will make use of the H2O's Grid Search functionality. H2O offers two types of grid search -- `Cartesian` and `RandomDiscrete`. Cartesian is the traditional, exhaustive, grid search over all the combinations of model parameters in the grid, whereas Random Grid Search will sample sets of model parameters randomly for some specified period of time (or maximum number of models).
```
# specify the grid
gbm_params = {
'max_depth': [3, 5, 9],
'sample_rate': [0.8, 1.0],
'col_sample_rate': [0.2, 0.5, 1.0]}
```
If we wish to specify model parameters that are not part of our grid, we pass them along to the grid via the `H2OGridSearch.train()` method. See example below.
```
from h2o.grid.grid_search import H2OGridSearch
gbm_tuned = H2OGridSearch(
grid_id = 'gbm_tuned1',
hyper_params = gbm_params,
model = H2OGradientBoostingEstimator)
gbm_tuned.train(
y = label_col,
x = input_cols,
training_frame = train,
validation_frame = valid,
# nfolds = 5, # alternatively, we can use N-fold cross-validation
ntrees = 100,
stopping_rounds = 3,
stopping_metric = 'auc',
score_tree_interval = 5,
stopping_tolerance = 0.0005) # we can specify other parameters like early stopping here
```
To compare the model performance among all the models in a grid, sorted by a particular metric (e.g. AUC), we can use the `get_grid` method.
```
gbm_tuned = gbm_tuned.get_grid(
sort_by = 'auc', decreasing = True)
gbm_tuned
```
Instead of running a grid search, the example below shows the code modification needed to run a random search.
In addition to the hyperparameter dictionary, we will need specify the `search_criteria` as `RandomDiscrete'` with a number for `max_models`, which is equivalent to the number of iterations to run for the random search. This example is set to run fairly quickly, we can increase `max_models` to cover more of the hyperparameter space. Also, we can expand the hyperparameter space of each of the algorithms by modifying the hyperparameter list below.
```
# specify the grid and search criteria
gbm_params = {
'max_depth': [3, 5, 9],
'sample_rate': [0.8, 1.0],
'col_sample_rate': [0.2, 0.5, 1.0]}
# note that in addition to max_models
# we can specify max_runtime_secs
# to run as many model as we can
# for X amount of seconds
search_criteria = {
'max_models': 5,
'strategy': 'RandomDiscrete'}
# train the hyperparameter searched model
gbm_tuned = H2OGridSearch(
grid_id = 'gbm_tuned2',
hyper_params = gbm_params,
search_criteria = search_criteria,
model = H2OGradientBoostingEstimator)
gbm_tuned.train(
y = label_col,
x = input_cols,
training_frame = train,
validation_frame = valid,
ntrees = 100)
# evaluate the model performance
gbm_tuned = gbm_tuned.get_grid(
sort_by = 'auc', decreasing = True)
gbm_tuned
```
Lastly, let's extract the top model, as determined by the validation data's AUC score from the grid and use it to evaluate the model performance on a test set, so we get an honest estimate of the top model performance.
```
# our model is reordered based on the sorting done above;
# hence we can retrieve the first model id to retrieve
# the best performing model we currently have
gbm_best = gbm_tuned.models[0]
gbm_best_performance = gbm_best.model_performance(test)
gbm_best_performance.auc()
# saving and loading the model
model_path = h2o.save_model(
model = gbm_best, path = 'h2o_gbm', force = True)
saved_model = h2o.load_model(model_path)
gbm_best_performance = saved_model.model_performance(test)
gbm_best_performance.auc()
# generate the prediction on the test set, notice that
# it will generate the predicted probability
# along with the actual predicted class;
#
# we can extract the predicted probability for the
# positive class and dump it back to pandas using
# the syntax below if necessary:
#
# gbm_test_pred['p1'].as_data_frame(use_pandas = True)
gbm_test_pred = gbm_best.predict(test)
gbm_test_pred
```
## Model Interpretation
After building our predictive model, often times we would like to inspect which variables/features were contributing the most. This interpretation process allows us the double-check to ensure what the model is learning makes intuitive sense and enable us to explain the results to non-technical audiences. With h2o's tree-base models, we can access the `varimp` attribute to get the top important features.
```
gbm_best.varimp(use_pandas = True).head()
```
We can return the variable importance as a pandas dataframe, hopefully the table should make intuitive sense, where the first column is the feature/variable and the rest of the columns are the feature's importance represented under different scale. We'll be working with the last column, where the feature importance has been normalized to sum up to 1. And note the results are already sorting in decreasing order, where the more important the feature the earlier the feature will appear in the table.
```
# we can drop the use_pandas argument and the
# result will be a list of tuple
gbm_best.varimp()[:4]
```
We can also visualize this information with a bar chart.
```
def plot_varimp(h2o_model, n_features = None):
"""Plot variable importance for H2O tree-based models"""
importances = h2o_model.varimp()
feature_labels = [tup[0] for tup in importances]
feature_imp = [tup[3] for tup in importances]
# specify bar centers on the y axis, but flip the order so largest bar appears at top
pos = range(len(feature_labels))[::-1]
if n_features is None:
n_features = min(len(feature_imp), 10)
fig, ax = plt.subplots(1, 1)
plt.barh(pos[:n_features], feature_imp[:n_features],
align = 'center', height = 0.8, color = '#1F77B4', edgecolor = 'none')
# Hide the right and top spines, color others grey
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.spines['bottom'].set_color('#7B7B7B')
ax.spines['left'].set_color('#7B7B7B')
# Only show ticks on the left and bottom spines
ax.yaxis.set_ticks_position('left')
ax.xaxis.set_ticks_position('bottom')
plt.yticks(pos[:n_features], feature_labels[:n_features])
plt.ylim([min(pos[:n_features]) - 1, max(pos[:n_features]) + 1])
title_fontsize = 14
algo = h2o_model._model_json['algo']
if algo == 'gbm':
plt.title('Variable Importance: H2O GBM', fontsize=title_fontsize)
elif algo == 'drf':
plt.title('Variable Importance: H2O RF', fontsize=title_fontsize)
elif algo == 'xgboost':
plt.title('Variable Importance: H2O XGBoost', fontsize=title_fontsize)
plt.show()
plt.rcParams['figure.figsize'] = 10, 8
plot_varimp(gbm_best)
```
Another type of feature interpretation functionality that we can leverage in h2o is the partial dependence plot. Partial dependence plots is a model-agnostic machine learning interpretation tool. Please consider walking through another [resource](http://nbviewer.jupyter.org/github/ethen8181/machine-learning/blob/master/model_selection/partial_dependence/partial_dependence.ipynb) if you're not familiar with what it's doing.
```
partial_dep = gbm_best.partial_plot(data, cols=['annual_inc'], plot=False)
partial_dep[0].as_data_frame()
```
We use the `.partial_plot` method for a h2o estimator, in this case our gbm model, then we pass in the data and the column that we wish to calculate the partial dependence for as the input argument. The result we get back will be the partial dependence table as shown above. To make the process easier, we'll leverage a [helper function](https://github.com/ethen8181/machine-learning/blob/master/big_data/h2o/h2o_explainers.py) on top of the h2o `.partial_plot` method so we can also get a visualization for ease of interpretation.
```
from h2o_explainers import H2OPartialDependenceExplainer
pd_explainer = H2OPartialDependenceExplainer(gbm_best)
pd_explainer.fit(data, 'annual_inc')
pd_explainer.plot()
plt.show()
```
Based on the visualization above, we can see that the higher your annual income, the lower the likelihood it is for your loan to be bad, but after a certain point, then this feature has no affect on the model.
As for the feature `term` we can see that people having a 36 months loan term is less likely to have a bad loan compared to the ones that have a 60 months loan term.
```
pd_explainer.fit(data, 'term')
pd_explainer.plot()
plt.show()
# remember to shut down the h2o cluster once we're done
h2o.cluster().shutdown(prompt = False)
```
# Reference
- [H2O Sphinx Documentation](http://docs.h2o.ai/h2o/latest-stable/h2o-docs/index.html)
- [H2O Web Page Documentation](http://docs.h2o.ai/)
- [Notebook: H2O Decision Tree Ensembles](http://nbviewer.jupyter.org/github/h2oai/h2o-tutorials/blob/24e1b11fa019406cd702a3a491b9414fbfd8c8c0/training/h2o_algos/src/py/decision_tree_ensembles.ipynb)
- [Notebook: H2O GBM Tuning Tutorial for Python](http://nbviewer.jupyter.org/github/h2oai/h2o-3/blob/master/h2o-docs/src/product/tutorials/gbm/gbmTuning.ipynb)
- [Quora: Why would one use H2O.ai over scikit-learn machine learning tool?](https://www.quora.com/Why-would-one-use-H2O-ai-over-scikit-learn-machine-learning-tool)
- [Github: Tutorials and training material for the H2O Machine Learning Platform](https://github.com/h2oai/h2o-tutorials)
- [Github: A minimal benchmark for scalability, speed and accuracy of commonly used open source implementations](https://github.com/szilard/benchm-ml)
| github_jupyter |
# Aggregation and Grouping
An essential piece of analysis of large data is efficient summarization: computing aggregations like ``sum()``, ``mean()``, ``median()``, ``min()``, and ``max()``, in which a single number gives insight into the nature of a potentially large dataset.
In this section, we'll explore aggregations in Pandas, from simple operations akin to what we've seen on NumPy arrays, to more sophisticated operations based on the concept of a ``groupby``.
## Planets Data
Here we will use the Planets dataset, available via the [Seaborn package](http://seaborn.pydata.org/).
It gives information on planets that astronomers have discovered around other stars (known as *extrasolar planets* or *exoplanets* for short). It can be downloaded with a simple Seaborn command:
```
import seaborn as sns
planets = sns.load_dataset('planets')
planets.shape
planets.head()
print("This has some details on the {} extrasolar planets discovered up to {}.".format(planets.shape[0],max(planets.year)))
```
## Simple Aggregation in Pandas
Earlier, we explored some of the data aggregations available for NumPy arrays.
As with a one-dimensional NumPy array, for a Pandas ``Series`` the aggregates return a single value:
```
import numpy as np
import pandas as pd
rng = np.random.RandomState(42)
ser = pd.Series(rng.rand(5))
ser
ser.sum()
ser.mean()
```
For a ``DataFrame``, by default the aggregates return results within each column:
```
df = pd.DataFrame({'A': rng.rand(5),
'B': rng.rand(5)})
df
df.mean()
```
By specifying the ``axis`` argument, you can instead aggregate within each row:
```
df.mean(axis='columns')
```
Pandas ``Series`` and ``DataFrame``s include a wide range of aggregations; in addition, there is a convenient method ``describe()`` that computes several common aggregates for each column and returns the result.
Let's use this on the Planets data, for now dropping rows with missing values:
```
planets.dropna().describe()
```
This can be a useful way to begin understanding the overall properties of a dataset.
For example, we see in the ``year`` column that although exoplanets were discovered as far back as 1989, half of all known expolanets were not discovered until 2010 or after.
This is largely thanks to the *Kepler* mission, which is a space-based telescope specifically designed for finding eclipsing planets around other stars.
The following table summarizes some other built-in Pandas aggregations:
| Aggregation | Description |
|--------------------------|---------------------------------|
| ``count()`` | Total number of items |
| ``mean()``, ``median()`` | Mean and median |
| ``min()``, ``max()`` | Minimum and maximum |
| ``std()``, ``var()`` | Standard deviation and variance |
| ``mad()`` | Mean absolute deviation |
| ``prod()`` | Product of all items |
| ``sum()`` | Sum of all items |
These methods are both both ``DataFrame`` and ``Series`` objects.
## Your turn
Using the `planets` DataFrame try out the different aggregation functions:
```
# Your code goes here
```
To go deeper into the data, however, simple aggregates are often not enough.
The next level of data summarization is the ``groupby`` operation, which allows you to quickly and efficiently compute aggregates on subsets of data.
## GroupBy: Split, Apply, Combine
Simple aggregations can give you a flavor of your dataset, but often we would prefer to aggregate conditionally on some label or index: this is implemented in the so-called ``groupby`` operation.
The name "group by" comes from a command in the SQL database language, but it is perhaps more illuminative to think of it in the terms first coined by Hadley Wickham of Rstats fame: *split, apply, combine*.
### Split, apply, combine
A canonical example of this split-apply-combine operation, where the "apply" is a summation aggregation, is illustrated in this figure:
<img src="https://github.com/soltaniehha/Business-Analytics/blob/master/figs/06-02split-apply-combine.png?raw=true" width="800" align="center"/>
This makes clear what the ``groupby`` accomplishes:
- The *split* step involves breaking up and grouping a ``DataFrame`` depending on the value of the specified key.
- The *apply* step involves computing some function, usually an aggregate, transformation, or filtering, within the individual groups.
- The *combine* step merges the results of these operations into an output array.
While this could certainly be done manually using some combination of the masking, aggregation, and merging commands covered earlier, an important realization is that *the intermediate splits do not need to be explicitly instantiated*. Rather, the ``GroupBy`` can (often) do this in a single pass over the data, updating the sum, mean, count, min, or other aggregate for each group along the way.
The power of the ``GroupBy`` is that it abstracts away these steps: the user need not think about *how* the computation is done under the hood, but rather thinks about the *operation as a whole*.
As a concrete example, let's take a look at using Pandas for the computation shown in this diagram.
We'll start by creating the input ``DataFrame``:
```
df = pd.DataFrame({'key': ['A', 'B', 'C', 'A', 'B', 'C'],
'data': range(6)})
df
```
The most basic split-apply-combine operation can be computed with the ``groupby()`` method of ``DataFrame``s, passing the name of the desired key column:
```
df.groupby('key')
```
Notice that what is returned is not a set of ``DataFrame``s, but a ``DataFrameGroupBy`` object.
This object is where the magic is: you can think of it as a special view of the ``DataFrame``, which is poised to dig into the groups but does no actual computation until the aggregation is applied.
This "lazy evaluation" approach means that common aggregates can be implemented very efficiently in a way that is almost transparent to the user.
To produce a result, we can apply an aggregate to this ``DataFrameGroupBy`` object, which will perform the appropriate apply/combine steps to produce the desired result:
```
df.groupby('key').sum()
```
The ``sum()`` method is just one possibility here; you can apply virtually any common Pandas or NumPy aggregation function, as well as virtually any valid ``DataFrame`` operation, as we will see in the following discussion.
### The GroupBy object
The ``GroupBy`` object is a very flexible abstraction.
In many ways, you can simply treat it as if it's a collection of ``DataFrame``s, and it does the difficult things under the hood. Let's see some examples using the Planets data.
Perhaps the most important operations made available by a ``GroupBy`` are *aggregate*, *filter*, *transform*, and *apply*.
We'll discuss each of these more fully in the "Aggregate, Filter, Transform, Apply" section below, but before that let's introduce some of the other functionality that can be used with the basic ``GroupBy`` operation.
#### Column indexing
The ``GroupBy`` object supports column indexing in the same way as the ``DataFrame``, and returns a modified ``GroupBy`` object.
For example:
```
planets.groupby('method')
planets.groupby('method')['orbital_period']
```
Here we've selected a particular ``Series`` group from the original ``DataFrame`` group by reference to its column name.
As with the ``GroupBy`` object, no computation is done until we call some aggregate on the object:
```
planets.groupby('method')['orbital_period'].median()
```
This gives an idea of the general scale of orbital periods (in days) that each method is sensitive to.
#### Iteration over groups
The ``GroupBy`` object supports direct iteration over the groups, returning each group as a ``Series`` or ``DataFrame``:
```
for (method, group) in planets.groupby('method'):
print("{0:30s} shape={1}".format(method, group.shape))
```
This can be useful for doing certain things manually, though it is often much faster to use the built-in ``apply`` functionality, which we will discuss momentarily.
#### Dispatch methods
Through some Python class magic, any method not explicitly implemented by the ``GroupBy`` object will be passed through and called on the groups, whether they are ``DataFrame`` or ``Series`` objects.
For example, you can use the ``describe()`` method of ``DataFrame``s to perform a set of aggregations that describe each group in the data:
```
planets.groupby('method')['year'].describe()
```
Looking at this table helps us to better understand the data: for example, the vast majority of planets have been discovered by the Radial Velocity and Transit methods, though the latter only became common (due to new, more accurate telescopes) in the last decade.
The newest methods seem to be Transit Timing Variation and Orbital Brightness Modulation, which were not used to discover a new planet until 2011.
Since `describe()` method returns a DataFrame we can apply the `sort_values()` method to sort based on one of the columns. For instance we can sort by min year to check out the methods based on their history:
```
planets.groupby('method')['year'].describe().sort_values('min')
```
This is just one example of the utility of dispatch methods.
Notice that they are applied *to each individual group*, and the results are then combined within ``GroupBy`` and returned.
Again, any valid ``DataFrame``/``Series`` method can be used on the corresponding ``GroupBy`` object, which allows for some very flexible and powerful operations!
### Aggregate, filter, transform, apply
The preceding discussion focused on aggregation for the combine operation, but there are more options available.
In particular, ``GroupBy`` objects have ``aggregate()``, ``filter()``, ``transform()``, and ``apply()`` methods that efficiently implement a variety of useful operations before combining the grouped data.
For the purpose of the following subsections, we'll use this ``DataFrame``:
```
rng = np.random.RandomState(0)
df = pd.DataFrame({'key': ['A', 'B', 'C', 'A', 'B', 'C'],
'data1': range(6),
'data2': rng.randint(0, 10, 6)})
df
```
#### Aggregation
We're now familiar with ``GroupBy`` aggregations with ``sum()``, ``median()``, and the like, but the ``aggregate()`` method allows for even more flexibility.
It can take a string, a function, or a list thereof, and compute all the aggregates at once.
Here is a quick example combining all these:
```
agg1 = df.groupby('key').aggregate(['min', np.median, max])
agg1
```
The result is a multi-index DataFrame. We can access subsets of this DataFrame by a format like this:
```
agg1['data1']
agg1['data1']['median']
```
Another useful pattern is to pass a dictionary mapping column names to operations to be applied on that column:
```
df.groupby('key').agg({'data1': 'min',
'data2': 'max'})
```
#### Filtering
A filtering operation allows you to drop data based on the group properties.
For example, we might want to keep all groups in which the average is larger than some critical value:
```
df2 = df[['key', 'data2']]
df2
df2.groupby('key').mean()
def filter_func(x):
return x['data2'].mean() > 5
df2.groupby('key').filter(filter_func)
```
The filter function should return a Boolean value specifying whether the group passes the filtering. Here because group A and B do not have an average greater than 5, they are dropped from the result.
#### Transformation
While aggregation must return a reduced version of the data, transformation can return some transformed version of the full data to recombine.
For such a transformation, the output is the same shape as the input.
A common example is to center the data by subtracting the group-wise mean:
```
df2.groupby('key').transform(lambda x: x - x.mean())
```
#### The apply() method
The ``apply()`` method lets you apply an arbitrary function to the group results.
The function should take a ``DataFrame``, and return either a Pandas object (e.g., ``DataFrame``, ``Series``) or a scalar; the combine operation will be tailored to the type of output returned.
For example, here is an ``apply()`` that normalizes the first column by the sum of the second:
```
df
def norm_by_data2(x):
# x is a DataFrame of group values
x['data1'] /= x['data2'].sum()
return x
df.groupby('key').apply(norm_by_data2)
```
``apply()`` within a ``GroupBy`` is quite flexible: the only criterion is that the function takes a ``DataFrame`` and returns a Pandas object or scalar; what you do in the middle is up to you!
### Specifying the split key
In the simple examples presented before, we split the ``DataFrame`` on a single column name.
This is just one of many options by which the groups can be defined, and we'll go through some other options for group specification here.
#### A list, array, series, or index providing the grouping keys
The key can be any series or list with a length matching that of the ``DataFrame``. For example:
```
df
L = [0, 1, 0, 1, 2, 0]
df.groupby(L).sum()
```
#### A dictionary or series mapping index to group
Another method is to provide a dictionary that maps index values to the group keys:
```
df3 = df.set_index('key')
df3
mapping = {'A': 'vowel', 'B': 'consonant', 'C': 'consonant'}
df3.groupby(mapping).sum()
```
#### Any Python function
Similar to mapping, you can pass any Python function that will input the index value and output the group:
```
df3.groupby(str.lower).mean()
```
#### A list of valid keys
Further, any of the preceding key choices can be combined to group on a multi-index:
```
df3.groupby([str.lower, mapping]).mean()
```
### Grouping example
As an example of this, in a couple lines of Python code we can put all these together and count discovered planets by method and by decade:
```
decade = 10 * (planets['year'] // 10)
decade = decade.astype(str) + 's'
planets.groupby(['method', decade])['number'].sum().unstack().fillna(0)
```
This shows the power of combining many of the operations we've discussed up to this point when looking at realistic datasets.
We immediately gain a coarse understanding of when and how planets have been discovered over the past several decades!
Here I would suggest digging into these few lines of code, and evaluating the individual steps to make sure you understand exactly what they are doing to the result.
It's certainly a somewhat complicated example, but understanding these pieces will give you the means to similarly explore your own data.
Let's visualize this summary as a heatmap:
```
summary = planets.groupby(['method', decade])['number'].sum().unstack()
import seaborn as sns
sns.set(rc={'figure.figsize':(14,8)})
sns.heatmap(summary, center=0, annot=True, linewidths=.5, fmt='.0f')
```
| github_jupyter |
```
# Copyright 2021 NVIDIA Corporation. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
<img src="http://developer.download.nvidia.com/compute/machine-learning/frameworks/nvidia_logo.png" style="width: 90px; float: right;">
# Creating Multi-Modal Movie Feature Store
Finally, with both the text and image features ready, we now put the multi-modal movie features into a unified feature store.
If you have downloaded the real data and proceeded through the feature extraction process in notebooks 03-05, then proceed to create the feature store. Else, skip to the `Synthetic data` section below to create random features.
## Real data
```
import pickle
with open('movies_poster_features.pkl', 'rb') as f:
poster_feature = pickle.load(f)["feature_dict"]
len(poster_feature)
with open('movies_synopsis_embeddings-1024.pkl', 'rb') as f:
text_feature = pickle.load(f)["embeddings"]
len(text_feature)
import pandas as pd
links = pd.read_csv("./data/ml-25m/links.csv", dtype={"imdbId": str})
links.shape
links.head()
poster_feature['0105812'].shape
import numpy as np
feature_array = np.zeros((len(links), 1+2048+1024))
for i, row in links.iterrows():
feature_array[i,0] = row['movieId']
if row['imdbId'] in poster_feature:
feature_array[i,1:2049] = poster_feature[row['imdbId']]
if row['movieId'] in text_feature:
feature_array[i,2049:] = text_feature[row['movieId']]
dtype= {**{'movieId': np.int64},**{x: np.float32 for x in ['poster_feature_%d'%i for i in range(2048)]+['text_feature_%d'%i for i in range(1024)]}}
len(dtype)
feature_df = pd.DataFrame(feature_array, columns=['movieId']+['poster_feature_%d'%i for i in range(2048)]+['text_feature_%d'%i for i in range(1024)])
feature_df.head()
feature_df.shape
!pip install pyarrow
feature_df.to_parquet('feature_df.parquet')
```
## Synthetic data
If you have not extrated image and text features from real data, proceed with this section to create synthetic features.
```
import pandas as pd
links = pd.read_csv("./data/ml-25m/links.csv", dtype={"imdbId": str})
import numpy as np
feature_array = np.random.rand(links.shape[0], 3073)
feature_array[:,0] = links['movieId'].values
feature_df = pd.DataFrame(feature_array, columns=['movieId']+['poster_feature_%d'%i for i in range(2048)]+['text_feature_%d'%i for i in range(1024)])
feature_df.to_parquet('feature_df.parquet')
feature_df.head()
```
| github_jupyter |
### Hydrogen Atom
```
import numpy as np
from IPython.display import Image
Image(url='img/Hydrogen_GIF.gif')
```
#### Radius of nth orbit $$ r_n= \frac{n^2h^2\varepsilon_0}{\pi m e^2}=(0.528)n^2Angstrom$$
#### Velocity of electron in nth orbit $$ v_n=\frac{e^2}{2nh\varepsilon_0} = \frac{2.2*10^6}{n}m/s$$
#### Total energy of electron in nth orbit $$ E_n=\frac{-me^4}{8n^2h^2\varepsilon_0^2}=-\frac{13.6}{n^2}eV$$
#### Wave Number $$ \nu =\frac{me^4}{8n^2h^3\varepsilon_0^2}(\frac{1}{n_1^2}-\frac{1}{n_2^2})=R(\frac{1}{n_1^2}-\frac{1}{n_2^2})$$
where R is the Rydberg Constant and $ R = 1.0973*10^7 /m$
#### Wavelength $$ \lambda=\frac{1}{\nu}$$
```
def Hydrogen_radius(n):
f = n**2*k
return f
k=0.528
Hydrogen_radius(1),Hydrogen_radius(2),Hydrogen_radius(3),Hydrogen_radius(4),Hydrogen_radius(5),Hydrogen_radius(6),Hydrogen_radius(7),Hydrogen_radius(8),Hydrogen_radius(9),Hydrogen_radius(10),Hydrogen_radius(11)
def velocity_of_electron(n):
v=k1/n
return v
k1=2.2*10**6
velocity_of_electron(1),velocity_of_electron(2),velocity_of_electron(3),velocity_of_electron(4),velocity_of_electron(5),velocity_of_electron(6),velocity_of_electron(7),velocity_of_electron(8),velocity_of_electron(9),velocity_of_electron(10),velocity_of_electron(11)
def Total_Energy(n):
E=-k2/n**2
return E
k2=13.6
Total_Energy(1), Total_Energy(2), Total_Energy(3), Total_Energy(4), Total_Energy(5), Total_Energy(6), Total_Energy(7), Total_Energy(8), Total_Energy(9),Total_Energy(10),Total_Energy(11)
def Wave_Number(n1,n2):
g=R*(1/(n1)**2-1/(n2)**2)
return g
R=1.097*10**7
Wave_Number(1,2),Wave_Number(1,3),Wave_Number(1,4),Wave_Number(1,5),Wave_Number(1,6),Wave_Number(1,7),Wave_Number(1,8),Wave_Number(1,9),Wave_Number(1,10),Wave_Number(1,11), Wave_Number(1,12)
def Wave_Length(n1,n2):
h=(R*(1/(n1)**2-1/(n2)**2))**-1
return h
R=1.097*10**7
Wave_Length(1,2),Wave_Length(1,3),Wave_Length(1,4),Wave_Length(1,5),Wave_Length(1,6),Wave_Length(1,7),Wave_Length(1,8),Wave_Length(1,9),Wave_Length(1,10),Wave_Length(1,11),Wave_Length(1,12)
class Hydrogen_atom():
def __init__(self,value_k,value_k1,value_k2,value_R, info,info1):
self.k=value_k
self.k1=value_k1
self.k2=value_k2
self.R=value_R
self.info=info
self.info1=info1
def Hydrogen_radius(self,n):
rn=self.k*n**2
#print(self.info)
return rn
def velocity_of_electron(self,n):
vn=self.k1/n
#print(self.info1)
return vn
def Total_Energy(self, n):
En=-self.k2/n**2
return En
def Wave_Number(self,n1,n2):
f=self.R*(1/(n1)**2-1/(n2)**2)
return f
def Wave_Length(self, n1,n2):
L=(self.R*(1/(n1)**2-1/(n2)**2))**-1
return L
H=Hydrogen_atom(value_k=0.528,value_k1=2.2*10**6,value_k2=13.6,value_R=1.097*10**7,info='Wow', info1='Jaw Dropping!!')
H.Hydrogen_radius(1),H.velocity_of_electron(1),H.Total_Energy(1),H.Wave_Number(1,2), H.Wave_Length(1,2)
N=[]
R=[]
V=[]
E=[]
F=[]
G=[]
for n in range(1,70):
rr=H.Hydrogen_radius(n)
vv=H.velocity_of_electron(n)
ee=H.Total_Energy(n)
for n1 in range(1,70):
for n2 in range(1,70):
if n1<n2:
ff=H.Wave_Number(n1,n2)
gg=H.Wave_Length(n1,n2)
N.append(n)
R.append(rr)
V.append(vv)
E.append(ee)
F.append(ff)
G.append(gg)
print(N,R,V,E, F,G)
# construction of a dictionary
data={}
data.update({"PrincipalQuantumNumber":N, "radius":R, "Velocity":V, "Energy":E, "WaveNumber":F, "wavelength":G})
print(data)
import pandas as pd
DF=pd.DataFrame(data)
#print(DF)
DF.to_csv("Hydrogen_atom.csv")
DF.head()
df=pd.read_csv("Hydrogen_atom.csv")
df
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
sns.set()
import random as random
import json as json
import pandas as pd
import numpy as np
plt.plot(df.PrincipalQuantumNumber,df.radius,label='Radius of Orbit', color='r')
plt.plot(df.PrincipalQuantumNumber,df.Velocity,label='Velocity of Electron', color='b')
plt.plot(df.PrincipalQuantumNumber,-df.Energy,label='Total Energy', color='yellow')
plt.plot(df.PrincipalQuantumNumber,df.WaveNumber,label='Wave Number', color='magenta')
plt.plot(df.PrincipalQuantumNumber,df.wavelength,label='Wave Length', color='cyan')
plt.title('Variation of H-functions with n')
plt.xlabel('Principal Quantum Number')
plt.grid(True)
plt.legend()
plt.show()
plt.semilogy(df.PrincipalQuantumNumber,df.radius,label='Radius of Orbit', color='r')
plt.semilogy(df.PrincipalQuantumNumber,df.Velocity,label='Velocity of Electron', color='violet')
plt.semilogy(df.PrincipalQuantumNumber,-df.Energy,label='Total Energy', color='yellow')
plt.semilogy(df.PrincipalQuantumNumber,df.WaveNumber,label='Wave Number', color='green')
plt.semilogy(df.PrincipalQuantumNumber,df.wavelength,label='Wave Length', color='cyan')
plt.xlabel('Principal Quantum Number')
plt.title('Variation of H-functions with n')
plt.grid(True)
#plt.legend()
plt.loglog(df.PrincipalQuantumNumber,df.radius,label='Radius of Orbit', color='r')
plt.loglog(df.PrincipalQuantumNumber,df.Velocity,label='Velocity of Electron', color='violet')
plt.loglog(df.PrincipalQuantumNumber,-df.Energy,label='Total Energy', color='g')
plt.semilogy(df.PrincipalQuantumNumber,df.WaveNumber,label='Wave Number', color='magenta')
plt.semilogy(df.PrincipalQuantumNumber,df.wavelength,label='Wave Length',color='cyan')
plt.title('Variation of H-functions with n')
plt.xlabel('Principal Quantum Number')
plt.grid(True)
plt.figure(figsize=[18,8])
#plt.legend()
```
### Hydrogen Wave functions
#### Higher(quantum interpretation) level reference: https://docs.sympy.org/latest/modules/physics/index.html
## what is JSON?
#### JavaScript Object Notation (JSON) is a standard text-based format for representing structured data based on JavaScript object syntax. It is commonly used for transmitting data in web applications (e.g., sending some data from the server to the client, so it can be displayed on a web page, or vice versa).
JSON is a text-based data format following JavaScript object syntax, which was popularized by Douglas Crockford. Even though it closely resembles JavaScript object literal syntax, it can be used independently from JavaScript, and many programming environments feature the ability to read (parse) and generate JSON.
JSON exists as a string — useful when you want to transmit data across a network. It needs to be converted to a native JavaScript object when you want to access the data. This is not a big issue — JavaScript provides a global JSON object that has methods available for converting between the two.
## What is seaborn?
Seaborn is a Python data visualization library based on matplotlib. It provides a high-level interface for drawing attractive and informative statistical graphics.
| github_jupyter |
# Tutorial 6 - Managing simulation outputs
In the previous tutorials we have interacted with the outputs of the simulation via the default dynamic plot. However, usually we need to access the output data to manipulate it or transfer to another software which is the topic of this notebook.
We start by building and solving our model as shown in previous notebooks:
```
%pip install pybamm -q # install PyBaMM if it is not installed
import pybamm
model = pybamm.lithium_ion.SPMe()
sim = pybamm.Simulation(model)
sim.solve([0, 3600])
```
## Accessing solution variables
We can now access the solved variables directly to visualise or create our own plots. We first extract the solution object:
```
solution = sim.solution
```
and now we can create a post-processed variable (for a list of all the available variables see [Tutorial 3](./Tutorial%203%20-%20Basic%20plotting.ipynb))
```
t = solution["Time [s]"]
V = solution["Terminal voltage [V]"]
```
One option is to visualise the data set returned by the solver directly
```
V.entries
```
which correspond to the data at the times
```
t.entries
```
In addition, post-processed variables can be called at any time (by interpolation)
```
V([200, 400, 780, 1236]) # times in seconds
```
## Saving the simulation and output data
In some cases simulations might take a long time to run so it is advisable to save in your computer so it can be analysed later without re-running the simulation. You can save the whole simulation doing:
```
sim.save("SPMe.pkl")
```
If you now check the root directory of your notebooks you will notice that a new file called `"SPMe.pkl"` has appeared. We can load the stored simulation doing
```
sim2 = pybamm.load("SPMe.pkl")
```
which allows the same manipulation as the original simulation would allow
```
sim2.plot()
```
Alternatively, we can just save the solution of the simulation in a similar way
```
sol = sim.solution
sol.save("SPMe_sol.pkl")
```
and load it in a similar way too
```
sol2 = pybamm.load("SPMe_sol.pkl")
pybamm.dynamic_plot(sol2)
```
Another option is to just save the data for some variables
```
sol.save_data("sol_data.pkl", ["Current [A]", "Terminal voltage [V]"])
```
or save in csv or mat format
```
sol.save_data("sol_data.csv", ["Current [A]", "Terminal voltage [V]"], to_format="csv")
# matlab needs names without spaces
sol.save_data("sol_data.mat", ["Current [A]", "Terminal voltage [V]"], to_format="matlab",
short_names={"Current [A]": "I", "Terminal voltage [V]": "V"})
```
In this notebook we have shown how to extract and store the outputs of PyBaMM's simulations. Next, in [Tutorial 7](./Tutorial%207%20-%20Model%20options.ipynb) we will show how to change the model options.
Before finishing we will remove the data files we saved so that we leave the directory as we found it
```
import os
os.remove("SPMe.pkl")
os.remove("SPMe_sol.pkl")
os.remove("sol_data.pkl")
os.remove("sol_data.csv")
os.remove("sol_data.mat")
```
## References
The relevant papers for this notebook are:
```
pybamm.print_citations()
```
| github_jupyter |
<a href="https://cognitiveclass.ai"><img src = "https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/Logos/organization_logo/organization_logo.png" width = 400> </a>
<h1 align=center><font size = 5>Classification Models with Keras</font></h1>
## Introduction
In this lab, we will learn how to use the Keras library to build models for classificaiton problems. We will use the popular MNIST dataset, a dataset of images, for a change.
The <strong>MNIST database</strong>, short for Modified National Institute of Standards and Technology database, is a large database of handwritten digits that is commonly used for training various image processing systems. The database is also widely used for training and testing in the field of machine learning.
The MNIST database contains 60,000 training images and 10,000 testing images of digits written by high school students and employees of the United States Census Bureau.
Also, this way, will get to compare how conventional neural networks compare to convolutional neural networks, that we will build in the next module.
<h2>Classification Models with Keras</h2>
<h3>Objective for this Notebook<h3>
<h5> 1. Use of MNIST database for training various image processing systems</h5>
<h5> 2. Build a Neural Network </h5>
<h5> 3. Train and Test the Network. </h5>
<p>This link will be used by your peers to assess your project. In your web app, your peers will be able to upload an image, which will then be classified using your custom classifier you connected to the web app. Your project will be graded by how accurately your app can classify <b>Fire</b>, <b>Smoke</b> and <b>Neutral (No Fire or Smoke)</b>.<p>
## Table of Contents
<div class="alert alert-block alert-info" style="margin-top: 20px">
<font size = 3>
1. <a href="#item312">Import Keras and Packages</a>
2. <a href="#item322">Build a Neural Network</a>
3. <a href="#item332">Train and Test the Network</a>
</font>
</div>
<a id='item312'></a>
## Import Keras and Packages
Let's start by importing Keras and some of its modules.
```
import tensorflow.keras
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.utils import to_categorical
```
Since we are dealing we images, let's also import the Matplotlib scripting layer in order to view the images.
```
import matplotlib.pyplot as plt
```
The Keras library conveniently includes the MNIST dataset as part of its API. You can check other datasets within the Keras library [here](https://keras.io/datasets?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ).
So, let's load the MNIST dataset from the Keras library. The dataset is readily divided into a training set and a test set.
```
# import the data
from keras.datasets import mnist
# read the data
(X_train, y_train), (X_test, y_test) = mnist.load_data()
```
Let's confirm the number of images in each set. According to the dataset's documentation, we should have 60000 images in X_train and 10000 images in the X_test.
```
X_train.shape
```
The first number in the output tuple is the number of images, and the other two numbers are the size of the images in datset. So, each image is 28 pixels by 28 pixels.
Let's visualize the first image in the training set using Matplotlib's scripting layer.
```
plt.imshow(X_train[0])
```
With conventional neural networks, we cannot feed in the image as input as is. So we need to flatten the images into one-dimensional vectors, each of size 1 x (28 x 28) = 1 x 784.
```
# flatten images into one-dimensional vector
num_pixels = X_train.shape[1] * X_train.shape[2] # find size of one-dimensional vector
X_train = X_train.reshape(X_train.shape[0], num_pixels).astype('float32') # flatten training images
X_test = X_test.reshape(X_test.shape[0], num_pixels).astype('float32') # flatten test images
```
Since pixel values can range from 0 to 255, let's normalize the vectors to be between 0 and 1.
```
# normalize inputs from 0-255 to 0-1
X_train = X_train / 255
X_test = X_test / 255
```
Finally, before we start building our model, remember that for classification we need to divide our target variable into categories. We use the to_categorical function from the Keras Utilities package.
```
# one hot encode outputs
y_train = to_categorical(y_train)
y_test = to_categorical(y_test)
num_classes = y_test.shape[1]
print(num_classes)
```
<a id='item322'></a>
## Build a Neural Network
```
# define classification model
def classification_model():
# create model
model = Sequential()
model.add(Dense(num_pixels, activation='relu', input_shape=(num_pixels,)))
model.add(Dense(100, activation='relu'))
model.add(Dense(num_classes, activation='softmax'))
# compile model
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
return model
```
<a id='item332'></a>
## Train and Test the Network
```
# build the model
model = classification_model()
# fit the model
model.fit(X_train, y_train, validation_data=(X_test, y_test), epochs=10, verbose=2)
# evaluate the model
scores = model.evaluate(X_test, y_test, verbose=0)
```
Let's print the accuracy and the corresponding error.
```
print('Accuracy: {}% \n Error: {}'.format(scores[1], 1 - scores[1]))
```
Just running 10 epochs could actually take over 20 minutes. But enjoy the results as they are getting generated.
Sometimes, you cannot afford to retrain your model everytime you want to use it, especially if you are limited on computational resources and training your model can take a long time. Therefore, with the Keras library, you can save your model after training. To do that, we use the save method.
```
model.save('classification_model.h5')
```
Since our model contains multidimensional arrays of data, then models are usually saved as .h5 files.
When you are ready to use your model again, you use the load_model function from <strong>keras.models</strong>.
```
from tensorflow.keras.models import load_model
pretrained_model = load_model('classification_model.h5')
```
### Thank you for completing this lab!
This notebook was created by [Alex Aklson](https://www.linkedin.com/in/aklson?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ). I hope you found this lab interesting and educational. Feel free to contact me if you have any questions!
## Change Log
| Date (YYYY-MM-DD) | Version | Changed By | Change Description |
| ----------------- | ------- | ---------- | ----------------------------------------------------------- |
| 2020-09-21 | 2.0 | Srishti | Migrated Lab to Markdown and added to course repo in GitLab |
<hr>
## <h3 align="center"> © IBM Corporation 2020. All rights reserved. <h3/>
This notebook is part of a course on **Coursera** called _Introduction to Deep Learning & Neural Networks with Keras_. If you accessed this notebook outside the course, you can take this course online by clicking [here](https://cocl.us/DL0101EN_Coursera_Week3_LAB2).
<hr>
Copyright © 2019 [IBM Developer Skills Network](https://cognitiveclass.ai?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ).
| github_jupyter |
```
import numpy as np
import pandas as pd
def getNAs(df):
colNames = []
percentNA = []
for i in df.columns:
colNames.append(i)
numNA = df[i].isna().sum()
percent = (numNA/len(df))*100
percentNA.append(percent)
colNames = pd.DataFrame(colNames)
colNames = colNames.rename(columns={0: "label"})
percentNA = pd.DataFrame(percentNA)
percentNA = percentNA.rename(columns={0: "numNA"})
d = pd.concat([colNames,percentNA], axis = 1).sort_values(by=['numNA'], ascending = False)
return d
def remove_columns(df, threshold_percent):
'''drop columns by a threshold (percentage of na)'''
data = getNAs(df)
column_names = data[data.numNA >= threshold_percent].label
clean_data = df.drop(column_names, axis = 1)
return clean_data
#convert all categories to lowercase, there is an issue with factor levels of y, n, N, Y for yes/no
def entry_to_lowercase(df):
for i in df.columns:
if (df[i].dtype == "O"):
df[i] = semi_clean_crop[i].str.lower()
return df
### identify which are categorical and which are numeric by looking at the factor levels
## create dataframe with variable name, levels, number of categories and data types
def id_data_types(df):
names = pd.DataFrame()
category = pd.DataFrame()
data_type = pd.DataFrame()
numUni = pd.DataFrame()
for i in df.columns:
names = names.append({"variable": i},ignore_index = True)
category = category.append({"category": list(df[i].unique())}, ignore_index = True)
numUni = numUni.append({"numUnique": len(df[i].unique())}, ignore_index = True)
data_type = data_type.append({"data_type": df[i].dtype}, ignore_index = True)
view_data = pd.concat([names, category, numUni,data_type],axis =1)
return (view_data)
def replace_na_with_NaN(df):
for i in df.columns:
df[i] = df[i].replace('na', np.NaN)
return df
rhomis_data = pd.read_csv("/Users/anab/Documents/MS_UCDavis/STA208/project/RHoMIS_Full_Data.csv", engine = "python")
rh_indic = pd.read_csv("/Users/anab/Documents/MS_UCDavis/STA208/project/STA_208/data/RHoMIS_Indicators.csv", engine='python')
## final dataset, keep and drop these columns
rhomis_data = rhomis_data[["crop_count", "crop_name_1", "crop_harvest_1", "crop_intercrop_1"]]
rh_indic = rh_indic.drop(['ITERATION','GPS_LAT', 'GPS_LON', 'GPS_ALT', 'Altitude','WorstFoodSecMonth',
'BestFoodSecMonth','FIES_Score', 'currency_conversion_factor', 'GHGEmissions' ],axis = 1)
bigData = pd.concat([rhomis_data,rh_indic ],axis = 1)
bigData = remove_columns(bigData, 50)
bigData.head(3)
bigData = replace_na_with_NaN(bigData)
bigData.dtypes
a = bigData.select_dtypes('object')
id_data_types(a.select_dtypes('object'))
bigData.crop_count = bigData.crop_count.fillna(-500)
bigData.crop_count = bigData.crop_count.astype(int)
bigData.crop_count = bigData.crop_count.replace(-500, np.nan)
```
| github_jupyter |
[](https://colab.research.google.com/github/Picsell-ia/training/blob/master/Object_Detection_TF1_easy.ipynb)
# Prerequisites
We assume that you have a working python3.6+ installation on your computer.
```
!git clone https://github.com/Picsell-ia/training/
%cd training
!pip install -r requirements.txt
```
# Object Detection simplified
In this guide we define a simple wrapper function of the more in-depth object-detecton guide.
```
import sys
sys.path.append("slim")
from picsellia import Client
import picsell_utils
import tensorflow as tf
def wrapper_function(api_token, project_token, model_name, batch_size, nb_steps,
learning_rate=None, annotation_type="rectangle"):
clt = Client(api_token)
clt.checkout_project(project_token=project_token)
clt.checkout_network(model_name)
clt.train_test_split()
clt.dl_pictures()
picsell_utils.create_record_files(label_path=clt.label_path, record_dir=clt.record_dir,
tfExample_generator=clt.tf_vars_generator,
annotation_type=annotation_type)
picsell_utils.edit_config(model_selected=clt.model_selected,
config_output_dir=clt.config_dir,
record_dir=clt.record_dir,
label_map_path=clt.label_path,
num_steps=nb_steps,
batch_size=batch_size,
learning_rate=learning_rate,
annotation_type=annotation_type,
eval_number=len(clt.eval_list))
picsell_utils.train(ckpt_dir=clt.checkpoint_dir,
conf_dir=clt.config_dir)
dict_log = picsell_utils.tfevents_to_dict(path=clt.checkpoint_dir)
metrics = picsell_utils.evaluate(clt.metrics_dir, clt.config_dir, clt.checkpoint_dir)
picsell_utils.export_infer_graph(ckpt_dir=clt.checkpoint_dir,
exported_model_dir=clt.exported_model_dir,
pipeline_config_path=clt.config_dir)
picsell_utils.infer(clt.record_dir,
exported_model_dir=clt.exported_model_dir,
label_map_path=clt.label_path,
results_dir=clt.results_dir, from_tfrecords=True)
clt.send_everything(dict_log, metrics)
api_token = "your_api_token"
project_token = "your_project_token"
model_name = "your_model_name"
wrapper_function(api_token, project_token, model_name,
batch_size=10, nb_steps=1000)
```
| github_jupyter |
```
import numpy
from galpy.potential import LogarithmicHaloPotential
from galpy.orbit import Orbit
from galpy.util import bovy_plot, bovy_coords, bovy_conversion
%pylab inline
```
# Initial conditions for $N$-body simulations to create the impact we want
Setup the potential and coordinate system
```
lp= LogarithmicHaloPotential(normalize=1.,q=0.9)
R0, V0= 8., 220.
```
Functions for converting coordinates between rectangular to cylindrical:
```
def rectangular_to_cylindrical(xv):
R,phi,Z= bovy_coords.rect_to_cyl(xv[:,0],xv[:,1],xv[:,2])
vR,vT,vZ= bovy_coords.rect_to_cyl_vec(xv[:,3],xv[:,4],xv[:,5],R,phi,Z,cyl=True)
out= numpy.empty_like(xv)
# Preferred galpy arrangement of cylindrical coordinates
out[:,0]= R
out[:,1]= vR
out[:,2]= vT
out[:,3]= Z
out[:,4]= vZ
out[:,5]= phi
return out
def cylindrical_to_rectangular(xv):
# Using preferred galpy arrangement of cylindrical coordinates
X,Y,Z= bovy_coords.cyl_to_rect(xv[:,0],xv[:,5],xv[:,3])
vX,vY,vZ= bovy_coords.cyl_to_rectvec(xv[:,1],xv[:,2],xv[:,4],xv[:,5])
out= numpy.empty_like(xv)
out[:,0]= X
out[:,1]= Y
out[:,2]= Z
out[:,3]= vX
out[:,4]= vY
out[:,5]= vZ
return out
```
At the time of impact, the phase-space coordinates of the GC can be computed using orbit integration:
```
xv_prog_init= numpy.array([30.,0.,0.,0.,105.74895,105.74895])
RvR_prog_init= rectangular_to_cylindrical(xv_prog_init[:,numpy.newaxis].T)[0,:]
prog_init= Orbit([RvR_prog_init[0]/R0,RvR_prog_init[1]/V0,RvR_prog_init[2]/V0,
RvR_prog_init[3]/R0,RvR_prog_init[4]/V0,RvR_prog_init[5]],ro=R0,vo=V0)
times= numpy.linspace(0.,10./bovy_conversion.time_in_Gyr(V0,R0),10001)
prog_init.integrate(times,lp)
xv_prog_impact= [prog_init.x(times[-1]),prog_init.y(times[-1]),prog_init.z(times[-1]),
prog_init.vx(times[-1]),prog_init.vy(times[-1]),prog_init.vz(times[-1])]
```
The DM halo at the time of impact is at the following location:
```
xv_dm_impact= numpy.array([-13.500000,2.840000,-1.840000,6.82200571,132.7700529,149.4174464])
RvR_dm_impact= rectangular_to_cylindrical(xv_dm_impact[:,numpy.newaxis].T)[0,:]
dm_impact= Orbit([RvR_dm_impact[0]/R0,RvR_dm_impact[1]/V0,RvR_dm_impact[2]/V0,
RvR_dm_impact[3]/R0,RvR_dm_impact[4]/V0,RvR_dm_impact[5]],ro=R0,vo=V0)
dm_impact= dm_impact.flip()
times= numpy.linspace(0.,10./bovy_conversion.time_in_Gyr(V0,R0),1001)
dm_impact.integrate(times,lp)
```
The orbits over the past 10 Gyr for both objects are:
```
prog_init.plot()
dm_impact.plot(overplot=True)
plot(RvR_dm_impact[0],RvR_dm_impact[3],'ro')
xlim(0.,35.)
ylim(-20.,20.)
```
## Initial condition for the King cluster
We start the King cluster at 10.25 WD time units, which corresponds to 10.25x0.9777922212082034 Gyr. The phase-space coordinates of the cluster are then:
```
prog_backward= prog_init.flip()
ts= numpy.linspace(0.,(10.25*0.9777922212082034-10.)/bovy_conversion.time_in_Gyr(V0,R0),1001)
prog_backward.integrate(ts,lp)
print [prog_backward.x(ts[-1]),prog_backward.y(ts[-1]),prog_backward.z(ts[-1]),
-prog_backward.vx(ts[-1]),-prog_backward.vy(ts[-1]),-prog_backward.vz(ts[-1])]
```
## Initial conditions for the Plummer DM subhalo
## Starting 0.125 time units ago
```
dm_impact= Orbit([RvR_dm_impact[0]/R0,RvR_dm_impact[1]/V0,RvR_dm_impact[2]/V0,
RvR_dm_impact[3]/R0,RvR_dm_impact[4]/V0,RvR_dm_impact[5]],ro=R0,vo=V0)
dm_impact= dm_impact.flip()
ts= numpy.linspace(0.,0.125*0.9777922212082034/bovy_conversion.time_in_Gyr(V0,R0),10001)
dm_impact.integrate(ts,lp)
print [dm_impact.x(ts[-1]),dm_impact.y(ts[-1]),dm_impact.z(ts[-1]),
-dm_impact.vx(ts[-1]),-dm_impact.vy(ts[-1]),-dm_impact.vz(ts[-1])]
```
## Starting 0.25 time units ago
```
dm_impact= Orbit([RvR_dm_impact[0]/R0,RvR_dm_impact[1]/V0,RvR_dm_impact[2]/V0,
RvR_dm_impact[3]/R0,RvR_dm_impact[4]/V0,RvR_dm_impact[5]],ro=R0,vo=V0)
dm_impact= dm_impact.flip()
ts= numpy.linspace(0.,0.25*0.9777922212082034/bovy_conversion.time_in_Gyr(V0,R0),10001)
dm_impact.integrate(ts,lp)
print [dm_impact.x(ts[-1]),dm_impact.y(ts[-1]),dm_impact.z(ts[-1]),
-dm_impact.vx(ts[-1]),-dm_impact.vy(ts[-1]),-dm_impact.vz(ts[-1])]
```
## Starting 0.375 time units ago
```
dm_impact= Orbit([RvR_dm_impact[0]/R0,RvR_dm_impact[1]/V0,RvR_dm_impact[2]/V0,
RvR_dm_impact[3]/R0,RvR_dm_impact[4]/V0,RvR_dm_impact[5]],ro=R0,vo=V0)
dm_impact= dm_impact.flip()
ts= numpy.linspace(0.,0.375*0.9777922212082034/bovy_conversion.time_in_Gyr(V0,R0),10001)
dm_impact.integrate(ts,lp)
print [dm_impact.x(ts[-1]),dm_impact.y(ts[-1]),dm_impact.z(ts[-1]),
-dm_impact.vx(ts[-1]),-dm_impact.vy(ts[-1]),-dm_impact.vz(ts[-1])]
```
## Starting 0.50 time units ago
```
dm_impact= Orbit([RvR_dm_impact[0]/R0,RvR_dm_impact[1]/V0,RvR_dm_impact[2]/V0,
RvR_dm_impact[3]/R0,RvR_dm_impact[4]/V0,RvR_dm_impact[5]],ro=R0,vo=V0)
dm_impact= dm_impact.flip()
ts= numpy.linspace(0.,0.50*0.9777922212082034/bovy_conversion.time_in_Gyr(V0,R0),10001)
dm_impact.integrate(ts,lp)
print [dm_impact.x(ts[-1]),dm_impact.y(ts[-1]),dm_impact.z(ts[-1]),
-dm_impact.vx(ts[-1]),-dm_impact.vy(ts[-1]),-dm_impact.vz(ts[-1])]
```
## Initial conditions for the Plummer DM subhalo with $\lambda$ scaled interaction velocities
To test the impulse approximation, we want to simulate interactions where the relative velocity ${\bf w}$ is changed by a factor of $\lambda$: ${\bf w} \rightarrow \lambda {\bf w}$. We start by computing the relative velocity for the impacts above and define a function that returns a dark-matter velocity after scaling the relative velocity by $\lambda$:
```
v_gc= numpy.array([xv_prog_impact[3],xv_prog_impact[4],xv_prog_impact[5]])
v_dm= numpy.array([6.82200571,132.7700529,149.4174464])
w_base= v_dm-v_gc
def v_dm_scaled(lam):
return w_base*lam+v_gc
```
## Starting 0.25 time units ago, scaled down by 0.5
```
lam= 0.5
xv_dm_impact= numpy.array([-13.500000,2.840000,-1.840000,v_dm_scaled(lam)[0],v_dm_scaled(lam)[1],v_dm_scaled(lam)[2]])
RvR_dm_impact= rectangular_to_cylindrical(xv_dm_impact[:,numpy.newaxis].T)[0,:]
dm_impact= Orbit([RvR_dm_impact[0]/R0,RvR_dm_impact[1]/V0,RvR_dm_impact[2]/V0,
RvR_dm_impact[3]/R0,RvR_dm_impact[4]/V0,RvR_dm_impact[5]],ro=R0,vo=V0)
dm_impact= dm_impact.flip()
ts= numpy.linspace(0.,0.25*0.9777922212082034/bovy_conversion.time_in_Gyr(V0,R0),10001)
dm_impact.integrate(ts,lp)
print [dm_impact.x(ts[-1]),dm_impact.y(ts[-1]),dm_impact.z(ts[-1]),
-dm_impact.vx(ts[-1]),-dm_impact.vy(ts[-1]),-dm_impact.vz(ts[-1])]
```
| github_jupyter |
# Growth models
This notebook focuses on the development and exploration of growth models and their properties.
```
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import curve_fit
import pandas as pd
# Logistic
def model(ts: np.ndarray, mu: float, K: float, c0: float, lag: float) -> np.ndarray:
return np.array([K / (1. + (K-c0)/c0*np.exp(mu * (lag-t))) if t > lag else c0 for t in ts])
# Gompertz
def model(ts: np.ndarray, mu: float, K: float, c0: float, lag: float) -> np.ndarray:
return np.array([K*np.exp(np.log(c0/K)*np.exp(mu * (lag-t))) if t > lag else c0 for t in ts])
r_true = 0.015
K_true = 2
c0_true = 0.002
lag_true = 200
sig = 0.05
n = 101
ts = np.linspace(0, 1200, n)
xs = model(ts, r_true, K_true, c0_true, lag_true)
ys = xs * (1 + sig*np.random.randn(n))
plt.scatter(ts, xs)
mle, cov = curve_fit(model, ts, ys, p0 = [0.02, 2, 0.01, 100])
r = mle[0]
K = mle[1]
c0 = mle[2]
lag = mle[3]
df = pd.DataFrame(mle, columns=['MLE'])
df.insert(0, 'Names', ['r', 'K', 'c0', 'lag'])
print(df)
ax = plt.subplot()
ax.scatter(ts, ys, c='k', s=2, label='Data')
ax.plot(ts, model(ts, r, K, c0, lag), c='r', label='Model')
ax.plot(ts, model(ts, r, K, c0 / 100, lag), c='b', label='Model (smaller c0)')
ax.plot(ts, model(ts, r, K, c0 * 100, lag), c='g', label='Model (larger c0)')
plt.legend()
```
Try fitting {r, K, c0} with lag fixed. As you vary lag, the optimizer just finds a different value for c0 that makes the model fit the data just as well.
```
lag = 100.0
def objective1(t, r, K, c0):
return model(t, r, K, c0, lag)
mle, cov = curve_fit(objective1, ts, ys, p0 = [0.02, 2, 0.01], bounds=([0,0,0], [0.1,3,0.1]))
r = mle[0]
K = mle[1]
c0 = mle[2]
df = pd.DataFrame(mle, columns=['MLE'])
df.insert(0, 'Names', ['r', 'K', 'c0'])
print(df)
ax = plt.subplot()
ax.scatter(ts, ys, c='k', s=2, label='Data')
ax.plot(ts, model(ts, r, K, c0, lag), c='r', label='Model')
plt.legend()
```
Save the data
```
X = np.stack([ts, ys]).T
np.savetxt('logistic_example_1.csv', X, delimiter=',')
```
Evaluate the objective function over a patch of values in the 2d subspace spanned by c0 and lag.
```
n = 151
c0s = np.logspace(-5, -1, n)
lags = np.linspace(0, 300, n)
err = np.zeros((n, n))
for i, c0 in enumerate(c0s):
for j, lag in enumerate(lags):
err[i][j] = np.linalg.norm(ys - model(ts, r_true, K_true, c0, lag))
```
Plot the surface, and compare with the true optimum
```
plt.style.use("dark_background")
plt.contourf(c0s, lags, err, 20)
plt.scatter([c0_true], [lag_true], c='r', label='True parameters')
plt.xscale('log')
plt.xlabel('c0')
plt.ylabel('lag')
plt.colorbar(label='Objective')
plt.legend()
```
| github_jupyter |
```
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
```
# Bi-LSTM Component
```
class BidirectionalLSTM(nn.Module):
def __init__(self, nIn, nHidden, nOut):
super(BidirectionalLSTM, self).__init__()
self.rnn = nn.LSTM(nIn, nHidden, bidirectional=True)
self.embedding = nn.Linear(nHidden * 2, nOut)
def forward(self, input):
recurrent, _ = self.rnn(input)
T, b, h = recurrent.size()
t_rec = recurrent.view(T * b, h)
output = self.embedding(t_rec) # [T * b, nOut]
output = output.view(T, b, -1)
return output
```
# R-CNN Component
```
class R_CNN(nn.Module):
def __init__(self):
super(R_CNN, self).__init__()
in_nc = 3
nf = 64
hdn = 300
nclass = 23 #dekhabet class
self.convs = nn.Sequential(
nn.Conv2d(in_nc, nf, 3, 1, 1),
nn.LeakyReLU(0.2, True),
nn.MaxPool2d(2, 2), #64 filters, 32*64
nn.Conv2d(nf, nf*2, 3, 1, 1),
nn.LeakyReLU(0.2, True),
nn.MaxPool2d(2, 2), #128 filters, 16*32
nn.Conv2d(nf*2, nf*4, 3, 1, 1),
nn.BatchNorm2d(nf*4),
nn.Conv2d(nf*4, nf*4, 3, 1, 1),
nn.LeakyReLU(0.2, True),
nn.MaxPool2d(2,2), #256 filters, 40*16
nn.Conv2d(nf*4, nf*4, 3, 1, 1),
nn.LeakyReLU(0.2, True),
nn.MaxPool2d((2, 2)),
nn.Conv2d(nf*4, nf*8, 3, 1, 1),
nn.BatchNorm2d(nf*8),
nn.Conv2d(nf*8, nf*8, 3, 1, 1),
nn.LeakyReLU(0.2, True),
nn.MaxPool2d((2, 1)),
nn.Conv2d(nf*8, nf*8, 3, 1, 1),
nn.LeakyReLU(0.2, True),
nn.MaxPool2d((2, 1)),
nn.Conv2d(nf*8, nf*8, 2, 1, 0),
)
self.bilstm = nn.Sequential(
BidirectionalLSTM(nf*8, hdn, hdn),
BidirectionalLSTM(hdn, hdn, nclass),
)
self.lgsftMx = nn.LogSoftmax(dim=2)
def forward(self, x):
out = self.convs(x)
out = out.squeeze(2)
out = out.permute(2, 0, 1) #ctc expects [width,batch,label]
out = self.bilstm(out)
out = F.log_softmax(out, dim=2)
return out
```
# Initiate Model And Loss
```
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
model = R_CNN()
model = model.to(device)
criterion = nn.CTCLoss().to(device)
optimizer = torch.optim.Adam(model.parameters(), lr=0.0001, betas=(0.5, 0.999))
```
# Getting The Data
```
input = torch.randn(5, 3, 128, 256)
input = input.to(device)
target = [[5,1,5,3,0,2,1,7,20,11],
[5,1,5,3,0,2,1,7,20,11],
[5,1,5,3,0,2,1,7,20,11],
[5,1,5,3,0,2,1,7,20,11],
[5,1,5,3,0,2,1,7,20,11]]
target = torch.FloatTensor(target)
target = target.to(device)
T = 15 # Input sequence length
C = 22 # Number of classes (including blank)
N = 5 # Batch size
S = 9 # Target sequence length of longest target in batch
S_min = 2 # Minimum target length, for demonstration purposes
# target = torch.randint(low=1, high=23, size=(N, S), dtype=torch.long)
target_lengths = torch.randint(low=S_min, high=S, size=(N,))
# input = torch.randn(T, N, C).log_softmax(2).detach().requires_grad_()
print(target)
print(target_lengths)
```
# Training
```
pred = model(input)
print(pred.shape)
preds_size = Variable(torch.LongTensor([pred.size(0)] * 5))
cost = criterion(pred, target, preds_size, target_lengths)
print(cost)
total_step = len(trainloader_pixel)
ctc_loss_list = []
acc_list = []
batch_size= 25
num_epochs = 2500
for epoch in range(num_epochs):
trainiter = iter(trainiter)
for i in range(5):
spectros, lbls, lbl_lens = trainIter_pixel.next()
spectros = spectos.to(device)
lbls = lbls.to(device)
lbl_lens.to(device)
pred = model(spectros)
preds_size = Variable(torch.LongTensor([pred.size(0)] * batch_size))
cost = criterion(pred, lbls, preds_size, lbl_lens)/batch_size
#backprop and optimize!
cost.backward()
optimizer.step()
if (epoch+1) % 100 == 0:
print('Epoch No [{}/{}] {:.4f}'.format(epoch+1,num_epochs,d_loss.item()))
ctc_loss_list.append(d_loss.item())
if (epoch+1) % 1000 == 0:
print('Epoch No {} reached saving model'.format(epoch+1)
torch.save(model.state_dict(), 'outputModel/KDNet_epoch_{}.pkl'.format(epoch+1))
```
| github_jupyter |
# PyTorch Tutorial #01
# Simple Linear Model
## Introduction
This tutorial demonstrates the basic workflow of using PyTorch with a simple linear model. After loading the so-called MNIST data-set with images of hand-written digits, we define and optimize a simple mathematical model in PyTorch. The results are then plotted and discussed.
You should be familiar with basic linear algebra, Python and the Jupyter Notebook editor. It also helps if you have a basic understanding of Machine Learning and classification.
## Imports
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from sklearn.metrics import confusion_matrix
from tqdm import tqdm_notebook,trange
import torch
from torch.nn import Parameter
from torch import nn
from torch.nn import functional as F
from torchvision import datasets, transforms
from torchvision.datasets import MNIST
from torch.utils.data import DataLoader
from torch.optim import SGD
torch.__version__
```
## Load Data
For loading and preprocessing data we will using a Library called `torchvison`
It will automatically download and preprocess the images. Preprocessing is done based on the `transform_func`.
Transforms are common image transforms. They can be chained together using `Compose`
Here we are Applying 2 transformation to images
1. Reading and Converting images into Pytorch Tensors
2. Normalise the images based on the mean and std
```
# Transformer function for image preprocessing
transforms_func = transforms.Compose([transforms.ToTensor(),transforms.Normalize((0.1307,), (0.3081,))])
# mnist train_set
mnist_train = MNIST('./data',train=True,download=True,transform=transforms_func)
# mnist test_set
mnist_test = MNIST('./data',train=False,transform=transforms_func)
```
## Creating a validation set
Here we are spliting the training data to `mnist_train` and `mnist_valid`
A validation set can be used to check the model is overfittig or not on th e training set and accordingly stop training process or save intermediate model weights
```
train_len = int(0.9*mnist_train.__len__())
valid_len = mnist_train.__len__() - train_len
mnist_train, mnist_valid = torch.utils.data.random_split(mnist_train, lengths=[train_len, valid_len])
```
The MNIST data-set has now been loaded and consists of 70.000 images and class-numbers for the images. The data-set is split into 3 mutually exclusive sub-sets. We will only use the training and test-sets in this tutorial.
```
print("Size of:")
print("- Training-set:\t\t{}".format(mnist_train.__len__()))
print("- Validation-set:\t{}".format(mnist_valid.__len__()))
print("- Test-set:\t\t{}".format(mnist_test.__len__()))
# The images are stored in one-dimensional arrays of this length.
img_size_flat = 784 # 28 x 28
# Tuple with height and width of images used to reshape arrays.
img_shape = (28,28)
# Number of classes, one class for each of 10 digits.
num_classes = 10
```
## Helper-function for plotting images
Function used to plot 9 images in a 3x3 grid, and writing the true and predicted classes below each image.
```
def plot_images(images, cls_true, cls_pred=None):
assert len(images) == len(cls_true) == 9
# Create figure with 3x3 sub-plots.
fig, axes = plt.subplots(3, 3)
fig.subplots_adjust(hspace=0.3, wspace=0.3)
for i, ax in enumerate(axes.flat):
# Plot image.
ax.imshow(images[i].reshape(img_shape), cmap='binary')
# Show true and predicted classes.
if cls_pred is None:
xlabel = "True: {0}".format(cls_true[i])
else:
xlabel = "True: {0}, Pred: {1}".format(cls_true[i], cls_pred[i])
ax.set_xlabel(xlabel)
# Remove ticks from the plot.
ax.set_xticks([])
ax.set_yticks([])
# Ensure the plot is shown correctly with multiple plots
# in a single Notebook cell.
plt.show()
```
## Plot a few images to see if data is correct
```
# Get the first images from the test-set.
images = mnist_test.test_data[0:9]
# Get the true classes for those images.
cls_true = mnist_test.test_labels[0:9]
# Plot the images and labels using our helper-function above.
plot_images(images=images, cls_true=cls_true)
```
# PyTorch
PyTorch is a Python package that provides two high-level features:
1. Tensor computation (like NumPy) with strong GPU acceleration
2. Deep neural networks built on a tape-based autograd system
You can reuse your favorite Python packages such as NumPy, SciPy and Cython to extend PyTorch when needed.
## More about PyTorch
At a granular level, PyTorch is a library that consists of the following components:
| Component | Description |
| ---- | --- |
| **torch** | a Tensor library like NumPy, with strong GPU support |
| **torch.autograd** | a tape-based automatic differentiation library that supports all differentiable Tensor operations in torch |
| **torch.nn** | a neural networks library deeply integrated with autograd designed for maximum flexibility |
| **torch.multiprocessing** | Python multiprocessing, but with magical memory sharing of torch Tensors across processes. Useful for data loading and Hogwild training |
| **torch.utils** | DataLoader, Trainer and other utility functions for convenience |
| **torch.legacy(.nn/.optim)** | legacy code that has been ported over from torch for backward compatibility reasons |
Usually one uses PyTorch either as:
- a replacement for NumPy to use the power of GPUs.
- a deep learning research platform that provides maximum flexibility and speed
### Dynamic Neural Networks: Tape-Based Autograd
PyTorch has a unique way of building neural networks: using and replaying a tape recorder.
Most frameworks such as TensorFlow, Theano, Caffe and CNTK have a static view of the world.
One has to build a neural network, and reuse the same structure again and again.
Changing the way the network behaves means that one has to start from scratch.
With PyTorch, we use a technique called reverse-mode auto-differentiation, which allows you to
change the way your network behaves arbitrarily with zero lag or overhead. Our inspiration comes
from several research papers on this topic, as well as current and past work such as
[torch-autograd](https://github.com/twitter/torch-autograd),
[autograd](https://github.com/HIPS/autograd),
[Chainer](http://chainer.org), etc.
While this technique is not unique to PyTorch, it's one of the fastest implementations of it to date.
You get the best of speed and flexibility for your crazy research.
# Recommended Reading
https://pytorch.org/tutorials/beginner/blitz/tensor_tutorial.html
https://pytorch.org/tutorials/beginner/blitz/autograd_tutorial.html
# Neural Network
Neural networks can be constructed using the torch.nn package.
A typical training procedure for a neural network is as follows:
- Define the neural network that has some learnable parameters (or weights)
- Iterate over a dataset of inputs
- Process input through the network
- Compute the loss (how far is the output from being correct)
- Propagate gradients back into the network’s parameters
- Update the weights of the network, typically using a simple update rule:
`weight = weight - learning_rate * gradient`
# Model
We will define networks as a subclass of nn.Module
You just have to define the forward function, and the backward function (where gradients are computed) is automatically defined for you using autograd. You can use any of the Tensor operations in the forward function.
The netowork we will be definning here will be containing a single Linear Layer.
The linear layer will contain 2 variables `weights` an `bias`, That must be changed by PyTorch so as to make the model perform better on the traning data
This simple mathematical model multiplies the images variable x with the weights and then adds the biases.
The result is a matrix of shape [num_images, num_classes] because x has shape [num_images, img_size_flat] and weights has shape [img_size_flat, num_classes], so the multiplication of those two matrices is a matrix with shape [num_images, num_classes] and then the biases vector is added to each row of that matrix.
### __init__( ) function
1. The first variable that must be optimized is called weights and is defined here as a Pytorch variable that must be initialized with zeros and whose shape is `[img_size_flat, num_classes]`, so it is a 2-dimensional tensor (or matrix) with img_size_flat rows and num_classes columns.
2. The second variable that must be optimized is called biases and is defined as a 1-dimensional tensor (or vector) of length num_classes.
3. Here in the __init__() function both weight and bias Tensors wrapped in as `Parameter`.Parameters are `Tensor` subclasses, that have a very special property when used with Module s - when they’re assigned as Module attributes they are automatically added to the list of its parameters, and will appear e.g. in `parameters()` iterator.
### froward( ) function
1. The forward function will taking input of shape `[num_images, img_size_flat]` and multiplying with `weight` of shape `[num_images, num_classes]` and the `bias` is added to final result. We can use torch built in func call `torch.addmm()` for performing the above operation.It is similar to `tf.nn.xw_plus_b` in `Tensorflow`
2. The `out` in first line of `forward()` will have shape `[num_images, num_classes]`
3. However, these estimates are a bit rough and difficult to interpret because the numbers may be very small or large, so we want to normalize them so that each row of the `out` matrix sums to one, and each element is limited between zero and one. This is calculated using the so-called softmax function and the result is stored in `out` it self.
```N.B The module `softmax` doesn’t work directly with NLLLoss, which expects the Log to be computed between the Softmax and itself. we will be Using LogSoftmax instead (it’s faster and has better numerical properties).```
Softmax has takes input Matrix and `dim`, A dimension along which Softmax will be computed (so every slice along dim will sum to 1).
### get_weights( )
1. This Function used to get the weights of the Network, To plot it understand what the model actually learned
```
class LinearModel(nn.Module):
def __init__(self):
super(LinearModel, self).__init__()
self.weight = Parameter(torch.zeros((784, 10),dtype=torch.float32,requires_grad=True))
self.bias = Parameter(torch.zeros((10),dtype=torch.float32,requires_grad=True))
def get_weights(self):
return self.weight
def forward(self,x):
out = torch.addmm(self.bias, x, self.weight)
out = F.log_softmax(out,dim=1)
return out
```
# Training Network
Function to train the Network
- Input:
- model : model object
- device : `cpu` or `cuda`
- train_loader : Data loader. Combines a dataset and a sampler, and provides single- or multi-process iterators over the dataset.
- optimizer : the function we are going to use for adjusting model parametrs
### Step by step
1. `model.train()`tells your model that you are training the model. So effectively layers like dropout, batchnorm etc. which behave different on the train and test procedures know what is going on and hence can behave accordingly.
2. `for loop` it will iterate through train_loader , it will give you 2 output `data` and `target`. The size of `data` and `target` will be depend on the `batch_size` that you have provided while creating the `DataLoader` function for `train dataset` . `data` shape `[batch_size,row,columns]`,`target` shape `[batch_size]`
3. Here the `data` will be of shape `[batch_size,row,columns]` ,we will convert `data` in to `[batch_size,row x columns]` (Flattening the images to fit into Linear layer)
4. Moving the the `data` and `target` to devices based on our choice and machine specifictaion.
5. By calling `optimizer.zero_grad` we will set Gradient buffers to zero.Else the gradients will get accumulated to existing gradients.
6. Input the `data` to `model` and get outputs, The output will be of shape `[batch_size,num_classes]`
7. `Loss function` : A loss function takes the (output, target) pair of inputs, and computes a value that estimates how far away the output is from the target . We will be using The negative log likelihood loss. It is useful to train a classification problem with C classes.
`nll_loss` function calculate the difference between the `output` to `target` , `output` of shape `[batch_size,num_classes]` and `target` of shape `[batch_size]` where each value is `0 ≤ targets[i] ≤ num_classes−1`
8. when we call `loss.backward()`, the whole graph is differentiated w.r.t. the loss, and all Tensors in the graph that has `requires_grad=True` will have their `.grad` Tensor accumulated with the gradient.
9. To backpropagate the error all we have to do is to `loss.backward()`
10. `optimizer.step()` is performs a parameter update based on the current gradient (stored in .grad attribute of a parameter) and the update rule.
```
def train(model,device,train_loader,optimizer):
model.train()
correct = 0
for data,target in tqdm_notebook(train_loader,total=train_loader.__len__()):
data = torch.reshape(data,(-1,784))
data, target = data.to(device), target.to(device)
optimizer.zero_grad()
output = model(data)
loss = F.nll_loss(output, target)
loss.backward()
optimizer.step()
pred = output.max(1, keepdim=True)[1]
correct += pred.eq(target.view_as(pred)).sum().item()
print('Accuracy: {}/{} ({:.0f}%)\n'.format(correct, len(train_loader.dataset),100. * correct / len(train_loader.dataset)))
```
# Testing Network
Function to test the Network
- Input:
- model : model object
- device : `cpu` or `cuda`
- test_loader : Data loader. Combines a dataset and a sampler, and provides single- or multi-process iterators over the dataset.
### Step by step
1. `model.eval()` tells your model that you are testing the model.so that Regularization Layers like `Droupout` and `BatchNormalization` get disbaled ,which behave different on the train and test procedures.
2. The wrapper `with torch.no_grad()` temporarily set all the requires_grad flag to false. Because we don't need to compute gradinets while performaing inference on the network and It will reduce memory usage and speed up computations
3. The nex three line are same as Train function
4. we will calculate the test_loss across the complete dataset
5. we will also calcualte the `accuracy` of model by comapring with target
```
def test(model, device, test_loader):
model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for i,(data, target) in tqdm_notebook(enumerate(test_loader),total=test_loader.__len__()):
data, target = data.to(device), target.to(device)
data = torch.reshape(data,(-1,784))
output = model(data)
test_loss += F.nll_loss(output, target, reduction='sum').item() # sum up batch loss
pred = output.max(1, keepdim=True)[1] # get the index of the max log-probability
correct += pred.eq(target.view_as(pred)).sum().item()
test_loss /= len(test_loader.dataset)
print('\nTest set: Average loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)\n'.format(
test_loss, correct, len(test_loader.dataset),
100. * correct / len(test_loader.dataset)))
```
## Function to get predicted class for all the test set
```
def get_incorrect_samples(model, device, test_loader):
model.eval()
prediction = []
correct = []
with torch.no_grad():
for i,(data, target) in tqdm_notebook(enumerate(test_loader),total=test_loader.__len__()):
data, target = data.to(device), target.to(device)
correct.extend(target)
data = torch.reshape(data,(-1,784))
output = model(data)
pred = output.max(1, keepdim=True)[1]
prediction.extend(pred)
out = torch.tensor(test_loader.dataset.test_labels).eq(torch.tensor(prediction).view_as(test_loader.dataset.test_labels))
return out.numpy(),np.asarray(prediction)
```
### Checking Device Availabilty
```
device = "cuda" if torch.cuda.is_available() else "cpu"
device
```
## DataLoader function for Training Set and Test Set
Data loader. Combines a dataset and a sampler, and provides single- or multi-process iterators over the dataset.
```
kwargs = {'num_workers': 1, 'pin_memory': True} if device=='cuda' else {}
train_loader = DataLoader(mnist_train,batch_size=64,shuffle=True,**kwargs)
test_loader = DataLoader(mnist_test,batch_size=1024,shuffle=False,**kwargs)
```
# Model
Model object is created and Transfered to `device` accoring to availabilty of `GPU`
```
model = LinearModel().to(device)
```
## Optimizer
We will be using `stochastic gradient descent|(SGD)` optimizer
`SGD` takes `model parameters` the we want to optimze and a learning_rate `lr` in which the model parametrs get updated
```
optimizer = SGD(model.parameters(),lr=0.5)
```
model parameters contains `weight` and `bias` that we defined inside our model , So while training the `weight` and `bias` will get updated according to our training set
```
list(model.parameters())
```
## Training
`epoch` is 10 .So in training the model will get iterated through the complete Training set 10 times .
After each epoch we will run `test` and check how well the model is performing on unknown data
If the Training Accuracy is going Up and Testing Accuracy is going down we can say that model is overfitting on the train set.
There are Techinques like `EarlyStopping` and usage of validatoin datset that we will discuss later
```
epochs = 2
for epoch in range(epochs):
train(model,device,train_loader,optimizer)
test(model,device,test_loader)
```
## Helper function to Plot Weights of the model
```
def plot_weights():
# Get the values for the weights from the TensorFlow variable.
w = model.get_weights()
w = w.detach().numpy()
img_shape = (28,28)
# Get the lowest and highest values for the weights.
# This is used to correct the colour intensity across
# the images so they can be compared with each other.
w_min = np.min(w)
w_max = np.max(w)
# Create figure with 3x4 sub-plots,
# where the last 2 sub-plots are unused.
fig, axes = plt.subplots(3, 4)
fig.subplots_adjust(hspace=0.3, wspace=0.3)
for i, ax in enumerate(axes.flat):
# Only use the weights for the first 10 sub-plots.
if i<10:
# Get the weights for the i'th digit and reshape it.
# Note that w.shape == (img_size_flat, 10)
image = w[:, i].reshape(img_shape)
# Set the label for the sub-plot.
ax.set_xlabel("Weights: {0}".format(i))
# Plot the image.
ax.imshow(image, vmin=w_min, vmax=w_max, cmap='seismic')
# Remove ticks from each sub-plot.
ax.set_xticks([])
ax.set_yticks([])
# Ensure the plot is shown correctly with multiple plots
# in a single Notebook cell.
plt.show()
plot_weights()
def plot_example_errors():
# Use TensorFlow to get a list of boolean values
# whether each test-image has been correctly classified,
# and a list for the predicted class of each image.
#correct, cls_pred = session.run([correct_prediction, y_pred_cls],
#feed_dict=feed_dict_test)
# Negate the boolean array.
correct,cls_pred = get_incorrect_samples(model,'cpu',test_loader)
incorrect = [i for i,x in enumerate(correct) if x==0]
# Get the images from the test-set that have been
# incorrectly classified.
images = mnist_test.test_data.numpy()[incorrect]
# import pdb;pdb.set_trace()
# Get the predicted classes for those images.
cls_pred = cls_pred[incorrect]
# Get the true classes for those images.
cls_true = mnist_test.test_labels.numpy()[incorrect]
# Plot the first 9 images.
plot_images(images=images[0:9],
cls_true=cls_true[0:9],
cls_pred=cls_pred[0:9])
plot_example_errors()
```
## Helper function to Print Confusion Matrix
```
def print_confusion_matrix():
# Get the true classifications for the test-set.
cls_true = test_loader.dataset.test_labels
# Get the predicted classifications for the test-set.
# cls_pred = session.run(y_pred_cls, feed_dict=feed_dict_test)
_,cls_pred = get_incorrect_samples(model,'cpu',test_loader)
# Get the confusion matrix using sklearn.
cm = confusion_matrix(y_true=cls_true,
y_pred=cls_pred)
# Print the confusion matrix as text.
print(cm)
# Plot the confusion matrix as an image.
plt.imshow(cm, interpolation='nearest', cmap=plt.cm.Blues)
# Make various adjustments to the plot.
plt.tight_layout()
plt.colorbar()
tick_marks = np.arange(num_classes)
plt.xticks(tick_marks, range(num_classes))
plt.yticks(tick_marks, range(num_classes))
plt.xlabel('Predicted')
plt.ylabel('True')
# Ensure the plot is shown correctly with multiple plots
# in a single Notebook cell.
plt.show()
print_confusion_matrix()
```
## Exercises
These are a few suggestions for exercises that may help improve your skills with PyTorch. It is important to get hands-on experience with PyTorch in order to learn how to use it properly.
You may want to backup this Notebook before making any changes.
- Change the learning-rate for the optimizer.
- Change the optimizer to e.g. AdagradOptimizer or AdamOptimizer.
- Change the batch-size to e.g. 1 or 1000.
- How do these changes affect the performance?
- Do you think these changes will have the same effect (if any) on other classification problems and mathematical models?
- Do you get the exact same results if you run the Notebook multiple times without changing any parameters? Why or why not?
- Change the function plot_example_errors() so it also prints the logits and y_pred values for the mis-classified examples.
- Use F.CrossEntropyLoss() instead of F.nll_loss. This may require several changes to multiple places in the source-code. Discuss the advantages and disadvantages of using the two methods.
- Remake the program yourself without looking too much at this source-code.
- Explain to a friend how the program works.
| github_jupyter |
## Dependencies
```
import json, warnings, shutil, glob
from jigsaw_utility_scripts import *
from scripts_step_lr_schedulers import *
from transformers import TFXLMRobertaModel, XLMRobertaConfig
from tensorflow.keras.models import Model
from tensorflow.keras import optimizers, metrics, losses, layers
SEED = 0
seed_everything(SEED)
warnings.filterwarnings("ignore")
```
## TPU configuration
```
strategy, tpu = set_up_strategy()
print("REPLICAS: ", strategy.num_replicas_in_sync)
AUTO = tf.data.experimental.AUTOTUNE
```
# Load data
```
database_base_path = '/kaggle/input/jigsaw-data-split-roberta-192-ratio-2-upper/'
k_fold = pd.read_csv(database_base_path + '5-fold.csv')
valid_df = pd.read_csv("/kaggle/input/jigsaw-multilingual-toxic-comment-classification/validation.csv",
usecols=['comment_text', 'toxic', 'lang'])
print('Train samples: %d' % len(k_fold))
display(k_fold.head())
print('Validation samples: %d' % len(valid_df))
display(valid_df.head())
base_data_path = 'fold_1/'
# Unzip files
!tar -xvf /kaggle/input/jigsaw-data-split-roberta-192-ratio-2-upper/fold_1.tar.gz
```
# Model parameters
```
base_path = '/kaggle/input/jigsaw-transformers/XLM-RoBERTa/'
config = {
"MAX_LEN": 192,
"BATCH_SIZE": 128,
"EPOCHS": 5,
"LEARNING_RATE": 1e-5,
"ES_PATIENCE": None,
"base_model_path": base_path + 'tf-xlm-roberta-large-tf_model.h5',
"config_path": base_path + 'xlm-roberta-large-config.json'
}
with open('config.json', 'w') as json_file:
json.dump(json.loads(json.dumps(config)), json_file)
```
## Learning rate schedule
```
lr_min = 1e-7
lr_max = config['LEARNING_RATE']
step_size = len(k_fold[k_fold['fold_1'] == 'train']) // config['BATCH_SIZE']
total_steps = config['EPOCHS'] * step_size
warmup_steps = step_size * 1
decay = .9997
rng = [i for i in range(0, total_steps, config['BATCH_SIZE'])]
y = [exponential_schedule_with_warmup(tf.cast(x, tf.float32), warmup_steps=warmup_steps,
lr_start=lr_min, lr_max=lr_max, lr_min=lr_min, decay=decay) for x in rng]
fig, ax = plt.subplots(figsize=(20, 6))
plt.plot(rng, y)
print("Learning rate schedule: {:.3g} to {:.3g} to {:.3g}".format(y[0], max(y), y[-1]))
```
# Model
```
module_config = XLMRobertaConfig.from_pretrained(config['config_path'], output_hidden_states=False)
def model_fn(MAX_LEN):
input_ids = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='input_ids')
attention_mask = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='attention_mask')
base_model = TFXLMRobertaModel.from_pretrained(config['base_model_path'], config=module_config)
last_hidden_state, _ = base_model({'input_ids': input_ids, 'attention_mask': attention_mask})
x_avg = layers.GlobalAveragePooling1D()(last_hidden_state)
x_max = layers.GlobalMaxPooling1D()(last_hidden_state)
x = layers.Concatenate()([x_avg, x_max])
x = layers.Dropout(0.3)(x)
output = layers.Dense(1, activation='sigmoid', name='output')(x)
model = Model(inputs=[input_ids, attention_mask], outputs=output)
return model
```
# Train
```
# Load data
x_train = np.load(base_data_path + 'x_train.npy')
y_train = np.load(base_data_path + 'y_train_int.npy').reshape(x_train.shape[1], 1).astype(np.float32)
x_valid_ml = np.load(database_base_path + 'x_valid.npy')
y_valid_ml = np.load(database_base_path + 'y_valid.npy').reshape(x_valid_ml.shape[1], 1).astype(np.float32)
#################### ADD TAIL ####################
# x_train = np.hstack([x_train, np.load(base_data_path + 'x_train_tail.npy')])
# y_train = np.vstack([y_train, y_train])
step_size = x_train.shape[1] // config['BATCH_SIZE']
valid_step_size = x_valid_ml.shape[1] // config['BATCH_SIZE']
# Build TF datasets
train_dist_ds = strategy.experimental_distribute_dataset(get_training_dataset(x_train, y_train, config['BATCH_SIZE'], AUTO, seed=SEED))
valid_dist_ds = strategy.experimental_distribute_dataset(get_validation_dataset(x_valid_ml, y_valid_ml, config['BATCH_SIZE'], AUTO, repeated=True, seed=SEED))
train_data_iter = iter(train_dist_ds)
valid_data_iter = iter(valid_dist_ds)
# Step functions
@tf.function
def train_step(data_iter):
def train_step_fn(x, y):
with tf.GradientTape() as tape:
probabilities = model(x, training=True)
loss = loss_fn(y, probabilities)
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
train_auc.update_state(y, probabilities)
train_loss.update_state(loss)
for _ in tf.range(step_size):
strategy.experimental_run_v2(train_step_fn, next(data_iter))
@tf.function
def valid_step(data_iter):
def valid_step_fn(x, y):
probabilities = model(x, training=False)
loss = loss_fn(y, probabilities)
valid_auc.update_state(y, probabilities)
valid_loss.update_state(loss)
for _ in tf.range(valid_step_size):
strategy.experimental_run_v2(valid_step_fn, next(data_iter))
# Train model
with strategy.scope():
model = model_fn(config['MAX_LEN'])
optimizer = optimizers.Adam(learning_rate=lambda:
exponential_schedule_with_warmup(tf.cast(optimizer.iterations, tf.float32),
warmup_steps=warmup_steps, lr_start=lr_min,
lr_max=lr_max, lr_min=lr_min, decay=decay))
loss_fn = losses.binary_crossentropy
train_auc = metrics.AUC()
valid_auc = metrics.AUC()
train_loss = metrics.Sum()
valid_loss = metrics.Sum()
metrics_dict = {'loss': train_loss, 'auc': train_auc,
'val_loss': valid_loss, 'val_auc': valid_auc}
history = custom_fit(model, metrics_dict, train_step, valid_step, train_data_iter, valid_data_iter,
step_size, valid_step_size, config['BATCH_SIZE'], config['EPOCHS'],
config['ES_PATIENCE'], save_last=True)
# Make predictions
# x_train = np.load(base_data_path + 'x_train.npy')
# x_valid = np.load(base_data_path + 'x_valid.npy')
x_valid_ml = np.load(database_base_path + 'x_valid.npy')
# train_preds = model.predict(get_test_dataset(x_train, config['BATCH_SIZE'], AUTO))
# valid_preds = model.predict(get_test_dataset(x_valid, config['BATCH_SIZE'], AUTO))
valid_ml_preds = model.predict(get_test_dataset(x_valid_ml, config['BATCH_SIZE'], AUTO))
# k_fold.loc[k_fold['fold_1'] == 'train', 'pred_1'] = np.round(train_preds)
# k_fold.loc[k_fold['fold_1'] == 'validation', 'pred_1'] = np.round(valid_preds)
valid_df['pred_1'] = valid_ml_preds
### Delete data dir
shutil.rmtree(base_data_path)
```
## Model loss graph
```
sns.set(style="whitegrid")
plot_metrics(history)
```
# Model evaluation
```
# display(evaluate_model(k_fold, 1, label_col='toxic_int').style.applymap(color_map))
```
# Confusion matrix
```
# train_set = k_fold[k_fold['fold_1'] == 'train']
# validation_set = k_fold[k_fold['fold_1'] == 'validation']
# plot_confusion_matrix(train_set['toxic_int'], train_set['pred_1'],
# validation_set['toxic_int'], validation_set['pred_1'])
```
# Model evaluation by language
```
display(evaluate_model_lang(valid_df, 1).style.applymap(color_map))
```
# Visualize predictions
```
pd.set_option('max_colwidth', 120)
print('English validation set')
display(k_fold[['comment_text', 'toxic'] + [c for c in k_fold.columns if c.startswith('pred')]].head(10))
print('Multilingual validation set')
display(valid_df[['comment_text', 'toxic'] + [c for c in valid_df.columns if c.startswith('pred')]].head(10))
```
# Test set predictions
```
x_test = np.load(database_base_path + 'x_test.npy')
test_preds = model.predict(get_test_dataset(x_test, config['BATCH_SIZE'], AUTO))
submission = pd.read_csv('/kaggle/input/jigsaw-multilingual-toxic-comment-classification/sample_submission.csv')
submission['toxic'] = test_preds
submission.to_csv('submission.csv', index=False)
display(submission.describe())
display(submission.head(10))
```
| github_jupyter |
# Some common distributions to know
```
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
%matplotlib inline
```
## Discrete distributions
The binomial distribution
$$f(k|n,\theta) = \binom{n}{k}\theta^k(1-\theta)^{n-k}$$
e.g. Toss a coin n times
```
from scipy.stats import binom
n,theta = 100, 0.5
mean, var, skew, kurt = binom.stats(n, theta, moments='mvsk')
fig, ax = plt.subplots(1, 1)
x = np.arange(binom.ppf(0.01, n, theta), binom.ppf(0.99, n, theta))
ax.vlines(x, 0, binom.pmf(x, n, theta), colors='b', lw=5, alpha=0.5)
plt.ylabel('Mass')
plt.xlabel('k')
plt.xlim(25,75)
plt.ylim(0.0 ,.1)
```
The bernoulli distribution
\begin{align*}
f(x|\theta) =
\begin{cases}
\theta & \text{if $x=1$} \\
1-\theta & \text{if $x=0$}
\end{cases}
\end{align*}
e.g. Toss a coin once
```
from scipy.stats import bernoulli
theta = 0.5
mean, var, skew, kurt = bernoulli.stats(theta, moments='mvsk')
fig, ax = plt.subplots(1, 1)
x = np.arange(0,1.1)
ax.vlines(x, 0, bernoulli.pmf(x, theta), colors='b', lw=5, alpha=0.5)
plt.ylabel('Mass')
plt.xlabel('x')
plt.xlim(-0.1 ,1.1)
plt.ylim(0.0 ,1)
#from scipy.stats import multinomial $ waiting until scipy 0.19 is released
x = np.arange(0,6)
theta = [1/6, 1/6, 1/6, 1/6, 1/6, 1/6]
n = 100 # number of trials
mean, var, skew, kurt = multinomial.stats(theta, moments='mvsk')
fig, ax = plt.subplots(1, 1)
ax.vlines(x, 0, multinomial.pmf(x, theta), colors='b', lw=5, alpha=0.5)
plt.ylabel('Mass')
plt.xlabel('x')
plt.xlim(-0.1 ,1.1)
plt.ylim(0.0 ,1)
```
The poisson distribution
$$f(x|\theta) = e^{-\lambda} \frac{\lambda^x}{x!}$$
e.g. rare events, radioactive decay, Trump saying something coherent
```
from scipy.stats import poisson
lambda_ = 0.6
mean, var, skew, kurt = poisson.stats(lambda_, moments='mvsk')
fig, ax = plt.subplots(1, 1)
x = np.arange(0,3)
ax.vlines(x, 0, poisson.pmf(x, lambda_), colors='b', lw=5, alpha=0.5)
plt.ylabel('Mass')
plt.xlabel('x')
plt.xlim(-0.1 ,3)
plt.ylim(0.0 ,1)
```
The emperical distribution
$$f(A) = \frac{1}{N}\sum^{N}_{i=1}\delta_{x_{i}}(A)$$
\begin{align*}
\delta_{x_{i}}(A) =
\begin{cases}
0 & \text{if $x\notin A$} \\
1 & \text{if $x\in A$}
\end{cases}
\end{align*}
## Continuous Distributions
Gaussian (normal)
$$f(x|\mu, \sigma^2) = \frac{1}{\sqrt{2\pi\sigma^2}}e^{-\frac{1}{2\sigma^2}(x-\mu)^2}$$
```
from scipy.stats import norm
fig, ax = plt.subplots(1, 1)
mean, var, skew, kurt = norm.stats(moments='mvsk')
x = np.linspace(norm.ppf(0.01),norm.ppf(0.99), 100)
ax.plot(x, norm.pdf(x),
'b-', lw=3, alpha=0.6, label='norm pdf')
plt.xlim(-3 ,3)
plt.ylim(0.0 ,1)
plt.ylabel('Density')
plt.xlabel('x')
```
Students t (special cases cauchy lorentz)
$$ f(x|v) = \frac{\Gamma(\frac{v+1}{2})}{\sqrt{v\pi}\Gamma(\frac{v}{2})} \Big( 1+\frac{x^2}{v} \Big)^{-\frac{v+1}{2}}, v= df $$
e.g. scienceing
```
from scipy.stats import t
fig, ax = plt.subplots(1, 1)
df = 3
mean, var, skew, kurt = t.stats(df, moments='mvsk')
x = np.linspace(t.ppf(0.01, df),
t.ppf(0.99, df), 100)
ax.plot(x, t.pdf(x, df),
'b-', lw=3, alpha=0.6, label='t pdf')
plt.ylabel('Density')
plt.xlabel('x')
```
Laplace
$$ f(x|\mu,b) = \frac{1}{2b}e^{\big(-\frac{|x-\mu|)}{b}\big)}$$
e.g. like normal but with more sparsity, brownian motion
```
from scipy.stats import laplace
mean, var, skew, kurt = laplace.stats(moments='mvsk')
fig, ax = plt.subplots(1, 1)
x = np.linspace(laplace.ppf(0.01),
laplace.ppf(0.99), 100)
ax.plot(x, laplace.pdf(x),
'b-', lw=3, alpha=0.6, label='laplace pdf')
plt.ylabel('Density')
plt.xlabel('x')
```
Gamma
$$ f(x|a,b) = \frac{b^a}{\Gamma(a)}x^{a-1}e^{-xb} $$, where the shape a >0, and the rate b >0
e.g.
```
from scipy.stats import gamma
fig, ax = plt.subplots(1, 1)
a = 2
mean, var, skew, kurt = gamma.stats(a, moments='mvsk')
x = np.linspace(gamma.ppf(0.01, a),
gamma.ppf(0.99, a), 100)
ax.plot(x, gamma.pdf(x, a),
'b-', lw=3, alpha=0.6, label='gamma pdf')
plt.ylabel('Density')
plt.xlabel('x')
```
The beta distribution
$$f(x|a,b) = \frac{1}{B(a,b)}x^{a-1}(1-x)^{b-1}$$
$$B(a,b) = \frac{\Gamma(a)\Gamma(b)}{\Gamma(a+b)}$$
```
from scipy.stats import beta
a, b = 2, 0.8
mean, var, skew, kurt = beta.stats(a, b, moments='mvsk')
fig, ax = plt.subplots(1, 1)
x = np.linspace(beta.ppf(0.01, a, b),
beta.ppf(0.99, a, b), 100)
ax.plot(x, beta.pdf(x, a, b),
'b-', lw=3, alpha=0.6, label='beta pdf')
plt.ylabel('Density')
plt.xlabel('x')
```
pareto
\begin{align*}
f(x| k,m) =
\begin{cases}
\frac{kx_m^k}{x^{k+1}} & \text{if $x \ge x_m$} \\
0 & \text{if $x < x_m$}
\end{cases}
\end{align*}
```
from scipy.stats import pareto
fig, ax = plt.subplots(1, 1)
b = 2.62
mean, var, skew, kurt = pareto.stats(b, moments='mvsk')
x = np.linspace(pareto.ppf(0.01, b),
pareto.ppf(0.99, b), 100)
ax.plot(x, pareto.pdf(x, b),
'b-', lw=3, alpha=0.6, label='pareto pdf')
plt.ylabel('Density')
plt.xlabel('x')
```
The multivariate Gaussian
$$f(x|\mu, \Sigma) = \frac{1}{(2\pi)^{d/2}|\Sigma|^{1/2}}e^{\frac{1}{2}(x-\mu)^T\Sigma^{-1}(x-\mu)}$$
```
from scipy.stats import multivariate_normal
mean, cov = [0, 1], [(1, .5), (.5, 1)]
x, y = np.random.multivariate_normal(mean, cov, 1000).T
with sns.axes_style("white"):
sns.jointplot(x=x, y=y, kind="hex", color="b");
```
The Dirichlet distribution
$$f(x|\alpha) = \frac{1}{B(\alpha)}\prod_{k=1}^{K}x_k^{\alpha_k-1}I(x\in S_k)$$
$$B(\alpha) = \frac{\prod_{k=1}^{K}\Gamma(\alpha_k)}{\Gamma(\alpha_0)}$$
e.g. multivariate generalization of beta distribution
```
#The code below to visualize was taken from Thomas boggs elegant contours here:http://blog.bogatron.net/blog/2014/02/02/visualizing-dirichlet-distributions/
import matplotlib.tri as tri
_corners = np.array([[0, 0], [1, 0], [0.5, 0.75**0.5]])
_triangle = tri.Triangulation(_corners[:, 0], _corners[:, 1])
_midpoints = [(_corners[(i + 1) % 3] + _corners[(i + 2) % 3]) / 2.0 \
for i in range(3)]
def xy2bc(xy, tol=1.e-3):
'''Converts 2D Cartesian coordinates to barycentric.
Arguments:
`xy`: A length-2 sequence containing the x and y value.
'''
s = [(_corners[i] - _midpoints[i]).dot(xy - _midpoints[i]) / 0.75 \
for i in range(3)]
return np.clip(s, tol, 1.0 - tol)
class Dirichlet(object):
def __init__(self, alpha):
'''Creates Dirichlet distribution with parameter `alpha`.'''
from math import gamma
from operator import mul
self._alpha = np.array(alpha)
self._coef = gamma(np.sum(self._alpha)) / \
reduce(mul, [gamma(a) for a in self._alpha])
def pdf(self, x):
'''Returns pdf value for `x`.'''
from operator import mul
return self._coef * reduce(mul, [xx ** (aa - 1)
for (xx, aa)in zip(x, self._alpha)])
def sample(self, N):
'''Generates a random sample of size `N`.'''
return np.random.dirichlet(self._alpha, N)
def draw_pdf_contours(dist, border=False, nlevels=200, subdiv=8, **kwargs):
'''Draws pdf contours over an equilateral triangle (2-simplex).
Arguments:
`dist`: A distribution instance with a `pdf` method.
`border` (bool): If True, the simplex border is drawn.
`nlevels` (int): Number of contours to draw.
`subdiv` (int): Number of recursive mesh subdivisions to create.
kwargs: Keyword args passed on to `plt.triplot`.
'''
from matplotlib import ticker, cm
import math
refiner = tri.UniformTriRefiner(_triangle)
trimesh = refiner.refine_triangulation(subdiv=subdiv)
pvals = [dist.pdf(xy2bc(xy)) for xy in zip(trimesh.x, trimesh.y)]
plt.tricontourf(trimesh, pvals, nlevels, **kwargs)
plt.axis('equal')
plt.xlim(0, 1)
plt.ylim(0, 0.75**0.5)
plt.axis('off')
if border is True:
plt.hold(1)
plt.triplot(_triangle, linewidth=1)
def plot_points(X, barycentric=True, border=True, **kwargs):
'''Plots a set of points in the simplex.
Arguments:
`X` (ndarray): A 2xN array (if in Cartesian coords) or 3xN array
(if in barycentric coords) of points to plot.
`barycentric` (bool): Indicates if `X` is in barycentric coords.
`border` (bool): If True, the simplex border is drawn.
kwargs: Keyword args passed on to `plt.plot`.
'''
if barycentric is True:
X = X.dot(_corners)
plt.plot(X[:, 0], X[:, 1], 'k.', ms=1, **kwargs)
plt.axis('equal')
plt.xlim(0, 1)
plt.ylim(0, 0.75**0.5)
plt.axis('off')
if border is True:
plt.hold(1)
plt.triplot(_triangle, linewidth=1)
if __name__ == '__main__':
f = plt.figure(figsize=(8, 6))
alphas = [[0.999] * 3,
[5] * 3,
[2, 5, 15]]
for (i, alpha) in enumerate(alphas):
plt.subplot(2, len(alphas), i + 1)
dist = Dirichlet(alpha)
draw_pdf_contours(dist)
title = r'$\alpha$ = (%.3f, %.3f, %.3f)' % tuple(alpha)
plt.title(title, fontdict={'fontsize': 8})
plt.subplot(2, len(alphas), i + 1 + len(alphas))
plot_points(dist.sample(5000))
print 'Wrote plots to "dirichlet_plots.png".'
draw_pdf_contours(Dirichlet([5, 5, 5]))
```
The multinomial distribution
$$f(x|n,\theta) = \binom{n}{x_1 \ldots x_K}\prod^{K}_{j=1}\theta^{x_j}_j$$
e.g. Roll a K-sided die n times
| github_jupyter |
<div class="alert alert-block alert-info" style="margin-top: 20px">
<a href="https://cocl.us/corsera_da0101en_notebook_top">
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DA0101EN/Images/TopAd.png" width="750" align="center">
</a>
</div>
<a href="https://www.bigdatauniversity.com"><img src = "https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DA0101EN/Images/CCLog.png" width = 300, align = "center"></a>
<h1 align=center><font size = 5>Data Analysis with Python</font></h1>
Exploratory Data Analysis
<h3>Welcome!</h3>
In this section, we will explore several methods to see if certain characteristics or features can be used to predict car price.
<h2>Table of content</h2>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<ol>
<li><a href="#import_data">Import Data from Module</a></li>
<li><a href="#pattern_visualization">Analyzing Individual Feature Patterns using Visualization</a></li>
<li><a href="#discriptive_statistics">Descriptive Statistical Analysis</a></li>
<li><a href="#basic_grouping">Basics of Grouping</a></li>
<li><a href="#correlation_causation">Correlation and Causation</a></li>
<li><a href="#anova">ANOVA</a></li>
</ol>
Estimated Time Needed: <strong>30 min</strong>
</div>
<hr>
<h3>What are the main characteristics which have the most impact on the car price?</h3>
<h2 id="import_data">1. Import Data from Module 2</h2>
<h4>Setup</h4>
Import libraries
```
import pandas as pd
import numpy as np
```
load data and store in dataframe df:
This dataset was hosted on IBM Cloud object click <a href="https://cocl.us/DA101EN_object_storage">HERE</a> for free storage
```
path='https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DA0101EN-SkillsNetwork/labs/Data%20files/automobileEDA.csv'
df = pd.read_csv(path)
df.head()
```
<h2 id="pattern_visualization">2. Analyzing Individual Feature Patterns using Visualization</h2>
To install seaborn we use the pip which is the python package manager.
```
%%capture
! pip install seaborn
```
Import visualization packages "Matplotlib" and "Seaborn", don't forget about "%matplotlib inline" to plot in a Jupyter notebook.
```
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
```
<h4>How to choose the right visualization method?</h4>
<p>When visualizing individual variables, it is important to first understand what type of variable you are dealing with. This will help us find the right visualization method for that variable.</p>
```
# list the data types for each column
print(df['peak-rpm'].dtypes)
```
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h3>Question #1:</h3>
<b>What is the data type of the column "peak-rpm"? </b>
</div>
Double-click <b>here</b> for the solution.
<!-- The answer is below:
float64
-->
for example, we can calculate the correlation between variables of type "int64" or "float64" using the method "corr":
```
df.corr()
```
The diagonal elements are always one; we will study correlation more precisely Pearson correlation in-depth at the end of the notebook.
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h1> Question #2: </h1>
<p>Find the correlation between the following columns: bore, stroke,compression-ratio , and horsepower.</p>
<p>Hint: if you would like to select those columns use the following syntax: df[['bore','stroke' ,'compression-ratio','horsepower']]</p>
</div>
```
# Write your code below and press Shift+Enter to execute
corr_1=df[['bore', 'stroke', 'compression-ratio', 'horsepower']].corr()
corr_1
```
Double-click <b>here</b> for the solution.
<!-- The answer is below:
df[['bore', 'stroke', 'compression-ratio', 'horsepower']].corr()
-->
<h2>Continuous numerical variables:</h2>
<p>Continuous numerical variables are variables that may contain any value within some range. Continuous numerical variables can have the type "int64" or "float64". A great way to visualize these variables is by using scatterplots with fitted lines.</p>
<p>In order to start understanding the (linear) relationship between an individual variable and the price. We can do this by using "regplot", which plots the scatterplot plus the fitted regression line for the data.</p>
Let's see several examples of different linear relationships:
<h4>Positive linear relationship</h4>
Let's find the scatterplot of "engine-size" and "price"
```
# Engine size as potential predictor variable of price
sns.regplot(x="engine-size", y="price", data=df)
plt.ylim(0,)
```
<p>As the engine-size goes up, the price goes up: this indicates a positive direct correlation between these two variables. Engine size seems like a pretty good predictor of price since the regression line is almost a perfect diagonal line.</p>
We can examine the correlation between 'engine-size' and 'price' and see it's approximately 0.87
```
df[["engine-size", "price"]].corr()
```
Highway mpg is a potential predictor variable of price
```
u=sns.regplot(x="highway-mpg", y="price", data=df, x_estimator=np.mean)
print(u)
from scipy import stats
slope, intercept, r_value, p_value, std_err = stats.linregress(df['highway-mpg'], df['price'] )
print(slope, intercept, r_value, p_value, std_err)
```
<p>As the highway-mpg goes up, the price goes down: this indicates an inverse/negative relationship between these two variables. Highway mpg could potentially be a predictor of price.</p>
We can examine the correlation between 'highway-mpg' and 'price' and see it's approximately -0.704
```
df[['highway-mpg', 'price']].corr()
```
<h3>Weak Linear Relationship</h3>
Let's see if "Peak-rpm" as a predictor variable of "price".
```
sns.regplot(x="peak-rpm", y="price", data=df)
stats.linregress(df["peak-rpm"],df["price"])
```
<p>Peak rpm does not seem like a good predictor of the price at all since the regression line is close to horizontal. Also, the data points are very scattered and far from the fitted line, showing lots of variability. Therefore it's it is not a reliable variable.</p>
We can examine the correlation between 'peak-rpm' and 'price' and see it's approximately -0.101616
```
df[['peak-rpm','price']].corr()
```
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h1> Question 3 a): </h1>
<p>Find the correlation between x="stroke", y="price".</p>
<p>Hint: if you would like to select those columns use the following syntax: df[["stroke","price"]] </p>
</div>
```
# Write your code below and press Shift+Enter to execute
df[["stroke", "price"]].corr()
```
Double-click <b>here</b> for the solution.
<!-- The answer is below:
#The correlation is 0.0823, the non-diagonal elements of the table.
#code:
df[["stroke","price"]].corr()
-->
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h1>Question 3 b):</h1>
<p>Given the correlation results between "price" and "stroke" do you expect a linear relationship?</p>
<p>Verify your results using the function "regplot()".</p>
</div>
```
# Write your code below and press Shift+Enter to execute
sns.regplot(x="price", y="stroke", data=df)
```
Double-click <b>here</b> for the solution.
<!-- The answer is below:
#There is a weak correlation between the variable 'stroke' and 'price.' as such regression will not work well. We #can see this use "regplot" to demonstrate this.
#Code:
sns.regplot(x="stroke", y="price", data=df)
-->
<h3>Categorical variables</h3>
<p>These are variables that describe a 'characteristic' of a data unit, and are selected from a small group of categories. The categorical variables can have the type "object" or "int64". A good way to visualize categorical variables is by using boxplots.</p>
Let's look at the relationship between "body-style" and "price".
```
sns.boxplot(x="body-style", y="price", data=df)
```
<p>We see that the distributions of price between the different body-style categories have a significant overlap, and so body-style would not be a good predictor of price. Let's examine engine "engine-location" and "price":</p>
```
sns.boxplot(x="engine-location", y="price", data=df)
```
<p>Here we see that the distribution of price between these two engine-location categories, front and rear, are distinct enough to take engine-location as a potential good predictor of price.</p>
Let's examine "drive-wheels" and "price".
```
# drive-wheels
sns.boxplot(x="drive-wheels", y="price", data=df)
```
<p>Here we see that the distribution of price between the different drive-wheels categories differs; as such drive-wheels could potentially be a predictor of price.</p>
<h2 id="discriptive_statistics">3. Descriptive Statistical Analysis</h2>
<p>Let's first take a look at the variables by utilizing a description method.</p>
<p>The <b>describe</b> function automatically computes basic statistics for all continuous variables. Any NaN values are automatically skipped in these statistics.</p>
This will show:
<ul>
<li>the count of that variable</li>
<li>the mean</li>
<li>the standard deviation (std)</li>
<li>the minimum value</li>
<li>the IQR (Interquartile Range: 25%, 50% and 75%)</li>
<li>the maximum value</li>
<ul>
We can apply the method "describe" as follows:
```
df.describe()
```
The default setting of "describe" skips variables of type object. We can apply the method "describe" on the variables of type 'object' as follows:
```
df.describe(include=['object'])
```
<h3>Value Counts</h3>
<p>Value-counts is a good way of understanding how many units of each characteristic/variable we have. We can apply the "value_counts" method on the column 'drive-wheels'. Don’t forget the method "value_counts" only works on Pandas series, not Pandas Dataframes. As a result, we only include one bracket "df['drive-wheels']" not two brackets "df[['drive-wheels']]".</p>
```
df['drive-wheels'].value_counts()
```
We can convert the series to a Dataframe as follows :
```
df['drive-wheels'].value_counts().to_frame()
```
Let's repeat the above steps but save the results to the dataframe "drive_wheels_counts" and rename the column 'drive-wheels' to 'value_counts'.
```
drive_wheels_counts = df['drive-wheels'].value_counts().to_frame()
drive_wheels_counts.rename(columns={'drive-wheels': 'value_counts'}, inplace=True)
drive_wheels_counts
```
Now let's rename the index to 'drive-wheels':
```
drive_wheels_counts.index.name = 'drive-wheels'
drive_wheels_counts
```
We can repeat the above process for the variable 'engine-location'.
```
# engine-location as variable
engine_loc_counts = df['engine-location'].value_counts().to_frame()
engine_loc_counts.rename(columns={'engine-location': 'value_counts'}, inplace=True)
engine_loc_counts.index.name = 'engine-location'
engine_loc_counts.head(10)
```
<p>Examining the value counts of the engine location would not be a good predictor variable for the price. This is because we only have three cars with a rear engine and 198 with an engine in the front, this result is skewed. Thus, we are not able to draw any conclusions about the engine location.</p>
<h2 id="basic_grouping">4. Basics of Grouping</h2>
<p>The "groupby" method groups data by different categories. The data is grouped based on one or several variables and analysis is performed on the individual groups.</p>
<p>For example, let's group by the variable "drive-wheels". We see that there are 3 different categories of drive wheels.</p>
```
df['drive-wheels'].unique()
```
<p>If we want to know, on average, which type of drive wheel is most valuable, we can group "drive-wheels" and then average them.</p>
<p>We can select the columns 'drive-wheels', 'body-style' and 'price', then assign it to the variable "df_group_one".</p>
```
df_group_one = df[['drive-wheels','body-style','price']]
```
We can then calculate the average price for each of the different categories of data.
```
# grouping results
df_group_one = df_group_one.groupby(['drive-wheels'],as_index=False).mean()
df_group_one
```
<p>From our data, it seems rear-wheel drive vehicles are, on average, the most expensive, while 4-wheel and front-wheel are approximately the same in price.</p>
<p>You can also group with multiple variables. For example, let's group by both 'drive-wheels' and 'body-style'. This groups the dataframe by the unique combinations 'drive-wheels' and 'body-style'. We can store the results in the variable 'grouped_test1'.</p>
```
# grouping results
df_gptest = df[['drive-wheels','body-style','price']]
grouped_test1 = df_gptest.groupby(['drive-wheels','body-style'],as_index=False).mean()
grouped_test1
```
<p>This grouped data is much easier to visualize when it is made into a pivot table. A pivot table is like an Excel spreadsheet, with one variable along the column and another along the row. We can convert the dataframe to a pivot table using the method "pivot " to create a pivot table from the groups.</p>
<p>In this case, we will leave the drive-wheel variable as the rows of the table, and pivot body-style to become the columns of the table:</p>
```
grouped_pivot = grouped_test1.pivot(index='drive-wheels',columns='body-style')
grouped_pivot
```
<p>Often, we won't have data for some of the pivot cells. We can fill these missing cells with the value 0, but any other value could potentially be used as well. It should be mentioned that missing data is quite a complex subject and is an entire course on its own.</p>
```
grouped_pivot = grouped_pivot.fillna(0) #fill missing values with 0
grouped_pivot
```
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h1>Question 4:</h1>
<p>Use the "groupby" function to find the average "price" of each car based on "body-style" ? </p>
</div>
```
# Write your code below and press Shift+Enter to execute
df_bs=df[['body-style','price']]
avg_bs=df_bs.groupby(['body-style'], as_index=False).mean()
avg_bs.head()
```
Double-click <b>here</b> for the solution.
<!-- The answer is below:
# grouping results
df_gptest2 = df[['body-style','price']]
grouped_test_bodystyle = df_gptest2.groupby(['body-style'],as_index= False).mean()
grouped_test_bodystyle
-->
If you did not import "pyplot" let's do it again.
```
import matplotlib.pyplot as plt
%matplotlib inline
```
<h4>Variables: Drive Wheels and Body Style vs Price</h4>
Let's use a heat map to visualize the relationship between Body Style vs Price.
```
#use the grouped results
plt.pcolor(grouped_pivot, cmap='RdBu')
plt.colorbar()
plt.show()
```
<p>The heatmap plots the target variable (price) proportional to colour with respect to the variables 'drive-wheel' and 'body-style' in the vertical and horizontal axis respectively. This allows us to visualize how the price is related to 'drive-wheel' and 'body-style'.</p>
<p>The default labels convey no useful information to us. Let's change that:</p>
```
fig, ax = plt.subplots()
im = ax.pcolor(grouped_pivot, cmap='RdBu')
#label names
row_labels = grouped_pivot.columns.levels[1]
col_labels = grouped_pivot.index
#move ticks and labels to the center
ax.set_xticks(np.arange(grouped_pivot.shape[1]) + 0.5, minor=False)
ax.set_yticks(np.arange(grouped_pivot.shape[0]) + 0.5, minor=False)
#insert labels
ax.set_xticklabels(row_labels, minor=False)
ax.set_yticklabels(col_labels, minor=False)
#rotate label if too long
plt.xticks(rotation=90)
fig.colorbar(im)
plt.show()
```
<p>Visualization is very important in data science, and Python visualization packages provide great freedom. We will go more in-depth in a separate Python Visualizations course.</p>
<p>The main question we want to answer in this module, is "What are the main characteristics which have the most impact on the car price?".</p>
<p>To get a better measure of the important characteristics, we look at the correlation of these variables with the car price, in other words: how is the car price dependent on this variable?</p>
<h2 id="correlation_causation">5. Correlation and Causation</h2>
<p><b>Correlation</b>: a measure of the extent of interdependence between variables.</p>
<p><b>Causation</b>: the relationship between cause and effect between two variables.</p>
<p>It is important to know the difference between these two and that correlation does not imply causation. Determining correlation is much simpler the determining causation as causation may require independent experimentation.</p>
<p3>Pearson Correlation</p>
<p>The Pearson Correlation measures the linear dependence between two variables X and Y.</p>
<p>The resulting coefficient is a value between -1 and 1 inclusive, where:</p>
<ul>
<li><b>1</b>: Total positive linear correlation.</li>
<li><b>0</b>: No linear correlation, the two variables most likely do not affect each other.</li>
<li><b>-1</b>: Total negative linear correlation.</li>
</ul>
<p>Pearson Correlation is the default method of the function "corr". Like before we can calculate the Pearson Correlation of the of the 'int64' or 'float64' variables.</p>
```
df.corr()
```
sometimes we would like to know the significant of the correlation estimate.
<b>P-value</b>:
<p>What is this P-value? The P-value is the probability value that the correlation between these two variables is statistically significant. Normally, we choose a significance level of 0.05, which means that we are 95% confident that the correlation between the variables is significant.</p>
By convention, when the
<ul>
<li>p-value is $<$ 0.001: we say there is strong evidence that the correlation is significant.</li>
<li>the p-value is $<$ 0.05: there is moderate evidence that the correlation is significant.</li>
<li>the p-value is $<$ 0.1: there is weak evidence that the correlation is significant.</li>
<li>the p-value is $>$ 0.1: there is no evidence that the correlation is significant.</li>
</ul>
We can obtain this information using "stats" module in the "scipy" library.
```
from scipy import stats
```
<h3>Wheel-base vs Price</h3>
Let's calculate the Pearson Correlation Coefficient and P-value of 'wheel-base' and 'price'.
```
pearson_coef, p_value = stats.pearsonr(df['wheel-base'], df['price'])
print("The Pearson Correlation Coefficient is", pearson_coef, " with a P-value of P =", p_value)
```
<h5>Conclusion:</h5>
<p>Since the p-value is $<$ 0.001, the correlation between wheel-base and price is statistically significant, although the linear relationship isn't extremely strong (~0.585)</p>
<h3>Horsepower vs Price</h3>
Let's calculate the Pearson Correlation Coefficient and P-value of 'horsepower' and 'price'.
```
pearson_coef, p_value = stats.pearsonr(df['horsepower'], df['price'])
print("The Pearson Correlation Coefficient is", pearson_coef, " with a P-value of P = ", p_value)
```
<h5>Conclusion:</h5>
<p>Since the p-value is $<$ 0.001, the correlation between horsepower and price is statistically significant, and the linear relationship is quite strong (~0.809, close to 1)</p>
<h3>Length vs Price</h3>
Let's calculate the Pearson Correlation Coefficient and P-value of 'length' and 'price'.
```
pearson_coef, p_value = stats.pearsonr(df['length'], df['price'])
print("The Pearson Correlation Coefficient is", pearson_coef, " with a P-value of P = ", p_value)
```
<h5>Conclusion:</h5>
<p>Since the p-value is $<$ 0.001, the correlation between length and price is statistically significant, and the linear relationship is moderately strong (~0.691).</p>
<h3>Width vs Price</h3>
Let's calculate the Pearson Correlation Coefficient and P-value of 'width' and 'price':
```
pearson_coef, p_value = stats.pearsonr(df['width'], df['price'])
print("The Pearson Correlation Coefficient is", pearson_coef, " with a P-value of P =", p_value )
```
##### Conclusion:
Since the p-value is < 0.001, the correlation between width and price is statistically significant, and the linear relationship is quite strong (~0.751).
### Curb-weight vs Price
Let's calculate the Pearson Correlation Coefficient and P-value of 'curb-weight' and 'price':
```
pearson_coef, p_value = stats.pearsonr(df['curb-weight'], df['price'])
print( "The Pearson Correlation Coefficient is", pearson_coef, " with a P-value of P = ", p_value)
```
<h5>Conclusion:</h5>
<p>Since the p-value is $<$ 0.001, the correlation between curb-weight and price is statistically significant, and the linear relationship is quite strong (~0.834).</p>
<h3>Engine-size vs Price</h3>
Let's calculate the Pearson Correlation Coefficient and P-value of 'engine-size' and 'price':
```
pearson_coef, p_value = stats.pearsonr(df['engine-size'], df['price'])
print("The Pearson Correlation Coefficient is", pearson_coef, " with a P-value of P =", p_value)
```
<h5>Conclusion:</h5>
<p>Since the p-value is $<$ 0.001, the correlation between engine-size and price is statistically significant, and the linear relationship is very strong (~0.872).</p>
<h3>Bore vs Price</h3>
Let's calculate the Pearson Correlation Coefficient and P-value of 'bore' and 'price':
```
pearson_coef, p_value = stats.pearsonr(df['bore'], df['price'])
print("The Pearson Correlation Coefficient is", pearson_coef, " with a P-value of P = ", p_value )
```
<h5>Conclusion:</h5>
<p>Since the p-value is $<$ 0.001, the correlation between bore and price is statistically significant, but the linear relationship is only moderate (~0.521).</p>
We can relate the process for each 'City-mpg' and 'Highway-mpg':
<h3>City-mpg vs Price</h3>
```
pearson_coef, p_value = stats.pearsonr(df['city-mpg'], df['price'])
print("The Pearson Correlation Coefficient is", pearson_coef, " with a P-value of P = ", p_value)
```
<h5>Conclusion:</h5>
<p>Since the p-value is $<$ 0.001, the correlation between city-mpg and price is statistically significant, and the coefficient of ~ -0.687 shows that the relationship is negative and moderately strong.</p>
<h3>Highway-mpg vs Price</h3>
```
pearson_coef, p_value = stats.pearsonr(df['highway-mpg'], df['price'])
print( "The Pearson Correlation Coefficient is", pearson_coef, " with a P-value of P = ", p_value )
```
##### Conclusion:
Since the p-value is < 0.001, the correlation between highway-mpg and price is statistically significant, and the coefficient of ~ -0.705 shows that the relationship is negative and moderately strong.
<h2 id="anova">6. ANOVA</h2>
<h3>ANOVA: Analysis of Variance</h3>
<p>The Analysis of Variance (ANOVA) is a statistical method used to test whether there are significant differences between the means of two or more groups. ANOVA returns two parameters:</p>
<p><b>F-test score</b>: ANOVA assumes the means of all groups are the same, calculates how much the actual means deviate from the assumption, and reports it as the F-test score. A larger score means there is a larger difference between the means.</p>
<p><b>P-value</b>: P-value tells how statistically significant is our calculated score value.</p>
<p>If our price variable is strongly correlated with the variable we are analyzing, expect ANOVA to return a sizeable F-test score and a small p-value.</p>
<h3>Drive Wheels</h3>
<p>Since ANOVA analyzes the difference between different groups of the same variable, the groupby function will come in handy. Because the ANOVA algorithm averages the data automatically, we do not need to take the average before hand.</p>
<p>Let's see if different types 'drive-wheels' impact 'price', we group the data.</p>
Let's see if different types 'drive-wheels' impact 'price', we group the data.
```
grouped_test2=df_gptest[['drive-wheels', 'price']].groupby(['drive-wheels'])
grouped_test2.head(2)
df_gptest
```
We can obtain the values of the method group using the method "get_group".
```
grouped_test2.get_group('4wd')['price']
```
we can use the function 'f_oneway' in the module 'stats' to obtain the <b>F-test score</b> and <b>P-value</b>.
```
# ANOVA
f_val, p_val = stats.f_oneway(grouped_test2.get_group('fwd')['price'], grouped_test2.get_group('rwd')['price'], grouped_test2.get_group('4wd')['price'])
print( "ANOVA results: F=", f_val, ", P =", p_val)
```
This is a great result, with a large F test score showing a strong correlation and a P value of almost 0 implying almost certain statistical significance. But does this mean all three tested groups are all this highly correlated?
#### Separately: fwd and rwd
```
f_val, p_val = stats.f_oneway(grouped_test2.get_group('fwd')['price'], grouped_test2.get_group('rwd')['price'])
print( "ANOVA results: F=", f_val, ", P =", p_val )
```
Let's examine the other groups
#### 4wd and rwd
```
f_val, p_val = stats.f_oneway(grouped_test2.get_group('4wd')['price'], grouped_test2.get_group('rwd')['price'])
print( "ANOVA results: F=", f_val, ", P =", p_val)
```
<h4>4wd and fwd</h4>
```
f_val, p_val = stats.f_oneway(grouped_test2.get_group('4wd')['price'], grouped_test2.get_group('fwd')['price'])
print("ANOVA results: F=", f_val, ", P =", p_val)
```
<h3>Conclusion: Important Variables</h3>
<p>We now have a better idea of what our data looks like and which variables are important to take into account when predicting the car price. We have narrowed it down to the following variables:</p>
Continuous numerical variables:
<ul>
<li>Length</li>
<li>Width</li>
<li>Curb-weight</li>
<li>Engine-size</li>
<li>Horsepower</li>
<li>City-mpg</li>
<li>Highway-mpg</li>
<li>Wheel-base</li>
<li>Bore</li>
</ul>
Categorical variables:
<ul>
<li>Drive-wheels</li>
</ul>
<p>As we now move into building machine learning models to automate our analysis, feeding the model with variables that meaningfully affect our target variable will improve our model's prediction performance.</p>
<h1>Thank you for completing this notebook</h1>
<div class="alert alert-block alert-info" style="margin-top: 20px">
```
<p><a href="https://cocl.us/corsera_da0101en_notebook_bottom"><img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DA0101EN/Images/BottomAd.png" width="750" align="center"></a></p>
```
</div>
<h3>About the Authors:</h3>
This notebook was written by <a href="https://www.linkedin.com/in/mahdi-noorian-58219234/" target="_blank">Mahdi Noorian PhD</a>, <a href="https://www.linkedin.com/in/joseph-s-50398b136/" target="_blank">Joseph Santarcangelo</a>, Bahare Talayian, Eric Xiao, Steven Dong, Parizad, Hima Vsudevan and <a href="https://www.linkedin.com/in/fiorellawever/" target="_blank">Fiorella Wenver</a> and <a href=" https://www.linkedin.com/in/yi-leng-yao-84451275/ " target="_blank" >Yi Yao</a>.
<p><a href="https://www.linkedin.com/in/joseph-s-50398b136/" target="_blank">Joseph Santarcangelo</a> is a Data Scientist at IBM, and holds a PhD in Electrical Engineering. His research focused on using Machine Learning, Signal Processing, and Computer Vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.</p>
| Date (YYYY-MM-DD) | Version | Changed By | Change Description |
| ----------------- | ------- | ---------- | --------------------- |
| 2020-07-29 | 0 | Nayef | Upload file to Gitlab |
| | | | |
<hr>
<p>Copyright © 2018 IBM Developer Skills Network. This notebook and its source code are released under the terms of the <a href="https://cognitiveclass.ai/mit-license/">MIT License</a>.</p>
| github_jupyter |
**Chapter 12 – Custom Models and Training with TensorFlow**
_This notebook contains all the sample code in chapter 12._
# Setup
First, let's import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures. We also check that Python 3.5 or later is installed (although Python 2.x may work, it is deprecated so we strongly recommend you use Python 3 instead), as well as Scikit-Learn ≥0.20 and TensorFlow ≥2.0-preview.
```
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
# TensorFlow ≥2.0-preview is required
import tensorflow as tf
from tensorflow import keras
assert tf.__version__ >= "2.0"
# Common imports
import numpy as np
import os
# to make this notebook's output stable across runs
np.random.seed(42)
# To plot pretty figures
%matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "deep"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
```
## Tensors and operations
### Tensors
```
tf.constant([[1., 2., 3.], [4., 5., 6.]]) # matrix
tf.constant(42) # scalar
t = tf.constant([[1., 2., 3.], [4., 5., 6.]])
t
t.shape
t.dtype
```
### Indexing
```
t[:, 1:]
t[..., 1, tf.newaxis]
```
### Ops
```
t + 10
tf.square(t)
t @ tf.transpose(t)
```
### Using `keras.backend`
```
from tensorflow import keras
K = keras.backend
K.square(K.transpose(t)) + 10
```
### From/To NumPy
```
a = np.array([2., 4., 5.])
tf.constant(a)
t.numpy()
np.array(t)
tf.square(a)
np.square(t)
```
### Conflicting Types
```
try:
tf.constant(2.0) + tf.constant(40)
except tf.errors.InvalidArgumentError as ex:
print(ex)
try:
tf.constant(2.0) + tf.constant(40., dtype=tf.float64)
except tf.errors.InvalidArgumentError as ex:
print(ex)
t2 = tf.constant(40., dtype=tf.float64)
tf.constant(2.0) + tf.cast(t2, tf.float32)
```
### Strings
```
tf.constant(b"hello world")
tf.constant("café")
u = tf.constant([ord(c) for c in "café"])
u
b = tf.strings.unicode_encode(u, "UTF-8")
tf.strings.length(b, unit="UTF8_CHAR")
tf.strings.unicode_decode(b, "UTF-8")
```
### String arrays
```
p = tf.constant(["Café", "Coffee", "caffè", "咖啡"])
tf.strings.length(p, unit="UTF8_CHAR")
r = tf.strings.unicode_decode(p, "UTF8")
r
print(r)
```
### Ragged tensors
```
print(r[1])
print(r[1:3])
r2 = tf.ragged.constant([[65, 66], [], [67]])
print(tf.concat([r, r2], axis=0))
r3 = tf.ragged.constant([[68, 69, 70], [71], [], [72, 73]])
print(tf.concat([r, r3], axis=1))
tf.strings.unicode_encode(r3, "UTF-8")
r.to_tensor()
```
### Sparse tensors
```
s = tf.SparseTensor(indices=[[0, 1], [1, 0], [2, 3]],
values=[1., 2., 3.],
dense_shape=[3, 4])
print(s)
tf.sparse.to_dense(s)
s2 = s * 2.0
try:
s3 = s + 1.
except TypeError as ex:
print(ex)
s4 = tf.constant([[10., 20.], [30., 40.], [50., 60.], [70., 80.]])
tf.sparse.sparse_dense_matmul(s, s4)
s5 = tf.SparseTensor(indices=[[0, 2], [0, 1]],
values=[1., 2.],
dense_shape=[3, 4])
print(s5)
try:
tf.sparse.to_dense(s5)
except tf.errors.InvalidArgumentError as ex:
print(ex)
s6 = tf.sparse.reorder(s5)
tf.sparse.to_dense(s6)
```
### Sets
```
set1 = tf.constant([[2, 3, 5, 7], [7, 9, 0, 0]])
set2 = tf.constant([[4, 5, 6], [9, 10, 0]])
tf.sparse.to_dense(tf.sets.union(set1, set2))
tf.sparse.to_dense(tf.sets.difference(set1, set2))
tf.sparse.to_dense(tf.sets.intersection(set1, set2))
```
### Variables
```
v = tf.Variable([[1., 2., 3.], [4., 5., 6.]])
v.assign(2 * v)
v[0, 1].assign(42)
v[:, 2].assign([0., 1.])
try:
v[1] = [7., 8., 9.]
except TypeError as ex:
print(ex)
v.scatter_nd_update(indices=[[0, 0], [1, 2]],
updates=[100., 200.])
sparse_delta = tf.IndexedSlices(values=[[1., 2., 3.], [4., 5., 6.]],
indices=[1, 0])
v.scatter_update(sparse_delta)
```
### Tensor Arrays
```
array = tf.TensorArray(dtype=tf.float32, size=3)
array = array.write(0, tf.constant([1., 2.]))
array = array.write(1, tf.constant([3., 10.]))
array = array.write(2, tf.constant([5., 7.]))
array.read(1)
array.stack()
mean, variance = tf.nn.moments(array.stack(), axes=0)
mean
variance
```
## Custom loss function
Let's start by loading and preparing the California housing dataset. We first load it, then split it into a training set, a validation set and a test set, and finally we scale it:
```
from sklearn.datasets import fetch_california_housing
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
housing = fetch_california_housing()
X_train_full, X_test, y_train_full, y_test = train_test_split(
housing.data, housing.target.reshape(-1, 1), random_state=42)
X_train, X_valid, y_train, y_valid = train_test_split(
X_train_full, y_train_full, random_state=42)
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_valid_scaled = scaler.transform(X_valid)
X_test_scaled = scaler.transform(X_test)
def huber_fn(y_true, y_pred):
error = y_true - y_pred
is_small_error = tf.abs(error) < 1
squared_loss = tf.square(error) / 2
linear_loss = tf.abs(error) - 0.5
return tf.where(is_small_error, squared_loss, linear_loss)
plt.figure(figsize=(8, 3.5))
z = np.linspace(-4, 4, 200)
plt.plot(z, huber_fn(0, z), "b-", linewidth=2, label="huber($z$)")
plt.plot(z, z**2 / 2, "b:", linewidth=1, label=r"$\frac{1}{2}z^2$")
plt.plot([-1, -1], [0, huber_fn(0., -1.)], "r--")
plt.plot([1, 1], [0, huber_fn(0., 1.)], "r--")
plt.gca().axhline(y=0, color='k')
plt.gca().axvline(x=0, color='k')
plt.axis([-4, 4, 0, 4])
plt.grid(True)
plt.xlabel("$z$")
plt.legend(fontsize=14)
plt.title("Huber loss", fontsize=14)
plt.show()
input_shape = X_train.shape[1:]
model = keras.models.Sequential([
keras.layers.Dense(30, activation="selu", kernel_initializer="lecun_normal",
input_shape=input_shape),
keras.layers.Dense(1),
])
model.compile(loss=huber_fn, optimizer="nadam", metrics=["mae"])
model.fit(X_train_scaled, y_train, epochs=2,
validation_data=(X_valid_scaled, y_valid))
```
## Saving/Loading Models with Custom Objects
```
model.save("my_model_with_a_custom_loss.h5")
model = keras.models.load_model("my_model_with_a_custom_loss.h5",
custom_objects={"huber_fn": huber_fn})
model.fit(X_train_scaled, y_train, epochs=2,
validation_data=(X_valid_scaled, y_valid))
def create_huber(threshold=1.0):
def huber_fn(y_true, y_pred):
error = y_true - y_pred
is_small_error = tf.abs(error) < threshold
squared_loss = tf.square(error) / 2
linear_loss = threshold * tf.abs(error) - threshold**2 / 2
return tf.where(is_small_error, squared_loss, linear_loss)
return huber_fn
model.compile(loss=create_huber(2.0), optimizer="nadam", metrics=["mae"])
model.fit(X_train_scaled, y_train, epochs=2,
validation_data=(X_valid_scaled, y_valid))
model.save("my_model_with_a_custom_loss_threshold_2.h5")
model = keras.models.load_model("my_model_with_a_custom_loss_threshold_2.h5",
custom_objects={"huber_fn": create_huber(2.0)})
model.fit(X_train_scaled, y_train, epochs=2,
validation_data=(X_valid_scaled, y_valid))
class HuberLoss(keras.losses.Loss):
def __init__(self, threshold=1.0, **kwargs):
self.threshold = threshold
super().__init__(**kwargs)
def call(self, y_true, y_pred):
error = y_true - y_pred
is_small_error = tf.abs(error) < self.threshold
squared_loss = tf.square(error) / 2
linear_loss = self.threshold * tf.abs(error) - self.threshold**2 / 2
return tf.where(is_small_error, squared_loss, linear_loss)
def get_config(self):
base_config = super().get_config()
return {**base_config, "threshold": self.threshold}
model = keras.models.Sequential([
keras.layers.Dense(30, activation="selu", kernel_initializer="lecun_normal",
input_shape=input_shape),
keras.layers.Dense(1),
])
model.compile(loss=HuberLoss(2.), optimizer="nadam", metrics=["mae"])
model.fit(X_train_scaled, y_train, epochs=2,
validation_data=(X_valid_scaled, y_valid))
model.save("my_model_with_a_custom_loss_class.h5")
#model = keras.models.load_model("my_model_with_a_custom_loss_class.h5", # TODO: check PR #25956
# custom_objects={"HuberLoss": HuberLoss})
model.fit(X_train_scaled, y_train, epochs=2,
validation_data=(X_valid_scaled, y_valid))
#model = keras.models.load_model("my_model_with_a_custom_loss_class.h5", # TODO: check PR #25956
# custom_objects={"HuberLoss": HuberLoss})
model.loss.threshold
```
## Other Custom Functions
```
def my_softplus(z): # return value is just tf.nn.softplus(z)
return tf.math.log(tf.exp(z) + 1.0)
def my_glorot_initializer(shape, dtype=tf.float32):
stddev = tf.sqrt(2. / (shape[0] + shape[1]))
return tf.random.normal(shape, stddev=stddev, dtype=dtype)
def my_l1_regularizer(weights):
return tf.reduce_sum(tf.abs(0.01 * weights))
def my_positive_weights(weights): # return value is just tf.nn.relu(weights)
return tf.where(weights < 0., tf.zeros_like(weights), weights)
layer = keras.layers.Dense(1, activation=my_softplus,
kernel_initializer=my_glorot_initializer,
kernel_regularizer=my_l1_regularizer,
kernel_constraint=my_positive_weights)
model = keras.models.Sequential([
keras.layers.Dense(30, activation="selu", kernel_initializer="lecun_normal",
input_shape=input_shape),
keras.layers.Dense(1, activation=my_softplus,
kernel_regularizer=my_l1_regularizer,
kernel_constraint=my_positive_weights,
kernel_initializer=my_glorot_initializer),
])
model.compile(loss="mse", optimizer="nadam", metrics=["mae"])
model.fit(X_train_scaled, y_train, epochs=2,
validation_data=(X_valid_scaled, y_valid))
model.save("my_model_with_many_custom_parts.h5")
# TODO:
"""
model = keras.models.load_model(
"my_model_with_many_custom_parts.h5",
custom_objects={
"my_l1_regularizer": my_l1_regularizer(0.01),
"my_positive_weights": my_positive_weights,
"my_glorot_initializer": my_glorot_initializer,
"my_softplus": my_softplus,
})
"""
class MyL1Regularizer(keras.regularizers.Regularizer):
def __init__(self, factor):
self.factor = factor
def __call__(self, weights):
return tf.reduce_sum(tf.abs(self.factor * weights))
def get_config(self):
return {"factor": self.factor}
model = keras.models.Sequential([
keras.layers.Dense(30, activation="selu", kernel_initializer="lecun_normal",
input_shape=input_shape),
keras.layers.Dense(1, activation=my_softplus,
kernel_regularizer=MyL1Regularizer(0.01),
kernel_constraint=my_positive_weights,
kernel_initializer=my_glorot_initializer),
])
model.compile(loss="mse", optimizer="nadam", metrics=["mae"])
model.fit(X_train_scaled, y_train, epochs=2,
validation_data=(X_valid_scaled, y_valid))
model.save("my_model_with_many_custom_parts.h5")
# TODO: check https://github.com/tensorflow/tensorflow/issues/26061
"""
model = keras.models.load_model(
"my_model_with_many_custom_parts.h5",
custom_objects={
"MyL1Regularizer": MyL1Regularizer,
"my_positive_weights": my_positive_weights,
"my_glorot_initializer": my_glorot_initializer,
"my_softplus": my_softplus,
})
"""
```
## Custom Metrics
```
model = keras.models.Sequential([
keras.layers.Dense(30, activation="selu", kernel_initializer="lecun_normal",
input_shape=input_shape),
keras.layers.Dense(1),
])
model.compile(loss="mse", optimizer="nadam", metrics=[create_huber(2.0)])
model.fit(X_train_scaled, y_train, epochs=2)
```
**Warning**: if you use the same function as the loss and a metric, you may be surprised to see different results. This is generally just due to floating point precision errors: even though the mathematical equations are equivalent, the operations are not run in the same order, which can lead to small differences. Moreover, when using sample weights, there's more than just precision errors:
* the loss since the start of the epoch is the mean of all batch losses seen so far. Each batch loss is the sum of the weighted instance losses divided by the _batch size_ (not the sum of weights, so the batch loss is _not_ the weighted mean of the losses).
* the metric since the start of the epoch is equal to the sum of weighted instance losses divided by sum of all weights seen so far. In other words, it is the weighted mean of all the instance losses. Not the same thing.
If you do the math, you will find that metric = loss * mean of sample weights (plus some floating point precision error).
```
model.compile(loss=create_huber(2.0), optimizer="nadam", metrics=[create_huber(2.0)])
sample_weight = np.random.rand(len(y_train))
history = model.fit(X_train_scaled, y_train, epochs=2, sample_weight=sample_weight)
history.history["loss"][0], history.history["huber_fn"][0] * sample_weight.mean()
```
### Streaming metrics
```
precision = keras.metrics.Precision()
precision([0, 1, 1, 1, 0, 1, 0, 1], [1, 1, 0, 1, 0, 1, 0, 1])
precision([0, 1, 0, 0, 1, 0, 1, 1], [1, 0, 1, 1, 0, 0, 0, 0])
precision.result()
precision.variables
precision.reset_states()
```
Creating a streaming metric:
```
class HuberMetric(keras.metrics.Metric):
def __init__(self, threshold=1.0, **kwargs):
super().__init__(**kwargs) # handles base args (e.g., dtype)
self.threshold = threshold
self.huber_fn = create_huber(threshold)
self.total = self.add_weight("total", initializer="zeros")
self.count = self.add_weight("count", initializer="zeros")
def update_state(self, y_true, y_pred, sample_weight=None):
metric = self.huber_fn(y_true, y_pred)
self.total.assign_add(tf.reduce_sum(metric))
self.count.assign_add(tf.cast(tf.size(y_true), tf.float32))
def result(self):
return self.total / self.count
def get_config(self):
base_config = super().get_config()
return {**base_config, "threshold": self.threshold}
m = HuberMetric(2.)
# total = 2 * |10 - 2| - 2²/2 = 14
# count = 1
# result = 14 / 1 = 14
m(tf.constant([[2.]]), tf.constant([[10.]]))
# total = total + (|1 - 0|² / 2) + (2 * |9.25 - 5| - 2² / 2) = 14 + 7 = 21
# count = count + 2 = 3
# result = total / count = 21 / 3 = 7
m(tf.constant([[0.], [5.]]), tf.constant([[1.], [9.25]]))
m.result()
m.variables
m.reset_states()
m.variables
```
Let's check that the `HuberMetric` class works well:
```
model = keras.models.Sequential([
keras.layers.Dense(30, activation="selu", kernel_initializer="lecun_normal",
input_shape=input_shape),
keras.layers.Dense(1),
])
model.compile(loss=create_huber(2.0), optimizer="nadam", metrics=[HuberMetric(2.0)])
model.fit(X_train_scaled, y_train, epochs=2)
model.save("my_model_with_a_custom_metric.h5")
#model = keras.models.load_model("my_model_with_a_custom_metric.h5", # TODO: check PR #25956
# custom_objects={"huber_fn": create_huber(2.0),
# "HuberMetric": HuberMetric})
model.fit(X_train_scaled, y_train, epochs=2)
model.metrics[0].threshold
```
Looks like it works fine! More simply, we could have created the class like this:
```
class HuberMetric(keras.metrics.Mean):
def __init__(self, threshold=1.0, name='HuberMetric', dtype=None):
self.threshold = threshold
self.huber_fn = create_huber(threshold)
super().__init__(name=name, dtype=dtype)
def update_state(self, y_true, y_pred, sample_weight=None):
metric = self.huber_fn(y_true, y_pred)
super(HuberMetric, self).update_state(metric, sample_weight)
def get_config(self):
base_config = super().get_config()
return {**base_config, "threshold": self.threshold}
```
This class handles shapes better, and it also supports sample weights.
```
model = keras.models.Sequential([
keras.layers.Dense(30, activation="selu", kernel_initializer="lecun_normal",
input_shape=input_shape),
keras.layers.Dense(1),
])
model.compile(loss=keras.losses.Huber(2.0), optimizer="nadam", weighted_metrics=[HuberMetric(2.0)])
sample_weight = np.random.rand(len(y_train))
history = model.fit(X_train_scaled, y_train, epochs=2, sample_weight=sample_weight)
history.history["loss"][0], history.history["HuberMetric"][0] * sample_weight.mean()
model.save("my_model_with_a_custom_metric_v2.h5")
#model = keras.models.load_model("my_model_with_a_custom_metric_v2.h5", # TODO: check PR #25956
# custom_objects={"HuberMetric": HuberMetric})
model.fit(X_train_scaled, y_train, epochs=2)
model.metrics[0].threshold
```
## Custom Layers
```
exponential_layer = keras.layers.Lambda(lambda x: tf.exp(x))
exponential_layer([-1., 0., 1.])
```
Adding an exponential layer at the output of a regression model can be useful if the values to predict are positive and with very different scales (e.g., 0.001, 10., 10000):
```
model = keras.models.Sequential([
keras.layers.Dense(30, activation="relu", input_shape=input_shape),
keras.layers.Dense(1),
exponential_layer
])
model.compile(loss="mse", optimizer="nadam")
model.fit(X_train_scaled, y_train, epochs=5,
validation_data=(X_valid_scaled, y_valid))
model.evaluate(X_test_scaled, y_test)
class MyDense(keras.layers.Layer):
def __init__(self, units, activation=None, **kwargs):
super().__init__(**kwargs)
self.units = units
self.activation = keras.activations.get(activation)
def build(self, batch_input_shape):
self.kernel = self.add_weight(
name="kernel", shape=[batch_input_shape[-1], self.units],
initializer="glorot_normal")
self.bias = self.add_weight(
name="bias", shape=[self.units], initializer="zeros")
super().build(batch_input_shape) # must be at the end
def call(self, X):
return self.activation(X @ self.kernel + self.bias)
def compute_output_shape(self, batch_input_shape):
return tf.TensorShape(batch_input_shape.as_list()[:-1] + [self.units])
def get_config(self):
base_config = super().get_config()
return {**base_config, "units": self.units,
"activation": keras.activations.serialize(self.activation)}
model = keras.models.Sequential([
MyDense(30, activation="relu", input_shape=input_shape),
MyDense(1)
])
model.compile(loss="mse", optimizer="nadam")
model.fit(X_train_scaled, y_train, epochs=2,
validation_data=(X_valid_scaled, y_valid))
model.evaluate(X_test_scaled, y_test)
model.save("my_model_with_a_custom_layer.h5")
model = keras.models.load_model("my_model_with_a_custom_layer.h5",
custom_objects={"MyDense": MyDense})
class MyMultiLayer(keras.layers.Layer):
def call(self, X):
X1, X2 = X
return X1 + X2, X1 * X2
def compute_output_shape(self, batch_input_shape):
batch_input_shape1, batch_input_shape2 = batch_input_shape
return [batch_input_shape1, batch_input_shape2]
inputs1 = keras.layers.Input(shape=[2])
inputs2 = keras.layers.Input(shape=[2])
outputs1, outputs2 = MyMultiLayer()((inputs1, inputs2))
```
Let's create a layer with a different behavior during training and testing:
```
class AddGaussianNoise(keras.layers.Layer):
def __init__(self, stddev, **kwargs):
super().__init__(**kwargs)
self.stddev = stddev
def call(self, X, training=None):
if training is None:
training = keras.backend.learning_phase()
if training:
noise = tf.random.normal(tf.shape(X), stddev=self.stddev)
return X + noise
else:
return X
def compute_output_shape(self, batch_input_shape):
return batch_input_shape
model.compile(loss="mse", optimizer="nadam")
model.fit(X_train_scaled, y_train, epochs=2,
validation_data=(X_valid_scaled, y_valid))
model.evaluate(X_test_scaled, y_test)
```
## Custom Models
```
X_new_scaled = X_test_scaled
class ResidualBlock(keras.layers.Layer):
def __init__(self, n_layers, n_neurons, **kwargs):
super().__init__(**kwargs)
self.n_layers = n_layers # not shown in the book
self.n_neurons = n_neurons # not shown
self.hidden = [keras.layers.Dense(n_neurons, activation="elu",
kernel_initializer="he_normal")
for _ in range(n_layers)]
def call(self, inputs):
Z = inputs
for layer in self.hidden:
Z = layer(Z)
return inputs + Z
def get_config(self): # not shown
base_config = super().get_config() # not shown
return {**base_config, # not shown
"n_layers": self.n_layers, "n_neurons": n_neurons} # not shown
class ResidualRegressor(keras.models.Model):
def __init__(self, output_dim, **kwargs):
super().__init__(**kwargs)
self.output_dim = output_dim # not shown in the book
self.hidden1 = keras.layers.Dense(30, activation="elu",
kernel_initializer="he_normal")
self.block1 = ResidualBlock(2, 30)
self.block2 = ResidualBlock(2, 30)
self.out = keras.layers.Dense(output_dim)
def call(self, inputs):
Z = self.hidden1(inputs)
for _ in range(1 + 3):
Z = self.block1(Z)
Z = self.block2(Z)
return self.out(Z)
def get_config(self): # not shown
base_config = super().get_config() # not shown
return {**base_config, # not shown
"output_dim": self.output_dim} # not shown
model = ResidualRegressor(1)
model.compile(loss="mse", optimizer="nadam")
history = model.fit(X_train_scaled, y_train, epochs=5)
score = model.evaluate(X_test_scaled, y_test)
y_pred = model.predict(X_new_scaled)
#TODO: check that persistence ends up working in TF2
#model.save("my_custom_model.h5")
#model = keras.models.load_model("my_custom_model.h5",
# custom_objects={
# "ResidualBlock": ResidualBlock,
# "ResidualRegressor": ResidualRegressor
# })
```
We could have defined the model using the sequential API instead:
```
block1 = ResidualBlock(2, 30)
model = keras.models.Sequential([
keras.layers.Dense(30, activation="elu", kernel_initializer="he_normal"),
block1, block1, block1, block1,
ResidualBlock(2, 30),
keras.layers.Dense(1)
])
model.compile(loss="mse", optimizer="nadam")
history = model.fit(X_train_scaled, y_train, epochs=5)
score = model.evaluate(X_test_scaled, y_test)
y_pred = model.predict(X_new_scaled)
```
## Losses and Metrics Based on Model Internals
TODO: check https://github.com/tensorflow/tensorflow/issues/26260
```python
class ReconstructingRegressor(keras.models.Model):
def __init__(self, output_dim, **kwargs):
super().__init__(**kwargs)
self.hidden = [keras.layers.Dense(30, activation="selu",
kernel_initializer="lecun_normal")
for _ in range(5)]
self.out = keras.layers.Dense(output_dim)
self.reconstruction_mean = keras.metrics.Mean(name="reconstruction_error")
def build(self, batch_input_shape):
n_inputs = batch_input_shape[-1]
self.reconstruct = keras.layers.Dense(n_inputs)
super().build(batch_input_shape)
@tf.function
def call(self, inputs, training=None):
if training is None:
training = keras.backend.learning_phase()
Z = inputs
for layer in self.hidden:
Z = layer(Z)
reconstruction = self.reconstruct(Z)
recon_loss = tf.reduce_mean(tf.square(reconstruction - inputs))
self.add_loss(0.05 * reconstruction_loss)
if training:
result = self.reconstruction_mean(recon_loss)
self.add_metric(result)
return self.out(Z)
model = ReconstructingRegressor(1)
model.build(tf.TensorShape([None, 8])) # <= Fails if this line is removed
model.compile(loss="mse", optimizer="nadam")
history = model.fit(X, y, epochs=2)
```
```
class ReconstructingRegressor(keras.models.Model):
def __init__(self, output_dim, **kwargs):
super().__init__(**kwargs)
self.hidden = [keras.layers.Dense(30, activation="selu",
kernel_initializer="lecun_normal")
for _ in range(5)]
self.out = keras.layers.Dense(output_dim)
def build(self, batch_input_shape):
n_inputs = batch_input_shape[-1]
self.reconstruct = keras.layers.Dense(n_inputs)
super().build(batch_input_shape)
def call(self, inputs):
Z = inputs
for layer in self.hidden:
Z = layer(Z)
reconstruction = self.reconstruct(Z)
recon_loss = tf.reduce_mean(tf.square(reconstruction - inputs))
self.add_loss(0.05 * recon_loss)
return self.out(Z)
model = ReconstructingRegressor(1)
model.build(tf.TensorShape([None, 8])) # TODO: check https://github.com/tensorflow/tensorflow/issues/26274
model.compile(loss="mse", optimizer="nadam")
history = model.fit(X_train_scaled, y_train, epochs=2)
y_pred = model.predict(X_test_scaled)
```
## Computing Gradients Using Autodiff
```
def f(w1, w2):
return 3 * w1 ** 2 + 2 * w1 * w2
w1, w2 = 5, 3
eps = 1e-6
(f(w1 + eps, w2) - f(w1, w2)) / eps
(f(w1, w2 + eps) - f(w1, w2)) / eps
w1, w2 = tf.Variable(5.), tf.Variable(3.)
with tf.GradientTape() as tape:
z = f(w1, w2)
gradients = tape.gradient(z, [w1, w2])
gradients
with tf.GradientTape() as tape:
z = f(w1, w2)
dz_dw1 = tape.gradient(z, w1)
try:
dz_dw2 = tape.gradient(z, w2)
except RuntimeError as ex:
print(ex)
with tf.GradientTape(persistent=True) as tape:
z = f(w1, w2)
dz_dw1 = tape.gradient(z, w1)
dz_dw2 = tape.gradient(z, w2) # works now!
del tape
dz_dw1, dz_dw2
c1, c2 = tf.constant(5.), tf.constant(3.)
with tf.GradientTape() as tape:
z = f(c1, c2)
gradients = tape.gradient(z, [c1, c2])
gradients
with tf.GradientTape() as tape:
tape.watch(c1)
tape.watch(c2)
z = f(c1, c2)
gradients = tape.gradient(z, [c1, c2])
gradients
with tf.GradientTape() as tape:
z1 = f(w1, w2 + 2.)
z2 = f(w1, w2 + 5.)
z3 = f(w1, w2 + 7.)
tape.gradient([z1, z2, z3], [w1, w2])
with tf.GradientTape(persistent=True) as tape:
z1 = f(w1, w2 + 2.)
z2 = f(w1, w2 + 5.)
z3 = f(w1, w2 + 7.)
tf.reduce_sum(tf.stack([tape.gradient(z, [w1, w2]) for z in (z1, z2, z3)]), axis=0)
del tape
with tf.GradientTape(persistent=True) as hessian_tape:
with tf.GradientTape() as jacobian_tape:
z = f(w1, w2)
jacobians = jacobian_tape.gradient(z, [w1, w2])
hessians = [hessian_tape.gradient(jacobian, [w1, w2])
for jacobian in jacobians]
del hessian_tape
jacobians
hessians
def f(w1, w2):
return 3 * w1 ** 2 + tf.stop_gradient(2 * w1 * w2)
with tf.GradientTape() as tape:
z = f(w1, w2)
tape.gradient(z, [w1, w2])
x = tf.Variable(100.)
with tf.GradientTape() as tape:
z = my_softplus(x)
tape.gradient(z, [x])
tf.math.log(tf.exp(tf.constant(30., dtype=tf.float32)) + 1.)
x = tf.Variable([100.])
with tf.GradientTape() as tape:
z = my_softplus(x)
tape.gradient(z, [x])
@tf.custom_gradient
def my_better_softplus(z):
exp = tf.exp(z)
def my_softplus_gradients(grad):
return grad / (1 + 1 / exp)
return tf.math.log(exp + 1), my_softplus_gradients
def my_better_softplus(z):
return tf.where(z > 30., z, tf.math.log(tf.exp(z) + 1.))
x = tf.Variable([1000.])
with tf.GradientTape() as tape:
z = my_better_softplus(x)
z, tape.gradient(z, [x])
```
# Computing Gradients Using Autodiff
```
l2_reg = keras.regularizers.l2(0.05)
model = keras.models.Sequential([
keras.layers.Dense(30, activation="elu", kernel_initializer="he_normal",
kernel_regularizer=l2_reg),
keras.layers.Dense(1, kernel_regularizer=l2_reg)
])
def random_batch(X, y, batch_size=32):
idx = np.random.randint(len(X), size=batch_size)
return X[idx], y[idx]
def print_status_bar(iteration, total, loss, metrics=None):
metrics = " - ".join(["{}: {:.4f}".format(m.name, m.result())
for m in [loss] + (metrics or [])])
end = "" if iteration < total else "\n"
print("\r{}/{} - ".format(iteration, total) + metrics,
end=end)
import time
mean_loss = keras.metrics.Mean(name="loss")
mean_square = keras.metrics.Mean(name="mean_square")
for i in range(1, 50 + 1):
loss = 1 / i
mean_loss(loss)
mean_square(i ** 2)
print_status_bar(i, 50, mean_loss, [mean_square])
time.sleep(0.05)
```
A fancier version with a progress bar:
```
def progress_bar(iteration, total, size=30):
running = iteration < total
c = ">" if running else "="
p = (size - 1) * iteration // total
fmt = "{{:-{}d}}/{{}} [{{}}]".format(len(str(total)))
params = [iteration, total, "=" * p + c + "." * (size - p - 1)]
return fmt.format(*params)
progress_bar(3500, 10000, size=6)
def print_status_bar(iteration, total, loss, metrics=None, size=30):
metrics = " - ".join(["{}: {:.4f}".format(m.name, m.result())
for m in [loss] + (metrics or [])])
end = "" if iteration < total else "\n"
print("\r{} - {}".format(progress_bar(iteration, total), metrics), end=end)
mean_loss = keras.metrics.Mean(name="loss")
mean_square = keras.metrics.Mean(name="mean_square")
for i in range(1, 50 + 1):
loss = 1 / i
mean_loss(loss)
mean_square(i ** 2)
print_status_bar(i, 50, mean_loss, [mean_square])
time.sleep(0.05)
n_epochs = 5
batch_size = 32
n_steps = len(X_train) // batch_size
optimizer = keras.optimizers.Nadam(lr=0.01)
loss_fn = keras.losses.mean_squared_error
mean_loss = keras.metrics.Mean()
metrics = [keras.metrics.MeanAbsoluteError()]
for epoch in range(1, n_epochs + 1):
print("Epoch {}/{}".format(epoch, n_epochs))
for step in range(1, n_steps + 1):
X_batch, y_batch = random_batch(X_train_scaled, y_train)
with tf.GradientTape() as tape:
y_pred = model(X_batch)
main_loss = tf.reduce_mean(loss_fn(y_batch, y_pred))
loss = tf.add_n([main_loss] + model.losses)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
for variable in model.variables:
if variable.constraint is not None:
variable.assign(variable.constraint(variable))
mean_loss(loss)
for metric in metrics:
metric(y_batch, y_pred)
print_status_bar(step * batch_size, len(y_train), mean_loss, metrics)
print_status_bar(len(y_train), len(y_train), mean_loss, metrics)
for metric in [mean_loss] + metrics:
metric.reset_states()
try:
from tqdm import tnrange
from collections import OrderedDict
with tnrange(1, n_epochs + 1, desc="All epochs") as epochs:
for epoch in epochs:
with tnrange(1, n_steps + 1, desc="Epoch {}/{}".format(epoch, n_epochs)) as steps:
for step in steps:
X_batch, y_batch = random_batch(X_train_scaled, y_train)
with tf.GradientTape() as tape:
y_pred = model(X_batch)
main_loss = tf.reduce_mean(loss_fn(y_batch, y_pred))
loss = tf.add_n([main_loss] + model.losses)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
for variable in model.variables:
if variable.constraint is not None:
variable.assign(variable.constraint(variable))
status = OrderedDict()
mean_loss(loss)
status["loss"] = mean_loss.result().numpy()
for metric in metrics:
metric(y_batch, y_pred)
status[metric.name] = metric.result().numpy()
steps.set_postfix(status)
for metric in [mean_loss] + metrics:
metric.reset_states()
except ImportError as ex:
print("To run this cell, please install tqdm, ipywidgets and restart Jupyter")
```
## TensorFlow Functions
```
def cube(x):
return x ** 3
cube(2)
cube(tf.constant(2.0))
tf_cube = tf.function(cube)
tf_cube
tf_cube(2)
tf_cube(tf.constant(2.0))
```
### TF Functions and Concrete Functions
```
concrete_function = tf_cube.get_concrete_function(tf.constant(2.0))
concrete_function.graph
concrete_function(tf.constant(2.0))
concrete_function is tf_cube.get_concrete_function(tf.constant(2.0))
```
### Exploring Function Definitions and Graphs
```
concrete_function.graph
ops = concrete_function.graph.get_operations()
ops
pow_op = ops[2]
list(pow_op.inputs)
pow_op.outputs
concrete_function.graph.get_operation_by_name('x')
concrete_function.graph.get_tensor_by_name('Identity:0')
concrete_function.function_def.signature
```
### How TF Functions Trace Python Functions to Extract Their Computation Graphs
```
@tf.function
def tf_cube(x):
print("print:", x)
return x ** 3
result = tf_cube(tf.constant(2.0))
result
result = tf_cube(2)
result = tf_cube(3)
result = tf_cube(tf.constant([[1., 2.]])) # New shape: trace!
result = tf_cube(tf.constant([[3., 4.], [5., 6.]])) # New shape: trace!
result = tf_cube(tf.constant([[7., 8.], [9., 10.], [11., 12.]])) # no trace
```
It is also possible to specify a particular input signature:
```
@tf.function(input_signature=[tf.TensorSpec([None, 28, 28], tf.float32)])
def shrink(images):
print("Tracing", images)
return images[:, ::2, ::2] # drop half the rows and columns
img_batch_1 = tf.random.uniform(shape=[100, 28, 28])
img_batch_2 = tf.random.uniform(shape=[50, 28, 28])
preprocessed_images = shrink(img_batch_1) # Traces the function.
preprocessed_images = shrink(img_batch_2) # Reuses the same concrete function.
img_batch_3 = tf.random.uniform(shape=[2, 2, 2])
try:
preprocessed_images = shrink(img_batch_3) # rejects unexpected types or shapes
except ValueError as ex:
print(ex)
```
### Using Autograph To Capture Control Flow
A "static" `for` loop using `range()`:
```
@tf.function
def add_10(x):
for i in range(10):
x += 1
return x
add_10(tf.constant(5))
add_10.get_concrete_function(tf.constant(5)).graph.get_operations()
```
A "dynamic" loop using `tf.while_loop()`:
```
@tf.function
def add_10(x):
condition = lambda i, x: tf.less(i, 10)
body = lambda i, x: (tf.add(i, 1), tf.add(x, 1))
final_i, final_x = tf.while_loop(condition, body, [tf.constant(0), x])
return final_x
add_10(tf.constant(5))
add_10.get_concrete_function(tf.constant(5)).graph.get_operations()
```
A "dynamic" `for` loop using `tf.range()` (captured by autograph):
```
@tf.function
def add_10(x):
for i in tf.range(10):
x = x + 1
return x
add_10.get_concrete_function(tf.constant(0)).graph.get_operations()
```
### Handling Variables and Other Resources in TF Functions
```
counter = tf.Variable(0)
@tf.function
def increment(counter, c=1):
return counter.assign_add(c)
increment(counter)
increment(counter)
function_def = increment.get_concrete_function(counter).function_def
function_def.signature.input_arg[0]
counter = tf.Variable(0)
@tf.function
def increment(c=1):
return counter.assign_add(c)
increment()
increment()
function_def = increment.get_concrete_function().function_def
function_def.signature.input_arg[0]
class Counter:
def __init__(self):
self.counter = tf.Variable(0)
@tf.function
def increment(self, c=1):
return self.counter.assign_add(c)
c = Counter()
c.increment()
c.increment()
@tf.function
def add_10(x):
for i in tf.range(10):
x += 1
return x
tf.autograph.to_code(add_10.python_function, experimental_optional_features=None)
# TODO: experimental_optional_features is needed to have the same behavior as @tf.function,
# check that this is not needed when TF2 is released
def display_tf_code(func, experimental_optional_features=None):
from IPython.display import display, Markdown
if hasattr(func, "python_function"):
func = func.python_function
code = tf.autograph.to_code(func, experimental_optional_features=experimental_optional_features)
display(Markdown('```python\n{}\n```'.format(code)))
display_tf_code(add_10)
```
## Using TF Functions with tf.keras (or Not)
By default, tf.keras will automatically convert your custom code into TF Functions, no need to use
`tf.function()`:
```
# Custom loss function
def my_mse(y_true, y_pred):
print("Tracing loss my_mse()")
return tf.reduce_mean(tf.square(y_pred - y_true))
# Custom metric function
def my_mae(y_true, y_pred):
print("Tracing metric my_mae()")
return tf.reduce_mean(tf.abs(y_pred - y_true))
# Custom layer
class MyDense(keras.layers.Layer):
def __init__(self, units, activation=None, **kwargs):
super().__init__(**kwargs)
self.units = units
self.activation = keras.activations.get(activation)
def build(self, input_shape):
self.kernel = self.add_weight(name='kernel',
shape=(input_shape[1], self.units),
initializer='uniform',
trainable=True)
self.biases = self.add_weight(name='bias',
shape=(self.units,),
initializer='zeros',
trainable=True)
super().build(input_shape)
def call(self, X):
print("Tracing MyDense.call()")
return self.activation(X @ self.kernel + self.biases)
# Custom model
class MyModel(keras.models.Model):
def __init__(self, **kwargs):
super().__init__(**kwargs)
self.hidden1 = MyDense(30, activation="relu")
self.hidden2 = MyDense(30, activation="relu")
self.output_ = MyDense(1)
def call(self, input):
print("Tracing MyModel.call()")
hidden1 = self.hidden1(input)
hidden2 = self.hidden2(hidden1)
concat = keras.layers.concatenate([input, hidden2])
output = self.output_(concat)
return output
model = MyModel()
model.compile(loss=my_mse, optimizer="nadam", metrics=[my_mae])
model.fit(X_train_scaled, y_train, epochs=2,
validation_data=(X_valid_scaled, y_valid))
model.evaluate(X_test_scaled, y_test)
```
You can turn this off by creating the model with `dynamic=True` (or calling `super().__init__(dynamic=True, **kwargs)` in the model's constructor):
```
model = MyModel(dynamic=True)
model.compile(loss=my_mse, optimizer="nadam", metrics=[my_mae])
```
Not the custom code will be called at each iteration. Let's fit, validate and evaluate with tiny datasets to avoid getting too much output:
```
model.fit(X_train_scaled[:64], y_train[:64], epochs=1,
validation_data=(X_valid_scaled[:64], y_valid[:64]), verbose=0)
model.evaluate(X_test_scaled[:64], y_test[:64], verbose=0)
```
Alternatively, you can compile a model with `run_eagerly=True`:
```
model = MyModel()
model.compile(loss=my_mse, optimizer="nadam", metrics=[my_mae], run_eagerly=True)
model.fit(X_train_scaled[:64], y_train[:64], epochs=1,
validation_data=(X_valid_scaled[:64], y_valid[:64]), verbose=0)
model.evaluate(X_test_scaled[:64], y_test[:64], verbose=0)
```
## Custom Optimizers
Defining custom optimizers is not very common, but in case you are one of the happy few who gets to write one, here is an example:
```
class MyMomentumOptimizer(keras.optimizers.Optimizer):
def __init__(self, learning_rate=0.001, momentum=0.9, name="MyMomentumOptimizer", **kwargs):
"""Call super().__init__() and use _set_hyper() to store hyperparameters"""
super().__init__(name, **kwargs)
self._set_hyper("learning_rate", kwargs.get("lr", learning_rate)) # handle lr=learning_rate
self._set_hyper("decay", self._initial_decay) #
self._set_hyper("momentum", momentum)
def _create_slots(self, var_list):
"""For each model variable, create the optimizer variable associated with it.
TensorFlow calls these optimizer variables "slots".
For momentum optimization, we need one momentum slot per model variable.
"""
for var in var_list:
self.add_slot(var, "momentum")
@tf.function
def _resource_apply_dense(self, grad, var):
"""Update the slots and perform one optimization step for one model variable
"""
var_dtype = var.dtype.base_dtype
lr_t = self._decayed_lr(var_dtype) # handle learning rate decay
momentum_var = self.get_slot(var, "momentum")
momentum_hyper = self._get_hyper("momentum", var_dtype)
momentum_var.assign(momentum_var * momentum_hyper - (1. - momentum_hyper)* grad)
var.assign_add(momentum_var * lr_t)
def _resource_apply_sparse(self, grad, var):
raise NotImplementedError
def get_config(self):
base_config = super().get_config()
return {
**base_config,
"learning_rate": self._serialize_hyperparameter("learning_rate"),
"decay": self._serialize_hyperparameter("decay"),
"momentum": self._serialize_hyperparameter("momentum"),
}
model = keras.models.Sequential([keras.layers.Dense(1, input_shape=[8])])
model.compile(loss="mse", optimizer=MyMomentumOptimizer())
model.fit(X_train_scaled, y_train, epochs=5)
```
| github_jupyter |
# Parcels plotting methods
**Disclaimer: This tutorial demonstrates the simple plotting functionality included in Parcels. For high quality analysis it is recommended to create your own code. [This tutorial](https://nbviewer.jupyter.org/github/OceanParcels/parcels/blob/master/parcels/examples/tutorial_output.ipynb) shows how to start a more advanced analysis.**
The `show()` method of the `Particelset` class is capable of plotting the particle locations and velocity fields in scalar and vector form. In this notebook, we demonstrate these capabilities using the GlobCurrent dataset. We begin by importing the relevant modules.
```
%matplotlib inline
from parcels import FieldSet, ParticleSet, JITParticle, AdvectionRK4
from datetime import timedelta, datetime
```
We then instatiate a `FieldSet` with the velocity field data from GlobCurrent dataset.
```
filenames = {'U': "GlobCurrent_example_data/20*.nc",
'V': "GlobCurrent_example_data/20*.nc"}
variables = {'U': 'eastward_eulerian_current_velocity',
'V': 'northward_eulerian_current_velocity'}
dimensions = {'lat': 'lat',
'lon': 'lon',
'time': 'time'}
fieldset = FieldSet.from_netcdf(filenames, variables, dimensions)
```
Next, we instantiate a `ParticeSet` composed of `JITParticles`:
```
pset = ParticleSet.from_line(fieldset=fieldset, size=5, pclass=JITParticle,
start=(31, -31), finish=(32, -31), time=datetime(2002, 1, 1))
```
Given this `ParticleSet`, we can now explore the different features of the `show()` method. To start, let's simply call `show()` with no arguments.
```
pset.show()
```
Then, let's advect the particles starting on January 1, 2002 for a week.
```
pset.execute(AdvectionRK4, runtime=timedelta(days=7), dt=timedelta(minutes=5))
```
If we call `show()` again, we will see that the particles have been advected:
```
pset.show()
```
To plot without the continents on the same plot, add `land=False`.
```
pset.show(land=False)
```
To set the domain of the plot, we specify the domain argument. The format `domain` expects a dictionary with entries `{'S', 'N', 'E', 'W'}` for South, North, East and West extent, respectively. Note that the plotted domain is found by interpolating the user-specified domain onto the velocity grid. For instance,
```
pset.show(domain={'N':-31, 'S':-35, 'E':33, 'W':26})
```
We can also easily display a scalar contour plot of a single component of the velocity vector field. This is done by setting the `field` argument equal to the desired scalar velocity field.
```
pset.show(field=fieldset.U)
```
To plot the scalar U velocity field at a different date and time, we set the argument `show_time` equal to a `datetime` or `timedelta` object or simply the number of seconds since the time origin. For instance, let's view the U field on January, 10, 2002 at 2 PM.
```
pset.show(field=fieldset.U, show_time=datetime(2002, 1, 10, 2))
```
Note that the particle locations do not change, but remain at the locations corresponding to the end of the last integration. To remove them from the plot, we set the argument `with_particles` equal to `False`.
```
pset.show(field=fieldset.U, show_time=datetime(2002, 1, 10, 2), with_particles=False)
```
By setting the `field` argument equal to `vector`, we can display the velocity in full vector form.
```
pset.show(field='vector')
```
The normalized vector field is colored by speed. To control the maximum speed value on the colorbar, set the `vmax` argument equal to the desired value.
```
pset.show(field='vector', vmax=3.0, domain={'N':-31, 'S':-39, 'E':33, 'W':18})
```
We can change the projection of the plot by providing one of the [projections](https://scitools.org.uk/cartopy/docs/v0.15/crs/projections.html) from `cartopy`. For example, to plot on a Robinson projection , we use `projection=cartopy.crs.Robinson()`. Note that not all projections support gridlines, so these may not be shown.
```
try: # Within a try/pass for unit testing on machines without cartopy installed
import cartopy
pset.show(field='vector', vmax=3.0, domain={'N':-31, 'S':-39, 'E':33, 'W':18}, projection=cartopy.crs.Robinson())
except:
pass
```
If we want to save the file rather than show it, we set the argument `savefile` equal to the `'path/to/save/file'`.
```
pset.show(field='vector', vmax=3.0, domain={'N':-31, 'S':-39, 'E':33, 'W':18}, land=True, savefile='particles')
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Module-1:-Overview" data-toc-modified-id="Module-1:-Overview-1"><span class="toc-item-num">1 </span>Module 1: Overview</a></span></li><li><span><a href="#Printing" data-toc-modified-id="Printing-2"><span class="toc-item-num">2 </span>Printing</a></span></li><li><span><a href="#Creating-variables" data-toc-modified-id="Creating-variables-3"><span class="toc-item-num">3 </span>Creating variables</a></span></li><li><span><a href="#Variable-types" data-toc-modified-id="Variable-types-4"><span class="toc-item-num">4 </span>Variable types</a></span></li><li><span><a href="#Calculations-with-variables" data-toc-modified-id="Calculations-with-variables-5"><span class="toc-item-num">5 </span>Calculations with variables</a></span></li><li><span><a href="#Built-in-constants-and-mathematical-functions" data-toc-modified-id="Built-in-constants-and-mathematical-functions-6"><span class="toc-item-num">6 </span>Built-in constants and mathematical functions</a></span></li><li><span><a href="#Last-exercises" data-toc-modified-id="Last-exercises-7"><span class="toc-item-num">7 </span>Last exercises</a></span></li></ul></div>
> All content here is under a Creative Commons Attribution [CC-BY 4.0](https://creativecommons.org/licenses/by/4.0/) and all source code is released under a [BSD-2 clause license](https://en.wikipedia.org/wiki/BSD_licenses).
>
>Please reuse, remix, revise, and [reshare this content](https://github.com/kgdunn/python-basic-notebooks) in any way, keeping this notice.
# Module 1: Overview
This tutorial covers several topics that are important to the beginning Python user, especially if you are coming from using another programming language.
1. Printing
2. Creating variables
3. Types of variables
3. Calculations with variables
4. Built-in constants and mathematical functions
6. Exercises
## Preparing for this module
* Have a basic Python installation that works. You can follow the instructions below, which should generate the output shown.
# Printing
* The ``print(...)`` function sends output to the screen. It is useful for debugging.
* What is the equivalent way to do it in MATLAB, or in Java, or another language that you know already?
* The ``print(...)`` function can use 'strings' or "strings", in other words with single or double quotes.
Print the following text to the screen:
> Hi, my name is \_\_\_\_\_
Now try this:
> The Python language was created by Guido van Rossum in the 1980's when he worked at the Centrum Wiskunde & Informatica in Amsterdam. Yes, he's from the Netherlands, but has moved to the US and worked for Google, but now at Dropbox.
You can verify the above by typing just the following two commands separately in Python:
* ``credits``
* ``copyright``
```
print('Hi, my name is Kevin.')
```
```python
long_string = """If you really want to write paragraphs,
and paragraphs of text, you do it with the triple quotes. Try it"""
print(long_string)
```
* Verify how the above variable ``long_string`` will be printed. Does Python put a line break where you expect it?
* Can you use single quotes instead of double quotes ?
You can also create longer strings in Python using the bracket construction. Try this:
```python
print('Here is my first line.',
'Then the second.',
'And finally a third.',
'But did you expect that?')
```
The reason for this is stylistic. Python, unlike other languages, has some recommended rules,
which we will introduce throughout these modules. One of these rules is that you don't exceed 79 characters per line.
It helps to keep your code width narrow: you can then put two or three code files side-by-side on a widescreen monitor.
Python also has escape characters. Discover it:
* Try to print just the backslash character on its own: `print('\')`
* Try this instead: `print('\\')`
The "\\" on its own is an *escape character*. Google that term.
What are the escape characters for:
* a tab?
* a new line?
1. Try to print this: ``print('The files are in C:\Data\dirnew\excel')``.
2. Try to print this: ``print('The files are in C:\temp\newdir\excel')``.
> Why does it create such an unexpected mess?
>
> There are two different ways to quickly fix the code to show what is intended.
> Try using escape characters first; we will show a more efficient way later.
# Creating variables
We already saw above how a variable was created: ``long_string = """ .... """``.
You've done this plenty of times in other programming languages; almost always with an "=". We prefer to refer to "=" as the "assignment" operator; not "equals".
What goes on the left hand side of the assignment must be a '*valid variable name*'.
Which of the following are valid variables, or valid ways to create variables in Python?
```python
my_integer = 3.1428571
_my_float = 3.1428571 # variables like this have a special use in Python
__my_float__ = 3.1428571 # variables like this have a special use in Python
€value = 42.95
cost_in_€ = 42.95
cost_in_dollars = 42.95
42.95 = cost_in_dollars
dollar.price = 42.95
favourite#tag = '#like4like'
favourite_hashtag = '#일상'
x = y = z = 1
a, b, c = 1, 2, 3 # tuple assignment
a, b, c = (1, 2, 3)
i, f, s = 42, 12.94, 'spam'
from = 'my lover'
raise = 'your glass'
pass = 'you shall not'
fail = 'you will not'
True = 'not False'
pay day = 'Thursday'
NA = 'not applicable' # for R users
a = 42; # for MATLAB users: semi-colons are never required
A = 13 # like most languages, Python is also case-sensitive
```
What's the most interesting idea/concept you learned from this?
In the software editor you are using, is there a space where you can see the variable's current value? \[This is called the *Watch* window in most graphical editors\]

# Variable types
Each variable you create has a ``type``, which is automatically detected based on what is on the right hand side of the "=" sign. This is similar to MATLAB, but very different from C++ where you require this type of code:
```c
int a, b; // first declare your variables
float result;
a = 5; // then you get to use them
b = 2;
result = a / b; // you will get an unexpected value if you had defined "result" as "int"
```
In Python you have _dynamic typing_, where you simply write:
```python
a, b = 5, 2
result = a / b
```
and Python figures out the most appropriate type from the context.
Run those 2 lines of Python code. Then add this below. What is the output you see?
```python
type(a)
type(result)
```
Each variable always has a **type**. Usually you know what the type is, because you created the variable yourself at some point.
But on occasion you use someone else's code and you get back an answer that you don't know the type of. Then it is useful to check it with the ``type(...)`` function.
Try these lines in Python:
```python
type(99)
type(99.)
type(9E9)
type('It\'s raining cats and dogs today!') # How can you rewrite this line better?
type(r'Brexit will cost you £8E8. Thank you.')
type(['this', 'is', 'a', 'vector', 'of', 7, 'values'])
type([])
type(4 > 5)
type(True)
type(False)
type(None)
type({'this': 'is what is called a', 'dictionary': 'variable!'})
type(('this', 'is', 'called', 'a', 'tuple'))
type(())
type(None)
type(print) # anything that can be assigned to a variable name can be 'type checked'
```
You can convert most variables to a string, as follows: ``str(...)``
Try these conversions to make sure you get what you expect:
```python
str(45)
type(str(45))
str(92.5)
str(None)
str(print)
```
# Calculations with variables
The next step is to perform some calculations with the variables.
The standard expressions exist in Python:
| Operation | Symbol |
|----------------|--------|
| Addition | + |
| Subtraction | - |
| Multiplication | * |
| Division | / |
| Power of | ** |
| Modulo | % |
Please note: "power of" is not with the ^ operator, and can mislead you. Try this:
* ``print(2 ^ 4)``
* ``print(2**4)``
If you are not familiar with the modulo operator, you can use it as follows: if you have a variable ``x`` and you want to check if it is exactly divisible by, for example, a number 5, then you can check if the modulus, the remainder, is zero.
* ``x % 5``
* ``7 % 5`` evaluates to 2, since there's a remainder of 2 when 7 is divided by 5
* ``10 % 5`` evaluates to 0, since there's no remainder after 10 is divided by 5
Given the above, use Python as a calculator to find the values of these expressions:
If ``a = 5`` and ``b = 9``
* ``a / b``
* What type is the result of the above expression?
* ``a * b``
* What type is the result of the above expression?
The distance $d$ travelled by an object falling for time $t$, given in seconds, is $$d=\frac {1}{2}gt^{2}$$ where $g$ is the gravitational constant = $9.8\, \text{m.s}^{-2}$. Calculate the distance that you will travel in free-fall gravity in 10 seconds:
```python
t = ____ # seconds
d = ____ # meters
print('The distance fallen is ' + str(d) + ' meters.')
```
Try it now the other way around: the time taken for an object to fall is: $$ t= \sqrt {\frac {2d}{g}}$$
We will introduce the ``sqrt`` function in the next section, but you can also calculate square root using a power of 0.5, as in $\sqrt{x} = x^{0.5}$.
Using that knowledge, how long will it take for an object to fall from the top of the building you are currently in:
```python
# Creates a string value in variable 'd'. Verify that it is a string type.
d = input('The height of the building, in meters, which I am currently in is: ')
d = float(d) # convert the string variable to a floating point value
t = ____ # seconds
# You might also want to investigate the "round" function at this point
# to improve the output for variable t.
print('The time for an object to fall from a building',
'of ' + str(d) + ' meters tall is ' + str(t) + \
' seconds.')
```
You wish to automate your workflow to write a standardized statistical report. It should appear as follows on the screen:
> There were <b>4</b> outliers detected in the data set using Grubbs' test at the **99**% confidence level. These outliers will not be used in the subsequent calculations. The regression trend of **45.9** mg/day was detected for this product, with a p-value of **0.00341**. This indicates that there is an important **rising** trend over time.
The 5 pieces in bold should be replaced with variables, so the above paragraph can be automatically written in the future with different variable values.
Write the Python code, creating the 5 variables. Print your paragraph of text to the screen. The last variable can either be 'rising' or 'falling'. (In the next class we will see how you can use an if-else statement to write ``'rising'`` or ``'falling'`` depending on the sign of the regression slope.
Python, like other languages, has the order of operations rules (same as the PEMDAS rules you might have learned in school):
1. **P**arentheses (round brackets)
2. **E**xponentiation or powers, left to right
3. **M**ultiplication and <b>D</b>ivision, left to right
4. <b>A</b>ddition and **S**ubtraction, left to right
So what is the result of these statements?
```python
a = 1 + 3 ** 2 * 4 / 2
b = 1 + 3 ** (2 * 4) / 2
c = (1 + 3) ** 2 * 4 / 2
```
While it is good to know these rules, the general advice is to always use brackets to clearly show your actual intention.
> Never leave the reader of your code guessing: someone will have to maintain your code after you; including yourself, a few years later 😉
Write code for the following:
Divide the sum of <i>a</i> and _b_ by the product of <i>c</i> and *d*, and store the result in <i>x</i>.
The above operators return results which are either ``int`` or ``float``.
There are another set of operators which return ***bool***ean values: ``True`` or ``False``. We will use these frequently when we make decisions in our code. For example:
> if \_\_<condition> \_\_ then \_\_<action\>\_\_
We cover **if-statements** in the [next module](https://yint.org/pybasic02).
But for now, try out these ``<condition>`` statements:
```python
3 < 5
5 < 3
4 <= 4
4 <= 9.2
5 == 5
5. == 5 # float on the left, and int on the right. Does it matter?
5. != 5 # does that make sense?
True != False
False < True
```
Related to these operators are some others, which you can use to combine up: ``and`` and ``not`` and ``or``
Try these out. What do you get in each case?
```python
True and not False
True and not(False)
True and True
not(False) or False
```
In the quadratic equation $$ax^2 + bx + c=0$$ the short-cut solution is given by $$ x= -\frac{b}{2a}$$
but only if $b^2 - 4ac=0$ and $a \neq 0$.
Verify if you can use this short-cut solution for these combinations:
* ``a=3, b=-1, c=2 # use tuple-assignment to create these 3 variables in 1 line of code``
* ``a=0, b=-1, c=2``
* ``a=3, b=6, c=3``
Write the single line of Python code that will return ``True`` if you can use the shortcut, or ``False`` if you cannot.
# Built-in constants and mathematical functions
You will certainly need to calculate logs, exponentials, square roots, or require the value of $e$ or $\pi$ at some point.
In this last section we get a bit ahead, and load a Python library to provide this for us. Libraries - we will see later - are a collection of functions and variables that pre-package useful tools. Libraries can be large collections of code, and are for special purposes, so they are not loaded automatically when you launch Python.
In MATLAB you can think of *Toolboxes* as being equivalent; in R you have *Packages*; in C++ and Java you also use the word *Library* for the equivalent concept.
In Python, there are several libraries that come standard, and one is the ``math`` library. Use the ``import`` command to load the library. The ``math`` library can be used as follows:
```python
import math
radius = 3 # cm
area_of_circle = math.pi * radius**2
print('The area of the circle is ' + str(area_of_circle))
```
Now that you know how to use the ``math`` library, it is worth searching what else is in it:
> https://www.google.com/search?q=python+math+library
All built-in Python libraries are documented in the same way. Searching this way usually brings up the link near the top. Make sure you look at the documentation for Python version 3.x.
Now that you have the documentation ready, use functions from that library to calculate:
* the *ceiling* of a number, for example ``a = 3.7``
* the *floor* of a number, for example ``b = 3.7``
* the *absolute* value of ``c = -2.9``
* the log to the base $e$ of ``d = 100``
* the log to the base 10 of ``e = 100``
* the Golden ratio ${\dfrac {1+{\sqrt {5}}}{2}}$
* check that the factorial n=9, in other words $n!$ is equal to 362880
* and see that Stirling's approximation, $n! \approx \sqrt{2\pi n} \cdot n^n e^{-n}$ for a factorial matches closely [you will use 4 different methods from the ``math`` library to calculate this!]
```python
<put your code here>
print('The true value of 9! is ' + ___ + ', while the Stirling approximation is ' + ___)
```
* verify that the cosine of ``g`` = $2\pi$ radians is indeed 1.0
# Last exercises
1. The population of a country could be approximated by the formula $$ p(t) = \dfrac{197 273 000}{1 + e^{− 0.03134(t − 1913)}}$$
where the time $t$ is in years.
* What is the population in 1913?
* What is the population in 2013?
* Does the population grow and grow without bounds, or does it reach steady state (it stabilizes at some constant value eventually)
1. Explain to your partner whom you are working with, what some of the benefits are of writing your code in Python files.
```
# IGNORE this. Execute this cell to load the notebook's style sheet.
from IPython.core.display import HTML
css_file = './images/style.css'
HTML(open(css_file, "r").read())
```
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W3D3_NetworkCausality/W3D3_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Neuromatch Academy 2020: Week 3 Day 3, Tutorial 1
# Causality Day: Interventions
#Tutorial Objectives
We list our overall day objectives below, with the sections we will focus on in this notebook in bold:
1. **Master definitions of causality**
2. **Understand that estimating causality is possible**
3. Learn 5 different methods and understand when they fail
1. **perturbations**
2. correlations
3. simultaneous fitting/regression
4. Granger causality
5. instrumental variables
### Tutorial setting
The methods we'll learn are very general.
They apply to analyzing fMRI data: **when is functional connectivity causal connectivity**?
They apply to sensory analyses: **what about stimuli causes neurons to spike?**
They apply to motor control: **How does this brain area cause changes in movement?**
Causal questions are everywhere!
### Tutorial 1 Objectives:
1. Simulate a system we can discuss
2. Understand perturbation as a method of estimating causality
# Setup
```
import matplotlib.pyplot as plt # import plotting modules
import numpy as np # import numpy for matrix manipulation
#@title Figure Settings
%matplotlib inline
fig_w, fig_h = (8, 6)
plt.rcParams.update({'figure.figsize': (fig_w, fig_h)})
%config InlineBackend.figure_format = 'retina'
#@title Helper functions
def sigmoid(x):
"""
Compute sigmoid nonlinearity element-wise on x.
Args:
x (np.ndarray): the numpy data array we want to transform
Returns
(np.ndarray): x with sigmoid nonlinearity applied
"""
return 1 / (1 + np.exp(-x))
def create_connectivity(n_neurons, random_state=42):
"""
Generate our nxn causal connectivity matrix.
Args:
n_neurons (int): the number of neurons in our system.
random_state (int): random seed for reproducibility
Returns:
A (np.ndarray): our 0.1 sparse connectivity matrix
"""
np.random.seed(random_state)
A_0 = np.random.choice([0,1], size=(n_neurons, n_neurons), p=[0.9, 0.1])
# set the timescale of the dynamical system to about 100 steps
_, s_vals , _ = np.linalg.svd(A_0)
A = A_0 / (1.01 * s_vals[0])
# _, s_val_test, _ = np.linalg.svd(A)
# assert s_val_test[0] < 1, "largest singular value >= 1"
return A
def see_neurons(A, ax):
"""
Visualizes the connectivity matrix.
Args:
A (np.ndarray): the connectivity matrix of shape (n_neurons, n_neurons)
ax (plt.axis): the matplotlib axis to display on
Returns:
Nothing, but visualizes A.
"""
n = len(A)
ax.set_aspect('equal')
thetas = np.linspace(0,np.pi*2,n,endpoint=False )
x,y = np.cos(thetas),np.sin(thetas),
ax.scatter(x,y,c='k',s=150)
#renormalize
A = A / A.max()
for i in range(n):
for j in range(n):
if A[i,j]>0:
ax.arrow(x[i],y[i],x[j]-x[i],y[j]-y[i],color='k',alpha=A[i,j],head_width=.15,
width = A[i,j]/25,shape='right', length_includes_head=True)
ax.axis('off')
def plot_connectivity_matrix(A, ax=None):
"""Plot the (weighted) connectivity matrix A as a heatmap."""
if ax is None:
ax = plt.gca()
lim = np.abs(A).max()
ax.imshow(A, vmin=-lim, vmax=lim, cmap="coolwarm")
```
---
# Defining and estimating causality
Let's think carefully about the statement "**A causes B**". To be concrete, let's take two neurons. What does it mean to say that neuron $A$ causes neuron $B$ to fire?
The *interventional* definition of causality says that:
$$
(A \text{ causes } B) \Leftrightarrow ( \text{ If we force }A \text { to be different, then }B\text{ changes})
$$
To determine if $A$ causes $B$ to fire, we can inject current into neuron $A$ and see what happens to $B$.
**A mathematical definition of causality**:
Over many trials, the average causal effect $\delta_{A\to B}$ of neuron $A$ upon neuron $B$ is the average change in neuron $B$'s activity when we set $A=1$ versus when we set $A=0$.
$$
\delta_{A\to B} = \mathbb{E}[B | A=1] - \mathbb{E}[B | A=0]
$$
Note that this is an average effect. While one can get more sophisticated about conditional effects ($A$ only effects $B$ when it's not refractory, perhaps), today we will only consider average effects.
**Relation to a randomized controlled trial (RCT)**:
The logic we just described is the logic of a randomized control trial (RCT). If you randomly give 100 people a drug and 100 people a placebo, the effect is the difference in outcomes.
## Exercise 1 (Warm-up): a randomized controlled trial for two neurons
Let's actually code that out.
**Setting**: Neuron $A$ synapses on Neuron $B$
**Goal**: Perturb $A$ and confirm that $B$ changes.
Let's have that $B = A + \epsilon$, where $\epsilon$ is standard normal noise $\epsilon\sim\mathcal{N}(0,1)$.
```
def neuron_B(activity_of_A):
"""Model activity of neuron B as neuron A activity + noise
Args:
activity_of_A (ndarray): An array of shape (T,) containing the neural activity of neuron A
Returns:
ndarray: activity of neuron B
"""
noise = np.random.randn(*activity_of_A.shape)
return activity_of_A + noise
# Neuron A activity of zeros
A_0 = np.zeros(5000)
# Neuron A activity of ones
A_1 = np.ones(5000)
###########################################################################
## TODO for students: Estimate the causal effect of A upon B by getting the
## difference in mean of B when A=0 vs. A=1
###########################################################################
# diff_in_means =
# print(diff_in_means)
# to_remove solution
def neuron_B(activity_of_A):
"""Model activity of neuron B as neuron A activity + noise
Args:
activity_of_A (ndarray): An array of shape (T,) containing the neural activity of neuron A
Returns:
ndarray: activity of neuron B
"""
noise = np.random.randn(*activity_of_A.shape)
return activity_of_A + noise
A_0 = np.zeros(5000)
A_1 = np.ones(5000)
diff_in_means = neuron_B(A_1).mean() - neuron_B(A_0).mean()
print(diff_in_means)
```
The result should be close to $1$. Is this what you get?
# Simulating a system of neurons
```
#@title Video
# Insert the ID of the corresponding youtube video
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="3oHhuzUZRfA", width=854, height=480, fs=1)
print("Video available at https://youtu.be/" + video.id)
video
```
Can we still estimate causal effects when the neurons are in big networks? This is the main question we will ask today. Let's first create our system, and the rest of today will be spend analyzing it.
## Today's model
Our system has N interconnected neurons that affect each other over time. Each neuron at time $t+1$ is a function of the activity of the other neurons from the previous time $t$.
Neurons affect each other with a linear function with added noise, which is then passed through a nonlinearity:
$$
\vec{x}_{t+1} = \sigma(A\vec{x}_t + \epsilon_t),
$$
- $\vec{x}_t$ is an $n$-dimensional vector representing our $n$-neuron system at timestep $t$
- $\sigma$ is a sigmoid nonlinearity
- $A$ is our $n \times n$ *causal ground truth connectivity matrix* (more on this later)
- $\epsilon_t$ is random noise: $\epsilon_t \sim N(\vec{0}, I_n)$
- $\vec{x}_0$ is initialized to $\vec{0}$
### Notes on $A$
$A$ is a connectivity matrix, so the element $A_{ij}$ represents the causal effect of neuron $i$ on neuron $j$.
We'll set it up so that neurons on average receive connections from only 10% of the whole population.
## Visualize true connectivity
We will create a connectivity matrix between 6 neurons and visualize it in two different ways: as a graph with directional edges between connected neurons and as an image of the connectivity matrix
```
## Intializes the system
n_neurons = 6
A = create_connectivity(n_neurons) # we are invoking a helper function that generates our nxn causal connectivity matrix.
# Let's plot it!
fig, axs = plt.subplots(1,2, figsize=(10,5))
see_neurons(A,axs[0]) # we are invoking a helper function that visualizes the connectivity matrix
plot_connectivity_matrix(A)
plt.xlabel("Neuron connectivity")
plt.ylabel("Neuron connectivity")
fig.suptitle("Connectivity Matrix")
plt.show()
```
## Exercise 2: System Simulation
In this exercise we're going to simulate the system. Please complete the following function so that at every timestep the activity vector $x$ is updated according to:
$$
\vec{x}_{t+1} = \sigma(A\vec{x}_t + \epsilon_t).
$$
Make sure your noise is N-dimensional, with one element per neuron.
```
def simulate_neurons(A, timesteps, random_state=42):
"""
Simulates a dynamical system for the specified number of neurons and timesteps.
Args:
A (np.array): the connectivity matrix
timesteps (int): the number of timesteps to simulate our system.
random_state (int): random seed for reproducibility
Returns:
- X has shape (n_neurons, timeteps). A schematic:
___t____t+1___
| 0 1 |
| 1 0 |
n_neurons | 0 -> 1 |
| 0 0 |
|___1____0_____|
"""
np.random.seed(random_state)
n_neurons = len(A)
X = np.zeros((n_neurons, timesteps))
for t in range(timesteps-1):
########################################################################
## TODO: Fill in the update rule for our dynamical system.
## Function Hints:
## epsilon: multivariate normal with mean zero and unit variance ->
## np.random.multivariate_normal(np.zeros(n), np.eye(n_neurons))
## Fill in function and remove
raise NotImplementedError("Please make the dynamical system.")
########################################################################
epsilon = ...
X[:, t+1] = sigmoid(A.dot(...) + epsilon) # we are using helper function sigmoid
assert epsilon.shape == (n_neurons,)
return X
## Uncomment to test it
# timesteps = 5000 # Simulate for 5000 timesteps.
# # Simulate our dynamical system for the given amount of time
# X = simulate_neurons(A, timesteps)
# f, ax = plt.subplots()
# ax.imshow(X[:,:10])
# ax.set(xlabel='Timestep', ylabel='Neuron', title='Simulated Neural Activity')
# to_remove solution
def simulate_neurons(A, timesteps, random_state=42):
"""
Simulates a dynamical system for the specified number of neurons and timesteps.
Args:
A (np.array): the connectivity matrix
timesteps (int): the number of timesteps to simulate our system.
random_state (int): random seed for reproducibility
Returns:
- X has shape (n_neurons, timeteps). A schematic:
___t____t+1___
| 0 1 |
| 1 0 |
n_neurons | 0 -> 1 |
| 0 0 |
|___1____0_____|
"""
np.random.seed(random_state)
n_neurons = len(A)
X = np.zeros((n_neurons, timesteps))
for t in range(timesteps-1):
# solution
epsilon = np.random.multivariate_normal(np.zeros(n_neurons), np.eye(n_neurons))
X[:, t+1] = sigmoid(A.dot(X[:,t]) + epsilon) # we are using helper function sigmoid
assert epsilon.shape == (n_neurons,)
return X
### Now test it
timesteps = 5000 # Simulate for 5000 timesteps.
# Simulate our dynamical system for the given amount of time
X = simulate_neurons(A, timesteps)
with plt.xkcd():
f, ax = plt.subplots()
ax.imshow(X[:,:10])
ax.set(xlabel='Timestep', ylabel='Neuron', title='Simulated Neural Activity')
```
# Recovering connectivity through perturbation
```
#@title Video
# Insert the ID of the corresponding youtube video
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="zlRhaGBzkug", width=854, height=480, fs=1)
print("Video available at https://youtu.be/" + video.id)
video
```
## Random perturbation in our system of neurons
We want to get the causal effect of each neuron upon each other neuron. The ground truth of the causal effects is the connectivity matrix $A$.
Remember that we would like to calculate:
$$
\delta_{A\to B} = \mathbb{E}[B | A=1] - \mathbb{E}[B | A=0]
$$
We'll do this by randomly setting the system state to 0 or 1 and observing the outcome after one timestep. If we do this $N$ times, the effect of neuron $i$ upon neuron $j$ is:
$$
\delta_{x^i\to x^j} \approx \frac1N \sum_i^N[x_{t+1}^j | x^i_t=1] - \frac1N \sum_i^N[x_{t+1}^j | x^i_t=0]]
$$
This is just the average difference of the activity of neuron $j$ in the two conditions.
**Coding strategy**: We are going to calculate the above equation, but imagine it like *intervening* in activity every other timestep.
```
def simulate_neurons_perturb(A, timesteps, perturb_freq=2):
"""
Simulates a dynamical system for the specified number of neurons and timesteps,
BUT every other timestep the activity is clamped to a random pattern of 1s and 0s
Args:
A (np.array): the true connectivity matrix
timesteps (int): the number of timesteps to simulate our system.
perturb_freq (int): the perturbation frequency (2 means perturb every other timestep)
random_state (int): random seed for reproducibility
Returns:
The results of the simulated system.
- X has shape (n_neurons, timeteps)
"""
n_neurons = len(A)
X = np.zeros((n_neurons, timesteps))
for t in range(timesteps-1):
if t % perturb_freq == 0:
X[:,t] = np.random.choice([0,1], size=n_neurons)
epsilon = np.random.multivariate_normal(np.zeros(n_neurons), np.eye(n_neurons))
X[:, t+1] = sigmoid(A.dot(X[:,t]) + epsilon) # we are using helper function sigmoid
return X
```
Take a look at the the update equation. While the rest of the function is the same as the ``simulate_neurons`` function in the previous exercise, every time step we now additionally include:
```
if t % perturb_freq == 0:
X[:,t] = np.random.choice([0,1], size=n_neurons)
```
Pretty serious perturbation, huh? You don't want that going on in your brain.
**Now visually compare the dynamics:** Run this next cell and see if you can spot how the dynamics have changed.
```
timesteps = 5000 # Simulate for 5000 timesteps.
# Simulate our dynamical system for the given amount of time
X_perturbed = simulate_neurons_perturb(A, timesteps)
# Plot our standard versus perturbed dynamics
fig, axs = plt.subplots(1,2, figsize=(15,4))
axs[0].imshow(X[:,:10])
axs[1].imshow(X_perturbed[:,:10])
axs[0].set_ylabel("Neuron",fontsize=15)
axs[1].set_xlabel("Timestep",fontsize=15)
axs[0].set_xlabel("Timestep",fontsize=15);
axs[0].set_title("Standard dynamics",fontsize=15)
axs[1].set_title("Perturbed dynamics",fontsize=15);
```
## Exercise 3: Using perturbed dynamics to recover connectivity
From the above perturbed dynamics, write a function that recovers the causal effect of all neurons upon a single neuron chosen. Remember from above you're calculating:
$$
\delta_{x^i\to x^j} \approx \frac1N \sum_i^N[x_{t+1}^j | x^i_t=1] - \frac1N \sum_i^N[x_{t+1}^j | x^i_t=0]]
$$
We wrote a code skeleton for you. It iterates through all neurons and gets the effect of `neuron_idx` upon `selected_neuron`.
```
def get_perturbed_connectivity_single_neuron(perturbed_X, perturb_freq, selected_neuron):
"""
Computes the connectivity matrix for the selected neuron using differences in means.
Args:
perturbed_X (np.ndarray): the perturbed dynamical system matrix of shape (n_neurons, timesteps)
perturb_freq (int): the perturbation frequency (2 means perturb every other timestep)
selected_neuron (int): the index of the neuron we want to estimate connectivity for
Returns:
estimated_connectivity (np.ndarray): estimated connectivity for the selected neuron, of shape (n_neurons,)
"""
neuron_perturbations = perturbed_X[selected_neuron, ::perturb_freq] # extract the perturbations of neuron 1
all_neuron_output = perturbed_X[:, 1::perturb_freq] # extract the observed outcomes of all the neurons
estimated_connectivity = np.zeros(n_neurons) # our stored estimated connectivity matrix
for neuron_idx in range(n_neurons):
selected_neuron_output = all_neuron_output[neuron_idx, :]
one_idx = np.argwhere(neuron_perturbations == 1)
zero_idx = np.argwhere(neuron_perturbations == 0)
########################################################################
## TODO: Insert your code here to compute the neuron activation from perturbations.
## Recall from above that this is E[Y | T=1] - E[Y | T=0].
## Y in this case is selected_neuron_output.
## T=1 and T=0 in this case are one_idx and zero_idx, respectively.
##
## Function Hints:
## expected value -> np.mean()
# Fill out function and remove
raise NotImplementedError("Complete the exercise of causal effects")
########################################################################
difference_in_means = ...
estimated_connectivity[neuron_idx] = difference_in_means
return estimated_connectivity
# ## Uncomment to test
# # Initialize the system
# n_neurons = 6
# timesteps = 5000 # Simulate for 5000 timesteps.
# perturb_freq = 2 # perturb the system every other time step
# # Simulate our perturbed dynamical system for the given amount of time
# perturbed_X = simulate_neurons_perturb(A, timesteps, perturb_freq=perturb_freq)
# # we'll measure the connectivity of neuron 1
# selected_neuron = 1
# estimated_connectivity = get_perturbed_connectivity_single_neuron(perturbed_X, perturb_freq, selected_neuron)
# #Now plot
# fig, axs = plt.subplots(1,2, figsize=(10,5))
# plot_connectivity_matrix(np.expand_dims(estimated_connectivity, axis=1), ax=axs[0])
# axs[0].set(title="Estimated connectivity", ylabel="Neuron")
# plot_connectivity_matrix(A[:, [selected_neuron]], ax=axs[1])
# axs[1].set(title="True connectivity")
# to_remove solution
def get_perturbed_connectivity_single_neuron(perturbed_X, perturb_freq, selected_neuron):
"""
Computes the connectivity matrix for the selected neuron using differences in means.
Args:
perturbed_X (np.ndarray): the perturbed dynamical system matrix of shape (n_neurons, timesteps)
perturb_freq (int): the perturbation frequency (2 means perturb every other timestep)
selected_neuron (int): the index of the neuron we want to estimate connectivity for
Returns:
estimated_connectivity (np.ndarray): estimated connectivity for the selected neuron, of shape (n_neurons,)
"""
neuron_perturbations = perturbed_X[selected_neuron, ::perturb_freq] # extract the perturbations of neuron 1
all_neuron_output = perturbed_X[:, 1::perturb_freq] # extract the observed outcomes of all the neurons
estimated_connectivity = np.zeros(n_neurons) # our stored estimated connectivity matrix
for neuron_idx in range(n_neurons):
selected_neuron_output = all_neuron_output[neuron_idx, :]
one_idx = np.argwhere(neuron_perturbations == 1)
zero_idx = np.argwhere(neuron_perturbations == 0)
difference_in_means = np.mean(selected_neuron_output[one_idx]) - np.mean(selected_neuron_output[zero_idx])
estimated_connectivity[neuron_idx] = difference_in_means
return estimated_connectivity
# Initialize the system
n_neurons = 6
timesteps = 5000 # Simulate for 5000 timesteps.
perturb_freq = 2 # perturb the system every other time step
# Simulate our perturbed dynamical system for the given amount of time
perturbed_X = simulate_neurons_perturb(A, timesteps, perturb_freq=perturb_freq)
# we'll measure the connectivity of neuron 1
selected_neuron = 1
estimated_connectivity = get_perturbed_connectivity_single_neuron(perturbed_X, perturb_freq, selected_neuron)
#Now plot
with plt.xkcd():
fig, axs = plt.subplots(1,2, figsize=(10,5))
plot_connectivity_matrix(np.expand_dims(estimated_connectivity, axis=1), ax=axs[0])
axs[0].set(title="Estimated connectivity", ylabel="Neuron")
plot_connectivity_matrix(A[:, [selected_neuron]], ax=axs[1])
axs[1].set(title="True connectivity")
```
Let's quantify how close these two are by correlating the estimated and true connectivity matrices:
```
# We should see almost perfect correlation between our estimates and the true connectivity
np.corrcoef(A[:, selected_neuron], estimated_connectivity)[1,0]
```
## Measuring how perturbations recover the entire connectivity matrix
Nice job! You just estimated connectivity for a single neuron.
We're now going to use the same strategy for all neurons at once. We coded up this function for you. Don't worry about how it works. (If you're curious and have extra time, scroll to the explanation at the bottom).
```
#@title Define perturbation estimator for all neurons
def get_perturbed_connectivity_all_neurons(perturbed_X, perturb_freq):
"""
Estimates the connectivity matrix of perturbations through stacked correlations.
Args:
perturbed_X (np.ndarray): the simulated dynamical system X of shape (n_neurons, timesteps)
perturb_freq (int): the perturbation frequency (2 means perturb every other timestep)
Returns:
R (np.ndarray): the estimated connectivity matrix of shape (n_neurons, n_neurons)
"""
# select perturbations (P) and outcomes (O)
P = perturbed_X[:, ::perturb_freq]
O = perturbed_X[:, 1::perturb_freq]
# stack perturbations and outcomes into a 2n by (timesteps / 2) matrix
S = np.concatenate([P, O], axis=0)
# select the perturbation -> outcome block of correlation matrix (upper right)
R = np.corrcoef(S)[:n_neurons, n_neurons:]
return R
# Initializes the system
n_neurons = 6
A = create_connectivity(n_neurons) # we are invoking a helper function that generates our nxn causal connectivity matrix.
timesteps = 5000 # Simulate for 5000 timesteps.
perturb_freq = 2 # perturb the system every other time step
# Simulate our perturbed dynamical system for the given amount of time
perturbed_X = simulate_neurons_perturb(A, timesteps, perturb_freq)
# Get our estimated connectivity matrix
R = get_perturbed_connectivity_all_neurons(perturbed_X, perturb_freq)
# Let's visualize the true connectivity and estimated connectivity together
fig, axs = plt.subplots(1,2, figsize=(10,5))
see_neurons(A, axs[0]) # we are invoking a helper function that visualizes the connectivity matrix
plot_connectivity_matrix(A, ax=axs[1])
plt.title("True connectivity matrix A")
plt.show()
fig, axs = plt.subplots(1,2, figsize=(10,5))
see_neurons(R.T,axs[0]) # we are invoking a helper function that visualizes the connectivity matrix
plot_connectivity_matrix(R.T, ax=axs[1])
plt.title("Estimated connectivity matrix R")
```
**Quantifying success**: We can calculate the correlation coefficient between the elements of the two matrices
```
np.corrcoef(A.transpose().flatten(), R.flatten())[1,0]
```
We do a good job recovering the true causality of the system!
## Optional Note: how we compute the estimated connectivity matrix
**This is an optional explanation of what the code is doing in `connectivity_from_perturbed_simulation()`**
First, we compute an estimated connectivity matrix $R$. We extract
perturbation matrix $P$ and outcomes matrix $O$:
$$
P = \begin{bmatrix}
\mid & \mid & ... & \mid \\
x_0 & x_2 & ... & x_T \\
\mid & \mid & ... & \mid
\end{bmatrix}_{n \times T/2}
$$
$$
O = \begin{bmatrix}
\mid & \mid & ... & \mid \\
x_1 & x_3 & ... & x_{T-1} \\
\mid & \mid & ... & \mid
\end{bmatrix}_{n \times T/2}
$$
And calculate the correlation of matrix $S$, which is $P$ and $O$ stacked on each other:
$$
S = \begin{bmatrix}
P \\
O
\end{bmatrix}_{2n \times T/2}
$$
We then extract $R$ as the upper right $n \times n$ block of $corr(S)$:
This is because the upper right block corresponds to the estimated perturbation effect on outcomes for each pair of neurons in our system.
# Summary
In this tutorial, we learned about how to define and estimate causality using pertubations. In particular we:
1) Learned how to simulate a system of connected neurons
2) Learned how to estimate the connectivity between neurons by directly perturbing neural activity
| github_jupyter |
Loo site.
# Process input data
> This part mainly load dataset, and construct train & test dataset.
```
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow.keras import layers
from sklearn.metrics import r2_score, mean_squared_error, mean_absolute_error
from scipy import stats
from scipy.stats import pearsonr
from CausalLSTM.data import load_and_qc_data
np.random.seed(1)
tf.compat.v1.set_random_seed(13)
site_name = 'Blo'
ROOT = '/hard/lilu/Fluxnet/'#'/Users/lewlee/Desktop/'
PATH = ROOT + 'FLX_US-Blo_FLUXNET2015_FULLSET_DD_1997-2007_1-3.csv'
#'FLX_FI-Sod_FLUXNET2015_FULLSET_DD_2001-2014_1-3.csv'
#'FLX_NL-Loo_FLUXNET2015_FULLSET_DD_1996-2013_1-3.csv'
#'FLX_AU-DaP_FLUXNET2015_FULLSET_DD_2007-2013_2-3.csv'
#'FLX_CH-Lae_FLUXNET2015_FULLSET_DD_2004-2014_1-3.csv'
#'FLX_ZA-Kru_FLUXNET2015_FULLSET_DD_2000-2010_1-3.csv'
#'FLX_US-Blo_FLUXNET2015_FULLSET_DD_1997-2007_1-3.csv'
#'FLX_NL-Loo_FLUXNET2015_FULLSET_DD_1996-2013_1-3.csv'
#'LW_IN_F_MDS'
df, label = load_and_qc_data(
PATH=PATH,
feature_params=['TA_F_MDS','SW_IN_F_MDS','P_F','PA_F','VPD_F_MDS','CO2_F_MDS','WS_F','TS_F_MDS_1','SWC_F_MDS_1'],#'SW_OUT','LW_OUT',
label_params=['SWC_F_MDS_1'],
qc_params=['SWC_F_MDS_1_QC']
)
```
Prepare int forcing data of fluxnet data of CoLM
```
print('The simple describe of dataset is {}:'.format(len(label)))
print('The num of nan is {}'.format(np.sum(np.isnan(label))))
df.describe().transpose()
# select manully
fig = plt.figure(figsize=(40,5))
n = len(df)
ax = plt.subplot(111)
plt.plot(np.arange(n), df['SWC_F_MDS_1'])
ax1 = ax.twinx()
ax1.bar(x=np.arange(n), height=df['P_F'], color='red')
from CausalLSTM.tree_causality import CausalTree
ct = CausalTree(num_features=9,
name_features=['TA_F_MDS','SW_IN_F_MDS','P_F','PA_F','VPD_F_MDS','CO2_F_MDS','WS_F','TS_F_MDS_1','SWC_F_MDS_1'],
corr_thresold=0.5,
mic_thresold=0.5,
flag=[1,1,0])
adjacency_matrix, tree, children, child_input_idx, child_state_idx = ct(np.array(df))
print(adjacency_matrix[:, :])
print(tree)
print(children)
print(child_input_idx)
print(child_state_idx)
column_indices = {name: i for i, name in enumerate(df.columns)}
train_df = df[0:int(n*0.8)]
test_df = df[int(n*0.8):]
num_features = df.shape[1]
print(column_indices)
print('length of train dataset is {}'.format(len(train_df)))
print('length of test dataset is {}'.format(len(test_df)))
train_mean = train_df.mean()
train_std = train_df.std()
train_df = (train_df - train_mean) / train_std
test_df = (test_df - train_mean) / train_std
df_std = (df - train_mean) / train_std
df_std = df_std.melt(var_name='Column', value_name='Normalized')
plt.figure(figsize=(12, 6))
ax = sns.violinplot(x='Column', y='Normalized', data=df_std)
_ = ax.set_xticklabels(df.keys(), rotation=90)
def generate(inputs, outputs,
len_input, len_output, window_size):
"""Generate inputs and outputs for SMNET."""
# caculate the last time point to generate batch
end_idx = inputs.shape[0] - len_input - len_output - window_size
# generate index of batch start point in order
batch_start_idx = range(end_idx)
# get batch_size
batch_size = len(batch_start_idx)
# generate inputs
input_batch_idx = [
(range(i, i + len_input)) for i in batch_start_idx]
inputs = np.take(inputs, input_batch_idx, axis=0). \
reshape(batch_size, len_input,
inputs.shape[1])
# generate outputs
output_batch_idx = [
(range(i + len_input + window_size, i + len_input + window_size +
len_output)) for i in batch_start_idx]
outputs = np.take(outputs, output_batch_idx, axis=0). \
reshape(batch_size, len_output,
outputs.shape[1])
return inputs, outputs
train_x , train_y = generate(train_df.values[:,:], train_df.values[:,-1][:, np.newaxis], 10, 1, 7)
test_x, test_y = generate(test_df.values[:,:], test_df.values[:,-1][:, np.newaxis], 10, 1, 7)
print('the shape of train dataset is {}'.format(train_x.shape))
print('the shape of test dataset is {}'.format(test_x.shape))
```
## Casuality LSTM vs LSTM
> **tensorflow edition**
```
train_x = train_x[:3000]
train_y = train_y[:3000]
test_x = test_x[:700]
test_y = test_y[:700]
batch_size = 50
from sklearn.ensemble import RandomForestRegressor
model = RandomForestRegressor()
model.fit(train_x.reshape(-1, 90), train_y.reshape(-1,1))
y_pred_rf = model.predict(test_x.reshape(-1, 90))
print('r2 of test dataset is {}'.format(
r2_score(np.squeeze(y_pred_rf), np.squeeze(test_y))))
from tensorflow.keras.callbacks import ModelCheckpoint
from tensorflow.keras.callbacks import ReduceLROnPlateau
checkpoint = ModelCheckpoint(filepath='/Users/lewlee/Desktop/log/',
monitor = 'val_loss',
save_best_only='True',
save_weights_only='True')
lr = ReduceLROnPlateau(monitor='val_loss',
factor=0.1,
patience=10,
verbose=0,
mode='auto',
epsilon=0.0001,
cooldown=0,
min_lr=0)
model = LSTM(16, batch_size)
model.compile(optimizer=tf.keras.optimizers.Adam(),loss=['mse'])
history = model.fit(train_x, np.squeeze(train_y), batch_size=batch_size, epochs=50, validation_split=0.2,callbacks=[checkpoint,lr])
y_pred_lstm = model.predict(test_x, batch_size=batch_size)
print('r2 of test dataset is {}'.format(r2_score(np.squeeze(y_pred_lstm), np.squeeze(test_y))))
import matplotlib.pyplot as plt
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
model = CausalLSTM(num_nodes=len(children),
num_hiddens=16,
children=children,
child_input_idx=child_input_idx,
child_state_idx=child_state_idx,
input_len=10,
batch_size=batch_size)
model.compile(optimizer=tf.keras.optimizers.Adam(),loss=['mse'])
history = model.fit(train_x, np.squeeze(train_y), batch_size=batch_size, epochs=50,validation_split=0.2,callbacks=[checkpoint,lr])
y_pred_clstm = model.predict(test_x, batch_size=batch_size)
print('r2 of test dataset is {}'.format(r2_score(np.squeeze(y_pred_clstm), np.squeeze(test_y))))
import matplotlib.pyplot as plt
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
def unnormalized(inputs):
return inputs*train_std[-1]+train_mean[-1]
y_pred_lstm = np.squeeze(unnormalized(y_pred_lstm))
y_pred_clstm = np.squeeze(unnormalized(y_pred_clstm))
y_pred_rf = np.squeeze(unnormalized(y_pred_rf))
y_test = np.squeeze(unnormalized(test_y))
OUT_PATH = '/work/lilu/CausalLSTM/figures/'
import os
if not os.path.exists(OUT_PATH + site_name):
os.mkdir(OUT_PATH + site_name)
import netCDF4 as nc
dataset = nc.Dataset('/Users/lewlee/Desktop/ZA-Kru_2D_Fluxes_2001-2010.nc')
soil_moisture = dataset.variables['f_wliq_soisno'][:]
sm_lv1 = soil_moisture[-740:, 5, 0, 0][:700] # 0-0.0175m
sm_lv2 = soil_moisture[-740:, 6, 0, 0][:700] # 0.0175-(0.0175+0.0173)m
sm_lv3 = soil_moisture[-740:, 7, 0, 0][:700] # (0.0175+0.0173)-(0.0175+0.0173+0.0283)m
sm = (sm_lv1*1.75+sm_lv2*1.73+sm_lv3*2.83)/(1.75+1.73+2.83)
plt.figure(figsize=(20,5))
ax = plt.subplot(111)
plt.plot(y_pred_lstm,linewidth=3)
plt.plot(y_pred_clstm, linewidth=3)
plt.plot(y_test, linewidth=5)
plt.legend(['LSTM','Causal LSTM','Observation'])
ax.set_xlabel('Time (day)', fontsize=20)
ax.set_ylabel('Soil moisture (Volumetric)', fontsize=20)
ax1 = ax.twinx()
test_df_ = test_df['P_F']*train_std['P_F']+train_mean['P_F']
test_df_ = np.array(test_df_[:1000])
ax1.bar(x=np.arange(len(np.array(test_df_))), height=np.array(test_df_), color='red')
ax1.set_ylim(0,max(test_df_)+20)
ax1.set_xlabel('Time (day)', fontsize=20)
ax1.set_ylabel('Precipitation (mm)', fontsize=20)
plt.savefig(OUT_PATH + site_name +'/time_series_'+site_name+'.pdf')
def linear_(x,y):
a, b = np.polyfit(x, y, deg=1)
y_est = a * x + b
y_err = x.std() * np.sqrt(1/len(x) + (x - x.mean())**2 / np.sum((x - x.mean())**2))
return y_est, y_err, a, b
y_est_lstm, y_err_lstm, a_lstm, b_lstm = linear_(np.squeeze(y_pred_lstm), np.squeeze(y_test))
y_est_clstm, y_err_clstm, a_clstm, b_clstm = linear_(np.squeeze(y_pred_clstm), np.squeeze(y_test))
min_, max_ = np.min(y_pred_lstm), np.max(y_pred_lstm)
plt.figure(figsize=(8,8))
ax1 = plt.subplot(221)
ax1.spines['top'].set_linewidth(2)
ax1.spines['right'].set_linewidth(2)
ax1.spines['left'].set_linewidth(2)
ax1.spines['bottom'].set_linewidth(2)
ax1.plot((0, 1), (0, 1), transform=ax1.transAxes, ls='--',c='k', label="1:1 line")
ax1.scatter(y_pred_lstm, y_test)
ax1.plot(y_pred_lstm, y_est_lstm, '-', color='red')
ax1.fill_between(y_pred_lstm, y_est_lstm - y_err_lstm, y_est_lstm + y_err_lstm, alpha=0.2)
plt.xlim(min_-10,max_+10)
plt.ylim(min_-10,max_+10)
ax1.set_xlabel('LSTM',fontsize=20)
ax1.set_ylabel('Fluxnet-'+site_name,fontsize=20)
plt.text(min_-8,max_+8, '$R=%.2f$' % (pearsonr(np.squeeze(y_pred_lstm), np.squeeze(y_test))[0]))
plt.text(min_-8,max_+6, '$RMSE=%.2f$' % (np.sqrt(mean_squared_error(np.squeeze(y_pred_lstm), np.squeeze(y_test)))))
plt.text(min_-8,max_+4, '$MAE=%.2f$' % (np.sqrt(mean_absolute_error(np.squeeze(y_pred_lstm), np.squeeze(y_test)))))
plt.text(min_-8,max_+2, '$Y = %.1fX + %.2f$' % (a_lstm, b_lstm))
ax2 = plt.subplot(222)
ax2.spines['top'].set_linewidth(2)
ax2.spines['right'].set_linewidth(2)
ax2.spines['left'].set_linewidth(2)
ax2.spines['bottom'].set_linewidth(2)
ax2.scatter(y_pred_clstm, y_test)
plt.xlim(min_-10,max_+10)
plt.ylim(min_-10,max_+10)
ax2.plot((0, 1), (0, 1), transform=ax2.transAxes, ls='--',c='k', label="1:1 line")
ax2.plot(y_pred_clstm, y_est_clstm, '-', color='red')
ax2.fill_between(y_pred_clstm, y_est_clstm - y_err_clstm, y_est_clstm + y_err_clstm, alpha=0.2)
ax2.set_xlabel('Causal LSTM',fontsize=20)
plt.text(min_-8,max_+8, '$R=%.2f$' % (pearsonr(np.squeeze(y_pred_clstm), np.squeeze(y_test))[0]))
plt.text(min_-8,max_+6, '$RMSE=%.2f$' % (np.sqrt(mean_squared_error(np.squeeze(y_pred_clstm), np.squeeze(y_test)))))
plt.text(min_-8,max_+4, '$MAE=%.2f$' % (np.sqrt(mean_absolute_error(np.squeeze(y_pred_clstm), np.squeeze(y_test)))))
plt.text(min_-8,max_+2, '$Y = %.1fX + %.2f$' % (a_clstm, b_clstm))
plt.savefig(OUT_PATH + site_name +'/scatter_'+site_name+'.pdf')
```
| github_jupyter |
# 7. Поиск оценок. Метод моментов. Правдоподобие.
# Теория
* [Метод моментов](https://nsu.ru/mmf/tvims/chernova/ms/lec/node12.html)
* [Максимального правдоподобия](https://nsu.ru/mmf/tvims/chernova/ms/lec/node14.html)
## Метод моментов.
Пусть у нас есть выборка $X_1, \ldots, X_n$ из распределения с параметром $\theta$.
Если мы сможем подобрать такую функцию $g$, такую что
$$\mathbb{E}(g(X_i)) = h(\theta)$$
где $h$ - обратима, то оценка методом моментов находится как
$$\hat\theta = h^{-1}(\overline{g(X)})$$
Аналогично происходит, если у нас несколько параметров $\theta = (\mu, \sigma)$.
Тогда мы составляем систему из нескольких различных функций $g$(по количеству параметров) и решаем ее:
$$\begin{cases}
\mathbb{E}(g_1(X_i)) = h_1(\theta) \\
\mathbb{E}(g_2(X_i)) = h_2(\theta) \\
\ldots \\
\mathbb{E}(g_n(X_i)) = h_n(\theta)
\end{cases}$$
Метод называется методом моментов, так как за функции $g_i$ обычно берутся функции моментов
$$g_k(x) = x^k$$
Так как функции $h^{-1}$ - непрерывные, то оценки полученные методом моментов - состоятельные
### Метод максимального правдоподобия
Пусть у нас есть выборка $\textbf X^n = \{X_1, \ldots, X_n\}$ из распределения $X$ с параметром $\theta$.
Идея: давайте посчитаем вероятность того, что у нас выпала такая последовательность результатов и назовем
эту величину правдоподобием.
$$Likelihood = L(X^n; \theta) = \begin{cases}
\prod_{i=1}^{n}\rho_{X}(X_i), & \text{Для непрерывных распределений} \\
\prod_{i=1}^{n}P(X = X_i), & \text{Для дискретных распределений} \\
\end{cases}$$
Распределение $X$ как-то зависит от параметра $\theta$ и мы хотим максимизировать правдоподобие по параметру $\theta$
Итого оценка максимального правдоподобия:
$$\hat\theta = argmax_{\theta}L(\textbf{X}^n; \theta)$$
Понятное дело, что $\theta$ может быть вектором параметров ( Например $\theta = (\mu, \sigma)$ для $\mathcal(\mu, \sigma^2)$).
Тогда функцию нужно максимизировать по нескольким параметрам.
# На паре
1. Пусть выборка $X_1, ..., X_n$ порождена распределением $U[0, \theta]$.
Оцените параметр $\theta$ с помощью:
* Метода моментов (функцией $g(x) = x$ и $g(x) = x^k$)
* Метода максимального правдоподобия
2. Пусть выборка $X_1, \ldots, X_n$ порождена распределением $\mathcal{N}(\mu, \sigma^2)$.
С помощью метода моментов найдите оценки параметров $\hat \mu, \hat\sigma$.
3. Используя метод моментов и правдоподобием, постройте оценку $\lambda > 1$ по выборке из распределения Пуассона с параметром $\lambda$.
4. Пусть выборка $X_1, \ldots, X_n$ порождена распределением с плотностью
$$f(x) = \begin{cases}
\frac{\beta \alpha^{\beta}}{x^{\beta + 1}}, x \ge \alpha \\
0, x < \alpha
\end{cases}$$
Здесь $\alpha > 0$ и $\beta > 2$. С помощью **любого** метода постройте оценку параметров $\alpha$ и $\beta$.
## Домашка
1. (1) Найти оценку **методом моментов** параметра $p \in (0, 1)$ геометрического распределения.
2. (1) Пусть выборка $X_1, \ldots, X_n$ порождена распределением:
$$\begin{cases}
P(X_i = 1) = p_1\\
P(X_i = 2) = p_2\\
P(X_i = 3) = p_3\\
\end{cases}$$
$p_1 + p_2 + p_3 = 1$. Постройте оценку параметров $p_1, p_2, p_3$ **методом максимального правдоподобия**.
3. (1)Пусть дана выборка из распределения с плотностью
$$f_{\alpha}(y) =
\begin{cases}
3y^2\alpha^{-3}e^{-(\frac{y}{\alpha})^3} & y\geq 0\\
0 & y < 0
\end{cases}$$
Построить оценку параметра $\alpha > 0$ с помощью **метода моментов** используя k-ый момент $g(y) = y^k$
4. (1) Пусть выборка $X_1, ..., X_n$ порождена равномерным на отрезке $[\theta; 2\theta]$ распределением. Постройте оценку параметра $\theta$ **Методом моментов**.
5. (1) Пусть выборка $X_1, \ldots, X_n$ порождена распределением с плотностью $f(x)$: $$ f(x) = \frac{1}{2\sigma} e^{-\frac{\left|x - \mu\right|}{\sigma}}$$
Постройте оценку **методом максимального правдоподобия** для вектора параметров $\left(\mu, \sigma\right)$.
# Гробы
1. Используя метод моментов, оцените параметр $\theta$ равномерного распределения на отрезке:
* $\left[\theta - 1; \theta + 1\right]$, $\theta \in R$
* $\left[-\theta; \theta\right]$, $\theta > 0$.
2. Пусть выборка $X_1, ..., X_n$ порождена распределением с плотностью $f_{\theta}(x)$:
$f_{\theta}(x) = f(x - \theta)$,
где функция $f(x)$ имеет единственный максимум в точке $x = 0$. Постройте оценку максимального правдоподобия
$\hat \theta$ параметра сдвига $\theta$ по одному наблюдению $X_1$.
3. Постройте оценки параметров с помощью метода моментов и метода правдоподобия для Гамма-распределения с двумя параметрами:
$$f_{k, \theta}(y) =
\begin{cases}
x^{k-1}\frac{e^{-\frac{x}{\theta}}}{\theta^k\Gamma(k)} & x\geq 0\\
0 & x < 0
\end{cases}$$
4. Пусть выборка $X_1, ..., X_n$ порождена распределением Коши. Доказать, что медиана - оценка метода максимального прадободобия.
(P.S. - Не забудьте, что у распределения Коши не существует матожидания:))
5. Пусть выборка $X_1, ..., X_n$ порождена распределением:
$$\begin{cases}
P(X_i = 1) = p_1\\
P(X_i = 2) = p_2\\
P(X_i = 3) = p_3\\
P(X_i = 4) = p_4\\
\end{cases}$$
$p_1 + p_2 + p_3 + p_4 = 1$. Постройте оценку параметров $p_1, p_2, p_3, p_4$ методом максимального правдоподобия.
-----------
| github_jupyter |
```
%matplotlib inline
```
# Extract data from mongoDB and generate geometries, write back
This notebook goes through raw OSM data (raw ways and nodes) and writes back all Polygons and Linestrings to a derived ways table, including lengths and areas.
**Derived ways is dropped and re-inserted!**
Created on: 2016-10-27
Last update: 2016-12-09
Contact: michael.szell@moovel.com, michael.szell@gmail.com (Michael Szell)
## Preliminaries
### Parameters
```
cityname = "boston"
```
### Imports
```
# preliminaries
from __future__ import unicode_literals
import sys
import csv
import os
import math
import pprint
pp = pprint.PrettyPrinter(indent=4)
from collections import defaultdict
import time
import datetime
import numpy as np
from scipy import stats
import pyprind
import itertools
import logging
from ast import literal_eval as make_tuple
from collections import OrderedDict
import json
from shapely.geometry import mapping, shape, LineString, LinearRing, Polygon, MultiPolygon
import shapely
import shapely.ops as ops
from functools import partial
import pyproj
from scipy import spatial
from haversine import haversine
import pymongo
from pymongo import MongoClient
# plotting stuff
import matplotlib.pyplot as plt
```
### Create folder structure
```
if not os.path.exists("citydata"):
os.makedirs("citydata")
if not os.path.exists("logs"):
os.makedirs("logs")
if not os.path.exists("output"):
os.makedirs("output")
if not os.path.exists("output/" + cityname + "/carin"):
os.makedirs("output/" + cityname + "/carin")
if not os.path.exists("output/" + cityname + "/carout"):
os.makedirs("output/" + cityname + "/carout")
if not os.path.exists("output/" + cityname + "/bikein"):
os.makedirs("output/" + cityname + "/bikein")
if not os.path.exists("output/" + cityname + "/bikeout"):
os.makedirs("output/" + cityname + "/bikeout")
```
### DB Connection
```
client = MongoClient()
db_raw = client[cityname+'_raw']
nodes_raw = db_raw['nodes']
cursor = nodes_raw.find({})
numnodes = cursor.count()
ways_raw = db_raw['ways']
cursor = ways_raw.find({})
numways = cursor.count()
db_derived = client[cityname+'_derived']
db_derived.drop_collection('ways')
db_derived = client[cityname+'_derived']
ways_derived = db_derived['ways']
```
## Polygons and Linestrings from raw to derived
```
bar = pyprind.ProgBar(numways, bar_char='█', update_interval=1)
nodesinserted = 0
nodestotal = 0
for i,way in enumerate(ways_raw.find()):
bar.update(item_id = i)
tempgeojson = {}
tempgeojson["_id"] = way["_id"]
try:
tempgeojson["properties"] = way['tags']
except:
tempgeojson["properties"] = {}
tempgeojson["type"] = "Feature"
tempgeojson["geometry"] = {"type":"", "coordinates":[]}
coords = []
for nodeid in way["nodes"]:
nodestotal += 1
for n in nodes_raw.find({"_id": nodeid}):
nodesinserted += 1
coords.append([n["loc"]["coordinates"][0], n["loc"]["coordinates"][1]])
tempgeojson["geometry"]["coordinates"] = coords
if way["nodes"][0] == way["nodes"][-1]:
tempgeojson["geometry"]["type"] = "Polygon"
else:
tempgeojson["geometry"]["type"] = "LineString"
ways_derived.insert_one(tempgeojson)
nodesmissing = nodestotal - nodesinserted
print("Done. Nodes missing: "+ str(nodesmissing) + ", out of " + str(nodestotal))
```
## Calculate lengths and areas of ways and save back to mongoDB
```
client = MongoClient()
db_derived = client[cityname+'_derived']
ways_derived = db_derived['ways']
cursor = ways_derived.find({"geometry.type": "LineString"})
numLineStrings = cursor.count()
bar = pyprind.ProgBar(numLineStrings, bar_char='█', update_interval=1)
for i,way in enumerate(ways_derived.find({"geometry.type": "LineString"})):
bar.update(item_id = i)
npway = np.asarray(way["geometry"]["coordinates"])
distances = [1000*haversine(npway[i][::-1], npway[i+1][::-1]) for i in range(npway.shape[0]-1)]
ways_derived.update_one({'_id': way["_id"]}, {"$set": {"properties_derived.length": sum(distances)}}, upsert=False)
cursor = ways_derived.find({"geometry.type": "Polygon"})
numPolygons = cursor.count()
bar = pyprind.ProgBar(numPolygons, bar_char='█', update_interval=1)
for i,way in enumerate(ways_derived.find({"geometry.type": "Polygon"})):
bar.update(item_id = i)
npway = np.asarray(way["geometry"]["coordinates"])
distances = [1000*haversine(npway[i][::-1], npway[i+1][::-1]) for i in range(npway.shape[0]-1)]
ways_derived.update_one({'_id': way["_id"]}, {"$set": {"properties_derived.length": sum(distances)}}, upsert=False)
# Following area calculating code from: http://gis.stackexchange.com/questions/127607/area-in-km-from-polygon-of-coordinates
try: # IllegalArgumentException: Invalid number of points in LinearRing found 3 - must be 0 or >= 4
geom = Polygon(npway)
geom_area = ops.transform(
partial(
pyproj.transform,
pyproj.Proj(init='EPSG:4326'),
pyproj.Proj(
proj='aea',
lat1=geom.bounds[1],
lat2=geom.bounds[3])),
geom)
# Export the area in m^2
ways_derived.update_one({'_id': way["_id"]}, {"$set": {"properties_derived.area": geom_area.area}}, upsert=False)
except:
print("Something went wrong: " + str(i))
pass
```
| github_jupyter |
# Similarity Based Methods
## Index Based
### Resource Allocation
```
import networkx as nx
edges = [[1,3],[2,3],[2,4],[4,5],[5,6],[5,7]]
G = nx.from_edgelist(edges)
preds = nx.resource_allocation_index(G,[(1,2),(2,5),(3,4)])
print(list(preds))
draw_graph(G)
```
### Jaccard Coefficient
```
import networkx as nx
edges = [[1,3],[2,3],[2,4],[4,5],[5,6],[5,7]]
G = nx.from_edgelist(edges)
preds = nx.jaccard_coefficient(G,[(1,2),(2,5),(3,4)])
print(list(preds))
draw_graph(G)
```
## Community Based
### Community Common Neighbor
```
import networkx as nx
edges = [[1,3],[2,3],[2,4],[4,5],[5,6],[5,7]]
G = nx.from_edgelist(edges)
G.nodes[1]["community"] = 0
G.nodes[2]["community"] = 0
G.nodes[3]["community"] = 0
G.nodes[4]["community"] = 1
G.nodes[5]["community"] = 1
G.nodes[6]["community"] = 1
G.nodes[7]["community"] = 1
preds = nx.cn_soundarajan_hopcroft(G,[(1,2),(2,5),(3,4)])
print(list(preds))
draw_graph(G)
```
### Community Common Neighbor
```
import networkx as nx
edges = [[1,3],[2,3],[2,4],[4,5],[5,6],[5,7]]
G = nx.from_edgelist(edges)
G.nodes[1]["community"] = 0
G.nodes[2]["community"] = 0
G.nodes[3]["community"] = 0
G.nodes[4]["community"] = 1
G.nodes[5]["community"] = 1
G.nodes[6]["community"] = 1
G.nodes[7]["community"] = 1
preds = nx.ra_index_soundarajan_hopcroft(G,[(1,2),(2,5),(3,4)])
print(list(preds))
draw_graph(G)
```
## Embedding based
```
import networkx as nx
import pandas as pd
edgelist = pd.read_csv("cora.cites", sep='\t', header=None, names=["target", "source"])
G = nx.from_pandas_edgelist(edgelist)
draw_graph(G)
from stellargraph.data import EdgeSplitter
edgeSplitter = EdgeSplitter(G)
graph_test, samples_test, labels_test = edgeSplitter.train_test_split(
p=0.1, method="global"
)
edgeSplitter = EdgeSplitter(graph_test, G)
graph_train, samples_train, labels_train = edgeSplitter.train_test_split(
p=0.1, method="global"
)
from node2vec import Node2Vec
from node2vec.edges import HadamardEmbedder
node2vec = Node2Vec(graph_train)
model = node2vec.fit()
edges_embs = HadamardEmbedder(keyed_vectors=model.wv)
train_embeddings = [edges_embs[str(x[0]),str(x[1])] for x in samples_train]
test_embeddings = [edges_embs[str(x[0]),str(x[1])] for x in samples_test]
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier(n_estimators=1000)
rf.fit(train_embeddings, labels_train);
from sklearn import metrics
y_pred = rf.predict(test_embeddings)
print('Precision:', metrics.precision_score(labels_test, y_pred))
print('Recall:', metrics.recall_score(labels_test, y_pred))
print('F1-Score:', metrics.f1_score(labels_test, y_pred))
import matplotlib.pyplot as plt
def draw_graph(G, node_names={}, node_size=500):
pos_nodes = nx.spring_layout(G)
nx.draw(G, pos_nodes, with_labels=True, node_size=node_size, edge_color='gray', arrowsize=30)
pos_attrs = {}
for node, coords in pos_nodes.items():
pos_attrs[node] = (coords[0], coords[1] + 0.08)
#nx.draw_networkx_labels(G, pos_attrs, font_family='serif', font_size=20)
plt.axis('off')
axis = plt.gca()
axis.set_xlim([1.2*x for x in axis.get_xlim()])
axis.set_ylim([1.2*y for y in axis.get_ylim()])
plt.show()
```
| github_jupyter |
```
# NOTE: PLEASE MAKE SURE YOU ARE RUNNING THIS IN A PYTHON3 ENVIRONMENT
import tensorflow as tf
print(tf.__version__)
# This is needed for the iterator over the data
# But not necessary if you have TF 2.0 installed
# !pip install tensorflow-gpu
# tf.enable_eager_execution()
!pip install -q tensorflow-datasets
import tensorflow_datasets as tfds
imdb, info = tfds.load("imdb_reviews", with_info=True, as_supervised=True)
import numpy as np
train_data, test_data = imdb['train'], imdb['test']
training_sentences = []
training_labels = []
testing_sentences = []
testing_labels = []
# str(s.tonumpy()) is needed in Python3 instead of just s.numpy()
for s,l in train_data:
training_sentences.append(str(s.numpy()))
training_labels.append(l.numpy())
for s,l in test_data:
testing_sentences.append(str(s.numpy()))
testing_labels.append(l.numpy())
training_labels_final = np.array(training_labels)
testing_labels_final = np.array(testing_labels)
vocab_size = 10000
embedding_dim = 16
max_length = 120
trunc_type='post'
oov_tok = "<OOV>"
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
tokenizer = Tokenizer(num_words = vocab_size, oov_token=oov_tok)
tokenizer.fit_on_texts(training_sentences)
word_index = tokenizer.word_index
sequences = tokenizer.texts_to_sequences(training_sentences)
padded = pad_sequences(sequences,maxlen=max_length, truncating=trunc_type)
testing_sequences = tokenizer.texts_to_sequences(testing_sentences)
testing_padded = pad_sequences(testing_sequences,maxlen=max_length)
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
def decode_review(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
print(decode_review(padded[1]))
print(training_sentences[1])
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim, input_length=max_length),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(6, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
model.summary()
num_epochs = 10
model.fit(padded, training_labels_final, epochs=num_epochs, validation_data=(testing_padded, testing_labels_final))
e = model.layers[0]
weights = e.get_weights()[0]
print(weights.shape) # shape: (vocab_size, embedding_dim)
import io
out_v = io.open('vecs.tsv', 'w', encoding='utf-8')
out_m = io.open('meta.tsv', 'w', encoding='utf-8')
for word_num in range(1, vocab_size):
word = reverse_word_index[word_num]
embeddings = weights[word_num]
out_m.write(word + "\n")
out_v.write('\t'.join([str(x) for x in embeddings]) + "\n")
out_v.close()
out_m.close()
try:
from google.colab import files
except ImportError:
pass
else:
files.download('vecs.tsv')
files.download('meta.tsv')
sentence = "I really think this is amazing. honest."
sequence = tokenizer.texts_to_sequences(sentence)
print(sequence)
```
| github_jupyter |
# RadarCOVID-Report
## Data Extraction
```
import datetime
import json
import logging
import os
import shutil
import tempfile
import textwrap
import uuid
import matplotlib.ticker
import numpy as np
import pandas as pd
import seaborn as sns
%matplotlib inline
current_working_directory = os.environ.get("PWD")
if current_working_directory:
os.chdir(current_working_directory)
sns.set()
matplotlib.rcParams["figure.figsize"] = (15, 6)
extraction_datetime = datetime.datetime.utcnow()
extraction_date = extraction_datetime.strftime("%Y-%m-%d")
extraction_previous_datetime = extraction_datetime - datetime.timedelta(days=1)
extraction_previous_date = extraction_previous_datetime.strftime("%Y-%m-%d")
extraction_date_with_hour = datetime.datetime.utcnow().strftime("%Y-%m-%d@%H")
```
### Constants
```
spain_region_country_name = "Spain"
spain_region_country_code = "ES"
backend_generation_days = 7 * 2
daily_summary_days = 7 * 4 * 3
daily_plot_days = 7 * 4
tek_dumps_load_limit = daily_summary_days + 1
```
### Parameters
```
active_region_parameter = os.environ.get("RADARCOVID_REPORT__ACTIVE_REGION")
if active_region_parameter:
active_region_country_code, active_region_country_name = \
active_region_parameter.split(":")
else:
active_region_country_code, active_region_country_name = \
spain_region_country_code, spain_region_country_name
```
### COVID-19 Cases
```
confirmed_df = pd.read_csv("https://covid19tracking.narrativa.com/csv/confirmed.csv")
radar_covid_countries = {active_region_country_name}
confirmed_df = confirmed_df[confirmed_df["Country_EN"].isin(radar_covid_countries)]
confirmed_df = confirmed_df[pd.isna(confirmed_df.Region)]
confirmed_df.head()
confirmed_country_columns = list(filter(lambda x: x.startswith("Country_"), confirmed_df.columns))
confirmed_regional_columns = confirmed_country_columns + ["Region"]
confirmed_df.drop(columns=confirmed_regional_columns, inplace=True)
confirmed_df.head()
confirmed_df = confirmed_df.sum().to_frame()
confirmed_df.tail()
confirmed_df.reset_index(inplace=True)
confirmed_df.columns = ["sample_date_string", "cumulative_cases"]
confirmed_df.sort_values("sample_date_string", inplace=True)
confirmed_df["new_cases"] = confirmed_df.cumulative_cases.diff()
confirmed_df["covid_cases"] = confirmed_df.new_cases.rolling(7).mean().round()
confirmed_df.tail()
extraction_date_confirmed_df = \
confirmed_df[confirmed_df.sample_date_string == extraction_date]
extraction_previous_date_confirmed_df = \
confirmed_df[confirmed_df.sample_date_string == extraction_previous_date].copy()
if extraction_date_confirmed_df.empty and \
not extraction_previous_date_confirmed_df.empty:
extraction_previous_date_confirmed_df["sample_date_string"] = extraction_date
extraction_previous_date_confirmed_df["new_cases"] = \
extraction_previous_date_confirmed_df.covid_cases
extraction_previous_date_confirmed_df["cumulative_cases"] = \
extraction_previous_date_confirmed_df.new_cases + \
extraction_previous_date_confirmed_df.cumulative_cases
confirmed_df = confirmed_df.append(extraction_previous_date_confirmed_df)
confirmed_df["covid_cases"] = confirmed_df.covid_cases.fillna(0).astype(int)
confirmed_df.tail()
confirmed_df[["new_cases", "covid_cases"]].plot()
```
### Extract API TEKs
```
from Modules.ExposureNotification import exposure_notification_io
raw_zip_path_prefix = "Data/TEKs/Raw/"
fail_on_error_backend_identifiers = [active_region_country_code]
multi_region_exposure_keys_df = \
exposure_notification_io.download_exposure_keys_from_backends(
generation_days=backend_generation_days,
fail_on_error_backend_identifiers=fail_on_error_backend_identifiers,
save_raw_zip_path_prefix=raw_zip_path_prefix)
multi_region_exposure_keys_df["region"] = multi_region_exposure_keys_df["backend_identifier"]
multi_region_exposure_keys_df.rename(
columns={
"generation_datetime": "sample_datetime",
"generation_date_string": "sample_date_string",
},
inplace=True)
multi_region_exposure_keys_df.head()
early_teks_df = multi_region_exposure_keys_df[
multi_region_exposure_keys_df.rolling_period < 144].copy()
early_teks_df["rolling_period_in_hours"] = early_teks_df.rolling_period / 6
early_teks_df[early_teks_df.sample_date_string != extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
early_teks_df[early_teks_df.sample_date_string == extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
multi_region_exposure_keys_df = multi_region_exposure_keys_df[[
"sample_date_string", "region", "key_data"]]
multi_region_exposure_keys_df.head()
active_regions = \
multi_region_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
active_regions
multi_region_summary_df = multi_region_exposure_keys_df.groupby(
["sample_date_string", "region"]).key_data.nunique().reset_index() \
.pivot(index="sample_date_string", columns="region") \
.sort_index(ascending=False)
multi_region_summary_df.rename(
columns={"key_data": "shared_teks_by_generation_date"},
inplace=True)
multi_region_summary_df.rename_axis("sample_date", inplace=True)
multi_region_summary_df = multi_region_summary_df.fillna(0).astype(int)
multi_region_summary_df = multi_region_summary_df.head(backend_generation_days)
multi_region_summary_df.head()
multi_region_without_active_region_exposure_keys_df = \
multi_region_exposure_keys_df[multi_region_exposure_keys_df.region != active_region_country_code]
multi_region_without_active_region = \
multi_region_without_active_region_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
multi_region_without_active_region
exposure_keys_summary_df = multi_region_exposure_keys_df[
multi_region_exposure_keys_df.region == active_region_country_code]
exposure_keys_summary_df.drop(columns=["region"], inplace=True)
exposure_keys_summary_df = \
exposure_keys_summary_df.groupby(["sample_date_string"]).key_data.nunique().to_frame()
exposure_keys_summary_df = \
exposure_keys_summary_df.reset_index().set_index("sample_date_string")
exposure_keys_summary_df.sort_index(ascending=False, inplace=True)
exposure_keys_summary_df.rename(columns={"key_data": "shared_teks_by_generation_date"}, inplace=True)
exposure_keys_summary_df.head()
```
### Dump API TEKs
```
tek_list_df = multi_region_exposure_keys_df[
["sample_date_string", "region", "key_data"]].copy()
tek_list_df["key_data"] = tek_list_df["key_data"].apply(str)
tek_list_df.rename(columns={
"sample_date_string": "sample_date",
"key_data": "tek_list"}, inplace=True)
tek_list_df = tek_list_df.groupby(
["sample_date", "region"]).tek_list.unique().reset_index()
tek_list_df["extraction_date"] = extraction_date
tek_list_df["extraction_date_with_hour"] = extraction_date_with_hour
tek_list_path_prefix = "Data/TEKs/"
tek_list_current_path = tek_list_path_prefix + f"/Current/RadarCOVID-TEKs.json"
tek_list_daily_path = tek_list_path_prefix + f"Daily/RadarCOVID-TEKs-{extraction_date}.json"
tek_list_hourly_path = tek_list_path_prefix + f"Hourly/RadarCOVID-TEKs-{extraction_date_with_hour}.json"
for path in [tek_list_current_path, tek_list_daily_path, tek_list_hourly_path]:
os.makedirs(os.path.dirname(path), exist_ok=True)
tek_list_df.drop(columns=["extraction_date", "extraction_date_with_hour"]).to_json(
tek_list_current_path,
lines=True, orient="records")
tek_list_df.drop(columns=["extraction_date_with_hour"]).to_json(
tek_list_daily_path,
lines=True, orient="records")
tek_list_df.to_json(
tek_list_hourly_path,
lines=True, orient="records")
tek_list_df.head()
```
### Load TEK Dumps
```
import glob
def load_extracted_teks(mode, region=None, limit=None) -> pd.DataFrame:
extracted_teks_df = pd.DataFrame(columns=["region"])
paths = list(reversed(sorted(glob.glob(tek_list_path_prefix + mode + "/RadarCOVID-TEKs-*.json"))))
if limit:
paths = paths[:limit]
for path in paths:
logging.info(f"Loading TEKs from '{path}'...")
iteration_extracted_teks_df = pd.read_json(path, lines=True)
extracted_teks_df = extracted_teks_df.append(
iteration_extracted_teks_df, sort=False)
extracted_teks_df["region"] = \
extracted_teks_df.region.fillna(spain_region_country_code).copy()
if region:
extracted_teks_df = \
extracted_teks_df[extracted_teks_df.region == region]
return extracted_teks_df
daily_extracted_teks_df = load_extracted_teks(
mode="Daily",
region=active_region_country_code,
limit=tek_dumps_load_limit)
daily_extracted_teks_df.head()
exposure_keys_summary_df_ = daily_extracted_teks_df \
.sort_values("extraction_date", ascending=False) \
.groupby("sample_date").tek_list.first() \
.to_frame()
exposure_keys_summary_df_.index.name = "sample_date_string"
exposure_keys_summary_df_["tek_list"] = \
exposure_keys_summary_df_.tek_list.apply(len)
exposure_keys_summary_df_ = exposure_keys_summary_df_ \
.rename(columns={"tek_list": "shared_teks_by_generation_date"}) \
.sort_index(ascending=False)
exposure_keys_summary_df = exposure_keys_summary_df_
exposure_keys_summary_df.head()
```
### Daily New TEKs
```
tek_list_df = daily_extracted_teks_df.groupby("extraction_date").tek_list.apply(
lambda x: set(sum(x, []))).reset_index()
tek_list_df = tek_list_df.set_index("extraction_date").sort_index(ascending=True)
tek_list_df.head()
def compute_teks_by_generation_and_upload_date(date):
day_new_teks_set_df = tek_list_df.copy().diff()
try:
day_new_teks_set = day_new_teks_set_df[
day_new_teks_set_df.index == date].tek_list.item()
except ValueError:
day_new_teks_set = None
if pd.isna(day_new_teks_set):
day_new_teks_set = set()
day_new_teks_df = daily_extracted_teks_df[
daily_extracted_teks_df.extraction_date == date].copy()
day_new_teks_df["shared_teks"] = \
day_new_teks_df.tek_list.apply(lambda x: set(x).intersection(day_new_teks_set))
day_new_teks_df["shared_teks"] = \
day_new_teks_df.shared_teks.apply(len)
day_new_teks_df["upload_date"] = date
day_new_teks_df.rename(columns={"sample_date": "generation_date"}, inplace=True)
day_new_teks_df = day_new_teks_df[
["upload_date", "generation_date", "shared_teks"]]
day_new_teks_df["generation_to_upload_days"] = \
(pd.to_datetime(day_new_teks_df.upload_date) -
pd.to_datetime(day_new_teks_df.generation_date)).dt.days
day_new_teks_df = day_new_teks_df[day_new_teks_df.shared_teks > 0]
return day_new_teks_df
shared_teks_generation_to_upload_df = pd.DataFrame()
for upload_date in daily_extracted_teks_df.extraction_date.unique():
shared_teks_generation_to_upload_df = \
shared_teks_generation_to_upload_df.append(
compute_teks_by_generation_and_upload_date(date=upload_date))
shared_teks_generation_to_upload_df \
.sort_values(["upload_date", "generation_date"], ascending=False, inplace=True)
shared_teks_generation_to_upload_df.tail()
today_new_teks_df = \
shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.upload_date == extraction_date].copy()
today_new_teks_df.tail()
if not today_new_teks_df.empty:
today_new_teks_df.set_index("generation_to_upload_days") \
.sort_index().shared_teks.plot.bar()
generation_to_upload_period_pivot_df = \
shared_teks_generation_to_upload_df[
["upload_date", "generation_to_upload_days", "shared_teks"]] \
.pivot(index="upload_date", columns="generation_to_upload_days") \
.sort_index(ascending=False).fillna(0).astype(int) \
.droplevel(level=0, axis=1)
generation_to_upload_period_pivot_df.head()
new_tek_df = tek_list_df.diff().tek_list.apply(
lambda x: len(x) if not pd.isna(x) else None).to_frame().reset_index()
new_tek_df.rename(columns={
"tek_list": "shared_teks_by_upload_date",
"extraction_date": "sample_date_string",}, inplace=True)
new_tek_df.tail()
estimated_shared_diagnoses_df = daily_extracted_teks_df.copy()
estimated_shared_diagnoses_df["new_sample_extraction_date"] = \
pd.to_datetime(estimated_shared_diagnoses_df.sample_date) + datetime.timedelta(1)
estimated_shared_diagnoses_df["extraction_date"] = pd.to_datetime(estimated_shared_diagnoses_df.extraction_date)
estimated_shared_diagnoses_df["sample_date"] = pd.to_datetime(estimated_shared_diagnoses_df.sample_date)
estimated_shared_diagnoses_df.head()
# Sometimes TEKs from the same day are uploaded, we do not count them as new TEK devices:
same_day_tek_list_df = estimated_shared_diagnoses_df[
estimated_shared_diagnoses_df.sample_date == estimated_shared_diagnoses_df.extraction_date].copy()
same_day_tek_list_df = same_day_tek_list_df[["extraction_date", "tek_list"]].rename(
columns={"tek_list": "same_day_tek_list"})
same_day_tek_list_df.head()
shared_teks_uploaded_on_generation_date_df = same_day_tek_list_df.rename(
columns={
"extraction_date": "sample_date_string",
"same_day_tek_list": "shared_teks_uploaded_on_generation_date",
})
shared_teks_uploaded_on_generation_date_df.shared_teks_uploaded_on_generation_date = \
shared_teks_uploaded_on_generation_date_df.shared_teks_uploaded_on_generation_date.apply(len)
shared_teks_uploaded_on_generation_date_df.head()
shared_teks_uploaded_on_generation_date_df["sample_date_string"] = \
shared_teks_uploaded_on_generation_date_df.sample_date_string.dt.strftime("%Y-%m-%d")
shared_teks_uploaded_on_generation_date_df.head()
estimated_shared_diagnoses_df = estimated_shared_diagnoses_df[
estimated_shared_diagnoses_df.new_sample_extraction_date == estimated_shared_diagnoses_df.extraction_date]
estimated_shared_diagnoses_df.head()
same_day_tek_list_df["extraction_date"] = \
same_day_tek_list_df.extraction_date + datetime.timedelta(1)
estimated_shared_diagnoses_df = \
estimated_shared_diagnoses_df.merge(same_day_tek_list_df, how="left", on=["extraction_date"])
estimated_shared_diagnoses_df["same_day_tek_list"] = \
estimated_shared_diagnoses_df.same_day_tek_list.apply(lambda x: [] if x is np.nan else x)
estimated_shared_diagnoses_df.head()
estimated_shared_diagnoses_df.set_index("extraction_date", inplace=True)
estimated_shared_diagnoses_df["shared_diagnoses"] = estimated_shared_diagnoses_df.apply(
lambda x: len(set(x.tek_list).difference(x.same_day_tek_list)), axis=1).copy()
estimated_shared_diagnoses_df.reset_index(inplace=True)
estimated_shared_diagnoses_df.rename(columns={
"extraction_date": "sample_date_string"}, inplace=True)
estimated_shared_diagnoses_df = estimated_shared_diagnoses_df[["sample_date_string", "shared_diagnoses"]]
estimated_shared_diagnoses_df["sample_date_string"] = estimated_shared_diagnoses_df.sample_date_string.dt.strftime("%Y-%m-%d")
estimated_shared_diagnoses_df.head()
```
### Hourly New TEKs
```
hourly_extracted_teks_df = load_extracted_teks(
mode="Hourly", region=active_region_country_code, limit=25)
hourly_extracted_teks_df.head()
hourly_new_tek_count_df = hourly_extracted_teks_df \
.groupby("extraction_date_with_hour").tek_list. \
apply(lambda x: set(sum(x, []))).reset_index().copy()
hourly_new_tek_count_df = hourly_new_tek_count_df.set_index("extraction_date_with_hour") \
.sort_index(ascending=True)
hourly_new_tek_count_df["new_tek_list"] = hourly_new_tek_count_df.tek_list.diff()
hourly_new_tek_count_df["new_tek_count"] = hourly_new_tek_count_df.new_tek_list.apply(
lambda x: len(x) if not pd.isna(x) else 0)
hourly_new_tek_count_df.rename(columns={
"new_tek_count": "shared_teks_by_upload_date"}, inplace=True)
hourly_new_tek_count_df = hourly_new_tek_count_df.reset_index()[[
"extraction_date_with_hour", "shared_teks_by_upload_date"]]
hourly_new_tek_count_df.head()
hourly_estimated_shared_diagnoses_df = hourly_extracted_teks_df.copy()
hourly_estimated_shared_diagnoses_df["new_sample_extraction_date"] = \
pd.to_datetime(hourly_estimated_shared_diagnoses_df.sample_date) + datetime.timedelta(1)
hourly_estimated_shared_diagnoses_df["extraction_date"] = \
pd.to_datetime(hourly_estimated_shared_diagnoses_df.extraction_date)
hourly_estimated_shared_diagnoses_df = hourly_estimated_shared_diagnoses_df[
hourly_estimated_shared_diagnoses_df.new_sample_extraction_date ==
hourly_estimated_shared_diagnoses_df.extraction_date]
hourly_estimated_shared_diagnoses_df = \
hourly_estimated_shared_diagnoses_df.merge(same_day_tek_list_df, how="left", on=["extraction_date"])
hourly_estimated_shared_diagnoses_df["same_day_tek_list"] = \
hourly_estimated_shared_diagnoses_df.same_day_tek_list.apply(lambda x: [] if x is np.nan else x)
hourly_estimated_shared_diagnoses_df["shared_diagnoses"] = hourly_estimated_shared_diagnoses_df.apply(
lambda x: len(set(x.tek_list).difference(x.same_day_tek_list)), axis=1)
hourly_estimated_shared_diagnoses_df = \
hourly_estimated_shared_diagnoses_df.sort_values("extraction_date_with_hour").copy()
hourly_estimated_shared_diagnoses_df["shared_diagnoses"] = hourly_estimated_shared_diagnoses_df \
.groupby("extraction_date").shared_diagnoses.diff() \
.fillna(0).astype(int)
hourly_estimated_shared_diagnoses_df.set_index("extraction_date_with_hour", inplace=True)
hourly_estimated_shared_diagnoses_df.reset_index(inplace=True)
hourly_estimated_shared_diagnoses_df = hourly_estimated_shared_diagnoses_df[[
"extraction_date_with_hour", "shared_diagnoses"]]
hourly_estimated_shared_diagnoses_df.head()
hourly_summary_df = hourly_new_tek_count_df.merge(
hourly_estimated_shared_diagnoses_df, on=["extraction_date_with_hour"], how="outer")
hourly_summary_df.set_index("extraction_date_with_hour", inplace=True)
hourly_summary_df = hourly_summary_df.fillna(0).astype(int).reset_index()
hourly_summary_df["datetime_utc"] = pd.to_datetime(
hourly_summary_df.extraction_date_with_hour, format="%Y-%m-%d@%H")
hourly_summary_df.set_index("datetime_utc", inplace=True)
hourly_summary_df = hourly_summary_df.tail(-1)
hourly_summary_df.head()
```
### Data Merge
```
result_summary_df = exposure_keys_summary_df.merge(
new_tek_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
shared_teks_uploaded_on_generation_date_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
estimated_shared_diagnoses_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = confirmed_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df["sample_date"] = pd.to_datetime(result_summary_df.sample_date_string)
result_summary_df.set_index("sample_date", inplace=True)
result_summary_df.drop(columns=["sample_date_string"], inplace=True)
result_summary_df.sort_index(ascending=False, inplace=True)
result_summary_df.head()
with pd.option_context("mode.use_inf_as_na", True):
result_summary_df = result_summary_df.fillna(0).astype(int)
result_summary_df["teks_per_shared_diagnosis"] = \
(result_summary_df.shared_teks_by_upload_date / result_summary_df.shared_diagnoses).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case"] = \
(result_summary_df.shared_diagnoses / result_summary_df.covid_cases).fillna(0)
result_summary_df.head(daily_plot_days)
weekly_result_summary_df = result_summary_df \
.sort_index(ascending=True).fillna(0).rolling(7).agg({
"covid_cases": "sum",
"shared_teks_by_generation_date": "sum",
"shared_teks_by_upload_date": "sum",
"shared_diagnoses": "sum"
}).sort_index(ascending=False)
with pd.option_context("mode.use_inf_as_na", True):
weekly_result_summary_df = weekly_result_summary_df.fillna(0).astype(int)
weekly_result_summary_df["teks_per_shared_diagnosis"] = \
(weekly_result_summary_df.shared_teks_by_upload_date / weekly_result_summary_df.shared_diagnoses).fillna(0)
weekly_result_summary_df["shared_diagnoses_per_covid_case"] = \
(weekly_result_summary_df.shared_diagnoses / weekly_result_summary_df.covid_cases).fillna(0)
weekly_result_summary_df.head()
last_7_days_summary = weekly_result_summary_df.to_dict(orient="records")[0]
last_7_days_summary
```
## Report Results
```
display_column_name_mapping = {
"sample_date": "Sample\u00A0Date\u00A0(UTC)",
"datetime_utc": "Timestamp (UTC)",
"upload_date": "Upload Date (UTC)",
"generation_to_upload_days": "Generation to Upload Period in Days",
"region": "Backend Region",
"covid_cases": "COVID-19 Cases (7-day Rolling Average)",
"shared_teks_by_generation_date": "Shared TEKs by Generation Date",
"shared_teks_by_upload_date": "Shared TEKs by Upload Date",
"shared_diagnoses": "Shared Diagnoses (Estimation)",
"teks_per_shared_diagnosis": "TEKs Uploaded per Shared Diagnosis",
"shared_diagnoses_per_covid_case": "Usage Ratio (Fraction of Cases Which Shared Diagnosis)",
"shared_teks_uploaded_on_generation_date": "Shared TEKs Uploaded on Generation Date",
}
summary_columns = [
"covid_cases",
"shared_teks_by_generation_date",
"shared_teks_by_upload_date",
"shared_teks_uploaded_on_generation_date",
"shared_diagnoses",
"teks_per_shared_diagnosis",
"shared_diagnoses_per_covid_case",
]
```
### Daily Summary Table
```
result_summary_df_ = result_summary_df.copy()
result_summary_df = result_summary_df[summary_columns]
result_summary_with_display_names_df = result_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
result_summary_with_display_names_df
```
### Daily Summary Plots
```
result_plot_summary_df = result_summary_df.head(daily_plot_days)[summary_columns] \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
summary_ax_list = result_plot_summary_df.sort_index(ascending=True).plot.bar(
title=f"Daily Summary",
rot=45, subplots=True, figsize=(15, 22), legend=False)
ax_ = summary_ax_list[-1]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.95)
ax_.yaxis.set_major_formatter(matplotlib.ticker.PercentFormatter(1.0))
_ = ax_.set_xticklabels(sorted(result_plot_summary_df.index.strftime("%Y-%m-%d").tolist()))
```
### Daily Generation to Upload Period Table
```
display_generation_to_upload_period_pivot_df = \
generation_to_upload_period_pivot_df \
.head(backend_generation_days)
display_generation_to_upload_period_pivot_df \
.head(backend_generation_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping)
import matplotlib.pyplot as plt
fig, generation_to_upload_period_pivot_table_ax = plt.subplots(
figsize=(10, 1 + 0.5 * len(display_generation_to_upload_period_pivot_df)))
generation_to_upload_period_pivot_table_ax.set_title(
"Shared TEKs Generation to Upload Period Table")
sns.heatmap(
data=display_generation_to_upload_period_pivot_df
.rename_axis(columns=display_column_name_mapping)
.rename_axis(index=display_column_name_mapping),
fmt=".0f",
annot=True,
ax=generation_to_upload_period_pivot_table_ax)
generation_to_upload_period_pivot_table_ax.get_figure().tight_layout()
```
### Hourly Summary Plots
```
hourly_summary_ax_list = hourly_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.plot.bar(
title=f"Last 24h Summary",
rot=45, subplots=True, legend=False)
ax_ = hourly_summary_ax_list[-1]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.9)
_ = ax_.set_xticklabels(sorted(hourly_summary_df.index.strftime("%Y-%m-%d@%H").tolist()))
```
### Publish Results
```
def get_temporary_image_path() -> str:
return os.path.join(tempfile.gettempdir(), str(uuid.uuid4()) + ".png")
def save_temporary_plot_image(ax):
if isinstance(ax, np.ndarray):
ax = ax[0]
media_path = get_temporary_image_path()
ax.get_figure().savefig(media_path)
return media_path
def save_temporary_dataframe_image(df):
import dataframe_image as dfi
media_path = get_temporary_image_path()
dfi.export(df, media_path)
return media_path
github_repository = os.environ.get("GITHUB_REPOSITORY")
if github_repository is None:
github_repository = "pvieito/Radar-STATS"
github_project_base_url = "https://github.com/" + github_repository
display_formatters = {
display_column_name_mapping["teks_per_shared_diagnosis"]: lambda x: f"{x:.2f}",
display_column_name_mapping["shared_diagnoses_per_covid_case"]: lambda x: f"{x:.2%}",
}
daily_summary_table_html = result_summary_with_display_names_df \
.head(daily_plot_days) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.to_html(formatters=display_formatters)
multi_region_summary_table_html = multi_region_summary_df \
.head(daily_plot_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(formatters=display_formatters)
extraction_date_result_summary_df = \
result_summary_df[result_summary_df.index == extraction_date]
extraction_date_result_hourly_summary_df = \
hourly_summary_df[hourly_summary_df.extraction_date_with_hour == extraction_date_with_hour]
covid_cases = \
extraction_date_result_summary_df.covid_cases.sum()
shared_teks_by_generation_date = \
extraction_date_result_summary_df.shared_teks_by_generation_date.sum()
shared_teks_by_upload_date = \
extraction_date_result_summary_df.shared_teks_by_upload_date.sum()
shared_diagnoses = \
extraction_date_result_summary_df.shared_diagnoses.sum()
teks_per_shared_diagnosis = \
extraction_date_result_summary_df.teks_per_shared_diagnosis.sum()
shared_diagnoses_per_covid_case = \
extraction_date_result_summary_df.shared_diagnoses_per_covid_case.sum()
shared_teks_by_upload_date_last_hour = \
extraction_date_result_hourly_summary_df.shared_teks_by_upload_date.sum().astype(int)
shared_diagnoses_last_hour = \
extraction_date_result_hourly_summary_df.shared_diagnoses.sum().astype(int)
summary_plots_image_path = save_temporary_plot_image(
ax=summary_ax_list)
summary_table_image_path = save_temporary_dataframe_image(
df=result_summary_with_display_names_df)
hourly_summary_plots_image_path = save_temporary_plot_image(
ax=hourly_summary_ax_list)
multi_region_summary_table_image_path = save_temporary_dataframe_image(
df=multi_region_summary_df)
generation_to_upload_period_pivot_table_image_path = save_temporary_plot_image(
ax=generation_to_upload_period_pivot_table_ax)
```
### Save Results
```
report_resources_path_prefix = "Data/Resources/Current/RadarCOVID-Report-"
result_summary_df.to_csv(
report_resources_path_prefix + "Summary-Table.csv")
result_summary_df.to_html(
report_resources_path_prefix + "Summary-Table.html")
hourly_summary_df.to_csv(
report_resources_path_prefix + "Hourly-Summary-Table.csv")
multi_region_summary_df.to_csv(
report_resources_path_prefix + "Multi-Region-Summary-Table.csv")
generation_to_upload_period_pivot_df.to_csv(
report_resources_path_prefix + "Generation-Upload-Period-Table.csv")
_ = shutil.copyfile(
summary_plots_image_path,
report_resources_path_prefix + "Summary-Plots.png")
_ = shutil.copyfile(
summary_table_image_path,
report_resources_path_prefix + "Summary-Table.png")
_ = shutil.copyfile(
hourly_summary_plots_image_path,
report_resources_path_prefix + "Hourly-Summary-Plots.png")
_ = shutil.copyfile(
multi_region_summary_table_image_path,
report_resources_path_prefix + "Multi-Region-Summary-Table.png")
_ = shutil.copyfile(
generation_to_upload_period_pivot_table_image_path,
report_resources_path_prefix + "Generation-Upload-Period-Table.png")
```
### Publish Results as JSON
```
summary_results_api_df = result_summary_df.reset_index()
summary_results_api_df["sample_date_string"] = \
summary_results_api_df["sample_date"].dt.strftime("%Y-%m-%d")
summary_results = dict(
extraction_datetime=extraction_datetime,
extraction_date=extraction_date,
extraction_date_with_hour=extraction_date_with_hour,
last_hour=dict(
shared_teks_by_upload_date=shared_teks_by_upload_date_last_hour,
shared_diagnoses=shared_diagnoses_last_hour,
),
today=dict(
covid_cases=covid_cases,
shared_teks_by_generation_date=shared_teks_by_generation_date,
shared_teks_by_upload_date=shared_teks_by_upload_date,
shared_diagnoses=shared_diagnoses,
teks_per_shared_diagnosis=teks_per_shared_diagnosis,
shared_diagnoses_per_covid_case=shared_diagnoses_per_covid_case,
),
last_7_days=last_7_days_summary,
daily_results=summary_results_api_df.to_dict(orient="records"))
summary_results = \
json.loads(pd.Series([summary_results]).to_json(orient="records"))[0]
with open(report_resources_path_prefix + "Summary-Results.json", "w") as f:
json.dump(summary_results, f, indent=4)
```
### Publish on README
```
with open("Data/Templates/README.md", "r") as f:
readme_contents = f.read()
readme_contents = readme_contents.format(
extraction_date_with_hour=extraction_date_with_hour,
github_project_base_url=github_project_base_url,
daily_summary_table_html=daily_summary_table_html,
multi_region_summary_table_html=multi_region_summary_table_html)
with open("README.md", "w") as f:
f.write(readme_contents)
```
### Publish on Twitter
```
enable_share_to_twitter = os.environ.get("RADARCOVID_REPORT__ENABLE_PUBLISH_ON_TWITTER")
github_event_name = os.environ.get("GITHUB_EVENT_NAME")
if enable_share_to_twitter and github_event_name == "schedule":
import tweepy
twitter_api_auth_keys = os.environ["RADARCOVID_REPORT__TWITTER_API_AUTH_KEYS"]
twitter_api_auth_keys = twitter_api_auth_keys.split(":")
auth = tweepy.OAuthHandler(twitter_api_auth_keys[0], twitter_api_auth_keys[1])
auth.set_access_token(twitter_api_auth_keys[2], twitter_api_auth_keys[3])
api = tweepy.API(auth)
summary_plots_media = api.media_upload(summary_plots_image_path)
summary_table_media = api.media_upload(summary_table_image_path)
generation_to_upload_period_pivot_table_image_media = api.media_upload(generation_to_upload_period_pivot_table_image_path)
media_ids = [
summary_plots_media.media_id,
summary_table_media.media_id,
generation_to_upload_period_pivot_table_image_media.media_id,
]
status = textwrap.dedent(f"""
#RadarCOVID Report – {extraction_date_with_hour}
Today:
- Uploaded TEKs: {shared_teks_by_upload_date:.0f} ({shared_teks_by_upload_date_last_hour:+d} last hour)
- Shared Diagnoses: ≤{shared_diagnoses:.0f} ({shared_diagnoses_last_hour:+d} last hour)
- TEKs per Diagnosis: ≥{teks_per_shared_diagnosis:.1f}
- Usage Ratio: ≤{shared_diagnoses_per_covid_case:.2%}
Week:
- Shared Diagnoses: ≤{last_7_days_summary["shared_diagnoses"]:.0f}
- Usage Ratio: ≤{last_7_days_summary["shared_diagnoses_per_covid_case"]:.2%}
More Info: {github_project_base_url}#documentation
""")
status = status.encode(encoding="utf-8")
api.update_status(status=status, media_ids=media_ids)
```
| github_jupyter |
# Aula 3: Python - um pouco mais - avançado
**O que veremos hoje?**
- como declarar funções e importar pacotes em python
- pacote para manipulação numérica de informações
- pacote para visualização de dados (1D)
**material de apoio**
- numpy guia de usuário: https://numpy.org/doc/stable/numpy-user.pdf
- numpy: https://ncar-hackathons.github.io/scientific-computing/numpy/intro.html
- numpy + scipy: https://github.com/gustavo-marques/hands-on-examples/blob/master/scientific-computing/numpy_scipy.ipynb
- matplotlib documentação: https://matplotlib.org/3.3.2/contents.html
-
# Python - um pouco mais - avançado
Quando definimos um algoritmo para solucionar um problema, é comum na programação estabelecermos pequenos problemas que possam ser resolvidos. Tal qual já falamos, seria uma espécie de passo a passo para se chegar na solução de um problema.
Em python podemos programar estes pequenos problemas em pequeno (não tanto às vezes) códigos, aplicando os conceitos de funções e métodos.
Funções seguem o estilo de programação procedural, ou seja, diversas funções sendo requisitadas quando necessário. Enquanto que métodos estão associados à programação orientada a objetos, uma ténica de programação abstrata. Neste curso trataremos somente sobre funções, por questões de tempo.
# Funções (functions)
**Definição**
É nada mais do que um bloco de código independente, que realiza uma ou mais ações específicas, retornando ou não um valor/variável quando requisitada. As funções possuem um nome, utilizado para chamá-las, e podemos transmitir informações para elas na forma de ```argumentos```.
**Sintaxe**

Vamos destrinchar brevemente uma função.
Utilizamos o termo ```def``` para indicar ao python que vamos começar a definir uma nova função e, logo em seguida, dizemos o nome desta função. Dentro dos parênteses colocamos os argumentos, que podem ter qualquer nome. Para entender argumentos, vamos pensar que são variáveis que passarem para a função trabalhar. Isto é necessário uma vez que, por se tratarem de blocos de códigos independentes, as funções não possuem acesso às variáveis gerais de um programa. Por fim, ao final da definição do nome da função e seus argumentos, inserimos ```:``` para indicarmos um novo bloco de códigos.
Logo abaixo, entre aspas triplas, colocamos uma ```docstring``` que nada mais é do que uma documentação da função, indicando sua utilidade, explicando os argumentos que ela precisa receber para funcionar adequadamente e outras informações úteis. Isto é muito importante, pois nos ajudará no futuro quando tivermos diversas funções e não lembrarmos para que todas servem. O docstring vai nos salvar um tempo precioso.
Por fim, funções podem ou não retornar valores quando requisitadas. Pode ser o resultado de um cálculo, uma mensagem dizendo que o código rodou bem ou não ou simplesmente nada (por exemplo, uma função para plotar e salvar uma figura).
Convencionalmente, definimos as funções logo após importar pacotes, ou seja, no início do código. Mas veremos exemplos mais avançados e complexos ao longo das aulas.
```
def K_para_C(T_em_K):
# sequencia de bloco de códigos identados
print('Estamos dentro de uma função!')
# conversão
T_em_C = T_em_K - 273.15
return T_em_C
temperatura = K_para_C(273.15)
print(temperatura)
```
Podemos deixar uma função mais completa adicionando informações sobre como ela funciona. Para isso, usamos as ```docstrings```:
```
def K_para_C(T_em_K):
"""
Este conjunto de texto entre aspas triplas se chamada docstring. É usada para passar informações
importantes sobre a função, como: valores que ela recebe e retorna, o que ela faz, se é baseada em algum
artigo científico, por exemplo, e muito mais. Quando tivermos dúvida sobre alguma função, podemos buscar
o docstring dela utilizando o comando: K_para_C?
No nosso caso, essa é uma função para converter uma temperatura de Kelvin para Celsius
parameters
----------
T_em_K: float
Temperatura em Kelvin para ser convertida.
returns
-------
T_em_C: float
Temperatura em graus Celsius.
"""
# sequencia de bloco de códigos identados
print('Estamos dentro de uma função!')
# conversão
T_em_C = T_em_K - 273.15
return T_em_C
```
# Pacotes (packages ou libraries)
São conjuntos de funções específicas para resolver um problema, ou realizar análise de dados. Na verdade, utilizamos diversos pacotes ao programar em python, como veremos mais a frente.
Podemos criar nosso próprio pacote, gerando instaladores e etc, mas o mais usual é instalarmos estes pacotes através de sistemas de gerenciamento de pacotes como o ```pip``` ou o próprio ```conda/anaconda```.
---------------
# Pacote Numérico: NumPy
- background de qualquer pacote científico hoje em dia
- utilizado para operações numéricas (matriciais ou escalares)
- alta performance
**Instalação**
```bash
pip install numpy
```
ou
```bash
conda install numpy
```
**Importar** pacote:
Para utilizar o pacote NumPy, precisamos importa-lo em nosso código. Fazemos isso através do comando ```import``` e, no caso abaixo, adicionamos um apelido para o numpy. Isso nos facilitará para utilizar as funções deste pacote.
```python
import numpy as np
```
Se pudéssemos traduzir o comando acima em português, o faríamos da seguinte forma: ```importe numpy como np```.
**Conceitos**
Lembrando alguns conceitos matemáticos:
- Vetores (N,):
$V = \begin{bmatrix}
1 & 2 & 3
\end{bmatrix}$
- Matrizes (NxM, linhasxcolunas):
$M = \begin{bmatrix}
1 & 2 & 3\\
4 & 5 & 6
\end{bmatrix}$
Vamos ver alguns exemplos e entender na prática a funcionalidade do NumPy.
```
# primeiro importamos e só o precisamos fazer uma vez em cada código
import numpy as np
V = np.array([1, 2, 3])
M = np.array([[1, 2, 3], [4, 5, 6]])
M
```
Apesar de falarmos em vetores e matrizes, para o numpy é tudo a mesma coisa. O que diferencia é apenas a dimensão.
Nota: ```ndarray``` significa n-dimensional array (matriz com n-dimensões)
```
# tipo
type(M), type(V)
```
Como descobrir a quantidade de dimensões?
```
V.ndim, M.ndim
```
Podemos verificar o formato da matriz que estamos trabalhando, utilizando o método .shape, ou o tamanho geral da matriz, com o método .size:
```
V.shape, V.size
M.shape, M.size
```
**Utilidade**
Os conceitos citados acima parecem muito com as listas que vimos anteriormente. Porém, neste caso, utilizar numpy nos permitirá trabalhar com operações matriciais, além de ter uma performance, em termos de memória, bem melhor.
Além disso, uma vez que criamos uma matriz com o numpy, não poderemos atribuir nenhum valor que não seja do mesmo tipo do que a matriz foi criada:
```
V[0] = 'teste'
# mas porque este funciona?
A = np.array(['teste', 0])
A[0] = 0
```
Podemos também indicar o tipo de matriz que queremos, ao cria-la, utilizando o argumento ```dtype``` e inserindo alguma das opções que já nos é conhecida (int, float, bool e complex):
```
c = np.array([1, 2, 3], dtype=complex)
c
```
**Principais funções disponíveis no Numpy**
```
# criar um vetor ordenado (crescente) de 1 a 1
x = np.arange(0, 100, 1)
x
# podemos criar o mesmo vetor, mas na ordem descrescente
x_inverso = np.arange(100, 0, -1)
x_inverso
# criar um vetor de 0 a 100, com de 10 intervalos
y = np.linspace(0, 100, 10)
y
# criar uma grade
x,y = np.mgrid[0:5, 0:5]
x
# utilizando números aleatórios
np.random.rand(3,3)
```
Outros métodos:
cumsum, dot, det, sort, max, min, argmax, argmin, sqrt, e outros.
Testem o que estes métodos fazem.
Lembre que parar usar você deve:
```python
x = np.arange(0,10,1)
soma = x.sum()
```
---------------
Mesclando numpy e visualização de dados
# Pacote de visualização: Matplotlib
- base para qualquer visualização de dados
- é um saco importar, porém com o tempo fica menor pior
- muito poderoso em termos de controle dos elementos do gráfico e da estrutura em si, isto é:
- estruturas complexas de eixos de visualização podem ser manipuladas por este pacote
**Importar**
```python
import matplotlib.pyplot as plt
```
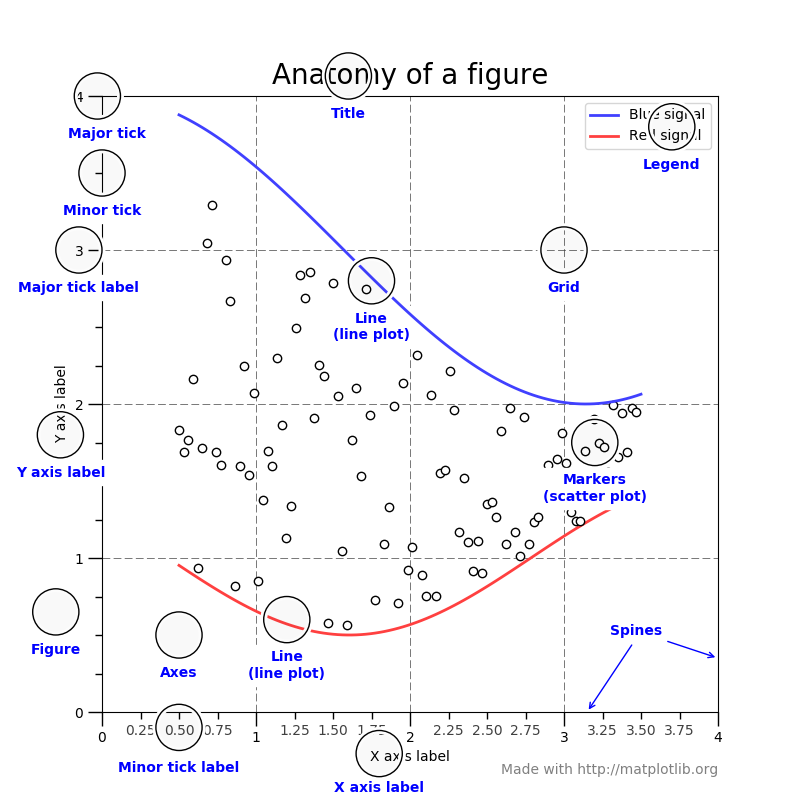
Fonte: Prof Storopoli [https://github.com/storopoli/ciencia-de-dados]
```
import matplotlib.pyplot as plt
# quando usando notebook, use o comando abaixo para exibir as figuras no proprio arquivo:
%matplotlib inline
```
Podemos plotar um gráfico único em uma imagem, utilizando:
```
# iniciamos (instanciar) uma nova figura
fig = plt.figure()
# note que, como nada foi plotado efetivamente, nada aparece aqui embaixo.
```
Refazendo o comando e inserindo um plot 1D simples:
```
fig = plt.figure()
plt.plot([1,2,3])
```
Ainda temos a opção de plotar diversos gráficos em uma mesma figura. Para isso, usamos um outro método do matplotlib:
```
fig,axes = plt.subplots(nrows=1, ncols=2)
```
Diferente do primeiro método, este segundo já nos gera dois eixos para plotarmos as informações que quiseremos. Note que a variável ```axes``` neste caso e uma matriz 1D com shape ```(1,2)```, contendo estes eixos.
Assim para de fato inserirmos alguma informação na figura com subplots, fazemos:
```
fig,axes = plt.subplots(1,2)
# exibindo o tipo de axes
print(type(axes))
# plotando no gráfico da esquerda (1o gráfico)
ax = axes[0]
ax.plot([1,2,3])
# plotando no gráfico da direita (2o gráfico)
axes[1].plot([3,2,1])
```
Ainda podemos deixar a figura mais organizada. Note que ambos os subplots compartilham da mesmo eixo y (ordenadas), com mesmo limite de valores. Podemos dizer para elas compartilharem este mesmo eixo com o argumento ```sharey```. Isto também é válido para as abscissas (eixo x), porém com ```sharex```.
Vejam:
```
fig,axes = plt.subplots(nrows=1, ncols=2, sharey=True)
# exibindo o tipo de axes
print(type(axes))
# plotando no gráfico da esquerda (1o gráfico)
ax = axes[0]
ax.plot([1,2,3])
# plotando no gráfico da direita (2o gráfico)
axes[1].plot([3,2,1])
```
E ainda podemos especificar o tamanho da nossa figura com o argumento ```figsize```. Este argumento também funciona em ```plt.figure()```.
```
fig,axes = plt.subplots(nrows=1, ncols=2, sharey=True, figsize=(15,5))
# plotando no gráfico da esquerda (1o gráfico)
ax = axes[0]
ax.plot([1,2,3])
# plotando no gráfico da direita (2o gráfico)
axes[1].plot([3,2,1])
```
### Customizando os gráficos
Podemos ainda trocar as cores, os marcadores dentro de uma figura,
usando comando simples, como:
```
fig,axes = plt.subplots(nrows=1, ncols=2, sharey=True, figsize=(15,5))
# plotando no gráfico da esquerda (1o gráfico)
ax = axes[0]
ax.plot([1,2,3], 'r-')
# plotando no gráfico da direita (2o gráfico)
axes[1].plot([3,2,1], 'g-o')
```
Aproveitando que falamos de função no início, vamos montar uma função que reproduza as figuras feitas acima:
```
def plot_simples():
fig,axes = plt.subplots(nrows=1, ncols=2, sharey=True, figsize=(15,5))
# plotando no gráfico da esquerda (1o gráfico)
ax = axes[0]
ax.plot([1,2,3], 'r-')
# plotando no gráfico da direita (2o gráfico)
axes[1].plot([3,2,1], 'g-o')
# retornamos fig e axes pq precisaremos no futuro
return fig,axes
```
Agora, usando nossa função, vamos fazer mais customizações na nossa figura:
```
import numpy as np
# usando a função. Note que atribuimos o retorno da função a duas variáveis
fig,axes = plot_simples()
# inserindo rótulos ao eixo das abscissas (x) e ordenada (y):
axes[0].set_xlabel('Abscissa')
axes[0].set_ylabel('Ordenada')
axes[1].set_xlabel('Abscissa')
# adicionando um título para cada subplot
axes[0].set_title('Figura da esquerda')
axes[1].set_title('Figura da direita')
# podemos configurar o intervalo discreto das abscissas e ordenadas
axes[0].set_xticks(np.arange(0,3,1))
# podemos ainda trocar os rótulos de cada tick
axes[0].set_xticklabels(['primeiro', 'segundo', 'terceiro'])
# configurar os limites
axes[1].set_xlim([0,10])
# inserindo outros elementos:
axes[0].grid('--', alpha=.3)
axes[1].grid('--', alpha=.3)
```
**Tipos de gráficos para 1D**
- linha: ```plt.plot()```
- barra: ```plt.barh()```
- histograma: ```plt.hist()```
Podemos plotar, de forma semelhante, histogramas ou gráficos de barra ou histograma:
```
fig,axes = plt.subplots(1,2,figsize=(15,5))
# grafico de barras horizontal
axes[0].barh(x,y)
# histograma para x
_,_,_ = axes[1].hist(x,5)
# bonus: personalizacao dos subplots usando apenas uma linha com compreensão de listas (list comprehension)
_ = [ax.grid('--', alpha=.3) for ax in axes]
```
Se tiver dúvidas de como usar uma função, você pode consultar a documentação do matplotlib ou, aqui mesmo, pedir por ajuda com:
```
# precisando de ajuda?
help(plt.plot)
```
Enfim, para **salvar** uma figura, utilizamos o método ```plt.savefig()```:
- formatos disponíveis: pdf, png, jpg, tif, svg
- dpi: qualidade da figura salva
- bbox_to_inches: use ```tight``` para remover espaços em branco ao redor
```
# usando a nossa função
fig,ax = plot_simples()
# salvando a figura
plt.savefig('nome_da_figure.png', dpi=150, bbox_to_inches='tight')
```
Exercício:
Utilizando o dicionário criado com a lista de espécies, plote um gráfico de barras horizontais, utilizando ```plt.barh()```.
**dica**: use ```list()``` para converter as chaves e valores do dicionário para uma lista.
```
# espaço reservado para tentar resolver o exercício
```
É claro que existem diversas formas de visualização de gráficos de uma dimensão no python. Apresentamos dois tipos bem básicos, porém muito utilizados no dia a dia de um cientista.
Para mais exemplo, navegue pela documentação do matplotlib. Ao longo do curso iremos explorar diversos formatos de visualização e explicaremos cada caso conforme uso.
Exercícios de casa: perfis verticais
Arquivos com temperatura e salinidade climatológica do World Ocean Atlas (WOA) serão fornecidos para uma região específica do litoral Norte de São Paulo: Ubatuba.
1. carregue os dados com numpy usando genfromtxt ou loadtxt(sep=','), sendo uma variável para cada propriedade
```python
temperatura = np.loadtxt('../dados/salinidade_woa2018_ubatuba_60m.csv', delimiter=',')
```
2. explore a estrutura da matriz que virá. Por exemplo, identifique:
- o que é cada coluna? E cada linha?
- como acessá-los pelo indexamento de matrizes?
Esteja familiarizado com a matriz fornecida antes de prosseguir para a visualização, assim erros que poderão assustar serão evitados.
```
# codigo para baixar o arquivo, caso você esteja rodando este notebook no Google Colab
!wget --directory-prefix=../dados/ https://raw.githubusercontent.com/nilodna/python-basico/feature_iojr-shortcourse/dados/temperatura_woa2018_ubatuba_60m.csv
!wget --directory-prefix=../dados/ https://raw.githubusercontent.com/nilodna/python-basico/feature_iojr-shortcourse/dados/salinidade_woa2018_ubatuba_60m.csv
# explorando matrizes de temperatura e salinidade
```
Nível 1:
- faça subplots de perfis verticais para o mês de janeiro. Cada subplot será uma propriedade (temperatura e salinidade)
Nível 2:
- Faça uma média dos meses de verão e meses de inverno
- plot as médias em dois subplots, para cada propriedade
Nível 3:
- plote todos os meses da climatologia em cada figura (uma para cada propriedade como feito nos anteriores).
- mantenha consistência em cores ou marcadores para cada mês
Bônus:
- plote em um só gráfico a temperatura e salinidade para um mês a sua escolha.
- sabendo que os limites serão diferentes, adicione um segundo eixo (eixo gêmeo) à figura usando ```ax.twinx()```.
**dicas**
- use ```:``` para acessar todos os valores de uma dimensão da matriz
- estilize seu perfil vertical com gradeamento (grid), rotulos (labels) nos eixos. legenda (legend), etc...
| github_jupyter |
```
import pandas as pd
import numpy as np
train = pd.read_csv( "../data/processed/train_1.csv")
test = pd.read_csv("../data/processed/test_1.csv")
validation = pd.read_csv("../data/processed/validation_1.csv")
from sklearn.model_selection import train_test_split
X = train['review']
y = train['sentiment']
from sklearn.feature_extraction.text import CountVectorizer
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.models import Sequential
from keras.layers import Dense, Embedding, LSTM, SpatialDropout1D
from sklearn.model_selection import train_test_split
from keras.utils.np_utils import to_categorical
max_features = 2000
tokenizer = Tokenizer(num_words=max_features, split=' ')
tokenizer.fit_on_texts(X.values)
X = tokenizer.texts_to_sequences(X.values)
X = pad_sequences(X)
embed_dim = 128
lstm_out = 196
model = Sequential()
model.add(Embedding(max_features, embed_dim,input_length = X.shape[1]))
model.add(SpatialDropout1D(0.4))
model.add(LSTM(lstm_out, dropout=0.2, recurrent_dropout=0.2))
model.add(Dense(2,activation='softmax'))
model.compile(loss = 'categorical_crossentropy', optimizer='adam',metrics = ['accuracy'])
print(model.summary())
Y = pd.get_dummies(train['sentiment']).values
X_train, X_test, Y_train, Y_test = train_test_split(X,Y, test_size = 0.33, random_state = 42)
print(X_train.shape,Y_train.shape)
print(X_test.shape,Y_test.shape)
batch_size = 32
model.fit(X_train, Y_train, epochs = 10, batch_size=batch_size, verbose = 2)
validation_size = 1500
X_validate = X_test[-validation_size:]
Y_validate = Y_test[-validation_size:]
X_test = X_test[:-validation_size]
Y_test = Y_test[:-validation_size]
score,acc = model.evaluate(X_test, Y_test, verbose = 2, batch_size = batch_size)
print("score: %.2f" % (score))
print("acc: %.2f" % (acc))
pos_cnt, neg_cnt, pos_correct, neg_correct = 0, 0, 0, 0
for x in range(len(X_validate)):
result = model.predict(X_validate[x].reshape(1,X_test.shape[1]),batch_size=1,verbose = 2)[0]
if np.argmax(result) == np.argmax(Y_validate[x]):
if np.argmax(Y_validate[x]) == 0:
neg_correct += 1
else:
pos_correct += 1
if np.argmax(Y_validate[x]) == 0:
neg_cnt += 1
else:
pos_cnt += 1
print("pos_acc", pos_correct/pos_cnt*100, "%")
print("neg_acc", neg_correct/neg_cnt*100, "%")
#load embeddings
import codecs
from tqdm import tqdm
print('loading word embeddings...')
embeddings_index = {}
f = codecs.open('../data/external/wiki.id.vec', encoding='utf-8')
for line in tqdm(f):
values = line.rstrip().rsplit(' ')
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
f.close()
print('found %s word vectors' % len(embeddings_index))
from nltk.tokenize import RegexpTokenizer
from keras.preprocessing import sequence
import keras
from keras import optimizers
from keras import backend as K
from keras import regularizers
from keras.models import Sequential
from keras.layers import Dense, Activation, Dropout, Flatten
from keras.layers import Embedding, Conv1D, MaxPooling1D, GlobalMaxPooling1D
from keras.utils import plot_model
from keras.preprocessing import sequence
from keras.preprocessing.text import Tokenizer
from keras.callbacks import EarlyStopping
raw_docs_train = train['review'].tolist()
raw_docs_test = test['review'].tolist()
MAX_NB_WORDS = 100000
tokenizer = RegexpTokenizer(r'\w+')
train['doc_len'] = train['review'].apply(lambda words: len(words.split(" ")))
max_seq_len = np.round(train['doc_len'].mean() + train['doc_len'].std()).astype(int)
print("pre-processing train data...")
processed_docs_train = []
for doc in tqdm(raw_docs_train):
tokens = tokenizer.tokenize(doc)
filtered = [word for word in tokens]
processed_docs_train.append(" ".join(filtered))
#end for
processed_docs_test = []
for doc in tqdm(raw_docs_test):
tokens = tokenizer.tokenize(doc)
filtered = [word for word in tokens]
processed_docs_test.append(" ".join(filtered))
#end for
print("tokenizing input data...")
tokenizer = Tokenizer(num_words=MAX_NB_WORDS, lower=True, char_level=False)
tokenizer.fit_on_texts(processed_docs_train + processed_docs_test) #leaky
word_seq_train = tokenizer.texts_to_sequences(processed_docs_train)
word_seq_test = tokenizer.texts_to_sequences(processed_docs_test)
word_index = tokenizer.word_index
print("dictionary size: ", len(word_index))
#pad sequences
word_seq_train = sequence.pad_sequences(word_seq_train, maxlen=max_seq_len)
word_seq_test = sequence.pad_sequences(word_seq_test, maxlen=max_seq_len)
#training params
batch_size = 256
num_epochs = 8
#model parameters
num_filters = 64
embed_dim = 300
weight_decay = 1e-4
print('preparing embedding matrix...')
words_not_found = []
nb_words = min(MAX_NB_WORDS, len(word_index))
embedding_matrix = np.zeros((nb_words, embed_dim))
for word, i in word_index.items():
if i >= nb_words:
continue
embedding_vector = embeddings_index.get(word)
if (embedding_vector is not None) and len(embedding_vector) > 0:
# words not found in embedding index will be all-zeros.
embedding_matrix[i] = embedding_vector
else:
words_not_found.append(word)
print('number of null word embeddings: %d' % np.sum(np.sum(embedding_matrix, axis=1) == 0))
print("sample words not found: ", np.random.choice(words_not_found, 10))
#CNN architecture
print("training CNN ...")
num_classes = 2
model = Sequential()
model.add(Embedding(nb_words, embed_dim,
weights=[embedding_matrix], input_length=max_seq_len, trainable=False))
model.add(Conv1D(num_filters, 7, activation='relu', padding='same'))
model.add(MaxPooling1D(2))
model.add(Conv1D(num_filters, 7, activation='relu', padding='same'))
model.add(GlobalMaxPooling1D())
model.add(Dropout(0.5))
model.add(Dense(32, activation='relu', kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Dense(num_classes, activation='sigmoid')) #multi-label (k-hot encoding)
adam = optimizers.Adam(lr=0.001, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0)
model.compile(loss='binary_crossentropy', optimizer=adam, metrics=['accuracy'])
model.summary()
#define callbacks
early_stopping = EarlyStopping(monitor='val_loss', min_delta=0.01, patience=4, verbose=1)
callbacks_list = [early_stopping]
```
| github_jupyter |
```
import warnings
warnings.filterwarnings('ignore')
import os
from matplotlib import pyplot as plt
%matplotlib inline
import shutil
import glob
from pathlib import Path
import pandas as pd
import numpy as np
import time
from PIL import Image
import cv2
from numpy import asarray
from keras.utils import to_categorical
import os
import seaborn as sns
from sklearn.metrics import confusion_matrix,classification_report
from sklearn.utils import class_weight
# get the file path and name
df_train = pd.DataFrame([file_path for file_path in Path('videos_faces/train').glob('**/*.jpg')], columns=['file'])
df_train["root"] = df_train["file"].apply(lambda x: os.path.split(os.path.split(x)[0])[1])
df_train['basefile'] = df_train['file'].apply(lambda x: os.path.basename(x))
df_train['sequence'] = df_train['basefile'].apply(lambda x: int(x[x.find('_')+1:-4]))
df_train['basename'] = df_train['basefile'].apply(lambda x: x[:x.find('_')])
df_train.sort_values(["root", "basename", "sequence"], inplace = True)
df_test = pd.DataFrame([file_path for file_path in Path('videos_faces/test').glob('**/*.jpg')], columns=['file'])
df_test["root"] = df_test["file"].apply(lambda x: os.path.split(os.path.split(x)[0])[1])
df_test['basefile'] = df_test['file'].apply(lambda x: os.path.basename(x))
df_test['sequence'] = df_test['basefile'].apply(lambda x: int(x[x.find('_')+1:-4]))
df_test['basename'] = df_test['basefile'].apply(lambda x: x[:x.find('_')])
df_test.sort_values(["root", "basename", "sequence"], inplace = True)
df_val = pd.DataFrame([file_path for file_path in Path('videos_faces/validation').glob('**/*.jpg')], columns=['file'])
df_val["root"] = df_val["file"].apply(lambda x: os.path.split(os.path.split(x)[0])[1])
df_val['basefile'] = df_val['file'].apply(lambda x: os.path.basename(x))
df_val['sequence'] = df_val['basefile'].apply(lambda x: int(x[x.find('_')+1:-4]))
df_val['basename'] = df_val['basefile'].apply(lambda x: x[:x.find('_')])
df_val.sort_values(["root", "basename", "sequence"], inplace = True)
def del_files(file_list):
for f in file_list:
try:
os.remove(f)
except:
continue
# resequence the faces
df_train['face_seq']=df_train.groupby('basename').cumcount()
df_test['face_seq']=df_test.groupby('basename').cumcount()
df_val['face_seq']=df_val.groupby('basename').cumcount()
# Remove just the files that are too long, in essence the last one or two out of 20
# get list of files and delete
train_del_files = df_train[df_train['face_seq'] > 16]['file'].to_list()
del_files(train_del_files)
test_del_files = df_test[df_test['face_seq'] > 16]['file'].to_list()
del_files(test_del_files)
val_del_files = df_val[df_val['face_seq'] > 16]['file'].to_list()
del_files(val_del_files)
# remove from dataframe
df_train.drop(df_train[df_train['face_seq'] > 16].index, inplace = True)
df_test.drop(df_test[df_test['face_seq'] > 16].index, inplace = True)
df_val.drop(df_val[df_val['face_seq'] > 16].index, inplace = True)
# check minimum number of frames per basename again - should be 17
print("train", df_train.groupby(['basename']).size().min())
print("test",df_test.groupby(['basename']).size().min())
print("val",df_val.groupby(['basename']).size().min())
# reset the indexes
df_train.reset_index(drop=True, inplace=True)
df_test.reset_index(drop=True, inplace=True)
df_val.reset_index(drop=True, inplace=True)
# Now all the dataframes are in order and all the files are in order
# we can load the files as arrays and create the model
df_train.head()
# create label
df_train['label'] = df_train['root'].str.replace('b','').astype(int)
df_test['label'] = df_test['root'].str.replace('b','').astype(int)
df_val['label'] = df_val['root'].str.replace('b','').astype(int)
# Need to load images into a numpy array - cannot find how to use generator for LSTMCNN
files = df_val['file'].to_list()
x_val = np.array([np.array(Image.open(fname).resize((160,160))) for fname in files])
#x_val.dump('videos_faces/x_val.npy')
files = df_train['file'].to_list()
x_train = np.array([np.array(Image.open(fname).resize((160,160))) for fname in files])
#x_train.dump('videos_faces/x_train.npy')
files = df_test['file'].to_list()
x_test = np.array([np.array(Image.open(fname).resize((160,160))) for fname in files])
#x_train.dump('videos_faces/x_train.npy')
# Get the labels and one-hot encode
y_train = df_train['label'].to_numpy()
y_train = to_categorical(y_train, num_classes=4)
#y_train.dump('videos_faces/y_train.npy')
y_val = df_val['label'].to_numpy()
y_val = to_categorical(y_val, num_classes=4)
#y_val.dump('videos_faces/y_val.npy')
y_test = df_test['label'].to_numpy()
y_test = to_categorical(y_test, num_classes=4)
#y_val.dump('videos_faces/y_val.npy')
# These are good for CNN, then can sequence out features for LSTM
# But we are going to do a CNNLSTM
print(x_train.shape, y_train.shape)
print(x_val.shape, y_val.shape)
print(x_test.shape, y_test.shape)
print("train", x_train.shape[0] / 17)
print("val", x_val.shape[0] / 17)
print("test", x_test.shape[0] / 17)
# Reshape to get sequences together
# reshape
x_train_lstm = x_train.reshape(5356, 17, 160, 160, 3)
x_val_lstm = x_val.reshape(1429, 17, 160, 160, 3)
x_test_lstm = x_test.reshape(1784, 17, 160, 160, 3)
y_train_lstm = y_train[1::17]
y_val_lstm = y_val[1::17]
y_test_lstm = y_test[1::17]
#LSTM Shapes
print(x_train_lstm.shape, y_train_lstm.shape)
print(x_val_lstm.shape, y_val_lstm.shape)
print(x_test_lstm.shape, y_test_lstm.shape)
import tensorflow as tf
print(tf.config.experimental.list_physical_devices('GPU'))
import tensorflow as tf
gpus = tf.config.experimental.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(gpus[0], True)
import tensorflow as tf
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
# Restrict TensorFlow to only allocate 1GB of memory on the first GPU
try:
tf.config.experimental.set_virtual_device_configuration(
gpus[0],
[tf.config.experimental.VirtualDeviceConfiguration(memory_limit=2048)])
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPUs,", len(logical_gpus), "Logical GPUs")
except RuntimeError as e:
# Virtual devices must be set before GPUs have been initialized
print(e)
from keras import layers, Sequential
from keras.layers import Dense, Flatten, GlobalAveragePooling2D, ConvLSTM2D, Dropout
from keras.models import Model
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ModelCheckpoint, EarlyStopping
from keras.optimizers import RMSprop, SGD, Adam
from keras.applications import MobileNetV2
#################
# CNN-LSTM. #
#################
lr = 0.0001
decay = 1e-6
img_height , img_width = 160, 160
seq_len = 17
model = Sequential()
model.add(ConvLSTM2D(filters = 16, kernel_size = (3, 3), return_sequences = False, data_format = "channels_last", input_shape = (seq_len, img_height, img_width, 3)))
model.add(Dropout(0.2))
model.add(Flatten())
model.add(Dense(256, activation="relu"))
model.add(Dropout(0.3))
model.add(Dense(4, activation = "softmax"))
model.summary()
# checkpoint callback
timestr = time.strftime("%Y%m%d-%H%M%S")
dir_name = '/host/efs/models/lstm_17/'
model_name = 'ConvLSTM2D'
best_model_file = dir_name + model_name + '_' + timestr + '_{epoch}.hdf5'
checkpoint = ModelCheckpoint(best_model_file,
monitor='accuracy',
verbose=1,
save_best_only=True,
mode='max')
# early stopping callback
early_stopping = EarlyStopping(monitor='val_loss', patience=4, verbose=1, mode='auto')
callbacks = [checkpoint, early_stopping]
model.compile(optimizer=Adam(lr=lr, decay=decay),
loss="categorical_crossentropy",
metrics =["accuracy"])
history = model.fit(
x=x_train_lstm,
y=y_train_lstm,
batch_size=8,
epochs=5,
verbose=1,
callbacks=callbacks,
validation_data=(x_val_lstm, y_val_lstm),
)
# Show graphs
print(history.history.keys())
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='upper left')
plt.show()
print(history.history.keys())
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='upper left')
plt.show()
# test on unseen data
y_pred = model.predict(x_test_lstm)
# to get around one hot encoding
y_pred = np.argmax(y_pred, axis = 1)
y_test = np.argmax(y_test_lstm, axis = 1)
# print the report
print(classification_report(y_test, y_pred))
class_labels = [0,1,2,3]
cm = confusion_matrix(y_test, y_pred, class_labels)
ax= plt.subplot()
sns.heatmap(cm, annot=True, ax = ax, fmt='g', cmap='Greens'); #annot=True to annotate cells
# labels, title and ticks
ax.set_xlabel('Predicted labels');ax.set_ylabel('True labels');
ax.set_title('Confusion Matrix');
ax.xaxis.set_ticklabels(class_labels); ax.yaxis.set_ticklabels(class_labels);
```
| github_jupyter |
```
import os
os.environ['THEANO_FLAGS'] = "device=gpu1"
import theano
from keras.models import Sequential
from keras.layers.core import Flatten, Dense, Dropout
from keras.layers.convolutional import Convolution2D, MaxPooling2D, ZeroPadding2D
from keras.optimizers import SGD
from keras.applications.resnet50 import ResNet50
from keras.models import Model
from keras import optimizers
from keras.preprocessing.image import ImageDataGenerator
from keras.models import load_model
import keras
import matplotlib.pyplot as plt
import pickle
class History(keras.callbacks.Callback):
def __init__(self,):
self.history = {'loss':[], 'val_loss':[], 'fbeta_score':[], 'val_fbeta_score':[], 'acc':[], 'val_acc':[]}
dataset_index = 4
model = ResNet50(include_top=False, weights='imagenet', input_shape=(3, 224, 224))
#model.summary()
model.layers[-1].outbound_nodes = []
model.outputs = [model.layers[-1].output]
output = model.get_layer('avg_pool').output
output = Flatten()(output)
output = Dense(output_dim=128, activation='relu')(output) # your newlayer Dense(...)
output = Dropout(0.5)(output)
output = Dense(output_dim=1, activation='sigmoid')(output)
new_model = Model(model.input, output)
#new_model.summary()
for layer in new_model.layers[:-4]:
layer.trainable = False
model = new_model
model.compile(loss='binary_crossentropy',
optimizer=optimizers.SGD(lr=1e-3, momentum=0.9),
metrics=['fbeta_score', 'accuracy'])
batch_size = 64
nb_classes = 2
nb_epoch = 50#change to 50 afterwards
nb_eval = 4#change to 4 afterwards
data_augmentation = True
# input image dimensions
img_rows, img_cols = 224, 224
# The CIFAR10 images are RGB.
img_channels = 3
import retina_dataset
# The data, shuffled and split between train and test sets:
(X_train, Y_train), (X_test, Y_test) = retina_dataset.load_data(dataset_index, 224)
print('X_train shape:', X_train.shape)
print(X_train.shape[0], 'train samples')
print(X_test.shape[0], 'test samples')
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
X_train /= 255
X_test /= 255
if not data_augmentation:
print('Not using data augmentation.')
model.fit(X_train, Y_train,
batch_size=batch_size,
nb_epoch=nb_epoch,
validation_data=(X_test, Y_test),
shuffle=True)
else:
print('Using real-time data augmentation.')
# This will do preprocessing and realtime data augmentation:
datagen = ImageDataGenerator(
featurewise_center=False, # set input mean to 0 over the dataset
samplewise_center=False, # set each sample mean to 0
featurewise_std_normalization=False, # divide inputs by std of the dataset
samplewise_std_normalization=False, # divide each input by its std
zca_whitening=False, # apply ZCA whitening
rotation_range=25, # randomly rotate images in the range (degrees, 0 to 180)
width_shift_range=0.3, # randomly shift images horizontally (fraction of total width)
height_shift_range=0.3, # randomly shift images vertically (fraction of total height)
horizontal_flip=True, # randomly flip images
vertical_flip=False) # randomly flip images
hist = History()
for _ in range(nb_eval):
print('Eval: '+str(_))
# Compute quantities required for featurewise normalization
# (std, mean, and principal components if ZCA whitening is applied).
datagen.fit(X_train)
# Fit the model on the batches generated by datagen.flow().
new_hist = model.fit_generator(datagen.flow(X_train, Y_train ,
batch_size=batch_size),
samples_per_epoch=X_train.shape[0],
nb_epoch=nb_epoch,
validation_data=(X_test, Y_test))
hist.history['loss'] += new_hist.history['loss']
hist.history['val_loss'] += new_hist.history['val_loss']
hist.history['fbeta_score'] += new_hist.history['fbeta_score']
hist.history['val_fbeta_score'] += new_hist.history['val_fbeta_score']
hist.history['acc'] += new_hist.history['acc']
hist.history['val_acc'] += new_hist.history['val_acc']
model.save('models/resnet_pretrained_for_dataset'+str(dataset_index)+'_lr=-3_dense=128.h5')
model.save_weights('models/resnet_pretrained_weights_for_dataset'+str(dataset_index)+'_lr=-3_dense=128.h5')
plt.plot(hist.history['fbeta_score'])
plt.plot(hist.history['val_fbeta_score'])
plt.title('model f1 score')
plt.ylabel('f1')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
plt.plot(hist.history['loss'])
plt.plot(hist.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
plt.plot(hist.history['acc'])
plt.plot(hist.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('acc')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
with open('models/resnet_pretrained_for_dataset'+str(dataset_index)+'_lr=-3_dense=128_trainning_history', 'w') as f: # Python 3: open(..., 'wb')
pickle.dump(hist, f)
import pickle
with open('models/resnet_pretrained_for_dataset'+str(dataset_index)+'_lr=-3_dense=128_trainning_history', 'w') as f: # Python 3: open(..., 'wb')
pickle.dump(hist, f)
```
| github_jupyter |
Nota basada en [liga](https://www.dropbox.com/s/s4ch0ww1687pl76/3.2.2.Factorizaciones_matriciales_SVD_Cholesky_QR.pdf?dl=0)
# Definiciones generales
En lo que sigue se supone una matriz cuadrada $A \in \mathbb{R}^{nxn}$.
## Eigenvalor (valor propio o característico)
El número $\lambda$ (real o complejo) se denomina *eigenvalor* de A si existe $v \in \mathbb{C}^n - \{0\}$ tal que $Av = \lambda v$. El vector $v$ se nombra eigenvector (vector propio o característico) de $A$ correspondiente al eigenvalor $\lambda$.
**Obs:**
* Una matriz con componentes reales puede tener eigenvalores y eigenvectores con valores en $\mathbb{C}$ o $\mathbb{C}^n$ respectivamente.
* El conjunto de eigenvalores se le llama **espectro de una matriz**.
* $A$ siempre tiene al menos un eigenvalor con eigenvector asociado.
**Nota:** Si A es simétrica entonces tiene eigenvalores reales y aún más: $A$ tiene eigenvectores reales linealmente independientes y forman un conjunto ortonormal. Entonces $A$ se escribe como un producto de tres matrices nombrado descomposición espectral: $$A = Q \Lambda Q^T$$ donde: $Q$ es una matriz ortogonal cuyas columnas son eigenvectores de $A$ y $\Lambda$ es una matriz diagonal con eigenvalores de $A$.
# Valores y vectores singulares de una matriz
En lo que sigue se supone $A \in \mathbb{R}^{mxn}$.
## Valor singular
El número $\sigma$ se denomina valor *singular* de $A$ si $\sigma = \sqrt{\lambda_{A^TA}} = \sqrt{\lambda_{AA^T}}$ donde: $\lambda_{A^TA}$ y $\lambda_{AA^T}$ es eigenvalor de $A^TA$ y $AA^T$ respectivamente .
**Obs:** la definición se realiza sobre $A^TA$ o $AA^T$ pues éstas matrices tienen el mismo espectro y además sus eigenvalores son reales y positivos por lo que $\sigma \in \mathbb{R}$ y de hecho $\sigma \geq 0$ (la raíz cuadrada se calcula para un eigenvalor no negativo).
## Vector singular izquierdo, vector singular derecho
Asociado con cada valor singular $\sigma$ existen vectores singulares $u,v$ que cumplen con la igualdad: $$Av = \sigma u .$$ Al vector $u$ se le nombra vector singular *izquierdo* y al vector $v$ se le nombra vector singular *derecho*.
## Descomposición en valores singulares (SVD)
Si $A \in \mathbb{R}^{mxn}$ entonces existen $U \in \mathbb{R}^{mxm}, V \in \mathbb{R}^{nxn}$ ortogonales tales que: $A = U\Sigma V^T$ con $\Sigma = diag(\sigma_1, \sigma_2, \dots, \sigma_p) \in \mathbb{R}^{mxn}$, $p = \min\{m,n\}$ y $\sigma_1 \geq \sigma_2 \geq \dots \geq \sigma_p \geq 0$.
Ver [3.3.c.Factorizacion_QR](https://github.com/ITAM-DS/analisis-numerico-computo-cientifico/blob/master/temas/III.computo_matricial/3.3.c.Factorizacion_QR.ipynb) para definición de matriz ortogonal.
**Obs:** La notación $\sigma_1$ hace referencia al valor singular más grande de A, $\sigma_2$ al segundo valor singular más grande de A y así sucesivamente.
**Obs2:** La SVD que se definió arriba es nombrada *SVD full*, hay una forma **truncada** en la que $U \in \mathbb{R}^{mxk}$, $V \in \mathbb{R}^{nxk}$ y $\Sigma \in \mathbb{R}^{kxk}$.
Existen diferentes propiedades de los valores y vectores singulares, aquí se enlistan algunas:
* Si $rank(A) = r$ entonces $r \leq p$ y $\sigma_1 \geq \sigma_2 \geq \dots \geq \sigma_r > \sigma_{r+1} = \sigma_{r+2} = \dots = \sigma_p = 0$.
* Si $rank(A) = r$ entonces $A = \displaystyle \sum_{i=0}^r \sigma_i u_i v_i^T$ con $u_i$ $i$-ésima columna de U y $v_i$ $i$-ésima columna de V.
* Geométricamente los valores singulares de una matriz $A \in \mathbb{R}^{mxn}$ son las longitudes de los semiejes del hiperelipsoide $E$ definido por $E = \{Ax : ||x|| \leq 1, \text{ con } ||\cdot || \text{ norma Euclidiana}\}$ y los vectores $u_i$ son direcciones de estos semiejes; los vectores $vi$'s tienen norma igual a $1$ por lo que se encuentran en una circunferencia de radio igual a $1$ y como $Av_i = \sigma u_i$ entonces $A$ mapea los vectores $v_i$'s a los semiejes $u_i$'s respectivamente:
<img src="https://dl.dropboxusercontent.com/s/1yqoe4qibyyej53/svd_2.jpg?dl=0" heigth="700" width="700">
* La SVD da bases ortogonales para los $4$ espacios fundamentales de una matriz: espacio columna, espacio nulo izquierdo, espacio nulo y espacio renglón:
<img src="https://dl.dropboxusercontent.com/s/g9giy9nz9yjh4ug/svd.jpg?dl=0" heigth="600" width="600">
* Si $t < r$ y $r=rank(A)$ entonces $A_t = \displaystyle \sum_{i=0}^t \sigma_i u_i v_i^T$ es una matriz de entre todas las matrices con $rank$ igual a t, que es más *cercana* a A (la cercanía se mide con una norma **matricial**).
Entre las aplicaciones de la SVD se encuentran:
* Procesamiento de imágenes y señales.
* Sistemas de recomendación (Netflix).
* Mínimos cuadrados.
* Componentes principales.
* Reconstrucción de imágenes.
# Método de Jacobi para calcular la SVD
La idea del método de Jacobi *one sided* consiste en multiplicar a la matriz $A \in \mathbb{R}^{m \times n}$ por la derecha de forma repetida por matrices ortogonales de nombre **rotaciones Givens** hasta que se converja a $U \Sigma$.
## Rotaciones
Si $u, v \in \mathbb{R}^2-\{0\}$ con $\ell = ||u||_2 = ||v||_2$ y se desea rotar al vector $u$ en sentido contrario a las manecillas del reloj por un ángulo $\theta$ para llevarlo a la dirección de $v$:
<img src="https://dl.dropboxusercontent.com/s/vq8eu0yga2x7cb2/rotation_1.png?dl=0" heigth="500" width="500">
A partir de las relaciones anteriores como $cos(\phi)=\frac{u_1}{\ell}, sen(\phi)=\frac{u_2}{\ell}$ se tiene: $v_1 = (cos\theta)u_1-(sen\theta)u_2$, $v_2=(sen\theta)u_1+(cos\theta)u_2$ equivalentemente:
$$\begin{array}{l}
\left[\begin{array}{c}
v_1\\
v_2
\end{array}
\right]
=
\left[ \begin{array}{cc}
cos\theta & -sen\theta\\
sen\theta & cos\theta
\end{array}
\right] \cdot
\left[\begin{array}{c}
u_1\\
u_2
\end{array}
\right]
\end{array}
$$
**Comentarios:**
* La matriz $R_O$ se llama matriz de **rotación** o **rotaciones Givens**, es una matriz ortogonal pues $R_O^TR_O=I_2$.
* La multiplicación $v=R_Ou$ es una rotación en sentido contrario a las manecillas del reloj. La multiplicación $u=R_O^Tv$ es una rotación en sentido de las manecillas del reloj y el ángulo asociado es $-\theta$.
* Obsérvese que $det(R_O)=1$.
**Ejemplo:**
Rotar al vector $v=(1,1)^T$ un ángulo de $45^o$ en sentido contrario a las manecillas del reloj.
**Solución:**
```
import numpy as np
import math
v=np.array([1,1])
```
La matriz $R_O$ es: $R_O = \left[ \begin{array}{cc}
cos(\frac{\pi}{4}) & -sen(\frac{\pi}{4})\\
sen(\frac{\pi}{4}) & cos(\frac{\pi}{4})
\end{array}
\right ]
$
```
theta=math.pi/4
RO=np.array([[math.cos(theta), -math.sin(theta)],
[math.sin(theta), math.cos(theta)]])
RO
RO@v
```
Obsérvese en el ejemplo anterior que se preservó su la longitud de $v$:
```
np.linalg.norm(v)
```
* Esto se cumple para todas las matrices ortogonales. Son **isometrías** bajo la norma $2$ o Euclidiana: $||R_0v||_2=||v||_2$
* Obsérvese en el ejemplo anterior que se hizo cero la entrada $v_1$ de $v$. Las matrices de rotación se utilizan para hacer ceros en entradas de un vector. Por ejemplo si $v=(v_1,v_2)^T$ y se desea hacer cero la entrada $v_2$ se puede utilizar la matriz de rotación:
$$R_O = \left[ \begin{array}{cc}
\frac{v_1}{\sqrt{v_1^2+v_2^2}} & \frac{v_2}{\sqrt{v_1^2+v_2^2}}\\
-\frac{v_2}{\sqrt{v_1^2+v_2^2}} & \frac{v_1}{\sqrt{v_1^2+v_2^2}}
\end{array}
\right ]
$$
pues:
$$\begin{array}{l}
\left[ \begin{array}{cc}
\frac{v_1}{\sqrt{v_1^2+v_2^2}} & \frac{v_2}{\sqrt{v_1^2+v_2^2}}\\
-\frac{v_2}{\sqrt{v_1^2+v_2^2}} & \frac{v_1}{\sqrt{v_1^2+v_2^2}}
\end{array}
\right ] \cdot
\left[\begin{array}{c}
v_1\\
v_2
\end{array}
\right]=
\left[ \begin{array}{c}
\frac{v_1^2+v_2^2}{\sqrt{v_1^2+v_2^2}}\\
\frac{-v_1v_2+v_1v_2}{\sqrt{v_1^2+v_2^2}}
\end{array}
\right ]
=
\left[ \begin{array}{c}
\frac{v_1^2+v_2^2}{\sqrt{v_1^2+v_2^2}}\\
0
\end{array}
\right ]=
\left[ \begin{array}{c}
||v||_2\\
0
\end{array}
\right ]
\end{array}
$$
Y definiendo $cos(\theta)=\frac{v_1}{\sqrt{v_1^2+v_2^2}}, sen(\theta)=\frac{v_2}{\sqrt{v_1^2+v_2^2}}$ se tiene :
$$
R_O=\left[ \begin{array}{cc}
cos\theta & sen\theta\\
-sen\theta & cos\theta
\end{array}
\right]
$$
que en el ejemplo anterior como $v=(1,1)^T$ entonces: $cos(\theta)=\frac{1}{\sqrt{2}}, sen(\theta)=\frac{1}{\sqrt{2}}$ por lo que $\theta=\frac{\pi}{4}$ y:
$$
R_O=\left[ \begin{array}{cc}
\frac{1}{\sqrt{2}} & \frac{1}{\sqrt{2}}\\
-\frac{1}{\sqrt{2}} & \frac{1}{\sqrt{2}}
\end{array}
\right]
$$
Para hacer cero la entrada $v_1$ de $v$ hay que usar:
$$\begin{array}{l}
R_O=\left[ \begin{array}{cc}
cos\theta & -sen\theta\\
sen\theta & cos\theta
\end{array}
\right]
=\left[ \begin{array}{cc}
\frac{1}{\sqrt{2}} & -\frac{1}{\sqrt{2}}\\
\frac{1}{\sqrt{2}} & \frac{1}{\sqrt{2}}
\end{array}
\right]
\end{array}
$$
## Algoritmo de Jacobi one sided
El producto de las rotaciones Givens construye a la matriz ortogonal $V \in \mathbb{R}^{n \times n}$:
$$AV \rightarrow W \in \mathbb{R}^{m \times n}$$
**Comentarios:**
* Las normas Euclidianas de las columnas de $W$ construyen a los valores singulares $\sigma_i \forall i=1,\dots,r$:
$$W = [U_1 \quad 0]\left[ \begin{array}{cc}
\Sigma & 0\\
0 & 0
\end{array}
\right]$$
con $U_1 \in \mathbb{R}^{m \times r}$ matriz con columnas ortonormales: $U_1^TU_1=I_r$ y $\Sigma = diag(\sigma_1,\dots, \sigma_r)$ matriz diagonal.
**Algoritmo one sided Jacobi**
Entrada:
* matriz $A \in \mathbb{R}^{m \times n}$: matriz a la que se le calculará su SVD.
* $TOL$ controla la convergencia del método.
* $maxsweeps$ número máximo de sweeps (descritos en los comentarios de abajo).
Salida:
* matrices $V \in \mathbb{R}^{n \times n}$ ortogonal, $W \in \mathbb{R}^{m \times n}$ representada en el algoritmo por $A^{(k)}$ para un valor de $k$ controlado por la convergencia (descrita en los comentarios de abajo).
Nota: se utilizará la notación $A^{(k)}=[a_1^{(k)} a_2^{(k)} \cdots a_n^{(k)}]$ con cada $a_i^{(k)}$ como $i$-ésima columna de $A$ y $k$ representa el *sweep*.
Definir $A^{(0)}=A$, $V^{(0)}=I_n$ (*sweep* $=0$).
* Mientras no se haya llegado al número máximo de sweeps ($k \leq maxsweeps$) o el número de columnas ortogonales sea menor a $\frac{n(n-1)}{2}$:
* Para todos los pares con índices $i<j$ generados con alguna metodología (descrita en la sección de abajo) y para k desde 0 hasta convergencia:
* Revisar si las columnas $a_i^{(k)}, a_j^{(k)}$ de $A^{(k)}$ son ortogonales (el chequeo se describe en los comentarios). Si son ortogonales se incrementa por uno la variable $num\text{_}columnas\text{_}ortogonales$. Si no son ortogonales:
* Calcular $\left[ \begin{array}{cc}
a & c\\
c & b
\end{array}
\right]$ la submatriz $(i,j)$ de $A^{T(0)}A^{(0)}$ donde: $a = ||a_i^{(k)}||_2^2, b=||a_j^{(k)}||_2^2, c=a_i^{T(k)}a_j^{(k)}$.
* Calcular la rotación Givens que diagonaliza $\left[ \begin{array}{cc}
a & c\\
c & b
\end{array}
\right]$ definiendo: $\xi = \frac{b-a}{2c}, t=\frac{signo(\xi)}{|\xi| + \sqrt{1+\xi^2}}, cs = \frac{1}{\sqrt{1+t^2}}, sn = cs*t$. Recordar que $signo(a) = \begin{cases}
1 &\text{ si } a \geq 0 ,\\
-1 &\text{ si } a < 0
\end{cases}$.
* Actualizar las columnas $i,j$ de $A^{(k)}$. Para $\ell$ de $1$ a $n$:
* $temp = A^{(k)}_{\ell i}$
* $A_{\ell i}^{(k)} = cs*temp - sn*A_{\ell j}^{(k)}$
* $A_{\ell j}^{(k)} = sn*temp + cs*A_{\ell j}^{(k)}$
* Actualizar a la matriz $V^{(k)}$. Para $\ell$ de $1$ a $n$:
* $temp = V_{\ell i}^{(k)}$
* $V_{\ell i}^{(k)} = cs*temp - sn*V_{\ell j}^{(k)}$
* $V_{\ell j}^{(k)} = sn*temp + cs*V_{\ell j}^{(k)}$
* Incrementar por uno la variable $k$ que cuenta el número de sweeps.
**Comentarios:**
* Las rotaciones se realizan en una secuencia con nombre *sweep*. Cada *sweep* consiste de como máximo $\frac{n(n-1)}{2}$ rotaciones (pues depende de cuántas columnas son o no ortogonales) y en cada *sweep* se ortogonalizan $2$ columnas. El número de *sweeps* a realizar se controla con la variable $maxsweeps$ y con la variable $num\text{_}columnas\text{_}ortogonales$ que va contando en cada sweep el número de columnas ortogonales.
* La convergencia del algoritmo anterior involucra dos aspectos:
* El número de columnas ortogonales que tenemos en un *sweep*: si en un *sweep* tal número es $\frac{n(n-1)}{2}$ (almacenado en la variable $num\text{_}columnas\text{_}ortogonales$) entonces el algoritmo termina.
* ¿Cómo reviso si las columnas $i,j$ de $A^{(k)}$ son ortogonales? si se cumple que
$$\frac{a_i^{T (k)}a_j^{(k)}}{||a_i^{(k)}||_2||a_j^{(k)}||_2} < TOL$$
con $TOL$ un valor menor o igual a $10^{-8}$ entonces son ortogonales las columnas $a_i^{(k)}, a_j^{(k)}$ de $A^{(k)}$.
* El ángulo $\theta$ se elige de acuerdo a $2$ criterios:
1) El ángulo es $0$ si $a_i^{T(k)}a_j^{(k)}=0$ y por lo tanto las columnas $i,j$ son ortogonales y no se hace rotación.
2) $\theta \in (\frac{-\pi}{4}, \frac{pi}{4})$ tal que $a_i^{T(k+1)}a_j^{(k+1)}=0$ y para este caso se calculan $\xi, t, cs,sn$ (las variables $cs, sn$ hacen referencia a $cos(\theta), sen(\theta)$.
* Las actualizaciones para $A,V$ en el algoritmo son de la forma: $A^{(k+1)} = A^{(k)}U^{(k)}, V^{(k+1)} = V^{(k)}U^{(k)}$ para $k>0$ donde: $U^{(k)}$ son matrices de rotación del plano $(i,j)$. Esto es, una identidad pero con elementos:
$$u_{ii}^{(k)} = cos(\theta), \quad u_{ij}^{(k)} = sen(\theta)$$
$$u_{ji}^{(k)}=-sen(\theta), \quad u_{jj}^{(k)}=cos(\theta).$$
* La multiplicación $A^{(k)}U^{(k)}$ sólo involucra a $2$ columnas de $A^{(k)}$:
$$(a_i^{(k+1)} a_j^{(k+1)}) = (a_i^{(k)} a_j^{(k)})\left[ \begin{array}{cc}
cos(\theta) & sen(\theta)\\
-sen(\theta) & cos(\theta)
\end{array}
\right]$$
* Al finalizar el algoritmo los valores singulares calculados son las normas Euclidianas de cada columna de $A^{(k)}$. Las columnas normalizadas de $A^{(k)}$ son las columnas de $U$.
**¿Cómo elijo los pares de columnas (i,j) a las que se les aplicará un proceso de ortogonalización?**
Hay distintas metodologías para la elección de las columnas $a_i^{(k)}, a_j^{(k)}$ de $A^{(k)}$. Se presenta a continuación una de ellas:
* Ordenamiento cíclico por renglones: un *sweep* involucra trabajar en los pares de columnas $(1,2), (1,3), \dots, (1,n), (2,3), (2,4), \dots, (n-1,n)$. Este ordenamiento siempre converge si $|\theta| \leq \frac{\pi}{4}$.
## ¿Dado el sistema $Ax=b$, $A \in \mathbb{R}^{n \times n}$ cómo se resuelve con la factorización $SVD$?
Paso 1: encontrar factores $U, \Sigma, V$ tales que $A=U \Sigma V^T$.
Paso 2: resolver el sistema diagonal $\Sigma d = U^Tb$.
Paso 3: realizar la multiplicación matriz vector para obtener $x$: $x=Vd$.
**Referencias:**
* Ver [4_SVD_y_reconstruccion_de_imagenes](https://github.com/ITAM-DS/Propedeutico/blob/master/Python/clases/3_algebra_lineal/4_SVD_y_reconstruccion_de_imagenes.ipynb) para definiciones de eigenvalores, eigenvectores y la descomposición en valores singulares o DVS (o SVD por sus siglas en inglés)
| github_jupyter |
This notebook demonstrates sample usage of dicompyler-core.
### Load some useful imports:
```
%matplotlib inline
import os
import numpy as np
from dicompylercore import dicomparser, dvh, dvhcalc
import matplotlib.pyplot as plt
import urllib.request
import os.path
```
### Download some example data:
```
%mkdir -p example_data
repo_url = 'https://github.com/dicompyler/dicompyler-core/blob/master/tests/testdata/file?raw=true'
# files = ['ct.0.dcm', 'rtss.dcm', 'rtplan.dcm', 'rtdose.dcm']
files = ['example_data/{}'.format(y) for y in ['rtss.dcm', 'rtdose.dcm']]
file_urls = [repo_url.replace('file', x) for x in files]
# Only download if the data is not present
[urllib.request.urlretrieve(x, y) for x, y in zip(file_urls, files) if not os.path.exists(y)]
```
### DICOM data can be easily accessed using convenience functions using the `dicompylercore.dicomparser.DicomParser` class:
```
rtss_dcm = files[0]
rtdose_dcm = files[1]
rtss = dicomparser.DicomParser(rtss_dcm)
rtdose = dicomparser.DicomParser(rtdose_dcm)
```
### Get a list of structures:
```
key = 5
structures = rtss.GetStructures()
structures[key]
```
### Iterate through slices of the dose grid:
```
planes = \
(np.array(rtdose.ds.GridFrameOffsetVector) \
* rtdose.ds.ImageOrientationPatient[0]) \
+ rtdose.ds.ImagePositionPatient[2]
dd = rtdose.GetDoseData()
from ipywidgets import FloatSlider, interactive
w = FloatSlider(
value=0.56,
min=planes[0],
max=planes[-1],
step=np.diff(planes)[0],
description='Slice Position (mm):',
)
def showdose(z):
plt.imshow(rtdose.GetDoseGrid(z) * dd['dosegridscaling'],
vmin=0,
vmax=dd['dosemax'] * dd['dosegridscaling'])
interactive(showdose, z=w)
```
### Access DVH data using the `dicompylercore.dvh.DVH` class:
```
heart = rtdose.GetDVHs()[key]
heart.name = structures[key]['name']
heart.describe()
heart.relative_volume.plot()
```
### Set the Rx dose to show volume statistics and relative dose:
```
heart.rx_dose = 14
heart.relative_dose().describe()
```
### Chain functions to view the DVH data in various formats:
```
tumorbed = rtdose.GetDVHs()[9]
tumorbed.name = structures[9]['name']
tumorbed.rx_dose = 14
tumorbed.relative_volume.differential.plot()
tumorbed.relative_dose().differential.absolute_dose().cumulative.plot()
```
### Access DVH statistics in multiple ways:
```
lung = rtdose.GetDVHs()[6]
lung.name = structures[6]['name']
lung.rx_dose = 14
lung.plot()
lung.max
lung.V5Gy
lung.relative_volume.V5Gy
lung.D2cc
lung.relative_dose().D2cc
lung.D1cc == lung.statistic('D1cc') == lung.dose_constraint(1, 'cc')
```
### Plot all DVHs found in a DICOM RT Dose DVH sequence:
```
plt.figure(figsize=(10, 6))
plt.axis([0, 20, 0, 100])
for s in structures.values():
if not s['empty']:
dvh.DVH.from_dicom_dvh(rtdose.ds, s['id'], name=s['name'], color=s['color']).relative_volume.plot()
```
### Calculate a DVH from a DICOM RT Structure Set & Dose via the `dicompylercore.dvhcalc.get_dvh` function:
```
b = dvhcalc.get_dvh(rtss_dcm, rtdose_dcm, key)
b.plot()
def compare_dvh(key=1):
structure = rtss.GetStructures()[key]
orig = dvh.DVH.from_dicom_dvh(rtdose.ds, key, name=structure['name'] + ' Orig')
calc = dvhcalc.get_dvh(rtss_dcm, rtdose_dcm, key)
calc.name = structure['name'] + ' Calc'
orig.compare(calc)
compare_dvh(key)
```
| github_jupyter |
# Exercise 9.02
Import Libraries & Process Data
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from tensorflow import random
dataset_training = pd.read_csv('../GOOG_train.csv')
dataset_training.head()
training_data = dataset_training[['Open']].values
training_data
from sklearn.preprocessing import MinMaxScaler
sc = MinMaxScaler(feature_range = (0, 1))
training_data_scaled = sc.fit_transform(training_data)
training_data_scaled
```
Create Data Time Stamps & Rehape the Data
```
X_train = []
y_train = []
for i in range(60, 1258):
X_train.append(training_data_scaled[i-60:i, 0])
y_train.append(training_data_scaled[i, 0])
X_train, y_train = np.array(X_train), np.array(y_train)
X_train = np.reshape(X_train, (X_train.shape[0], X_train.shape[1], 1))
X_train
```
Create & Compile an RNN Architecure
```
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM
from keras.layers import Dropout
seed = 1
np.random.seed(seed)
random.set_seed(seed)
model = Sequential()
model.add(LSTM(units = 100, return_sequences = True, input_shape = (X_train.shape[1], 1)))
# Adding a second LSTM layer
model.add(LSTM(units = 100, return_sequences = True))
# Adding a third LSTM layer
model.add(LSTM(units = 100, return_sequences = True))
# Adding a fourth LSTM layer
model.add(LSTM(units = 100))
# Adding the output layer
model.add(Dense(units = 1))
# Compiling the RNN
model.compile(optimizer = 'adam', loss = 'mean_squared_error')
# Fitting the RNN to the Training set
model.fit(X_train, y_train, epochs = 100, batch_size = 32)
```
Prepare the Test Data , Concatenate Test & Train Datasets
```
dataset_testing = pd.read_csv("../GOOG_test.csv")
actual_stock_price = dataset_testing[['Open']].values
actual_stock_price
total_data = pd.concat((dataset_training['Open'], dataset_testing['Open']), axis = 0)
inputs = total_data[len(total_data) - len(dataset_testing) - 60:].values
inputs = inputs.reshape(-1,1)
inputs = sc.transform(inputs)
X_test = []
for i in range(60, 81):
X_test.append(inputs[i-60:i, 0])
X_test = np.array(X_test)
X_test = np.reshape(X_test, (X_test.shape[0], X_test.shape[1], 1))
predicted_stock_price = model.predict(X_test)
predicted_stock_price = sc.inverse_transform(predicted_stock_price)
```
Visualize the Results
```
# Visualising the results
plt.plot(actual_stock_price, color = 'green', label = 'Real Alphabet Stock Price',ls='--')
plt.plot(predicted_stock_price, color = 'red', label = 'Predicted Alphabet Stock Price',ls='-')
plt.title('Predicted Stock Price')
plt.xlabel('Time in days')
plt.ylabel('Real Stock Price')
plt.legend()
plt.show()
```
| github_jupyter |
```
%matplotlib inline
```
Autograd: Automatic Differentiation
===================================
Central to all neural networks in PyTorch is the ``autograd`` package.
Let’s first briefly visit this, and we will then go to training our
first neural network.
The ``autograd`` package provides automatic differentiation for all operations
on Tensors. It is a define-by-run framework, which means that your backprop is
defined by how your code is run, and that every single iteration can be
different.
Let us see this in more simple terms with some examples.
Tensor
--------
``torch.Tensor`` is the central class of the package. If you set its attribute
``.requires_grad`` as ``True``, it starts to track all operations on it. When
you finish your computation you can call ``.backward()`` and have all the
gradients computed automatically. The gradient for this tensor will be
accumulated into ``.grad`` attribute.
To stop a tensor from tracking history, you can call ``.detach()`` to detach
it from the computation history, and to prevent future computation from being
tracked.
To prevent tracking history (and using memory), you can also wrap the code block
in ``with torch.no_grad():``. This can be particularly helpful when evaluating a
model because the model may have trainable parameters with `requires_grad=True`,
but for which we don't need the gradients.
There’s one more class which is very important for autograd
implementation - a ``Function``.
``Tensor`` and ``Function`` are interconnected and build up an acyclic
graph, that encodes a complete history of computation. Each tensor has
a ``.grad_fn`` attribute that references a ``Function`` that has created
the ``Tensor`` (except for Tensors created by the user - their
``grad_fn is None``).
If you want to compute the derivatives, you can call ``.backward()`` on
a ``Tensor``. If ``Tensor`` is a scalar (i.e. it holds a one element
data), you don’t need to specify any arguments to ``backward()``,
however if it has more elements, you need to specify a ``gradient``
argument that is a tensor of matching shape.
```
import torch
```
Create a tensor and set requires_grad=True to track computation with it
```
x = torch.ones(2, 2, requires_grad=True)
print(x)
```
Do an operation of tensor:
```
y = x + 2
print(y)
```
``y`` was created as a result of an operation, so it has a ``grad_fn``.
```
print(y.grad_fn)
```
Do more operations on y
```
z = y * y * 3
out = z.mean()
print(z, out)
```
``.requires_grad_( ... )`` changes an existing Tensor's ``requires_grad``
flag in-place. The input flag defaults to ``False`` if not given.
```
a = torch.randn(2, 2)
a = ((a * 3) / (a - 1))
print(a.requires_grad)
a.requires_grad_(True)
print(a.requires_grad)
b = (a * a).sum()
print(b.grad_fn)
```
Gradients
---------
Let's backprop now
Because ``out`` contains a single scalar, ``out.backward()`` is
equivalent to ``out.backward(torch.tensor(1))``.
```
out.backward()
```
print gradients d(out)/dx
```
print(x.grad)
```
You should have got a matrix of ``4.5``. Let’s call the ``out``
*Tensor* “$o$”.
We have that $o = \frac{1}{4}\sum_i z_i$,
$z_i = 3(x_i+2)^2$ and $z_i\bigr\rvert_{x_i=1} = 27$.
Therefore,
$\frac{\partial o}{\partial x_i} = \frac{3}{2}(x_i+2)$, hence
$\frac{\partial o}{\partial x_i}\bigr\rvert_{x_i=1} = \frac{9}{2} = 4.5$.
You can do many crazy things with autograd!
```
x = torch.randn(3, requires_grad=True)
y = x * 2
while y.data.norm() < 1000:
y = y * 2
print(y)
gradients = torch.tensor([0.1, 1.0, 0.0001], dtype=torch.float)
y.backward(gradients)
print(x.grad)
```
You can also stop autograd from tracking history on Tensors
with ``.requires_grad=True`` by wrapping the code block in
``with torch.no_grad()``:
```
print(x.requires_grad)
print((x ** 2).requires_grad)
with torch.no_grad():
print((x ** 2).requires_grad)
```
**Read Later:**
Documentation of ``autograd`` and ``Function`` is at
https://pytorch.org/docs/autograd
| github_jupyter |
# Explore the GPyTorch and Celerité API
We start by exploring the APIs of the two packages `GPyTorch` and `celerite`. They are both packages for scalable Gaussian Processes with different strategies for doing the scaling.
```
import gpytorch
import celerite
gpytorch.__version__, celerite.__version__
```
We'll need some other standard and astronomy-specific imports and configurations.
```
import numpy as np
import matplotlib.pyplot as plt
import astropy.units as u
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
```
Let's draw synthetic time series "data" with a Gaussian process from celerite. This approach is useful, since we know the answer: and the kernel that generated the data and its parameter values. We'll pick Matérn kernels, since both frameworks offer them out-of-the-box. Technically, the celerite Matern is an approximation, but we'll be sure to make draws with parameter values where the approximation will be near-exact.
## Matérn 3/2 with celerite.
This kernel is characterized by two parameters:
$k(\tau) = \sigma^2\,\left(1+
\frac{\sqrt{3}\,\tau}{\rho}\right)\,
\exp\left(-\frac{\sqrt{3}\,\tau}{\rho}\right)$
Here are the inputs for `celerite`:
> Args:
- log_sigma (float): The log of the parameter :math:`\sigma`.
- log_rho (float): The log of the parameter :math:`\rho`.
- eps (Optional[float]): The value of the parameter :math:`\epsilon`.
(default: `0.01`)
```
from celerite import terms
true_rho = 1.5
true_sigma = 1.2
true_log_sigma = np.log(true_sigma)
true_log_rho = np.log(true_rho) # Has units of time, so 1/f
kernel_matern = terms.Matern32Term(log_sigma=true_log_sigma, log_rho=true_log_rho, eps=0.00001)
t_vec = np.linspace(0, 40, 500)
gp = celerite.GP(kernel_matern, mean=0, fit_mean=True)
gp.compute(t_vec)
y_true = gp.sample()
noise = np.random.normal(0, 0.3, size=len(y_true))
y_obs = y_true + noise
plt.plot(t_vec, y_obs, label='Noisy observation')
plt.plot(t_vec, y_true, label='"Truth"')
plt.xlabel('$t$')
plt.ylabel('$y$')
plt.legend();
```
Ok, we have a dataset to work with.
## Now with GPyTorch and RBF kernel
```
import torch
t_ten = torch.from_numpy(t_vec)
y_ten = torch.from_numpy(y_obs)
train_x = t_ten.to(torch.float32)
train_y = y_ten.to(torch.float32)
# We will use the simplest form of GP model, exact inference
class ExactGPModel(gpytorch.models.ExactGP):
def __init__(self, train_x, train_y, likelihood):
super(ExactGPModel, self).__init__(train_x, train_y, likelihood)
self.mean_module = gpytorch.means.ConstantMean()
self.covar_module = gpytorch.kernels.ScaleKernel(gpytorch.kernels.MaternKernel(nu=3/2))
def forward(self, x):
mean_x = self.mean_module(x)
covar_x = self.covar_module(x)
return gpytorch.distributions.MultivariateNormal(mean_x, covar_x)
# initialize likelihood and model
likelihood = gpytorch.likelihoods.GaussianLikelihood()
model = ExactGPModel(train_x, train_y, likelihood)
```
### Train the model.
```
# Find optimal model hyperparameters
model.train()
likelihood.train()
model.state_dict()
with gpytorch.settings.max_cg_iterations(5000):
# Use the adam optimizer
optimizer = torch.optim.Adam([
{'params': model.parameters()}, # Includes GaussianLikelihood parameters
], lr=0.1)
# "Loss" for GPs - the marginal log likelihood
mll = gpytorch.mlls.ExactMarginalLogLikelihood(likelihood, model)
training_iter = 300
for i in range(training_iter):
# Zero gradients from previous iteration
optimizer.zero_grad()
# Output from model
output = model(train_x)
# Calc loss and backprop gradients
loss = -mll(output, train_y)
loss.backward()
if (i % 20) == 0:
print('Iter %d/%d - Loss: %.3f lengthscale: %.3f noise: %.6f' % (
i + 1, training_iter, loss.item(),
model.covar_module.base_kernel.raw_lengthscale.item(),
model.likelihood.noise.item()
))
#print(list(model.parameters()))
optimizer.step()
```
### How did it do?
```
# Get into evaluation (predictive posterior) mode
model.eval()
likelihood.eval()
# Test points are regularly spaced along [0,1]
# Make predictions by feeding model through likelihood
with torch.no_grad(), gpytorch.settings.fast_pred_var(), gpytorch.settings.max_cg_iterations(9000):
test_x = torch.linspace(0, 40, 501, dtype=torch.float32)
observed_pred = likelihood(model(test_x))
with torch.no_grad():
# Initialize plot
f, ax = plt.subplots(1, 1, figsize=(22, 9))
# Get upper and lower confidence bounds
lower, upper = observed_pred.confidence_region()
# Plot training data as black stars
ax.plot(train_x.numpy(), train_y.numpy(), 'k.', alpha=0.5)
# Plot predictive means as blue line
ax.plot(t_vec, y_true, lw=4)
ax.plot(test_x.numpy(), observed_pred.mean.numpy(), lw=4)
# Shade between the lower and upper confidence bounds
ax.fill_between(test_x.numpy(), lower.numpy(), upper.numpy(), alpha=0.5, color='#2ecc71')
ax.legend(['Observed Data', 'Truth', 'Mean', '2 $\sigma$ Confidence'])
```
Nice!
What are the four parameters?
```
model.mean_module.constant
likelihood.raw_noise
model.covar_module.raw_outputscale
model.covar_module.base_kernel.raw_lengthscale
```
| github_jupyter |
<a href="https://colab.research.google.com/github/bilalProgTech/mtech-data-science/blob/master/AISem3/CW/AI_Sem_3_A1_BilalHungund_D013.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Problem Statement - Home Credit Risk¶
Many people struggle to get loans due to insufficient or non-existent credit histories. And, unfortunately, this population is often taken advantage of by untrustworthy lenders.
Home Credit strives to broaden financial inclusion for the unbanked population by providing a positive and safe borrowing experience. In order to make sure this underserved population has a positive loan experience, Home Credit makes use of a variety of alternative data--including telco and transactional information--to predict their clients' repayment abilities.
While Home Credit is currently using various statistical and machine learning methods to make these predictions, they're challenging Kagglers to help them unlock the full potential of their data. Doing so will ensure that clients capable of repayment are not rejected and that loans are given with a principal, maturity, and repayment calendar that will empower their clients to be successful.
# Link Reference
https://www.kaggle.com/c/home-credit-default-risk/
# Approaches to predict the home credit risk
* LGBMClassifier (Ensemble Technique)
# Leaderboard Accuracy
Private Score: 0.77379 <br>
Public Score: 0.77186
# Downloading the data
```
from google.colab import drive
drive.mount('/content/gdrive')
import os
os.environ['KAGGLE_CONFIG_DIR'] = '/content/gdrive/My Drive/Kaggle'
!pwd
! kaggle competitions download -c home-credit-default-risk
!unzip \*.zip && rm *.zip
```
# Uploading the data
```
import pandas as pd
import numpy as np
pos_cash_balance = pd.read_csv('/content/POS_CASH_balance.csv')
train = pd.read_csv('/content/application_train.csv')
test = pd.read_csv('/content/application_test.csv')
bureau = pd.read_csv('/content/bureau.csv')
bureau_balance = pd.read_csv('/content/bureau_balance.csv')
cc_balance = pd.read_csv('/content/credit_card_balance.csv')
ins_payment = pd.read_csv('/content/installments_payments.csv')
prev_app = pd.read_csv('/content/previous_application.csv')
pos_cash_balance.head()
train.head()
bureau.head()
bureau_balance.head()
cc_balance.head()
ins_payment.head()
prev_app.head()
train.shape, test.shape, bureau.shape, bureau_balance.shape, cc_balance.shape, ins_payment.shape, prev_app.shape, pos_cash_balance.shape
```
# Preprocessing bureau balance and bureau data
```
operation_mean = ['mean']
operation_sum = ['sum']
operation_count = ['count']
bb_op = {'MONTHS_BALANCE': operation_mean, 'STATUS': operation_count}
bb_grouping = bureau_balance.groupby(['SK_ID_BUREAU'])
bb_groupby = bb_grouping.agg(bb_op)
bb_groupby.head()
bb_groupby.columns = ['BB_' + '_'.join(col).strip() for col in bb_groupby.columns.values]
bb_groupby.reset_index(inplace=True)
bb_groupby.head()
bureau = pd.merge(bureau, bb_groupby)
bureau.head()
bureau.isnull().sum() / bureau.shape[0]
bureau.describe()
bureau = bureau.drop(['AMT_CREDIT_MAX_OVERDUE'], axis=1)
bureau.fillna(0, inplace=True)
bureau.head()
bureau.columns
bureau_op = {'CREDIT_ACTIVE': operation_count, 'CREDIT_CURRENCY': operation_count,
'DAYS_CREDIT': operation_mean, 'CREDIT_DAY_OVERDUE': operation_mean,
'DAYS_CREDIT_ENDDATE': operation_mean, 'DAYS_ENDDATE_FACT': operation_mean,
'CNT_CREDIT_PROLONG': operation_mean, 'AMT_CREDIT_SUM': operation_mean,
'AMT_CREDIT_SUM_DEBT': operation_mean, 'AMT_CREDIT_SUM_LIMIT': operation_mean,
'AMT_CREDIT_SUM_OVERDUE': operation_mean, 'CREDIT_TYPE': operation_count,
'DAYS_CREDIT_UPDATE': operation_mean, 'AMT_ANNUITY': operation_mean,
'BB_MONTHS_BALANCE_mean': operation_mean, 'BB_STATUS_count': operation_sum}
bureau_grouping = bureau.groupby(['SK_ID_CURR'])
bureau_groupby = bureau_grouping.agg(bureau_op)
bureau_groupby.head()
bureau_groupby.columns = ['B_' + '_'.join(col).strip() for col in bureau_groupby.columns.values]
bureau_groupby.reset_index(inplace=True)
bureau_groupby.head()
```
# Preprocessing behavioural data of client
1. POS Cash Balance
2. Credit Card Balance
3. Installments Payments
4. Previous Application
```
pos_cash_balance.head()
pos_cash_balance_op = {'NAME_CONTRACT_STATUS': operation_count, 'MONTHS_BALANCE': operation_mean,
'CNT_INSTALMENT': operation_mean, 'CNT_INSTALMENT_FUTURE': operation_mean,
'SK_DPD': operation_mean, 'SK_DPD_DEF': operation_mean}
pos_cash_balance_grouping = pos_cash_balance.groupby(['SK_ID_PREV', 'SK_ID_CURR'])
pos_cash_balance_groupby = pos_cash_balance_grouping.agg(pos_cash_balance_op)
pos_cash_balance_groupby.head()
pos_cash_balance_groupby.columns = ['POS_' + '_'.join(col).strip() for col in pos_cash_balance_groupby.columns.values]
pos_cash_balance_groupby.reset_index(inplace=True)
pos_cash_balance_groupby.head()
cc_balance.isnull().sum() / cc_balance.shape[0]
cc_balance.head()
cc_balance_op = {'NAME_CONTRACT_STATUS': operation_count, 'MONTHS_BALANCE': operation_mean,
'AMT_TOTAL_RECEIVABLE': operation_mean, 'CNT_DRAWINGS_CURRENT': operation_mean,
'AMT_PAYMENT_TOTAL_CURRENT': operation_mean, 'AMT_CREDIT_LIMIT_ACTUAL': operation_mean,
'AMT_BALANCE': operation_mean, 'SK_DPD': operation_mean, 'SK_DPD_DEF': operation_mean}
cc_balance_grouping = cc_balance.groupby(['SK_ID_PREV', 'SK_ID_CURR'])
cc_balance_groupby = cc_balance_grouping.agg(cc_balance_op)
cc_balance_groupby.head()
cc_balance_groupby.columns = ['CC_' + '_'.join(col).strip() for col in cc_balance_groupby.columns.values]
cc_balance_groupby.reset_index(inplace=True)
cc_balance_groupby.head()
ins_payment.head()
ins_payment_op = {'NUM_INSTALMENT_VERSION': operation_mean,
'NUM_INSTALMENT_NUMBER': operation_mean, 'DAYS_INSTALMENT': operation_mean,
'DAYS_ENTRY_PAYMENT': operation_mean, 'AMT_INSTALMENT': operation_mean,
'AMT_PAYMENT': operation_mean}
ins_payment_grouping = ins_payment.groupby(['SK_ID_PREV', 'SK_ID_CURR'])
ins_payment_groupby = ins_payment_grouping.agg(ins_payment_op)
ins_payment_groupby.head()
ins_payment_groupby.columns = ['IP_' + '_'.join(col).strip() for col in ins_payment_groupby.columns.values]
ins_payment_groupby.reset_index(inplace=True)
ins_payment_groupby.head()
prev_app.shape
prev_app = pd.merge(prev_app, ins_payment_groupby, on=['SK_ID_CURR', 'SK_ID_PREV'], how='left')
prev_app = pd.merge(prev_app, cc_balance_groupby, on=['SK_ID_CURR', 'SK_ID_PREV'], how='left')
prev_app = pd.merge(prev_app, pos_cash_balance_groupby, on=['SK_ID_CURR', 'SK_ID_PREV'], how='left')
print(prev_app.shape)
prev_app.head()
prev_app.dtypes
pos_in_prevapp = prev_app[prev_app.columns[pd.Series(prev_app.columns).str.startswith('POS_')]]
floats = pos_in_prevapp.select_dtypes('float64').columns
prev_app[floats] = prev_app[floats].fillna(0)
ip_in_prevapp = prev_app[prev_app.columns[pd.Series(prev_app.columns).str.startswith('IP_')]]
floats = ip_in_prevapp.select_dtypes('float64').columns
prev_app[floats] = prev_app[floats].fillna(0)
cc_in_prevapp = prev_app[prev_app.columns[pd.Series(prev_app.columns).str.startswith('CC_')]]
floats = cc_in_prevapp.select_dtypes('float64').columns
prev_app[floats] = prev_app[floats].fillna(0)
objects = prev_app.select_dtypes('object').columns
prev_app[objects] = prev_app[objects].fillna('Unknown')
floats = prev_app.select_dtypes('float64').columns
prev_app[floats] = prev_app[floats].fillna(0)
ints = prev_app.select_dtypes('int64').columns
prev_app[ints] = prev_app[ints].fillna(0)
prev_app.head()
prev_app_op = {'AMT_ANNUITY': operation_mean,
'AMT_APPLICATION': operation_mean,'AMT_CREDIT': operation_mean,
'AMT_DOWN_PAYMENT': operation_mean,'AMT_GOODS_PRICE': operation_mean,
'HOUR_APPR_PROCESS_START': operation_mean, 'NFLAG_LAST_APPL_IN_DAY': operation_mean,
'RATE_DOWN_PAYMENT': operation_mean,'RATE_INTEREST_PRIMARY': operation_mean,
'RATE_INTEREST_PRIVILEGED': operation_mean,
'DAYS_DECISION': operation_mean,'SELLERPLACE_AREA': operation_mean,
'CNT_PAYMENT': operation_mean,
'DAYS_FIRST_DRAWING': operation_mean,'DAYS_FIRST_DUE': operation_mean,
'DAYS_LAST_DUE_1ST_VERSION': operation_mean,'DAYS_LAST_DUE': operation_mean,
'DAYS_TERMINATION': operation_mean,'NFLAG_INSURED_ON_APPROVAL': operation_mean,
'IP_NUM_INSTALMENT_VERSION_mean': operation_mean,'IP_NUM_INSTALMENT_NUMBER_mean': operation_mean,
'IP_DAYS_INSTALMENT_mean': operation_mean,'IP_DAYS_ENTRY_PAYMENT_mean': operation_mean,
'IP_AMT_INSTALMENT_mean': operation_mean,'IP_AMT_PAYMENT_mean': operation_mean,'CC_MONTHS_BALANCE_mean': operation_mean,
'CC_AMT_TOTAL_RECEIVABLE_mean': operation_mean,'CC_CNT_DRAWINGS_CURRENT_mean': operation_mean,
'CC_AMT_PAYMENT_TOTAL_CURRENT_mean': operation_mean,'CC_AMT_CREDIT_LIMIT_ACTUAL_mean': operation_mean,
'CC_AMT_BALANCE_mean': operation_mean,'CC_SK_DPD_mean': operation_mean,
'CC_SK_DPD_DEF_mean': operation_mean,
'POS_MONTHS_BALANCE_mean': operation_mean,'POS_CNT_INSTALMENT_mean': operation_mean,
'POS_CNT_INSTALMENT_FUTURE_mean': operation_mean,'POS_SK_DPD_mean': operation_mean,
'POS_SK_DPD_DEF_mean': operation_mean, 'NAME_CONTRACT_TYPE': operation_count,
'WEEKDAY_APPR_PROCESS_START': operation_count,
'FLAG_LAST_APPL_PER_CONTRACT': operation_count,'NAME_CASH_LOAN_PURPOSE': operation_count,
'NAME_CONTRACT_STATUS': operation_count,'NAME_PAYMENT_TYPE': operation_count,
'NAME_PAYMENT_TYPE': operation_count,
'CODE_REJECT_REASON': operation_count,'NAME_TYPE_SUITE': operation_count,
'NAME_CLIENT_TYPE': operation_count,'NAME_GOODS_CATEGORY': operation_count,
'NAME_PORTFOLIO': operation_count,'NAME_PRODUCT_TYPE': operation_count,
'CHANNEL_TYPE': operation_count,
'NAME_SELLER_INDUSTRY': operation_count,
'NAME_YIELD_GROUP': operation_count,
'CC_NAME_CONTRACT_STATUS_count': operation_sum, 'POS_NAME_CONTRACT_STATUS_count': operation_sum}
prev_app_grouping = prev_app.groupby(['SK_ID_CURR'])
prev_app_groupby = prev_app_grouping.agg(prev_app_op)
prev_app_groupby.head()
prev_app_groupby.columns = ['PA_' + '_'.join(col).strip() for col in prev_app_groupby.columns.values]
prev_app_groupby.reset_index(inplace=True)
prev_app_groupby.head()
count_var = prev_app_groupby[prev_app_groupby.columns[pd.Series(prev_app_groupby.columns).str.contains('_count')]].columns
prev_app_groupby[count_var] = prev_app_groupby[count_var].astype('int64')
prev_app_groupby.head()
prev_app_groupby.isnull().sum()
bureau_groupby.isnull().sum()
combine = train.append(test)
combine.shape, train.shape, test.shape
```
# Merging all the data into train and test set
```
combine = pd.merge(combine, bureau_groupby, how='left', on=['SK_ID_CURR'])
combine = pd.merge(combine, prev_app_groupby, how='left', on=['SK_ID_CURR'])
combine.shape, prev_app_groupby.shape, bureau_groupby.shape
print(len(combine.select_dtypes('object').columns))
objects = combine.select_dtypes('object').columns
combine[objects] = combine[objects].fillna('Unknown')
combine.select_dtypes('object').columns
print(len(combine.select_dtypes('float64').columns))
float64 = combine.select_dtypes('float64').columns[1:]
combine[float64] = combine[float64].fillna(combine[float64].mean())
combine.select_dtypes('float64').columns
print(len(combine.select_dtypes('int64').columns))
int64 = combine.select_dtypes('int64').columns
combine[int64] = combine[int64].fillna(combine[int64].mean())
combine.select_dtypes('int64').columns
count_var = combine[combine.columns[pd.Series(combine.columns).str.contains('_count')]].columns
combine[count_var] = combine[count_var].astype('int64')
combine.isnull().sum().sum(), combine.shape
combine = pd.get_dummies(combine)
combine.shape
X = combine[combine['TARGET'].isnull()!=True].drop(['TARGET', 'SK_ID_CURR'], axis=1)
y = combine[combine['TARGET'].isnull()!=True]['TARGET'].reset_index(drop=True)
X_test = combine[combine['TARGET'].isnull()==True].drop(['TARGET','SK_ID_CURR'], axis=1)
X.shape, y.shape, X_test.shape
```
# Data Modelling
```
from sklearn.model_selection import KFold, StratifiedKFold
from sklearn.metrics import confusion_matrix, recall_score, accuracy_score, precision_score, roc_auc_score, log_loss
from sklearn.ensemble import RandomForestClassifier
from lightgbm import LGBMClassifier
err_as = []
err_rs = []
err_ps = []
err_roc = []
err_ll = []
y_pred_tot_lgm = []
features = X.columns
feature_importance_df = pd.DataFrame()
fold = StratifiedKFold(n_splits=5)
i = 1
for train_index, test_index in fold.split(X, y):
x_train, x_val = X.iloc[train_index], X.iloc[test_index]
y_train, y_val = y[train_index], y[test_index]
m = LGBMClassifier(max_depth=5,
learning_rate=0.05,
n_estimators=5000,
min_child_weight=0.01,
colsample_bytree=0.5,
random_state=1994)
m.fit(x_train, y_train,
eval_set=[(x_train,y_train),(x_val, y_val)],
early_stopping_rounds=200,
eval_metric='auc',
verbose=200)
pred_y = m.predict(x_val)
prob_pred = m.predict_proba(x_val)[:,1]
fold_importance_df = pd.DataFrame()
fold_importance_df["Feature"] = features
fold_importance_df["importance"] = m.feature_importances_
fold_importance_df["fold"] = i + 1
feature_importance_df = pd.concat([feature_importance_df, fold_importance_df], axis=0)
print("Fold ",i, " Accuracy: ",(accuracy_score(pred_y, y_val)))
print("Fold ",i, " Recall: ",(recall_score(pred_y, y_val)))
print("Fold ",i, " Precision: ",(precision_score(pred_y, y_val)))
print("Fold ",i, " ROC AUC: ",(roc_auc_score(y_val, prob_pred)))
print("Fold ",i, " Logloss: ",(log_loss(y_val, prob_pred)))
print(confusion_matrix(pred_y, y_val))
err_as.append(accuracy_score(pred_y, y_val))
err_rs.append(recall_score(pred_y, y_val))
err_ps.append(precision_score(pred_y, y_val))
err_roc.append(roc_auc_score(y_val, prob_pred))
err_ll.append(log_loss(y_val, prob_pred))
pred_test = m.predict_proba(X_test)[:,1]
i = i + 1
y_pred_tot_lgm.append(pred_test)
print('Mean Accuracy Score on CV-5: ', np.mean(err_as, 0))
print('Mean Precision Score on CV-5: ', np.mean(err_ps, 0))
print('Mean Recall Score on CV-5: ', np.mean(err_rs, 0))
print('Mean ROC AUC Score on CV-5: ', np.mean(err_roc, 0))
print('Mean Logloss Score on CV-5: ', np.mean(err_ll, 0))
```
# Feature Engineering
```
all_feat = feature_importance_df[["Feature",
"importance"]].groupby("Feature").mean().sort_values(by="importance",
ascending=False)
all_feat.reset_index(inplace=True)
important_feat = list(all_feat['Feature'])
all_feat.head(20)
df = X[important_feat]
corr_matrix = df.corr().abs()
upper = corr_matrix.where(np.triu(np.ones(corr_matrix.shape), k=1).astype(np.bool))
high_cor = [column for column in upper.columns if any(upper[column] > 0.98)]
print(len(high_cor))
print(high_cor)
features = [i for i in important_feat if i not in high_cor]
print(len(features))
print(features)
```
# Applying LGBM Classifier with Feature Engineering
```
X1 = X[features]
X_test1 = X_test[features]
err_as = []
err_rs = []
err_ps = []
err_roc = []
err_ll = []
y_pred_tot_lgm_1 = []
fold = StratifiedKFold(n_splits=5)
i = 1
for train_index, test_index in fold.split(X1, y):
x_train, x_val = X1.iloc[train_index], X1.iloc[test_index]
y_train, y_val = y[train_index], y[test_index]
m = LGBMClassifier(max_depth=5,
learning_rate=0.05,
n_estimators=5000,
min_child_weight=0.01,
colsample_bytree=0.5,
random_state=1994)
m.fit(x_train, y_train,
eval_set=[(x_train,y_train),(x_val, y_val)],
early_stopping_rounds=200,
eval_metric='auc',
verbose=200)
pred_y = m.predict(x_val)
prob_pred = m.predict_proba(x_val)[:,1]
print("Fold ",i, " Accuracy: ",(accuracy_score(pred_y, y_val)))
print("Fold ",i, " Recall: ",(recall_score(pred_y, y_val)))
print("Fold ",i, " Precision: ",(precision_score(pred_y, y_val)))
print("Fold ",i, " ROC AUC: ",(roc_auc_score(y_val, prob_pred)))
print("Fold ",i, " Logloss: ",(log_loss(y_val, prob_pred)))
print(confusion_matrix(pred_y, y_val))
err_as.append(accuracy_score(pred_y, y_val))
err_rs.append(recall_score(pred_y, y_val))
err_ps.append(precision_score(pred_y, y_val))
err_roc.append(roc_auc_score(y_val, prob_pred))
err_ll.append(log_loss(y_val, prob_pred))
pred_test = m.predict_proba(X_test1)[:,1]
i = i + 1
y_pred_tot_lgm_1.append(pred_test)
print('Mean Accuracy Score on CV-5: ', np.mean(err_as, 0))
print('Mean Precision Score on CV-5: ', np.mean(err_ps, 0))
print('Mean Recall Score on CV-5: ', np.mean(err_rs, 0))
print('Mean ROC AUC Score on CV-5: ', np.mean(err_roc, 0))
print('Mean Logloss Score on CV-5: ', np.mean(err_ll, 0))
```
# Submission file of test set for competition
```
submission = pd.DataFrame()
submission['SK_ID_CURR'] = test['SK_ID_CURR']
submission['TARGET'] = np.mean(y_pred_tot_lgm, 0)
submission.head()
submission.to_csv('submission.csv', index=False)
```
| github_jupyter |
# Guide FAQ
Hi guys, in this lecture I’m going to elaborate a little bit more on the nature of this course, an ‘FAQ’ of sorts.
## “Why doesn’t this guide cover sets, dicts, tuples?”
I’ve left out lots of things for a variety of reasons, the main three reasons being:
1. I do not have an infinite amount of time to spend on this project.
1. Syntax discussion is, though necessary, really boring to teach.
1. You must learn to think for yourselves!
Every time I increase the ‘scope’ of this project ‘quality’ is going to suffer; more lectures means more typos and bugs to catch with less time (per lecture) to catch them in!
And then there is the third (and most important) point; programming is about *self-learning* as opposed to being *spoon-fed* material. On numerous occasions throughout this guide I will encourage you to learn for yourselves, my job is to give you a set of tools to teach yourself with!
In short, this guide was never intended to be fully comprehensive and if you find yourself wanting to know how ‘X’ works (e.g. Sets, Tuples, Dicts) then the answer is merely a google away.
## “What is the 'Zen of Python' and why should I care?”
```
import this
```
The Zen of Python is a poem by Tim Peters, each line of which expresses something fundamental about the nature/design of Python. When I started to write this guide from very early on I knew I wanted to teach more than just syntax (anyone can do that), I wanted to write a guide that also instilled some of Python’s philosophy and ethos, without the discussion turning into abstract/pretentious drivel. Such discussion also had to be suitable for beginners as well.
In the end, I felt that the covering the ‘Zen of Python’ helps achieve these aims succinctly. By the end of this guide you should be able to understand most of what Tim Peter’s was waffling on about!
## “This guide is really good, can I use it?”
I’m pretty much okay with anyone taking this body of work and doing whatever they want with it *(that’s why I released it with an “MIT” license)*. So long as you give credit and don’t try to sell it for a profit it I don’t really care what you do.
## “This guide sucks, this is wrong, that is wrong …”
Then please give me your suggestions for improvement. :)
## "Why did you write this guide?"
Why? Why anything?
I guess there are two main reasons why I started this guide:
1. I enjoy teaching
1. I thought a big shiny project on Github might get me a job.
Personally I've been trying to get an entry level software job for about six months now, 40 applications a month and only two telephone interviews to show for it. I guess being a 29 year old dude with no relevant experience (or a computer science degree) probably means my CV doesn't really get past the *"HR filter"*.
When I started writing this guide I did so with the hope that maybe-- just maybe-- it would help get me a foot in the door, so to speak. Time will tell I guess.
### UPDATE JAN 2020
I wrote the paragraph above just about three years ago. I kindof abandoned this guide and have only recently come back to it.
Its perhaps worth pointing out that I did in fact end up landing a junior developer job at a small medical start-up writing mostly C# and Python. So now that I am older and wiser I've decided to update this guide to better reflect how I feel about code nowadays.
Anyway, I think I can say with hindsight that writing this guide probably wasn't the best use of my time; I had this idea that it would be a cool project to show potential employers what I could do, but I don't think they cared, or even looked at it. I probably would have been better off solving algorithms puzzles all day. Oh well, we live and we learn.
| github_jupyter |
```
import drama as drm
import numpy as np
import matplotlib.pylab as plt
from matplotlib import gridspec
import os
import glob
import h5py
import scipy.io as sio
%matplotlib inline
# import warnings
# warnings.filterwarnings('error')
fils = sorted(glob.glob('../data/*.mat'), key=os.path.getsize)[1:18]
n_files = len(fils)
file_names = [i.split('/')[-1][:-4] for i in fils]
print (file_names)
for i in range(len(fils)):
print (file_names[i])
try:
data = sio.loadmat(fils[i])
X = data['X'].astype(float)
y = data['y'].astype(float)
except:
data = h5py.File(fils[i])
X = np.array(data['X']).T.astype(float)
y = np.array(data['y']).T.astype(float)
iinds = np.argwhere(y[:,0]==0)[:,0]
oinds = np.argwhere(y[:,0]==1)[:,0]
nhalf = iinds.shape[0]//2
np.random.shuffle(iinds)
np.random.shuffle(oinds)
n_train = 3
X_train = np.concatenate([X[iinds[:nhalf]],X[oinds[:n_train]]],axis=0)
y_train = np.concatenate([y[iinds[:nhalf]],y[oinds[:n_train]]],axis=0)
X_test = np.concatenate([X[iinds[nhalf:]],X[oinds[n_train:]]],axis=0)
y_test = np.concatenate([y[iinds[nhalf:]],y[oinds[n_train:]]],axis=0)
X_train = X_train/X_train.max()
X_test = X_test/X_test.max()
res = drm.unsupervised_outlier_finder_all(X_train)
auc = []
mcc = []
rws = []
auc_b = -100
mcc_b = -100
rws_b = -100
for i in range(50):
for j in ['real','latent']:
o1 = res[j][i]
auc = drm.roc_auc_score(y_train==1, o1)
mcc = drm.MCC(y_train==1, o1)
rws = drm.rws_score(y_train==1, o1)
if auc_b<auc:
auc_b = auc
auc_set = [j,res['drt'][i],res['metric'][i]]
if mcc_b<mcc:
mcc_b = mcc
mcc_set = [j,res['drt'][i],res['metric'][i]]
if rws_b<rws:
rws_b = rws
rws_set = [j,res['drt'][i],res['metric'][i]]
res = drm.get_outliers(X_test,auc_set[1],auc_set[2],clustering=None,z_dim=2,space=auc_set[0])
o1 = res[auc_set[0]][auc_set[2]]
res = drm.get_outliers(X_test,mcc_set[1],mcc_set[2],clustering=None,z_dim=2,space=mcc_set[0])
o2 = res[mcc_set[0]][mcc_set[2]]
res = drm.get_outliers(X_test,rws_set[1],rws_set[2],clustering=None,z_dim=2,space=rws_set[0])
o3 = res[rws_set[0]][rws_set[2]]
acc = drm.roc_auc_score(y_test==1, o1)
mcc = drm.MCC(y_test==1, o1)
rws = drm.rws_score(y_test==1, o1)
print(acc,mcc,rws)
result = []
lof_all = np.zeros((n_files,3))
ifr_all = np.zeros((n_files,3))
for i in range(len(fils)):
print (file_names[i])
try:
data = sio.loadmat(fils[i])
X = data['X'].astype(float)
y = data['y'].astype(float)
except:
data = h5py.File(fils[i])
X = np.array(data['X']).T.astype(float)
y = np.array(data['y']).T.astype(float)
res = drm.unsupervised_outlier_finder_all(X)
arr,drts,metrs = drm.result_array(res,y,'real')
result.append(arr)
df = drm.sk_check(X,X,y,[1])
for j,scr in enumerate(['AUC','MCC','RWS']):
lof_all[i,j] = df[scr][0]
ifr_all[i,j] = df[scr][1]
result = np.array(result)
drm.plot_table(np.mean(result,axis=0),drts,metrs)
def result,lof_all
auc = np.sum((result[:, :, :, 0].T>lof_all[:, 0]) & (result[:, :, :, 0].T>ifr_all[:, 0]),axis=-1).T
mcc = np.sum((result[:, :, :, 1].T>lof_all[:, 1]) & (result[:, :, :, 1].T>ifr_all[:, 1]),axis=-1).T
rws = np.sum((result[:, :, :, 2].T>lof_all[:, 2]) & (result[:, :, :, 2].T>ifr_all[:, 2]),axis=-1).T
fig = plt.figure(figsize=(20,10))
plt.clf()
ax = fig.add_subplot(111)
ax.set_aspect('auto')
res = ax.imshow(auc, cmap=plt.cm.jet,interpolation='nearest')
width, height = auc.shape
for x in xrange(width):
for y in xrange(height):
ax.annotate('AUC: {:d}\n MCC: {:d}\n RWS: {:d}'.format(auc[x][y],mcc[x][y],rws[x][y]), xy=(y, x),
horizontalalignment='center',
verticalalignment='center',fontsize=18);
plt.xticks(range(10),['cityblock','L2','L4','braycurtis',
'canberra','chebyshev','correlation','mahalanobis','wL2','wL4'],fontsize=15)
plt.yticks(range(5), ['NMF','FastICA','PCA','AE','VAE'],fontsize=15)
plt.title('Number of successes (LOF and i-forest) out of 20 data set',fontsize=25)
plt.annotate('** Colors depend on AUC.', (0,0), (0, -30), xycoords='axes fraction',
textcoords='offset points', va='top',fontsize=15)
# plt.savefig('AND_success.jpg',dpi=150,bbox_inches='tight')
```
| github_jupyter |
# Mask R-CNN Demo
A quick intro to using the pre-trained model to detect and segment objects.
```
import os
import sys
import random
import math
import numpy as np
import skimage.io
import matplotlib
import matplotlib.pyplot as plt
# Root directory of the project
ROOT_DIR = os.path.abspath("../")
# Import Mask RCNN
sys.path.append(ROOT_DIR) # To find local version of the library
from mrcnn import utils
import mrcnn.model as modellib
from mrcnn import visualize
# Import stone config
sys.path.append(os.path.join(ROOT_DIR, "samples/stone/")) # To find local version
import stone
%matplotlib inline
# Directory to save logs and trained model
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
# Local path to trained weights file
STONE_MODEL_PATH = os.path.join(MODEL_DIR, "mask_rcnn_stone_0001.h5")
# Download stone trained weights from Releases if needed
if not os.path.exists(STONE_MODEL_PATH):
utils.download_trained_weights(STONE_MODEL_PATH)
# When running the Demo on the images in which the model is trained, use the second directory.
# When running the Demo on Random images, uncomment the first direcotry and comment the second directory.
# Directory to run the dataset images
IMAGE_DIR = os.path.join(ROOT_DIR, "images/Random_test_Images")
# Directory of images to run detection on
#IMAGE_DIR = os.path.join(ROOT_DIR, "images/Dataset_images")
```
## Configurations
We'll be using a model trained on the MS-COCO dataset. The configurations of this model are in the ```CocoConfig``` class in ```coco.py```.
For inferencing, modify the configurations a bit to fit the task. To do so, sub-class the ```CocoConfig``` class and override the attributes you need to change.
```
class InferenceConfig(stone.StoneConfig):
# Set batch size to 1 since we'll be running inference on
# one image at a time. Batch size = GPU_COUNT * IMAGES_PER_GPU
GPU_COUNT = 1
IMAGES_PER_GPU = 1
config = InferenceConfig()
config.display()
```
## Create Model and Load Trained Weights
```
# Create model object in inference mode.
model = modellib.MaskRCNN(mode="inference", model_dir=MODEL_DIR, config=config)
# Load weights trained on MS-COCO
model.load_weights(STONE_MODEL_PATH, by_name=True)
```
## Class Names
The model classifies objects and returns class IDs, which are integer value that identify each class. Some datasets assign integer values to their classes and some don't. For example, in the MS-COCO dataset, the 'person' class is 1 and 'teddy bear' is 88. The IDs are often sequential, but not always. The COCO dataset, for example, has classes associated with class IDs 70 and 72, but not 71.
To improve consistency, and to support training on data from multiple sources at the same time, our ```Dataset``` class assigns it's own sequential integer IDs to each class. For example, if you load the COCO dataset using our ```Dataset``` class, the 'person' class would get class ID = 1 (just like COCO) and the 'teddy bear' class is 78 (different from COCO). Keep that in mind when mapping class IDs to class names.
To get the list of class names, you'd load the dataset and then use the ```class_names``` property like this.
```
# Load COCO dataset
dataset = coco.CocoDataset()
dataset.load_coco(COCO_DIR, "train")
dataset.prepare()
# Print class names
print(dataset.class_names)
```
We don't want to require you to download the COCO dataset just to run this demo, so we're including the list of class names below. The index of the class name in the list represent its ID (first class is 0, second is 1, third is 2, ...etc.)
```
# COCO Class names
# Index of the class in the list is its ID. For example, to get ID of
# the teddy bear class, use: class_names.index('teddy bear')
class_names = ['BG', 'stone']
```
## Run Object Detection
```
# Load a random image from the images folder
file_names = next(os.walk(IMAGE_DIR))[2]
image = skimage.io.imread(os.path.join(IMAGE_DIR, random.choice(file_names)))
# Run detection
results = model.detect([image], verbose=1)
# Visualize results
r = results[0]
visualize.display_instances(image, r['rois'], r['masks'], r['class_ids'],
class_names, r['scores'])
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#@title MIT License
#
# Copyright (c) 2017 François Chollet
#
# Permission is hereby granted, free of charge, to any person obtaining a
# copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
```
# Regression: Predict fuel efficiency
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/beta/tutorials/keras/basic_regression"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r2/tutorials/keras/basic_regression.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r2/tutorials/keras/basic_regression.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/site/en/r2/tutorials/keras/basic_regression.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
In a *regression* problem, we aim to predict the output of a continuous value, like a price or a probability. Contrast this with a *classification* problem, where we aim to select a class from a list of classes (for example, where a picture contains an apple or an orange, recognizing which fruit is in the picture).
This notebook uses the classic [Auto MPG](https://archive.ics.uci.edu/ml/datasets/auto+mpg) Dataset and builds a model to predict the fuel efficiency of late-1970s and early 1980s automobiles. To do this, we'll provide the model with a description of many automobiles from that time period. This description includes attributes like: cylinders, displacement, horsepower, and weight.
This example uses the `tf.keras` API, see [this guide](https://www.tensorflow.org/guide/keras) for details.
```
# Use seaborn for pairplot
!pip install seaborn
from __future__ import absolute_import, division, print_function, unicode_literals
import pathlib
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
!pip install tensorflow==2.0.0-beta0
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
print(tf.__version__)
```
## The Auto MPG dataset
The dataset is available from the [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/).
### Get the data
First download the dataset.
```
dataset_path = keras.utils.get_file("auto-mpg.data", "http://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/auto-mpg.data")
dataset_path
```
Import it using pandas
```
column_names = ['MPG','Cylinders','Displacement','Horsepower','Weight',
'Acceleration', 'Model Year', 'Origin']
raw_dataset = pd.read_csv(dataset_path, names=column_names,
na_values = "?", comment='\t',
sep=" ", skipinitialspace=True)
dataset = raw_dataset.copy()
dataset.tail()
```
### Clean the data
The dataset contains a few unknown values.
```
dataset.isna().sum()
```
To keep this initial tutorial simple drop those rows.
```
dataset = dataset.dropna()
```
The `"Origin"` column is really categorical, not numeric. So convert that to a one-hot:
```
origin = dataset.pop('Origin')
dataset['USA'] = (origin == 1)*1.0
dataset['Europe'] = (origin == 2)*1.0
dataset['Japan'] = (origin == 3)*1.0
dataset.tail()
```
### Split the data into train and test
Now split the dataset into a training set and a test set.
We will use the test set in the final evaluation of our model.
```
train_dataset = dataset.sample(frac=0.8,random_state=0)
test_dataset = dataset.drop(train_dataset.index)
```
### Inspect the data
Have a quick look at the joint distribution of a few pairs of columns from the training set.
```
sns.pairplot(train_dataset[["MPG", "Cylinders", "Displacement", "Weight"]], diag_kind="kde")
```
Also look at the overall statistics:
```
train_stats = train_dataset.describe()
train_stats.pop("MPG")
train_stats = train_stats.transpose()
train_stats
```
### Split features from labels
Separate the target value, or "label", from the features. This label is the value that you will train the model to predict.
```
train_labels = train_dataset.pop('MPG')
test_labels = test_dataset.pop('MPG')
```
### Normalize the data
Look again at the `train_stats` block above and note how different the ranges of each feature are.
It is good practice to normalize features that use different scales and ranges. Although the model *might* converge without feature normalization, it makes training more difficult, and it makes the resulting model dependent on the choice of units used in the input.
Note: Although we intentionally generate these statistics from only the training dataset, these statistics will also be used to normalize the test dataset. We need to do that to project the test dataset into the same distribution that the model has been trained on.
```
def norm(x):
return (x - train_stats['mean']) / train_stats['std']
normed_train_data = norm(train_dataset)
normed_test_data = norm(test_dataset)
```
This normalized data is what we will use to train the model.
Caution: The statistics used to normalize the inputs here (mean and standard deviation) need to be applied to any other data that is fed to the model, along with the one-hot encoding that we did earlier. That includes the test set as well as live data when the model is used in production.
## The model
### Build the model
Let's build our model. Here, we'll use a `Sequential` model with two densely connected hidden layers, and an output layer that returns a single, continuous value. The model building steps are wrapped in a function, `build_model`, since we'll create a second model, later on.
```
def build_model():
model = keras.Sequential([
layers.Dense(64, activation='relu', input_shape=[len(train_dataset.keys())]),
layers.Dense(64, activation='relu'),
layers.Dense(1)
])
optimizer = tf.keras.optimizers.RMSprop(0.001)
model.compile(loss='mse',
optimizer=optimizer,
metrics=['mae', 'mse'])
return model
model = build_model()
```
### Inspect the model
Use the `.summary` method to print a simple description of the model
```
model.summary()
```
Now try out the model. Take a batch of `10` examples from the training data and call `model.predict` on it.
```
example_batch = normed_train_data[:10]
example_result = model.predict(example_batch)
example_result
```
It seems to be working, and it produces a result of the expected shape and type.
### Train the model
Train the model for 1000 epochs, and record the training and validation accuracy in the `history` object.
```
# Display training progress by printing a single dot for each completed epoch
class PrintDot(keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs):
if epoch % 100 == 0: print('')
print('.', end='')
EPOCHS = 1000
history = model.fit(
normed_train_data, train_labels,
epochs=EPOCHS, validation_split = 0.2, verbose=0,
callbacks=[PrintDot()])
```
Visualize the model's training progress using the stats stored in the `history` object.
```
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
hist.tail()
def plot_history(history):
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
plt.figure()
plt.xlabel('Epoch')
plt.ylabel('Mean Abs Error [MPG]')
plt.plot(hist['epoch'], hist['mae'],
label='Train Error')
plt.plot(hist['epoch'], hist['val_mae'],
label = 'Val Error')
plt.ylim([0,5])
plt.legend()
plt.figure()
plt.xlabel('Epoch')
plt.ylabel('Mean Square Error [$MPG^2$]')
plt.plot(hist['epoch'], hist['mse'],
label='Train Error')
plt.plot(hist['epoch'], hist['val_mse'],
label = 'Val Error')
plt.ylim([0,20])
plt.legend()
plt.show()
plot_history(history)
```
This graph shows little improvement, or even degradation in the validation error after about 100 epochs. Let's update the `model.fit` call to automatically stop training when the validation score doesn't improve. We'll use an *EarlyStopping callback* that tests a training condition for every epoch. If a set amount of epochs elapses without showing improvement, then automatically stop the training.
You can learn more about this callback [here](https://www.tensorflow.org/versions/master/api_docs/python/tf/keras/callbacks/EarlyStopping).
```
model = build_model()
# The patience parameter is the amount of epochs to check for improvement
early_stop = keras.callbacks.EarlyStopping(monitor='val_loss', patience=10)
history = model.fit(normed_train_data, train_labels, epochs=EPOCHS,
validation_split = 0.2, verbose=0, callbacks=[early_stop, PrintDot()])
plot_history(history)
```
The graph shows that on the validation set, the average error is usually around +/- 2 MPG. Is this good? We'll leave that decision up to you.
Let's see how well the model generalizes by using the **test** set, which we did not use when training the model. This tells us how well we can expect the model to predict when we use it in the real world.
```
loss, mae, mse = model.evaluate(normed_test_data, test_labels, verbose=0)
print("Testing set Mean Abs Error: {:5.2f} MPG".format(mae))
```
### Make predictions
Finally, predict MPG values using data in the testing set:
```
test_predictions = model.predict(normed_test_data).flatten()
plt.scatter(test_labels, test_predictions)
plt.xlabel('True Values [MPG]')
plt.ylabel('Predictions [MPG]')
plt.axis('equal')
plt.axis('square')
plt.xlim([0,plt.xlim()[1]])
plt.ylim([0,plt.ylim()[1]])
_ = plt.plot([-100, 100], [-100, 100])
```
It looks like our model predicts reasonably well. Let's take a look at the error distribution.
```
error = test_predictions - test_labels
plt.hist(error, bins = 25)
plt.xlabel("Prediction Error [MPG]")
_ = plt.ylabel("Count")
```
It's not quite gaussian, but we might expect that because the number of samples is very small.
## Conclusion
This notebook introduced a few techniques to handle a regression problem.
* Mean Squared Error (MSE) is a common loss function used for regression problems (different loss functions are used for classification problems).
* Similarly, evaluation metrics used for regression differ from classification. A common regression metric is Mean Absolute Error (MAE).
* When numeric input data features have values with different ranges, each feature should be scaled independently to the same range.
* If there is not much training data, one technique is to prefer a small network with few hidden layers to avoid overfitting.
* Early stopping is a useful technique to prevent overfitting.
| github_jupyter |
Solution by Eliott Rosenberg (enr27@cornell.edu)
```
import numpy as np
# Importing standard Qiskit libraries
from qiskit import QuantumCircuit, transpile, Aer, IBMQ
from qiskit.tools.jupyter import *
from qiskit.visualization import *
from ibm_quantum_widgets import *
# Loading your IBM Quantum account(s)
provider = IBMQ.load_account()
```
First, we use the provided code to create the qmolecule object for LiH.
```
import numpy as np
from qiskit_nature.drivers import PySCFDriver
molecule = 'Li 0.0 0.0 0.0; H 0.0 0.0 1.5474'
driver = PySCFDriver(atom=molecule)
qmolecule = driver.run()
```
Next, we need to generate a qubit Hamiltonian corresponding to this problem. We want to simplify this Hamiltonian as much as possible. That is, we want to minimize the number of qubits needed to represent the Hamiltonian since this will mean that we need fewer CNOT gates to entangle the qubits.
To do this, we will use the ParityMapper with `two_qubit_reduction=True`. We will also freeze the core (non-valence) electrons and remove unoccupied orbitals. We further identify the $Z_2$ symmetries and keep only the sector that contains the ground state.
Relevant documentation pages:
https://qiskit.org/documentation/nature/stubs/qiskit_nature.converters.second_quantization.QubitConverter.html
https://qiskit.org/documentation/nature/stubs/qiskit_nature.transformers.FreezeCoreTransformer.html
```
from qiskit_nature.problems.second_quantization.electronic import ElectronicStructureProblem
from qiskit_nature.transformers import FreezeCoreTransformer
from qiskit_nature.mappers.second_quantization import ParityMapper, BravyiKitaevMapper, JordanWignerMapper
from qiskit_nature.converters.second_quantization.qubit_converter import QubitConverter
# unoccupied orbitals that will be removed.
#This is following https://qiskit.org/textbook/ch-applications/vqe-molecules.html#Running-VQE-on-a-Statevector-Simulator,
#in which these two orbitals are removed.
# You can confirm that removing these orbitals only has a small effect on the ground state energy by
# setting remove_list to [] and seeing how this affects the exact ground state energy.
remove_list = [3,4]
# freeze_core = True means that we are not treating the core electron as part of our quantum system
transformer = [FreezeCoreTransformer(freeze_core=True,remove_orbitals=remove_list)]
problem = ElectronicStructureProblem(driver,q_molecule_transformers=transformer)
# Generate the second-quantized operators
second_q_ops = problem.second_q_ops()
# Hamiltonian
main_op = second_q_ops[0]
mapper = ParityMapper()
# The Hamiltonian has additional Z2 symmetries. We can reduce the problem size by working in a particular
# eigenspace of these symmetry operators. We just have to make sure that the eigenspace we pick
# contains the ground state. You can confirm that it contains the ground state by getting rid of
# z2symmetry_reduction=[1,1] and seeing that this doesn't affect the ground state energy.
converter = QubitConverter(mapper=mapper, two_qubit_reduction=True, z2symmetry_reduction=[1,1])
num_particles = (problem.molecule_data_transformed.num_alpha,
problem.molecule_data_transformed.num_beta)
qubit_op = converter.convert(main_op, num_particles=num_particles)
```
We can see what the qubit Hamiltonian looks like now. With all of these simplifications, we have reduced it to just 4 qubits.
```
print(qubit_op)
```
We can also compute the exact ground state energy so that we know what we're targeting and to confirm that our various simplifications haven't changed the ground state energy too much.
```
# exact ground state energy:
from qiskit_nature.algorithms.ground_state_solvers.minimum_eigensolver_factories import NumPyMinimumEigensolverFactory
from qiskit_nature.algorithms.ground_state_solvers import GroundStateEigensolver
import numpy as np
def exact_diagonalizer(problem, converter):
solver = NumPyMinimumEigensolverFactory()
calc = GroundStateEigensolver(converter, solver)
result = calc.solve(problem)
return result
result_exact = exact_diagonalizer(problem, converter)
exact_energy = np.real(result_exact.eigenenergies[0])
print("Exact electronic energy", exact_energy)
print(result_exact)
```
We see that, with the frozen core and the removed orbitals, the ground state energy is -1.08870601573474. For comparison, if we had not removed the orbitals, the ground state energy would have been -1.08978239634873, so removing the orbitals leads to an error of 0.001, within the chemical accuracy. Adding on the extracted energy from the core, these give -8.907396311 and -8.908472692, respectively, whereas the exact ground state energy, treating the core quantum mechanically, is -8.90869711642421. We see that freezing the core is a very good approximation, causing only a 0.0002 discrepancy, an order of magnitude smaller than the chemical accuracy.
Next, we are instructed to begin in the Hartee-Fock initial state:
```
# initial state:
from qiskit_nature.circuit.library import HartreeFock
num_particles = (problem.molecule_data_transformed.num_alpha,
problem.molecule_data_transformed.num_beta)
num_spin_orbitals = 2 * problem.molecule_data_transformed.num_molecular_orbitals
init_state = HartreeFock(num_spin_orbitals, num_particles, converter)
print(init_state)
```
Next, we construct an ansatz. It uses 3 CNOTs to entangle the 4 qubits and has lots of tunable parameters. The UGate is a general 1-qubit unitary. If you change optimize_externally to True, then we import the optimial parameters, which I found using an analytic gradient optimizer that I wrote independently of this challenge. (In this case, we tack on an Rz gate so that there is one parameter for the qiskit optimizer to optimize.)
```
num_qubits = init_state.num_qubits
from qiskit.circuit import Parameter, QuantumCircuit
from qiskit.circuit.library import TwoLocal
ansatz = QuantumCircuit(num_qubits)
optimize_externally = True
if optimize_externally:
theta = np.genfromtxt('theta1.csv')
whichParameter = 0
for q in range(num_qubits):
ansatz.u(theta[whichParameter],theta[whichParameter+1],theta[whichParameter+2],q)
whichParameter += 3
for q in range(num_qubits-1):
ansatz.cx(q,q+1)
ansatz.u(theta[whichParameter],theta[whichParameter+1],theta[whichParameter+2],q)
whichParameter += 3
ansatz.u(theta[whichParameter],theta[whichParameter+1],theta[whichParameter+2],q+1)
whichParameter += 3
ansatz.rz(Parameter('th'),0)
else:
whichParameter = 0
for q in range(num_qubits):
ansatz.u(Parameter('th'+str(whichParameter)),Parameter('th'+str(whichParameter+1)),Parameter('th'+str(whichParameter+2)),q)
whichParameter += 3
for q in range(num_qubits-1):
ansatz.cx(q,q+1)
ansatz.u(Parameter('th'+str(whichParameter)),Parameter('th'+str(whichParameter+1)),Parameter('th'+str(whichParameter+2)),q)
whichParameter += 3
ansatz.u(Parameter('th'+str(whichParameter)),Parameter('th'+str(whichParameter+1)),Parameter('th'+str(whichParameter+2)),q+1)
whichParameter += 3
ansatz.compose(init_state, front=True, inplace=True)
ansatz.draw()
```
Finally, we optimize the ansatz:
```
# backend
from qiskit import Aer
backend = Aer.get_backend('statevector_simulator')
# optimizer
from qiskit.algorithms.optimizers import COBYLA, L_BFGS_B, SPSA, SLSQP
optimizer = COBYLA(maxiter=5000)
# optimize
from qiskit.algorithms import VQE
from IPython.display import display, clear_output
# Print and save the data in lists
def callback(eval_count, parameters, mean, std):
# Overwrites the same line when printing
display("Evaluation: {}, Energy: {}, Std: {}".format(eval_count, mean, std))
clear_output(wait=True)
counts.append(eval_count)
values.append(mean)
params.append(parameters)
deviation.append(std)
counts = []
values = []
params = []
deviation = []
# Set initial parameters of the ansatz
# We choose a fixed small displacement
# So all participants start from similar starting point
try:
initial_point = [0.01] * len(ansatz.ordered_parameters)
except:
initial_point = [0.01] * ansatz.num_parameters
# use my initial point instead:
#initial_point = theta
algorithm = VQE(ansatz,
optimizer=optimizer,
quantum_instance=backend,
callback=callback,
initial_point=initial_point)
result = algorithm.compute_minimum_eigenvalue(qubit_op)
print(result)
```
We see that this converges to within chemical accuracy, so we're done and can submit.
```
# Check your answer using following code
from qc_grader import grade_ex5
freeze_core = True # change to True if you freezed core electrons
grade_ex5(ansatz,qubit_op,result,freeze_core)
```
| github_jupyter |
```
%matplotlib inline
import sys
import os
import glob
import csv
import librosa
import librosa.display
import pretty_midi
import numpy as np
from scipy.spatial.distance import cdist
import matplotlib.pyplot as plt
import pandas as pd
from multiprocessing import Pool
from tqdm import tqdm, trange
import pickle
```
## 1. Audio Synchronization Baseline
### 1.1 Get chroma features from midi
```
synth_midi_path = 'synth_midi'
midi_path = 'midi'
piece = 'debussy_childrencorner6'
midi_file1 = synth_midi_path + '/sharpeye/' + piece + '_v1.mid'
midi_file2 = midi_path + '/' + piece + '.mid'
sr = 22050
hop_size = 0.025
window_len = 0.025
mid1 = pretty_midi.PrettyMIDI(midi_file1)
mid2 = pretty_midi.PrettyMIDI(midi_file2)
audio1 = mid1.synthesize()
audio2 = mid2.synthesize()
chroma1 = librosa.feature.mfcc(audio1, sr, hop_length=int(hop_size*sr), n_fft=int(window_len*sr))
chroma2 = librosa.feature.mfcc(audio2, sr, hop_length=int(hop_size*sr), n_fft=int(window_len*sr))
```
### 1.2 DTW on chroma feature
```
def alignAudio(M1, M2):
# Get cost metric
C = cdist(M1, M2, 'seuclidean', V=None)
# DTW
steps = np.array([1,1,1,2,2,1]).reshape((3,2))
weights = np.array([2,3,3])
D, wp = librosa.core.dtw(C=C, step_sizes_sigma=steps, weights_mul=weights)
return wp[::-1,:].transpose()
wp = alignAudio(np.transpose(chroma1), np.transpose(chroma2))
```
### 1.3 Calculate Error
```
def getMidiRefLocs(annot_file):
timeStamps = []
with open(annot_file, newline='') as csvfile:
spamreader = csv.reader(csvfile, delimiter=',', quotechar='|')
for row in spamreader:
if row[0] != '-':
timeStamps.append(float(row[0]))
else:
timeStamps.append(float('inf'))
timeStamps = np.array(timeStamps)
return timeStamps
def getSheetRefLocs(scoreid, changeDPI = False):
hyp_file = 'hyp_align/'+scoreid+'.pkl'
dhyp = pickle.load(open(hyp_file, 'rb'))
striplens = dhyp['striplens']
# get annotation file
annot_dir = 'annot_data'
piece = scoreid.split('_')
annot_file_beats = '%s/%s_%s_beats.csv' % (annot_dir, piece[0], piece[1])
df_all = pd.read_csv(annot_file_beats)
# calculate global pixel position
scoreid = piece[1]+'_'+piece[2]
df = df_all.loc[df_all.score == scoreid]
pixelOffset = np.cumsum([0] + striplens) # cumulative pixel offset for each strip
stripsPerPage = [df.loc[df.page == i,'strip'].max() for i in range(df.page.max()+1) ]
stripOffset = np.cumsum([0] + stripsPerPage)
stripIdx = stripOffset[df.page] + df.strip - 1 # cumulative strip index
if changeDPI:
hpixlocs = pixelOffset[stripIdx] + (df.hpixel * 400 // (72*4))
else:
hpixlocs = pixelOffset[stripIdx] + df.hpixel
return hpixlocs.values
synth_timestamps = mid1.get_beats()
perf_timestamps1 = mid2.get_beats()
def calcPredErrors(wp, perf_timestamps, synth_timestamps):
all_errs = []
for i, beat_time in enumerate(perf_timestamps):
frame_id2 = beat_time // hop_size
wp_id2 = np.argmin([abs(x-frame_id2) for x in wp[1]])
frame_id1 = wp[0][wp_id2]
all_errs.append((synth_timestamps[i] - (hop_size*frame_id1)) * 1000) # in ms
return all_errs
def calcErrorStats(errs_raw, tols, isSingle = False):
if isSingle:
errs = errs_raw
else:
errs = np.array([err for sublist in errs_raw for err in sublist])
errs = errs[~np.isnan(errs)] # when beat is not annotated, value is nan
errorRates = []
for tol in tols:
toAdd = np.sum(np.abs(errs) > tol) * 1.0 / len(errs)
errorRates.append(toAdd)
return errorRates
errs1 = calcPredErrors(wp, perf_timestamps1, synth_timestamps)
tols = np.arange(5000)
errorRates1 = calcErrorStats(errs1, tols, True)
plt.plot(tols, 100.0*np.array(errorRates1), 'k-', label='auto-annot')
plt.xlabel('Error Tolerance (milliseconds)')
plt.ylabel('Error Rate (%)')
plt.gca().set_ylim([0,100])
plt.legend()
plt.show()
```
### 1.4 Run experiment on the whole dataset
```
synth_midi_path = 'synth_midi/'
midi_path = 'midi/'
annot_dir = 'annot_data/'
pieces = ['brahms_op116no6', 'brahms_op117no2',
'chopin_op30no2', 'chopin_op63no3', 'chopin_op68no3',
'clementi_op36no1mv3', 'clementi_op36no2mv3', 'clementi_op36no3mv3',
'debussy_childrencorner1', 'debussy_childrencorner3', 'debussy_childrencorner6',
'mendelssohn_op19no2', 'mendelssohn_op62no3', 'mendelssohn_op62no5',
'mozart_kv311mv3', 'mozart_kv333mv3',
'schubert_op90no1', 'schubert_op90no3', 'schubert_op94no2',
'tchaikovsky_season01', 'tchaikovsky_season06', 'tchaikovsky_season08']
def calcSingleError(mid_pair, perf_timestamps, synth_timestamps):
mid1 = mid_pair[0]
mid2 = mid_pair[1]
audio1 = mid1.synthesize()
audio2 = mid2.synthesize()
chroma1 = librosa.feature.mfcc(audio1, sr, hop_length=int(hop_size*sr), n_fft=int(window_len*sr))
chroma2 = librosa.feature.mfcc(audio2, sr, hop_length=int(hop_size*sr), n_fft=int(window_len*sr))
wp = alignAudio(np.transpose(chroma1), np.transpose(chroma2))
if len(synth_timestamps) != len(perf_timestamps):
minLen = min(len(synth_timestamps), len(perf_timestamps))
synth_timestamps = synth_timestamps[:minLen]
perf_timestamps = perf_timestamps[:minLen]
errs = calcPredErrors(wp, perf_timestamps, synth_timestamps)
return errs, wp
def runExperiment(program, pieces_list):
allErrs_time = []
allErrs_pixel = []
for piece in pieces_list:
all_sheets = sorted(glob.glob('score_data/prepped_pdf/%s*' % piece))
real_midis = sorted(glob.glob(midi_path+'%s*' % piece))
perf_timestamps = getMidiRefLocs(annot_dir + 'midi/' + piece + '.csv')
if program == 'sharpeye':
synth_annot_files = sorted(glob.glob(annot_dir+'synth_midi/'+'%s*_se.csv' % piece.split('_')[1]))
elif program == 'photoscore':
synth_annot_files = sorted(glob.glob(annot_dir+'synth_midi/'+'%s*_ps.csv' % piece.split('_')[1]))
for i in range(len(real_midis)):
mid2 = pretty_midi.PrettyMIDI(real_midis[i])
for j in range(len(all_sheets)):
scoreid = all_sheets[j].split('/')[-1].split('.')[0]
sheet_annot = getSheetRefLocs(scoreid)
synth_file = synth_midi_path+program+'/'+scoreid+'.mid'
synth_name = synth_file.split('/')[-1].split('.')[0]
synth_name = synth_name.split('_')[1] + '_' + synth_name.split('_')[2]
if program == 'sharpeye':
synth_annot_file = annot_dir + 'synth_midi/' + synth_name + '_se.csv'
elif program == 'photoscore':
synth_annot_file = annot_dir + 'synth_midi/' + synth_name + '_ps.csv'
if synth_annot_file in synth_annot_files and (program != 'photoscore' or scoreid != 'chopin_op68no3_v6'):
mid1 = pretty_midi.PrettyMIDI(synth_file)
print(real_midis[i], synth_file)
synth_timestamps = getMidiRefLocs(synth_annot_file)
err_t, wp = calcSingleError([mid1, mid2], perf_timestamps, synth_timestamps)
allErrs_time.append(err_t)
hypPixels = np.interp(perf_timestamps, wp[:,1], wp[:,0])
minLen_p = min(len(hypPixels), len(sheet_annot))
allErrs_pixel.append(hypPixels[:minLen_p] - sheet_annot[:minLen_p])
else:
allErrs_pixel.append([float('inf')]*len(sheet_annot))
allErrs_time.append([float('inf')]*len(perf_timestamps))
return allErrs_pixel, allErrs_time
def alignAll(program, pieces_list):
allwp = {}
for piece in pieces_list:
synth_midis = [os.path.basename(elem) for elem in sorted(glob.glob(synth_midi_path+program+'/%s*' % piece))]
real_midis = [os.path.basename(elem) for elem in sorted(glob.glob(midi_path+'/*%s*' % piece))]
for i in trange(len(real_midis)):
real_midi_name = real_midis[i]
#real_full_path = midi_path + '/' + piece + '/' + real_midi_name
real_full_path = midi_path + '/' + real_midi_name
mid2 = pretty_midi.PrettyMIDI(real_full_path)
audio2 = mid2.synthesize()
chroma2 = librosa.feature.mfcc(audio2, sr, hop_length=int(hop_size*sr), n_fft=int(window_len*sr))
for j in range(len(synth_midis)):
synth_midi_name = synth_midis[j]
synth_full_path = synth_midi_path+'/'+program+'/'+synth_midi_name
mid1 = pretty_midi.PrettyMIDI(synth_full_path)
audio1 = mid1.synthesize()
chroma1 = librosa.feature.mfcc(audio1, sr, hop_length=int(hop_size*sr), n_fft=int(window_len*sr))
wp = alignAudio(np.transpose(chroma1), np.transpose(chroma2))
allwp[(real_full_path, synth_full_path)] = wp
with open('results/audioalign_nonmzk_'+program+'.pkl','wb') as f:
pickle.dump(allwp, f)
alignAll('photoscore', pieces)
alignAll('sharpeye', pieces)
allErrs_se_as_pix, allErrs_se_as_t = runExperiment('sharpeye', pieces)
allErrs_ps_as_pix, allErrs_ps_as_t = runExperiment('photoscore', pieces)
with open('results/errorData_real_as.pkl','wb') as f:
pickle.dump([allErrs_ps_as_pix, allErrs_ps_as_t, allErrs_se_as_pix, allErrs_se_as_t],f)
```
## 2. Midi-Beat-Matching
```
def midiBeatMatch(program, pieces_list):
allErrs_pixel = []
allErrs_time = []
for piece in pieces_list:
perf_timestamps = getMidiRefLocs(annot_dir + 'midi/' + piece + '.csv')
all_sheets = sorted(glob.glob('score_data/prepped_pdf/%s*' % piece))
if program == 'sharpeye':
synth_annot_files = sorted(glob.glob(annot_dir+'synth_midi/'+'%s*_se.csv' % piece.split('_')[1]))
elif program == 'photoscore':
synth_annot_files = sorted(glob.glob(annot_dir+'synth_midi/'+'%s*_ps.csv' % piece.split('_')[1]))
print(synth_annot_files)
for j in range(len(all_sheets)):
scoreid = all_sheets[j].split('/')[-1].split('.')[0]
sheet_annot = getSheetRefLocs(scoreid)
synth_file = synth_midi_path+program+'/'+scoreid+'.mid'
synth_name = synth_file.split('/')[-1].split('.')[0]
synth_name = synth_name.split('_')[1] + '_' + synth_name.split('_')[2]
if program == 'sharpeye':
synth_annot_file = annot_dir + 'synth_midi/' + synth_name + '_se.csv'
elif program == 'photoscore':
synth_annot_file = annot_dir + 'synth_midi/' + synth_name + '_ps.csv'
print(synth_annot_file)
if synth_annot_file in synth_annot_files and (program != 'photoscore' or scoreid != 'chopin_op68no3_v6'):
mid1 = pretty_midi.PrettyMIDI(synth_file)
start_time = mid1.estimate_beat_start(candidates=10, tolerance=0.025)
auto_beat = mid1.get_beats()
synth_timestamps = getMidiRefLocs(synth_annot_file)
print(auto_beat[0:10])
print(synth_timestamps[0:10])
minLen_t = min(len(synth_timestamps), len(auto_beat))
allErrs_time.append((np.array(synth_timestamps[:minLen_t]) - np.array(auto_beat[:minLen_t])) * 1000)
minLen_p = min(minLen_t, len(sheet_annot))
hypPixels = np.interp(auto_beat, synth_timestamps[:minLen_p], sheet_annot[:minLen_p])
allErrs_pixel.append(hypPixels[:minLen_p] - sheet_annot[:minLen_p])
else:
allErrs_pixel.append([float('inf')*len(sheet_annot)])
allErrs_time.append([float('inf')]*len(perf_timestamps))
return allErrs_pixel, allErrs_time
allErrs_se_bm_pix, allErrs_se_bm_t = midiBeatMatch('sharpeye', pieces)
allErrs_ps_bm_pix, allErrs_ps_bm_t = midiBeatMatch('photoscore', pieces)
with open('results/errorData_real_bm.pkl','wb') as f:
pickle.dump([allErrs_ps_bm_pix, allErrs_ps_bm_t, allErrs_se_bm_pix, allErrs_se_bm_t],f)
```
## 3. Compare Error to Bootleg System
```
[allErrs_ps_bm_pix, allErrs_ps_bm_t, allErrs_se_bm_pix, allErrs_se_bm_t] = pickle.load(open('results/errorData_real_bm.pkl', 'rb'))
[pixel_errs_bs, pixel_errs_b1, time_errs_bs, time_errs_b1] = pickle.load(open('results/errorData_real_bootleg.pkl', 'rb'))
[allErrs_ps_as_pix, allErrs_ps_as_t, allErrs_se_as_pix, allErrs_se_as_t] = pickle.load(open('results/errorData_real_as.pkl','rb'))
tols = np.arange(2001)
plt.plot(tols, 100.0*np.array(calcErrorStats(time_errs_b1, tols)), 'k-', label='GL')
plt.plot(tols, 100.0*np.array(calcErrorStats(allErrs_se_bm_t, tols)), 'g-.', label='MBM-se')
plt.plot(tols, 100.0*np.array(calcErrorStats(allErrs_ps_bm_t, tols)), 'r-.', label='MBM-ps')
plt.plot(tols, 100.0*np.array(calcErrorStats(allErrs_se_as_t, tols)), 'g--', label='AS-se')
plt.plot(tols, 100.0*np.array(calcErrorStats(allErrs_ps_as_t, tols)), 'r--', label='AS-ps')
plt.plot(tols, 100.0*np.array(calcErrorStats(time_errs_bs, tols)), 'g-', label='BS')
plt.xlabel('Error Tolerance (milliseconds)')
plt.ylabel('Error Rate (%)')
plt.gca().set_ylim([0,100])
plt.legend()
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.savefig('figs/error_curves(final).png', dpi=300, bbox_inches = 'tight')
plt.show()
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Probability Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
#@title Licensed under the Apache License, Version 2.0 (the "License"); { display-mode: "form" }
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# TFP 確率的レイヤー: 回帰
<table class="tfo-notebook-buttons" align="left">
<td><a target="_blank" href="https://www.tensorflow.org/probability/examples/Probabilistic_Layers_Regression"><img src="https://www.tensorflow.org/images/tf_logo_32px.png">TensorFlow.org で表示</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ja/probability/examples/Probabilistic_Layers_Regression.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png">Google Colab で実行</a></td>
<td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ja/probability/examples/Probabilistic_Layers_Regression.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">GitHub でソースを表示</a></td>
<td><a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ja/probability/examples/Probabilistic_Layers_Regression.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png">ノートブックをダウンロード</a></td>
</table>
この例では、TFP の「確率的レイヤー」を使用して回帰モデルを適合させる方法を示します。
### 依存関係と前提条件
```
#@title Import { display-mode: "form" }
from pprint import pprint
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import tensorflow.compat.v2 as tf
tf.enable_v2_behavior()
import tensorflow_probability as tfp
sns.reset_defaults()
#sns.set_style('whitegrid')
#sns.set_context('talk')
sns.set_context(context='talk',font_scale=0.7)
%matplotlib inline
tfd = tfp.distributions
```
### 迅速に作成
はじめる前に、このデモで GPU を使用していることを確認します。
[ランタイム] -> [ランタイムタイプの変更] -> [ハードウェアアクセラレータ] -> [GPU] を選択します。
次のスニペットは、GPU にアクセスできることを確認します。
```
if tf.test.gpu_device_name() != '/device:GPU:0':
print('WARNING: GPU device not found.')
else:
print('SUCCESS: Found GPU: {}'.format(tf.test.gpu_device_name()))
```
注意: 何らかの理由で GPU にアクセスできない場合でも、このコラボは機能します (トレーニングには時間がかかります)。
## 目的
TFP を使用して確率モデルを指定し、負の対数尤度を簡単に最小化できたら素晴らしいと思いませんか?
```
negloglik = lambda y, rv_y: -rv_y.log_prob(y)
```
このコラボでは(線形回帰問題のコンテキストで)その方法を紹介します。
```
#@title Synthesize dataset.
w0 = 0.125
b0 = 5.
x_range = [-20, 60]
def load_dataset(n=150, n_tst=150):
np.random.seed(43)
def s(x):
g = (x - x_range[0]) / (x_range[1] - x_range[0])
return 3 * (0.25 + g**2.)
x = (x_range[1] - x_range[0]) * np.random.rand(n) + x_range[0]
eps = np.random.randn(n) * s(x)
y = (w0 * x * (1. + np.sin(x)) + b0) + eps
x = x[..., np.newaxis]
x_tst = np.linspace(*x_range, num=n_tst).astype(np.float32)
x_tst = x_tst[..., np.newaxis]
return y, x, x_tst
y, x, x_tst = load_dataset()
```
### ケース 1: 不確実性なし
```
# Build model.
model = tf.keras.Sequential([
tf.keras.layers.Dense(1),
tfp.layers.DistributionLambda(lambda t: tfd.Normal(loc=t, scale=1)),
])
# Do inference.
model.compile(optimizer=tf.optimizers.Adam(learning_rate=0.01), loss=negloglik)
model.fit(x, y, epochs=1000, verbose=False);
# Profit.
[print(np.squeeze(w.numpy())) for w in model.weights];
yhat = model(x_tst)
assert isinstance(yhat, tfd.Distribution)
#@title Figure 1: No uncertainty.
w = np.squeeze(model.layers[-2].kernel.numpy())
b = np.squeeze(model.layers[-2].bias.numpy())
plt.figure(figsize=[6, 1.5]) # inches
#plt.figure(figsize=[8, 5]) # inches
plt.plot(x, y, 'b.', label='observed');
plt.plot(x_tst, yhat.mean(),'r', label='mean', linewidth=4);
plt.ylim(-0.,17);
plt.yticks(np.linspace(0, 15, 4)[1:]);
plt.xticks(np.linspace(*x_range, num=9));
ax=plt.gca();
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
ax.spines['left'].set_position(('data', 0))
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
#ax.spines['left'].set_smart_bounds(True)
#ax.spines['bottom'].set_smart_bounds(True)
plt.legend(loc='center left', fancybox=True, framealpha=0., bbox_to_anchor=(1.05, 0.5))
plt.savefig('/tmp/fig1.png', bbox_inches='tight', dpi=300)
```
### ケース 2: 偶然性の不確実性
```
# Build model.
model = tf.keras.Sequential([
tf.keras.layers.Dense(1 + 1),
tfp.layers.DistributionLambda(
lambda t: tfd.Normal(loc=t[..., :1],
scale=1e-3 + tf.math.softplus(0.05 * t[...,1:]))),
])
# Do inference.
model.compile(optimizer=tf.optimizers.Adam(learning_rate=0.01), loss=negloglik)
model.fit(x, y, epochs=1000, verbose=False);
# Profit.
[print(np.squeeze(w.numpy())) for w in model.weights];
yhat = model(x_tst)
assert isinstance(yhat, tfd.Distribution)
#@title Figure 2: Aleatoric Uncertainty
plt.figure(figsize=[6, 1.5]) # inches
plt.plot(x, y, 'b.', label='observed');
m = yhat.mean()
s = yhat.stddev()
plt.plot(x_tst, m, 'r', linewidth=4, label='mean');
plt.plot(x_tst, m + 2 * s, 'g', linewidth=2, label=r'mean + 2 stddev');
plt.plot(x_tst, m - 2 * s, 'g', linewidth=2, label=r'mean - 2 stddev');
plt.ylim(-0.,17);
plt.yticks(np.linspace(0, 15, 4)[1:]);
plt.xticks(np.linspace(*x_range, num=9));
ax=plt.gca();
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
ax.spines['left'].set_position(('data', 0))
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
#ax.spines['left'].set_smart_bounds(True)
#ax.spines['bottom'].set_smart_bounds(True)
plt.legend(loc='center left', fancybox=True, framealpha=0., bbox_to_anchor=(1.05, 0.5))
plt.savefig('/tmp/fig2.png', bbox_inches='tight', dpi=300)
```
### ケース 3: 認識論的不確実性
```
# Specify the surrogate posterior over `keras.layers.Dense` `kernel` and `bias`.
def posterior_mean_field(kernel_size, bias_size=0, dtype=None):
n = kernel_size + bias_size
c = np.log(np.expm1(1.))
return tf.keras.Sequential([
tfp.layers.VariableLayer(2 * n, dtype=dtype),
tfp.layers.DistributionLambda(lambda t: tfd.Independent(
tfd.Normal(loc=t[..., :n],
scale=1e-5 + tf.nn.softplus(c + t[..., n:])),
reinterpreted_batch_ndims=1)),
])
# Specify the prior over `keras.layers.Dense` `kernel` and `bias`.
def prior_trainable(kernel_size, bias_size=0, dtype=None):
n = kernel_size + bias_size
return tf.keras.Sequential([
tfp.layers.VariableLayer(n, dtype=dtype),
tfp.layers.DistributionLambda(lambda t: tfd.Independent(
tfd.Normal(loc=t, scale=1),
reinterpreted_batch_ndims=1)),
])
# Build model.
model = tf.keras.Sequential([
tfp.layers.DenseVariational(1, posterior_mean_field, prior_trainable, kl_weight=1/x.shape[0]),
tfp.layers.DistributionLambda(lambda t: tfd.Normal(loc=t, scale=1)),
])
# Do inference.
model.compile(optimizer=tf.optimizers.Adam(learning_rate=0.01), loss=negloglik)
model.fit(x, y, epochs=1000, verbose=False);
# Profit.
[print(np.squeeze(w.numpy())) for w in model.weights];
yhat = model(x_tst)
assert isinstance(yhat, tfd.Distribution)
#@title Figure 3: Epistemic Uncertainty
plt.figure(figsize=[6, 1.5]) # inches
plt.clf();
plt.plot(x, y, 'b.', label='observed');
yhats = [model(x_tst) for _ in range(100)]
avgm = np.zeros_like(x_tst[..., 0])
for i, yhat in enumerate(yhats):
m = np.squeeze(yhat.mean())
s = np.squeeze(yhat.stddev())
if i < 25:
plt.plot(x_tst, m, 'r', label='ensemble means' if i == 0 else None, linewidth=0.5)
avgm += m
plt.plot(x_tst, avgm/len(yhats), 'r', label='overall mean', linewidth=4)
plt.ylim(-0.,17);
plt.yticks(np.linspace(0, 15, 4)[1:]);
plt.xticks(np.linspace(*x_range, num=9));
ax=plt.gca();
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
ax.spines['left'].set_position(('data', 0))
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
#ax.spines['left'].set_smart_bounds(True)
#ax.spines['bottom'].set_smart_bounds(True)
plt.legend(loc='center left', fancybox=True, framealpha=0., bbox_to_anchor=(1.05, 0.5))
plt.savefig('/tmp/fig3.png', bbox_inches='tight', dpi=300)
```
### ケース 4: 偶然性の不確実性と認識論的不確実性
```
# Build model.
model = tf.keras.Sequential([
tfp.layers.DenseVariational(1 + 1, posterior_mean_field, prior_trainable, kl_weight=1/x.shape[0]),
tfp.layers.DistributionLambda(
lambda t: tfd.Normal(loc=t[..., :1],
scale=1e-3 + tf.math.softplus(0.01 * t[...,1:]))),
])
# Do inference.
model.compile(optimizer=tf.optimizers.Adam(learning_rate=0.01), loss=negloglik)
model.fit(x, y, epochs=1000, verbose=False);
# Profit.
[print(np.squeeze(w.numpy())) for w in model.weights];
yhat = model(x_tst)
assert isinstance(yhat, tfd.Distribution)
#@title Figure 4: Both Aleatoric & Epistemic Uncertainty
plt.figure(figsize=[6, 1.5]) # inches
plt.plot(x, y, 'b.', label='observed');
yhats = [model(x_tst) for _ in range(100)]
avgm = np.zeros_like(x_tst[..., 0])
for i, yhat in enumerate(yhats):
m = np.squeeze(yhat.mean())
s = np.squeeze(yhat.stddev())
if i < 15:
plt.plot(x_tst, m, 'r', label='ensemble means' if i == 0 else None, linewidth=1.)
plt.plot(x_tst, m + 2 * s, 'g', linewidth=0.5, label='ensemble means + 2 ensemble stdev' if i == 0 else None);
plt.plot(x_tst, m - 2 * s, 'g', linewidth=0.5, label='ensemble means - 2 ensemble stdev' if i == 0 else None);
avgm += m
plt.plot(x_tst, avgm/len(yhats), 'r', label='overall mean', linewidth=4)
plt.ylim(-0.,17);
plt.yticks(np.linspace(0, 15, 4)[1:]);
plt.xticks(np.linspace(*x_range, num=9));
ax=plt.gca();
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
ax.spines['left'].set_position(('data', 0))
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
#ax.spines['left'].set_smart_bounds(True)
#ax.spines['bottom'].set_smart_bounds(True)
plt.legend(loc='center left', fancybox=True, framealpha=0., bbox_to_anchor=(1.05, 0.5))
plt.savefig('/tmp/fig4.png', bbox_inches='tight', dpi=300)
```
### ケース 5: 関数的不確実性
```
#@title Custom PSD Kernel
class RBFKernelFn(tf.keras.layers.Layer):
def __init__(self, **kwargs):
super(RBFKernelFn, self).__init__(**kwargs)
dtype = kwargs.get('dtype', None)
self._amplitude = self.add_variable(
initializer=tf.constant_initializer(0),
dtype=dtype,
name='amplitude')
self._length_scale = self.add_variable(
initializer=tf.constant_initializer(0),
dtype=dtype,
name='length_scale')
def call(self, x):
# Never called -- this is just a layer so it can hold variables
# in a way Keras understands.
return x
@property
def kernel(self):
return tfp.math.psd_kernels.ExponentiatedQuadratic(
amplitude=tf.nn.softplus(0.1 * self._amplitude),
length_scale=tf.nn.softplus(5. * self._length_scale)
)
# For numeric stability, set the default floating-point dtype to float64
tf.keras.backend.set_floatx('float64')
# Build model.
num_inducing_points = 40
model = tf.keras.Sequential([
tf.keras.layers.InputLayer(input_shape=[1]),
tf.keras.layers.Dense(1, kernel_initializer='ones', use_bias=False),
tfp.layers.VariationalGaussianProcess(
num_inducing_points=num_inducing_points,
kernel_provider=RBFKernelFn(),
event_shape=[1],
inducing_index_points_initializer=tf.constant_initializer(
np.linspace(*x_range, num=num_inducing_points,
dtype=x.dtype)[..., np.newaxis]),
unconstrained_observation_noise_variance_initializer=(
tf.constant_initializer(np.array(0.54).astype(x.dtype))),
),
])
# Do inference.
batch_size = 32
loss = lambda y, rv_y: rv_y.variational_loss(
y, kl_weight=np.array(batch_size, x.dtype) / x.shape[0])
model.compile(optimizer=tf.optimizers.Adam(learning_rate=0.01), loss=loss)
model.fit(x, y, batch_size=batch_size, epochs=1000, verbose=False)
# Profit.
yhat = model(x_tst)
assert isinstance(yhat, tfd.Distribution)
#@title Figure 5: Functional Uncertainty
y, x, _ = load_dataset()
plt.figure(figsize=[6, 1.5]) # inches
plt.plot(x, y, 'b.', label='observed');
num_samples = 7
for i in range(num_samples):
sample_ = yhat.sample().numpy()
plt.plot(x_tst,
sample_[..., 0].T,
'r',
linewidth=0.9,
label='ensemble means' if i == 0 else None);
plt.ylim(-0.,17);
plt.yticks(np.linspace(0, 15, 4)[1:]);
plt.xticks(np.linspace(*x_range, num=9));
ax=plt.gca();
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
ax.spines['left'].set_position(('data', 0))
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
#ax.spines['left'].set_smart_bounds(True)
#ax.spines['bottom'].set_smart_bounds(True)
plt.legend(loc='center left', fancybox=True, framealpha=0., bbox_to_anchor=(1.05, 0.5))
plt.savefig('/tmp/fig5.png', bbox_inches='tight', dpi=300)
```
| github_jupyter |
# Acquire data & data structures
```
import requests
import pandas as pd
```
## Internet
#### Check the response status code
- status code 200: the request response cycle was successful
- any other status code: it didn't work (e.g., 404 = page not found)
- Convert content to utf-8 if necessary
```
def connect(url, decode='utf-8'):
response = requests.get(url)
if response.status_code == 200:
print('successfully connected, response code: {}'.format(response.status_code))
else:
print('connection failed')
return response.content.decode(decode)
url = 'http://www.lauthom.nl/search/tools'
content = connect(url)
content[:500]
```
### JSON
```
import json
```
### json.loads recursively decodes a string in JSON format into equivalent python objects
- data_string's outermost element is converted into a python list
- the first element of that list is converted into a dictionary
- the key of that dictionary is converted into a string
- the value of that dictionary is converted into a list of two integer elements
```
data_string = '[{"b": [2, 4], "c": 3.0, "a": "A"}]'
python_data = json.loads(data_string)
print('{}\n{}\n{}\n{}\n{}\n{}'.format(type(data_string), type(python_data), python_data, python_data[0], python_data[0]['b'], python_data[0]['b'][1]))
```
### json.dumps and json.loads
```
JSON_string = "JSON throws exception when not in correct format"
print(JSON_string)
# Stringify strings
JSON_stringified = json.dumps(JSON_string)
print(JSON_stringified)
# Correct
json.loads(JSON_stringified)
# JSONDecodeError
# json.loads(JSON_string)
```
### requests & JSON
```
address = 'Amsterdam, Netherlands'
url = 'https://maps.googleapis.com/maps/api/geocode/json?address={}'.format(address)
response = requests.get(url).json()
type(response), response
```
### Get JSON formatted content
```
def get_json(url, decode='utf-8'):
try:
response = requests.get(url)
if not response.status_code == 200:
print('HTTP error, response code: {}'.format(response.status_code))
else:
try:
response_data = response.json()
except:
print("response not in valid JSON format")
except:
print('something went wrong with requests.get')
return response_data
response_data = get_json(url)
response_data
```
### Get address, latitude, longitude
```
def get_lat_lng(url):
response = get_json(url)
result = response['results'][0]
formatted_address = result['formatted_address']
lat = result['geometry']['location']['lat']
lng = result['geometry']['location']['lng']
return formatted_address, lat, lng
get_lat_lng(url)
address = 'London Business School'
url = 'https://maps.googleapis.com/maps/api/geocode/json?address={}'.format(address)
get_lat_lng(url)
```
### Get list of addresses with lat, lon
```
def get_lat_lng_list(url):
response = get_json(url)
result_list = []
for result in response['results']:
formatted_address = result['formatted_address']
lat = result['geometry']['location']['lat']
lng = result['geometry']['location']['lng']
result_list.append((formatted_address, lat, lng))
return result_list
address = 'Baker Street'
url = 'https://maps.googleapis.com/maps/api/geocode/json?address={}'.format(address)
get_lat_lng_list(url)
```
## XML
- library lxml - deals with converting an XML-string to python objects and vice versa
```
from lxml import etree
data_string = """
<Bookstore>
<Book ISBN="ISBN-13:978-1599620787" Price="15.23" Weight="1.5">
<Title>New York Deco</Title>
<Authors>
<Author Residence="New York City">
<First_Name>Richard</First_Name>
<Last_Name>Berenholtz</Last_Name>
</Author>
</Authors>
</Book>
<Book ISBN="ISBN-13:978-1579128562" Price="15.80">
<Remark>
Five Hundred Buildings of New York and over one million other books are available for Amazon Kindle.
</Remark>
<Title>Five Hundred Buildings of New York</Title>
<Authors>
<Author Residence="Beijing">
<First_Name>Bill</First_Name>
<Last_Name>Harris</Last_Name>
</Author>
<Author Residence="New York City">
<First_Name>Jorg</First_Name>
<Last_Name>Brockmann</Last_Name>
</Author>
</Authors>
</Book>
</Bookstore>
"""
root = etree.XML(data_string)
root.tag, type(root.tag)
print(etree.tostring(root, pretty_print=True).decode("utf-8"))
```
#### Iterating over complete XML tree
```
for element in root.iter():
print(element)
```
#### Iterate over children in subtree, accessing tags
```
for child in root:
print(child, child.tag)
```
#### Iterate to get specific tags and data
1. author tags are accessed
2. For each author tag, the .find function accesses the First_Name and Last_Name tags
3. The .find function only looks at the children, not other descendants, so be careful!
4. The .text attribute prints the text in a leaf node
```
for element in root.iter('Author'):
print(element.find('First_Name').text, element.find('Last_Name').text)
```
#### Filter values of attributes
e.g. find the first name of the author of a book that weighs 1.5 oz
```
root.find('Book[@Weight="1.5"]/Authors/Author/First_Name').text
```
## Exchange rates from XE.com
```
url = 'https://www.xe.com/currencyconverter/convert/?Amount=1&From=USD&To=EUR'
```
### BeautifulSoup
```
from bs4 import BeautifulSoup
def result_page(url, keywords=''):
response = requests.get(url + keywords)
if not response.status_code == 200:
return None
return BeautifulSoup(response.content, 'lxml')
def get_data(url, keywords='', selector=''):
rate_list = []
try:
results_page = result_page(url, keywords)
rates = results_page.find_all('td', class_='rateCell')
for rate in rates:
rate_ = rate.get_text()
try:
currency = rate.find('a').get('rel')[0][:7]
rate_list.append((currency, rate_))
except:
currency = ''
return rate_list
except:
return None
pd.DataFrame(get_data(url))
```
| github_jupyter |
# Lab 4: Functions and Functional Programming (Part 2)
Look at you go! Congratulations on making it to the second part of the lab!
These assignments are *absolutely not required*! Even if you're here, you may find it more valuable to skim the problems here and attempt the problems that are most interesting to you - and that's perfectly fine. Don't feel any need to complete them in sequential order at this point.
I'm honestly WAYYYY more psyched about the functional programming stuff than the functions stuff (don't tell anyone 👀) so let's start there!
## Building Decorators
### Automatic Caching
In class, we wrote a decorator `memoize` that will automatically caches any calls to the decorated function. You can assume that all arguments passed to the decorated function will always be hashable types.
```Python
def memoize(function):
cache = {}
def memoized_fn(*args):
if args not in cache:
cache[args] = function(*args)
return cache[args]
return memoized_fn
```
We saw how one use case for this in class:
```Python
@memoize
def fib(n):
return fib(n-1) + fib(n-2) if n > 2 else 1
fib(10) # 55 (takes a moment to execute)
fib(10) # 55 (returns immediately)
fib(100) # doesn't take forever
fib(400) # doesn't raise RuntimeError
```
#### Cache Options (Challenge)
Add `maxsize` and `eviction_policy` keyword arguments, with reasonable defaults (perhaps `maxsize=None` as a sentinel), to your `cache` decorator. `eviction_policy` should be one of `'LRU'`, `'MRU'`, or `'random'`. It can be tricky to figure out how to construct a decorator with arguments.
Also, add function attributes called `.cache_info` and `.cache_clear` which can be called to get aggregate statistics about the cache and clear the cache, respectively.
*Note*: This caching decorator (with arguments!) is actually implemented as part of the language in `functools.lru_cache`
```
def memoize(???):
pass
@memoize(???)
def fib(n):
return fib(n-1) + fib(n-2) if n > 2 else 1
```
### Better Debugging Decorator
The `debug` decorator we wrote in class isn't very good. It doesn't tell us which function is being called, and it just dumps a tuple of positional arguments and a dictionary of keyword arguments - it doesn't even know what the names of the positional arguments are! If the default arguments aren't overridden, it won't show us their value either.
Use function attributes to improve our `debug` decorator into a `print_args` decorator that is "as good as you can make it."
```Python
def print_args(function):
def wrapper(*args, **kwargs):
# (1) You could do something here
retval = function(*args, **kwargs)
# (2) You could also do something here
return retval
return wrapper
```
*Hint: Consider using the attributes `fn.__name__` and `fn.__code__`. You'll have to investigate these attributes, but I will say that the `fn.__code__` code object contains a number of useful attributes - for instance, `fn.__code__.co_varnames`. Check it out! More information on function attributes is available in the latter half of Lab 3.*
#### Note
There are a lot of subtleties to this function, since functions can be called in a number of different ways. How does your `print_args` handle keyword arguments or even keyword-only arguments? Variadic positional arguments? Variadic keyword arguments? For more customization, look at `fn.__defaults__`, `fn.__kwdefaults__`, as well as other attributes of `fn.__code__`.
```
def print_args(???):
pass
@print_args(???)
def is_prime(n):
for i in range(2, n):
if n % i == 0:
return False
return True
print(is_prime(198239813))
print(is_prime(4028769383))
@print_args(???)
def stylize_quote(quote, **kwargs):
print('> {}'.format(quote))
print('-'*(len(quote) + 2))
for k, v in kwargs.items():
print('{k}: {v}'.format(k=k, v=v))
stylize_quote('Doth mother know you weareth her drapes?', speaker='Iron Man', year='2012', movie='The Avengers')
@print_args(???)
def draw_table(num_rows, num_cols):
sep = '+' + '+'.join(['-'] * num_cols) + '+'
line = '|' + '|'.join([' '] * num_cols) + '|'
for _ in range(num_rows):
print(sep)
print(line)
print(sep)
draw_table(10, 10)
draw_table(3, 8)
```
### Dynamic Type Checker (challenge)
Functions in Python can be optionally annotated by semantically-useless but structurally-valuable type annotations. For example:
```Python
def foo(a: int, b: str) -> bool:
return b[a] == 'X'
foo.__annotations__ # => {'a': int, 'b': str, 'return': bool}
```
Write a runtime type checker, implemented as a decorator, that enforces that the types of arguments and the return value are valid.
```Python
def enforce_types(function):
pass # Your implementation here
```
For example:
```Python
@enforce_types
def foo(a: int, b: str) -> bool:
if a == -1:
return 'Gotcha!'
return b[a] == 'X'
foo(3, 'abcXde') # => True
foo(2, 'python') # => False
foo(1, 4) # prints "Invalid argument type for b: expected str, received int
foo(-1, '') # prints "Invalid return type: expected bool, received str
```
There are lots of nuances to this function. What happens if some annotations are missing? How are keyword arguments and variadic arguments handled? What happens if the expected type of a parameter is not a primitive type? Can you annotate a function to describe that a parameter should be a list of strings? A tuple of (str, bool) pairs? A dictionary mapping strings to lists of integers? Read more about [advanced type hints](https://docs.python.org/3/library/typing.html) from the documentation.
As you make progress, show your decorator to a member of the course staff.
```
def enforce_types(function):
pass # Your implementation here
@enforce_types
def foo(a: int, b: str) -> bool:
if a == -1:
return 'Gotcha!'
return b[a] == 'X'
foo(3, 'abcXde') # => True
foo(2, 'python') # => False
foo(1, 4) # prints "Invalid argument type for b: expected str, received int
foo(-1, '') # prints "Invalid return type: expected bool, received str
```
## Nested Functions and Closures
In class, we saw that a function can be defined within the scope of another function. Recall from Week 3 that functions introduce new scopes via a new local symbol table. An inner function is only in scope inside of the outer function, so this type of function definition is usually only used when the inner function is being returned to the outside world.
```Python
def outer():
def inner(a):
return a
return inner
f = outer()
print(f) # <function outer.<locals>.inner at 0x1044b61e0>
print(f(10)) # => 10
f2 = outer()
print(f2) # <function outer.<locals>.inner at 0x1044b6268> (Different from above!)
print(f2(11)) # => 11
```
Why are the memory addresses different for `f` and `f2`? Discuss with a partner.
```
def outer():
def inner(a):
return a
return inner
f = outer()
print(f) # <function outer.<locals>.inner at 0x1044b61e0>
print(f(10)) # => 10
f2 = outer()
print(f2) # <function outer.<locals>.inner at 0x1044b6268> (Different from above!)
print(f2(11)) # => 11
```
### Closure
As we saw above, the definition of the inner function occurs during the execution of the outer function. This implies that a nested function has access to the environment in which it was defined. Therefore, it is possible to return an inner function that remembers contents of the outer function, even after the outer function has completed execution. This model is referred to as a closure.
```Python
def make_adder(n):
def add_n(m): # Captures the outer variable `n` in a closure
return m + n
return add_n
add1 = make_adder(1)
print(add1) # <function make_adder.<locals>.add_n at 0x103edf8c8>
add1(4) # => 5
add1(5) # => 6
add2 = make_adder(2)
print(add2) # <function make_adder.<locals>.add_n at 0x103ecbf28>
add2(4) # => 6
add2(5) # => 7
```
The information in a closure is available in the function's `__closure__` attribute. For example:
```Python
closure = add1.__closure__
cell0 = closure[0]
cell0.cell_contents # => 1 (this is the n = 1 passed into make_adder)
```
As another example, consider the function:
```Python
def foo(a, b, c=-1, *d, e=-2, f=-3, **g):
def wraps():
print(a, c, e, g)
return wraps
```
The `print` call induces a closure of `wraps` over `a`, `c`, `e`, `g` from the enclosing scope of `foo`. Or, you can imagine that wraps "knows" that it will need `a`, `c`, `e`, and `g` from the enclosing scope, so at the time `wraps` is defined, Python takes a "screenshot" of these variables from the enclosing scope and stores references to the underlying objects in the `__closure__` attribute of the `wraps` function.
```Python
w = foo(1, 2, 3, 4, 5, e=6, f=7, y=2, z=3)
list(map(lambda cell: cell.cell_contents, w.__closure__))
# => [1, 3, 6, {'y': 2, 'z': 3}]
```
What happens in the following situation? Why?
```Python
def outer(l):
def inner(n):
return l * n
return inner
l = [1, 2, 3]
f = outer(l)
print(f(3)) # => ??
l.append(4)
print(f(3)) # => ??
```
```
def outer(l):
def inner(n):
return l * n
return inner
l = [1, 2, 3]
f = outer(l)
print(f(3)) # => ??
l.append(4)
print(f(3)) # => ??
```
## Functions
Alright, back to functions! This stuff is really fun too! Let's start with an optional problem that puts together all of the things we've learned about functions so far.
### *Optional: Putting it all together*
*If you feel confident that you understand how function calling works, you can skip this section. We suggest that you work through it if you'd like more practice, but the final decision is up to you.*
Often, however, we don't just see keyword arguments of variadic parameter lists in isolated situations. The following function definition, which incorporates positional parameters, keyword parameters, variadic positional parameters, keyword-only default parameters and variadic keyword parameters, is valid Python code.
```Python
def all_together(x, y, z=1, *nums, indent=True, spaces=4, **options):
print("x:", x)
print("y:", y)
print("z:", z)
print("nums:", nums)
print("indent:", indent)
print("spaces:", spaces)
print("options:", options)
```
For each of the following function calls, predict whether the call is valid or not. If it is valid, what will the output be? If it is invalid, what is the cause of the error?
```Python
all_together(2)
all_together(2, 5, 7, 8, indent=False)
all_together(2, 5, 7, 6, indent=None)
all_together()
all_together(indent=True, 3, 4, 5)
all_together(**{'indent': False}, scope='maximum')
all_together(dict(x=0, y=1), *range(10))
all_together(**dict(x=0, y=1), *range(10))
all_together(*range(10), **dict(x=0, y=1))
all_together([1, 2], {3:4})
all_together(8, 9, 10, *[2, 4, 6], x=7, spaces=0, **{'a':5, 'b':'x'})
all_together(8, 9, 10, *[2, 4, 6], spaces=0, **{'a':[4,5], 'b':'x'})
all_together(8, 9, *[2, 4, 6], *dict(z=1), spaces=0, **{'a':[4,5], 'b':'x'})
```
```
# Before running me, predict which of these calls will be invalid and which will be valid!
# For valid calls, what is the output?
# For invalid calls, why is it invalid?
def all_together(x, y, z=1, *nums, indent=True, spaces=4, **options):
print("x:", x)
print("y:", y)
print("z:", z)
print("nums:", nums)
print("indent:", indent)
print("spaces:", spaces)
print("options:", options)
# Uncomment the ones you want to run!
# all_together(2)
# all_together(2, 5, 7, 8, indent=False)
# all_together(2, 5, 7, 6, indent=None)
# all_together()
# all_together(indent=True, 3, 4, 5)
# all_together(**{'indent': False}, scope='maximum')
# all_together(dict(x=0, y=1), *range(10))
# all_together(**dict(x=0, y=1), *range(10))
# all_together(*range(10), **dict(x=0, y=1))
# all_together([1, 2], {3:4})
# all_together(8, 9, 10, *[2, 4, 6], x=7, spaces=0, **{'a':5, 'b':'x'})
# all_together(8, 9, 10, *[2, 4, 6], spaces=0, **{'a':[4,5], 'b':'x'})
# all_together(8, 9, *[2, 4, 6], *dict(z=1), spaces=0, **{'a':[4,5], 'b':'x'})
```
Write at least two more instances of function calls, not listed above, and predict their output. Are they valid or invalid? Check your hypothesis.
```
# Write two more function calls.
# all_together(...)
# all_together(...)
```
### Default Mutable Arguments - A Dangerous Game
A function's default values are evaluated at the point of function definition in the defining scope. For example:
```
x = 5
def square(num=x):
return num * num
x = 6
print(square()) # => 25, not 36
print(square(x)) # => 36
```
**Warning: A function's default values are evaluated *only once*, when the function definition is encountered. This is important when the default value is a mutable object such as a list or dictionary**
Predict what the following code will do, then run it to test your hypothesis:
```Python
def append_twice(a, lst=[]):
lst.append(a)
lst.append(a)
return lst
# Works well when the keyword is provided
print(append_twice(1, lst=[4])) # => [4, 1, 1]
print(append_twice(11, lst=[2, 3, 5, 7])) # => [2, 3, 5, 7, 11, 11]
# But what happens here?
print(append_twice(1))
print(append_twice(2))
print(append_twice(3))
```
```
# Something fishy is going on here. Can you deduce what is happening?
def append_twice(a, lst=[]):
lst.append(a)
lst.append(a)
return lst
# Works well when the keyword is provided
print(append_twice(1, lst=[4])) # => [4, 1, 1]
print(append_twice(11, lst=[2, 3, 5, 7])) # => [2, 3, 5, 7, 11, 11]
# But what happens here?
print(append_twice(1))
print(append_twice(2))
print(append_twice(3))
```
After you run the code, you should see the following printed to the screen:
```
[1, 1]
[1, 1, 2, 2]
[1, 1, 2, 2, 3, 3]
```
Discuss with a partner why this is happening.
If you don’t want the default value to be shared between subsequent calls, you can use a sentinel value as the default value (to signal that no keyword argument was explicitly provided by the caller). If so, your function may look something like:
```Python
def append_twice(a, lst=None):
if lst is None:
lst = []
lst.append(a)
lst.append(a)
return lst
```
Discuss with a partner whether you think this solution feels better or worse.
```
def append_twice(a, lst=None):
if lst is None:
lst = []
lst.append(a)
lst.append(a)
return lst
```
## Investigating Function Objects
In Monday's class, we mentioned that functions are objects, and that they might have interesting attributes to explore. We'll poke around several of these attributes more in depth here.
Usually, this information isn't particularly useful for practitioners (you'll rarely want to hack around with the internals of functions), but even seeing that you *can* in Python is very cool.
In this section, there is no code to write. Instead, you will be reading and running code and observing the output. Nevertheless, we encourage you to play around with the code cells to experiment and explore on your own.
#### Default Values (`__defaults__` and `__kwdefaults__`)
As stated earlier, any default values (either normal default arguments or the keyword-only default arguments that follow a variadic positional argument parameter) are bound to the function object at the time of function definition. Consider our `all_together` function from earlier, and run the following code. Why might the `__defaults__` attribute be a tuple, but the `__kwdefaults__` attribute be a dictionary?
```
def all_together(x, y, z=1, *nums, indent=True, spaces=4, **options): pass
all_together.__defaults__ # => (1, )
all_together.__kwdefaults__ # => {'indent':True, 'spaces':4}
```
#### Documentation (`__doc__`)
The first string literal in any function, if it comes before any expression, is bound to the function's `__doc__` attribute.
```
def my_function():
"""Summary line: do nothing, but document it.
Description: No, really, it doesn't do anything.
"""
pass
print(my_function.__doc__)
# Summary line: Do nothing, but document it.
#
# Description: No, really, it doesn't do anything.
```
As stated in lecture, lots of tools use these documentation strings to great advantage. For example, the builtin `help` function displays information from docstrings, and many API-documentation-generation tools like [Sphynx](http://www.sphinx-doc.org/en/stable/) or [Epydoc](http://epydoc.sourceforge.net/) use information contained in the docstring to form smart references and hyperlinks on documentation websites.
Furthermore, the [doctest](https://docs.python.org/3/library/doctest.html) standard library module, in it's own words, "searches [the documentation string] for pieces of text that look like interactive Python sessions, and then executes those sessions to verify that they work exactly as shown." Cool!
#### Code Object (`__code__`)
In CPython, the reference implementation of Python used by many people (including us), functions are byte-compiled into executable Python code, or _bytecode_, when defined. This code object, which represents the bytecode and some administrative information, is bound to the `__code__` attribute, and has a ton of interesting properties, best illustrated by example. Code objects are immutable and contain no references to immutable objects.
```Python
def all_together(x, y, z=1, *nums, indent=True, spaces=4, **options):
"""A useless comment"""
print(x + y * z)
print(sum(nums))
for k, v in options.items():
if indent:
print("{}\t{}".format(k, v))
else:
print("{}{}{}".format(k, " " * spaces, v))
code = all_together.__code__
```
| Attribute | Sample Value | Explanation |
| --- | --- | --- |
| `code.co_argcount` | `3` | number of positional arguments (including arguments with default values) |
| `code.co_cellvars` | `()` | tuple containing the names of local variables that are referenced by nested functions |
| `code.co_code` | `b't\x00\x00...\x04S\x00'` | string representing the sequence of bytecode instructions |
| `code.co_consts` | `('A useless comment', '{}\t{}', '{}{}{}', ' ', None)` | tuple containing the literals used by the bytecode - our `None` is from the implicit `return None` at the end |
| `code.co_filename` | `filename` or `<stdin>` or `<ipython-input-#-xxx>` | file in which the function was defined |
| `code.co_firstlineno` | `1` | line of the file the first line of the function appears |
| `code.co_flags` | `79` | AND of compiler-specific binary flags whose internal meaning is (mostly) opaque to us |
| `code.co_freevars` | `()` | tuple containing the names of free variables |
| `code.co_kwonlyargcount` | `2` | number of keyword-only arguments |
| `code.co_lnotab` | `b'\x00\x02\x10\x01\x0c\x01\x12\x01\x04\x01\x12\x02'` | string encoding the mapping from bytecode offsets to line numbers |
| `code.co_name` | `"all_together"` | the function name |
| `code.co_names` | `('print', 'sum', 'items', 'format')` | tuple containing the names used by the bytecode |
| `code.co_nlocals` | `9` | number of local variables used by the function (including arguments) |
| `code.co_stacksize` | `7` | required stack size (including local variables) |
| `code.co_varnames` | `('x', 'y', 'z', 'indent', 'spaces', 'nums', 'options', 'k', 'v')` | tuple containing the names of the local variables (starting with the argument names) |
More info on this, and on all types in Python, can be found at the [data model reference](https://docs.python.org/3/reference/datamodel.html#the-standard-type-hierarchy). For code objects, you have to scroll down to "Internal Types."
```
def all_together(x, y, z=1, *nums, indent=True, spaces=4, **options):
"""A useless comment"""
print(x + y * z)
print(sum(nums))
for k, v in options.items():
if indent:
print("{}\t{}".format(k, v))
else:
print("{}{}{}".format(k, " " * spaces, v))
code = all_together.__code__
print(code.co_argcount)
print(code.co_cellvars)
print(code.co_code)
print(code.co_consts)
print(code.co_filename)
print(code.co_firstlineno)
print(code.co_flags)
print(code.co_freevars)
print(code.co_kwonlyargcount)
print(code.co_lnotab)
print(code.co_name)
print(code.co_names)
print(code.co_nlocals)
print(code.co_stacksize)
print(code.co_varnames)
```
##### Security
As we briefly mentioned in class, this can lead to a pretty glaring security vulnerability. Namely, the code object on a given function can be hot-swapped for the code object of another (perhaps malicious function) at runtime!
```
def nice(): print("You're awesome!")
def mean(): print("You're... not awesome. OOOOH")
# Overwrite the code object for nice
nice.__code__ = mean.__code__
print(nice()) # prints "You're... not awesome. OOOOH"
```
##### `dis` module
The `dis` module, for "disassemble," exports a `dis` function that allows us to disassemble Python byte code (at least, for Python distributions implemented in CPython for existing versions). The disassembled code isn't exactly normal assembly code, but rather is a specialized Python syntax
```Python
def gcd(a, b):
while b:
a, b = b, a % b
return a
import dis
dis.dis(gcd)
"""
2 0 SETUP_LOOP 27 (to 30)
>> 3 LOAD_FAST 1 (b)
6 POP_JUMP_IF_FALSE 29
3 9 LOAD_FAST 1 (b)
12 LOAD_FAST 0 (a)
15 LOAD_FAST 1 (b)
18 BINARY_MODULO
19 ROT_TWO
20 STORE_FAST 0 (a)
23 STORE_FAST 1 (b)
26 JUMP_ABSOLUTE 3
>> 29 POP_BLOCK
4 >> 30 LOAD_FAST 0 (a)
33 RETURN_VALUE
"""
```
Details on the instructions themselves can be found [here](https://docs.python.org/3/library/dis.html#python-bytecode-instructions).
You can read more about the `dis` module [here](https://docs.python.org/3/library/dis.html).
```
def gcd(a, b):
while b:
a, b = b, a % b
return a
import dis
dis.dis(gcd)
```
#### Parameter Annotations (`__annotations__`)
Python allows us to add type annotations on functions arguments and return values. This leads to a world of complex possibilities and is still fairly controversial in the Python ecosystem. Nevertheless, it can be used to communicate to your clients expectations for the types of arguments.
Importantly, Python doesn't actually do anything with these annotations and will not check that supplied arguments conform to the type hint specified. This language feature is only made available through the collection of function annotations.
```
def annotated(a: int, b: str) -> list:
return [a, b]
print(annotated.__annotations__)
# => {'b': <class 'str'>, 'a': <class 'int'>, 'return': <class 'list'>}
```
This information can be used to build some really neat runtime type-checkers for Python!
For more info, check out [PEP 3107](https://www.python.org/dev/peps/pep-3107/) on function annotations or [PEP 484](https://www.python.org/dev/peps/pep-0484/) on type hinting (which was introduced in Python 3.5)
#### Call (`__call__`)
All Python functions have a `__call__` attribute, which is the actual object called when you use parentheses to "call" a function. That is,
```
def greet(): print("Hello world!")
greet() # "Hello world!"
# is just syntactic sugar for
greet.__call__() # "Hello world!"
```
This means that any object (including instances of custom classes) with a `__call__` method can use the parenthesized function call syntax! For example, we can construct a callable `Polynomial` class. We haven't talked about class syntax yet, so feel free to skip this example.
```Python
class Polynomial:
def __init__(self, coeffs):
"""Store the coefficients..."""
def __call__(self, x):
"""Compute f(x)..."""
# The polynomial f(x) = 4 + 4 * x + 4 * x ** 2
f = Polynomial(4, 4, 1)
f(5) # Really, this is f.__call__(5)
```
We'll see a lot more about using these so-called "magic methods" to exploit Python's apparent operators (like function calling, `+` (`__add__`) or `*` (`__mul__`), etc) in Week 5.
#### Name Information (`__module__`, `__name__`, and `__qualname__`)
Python functions also store some name information about a function, generally for the purposes of friendly printing.
`__module__` refers to the module that was active at the time the function was defined. Any functions defined in the interactive interpreter, or run as as a script, will have `__module__ == '__main__'`, but imported modules will have their `__module__` attribute set to the module name. For example, `math.sqrt.__module__` is `"math"`.
`__name__` is the function's name. Nothing special here.
`__qualname__`, which stands for "qualified name," only differs from `__name__` when you're dealing with nested functions, which we'll talk about more Week 4.
#### Closure (`__closure__`)
If you're familiar with closures in other languages, Python closures work almost the exact same way. Closures really only arise when dealing with nested functions, so we'll see more Week 4. This bit of text is just to give you a teaser for what's coming soon - yes, Python has closures!
#### `inspect` module
As a brief note, all of this mucking around with the internals of Python functions can't be good for our health. Luckily, there's a standard library module for this! The `inspect` module gives us a lot of nice tools for interacting not only with the internals of functions, but also the internals of a lot of other types as well. Check out [the documentation](https://docs.python.org/3/library/inspect.html) for some nice examples.
```
import inspect
def all_together(x, y, z=1, *nums, indent=True, spaces=4, **options): pass
print(inspect.getfullargspec(all_together))
```
## Finished Early?
Wow! Uh... this is all we've got for you. So at this point, feel free to call a TA over, have them sign off on your work, and then you're free to go!
If you'd still like to stay in lab, though, and you didn't get a chnace to read through the following documents last week, though, now is a perfectly good time to peruse them: scan through [PEP 8](https://www.python.org/dev/peps/pep-0008/), Python's official style guide, as well as [PEP 257](https://www.python.org/dev/peps/pep-0257/), Python's suggestions for docstring conventions, if you didn't get a chance to read them last week.
> With 🦄s by @psarin and @coopermj
| github_jupyter |

```
%local
from aws.finspace.cluster import FinSpaceClusterManager
# if this was already run, no need to run again
if 'finspace_clusters' not in globals():
finspace_clusters = FinSpaceClusterManager()
finspace_clusters.auto_connect()
else:
print(f'connected to cluster: {finspace_clusters.get_connected_cluster_id()}')
```
## Configure Spark for Snowflake
This ensures the cluster gets the maven packages deployed to it so the cluster can communicate with Snowflake. The '-f' argument below will force any running spark session to restart.
```
%%configure -f
{
"conf": {
"spark.jars.packages": "net.snowflake:snowflake-jdbc:3.13.5,net.snowflake:spark-snowflake_2.11:2.9.0-spark_2.4"
}
}
%local
import configparser
# read the config
config = configparser.ConfigParser()
config.read("snowflake.ini")
# values from config
snowflake_user=config['snowflake']['user']
snowflake_password=config['snowflake']['password']
snowflake_account=config['snowflake']['account']
snowflake_database=config['snowflake']['database']
snowflake_warehouse=config['snowflake']['warehouse']
print(f"""snowflake_user={snowflake_user}
snowflake_password={snowflake_password}
snowflake_account={snowflake_account}
snowflake_database={snowflake_database}
snowflake_warehouse={snowflake_warehouse}
""")
%send_to_spark -i snowflake_user
%send_to_spark -i snowflake_password
%send_to_spark -i snowflake_account
%send_to_spark -i snowflake_database
%send_to_spark -i snowflake_warehouse
# Snowflake options for the spark data source
# username and password should be protected, admitedly in the clear for convenience
sfOptions = {
"sfURL" : f"{snowflake_account}.snowflakecomputing.com",
"sfUser" : snowflake_user,
"sfPassword" : snowflake_password,
"sfDatabase" : snowflake_database,
"sfWarehouse" : snowflake_warehouse,
"autopushdown": "on",
"keep_column_case": "on"
}
# class name for the snowflake spark data source
SNOWFLAKE_SOURCE_NAME = "net.snowflake.spark.snowflake"
```
# Python Helper Classes
These are the FinSpace helper classes found in the samples and examples github
```
# %load finspace.py
import datetime
import time
import boto3
import os
import pandas as pd
import urllib
from urllib.parse import urlparse
from botocore.config import Config
from boto3.session import Session
# Base FinSpace class
class FinSpace:
def __init__(
self,
config=Config(retries={'max_attempts': 3, 'mode': 'standard'}),
boto_session: Session = None,
dev_overrides: dict = None,
service_name = 'finspace-data'):
"""
To configure this class object, simply instantiate with no-arg if hitting prod endpoint, or else override it:
e.g.
`hab = FinSpaceAnalyticsManager(region_name = 'us-east-1',
dev_overrides = {'hfs_endpoint': 'https://39g32x40jk.execute-api.us-east-1.amazonaws.com/alpha'})`
"""
self.hfs_endpoint = None
self.region_name = None
if dev_overrides is not None:
if 'hfs_endpoint' in dev_overrides:
self.hfs_endpoint = dev_overrides['hfs_endpoint']
if 'region_name' in dev_overrides:
self.region_name = dev_overrides['region_name']
else:
if boto_session is not None:
self.region_name = boto_session.region_name
else:
self.region_name = self.get_region_name()
self.config = config
self._boto3_session = boto3.session.Session(region_name=self.region_name) if boto_session is None else boto_session
print(f"service_name: {service_name}")
print(f"endpoint: {self.hfs_endpoint}")
print(f"region_name: {self.region_name}")
self.client = self._boto3_session.client(service_name, endpoint_url=self.hfs_endpoint, config=self.config)
@staticmethod
def get_region_name():
req = urllib.request.Request("http://169.254.169.254/latest/meta-data/placement/region")
with urllib.request.urlopen(req) as response:
return response.read().decode("utf-8")
# --------------------------------------
# Utility Functions
# --------------------------------------
@staticmethod
def get_list(all_list: dir, name: str):
"""
Search for name found in the all_list dir and return that list of things.
Removes repetitive code found in functions that call boto apis then search for the expected returned items
:param all_list: list of things to search
:type: dir:
:param name: name to search for in all_lists
:type: str
:return: list of items found in name
"""
r = []
# is the given name found, is found, add to list
if name in all_list:
for s in all_list[name]:
r.append(s)
# return the list
return r
# --------------------------------------
# Classification Functions
# --------------------------------------
def list_classifications(self):
"""
Return list of all classifications
:return: all classifications
"""
all_list = self.client.list_classifications(sort='NAME')
return self.get_list(all_list, 'classifications')
def classification_names(self):
"""
Get the classifications names
:return list of classifications names only
"""
classification_names = []
all_classifications = self.list_classifications()
for c in all_classifications:
classification_names.append(c['name'])
return classification_names
def classification(self, name: str):
"""
Exact name search for a classification of the given name
:param name: name of the classification to find
:type: str
:return
"""
all_classifications = self.list_classifications()
existing_classification = next((c for c in all_classifications if c['name'].lower() == name.lower()), None)
if existing_classification:
return existing_classification
def describe_classification(self, classification_id: str):
"""
Calls the describe classification API function and only returns the taxonomy portion of the response.
:param classification_id: the GUID of the classification to get description of
:type: str
"""
resp = None
taxonomy_details_resp = self.client.describe_taxonomy(taxonomyId=classification_id)
if 'taxonomy' in taxonomy_details_resp:
resp = taxonomy_details_resp['taxonomy']
return (resp)
def create_classification(self, classification_definition):
resp = self.client.create_taxonomy(taxonomyDefinition=classification_definition)
taxonomy_id = resp["taxonomyId"]
return (taxonomy_id)
def delete_classification(self, classification_id):
resp = self.client.delete_taxonomy(taxonomyId=classification_id)
if resp['ResponseMetadata']['HTTPStatusCode'] != 200:
return resp
return True
# --------------------------------------
# Attribute Set Functions
# --------------------------------------
def list_attribute_sets(self):
"""
Get list of all dataset_types in the system
:return: list of dataset types
"""
resp = self.client.list_dataset_types()
results = resp['datasetTypeSummaries']
while "nextToken" in resp:
resp = self.client.list_dataset_types(nextToken=resp['nextToken'])
results.extend(resp['datasetTypeSummaries'])
return (results)
def attribute_set_names(self):
"""
Get the list of all dataset type names
:return list of all dataset type names
"""
dataset_type_names = []
all_dataset_types = self.list_dataset_types()
for c in all_dataset_types:
dataset_type_names.append(c['name'])
return dataset_type_names
def attribute_set(self, name: str):
"""
Exact name search for a dataset type of the given name
:param name: name of the dataset type to find
:type: str
:return
"""
all_dataset_types = self.list_dataset_types()
existing_dataset_type = next((c for c in all_dataset_types if c['name'].lower() == name.lower()), None)
if existing_dataset_type:
return existing_dataset_type
def describe_attribute_set(self, attribute_set_id: str):
"""
Calls the describe dataset type API function and only returns the dataset type portion of the response.
:param attribute_set_id: the GUID of the dataset type to get description of
:type: str
"""
resp = None
dataset_type_details_resp = self.client.describe_dataset_type(datasetTypeId=attribute_set_id)
if 'datasetType' in dataset_type_details_resp:
resp = dataset_type_details_resp['datasetType']
return (resp)
def create_attribute_set(self, attribute_set_def):
resp = self.client.create_dataset_type(datasetTypeDefinition=attribute_set_def)
att_id = resp["datasetTypeId"]
return (att_id)
def delete_attribute_set(self, attribute_set_id: str):
resp = self.client.delete_attribute_set(attributeSetId=attribute_set_id)
if resp['ResponseMetadata']['HTTPStatusCode'] != 200:
return resp
return True
def associate_attribute_set(self, att_name: str, att_values: list, dataset_id: str):
# get the attribute set by name, will need its id
att_set = self.attribute_set(att_name)
# get the dataset's information, will need the arn
dataset = self.describe_dataset_details(dataset_id=dataset_id)
# disassociate any existing relationship
try:
self.client.dissociate_dataset_from_dataset_type(datasetArn=dataset['arn'],
datasetTypeId=att_set['id'])
except:
print("Nothing to disassociate")
self.client.associate_dataset_with_dataset_type(datasetArn=dataset['arn'], datasetTypeId=att_set['id'])
ret = self.client.update_dataset_type_context(datasetArn=dataset['arn'], datasetTypeId=att_set['id'],
values=att_values)
return ret
# --------------------------------------
# Permission Group Functions
# --------------------------------------
def list_permission_groups(self, max_results: int):
all_perms = self.client.list_permission_groups(MaxResults=max_results)
return (self.get_list(all_perms, 'permissionGroups'))
def permission_group(self, name):
all_groups = self.list_permission_groups(max_results = 100)
existing_group = next((c for c in all_groups if c['name'].lower() == name.lower()), None)
if existing_group:
return existing_group
def describe_permission_group(self, permission_group_id: str):
resp = None
perm_resp = self.client.describe_permission_group(permissionGroupId=permission_group_id)
if 'permissionGroup' in perm_resp:
resp = perm_resp['permissionGroup']
return (resp)
# --------------------------------------
# Dataset Functions
# --------------------------------------
def describe_dataset_details(self, dataset_id: str):
"""
Calls the describe dataset details API function and only returns the dataset details portion of the response.
:param dataset_id: the GUID of the dataset to get description of
:type: str
"""
resp = None
dataset_details_resp = self.client.describe_dataset_details(datasetId=dataset_id)
if 'dataset' in dataset_details_resp:
resp = dataset_details_resp["dataset"]
return (resp)
def create_dataset(self, name: str, description: str, permission_group_id: str, dataset_permissions: [], kind: str,
owner_info, schema):
"""
Create a dataset
Warning, dataset names are not unique, be sure to check for the same name dataset before creating a new one
:param name: Name of the dataset
:type: str
:param description: Description of the dataset
:type: str
:param permission_group_id: permission group for the dataset
:type: str
:param dataset_permissions: permissions for the group on the dataset
:param kind: Kind of dataset, choices: TABULAR
:type: str
:param owner_info: owner information for the dataset
:param schema: Schema of the dataset
:return: the dataset_id of the created dataset
"""
if dataset_permissions:
request_dataset_permissions = [{"permission": permissionName} for permissionName in dataset_permissions]
else:
request_dataset_permissions = []
response = self.client.create_dataset(name=name,
permissionGroupId = permission_group_id,
datasetPermissions = request_dataset_permissions,
kind=kind,
description = description.replace('\n', ' '),
ownerInfo = owner_info,
schema = schema)
return response["datasetId"]
def ingest_from_s3(self,
s3_location: str,
dataset_id: str,
change_type: str,
wait_for_completion: bool = True,
format_type: str = "CSV",
format_params: dict = {'separator': ',', 'withHeader': 'true'}):
"""
Creates a changeset and ingests the data given in the S3 location into the changeset
:param s3_location: the source location of the data for the changeset, will be copied into the changeset
:stype: str
:param dataset_id: the identifier of the containing dataset for the changeset to be created for this data
:type: str
:param change_type: What is the kind of changetype? "APPEND", "REPLACE" are the choices
:type: str
:param wait_for_completion: Boolean, should the function wait for the operation to complete?
:type: str
:param format_type: format type, CSV, PARQUET, XML, JSON
:type: str
:param format_params: dictionary of format parameters
:type: dict
:return: the id of the changeset created
"""
create_changeset_response = self.client.create_changeset(
datasetId=dataset_id,
changeType=change_type,
sourceType='S3',
sourceParams={'s3SourcePath': s3_location},
formatType=format_type.upper(),
formatParams=format_params
)
changeset_id = create_changeset_response['changeset']['id']
if wait_for_completion:
self.wait_for_ingestion(dataset_id, changeset_id)
return changeset_id
def describe_changeset(self, dataset_id: str, changeset_id: str):
"""
Function to get a description of the the givn changeset for the given dataset
:param dataset_id: identifier of the dataset
:type: str
:param changeset_id: the idenfitier of the changeset
:type: str
:return: all information about the changeset, if found
"""
describe_changeset_resp = self.client.describe_changeset(datasetId=dataset_id, id=changeset_id)
return describe_changeset_resp['changeset']
def create_as_of_view(self, dataset_id: str, as_of_date: datetime, destination_type: str,
partition_columns: list = [], sort_columns: list = [], destination_properties: dict = {},
wait_for_completion: bool = True):
"""
Creates an 'as of' static view up to and including the requested 'as of' date provided.
:param dataset_id: identifier of the dataset
:type: str
:param as_of_date: as of date, will include changesets up to this date/time in the view
:type: datetime
:param destination_type: destination type
:type: str
:param partition_columns: columns to partition the data by for the created view
:type: list
:param sort_columns: column to sort the view by
:type: list
:param destination_properties: destination properties
:type: dict
:param wait_for_completion: should the function wait for the system to create the view?
:type: bool
:return str: GUID of the created view if successful
"""
create_materialized_view_resp = self.client.create_materialized_snapshot(
datasetId=dataset_id,
asOfTimestamp=as_of_date,
destinationType=destination_type,
partitionColumns=partition_columns,
sortColumns=sort_columns,
autoUpdate=False,
destinationProperties=destination_properties
)
view_id = create_materialized_view_resp['id']
if wait_for_completion:
self.wait_for_view(dataset_id=dataset_id, view_id=view_id)
return view_id
def create_auto_update_view(self, dataset_id: str, destination_type: str,
partition_columns=[], sort_columns=[], destination_properties={},
wait_for_completion=True):
"""
Creates an auto-updating view of the given dataset
:param dataset_id: identifier of the dataset
:type: str
:param destination_type: destination type
:type: str
:param partition_columns: columns to partition the data by for the created view
:type: list
:param sort_columns: column to sort the view by
:type: list
:param destination_properties: destination properties
:type: str
:param wait_for_completion: should the function wait for the system to create the view?
:type: bool
:return str: GUID of the created view if successful
"""
create_materialized_view_resp = self.client.create_materialized_snapshot(
datasetId=dataset_id,
destinationType=destination_type,
partitionColumns=partition_columns,
sortColumns=sort_columns,
autoUpdate=True,
destinationProperties=destination_properties
)
view_id = create_materialized_view_resp['id']
if wait_for_completion:
self.wait_for_view(dataset_id=dataset_id, view_id=view_id)
return view_id
def wait_for_ingestion(self, dataset_id: str, changeset_id: str, sleep_sec=10):
"""
function that will continuously poll the changeset creation to ensure it completes or fails before returning.
:param dataset_id: GUID of the dataset
:type: str
:param changeset_id: GUID of the changeset
:type: str
:param sleep_sec: seconds to wait between checks
:type: int
"""
while True:
status = self.describe_changeset(dataset_id=dataset_id, changeset_id=changeset_id)['status']
if status == 'SUCCESS':
print(f"Changeset complete")
break
elif status == 'PENDING' or status == 'RUNNING':
print(f"Changeset status is still PENDING, waiting {sleep_sec} sec ...")
time.sleep(sleep_sec)
continue
else:
raise Exception(f"Bad changeset status: {status}, failing now.")
def wait_for_view(self, dataset_id: str, view_id: str, sleep_sec=10):
"""
function that will continuously poll the view creation to ensure it completes or fails before returning.
:param dataset_id: GUID of the dataset
:type: str
:param view_id: GUID of the view
:type: str
:param sleep_sec: seconds to wait between checks
:type: int
"""
while True:
list_views_resp = self.client.list_materialization_snapshots(datasetId=dataset_id, maxResults=100)
matched_views = list(filter(lambda d: d['id'] == view_id, list_views_resp['materializationSnapshots']))
if len(matched_views) != 1:
size = len(matched_views)
raise Exception(f"Unexpected error: found {size} views that match the view Id: {view_id}")
status = matched_views[0]['status']
if status == 'SUCCESS':
print(f"View complete")
break
elif status == 'PENDING' or status == 'RUNNING':
print(f"View status is still PENDING, continue to wait till finish...")
time.sleep(sleep_sec)
continue
else:
raise Exception(f"Bad view status: {status}, failing now.")
def list_changesets(self, dataset_id: str):
resp = self.client.list_changesets(datasetId=dataset_id, sortKey='CREATE_TIMESTAMP')
results = resp['changesets']
while "nextToken" in resp:
resp = self.client.list_changesets(datasetId=dataset_id, sortKey='CREATE_TIMESTAMP',
nextToken=resp['nextToken'])
results.extend(resp['changesets'])
return (results)
def list_views(self, dataset_id: str, max_results=50):
resp = self.client.list_materialization_snapshots(datasetId=dataset_id, maxResults=max_results)
results = resp['materializationSnapshots']
while "nextToken" in resp:
resp = self.client.list_materialization_snapshots(datasetId=dataset_id, maxResults=max_results,
nextToken=resp['nextToken'])
results.extend(resp['materializationSnapshots'])
return (results)
def list_datasets(self, max_results: int):
all_datasets = self.client.list_datasets(maxResults=max_results)
return (self.get_list(all_datasets, 'datasets'))
def list_dataset_types(self):
resp = self.client.list_dataset_types(sort='NAME')
results = resp['datasetTypeSummaries']
while "nextToken" in resp:
resp = self.client.list_dataset_types(sort='NAME', nextToken=resp['nextToken'])
results.extend(resp['datasetTypeSummaries'])
return (results)
@staticmethod
def get_execution_role():
"""
Convenience function from SageMaker to get the execution role of the user of the sagemaker studio notebook
:return: the ARN of the execution role in the sagemaker studio notebook
"""
import sagemaker as sm
e_role = sm.get_execution_role()
return (f"{e_role}")
def get_user_ingestion_info(self):
return (self.client.get_user_ingestion_info())
def upload_pandas(self, data_frame: pd.DataFrame):
import awswrangler as wr
resp = self.client.get_working_location(locationType='INGESTION')
upload_location = resp['s3Uri']
wr.s3.to_parquet(data_frame, f"{upload_location}data.parquet", index=False, boto3_session=self._boto3_session)
return upload_location
def ingest_pandas(self, data_frame: pd.DataFrame, dataset_id: str, change_type: str, wait_for_completion=True):
print("Uploading the pandas dataframe ...")
upload_location = self.upload_pandas(data_frame)
print("Data upload finished. Ingesting data ...")
return self.ingest_from_s3(upload_location, dataset_id, change_type, wait_for_completion, format_type='PARQUET')
def read_view_as_pandas(self, dataset_id: str, view_id: str):
"""
Returns a pandas dataframe the view of the given dataset. Views in FinSpace can be quite large, be careful!
:param dataset_id:
:param view_id:
:return: Pandas dataframe with all data of the view
"""
import awswrangler as wr # use awswrangler to read the table
# @todo: switch to DescribeMateriliazation when available in HFS
views = self.list_views(dataset_id=dataset_id, max_results=50)
filtered = [v for v in views if v['id'] == view_id]
if len(filtered) == 0:
raise Exception('No such view found')
if len(filtered) > 1:
raise Exception('Internal Server error')
view = filtered[0]
# 0. Ensure view is ready to be read
if (view['status'] != 'SUCCESS'):
status = view['status']
print(f'view run status is not ready: {status}. Returning empty.')
return
glue_db_name = view['destinationTypeProperties']['databaseName']
glue_table_name = view['destinationTypeProperties']['tableName']
# determine if the table has partitions first, different way to read is there are partitions
p = wr.catalog.get_partitions(table=glue_table_name, database=glue_db_name, boto3_session=self._boto3_session)
def no_filter(partitions):
if len(partitions.keys()) > 0:
return True
return False
df = None
if len(p) == 0:
df = wr.s3.read_parquet_table(table=glue_table_name, database=glue_db_name,
boto3_session=self._boto3_session)
else:
spath = wr.catalog.get_table_location(table=glue_table_name, database=glue_db_name,
boto3_session=self._boto3_session)
cpath = wr.s3.list_directories(f"{spath}/*", boto3_session=self._boto3_session)
read_path = f"{spath}/"
# just one? Read it
if len(cpath) == 1:
read_path = cpath[0]
df = wr.s3.read_parquet(read_path, dataset=True, partition_filter=no_filter,
boto3_session=self._boto3_session)
# Query Glue table directly with wrangler
return df
@staticmethod
def get_schema_from_pandas(df: pd.DataFrame):
"""
Returns the FinSpace schema columns from the given pandas dataframe.
:param df: pandas dataframe to interrogate for the schema
:return: FinSpace column schema list
"""
# for translation to FinSpace's schema
# 'STRING'|'CHAR'|'INTEGER'|'TINYINT'|'SMALLINT'|'BIGINT'|'FLOAT'|'DOUBLE'|'DATE'|'DATETIME'|'BOOLEAN'|'BINARY'
DoubleType = "DOUBLE"
FloatType = "FLOAT"
DateType = "DATE"
StringType = "STRING"
IntegerType = "INTEGER"
LongType = "BIGINT"
BooleanType = "BOOLEAN"
TimestampType = "DATETIME"
hab_columns = []
for name in dict(df.dtypes):
p_type = df.dtypes[name]
switcher = {
"float64": DoubleType,
"int64": IntegerType,
"datetime64[ns, UTC]": TimestampType,
"datetime64[ns]": DateType
}
habType = switcher.get(str(p_type), StringType)
hab_columns.append({
"dataType": habType,
"name": name,
"description": ""
})
return (hab_columns)
@staticmethod
def get_date_cols(df: pd.DataFrame):
"""
Returns which are the data columns found in the pandas dataframe.
Pandas does the hard work to figure out which of the columns can be considered to be date columns.
:param df: pandas dataframe to interrogate for the schema
:return: list of column names that can be parsed as dates by pandas
"""
date_cols = []
for name in dict(df.dtypes):
p_type = df.dtypes[name]
if str(p_type).startswith("date"):
date_cols.append(name)
return (date_cols)
def get_best_schema_from_csv(self, path, is_s3=True, read_rows=500, sep=','):
"""
Uses multiple reads of the file with pandas to determine schema of the referenced files.
Files are expected to be csv.
:param path: path to the files to read
:type: str
:param is_s3: True if the path is s3; False if filesystem
:type: bool
:param read_rows: number of rows to sample for determining schema
:param sep:
:return dict: schema for FinSpace
"""
#
# best efforts to determine the schema, sight unseen
import awswrangler as wr
# 1: get the base schema
df1 = None
if is_s3:
df1 = wr.s3.read_csv(path, nrows=read_rows, sep=sep)
else:
df1 = pd.read_csv(path, nrows=read_rows, sep=sep)
num_cols = len(df1.columns)
# with number of columns, try to infer dates
df2 = None
if is_s3:
df2 = wr.s3.read_csv(path, parse_dates=list(range(0, num_cols)), infer_datetime_format=True,
nrows=read_rows, sep=sep)
else:
df2 = pd.read_csv(path, parse_dates=list(range(0, num_cols)), infer_datetime_format=True, nrows=read_rows,
sep=sep)
date_cols = self.get_date_cols(df2)
# with dates known, parse the file fully
df = None
if is_s3:
df = wr.s3.read_csv(path, parse_dates=date_cols, infer_datetime_format=True, nrows=read_rows, sep=sep)
else:
df = pd.read_csv(path, parse_dates=date_cols, infer_datetime_format=True, nrows=read_rows, sep=sep)
schema_cols = self.get_schema_from_pandas(df)
return (schema_cols)
def s3_upload_file(self, source_file: str, s3_destination: str):
"""
Uploads a local file (full path) to the s3 destination given (expected form: s3://<bucket>/<prefix>/).
The filename will have spaces replaced with _.
:param source_file: path of file to upload
:param s3_destination: full path to where to save the file
:type: str
"""
hab_s3_client = self._boto3_session.client(service_name='s3')
o = urlparse(s3_destination)
bucket = o.netloc
prefix = o.path.lstrip('/')
fname = os.path.basename(source_file)
hab_s3_client.upload_file(source_file, bucket, f"{prefix}{fname.replace(' ', '_')}")
def list_objects(self, s3_location: str):
"""
lists the objects found at the s3_location. Strips out the boto API response header,
just returns the contents of the location. Internally uses the list_objects_v2.
:param s3_location: path, starting with s3:// to get the list of objects from
:type: str
"""
o = urlparse(s3_location)
bucket = o.netloc
prefix = o.path.lstrip('/')
results = []
hab_s3_client = self._boto3_session.client(service_name='s3')
paginator = hab_s3_client.get_paginator('list_objects_v2')
pages = paginator.paginate(Bucket=bucket, Prefix=prefix)
for page in pages:
if 'Contents' in page:
results.extend(page['Contents'])
return (results)
def list_clusters(self, status: str = None):
"""
Lists current clusters and their statuses
:param status: status to filter for
:return dict: list of clusters
"""
resp = self.client.list_clusters()
clusters = []
if 'clusters' not in resp:
return (clusters)
for c in resp['clusters']:
if status is None:
clusters.append(c)
else:
if c['clusterStatus']['state'] in status:
clusters.append(c)
return (clusters)
def get_cluster(self, cluster_id):
"""
Resize the given cluster to desired template
:param cluster_id: cluster id
"""
clusters = self.list_clusters()
for c in clusters:
if c['clusterId'] == cluster_id:
return (c)
return (None)
def update_cluster(self, cluster_id: str, template: str):
"""
Resize the given cluster to desired template
:param cluster_id: cluster id
:param template: target template to resize to
"""
cluster = self.get_cluster(cluster_id=cluster_id)
if cluster['currentTemplate'] == template:
print(f"Already using template: {template}")
return (cluster)
self.client.update_cluster(clusterId=cluster_id, template=template)
return (self.get_cluster(cluster_id=cluster_id))
def wait_for_status(self, clusterId: str, status: str, sleep_sec=10, max_wait_sec=900):
"""
Function polls service until cluster is in desired status.
:param clusterId: the cluster's ID
:param status: desired status for clsuter to reach
:
"""
total_wait = 0
while True and total_wait < max_wait_sec:
resp = self.client.list_clusters()
this_cluster = None
# is this the cluster?
for c in resp['clusters']:
if clusterId == c['clusterId']:
this_cluster = c
if this_cluster is None:
print(f"clusterId:{clusterId} not found")
return (None)
this_status = this_cluster['clusterStatus']['state']
if this_status.upper() != status.upper():
print(f"Cluster status is {this_status}, waiting {sleep_sec} sec ...")
time.sleep(sleep_sec)
total_wait = total_wait + sleep_sec
continue
else:
return (this_cluster)
def get_working_location(self, locationType='SAGEMAKER'):
resp = None
location = self.client.get_working_location(locationType=locationType)
if 's3Uri' in location:
resp = location['s3Uri']
return (resp)
# %load finspace_spark.py
import datetime
import time
import boto3
from botocore.config import Config
# FinSpace class with Spark bindings
class SparkFinSpace(FinSpace):
import pyspark
def __init__(
self,
spark: pyspark.sql.session.SparkSession = None,
config = Config(retries = {'max_attempts': 0, 'mode': 'standard'}),
dev_overrides: dict = None
):
FinSpace.__init__(self, config=config, dev_overrides=dev_overrides)
self.spark = spark # used on Spark cluster for reading views, creating changesets from DataFrames
def upload_dataframe(self, data_frame: pyspark.sql.dataframe.DataFrame):
resp = self.client.get_user_ingestion_info()
upload_location = resp['ingestionPath']
# data_frame.write.option('header', 'true').csv(upload_location)
data_frame.write.parquet(upload_location)
return upload_location
def ingest_dataframe(self, data_frame: pyspark.sql.dataframe.DataFrame, dataset_id: str, change_type: str, wait_for_completion=True):
print("Uploading data...")
upload_location = self.upload_dataframe(data_frame)
print("Data upload finished. Ingesting data...")
return self.ingest_from_s3(upload_location, dataset_id, change_type, wait_for_completion, format_type='parquet', format_params={})
def read_view_as_spark(
self,
dataset_id: str,
view_id: str
):
# TODO: switch to DescribeMatz when available in HFS
views = self.list_views(dataset_id=dataset_id, max_results=50)
filtered = [v for v in views if v['id'] == view_id]
if len(filtered) == 0:
raise Exception('No such view found')
if len(filtered) > 1:
raise Exception('Internal Server error')
view = filtered[0]
# 0. Ensure view is ready to be read
if (view['status'] != 'SUCCESS'):
status = view['status']
print(f'view run status is not ready: {status}. Returning empty.')
return
glue_db_name = view['destinationTypeProperties']['databaseName']
glue_table_name = view['destinationTypeProperties']['tableName']
# Query Glue table directly with catalog function of spark
return self.spark.table(f"`{glue_db_name}`.`{glue_table_name}`")
def get_schema_from_spark(self, data_frame: pyspark.sql.dataframe.DataFrame):
from pyspark.sql.types import StructType
# for translation to FinSpace's schema
# 'STRING'|'CHAR'|'INTEGER'|'TINYINT'|'SMALLINT'|'BIGINT'|'FLOAT'|'DOUBLE'|'DATE'|'DATETIME'|'BOOLEAN'|'BINARY'
DoubleType = "DOUBLE"
FloatType = "FLOAT"
DateType = "DATE"
StringType = "STRING"
IntegerType = "INTEGER"
LongType = "BIGINT"
BooleanType = "BOOLEAN"
TimestampType = "DATETIME"
hab_columns = []
items = [i for i in data_frame.schema]
switcher = {
"BinaryType" : StringType,
"BooleanType" : BooleanType,
"ByteType" : IntegerType,
"DateType" : DateType,
"DoubleType" : FloatType,
"IntegerType" : IntegerType,
"LongType" : IntegerType,
"NullType" : StringType,
"ShortType" : IntegerType,
"StringType" : StringType,
"TimestampType" : TimestampType,
}
for i in items:
# print( f"name: {i.name} type: {i.dataType}" )
habType = switcher.get( str(i.dataType), StringType)
hab_columns.append({
"dataType" : habType,
"name" : i.name,
"description" : ""
})
return( hab_columns )
# initialize the FinSpace helper
finspace = SparkFinSpace(spark=spark)
```
# Realized Volatility
This notebook will pull summarized data from FinSpace's catalog and then use the analytic function realized_volatility to compute realized volatility for a group of tickers and exchange event types.
```
#####----------------------------------------------------------
##### REPLACE WITH CORRECT IDS!
##### Dataset: "US Equity Time-Bar Summary - 1 min, 14 Symbols - Sample"
#####
#####----------------------------------------------------------
dataset_id = ''
view_id = ''
# import needed libraries
import pandas as pd
import matplotlib.pyplot as plt
import datetime as dt
import pyspark.sql.functions as F
import pyspark.sql.types as T
# import time series libraries
from aws.finspace.timeseries.spark.analytics import *
from aws.finspace.timeseries.spark.windows import *
from aws.finspace.timeseries.spark.util import string_to_timestamp_micros
sumDF = finspace.read_view_as_spark(dataset_id = dataset_id, view_id = view_id)
# Filter and select
sDate = dt.datetime(2020, 1, 1)
eDate = dt.datetime(2020, 3, 1)
df = ( sumDF.filter( sumDF.date.between(sDate, eDate) ) )
# sample the data
df.show(5, truncate=False)
```
# Spark Analytics
All our analytic functions have help, lets look at the signatures for the functions we will use

```
help(realized_volatility)
```
# Calculate Realized Volatility
Compute the realized volatility of the time series data.
```
tenor = 15
numStd = 2
# analytics to calculate
realVolDef = realized_volatility( tenor, "end", "vwap" )
# group the sets of values
partitionList = ["ticker", "eventtype"]
tsDF = df
tsDF = compute_analytics_on_features(tsDF, "realized_volatility", realVolDef, partition_col_list = partitionList).cache()
tsDF.printSchema()
```
# Realized Volatility Graph
Plot realized volatility
1. Calculations are performed on the cluster
2. Results are then collected to the driver as a pandas DataFrame
3. Plot image created from pandas data
4. Plot image is sent to the 'local' notebook for display
This is all done for you.
```
fTicker = 'AMZN'
# filter and bring data into a pandas dataframe for plotting
pltDF = ( tsDF
.filter(tsDF.eventtype == "TRADE NB")
.filter(tsDF.ticker == fTicker)
.select( 'end', 'realized_volatility' )
).toPandas().set_index('end')
pltDF.index = pltDF.index.strftime("%Y-%m-%d %H:%m")
fig, ax = plt.subplots(1, 1, figsize=(12, 6))
# Realized Volatility
pltDF[[ 'realized_volatility' ]].plot(figsize=(12,6))
# labels and other items to make the plot readable
plt.title(f"{fTicker} Realized Vol (tenor: {tenor}, 1 min bars)")
plt.ylabel('Realized Vol')
plt.xlabel('Date/Time')
plt.xticks(rotation=30)
plt.subplots_adjust(bottom=0.2)
%matplot plt
```
# So why that spike?
[Amazon soars after huge earnings beat](https://www.cnbc.com/2020/01/30/amazon-amzn-q4-2019-earnings.html) (CNBC).
- Amazon reported fourth-quarter results on Thursday that smashed analysts’ expectations.
- The company’s profits rebounded during the quarter, while revenue climbed 21% year over year.
- The outperforming results show Amazon’s big investments in one-day delivery are paying off.
# Incorporate Events from Factset
Factset has the event data: [Factset Events](https://app.snowflake.com/marketplace/listing/GZT0Z28ANXM)
```
# attribute name that contains the necessary catalog, schema and table data to get data from Snowflake.
snowflake_att_name = 'Snowflake Table Attributes'
#####----------------------------------------------------------
##### REPLACE WITH CORRECT IDS!
#####----------------------------------------------------------
# dataset IDs of the data as registered in the FinSpace catalog
sym_ticker_region = ''
sym_coverage = ''
sym_entity = ''
ce_events = ''
ce_reports = ''
ce_sec_entity_hist = ''
```
## Utility functions
Use the Snowflake Metadata in Finspace to query for the data in snowflake, creating a Spark dataframe based on the snowflake table.
```
def find_by_key_value(l, key, v):
for n in l:
if n[key] == v:
return n
def get_field_by_name(f_list, title, name = 'name'):
f = find_by_key_value(f_list, 'title', title)
if f is not None:
return f[name]
return None
def get_field_values(f_list, field):
for f in f_list:
if f['field'] == field:
return f['values']
def get_snowflake_query(dataset_id, att_name):
sfAttrSet = finspace.attribute_set(att_name)
if sfAttrSet is None:
print(f'Did not find the attribute set with name: {att_name}')
return None
# get the dataset details
dataset_details = finspace.client.describe_dataset_details(datasetId = dataset_id)
if dataset_details is None:
print(f'Did not find the dataset with id: {dataset_id}')
return None
# find the snowflake attributes related to the dataset
attributes = dataset_details['datasetTypeContexts']
sfAttr = find_by_key_value(attributes, 'id', sfAttrSet['id'])
if sfAttr is None:
print(f'Did not find the attribute set with name: {att_name} in the dataset with id: {dataset_id}')
return None
field_defs = sfAttr['definition']['fields']
catalog = get_field_by_name(field_defs, 'Catalog')
schema = get_field_by_name(field_defs, 'Schema')
table = get_field_by_name(field_defs, 'Table')
field_values = sfAttr['values']
return f'"{get_field_values(field_values, catalog)[0]}"."{get_field_values(field_values, schema)[0]}"."{get_field_values(field_values, table)[0]}"'
def get_dataframe_from_snowflake(dataset_id, att_name='Snowflake Table Attributes'):
# get the query for snowflake from the data in the attributeset
sf_query = get_snowflake_query(dataset_id = dataset_id, att_name = att_name)
# load the dataframe from snowflake using information from finspace
df = ( spark.read.format(SNOWFLAKE_SOURCE_NAME)
.options(**sfOptions)
.option('query', sf_query)
.load()
)
return df
```
## Create DataFrames from Snowflake Tables
These functions create Spark DataFrames from Snowflake tables, using the information about their location that was registered in FinSpace's catalog.
```
# symbols
sym_ticker_region_df = get_dataframe_from_snowflake(dataset_id = sym_ticker_region, att_name = snowflake_att_name)
sym_coverage_df = get_dataframe_from_snowflake(dataset_id = sym_coverage, att_name = snowflake_att_name)
sym_entity_df = get_dataframe_from_snowflake(dataset_id = sym_entity, att_name = snowflake_att_name)
# events
ce_sec_entity_hist_df = get_dataframe_from_snowflake(dataset_id = ce_sec_entity_hist, att_name = snowflake_att_name)
ce_reports_df = get_dataframe_from_snowflake(dataset_id = ce_reports, att_name = snowflake_att_name)
ce_events_df = get_dataframe_from_snowflake(dataset_id = ce_events, att_name = snowflake_att_name)
```
## Now the Factset data to the dataset with volatility
Spark dataframe operations to join the data on necessary keys
```
# first the symbol data....
symbol_df = ( sym_ticker_region_df
.join(sym_coverage_df, ['FSYM_ID'] ) # sym_ticker_region_df.FSYM_ID == sym_coverage_df.FSYM_ID)
.withColumnRenamed('FSYM_ID', 'FSYM_ID_DELETE')
.join(ce_sec_entity_hist_df, ce_sec_entity_hist_df.FSYM_ID == sym_coverage_df.FSYM_SECURITY_ID)
.join(sym_entity_df, ['FACTSET_ENTITY_ID']) #sym_entity_df.FACTSET_ENTITY_ID == ce_sec_entity_hist_df.FACTSET_ENTITY_ID)
).drop('FSYM_ID_DELETE')
# Data about Amazon
ticker = 'AMZN'
region = 'US'
symbol_df.filter(symbol_df.TICKER_REGION == f'{ticker}-{region}' ).show(truncate=False)
# Now pull in the events as well
events_df = ( symbol_df
.join(ce_reports_df, ['FACTSET_ENTITY_ID']) #ce_reports_df.FACTSET_ENTITY_ID == ce_sec_entity_hist_df.FACTSET_ENTITY_ID)
.join(ce_events_df, ['EVENT_ID']) #ce_events_df.EVENT_ID == ce_reports_df.EVENT_ID)
)
# what where the events over that same period?
print(f'{ticker}: {sDate} to {eDate}')
e_df = ( events_df
.filter( events_df.TICKER_REGION == f'{ticker}-{region}' )
.filter( events_df.EVENT_DATETIME_UTC.between(sDate, eDate) )
.orderBy( events_df.ENTITY_PROPER_NAME, events_df.EVENT_DATETIME_UTC.asc() )
)
e_df.select(e_df.TICKER_REGION, e_df.FSYM_ID, e_df.FACTSET_ENTITY_ID, e_df.ENTITY_PROPER_NAME, e_df.EVENT_DATETIME_UTC, e_df.EVENT_TYPE, e_df.TITLE).show(truncate=False)
```
# Plot with Events
Adding the events as labeled veritcal lines to the original plot.
```
fTicker = 'AMZN'
fRegion = 'US'
max_str_len = 30
# filter and bring data into a pandas dataframe for plotting
tradeDF = ( tsDF
.filter(tsDF.eventtype == "TRADE NB")
.filter(tsDF.ticker == fTicker)
.select( 'end', 'realized_volatility' )
).toPandas().set_index('end')
evtDF = ( events_df
.filter( events_df.TICKER_REGION == f'{fTicker}-{fRegion}' )
.filter(events_df.EVENT_DATETIME_UTC.between(sDate, eDate) )
.orderBy( events_df.ENTITY_PROPER_NAME, events_df.EVENT_DATETIME_UTC.asc() )
.select( ['EVENT_DATETIME_UTC', 'TICKER_REGION', 'ENTITY_PROPER_NAME', 'EVENT_TYPE', 'TITLE'] )
).toPandas().set_index('EVENT_DATETIME_UTC')
# dataframe for plotting, concat the two
pltDF = pd.concat( [tradeDF, evtDF], axis=0).sort_index()
# index as string
pltDF.index = pltDF.index.strftime("%Y-%m-%d %H:%m")
# for placing the labels on events
y_min = pltDF.realized_volatility.min()
y_max = pltDF.realized_volatility.max()
mid = (y_max - y_min) / 2
step = 0.1*mid
# the plot
fig, ax = plt.subplots(1, 1, figsize=(12, 6))
# Realized Volatility
pltDF[[ 'realized_volatility' ]].plot(figsize=(12,6), legend=None)
# labels and other items to make the plot readable
plt.title(f"{fTicker} Realized Vol (tenor: {tenor}, 1 min bars)")
plt.ylabel('Realized Vol')
plt.xlabel('Date/Time')
plt.xticks(rotation=30)
plt.subplots_adjust(bottom=0.2)
# get the locations of the events and plot vertical lines
row_indexes = pltDF.index.tolist()
events = pltDF.loc[pltDF.EVENT_TYPE.notnull()]
s = 2
for d in events.index:
idx = row_indexes.index(d)
plt.axvline(idx, linewidth=1, color='r', alpha=0.2)
e = events.loc[d]
t = f'({e.EVENT_TYPE}) {e.TITLE}'
t = (t[:max_str_len] + '..') if len(t) > max_str_len else t
y = y_max - (step * s)
s = s+1
plt.text(idx*1.01, y, t, rotation=0)
#
%matplot plt
```
# Save the Data to Snowflake
Lets save the data we created for the plot into Snowflake
```
# SOURCE DATAFRAME
# ---------------------------------------------
print('Source: Volatility DataFrame')
print(f'Rows: {tsDF.count():,}')
tsDF.show(5, False)
```
### WRITE to Snowflake
```
# INTO: Snowflake Target
# ---------------------------------------------
sfDatabaseName = snowflake_database
sfTableName = f'volatility_from_finspace'
print(f'Saving to Snowflake: {sfDatabaseName}..{sfTableName}')
( tsDF.write
.format(SNOWFLAKE_SOURCE_NAME)
.options(**sfOptions)
.option("dbtable", f"{sfDatabaseName}..{sfTableName}")
.mode('overwrite')
.save()
)
```
### READ AGAIN from Snowflake
```
# READ table from Snowflake again
# ---------------------------------------------
tsDF_SF = ( spark.read.format(SNOWFLAKE_SOURCE_NAME)
.options(**sfOptions)
.option("query", f"{sfDatabaseName}..{sfTableName}")
.load() )
print(f'Read from SNOWFLAKE: {sfDatabaseName}..{sfTableName}')
print(f'Rows: {tsDF_SF.count():,}')
tsDF_SF.show(5, False)
```
# Snapshot the Data
Take advantage of FinSpace's changsets and views to create a snapshot of snowflake table's data.
```
#####----------------------------------------------------------
##### REPLACE WITH CORRECT IDS!
#####----------------------------------------------------------
# EMPLOYEES Dataset in FinSpace
dataset_id = ''
# get the data from snowflake
df = get_dataframe_from_snowflake(dataset_id = dataset_id, att_name = snowflake_att_name)
# then add the changeset, to the dataset
changeset_id = finspace.ingest_dataframe(data_frame=df, dataset_id = dataset_id, change_type='REPLACE', wait_for_completion=True)
# Create an auto-updating View if one does not exist
existing_snapshots = finspace.list_views(dataset_id = dataset_id, max_results=100)
autoupdate_snapshot_id = None
# does one exist?
for ss in existing_snapshots:
if ss['autoUpdate'] == True:
autoupdate_snapshot_id = ss['id']
# if no auto-updating view, create it
if autoupdate_snapshot_id is None:
autoupdate_snapshot_id = finspace.create_auto_update_view(
dataset_id = dataset_id,
destination_type = "GLUE_TABLE",
partition_columns = [],
sort_columns = [],
wait_for_completion = False)
print( f"Created autoupdate_snapshot_id = {autoupdate_snapshot_id}" )
else:
print( f"Exists: autoupdate_snapshot_id = {autoupdate_snapshot_id}" )
```
## Employees Dataset
The dataset in FinSpace
[EMPLOYEES Dataset](https://ak6gkzahz2s26bwtlnx5p6.us-east-1.amazonfinspace.com/dataset/jeyitu0/data)
| github_jupyter |
```
import numpy as np
import gym
import sys
import pandas as pd
import itertools
import matplotlib.pyplot as plt
if "../" not in sys.path:
sys.path.append("../")
from lib.envs import plotting
from collections import defaultdict
from gym.envs.toy_text import discrete
plt.style.use('ggplot')
from collections import namedtuple
from matplotlib import pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
EpisodeStats = namedtuple("Stats",["episode_lengths", "episode_rewards"])
def plot_cost_to_go_mountain_car(env, estimator, num_tiles=20):
x = np.linspace(env.observation_space.low[0], env.observation_space.high[0], num=num_tiles)
y = np.linspace(env.observation_space.low[1], env.observation_space.high[1], num=num_tiles)
X, Y = np.meshgrid(x, y)
Z = np.apply_along_axis(lambda _: -np.max(estimator.predict(_)), 2, np.dstack([X, Y]))
fig = plt.figure(figsize=(10, 5))
ax = fig.add_subplot(111, projection='3d')
surf = ax.plot_surface(X, Y, Z, rstride=1, cstride=1,
cmap=matplotlib.cm.coolwarm, vmin=-1.0, vmax=1.0)
ax.set_xlabel('Position')
ax.set_ylabel('Velocity')
ax.set_zlabel('Value')
ax.set_title("Mountain \"Cost To Go\" Function")
fig.colorbar(surf)
plt.show()
def plot_value_function(V, title="Value Function"):
"""
Plots the value function as a surface plot.
"""
min_x = min(k[0] for k in V.keys())
max_x = max(k[0] for k in V.keys())
min_y = min(k[1] for k in V.keys())
max_y = max(k[1] for k in V.keys())
x_range = np.arange(min_x, max_x + 1)
y_range = np.arange(min_y, max_y + 1)
X, Y = np.meshgrid(x_range, y_range)
# Find value for all (x, y) coordinates
Z_noace = np.apply_along_axis(lambda _: V[(_[0], _[1], False)], 2, np.dstack([X, Y]))
Z_ace = np.apply_along_axis(lambda _: V[(_[0], _[1], True)], 2, np.dstack([X, Y]))
def plot_surface(X, Y, Z, title):
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111, projection='3d')
surf = ax.plot_surface(X, Y, Z, rstride=1, cstride=1,
cmap=matplotlib.cm.coolwarm, vmin=-1.0, vmax=1.0)
ax.set_xlabel('Player Sum')
ax.set_ylabel('Dealer Showing')
ax.set_zlabel('Value')
ax.set_title(title)
ax.view_init(ax.elev, -120)
fig.colorbar(surf)
plt.show()
plot_surface(X, Y, Z_noace, "{} (No Usable Ace)".format(title))
plot_surface(X, Y, Z_ace, "{} (Usable Ace)".format(title))
def plot_episode_stats(stats, smoothing_window=10, noshow=False):
# Plot the episode length over time
fig1 = plt.figure(figsize=(10,5))
plt.plot(stats.episode_lengths)
plt.xlabel("Episode")
plt.ylabel("Episode Length")
plt.title("Episode Length over Time")
if noshow:
plt.close(fig1)
else:
plt.show(fig1)
# Plot the episode reward over time
fig2 = plt.figure(figsize=(10,5))
rewards_smoothed = pd.Series(stats.episode_rewards).rolling(smoothing_window, min_periods=smoothing_window).mean()
plt.plot(rewards_smoothed)
plt.xlabel("Episode")
plt.ylabel("Episode Reward (Smoothed)")
plt.title("Episode Reward over Time (Smoothed over window size {})".format(smoothing_window))
if noshow:
plt.close(fig2)
else:
plt.show(fig2)
# Plot time steps and episode number
fig3 = plt.figure(figsize=(10,5))
plt.plot(np.cumsum(stats.episode_lengths), np.arange(len(stats.episode_lengths)))
plt.xlabel("Time Steps")
plt.ylabel("Episode")
plt.title("Episode per time step")
if noshow:
plt.close(fig3)
else:
plt.show(fig3)
return fig1, fig2, fig3
UP = 0
RIGHT = 1
DOWN = 2
LEFT = 3
class WindyGridworldEnv(discrete.DiscreteEnv):
metadata = {'render.modes': ['human', 'ansi']}
def _limit_coordinates(self, coord):
coord[0] = min(coord[0], self.shape[0] - 1)
coord[0] = max(coord[0], 0)
coord[1] = min(coord[1], self.shape[1] - 1)
coord[1] = max(coord[1], 0)
return coord
def _calculate_transition_prob(self, current, delta, winds):
new_position = np.array(current) + np.array(delta) + np.array([-1, 0]) * winds[tuple(current)]
new_position = self._limit_coordinates(new_position).astype(int)
new_state = np.ravel_multi_index(tuple(new_position), self.shape)
is_done = tuple(new_position) == (3, 7)
return [(1.0, new_state, -1.0, is_done)]
def __init__(self):
self.shape = (7, 10)
nS = np.prod(self.shape)
nA = 4
# Wind strength
winds = np.zeros(self.shape)
winds[:,[3,4,5,8]] = 1
winds[:,[6,7]] = 2
# Calculate transition probabilities
P = {}
for s in range(nS):
position = np.unravel_index(s, self.shape)
P[s] = { a : [] for a in range(nA) }
P[s][UP] = self._calculate_transition_prob(position, [-1, 0], winds)
P[s][RIGHT] = self._calculate_transition_prob(position, [0, 1], winds)
P[s][DOWN] = self._calculate_transition_prob(position, [1, 0], winds)
P[s][LEFT] = self._calculate_transition_prob(position, [0, -1], winds)
# We always start in state (3, 0)
isd = np.zeros(nS)
isd[np.ravel_multi_index((3,0), self.shape)] = 1.0
super(WindyGridworldEnv, self).__init__(nS, nA, P, isd)
def render(self, mode='human', close=False):
self._render(mode, close)
def _render(self, mode='human', close=False):
if close:
return
outfile = StringIO() if mode == 'ansi' else sys.stdout
for s in range(self.nS):
position = np.unravel_index(s, self.shape)
# print(self.s)
if self.s == s:
output = " x "
elif position == (3,7):
output = " T "
else:
output = " o "
if position[1] == 0:
output = output.lstrip()
if position[1] == self.shape[1] - 1:
output = output.rstrip()
output += "\n"
outfile.write(output)
outfile.write("\n")
env = WindyGridworldEnv()
print(env.reset())
env.render()
print(env.step(1))
env.render()
print(env.step(1))
env.render()
print(env.step(1))
env.render()
print(env.step(2))
env.render()
print(env.step(1))
env.render()
print(env.step(1))
env.render()
def make_epsilon_greedy_policy(Q, epsilon, nA):
"""
Creates an epsilon-greedy policy based on a given Q-function and epsilon.
Args:
Q: A dictionary that maps from state -> action-values.
Each value is a numpy array of length nA (see below)
epsilon: The probability to select a random action . float between 0 and 1.
nA: Number of actions in the environment.
Returns:
A function that takes the observation as an argument and returns
the probabilities for each action in the form of a numpy array of length nA.
"""
def policy_fn(observation):
A = np.ones(nA, dtype=float) * epsilon / nA
best_action = np.argmax(Q[observation])
A[best_action] += (1.0 - epsilon)
return A
return policy_fn
def sarsa(env, num_episodes, discount_factor=1.0, alpha=0.5, epsilon=0.1):
"""
SARSA algorithm: On-policy TD control. Finds the optimal epsilon-greedy policy.
Args:
env: OpenAI environment.
num_episodes: Number of episodes to run for.
discount_factor: Gamma discount factor.
alpha: TD learning rate.
epsilon: Chance the sample a random action. Float betwen 0 and 1.
Returns:
A tuple (Q, stats).
Q is the optimal action-value function, a dictionary mapping state -> action values.
stats is an EpisodeStats object with two numpy arrays for episode_lengths and episode_rewards.
"""
# The final action-value function.
# A nested dictionary that maps state -> (action -> action-value).
Q = defaultdict(lambda: np.zeros(env.action_space.n))
# Keeps track of useful statistics
stats = plotting.EpisodeStats(
episode_lengths=np.zeros(num_episodes),
episode_rewards=np.zeros(num_episodes))
# The policy we're following
policy = make_epsilon_greedy_policy(Q, epsilon, env.action_space.n)
for i_episode in range(num_episodes):
# Print out which episode we're on, useful for debugging.
if (i_episode + 1) % 100 == 0:
print("\rEpisode {}/{}.".format(i_episode + 1, num_episodes), end="")
sys.stdout.flush()
# Implement this!
return Q, stats
Q, stats = sarsa(env, 200)
```
| github_jupyter |
# Dataset
```
from datasets import load_dataset
import random
import torch
from torch import nn
from torch.utils.data import DataLoader
from tqdm.auto import tqdm
from transformers import AdamW, DistilBertTokenizerFast, DistilBertForSequenceClassification, get_scheduler
tokenizer = DistilBertTokenizerFast.from_pretrained('distilbert-base-uncased')
dataset = load_dataset("civil_comments")
class CivilCommentsDataset(torch.utils.data.Dataset):
"""
Builds split instance of the `civil_comments` dataset: https://huggingface.co/datasets/civil_comments.
"""
def __init__(self, encodings, labels):
self.encodings = encodings
self.labels = labels
def __getitem__(self, idx):
item = {key: torch.tensor(val[idx]) for key, val in self.encodings.items()}
item['labels'] = torch.tensor(self.labels[idx])
return item
def __len__(self):
return len(self.labels)
def build_data_split(split, num_data_points):
print(f"Generating {num_data_points} data points for {split} split...", end="", flush=True)
civil_idx = []
uncivil_idx = []
num_civil = num_data_points / 2
num_uncivil = num_data_points / 2
for i, data in enumerate(dataset[split]):
if data["toxicity"] < 0.5 and num_civil > 0:
civil_idx.append(i)
num_civil -= 1
elif data["toxicity"] > 0.5 and num_uncivil > 0:
uncivil_idx.append(i)
num_uncivil -= 1
if num_civil == 0 and num_uncivil == 0:
break
indexes = civil_idx + uncivil_idx
random.shuffle(indexes)
encodings = tokenizer(dataset[split][indexes]["text"], truncation=True, padding=True)
labels = dataset[split][indexes]["toxicity"]
print("done")
return encodings, labels
encodings, labels = build_data_split("train", 500)
train_dataset = CivilCommentsDataset(encodings, labels)
encodings, labels = build_data_split("validation", 500)
val_dataset = CivilCommentsDataset(encodings, labels)
```
# Model
```
model = DistilBertForSequenceClassification.from_pretrained(
'distilbert-base-uncased',
num_labels=1,
)
model.dropout.p = 0
model.add_module(module=nn.Sigmoid(), name="sigmoid")
for param in model.base_model.parameters():
param.requires_grad = False
train_data_loader = DataLoader(train_dataset, shuffle=True, batch_size=128)
eval_data_loader = DataLoader(val_dataset, batch_size=128)
optimizer = AdamW(model.parameters(), lr=1e-3)
num_epochs = 20
num_training_steps = num_epochs * len(train_data_loader)
lr_scheduler = get_scheduler(
"linear",
optimizer=optimizer,
num_warmup_steps=0,
num_training_steps=num_training_steps
)
device = torch.device("cuda") if torch.cuda.is_available() else torch.device("cpu")
model.to(device)
progress_bar = tqdm(range(num_training_steps))
def eval_mod():
mse_mean = []
acc_mean = []
for batch in eval_data_loader:
batch = {k: v.to(device) for k, v in batch.items()}
with torch.no_grad():
outputs = model(**batch)
labels = batch["labels"]
outputs = outputs.logits
mse_mean.append(torch.mean(torch.square(outputs - labels)))
acc_mean.append(
torch.mean(torch.eq(outputs.transpose(0, 1) > 0.5, labels > 0.5).float())
)
return torch.mean(torch.stack(mse_mean)), torch.mean(torch.stack(acc_mean))
```
# Main Program
```
for epoch in range(num_epochs):
losses = []
for batch in train_data_loader:
batch = {k: v.to(device) for k, v in batch.items()}
outputs = model(**batch)
loss = outputs.loss
losses.append(float(loss.data))
loss.backward()
optimizer.step()
lr_scheduler.step()
optimizer.zero_grad()
progress_bar.update(1)
mse_mean, accuracy_mean = eval_mod()
loss_mean = torch.mean(torch.tensor(losses))
print(f" After epoch {epoch} | Train Loss: {loss_mean:.2f}, Val MSE: {mse_mean:.2f}, Val Accuracy: {accuracy_mean:.2f}")
model.save_pretrained(f"./results/checkpoints/epoch-{epoch}")
model.save_pretrained("./results/final_model")
print("\nProgram complete")
```
| github_jupyter |
# Project 3: Implement SLAM
---
## Project Overview
In this project, you'll implement SLAM for robot that moves and senses in a 2 dimensional, grid world!
SLAM gives us a way to both localize a robot and build up a map of its environment as a robot moves and senses in real-time. This is an active area of research in the fields of robotics and autonomous systems. Since this localization and map-building relies on the visual sensing of landmarks, this is a computer vision problem.
Using what you've learned about robot motion, representations of uncertainty in motion and sensing, and localization techniques, you will be tasked with defining a function, `slam`, which takes in six parameters as input and returns the vector `mu`.
> `mu` contains the (x,y) coordinate locations of the robot as it moves, and the positions of landmarks that it senses in the world
You can implement helper functions as you see fit, but your function must return `mu`. The vector, `mu`, should have (x, y) coordinates interlaced, for example, if there were 2 poses and 2 landmarks, `mu` will look like the following, where `P` is the robot position and `L` the landmark position:
```
mu = matrix([[Px0],
[Py0],
[Px1],
[Py1],
[Lx0],
[Ly0],
[Lx1],
[Ly1]])
```
You can see that `mu` holds the poses first `(x0, y0), (x1, y1), ...,` then the landmark locations at the end of the matrix; we consider a `nx1` matrix to be a vector.
## Generating an environment
In a real SLAM problem, you may be given a map that contains information about landmark locations, and in this example, we will make our own data using the `make_data` function, which generates a world grid with landmarks in it and then generates data by placing a robot in that world and moving and sensing over some numer of time steps. The `make_data` function relies on a correct implementation of robot move/sense functions, which, at this point, should be complete and in the `robot_class.py` file. The data is collected as an instantiated robot moves and senses in a world. Your SLAM function will take in this data as input. So, let's first create this data and explore how it represents the movement and sensor measurements that our robot takes.
---
## Create the world
Use the code below to generate a world of a specified size with randomly generated landmark locations. You can change these parameters and see how your implementation of SLAM responds!
`data` holds the sensors measurements and motion of your robot over time. It stores the measurements as `data[i][0]` and the motion as `data[i][1]`.
#### Helper functions
You will be working with the `robot` class that may look familiar from the first notebook,
In fact, in the `helpers.py` file, you can read the details of how data is made with the `make_data` function. It should look very similar to the robot move/sense cycle you've seen in the first notebook.
```
import numpy as np
from helpers import make_data
# your implementation of slam should work with the following inputs
# feel free to change these input values and see how it responds!
# world parameters
num_landmarks = 5 # number of landmarks
N = 20 # time steps
world_size = 100.0 # size of world (square)
# robot parameters
measurement_range = 50.0 # range at which we can sense landmarks
motion_noise = 2.0 # noise in robot motion
measurement_noise = 2.0 # noise in the measurements
distance = 20.0 # distance by which robot (intends to) move each iteratation
# make_data instantiates a robot, AND generates random landmarks for a given world size and number of landmarks
data = make_data(N, num_landmarks, world_size, measurement_range, motion_noise, measurement_noise, distance)
```
### A note on `make_data`
The function above, `make_data`, takes in so many world and robot motion/sensor parameters because it is responsible for:
1. Instantiating a robot (using the robot class)
2. Creating a grid world with landmarks in it
**This function also prints out the true location of landmarks and the *final* robot location, which you should refer back to when you test your implementation of SLAM.**
The `data` this returns is an array that holds information about **robot sensor measurements** and **robot motion** `(dx, dy)` that is collected over a number of time steps, `N`. You will have to use *only* these readings about motion and measurements to track a robot over time and find the determine the location of the landmarks using SLAM. We only print out the true landmark locations for comparison, later.
In `data` the measurement and motion data can be accessed from the first and second index in the columns of the data array. See the following code for an example, where `i` is the time step:
```
measurement = data[i][0]
motion = data[i][1]
```
```
# print out some stats about the data
time_step = 0
print('Example measurements: \n', data[time_step][0])
print('\n')
print('Example motion: \n', data[time_step][1])
```
Try changing the value of `time_step`, you should see that the list of measurements varies based on what in the world the robot sees after it moves. As you know from the first notebook, the robot can only sense so far and with a certain amount of accuracy in the measure of distance between its location and the location of landmarks. The motion of the robot always is a vector with two values: one for x and one for y displacement. This structure will be useful to keep in mind as you traverse this data in your implementation of slam.
## Initialize Constraints
One of the most challenging tasks here will be to create and modify the constraint matrix and vector: omega and xi. In the second notebook, you saw an example of how omega and xi could hold all the values the define the relationships between robot poses `xi` and landmark positions `Li` in a 1D world, as seen below, where omega is the blue matrix and xi is the pink vector.
<img src='images/motion_constraint.png' width=50% height=50% />
In *this* project, you are tasked with implementing constraints for a 2D world. We are referring to robot poses as `Px, Py` and landmark positions as `Lx, Ly`, and one way to approach this challenge is to add *both* x and y locations in the constraint matrices.
<img src='images/constraints2D.png' width=50% height=50% />
You may also choose to create two of each omega and xi (one for x and one for y positions).
### TODO: Write a function that initializes omega and xi
Complete the function `initialize_constraints` so that it returns `omega` and `xi` constraints for the starting position of the robot. Any values that we do not yet know should be initialized with the value `0`. You may assume that our robot starts out in exactly the middle of the world with 100% confidence (no motion or measurement noise at this point). The inputs `N` time steps, `num_landmarks`, and `world_size` should give you all the information you need to construct intial constraints of the correct size and starting values.
*Depending on your approach you may choose to return one omega and one xi that hold all (x,y) positions *or* two of each (one for x values and one for y); choose whichever makes most sense to you!*
```
def initialize_constraints(N, num_landmarks, world_size):
''' This function takes in a number of time steps N, number of landmarks, and a world_size,
and returns initialized constraint matrices, omega and xi.'''
## Recommended: Define and store the size (rows/cols) of the constraint matrix in a variable
size = 2 * (N + num_landmarks)
## TODO: Define the constraint matrix, Omega, with two initial "strength" values
## for the initial x, y location of our robot
omega = np.zeros((size, size))
omega[0][0] = 1.0
omega[1][1] = 1.0
## TODO: Define the constraint *vector*, xi
## you can assume that the robot starts out in the middle of the world with 100% confidence
xi = np.zeros((size, 1))
xi[0][0] = world_size/2
xi[1][0] = world_size/2
return omega, xi
```
### Test as you go
It's good practice to test out your code, as you go. Since `slam` relies on creating and updating constraint matrices, `omega` and `xi` to account for robot sensor measurements and motion, let's check that they initialize as expected for any given parameters.
Below, you'll find some test code that allows you to visualize the results of your function `initialize_constraints`. We are using the [seaborn](https://seaborn.pydata.org/) library for visualization.
**Please change the test values of N, landmarks, and world_size and see the results**. Be careful not to use these values as input into your final smal function.
This code assumes that you have created one of each constraint: `omega` and `xi`, but you can change and add to this code, accordingly. The constraints should vary in size with the number of time steps and landmarks as these values affect the number of poses a robot will take `(Px0,Py0,...Pxn,Pyn)` and landmark locations `(Lx0,Ly0,...Lxn,Lyn)` whose relationships should be tracked in the constraint matrices. Recall that `omega` holds the weights of each variable and `xi` holds the value of the sum of these variables, as seen in Notebook 2. You'll need the `world_size` to determine the starting pose of the robot in the world and fill in the initial values for `xi`.
```
# import data viz resources
import matplotlib.pyplot as plt
from pandas import DataFrame
import seaborn as sns
%matplotlib inline
# define a small N and world_size (small for ease of visualization)
N_test = 5
num_landmarks_test = 2
small_world = 10
# initialize the constraints
initial_omega, initial_xi = initialize_constraints(N_test, num_landmarks_test, small_world)
# define figure size
plt.rcParams["figure.figsize"] = (10,7)
# display omega
sns.heatmap(DataFrame(initial_omega), cmap='Blues', annot=True, linewidths=.5)
# define figure size
plt.rcParams["figure.figsize"] = (1,7)
# display xi
sns.heatmap(DataFrame(initial_xi), cmap='Oranges', annot=True, linewidths=.5)
```
---
## SLAM inputs
In addition to `data`, your slam function will also take in:
* N - The number of time steps that a robot will be moving and sensing
* num_landmarks - The number of landmarks in the world
* world_size - The size (w/h) of your world
* motion_noise - The noise associated with motion; the update confidence for motion should be `1.0/motion_noise`
* measurement_noise - The noise associated with measurement/sensing; the update weight for measurement should be `1.0/measurement_noise`
#### A note on noise
Recall that `omega` holds the relative "strengths" or weights for each position variable, and you can update these weights by accessing the correct index in omega `omega[row][col]` and *adding/subtracting* `1.0/noise` where `noise` is measurement or motion noise. `Xi` holds actual position values, and so to update `xi` you'll do a similar addition process only using the actual value of a motion or measurement. So for a vector index `xi[row][0]` you will end up adding/subtracting one measurement or motion divided by their respective `noise`.
### TODO: Implement Graph SLAM
Follow the TODO's below to help you complete this slam implementation (these TODO's are in the recommended order), then test out your implementation!
#### Updating with motion and measurements
With a 2D omega and xi structure as shown above (in earlier cells), you'll have to be mindful about how you update the values in these constraint matrices to account for motion and measurement constraints in the x and y directions. Recall that the solution to these matrices (which holds all values for robot poses `P` and landmark locations `L`) is the vector, `mu`, which can be computed at the end of the construction of omega and xi as the inverse of omega times xi: $\mu = \Omega^{-1}\xi$
**You may also choose to return the values of `omega` and `xi` if you want to visualize their final state!**
```
## TODO: Complete the code to implement SLAM
## slam takes in 6 arguments and returns mu,
## mu is the entire path traversed by a robot (all x,y poses) *and* all landmarks locations
def slam(data, N, num_landmarks, world_size, motion_noise, measurement_noise):
## TODO: Use your initilization to create constraint matrices, omega and xi
omega, xi = initialize_constraints(N, num_landmarks, world_size)
## TODO: Iterate through each time step in the data
for i in range(len(data)):
# Obtain measurement which is a list of lists
measurement = data[i][0]
#Obtain number of measurements
num_measurements = len(measurement)
# NOTE: number of measurements can vary
#print("Measurement "+str(i)+" number "+str(num_measurements))
#print(measurement)
# Obtain motion which is always a list
motion = data[i][1]
#Obtain number of motion
num_motion = len(motion)
# NOTE: number of measurements can vary
#print("Motion "+str(i)+" number "+str(num_motion))
#print(motion)
# Obtain position; multiply by 2 due to x,y co-ordinates
pos = i*2
for j in range(num_measurements):
landmark = 2*(N + measurement[j][0])
# xy is due to the (x,y) co-ordinate system
for xy in range(2):
omega[pos + xy][pos + xy] += 1.0/measurement_noise
omega[landmark + xy][landmark + xy] += 1.0/measurement_noise
omega[pos + xy][landmark + xy] += -1.0/measurement_noise
omega[landmark + xy][pos + xy] += -1.0/measurement_noise
xi[pos + xy][0] += -measurement[j][1 + xy]/measurement_noise
xi[landmark + xy][0] += measurement[j][1 + xy]/measurement_noise
for xy in range(2):
omega[pos + xy][pos + xy] += 1.0/motion_noise
omega[pos + 2 + xy][pos + 2 + xy] += 1.0/motion_noise
omega[pos + 2 + xy][pos + xy] += -1.0/motion_noise
omega[pos + xy][pos + 2 + xy] += -1.0/motion_noise
xi[pos + xy][0] += -motion[xy]/motion_noise
xi[pos + 2 + xy][0] += motion[xy]/motion_noise
omega_inv = np.linalg.inv(np.matrix(omega))
mu = omega_inv*xi
return mu # return `mu`
```
## Helper functions
To check that your implementation of SLAM works for various inputs, we have provided two helper functions that will help display the estimated pose and landmark locations that your function has produced. First, given a result `mu` and number of time steps, `N`, we define a function that extracts the poses and landmarks locations and returns those as their own, separate lists.
Then, we define a function that nicely print out these lists; both of these we will call, in the next step.
```
# a helper function that creates a list of poses and of landmarks for ease of printing
# this only works for the suggested constraint architecture of interlaced x,y poses
def get_poses_landmarks(mu, N):
# create a list of poses
poses = []
for i in range(N):
poses.append((mu[2*i].item(), mu[2*i+1].item()))
# create a list of landmarks
landmarks = []
for i in range(num_landmarks):
landmarks.append((mu[2*(N+i)].item(), mu[2*(N+i)+1].item()))
# return completed lists
return poses, landmarks
def print_all(poses, landmarks):
print('\n')
print('Estimated Poses:')
for i in range(len(poses)):
print('['+', '.join('%.3f'%p for p in poses[i])+']')
print('\n')
print('Estimated Landmarks:')
for i in range(len(landmarks)):
print('['+', '.join('%.3f'%l for l in landmarks[i])+']')
```
## Run SLAM
Once you've completed your implementation of `slam`, see what `mu` it returns for different world sizes and different landmarks!
### What to Expect
The `data` that is generated is random, but you did specify the number, `N`, or time steps that the robot was expected to move and the `num_landmarks` in the world (which your implementation of `slam` should see and estimate a position for. Your robot should also start with an estimated pose in the very center of your square world, whose size is defined by `world_size`.
With these values in mind, you should expect to see a result that displays two lists:
1. **Estimated poses**, a list of (x, y) pairs that is exactly `N` in length since this is how many motions your robot has taken. The very first pose should be the center of your world, i.e. `[50.000, 50.000]` for a world that is 100.0 in square size.
2. **Estimated landmarks**, a list of landmark positions (x, y) that is exactly `num_landmarks` in length.
#### Landmark Locations
If you refer back to the printout of *exact* landmark locations when this data was created, you should see values that are very similar to those coordinates, but not quite (since `slam` must account for noise in motion and measurement).
```
# call your implementation of slam, passing in the necessary parameters
mu = slam(data, N, num_landmarks, world_size, motion_noise, measurement_noise)
# print out the resulting landmarks and poses
if(mu is not None):
# get the lists of poses and landmarks
# and print them out
poses, landmarks = get_poses_landmarks(mu, N)
print_all(poses, landmarks)
```
## Visualize the constructed world
Finally, using the `display_world` code from the `helpers.py` file (which was also used in the first notebook), we can actually visualize what you have coded with `slam`: the final position of the robot and the positon of landmarks, created from only motion and measurement data!
**Note that these should be very similar to the printed *true* landmark locations and final pose from our call to `make_data` early in this notebook.**
```
# import the helper function
from helpers import display_world
# Display the final world!
# define figure size
plt.rcParams["figure.figsize"] = (20,20)
# check if poses has been created
if 'poses' in locals():
# print out the last pose
print('Last pose: ', poses[-1])
# display the last position of the robot *and* the landmark positions
display_world(int(world_size), poses[-1], landmarks)
```
### Question: How far away is your final pose (as estimated by `slam`) compared to the *true* final pose? Why do you think these poses are different?
You can find the true value of the final pose in one of the first cells where `make_data` was called. You may also want to look at the true landmark locations and compare them to those that were estimated by `slam`. Ask yourself: what do you think would happen if we moved and sensed more (increased N)? Or if we had lower/higher noise parameters.
**Answer**:
* If the data collection (N, the number of sensing) is increased, the estimate is more accurated.
* But, As the number of measurements (N) increases, the noise accumulates and the accuracy decreases.
* Therefore, I think the relationship between accuracy and measurement is inversely proportional.
* If Landmarks are [[89, 61], [39, 66], [94, 55], [52, 64], [55, 50]],
* Robot: [x=89.24863, y=42.06814] ----------> After SLAM (Estimated): [x=90.51137187463564, y=42.918242069320854]
## Testing
To confirm that your slam code works before submitting your project, it is suggested that you run it on some test data and cases. A few such cases have been provided for you, in the cells below. When you are ready, uncomment the test cases in the next cells (there are two test cases, total); your output should be **close-to or exactly** identical to the given results. If there are minor discrepancies it could be a matter of floating point accuracy or in the calculation of the inverse matrix.
### Submit your project
If you pass these tests, it is a good indication that your project will pass all the specifications in the project rubric. Follow the submission instructions to officially submit!
```
# Here is the data and estimated outputs for test case 1
test_data1 = [[[[1, 19.457599255548065, 23.8387362100849], [2, -13.195807561967236, 11.708840328458608], [3, -30.0954905279171, 15.387879242505843]], [-12.2607279422326, -15.801093326936487]], [[[2, -0.4659930049620491, 28.088559771215664], [4, -17.866382374890936, -16.384904503932]], [-12.2607279422326, -15.801093326936487]], [[[4, -6.202512900833806, -1.823403210274639]], [-12.2607279422326, -15.801093326936487]], [[[4, 7.412136480918645, 15.388585962142429]], [14.008259661173426, 14.274756084260822]], [[[4, -7.526138813444998, -0.4563942429717849]], [14.008259661173426, 14.274756084260822]], [[[2, -6.299793150150058, 29.047830407717623], [4, -21.93551130411791, -13.21956810989039]], [14.008259661173426, 14.274756084260822]], [[[1, 15.796300959032276, 30.65769689694247], [2, -18.64370821983482, 17.380022987031367]], [14.008259661173426, 14.274756084260822]], [[[1, 0.40311325410337906, 14.169429532679855], [2, -35.069349468466235, 2.4945558982439957]], [14.008259661173426, 14.274756084260822]], [[[1, -16.71340983241936, -2.777000269543834]], [-11.006096015782283, 16.699276945166858]], [[[1, -3.611096830835776, -17.954019226763958]], [-19.693482634035977, 3.488085684573048]], [[[1, 18.398273354362416, -22.705102332550947]], [-19.693482634035977, 3.488085684573048]], [[[2, 2.789312482883833, -39.73720193121324]], [12.849049222879723, -15.326510824972983]], [[[1, 21.26897046581808, -10.121029799040915], [2, -11.917698965880655, -23.17711662602097], [3, -31.81167947898398, -16.7985673023331]], [12.849049222879723, -15.326510824972983]], [[[1, 10.48157743234859, 5.692957082575485], [2, -22.31488473554935, -5.389184118551409], [3, -40.81803984305378, -2.4703329790238118]], [12.849049222879723, -15.326510824972983]], [[[0, 10.591050242096598, -39.2051798967113], [1, -3.5675572049297553, 22.849456408289125], [2, -38.39251065320351, 7.288990306029511]], [12.849049222879723, -15.326510824972983]], [[[0, -3.6225556479370766, -25.58006865235512]], [-7.8874682868419965, -18.379005523261092]], [[[0, 1.9784503557879374, -6.5025974151499]], [-7.8874682868419965, -18.379005523261092]], [[[0, 10.050665232782423, 11.026385307998742]], [-17.82919359778298, 9.062000642947142]], [[[0, 26.526838150174818, -0.22563393232425621], [4, -33.70303936886652, 2.880339841013677]], [-17.82919359778298, 9.062000642947142]]]
## Test Case 1
##
# Estimated Pose(s):
# [50.000, 50.000]
# [37.858, 33.921]
# [25.905, 18.268]
# [13.524, 2.224]
# [27.912, 16.886]
# [42.250, 30.994]
# [55.992, 44.886]
# [70.749, 59.867]
# [85.371, 75.230]
# [73.831, 92.354]
# [53.406, 96.465]
# [34.370, 100.134]
# [48.346, 83.952]
# [60.494, 68.338]
# [73.648, 53.082]
# [86.733, 38.197]
# [79.983, 20.324]
# [72.515, 2.837]
# [54.993, 13.221]
# [37.164, 22.283]
# Estimated Landmarks:
# [82.679, 13.435]
# [70.417, 74.203]
# [36.688, 61.431]
# [18.705, 66.136]
# [20.437, 16.983]
### Uncomment the following three lines for test case 1 and compare the output to the values above ###
# mu_1 = slam(test_data1, 20, 5, 100.0, 2.0, 2.0)
# poses, landmarks = get_poses_landmarks(mu_1, 20)
# print_all(poses, landmarks)
mu_1 = slam(test_data1, 20, 5, 100.0, 2.0, 2.0)
poses, landmarks = get_poses_landmarks(mu_1, 20)
print_all(poses, landmarks)
# Here is the data and estimated outputs for test case 2
test_data2 = [[[[0, 26.543274387283322, -6.262538160312672], [3, 9.937396825799755, -9.128540360867689]], [18.92765331253674, -6.460955043986683]], [[[0, 7.706544739722961, -3.758467215445748], [1, 17.03954411948937, 31.705489938553438], [3, -11.61731288777497, -6.64964096716416]], [18.92765331253674, -6.460955043986683]], [[[0, -12.35130507136378, 2.585119104239249], [1, -2.563534536165313, 38.22159657838369], [3, -26.961236804740935, -0.4802312626141525]], [-11.167066095509824, 16.592065417497455]], [[[0, 1.4138633151721272, -13.912454837810632], [1, 8.087721200818589, 20.51845934354381], [3, -17.091723454402302, -16.521500551709707], [4, -7.414211721400232, 38.09191602674439]], [-11.167066095509824, 16.592065417497455]], [[[0, 12.886743222179561, -28.703968411636318], [1, 21.660953298391387, 3.4912891084614914], [3, -6.401401414569506, -32.321583037341625], [4, 5.034079343639034, 23.102207946092893]], [-11.167066095509824, 16.592065417497455]], [[[1, 31.126317672358578, -10.036784369535214], [2, -38.70878528420893, 7.4987265861424595], [4, 17.977218575473767, 6.150889254289742]], [-6.595520680493778, -18.88118393939265]], [[[1, 41.82460922922086, 7.847527392202475], [3, 15.711709540417502, -30.34633659912818]], [-6.595520680493778, -18.88118393939265]], [[[0, 40.18454208294434, -6.710999804403755], [3, 23.019508919299156, -10.12110867290604]], [-6.595520680493778, -18.88118393939265]], [[[3, 27.18579315312821, 8.067219022708391]], [-6.595520680493778, -18.88118393939265]], [[], [11.492663265706092, 16.36822198838621]], [[[3, 24.57154567653098, 13.461499960708197]], [11.492663265706092, 16.36822198838621]], [[[0, 31.61945290413707, 0.4272295085799329], [3, 16.97392299158991, -5.274596836133088]], [11.492663265706092, 16.36822198838621]], [[[0, 22.407381798735177, -18.03500068379259], [1, 29.642444125196995, 17.3794951934614], [3, 4.7969752441371645, -21.07505361639969], [4, 14.726069092569372, 32.75999422300078]], [11.492663265706092, 16.36822198838621]], [[[0, 10.705527984670137, -34.589764174299596], [1, 18.58772336795603, -0.20109708164787765], [3, -4.839806195049413, -39.92208742305105], [4, 4.18824810165454, 14.146847823548889]], [11.492663265706092, 16.36822198838621]], [[[1, 5.878492140223764, -19.955352450942357], [4, -7.059505455306587, -0.9740849280550585]], [19.628527845173146, 3.83678180657467]], [[[1, -11.150789592446378, -22.736641053247872], [4, -28.832815721158255, -3.9462962046291388]], [-19.841703647091965, 2.5113335861604362]], [[[1, 8.64427397916182, -20.286336970889053], [4, -5.036917727942285, -6.311739993868336]], [-5.946642674882207, -19.09548221169787]], [[[0, 7.151866679283043, -39.56103232616369], [1, 16.01535401373368, -3.780995345194027], [4, -3.04801331832137, 13.697362774960865]], [-5.946642674882207, -19.09548221169787]], [[[0, 12.872879480504395, -19.707592098123207], [1, 22.236710716903136, 16.331770792606406], [3, -4.841206109583004, -21.24604435851242], [4, 4.27111163223552, 32.25309748614184]], [-5.946642674882207, -19.09548221169787]]]
## Test Case 2
##
# Estimated Pose(s):
# [50.000, 50.000]
# [69.035, 45.061]
# [87.655, 38.971]
# [76.084, 55.541]
# [64.283, 71.684]
# [52.396, 87.887]
# [44.674, 68.948]
# [37.532, 49.680]
# [31.392, 30.893]
# [24.796, 12.012]
# [33.641, 26.440]
# [43.858, 43.560]
# [54.735, 60.659]
# [65.884, 77.791]
# [77.413, 94.554]
# [96.740, 98.020]
# [76.149, 99.586]
# [70.211, 80.580]
# [64.130, 61.270]
# [58.183, 42.175]
# Estimated Landmarks:
# [76.777, 42.415]
# [85.109, 76.850]
# [13.687, 95.386]
# [59.488, 39.149]
# [69.283, 93.654]
### Uncomment the following three lines for test case 2 and compare to the values above ###
# mu_2 = slam(test_data2, 20, 5, 100.0, 2.0, 2.0)
# poses, landmarks = get_poses_landmarks(mu_2, 20)
# print_all(poses, landmarks)
mu_2 = slam(test_data2, 20, 5, 100.0, 2.0, 2.0)
poses, landmarks = get_poses_landmarks(mu_2, 20)
print_all(poses, landmarks)
```
| github_jupyter |
```
from EnsemblePursuit.EnsemblePursuit import EnsemblePursuit
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
from scipy.stats import zscore
from scipy.ndimage import gaussian_filter, gaussian_filter1d
from sklearn.preprocessing import MinMaxScaler
data_path='/media/maria/DATA1/Documents/data_for_suite2p/TX39/'
dt=1
spks= np.load(data_path+'spks.npy')
print('Shape of the data matrix, neurons by timepoints:',spks.shape)
iframe = np.load(data_path+'iframe.npy') # iframe[n] is the microscope frame for the image frame n
ivalid = iframe+dt<spks.shape[-1] # remove timepoints outside the valid time range
iframe = iframe[ivalid]
S = spks[:, iframe+dt]
print(S.shape)
U=np.load('U.npy')
del spks
stim_ens_inds=np.nonzero(U[:,13])[0]
print(stim_ens_inds.shape)
stim_k=2.0
stim_theta=2.0
stim_weights=np.random.gamma(shape=stim_k,scale=stim_theta,size=(stim_ens_inds.shape[0],))
plt.hist(stim_weights)
plt.show()
beh_ens_inds=np.nonzero(U[:,8])[0]
print(beh_ens_inds.shape)
beh_k=0.5
beh_theta=1.0
beh_weights=np.random.gamma(shape=beh_k,scale=beh_theta,size=(beh_ens_inds.shape[0],))
plt.hist(beh_weights)
plt.show()
#stim_ens_inds=np.nonzero(U[:,13])[0].sum()
weights=np.hstack((stim_weights,beh_weights))
sc=MinMaxScaler()
weights=sc.fit_transform(weights.reshape(-1,1))
print(weights)
stim_inp=S[stim_ens_inds]
beh_imp=S[beh_ens_inds]
input_patterns=np.vstack((stim_inp,beh_imp))
sc=MinMaxScaler()
input_patterns=sc.fit_transform(input_patterns)
weights=weights.flatten()
v_lst=[np.dot(weights,input_patterns[:,0])]
for j in range(1,30560):
v_lst.append(np.dot(weights,input_patterns[:,j]))
plt.plot(v_lst[:1000],color='g')
plt.title('Plasticity OFF, output')
plt.show()
v_lst=np.array(v_lst)
def gain_function(x):
#x=np.array(x)
#x[x<0]= 5*np.tanh(x[x<0]/5)
r_0=0.5
r_max=10.0
#print(x)
#if x>0:
#return r_0*np.tanh(x/r_0)
#else:
#return (r_max-r_0)*np.tanh(x/(r_max-r_0))
#x[x>=0]=(4000-5)*np.tanh(x[x>=0]/(4000-5))
return x
def update_weights(pre_syn_activity_pattern,post_syn_activity_pattern,W,theta_BCM):
alpha = 0.01
#print('syn',pre_syn_activity_pattern.reshape(9479,1)@post_syn_activity_pattern.reshape(1,2))
W+= alpha*pre_syn_activity_pattern.reshape(1105,)*post_syn_activity_pattern.reshape(1,)*(post_syn_activity_pattern-theta_BCM)
W[W<0]=0
return W
def update_BCM_threshold(theta_BCM,activity_pattern):
theta_BCM_dt = 0.01
BCM_target = 2.0
tau_theta=0.1
#print(theta_BCM)
theta_BCM += theta_BCM_dt*(((activity_pattern/BCM_target)*activity_pattern - theta_BCM))
print(theta_BCM)
return theta_BCM
activity_patterns=input_patterns
theta_lst=[]
print(weights.shape)
print(activity_patterns.shape)
print(weights)
_weights=weights.flatten()
theta_BCM = np.random.normal(size=(1105,))
print(theta_BCM)
rate=np.array([[0]])
h=0.1
for t in range(0,1000):
inpt=np.dot(_weights,activity_patterns[:,t])
#print('inp',inpt)
dxdt=(-rate[-1]+gain_function(inpt))
#print('dxdt',dxdt)
rate=np.vstack((rate,(rate[-1]+h*dxdt)))
_weights=update_weights(activity_patterns[:,t],rate[-1],_weights,theta_BCM)
theta_BCM=update_BCM_threshold(theta_BCM,rate[-1])
theta_lst.append(theta_BCM)
#plt.plot(theta_lst)
#print(theta_BCM)
#print(theta_lst[0])
for j in range(0,5):
plt.plot(theta_lst[j][:100])
print(theta_lst[j])
plt.show()
plt.plot(rate,color='tab:green')
plt.title('BCM Plasticity ON, output')
v_lst=rate[1:10001].flatten()
def train_test_split(NT):
nsegs = 20
nt=NT
nlen = nt/nsegs
ninds = np.linspace(0,nt-nlen,nsegs).astype(int)
itest = (ninds[:,np.newaxis] + np.arange(0,nlen*0.25,1,int)).flatten()
itrain = np.ones(nt, np.bool)
itrain[itest] = 0
return itrain, itest
mov=np.load(data_path+'mov.npy')
mov = mov[:, :, ivalid]
ly, lx, nstim = mov.shape
#print(nstim)
NT = v_lst.shape[0]
NN=1
mov=mov[:,:,:NT]
print(NT)
itrain,itest=train_test_split(NT)
X = np.reshape(mov, [-1, NT]) # reshape to Npixels by Ntimepoints
X = X-0.5 # subtract the background
X = np.abs(X) # does not matter if a pixel is black (0) or white (1)
X = zscore(X, axis=1)/NT**.5 # z-score each pixel separately
npix = X.shape[0]
lam = 0.1
#ncomps = Sp.shape[0]
B0 = np.linalg.solve((X[:,itrain] @ X[:,itrain].T + lam * np.eye(npix)), (X[:,itrain] @ v_lst[itrain].T)) # get the receptive fields for each neuron
B0 = np.reshape(B0, (ly, lx, 1))
B0 = gaussian_filter(B0, [.5, .5, 0])
rf = B0[:,:,0]
rfmax = np.max(B0)
# rfmax = np.max(np.abs(rf))
plt.imshow(rf, aspect='auto', cmap = 'bwr', vmin = -rfmax, vmax = rfmax)
```
| github_jupyter |
```
%store
%store -r __importRegression
__importRegression
endMonth = 34
#fnameTest = '../input/validation/test_' + str(endMonth) + '.csv'
#fnameTrain = '../input/validation/train_' + str(endMonth) + '.csv'
#train = pd.read_csv(fnameTrain)
#test = pd.read_csv(fnameTest)
#train = pd.read_csv('../input/train.csv')
#test = pd.read_csv('../input/test.csv')
all_data = pd.read_csv("../input/all_data_1_2_3_4_5_12_cat.csv")
items = pd.read_csv("../input/items.csv")
categories = pd.read_csv("../input/item_categories.csv")
def reduce_mem_usage(props):
start_mem_usg = props.memory_usage().sum() / 1024**2
print("Memory usage of properties dataframe is :",start_mem_usg," MB")
NAlist = [] # Keeps track of columns that have missing values filled in.
for col in props.columns:
if props[col].dtype != object: # Exclude strings
# Print current column type
print("******************************")
print("Column: ",col)
print("dtype before: ",props[col].dtype)
# make variables for Int, max and min
IsInt = False
mx = props[col].max()
mn = props[col].min()
# Integer does not support NA, therefore, NA needs to be filled
if not np.isfinite(props[col]).all():
NAlist.append(col)
props[col].fillna(mn-1,inplace=True)
# test if column can be converted to an integer
asint = props[col].fillna(0).astype(np.int64)
result = (props[col] - asint)
result = result.sum()
if result > -0.01 and result < 0.01:
IsInt = True
# Make Integer/unsigned Integer datatypes
if IsInt:
if mn >= 0:
if mx < 255:
props[col] = props[col].astype(np.uint8)
elif mx < 65535:
props[col] = props[col].astype(np.uint16)
elif mx < 4294967295:
props[col] = props[col].astype(np.uint32)
else:
props[col] = props[col].astype(np.uint64)
else:
if mn > np.iinfo(np.int8).min and mx < np.iinfo(np.int8).max:
props[col] = props[col].astype(np.int8)
elif mn > np.iinfo(np.int16).min and mx < np.iinfo(np.int16).max:
props[col] = props[col].astype(np.int16)
elif mn > np.iinfo(np.int32).min and mx < np.iinfo(np.int32).max:
props[col] = props[col].astype(np.int32)
elif mn > np.iinfo(np.int64).min and mx < np.iinfo(np.int64).max:
props[col] = props[col].astype(np.int64)
# Make float datatypes 32 bit
else:
props[col] = props[col].astype(np.float32)
# Print new column type
print("dtype after: ",props[col].dtype)
print("******************************")
# Print final result
print("___MEMORY USAGE AFTER COMPLETION:___")
mem_usg = props.memory_usage().sum() / 1024**2
print("Memory usage is: ",mem_usg," MB")
print("This is ",100*mem_usg/start_mem_usg,"% of the initial size")
return props, NAlist
all_data, NAs = reduce_mem_usage(all_data)
items.head(15)
categories.head(15)
all_data.head()
sub_data = all_data.copy()
sub_data.head()
col_to_keep = ['shop_id', 'target_lag_1', 'target_lag_2', 'target_lag_3', 'target_lag_4', 'target_lag_5', 'target_lag_12']
col_to_drop = list(sub_data.columns.difference(col_to_keep))
col_to_drop
sub_data.drop(col_to_drop, axis=1, inplace=True)
sub_data.head()
sub_data.shop_id.nunique()
sub_data = sub_data.groupby('shop_id', as_index=False).sum()
sub_data.head()
cols = sub_data.columns.tolist()
cols
order = ['shop_id',
'target_lag_1',
'target_lag_2',
'target_lag_3',
'target_lag_4',
'target_lag_5',
'target_lag_12']
sub_data = sub_data[order]
sub_data.head()
sub_data = sub_data.T
sub_data.head()
sub_data.columns
sub_data.plot(y=sub_data.columns)
sub_data.drop('shop_id', axis=1, inplace=True)
sub_data.d
sub_data = sub_data.groupby(['shop_id', 'date_block_num'], as_index=False).agg({'target':sum})
sub_data.head()
sub_data.plot.scatter('shop_id', 'target')
```
labels = test.pop('item_cnt_day')
```
train.head()
```
Let's find total sales per shop
```
train_shop = train.copy()
train_shop.drop(['date', 'date_block_num', 'item_price', 'item_id'], axis=1, inplace=True)
train_shop.head()
train_shop = train_shop.groupby(['shop_id'], as_index=False).agg({'item_cnt_day':sum})
train_shop['shop_total_sales'] = train_shop.item_cnt_day
train_shop.drop('item_cnt_day', axis=1, inplace=True)
train_shop.head()
train_shop.plot(x='shop_id', y='shop_total_sales', kind='bar')
train = pd.merge(train, train_shop, left_on='shop_id', right_on='shop_id')
train.head()
train_items = train.copy()
train_items.drop(['date', 'date_block_num', 'shop_id', 'item_price', 'shop_total_sales'], axis=1, inplace=True)
train_items.head()
train_items = train_items.groupby(['item_id'], as_index=False).agg({'item_cnt_day':sum})
train_items['item_total_sales'] = train_items.item_cnt_day
train_items.drop('item_cnt_day', axis=1, inplace=True)
train_items.head()
train_items.describe()
train = pd.merge(train, train_items, left_on='item_id', right_on='item_id')
train.head(10)
```
Let's find sales per season.
- Fall: months 09, 10, 11
- Winter: months 12, 01, 02
- Spring: months 03, 04, 05
- Summer: months 06, 07, 08
```
train['month'] = train.date_block_num % 12 + 1
train.head(10)
def season(x):
if x in [9, 10, 11]:
return 1 # Fall
if x in [12, 1, 2]:
return 2 # Winter
if x in [3, 4, 5]:
return 3 # Spring
if x in [6, 7, 8]:
return 4 # Summer
train['season'] = train.month.apply(season)
train.head(10)
```
| github_jupyter |
# Visualizing the Titanic Disaster
### Introduction:
This exercise is based on the titanic Disaster dataset avaiable at [Kaggle](https://www.kaggle.com/c/titanic).
To know more about the variables check [here](https://www.kaggle.com/c/titanic/data)
### Step 1. Import the necessary libraries
```
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
%matplotlib inline
```
### Step 2. Import the dataset from this [address](https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/07_Visualization/Titanic_Desaster/train.csv)
### Step 3. Assign it to a variable titanic
```
url = 'https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/Visualization/Titanic_Desaster/train.csv'
titanic = pd.read_csv(url)
titanic.head()
```
### Step 4. Set PassengerId as the index
```
titanic.set_index('PassengerId').head()
```
### Step 5. Create a pie chart presenting the male/female proportion
```
# sum the instances of males and females
males = (titanic['Sex'] == 'male').sum()
females = (titanic['Sex'] == 'female').sum()
# put them into a list called proportions
proportions = [males, females]
# Create a pie chart
plt.pie(
# using proportions
proportions,
# with the labels being officer names
labels = ['Males', 'Females'],
# with no shadows
shadow = False,
# with colors
colors = ['blue','red'],
# with one slide exploded out
explode = (0.15 , 0),
# with the start angle at 90%
startangle = 90,
# with the percent listed as a fraction
autopct = '%1.1f%%'
)
# View the plot drop above
plt.axis('equal')
# Set labels
plt.title("Sex Proportion")
# View the plot
plt.tight_layout()
plt.show()
```
### Step 6. Create a scatterplot with the Fare payed and the Age, differ the plot color by gender
```
# creates the plot using
lm = sns.lmplot(x = 'Age', y = 'Fare', data = titanic, hue = 'Sex', fit_reg=False)
# set title
lm.set(title = 'Fare x Age')
# get the axes object and tweak it
axes = lm.axes
axes[0,0].set_ylim(-5,)
axes[0,0].set_xlim(-5,85)
```
### Step 7. How many people survived?
```
titanic.Survived.sum()
```
### Step 8. Create a histogram with the Fare payed
```
# sort the values from the top to the least value and slice the first 5 items
df = titanic.Fare.sort_values(ascending = False)
df
# create bins interval using numpy
binsVal = np.arange(0,600,10)
binsVal
# create the plot
plt.hist(df, bins = binsVal)
# Set the title and labels
plt.xlabel('Fare')
plt.ylabel('Frequency')
plt.title('Fare Payed Histrogram')
# show the plot
plt.show()
```
### BONUS: Create your own question and answer it.
| github_jupyter |
# Maarten Breddels
# Motivation
## Glue Jupyter
* Glue in the notebook
* Comes from Qt -> GUI Challenges for ipywidgets
```
import glue_jupyter as gj
import ipywidgets
import ipywidgets as widgets
data = gj.example_data_xyz()
app = gj.jglue(data=data)
app.histogram1d();
```
## Oliver Borderies - SocGen
* Olivier: I want to put widget into a modern component pages (Vue)
* Me: I want more of these React based MaterialUI widgets
* Olivier: but Vue is better
* Me: But React is more popular
* Me & Olivier: lets autogen MaterialUI & Vuetify widgets
* Mario: I can make it!
* QuantStack project
# Plan
* Wrap both
* MaterialUI (React based)
* Vuetify (Vue based)
* Both are Material Design component libraries
* All code is autogenerated
* ipyvuetify
* xvuetify?
* jlvuetify?
* jvuetify
* Enable modern Single Page Applications (SPA) with widget support.
* With kernel (voila)
* Without kernel, plain html (nbconvert?)
* (for free it renders on mobile)
# ipymaterialui
* `$ pip install ipymaterialui`
* Wraps MaterialUI
* React based
* Material Design
* ~15 widgets manually wrapped
* Mario is working on wrapping it all
* (future: ipyreact + cookiecutter/manual how to wrap new/existing React components)
```
import ipymaterialui as mui
text1 = "Jupyter"
text2 = "Jupyter Widgets"
text3 = "Material UI"
text4 = "React"
texts = [text1, text2, text3, text4]
# the baseclass is just a
chips = [mui.Chip(label=text) for text in texts]
chips_div = mui.Div(children=chips)
chips_div
# Nice looking lists, the 3rd acting like a button
list_items = [
mui.ListItem(children=[mui.ListItemText(primary=text1, secondary=text3)], divider=True),
mui.ListItem(children=[mui.ListItemText(primary=text2, secondary=text4)], divider=True),
mui.ListItem(children=[mui.ListItemText(primary=text3, secondary=text1)], divider=True, button=True),
mui.ListItem(children=[mui.ListItemText(primary=text4, secondary=text2)], divider=True)
]
mui.List(children=list_items)
# For the moment only list items can be used for popup menus
# This needs a more generic solution?
menuitems = [
mui.MenuItem(description=text1, value='1'),
mui.MenuItem(description=text2, value='2'),
mui.MenuItem(description=text3, value='3')
]
menu = mui.Menu(children=menuitems)
list_item_text = mui.ListItemText(primary=text4, secondary=text1, button=True)
list_item = mui.ListItem(children=[list_item_text], button=True, menu=menu)
list_item
```
# Sure nice, but Olivier wants Vue(tify)
# Ipyvuetify
* `$ pip install ipyvuetify`
* QuantStack/SocGen project (Olivier Borderier)
* Made by Mario Buikhuizen
* Wraps Vuetify
* Vue based
* Material Design
```
import ipyvuetify as v
import ipywidgets as widgets
from threading import Timer
lorum_ipsum = 'Lorem ipsum dolor sit amet, consectetur adipiscing elit, sed do eiusmod tempor incididunt ut labore et dolore magna aliqua. Ut enim ad minim veniam, quis nostrud exercitation ullamco laboris nisi ut aliquip ex ea commodo consequat. Duis aute irure dolor in reprehenderit in voluptate velit esse cillum dolore eu fugiat nulla pariatur. Excepteur sint occaecat cupidatat non proident, sunt in culpa qui officia deserunt mollit anim id est laborum.'
v.Layout(children=[
v.Btn(color='primary', children=['primary']),
v.Btn(color='error', children=['error']),
v.Btn(color='pink lighten-4', children=['custom']),
v.Btn(color='#654321', dark=True, children=['hex']),
v.Btn(color='#654321', disabled=True, children=['disabled']),
])
v.Layout(children=[
v.Btn(color='primary', flat=True, children=['flat']),
v.Btn(color='primary', flat=True, disabled=True, children=['flat']),
v.Btn(color='primary', round=True, children=['round']),
v.Btn(color='primary', round=True, disabled=True, children=['round']),
v.Btn(color='primary', depressed=True, children=['depressed']),
v.Btn(color='primary', flat=True, icon=True, children=[v.Icon(children=['thumb_up'])]),
v.Btn(color='primary', outline=True, children=['outline']),
])
v.Layout(children=[
v.Btn(color='primary', small=True, children=['small']),
v.Btn(color='primary', children=['normal']),
v.Btn(color='primary', large=True, children=['large']),
v.Btn(color='primary', small=True, fab=True, children=[v.Icon(children=['edit'])]),
v.Btn(color='primary', fab=True, children=[v.Icon(children=['edit'])]),
v.Btn(color='primary', fab=True, large=True, children=[v.Icon(children=['edit'])]),
])
def toggleLoading():
button2.loading = not button2.loading
button2.disabled = button2.loading
def on_loader_click(*args):
toggleLoading()
Timer(2.0, toggleLoading).start()
button2 = v.Btn(loading=False, children=['loader'])
button2.on_event('click', on_loader_click)
v.Layout(children=[button2])
toggle_single = v.BtnToggle(v_model=2, class_='mr-3', children=[
v.Btn(flat=True, children=[v.Icon(children=['format_align_left'])]),
v.Btn(flat=True, children=[v.Icon(children=['format_align_center'])]),
v.Btn(flat=True, children=[v.Icon(children=['format_align_right'])]),
v.Btn(flat=True, children=[v.Icon(children=['format_align_justify'])]),
])
toggle_multi = v.BtnToggle(v_model=[0,2], multiple=True, children=[
v.Btn(flat=True, children=[v.Icon(children=['format_bold'])]),
v.Btn(flat=True, children=[v.Icon(children=['format_italic'])]),
v.Btn(flat=True, children=[v.Icon(children=['format_underline'])]),
v.Btn(flat=True, children=[v.Icon(children=['format_color_fill'])]),
])
v.Layout(pa_1=True, children=[
toggle_single,
toggle_multi,
])
v.Layout(children=[
v.Btn(color='primary', children=[
v.Icon(left=True, children=['fingerprint']),
'Icon left'
]),
v.Btn(color='primary', children=[
'Icon right',
v.Icon(right=True, children=['fingerprint']),
]),
v.Tooltip(bottom=True, children=[
v.Btn(slot='activator', color='primary', children=[
'tooltip'
]),
'Insert tooltip text here'
])
])
def on_menu_click(widget, event, data):
if len(layout.children) == 1:
layout.children = layout.children + [info]
info.children=[f'Item {items.index(widget)+1} clicked']
items = [v.ListTile(children=[
v.ListTileTitle(children=[
f'Click me {i}'])])
for i in range(1, 5)]
for item in items:
item.on_event('click', on_menu_click)
menu = v.Menu(offset_y=True, children=[
v.Btn(slot='activator', color='primary', children=[
'menu',
v.Icon(right=True, children=[
'arrow_drop_down'
])
]),
v.List(children=items)
])
info = v.Chip()
layout = v.Layout(children=[
menu
])
layout
v.Dialog(v_model=False, width='500', children=[
v.Btn(slot="activator", color='success', dark=True, children=[
"Open dialog"
]),
v.Card(children=[
v.CardTitle(class_='headline gray lighten-2', primary_title=True, children=[
"Lorem ipsum"]),
v.CardText(children=[
lorum_ipsum])
])
])
slider = v.Slider(v_model=25)
slider2 = v.Slider(thumb_label=True, v_model=25)
slider3 = v.Slider(thumb_label='always', v_model=25)
widgets.jslink((slider, 'v_model'), (slider2, 'v_model'))
widgets.jslink((slider, 'v_model'), (slider3, 'v_model'))
v.Container(children=[
slider,
slider2,
slider3
])
select1=v.Select(label="Choose option", items=['Option a', 'Option b', 'Option c'])
v.Layout(children=[select1])
tab_list = [v.Tab(children=[f'Tab {i}']) for i in range(1,4)]
content_list = [v.TabItem(children=[lorum_ipsum]) for i in range(1,4)]
tabs = v.Tabs(
v_model=1,
children=tab_list + content_list)
tabs
vepc1 = v.ExpansionPanelContent(children=[
v.Html(tag='div', slot='header', children=['item1']),
v.Card(children=[
v.CardText(children=['First Text'])])])
vepc2 = v.ExpansionPanelContent(children=[
v.Html(tag='div', slot='header', children=['item2']),
v.Card(children=[
v.CardText(children=['Second Text'])])])
vep = v.ExpansionPanel(children=[vepc1, vepc2])
vl = v.Layout(children=[vep])
vl
import ipyvuetify as v
from traitlets import (Unicode, List, Bool, Any)
class MyApp(v.VuetifyTemplate):
dark = Bool(True).tag(sync=True)
drawers = Any(['Default (no property)', 'Permanent', 'Temporary']).tag(sync=True)
model = Any(None).tag(sync=True)
type = Unicode('default (no property)').tag(sync=True)
clipped = Bool(False).tag(sync=True)
floating = Bool(True).tag(sync=True)
mini = Bool(False).tag(sync=True)
inset = Bool(False).tag(sync=True)
template = Unicode('''
<template>
<v-app id="sandbox" :dark="dark">
<v-navigation-drawer
v-model="model"
:permanent="type === 'permanent'"
:temporary="type === 'temporary'"
:clipped="clipped"
:floating="floating"
:mini-variant="mini"
absolute
overflow
app
>
</v-navigation-drawer>
<v-toolbar :clipped-left="clipped" app absolute>
<v-toolbar-side-icon
v-if="type !== 'permanent'"
@click.stop="model = !model"
></v-toolbar-side-icon>
<v-toolbar-title>Vuetify</v-toolbar-title>
</v-toolbar>
<v-content>
<v-container fluid>
<v-layout align-center justify-center>
<v-flex xs10>
<v-card>
<v-card-text>
<v-layout row wrap>
<v-flex xs12 md6>
<span>Scheme</span>
<v-switch v-model="dark" primary label="Dark"></v-switch>
</v-flex>
<v-flex xs12 md6>
<span>Drawer</span>
<v-radio-group v-model="type" column>
<v-radio
v-for="drawer in drawers"
:key="drawer"
:label="drawer"
:value="drawer.toLowerCase()"
primary
></v-radio>
</v-radio-group>
<v-switch v-model="clipped" label="Clipped" primary></v-switch>
<v-switch v-model="floating" label="Floating" primary></v-switch>
<v-switch v-model="mini" label="Mini" primary></v-switch>
</v-flex>
<v-flex xs12 md6>
<span>Footer</span>
<v-switch v-model="inset" label="Inset" primary></v-switch>
</v-flex>
</v-layout>
</v-card-text>
<v-card-actions>
<v-spacer></v-spacer>
<v-btn flat>Cancel</v-btn>
<v-btn flat color="primary">Submit</v-btn>
</v-card-actions>
</v-card>
</v-flex>
</v-layout>
</v-container>
</v-content>
<v-footer :inset="inset" app>
<span class="px-3">© {{ new Date().getFullYear() }}</span>
</v-footer>
</v-app>
</template>''').tag(sync=True)
def vue_menu_click(self, data):
self.color = self.items[data]
self.button_text = self.items[data]
app = MyApp()
app
app.inset = True
app.dark = True
app.type = 'permanent'
```
# core ipywidgets vs ipyvuetify
* Composible vs verbosity
```
options = ['pepperoni', 'pineapple', 'anchovies']
v.RadioGroup(children=[v.Radio(label=k, value=k) for k in options], v_model=options[0])
widgets.RadioButtons(options=options, value=options[0], description='Pizza topping:')
v.Btn(color='primary', children=[
v.Icon(left=True, children=['fingerprint']),
'Icon left'
])
widgets.Button(color='primary', description='icon left', icon='home')
menu = v.Menu(offset_y=True, children=[
v.Btn(slot='activator', color='primary', children=[
'menu',
v.Icon(right=True, chi ldren=[
'arrow_drop_down'
])
]),
v.List(children=items)
])
menu
```
| github_jupyter |
# Partial dependence plots.
*From http://scikit-learn.org/stable/auto_examples/ensemble/plot_partial_dependence.html*
Partial dependence plots show the dependence between the target function and a set of ‘target’ features, marginalizing over the values of all other features (the complement features). Due to the limits of human perception the size of the target feature set must be small (usually, one or two) thus the target features are usually chosen among the most important features.
*From https://cran.r-project.org/web/packages/datarobot/vignettes/PartialDependence.html*:
Consider an arbitrary model obtained by fitting a pparticular structure (e.g., random forest, support vector machine, or linear regression model) to a given dataset $\mathcal{D}$. This dataset includes $N$ observations $y_k$ of a response variable $y$ for $k = 1,2,\cdots,N$, along with $p$ covariates denoted $x_{i,k}$ for $i=1,2,\cdots,p$ and $k=1,2,\cdots,N$. This model generates predictions of the form:
$$\hat{y}_k = F(x_{1,k},x_{2,k},\cdots,x_{p,k})$$
for some function $F(\cdots)$. In the case of a single covariate $x_j$, Friedman's partial dependence plots are obtained by computing the following average and plotting it over a useful range of $x$ values:
$$\phi_j(x) = \frac{1}{N}\sum_{k=1}^NF(x_{1,k},\cdots,x_{j-1,k},x,x_{j+1,k},\cdots,x_{p,k})$$
The idea is that the function $\phi_j(x)$ tells us howt he value of the variable $x_j$ influences the model predictions $\hat{y}_k$ after we have "averaged out" the influence of all other variables. For linear regression models, the resulting plots are simply straight lines whose slopes are equal to the model parameters. Sepcifically, for a linear model, the prediction defined above has the form:
$$\hat{y}_k = \sum_{i=1}^p a_ix_{i,j}$$
from which it follows that the partial dependence function is
$$\phi_j(x) = a_j x + \frac{1}{N}\sum_{k=1}^N\sum_{i\ne j}a_ix_{i,k} = a_j x + \sum_{i\ne j}a_i \bar{x}_i$$
where $\bar{x}_i$ is the average value of the $i^{th}$ covariate. The main advantage of these plots is that they can be constructed for any predictive model, regardless of its form or complexity.
The multivariate extension of the partial dependence plots just described is straightforward in principle, but several practical issues arise. First and most obviously, these plots are harder to interpret: the bivariate partial dependence function $\phi_{i,j}(x,y)$ for two covariates $x_i$ and $x_j$ is defined analogously to $\phi(x)$ by averaging over all other covariates, and this function is still relatively easy to plot and visualize, but higher-dimensional extensions are problematic. Also, these multivariate partial dependence plots have been criticized as being inadequate in the face of certain strong interactions **-- more on this below**.
*Back to the sklearn example*:
This example shows how to obtain partial dependence plots from a GradientBoostingRegressor trained on the California housing dataset. The example is taken from ESL.
The plot shows four one-way and one two-way partial dependence plots. The target variables for the one-way PDP are: median income (`MedInc`), avg. occupants per household (`AvgOccup`), median house age (`HouseAge`), and avg. rooms per household (`AveRooms`).
We can clearly see that the median house price shows a linear relationship with the median income (top left) and that the house price drops when the avg. occupants per household increases (top middle). The top right plot shows that the house age in a district does not have a strong influence on the (median) house price; so does the average rooms per household. The tick marks on the x-axis represent the deciles of the feature values in the training data.
Partial dependence plots with two target features enable us to visualize interactions among them. The two-way partial dependence plot shows the dependence of median house price on joint values of house age and avg. occupants per household. We can clearly see an interaction between the two features: For an avg. occupancy greater than two, the house price is nearly independent of the house age, whereas for values less than two there is a strong dependence on age.
```
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from sklearn.model_selection import train_test_split
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.ensemble.partial_dependence import plot_partial_dependence
from sklearn.ensemble.partial_dependence import partial_dependence
from sklearn.datasets.california_housing import fetch_california_housing
from sklearn.linear_model import LinearRegression
cal_housing = fetch_california_housing()
# split 80/20 train-test
X_train, X_test, y_train, y_test = train_test_split(cal_housing.data,
cal_housing.target,
test_size=0.2,
random_state=1)
names = cal_housing.feature_names
print("Training GBRT...")
boring_linear.fit(X_train, y_train)
clf.fit(X_train, y_train)
print(cal_housing.DESCR)
import pandas as pd
pd.Series(clf.feature_importances_, index=names)
# Note latitude and longitude are "important". Let's plot.
print('Convenience plot with ``partial_dependence_plots``')
features = [0, 5, 1, 2, (5, 1)]
fig, ax = plt.subplots(figsize=(12,6))
fig, ax = plot_partial_dependence(clf, X_train, features,
feature_names=names,
n_jobs=3, grid_resolution=50,
ax=ax)
fig.suptitle('Partial dependence of house value on nonlocation features\n'
'for the California housing dataset')
plt.subplots_adjust(top=0.9) # tight_layout causes overlap with suptitle
print('Custom 3d plot via ``partial_dependence``')
fig = plt.figure(figsize=(12,8))
target_feature = (1, 5)
pdp, axes = partial_dependence(clf, target_feature,
X=X_train,
grid_resolution=50)
XX, YY = np.meshgrid(axes[0], axes[1])
Z = pdp[0].reshape(list(map(np.size, axes))).T
ax = Axes3D(fig)
surf = ax.plot_surface(XX, YY, Z, rstride=1, cstride=1,
cmap=plt.cm.BuPu, edgecolor='k')
ax.set_xlabel(names[target_feature[0]])
ax.set_ylabel(names[target_feature[1]])
ax.set_zlabel('Partial dependence')
# pretty init view
ax.view_init(elev=22, azim=122)
plt.colorbar(surf)
plt.suptitle('Partial dependence of house value on median\n'
'age and average occupancy')
plt.subplots_adjust(top=0.9)
plt.show()
```
# When this fails
- Adapted from https://arxiv.org/pdf/1309.6392.pdf
```
size = 10000
X1 = np.random.uniform(-1,1,size)
X2 = np.random.uniform(-1,1,size)
X3 = np.random.uniform(-1,1,size)
eps = np.random.normal(0,0.5,size)
Y = 0.2*X1 - 5*X2 + 10*X2*np.where(X3>=0,1,0) + eps
%matplotlib inline
plt.scatter(X2,Y, color = C3)
Xs = np.stack([X1,X2,X3]).T
from sklearn.model_selection import GridSearchCV
clf = GradientBoostingRegressor(max_depth=3)
params = {'n_estimators':[50,100,200]}
grid = GridSearchCV(clf, param_grid = params, n_jobs = 5, cv = 5)
grid.fit(Xs,Y)
plot_partial_dependence(grid.best_estimator_, Xs, [1],
n_jobs=1, grid_resolution=25)
plt.xlim(-1,1)
plt.ylim(-6,6)
```
# This shows key caveat in Friedman's initial explanation of PDPs:
In general, the functional form of $\hat{F}_{z_{\setminus l}}(z_l)$ will depend on the particular values chosen for $z_{\setminus l}$. **If, however, this dependence is not to strong than the average function can represent a useful summary of the partial dependence of $\hat{F}(x)$ on the chosen variable subset $z_l$**.
In the special cases where the dependence of $\hat{F}(x)$ in $z_l$ is additive
$$\hat{F}(x) = \hat{F}_l(z_l) + \hat{F}_{\setminus l}(z_{\setminus l})$$,
or multiplicative
$$\hat{F}(x) = \hat{F}_l(z_l)\hat{F}_{\setminus l}(z_{\setminus l})$$
the *form* of $\hat{F}_{z\setminus l}(z_l)$ does not depend on the joint values of the complement varaibles $z_{\setminus l}$. Then $\bar{F}_l(z_l)$ provides a complete description of the nature of the variation of $\hat{F}(x)$ on the chosen input variable subset $z_l$.
| github_jupyter |
<center><h1>Web Scraping Kijiji - Streamlined Version</h1><h3>Using Python and Beautiful Soup</h3></center>
```
import pandas as pd
from IPython.display import HTML
from bs4 import BeautifulSoup
import urllib.request as request
from ipywidgets import interact
pd.set_option("display.max_rows",1000)
pd.set_option("display.max_columns",20)
pd.set_option("display.max_colwidth", 200)
base_url = 'http://www.kijiji.ca'
toronto_url = 'http://www.kijiji.ca/h-city-of-toronto/1700273'
html_kijiji = request.urlopen(toronto_url)
soup_kijiji = BeautifulSoup(html_kijiji, 'lxml')
div_categories = soup_kijiji.find_all('a', class_='category-selected')
categories = {}
for item in div_categories:
categories[item.get_text()] = base_url + item['href']
category_list = [key for key in categories.keys()]
pages = {'Page 1':'', 'Page 2':'page-2', 'Page 3':'page-3', 'Page 4':'page-4', 'Page 5':'page-5',
'Page 6':'page-6', 'Page 7':'page-7', 'Page 8':'page-8', 'Page 9':'page-9'}
@interact
def kijiji_listings(category = sorted(category_list), page = sorted(pages)):
if page == 'Page 1':
print(categories[category])
html_cars = request.urlopen(categories[category])
else:
url = categories[category]
last_forward_slash = url.rfind('/')
beginning_url = url[:last_forward_slash+1]
ending_url = url[forward_slash:]
print(beginning_url + pages[page] + ending_url)
html_cars = request.urlopen(beginning_url + pages[page] + ending_url)
soup_cars = BeautifulSoup(html_cars, 'lxml')
#tables = soup_cars.find_all('table', class_ = re.compile('regular-ad|top-'))
tables = soup_cars.find_all('table')
img_urls = []
for table in tables[1:]:
for row in table.find_all('td', class_='image'):
try:
img_urls.append("<img src='" + row.div.img['src'] + "'>")
except:
img_urls.append("<img src='" + row.img['src'] + "'>")
titles = []
for table in tables[1:]:
for row in table.find_all('td', class_='description'):
titles.append(row.a.get_text().strip())
comments = []
for table in tables[1:]:
for row in table.find_all('td', class_='description'):
comments.append(row.p.get_text().strip())
details = []
for table in tables[1:]:
for row in table.find_all('td', class_='description'):
for item in row.find_all('p', class_='details'):
details.append(item.get_text().strip())
prices = []
for table in tables[1:]:
for row in table.find_all('td', class_='price'):
try:
prices.append(float(row.get_text().replace('$','').replace(',','').strip()))
except:
prices.append(0.0)
try:
df = pd.DataFrame({'Price':prices, 'Image':img_urls, 'Title':titles, 'Comment':comments, 'Details':details})
# Arrange the columns in a certain order
df = df[['Image','Title','Comment','Details','Price']]
# Some category listings don't have a price and title, so this script would bomb unless we leave them out
except:
df = pd.DataFrame({'Image':img_urls, 'Comment':comments, 'Details':details})
# Arrange the columns in a certain order
df = df[['Image','Comment','Details']]
return HTML(df.to_html(escape=False)) # if escape is set to True, the images won't be rendered
```
<br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br>
| github_jupyter |
# Train ML model on Cloud AI Platform
This notebook shows how to:
* Export training code from [a Keras notebook](../06_feateng_keras/solution/taxifare_fc.ipynb) into a trainer file
* Create a Docker container based on a [DLVM container](https://cloud.google.com/ai-platform/deep-learning-containers/docs/kubernetes-container])
* Deploy training job to cluster
## TODO: Export the data from BigQuery to GCS
1. Navigate to [export_data.ipynb](export_data.ipynb)
2. Update 'your-gcs-project-here' to your GCP project name
3. Run all the notebook cells
## TODO: Edit notebook parameters
1. Navigate to [notebook_params.yaml](notebook_params.yaml)
2. Replace the bucket name with your own bucket containing your model (likely gcp-project with -ml at the end)
3. Save the notebook
4. Return to this notebook and continue
## Export code from notebook
This notebook extracts code from a notebook and creates a Python file suitable for use as model.py
```
import logging
import nbformat
import sys
import yaml
def write_parameters(cell_source, params_yaml, outfp):
with open(params_yaml, 'r') as ifp:
y = yaml.safe_load(ifp)
# print out all the lines in notebook
write_code(cell_source, 'PARAMS from notebook', outfp)
# print out YAML file; this will override definitions above
formats = [
'{} = {}', # for integers and floats
'{} = "{}"', # for strings
]
write_code(
'\n'.join([
formats[type(value) is str].format(key, value) for key, value in y.items()]),
'PARAMS from YAML',
outfp
)
def write_code(cell_source, comment, outfp):
lines = cell_source.split('\n')
if len(lines) > 0 and lines[0].startswith('%%'):
prefix = '#'
else:
prefix = ''
print("### BEGIN {} ###".format(comment), file=outfp)
for line in lines:
line = prefix + line.replace('print(', 'logging.info(')
if len(line) > 0 and (line[0] == '!' or line[0] == '%'):
print('#' + line, file=outfp)
else:
print(line, file=outfp)
print("### END {} ###\n".format(comment), file=outfp)
def convert_notebook(notebook_filename, params_yaml, outfp):
write_code('import logging', 'code added by notebook conversion', outfp)
with open(INPUT) as ifp:
nb = nbformat.reads(ifp.read(), nbformat.NO_CONVERT)
for cell in nb.cells:
if cell.cell_type == 'code':
if 'tags' in cell.metadata and 'display' in cell.metadata.tags:
logging.info('Ignoring cell # {} with display tag'.format(cell.execution_count))
elif 'tags' in cell.metadata and 'parameters' in cell.metadata.tags:
logging.info('Writing params cell # {}'.format(cell.execution_count))
write_parameters(cell.source, PARAMS, outfp)
else:
logging.info('Writing model cell # {}'.format(cell.execution_count))
write_code(cell.source, 'Cell #{}'.format(cell.execution_count), outfp)
import os
INPUT='../../06_feateng_keras/solution/taxifare_fc.ipynb'
PARAMS='./notebook_params.yaml'
OUTDIR='./container/trainer'
!mkdir -p $OUTDIR
OUTFILE=os.path.join(OUTDIR, 'model.py')
!touch $OUTDIR/__init__.py
with open(OUTFILE, 'w') as ofp:
#convert_notebook(INPUT, PARAMS, sys.stdout)
convert_notebook(INPUT, PARAMS, ofp)
#!cat $OUTFILE
```
## Try out model file
<b>Note</b> Once the training starts, __Interrupt the Kernel__ (from the notebook ribbon bar above). Because it processes the entire dataset, this will take a long time on the relatively small machine on which you are running Notebooks.
```
!python3 $OUTFILE
```
## Create Docker container
Package up the trainer file into a Docker container and submit the image.
```
%%writefile container/Dockerfile
FROM gcr.io/deeplearning-platform-release/tf2-cpu
#RUN python3 -m pip install --upgrade --quiet tf-nightly-2.0-preview
RUN python3 -m pip install --upgrade --quiet cloudml-hypertune
COPY trainer /trainer
CMD ["python3", "/trainer/model.py"]
%%writefile container/push_docker.sh
export PROJECT_ID=$(gcloud config list project --format "value(core.project)")
export IMAGE_REPO_NAME=serverlessml_training_container
#export IMAGE_TAG=$(date +%Y%m%d_%H%M%S)
#export IMAGE_URI=gcr.io/$PROJECT_ID/$IMAGE_REPO_NAME:$IMAGE_TAG
export IMAGE_URI=gcr.io/$PROJECT_ID/$IMAGE_REPO_NAME
echo "Building $IMAGE_URI"
docker build -f Dockerfile -t $IMAGE_URI ./
echo "Pushing $IMAGE_URI"
docker push $IMAGE_URI
!find container
```
<b>Note</b>: If you get a permissions error when running push_docker.sh from Notebooks, do it from CloudShell:
* Open [CloudShell](https://console.cloud.google.com/cloudshell) on the GCP Console
* ```git clone https://github.com/GoogleCloudPlatform/training-data-analyst```
* ```cd training-data-analyst/quests/serverlessml/07_caip/solution/container```
* ```bash push_docker.sh```
This next step takes 5 - 10 minutes to run
```
%%bash
cd container
bash push_docker.sh
```
## Deploy to AI Platform
Submit a training job using this custom container that we have just built. After you submit the job, [monitor it here](https://console.cloud.google.com/ai-platform/jobs).
```
%%bash
JOBID=serverlessml_$(date +%Y%m%d_%H%M%S)
REGION=us-central1
PROJECT_ID=$(gcloud config list project --format "value(core.project)")
BUCKET=$(gcloud config list project --format "value(core.project)")-ml
#IMAGE=gcr.io/deeplearning-platform-release/tf2-cpu
IMAGE=gcr.io/$PROJECT_ID/serverlessml_training_container
gcloud beta ai-platform jobs submit training $JOBID \
--staging-bucket=gs://$BUCKET --region=$REGION \
--master-image-uri=$IMAGE \
--master-machine-type=n1-standard-4 --scale-tier=CUSTOM
```
The training job will take 35 - 45 minutes to complete on the dataset. You can cancel the job once you confirm it started and have inspected the logs.
Copyright 2020 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| github_jupyter |
# Run pathways in FaIR
The pathways are generated elsewhere, imported here and then run.
```
import json
from multiprocessing import Pool
import platform
from climateforcing.utils import mkdir_p
import fair
import matplotlib.pyplot as pl
import numpy as np
import pandas as pd
from tqdm import tqdm
with open('../data_input/fair-1.6.2-ar6/fair-1.6.2-wg3-params.json') as f:
config_list = json.load(f)
emissions_in = {}
results_out = {}
WORKERS = 3 # set this based on your individual machine - allows parallelisation. nprocessors-1 is a sensible shout.
scenarios = ["ssp245_constant-2020-ch4", "ch4_30", "ch4_40", "ch4_50", "coal-phase-out"]
for scenario in scenarios:
emissions_in[scenario] = np.loadtxt('../data_output/fair_emissions_files/{}.csv'.format(scenario), delimiter=',')
```
## convenience function for running FaIR config with each emission species
```
def run_fair(args):
thisC, thisF, thisT, _, thisOHU, _, thisAF = fair.forward.fair_scm(**args)
return (thisC[:,0], thisC[:,1], thisT, thisF[:,1], np.sum(thisF, axis=1))
def fair_process(emissions):
updated_config = []
for i, cfg in enumerate(config_list):
updated_config.append({})
for key, value in cfg.items():
if isinstance(value, list):
updated_config[i][key] = np.asarray(value)
else:
updated_config[i][key] = value
updated_config[i]['emissions'] = emissions
updated_config[i]['diagnostics'] = 'AR6'
updated_config[i]["efficacy"] = np.ones(45)
updated_config[i]["gir_carbon_cycle"] = True
updated_config[i]["temperature_function"] = "Geoffroy"
updated_config[i]["aerosol_forcing"] = "aerocom+ghan2"
updated_config[i]["fixPre1850RCP"] = False
# updated_config[i]["scale"][43] = 0.6
updated_config[i]["F_solar"][270:] = 0
# multiprocessing is not working for me on Windows
if platform.system() == 'Windows':
shape = (361, len(updated_config))
c_co2 = np.ones(shape) * np.nan
c_ch4 = np.ones(shape) * np.nan
t = np.ones(shape) * np.nan
f_ch4 = np.ones(shape) * np.nan
f_tot = np.ones(shape) * np.nan
for i, cfg in tqdm(enumerate(updated_config), total=len(updated_config), position=0, leave=True):
c_co2[:,i], c_ch4[:,i], t[:,i], f_ch4[:,i], f_tot[:,i] = run_fair(updated_config[i])
else:
if __name__ == '__main__':
with Pool(WORKERS) as pool:
result = list(tqdm(pool.imap(run_fair, updated_config), total=len(updated_config), position=0, leave=True))
result_t = np.array(result).transpose(1,2,0)
c_co2, c_ch4, t, f_ch4, f_tot = result_t
temp_rebase = t - t[100:151,:].mean(axis=0)
return c_co2, c_ch4, temp_rebase, f_ch4, f_tot
```
## Do the runs
```
for scenario in tqdm(scenarios, position=0, leave=True):
results_out[scenario] = {}
(
results_out[scenario]['co2_concentrations'],
results_out[scenario]['ch4_concentrations'],
results_out[scenario]['temperatures'],
results_out[scenario]['ch4_effective_radiative_forcing'],
results_out[scenario]['effective_radiative_forcing']
) = fair_process(emissions_in[scenario])
```
## Save temperature outputs to analyse elsewhere
```
for scenario in scenarios:
for var in ['co2_concentrations', 'ch4_concentrations', 'temperatures', 'ch4_effective_radiative_forcing', 'effective_radiative_forcing']:
mkdir_p('../data_output/fair_{}/'.format(var))
df_out = pd.DataFrame(results_out[scenario][var][245:351,:])
df_out['year'] = np.arange(1995.5, 2101)
df_out.set_index('year', inplace=True)
df_out.to_csv('../data_output/fair_{}/{}.csv'.format(var, scenario), float_format="%6.4f")
```
| github_jupyter |
```
%load_ext sparkmagic.magics
```
Run the following cell to invoke the user interface for managing Spark.
```
%spark add -s cpu_session -l python -u http://node03.conductor.iccmop:8993 -a u -k config
#{"conf": {"spark.default.parallelism":30,"spark.cores.max":30,
# "spark.ego.gpu.app": "false"}}
%%spark -s cpu_session
from __future__ import print_function
import os
import time
import argparse
import sys
## SPARK
from pyspark import SparkConf, SparkContext
from pyspark.sql.session import SparkSession
# data paths
# Please use the absolute path of the file if you wish to run the example on a distributed mode
data_path = '/shared/kelvin/snapml'
filename = data_path + '/criteo.kaggle2014'
train_filename = filename + '-train.libsvm'
test_filename = filename + '-test.libsvm'
## snapML
os.environ["PYTHONPATH"] = '/opt/DL/snap-ml-spark/lib/'
os.environ["SPARK_PYTHON_DIR"] = '/var/conductor/livy-integration/spark-2.3.1-hadoop-2.7/python'
sys.path.append('/opt/DL/snap-ml-spark/lib/')
sys.path.append('/usr/lib64/python2.7/site-packages')
n_features_ = 1000000
snapml_regularizer = 10.0
from pyspark.ml.classification import LogisticRegression as sparkml_LogisticRegression
#train_filename = "file://" + train_filename
train_filename = "file://" + test_filename
test_filename = "file://" + test_filename
# Load training data
train_data = spark.read.format("libsvm").option("numFeatures", str(n_features_)).load(train_filename)
test_data = spark.read.format("libsvm").option("numFeatures", str(n_features_)).load(test_filename)
n_examples = train_data.count()
# Create sparkML lib Logistic Regression
sparkml_lr = sparkml_LogisticRegression(fitIntercept=False, regParam=snapml_regularizer/n_examples, standardization=False)
# Fit the model and time it
sparkml_t0 = time.time()
sparkml_lr_model = sparkml_lr.fit(train_data)
sparkml_time = time.time() - sparkml_t0
# Perform inference on test data
predictions = sparkml_lr_model.transform(test_data)
# Show predictions against test labels
predictions.select("rawPrediction", "prediction", "label", "features").show(10)
# Compute accuracy
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
evaluator = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction", metricName="accuracy")
sparkml_accuracy = evaluator.evaluate(predictions)
# Print off Spark result
print('Spark ML', evaluator.getMetricName(),'=', sparkml_accuracy,", time: %.2f" % sparkml_time, 's')
%spark delete -s cpu_session
%spark add -s gpu_session -l python -u http://node02.conductor.iccmop:8995 -a u -k config
#{"conf": {"spark.ego.gpu.app":"true","spark.ego.gpu.mode":"default","spark.default.parallelism":8}}
%%spark -s gpu_session
from __future__ import print_function
import os
import time
import argparse
import sys
## SPARK
sys.path.append('/opt/DL/snap-ml-spark/lib/')
# data paths
# Please use the absolute path of the file if you wish to run the example on a distributed mode
data_path = '/shared/kelvin/snapml'
filename = data_path + '/criteo.kaggle2014'
train_filename = filename + '-train.libsvm'
test_filename = filename + '-test.libsvm'
## snapML
os.environ["PYTHONPATH"] = '/opt/DL/snap-ml-spark/lib/'
os.environ["SPARK_PYTHON_DIR"] = '/var/conductor/livy-integration/spark-2.3.1-hadoop-2.7/python'
sys.path.append('/opt/DL/snap-ml-spark/lib/')
sys.path.append('/usr/lib64/python2.7/site-packages')
from pyspark import SparkConf, SparkContext
from pyspark.sql.session import SparkSession
from snap_ml_spark import DatasetReader
from snap_ml_spark import LogisticRegression as snapml_LogisticRegression
from snap_ml_spark.Metrics import accuracy, logisticLoss
n_features_ = 1000000
print('n_features: %.d' %n_features_)
# Load training data
train_data = DatasetReader().setFormat("libsvm").setNumFt(n_features_).load(train_filename)
count1 = train_data.count()
print('count1: %.d' %count1)
# Load test data
test_data = DatasetReader().setFormat("libsvm").setNumFt(n_features_).load(test_filename)
count2 = train_data.count()
##print('count2: %.d' %count2)
# Create snapML Logistic Regression
snapml_regularizer = 10.0
snapml_lr = snapml_LogisticRegression(max_iter=20, regularizer=snapml_regularizer, verbose=False, dual=True, use_gpu=True, n_threads=-1, class_weights=None)
# Fit the model and time it
snapml_t0 = time.time()
snapml_lr.fit(train_data)
snapml_time = time.time() - snapml_t0
# Perform inference on test data
pred = snapml_lr.predict(test_data)
# Compute accuracy
snapml_accuracy = accuracy(pred)
# Print off SnapML result
print('snapML accuracy: %.4f' %snapml_accuracy, ", time: %.2f" % snapml_time, 's')
%spark delete -s gpu_session
%spark cleanup
```
| github_jupyter |
# LiteBIRD Simulation Example
This notebook does a simple LiteBIRD simulation which you can use as a starting point for testing and customization. The toast_litebird package is here: https://github.com/hpc4cmb/toast-litebird and the documentation is here: https://hpc4cmb.github.io/toast-litebird/
First you must get access to the kernel that has the toast_litebird package. Open a jupyter terminal and do:
```
%> module use /global/common/software/litebird/cori/modulefiles
%> module load litebird
%> litebird-jupyter.sh
```
Now in this notebook select the the litebird kernel. You may have to shutdown this notebook and re-open to see the new kernel.
```
import os
import sys
import healpy as hp
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
import toast
from toast.mpi import MPI
from toast.utils import memreport
from toast import pipeline_tools
from toast_litebird import pipeline_tools as lbtools
# Capture C++ output in the jupyter cells
%reload_ext wurlitzer
```
## Select Detectors
We can use some command line tools to easily select detectors and dump them to a file for use in a pipeline. This command creates a full hardware model:
```
! lb_hardware_sim --overwrite
```
We can look at the details of the hardware file:
```
! lb_hardware_info hardware.toml.gz
```
Now we can select just some detectors
```
! lb_hardware_trim \
--hardware hardware.toml.gz \
--overwrite \
--out selected \
--telescopes LFT \
--match 'band:.*040'
! lb_hardware_info selected.toml.gz
```
Plot this
```
! lb_hardware_plot --hardware selected.toml.gz --out selected.pdf
```
The previous command makes a PDF file. We can display it:
```
from IPython.display import IFrame
IFrame("selected.pdf", width=600, height=300)
```
## Parameters
These arguments control the entire notebook
```
class args:
# Hardware model
hardware = "selected.toml.gz"
bands = "040"
thinfp = False
# Observations
obs_num = 1
start_time = 0
sample_rate = 30.0
obs_time_h = 23.0
gap_h = 1.0
# half-wave plate
hwp_rpm = 91.0
hwp_step_deg = None
hwp_step_time_s = None
# Scanning parameters
spin_period_min = 10.0
spin_angle_deg = 50.0 # This is "beta"
prec_period_min = 96.174
prec_angle_deg = 45.0 # This is "alpha"
# Pixelization
coord = "G"
nside = 512
mode = "IQU"
single_precision_pointing = False
nside_submap = 16
# Noise simulation
common_mode_noise = False
# Output directory
outdir = "litebird_out"
```
## Communicator
Since this is a serial notebook, this communicator will just have one process.
```
mpiworld, procs, rank = toast.mpi.get_world()
comm = toast.mpi.Comm(mpiworld)
memreport("After communicator creation", comm.comm_world)
```
## Focalplane
Load the hardware file and create the focalplane.
```
hw, telescope = lbtools.get_hardware(args, comm)
focalplane = lbtools.get_focalplane(args, comm, hw)
memreport("After focalplane creation", comm.comm_world)
```
## Create Observations
This uses the parameters at the top of the notebook to simulate regular spaced observations.
```
data = lbtools.create_observations(args, comm, focalplane, 1)
memreport("After creating observations", comm.comm_world)
```
## Pointing matrix
Here we translate the boresight quaternions into detector pointing (pixels numbers and Stokes weights).
```
pipeline_tools.expand_pointing(args, comm, data)
memreport("After expanding pointing", comm.comm_world)
```
Make a boolean hit map for diagnostics
```
npix = 12 * args.nside ** 2
hitmap = np.zeros(npix)
for obs in data.obs:
tod = obs["tod"]
for det in tod.local_dets:
pixels = tod.cache.reference("pixels_{}".format(det))
hitmap[pixels] = 1
hitmap[hitmap == 0] = hp.UNSEEN
hp.mollview(hitmap, nest=True, title="all hit pixels", cbar=False)
hp.graticule(22.5, verbose=False)
```
## Sky signal
Create a synthetic Gaussian map to scan as input signal
```
lmax = args.nside * 2
cls = np.zeros([4, lmax + 1])
cls[0] = 1e0
sim_map = hp.synfast(cls, args.nside, lmax=lmax, fwhm=np.radians(15), new=True)
plt.figure(figsize=[12, 8])
for i, m in enumerate(sim_map):
hp.mollview(sim_map[i], cmap="coolwarm", title="Input signal {}".format("IQU"[i]), sub=[1, 3, 1+i])
hp.write_map("sim_map.fits", hp.reorder(sim_map, r2n=True), nest=True, overwrite=True)
```
Scan the sky signal
```
all_name = "all_signal"
sky_name = "sky_signal"
# Clear any previous signal from the buffers
toast.tod.OpCacheClear(all_name).exec(data)
distmap = toast.map.DistPixels(
data,
nnz=len(args.mode),
dtype=np.float32,
)
distmap.read_healpix_fits("sim_map.fits")
toast.todmap.OpSimScan(distmap=distmap, out=all_name).exec(data)
# Copy the sky signal, just in case we need it later
toast.tod.OpCacheCopy(input=all_name, output=sky_name, force=True).exec(data)
memreport("After scanning sky signal", comm.comm_world)
```
## Noise
Simulate noise and make a copy of signal+noise in case we need it later
```
copy_name = "copy_signal"
toast.tod.OpSimNoise(out=all_name, realization=0).exec(data)
toast.tod.OpCacheCopy(input=all_name, output=copy_name, force=True).exec(data)
memreport("After simulating noise", comm.comm_world)
```
## Your own operator here
Here we define an empty operator you can work with
```
class MyOperator(toast.Operator):
def __init__(self, name="signal"):
""" Arguments:
name(str) : Cache prefix to operate on
"""
self._name = name
def exec(self, data):
# We loop here over all local data but do nothing with it.
for obs in data.obs:
tod = obs["tod"]
for det in tod.local_dets:
signal = tod.local_signal(det, self._name)
# Do operations in-place
signal *= 1.0
#signal[:] = (some other data)
```
Then we apply the operator to the data
```
toast.tod.OpCacheCopy(input=copy_name, output=all_name, force=True).exec(data)
MyOperator(name=all_name).exec(data)
memreport("After my operator", comm.comm_world)
```
Plot a short segment of the signal before and after the operator
```
tod = data.obs[0]["tod"]
times = tod.local_times()
fig = plt.figure(figsize=[12, 8])
for idet, det in enumerate(tod.local_dets):
cflags = tod.local_common_flags()
before = tod.local_signal(det, copy_name)
after = tod.local_signal(det, all_name)
ind = slice(0, 1000)
# Flag out turnarounds
good = cflags[ind] == 0
ax = fig.add_subplot(8, 8, 1 + idet)
ax.set_title(det)
ax.plot(times[ind][good], before[ind][good], '.', label="before")
ax.plot(times[ind][good], after[ind][good], '.', label="after")
ax.legend(bbox_to_anchor=(1.1, 1.00))
fig.subplots_adjust(hspace=0.6)
```
## Make a map
Destripe the signal and make a map. We use the nascent TOAST mapmaker because it can be run in serial mode without MPI. The TOAST mapmaker is still significantly slower so production runs should used `libMadam`.
```
# Always begin mapmaking by copying the simulated signal.
destriped_name = "destriped"
toast.tod.OpCacheCopy(
input=all_name,
output=destriped_name,
force=True
).exec(data)
mapmaker = toast.todmap.OpMapMaker(
nside=args.nside,
nnz=3,
name=destriped_name,
outdir=args.outdir,
outprefix="toast_test_",
baseline_length=10,
iter_max=15,
use_noise_prior=False,
)
mapmaker.exec(data)
memreport("After map making", comm.comm_world)
```
Plot a segment of the timelines
```
tod = data.obs[0]["tod"]
times = tod.local_times()
fig = plt.figure(figsize=[12, 8])
for idet, det in enumerate(tod.local_dets):
sky = tod.local_signal(det, sky_name)
full = tod.local_signal(det, all_name)
destriped = tod.local_signal(det, destriped_name)
ind = slice(0, 1000)
ax = fig.add_subplot(8, 8, 1 + idet)
ax.set_title(det)
ax.plot(times[ind], sky[ind], '.', label="sky", zorder=100)
ax.plot(times[ind], full[ind] - sky[ind], '.', label="noise")
ax.plot(times[ind], full[ind] - destriped[ind], '.', label="baselines")
ax.legend(bbox_to_anchor=(1.1, 1.00))
fig.subplots_adjust(hspace=0.6)
fig = plt.figure(figsize=[12, 8])
for idet, det in enumerate(tod.local_dets):
sky = tod.local_signal(det, sky_name)
full = tod.local_signal(det, copy_name)
destriped = tod.local_signal(det, destriped_name)
ax = fig.add_subplot(8, 8, 1 + idet)
ax.set_title(det)
#plt.plot(times[ind], sky[ind], '-', label="signal", zorder=100)
plt.plot(times, full - sky, '.', label="noise")
plt.plot(times, full - destriped, '.', label="baselines")
ax.legend(bbox_to_anchor=(1.1, 1.00))
fig.subplots_adjust(hspace=.6)
plt.figure(figsize=[16, 8])
hitmap = hp.read_map("litebird_out/toast_test_hits.fits", verbose=False)
hitmap[hitmap == 0] = hp.UNSEEN
hp.mollview(hitmap, sub=[2, 2, 1], title="hits")
binmap = hp.read_map("litebird_out/toast_test_binned.fits", verbose=False)
binmap[binmap == 0] = hp.UNSEEN
hp.mollview(binmap, sub=[2, 2, 2], title="binned map", cmap="coolwarm")
# Fix the plotting range for input signal and the destriped map
amp = 5.0
destriped = hp.read_map("litebird_out/toast_test_destriped.fits", verbose=False)
destriped[destriped == 0] = hp.UNSEEN
# Remove monopole
good = destriped != hp.UNSEEN
destriped[good] -= np.median(destriped[good])
hp.mollview(destriped, sub=[2, 2, 3], title="destriped map", cmap="coolwarm", min=-amp, max=amp)
inmap = hp.read_map("sim_map.fits", verbose=False)
inmap[hitmap == hp.UNSEEN] = hp.UNSEEN
hp.mollview(inmap, sub=[2, 2, 4], title="input map", cmap="coolwarm", min=-amp, max=amp)
# Plot the white noise covariance
plt.figure(figsize=[12, 8])
wcov = hp.read_map("litebird_out/toast_test_npp.fits", None)
wcov[:, wcov[0] == 0] = hp.UNSEEN
hp.mollview(wcov[0], sub=[3, 3, 1], title="II", cmap="coolwarm")
hp.mollview(wcov[1], sub=[3, 3, 2], title="IQ", cmap="coolwarm")
hp.mollview(wcov[2], sub=[3, 3, 3], title="IU", cmap="coolwarm")
hp.mollview(wcov[3], sub=[3, 3, 5], title="QQ", cmap="coolwarm")
hp.mollview(wcov[4], sub=[3, 3, 6], title="QU", cmap="coolwarm")
hp.mollview(wcov[5], sub=[3, 3, 9], title="UU", cmap="coolwarm")
```
## Filter & bin
A filter-and-bin mapmaker is easily created by combining TOAST filter operators and running the mapmaker without destriping:
```
filtered_name = "filtered"
toast.tod.OpCacheCopy(input=copy_name, output=filtered_name, force=True).exec(data)
toast.tod.OpPolyFilter(order=3, name=filtered_name).exec(data)
mapmaker = toast.todmap.OpMapMaker(
nside=args.nside,
nnz=len(args.mode),
name=filtered_name,
outdir=args.outdir,
outprefix="toast_test_filtered_",
baseline_length=None,
)
mapmaker.exec(data)
plt.figure(figsize=[16, 8])
binmap = hp.read_map("litebird_out/toast_test_binned.fits", verbose=False)
binmap[binmap == 0] = hp.UNSEEN
hp.mollview(binmap, sub=[1, 3, 1], title="binned map", cmap="coolwarm")
filtered_map = hp.read_map("litebird_out/toast_test_filtered_binned.fits", verbose=False)
filtered_map[filtered_map == 0] = hp.UNSEEN
hp.mollview(filtered_map, sub=[1, 3, 2], title="filtered map", cmap="coolwarm")
inmap = hp.read_map("sim_map.fits", verbose=False)
inmap[binmap == hp.UNSEEN] = hp.UNSEEN
hp.mollview(inmap, sub=[1, 3, 3], title="input map", cmap="coolwarm")
```
| github_jupyter |
# Numpy
Fundamental building block of scientific Python.
* Main attraction: Powerful and highly flexible array object; your new ubiquitous working unit.
* Set of most common mathematical utilities (constants, random numbers, linear algebra functions).
## Import
```
# imports
import numpy as np # It will be used a lot, so the shorthand is helpful.
import matplotlib.pyplot as plt # Same here.
%matplotlib inline
# these can be useful if you plan on using the respective functions a lot:
np.random.seed(42) # Seeding is important to replicate results when using random numbers.
rnd = np.random.random
sin = np.sin # Be careful to no write "sin = np.sin()"! Why?
cos = np.cos
RAD2DEG = 180.0/np.pi # Constants for quick conversion between radians (used by sin/cos) and degree
DEG2RAD = np.pi/180.0
```
## Numpy array basics
Every numpy array has some basic values that denote its format. Note that numpy array **cannot** change their size once they are created, but they **can** change their shape, i.e., an array will always hold the same number of elements, but their organization into rows and columns may change as desired.
* **ndarray.ndim:** The number of axes/dimensions of an array. The default matrix used for math problems is of dimensionality 2.
* **ndarray.shape:** A tuple of integers indicating the size of an array in each dimension. For a matrix with n rows and m columns, shape will be (n,m). The length of the shape tuple is therefore the rank, or number of dimensions, ndim.
* **ndarray.size:** The total number of elements of the array. This is equal to the product of the elements of shape.
* **ndarray.dtype:** The data type of the array elements. Defaults to 64 bit floating point values and can be set when the array is created.
(*see:* [Numpy basics](http://wiki.scipy.org/Tentative_NumPy_Tutorial#head-6a1bc005bd80e1b19f812e1e64e0d25d50f99fe2))
```
m = np.array([[1,2,3],
[4,5,6],
[7,8,9]], dtype=np.int32) # np.float32, np.float64, np.complex64, np.complex128
print m
print 'ndim: ', m.ndim, '\nshape:', m.shape, '\nsize: ', m.size, '\ndtype:', m.dtype
```
### Under the hood
* Numpy arrays believe in sharing is caring and will share their data with other arrays. Slicing does NOT return a new array, but instead a *view* on the data of another array:
```
s = m[1]
print 'BEFORE'
print s, 'slice', '\n'
print m, '\n'
s[0] = 0
print 'AFTER'
print s, 'slice' '\n'
print m, '\n'
```
* You can check whether an array actually owns its data by looking at its flags (you should understand *both* differences in the two flag settings):
```
print m.flags, '\n'
print s.flags
```
## Array creation
```
# helper function for examples below; plots the graphical depiction of a given numpy array
def showMatrix(X):
Y = np.array(np.array(X, ndmin=2)) # 1D -> 2D
vmin = min(np.min(Y), 0)
vmax = max(np.max(Y), 1)
plt.imshow(Y, interpolation='none', vmin=vmin, vmax=vmax, cmap=plt.cm.get_cmap('Blues'))
Z = np.zeros(9)
showMatrix(Z)
Z = np.zeros((5,9))
showMatrix(Z)
Z = np.ones(9)
showMatrix(Z)
Z = np.ones((5,9))
showMatrix(Z)
Z = np.array( [0,0,0,0,0,0,0,0,0] )
showMatrix(Z)
Z = np.array( [[0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0]] )
showMatrix(Z)
Z = np.arange(9) # the numpy arange function also allows floating point arguments
showMatrix(Z)
```
(*see also:* [linspace](http://wiki.scipy.org/Numpy_Example_List#linspace))
```
Z = np.arange(5*9).reshape(5,9)
showMatrix(Z)
```
- Reshape must not change the number of elements within the array.
- A vector of length ***n*** and a matrix of dimensions (1,***n***) ARE NOT THE SAME THING!
```
Z = np.random.uniform(0,1,9) # args: min, max, no. of elements
showMatrix(Z)
Z = np.random.uniform(0, 1, (5, 9))
showMatrix(Z)
```
(*see:* [Numpy array creation](http://www.labri.fr/perso/nrougier/teaching/numpy/numpy.html#creation) & [Numpy array reshaping](http://www.labri.fr/perso/nrougier/teaching/numpy/numpy.html#reshaping))
## Array slicing
```
# single element
Z = np.zeros((5, 9))
Z[1,1] = 1
showMatrix(Z)
# single row
Z = np.zeros((5, 9))
Z[1,:] = 1
showMatrix(Z)
# single column
Z = np.zeros((5, 9))
Z[:,1] = 1
showMatrix(Z)
# specific area
Z = np.zeros((5, 9))
Z[2:4,2:6] = 1 # for each dimension format is always: <from:to:step> (with step being optional)
showMatrix(Z)
# every second column
Z = np.zeros((5, 9))
Z[:,::2] = 1 # for each dimension format is always: <from:to:step> (with step being optional)
showMatrix(Z)
# indices can be negative
Z = np.arange(10)
print ">>> Z[-1]: ", Z[-1] # start indexing at the back
print ">>> Z[3:-3]:", Z[3:-3] # slice of array center
print ">>> Z[::-1]:", Z[::-1] # quickly reverse an array
```
(*see:* [Numpy array slicing](http://www.labri.fr/perso/nrougier/teaching/numpy/numpy.html#slicing))
## Broadcasting
Arithmetic operations applied to two Numpy arrays of different dimensions leads to 'broadcasting', i.e., filling up the missing values to allow the operation if possible. This includes:
* Adding/subtracting/etc. a single value to a matrix.
* Adding/subtracting/etc. a column/row vector to a matrix.
* Adding/subtracting/etc. a column and a row vector.
**NOTE:** Multiplying with \* WILL ALSO BE APPLIED elementwise! Use **[np.dot()](http://docs.scipy.org/doc/numpy/reference/generated/numpy.dot.html)** for actual matrix multiplication!
**FUN FACT:** Truth value checks will also applied elementwise.
(*see:* [Numpy broadcasting](http://www.labri.fr/perso/nrougier/teaching/numpy/numpy.html#broadcasting))
## Exercises
1. Select a tile-pattern subset of a 5x9 matrix like this:

2. ..and like this:

3. ..and also like this:

4. Adapt the code for No.3 so that it works with arrays of arbitrary dimensions.
5. Write the code that perfoms the operation depicted below ([source](http://www.labri.fr/perso/nrougier/teaching/numpy/numpy.html#broadcasting)). Parameterize your code and use the above utility function to plot the final matrix in dimensions 8x2 and 256x64.

6. Write a function that subtracts the mean from a given matrix (arbitrary dimensions).
7. Write a function that gradually weighs the rows of a given matrix from top to bottom (arbitrary dimensions).
8. Write one line that checks whether there are any values smaller than 0 within a given array.
9. Create a two dimensional array containing the values 0..9.
1. Reverse the order of the rows of the matrix using a single slice.
2. Reverse the order of the columns of the matrix using a single slice.
3. Reverse the order of both the rows and the columns of the matrix using a single slice.
10. Check the [documentation](http://docs.scipy.org/doc/): What is the difference between **np.max()** and **np.nanmax()**?
1. Think of two cases where it would be important to use one over the other!
2. Explain how you can find both functions using only the numpy documentation itself.
```
#-#-# EXC_NUMPY: YOUR CODE HERE #-#-#
```
## Links
* [Quick reference (types, array handling)](http://www.labri.fr/perso/nrougier/teaching/numpy/numpy.html#quick-references)
* [Tentative numpy tutorial](http://wiki.scipy.org/Tentative_NumPy_Tutorial)
| github_jupyter |
# Add loops to the designed bound states
### Imports
```
%load_ext lab_black
# Python standard library
from glob import glob
import os
import socket
import sys
# 3rd party library imports
import dask
import matplotlib.pyplot as plt
import pandas as pd
import pyrosetta
import numpy as np
import scipy
import seaborn as sns
from tqdm.auto import tqdm # jupyter compatible progress bar
tqdm.pandas() # link tqdm to pandas
# Notebook magic
# save plots in the notebook
%matplotlib inline
# reloads modules automatically before executing cells
%load_ext autoreload
%autoreload 2
print(f"running in directory: {os.getcwd()}") # where are we?
print(f"running on node: {socket.gethostname()}") # what node are we on?
```
### Set working directory to the root of the crispy_shifty repo
TODO set to projects dir
```
os.chdir("/home/pleung/projects/crispy_shifty")
# os.chdir("/projects/crispy_shifty")
```
### Add loops between chA and chB
TODO
```
from crispy_shifty.utils.io import gen_array_tasks
simulation_name = "01_loop_bound_states"
design_list_file = os.path.join(
os.getcwd(), "projects/crispy_shifties/00_design_bound_states/designed_states.list"
)
output_path = os.path.join(os.getcwd(), f"projects/crispy_shifties/{simulation_name}")
options = " ".join(
[
"out:level 200",
"corrections::beta_nov16 true",
"indexed_structure_store:fragment_store /net/databases/VALL_clustered/connect_chains/ss_grouped_vall_helix_shortLoop.h5",
]
)
gen_array_tasks(
distribute_func="crispy_shifty.protocols.looping.loop_bound_state",
design_list_file=design_list_file,
output_path=output_path,
queue="medium",
memory="4G",
nstruct=1,
nstruct_per_task=1,
options=options,
simulation_name=simulation_name,
)
!sbatch -a 1-$(cat /mnt/home/pleung/projects/crispy_shifty/projects/crispy_shifties/01_loop_bound_states/tasks.cmds | wc -l) /mnt/home/pleung/projects/crispy_shifty/projects/crispy_shifties/01_loop_bound_states/run.sh
```
### Collect scorefiles of designed bound states and concatenate
TODO change to projects dir
```
sys.path.insert(0, "~/projects/crispy_shifty") # TODO
from crispy_shifty.utils.io import collect_score_file
simulation_name = "01_loop_bound_states"
output_path = os.path.join(os.getcwd(), f"projects/crispy_shifties/{simulation_name}")
if not os.path.exists(os.path.join(output_path, "scores.json")):
collect_score_file(output_path, "scores")
```
### Load resulting concatenated scorefile
TODO change to projects dir
```
sys.path.insert(0, "~/projects/crispy_shifty") # TODO
from crispy_shifty.utils.io import parse_scorefile_linear
output_path = os.path.join(os.getcwd(), f"projects/crispy_shifties/{simulation_name}")
scores_df = parse_scorefile_linear(os.path.join(output_path, "scores.json"))
scores_df = scores_df.convert_dtypes()
```
### Setup for plotting
```
sns.set(
context="talk",
font_scale=1, # make the font larger; default is pretty small
style="ticks", # make the background white with black lines
palette="colorblind", # a color palette that is colorblind friendly!
)
```
### Data exploration
Gonna remove the Rosetta sfxn scoreterms for now
```
from crispy_shifty.protocols.design import beta_nov16_terms
scores_df = scores_df[
[term for term in scores_df.columns if term not in beta_nov16_terms]
]
print(len(scores_df))
scores_df.columns
```
### Save a list of outputs
```
simulation_name = "01_loop_bound_states"
output_path = os.path.join(os.getcwd(), f"projects/crispy_shifties/{simulation_name}")
with open(os.path.join(output_path, "looped_states.list"), "w") as f:
for path in tqdm(scores_df.index):
print(path, file=f)
```
### Prototyping blocks
test `loop_bound_state`
```
%%time
import pyrosetta
pyrosetta.init(
"-corrections::beta_nov16 -indexed_structure_store:fragment_store /net/databases/VALL_clustered/connect_chains/ss_grouped_vall_helix_shortLoop.h5 -precompute_ig true"
)
sys.path.insert(0, "~/projects/crispy_shifty/") # TODO projects
from crispy_shifty.protocols.looping import loop_bound_state
t = loop_bound_state(
None,
**{
'pdb_path': '/mnt/home/pleung/projects/crispy_shifty/projects/crispy_shifties/00_design_bound_states/decoys/0000/00_design_bound_states_8bb4790d4d0c4797ba0404775b3dcbab.pdb.bz2',
}
)
for i, tppose in enumerate(t):
tppose.pose.dump_pdb(f"{i}.pdb")
import pyrosetta.distributed.viewer as viewer
ppose = pyrosetta.distributed.io.pose_from_file("test.pdb")
view = viewer.init(ppose, window_size=(1600, 1200))
view.add(viewer.setStyle())
view.add(viewer.setStyle(colorscheme="whiteCarbon", radius=0.10))
view.add(viewer.setHydrogenBonds())
view.add(viewer.setHydrogens(polar_only=True))
view.add(viewer.setDisulfides(radius=0.25))
view()
```
| github_jupyter |
Welcome to the [Object Detection API](https://github.com/tensorflow/models/tree/master/research/object_detection). This notebook will walk you step by step through the process of using a pre-trained model to detect objects in an image.
> **Important**: This tutorial is to help you through the first step towards using [Object Detection API](https://github.com/tensorflow/models/tree/master/research/object_detection) to build models. If you just just need an off the shelf model that does the job, see the [TFHub object detection example](https://colab.sandbox.google.com/github/tensorflow/hub/blob/master/examples/colab/object_detection.ipynb).
### Install
Make sure you have `pycocotools` installed
```
!pip install pycocotools tf_slim
```
Get `tensorflow/models` or `cd` to parent directory of the repository.
```
import os
import pathlib
if "models" in pathlib.Path.cwd().parts:
while "models" in pathlib.Path.cwd().parts:
os.chdir('..')
elif not pathlib.Path('models').exists():
!git clone --depth 1 --branch r2.3.0 https://github.com/tensorflow/models
```
Compile protobufs and install the object_detection package
```
%%bash
cd models/research/
protoc object_detection/protos/*.proto --python_out=.
%%bash
cd models/research
pip install .
```
### Imports
```
import numpy as np
import os
import six.moves.urllib as urllib
import sys
import tarfile
import tensorflow as tf
import zipfile
from collections import defaultdict
from io import StringIO
from matplotlib import pyplot as plt
from PIL import Image
from IPython.display import display
```
Import the object detection module.
```
from object_detection.utils import ops as utils_ops
from object_detection.utils import label_map_util
from object_detection.utils import visualization_utils as vis_util
```
Patches:
```
# patch tf1 into `utils.ops`
utils_ops.tf = tf.compat.v1
# Patch the location of gfile
tf.gfile = tf.io.gfile
```
# Model preparation
## Variables
Any model exported using the `export_inference_graph.py` tool can be loaded here simply by changing the path.
By default we use an "SSD with Mobilenet" model here. See the [detection model zoo](https://github.com/tensorflow/models/blob/master/research/object_detection/g3doc/detection_model_zoo.md) for a list of other models that can be run out-of-the-box with varying speeds and accuracies.
## Loader
```
def load_model(model_name):
base_url = 'http://download.tensorflow.org/models/object_detection/'
model_file = model_name + '.tar.gz'
model_dir = tf.keras.utils.get_file(
fname=model_name,
origin=base_url + model_file,
untar=True)
model_dir = pathlib.Path(model_dir)/"saved_model"
model = tf.saved_model.load(str(model_dir))
model = model.signatures['serving_default']
return model
```
## Loading label map
Label maps map indices to category names, so that when our convolution network predicts `5`, we know that this corresponds to `airplane`. Here we use internal utility functions, but anything that returns a dictionary mapping integers to appropriate string labels would be fine
```
# List of the strings that is used to add correct label for each box.
PATH_TO_LABELS = 'models/research/object_detection/data/mscoco_label_map.pbtxt'
category_index = label_map_util.create_category_index_from_labelmap(PATH_TO_LABELS, use_display_name=True)
```
For the sake of simplicity we will test on 2 images:
```
# If you want to test the code with your images, just add path to the images to the TEST_IMAGE_PATHS.
PATH_TO_TEST_IMAGES_DIR = pathlib.Path('models/research/object_detection/test_images')
TEST_IMAGE_PATHS = sorted(list(PATH_TO_TEST_IMAGES_DIR.glob("*.jpg")))
TEST_IMAGE_PATHS
```
# Detection
Load an object detection model:
```
model_name = 'ssd_mobilenet_v1_coco_2017_11_17'
detection_model = load_model(model_name)
```
Check the model's input signature, it expects a batch of 3-color images of type uint8:
```
print(detection_model.inputs)
```
And returns several outputs:
```
detection_model.output_dtypes
detection_model.output_shapes
```
Add a wrapper function to call the model, and cleanup the outputs:
```
def run_inference_for_single_image(model, image):
image = np.asarray(image)
# The input needs to be a tensor, convert it using `tf.convert_to_tensor`.
input_tensor = tf.convert_to_tensor(image)
# The model expects a batch of images, so add an axis with `tf.newaxis`.
input_tensor = input_tensor[tf.newaxis,...]
# Run inference
output_dict = model(input_tensor)
# All outputs are batches tensors.
# Convert to numpy arrays, and take index [0] to remove the batch dimension.
# We're only interested in the first num_detections.
num_detections = int(output_dict.pop('num_detections'))
output_dict = {key:value[0, :num_detections].numpy()
for key,value in output_dict.items()}
output_dict['num_detections'] = num_detections
# detection_classes should be ints.
output_dict['detection_classes'] = output_dict['detection_classes'].astype(np.int64)
# Handle models with masks:
if 'detection_masks' in output_dict:
# Reframe the the bbox mask to the image size.
detection_masks_reframed = utils_ops.reframe_box_masks_to_image_masks(
output_dict['detection_masks'], output_dict['detection_boxes'],
image.shape[0], image.shape[1])
detection_masks_reframed = tf.cast(detection_masks_reframed > 0.5,
tf.uint8)
output_dict['detection_masks_reframed'] = detection_masks_reframed.numpy()
return output_dict
```
Run it on each test image and show the results:
```
def show_inference(model, image_path):
# the array based representation of the image will be used later in order to prepare the
# result image with boxes and labels on it.
image_np = np.array(Image.open(image_path))
# Actual detection.
output_dict = run_inference_for_single_image(model, image_np)
# Visualization of the results of a detection.
vis_util.visualize_boxes_and_labels_on_image_array(
image_np,
output_dict['detection_boxes'],
output_dict['detection_classes'],
output_dict['detection_scores'],
category_index,
instance_masks=output_dict.get('detection_masks_reframed', None),
use_normalized_coordinates=True,
line_thickness=8)
display(Image.fromarray(image_np))
for image_path in TEST_IMAGE_PATHS:
show_inference(detection_model, image_path)
```
## Instance Segmentation
```
model_name = "mask_rcnn_inception_resnet_v2_atrous_coco_2018_01_28"
masking_model = load_model(model_name)
```
The instance segmentation model includes a `detection_masks` output:
```
masking_model.output_shapes
for image_path in TEST_IMAGE_PATHS:
show_inference(masking_model, image_path)
```
| github_jupyter |
```
# Plot various data
import pandas as pd
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import axes3d
import math
figsize=(4.5, 3)
# Only read this one CSV.
# Change this, if you are interested in different ciphers
data = pd.read_csv("DHE-RSA-AES128-GCM-SHA256.csv")
data1C = data[data['numConns'] == 1]
#data1C
data1C.loc[[0]]['dataLen'] = 500
#data1C
# Normalize the data
for i,r in data1C.iterrows():
r['openssl'] /= r['numBytes']
r['denseMap'] /= r['numPkts']
r['tbb'] /= r['numPkts']
r['siphash'] /= r['numPkts']
#plt.plot(data1C['dataLen'], data1C['openssl'])
# Plot the cost of siphash
fig = plt.figure(figsize=figsize, dpi=300)
plt.plot(data1C['dataLen'], data1C['siphash'])
plt.semilogx()
plt.xlabel('Data Length')
plt.ylabel('Cycles')
plt.title("Cost of siphash by data length")
plt.grid(b=True)
plt.savefig('output/siphash.pdf',bbox_inches='tight')
plt.show()
# Plot the cost of TBB's shared map
tbbMean = data.groupby(['numConns'])['tbb'].mean()
steps = data['numConns'].unique()
[print("{" + str(x) + "," + str(tbbMean[x]) + "},", end='') for x in data['numConns'].unique()]
fig = plt.figure(figsize=figsize, dpi=300)
plt.plot(steps, tbbMean)
plt.ylim([0,40000])
plt.xlabel('#Connections')
plt.ylabel('Cycles')
plt.title("Cost of shared map by number of connections")
plt.grid(b=True)
plt.savefig('output/tbb.pdf',bbox_inches='tight')
plt.show()
# Plot the cost of the memory allocation
memMean = data.groupby(['numConns'])['memory'].mean()
#print(memMean)
memMeanPC = [memMean[x] / x for x in data['numConns'].unique()]
#[print("{" + str(x) + "," + str(memMean[x]) + "},", end='') for x in data['numConns'].unique()]
fig = plt.figure(figsize=figsize, dpi=300)
plt.plot(steps, memMean)
plt.ylim([0,15000])
plt.xlabel('#Connections')
plt.ylabel('Cycles')
plt.title("Cost of memory allocation by number of connections")
plt.grid(b=True)
plt.savefig('output/memory.pdf',bbox_inches='tight')
plt.show()
# Build logarithmic data
dataDense = data
for _,r in dataDense.iterrows():
r['denseMap'] /= r['numPkts']
r['dataLen'] = math.log10(r['dataLen'])
r['numConns'] = math.log(r['numConns'],2)
# Plot the denseMap cost in 3D
denseMapMean = dataDense.groupby(['numConns'])['denseMap'].mean()
steps = data['numConns'].unique()
import numpy as np
#X = dataDense['dataLen'].apply(math.log10)
X = [np.arange(2,8)] * 6
#Y = dataDense['numConns'].apply(lambda x: math.log(x,2))
Y = [[x] * 6 for x in range(1,7)]
#Z = dataDense['denseMap']
Z = dataDense.pivot(index='numConns', columns='dataLen', values='denseMap').as_matrix()
#fig = plt.figure(figsize=(16, 10), dpi=80)
fig = plt.figure(figsize=(7, 4), dpi=300)
#fig = plt.figure(figsize=figsize, dpi=300)
ax = fig.gca(projection='3d')
ax.set_xlabel('#Bytes 10^x')
ax.set_ylabel('#Connections 2^x')
ax.set_zlabel('Cycles')
ax.set_zlim(0,600)
#ax.scatter(X, Y, Z)
ax.plot_wireframe(X,Y,Z)
plt.title("Cost of state map by number of connections")
plt.savefig('output/denseMap.pdf',bbox_inches='tight')
plt.show()
#plt.plot(steps, denseMapMean)
#plt.ylim([0,500])
#plt.xlabel('#Connections')
#plt.ylabel('Cycles')
#plt.title("Cost of state map by number of connections")
#plt.show()
#
#dataDense
```
| github_jupyter |
# Working with imbalanced data
In machine learning it is quite usual to have to deal with imbalanced dataset. This is particularly true in online learning for tasks such as fraud detection and spam classification. In these two cases, which are binary classification problems, there are usually many more 0s than 1s, which generally hinders the performance of the classifiers we thrown at them.
As an example we'll use the credit card dataset available in `river`. We'll first use a `collections.Counter` to count the number of 0s and 1s in order to get an idea of the class balance.
```
import collections
from river import datasets
X_y = datasets.CreditCard()
counts = collections.Counter(y for _, y in X_y)
for c, count in counts.items():
print(f'{c}: {count} ({count / sum(counts.values()):.5%})')
```
## Baseline
The dataset is quite unbalanced. For each 1 there are about 578 0s. Let's now train a logistic regression with default parameters and see how well it does. We'll measure the ROC AUC score.
```
from river import linear_model
from river import metrics
from river import evaluate
from river import preprocessing
X_y = datasets.CreditCard()
model = (
preprocessing.StandardScaler() |
linear_model.LogisticRegression()
)
metric = metrics.ROCAUC()
evaluate.progressive_val_score(X_y, model, metric)
```
## Importance weighting
The performance is already quite acceptable, but as we will now see we can do even better. The first thing we can do is to add weight to the 1s by using the `weight_pos` argument of the `Log` loss function.
```
from river import optim
model = (
preprocessing.StandardScaler() |
linear_model.LogisticRegression(
loss=optim.losses.Log(weight_pos=5)
)
)
metric = metrics.ROCAUC()
evaluate.progressive_val_score(X_y, model, metric)
```
## Focal loss
The deep learning for object detection community has produced a special loss function for imbalaced learning called [focal loss](https://arxiv.org/pdf/1708.02002.pdf). We are doing binary classification, so we can plug the binary version of focal loss into our logistic regression and see how well it fairs.
```
model = (
preprocessing.StandardScaler() |
linear_model.LogisticRegression(loss=optim.losses.BinaryFocalLoss(2, 1))
)
metric = metrics.ROCAUC()
evaluate.progressive_val_score(X_y, model, metric)
```
## Under-sampling the majority class
Adding importance weights only works with gradient-based models (which includes neural networks). A more generic, and potentially more effective approach, is to use undersamplig and oversampling. As an example, we'll under-sample the stream so that our logistic regression encounter 20% of 1s and 80% of 0s. Under-sampling has the additional benefit of requiring less training steps, and thus reduces the total training time.
```
from river import imblearn
model = (
preprocessing.StandardScaler() |
imblearn.RandomUnderSampler(
classifier=linear_model.LogisticRegression(),
desired_dist={0: .8, 1: .2},
seed=42
)
)
metric = metrics.ROCAUC()
evaluate.progressive_val_score(X_y, model, metric)
```
The `RandomUnderSampler` class is a wrapper for classifiers. This is represented by a rectangle around the logistic regression bubble when we draw the model.
```
model.draw()
```
## Over-sampling the minority class
We can also attain the same class distribution by over-sampling the minority class. This will come at cost of having to train with more samples.
```
model = (
preprocessing.StandardScaler() |
imblearn.RandomOverSampler(
classifier=linear_model.LogisticRegression(),
desired_dist={0: .8, 1: .2},
seed=42
)
)
metric = metrics.ROCAUC()
evaluate.progressive_val_score(X_y, model, metric)
```
## Sampling with a desired sample size
The downside of both `RandomUnderSampler` and `RandomOverSampler` is that you don't have any control on the amount of data the classifier trains on. The number of samples is adjusted so that the target distribution can be attained, either by under-sampling or over-sampling. However, you can do both at the same time and choose how much data the classifier will see. To do so, we can use the `RandomSampler` class. In addition to the desired class distribution, we can specify how much data to train on. The samples will both be under-sampled and over-sampled in order to fit your constraints. This is powerful because it allows you to control both the class distribution and the size of the training data (and thus the training time). In the following example we'll set it so that the model will train with 1 percent of the data.
```
model = (
preprocessing.StandardScaler() |
imblearn.RandomSampler(
classifier=linear_model.LogisticRegression(),
desired_dist={0: .8, 1: .2},
sampling_rate=.01,
seed=42
)
)
metric = metrics.ROCAUC()
evaluate.progressive_val_score(X_y, model, metric)
```
## Hybrid approach
As you might have guessed by now, nothing is stopping you from mixing imbalanced learning methods together. As an example, let's combine `sampling.RandomUnderSampler` and the `weight_pos` parameter from the `optim.losses.Log` loss function.
```
model = (
preprocessing.StandardScaler() |
imblearn.RandomUnderSampler(
classifier=linear_model.LogisticRegression(
loss=optim.losses.Log(weight_pos=5)
),
desired_dist={0: .8, 1: .2},
seed=42
)
)
metric = metrics.ROCAUC()
evaluate.progressive_val_score(X_y, model, metric)
```
| github_jupyter |
# Training Neural Networks
The network we built in the previous part isn't so smart, it doesn't know anything about our handwritten digits. Neural networks with non-linear activations work like universal function approximators. There is some function that maps your input to the output. For example, images of handwritten digits to class probabilities. The power of neural networks is that we can train them to approximate this function, and basically any function given enough data and compute time.
<img src="assets/function_approx.png" width=500px>
At first the network is naive, it doesn't know the function mapping the inputs to the outputs. We train the network by showing it examples of real data, then adjusting the network parameters such that it approximates this function.
To find these parameters, we need to know how poorly the network is predicting the real outputs. For this we calculate a **loss function** (also called the cost), a measure of our prediction error. For example, the mean squared loss is often used in regression and binary classification problems
$$
\large \ell = \frac{1}{2n}\sum_i^n{\left(y_i - \hat{y}_i\right)^2}
$$
where $n$ is the number of training examples, $y_i$ are the true labels, and $\hat{y}_i$ are the predicted labels.
By minimizing this loss with respect to the network parameters, we can find configurations where the loss is at a minimum and the network is able to predict the correct labels with high accuracy. We find this minimum using a process called **gradient descent**. The gradient is the slope of the loss function and points in the direction of fastest change. To get to the minimum in the least amount of time, we then want to follow the gradient (downwards). You can think of this like descending a mountain by following the steepest slope to the base.
<img src='assets/gradient_descent.png' width=350px>
## Backpropagation
For single layer networks, gradient descent is straightforward to implement. However, it's more complicated for deeper, multilayer neural networks like the one we've built. Complicated enough that it took about 30 years before researchers figured out how to train multilayer networks.
Training multilayer networks is done through **backpropagation** which is really just an application of the chain rule from calculus. It's easiest to understand if we convert a two layer network into a graph representation.
<img src='assets/backprop_diagram.png' width=550px>
In the forward pass through the network, our data and operations go from bottom to top here. We pass the input $x$ through a linear transformation $L_1$ with weights $W_1$ and biases $b_1$. The output then goes through the sigmoid operation $S$ and another linear transformation $L_2$. Finally we calculate the loss $\ell$. We use the loss as a measure of how bad the network's predictions are. The goal then is to adjust the weights and biases to minimize the loss.
To train the weights with gradient descent, we propagate the gradient of the loss backwards through the network. Each operation has some gradient between the inputs and outputs. As we send the gradients backwards, we multiply the incoming gradient with the gradient for the operation. Mathematically, this is really just calculating the gradient of the loss with respect to the weights using the chain rule.
$$
\large \frac{\partial \ell}{\partial W_1} = \frac{\partial L_1}{\partial W_1} \frac{\partial S}{\partial L_1} \frac{\partial L_2}{\partial S} \frac{\partial \ell}{\partial L_2}
$$
**Note:** I'm glossing over a few details here that require some knowledge of vector calculus, but they aren't necessary to understand what's going on.
We update our weights using this gradient with some learning rate $\alpha$.
$$
\large W^\prime_1 = W_1 - \alpha \frac{\partial \ell}{\partial W_1}
$$
The learning rate $\alpha$ is set such that the weight update steps are small enough that the iterative method settles in a minimum.
## Losses in PyTorch
Let's start by seeing how we calculate the loss with PyTorch. Through the `nn` module, PyTorch provides losses such as the cross-entropy loss (`nn.CrossEntropyLoss`). You'll usually see the loss assigned to `criterion`. As noted in the last part, with a classification problem such as MNIST, we're using the softmax function to predict class probabilities. With a softmax output, you want to use cross-entropy as the loss. To actually calculate the loss, you first define the criterion then pass in the output of your network and the correct labels.
Something really important to note here. Looking at [the documentation for `nn.CrossEntropyLoss`](https://pytorch.org/docs/stable/nn.html#torch.nn.CrossEntropyLoss),
> This criterion combines `nn.LogSoftmax()` and `nn.NLLLoss()` in one single class.
>
> The input is expected to contain scores for each class.
This means we need to pass in the raw output of our network into the loss, not the output of the softmax function. This raw output is usually called the *logits* or *scores*. We use the logits because softmax gives you probabilities which will often be very close to zero or one but floating-point numbers can't accurately represent values near zero or one ([read more here](https://docs.python.org/3/tutorial/floatingpoint.html)). It's usually best to avoid doing calculations with probabilities, typically we use log-probabilities.
```
import torch
from torch import nn
import torch.nn.functional as F
from torchvision import datasets, transforms
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,)),
])
# Download and load the training data
trainset = datasets.MNIST('~/.pytorch/MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
```
### Note
If you haven't seen `nn.Sequential` yet, please finish the end of the Part 2 notebook.
```
# Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10))
# Define the loss
criterion = nn.CrossEntropyLoss()
# Get our data
images, labels = next(iter(trainloader))
# Flatten images
images = images.view(images.shape[0], -1)
# Forward pass, get our logits
logits = model(images)
# Calculate the loss with the logits and the labels
loss = criterion(logits, labels)
print(loss)
```
In my experience it's more convenient to build the model with a log-softmax output using `nn.LogSoftmax` or `F.log_softmax` ([documentation](https://pytorch.org/docs/stable/nn.html#torch.nn.LogSoftmax)). Then you can get the actual probabilities by taking the exponential `torch.exp(output)`. With a log-softmax output, you want to use the negative log likelihood loss, `nn.NLLLoss` ([documentation](https://pytorch.org/docs/stable/nn.html#torch.nn.NLLLoss)).
>**Exercise:** Build a model that returns the log-softmax as the output and calculate the loss using the negative log likelihood loss. Note that for `nn.LogSoftmax` and `F.log_softmax` you'll need to set the `dim` keyword argument appropriately. `dim=0` calculates softmax across the rows, so each column sums to 1, while `dim=1` calculates across the columns so each row sums to 1. Think about what you want the output to be and choose `dim` appropriately.
```
# TODO: Build a feed-forward network
model = nn.Sequential(nn.Linear(784,128),nn.ReLU(),
nn.Linear(128,64),nn.ReLU(),
nn.Linear(64,10),
nn.LogSoftmax(dim = 1))
# TODO: Define the loss
criterion = nn.NLLLoss()
### Run this to check your work
# Get our data
images, labels = next(iter(trainloader))
# Flatten images
images = images.view(images.shape[0], -1)
# Forward pass, get our logits
logits = model(images)
# Calculate the loss with the logits and the labels
loss = criterion(logits, labels)
print(loss)
```
## Autograd
Now that we know how to calculate a loss, how do we use it to perform backpropagation? Torch provides a module, `autograd`, for automatically calculating the gradients of tensors. We can use it to calculate the gradients of all our parameters with respect to the loss. Autograd works by keeping track of operations performed on tensors, then going backwards through those operations, calculating gradients along the way. To make sure PyTorch keeps track of operations on a tensor and calculates the gradients, you need to set `requires_grad = True` on a tensor. You can do this at creation with the `requires_grad` keyword, or at any time with `x.requires_grad_(True)`.
You can turn off gradients for a block of code with the `torch.no_grad()` content:
```python
x = torch.zeros(1, requires_grad=True)
>>> with torch.no_grad():
... y = x * 2
>>> y.requires_grad
False
```
Also, you can turn on or off gradients altogether with `torch.set_grad_enabled(True|False)`.
The gradients are computed with respect to some variable `z` with `z.backward()`. This does a backward pass through the operations that created `z`.
```
x = torch.randn(2,2, requires_grad=True)
print(x)
y = x**2
print(y)
y = x+3
print(y.grad_fn)
y = x*3
print(y.grad_fn)
y = x-3
print(y.grad_fn)
```
Below we can see the operation that created `y`, a power operation `PowBackward0`.
```
## grad_fn shows the function that generated this variable
print(y.grad_fn)
```
The autograd module keeps track of these operations and knows how to calculate the gradient for each one. In this way, it's able to calculate the gradients for a chain of operations, with respect to any one tensor. Let's reduce the tensor `y` to a scalar value, the mean.
```
z = y.mean()
print(z)
```
You can check the gradients for `x` and `y` but they are empty currently.
```
print(x.grad)
```
To calculate the gradients, you need to run the `.backward` method on a Variable, `z` for example. This will calculate the gradient for `z` with respect to `x`
$$
\frac{\partial z}{\partial x} = \frac{\partial}{\partial x}\left[\frac{1}{n}\sum_i^n x_i^2\right] = \frac{x}{2}
$$
```
z.backward()
print(x.grad)
print(x/2)
```
These gradients calculations are particularly useful for neural networks. For training we need the gradients of the cost with respect to the weights. With PyTorch, we run data forward through the network to calculate the loss, then, go backwards to calculate the gradients with respect to the loss. Once we have the gradients we can make a gradient descent step.
## Loss and Autograd together
When we create a network with PyTorch, all of the parameters are initialized with `requires_grad = True`. This means that when we calculate the loss and call `loss.backward()`, the gradients for the parameters are calculated. These gradients are used to update the weights with gradient descent. Below you can see an example of calculating the gradients using a backwards pass.
```
# Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
images, labels = next(iter(trainloader))
images = images.view(images.shape[0], -1)
logits = model(images)
loss = criterion(logits, labels)
print('Before backward pass: \n', model[0].weight.grad)
loss.backward()
print('After backward pass: \n', model[0].weight.grad)
```
## Training the network!
There's one last piece we need to start training, an optimizer that we'll use to update the weights with the gradients. We get these from PyTorch's [`optim` package](https://pytorch.org/docs/stable/optim.html). For example we can use stochastic gradient descent with `optim.SGD`. You can see how to define an optimizer below.
```
from torch import optim
# Optimizers require the parameters to optimize and a learning rate
optimizer = optim.SGD(model.parameters(), lr=0.01)
```
Now we know how to use all the individual parts so it's time to see how they work together. Let's consider just one learning step before looping through all the data. The general process with PyTorch:
* Make a forward pass through the network
* Use the network output to calculate the loss
* Perform a backward pass through the network with `loss.backward()` to calculate the gradients
* Take a step with the optimizer to update the weights
Below I'll go through one training step and print out the weights and gradients so you can see how it changes. Note that I have a line of code `optimizer.zero_grad()`. When you do multiple backwards passes with the same parameters, the gradients are accumulated. This means that you need to zero the gradients on each training pass or you'll retain gradients from previous training batches.
```
print('Initial weights - ', model[0].weight)
images, labels = next(iter(trainloader))
images.resize_(64, 784)
# Clear the gradients, do this because gradients are accumulated
optimizer.zero_grad()
# Forward pass, then backward pass, then update weights
output = model(images)
loss = criterion(output, labels)
loss.backward()
print('Gradient -', model[0].weight.grad)
# Take an update step and few the new weights
optimizer.step()
print('Updated weights - ', model[0].weight)
```
### Training for real
Now we'll put this algorithm into a loop so we can go through all the images. Some nomenclature, one pass through the entire dataset is called an *epoch*. So here we're going to loop through `trainloader` to get our training batches. For each batch, we'll doing a training pass where we calculate the loss, do a backwards pass, and update the weights.
>**Exercise:** Implement the training pass for our network. If you implemented it correctly, you should see the training loss drop with each epoch.
```
## Your solution here
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
optimizer = optim.SGD(model.parameters(), lr=0.003)
epochs = 5
for e in range(epochs):
running_loss = 0
for images, labels in trainloader:
# Flatten MNIST images into a 784 long vector
images = images.view(images.shape[0], -1)
# TODO: Training pass
optimizer.zero_grad()
output = model.forward(images)
loss = criterion(output, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
else:
print(f"Training loss: {running_loss/len(trainloader)}")
```
With the network trained, we can check out it's predictions.
```
%matplotlib inline
import helper
images, labels = next(iter(trainloader))
img = images[0].view(1, 784)
# Turn off gradients to speed up this part
with torch.no_grad():
logps = model(img)
# Output of the network are log-probabilities, need to take exponential for probabilities
ps = torch.exp(logps)
helper.view_classify(img.view(1, 28, 28), ps)
```
Now our network is brilliant. It can accurately predict the digits in our images. Next up you'll write the code for training a neural network on a more complex dataset.
| github_jupyter |
```
import qtm.encoding
import qtm.fubini_study
import qtm.nqubit
import qtm.constant
import qtm.base
import importlib
import qiskit
import numpy as np
import matplotlib.pyplot as plt
import sys
sys.path.insert(1, '../')
importlib.reload(qtm.base)
importlib.reload(qtm.constant)
importlib.reload(qtm.onequbit)
importlib.reload(qtm.nqubit)
# Init parameters
num_qubits = 5
# For arbitrary initial state
num_layers = 1
thetas_origin = np.random.uniform(
low=0, high=2*np.pi, size=num_qubits*num_layers*5)
# For determine GHZ state
theta = np.random.uniform(0, 2*np.pi)
# GHZ
thetas = thetas_origin.copy()
qc = qiskit.QuantumCircuit(num_qubits, num_qubits)
loss_values_ghz = []
thetass_ghz = []
for i in range(0, 100):
# fubini_study for binho_state is same for linear state
G = qtm.fubini_study.calculate_linear_state(qc.copy(), thetas, num_layers)
grad_loss = qtm.base.grad_loss(
qc,
qtm.nqubit.create_GHZchecker_binho,
thetas, r=1/2, s=np.pi/2, num_layers=num_layers, theta=theta)
thetas = np.real(thetas - qtm.constant.learning_rate *
(np.linalg.inv(G) @ grad_loss))
qc_copy = qtm.nqubit.create_GHZchecker_binho(
qc.copy(), thetas, num_layers, theta)
loss = qtm.base.loss_basis(qtm.base.measure(
qc_copy, list(range(qc_copy.num_qubits))))
loss_values_ghz.append(loss)
thetass_ghz.append(thetas)
# Plot loss value in 100 steps
plt.plot(loss_values_ghz, label='GHZ', linestyle='-')
plt.title('GHZchecker_binho')
plt.legend()
plt.xlabel("Iteration")
plt.ylabel("Loss value")
plt.savefig('GHZchecker_binho.png', format='png', dpi=600)
plt.show()
traces_ghz, fidelities_ghz = [], []
for thetas in thetass_ghz:
# Get |psi> = U_gen|000...>
qc = qiskit.QuantumCircuit(num_qubits, num_qubits)
qc = qtm.nqubit.create_binho_state(qc, thetas, num_layers=num_layers)
psi = qiskit.quantum_info.Statevector.from_instruction(qc)
rho_psi = qiskit.quantum_info.DensityMatrix(psi)
# Get |psi~> = U_target|000...>
qc1 = qiskit.QuantumCircuit(num_qubits, num_qubits)
qc1 = qtm.nqubit.create_ghz_state(qc1, theta=theta)
psi_hat = qiskit.quantum_info.Statevector.from_instruction(qc1)
rho_psi_hat = qiskit.quantum_info.DensityMatrix(psi_hat)
# Calculate the metrics
trace, fidelity = qtm.base.get_metrics(psi, psi_hat)
traces_ghz.append(trace)
fidelities_ghz.append(fidelity)
# W
thetas = thetas_origin.copy()
qc = qiskit.QuantumCircuit(num_qubits, num_qubits)
loss_values_w = []
thetass_w = []
for i in range(0, 100):
G = qtm.fubini_study.calculate_linear_state(qc.copy(), thetas, num_layers)
grad_loss = qtm.base.grad_loss(
qc,
qtm.nqubit.create_Wchecker_binho,
thetas, r=1/2, s=np.pi/2, num_layers=num_layers)
thetas = np.real(thetas - qtm.constant.learning_rate *
(np.linalg.inv(G) @ grad_loss))
qc_copy = qtm.nqubit.create_Wchecker_binho(qc.copy(), thetas, num_layers)
loss = qtm.base.loss_basis(qtm.base.measure(
qc_copy, list(range(qc_copy.num_qubits))))
loss_values_w.append(loss)
thetass_w.append(thetas)
# Plot loss value in 100 steps
plt.plot(loss_values_w, label='W', linestyle='-')
plt.title('Wchecker_binho')
plt.legend()
plt.xlabel("Iteration")
plt.ylabel("Loss value")
plt.savefig('Wchecker_binho.png', format='png', dpi=600)
plt.show()
import qtm.custom_gate
traces_w, fidelities_w = [], []
for thetas in thetass_w:
# Get |psi> = U_gen|000...>
qc = qiskit.QuantumCircuit(num_qubits, num_qubits)
qc = qtm.nqubit.create_binho_state(qc, thetas, num_layers=num_layers)
psi = qiskit.quantum_info.Statevector.from_instruction(qc)
rho_psi = qiskit.quantum_info.DensityMatrix(psi)
# Get |psi~> = U_target|000...>
qc1 = qiskit.QuantumCircuit(num_qubits, num_qubits)
qc1 = qtm.nqubit.create_w_state(qc1)
psi_hat = qiskit.quantum_info.Statevector.from_instruction(qc1)
rho_psi_hat = qiskit.quantum_info.DensityMatrix(psi_hat)
# Calculate the metrics
trace, fidelity = qtm.base.get_metrics(psi, psi_hat)
traces_w.append(trace)
fidelities_w.append(fidelity)
# Haar
thetas = thetas_origin.copy()
psi = psi / np.linalg.norm(psi)
encoder = qtm.encoding.Encoding(psi, 'amplitude_encoding')
loss_values_haar = []
thetass_haar = []
for i in range(0, 100):
qc = qiskit.QuantumCircuit(num_qubits, num_qubits)
G = qtm.fubini_study.calculate_linear_state(qc.copy(), thetas, num_layers)
qc = encoder.qcircuit
grad_loss = qtm.base.grad_loss(
qc,
qtm.nqubit.create_haarchecker_binho,
thetas, r=1/2, s=np.pi/2, num_layers=num_layers, encoder=encoder)
thetas = np.real(thetas - qtm.constant.learning_rate *
(np.linalg.inv(G) @ grad_loss))
qc_copy = qtm.nqubit.create_haarchecker_binho(
qc.copy(), thetas, num_layers, encoder)
loss = qtm.base.loss_basis(qtm.base.measure(
qc_copy, list(range(qc_copy.num_qubits))))
loss_values_haar.append(loss)
thetass_haar.append(thetas)
# Plot loss value in 100 steps
plt.plot(loss_values_haar, label='Haar', linestyle='-')
plt.title('Haarchecker_binho')
plt.legend()
plt.xlabel("Iteration")
plt.ylabel("Loss value")
plt.savefig('Haarchecker_binho.png', format='png', dpi=600)
plt.show()
# Plot loss value in 100 steps
plt.plot(loss_values_ghz, label='GHZ')
plt.plot(loss_values_w, label='W')
plt.plot(loss_values_haar, label='Haar')
plt.title('Compare init state')
plt.legend()
plt.xlabel("Iteration")
plt.ylabel("Loss value")
plt.savefig('Compare_init_state.png', format='png', dpi=600)
plt.show()
traces_haar, fidelities_haar = [], []
for thetas in thetass_haar:
# Get |psi> = U_gen|000...>
qc = qiskit.QuantumCircuit(num_qubits, num_qubits)
qc = qtm.nqubit.create_binho_state(qc, thetas, num_layers=num_layers)
psi = qiskit.quantum_info.Statevector.from_instruction(qc)
rho_psi = qiskit.quantum_info.DensityMatrix(psi)
# Get |psi~> = U_target|000...>
qc1 = encoder.qcircuit
psi_hat = qiskit.quantum_info.Statevector.from_instruction(qc1)
rho_psi_hat = qiskit.quantum_info.DensityMatrix(psi_hat)
# Calculate the metrics
trace, fidelity = qtm.base.get_metrics(psi, psi_hat)
traces_haar.append(trace)
fidelities_haar.append(fidelity)
plt.plot(traces_ghz, label='trace_ghz', color='blue')
plt.plot(fidelities_ghz, label='fidelity_ghz', linestyle='-.', color='blue')
plt.plot(traces_w, label='trace_w', color='orange')
plt.plot(fidelities_w, label='fidelity_w', linestyle='-.', color='orange')
plt.plot(traces_haar, label='trace_haar', color='g')
plt.plot(fidelities_haar, label='fidelity_haar', linestyle='-.', color='g')
plt.xlabel("Iteration")
plt.ylabel("Value")
plt.legend()
plt.savefig('Compare_init_state_trace_fidelity.png', format='png', dpi=600)
plt.show()
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import glob
import nibabel as nib
import os
import time
import pandas as pd
import numpy as np
import cv2
from skimage.transform import resize
from mricode.utils import log_textfile, createPath, data_generator
from mricode.utils import copy_colab
from mricode.utils import return_iter
from mricode.utils import return_csv
from mricode.config import config
from mricode.models.SimpleCNN_small import SimpleCNN
from mricode.models.DenseNet_NoDict import MyDenseNet
import tensorflow as tf
from tensorflow.keras.layers import Conv3D
from tensorflow import nn
from tensorflow.python.ops import nn_ops
from tensorflow.python.framework import tensor_shape
from tensorflow.python.keras.engine.base_layer import InputSpec
from tensorflow.python.keras.utils import conv_utils
tf.__version__
tf.test.is_gpu_available()
path_output = './output/'
path_tfrecords = '/data2/res64/down/'
path_csv = '/data2/csv/'
filename_res = {'train': 'intell_residual_train.csv', 'val': 'intell_residual_valid.csv', 'test': 'intell_residual_test.csv'}
filename_final = filename_res
sample_size = 'site16_allimages'
batch_size = 8
onlyt1 = False
Model = SimpleCNN
#Model = MyDenseNet
versionkey = 'down256' #down256, cropped128, cropped64, down64
modelname = 'simplecnnsmall__allimages_' + versionkey
createPath(path_output + modelname)
train_df, val_df, test_df, norm_dict = return_csv(path_csv, filename_final, False)
train_iter = config[versionkey]['iter_train']
val_iter = config[versionkey]['iter_val']
test_iter = config[versionkey]['iter_test']
t1_mean = config[versionkey]['norm']['t1'][0]
t1_std= config[versionkey]['norm']['t1'][1]
t2_mean=config[versionkey]['norm']['t2'][0]
t2_std=config[versionkey]['norm']['t2'][1]
ad_mean=config[versionkey]['norm']['ad'][0]
ad_std=config[versionkey]['norm']['ad'][1]
fa_mean=config[versionkey]['norm']['fa'][0]
fa_std=config[versionkey]['norm']['fa'][1]
md_mean=config[versionkey]['norm']['md'][0]
md_std=config[versionkey]['norm']['md'][1]
rd_mean=config[versionkey]['norm']['rd'][0]
rd_std=config[versionkey]['norm']['rd'][1]
norm_dict
cat_cols = {'female': 2, 'race.ethnicity': 5, 'high.educ_group': 4, 'income_group': 8, 'married': 6}
num_cols = [x for x in list(val_df.columns) if '_norm' in x]
def calc_loss_acc(out_loss, out_acc, y_true, y_pred, cat_cols, num_cols, norm_dict):
for col in num_cols:
tmp_col = col
tmp_std = norm_dict[tmp_col.replace('_norm','')]['std']
tmp_y_true = tf.cast(y_true[col], tf.float32).numpy()
tmp_y_pred = np.squeeze(y_pred[col].numpy())
if not(tmp_col in out_loss):
out_loss[tmp_col] = np.sum(np.square(tmp_y_true-tmp_y_pred))
else:
out_loss[tmp_col] += np.sum(np.square(tmp_y_true-tmp_y_pred))
if not(tmp_col in out_acc):
out_acc[tmp_col] = np.sum(np.square((tmp_y_true-tmp_y_pred)*tmp_std))
else:
out_acc[tmp_col] += np.sum(np.square((tmp_y_true-tmp_y_pred)*tmp_std))
for col in list(cat_cols.keys()):
tmp_col = col
if not(tmp_col in out_loss):
out_loss[tmp_col] = tf.keras.losses.SparseCategoricalCrossentropy()(tf.squeeze(y_true[col]), tf.squeeze(y_pred[col])).numpy()
else:
out_loss[tmp_col] += tf.keras.losses.SparseCategoricalCrossentropy()(tf.squeeze(y_true[col]), tf.squeeze(y_pred[col])).numpy()
if not(tmp_col in out_acc):
out_acc[tmp_col] = tf.reduce_sum(tf.dtypes.cast((y_true[col] == tf.argmax(y_pred[col], axis=-1)), tf.float32)).numpy()
else:
out_acc[tmp_col] += tf.reduce_sum(tf.dtypes.cast((y_true[col] == tf.argmax(y_pred[col], axis=-1)), tf.float32)).numpy()
return(out_loss, out_acc)
def format_output(out_loss, out_acc, n, cols, print_bl=False):
loss = 0
acc = 0
output = []
for col in cols:
output.append([col, out_loss[col]/n, out_acc[col]/n])
loss += out_loss[col]/n
acc += out_acc[col]/n
df = pd.DataFrame(output)
df.columns = ['name', 'loss', 'acc']
if print_bl:
print(df)
return(loss, acc, df)
@tf.function
def train_step(X, y, model, optimizer, cat_cols, num_cols):
with tf.GradientTape() as tape:
predictions = model(X)
i = 0
loss = tf.keras.losses.MSE(tf.cast(y[num_cols[i]], tf.float32), tf.squeeze(predictions[num_cols[i]]))
for i in range(1,len(num_cols)):
loss += tf.keras.losses.MSE(tf.cast(y[num_cols[i]], tf.float32), tf.squeeze(predictions[num_cols[i]]))
for col in list(cat_cols.keys()):
loss += tf.keras.losses.SparseCategoricalCrossentropy()(tf.squeeze(y[col]), tf.squeeze(predictions[col]))
gradients = tape.gradient(loss, model.trainable_variables)
mean_std = [x.name for x in model.non_trainable_variables if ('batch_norm') in x.name and ('mean' in x.name or 'variance' in x.name)]
with tf.control_dependencies(mean_std):
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
return(y, predictions, loss)
@tf.function
def test_step(X, y, model):
predictions = model(X)
return(y, predictions)
def epoch(data_iter, df, model, optimizer, cat_cols, num_cols, norm_dict):
out_loss = {}
out_acc = {}
n = 0.
n_batch = 0.
total_time_dataload = 0.
total_time_model = 0.
start_time = time.time()
for batch in data_iter:
total_time_dataload += time.time() - start_time
start_time = time.time()
t1 = (tf.cast(batch['t1'], tf.float32)-t1_mean)/t1_std
if False:
ad = batch['ad']
ad = tf.where(tf.math.is_nan(ad), tf.zeros_like(ad), ad)
ad = (ad-ad_mean)/ad_std
fa = batch['fa']
fa = tf.where(tf.math.is_nan(fa), tf.zeros_like(fa), fa)
fa = (fa-fa_mean)/fa_std
md = batch['md']
md = tf.where(tf.math.is_nan(md), tf.zeros_like(md), md)
md = (md-md_mean)/md_std
rd = batch['rd']
rd = tf.where(tf.math.is_nan(rd), tf.zeros_like(rd), rd)
rd = (rd-rd_mean)/rd_std
subjectid = decoder(batch['subjectid'])
y = get_labels(df, subjectid, list(cat_cols.keys())+num_cols)
#X = tf.concat([t1], axis=4)
X = tf.concat([t1], axis=4)
if optimizer != None:
y_true, y_pred, loss = train_step(X, y, model, optimizer, cat_cols, num_cols)
else:
y_true, y_pred = test_step(X, y, model)
out_loss, out_acc = calc_loss_acc(out_loss, out_acc, y_true, y_pred, cat_cols, num_cols, norm_dict)
n += X.shape[0]
n_batch += 1
if (n_batch % 10) == 0:
log_textfile(path_output + modelname + '/log' + '.log', str(n_batch))
total_time_model += time.time() - start_time
start_time = time.time()
return (out_loss, out_acc, n, total_time_model, total_time_dataload)
def get_labels(df, subjectid, cols = ['nihtbx_fluidcomp_uncorrected_norm']):
subjects_df = pd.DataFrame(subjectid)
result_df = pd.merge(subjects_df, df, left_on=0, right_on='subjectkey', how='left')
output = {}
for col in cols:
output[col] = np.asarray(result_df[col].values)
return output
def best_val(df_best, df_val, df_test, e):
df_best = pd.merge(df_best, df_val, how='left', left_on='name', right_on='name')
df_best = pd.merge(df_best, df_test, how='left', left_on='name', right_on='name')
df_best.loc[df_best['best_loss_val']>=df_best['cur_loss_val'], 'best_loss_epochs'] = e
df_best.loc[(df_best['best_acc_val']<=df_best['cur_acc_val'])&(df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'best_acc_epochs'] = e
df_best.loc[df_best['best_loss_val']>=df_best['cur_loss_val'], 'best_loss_test'] = df_best.loc[df_best['best_loss_val']>=df_best['cur_loss_val'], 'cur_loss_test']
df_best.loc[df_best['best_loss_val']>=df_best['cur_loss_val'], 'best_loss_val'] = df_best.loc[df_best['best_loss_val']>=df_best['cur_loss_val'], 'cur_loss_val']
df_best.loc[(df_best['best_acc_val']<=df_best['cur_acc_val'])&(df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'best_acc_test'] = df_best.loc[(df_best['best_acc_val']<=df_best['cur_acc_val'])&(df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'cur_acc_test']
df_best.loc[(df_best['best_acc_val']<=df_best['cur_acc_val'])&(df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'best_acc_val'] = df_best.loc[(df_best['best_acc_val']<=df_best['cur_acc_val'])&(df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'cur_acc_val']
df_best.loc[(df_best['best_acc_val']>=df_best['cur_acc_val'])&(~df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'best_acc_test'] = df_best.loc[(df_best['best_acc_val']>=df_best['cur_acc_val'])&(~df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'cur_acc_test']
df_best.loc[(df_best['best_acc_val']>=df_best['cur_acc_val'])&(~df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'best_acc_val'] = df_best.loc[(df_best['best_acc_val']>=df_best['cur_acc_val'])&(~df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'cur_acc_val']
df_best = df_best.drop(['cur_loss_val', 'cur_acc_val', 'cur_loss_test', 'cur_acc_test'], axis=1)
return(df_best)
decoder = np.vectorize(lambda x: x.decode('UTF-8'))
template = 'Epoch {0}, Loss: {1:.3f}, Accuracy: {2:.3f}, Val Loss: {3:.3f}, Val Accuracy: {4:.3f}, Time Model: {5:.3f}, Time Data: {6:.3f}'
for col in [0]:
log_textfile(path_output + modelname + '/log' + '.log', cat_cols)
log_textfile(path_output + modelname + '/log' + '.log', num_cols)
loss_object = tf.keras.losses.SparseCategoricalCrossentropy()
optimizer = tf.keras.optimizers.Adam(lr = 0.001)
model = Model(cat_cols, num_cols)
df_best = None
for e in range(20):
log_textfile(path_output + modelname + '/log' + '.log', 'Epochs: ' + str(e))
loss = tf.Variable(0.)
acc = tf.Variable(0.)
val_loss = tf.Variable(0.)
val_acc = tf.Variable(0.)
test_loss = tf.Variable(0.)
test_acc = tf.Variable(0.)
tf.keras.backend.set_learning_phase(True)
train_out_loss, train_out_acc, n, time_model, time_data = epoch(train_iter, train_df, model, optimizer, cat_cols, num_cols, norm_dict)
tf.keras.backend.set_learning_phase(False)
val_out_loss, val_out_acc, n, _, _ = epoch(val_iter, val_df, model, None, cat_cols, num_cols, norm_dict)
test_out_loss, test_out_acc, n, _, _ = epoch(test_iter, test_df, model, None, cat_cols, num_cols, norm_dict)
loss, acc, _ = format_output(train_out_loss, train_out_acc, n, list(cat_cols.keys())+num_cols)
val_loss, val_acc, df_val = format_output(val_out_loss, val_out_acc, n, list(cat_cols.keys())+num_cols, print_bl=False)
test_loss, test_acc, df_test = format_output(test_out_loss, test_out_acc, n, list(cat_cols.keys())+num_cols, print_bl=False)
df_val.columns = ['name', 'cur_loss_val', 'cur_acc_val']
df_test.columns = ['name', 'cur_loss_test', 'cur_acc_test']
if e == 0:
df_best = pd.merge(df_test, df_val, how='left', left_on='name', right_on='name')
df_best['best_acc_epochs'] = 0
df_best['best_loss_epochs'] = 0
df_best.columns = ['name', 'best_loss_test', 'best_acc_test', 'best_loss_val', 'best_acc_val', 'best_acc_epochs', 'best_loss_epochs']
df_best = best_val(df_best, df_val, df_test, e)
print(df_best[['name', 'best_loss_test', 'best_acc_test']])
print(df_best[['name', 'best_loss_val', 'best_acc_val']])
log_textfile(path_output + modelname + '/log' + '.log', template.format(e, loss, acc, val_loss, val_acc, time_model, time_data))
if e in [10, 15]:
optimizer.lr = optimizer.lr/3
log_textfile(path_output + modelname + '/log' + '.log', 'Learning rate: ' + str(optimizer.lr))
df_best.to_csv(path_output + modelname + '/df_best' + str(e) + '.csv')
df_best.to_csv(path_output + modelname + '/df_best' + '.csv')
model.save_weights(path_output + modelname + '/checkpoints/' + str(e) + '/')
error
test_loss, test_acc, df_test = format_output(test_out_loss, test_out_acc, n, list(cat_cols.keys())+num_cols, print_bl=False)
df_test.to_csv('final_output_all.csv')
inputs = tf.keras.Input(shape=(64,64,64,2), name='inputlayer123')
a = model(inputs)['female']
mm = tf.keras.models.Model(inputs=inputs, outputs=a)
from tf_explain.core.smoothgrad import SmoothGrad
import pickle
explainer = SmoothGrad()
output_grid = {}
output_n = {}
for i in range(2):
output_grid[i] = np.zeros((64,64,64))
output_n[i] = 0
counter = 0
for batch in test_iter:
counter+=1
print(counter)
t1 = (tf.cast(batch['t1'], tf.float32)-t1_mean)/t1_std
t2 = (batch['t2']-t2_mean)/t2_std
X = tf.concat([t1, t2], axis=4)
subjectid = decoder(batch['subjectid'])
y = get_labels(test_df, subjectid, list(cat_cols.keys())+num_cols)
y_list = list(y['female'])
for i in range(X.shape[0]):
X_i = X[i]
X_i = tf.expand_dims(X_i, axis=0)
y_i = y_list[i]
grid = explainer.explain((X_i, _), mm, y_i, 20, 1.)
output_grid[y_i] += grid
output_n[y_i] += 1
pickle.dump([output_grid, output_n], open( "smoothgrad_female_all.p", "wb" ) )
#output_grid, output_n = pickle.load(open( "smoothgrad_female.p", "rb" ))
def apply_grey_patch(image, top_left_x, top_left_y, top_left_z, patch_size):
"""
Replace a part of the image with a grey patch.
Args:
image (numpy.ndarray): Input image
top_left_x (int): Top Left X position of the applied box
top_left_y (int): Top Left Y position of the applied box
patch_size (int): Size of patch to apply
Returns:
numpy.ndarray: Patched image
"""
patched_image = np.array(image, copy=True)
patched_image[
top_left_x : top_left_x + patch_size, top_left_y : top_left_y + patch_size, top_left_z : top_left_z + patch_size, :
] = 0
return patched_image
import math
def get_sensgrid(image, mm, class_index, patch_size):
sensitivity_map = np.zeros((
math.ceil(image.shape[0] / patch_size),
math.ceil(image.shape[1] / patch_size),
math.ceil(image.shape[2] / patch_size)
))
for index_z, top_left_z in enumerate(range(0, image.shape[2], patch_size)):
patches = [
apply_grey_patch(image, top_left_x, top_left_y, top_left_z, patch_size)
for index_x, top_left_x in enumerate(range(0, image.shape[0], patch_size))
for index_y, top_left_y in enumerate(range(0, image.shape[1], patch_size))
]
coordinates = [
(index_y, index_x)
for index_x, _ in enumerate(range(0, image.shape[0], patch_size))
for index_y, _ in enumerate(range(0, image.shape[1], patch_size))
]
predictions = mm.predict(np.array(patches), batch_size=1)
target_class_predictions = [prediction[class_index] for prediction in predictions]
for (index_y, index_x), confidence in zip(coordinates, target_class_predictions):
sensitivity_map[index_y, index_x, index_z] = 1 - confidence
sm = resize(sensitivity_map, (64,64,64))
heatmap = (sm - np.min(sm)) / (sm.max() - sm.min())
return(heatmap)
output_grid = {}
output_n = {}
for i in range(2):
output_grid[i] = np.zeros((64,64,64))
output_n[i] = 0
counter = 0
for batch in test_iter:
counter+=1
print(counter)
t1 = (tf.cast(batch['t1'], tf.float32)-t1_mean)/t1_std
t2 = (batch['t2']-t2_mean)/t2_std
X = tf.concat([t1, t2], axis=4)
subjectid = decoder(batch['subjectid'])
y = get_labels(test_df, subjectid, list(cat_cols.keys())+num_cols)
y_list = list(y['female'])
for i in range(X.shape[0]):
print(i)
X_i = X[i]
y_i = y_list[i]
grid = get_sensgrid(X_i, mm, y_i, 4)
output_grid[y_i] += grid
output_n[y_i] += 1
if counter==6:
break
pickle.dump([output_grid, output_n], open( "heatmap_female_all.p", "wb" ) )
error
batch = next(iter(train_iter))
t1 = (tf.cast(batch['t1'], tf.float32)-t1_mean)/t1_std
t2 = (batch['t2']-t2_mean)/t2_std
ad = batch['ad']
ad = tf.where(tf.math.is_nan(ad), tf.zeros_like(ad), ad)
ad = (ad-ad_mean)/ad_std
fa = batch['fa']
fa = tf.where(tf.math.is_nan(fa), tf.zeros_like(fa), fa)
fa = (fa-fa_mean)/fa_std
md = batch['md']
md = tf.where(tf.math.is_nan(md), tf.zeros_like(md), md)
md = (md-md_mean)/md_std
rd = batch['rd']
rd = tf.where(tf.math.is_nan(rd), tf.zeros_like(rd), rd)
rd = (rd-rd_mean)/rd_std
#subjectid = decoder(batch['subjectid'])
#y = get_labels(df, subjectid, list(cat_cols.keys())+num_cols)
#X = tf.concat([t1, t2, ad, fa, md, rd], axis=4)
X = tf.concat([t1, t2], axis=4)
tf.keras.backend.set_learning_phase(True)
model(X)['female']
tf.keras.backend.set_learning_phase(False)
model(X)['female']
mean_std = [x.name for x in model.non_trainable_variables if ('batch_norm') in x.name and ('mean' in x.name or 'variance' in x.name)]
model = Model(cat_cols, num_cols)
model.non_trainable_variables
```
| github_jupyter |

<a href="https://hub.callysto.ca/jupyter/hub/user-redirect/git-pull?repo=https%3A%2F%2Fgithub.com%2Fcallysto%2Fcallysto-sample-notebooks&branch=master&subPath=notebooks/Social_Sciences/Humanities/Shakespeare_and_Statistics.ipynb&depth=1" target="_parent"><img src="https://raw.githubusercontent.com/callysto/curriculum-notebooks/master/open-in-callysto-button.svg?sanitize=true" width="123" height="24" alt="Open in Callysto"/></a>
# Shakespeare and Statistics

Can art and science be combined? Natural language processing allows us to use the same statistical skills you might learn in a science class, such as counting up members of a population and looking at their distribution, to gain insight into the written word. Here's a really simple example of what you can do. Let's consider the following question:
## What are the top 20 phrases in Shakespeare's Macbeth?
Normally, when we study Shakespeare, we critically read his plays and study the plot, characters, and themes. While this is definitely interesting stuff, we can gain very different insights by taking a multidisciplinary approach.
This is something we would probably never do if we had to do it by hand. Imagine getting out your clipboard, writing down every different word or phrase you come across and then counting how many times that same word or phrase reappears. Check out how quickly it can be done using Callysto and the free, open tools it brings with it...
## Getting started
First, we have to load a few things... This handles a lot of setup behind the scenes so that you don't have to have it in the notebook distracting students from what you really want to show. It also brings in a number helpers that we've made to avoid some of the technical details of grabbing text from a website and processing it. You could choose to include these details in this notebook or look at the files we're loading if you got really curious, but you don't have to.
Simply run this block to continue.
```
from notebook_code.shakespeare import *
```
## Finding the text
The helper we're using to get the right file from the [gutenberg.org](http://www.gutenberg.org) website needs a number. This is the same number that the folks at Gutenberg use to organize all the works available on the site. Let's track that down, starting with a google search.
<br/>
<br/>
<br/>


<br/>
<br/>
<br/>
We may see more than one result, as there may be multiple versions of popular works. Just make sure to click a link that's from [gutenberg.org](http://www.gutenberg.org) and you should end up seeing a page like this:
<br/>
<br/>
<br/>

<br/>
<br/>
<br/>
Next, click that **Bibrec** tab, and you'll see a page like this:
<br/>
<br/>
<br/>

<br/>
<br/>
<br/>
On this page is the **EBook-No.** followed by a number. In this case, it's **1129**. That's the one we're looking for!
## Loading the text
Now we grab it using `get_gutenberg_text`. The following line goes to the site, fetches the text file with the **EBook-No.** *1129*, and stores it in `macbeth`. We can refer to it by using `macbeth` at any point from here on in.
```
macbeth = get_gutenberg_text(1129)
```
For example, we can just print it out to see that we've grabbed the correct document.
```
print(macbeth)
```
Looks good! But that's a lot of reading to do. And a lot of phrases to count and keep track of. Here's where some more of those helpers come into play.
## Crunching the text
`noun_phrases` will grab groups of words that have been identified as phrases containing nouns. This isn't always 100% correct. English can be a challenging language even for machines, and sometimes even the files on [gutenberg.org](http://www.gutenberg.org) contain errors that make it even harder, but it can usually do a pretty good job.
```
macbeth_phrases = noun_phrases(macbeth)
```
We can print these phrases out to see what they look like just like we printed out the `macbeth` text above.
```
print(macbeth_phrases)
```
What you're seeing is no longer raw text. It's now a list of strings. How long is the list? Let's find out. `len` is short for "length", and it will tell you how many items are in any list.
```
len(macbeth_phrases)
```
Looks like we have over 3000 noun phrases. We don't yet know how many of them are repeated.
## Counting everything up
Here's where this starts to look like a real science project! Let's use `count_unique` to get us a table of phrases and how many times they occur. They'll come back ordered from most frequent to least frequent. There will probably be a lot of them, so we'll add `[0:20]` which means to give us the top twenty. In these lists, the zero-th item is always the first item.
```
macbeth_counts = count_unique(macbeth_phrases)[0:20]
```
Just like before, we can print out `macbeth_counts` to see what's inside, or we can refer to it without `print` and Callysto will automatically show a nicer table version:
```
macbeth_counts
```
There we have it! The top 20 phrases in Macbeth! Let's put those in a plot.
## Plotting the results
You can do almost any kind of plot or other visual representation of observations like this you could think of in Callysto. But you can also create helpers to easily produce plots with common datasets. We've created `plot_text_counts` which will take in any table like the above, with a `text` and `count` column, and produce a horizontal bar plot, ordered from most to least frequent word.
plot_text_counts(macbeth_counts)
Surprise, surprise. *Macbeth* is the top phrase in Macbeth. Our main character is mentioned more than twice the number of times as the next most frequent phrase, *Macduff*, and more than three times the frequency that *Lady Macbeth* is mentioned.
## Thinking about the results
One of the first things we might realize from this simple investigation is the importance of proper nouns. Phrases containing the main characters occur far more frequently than other phrases, and the main character of the play is mentioned far more times than any other characters.
Are these observations particular to Macbeth? Or to Shakespeare's plays? Or are they more universal?
Now that we've gone through Macbeth, how hard could it be to look at a few others?
**Hamlet** can be found on [gutenberg.org](http://www.gutenberg.org) under **EBook-No.** *1524*, and we already know from the above how to plot that out. In fact, there were really only 4 important lines.
## Looking at Hamlet
Run the following block to download **Hamlet**, pull out all the noun phrases, count them up, and plot them out.
```
hamlet = get_gutenberg_text(1524)
hamlet_phrases = noun_phrases(hamlet)
hamlet_counts = count_unique(hamlet_phrases)[0:20]
plot_text_counts(hamlet_counts)
```
## Comparing other texts
We could keep doing this for other texts. But if you think about it, we're really just doing the same steps over and over and over again. We're copying those same 4 lines and the only thing we really need to change about them is the **EBook-No.** we give to `get_gutenberg_text`.
What if you collected all of the **EBook-No.** numbers for texts you wanted to look at and put them in a list, with some labels to help you identify them? Here's how you can do that:
```
choices = { 'Shakespeare - Macbeth': 1129,
'Shakespeare - Hamlet': 1524,
'Shakespeare - Romeo and Juliet': 1112,
'Charles Dickens - A Christmas Carol': 19337 }
```
Now we'll make our recipe. This could be hidden and made into a helper like we've been using so far, but we want to show how you can fit together what you've learned.
```
def plot_gutenberg(ebook_no):
text = get_gutenberg_text(ebook_no)
phrases = noun_phrases(text)
counts = count_unique(phrases)[0:20]
plot_text_counts(counts)
```
Notice that the main difference is that you've wrapped these lines inside something called `plot_gutenberg` which accepts a value it stores in `ebook_no`. Then, instead of the actual **EBook-No.**, you pass `ebook_no` into `get_gutenberg_text`.
You could then use `plot_gutenberg` on any **EBook-No.**
```
plot_gutenberg(1129)
```
Or, you could use the `choices` list that we defined above to do the same thing in a more readable way. Now we're not just plotting out **EBook-No.** *1129*. We're plotting out Shakespeare's *Macbeth*!
```
plot_gutenberg(choices['Shakespeare - Macbeth'])
```
Let's do one last thing. Why keep typing and running even one line when you can make it interactive!
But before we do that, we need to deal with one little problem. We still have to wait a few seconds to see the results when we run `plot_gutenberg`, and if we're going to make it even easier to switch back and forth between different works, we don't want to have to always wait to see our plot.
There are many ways to tackle this, but one quick way is to do the work that takes the most time up front and store the results. Let's create a new recipe called `count_noun_phrases` that'll do everything that `plot_gutenberg` did, *except* draw the final plot. Then we'll store it. We'll use the same `ebook_no` that the folks at Gutenberg do so we can easily find it.
This might take a minute or two, so be patient! It's going to go through everything in `choices`, grab the text, find the noun phrases, and count them up, then store all that in `stored_counts`. That's a lot of work!
```
stored_counts = {}
def count_noun_phrases(ebook_no):
text = get_gutenberg_text(ebook_no)
phrases = noun_phrases(text)
counts = count_unique(phrases)[0:20]
return counts
for ebook_no in choices.values():
stored_counts[ebook_no] = count_noun_phrases(ebook_no)
```
Now let's redefine what `plot_gutenberg` does. It no longer needs to crunch the data every time. It can just grab it from `stored_counts`.
```
def plot_gutenberg(ebook_no):
plot_text_counts(stored_counts[ebook_no])
```
## Building our text selector
Time for the closing act! This is where things get a bit fancier. Don't worry if you don't fully understand what's going on in this next block. Right now, just see if you can spot some similar names, and note that with just a couple more lines, we're going to make it really easy to browse the top 20 phrases of any number of works from [gutenberg.org](http://www.gutenberg.org)!
```
selector = widgets.Dropdown(options=choices, description='Select Text:')
interact(plot_gutenberg, ebook_no=selector)
```
## And we're done!
Plus, we may have even gotten some new insights. For example, you'll see a play from Charles Dickens in the list of choices. Sure enough, we find Scrooge, the main character of *A Christmas Carol*, right at the top, followed by a few other main characters in the top 20. So it's not just Shakespeare who likes to mention his main character's name a lot. Perhaps it's more common.
If you've already read some of these, you may also find it interesting that the top 20 most frequent noun phrases also capture a lot of what's unique about that particular work.
## Experimenting
This is only scratching the surface. You could remove the proper nouns on purpose to observe how language and style might change between different writers during different times in history. With a few changes you could even run this on news articles or any other text you could get your hands on.
One really easy thing to do is simply add more choices to your selector. Feel free to try this out on your own selection of Gutenberg texts. Once you find the right **EBook-No.**, just add it to the list of `choices`. Remember, you'll have to rerun the step above that crunches the noun phrases and stores them as well before rerunning the last step that makes our selector.
[](https://github.com/callysto/curriculum-notebooks/blob/master/LICENSE.md)
| github_jupyter |
<!--NOTEBOOK_HEADER-->
*This notebook contains material from [cbe61622](https://jckantor.github.io/cbe61622);
content is available [on Github](https://github.com/jckantor/cbe61622.git).*
<!--NAVIGATION-->
< [A.0 Python Source Library](https://jckantor.github.io/cbe61622/A.00-Appendices.html) | [Contents](toc.html) | [A.2 Downloading Python source files from github](https://jckantor.github.io/cbe61622/A.02-Downloading_Python_source_files_from_github.html) ><p><a href="https://colab.research.google.com/github/jckantor/cbe61622/blob/master/docs/A.01-Resources.ipynb"> <img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a><p><a href="https://jckantor.github.io/cbe61622/A.01-Resources.ipynb"> <img align="left" src="https://img.shields.io/badge/Github-Download-blue.svg" alt="Download" title="Download Notebook"></a>
# A.1 Resources
## A.1.1 Data Acquistion, Instrumentation, and Control Companies
* [National Instruments](https://www.ni.com/en-us.html). Full line vendor of industrial and laboratory grade instruments and software, including LabView.
## A.1.2 IoT Devices and Services
* [Adafruit](https://www.adafruit.com/about). Full range of devices, sensors, and software (including CircuitPython).
* [DFRobot](dfrobot.com). Producer of open source hardware and software products, including a full range of sensors, with a strong focus on developers and STEM education.
* [National Control Devices](https://ncd.io/). A producer of IoT devices, software, and services for industrial and military applications. Includes feather boards for 5 volt expansion of I2C communications.
* [Particle.io](particle.io). A producer of IoT devices and services with a larger developer community.
* [SeeedStudio](seeedstudio.com). A producer of a IoT devices, sensors, and development kits. Produces a wide range of ["Grove"](https://wiki.seeedstudio.com/Grove_System/) sensors which use standardized connector and modules to simplify development of IoT applications.
## A.1.3 Software Platforms
* [Arduino](https://www.arduino.cc/)
* [AWS IoT](https://aws.amazon.com/iot/)
* [Blynk](https://blynk.io/). Hardware agnostic platform for creating mobile IoT applications.
* [Internet of Things (IoT) of IBM Cloud](https://www.ibm.com/cloud/internet-of-things)
* [Node-RED](https://nodered.org/). An open-source, graphical editor for creating event-driven applications to run on [Node.js](https://nodejs.org/en/). Applications can be run on Raspberry Pi or in the cloud.
* [PlatformIO](https://platformio.org/).
<!--NAVIGATION-->
< [A.0 Python Source Library](https://jckantor.github.io/cbe61622/A.00-Appendices.html) | [Contents](toc.html) | [A.2 Downloading Python source files from github](https://jckantor.github.io/cbe61622/A.02-Downloading_Python_source_files_from_github.html) ><p><a href="https://colab.research.google.com/github/jckantor/cbe61622/blob/master/docs/A.01-Resources.ipynb"> <img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a><p><a href="https://jckantor.github.io/cbe61622/A.01-Resources.ipynb"> <img align="left" src="https://img.shields.io/badge/Github-Download-blue.svg" alt="Download" title="Download Notebook"></a>
| github_jupyter |
```
%matplotlib inline
```
# Wikipedia principal eigenvector
A classical way to assert the relative importance of vertices in a
graph is to compute the principal eigenvector of the adjacency matrix
so as to assign to each vertex the values of the components of the first
eigenvector as a centrality score:
https://en.wikipedia.org/wiki/Eigenvector_centrality
On the graph of webpages and links those values are called the PageRank
scores by Google.
The goal of this example is to analyze the graph of links inside
wikipedia articles to rank articles by relative importance according to
this eigenvector centrality.
The traditional way to compute the principal eigenvector is to use the
power iteration method:
https://en.wikipedia.org/wiki/Power_iteration
Here the computation is achieved thanks to Martinsson's Randomized SVD
algorithm implemented in scikit-learn.
The graph data is fetched from the DBpedia dumps. DBpedia is an extraction
of the latent structured data of the Wikipedia content.
```
# Author: Olivier Grisel <olivier.grisel@ensta.org>
# License: BSD 3 clause
from bz2 import BZ2File
import os
from datetime import datetime
from pprint import pprint
from time import time
import numpy as np
from scipy import sparse
from joblib import Memory
from sklearn.decomposition import randomized_svd
from urllib.request import urlopen
print(__doc__)
# #############################################################################
# Where to download the data, if not already on disk
redirects_url = "http://downloads.dbpedia.org/3.5.1/en/redirects_en.nt.bz2"
redirects_filename = redirects_url.rsplit("/", 1)[1]
page_links_url = "http://downloads.dbpedia.org/3.5.1/en/page_links_en.nt.bz2"
page_links_filename = page_links_url.rsplit("/", 1)[1]
resources = [
(redirects_url, redirects_filename),
(page_links_url, page_links_filename),
]
for url, filename in resources:
if not os.path.exists(filename):
print("Downloading data from '%s', please wait..." % url)
opener = urlopen(url)
open(filename, 'wb').write(opener.read())
print()
# #############################################################################
# Loading the redirect files
memory = Memory(cachedir=".")
def index(redirects, index_map, k):
"""Find the index of an article name after redirect resolution"""
k = redirects.get(k, k)
return index_map.setdefault(k, len(index_map))
DBPEDIA_RESOURCE_PREFIX_LEN = len("http://dbpedia.org/resource/")
SHORTNAME_SLICE = slice(DBPEDIA_RESOURCE_PREFIX_LEN + 1, -1)
def short_name(nt_uri):
"""Remove the < and > URI markers and the common URI prefix"""
return nt_uri[SHORTNAME_SLICE]
def get_redirects(redirects_filename):
"""Parse the redirections and build a transitively closed map out of it"""
redirects = {}
print("Parsing the NT redirect file")
for l, line in enumerate(BZ2File(redirects_filename)):
split = line.split()
if len(split) != 4:
print("ignoring malformed line: " + line)
continue
redirects[short_name(split[0])] = short_name(split[2])
if l % 1000000 == 0:
print("[%s] line: %08d" % (datetime.now().isoformat(), l))
# compute the transitive closure
print("Computing the transitive closure of the redirect relation")
for l, source in enumerate(redirects.keys()):
transitive_target = None
target = redirects[source]
seen = {source}
while True:
transitive_target = target
target = redirects.get(target)
if target is None or target in seen:
break
seen.add(target)
redirects[source] = transitive_target
if l % 1000000 == 0:
print("[%s] line: %08d" % (datetime.now().isoformat(), l))
return redirects
# disabling joblib as the pickling of large dicts seems much too slow
#@memory.cache
def get_adjacency_matrix(redirects_filename, page_links_filename, limit=None):
"""Extract the adjacency graph as a scipy sparse matrix
Redirects are resolved first.
Returns X, the scipy sparse adjacency matrix, redirects as python
dict from article names to article names and index_map a python dict
from article names to python int (article indexes).
"""
print("Computing the redirect map")
redirects = get_redirects(redirects_filename)
print("Computing the integer index map")
index_map = dict()
links = list()
for l, line in enumerate(BZ2File(page_links_filename)):
split = line.split()
if len(split) != 4:
print("ignoring malformed line: " + line)
continue
i = index(redirects, index_map, short_name(split[0]))
j = index(redirects, index_map, short_name(split[2]))
links.append((i, j))
if l % 1000000 == 0:
print("[%s] line: %08d" % (datetime.now().isoformat(), l))
if limit is not None and l >= limit - 1:
break
print("Computing the adjacency matrix")
X = sparse.lil_matrix((len(index_map), len(index_map)), dtype=np.float32)
for i, j in links:
X[i, j] = 1.0
del links
print("Converting to CSR representation")
X = X.tocsr()
print("CSR conversion done")
return X, redirects, index_map
# stop after 5M links to make it possible to work in RAM
X, redirects, index_map = get_adjacency_matrix(
redirects_filename, page_links_filename, limit=5000000)
names = {i: name for name, i in index_map.items()}
print("Computing the principal singular vectors using randomized_svd")
t0 = time()
U, s, V = randomized_svd(X, 5, n_iter=3)
print("done in %0.3fs" % (time() - t0))
# print the names of the wikipedia related strongest components of the
# principal singular vector which should be similar to the highest eigenvector
print("Top wikipedia pages according to principal singular vectors")
pprint([names[i] for i in np.abs(U.T[0]).argsort()[-10:]])
pprint([names[i] for i in np.abs(V[0]).argsort()[-10:]])
def centrality_scores(X, alpha=0.85, max_iter=100, tol=1e-10):
"""Power iteration computation of the principal eigenvector
This method is also known as Google PageRank and the implementation
is based on the one from the NetworkX project (BSD licensed too)
with copyrights by:
Aric Hagberg <hagberg@lanl.gov>
Dan Schult <dschult@colgate.edu>
Pieter Swart <swart@lanl.gov>
"""
n = X.shape[0]
X = X.copy()
incoming_counts = np.asarray(X.sum(axis=1)).ravel()
print("Normalizing the graph")
for i in incoming_counts.nonzero()[0]:
X.data[X.indptr[i]:X.indptr[i + 1]] *= 1.0 / incoming_counts[i]
dangle = np.asarray(np.where(np.isclose(X.sum(axis=1), 0),
1.0 / n, 0)).ravel()
scores = np.full(n, 1. / n, dtype=np.float32) # initial guess
for i in range(max_iter):
print("power iteration #%d" % i)
prev_scores = scores
scores = (alpha * (scores * X + np.dot(dangle, prev_scores))
+ (1 - alpha) * prev_scores.sum() / n)
# check convergence: normalized l_inf norm
scores_max = np.abs(scores).max()
if scores_max == 0.0:
scores_max = 1.0
err = np.abs(scores - prev_scores).max() / scores_max
print("error: %0.6f" % err)
if err < n * tol:
return scores
return scores
print("Computing principal eigenvector score using a power iteration method")
t0 = time()
scores = centrality_scores(X, max_iter=100)
print("done in %0.3fs" % (time() - t0))
pprint([names[i] for i in np.abs(scores).argsort()[-10:]])
```
| github_jupyter |
# FloPy
## MODPATH 7 create simulation example
This notebook demonstrates how to create a simple forward and backward MODPATH 7 simulation using the `.create_mp7()` method. The notebooks also shows how to create subsets of endpoint output and plot MODPATH results on ModelMap objects.
```
import sys
import os
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import platform
# run installed version of flopy or add local path
try:
import flopy
except:
fpth = os.path.abspath(os.path.join('..', '..'))
sys.path.append(fpth)
import flopy
print(sys.version)
print('numpy version: {}'.format(np.__version__))
print('matplotlib version: {}'.format(mpl.__version__))
print('flopy version: {}'.format(flopy.__version__))
if not os.path.exists("data"):
os.mkdir("data")
# define executable names
mpexe = "mp7"
mfexe = "mf6"
if platform.system() == "Windows":
mpexe += ".exe"
mfexe += ".exe"
```
### Flow model data
```
nper, nstp, perlen, tsmult = 1, 1, 1., 1.
nlay, nrow, ncol = 3, 21, 20
delr = delc = 500.
top = 400.
botm = [220., 200., 0.]
laytyp = [1, 0, 0]
kh = [50., 0.01, 200.]
kv = [10., 0.01, 20.]
wel_loc = (2, 10, 9)
wel_q = -150000.
rch = 0.005
riv_h = 320.
riv_z = 317.
riv_c = 1.e5
def get_nodes(locs):
nodes = []
for k, i, j in locs:
nodes.append(k * nrow * ncol + i * ncol + j)
return nodes
```
### MODPATH 7 using MODFLOW 6
#### Create and run MODFLOW 6
```
ws = os.path.join('data', 'mp7_ex1_cs')
nm = 'ex01_mf6'
# Create the Flopy simulation object
sim = flopy.mf6.MFSimulation(sim_name=nm, exe_name=mfexe,
version='mf6', sim_ws=ws)
# Create the Flopy temporal discretization object
pd = (perlen, nstp, tsmult)
tdis = flopy.mf6.modflow.mftdis.ModflowTdis(sim, pname='tdis',
time_units='DAYS', nper=nper,
perioddata=[pd])
# Create the Flopy groundwater flow (gwf) model object
model_nam_file = '{}.nam'.format(nm)
gwf = flopy.mf6.ModflowGwf(sim, modelname=nm,
model_nam_file=model_nam_file, save_flows=True)
# Create the Flopy iterative model solver (ims) Package object
ims = flopy.mf6.modflow.mfims.ModflowIms(sim, pname='ims',
complexity='SIMPLE',
outer_hclose=1e-6,
inner_hclose=1e-6,
rcloserecord=1e-6)
# create gwf file
dis = flopy.mf6.modflow.mfgwfdis.ModflowGwfdis(gwf, pname='dis', nlay=nlay,
nrow=nrow, ncol=ncol,
length_units='FEET',
delr=delr, delc=delc,
top=top,
botm=botm)
# Create the initial conditions package
ic = flopy.mf6.modflow.mfgwfic.ModflowGwfic(gwf, pname='ic', strt=top)
# Create the node property flow package
npf = flopy.mf6.modflow.mfgwfnpf.ModflowGwfnpf(gwf, pname='npf',
icelltype=laytyp, k=kh,
k33=kv)
# recharge
flopy.mf6.modflow.mfgwfrcha.ModflowGwfrcha(gwf, recharge=rch)
# wel
wd = [(wel_loc, wel_q)]
flopy.mf6.modflow.mfgwfwel.ModflowGwfwel(gwf, maxbound=1,
stress_period_data={0: wd})
# river
rd = []
for i in range(nrow):
rd.append([(0, i, ncol - 1), riv_h, riv_c, riv_z])
flopy.mf6.modflow.mfgwfriv.ModflowGwfriv(gwf, stress_period_data={0: rd})
# Create the output control package
headfile = '{}.hds'.format(nm)
head_record = [headfile]
budgetfile = '{}.cbb'.format(nm)
budget_record = [budgetfile]
saverecord = [('HEAD', 'ALL'),
('BUDGET', 'ALL')]
oc = flopy.mf6.modflow.mfgwfoc.ModflowGwfoc(gwf, pname='oc',
saverecord=saverecord,
head_filerecord=head_record,
budget_filerecord=budget_record)
# Write the datasets
sim.write_simulation()
# Run the simulation
success, buff = sim.run_simulation()
assert success, 'mf6 model did not run'
```
Get locations to extract data
```
nodew = get_nodes([wel_loc])
cellids = gwf.riv.stress_period_data.get_data()[0]['cellid']
nodesr = get_nodes(cellids)
```
#### Create and run MODPATH 7
Forward tracking
```
# create modpath files
mpnamf = nm + '_mp_forward'
# create basic forward tracking modpath simulation
mp = flopy.modpath.Modpath7.create_mp7(modelname=mpnamf, trackdir='forward', flowmodel=gwf, model_ws=ws,
rowcelldivisions=1, columncelldivisions=1, layercelldivisions=1,
exe_name=mpexe)
# write modpath datasets
mp.write_input()
# run modpath
mp.run_model()
```
Backward tracking from well and river locations
```
# create modpath files
mpnamb = nm + '_mp_backward'
# create basic forward tracking modpath simulation
mp = flopy.modpath.Modpath7.create_mp7(modelname=mpnamb, trackdir='backward', flowmodel=gwf, model_ws=ws,
rowcelldivisions=5, columncelldivisions=5, layercelldivisions=5,
nodes=nodew+nodesr, exe_name=mpexe)
# write modpath datasets
mp.write_input()
# run modpath
mp.run_model()
```
#### Load and Plot MODPATH 7 output
##### Forward Tracking
Load forward tracking pathline data
```
fpth = os.path.join(ws, mpnamf + '.mppth')
p = flopy.utils.PathlineFile(fpth)
pw = p.get_destination_pathline_data(dest_cells=nodew)
pr = p.get_destination_pathline_data(dest_cells=nodesr)
```
Load forward tracking endpoint data
```
fpth = os.path.join(ws, mpnamf + '.mpend')
e = flopy.utils.EndpointFile(fpth)
```
Get forward particles that terminate in the well
```
well_epd = e.get_destination_endpoint_data(dest_cells=nodew)
```
Get particles that terminate in the river boundaries
```
riv_epd = e.get_destination_endpoint_data(dest_cells=nodesr)
```
Well and river forward tracking pathlines
```
colors = ['green', 'orange', 'red']
f, axes = plt.subplots(ncols=3, nrows=2, sharey=True, sharex=True, figsize=(15, 10))
axes = axes.flatten()
idax = 0
for k in range(nlay):
ax = axes[idax]
ax.set_aspect('equal')
ax.set_title('Well pathlines - Layer {}'.format(k+1))
mm = flopy.plot.PlotMapView(model=gwf, ax=ax)
mm.plot_grid(lw=0.5)
mm.plot_pathline(pw, layer=k, color=colors[k], lw=0.75)
idax += 1
for k in range(nlay):
ax = axes[idax]
ax.set_aspect('equal')
ax.set_title('River pathlines - Layer {}'.format(k+1))
mm = flopy.plot.PlotMapView(model=gwf, ax=ax)
mm.plot_grid(lw=0.5)
mm.plot_pathline(pr, layer=k, color=colors[k], lw=0.75)
idax += 1
plt.tight_layout();
```
Forward tracking endpoints captured by the well and river
```
f, axes = plt.subplots(ncols=2, nrows=1, sharey=True, figsize=(10, 5))
axes = axes.flatten()
ax = axes[0]
ax.set_aspect('equal')
ax.set_title('Well recharge area')
mm = flopy.plot.PlotMapView(model=gwf, ax=ax)
mm.plot_grid(lw=0.5)
mm.plot_endpoint(well_epd, direction='starting', colorbar=True, shrink=0.5);
ax = axes[1]
ax.set_aspect('equal')
ax.set_title('River recharge area')
mm = flopy.plot.PlotMapView(model=gwf, ax=ax)
mm.plot_grid(lw=0.5)
mm.plot_endpoint(riv_epd, direction='starting', colorbar=True, shrink=0.5);
```
##### Backward tracking
Load backward tracking pathlines
```
fpth = os.path.join(ws, mpnamb + '.mppth')
p = flopy.utils.PathlineFile(fpth)
pwb = p.get_destination_pathline_data(dest_cells=nodew)
prb = p.get_destination_pathline_data(dest_cells=nodesr)
```
Load backward tracking endpoints
```
fpth = os.path.join(ws, mpnamb + '.mpend')
e = flopy.utils.EndpointFile(fpth)
ewb = e.get_destination_endpoint_data(dest_cells=nodew, source=True)
erb = e.get_destination_endpoint_data(dest_cells=nodesr, source=True)
```
Well backward tracking pathlines
```
f, axes = plt.subplots(ncols=2, nrows=1, figsize=(10, 5))
ax = axes[0]
ax.set_aspect('equal')
ax.set_title('Well recharge area')
mm = flopy.plot.PlotMapView(model=gwf, ax=ax)
mm.plot_grid(lw=0.5)
mm.plot_pathline(pwb, layer='all', color='blue', lw=0.5, linestyle=':', label='captured by wells')
mm.plot_endpoint(ewb, direction='ending') #, colorbar=True, shrink=0.5);
ax = axes[1]
ax.set_aspect('equal')
ax.set_title('River recharge area')
mm = flopy.plot.PlotMapView(model=gwf, ax=ax)
mm.plot_grid(lw=0.5)
mm.plot_pathline(prb, layer='all', color='green', lw=0.5, linestyle=':', label='captured by rivers')
plt.tight_layout();
```
| github_jupyter |
# SQL Aggregation and Join

```
import pandas as pd
import sqlite3
conn = sqlite3.connect("data/flights.db")
cur = conn.cursor()
```
# Objectives
- Use SQL aggregation functions with GROUP BY
- Use HAVING for group filtering
- Use SQL JOIN to combine tables using keys
# Aggregating Functions
> A SQL **aggregating function** takes in many values and returns one value.
We have already seen some SQL aggregating functions like `COUNT()`. There are also others, like SUM(), AVG(), MIN(), and MAX().
## Example Simple Aggregations
```
# Max value for longitude
pd.read_sql('''
SELECT
-- Note we have to cast to a numerical value first
MAX(CAST(longitude AS REAL))
FROM airports
''', conn)
# Max value for id in table
pd.read_sql('''
SELECT
MAX(CAST(id AS integer))
FROM
airports
''', conn)
# Effectively counts all the inactive airlines
pd.read_sql('''
SELECT COUNT()
FROM airlines
WHERE active='N'
''', conn)
```
We can also give aliases to our aggregations:
```
# Effectively counts all the active airlines
pd.read_sql('''
SELECT
COUNT() AS number_of_active_airlines
FROM
airlines
WHERE
active='Y'
''', conn)
```
# Grouping in SQL
We can go deeper and use aggregation functions on _groups_ using the `GROUP BY` clause.
The `GROUP BY` clause will group one or more columns together with the same values as one group to perform aggregation functions on.
## Example `GROUP BY` Statements
Let's say we want to know how many active and non-active airlines there are.
### Without `GROUP BY`
Let's first start with just seeing how many airlines there are:
```
df_results = pd.read_sql('''
SELECT
-- Reminder that this counts the number of rows before the SELECT
COUNT() AS number_of_airlines
FROM
airlines
''', conn)
df_results
```
One way for us to get the counts for each is to create two queries that will filter each kind of airline (active vs non-active) and count those values:
```
df_active = pd.read_sql('''
SELECT
COUNT() AS number_of_active_airlines
FROM
airlines
WHERE
active='Y'
''', conn)
df_not_active = pd.read_sql('''
SELECT
COUNT() AS number_of_not_active_airlines
FROM
airlines
WHERE
active='N'
''', conn)
display(df_active)
display(df_not_active)
```
This works but it's inefficient.
### With `GROUP BY`
Instead, we can tell the SQL server to do the work for us by grouping values we care about for us!
```
df_results = pd.read_sql('''
SELECT
COUNT() AS number_of_airlines
FROM
airlines
GROUP BY
active
''', conn)
df_results
```
This is great! And if you look closely, you can observe we have _three_ different groups instead of our expected two!
Let's also print out the `airlines.active` value for each group/aggregation so we know what we're looking at:
```
df_results = pd.read_sql('''
SELECT
airlines.active,
COUNT() AS number_of_airlines
FROM
airlines
GROUP BY
airlines.active
''', conn)
df_results
```
## Group Task
- Which countries have the highest numbers of active airlines? Return the top 10.
```
pd.read_sql('''
SELECT
*
FROM
airlines
''', conn)
```
<details>
<summary><b>Possible Solution</b></summary>
``` sql
pd.read_sql('''
SELECT
COUNT() AS num,
country
FROM
airlines
WHERE
active='Y'
GROUP BY
country
ORDER BY
num DESC
LIMIT 10
''', conn)```
</details>
> Note that the `GROUP BY` clause is considered _before_ the `ORDER BY` and `LIMIT` clauses
## Exercise: Grouping
- Run a query that will return the number of airports by time zone. Each row should have a number of airports and a time zone.
```
# Your code here
```
<details>
<summary><b>Possible Solution</b></summary>
``` sql
pd.read_sql('''
SELECT
airports.timezone
,COUNT() AS num_of_airports
FROM
airports
GROUP BY
airports.timezone
ORDER BY
num_of_airports DESC
''', conn) ```
</details>
# Filtering Groups with `HAVING`
We showed that you can filter tables with `WHERE`. We can similarly filter _groups/aggregations_ using `HAVING` clauses.
## Examples of Using `HAVING`
### Simple Filtering - Number of Airports in a Country
Let's come back to the aggregation of active airports:
```
pd.read_sql('''
SELECT
COUNT() AS num,
country
FROM
airlines
WHERE
active='Y'
GROUP BY
country
ORDER BY
num DESC
''', conn)
```
We can see we have a lot of results. But maybe we only want to keep the countries that have more than $30$ active airlines:
```
pd.read_sql('''
SELECT
country,
COUNT() AS num
FROM
airlines
WHERE
active='Y'
GROUP BY
country
HAVING
num > 30
ORDER BY
num DESC
''', conn)
```
## Filtering Different Aggregations - Airport Altitudes
We can also filter on other aggregations. For example, let's say we want to investigate the `airports` table.
Specifically, we want to know the height of the _highest airport_ in a country given that it has _at least $100$ airports_.
### Looking at the `airports` Table
```
df_airports = pd.read_sql('''
SELECT
*
FROM
airports
''', conn)
df_airports.head()
```
### Looking at the Highest Airport
Let's first get the highest altitude for each airport:
```
pd.read_sql('''
SELECT
airports.country
,MAX(
CAST(airports.altitude AS REAL)
) AS highest_airport_in_country
FROM
airports
GROUP BY
airports.country
ORDER BY
airports.country
''', conn)
```
### Looking at the Number of Airports Too
We can also get the number of airports for each country.
```
pd.read_sql('''
SELECT
airports.country
,MAX(
CAST(airports.altitude AS REAL)
) AS highest_airport_in_country
,COUNT() AS number_of_airports_in_country
FROM
airports
GROUP BY
airports.country
ORDER BY
airports.country
''', conn)
```
### Filtering on Aggregations
> Recall:
>
> We want to know the height of the _highest airport_ in a country given that it has _at least $100$ airports_.
```
pd.read_sql('''
SELECT
airports.country
,MAX(
CAST(airports.altitude AS REAL)
) AS highest_airport_in_country
-- Note we don't have to include this in our SELECT
,COUNT() AS number_of_airports_in_country
FROM
airports
GROUP BY
airports.country
HAVING
COUNT() >= 100
ORDER BY
airports.country
''', conn)
```
# Joins
The biggest advantage in using a relational database (like we've been with SQL) is that you can create **joins**.
> By using **`JOIN`** in our query, we can connect different tables using their _relationships_ to other tables.
>
> Usually we use a key (*foreign key*) to tell us how the two tables are related.
There are different types of joins and each has their different use case.
## `INNER JOIN`
> An **inner join** will join two tables together and only keep rows if the _key is in both tables_

Example of an inner join:
``` sql
SELECT
table1.column_name,
table2.different_column_name
FROM
table1
INNER JOIN table2
ON table1.shared_column_name = table2.shared_column_name
```
### Code Example for Inner Joins
Let's say we want to look at the different airplane routes
```
pd.read_sql('''
SELECT
*
FROM
routes
''', conn)
```
This is great but notice the `airline_id` column. It'd be nice to have some more information about the airlines associated with these routes.
We can do an **inner join** to get this information!
#### Inner Join Routes & Airline Data
```
pd.read_sql('''
SELECT
*
FROM
routes
INNER JOIN airlines
ON routes.airline_id = airlines.id
''', conn)
```
We can also specify that we want to retain only certain columns in the `SELECT` clause:
```
pd.read_sql('''
SELECT
routes.source AS departing
,routes.dest AS destination
,routes.stops AS stops_before_destination
,airlines.name AS airline
FROM
routes
INNER JOIN airlines
ON routes.airline_id = airlines.id
''', conn)
```
#### Note: Losing Data with Inner Joins
Since data rows are kept only if _both_ tables have the key, some data can be lost
```
df_all_routes = pd.read_sql('''
SELECT
*
FROM
routes
''', conn)
df_routes_after_join = pd.read_sql('''
SELECT
*
FROM
routes
INNER JOIN airlines
ON routes.airline_id = airlines.id
''', conn)
# Look at how the number of rows are different
df_all_routes.shape, df_routes_after_join.shape
```
If you want to keep your data from at least one of your tables, you should use a left join instead of an inner join.
## `LEFT JOIN`
> A **left join** will join two tables together and but will keep all data from the first (left) table using the key provided.

Example of a left and right join:
```sql
SELECT
table1.column_name,
table2.different_column_name
FROM
table1
LEFT JOIN table2
ON table1.shared_column_name = table2.shared_column_name
```
### Code Example for Left Join
Recall our example using an inner join and how it lost some data since the key wasn't in both the `routes` _and_ `airlines` tables.
```
df_all_routes = pd.read_sql('''
SELECT
*
FROM
routes
''', conn)
# This will lose some data (some routes not included)
df_routes_after_inner_join = pd.read_sql('''
SELECT
*
FROM
routes
INNER JOIN airlines
ON routes.airline_id = airlines.id
''', conn)
# The number of rows are different
df_all_routes.shape, df_routes_after_inner_join.shape
```
If wanted to ensure we always had every route even if the key in `airlines` was not found, we could replace our `INNER JOIN` with a `LEFT JOIN`:
```
# This will include all the data from routes
df_routes_after_left_join = pd.read_sql('''
SELECT
*
FROM
routes
LEFT JOIN airlines
ON routes.airline_id = airlines.id
''', conn)
df_routes_after_left_join.shape
```
## Exercise: Joins
Which airline has the most routes listed in our database?
```
# Your code here
```
<details>
<summary><b>Possible Solution</b></summary>
```sql
SELECT
airlines.name AS airline,
COUNT() AS number_of_routes
-- We first need to get all the relevant info via a join
FROM
routes
-- LEFT JOIN since we want all routes (even if airline id is unknown)
LEFT JOIN airlines
ON routes.airline_id = airlines.id
-- We need to group by airline's ID
GROUP BY
airlines.id
ORDER BY
number_of_routes DESC
```
</details>
# Level Up: Execution Order
```SQL
SELECT
COUNT(table2.col2) AS my_new_count
,table1.col2
FROM
table1
JOIN table2
ON table1.col1 = table2.col2
WHERE
table1.col1 > 0
GROUP BY
table2.col1
```
1. `From`
2. `Where`
3. `Group By`
4. `Having`
5. `Select`
6. `Order By`
7. `Limit`
| github_jupyter |
# solution
```
# python3
import sys, threading
sys.setrecursionlimit(10**6) # max depth of recursion
threading.stack_size(2**27) # new thread will get stack of such size
class TreeOrders:
def read(self):
self.n = int(sys.stdin.readline())
# self.n = int(input())
self.key = [0 for i in range(self.n)]
self.left = [0 for i in range(self.n)]
self.right = [0 for i in range(self.n)]
for i in range(self.n):
[a, b, c] = map(int, sys.stdin.readline().split())
# [a, b, c] = map(int, input().split())
self.key[i] = a
self.left[i] = b
self.right[i] = c
def inOrder(self):
cur_id = 0
stack = []
while True:
if cur_id != -1:
stack.append(cur_id)
cur_id = self.left[cur_id]
elif stack:
cur_id = stack.pop()
yield self.key[cur_id]
cur_id = self.right[cur_id]
else:
break
# self.result = []
# # Finish the implementation
# # You may need to add a new recursive method to do that
# return self.result
def preOrder(self):
cur_id = 0
stack = []
while True:
if cur_id != -1:
yield self.key[cur_id]
stack.append(cur_id)
cur_id = self.left[cur_id]
elif stack:
cur_id = stack.pop()
cur_id = self.right[cur_id]
else:
break
# self.result = []
# # Finish the implementation
# # You may need to add a new recursive method to do that
# return self.result
def postOrder(self):
stack1 = [0]
stack2 = []
while stack1:
cur_id = stack1.pop()
stack2.append(self.key[cur_id])
left_id = self.left[cur_id]
right_id = self.right[cur_id]
if left_id != -1:
stack1.append(left_id)
if right_id != -1:
stack1.append(right_id)
while stack2:
yield stack2.pop()
# self.result = []
# # Finish the implementation
# # You may need to add a new recursive method to do that
# return self.result
def main():
tree = TreeOrders()
tree.read()
print(" ".join(str(x) for x in tree.inOrder()))
print(" ".join(str(x) for x in tree.preOrder()))
print(" ".join(str(x) for x in tree.postOrder()))
threading.Thread(target=main).start()
```
# starter file
```
# python3
import sys, threading
sys.setrecursionlimit(10**6) # max depth of recursion
threading.stack_size(2**27) # new thread will get stack of such size
class TreeOrders:
def read(self):
self.n = int(sys.stdin.readline())
self.key = [0 for i in range(self.n)]
self.left = [0 for i in range(self.n)]
self.right = [0 for i in range(self.n)]
for i in range(self.n):
[a, b, c] = map(int, sys.stdin.readline().split())
self.key[i] = a
self.left[i] = b
self.right[i] = c
def inOrder(self):
self.result = []
# Finish the implementation
# You may need to add a new recursive method to do that
return self.result
def preOrder(self):
self.result = []
# Finish the implementation
# You may need to add a new recursive method to do that
return self.result
def postOrder(self):
self.result = []
# Finish the implementation
# You may need to add a new recursive method to do that
return self.result
def main():
tree = TreeOrders()
tree.read()
print(" ".join(str(x) for x in tree.inOrder()))
print(" ".join(str(x) for x in tree.preOrder()))
print(" ".join(str(x) for x in tree.postOrder()))
threading.Thread(target=main).start()
```
| github_jupyter |
# pystac-client Introduction
This notebook shows basic use of pystac-client to open an API, iterate through Collections and Items, and perform simple spatio-temporal searches.
```
from pystac_client import Client
# set pystac_client logger to DEBUG to see API calls
import logging
logging.basicConfig()
logger = logging.getLogger('pystac_client')
logger.setLevel(logging.DEBUG)
```
# Client
We first connect to an API by retrieving the root catalog, or landing page, of the API with the `Client.open` function.
```
# STAC API root URL
URL = 'https://planetarycomputer.microsoft.com/api/stac/v1'
# custom headers
headers = []
cat = Client.open(URL, headers=headers)
cat
```
# CollectionClient
As with a static catalog the `get_collections` function will iterate through the Collections in the Catalog. Notice that because this is an API it can get all the Collections through a single call, rather than having to fetch each one individually.
```
for collection in cat.get_collections():
print(collection)
collection = cat.get_collection('aster-l1t')
collection
```
# Items
The main functions for getting items return iterators, where pystac-client will handle retrieval of additional pages when needed. Note that one request is made for the first ten items, then a second request for the next ten.
```
items = collection.get_items()
# flush stdout so we can see the exact order that things happen
def get_ten_items(items):
for i, item in enumerate(items):
print(f"{i}: {item}", flush=True)
if i == 9:
return
print('First page', flush=True)
get_ten_items(items)
print('Second page', flush=True)
get_ten_items(items)
```
# API Search
If the Catalog is an API, we have the ability to search for items based on spatio-temporal properties.
```
# AOI around Delfzijl, in northern Netherlands
geom = {
"type": "Polygon",
"coordinates": [
[
[
6.42425537109375,
53.174765470134616
],
[
7.344360351562499,
53.174765470134616
],
[
7.344360351562499,
53.67393435835391
],
[
6.42425537109375,
53.67393435835391
],
[
6.42425537109375,
53.174765470134616
]
]
]
}
# limit sets the # of items per page so we can see multiple pages getting fetched
search = cat.search(
collections = "aster-l1t",
intersects = geom,
datetime = "2000-01-01/2010-12-31",
max_items = 15,
limit = 5
)
# PySTAC ItemCollection
items = search.get_all_items()
# Dictionary (GeoJSON FeatureCollection)
item_json = items.to_dict()
len(items)
# note that this will work in JupyterLab, but not in a Jupyter Notebook
import IPython.display
IPython.display.JSON(item_json)
# this cell can be used in Jupyter Notebook. Use above if using JupyterLab
import json
import uuid
from IPython.display import display_javascript, display_html, display
class RenderJSON(object):
def __init__(self, json_data):
if isinstance(json_data, dict) or isinstance(json_data, list):
self.json_str = json.dumps(json_data)
else:
self.json_str = json_data
self.uuid = str(uuid.uuid4())
def _ipython_display_(self):
display_html('<div id="{}" style="height: 600px; width:100%;font: 12px/18px monospace !important;"></div>'.format(self.uuid), raw=True)
display_javascript("""
require(["https://rawgit.com/caldwell/renderjson/master/renderjson.js"], function() {
renderjson.set_show_to_level(2);
document.getElementById('%s').appendChild(renderjson(%s))
});
""" % (self.uuid, self.json_str), raw=True)
RenderJSON(item_json)
```
| github_jupyter |
[Index](Index.ipynb) - [Back](Widget Events.ipynb) - [Next](Custom Widget - Hello World.ipynb)
```
%%html
<style>
.example-container { background: #999999; padding: 2px; min-height: 100px; }
.example-container.sm { min-height: 50px; }
.example-box { background: #9999FF; width: 50px; height: 50px; text-align: center; vertical-align: middle; color: white; font-weight: bold; margin: 2px;}
.example-box.med { width: 65px; height: 65px; }
.example-box.lrg { width: 80px; height: 80px; }
</style>
from IPython.html import widgets
from IPython.display import display
```
# Widget Styling
## Basic styling
The widgets distributed with IPython can be styled by setting the following traits:
- width
- height
- fore_color
- back_color
- border_color
- border_width
- border_style
- font_style
- font_weight
- font_size
- font_family
The example below shows how a `Button` widget can be styled:
```
button = widgets.Button(
description='Hello World!',
width=100, # Integers are interpreted as pixel measurements.
height='2em', # em is valid HTML unit of measurement.
color='lime', # Colors can be set by name,
background_color='#0022FF', # and also by color code.
border_color='red')
display(button)
```
## Parent/child relationships
To display widget A inside widget B, widget A must be a child of widget B. Widgets that can contain other widgets have a **`children` attribute**. This attribute can be **set via a keyword argument** in the widget's constructor **or after construction**. Calling display on an **object with children automatically displays those children**, too.
```
from IPython.display import display
float_range = widgets.FloatSlider()
string = widgets.Text(value='hi')
container = widgets.Box(children=[float_range, string])
container.border_color = 'red'
container.border_style = 'dotted'
container.border_width = 3
display(container) # Displays the `container` and all of it's children.
```
### After the parent is displayed
Children **can be added to parents** after the parent has been displayed. The **parent is responsible for rendering its children**.
```
container = widgets.Box()
container.border_color = 'red'
container.border_style = 'dotted'
container.border_width = 3
display(container)
int_range = widgets.IntSlider()
container.children=[int_range]
```
## Fancy boxes
If you need to display a more complicated set of widgets, there are **specialized containers** that you can use. To display **multiple sets of widgets**, you can use an **`Accordion` or a `Tab` in combination with one `Box` per set of widgets** (as seen below). The "pages" of these widgets are their children. To set the titles of the pages, one can **call `set_title`**.
### Accordion
```
name1 = widgets.Text(description='Location:')
zip1 = widgets.BoundedIntText(description='Zip:', min=0, max=99999)
page1 = widgets.Box(children=[name1, zip1])
name2 = widgets.Text(description='Location:')
zip2 = widgets.BoundedIntText(description='Zip:', min=0, max=99999)
page2 = widgets.Box(children=[name2, zip2])
accord = widgets.Accordion(children=[page1, page2], width=400)
display(accord)
accord.set_title(0, 'From')
accord.set_title(1, 'To')
```
### TabWidget
```
name = widgets.Text(description='Name:', padding=4)
color = widgets.Dropdown(description='Color:', padding=4, options=['red', 'orange', 'yellow', 'green', 'blue', 'indigo', 'violet'])
page1 = widgets.Box(children=[name, color], padding=4)
age = widgets.IntSlider(description='Age:', padding=4, min=0, max=120, value=50)
gender = widgets.RadioButtons(description='Gender:', padding=4, options=['male', 'female'])
page2 = widgets.Box(children=[age, gender], padding=4)
tabs = widgets.Tab(children=[page1, page2])
display(tabs)
tabs.set_title(0, 'Name')
tabs.set_title(1, 'Details')
```
# Alignment
Most widgets have a **`description` attribute**, which allows a label for the widget to be defined.
The label of the widget **has a fixed minimum width**.
The text of the label is **always right aligned and the widget is left aligned**:
```
display(widgets.Text(description="a:"))
display(widgets.Text(description="aa:"))
display(widgets.Text(description="aaa:"))
```
If a **label is longer** than the minimum width, the **widget is shifted to the right**:
```
display(widgets.Text(description="a:"))
display(widgets.Text(description="aa:"))
display(widgets.Text(description="aaa:"))
display(widgets.Text(description="aaaaaaaaaaaaaaaaaa:"))
```
If a `description` is **not set** for the widget, the **label is not displayed**:
```
display(widgets.Text(description="a:"))
display(widgets.Text(description="aa:"))
display(widgets.Text(description="aaa:"))
display(widgets.Text())
```
## Flex boxes
Widgets can be aligned using the `FlexBox`, `HBox`, and `VBox` widgets.
### Application to widgets
Widgets display vertically by default:
```
buttons = [widgets.Button(description=str(i)) for i in range(3)]
display(*buttons)
```
### Using hbox
To make widgets display horizontally, you need to **child them to a `HBox` widget**.
```
container = widgets.HBox(children=buttons)
display(container)
```
By setting the width of the container to 100% and its `pack` to `center`, you can center the buttons.
```
container.width = '100%'
container.pack = 'center'
```
## Visibility
Sometimes it is necessary to **hide or show widgets** in place, **without having to re-display** the widget.
The `visible` property of widgets can be used to hide or show **widgets that have already been displayed** (as seen below). The `visible` property can be:
* `True` - the widget is displayed
* `False` - the widget is hidden, and the empty space where the widget would be is collapsed
* `None` - the widget is hidden, and the empty space where the widget would be is shown
```
w1 = widgets.Latex(value="First line")
w2 = widgets.Latex(value="Second line")
w3 = widgets.Latex(value="Third line")
display(w1, w2, w3)
w2.visible=None
w2.visible=False
w2.visible=True
```
### Another example
In the example below, a form is rendered, which conditionally displays widgets depending on the state of other widgets. Try toggling the student checkbox.
```
form = widgets.VBox()
first = widgets.Text(description="First Name:")
last = widgets.Text(description="Last Name:")
student = widgets.Checkbox(description="Student:", value=False)
school_info = widgets.VBox(visible=False, children=[
widgets.Text(description="School:"),
widgets.IntText(description="Grade:", min=0, max=12)
])
pet = widgets.Text(description="Pet's Name:")
form.children = [first, last, student, school_info, pet]
display(form)
def on_student_toggle(name, value):
if value:
school_info.visible = True
else:
school_info.visible = False
student.on_trait_change(on_student_toggle, 'value')
```
[Index](Index.ipynb) - [Back](Widget Events.ipynb) - [Next](Custom Widget - Hello World.ipynb)
| github_jupyter |
<a href="https://colab.research.google.com/github/rakehsaleem/Crack-Detection-TF-1.x.x/blob/master/Mask_R_CNN_colab.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# **Mask R-CNN instance segmentation with custom dataset in Google Colab**
Jupyter notebook providing steps to train a **Mask R-CNN** model with custom dataset. Requirements are only dataset images and annotations file which you can get it from [here](https://github.com/rakehsaleem/Crack-Detection-TF-1.x.x/releases/tag/v1.0)
**Colab Runtime type: Python3, GPU enabled.**
#**Making Dataset**
I generated dataset annotations with [VGG Image Annotator](https://www.robots.ox.ac.uk/~vgg/software/via/).
Notebook train a model for one class object detection. It is possible to slightly modify notebook to train model for multiple classes.
Before running notebook, we need to create dataset:
1. Collect various pictures of objects to detect
2. Create annotation files in VGG
3. Create image.zip file having structure defined below
4. Upload the zip file in your Google Drive
Zip file structure:
```
images.zip
|- "train" directory
|- jpg image files of training data
|- "via_region_data.json" annotations file of training data
|- "val" directory
|- jpg image files of validation data
|- "via_region_data.json" annotations file of validation data
```
Check my image.zip file as dataset example.
# Install required packages
```
%cd
!git clone --quiet https://github.com/rakehsaleem/Crack-Detection-TF-1.x.x.git
%cd ~/Crack-Detection-TF-1.x.x
!pip uninstall -y tensorflow
!pip install tensorflow==1.14.0
!pip install keras==2.2.4
import tensorflow as tf
print(tf.__version__)
!pip install -q PyDrive
!pip install -r requirements.txt
!python setup.py install
```
**Dataset Preparation and set up locations**
```
%cd ~/Crack-Detection-TF-1.x.x
fileId = '12bCRgVoRaegFY7xQ2h6qmFrMKz8AtLZt'
import os
from zipfile import ZipFile
from shutil import copy
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
os.makedirs('dataset')
os.chdir('dataset')
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
fileName = fileId + '.zip'
downloaded = drive.CreateFile({'id': fileId})
downloaded.GetContentFile(fileName)
ds = ZipFile(fileName)
ds.extractall()
os.remove(fileName)
print('Extracted zip file ' + fileName)
%cd ~/Crack-Detection-TF-1.x.x
!cp ~/Crack-Detection-TF-1.x.x/crack/crack.py ./crack.py
!sed -i -- 's/epochs=30/epochs=5/g' crack.py
```
# Training Starts
If for some reason you get error related to keras, locate the file with error and go to file line to modify these two lines:
```
original_keras_version = f.attrs['keras_version'].decode('utf8')
original_backend = f.attrs['backend'].decode('utf8')
to
original_keras_version = f.attrs['keras_version']
original_backend = f.attrs['backend']
```
```
%cd ~/Crack-Detection-TF-1.x.x
!python crack.py train --dataset=dataset/Dataset3 --weights=coco
```
| github_jupyter |
<a href="https://practicalai.me"><img src="https://raw.githubusercontent.com/practicalAI/images/master/images/rounded_logo.png" width="100" align="left" hspace="20px" vspace="20px"></a>
<img src="https://cdn4.iconfinder.com/data/icons/data-analysis-flat-big-data/512/data_cleaning-512.png" width="90px" vspace="10px" align="right">
<div align="left">
<h1>Preprocessing</h1>
In this lesson, we will explore preprocessing and data loading utilities in Tensorflow + Keras, mainly focused on text-based data.
<table align="center">
<td>
<img src="https://raw.githubusercontent.com/practicalAI/images/master/images/rounded_logo.png" width="25"><a target="_blank" href="https://practicalai.me"> View on practicalAI</a>
</td>
<td>
<img src="https://raw.githubusercontent.com/practicalAI/images/master/images/colab_logo.png" width="25"><a target="_blank" href="https://colab.research.google.com/github/practicalAI/practicalAI/blob/master/notebooks/09_Preprocessing.ipynb"> Run in Google Colab</a>
</td>
<td>
<img src="https://raw.githubusercontent.com/practicalAI/images/master/images/github_logo.png" width="22"><a target="_blank" href="https://github.com/practicalAI/practicalAI/blob/master/notebooks/09_Preprocessing.ipynb"> View code on GitHub</a>
</td>
</table>
# Overview
* **Tokenizer**: data processing unit to convert text data to tokens
* **LabelEncoder**: convert text labels to tokens
# Set up
```
# Use TensorFlow 2.x
%tensorflow_version 2.x
import os
import numpy as np
import tensorflow as tf
# Arguments
SEED = 1234
DATA_FILE = 'news.csv'
SHUFFLE = True
INPUT_FEATURE = 'title'
OUTPUT_FEATURE = 'category'
LOWER = True
CHAR_LEVEL = False
TRAIN_SIZE = 0.7
VAL_SIZE = 0.15
TEST_SIZE = 0.15
NUM_EPOCHS = 10
BATCH_SIZE = 32
# Set seed for reproducability
np.random.seed(SEED)
tf.random.set_seed(SEED)
```
# Load data
We will download the [AG News dataset](http://www.di.unipi.it/~gulli/AG_corpus_of_news_articles.html), which consists of 120000 text samples from 4 unique classes ('Business', 'Sci/Tech', 'Sports', 'World')
```
import pandas as pd
import re
import urllib
# Upload data from GitHub to notebook's local drive
url = "https://raw.githubusercontent.com/practicalAI/practicalAI/master/data/news.csv"
response = urllib.request.urlopen(url)
html = response.read()
with open(DATA_FILE, 'wb') as fp:
fp.write(html)
# Load data
df = pd.read_csv(DATA_FILE, header=0)
X = df[INPUT_FEATURE].values
y = df[OUTPUT_FEATURE].values
df.head(5)
```
# Preprocess data
```
def preprocess_text(text):
"""Common text preprocessing steps."""
# Remove unwanted characters
text = re.sub(r"[^0-9a-zA-Z?.!,¿]+", " ", text)
# Add space between words and punctuations
text = re.sub(r"([?.!,¿])", r" \1 ", text)
text = re.sub(r'[" "]+', " ", text)
# Remove whitepsaces
text = text.rstrip().strip()
return text
# Preprocess the titles
df.title = df.title.apply(preprocess_text)
df.head(5)
```
<img height="45" src="http://bestanimations.com/HomeOffice/Lights/Bulbs/animated-light-bulb-gif-29.gif" align="left" vspace="5px" hspace="10px">
If you have preprocessing steps like standardization, etc. that are calculated, you need to separate the training and test set first before spplying those operations. This is because we cannot apply any knowledge gained from the test set accidentally during preprocessing/training. However for preprocessing steps like the function above where we aren't learning anything from the data itself, we can perform before splitting the data.
# Split data
```
import collections
from sklearn.model_selection import train_test_split
```
### Components
```
def train_val_test_split(X, y, val_size, test_size, shuffle):
"""Split data into train/val/test datasets."""
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=test_size, stratify=y, shuffle=shuffle)
X_train, X_val, y_train, y_val = train_test_split(
X_train, y_train, test_size=val_size, stratify=y_train, shuffle=shuffle)
return X_train, X_val, X_test, y_train, y_val, y_test
```
### Operations
```
# Create data splits
X_train, X_val, X_test, y_train, y_val, y_test = train_val_test_split(
X=X, y=y, val_size=VAL_SIZE, test_size=TEST_SIZE, shuffle=SHUFFLE)
class_counts = dict(collections.Counter(y))
print (f"X_train: {X_train.shape}, y_train: {y_train.shape}")
print (f"X_val: {X_val.shape}, y_val: {y_val.shape}")
print (f"X_test: {X_test.shape}, y_test: {y_test.shape}")
print (f"X_train[0]: {X_train[0]}")
print (f"y_train[0]: {y_train[0]}")
print (f"Classes: {class_counts}")
```
# Tokenizer
```
from tensorflow.keras.preprocessing.text import Tokenizer
```
### Operations
```
# Input vectorizer
X_tokenizer = Tokenizer(lower=LOWER,
char_level=CHAR_LEVEL,
oov_token='<UNK>')
# Fit only on train data
X_tokenizer.fit_on_texts(X_train)
vocab_size = len(X_tokenizer.word_index) + 1
print (f"# tokens: {vocab_size}")
# Convert text to sequence of tokens
print (f"X_train[0]: {X_train[0]}")
X_train = np.array(X_tokenizer.texts_to_sequences(X_train))
X_val = np.array(X_tokenizer.texts_to_sequences(X_val))
X_test = np.array(X_tokenizer.texts_to_sequences(X_test))
print (f"X_train[0]: {X_train[0]}")
print (f"len(X_train[0]): {len(X_train[0])} characters")
```
<img height="45" src="http://bestanimations.com/HomeOffice/Lights/Bulbs/animated-light-bulb-gif-29.gif" align="left" vspace="5px" hspace="10px">
Checkout other preprocessing functions in the [official documentation](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/preprocessing).
# LabelEncoder
```
import json
from sklearn.preprocessing import LabelEncoder
```
### Operations
```
# Output vectorizer
y_tokenizer = LabelEncoder()
# Fit on train data
y_tokenizer = y_tokenizer.fit(y_train)
num_classes = len(y_tokenizer.classes_)
print (f"# classes: {num_classes}")
# Convert labels to tokens
print (f"y_train[0]: {y_train[0]}")
y_train = y_tokenizer.transform(y_train)
y_val = y_tokenizer.transform(y_val)
y_test = y_tokenizer.transform(y_test)
print (f"y_train[0]: {y_train[0]}")
# Class weights
counts = collections.Counter(y_train)
class_weights = {_class: 1.0/count for _class, count in counts.items()}
print (f"class counts: {counts},\nclass weights: {class_weights}")
```
<img height="45" src="http://bestanimations.com/HomeOffice/Lights/Bulbs/animated-light-bulb-gif-29.gif" align="left" vspace="5px" hspace="10px">
Checkout the complete list of sklearn preprocessing functions in the [official documentation](https://scikit-learn.org/stable/modules/classes.html#module-sklearn.preprocessing).
# Padding
Our inputs are all of varying length but we need each batch to be uniformly shaped. Therefore, we will use padding to make all the inputs in the batch the same length. Our padding index will be 0 (note that X_tokenizer starts at index 1).
```
from tensorflow.keras.preprocessing.sequence import pad_sequences
sample_X = np.array([[3, 89, 45]])
max_seq_len = 10
padded_sample_X = pad_sequences(sample_X, padding="post", maxlen=max_seq_len)
print (f"{sample_X} → {padded_sample_X}")
```
We will put all of these preprocessing utilities to use in the subsequent lessons.
---
<div align="center">
Subscribe to our <a href="https://practicalai.me/#newsletter">newsletter</a> and follow us on social media to get the latest updates!
<a class="ai-header-badge" target="_blank" href="https://github.com/practicalAI/practicalAI">
<img src="https://img.shields.io/github/stars/practicalAI/practicalAI.svg?style=social&label=Star"></a>
<a class="ai-header-badge" target="_blank" href="https://www.linkedin.com/company/practicalai-me">
<img src="https://img.shields.io/badge/style--5eba00.svg?label=LinkedIn&logo=linkedin&style=social"></a>
<a class="ai-header-badge" target="_blank" href="https://twitter.com/practicalAIme">
<img src="https://img.shields.io/twitter/follow/practicalAIme.svg?label=Follow&style=social">
</a>
</div>
</div>
| github_jupyter |
# Load
```
import pandas as pd
import numpy as np
gold = pd.read_csv(r"data/annual_gold_rate.csv")
display(gold.head())
```
# Clean
We're only interested in the USD gold prices, so we'll drop the other currency columns.
```
gold = gold[["Date", "USD"]]
gold.index = gold["Date"]
gold = gold.drop(columns=["Date"])
gold.index = pd.to_datetime(gold.index, format="%Y")
gold.index.freq = gold.index.inferred_freq
```
# EDA
```
import matplotlib.pyplot as plt
plt.figure(figsize=(12,6))
plt.plot(gold.index, gold["USD"])
plt.title("Annual Gold Price")
plt.xlabel("Date")
plt.ylabel("Price")
plt.grid()
plt.show()
```
## Decomposition/Anomaly Detection
We'll use STL Decomposition to look for anomalies in the data.
```
from statsmodels.tsa.seasonal import STL
stl = STL(gold["USD"], robust=True, period=12)
res = stl.fit()
#seasonality
season_x = res.seasonal.index
season_y = res.seasonal.values
plt.figure(figsize=(12,6))
plt.title("Gold Price (USD) per Ounce Seasonality")
plt.plot(season_x, season_y, c="teal")
plt.xlabel("Date")
plt.ylabel("Price")
plt.show()
#trend
trend_x = res.trend.index
trend_y = res.trend.values
#plot actual vs. trend
plt.figure(figsize=(12,6))
plt.title("Trend vs Observed Annual Gold Prices (USD) per Ounce")
plt.plot(gold["USD"], c="black", label="Actual Data")
plt.plot(trend_x, trend_y, c="g", label="Trend")
plt.xlabel("Date")
plt.ylabel("Price")
plt.legend()
plt.show()
#residual
res_x = res.resid.index
res_y = res.resid.values
#plot residual
plt.figure(figsize=(12,6))
plt.title("Annual Gold Prices (USD) per Ounce Noise")
plt.plot(res_x, res_y, ls="--", c='black')
plt.plot_date(res_x, res_y, color='g', label="Normal Data Points")
low_outlier_index = np.where(res.resid.values <= -150)
plt.plot_date(res_x[low_outlier_index], res_y[low_outlier_index], c="red")
high_outlier_index = np.where(res.resid.values >= 150)
plt.plot_date(res_x[high_outlier_index], res_y[high_outlier_index], c="red", label="Anomalies")
plt.xlabel("Date")
plt.ylabel("Price")
plt.legend()
plt.show()
```
# Model
For the model, we'll use Holt' Linear Trend Method (Double Exponential Smoothing) since the data appears to have high variation.
```
from statsmodels.tsa.holtwinters import ExponentialSmoothing
train = gold.iloc[:31]
test = gold.iloc[31:]
holts = ExponentialSmoothing(train, trend="mul")
holts_fit = holts.fit(optimized=True)
train_pred = holts_fit.predict(start=0)
test_pred = holts_fit.forecast((len(gold)-len(train)))
from sklearn.metrics import mean_squared_error
from math import sqrt
#observed vs predicted
plt.figure(figsize=(15,6))
plt.plot(train_pred, color="darkgreen", label="predicted train", ls="--")
plt.plot(test_pred, color="crimson", label="predicted test", ls="-.")
plt.plot(train, color="blue", label="train")
plt.plot(test, color="orange", label="test")
plt.title("Observed vs. Predicted Train/Test Gold Price (USD) per Ounce")
plt.legend()
plt.show()
mse = mean_squared_error(test, test_pred)
scatter_index = (sqrt(mse))/(test.mean())[0].round(3)
print("Scatter Index:", scatter_index)
#forecast
forecast = holts_fit.forecast((len(gold)-len(train))+5)
plt.figure(figsize=(15,6))
plt.plot(gold.index, gold["USD"], color="black", label="Observed")
plt.plot(forecast[(len(gold)-len(train)):], color="green", ls="--", label="Forecasted")
plt.title("Forecasted Annaual Gold Price (USD) per Ounce")
plt.legend()
plt.show()
print(forecast[(len(gold)-len(train)):])
```
| github_jupyter |
```
%matplotlib inline
# import statements
import numpy as np
import matplotlib.pyplot as plt #for figures
from mpl_toolkits.basemap import Basemap #to render maps
import math
import json #to write dict with parameters
import GrowYourIC
from GrowYourIC import positions, geodyn, geodyn_trg, geodyn_static, plot_data, data
plt.rcParams['figure.figsize'] = (8.0, 3.0) #size of figures
cm = plt.cm.get_cmap('viridis')
cm2 = plt.cm.get_cmap('winter')
age_ic_dim = 1e9 #in years
rICB_dim = 1221. #in km
translation_velocity_dim = 4.e-10
time_translation = rICB_dim*1e3/translation_velocity_dim /(np.pi*1e7)
maxAge = 2.*time_translation/1e6
units = None #we give them already dimensionless parameters.
rICB = 1.
age_ic = 1.
omega = 0.
omega_2_dim = 0.45 #degree/Myears
omega_2 = omega_2_dim*np.pi/180*age_ic_dim*1e-6
velocity_amplitude = translation_velocity_dim*age_ic_dim*np.pi*1e7/rICB_dim/1e3
velocity_center = [0., 100.]#center of the eastern hemisphere
center = [0,-80] #center of the western hemisphere
velocity = geodyn_trg.translation_velocity(velocity_center, velocity_amplitude)
exponent_growth = 0.1
proxy_type = "age"#"growth rate"
proxy_name = "age (Myears)" #growth rate (km/Myears)"
proxy_lim = [0, 220] #or None
proxy_lim2 = [0, maxAge] #or None
print("=== Model 1 ===")
print("The translation recycles the inner core material in {0:.2e} million years.".format(maxAge))
print("Translation velocity is {0:.2e} km/years, {1:.2}.".format(translation_velocity_dim*np.pi*1e7/1e3, velocity_amplitude))
print("Rotation rate is {0:.2e} degree per Myears, {1:.2e}.".format(omega, omega))
print("===")
geodynModel = geodyn_trg.TranslationGrowthRotation() #can do all the models presented in the paper
parameters = dict({'units': units,
'rICB': rICB,
'tau_ic':age_ic,
'vt': velocity,
'exponent_growth': exponent_growth,
'omega': omega,
'proxy_type': proxy_type})
geodynModel.set_parameters(parameters)
geodynModel.define_units()
print("=== Model 2 ===")
print("The translation recycles the inner core material in {0:.2e} million years.".format(maxAge))
print("Translation velocity is {0:.2e} km/years, {1:.2}.".format(translation_velocity_dim*np.pi*1e7/1e3, velocity_amplitude))
print("Rotation rate is {0:.2e} degree per Myears, {1:.2e}.".format(omega_2_dim, omega_2))
print("===")
geodynModel2 = geodyn_trg.TranslationGrowthRotation() #can do all the models presented in the paper
parameters2 = dict({'units': units,
'rICB': rICB,
'tau_ic':age_ic,
'vt': velocity,
'exponent_growth': exponent_growth,
'omega': omega_2,
'proxy_type': proxy_type})
geodynModel2.set_parameters(parameters2)
geodynModel2.define_units()
## real data set - WD13
data_set = data.SeismicFromFile("../GrowYourIC/data/WD11.dat")
data_set.method = "bt_point"
proxy1 = geodyn.evaluate_proxy(data_set, geodynModel, proxy_type=proxy_type, verbose=False)
proxy2 = geodyn.evaluate_proxy(data_set, geodynModel2, proxy_type=proxy_type, verbose=False)
# random data set -
data_set_random = data.RandomData(3000)
data_set_random.method = "bt_point"
proxy_random1 = geodyn.evaluate_proxy(data_set_random, geodynModel, proxy_type=proxy_type, verbose=False)
proxy_random2 = geodyn.evaluate_proxy(data_set_random, geodynModel2, proxy_type=proxy_type, verbose=False)
# perfect repartition in depth (for meshgrid plots)
data_meshgrid = data.Equator_upperpart(150,150)
data_meshgrid.method = "bt_point"
proxy_meshgrid = geodyn.evaluate_proxy(data_meshgrid, geodynModel, proxy_type=proxy_type, verbose = False)
proxy_meshgrid2 = geodyn.evaluate_proxy(data_meshgrid, geodynModel2, proxy_type=proxy_type, verbose = False)
fig, ax = plt.subplots(3,1,figsize=(6, 6), sharex=True)
X, Y, Z = data_meshgrid.mesh_RPProxy(proxy_meshgrid)
sc = ax[0].contourf(Y, rICB_dim*(1.-X), Z, 100, cmap=cm, vmin=proxy_lim2[0], vmax=proxy_lim2[1])
sc2 = ax[0].contour(sc, levels=sc.levels[::15], colors = "k")
ax[0].set_ylim(-0, 120)
ax[0].set_xlim(-180,180)
cbar = fig.colorbar(sc, ax=ax[0])
cbar.set_clim(proxy_lim2[0],proxy_lim2[1])
cbar.set_label(proxy_name)
ax[0].set_ylabel("depth below ICB (km)")
ax[0].invert_yaxis()
r, t, p = data_set_random.extract_rtp("bottom_turning_point")
#fig, ax = plt.subplots(figsize=(8, 2))
sc=ax[1].scatter(p,rICB_dim*(1.-r), c=proxy_random1, s=10,cmap=cm, linewidth=0, vmin=proxy_lim2[0], vmax=proxy_lim2[1])
ax[1].set_ylim(-0,120)
ax[1].invert_yaxis()
ax[1].set_xlim(-180,180)
cbar = fig.colorbar(sc, ax=ax[1])
ax[1].set_ylabel("depth below ICB (km)")
cbar.set_label(proxy_name)
r, t, p = data_set.extract_rtp("bottom_turning_point")
#fig, ax = plt.subplots(figsize=(8, 2))
sc=ax[2].scatter(p,rICB_dim*(1.-r), c=proxy1, s=10,cmap=cm, linewidth=0, vmin=proxy_lim2[0], vmax=proxy_lim2[1])
ax[2].set_ylim(-0,120)
ax[2].invert_yaxis()
ax[2].set_xlim(-180,180)
cbar = fig.colorbar(sc, ax=ax[2])
ax[2].set_xlabel("longitude")
ax[2].set_ylabel("depth below ICB (km)")
cbar.set_label(proxy_name)
plt.savefig("Fig3_1.pdf")
#fig3, ax3 = plt.subplots(figsize=(8, 2))
fig, ax = plt.subplots(3,1,figsize=(6, 6), sharex=True)
X, Y, Z = data_meshgrid.mesh_RPProxy(proxy_meshgrid2)
sc = ax[0].contourf(Y, rICB_dim*(1.-X), Z, 100, cmap=cm, vmin=proxy_lim[0], vmax=proxy_lim[1])
sc2 = ax[0].contour(sc, levels=sc.levels[::15], colors = "k", vmin=proxy_lim[0], vmax=proxy_lim[1])
ax[0].set_ylim(-0, 120)
ax[0].invert_yaxis()
ax[0].set_xlim(-180,180)
cbar = fig.colorbar(sc,ax=ax[0])
cbar.set_clim(proxy_lim[0],proxy_lim[1])
cbar.set_label(proxy_name)
ax[0].set_ylabel("depth below ICB (km)")
r, t, p = data_set_random.extract_rtp("bottom_turning_point")
#fig, ax = plt.subplots(figsize=(8, 2))
sc=ax[1].scatter(p,rICB_dim*(1.-r), c=proxy_random2, s=10,cmap=cm, linewidth=0, vmin=proxy_lim[0], vmax=proxy_lim[1])
ax[1].set_ylim(-0,120)
ax[1].invert_yaxis()
ax[1].set_xlim(-180,180)
cbar = fig.colorbar(sc,ax=ax[1])
#if proxy_lim is not None:
# cbar.set_clim(0, maxAge)
ax[1].set_ylabel("depth below ICB (km)")
cbar.set_label(proxy_name)
r, t, p = data_set.extract_rtp("bottom_turning_point")
#fig, ax = plt.subplots(figsize=(8, 2))
sc=ax[2].scatter(p,rICB_dim*(1.-r), c=proxy2, s=10,cmap=cm, linewidth=0, vmin=proxy_lim[0], vmax=proxy_lim[1])
ax[2].set_ylim(-0,120)
fig.gca().invert_yaxis()
ax[2].set_xlim(-180,180)
cbar = fig.colorbar(sc,ax=ax[2])
#cbar.set_clim(0, 250)
ax[2].set_xlabel("longitude")
ax[2].set_ylabel("depth below ICB (km)")
cbar.set_label(proxy_name)
plt.savefig("Fig3_2.pdf")
```
| github_jupyter |
# Roemer Has It: The Hilbert Transform, Instantaneous Power/Phase/Frequency, and Negative Frequencies in Neuroscience
If you work with oscillations in the brain, or any sort of time-frequency analysis of neural (or other) time series, you've probably used the Hilbert transform at some point. Depending on your background and level of comfort with signal processing, it may be just a scipy or MATLAB function call to you, or you might understand the deeper mathematical theory behind it. What you might not appreciate (I certainly didn't), however, is one of the important assumptions behind it, specifically in the context of applying it to brain data. In particular, it's an important assumption in interpreting instantaneous oscillation power and phase in brain rhythms, especially if you've ever thought to yourself, "what does **instantaneous** power mean anyway?"
In the following post, I delve into the math behind the Hilbert Transform and provide some visualizations that will hopefully make these ideas clearer. When I was researching for this, seeing the complex sinusoids represented as vectors really helped me in understanding it better, as well as in seeing the elegance of the math. I will also clarify what scipy.signal.hilbert() is actually doing (hint: it's not computing the Hilbert Transform). You should have a basic exposure to frequency analysis (Fourier Transform) and filtering, i.e. know what they do. But no more than that is necessary, as this post will unpack some fundamentals of the Fourier Transform and complex sinusoids.
Note: the most confusing part about this post is probably the switching of representation between time domain and frequency domain, and at times, time-frequency domain. Be wary when this happens, and I will clearly signpost when I make a jump (thanks for the feedback, Tammy!)
I'm uploading this as a blog post mainly to see the swagged out markdown embedded code blocks, but notebook is identical and can be found [here](https://github.com/rdgao/roemerhasit/blob/master/RHI_Hilbert_Transform.ipynb).
# Instantaneous measures and the Hilbert Transform.
If you work with (M)EEG/ECoG/LFP, or even EMG, you may have computed instantaneous power. There's a long list of studies that looks at stimulus-evoked and time-resolved measures of, for example, oscillatory and high-frequency power (Crone 1998, first example that comes to mind.) If you're in the even more niche community of people for whom "phase-amplitude coupling" rings a bell (hello friends), then you're definitely familiar with these instantaneous measures.
Typically, how you arrive at these measures is through 1) narrowband filtering your signal, 2) get its complex representation via the Hilbert transform, and 3) compute squared magnitude/phase/phase difference. 1) and 2) can be combined via a complex wavelet filter, or obtained via FFT, but I will stick with the above because I want to write about the Hilbert Transform.
In particular, when I first started writing this post, I wanted to address two thoughts that always vaguely bothered me, but could never figure out why or where I got them from:
##### 1. "Instananeous" measures are suspicious, because how do you measure power/frequency/phase truly at an instant? Can you look at a single point on a sine wave (without the neighboring points) and determine the amplitude?
##### 2. The Hilbert Transform is mathematically well-defined only for narrowband signals, and breaks down when the signal is wideband (energy at many frequencies).
Turns out, both of the above were false beliefs. So I'm passing on my experience so you don't have to go through the struggle.
---
So what does the Hilbert Transform (HT) do? HT, in the most pragmatic sense, is a function that creates information. I know this sounds sacriligeous, but stay with me. If you read the docstring for scipy.signal.hilbert(), it says
```
"Compute the analytic signal, using the Hilbert Transform".
```
In other words, it turns a real-valued signal into a complex-valued signal - it's **analytic** form - which means turning a 1-D time series (real) into a 2-D time series (complex: real + imag).
For purpose of demonstration, I will use a snippet of rat hippocampus local field potential data (courtesy of CRCNS, dataset hc2), because of its strong theta oscillation. The following few steps may look familiar, as I will first be filtering the raw signal for its theta component, then computing the Hilbert Transform.
```
%matplotlib inline
import numpy as np
import scipy as sp
import matplotlib.pyplot as plt
plt.rcParams["axes.labelsize"]=20
plt.rcParams["font.size"]=15
plt.rcParams["font.family"] = "Arial"
from scipy import io, signal
# our open source lab package, free for all to use/contribute :)
# https://github.com/voytekresearch/neurodsp
import neurodsp as ndsp
# loading the test data
data = io.loadmat('data/sample_data_2.mat', squeeze_me=True)
x = data['x'][:20000]
fs = data['fs']
t = np.arange(len(x))/fs
# filtering and hilbert transform
x_filt = ndsp.filter(x, fs, 'bandpass', fc=(4,12), remove_edge_artifacts=False)
x_a = signal.hilbert(x_filt)
# plotting
plt.figure(figsize=(15,6))
plt.subplot(2,1,1)
plt.plot(t,x, 'k', label='Raw Signal', alpha=0.4, lw=1)
plt.plot(t,x_filt, 'r', label='Filtered Signal', lw=2)
plt.ylabel('Real Signal')
plt.legend()
plt.xlim((0,3));
plt.subplot(2,1,2)
plt.plot(t,x_filt, '--r', label='Filtered Signal', alpha=1, lw=1)
plt.plot(t,x_a.real, label='Hilbert Real', alpha=0.5, lw=2)
plt.plot(t,x_a.imag, label='Hilbert Imag', alpha=0.5, lw=2)
plt.xlabel('Time (s)')
plt.ylabel('Complex Signal')
plt.legend(loc='upper left')
plt.xlim((0,3));
# selecting 2 slices of data
t1 = 2650
t2 = 2700
yb = plt.ylim()
plt.plot([t1/fs]*2, yb, 'k--')
plt.plot([t2/fs]*2, yb, 'g--')
plt.tight_layout()
```
In the first plot, you can see the raw hippocampus LFP (black) and the filtered theta component overlaid. If you comment on how non-sinusoidal the oscillation looks without provocation, your IP will be banned forever (joking but also don't).
In the second plot are the filtered theta (dashed red), and the real (blue) and imaginary (orange) components of the analytic signal. Notice that the filtered signal and the real component of the analytic signal are completely overlapping, which is the first hint of what the Hilbert Transform does: it retrieves an imaginary component. This is what I meant by "creating information", but hold that protest still.
Also, notice that the imaginary component is a) almost identical to, and b) slightly lagging the real component - by exactly 90 degrees, or pi/2 rad, in fact. At this point, those of you familiar with this analysis pipeline should anticipate the next few functions calls, which computes the "instantaneous" power and phase from the filtered theta signal.
```
x_power = np.abs(x_a)**2
x_phase = np.angle(x_a)
# plotting
plt.figure(figsize=(15,6))
plt.subplot(2,1,1)
plt.plot(t,x_power, '.k', ms=1)
plt.yticks([])
plt.ylabel('Power')
plt.xlim((0,3));
plt.subplot(2,1,2)
plt.plot(t,x_phase, '.k', ms=1)
plt.ylabel('Phase')
plt.xlabel('Time (s)')
yb = plt.ylim()
plt.plot([t1/fs]*2, yb, 'k--')
plt.plot([t2/fs]*2, yb, 'g--')
plt.xlim((0,3))
plt.tight_layout()
```
Like I mentioned above, having worked on these methods for some time now, I've sometimes felt an air of mysticism, and sometimes, skepticism, about how instantaneous measures could be computed on a signal. This is because I had an intuitive notion of what instantaneous amplitude (power) and phase are: amplitude is roughly the signal envelope of the sinusoid, and phase is the, well, phase of the sinusoid in that instant.
Implictly, my notion of amplitude and phase is in accordance with the definition of a sinusoid as x(t) = Asin(wt+$\phi$), where A is the amplitude and $\phi$ is the phase lag. But given just a single point in a sinusoid-like real-valued signal, how does one decide what its instantaneous power is, if it's at any point in time other than the peak or trough, such as the two instants I've highlighted above in black and green?
But as you might have inferred from the above plots, that is actually a misconception. The instantaneous power is not computed on the filtered signal, but the analytic signal, which is the whole point of using the Hilbert Transform. Namely, it is the squared magnitude of the complex vector at an instant in time. Similarly, the instantaneous phase is the phase angle between the vector and the real axis (0 deg) in the complex plane. Mathematically, this refers to the complex sinusoid, i.e., x(t) = Ae^(iwt) in Euler form, or x(t) = A(cos(wt)+isin(wt)) in rectangular form.
**Warning**: now switching from temporal representation (time on x-axis) to the complex plane, where each dot is an instant (sample) in time.
```
plt.figure(figsize=(6,6))
plt.axhline(color='k', lw=1)
plt.axvline(color='k', lw=1)
plt.plot(x_a.real[:5000],x_a.imag[:5000], 'k.-', alpha=0.1, ms=2)
plt.quiver(0,0,x_a[t1].real,x_a[t1].imag, angles='xy', scale_units='xy', scale=1, color='k')
plt.quiver(0,0,x_a[t2].real,x_a[t2].imag, angles='xy', scale_units='xy', scale=1, color='g')
plt.xlabel('Real');plt.ylabel('Imaginary')
plt.box('off')
plt.title('Vector 1(k) phase: %.2f, vector 2(g) phase: %.2f'%(x_phase[t1]/(2*np.pi)*360, x_phase[t2]/(2*np.pi)*360));
```
In the above plot, I've traced out the first 5 seconds of the analytic signal in the complex domain, where each small dot represents a vector in an instant in time (with its arrow-tail removed), and the black and green arrows refer to those two moments marked in dashed lines in the previous plot. What you can see is that the signal traces out a relatively smooth circular trajectory, with a varying radius. At any moment in time, the radius squared is the signal power, and its angle from the positive real line is the phase: so the black arrow is at about -30 degrees (or 330), and the green one at 75 degrees.
Hopefully, this clears up any confusion on how instantaneous power and phase are computed. I have to emphasize here that these are not mystical or vaguely intuitive measures at all, as I had thought, but mathematically well-defined quantities of every complex signal. Which now brings us back to the original question: how does the Hilbert Transform create the imaginary component of a real signal, out of thin air?
# Your Hilbert Transform is a lie.
To understand how the Hilbert Transform produces the analytic signal, we have to delve into the Fourier Transform and the (complex) sinusoidal bases. But first, a clarification.
If you read through the [Wikipedia article on the Hilbert Transform](https://en.wikipedia.org/wiki/Hilbert_transform), it says:
```
The Hilbert transform has a particularly simple representation in the frequency domain: it imparts a phase shift of 90° to every Fourier component of a function.
```
Technically, the Hilbert Transform as defined mathematically is **not what scipy.signal.hilbert() or hilbert() in MATLAB does**. What those functions do is to return the analytic signal, which is a complex signal defined as:
x_a(t) = x(t) + iHT{x(t)} = Re(x_a(t)) + Im(x_a(t)).
In other words, the mathematical Hilbert Transform produces the imaginary component of the analytic signal, which is not what scipy.signal.hilbert() returns. See the full documentation here:

(Ignore the middle part of that formula for now.)
The conflict is somewhat confusing, but was implemented as such probably because the HT is used almost exclusively in the context of computing the analytic signal. From this point on, I will be precise and use "analytic signal" for x_a(t), and "Hilbert Transform" (noun) for y=HT{x(t)} (note, without the 'i' attached), "Hilbert Transform" (verb) for the operation HT{ }, and hilbert() for the scipy function. This is the mathematically correct usage, but contradicts our everyday colloquial usage in neuroscience (and more broadly, scientific computing).
EDIT: it is at this point that I went back and changed all my variable names from ```x_ht``` to ```x_a```.
Noting the Wikipedia quote, this is consistent with what we saw in the second subpanel of the very first figure above: the imaginary component was about 90 degrees phase-lagged to the filtered theta oscillation. However, this is not always the case. Let's see what happens when we compute the analytic signal of the raw wideband signal.
```
# plotting
plt.figure(figsize=(15,6))
plt.subplot(2,1,1)
plt.plot(t,signal.hilbert(x).real, label='Hilbert Real', lw=1)
plt.plot(t,signal.hilbert(x).imag, label='Hilbert Imag', lw=1)
plt.xlabel('Time (s)')
plt.legend()
plt.xlim((2,3));
```
**(Back to time domain!)**
I've zoomed in on a shorter segment of the signal for clarity. You can see that, roughly speaking, the imaginary component (orange) is still lagged about 90 deg compared to the real component, and this is due to the phase shift of the dominant theta frequency. However, it's not a literal time shift of the whole signal, since many of the more squiggly high-frequency ripple components are not exactly copied over but remain locked to the same theta phase (e.g., see ripple at 2.3s). As the Wikipedia article states, the Hilbert transform imparts a 90 degrees phase shift to **every** Fourier component, not just the strongest (theta) component. In other words, every complex Fourier coefficient in the raw broadband signal is delayed by 90 degrees.
Why does introducing a 90-degree lag produce the imaginary component of the signal? Finally, we are ready for the part that I enjoy the most: the relationship between the Hilbert Transform and the Fourier Transform, in vectors. Bonus, we make our own DIY Hilbert Transform.
### Negative Frequencies
Going back to the beginning of the tutorial, I mentioned that the Hilbert Transform creates information, which is in the form of the imaginary component of the analytic signal. That's actually a lie, technically. From the perspective of the time-domain signal, indeed, it seems to have generated a 2-dimensional (complex) signal from a 1-dimensional (real, scalar) signal. The information it "creates", however, **comes from a specific assumption**, and when viewed in the frequency (or Fourier) domain, it's actually not creating information at all, but splitting the original signal in two. I will illustrate this with a perfect sine wave first.
```
# create a sine wave for 5 seconds at 14Hz, with slight phase delay
t_trunc = t[:5000]
x_sin = np.cos(2*np.pi*14*t_trunc+0.5)
F_sin = np.fft.fft(x_sin, norm='ortho')
f_axis = np.fft.fftfreq(len(x_sin),1/fs)
plt.figure(figsize=(15,10))
# time domain plot
plt.subplot(4,1,1)
plt.plot(t_trunc,x_sin)
plt.xlabel('Time (s)')
plt.ylabel('Real Signal')
plt.xlim((0,3))
# frequency domain plots
plt.subplot(4,1,2)
plt.plot(f_axis, np.abs(F_sin)**2,'ko-')
plt.ylabel('Power')
plt.title('Fourier Power')
plt.xlim([-500,500])
plt.subplot(4,1,3)
plt.plot(f_axis, F_sin.real, 'ko-')
plt.title('Fourier Coeff. Real')
plt.xlim([-500,500])
plt.subplot(4,1,4)
plt.plot(f_axis, F_sin.imag, 'ko-', label='Fourier Imag')
plt.title('Fourier Coeff. Imag')
plt.xlim([-500,500])
plt.xlabel('Frequency (Hz)')
plt.tight_layout()
```
**Warning: jumping between time (1st plot) and frequency (2nd-4th plot) domain.**
The first plot shows the sine wave in time domain, and the next three show the Fourier power (squared amplitude), as well as the real and imaginary components of the Fourier coefficients. A few things will jump out at you. First, Fourier power is zero everywhere except two points, which are at 14 and -14 Hz. In neuroscience, it's almost always the case that power spectra are plotted with only the positive frequencies, i.e., the right half. This is because for any real-valued signal, as all brain recordings are, the power spectrum is symmetrical about 0Hz, so plotting the left side is just wasting space. **But sometimes we're so used to looking at the positive half of the spectrum that we forget the other half exists, or matters at all.**
Here's why it matters. The 14 Hz power we see **does not** represent the squared magnitude of our scalar sinusoid at f=14 Hz, i.e. x(t) = cos($2\pi$ft), which one might assume by looking just at the positive frequencies. Instead, it's the squared magnitude of the **complex sinusoid** at 14 Hz, in the form of z(t) = Aexp(i$2\pi$ft). This is because the basis functions for Fourier decomposition are not real sinusoids, but complex sinusoids.
Ugh, basis functions - it got a little gnarly, so I think you'll like this next part.
In rectangular form, the complex sinusoid looks like z(t) = Acos($2\pi$ft) + iAsin($2\pi$ft). Wait a minute, that's just our original signal (cos($2\pi$ft)) with an unwanted imaginary part tacked onto the end! The original signal is purely real, so how do we get rid of this imaginary sine component? The answer is in the negative frequency. Specifically, at -14Hz.
```
fc_pos = F_sin[np.where(f_axis==14)[0]][0]
fc_neg = F_sin[np.where(f_axis==-14)[0]][0]
plt.figure(figsize=(4,4))
plt.axhline(color='k', lw=1)
plt.axvline(color='k', lw=1)
plt.quiver(0,0,fc_pos.real,fc_pos.imag, angles='xy', scale_units='xy', scale=1, label='+ 14Hz', color='b',headwidth=5)
plt.quiver(0,0,fc_neg.real,fc_neg.imag, angles='xy', scale_units='xy', scale=1, label='- 14Hz', color='r',headwidth=5)
plt.quiver(0,0,(fc_pos+fc_neg).real,(fc_pos+fc_neg).imag, angles='xy', scale_units='xy', scale=1, label='Sum', color='k',headwidth=5)
plt.legend(loc='upper left')
plt.xlim((-65,65)); plt.ylim((-50,50))
plt.xlabel('Real');plt.ylabel('Imaginary')
plt.box('off')
```
**(Back to complex representation!)**
The three vectors above represent the complex sinusoids at 14Hz (blue) and -14Hz (red) in the complex plane, and their vector sum (black). They're not exactly like the black and green ones from a few plots above. Those previous ones represent two instants of the same complex sinusoidal function as it evolves in time, while these represent two complex sinusoids extracted from our made up signal, and their sum. There is one thing in common, however, which is that you could imagine the blue and red vectors here rotating about the origin as progression of time, which corresponds to phase advancing the time series. The only extra rule is that the **blue vector rotates counterclockwise**, since it's a positive frequency, and **the red vector rotates clockwise but at the same speed** (their frequency (14Hz) in radians).
It's no coincidence that the negative frequency vector is reflected about the real axis from the positive frequency - this is how the imaginary component is cancelled out to form our real-value only signal. Since the two colored vectors always mirror each other about the real axis as they rotate, their summation (black vector) will **always** fall exactly onto the real axis. As the colored vectors rotate about, the black vectors will shrink and grow, representing the value of the real sinusoidal function in time.
In math, this cancellation of imaginary components amounts to:
z(t) + z'(t) = A(cos($2\pi$ft) + isin($2\pi$ft)) + A(cos($2\pi$ft) - isin($2\pi$ft)) = 2Acos($2\pi$ft) = 2Ax(t),
and thus, we get back our signal (with a scaling constant of A=1/2 on the complex sinusoids). To further verify that this is accurate, we can check that the energy from the two sinusoids sum to be equal to the energy of the time series, which is just the sum of squares, or ```var(signal)*len(signal)```, in this case (Parseval's theorem).
```
print('Signal energy=%.2f, Fourier energy=%.2f'%(np.sum(x_sin**2), np.abs(fc_pos)**2+np.abs(fc_neg)**2))
```
# Hilbert Transform, revisted.
Armed with the concept of negative frequencies, let's work out what ```scipy.signal.hilbert()``` is doing. Following the perfect sinusoid example, the goal is to transform x(t) = cos($2\pi$ft) to its analytic signal, x_a(t) = x(t) + iy(t) = cos($2\pi$ft) + isin($2\pi$ft). Graphically, this amounts to getting the blue vector from the black vector. Without looking at the math, we should have an idea of what to do here, given the forward steps we took from the blue and red vector to the black vector - we simply need to erase the red vector and multiply the blue vector by 2. Turns out, this is exactly what scipy.signal.hilbert() does:

Briefly, the positive step function U retains the positive frequencies of the Fourier Transform, while multiplying by 2.
"The negative half of the frequency spectrum is zeroes out." In other words, deleting the negative frequency complex sinusoid(s) transforms the real-valued signal to the analytic signal. I thought this shortcut is honestly pretty cool, and makes a lot more sense after understanding the geometrical interpretation of the negative frequency sinusoids. Also, this probably points to the real reason for implementing scipy.signal.hilbert() as getting the analytic signal directly - it's a lot easier than getting the actual Hilbert Transform.
**More importantly, nothing about how the Hilbert is defined restricts its usage to a narrowband signal, debunking misconception #2 (mentioned at the beginning).** The complex (analytic) signal generated by HT is perfectly valid and well-defined at all points.
Finally, let's see if we can implement our own Hilbert Transform function by following these steps, with the **correct** name.
```
def analytical_sig(x):
Fx = np.fft.fft(x) # get fft
f_axis = np.fft.fftfreq(len(x))
Fx[np.where(f_axis<0)]=0. # zero out negative frequencies
return np.fft.ifft(Fx*2) # return 2x positive frequencies
x_a = signal.hilbert(x)
x_a2 = analytical_sig(x)
plt.figure(figsize=(15,3))
plt.plot(t,x_a.imag, alpha=0.5, label='scipy hilbert', lw=2)
plt.plot(t,x_a2.imag, 'k--', alpha=0.5, label='custom hilbert', lw=1)
plt.xlim((0,3))
plt.xlabel('Time (s)')
plt.legend()
```
Perfect overlap, passes the sniff test.
That is all. I hope this provided some intuition for what the Hilbert Transform is and isn't doing, debunked some (I hope) common misconceptions, as well as explained what instantaneous power/phase is in brain signals.
### Some last thoughts:
- Remember that x_a(t) = x(t) + jHT{x(t)}, so how do we actually compute the Hilbert Transform without going directly to the analytic signal, and how does the Hilbert Transform negate the negative frequency component? This [tutorial][1] does an in-depth explanation, which has some really simple but elegant complex math, as well as neat visualizations that should be accessible if you've been following up to this point.
- I think it's hilarious that scipy.signal.hilbert() not only does not return the Hilbert Transform, as its function name would suggest, it doesn't even compute the Hilbert Transform at all. The last part of the docstring basically says you can GET the Hilbert Transform of the signal yourself if you want to, by just taking the imaginary component, but why would you want to? There's gotta be a joke about programmers and engineers being practical here.
- I'm repeating myself here, but it's worth it: I had a misconception that the Hilbert Transform only "existed" for narrowband signals. This is not true, as we clearly demonstrated above. HT is a well defined mathematical operation regardless of the bandwidth of the signal. The issue here is more so in the context of neuroscientific data, i.e., how to interpret the Hilbert Transform of a signal that doesn't have a dominant oscillatory mode, specifically its instantaneous power and phase. If we are not careful here and just apply the method, we can extract a phase time series for all signals, but may or may not be valid. In the last 5 years, we've started to notice that oscillations are highly non-stationary, and sometimes not even present at all in the purport canonical bands (e.g., alpha). This gets at the broader question of bandpassing a signal for an oscillatory component when there is none, but that's discussion for another day. In short, the HT is not to blame, but narrowband filtering. (See [bycycle](https://voytekresearch.github.io/bycycle/auto_tutorials/plot_1_bycycle_philosophy.html#sphx-glr-auto-tutorials-plot-1-bycycle-philosophy-py) and [fooof](https://voytekresearch.github.io/fooof/) for more indepth exposition of the issues and our methods of dealing with them.)
- Actually, I will just say one more thing about that, and this is worth **emphasizing**. There is a fine but ideological difference between generative model and descriptive analysis. Using Fourier analysis to compute the band-limited energy does not mean you subscribe to the model that the signal is generated by sinusoids, as its simply a description (or representation) of the signal. The time series itself is another representation - you can represent that same signal in infinitely many ways. Using the Hilbert Transform is also not subscribing to a particular generative model - you don't have to presume a complex sinusoidal generator to describe it as one. However, there is a fine line you cross when you interpret the transformed representations as being generated by an implicit model, like claiming the band-limited phase time series as the phase of a sole oscillator at that frequency. This does subscribe to the model that there **is** an underlying oscillator in the signal, which is not at all implied by the descriptive analysis. Fourier analysis is very useful in many aspects, especially sparse representation of oscillatory signals. But it's not as useful in faithfully (and sparsely) describing non-sinusoidal (nonlinear) oscillations and non-oscillatory phenomena.
- I've actually never used complex sinusoidal or wavelet filters, only real-valued ones. But all the above holds, as filtering with a complex wavelet bypasses the need to apply the Hilbert Transform, since it's, in a sense, transformed (into the complex time series) as it is filtered. More details in the Mike X Cohen ref below.
- As with almost all neuroscientific time series analyses - (the late) Walter Freeman has written about it. I stumbled upon this as I was finishing up this post. Had I seen it earlier, I might not have even written it.
### Sources:
[1]:https://ccrma.stanford.edu/~jos/mdft/Analytic_Signals_Hilbert_Transform.html
[Phase Quadrature](https://ccrma.stanford.edu/~jos/mdft/Analytic_Signals_Hilbert_Transform.html)
[Complex Sinusoids](https://www.dsprelated.com/freebooks/filters/Plotting_Complex_Sinusoids_Circular.html)
[Symmetry](https://www.dsprelated.com/freebooks/mdft/Symmetry.html)
[Mike X Cohen lecture](https://www.youtube.com/watch?list=PLn0OLiymPak3jjr0hHI9OFXuQyPBQlFdk&v=VyLU8hlhI-I)
[scipy doc](https://docs.scipy.org/doc/scipy/reference/generated/scipy.signal.hilbert.html)
[Wikipedia](https://en.wikipedia.org/wiki/Hilbert_transform)
[Walter Freeman on HT for brainwaves](http://www.scholarpedia.org/article/Hilbert_transform_for_brain_waves)
### Gratitude
Thanks to Tammy, Scott, Paolo, and Brad for reading an earlier version of this and providing feedback.
| github_jupyter |
All the IPython Notebooks in this lecture series are available at https://github.com/rajathkumarmp/Python-Lectures
## Strings
Strings are ordered text based data which are represented by enclosing the same in single/double/triple quotes.
```
String0 = 'Taj Mahal is beautiful'
String1 = "Taj Mahal is beautiful"
String2 = '''Taj Mahal
is
beautiful'''
print (String0 , type(String0))
print (String1, type(String1))
print (String2, type(String2))
```
String Indexing and Slicing are similar to Lists which was explained in detail earlier.
```
print (String0[4])
print (String0[4:])
```
### Built-in Functions
**find( )** function returns the index value of the given data that is to found in the string. If it is not found it returns **-1**. Remember to not confuse the returned -1 for reverse indexing value.
```
print (String0.find('al'))
print (String0.find('am'))
```
The index value returned is the index of the first element in the input data.
```
print (String0[7])
```
One can also input **find( )** function between which index values it has to search.
```
print (String0.find('j',1))
print (String0.find('j',1,3))
```
**capitalize( )** is used to capitalize the first element in the string.
```
String3 = 'observe the first letter in this sentence.'
print (String3.capitalize())
```
**center( )** is used to center align the string by specifying the field width.
```
String0.center(70)
```
One can also fill the left out spaces with any other character.
```
String0.center(70,'-')
```
**zfill( )** is used for zero padding by specifying the field width.
```
String0.zfill(30)
```
**expandtabs( )** allows you to change the spacing of the tab character. '\t' which is by default set to 8 spaces.
```
s = 'h\te\tl\tl\to'
print (s)
print (s.expandtabs(1))
print (s.expandtabs())
```
**index( )** works the same way as **find( )** function the only difference is find returns '-1' when the input element is not found in the string but **index( )** function throws a ValueError
```
print (String0.index('Taj'))
print (String0.index('Mahal',0))
print (String0.index('Mahal',10,20))
```
**endswith( )** function is used to check if the given string ends with the particular char which is given as input.
```
print (String0.endswith('y'))
```
The start and stop index values can also be specified.
```
print (String0.endswith('l',0))
print (String0.endswith('M',0,5))
```
**count( )** function counts the number of char in the given string. The start and the stop index can also be specified or left blank. (These are Implicit arguments which will be dealt in functions)
```
print (String0.count('a',0))
print (String0.count('a',5,10))
```
**join( )** function is used add a char in between the elements of the input string.
```
','.join('1234')
```
'1234' is the input string and char ',' is added in between each element
**join( )** function can also be used to convert a list into a string.
```
a = list(String0)
print (a)
b = ''.join(a)
print (b)
```
Before converting it into a string **join( )** function can be used to insert any char in between the list elements.
```
c = '/'.join(a)[18:]
print (c)
```
**split( )** function is used to convert a string back to a list. Think of it as the opposite of the **join()** function.
```
d = c.split('/')
print (d)
```
In **split( )** function one can also specify the number of times you want to split the string or the number of elements the new returned list should conatin. The number of elements is always one more than the specified number this is because it is split the number of times specified.
```
e = c.split('/',3)
print (e)
print (len(e))
```
**lower( )** converts any capital letter to small letter.
```
print (String0)
print (String0.lower())
```
**upper( )** converts any small letter to capital letter.
```
String0.upper()
```
**replace( )** function replaces the element with another element.
```
String0.replace('Taj Mahal','Sheffield')
```
**strip( )** function is used to delete elements from the right end and the left end which is not required.
```
f = ' hello '
```
If no char is specified then it will delete all the spaces that is present in the right and left hand side of the data.
```
f.strip()
```
**strip( )** function, when a char is specified then it deletes that char if it is present in the two ends of the specified string.
```
f = ' ***----hello---******* '
f.strip('*')
```
The asterisk had to be deleted but is not. This is because there is a space in both the right and left hand side. So in strip function. The characters need to be inputted in the specific order in which they are present.
```
print (f.strip(' *'))
print (f.strip(' *-'))
```
**lstrip( )** and **rstrip( )** function have the same functionality as strip function but the only difference is **lstrip( )** deletes only towards the left side and **rstrip( )** towards the right.
```
print (f.lstrip(' *'))
print (f.rstrip(' *'))
```
<div class="alert alert-success">
<b>EXERCISE</b>: Create a list of strings, join them using a comma as separator, and replace a part of the string using replace
</div>
```
a=['1','2','3','4']
b=','.join(a)
b
b.replace('1','hello')
```
## Dictionaries
Dictionaries are more used like a database because here you can index a particular sequence with your user defined string.
To define a dictionary, equate a variable to { } or dict()
```
d0 = {}
d1 = dict()
print (type(d0), type(d1))
```
Dictionary works somewhat like a list but with an added capability of assigning it's own index style.
```
d0['One'] = 1
d0['OneTwo'] = 12
print (d0)
```
That is how a dictionary looks like. Now you are able to access '1' by the index value set at 'One'
```
print (d0['One'])
```
Two lists which are related can be merged to form a dictionary.
```
names = ['One', 'Two', 'Three', 'Four', 'Five']
numbers = [1, 2, 3, 4, 5]
```
**zip( )** function is used to combine two lists
```
d2 = zip(names,numbers)
print (d2)
```
The two lists are combined to form a single list and each elements are clubbed with their respective elements from the other list inside a tuple. Tuples because that is what is assigned and the value should not change.
Further, To convert the above into a dictionary. **dict( )** function is used.
```
a1 = dict(d2)
print (a1)
```
### Built-in Functions
**clear( )** function is used to erase the entire database that was created.
```
a1.clear()
print (a1)
```
Dictionary can also be built using loops.
```
for i in range(len(names)):
a1[names[i]] = numbers[i]
print (a1)
```
**values( )** function returns a list with all the assigned values in the dictionary.
```
a1.values()
```
**keys( )** function returns all the index or the keys to which contains the values that it was assigned to.
```
a1.keys()
```
**items( )** is returns a list containing both the list but each element in the dictionary is inside a tuple. This is same as the result that was obtained when zip function was used.
```
a1.items()
```
**pop( )** function is used to get the remove that particular element and this removed element can be assigned to a new variable. But remember only the value is stored and not the key. Because the is just a index value.
```
a2 = a1.pop('Four')
print (a1)
print (a2)
```
<div class="alert alert-success">
<b>EXERCISE</b>: Create a dictionary of countries with corresponding population size (invented), starting from two lists and using zip, then remove the first country and assign it to a new variable
</div>
```
countries = ['USA','UK']
population = [100,10]
result = dict(zip(countries,population))
usa = result.pop('USA')
print(result)
print(usa)
```
| github_jupyter |
```
#Importing all required packages
import pandas as pd
import numpy as np
import seaborn as sns
import re
import string
from string import punctuation
import nltk
from nltk.corpus import stopwords
nltk.download("stopwords")
from nltk import pos_tag
from nltk.tokenize import WhitespaceTokenizer
from nltk.stem import WordNetLemmatizer
nltk.download('averaged_perceptron_tagger')
nltk.download('wordnet')
nltk.download('vader_lexicon')
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
#Downloading dataset
df = pd.read_csv("Womens Clothing E-Commerce Reviews.csv")
#Dropping unnecessary columns
df = df.drop(['Title', 'Positive Feedback Count', 'Sr. No.'], axis=1)
#Dropping rows containing missing values, data cleaning
df.dropna(inplace = True)
df.head()
#Plotting the ratings on a graph - visualizing data
sns.set_style('whitegrid')
sns.countplot(x='Rating', data=df, palette='YlGnBu_r')
#Calculation of Polarity
df['Polarity Rating'] = df['Rating'].apply(lambda x: 'Positive' if x > 3 else('Neutral' if x == 3 else 'Negative'))
df.head()
#Plotting polarity on a graph - Visualizing data
sns.set_style('whitegrid')
sns.countplot(x='Polarity Rating', data=df, palette = 'summer')
from nltk.corpus import wordnet
def get_wordnet_pos(pos_tag):
if pos_tag.startswith('J'):
return wordnet.ADJ
elif pos_tag.startswith('V'):
return wordnet.VERB
elif pos_tag.startswith('N'):
return wordnet.NOUN
elif pos_tag.startswith('R'):
return wordnet.ADV
else:
return wordnet.NOUN
#Data Preprocessing
df_positive = df[df['Polarity Rating'] == 'Positive'][0:8000]
df_negative = df[df['Polarity Rating'] == 'Negative']
df_neutral = df[df['Polarity Rating'] == 'Neutral']
#Sample negative and neutral polarity dataset and create final dataframe
df_neutral_final = df_neutral.sample(8000, replace = True)
df_negative_final = df_negative.sample(8000, replace = True)
df = pd.concat([df_positive, df_negative_final, df_neutral_final], axis=0)
#Text Processing
def textProcessing(text):
text = text.lower()
#To remove punctuation
text = [word.strip(string.punctuation) for word in text.split(" ")]
#To remove words that contain numbers
text = [word for word in text if not any(c.isdigit() for c in word)]
#To remove stop words
stop = stopwords.words('english')
text = [x for x in text if x not in stop]
#To remove empty tokens
text = [t for t in text if len(t) > 0]
#Pos tag text
pos_tags = pos_tag(text)
#Lemmatize text
text = [WordNetLemmatizer().lemmatize(t[0], get_wordnet_pos(t[1])) for t in pos_tags]
#To remove words with only one letter
text = [t for t in text if len(t) > 1]
#Now, joining all
text = " ".join(text)
return text
df['Review'] = df['Review Text'].apply(textProcessing)
df.head()
from nltk.sentiment.vader import SentimentIntensityAnalyzer
sid = SentimentIntensityAnalyzer()
df['Sentiments'] = df['Review'].apply(lambda x: sid.polarity_scores(x))
df = pd.concat([df.drop(['Sentiments'], axis = 1), df['Sentiments'].apply(pd.Series)], axis = 1)
df.head()
df['Number of Characters'] = df['Review'].apply(lambda x: len(x))
df['Number of Words'] = df['Review'].apply(lambda x: len(x.split(" ")))
df.head()
from gensim.test.utils import common_texts
from gensim.models.doc2vec import Doc2Vec, TaggedDocument
documents = [TaggedDocument(doc,[i]) for i,doc in enumerate(df['Review'].apply(lambda x: x.split(" ")))]
model = Doc2Vec(documents, vector_size=5, window=2, min_count=1, workers=4)
doc2vec_df = df['Review'].apply(lambda x: model.infer_vector(x.split(" "))).apply(pd.Series)
doc2vec_df.columns = ["doc2vec_vector_" + str(x) for x in doc2vec_df.columns]
df = pd.concat([df, doc2vec_df], axis=1)
df.head()
from sklearn.feature_extraction.text import TfidfVectorizer
tfidf = TfidfVectorizer(min_df = 6)
tfidf_result = tfidf.fit_transform(df['Review']).toarray()
tfidf_df = pd.DataFrame(tfidf_result, columns=tfidf.get_feature_names())
tfidf_df.columns = ["word_" + str(x) for x in tfidf_df.columns]
tfidf_df.index = df.index
df = pd.concat([df, tfidf_df], axis=1)
df.head()
from wordcloud import WordCloud
import matplotlib.pyplot as plt
def show_wordcloud(data, title = None):
wordcloud = WordCloud(background_color = 'white', max_words = 200, max_font_size = 40, scale = 3,
random_state = 42).generate(str(data))
fig = plt.figure(1, figsize=(20,20))
plt.axis('off')
if title:
fig.suptitle(title, fontsize = 20)
fig.subplots_adjust(top = 2.3)
plt.imshow(wordcloud)
plt.show()
show_wordcloud(df['Review'])
df[df["Number of Words"] >= 5].sort_values("pos", ascending = False)[["Review", "pos"]].head(10)
df[df["Number of Words"] >= 5].sort_values("neg", ascending = False)[["Review", "neg"]].head(10)
```
| github_jupyter |
```
import healpy as hp
import numpy as np
%matplotlib inline
import matplotlib.pyplot as plt
import astropy.units as u
hp.disable_warnings()
```
# Handle white noise with healpy 3 not-uniform and partial sky coverage
> Simulate white noise maps and use hitmaps
- categories: [cosmology, python, healpy]
In this third notebook, we will handle a case of not-uniform and partial sky coverage.
```
# Number based on Simons Observatory SAT UHF1 array of detectors
net = 10. * u.Unit("uK * sqrt(s)")
```
5 years with a efficiency of 20%:
```
integration_time_total = 5 * u.year * .2
```
## Download a hitmap
We can download a simulated hitmap for a Simons Observatory band, for now however, we assume a uniform coverage.
```
hitmap_url = "https://portal.nersc.gov/project/sobs/so_mapsims_data/v0.2/healpix/ST0_UHF1_01_of_20.nominal_telescope_all_time_all_hmap.fits.gz"
!wget $hitmap_url
hitmap = hp.read_map("ST0_UHF1_01_of_20.nominal_telescope_all_time_all_hmap.fits.gz")
hitmap /= hitmap.max()
hp.mollview(hitmap)
```
## Generic partial sky survey
We have now a sky coverage which not uniform
```
nside = 512
npix = hp.nside2npix(nside)
hitmap = hitmap / hitmap.sum() * integration_time_total.to(u.s)
hitmap_plot = hitmap.value.copy()
hitmap_plot[hitmap == 0] = hp.UNSEEN
hp.mollview(hitmap_plot, unit=hitmap.unit)
variance_per_pixel = \
(net**2 / hitmap).decompose()
variance_per_pixel[np.isinf(variance_per_pixel)] = 0
m = np.random.normal(scale = np.sqrt(variance_per_pixel),
size=len(variance_per_pixel)) * np.sqrt(variance_per_pixel).unit
variance_per_pixel.max()
m.value[hitmap==0] = hp.UNSEEN
m = m.to(u.uK)
m.value[hitmap==0] = hp.UNSEEN
hp.mollview(m, unit=m.unit, min=-10, max=10, title="White noise map")
```
## Power spectrum
```
sky_fraction = hp.mask_good(m).sum()/len(m)
cl = hp.anafast(m) / sky_fraction
cl[100:].mean()
pixel_area = hp.nside2pixarea(nside)
white_noise_cl = (variance_per_pixel[variance_per_pixel>0].mean() * pixel_area).to(u.uK**2)
white_noise_cl_uniform = 1.5341266e-5 * u.uK**2
plt.figure(figsize=(6,4))
plt.loglog(cl, label="Map power spectrum", alpha=.7)
plt.hlines(white_noise_cl.value, 0, len(cl), color="blue",
label="White noise level")
plt.hlines(white_noise_cl_uniform.value, 0, len(cl),
label="White noise level uniform")
plt.title("Fullsky white noise spectrum")
plt.xlabel("$\ell$")
plt.ylabel("$C_\ell [\mu K ^ 2]$");
sky_fraction
```
## Pixel weighting in the power spectrum
When we have un-uniform noise across the map, it is advantageous to weight the pixels differently before taking the power spectrum in order to downweight the noisiest pixels.
If we weight by the square root of the number of hits per pixels, then we are normalizing the standard deviation per pixel to be the same across all pixels, in fact
we recover the same noise level we had when we were spreading the hits uniformly in the sky patch.
However, the optimal is actually to weight by the inverse variance, which means to weight by the hitmap, to estimate the value expected for this we need to weight the variance by the square of the hitmap (variance is in power so weighting the map by a quantity is equivalent to weighting the variance by its square).
```
cl_apodized = hp.anafast(m * np.sqrt(hitmap)) / np.mean(hitmap)
cl_apodized_inv_variance = hp.anafast(m * hitmap) / np.mean(hitmap**2)
cl_apodized_inv_variance[100:].mean() / white_noise_cl_uniform.value
white_noise_cl / white_noise_cl_uniform
white_noise_cl
white_noise_cl_uniform
np.average(variance_per_pixel, weights=hitmap) * pixel_area * 1e12
white_noise_cl_inv_variance = np.average(variance_per_pixel, weights=hitmap**2) * pixel_area * 1e12
plt.figure(figsize=(10,6))
plt.loglog(cl, label="White noise equally weighted", alpha=.7)
plt.hlines(white_noise_cl.value, 0, len(cl), color="blue", ls=":",
label="White noise level equally weighted")
plt.loglog(cl_apodized, label="White noise inverse weighted with stddev", alpha=.7)
plt.hlines(white_noise_cl_uniform.value, 0, len(cl), color="red", ls=":",
label="White noise level for map with uniform hits")
plt.loglog(cl_apodized_inv_variance, label="White noise inverse weighted with variance", alpha=.7)
plt.hlines(white_noise_cl_inv_variance.value, 0, len(cl), color="darkgreen", ls=":",
label="White noise level inverse weighted")
plt.title("Fullsky white noise spectrum")
plt.legend()
plt.xlabel("$\ell$")
plt.ylabel("$C_\ell [\mu K ^ 2]$");
```
| github_jupyter |
```
!pip install lightgbm
!pip install joblib
import pandas as pd
import numpy as np
import lightgbm as lgb
import gc
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix,classification_report,accuracy_score
from sklearn.preprocessing import StandardScaler
import joblib
np.random.seed(2020)
def lgb_model_age(train_x, test_x,train_y, test_y):
model = lgb.LGBMClassifier (objective = 'multiclass',
learning_rate=0.06787,
num_leaves =57,
colsample_bytree = 0.9712,
reg_alpha= 0.06883,
reg_lambda= 0.09217,
subsample = 0.9,
n_estimators = 10000)
model.fit(train_x,train_y,early_stopping_rounds=100,eval_set=[(train_x,train_y),(test_x,test_y)],verbose = 10)
# 模型存储
joblib.dump(model, 'w2v_lgb_age_2_0.2.pkl')
# 模型加载
# model = joblib.load('lgb_cough.pkl')
# # 模型预测
# y_t_pred = model.predict(test_x, num_iteration=model.best_iteration_)
y_t_pred = model.predict(test_x)
# print(model.get_score(importance_type='weight'))
cm = confusion_matrix(test_y, y_t_pred)
np.set_printoptions(precision=3) # 显示精度
cm_normalized = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis] # 将样本矩阵转化为比率
print('age************************************')
print('confusion_matrix is \n {:} \n '.format(cm_normalized))
print('test acc is \n {:} \n '.format(np.sum(test_y==y_t_pred)/len(test_y)))
print(classification_report(test_y,y_t_pred))
print('accuracy is %f , sen is %f,spe is %f ' % (accuracy_score(test_y, y_t_pred) * 100,
cm_normalized[0][0],cm_normalized[1][1] ))
return accuracy_score(test_y, y_t_pred)
def lgb_model_gender(train_x, test_x,train_y, test_y):
model = lgb.LGBMClassifier (objective = 'binary',
learning_rate=0.0376,
num_leaves = 22,
colsample_bytree = 0.23,
reg_alpha= 0.096,
reg_lambda= 0.0899,
subsample = 0.99,
n_estimators = 10000)
model.fit(train_x,train_y,early_stopping_rounds=100,eval_set=[(train_x,train_y),(test_x,test_y)],verbose = 10)
# 模型存储
joblib.dump(model, 'w2v_lgb_gender_2_0.2.pkl')
y_t_pred = model.predict(test_x)
# print(model.get_score(importance_type='weight'))
cm = confusion_matrix(test_y, y_t_pred)
np.set_printoptions(precision=3) # 显示精度
cm_normalized = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis] # 将样本矩阵转化为比率
print('gender************************************')
print('confusion_matrix is \n {:} \n '.format(cm_normalized))
print('test acc is \n {:} \n '.format(np.sum(test_y==y_t_pred)/len(test_y)))
print(classification_report(test_y,y_t_pred))
print('accuracy is %f , sen is %f,spe is %f ' % (accuracy_score(test_y, y_t_pred) * 100,
cm_normalized[0][0],cm_normalized[1][1] ))
return accuracy_score(test_y, y_t_pred)
def lgb_model_gender2(train_x, test_x,train_y, test_y):
# model = lgb.LGBMRegressor(n_estimators = 10000)
# model.fit(train_x,train_y,early_stopping_rounds=100,eval_set=[(train_x,train_y),(test_x,test_y)],verbose = 10)
# 模型存储
# joblib.dump(model, 'w2v_gender_reg_2.pkl')
model = joblib.load('w2v_gender_reg_2.pkl')
y_t_pred = model.predict(test_x)
y_t_pred = [2 if i >1.5 else 1 for i in y_t_pred]
# print(model.get_score(importance_type='weight'))
cm = confusion_matrix(test_y, y_t_pred)
np.set_printoptions(precision=3) # 显示精度
cm_normalized = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis] # 将样本矩阵转化为比率
print('gender************************************')
print('confusion_matrix is \n {:} \n '.format(cm_normalized))
print('test acc is \n {:} \n '.format(np.sum(test_y==y_t_pred)/len(test_y)))
print(classification_report(test_y,y_t_pred))
print('accuracy is %f , sen is %f,spe is %f ' % (accuracy_score(test_y, y_t_pred) * 100,
cm_normalized[0][0],cm_normalized[1][1] ))
return accuracy_score(test_y, y_t_pred)
def load_data():
# user
data = pd.read_csv('../w2v_feat_data/train_data.csv')
label = data[['age','gender']]
data = data.drop(['user_id','age','gender'],axis = 1)
return data,label
data,label = load_data()
#划分age的训练和测试数据
train_x, test_x, train_y, test_y = train_test_split(data, label, test_size=0.2,random_state=2020)
# stand = StandardScaler()
# stand.fit(train_age_x)
# train_age_x, test_age_x = stand.transform(train_age_x),stand.transform(test_age_x)
# 训练模型
acc_age = lgb_model_age(train_x, test_x,train_y['age'], test_y['age'])
# 训练模型
acc_gender = lgb_model_gender(train_x, test_x,train_y['gender'], test_y['gender'])
print(acc_age+acc_gender)
# acc_gender2 = lgb_model_gender2(train_x, test_x,train_y['gender'], test_y['gender'])
```
| github_jupyter |
# **Paper Information**
**TransGAN: Two Transformers Can Make One Strong GAN, and That Can Scale Up, CVPR 2021**, Yifan Jiang, Shiyu Chang, Zhangyang Wang
* Paper Link: https://arxiv.org/pdf/2102.07074v2.pdf
* Official Implementation: https://github.com/VITA-Group/TransGAN
* Paper Presentation by Ahmet Sarıgün : https://www.youtube.com/watch?v=xwrUkHiDoiY
**Project Group Members:**
* Ahmet Sarıgün, ahmet.sarigun@metu.edu.tr
* Dursun Bekci, bekci.dursun@metu.edu.tr
## **Paper Summary**
### **Introduction**
TransGAN is a transformer-based GAN model which can be considered as a pilot study as being completely free of convolutions. The architecture of TransGAN mainly consists of a memory-friendly transformer-based generator that progressively increases feature resolution, and correspondingly a patch-level discriminator that is also transformer-based. In training of the model, a series of techniques are combined in the original paper such as data augmentation, modified normalization, and relative position encoding to overcome the general training instability issues of the GAN. We implemented data augmentation [(Dosovitskiy et al., 2020)](https://arxiv.org/pdf/2010.11929.pdf), and relative position encoding in our work. In the original paper, performance of the model tested on different datasets such as STL-10, CIFAR-10, CelebA datasets and achieved competitive results compared to current state-of-the-art GANs using convolutions. In our project, we only tested our implementation on CIFAR10 dataset as we stated in our experimental result goals.
### **TransGAN Architecture**
The architecture pipeline of TransGAN is shown below in the figure taken from the original paper.
<img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/images/transgan.jpg">
Figure 1: The pipeline of the pure transform-based generator and discriminator of TransGAN.
### **Transformer Encoder as Basic Block**
We used the transformer encoder [(Vaswani et al., 2017)](https://arxiv.org/pdf/1706.03762.pdf) as our basic block as in the original paper. An encoder is a composition of two parts. The first part is constructed by a multi-head self-attention module and the second part is a feed-forward MLP with GELU non-linearity. We apply layer normalization [(Ba et al., 2016)](https://arxiv.org/pdf/1607.06450.pdf) before both of the two parts.Both parts employ residual connection.
$$Attention(Q, K, V ) = softmax(QK^T√d_k)V$$
<img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/images/vit.gif">
Credits for illustration of ViT: [@lucidrains](https://github.com/lucidrains)
### **Memory-friendly Generator**
In building the memory-friendly generator, TransGAN utilizes a common design philosophy in CNN-based GANs which iteratively upscale the resolution at multiple stages. Figure 1 (left) illustrates the memory-friendly generator which consists of multiple stages with several transformer blocks. At each stage, feature map resolution is gradually increased until it meets the target resolution *H × W*. The generator takes the random noise input and passes it through a multiple-layer perceptron (MLP). The output vector reshaped into a $H_0 × W_0$ resolution feature map (by default $H_0$ = $W_0$ = 8), each point a C-dimensional embedding. This “feature map" is next treated as a length-64 sequence of C-dimensional tokens, combined with the learnable positional encoding.
Then, transformer encoders take embedding tokens as inputs and calculate the correspondence between each token recursively. To synthesize higher resolution images, we insert an upsampling module after each stage, consisting of a pixelshuffle [(Shi et al., 2016)](https://arxiv.org/pdf/1609.05158.pdf) module.
### **Tokenized-input for Discriminator**
The authors design the discriminator as shown in Figure 1 (right) that it takes the patches of an image as inputs. Then, they split the input images $Y$ ∈ $R^{H×W×3}$ into 8x8 patches where each patch can be regarded as a "word". The patches are then converted to the 1D sequence of token embeddings through a linear flatten layer. After that, learnable position encoding is added, and tokens pass through the transformer encoder. Finally, tokens are taken by the classification head to output the real/fake prediction.
### **Training the Model**
In this section, we show our training code and training score for CIFAR-10 Dataset with the best performance hyperparameters that we found. We trained the largest model TransGAN-XL with data augmentation using different hyperparameters, and record the results in cifar/experiments folder.
## Importing Libraries
```
from __future__ import division
from __future__ import print_function
import time
import argparse
import numpy as np
import torch
import torch.nn.functional as F
import torch.optim as optim
from torchvision.utils import make_grid, save_image
from tensorboardX import SummaryWriter
from tqdm import tqdm
from copy import deepcopy
from utils import *
from models import *
from fid_score import *
from inception_score import *
!mkdir checkpoint
!mkdir generated_imgs
!pip install tensorboardX
!mkdir fid_stat
%cd fid_stat
!wget bioinf.jku.at/research/ttur/ttur_stats/fid_stats_cifar10_train.npz
%cd ..
```
## Hyperparameters for CIFAR-10 Dataset
Since Google Colab provides limited computational power, we decreased the generated_batch_size from 64 to 32, and we also run it for 10 epochs to show our pre-computed training scores. On our local GPU machine, we train the model with generated_batch_size is 64 and run for 200 epochs.
```
# training hyperparameters given by code author
lr_gen = 0.0001 #Learning rate for generator
lr_dis = 0.0001 #Learning rate for discriminator
latent_dim = 1024 #Latent dimension
gener_batch_size = 32 #Batch size for generator
dis_batch_size = 32 #Batch size for discriminator
epoch = 10 #Number of epoch
weight_decay = 1e-3 #Weight decay
drop_rate = 0.5 #dropout
n_critic = 5 #
max_iter = 500000
img_name = "img_name"
lr_decay = True
# architecture details by authors
image_size = 32 #H,W size of image for discriminator
initial_size = 8 #Initial size for generator
patch_size = 4 #Patch size for generated image
num_classes = 1 #Number of classes for discriminator
output_dir = 'checkpoint' #saved model path
dim = 384 #Embedding dimension
optimizer = 'Adam' #Optimizer
loss = "wgangp_eps" #Loss function
phi = 1 #
beta1 = 0 #
beta2 = 0.99 #
diff_aug = "translation,cutout,color" #data augmentation
```
## Training & Saving Model for CIFAR-10
As we mentioned above we run the training for 10 epochs due to limitation of Google Colab and showed the decrease in FID score from 253 to 138 in 10 epochs.
```
if torch.cuda.is_available():
dev = "cuda:0"
else:
dev = "cpu"
device = torch.device(dev)
generator= Generator(depth1=5, depth2=4, depth3=2, initial_size=8, dim=384, heads=4, mlp_ratio=4, drop_rate=0.5)#,device = device)
generator.to(device)
discriminator = Discriminator(diff_aug = diff_aug, image_size=32, patch_size=4, input_channel=3, num_classes=1,
dim=384, depth=7, heads=4, mlp_ratio=4,
drop_rate=0.5)
discriminator.to(device)
generator.apply(inits_weight)
discriminator.apply(inits_weight)
if optimizer == 'Adam':
optim_gen = optim.Adam(filter(lambda p: p.requires_grad, generator.parameters()), lr=lr_gen, betas=(beta1, beta2))
optim_dis = optim.Adam(filter(lambda p: p.requires_grad, discriminator.parameters()),lr=lr_dis, betas=(beta1, beta2))
elif optimizer == 'SGD':
optim_gen = optim.SGD(filter(lambda p: p.requires_grad, generator.parameters()),
lr=lr_gen, momentum=0.9)
optim_dis = optim.SGD(filter(lambda p: p.requires_grad, discriminator.parameters()),
lr=lr_dis, momentum=0.9)
elif optimizer == 'RMSprop':
optim_gen = optim.RMSprop(filter(lambda p: p.requires_grad, discriminator.parameters()),
lr=lr_dis, eps=1e-08, weight_decay=weight_decay, momentum=0, centered=False)
optim_dis = optim.RMSprop(filter(lambda p: p.requires_grad, discriminator.parameters()), lr=lr_dis, eps=1e-08, weight_decay=weight_decay, momentum=0, centered=False)
gen_scheduler = LinearLrDecay(optim_gen, lr_gen, 0.0, 0, max_iter * n_critic)
dis_scheduler = LinearLrDecay(optim_dis, lr_dis, 0.0, 0, max_iter * n_critic)
#RMSprop(params, lr=0.01, alpha=0.99, eps=1e-08, weight_decay=0, momentum=0, centered=False)
print("optimizer:",optimizer)
fid_stat = 'fid_stat/fid_stats_cifar10_train.npz'
writer=SummaryWriter()
writer_dict = {'writer':writer}
writer_dict["train_global_steps"]=0
writer_dict["valid_global_steps"]=0
def compute_gradient_penalty(D, real_samples, fake_samples, phi):
"""Calculates the gradient penalty loss for WGAN GP"""
# Random weight term for interpolation between real and fake samples
alpha = torch.Tensor(np.random.random((real_samples.size(0), 1, 1, 1))).to(real_samples.get_device())
# Get random interpolation between real and fake samples
interpolates = (alpha * real_samples + ((1 - alpha) * fake_samples)).requires_grad_(True)
d_interpolates = D(interpolates)
fake = torch.ones([real_samples.shape[0], 1], requires_grad=False).to(real_samples.get_device())
# Get gradient w.r.t. interpolates
gradients = torch.autograd.grad(
outputs=d_interpolates,
inputs=interpolates,
grad_outputs=fake,
create_graph=True,
retain_graph=True,
only_inputs=True,
)[0]
gradients = gradients.contiguous().view(gradients.size(0), -1)
gradient_penalty = ((gradients.norm(2, dim=1) - phi) ** 2).mean()
return gradient_penalty
def train(noise,generator, discriminator, optim_gen, optim_dis,
epoch, writer, schedulers, img_size=32, latent_dim = latent_dim,
n_critic = n_critic,
gener_batch_size=gener_batch_size, device="cuda:0"):
writer = writer_dict['writer']
gen_step = 0
generator = generator.train()
discriminator = discriminator.train()
transform = transforms.Compose([transforms.Resize(size=(img_size, img_size)),transforms.RandomHorizontalFlip(),transforms.ToTensor(),transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
train_set = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
train_loader = torch.utils.data.DataLoader(dataset=train_set, batch_size=30, shuffle=True)
for index, (img, _) in enumerate(train_loader):
global_steps = writer_dict['train_global_steps']
real_imgs = img.type(torch.cuda.FloatTensor)
noise = torch.cuda.FloatTensor(np.random.normal(0, 1, (img.shape[0], latent_dim)))#noise(img, latent_dim)#= args.latent_dim)
optim_dis.zero_grad()
real_valid=discriminator(real_imgs)
fake_imgs = generator(noise).detach()
#assert fake_imgs.size() == real_imgs.size(), f"fake_imgs.size(): {fake_imgs.size()} real_imgs.size(): {real_imgs.size()}"
fake_valid = discriminator(fake_imgs)
if loss == 'hinge':
loss_dis = torch.mean(nn.ReLU(inplace=True)(1.0 - real_valid)).to(device) + torch.mean(nn.ReLU(inplace=True)(1 + fake_valid)).to(device)
elif loss == 'wgangp_eps':
gradient_penalty = compute_gradient_penalty(discriminator, real_imgs, fake_imgs.detach(), phi)
loss_dis = -torch.mean(real_valid) + torch.mean(fake_valid) + gradient_penalty * 10 / (phi ** 2)
loss_dis.backward()
optim_dis.step()
writer.add_scalar("loss_dis", loss_dis.item(), global_steps)
if global_steps % n_critic == 0:
optim_gen.zero_grad()
if schedulers:
gen_scheduler, dis_scheduler = schedulers
g_lr = gen_scheduler.step(global_steps)
d_lr = dis_scheduler.step(global_steps)
writer.add_scalar('LR/g_lr', g_lr, global_steps)
writer.add_scalar('LR/d_lr', d_lr, global_steps)
gener_noise = torch.cuda.FloatTensor(np.random.normal(0, 1, (gener_batch_size, latent_dim)))
generated_imgs= generator(gener_noise)
fake_valid = discriminator(generated_imgs)
gener_loss = -torch.mean(fake_valid).to(device)
gener_loss.backward()
optim_gen.step()
writer.add_scalar("gener_loss", gener_loss.item(), global_steps)
gen_step += 1
#writer_dict['train_global_steps'] = global_steps + 1
if gen_step and index % 100 == 0:
sample_imgs = generated_imgs[:25]
img_grid = make_grid(sample_imgs, nrow=5, normalize=True, scale_each=True)
save_image(sample_imgs, f'generated_images/generated_img_{epoch}_{index % len(train_loader)}.jpg', nrow=5, normalize=True, scale_each=True)
tqdm.write("[Epoch %d] [Batch %d/%d] [D loss: %f] [G loss: %f]" %
(epoch+1, index % len(train_loader), len(train_loader), loss_dis.item(), gener_loss.item()))
def validate(generator, writer_dict, fid_stat):
writer = writer_dict['writer']
global_steps = writer_dict['valid_global_steps']
generator = generator.eval()
fid_score = get_fid(fid_stat, epoch, generator, num_img=5000, val_batch_size=60*2, latent_dim=1024, writer_dict=None, cls_idx=None)
print(f"FID score: {fid_score}")
writer.add_scalar('FID_score', fid_score, global_steps)
writer_dict['valid_global_steps'] = global_steps + 1
return fid_score
best = 1e4
for epoch in range(epoch):
lr_schedulers = (gen_scheduler, dis_scheduler) if lr_decay else None
train(noise, generator, discriminator, optim_gen, optim_dis,
epoch, writer, lr_schedulers,img_size=32, latent_dim = latent_dim,
n_critic = n_critic,
gener_batch_size=gener_batch_size)
checkpoint = {'epoch':epoch, 'best_fid':best}
checkpoint['generator_state_dict'] = generator.state_dict()
checkpoint['discriminator_state_dict'] = discriminator.state_dict()
score = validate(generator, writer_dict, fid_stat)
print(f'FID score: {score} - best ID score: {best} || @ epoch {epoch+1}.')
if epoch == 0 or epoch > 30:
if score < best:
save_checkpoint(checkpoint, is_best=(score<best), output_dir=output_dir)
print("Saved Latest Model!")
best = score
checkpoint = {'epoch':epoch, 'best_fid':best}
checkpoint['generator_state_dict'] = generator.state_dict()
checkpoint['discriminator_state_dict'] = discriminator.state_dict()
score = validate(generator, writer_dict, fid_stat) ####CHECK AGAIN
save_checkpoint(checkpoint,is_best=(score<best), output_dir=output_dir)
```
### **Experimental Result Goals vs. Achieved Results**
In this project, we aimed to reproduce qualitative results(generating image samples by CIFAR-10 Dataset) and quantitative results in Table 2 and Table 4 of the original paper that shown below.
<table>
<tr>
<td> <img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/results/table2.png" style="width: 400px;"/> </td>
<td> <img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/results/table4.png" style="width: 400px;"/> </td>
</tr></table>
Since we have limited computational resource and time for the training all size of TransGAN model on CIFAR-10 Dataset, we only trained the largest model with data augmentation, TransGAN-XL, for Table 4 results.
## Test Model and Results
In this section, we loaded pre-trained model and got the following qualitative and quantitative results.
### Qualitative Results
The following pictures show our generated images at different epoch numbers.
<table>
<tr>
<td style="text-align: center">0 Epoch</td>
<td style="text-align: center">40 Epoch</td>
<td style="text-align: center">100 Epoch</td>
<td style="text-align: center">200 Epoch</td>
</tr>
<trt>
<p align="center"><img width="30%" src="https://raw.githubusercontent.com/asarigun/TransGAN/main/images/atransgan_cifar.gif"></p>
</tr>
<tr>
<td> <img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/results/0.jpg" style="width: 400px;"/> </td>
<td> <img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/results/40.jpg" style="width: 400px;"/> </td>
<td> <img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/results/100.jpg" style="width: 400px;"/> </td>
<td> <img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/results/200.jpg" style="width: 400px;"/> </td>
</tr>
</table>
### Quantitative Results
As we mentioned above, due to the lack of computational resource, we did our experiments only with the largest model TransGAN-XL and get the following results. We had decided not to implement 'Co-Training with Self-Supervised Auxiliary Task' and 'Locality-Aware Initialization for Self-Attention' since they made only small differences as shown in the paper. The difference between our result and original paper result can be originated in using some different hyperparameters and abovementioned implementation differences. You can see our quantitative result, FID score 26.82, [here](https://github.com/asarigun/TransGAN/blob/main/results/wgangp_eps_optim_Adam_lr_gen_0_0001_lr_dis_0_0001_epoch_200.txt).
```
# downloading pretrained model
%cd checkpoint
!wget https://drive.google.com/file/d/134GJRMxXFEaZA0dF-aPpDS84YjjeXPdE/view?usp=sharing
%cd ..
# Loading Pretrained Model
from __future__ import division
from __future__ import print_function
import time
import argparse
import numpy as np
import torch
import torch.nn.functional as F
import torch.optim as optim
from torchvision.utils import make_grid, save_image
from tensorboardX import SummaryWriter
from tqdm import tqdm
from copy import deepcopy
from utils import *
from models import *
from fid_score import *
from inception_score import *
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
torch.backends.cudnn.enabled = True
torch.backends.cudnn.benchmark = True
def validate(generator, writer_dict, fid_stat, clean_dir=True):
epoch = 200
writer = writer_dict['writer']
generator.eval().cuda()
generator = torch.nn.DataParallel(generator.to("cuda:0"))
with torch.no_grad():
eval_iter = 20000 // 8
img_list = list()
for iter_idx in tqdm(range(eval_iter), desc='sample images'):
z = torch.cuda.FloatTensor(np.random.normal(0, 1, (64, 1024)))
gen_imgs = generator(z).to('cuda:0')
img_list.extend(list(gen_imgs))
fid_score = get_fid(fid_stat, epoch, generator, num_img=5000, val_batch_size=30*2, latent_dim=1024, writer_dict=None, cls_idx=None)
print(f"FID score: {fid_score}")
def main():
device = torch.device("cuda:0")
load_path = '/content/drive/TransGAN (1)/checkpoint /checkpoint.pth'
generator= Generator(depth1=5, depth2=4, depth3=2, initial_size=8, dim=384, heads=4, mlp_ratio=4, drop_rate=0.5)
generator.to(device)
fid_stat = 'fid_stat/fid_stats_cifar10_train.npz'
assert os.path.exists(fid_stat)
state_dict = torch.load(load_path)
from collections import OrderedDict
new_state_dict = OrderedDict()
for k, v in state_dict.items():
name = k[7:]
new_state_dict[name] = v
checkpoint_file = load_path
os.path.exists(checkpoint_file)
checkpoint = torch.load(checkpoint_file)
generator.load_state_dict(checkpoint, strict=False)
writer=SummaryWriter()
writer_dict = {'writer':writer}
fid_score = validate(generator, writer_dict, fid_stat, clean_dir=True)
main()
```
## Challenges and Discussions
Since the authors did not give detailed hyperparameters for each Transformers Block and Multi-Head Attention Mechanism on version 1, we needed to find the best hyperparameters. Also, in the training part, they did not give detailed hyperparameters such as droprate, weight decay, or batch normalization in version 1. But in the last version of the original paper, authors gave more detailed hyperparameters for training, therefore we got more reasonable results.
During the implementation, first we used Hinge loss and faced convergence problem in training. When we tried another loss function, WGAN-GP, that is mentioned in the last version of original paper, we achieved to overcome convergence problem and got better results.
As authors didn't share detailed training process in their previous version, we struggled to converge FID score during training. But in the latest version of the original paper, authors provided more details for training so that we achieved to converge FID score in the training.
Due to lack of computational resource, we only trained the largest model, TransGAN-XL in our project. We implemented data augmentation in our model as it is considered crucial for TransGAN in the original paper. We didn't implement 'Co-Training with Self-Supervised Auxiliary Task' and 'Locality-Aware Initialization for Self-Attention' since they make only small differences as shown in the paper.
## Citation
```
@article{jiang2021transgan,
title={TransGAN: Two Transformers Can Make One Strong GAN},
author={Jiang, Yifan and Chang, Shiyu and Wang, Zhangyang},
journal={arXiv preprint arXiv:2102.07074},
year={2021}
}
```
```
@article{dosovitskiy2020,
title={An Image is Worth 16x16 Words: Transformers for Image Recognition at Scale},
author={Dosovitskiy, Alexey and Beyer, Lucas and Kolesnikov, Alexander and Weissenborn, Dirk and Zhai, Xiaohua and Unterthiner, Thomas and Dehghani, Mostafa and Minderer, Matthias and Heigold, Georg and Gelly, Sylvain and Uszkoreit, Jakob and Houlsby, Neil},
journal={arXiv preprint arXiv:2010.11929},
year={2020}
}
```
```
@inproceedings{zhao2020diffaugment,
title={Differentiable Augmentation for Data-Efficient GAN Training},
author={Zhao, Shengyu and Liu, Zhijian and Lin, Ji and Zhu, Jun-Yan and Han, Song},
booktitle={Conference on Neural Information Processing Systems (NeurIPS)},
year={2020}
}
```
| github_jupyter |
# The Jupyter Notebook
and some python basics
This is a jupyter notebook, one of the environments in which you can run Python code.
It is comprised of "cells" that can be executed.
The cells can be one of two types, markdown or code. Markdown cells are just for static text and equations. Code cells execute python code. You can change the type with the combo box above or using keyboard shortcuts
Keyboard shortcuts are your friends. Learn the various shortcuts on your system, especially the "run cell" shortcut, usually ctrl-enter. alt-enter runs the cell and makes a new one after it!
When having trouble, tinker, and ask google (often solutions are on stack overflow)
```
1
'b'
type(1.0)
type("b")
int(0.232)
int(232.9)
s="eggs"
print(s)
print(type(s))
s[0]
s[0:4] # this is a comment
# this actually giving elements 0-3 of the string
```
**Let's step back: Where did python's name come from?**
a. A programmer who loved a snake?
b. Monte Python
c. Someone's favorite neck tatoo
When he began implementing Python, Guido van Rossum was also reading the published scripts from “Monty Python’s Flying Circus”, a BBC comedy series from the 1970s. Van Rossum thought he needed a name that was short, unique, and slightly mysterious, so he decided to call the language Python.
**Python was meant to be "fun":**
[Zen of Python and the guiding principles](https://en.wikipedia.org/wiki/Zen_of_Python)
Beautiful is better than ugly.
Explicit is better than implicit.
Simple is better than complex.
Complex is better than complicated.
Flat is better than nested.
Sparse is better than dense.
Readability counts.
Special cases aren't special enough to break the rules.
Although practicality beats purity.
Errors should never pass silently.
Unless explicitly silenced.
In the face of ambiguity, refuse the temptation to guess.
There should be one—and preferably only one—obvious way to do it.
Although that way may not be obvious at first unless you're Dutch.
Now is better than never.
Although never is often better than right now.
If the implementation is hard to explain, it's a bad idea.
If the implementation is easy to explain, it may be a good idea.
Namespaces are one honking great idea—let's do more of those!
# Math
**Operators**
Operators for integers:
`+ - * / // % **`
Operators for floats:
`+ - * / **`
Boolean expressions:
* keywords: `True` and `False` (note capitalization)
* `==` equals: `5 == 5` yields `True`
* `!=` does not equal: `5 != 5` yields `False`
* `>` greater than: `5 > 4` yields `True`
* `>=` greater than or equal: `5 >= 5` yields `True`
* Similarly, we have `<` and `<=`.
Logical operators:
* `and`, `or`, and `not`
* `True and False`
* `True or False`
* `not True`
```
sin(pi/2)
import math
math.sin(math.pi/2)
from math import sin
sin(math.pi/2)
10/5
10%5
print(10//5)
print(10%5)
print(10//4)
print(11%4)
```
# Basic Variables Overiew
```
t=55.0
print(f'{t} is a {type(t)}')
strings='hi mom'
type(strings)
t1=44.5
type(t1)
b1=True
type(b1)
```
# **Strings**
```
str1='test'
str2="case"
print(str1)
print(str2)
str3 = str1 + str2 + "!"
print(str3)
x=20
y='40'
print("twice {x} is {y}")
print(f"twice {x} is {y}")
print(f"twice {x} is {2*x}")
print(int(y))
print(type(float(y)))
str1="Hello, World!"
print(str1)
print(str1.upper())
print(str1.lower())
print(str1[0])
print(str1[len(str1)-1])
print(len(str1))
str1.replace('l','p')
str1.split()
```
# **Lists, Tuples, Sets, and Dict**
```
list1=[4, 5, 'hi mom', 6.34]
print(list1)
list1[0]
print('lenght of this list is', len(list1))
for x in list1:
print(x)
for x in range(len(list1)):
print(x)
print(list1[x])
list1.remove(4)
print(list1)
b=[]
for i in range(5):
b.append(i**2)
print(b)
x=['a', ['bb', ['ccc', 'ddd'], 'ee', 'ff'], 'g', ['hh', 'ii'], 'j']
type(x)
print(len(x))
print(x[0])
print(x[1])
print(x[1][1])
print(x[1][1][0])
tpl=(1, 4, 8)
print(tpl)
print(type(tpl))
print(tpl[1])
tpl[1]=5
st={0, 4, 6, 4}
print(type(st))
print(st)
print(st[0])
dct={}
dct[5]=55
dct['somekey']=999
dct['anotherkey']='hello'
print(dct)
print(dct['somekey'])
print(type(dct))
```
# **Looping and Flow Control**
```
xx=5
yy=True
if xx==5:
print('xx is 5')
elif xx==yy:
print('xx is equal to yy')
else:
print("nada")
xx=5 # comments
yy=True
# comment line
if xx!=5:
print('xx is not 5')
elif xx==yy:
print('xx is yy')
else:
print('nada')
print(xx != 5)
print(range(10))
for x in range(10):
print(x)
x=66
while x<100:
print(x)
x+=0.1*x # same as writing x = x + 0.1*x
n=64
print(range(2,n))
for num in range(2,n):
if num%2==0:
continue # sends you to the loop beginning
print(f'{num} is an odd number')
n=64
for x in range(2,n):
if n%x == 0:
print(f'{n} equals {x} * {n // x}')
if True==False:
pass # pass exits the loop, whereas continue goes back to beginning of loop
else:
print('true does not equal false')
```
# **Functions**
```
def rect_area(length, width):
return length*width
rect_area(5, 6)
def rect_area(length, width):
if length<=0 or width<=0:
raise ValueError("Length and Width must be postive and non-zero")
return length*width
print(rect_area(4,4))
print(rect_area(-3,4))
def f(x):
return x*x
def g(f,x):
return(f(f(x)))
g(f,5)
def g(a, x, b=0):
return a * x + b
print(g(2,5,1))
print(g(2,5,0))
print(g(2,5))
```
# **Exceptions**
```
5/0
try:
5/0
except ZeroDivisionError:
print("dont divide by zero buddy")
```
# **Classes**
```
class Materials:
def somefunc(self):
print("hello")
t=Materials()
t.somefunc()
# Class that holds fractions r = p / q
class Rational:
def __init__(self, p, q=1):
if q == 0:
raise ValueError('Denominator must not be zero')
if not isinstance(p, int):
raise ValueError('Numerator must be an integer')
if not isinstance(q, int):
raise ValueError('Denominator must be an integer')
g = math.gcd(p, q)
self.p = p // g # integer division
self.q = q // g
# method to convert rational to float
def __float__(self):
return self.p / self.q
# method to convert rational to string for printing
def __str__(self):
return f'{self.p}/{self.q}'
def __repr__(self):
return f'Rational({self.p}, {self.q})'
import math
a=Rational(4,6)
print(f'a = {a}')
print(f"float(a) = {float(a)}")
print(f"str(a) = {str(a)}")
print(f"repr(a) = {repr(a)}")
```
| github_jupyter |
# Layout
---------------------------------------
In this section, you can learn how to apply layout that you want to get to your network.
In cytoscape, layout networks is in two dimensions. First one is to apply layout algorithms. A variety of layout algorithms are available, including cyclic, tree, force-directed, edge-weight, and yFiles Organic layouts. Second, you can also use Manual Layout tools similar to other graphics application user interface.
In the following workflow, you can see how to apply layout algorithms programatically and save it.
## Table of contents
- Get list of available layout
- Apply layout
- Save Layout
# Network Data Preparation
---------------------------------------
To execute this examples, first, we have to import sample data.
```
# import library
library(RCy3)
library(igraph)
# first, delete existing windows to save memory:
deleteAllWindows(CytoscapeConnection())
# Load Data
gal.table <- read.table('../sampleData/galFiltered.sif',stringsAsFactors=FALSE)
# create graph class
g <- new ('graphNEL', edgemode='directed')
# Get NodesVec
gal.table.nodevec <- unique(c(gal.table[[1]], gal.table[[3]]))
# add nodes to graph
for(node in gal.table.nodevec){
g <- graph::addNode(node, g)
}
# get EdgeList
gal.table.fromvec = gal.table[[1]]
gal.table.tovec = gal.table[[3]]
for (index in 1:length(gal.table.fromvec)){
g <- graph::addEdge (gal.table.fromvec[[index]] ,gal.table.tovec[[index]], g)
}
# show it in cytescape
cw <- CytoscapeWindow('vignette', , graph=g, overwrite=TRUE)
displayGraph (cw)
layoutNetwork (cw, layout.name='degree-circle')
```
# Get list of available layout
---------------------------------------
First, let's begin to get the list of available layout. We can get it by using the following method.
### Method : getLayoutNames
We use two method in this section. 'getLayoutNames' let us know available layoutNames and we can apply one of them by 'layoutNetwork' method.
#### Usage
getLayoutNames(obj)
#### Arguments
- obj : a CytoscapeConnectionClass object.
```
# get available layout names
getLayoutNames(cw)
```
# Apply layout
---------------------------------------
Now, you get the available layout methods. By following method, you can apply method that you want.
### Method : layoutNetwork
#### Usage
layoutNetwork(obj, layout.name= grid )
#### Arguments
- obj : a CytoscapeWindowClass object.
- layout.name : a string, one of the values returned by getLayoutNames, ’grid’ by default.
```
# execute layout that you want
layoutNetwork(cw, layout.name= 'force-directed')
# This code is for save and show the network image.
file.name <- paste (getwd (), 'resultImage' ,'saveImageToShowLayoutExample' , sep= '/' )
image.type <- 'png'
resource.uri <- paste(cw@uri,
pluginVersion(cw), "networks", as.character(cw@window.id),
paste0("views/first.", image.type),
sep="/")
request.res <- GET(resource.uri, write_disk(paste0(file.name,".", image.type), overwrite = TRUE))
```

# Save Layout
---------------------------------------
When you use the below method, you can save the current layout (that is, node positions) to the specified file.
### Usage
saveLayout(obj, filename, timestamp.in.filename=FALSE)
### Arguments
- obj : a CytoscapeWindowClass object.
- filename : a string.
- timestamp.in.filename : logical
```
# TODO : I don't know why this method is not available. So I have to find it.
saveLayout (cw, layout2 , timestamp.in.filename=TRUE)
```
| github_jupyter |
# Exercise 6 - Convolutional Autoencoder
In this exercise we will construct a convolutional autoencoder for the sample of the CIFAR-10 dataset.
Import pickle, numpy, matplotlib as well as the *Model* class from **keras.models** and *Input* and *Conv2D*, *MaxPooling2D* and *UpSampling2D* from **keras.layers**.
```
import pickle
import numpy as np
import matplotlib.pyplot as plt
from keras.models import Model
from keras.layers import Input, Conv2D, MaxPooling2D, UpSampling2D
```
Load the data
```
with open('data_batch_1', 'rb') as f:
dat = pickle.load(f, encoding='bytes')
```
As this is an unsupervised learning method, we are only interested in the image data. Load the image data as per the previous exercise.
```
images = np.zeros((10000, 32, 32, 3), dtype='uint8')
for idx, img in enumerate(dat[b'data']):
images[idx, :, :, 0] = img[:1024].reshape((32, 32)) # Red
images[idx, :, :, 1] = img[1024:2048].reshape((32, 32)) # Green
images[idx, :, :, 2] = img[2048:].reshape((32, 32)) # Blue
```
As we are using a convolutional network we can use the images with only rescaling.
```
images = images / 255.
```
Define the convolutional autoencoder model. We will use the same shape input as an image.
```
input_layer = Input(shape=(32, 32, 3,))
```
Add a convolutional stage, with 32 layers or filters, a 3 x 3 weight matrix, a ReLU activation function and using **same** padding which means the output has the same length as the input image.
```
hidden_encoding = Conv2D(
32, # Number of layers or filters in the weight matrix
(3, 3), # Shape of the weight matrix
activation='relu',
padding='same', # How to apply the weights to the images
)(input_layer)
```
Add a max pooling layer to the encoder with a 2 x 2 kernel. Max Pooling looks at all the values in an image, scanning through with a 2 x 2 matrix. The maximum value in each 2 x 2 area is returned, thus reducing the size of the encoded layer by a half.
```
encoded = MaxPooling2D((2, 2))(hidden_encoding)
```
Add a decoding convolutional layer
```
hidden_decoding = Conv2D(
32, # Number of layers or filters in the weight matrix
(3, 3), # Shape of the weight matrix
activation='relu',
padding='same', # How to apply the weights to the images
)(encoded)
```
Now we need to return the image to its original size, for this we will upsample by the same size as Max Pooling.
```
upsample_decoding = UpSampling2D((2, 2))(hidden_decoding)
```
Add the final convolutional stage, using 3 layers for the RGB channels of the images.
```
decoded = Conv2D(
3, # Number of layers or filters in the weight matrix
(3, 3), # Shape of the weight matrix
activation='sigmoid',
padding='same', # How to apply the weights to the images
)(upsample_decoding)
```
Construct the model by passing the first and last layers of the network to the Model class.
```
autoencoder = Model(input_layer, decoded)
```
Display the structure of the model
```
autoencoder.summary()
```
Compile the autoencoder using a binary cross entropy loss function and adadelta gradient descent.
```
autoencoder.compile(loss='binary_crossentropy',
optimizer='adadelta')
```
Now let's fit the model, again we pass the images as the training data and as the desired output. Train for 20 epochs as convolutional networks take a lot longer to compute.
```
autoencoder.fit(images, images, epochs=20)
```
Calculate and store the output of the encoding stage for the first 5 samples.
```
encoder_output = Model(input_layer, encoded).predict(images[:5])
```
Each encoded image has a shape of 16 x 16 x 32 due to the number of filters selected for the convolutional stage. As such we cannot visualise them without modification. We will reshape them to be 256 x 32 in size for visualisation.
```
encoder_output = encoder_output.reshape((-1, 256, 32))
```
Get the output of the decoder for the 5 images
```
decoder_output = autoencoder.predict(images[:5])
```
Plot the original image, the mean encoder output and the decoder.
```
plt.figure(figsize=(10, 7))
for i in range(5):
plt.subplot(3, 5, i + 1)
plt.imshow(images[i], cmap='gray')
plt.axis('off')
plt.subplot(3, 5, i + 6)
plt.imshow(encoder_output[i], cmap='gray')
plt.axis('off')
plt.subplot(3, 5, i + 11)
plt.imshow(decoder_output[i])
plt.axis('off')
```
| github_jupyter |
# Дифференцируемое програмирование
**Разработчик: Александр Шевченко**
На этом семинаре будет реализовываться система распознавания рукописных слов, основанная на совмещении алгоритмов предсказания (динамическое программирование) и глубинного обучения. Мы будем использовать датасет Stanford OCR (http://ai.stanford.edu/~btaskar/ocr/), состоящий из слов на английском языке и изображений рукописных букв.
Для начала загрузим и подготовим данные. Для распаковки необходим gunzip. Пользователям Windows нужно скачать и распаковать датасет вручную.
```
!rm -rf letter.data
!wget http://ai.stanford.edu/~btaskar/ocr/letter.data.gz
!gunzip letter.data.gz
from utils import prepare_data
train_x, train_y, test_x, test_y, val_x, val_y, le = prepare_data()
```
Каждый элемент датасета содержит данные об одном слове. Списки $*\_x[i]$ содержат numpy массивы размера [word_len, 1, 32, 32], содержащие изображения рукописных букв. Списки $*\_y[i]$ содержат numpy массивы размера [word_len] с метками для каждого изображения.
```
import matplotlib.pyplot as plt
import seaborn as sb
import numpy as np
%matplotlib inline
```
Изображения выглядят следующим образом. Метки классов уже сконвертированы в числа для использования.
Обратите внимание, что в нашем датасете первые буквы в каждом слове обрезаны и не используются (это не баг, а сделано специально, потому что первая буква часто бывает заглавной и, соответственно, её вариабельность сильно выше).
```
sb.set()
fig, ax = plt.subplots(1, train_x[0].shape[0], figsize=(15,15))
ax = np.array(ax)
word = ''.join(le.inverse_transform(train_y[0]))
for idx in range(train_x[0].shape[0]):
ax[idx].set_title(word[idx])
ax[idx].axis('off')
ax[idx].imshow(train_x[0][idx,0,:,:])
plt.tight_layout()
```
### Score функция и правдоподобие
Мы будем использовать модель вида цепочка (то есть нас будут интересовать только связи между соседними буквами) со следующей score функцией:
$$
F(Y| X, \Theta) = \sum\limits_{i=0}^{L-1} U(x_i, y_i) + \sum\limits_{i=0}^{L-2} W(y_{i}, y_{i+1})
$$
$\Theta$ содержит параметры унарных $U$ и парных $W$ потенциалов. На этом семинаре для унарных потенциалов мы будем использовать простую нейросеть для классификации изображения (как для MNIST), а парные потенциалы параметризуем при помощи матрицы размера $26 \times 26$ (обратите внимание, что парные потенциалы зависят только от меток, т.е., не зависят непосредственно от изображений).
$U$ унарные потенциалы отвечают за совместимость метки $y_i$ и входного изображения буквы $x_i$. Парные потенциалы показывают насколько вероятно сочетание букв $(y_i, y_{i+1})$.
Используя score функцию $F$, мы можем задать распределение вероятностей над всеми возможными разметками последовательности $X$ (это распределение связано с графической моделью Conditional Random Field, CRF):
$$
P(Y| X,\Theta) = \frac{1}{Z(\Theta)} \exp\{F(Y| X, \Theta)\}
$$
### Предсказание (0.3 балла)
Для фиксированных значений параметров $\Theta$ предсказание может быть сделано, например, при помощи максимизации score функции $F$ (соответствует моде распределения над разметками). Для функций, связи между переменными в которых образуют граф вида цепочка (возможно для любого дерева), задача максимизации может быть решена точно за полиномиальное время при помощи метода динамического программирования.
Выведем конкретный алгоритм, используя подход динамического программирования для решения задачи максимизации score функции:
$$
\max_{Y} F(Y|X,\Theta) = \max_{Y} \left[ \sum\limits_{i=0}^{L-1} U(x_i, y_i) + \sum\limits_{i=0}^{L-2} W(y_{i}, y_{i+1}) \right]
$$
Выполняя алгебраические преобразования, задачу можно переписать следующим образом:
$$
\max_{Y} \left[ \sum\limits_{i=0}^{L-1} U(x_i, y_i) + \sum\limits_{i=0}^{L-2} W(y_{i}, y_{i+1}) \right] = \max_{y_0} \left[U(x_0, y_0) + \max_{y_1,...,y_{L-1}}\left( W(y_0, y_1) + \sum\limits_{i=1}^{L-1} U(x_i, y_i) + \sum\limits_{i=1}^{L-2} W(y_{i}, y_{i+1}) \right) \right]
$$
В качестве подзадач динамического программирования будем использовать внутренние максимумы. Обозначим через $V_j(y_j)$ такой максимум по переменным с индексами большими чем $j$, т.е.,
$$
V_j(y_j) = U(x_j, y_j) + \max_{y_{j+1},...,y_{L-1}}\left( W(y_j, y_{j+1}) + \sum\limits_{i=j+1}^{L-1} U(x_i, y_i) + \sum\limits_{i=j+1}^{L-2} W(y_{i}, y_{i+1}) \right).
$$
Динамическое программирование основано на интеративном вычислении $V_j(y_j)$ на основе ранее вычисленных значений. Используется следующая формула пересчёта:
$$
V_j(y_j) = U(x_j, y_j) + \max_{y_{j+1}}\left[ W(y_j, y_{j+1}) + V_{j+1}(y_{j+1}) \right].
$$
Инициализировать пересчёт можно так: $V_{L-1}(y_{L-1}) = U(x_{L-1}, y_{L-1})$. Значение score на наилучшей конфигурации (решение задачи) можно найти при помощи максимизации $\max_{y_0} [V_0(y_0)]$.
Используя сохраненные индексы максимумов в каждой из задач максимизации, можно сделать проход в обратном направлении и восстановить оптимальную разметку.
```
import torch
import torch.nn as nn
from torch.autograd import Variable
NUM_LABELS = 26
def dynamic_programming(U, W):
"""
Parameters:
U: unary potentials, torch tensor shape (len_word, NUM_LABELS)
W: pairwise potentials, torch tensor shape (NUM_LABELS, NUM_LABELS)
Returns:
arg_classes: argmaximum, torch long tensor shape (len_word)
"""
L = U.size(0)
V, argmax = torch.zeros(L, NUM_LABELS),\
torch.zeros(L, NUM_LABELS)
### code starts here ###
### code ends here ###
return arg_classes.long()
```
Если все реализовано верно, вы должны получить вывод: "nconsequential"
```
U = torch.load('unary_example.pth')
W = torch.load('pairwise_example.pth')
pred = dynamic_programming(U, W)
print(''.join(le.inverse_transform(int(i)) for i in pred))
```
Обратите внимание, что если делать предсказание только по унарным потенциалам, то алгоритм делает ошибки.
```
_, u_labels = U.max(1)
print(''.join(le.inverse_transform(int(i)) for i in u_labels))
```
## Настройка параметров $\Theta$ при помощи структурного метода опорных векторов (0.3 балла)
Для настройки параметров $\Theta$ будем использовать структурный метод опорных векторов (Structured SVM, SSVM). Интуитивно оптимизация данного функционала позволит обеспечить высокий score на правильных разметках и низкий score на неправильных. Функция потерь SSVM на одном объекте выборки $X$, $Y$ выглядит так:
$$
\max_{Y'} \left[\Delta(Y,Y') + F(Y',X,\Theta)\right] - F(Y,X,\Theta).
$$
Здесь $\Delta(Y,Y')$ - это функция, обобщающая отстут (margin) из классического SVM. Мы будет к качестве $\Delta$ использовать нормированное расстояние Хэмминга между последовательностями $Y$ и $Y'$, т.е. $\Delta(Y,Y') = \frac{1}{L}\sum\limits_{i=1}^{L} [y_i \neq y_i']$.
Задача максимизации, возникающая в рамках функции потерь, может быть решена при помощи уже реализованного алгоритма динамического программирования (добавление функции $\Delta$ в данном случае не усложняет задачу поскольку не менят структуру графа).
Для добавления $\Delta$ в score фунцию достаточно добавить $\frac{1}{L}$ ко всем унарным потенциалам, соответствующим неправильным меткам.
Процедура обучения (настройки параметров $\Theta$) состоит в минимизации функции потерь (усредненной по обучающей выборке) по $\Theta$ при помощи методов стохастической оптимизации. При обработке каждого элемента выборки нужно решать задачу максимизации $F+\Delta$. После нахождения оптимальной конфигурации (argmax) достаточно подставить полученные $Y'$ и вести оптимизацию по $\Theta$ опуская слагаемое отвечающее $\Delta$. На лекции методы этой группы назывались "структурным пулингом".
Сначала нужно реализовать решение задачи максимизации из функции потерь SSVM (loss-augmented inference) при помощи вызова ранее реализованного алгоритма динамического программирования. Для тестирования кода добавьте возможность умножения расстояния Хэмминга на вес weight.
```
def loss_aug_inference(U, W, target, weight=1.0):
"""
Parameters:
U: unary potentials, torch tensor shape (len_word, NUM_LABELS)
W: pairwise potentials, torch tensor shape (NUM_LABELS, NUM_LABELS)
target: true configuration, torch long tensor shape (len_word)
weight: (for debug) put more weight on the loss term
Returns:
arg_classes: argmaximum, torch long tensor shape (len_word)
"""
### code starts here ###
### code ends here ###
return arg_classes.long()
U = torch.load('unary_example.pth')
W = torch.load('pairwise_example.pth')
target = torch.LongTensor([13, 2, 14, 13, 18, 4, 16, 20, 4, 13, 19, 8, 0, 11])
pred = loss_aug_inference(U, W, target, weight=60.0)
correct = torch.LongTensor([13,2, 14, 13, 18, 5, 14, 21, 4, 13, 19, 8, 13, 2])
assert pred.eq(correct).sum() == correct.numel(), "Check your loss_aug_inference"
```
### Унарные потенциалы
Для извлечения унарных потенциалов будем использовать нейросеть вида LeNet. Для экономии времени семинара сеть обучена заранее (стандартная схема для MNIST, но с 26 классами).
Стоит так же заметить, что unary network без использования парных потенциалов достигает качества на валидации 0.92 (точность предсказания всех символов датасета).
```
import torch.nn.functional as F
class LeNet(nn.Module):
def __init__(self):
super(LeNet, self).__init__()
self.conv1 = nn.Conv2d(1, 10, 5)
self.conv2 = nn.Conv2d(10, 20, 5)
self.fc1 = nn.Linear(5 * 5 * 20, 140)
self.fc2 = nn.Linear(140, 26)
def forward(self, x):
x = F.max_pool2d(F.relu(self.conv1(x)), 2)
x = F.max_pool2d(F.relu(self.conv2(x)), 2)
x = x.view(-1, 5 * 5 * 20)
x = F.relu(self.fc1(x))
x = self.fc2(x)
return x
unary_net = LeNet()
unary_net.load_state_dict(torch.load('state_dict.pth'))
```
### Обучение
Для вычисления функции потерь SSVM необходимо реализовать подсчет score функции.
```
def score_function(Y, U, W):
"""
Parameters:
U: unary potentials, torch Variable shape (len_word, NUM_LABELS)
W: pairwise potentials, torch Variable shape (NUM_LABELS, NUM_LABELS)
Y: configuration, torch long tensor shape (len_word)
Returns:
value of score function
"""
### code starts here ###
### code ends here ###
return score_value
U = torch.load('unary_example.pth')
W = torch.load('pairwise_example.pth')
target = torch.LongTensor([13, 2, 14, 13, 18, 4, 16, 20, 4, 13, 19, 8, 0, 11])
s = score_function(target, U, W)
assert np.allclose(score_function(target, U, W), 175.58605), 'Check you score function'
```
Теперь нужно реализовать подсчёт функции потерь SSVM и вызов оптимизатора.
```
from torch.optim import Adam
from tqdm import trange
from IPython.display import clear_output
trace_values = []
torch.manual_seed(42)
W = Variable(torch.randn(NUM_LABELS, NUM_LABELS), requires_grad=True)
opt = Adam([W], lr=1e-2)
n_epoch = 5
for epoch in range(n_epoch):
mean_val = 0.
for i in trange(len(train_x)):
word, target = Variable(torch.from_numpy(train_x[i]).float()),\
Variable(torch.from_numpy(train_y[i]).long())
U = unary_net(word)
y_ = loss_aug_inference(U.data, W.data, target.data)
### code starts here ###
### code ends here ###
mean_val += loss.data[0] + 1. - y_.eq(target.data).sum() / U.size(0)
if i % 500 == 0 and i:
trace_values.append(mean_val / 500.)
mean_val = 0.
clear_output()
plt.title('SSVM loss')
plt.plot(np.arange(len(trace_values)), trace_values)
plt.show()
glob_acc = 0.
letters_num = np.sum([i.shape[0] for i in val_x])
for i in range(len(val_x)):
word, target = Variable(torch.from_numpy(val_x[i]).float()),\
torch.from_numpy(val_y[i]).long()
U, P = unary_net(word), W
pred = dynamic_programming(U.data, P.data)
eq_count = pred.eq(target).sum()
glob_acc += eq_count
glob_acc /= letters_num
print('global val accuracy {}'.format(glob_acc))
```
Точность на валидации должна получиться близкой к 0.965.
## Настройка параметров $\Theta$ при помощи метода максимального правдоподобия
Другим подходом к настройке параметров $\Theta$ является метод максимального правдоподобия.
Метод состоит в максимизации лог-правдоподобия на обучающей выборке. Правдоподобие задаётся следущим распределением вероятностей:
$$
P(Y| X,\Theta) = \frac{1}{Z(\Theta)} \exp\{F(Y| X, \Theta)\}
$$
### Вычисление нормировочной константы $Z$ (0.2 балла)
Для модели цепочки нормировочная константа может быть посчитана эффективно с использованием sum-product belief propagation (динамическое программирование)
$$
Z(\Theta) = \sum\limits_{Y'} \exp\{F(Y'| X, \Theta)\} = \sum\limits_{Y'} \exp\left\{\sum\limits_{i=0}^{L-1} U(x_i, y_i') + \sum\limits_{i=0}^{L-2} W(y_{i}', y_{i+1}'))\right\}
$$
$$
= \sum\limits_{y_0'} \exp\{U(x_0, y_0')\}\sum\limits_{y_1',...,y_{L-1}'}\exp\left\{W(y_{0}', y_{1}') + \sum\limits_{i=1}^{L-1} U(x_i, y_i') + \sum\limits_{i=2}^{L-1} W(y_{i}', y_{i+1}'))\right\}
$$
Определеним подзадачи динамического программирования (аналогично предсказанию, но сумма заменена на произведение, а максимум на сумму).
$$
V_j(y_j) = \exp\{U(x_j, y_j)\} \sum_{y_{j+1},...,y_{L-1}}\left( \exp\{W(y_j, y_{j+1})\} \prod\limits_{i=j+1}^{L-1} \exp\{U(x_i, y_i)\} \prod\limits_{i=j+1}^{L-2} \exp\{W(y_{i}, y_{i+1})\} \right).
$$
Динамическое программирование основано на интеративном вычислении $V_j(y_j)$ на основе ранее вычисленных значений. Используется следующая формула пересчёта:
$$
V_j(y_j) = \exp\{U(x_{j}, y_{j})\} \sum_{y_{j+1}}\left[ \exp\{W(y_j, y_{j+1})\} V_{j+1}(y_{j+1}) \right].
$$
Инициализировать пересчёт можно так: $V_{L-1}(y_{L-1}) = \exp\{U(x_{L-1}, y_{L-1})\}$. Окончательное значение нормировочной константы можно найти как $\sum_{y_0} [V_0(y_0)]$.
Для численно-устойчивой реализации необходимо использовать функцию log_sum_exp и производить вычисления в логарифмической шкале, т.е. найти $\log Z(\Theta)$.
HINT: для log_sum_exp используйте max trick:
$$
\log \sum\limits_{i=1}^N \exp\{x_i\} = \log \sum\limits_{i=1}^N \exp\{x_i - \max_{j}[x_j]\} + \max_{j}[x_j]
$$
Реализуйте подсчет нормировочной константы.
```
def log_sum_exp(vec, axis=0):
### code starts here ###
### code ends here ###
return result
def compute_log_partition(U, W):
"""
Parameters:
U: unary potentials, torch Variable shape (len_word, NUM_LABELS)
W: pairwise potentials, torch Variable shape (NUM_LABELS, NUM_LABELS)
Returns:
value of partition function
"""
### code starts here ###
### code ends here ###
return logZ
U = torch.load('unary_example.pth')
W = torch.load('pairwise_example.pth')
assert np.allclose(compute_log_partition(U, W), 175.63, rtol=1e-4, atol=1e-6), 'Check you compatability function'
```
### Обучение (0.2 балла)
Реализуйте подсчет negative loglikelihood и шаг оптимизации.
```
from torch.optim import Adam
from tqdm import trange
torch.manual_seed(42)
W = Variable(torch.randn(26, 26), requires_grad=True)
opt = Adam([W], lr=1e-2)
trace_values = []
n_epoch = 5
for epoch in range(n_epoch):
mean_val = 0.
for i in trange(len(train_x)):
word, target = Variable(torch.from_numpy(train_x[i]).float()),\
Variable(torch.from_numpy(train_y[i]).long())
U = unary_net(word)
logZ = compute_log_partition(U, W)
### code starts here ###
### code ends here ###
mean_val += loss.data[0]
if i % 500 == 0 and i:
trace_values.append(mean_val / 500.)
mean_val = 0.
clear_output()
plt.title('Negative loglikelihood loss')
plt.plot(np.arange(len(trace_values)), trace_values)
plt.show()
glob_acc = 0.
letters_num = np.sum([i.shape[0] for i in val_x])
for i in range(len(val_x)):
word, target = Variable(torch.from_numpy(val_x[i]).float()),\
torch.from_numpy(val_y[i]).long()
U, P = unary_net(word), W
pred = dynamic_programming(U.data, P.data)
eq_count = pred.eq(target).sum()
glob_acc += eq_count
glob_acc /= letters_num
print('global val accuracy {}'.format(glob_acc))
```
Точность на валидации должна получиться в районе 0.97.
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.