text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Numpy & Pandas
## Numpy and Arrays
Numpy is a powerful numeric library that is essential for anyone analyzing data with Python. Numpy is a huge package that can support a multitude of tasks. Numpy is also inextricably linked to SciPy, a powerful library for scientific computing with capabilities for fitting, linear algebra, machine learning, etc. Here we are just going to cover some of the basics of numpy, but I encourage you to check out the numpy documentation pages (https://numpy.org/doc/stable/) to get an idea of the variety of things you can do.
Arrays are a data type which is fundamental to Numpy. In some ways Numpy arrays are like Python lists:
- both are used for storing data/objects
- both are mutable
- items can be extracted from both using indexing and slicing
- both can be iterated over
However there are key aspects of arrays that make them very different:
- most operators act on the elements of an array instead of the array as a whole
- arrays can only hold data of a single type
- arrays can efficiently store large amounts of data in memory
```
import numpy as np
# create some sample lists
xlist = [1, 2, 3, 4]
ylist = [1, 4, 9, 16]
# create some sample arrays
x = np.array([1, 2, 3, 4])
y = np.array([1, 4, 9, 16])
```
First let's checkout the different behaviors between lists and arrays
```
print(xlist * 4)
print(x * 4)
print(x / 4)
print(xlist / 4)
```
Notice how the list was repeated 4 times, whereas each element of the array was multiplied by 4 and the result ended up being the same length.
Division works element-wise for arrays, but division is not defined and produces an error when performed on a list.
## Iterating, indexing, and slicing
Iterating over a 1D array looks just like iterating over a list
```
for val in xlist:
print(val)
for val in x:
print(val)
```
Iterating an N-dimensional array will iterate over slices along the first dimension.
```
y = np.zeros((5, 5))
for val in y:
print(val)
print()
# you could accomplish the same thing like this (but you probably shouldn't)
for i in range(y.shape[0]):
val = y[i, :]
print(val)
```
We can also select subsets of the array using conditionals.
```
xs = x[x < 2]
xs
```
Normally setting an array equal to another creates a view of the first array. In otherwords, editing either array will modify the other.
```
z = x
x[3] = 10
z
```
You can use the copy method if you really need a new copy of the array.
```
z = x.copy()
x[3] = 20
z
```
hstack and vstack are useful to stitch together multiple arrays
```
# hstack stitches together along the first dimension
hstack = np.hstack((x, z))
print(hstack)
print()
# vstack stitches along the last dimension
vstack = np.vstack((x, z))
print(vstack)
```
### Best practices
- If it's important that your code is fast, it's almost always better to avoid for loops. If I'm working on a complicated problem and I'm unsure whether to use a loop or array/DataFrame operations I usually write it up in a loop first so that I can conceptualize the problem a little easier then convert later to remove as many loops as possible.
- Loops are often more readable than list comprehensions.
## Pandas Tables
Pandas is a powerful data analysis package that provides tools for manipulating tabular data. This is particularly useful in many astronomical applications, such as spectroscopy, and time-series. Data is organized into rows and columns where the columns are named and recalled using arbitrary Python objects (strings are the most convenient). This is in contrast to Numpy arrays where columns can only be accessed using integer indices (however, see record arrays https://docs.scipy.org/doc/numpy-1.10.1/user/basics.rec.html).
Sorting, querying, merging, and aggregation are some of the most useful Pandas features, but this tutorial will only scratch the surface. See https://pandas.pydata.org/docs/ for the full documentation.
Pandas is most useful for dealing with heterogeneous and/or large datasets, when merging or complex queries are needed, or if you have metadata associated with columns (e.g. strings as labels).
The basic units/objects in Pandas are the Series and DataFrame objects.
```
import pandas as pd
# Lets create a sample Series object
x = [1.0, 2.0, 4.4, 4.5, 8.8, 9.1, 8.7, 2.3, 2.4, 3.1, 5.9]
s = pd.Series(x)
print(s)
```
We populated a Series starting from a list of floating point numbers. Notice that two columns are printed in the output. Every entry in a Series has a corresponding integer index, generally these indices are created automatically. The data type of the series is printed below the Series itself. Series objects can only store data of a single type, but any data type can be stored.
A Series is like a single column of data in a table. A DataFrame is the Pandas object that represents a full table. Each column in the table is a Series.
There are several ways to construct a Pandas DataFrame, including from Numpy arrays, Python dictionaries, a list of Series objects, reading from a CSV, reading from a URL, etc.
Let's first construct a single-column DataFrame from our series `s`.
```
df = pd.DataFrame(s, columns=['sample'])
df
```
Jupyter has special support for displaying DataFrames, simply typing the variable name of a DataFrame at the end of the cell will present a nicely formatted view of the table.
Let's add some more columns to our DataFrame.
```
df['sample_base'] = df['sample'] // 1
df['sample_plus1'] = df['sample'] + 1
df['sample_squared'] = df['sample']**2
df
```
Notice that we can access the values in a column using two different syntax.
We can also easily save/read Pandas DataFrames to/from disk
```
# write to a CSV file
df.to_csv('demo.csv')
# read back from the CSV we just saved
df = pd.read_csv('demo.csv')
# a variety of other formats are supported including JSON, ASCII, etc. Each format has its own read/write methods.
```
### Best Practices:
`numpy` is more memory efficient than `pandas`, but `pandas` is often helpful for organization and common data analysis tasks. For example, if I have a set of data that has 50 data points, each with time, radial velocity, error, S-index value, H-alpha value, and starname, a `pandas` `DataFrame` will probably be easier to keep track of than a multi-dimensional numpy array, or several 1D arrays. If `pandas` sounds like it will make your life harder rather than easier, it's probably not worth using.
Consider using `pandas` when your data are:
- heterogeneous (e.g. a mix of strings, ints, and floats)
- going to be combined with other similar data sets (e.g. I have one set of RV data, as described above, taken with the HIRES instrument, and another set from the APF instrument, and I want to extract all data taken for a given star).
# Activites
We'll break you into small groups. You will have 15 mins. Work on the following two activities as a group.
### Roles:
- navigator (person with most recent birthday)
- driver (person with farthest away birthday)
- moderator
### Product:
- Activity 1: plot of length vs time for both arrays and lists
- Activity 2: answer & justification for each scenario
## Activity #1
Let's see how much faster it is to work with Numpy arrays over Python lists.
```
import time
# First we'll create a long list
length = 10000000 # must be an int
x = list(range(length))
# now lets loop over all of the elements and add one then divide by two
# we will also use the time package to time how long it takes
t1 = time.time()
for i in range(len(x)):
x[i] = (x[i] + 1) / 2
t2 = time.time()
print("Updated {:d} elements in {:4.3f} seconds.".format(length, t2-t1))
```
1. Change the length of the list and keep track of how long the calculation takes as a function of that length.
1. Plot the time as a function of list length.
1. Now construct a Numpy array from the list `x` and perform the same calculation for several different array lengths.
1. Plot the calculation time as a function of array length and add this line to the plot created in step #2.
## Activity #2
Should you use a for loop in each of the following scenarios? Why or why not?
Scenario 1: I want to multiply each element in an array by 10.
Scenario 2: I'm writing a quicklook reduction pipleine that will run in real time (so it needs to be as fast as possible). I need to convolve each pixel in an image with the same kernel function.
Scenario 3: I'm writing an open-source data anlysis package that will be used and modified by many people. I have 10,000 images that I need to run the same set of functions on.
## (Optional) Activity #3
Lets load a couple files into a Pandas DataFrame and rearrange and merge them into a single file in a more useful format. `example_data/star_names.json` contains a list star names. The `primary_name` column is the primary ID for the star. For each unique `primary_name` there are many `other_names` associated with it. Each `primary_name`+`other_name` combination is stored in a separate row.
1. First load the file `example_data/star_names.json` into a Pandas DataFrame. The file is in JSON format so you might look into the `pandas.read_json` function.
1. Group the DataFrame on the `primary_name` column and create a custom aggregation function that takes all of the values in the `other_name` column that have the same `primary_name` and converts them into a single string deliminated with a pipe (`|`).
1. Load the `example_data/star_props.csv` file into a separate DataFrame and merge this with the grouped DataFrame from step #2.
1. Save the result as a new CSV file. The resulting file should look like `example_data/stars_merged.csv`. You may also load this file into Pandas to see what the final DataFrame should look like before saving to a CSV.
# More on Pandas Functionality
(Read through this section on your own)
Now sort by the `sample_squared` column
```
df = df.sort_values(by='sample_squared')
df
```
Notice that the indices were re-ordered as well. The indices retain information about the original ordering.
We can also select subsets of the data using conditionals similar to Numpy arrays.
```
q1 = df[df['sample'] <= 4]
q1
```
The `.groupby` method is used to create Pandas `DataFrameGroupBy` object which can be used to calculate statistics within the groups.
```
# groups that share a common sample_base field
g = df.groupby('sample_base')
# count number of rows within each group
print(g.count())
```
We can also merge DataFrames together using a common column.
Lets create a second DataFrame from the same original list of numbers and calculate the `sample_base` field again. We will also calculate a new column called `sample_sqrt`
```
df2 = pd.DataFrame(x, columns=['sample'])
df2['sample_base'] = df2['sample'] // 1
df2['sample_sqrt'] = np.sqrt(df2['sample'])
df2
```
Now we can add this new column into the original DataFrame by matching up the values on a shared column. In this case we want to match up on the original `sample` column.
Sometimes we have multiple DataFrames with one or more overlapping columns and we need to combine into a single DataFrame. This is where merging comes in.
Merging is a powerful and complex subject. I frequently find myself here: https://pandas.pydata.org/pandas-docs/stable/user_guide/merging.html to lookup various functionalities.
```
merged = pd.merge(df, df2, on='sample', suffixes=['_original', '_new'])
merged
```
If a column name appears in both DataFrames but is not the column that you are merging on, the strings defined in the `suffixes` argument will be appended to the end of the column names.
DataFrames can be written and read from files very easily. Many formats are supported but comma separated values (CSV) is the most commonly used in astronomy. The `read_csv` function can actually read a variety of text file formats by specifying the `delimiter` argument.
You can also load a CSV directly from a URL.
```
merged.to_csv('sample.csv')
!cat sample.csv
from_csv = pd.read_csv('sample.csv', index_col=0)
from_csv
```
| github_jupyter |
# An Introduction to Linear Learner with MNIST
_**Making a Binary Prediction of Whether a Handwritten Digit is a 0**_
1. [Introduction](#Introduction)
2. [Prerequisites and Preprocessing](#Prequisites-and-Preprocessing)
1. [Permissions and environment variables](#Permissions-and-environment-variables)
2. [Data ingestion](#Data-ingestion)
3. [Data inspection](#Data-inspection)
4. [Data conversion](#Data-conversion)
3. [Training the linear model](#Training-the-linear-model)
4. [Set up hosting for the model](#Set-up-hosting-for-the-model)
5. [Validate the model for use](#Validate-the-model-for-use)
## Introduction
Welcome to our example introducing Amazon SageMaker's Linear Learner Algorithm! Today, we're analyzing the [MNIST](https://en.wikipedia.org/wiki/MNIST_database) dataset which consists of images of handwritten digits, from zero to nine. We'll use the individual pixel values from each 28 x 28 grayscale image to predict a yes or no label of whether the digit is a 0 or some other digit (1, 2, 3, ... 9).
The method that we'll use is a linear binary classifier. Linear models are supervised learning algorithms used for solving either classification or regression problems. As input, the model is given labeled examples ( **`x`**, `y`). **`x`** is a high dimensional vector and `y` is a numeric label. Since we are doing binary classification, the algorithm expects the label to be either 0 or 1 (but Amazon SageMaker Linear Learner also supports regression on continuous values of `y`). The algorithm learns a linear function, or linear threshold function for classification, mapping the vector **`x`** to an approximation of the label `y`.
Amazon SageMaker's Linear Learner algorithm extends upon typical linear models by training many models in parallel, in a computationally efficient manner. Each model has a different set of hyperparameters, and then the algorithm finds the set that optimizes a specific criteria. This can provide substantially more accurate models than typical linear algorithms at the same, or lower, cost.
To get started, we need to set up the environment with a few prerequisite steps, for permissions, configurations, and so on.
## Prequisites and Preprocessing
### Permissions and environment variables
_This notebook was created and tested on an ml.m4.xlarge notebook instance._
Let's start by specifying:
- The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.
- The IAM role arn used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the boto regexp with a the appropriate full IAM role arn string(s).
```
bucket = '<your_s3_bucket_name_here>'
prefix = 'sagemaker/linear-mnist'
# Define IAM role
import boto3
import re
from sagemaker import get_execution_role
role = get_execution_role()
```
### Data ingestion
Next, we read the dataset from an online URL into memory, for preprocessing prior to training. This processing could be done *in situ* by Amazon Athena, Apache Spark in Amazon EMR, Amazon Redshift, etc., assuming the dataset is present in the appropriate location. Then, the next step would be to transfer the data to S3 for use in training. For small datasets, such as this one, reading into memory isn't onerous, though it would be for larger datasets.
```
%%time
import pickle, gzip, numpy, urllib.request, json
# Load the dataset
urllib.request.urlretrieve("http://deeplearning.net/data/mnist/mnist.pkl.gz", "mnist.pkl.gz")
with gzip.open('mnist.pkl.gz', 'rb') as f:
train_set, valid_set, test_set = pickle.load(f, encoding='latin1')
```
### Data inspection
Once the dataset is imported, it's typical as part of the machine learning process to inspect the data, understand the distributions, and determine what type(s) of preprocessing might be needed. You can perform those tasks right here in the notebook. As an example, let's go ahead and look at one of the digits that is part of the dataset.
```
%matplotlib inline
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (2,10)
def show_digit(img, caption='', subplot=None):
if subplot==None:
_,(subplot)=plt.subplots(1,1)
imgr=img.reshape((28,28))
subplot.axis('off')
subplot.imshow(imgr, cmap='gray')
plt.title(caption)
show_digit(train_set[0][30], 'This is a {}'.format(train_set[1][30]))
```
### Data conversion
Since algorithms have particular input and output requirements, converting the dataset is also part of the process that a data scientist goes through prior to initiating training. In this particular case, the Amazon SageMaker implementation of Linear Learner takes recordIO-wrapped protobuf, where the data we have today is a pickle-ized numpy array on disk.
Most of the conversion effort is handled by the Amazon SageMaker Python SDK, imported as `sagemaker` below.
```
import io
import numpy as np
import sagemaker.amazon.common as smac
vectors = np.array([t.tolist() for t in train_set[0]]).astype('float32')
labels = np.where(np.array([t.tolist() for t in train_set[1]]) == 0, 1, 0).astype('float32')
buf = io.BytesIO()
smac.write_numpy_to_dense_tensor(buf, vectors, labels)
buf.seek(0)
```
## Upload training data
Now that we've created our recordIO-wrapped protobuf, we'll need to upload it to S3, so that Amazon SageMaker training can use it.
```
import boto3
import os
key = 'recordio-pb-data'
boto3.resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'train', key)).upload_fileobj(buf)
s3_train_data = 's3://{}/{}/train/{}'.format(bucket, prefix, key)
print('uploaded training data location: {}'.format(s3_train_data))
```
Let's also setup an output S3 location for the model artifact that will be output as the result of training with the algorithm.
```
output_location = 's3://{}/{}/output'.format(bucket, prefix)
print('training artifacts will be uploaded to: {}'.format(output_location))
```
## Training the linear model
Once we have the data preprocessed and available in the correct format for training, the next step is to actually train the model using the data. Since this data is relatively small, it isn't meant to show off the performance of the Linear Learner training algorithm, although we have tested it on multi-terabyte datasets.
Again, we'll use the Amazon SageMaker Python SDK to kick off training, and monitor status until it is completed. In this example that takes between 7 and 11 minutes. Despite the dataset being small, provisioning hardware and loading the algorithm container take time upfront.
First, let's specify our containers. Since we want this notebook to run in all 4 of Amazon SageMaker's regions, we'll create a small lookup. More details on algorithm containers can be found in [AWS documentation](https://docs-aws.amazon.com/sagemaker/latest/dg/sagemaker-algo-docker-registry-paths.html).
```
containers = {'us-west-2': '174872318107.dkr.ecr.us-west-2.amazonaws.com/linear-learner:latest',
'us-east-1': '382416733822.dkr.ecr.us-east-1.amazonaws.com/linear-learner:latest',
'us-east-2': '404615174143.dkr.ecr.us-east-2.amazonaws.com/linear-learner:latest',
'eu-west-1': '438346466558.dkr.ecr.eu-west-1.amazonaws.com/linear-learner:latest'}
```
Next we'll kick off the base estimator, making sure to pass in the necessary hyperparameters. Notice:
- `feature_dim` is set to 784, which is the number of pixels in each 28 x 28 image.
- `predictor_type` is set to 'binary_classifier' since we are trying to predict whether the image is or is not a 0.
- `mini_batch_size` is set to 200. This value can be tuned for relatively minor improvements in fit and speed, but selecting a reasonable value relative to the dataset is appropriate in most cases.
```
import boto3
import sagemaker
sess = sagemaker.Session()
linear = sagemaker.estimator.Estimator(containers[boto3.Session().region_name],
role,
train_instance_count=1,
train_instance_type='ml.c4.xlarge',
output_path=output_location,
sagemaker_session=sess)
linear.set_hyperparameters(feature_dim=784,
predictor_type='binary_classifier',
mini_batch_size=200)
linear.fit({'train': s3_train_data})
```
## Set up hosting for the model
Now that we've trained our model, we can deploy it behind an Amazon SageMaker real-time hosted endpoint. This will allow out to make predictions (or inference) from the model dyanamically.
_Note, Amazon SageMaker allows you the flexibility of importing models trained elsewhere, as well as the choice of not importing models if the target of model creation is AWS Lambda, AWS Greengrass, Amazon Redshift, Amazon Athena, or other deployment target._
```
linear_predictor = linear.deploy(initial_instance_count=1,
instance_type='ml.m4.xlarge')
```
## Validate the model for use
Finally, we can now validate the model for use. We can pass HTTP POST requests to the endpoint to get back predictions. To make this easier, we'll again use the Amazon SageMaker Python SDK and specify how to serialize requests and deserialize responses that are specific to the algorithm.
```
from sagemaker.predictor import csv_serializer, json_deserializer
linear_predictor.content_type = 'text/csv'
linear_predictor.serializer = csv_serializer
linear_predictor.deserializer = json_deserializer
```
Now let's try getting a prediction for a single record.
```
result = linear_predictor.predict(train_set[0][30:31])
print(result)
```
OK, a single prediction works. We see that for one record our endpoint returned some JSON which contains `predictions`, including the `score` and `predicted_label`. In this case, `score` will be a continuous value between [0, 1] representing the probability we think the digit is a 0 or not. `predicted_label` will take a value of either `0` or `1` where (somewhat counterintuitively) `1` denotes that we predict the image is a 0, while `0` denotes that we are predicting the image is not of a 0.
Let's do a whole batch of images and evaluate our predictive accuracy.
```
import numpy as np
predictions = []
for array in np.array_split(test_set[0], 100):
result = linear_predictor.predict(array)
predictions += [r['predicted_label'] for r in result['predictions']]
predictions = np.array(predictions)
import pandas as pd
pd.crosstab(np.where(test_set[1] == 0, 1, 0), predictions, rownames=['actuals'], colnames=['predictions'])
```
As we can see from the confusion matrix above, we predict 931 images of 0 correctly, while we predict 44 images as 0s that aren't, and miss predicting 49 images of 0.
### (Optional) Delete the Endpoint
If you're ready to be done with this notebook, please run the delete_endpoint line in the cell below. This will remove the hosted endpoint you created and avoid any charges from a stray instance being left on.
```
import sagemaker
sagemaker.Session().delete_endpoint(linear_predictor.endpoint)
```
| github_jupyter |
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
fixed_band_indices = [3]
bandnames = ['u', 'g', 'r', 'i', 'z']
num_fixed_bands = len(fixed_band_indices)
# load stellar model grids
npzfile = np.load('./model_grids.npz')
stellarmodel_grid = npzfile['model_grid']
stellarmodel_grid = stellarmodel_grid[:, ::10, ::2, :] ## UNCOMMENT THAT
plotstrides = 1
gridsize1, gridsize2, numtypes = stellarmodel_grid.shape[1:4]
# logg and Teff grids
loggDA_vec=np.linspace(7., 9.5, gridsize1)
loggDB_vec=np.linspace(7., 9., gridsize1)
temp_vec = np.logspace(np.log10(3000.), np.log10(120000.), gridsize2)
# random linear grids for the GP prior. Will need to specify the logg and T grids! For the GP prior
dim1_grid = np.linspace(0, 1, stellarmodel_grid.shape[1])
dim2_grid = np.linspace(0, 1, stellarmodel_grid.shape[2])
# load data
npzfile = np.load('./WD_data_very_clean.npz')
obsmags = npzfile['obsmags']
nobj, num_bands = obsmags.shape
obsmags_covar_chol = npzfile['obsmags_covar_chol']
obsmags_covar_logdet = npzfile['obsmags_covar_logdet']
obsmags_covar = obsmags_covar_chol[:, :, :] * np.swapaxes(obsmags_covar_chol[:, :, :], 1, 2)
nobj, nobj // 100
stellarmodel_grid.shape
from matplotlib.colors import LogNorm
fig, axs = plt.subplots(3, 2, figsize=(10, 10))
axs = axs.ravel()
off = -1
for i in range(num_bands-1):
for j in range(i+1, num_bands-1):
off += 1
axs[off].set_xlabel('$'+bandnames[i]+'-'+bandnames[i+1]+'$')
axs[off].set_ylabel('$'+bandnames[j]+'-'+bandnames[j+1]+'$')
s = np.random.choice(nobj, 100, replace=False)
axs[off].errorbar(obsmags[s, i] - obsmags[s, i+1], obsmags[s, j] - obsmags[s, j+1],
xerr=np.sqrt(obsmags_covar[s, i, i] + obsmags_covar[s, i+1, i+1]),
yerr=np.sqrt(obsmags_covar[s, j, j] + obsmags_covar[s, j+1, j+1]),
fmt = 'o', ms=0, lw=1, c='r'
)
c = ['b', 'k']
for t in range(0, 2):
for g in range(0, stellarmodel_grid.shape[2], plotstrides):
mod1 = stellarmodel_grid[i, ::plotstrides, g, t] - stellarmodel_grid[i+1, ::plotstrides, g, t]
mod2 = stellarmodel_grid[j, ::plotstrides, g, t] - stellarmodel_grid[j+1, ::plotstrides, g, t]
axs[off].errorbar(mod1.ravel(), mod2.ravel(), c=c[t], fmt='-o', ms=2)
fig.tight_layout()
corrected_stellarmodel_grid = np.load('corrected_model_grids.npy')
corrections_dim1 = np.load('corrections_dim1.npy')
corrections_dim2 = np.load('corrections_dim2.npy')
# num_bands, gridsize1, numtypes
fig, axs = plt.subplots(num_bands, 2, figsize=(8, 8), sharex=False)
for i in range(num_bands):
axs[i, 0].plot(range(len(dim1_grid)), np.zeros(len(dim1_grid)), ls='--', c='k', label='Type 1')
axs[i, 1].plot(range(len(dim2_grid)), np.zeros(len(dim2_grid)), ls='--', c='k', label='Type 2')
axs[i, 0].plot(range(len(dim1_grid)), corrections_dim1[i, :, 0], label='Type 1')
axs[i, 0].plot(range(len(dim1_grid)), corrections_dim1[i, :, 1], label='Type 2')
axs[i, 1].plot(range(len(dim2_grid)), corrections_dim2[i, :, :])
axs[i, 0].set_ylabel('Corr in band '+str(i+1))
axs[i, 1].set_ylabel('Corr in band '+str(i+1))
axs[-1, 0].set_xlabel('Log g')
axs[-1, 1].set_xlabel('T')
axs[0, 0].legend()
fig.tight_layout()
# TODO: add actual log g and T grids!
# num_bands, gridsize1, numtypes
fig, axs = plt.subplots(num_bands, 2, figsize=(8, 8), sharex=False)
for i in range(num_bands):
axs[i, 0].plot(loggDA_vec, np.zeros(len(dim1_grid)), ls='--', c='k', label='Type 1')
axs[i, 1].plot(temp_vec, np.zeros(len(dim2_grid)), ls='--', c='k', label='Type 2')
axs[i, 0].plot(loggDA_vec, corrections_dim1[i, :, 0], label='Type 1')
axs[i, 0].plot(loggDB_vec, corrections_dim1[i, :, 1], label='Type 2')
axs[i, 1].plot(temp_vec, corrections_dim2[i, :, :])
axs[i, 0].set_ylabel('Corr in band '+str(i+1))
axs[i, 1].set_ylabel('Corr in band '+str(i+1))
axs[-1, 0].set_xlabel('Log g')
axs[-1, 1].set_xlabel('T')
axs[0, 0].legend()
fig.tight_layout()
fig, axs = plt.subplots(6, 2, figsize=(10, 20))
#axs = axs.ravel()
off = -1
def custom_range(x, factor=2):
m = np.mean(x)
s = np.std(x)
return [m - factor*s, m + factor * s]
for i in range(num_bands-1):
for j in range(i+1, num_bands-1):
off += 1
norm = None;#LogNorm()#
rr = [[-0.6, 1.5], [-0.6, 1]]
for t in range(0, 2):
rr = [custom_range(obsmags[:, i] - obsmags[:, i+1]), custom_range(obsmags[:, j] - obsmags[:, j+1])]
axs[off, t].hist2d(obsmags[:, i] - obsmags[:, i+1], obsmags[:, j] - obsmags[:, j+1],
30, norm=norm, range=rr, cmap='Greys', zorder=0)
axs[off, t].set_xlabel('$'+bandnames[i]+'-'+bandnames[i+1]+'$')
axs[off, t].set_ylabel('$'+bandnames[j]+'-'+bandnames[j+1]+'$')
for g in range(0, stellarmodel_grid.shape[2], plotstrides):
mod1 = stellarmodel_grid[i, ::plotstrides, g, t] - stellarmodel_grid[i+1, ::plotstrides, g, t]
mod2 = stellarmodel_grid[j, ::plotstrides, g, t] - stellarmodel_grid[j+1, ::plotstrides, g, t]
axs[off, t].plot(mod1.ravel(), mod2.ravel(), c='blue', zorder=1)
mod1 = corrected_stellarmodel_grid[i, ::plotstrides, g, t] - corrected_stellarmodel_grid[i+1, ::plotstrides, g, t]
mod2 = corrected_stellarmodel_grid[j, ::plotstrides, g, t] - corrected_stellarmodel_grid[j+1, ::plotstrides, g, t]
axs[off, t].plot(mod1.ravel(), mod2.ravel(), c='orange', zorder=2)
fig.tight_layout()
```
| github_jupyter |
Source: https://www.dataquest.io/blog/pandas-big-data/
```
import pandas as pd
import numpy as np
movie = pd.read_csv('/Users/ankushchoubey/Downloads/ml-latest-small/movies.csv')
movie.tail()
movie.info(memory_usage='deep')
gl_int = movie.select_dtypes(include=['int'])
converted_int = gl_int.apply(pd.to_numeric,downcast='unsigned')
def mem_usage(pandas_obj):
if isinstance(pandas_obj,pd.DataFrame):
usage_b = pandas_obj.memory_usage(deep=True).sum()
else: # we assume if not a df it's a series
usage_b = pandas_obj.memory_usage(deep=True)
usage_mb = usage_b / 1024 ** 2 # convert bytes to megabytes
reply = "{:03.2f} MB".format(usage_mb)
print(reply)
return reply
print(mem_usage(gl_int))
print(mem_usage(converted_int))
movie[converted_int.columns]= converted_int
movie.info(memory_usage='deep')
gl_obj = movie.select_dtypes(include=['object'])
converted_to_cat = gl_obj['genres'].astype('category')
movie['genres'] = converted_to_cat
movie.info(memory_usage='deep')
def reduce_mem_usage(df):
start_mem_usg = df.memory_usage().sum() / 1024**2
print("Memory usage of properties dataframe is :",start_mem_usg," MB")
NAlist = [] # Keeps track of columns that have missing values filled in.
conversion = {}
for col in df.columns:
if df[col].dtype != object: # Exclude strings
# Print current column type
print("******************************")
print("Column: ",col)
print("dtype before: ",df[col].dtype)
# make variables for Int, max and min
IsInt = False
mx = df[col].max()
mn = df[col].min()
# Integer does not support NA, therefore, NA needs to be filled
if not np.isfinite(df[col]).all():
NAlist.append(col)
df[col].fillna(mn-1,inplace=True)
# test if column can be converted to an integer
asint = df[col].fillna(0).astype(np.int64)
result = (df[col] - asint)
result = result.sum()
if result > -0.01 and result < 0.01:
IsInt = True
# Make Integer/unsigned Integer datatypes
if IsInt:
if mn >= 0:
if mx < 255:
df[col] = df[col].astype(np.uint8)
elif mx < 65535:
df[col] = df[col].astype(np.uint16)
elif mx < 4294967295:
df[col] = df[col].astype(np.uint32)
else:
df[col] = df[col].astype(np.uint64)
else:
if mn > np.iinfo(np.int8).min and mx < np.iinfo(np.int8).max:
df[col] = df[col].astype(np.int8)
elif mn > np.iinfo(np.int16).min and mx < np.iinfo(np.int16).max:
df[col] = df[col].astype(np.int16)
elif mn > np.iinfo(np.int32).min and mx < np.iinfo(np.int32).max:
df[col] = df[col].astype(np.int32)
elif mn > np.iinfo(np.int64).min and mx < np.iinfo(np.int64).max:
df[col] = df[col].astype(np.int64)
# Make float datatypes
else:
if mn > np.finfo(np.float16).min and mx < np.finfo(np.float16).max:
df[col] = df[col].astype(np.float16)
elif mn > np.finfo(np.float32).min and mx < np.finfo(np.float32).max:
df[col] = df[col].astype(np.float32)
elif mn > np.finfo(np.float64).min and mx < np.finfo(np.float64).max:
df[col] = df[col].astype(np.float64)
# Print new column type
print("dtype after: ",df[col].dtype)
print("******************************")
conversion[col]=str(df[col].dtype)
# Print final result
print("___MEMORY USAGE AFTER COMPLETION:___")
mem_usg = df.memory_usage().sum() / 1024**2
print("Memory usage is: ",mem_usg," MB")
print("This is ",100*mem_usg/start_mem_usg,"% of the initial size")
return df,conversion
a, NAlist = reduce_mem_usage(movie)
ratings_large = pd.read_csv('/Users/ankushchoubey/Downloads/ml-latest/ratings.csv')
d,v,conversion=reduce_mem_usage(ratings_large)
conversion
```
| github_jupyter |
```
# torchfactor package
from torchfactor.factorization.nmfnet import NMFNet
from torchfactor.experiment.experiment import Experiment
# other imports
import torch
import torchvision
import numpy as np
import matplotlib.pyplot as plt
class StackedMNISTDataset(torch.utils.data.Dataset):
"""A 1 element dataset that returns a column-wise stack of flattened images
for NMF training.
"""
def __init__(self, stacksize):
self.mnist = torchvision.datasets.MNIST(
"../../data/mnist", train=True,
transform=torchvision.transforms.Compose([torchvision.transforms.ToTensor()])
)
self.stacksize = stacksize
self.img_indices = np.random.randint(0, len(self.mnist), size=self.stacksize)
self.img_stack = torch.zeros((28*28, self.stacksize))
for i in range(self.img_indices.shape[0]):
self.img_stack[:,i] = self.mnist[self.img_indices[i]][0].flatten()
def __len__(self):
return 1
def __getitem__(self, index):
return self.img_stack
def show_recons(input_X, W, H):
stackrecons = W @ H
fig, ax = plt.subplots(10, 2, figsize=(10, 20))
for i in range(len(ax)):
img_recons = stackrecons[:,i].reshape((28,28))
ax[i, 0].imshow(input_X[:,i].reshape((28,28)))
ax[i, 0].set_title('Original Matrix {}'.format(i))
recon_loss = torch.nn.functional.mse_loss(torch.Tensor(stackrecons[:,i]),
torch.Tensor(input_X[:,i]))
ax[i, 1].imshow(img_recons)
ax[i, 1].set_title('W @ H Matrix -- Recon Error:{:2f}'.format(recon_loss))
plt.tight_layout()
plt.show()
def show_cols(W):
fig, ax = plt.subplots(5, 2, figsize=(10, 20))
for i in range(len(ax)):
ax[i, 0].imshow(W[:,2 * i].reshape((28,28)))
ax[i, 1].imshow(W[:,2 * i + 1].reshape((28, 28)))
plt.tight_layout()
plt.suptitle('Columns of W', fontsize=20, y=1.02)
plt.show()
imstack_dataset = StackedMNISTDataset(stacksize=200)
imstack_dataloader = torch.utils.data.DataLoader(
imstack_dataset, batch_size=1
)
### NMFNet with L2 Loss
num_comp = 20
net = NMFNet(28*28, num_comp)
optimizer = torch.optim.Adam(net.parameters(), lr=1e-2)
experiment = Experiment(
net=net, loss=torch.nn.MSELoss(), optimizer=optimizer,
train_dataloader=imstack_dataloader, validation_dataloader=imstack_dataloader,
inputs_are_ground_truth=True, has_labels=False
)
train_loss_over_epochs, val_loss_over_epochs, _ = experiment.run(
train_epochs=5000, train_validation_interval=1000
)
input_X = imstack_dataset[0]
W_net = net.get_W().detach().numpy()
H_net = net.get_H(input_X).detach().numpy()
show_recons(input_X, W_net, H_net)
show_cols(W_net)
### NMFNet with Hyrbid Loss
num_comp = 20
net = NMFNet(28*28, num_comp)
optimizer = torch.optim.Adam(net.parameters(), lr=1e-2)
experiment = Experiment(
net=net, loss=torch.nn.SmoothL1Loss(), optimizer=optimizer,
train_dataloader=imstack_dataloader, validation_dataloader=imstack_dataloader,
inputs_are_ground_truth=True, has_labels=False
)
train_loss_over_epochs, val_loss_over_epochs, _ = experiment.run(
train_epochs=5000, train_validation_interval=1000
)
input_X = imstack_dataset[0]
W_net = net.get_W().detach().numpy()
H_net = net.get_H(input_X).detach().numpy()
show_recons(input_X, W_net, H_net)
show_cols(W_net)
### Regular NMF
from sklearn.decomposition import NMF
model = NMF(n_components=20, init='random', random_state=0)
W = model.fit_transform(imstack_dataset[0])
H = model.components_
show_recons(imstack_dataset[0], W, H)
show_cols(W)
```
| github_jupyter |
# Milestone Project 2 - Walkthrough Steps Workbook
Below is a set of steps for you to follow to try to create the Blackjack Milestone Project game!
## Game Play
To play a hand of Blackjack the following steps must be followed:
1. Create a deck of 52 cards
2. Shuffle the deck
3. Ask the Player for their bet
4. Make sure that the Player's bet does not exceed their available chips
5. Deal two cards to the Dealer and two cards to the Player
6. Show only one of the Dealer's cards, the other remains hidden
7. Show both of the Player's cards
8. Ask the Player if they wish to Hit, and take another card
9. If the Player's hand doesn't Bust (go over 21), ask if they'd like to Hit again.
10. If a Player Stands, play the Dealer's hand. The dealer will always Hit until the Dealer's value meets or exceeds 17
11. Determine the winner and adjust the Player's chips accordingly
12. Ask the Player if they'd like to play again
## Playing Cards
A standard deck of playing cards has four suits (Hearts, Diamonds, Spades and Clubs) and thirteen ranks (2 through 10, then the face cards Jack, Queen, King and Ace) for a total of 52 cards per deck. Jacks, Queens and Kings all have a rank of 10. Aces have a rank of either 11 or 1 as needed to reach 21 without busting. As a starting point in your program, you may want to assign variables to store a list of suits, ranks, and then use a dictionary to map ranks to values.
## The Game
### Imports and Global Variables
** Step 1: Import the random module. This will be used to shuffle the deck prior to dealing. Then, declare variables to store suits, ranks and values. You can develop your own system, or copy ours below. Finally, declare a Boolean value to be used to control <code>while</code> loops. This is a common practice used to control the flow of the game.**
suits = ('Hearts', 'Diamonds', 'Spades', 'Clubs')
ranks = ('Two', 'Three', 'Four', 'Five', 'Six', 'Seven', 'Eight', 'Nine', 'Ten', 'Jack', 'Queen', 'King', 'Ace')
values = {'Two':2, 'Three':3, 'Four':4, 'Five':5, 'Six':6, 'Seven':7, 'Eight':8, 'Nine':9, 'Ten':10, 'Jack':10,
'Queen':10, 'King':10, 'Ace':11}
```
import random
suits = ('Hearts','Diamonds','Spades','Clubs')
ranks = ('Two','Three','Four','Five','Six','Seven','Eight','Nine','Ten','Jack','Queen','King','Ace')
values = {'Two' : 2,'Three' : 3,'Four':4,'Five':5,'Six':6,'Seven':7,'Eight':8,'Nine':9,'Ten':10,'Jack':10,'Queen':10,'King':10,'Ace':11}
playing = True
```
### Class Definitions
Consider making a Card class where each Card object has a suit and a rank, then a Deck class to hold all 52 Card objects, and can be shuffled, and finally a Hand class that holds those Cards that have been dealt to each player from the Deck.
**Step 2: Create a Card Class**<br>
A Card object really only needs two attributes: suit and rank. You might add an attribute for "value" - we chose to handle value later when developing our Hand class.<br>In addition to the Card's \_\_init\_\_ method, consider adding a \_\_str\_\_ method that, when asked to print a Card, returns a string in the form "Two of Hearts"
```
class Card:
suit =[]
rank =[]
value =[]
def __init__(self,suit,rank,value):
self.suit = suit
self.rank = rank
self.value = value
def __str__(self):
return "%s of %s and have value %s"%(self.rank,self.suit,self.value)
```
**Step 3: Create a Deck Class**<br>
Here we might store 52 card objects in a list that can later be shuffled. First, though, we need to *instantiate* all 52 unique card objects and add them to our list. So long as the Card class definition appears in our code, we can build Card objects inside our Deck \_\_init\_\_ method. Consider iterating over sequences of suits and ranks to build out each card. This might appear inside a Deck class \_\_init\_\_ method:
for suit in suits:
for rank in ranks:
In addition to an \_\_init\_\_ method we'll want to add methods to shuffle our deck, and to deal out cards during gameplay.<br><br>
OPTIONAL: We may never need to print the contents of the deck during gameplay, but having the ability to see the cards inside it may help troubleshoot any problems that occur during development. With this in mind, consider adding a \_\_str\_\_ method to the class definition.
```
class Deck:
def __init__(self):
self.deck = []
for suit in suits:
for rank in ranks:
value = values[rank]
self.deck.append(Card(suit,rank,value))
def __str__(self):
deck_comp = ''
for card in self.deck:
deck_comp += '\n' + card.__str__()
return 'the desk has \n'+deck_comp
def shuffle(self):
random.shuffle(self.deck)
def deal(self):
single_card = self.deck.pop()
return single_card
```
TESTING: Just to see that everything works so far, let's see what our Deck looks like!
```
test_deck = Deck()
test_deck.shuffle()
print(test_deck)
```
Great! Now let's move on to our Hand class.
**Step 4: Create a Hand Class**<br>
In addition to holding Card objects dealt from the Deck, the Hand class may be used to calculate the value of those cards using the values dictionary defined above. It may also need to adjust for the value of Aces when appropriate.
```
class Hand:
def __init__(self):
self.cards = [] # start with an empty list as we did in the Deck class
self.value = 0 # start with zero value
self.aces = 0 # add an attribute to keep track of aces
def add_card(self,card):
#card passed in from Deck.deal()
self.cards.append(card)
self.value += card.value
if card.rank == 'Ace':
self.aces += 1
def adjust_for_ace(self):
#if total value >21 and i still have an ace
# than change my ace to be 1 instead of an 11
while self.value>21 and self.ace:
self.value -= 10
self.aces -= 1
test_deck =Deck()
test_deck.shuffle()
#player
test_player = Hand()
pulled_card = test_deck.deal()
print(pulled_card)
test_player.add_card(pulled_card)
print(test_player.value)
test_player.add_card(test_deck.deal())
print(test_player.value)
```
**Step 5: Create a Chips Class**<br>
In addition to decks of cards and hands, we need to keep track of a Player's starting chips, bets, and ongoing winnings. This could be done using global variables, but in the spirit of object oriented programming, let's make a Chips class instead!
```
class Chips:
def __init__(self,total = 100):
self.total = total # This can be set to a default value or supplied by a user input
self.bet = 0
def win_bet(self):
self.total += self.bet
def lose_bet(self):
self.total -= self.bet
```
### Function Defintions
A lot of steps are going to be repetitive. That's where functions come in! The following steps are guidelines - add or remove functions as needed in your own program.
**Step 6: Write a function for taking bets**<br>
Since we're asking the user for an integer value, this would be a good place to use <code>try</code>/<code>except</code>. Remember to check that a Player's bet can be covered by their available chips.
```
def take_bet(Chips):
while True:
try:
chips.bet = int(input('Enter amount you would like to bet'))
break
except:
print('Enter only integer !!')
else:
if chips.bet > chips.total:
print('you dont have enough chips you have : {}'.format(chips.total))
else:
break
```
**Step 7: Write a function for taking hits**<br>
Either player can take hits until they bust. This function will be called during gameplay anytime a Player requests a hit, or a Dealer's hand is less than 17. It should take in Deck and Hand objects as arguments, and deal one card off the deck and add it to the Hand. You may want it to check for aces in the event that a player's hand exceeds 21.
```
def hit(deck,hand):
hand.add_card(deck.deal())
hand.adjust_for_ace()
```
**Step 8: Write a function prompting the Player to Hit or Stand**<br>
This function should accept the deck and the player's hand as arguments, and assign playing as a global variable.<br>
If the Player Hits, employ the hit() function above. If the Player Stands, set the playing variable to False - this will control the behavior of a <code>while</code> loop later on in our code.
```
def hit_or_stand(deck,hand):
global playing # to control an upcoming while loop
while True:
x = input('Hit or stand? Enter h or s')# hit #Hit #Stand
if x[0].lower() == 's':
print('Player Stand dealer turn')
playing = False
break
elif x[0].lower() == 'h':
print('Player hit')
hit(deck,hand)
playing = True
break
else :
print("wrong input,try again!!")
continue
```
**Step 9: Write functions to display cards**<br>
When the game starts, and after each time Player takes a card, the dealer's first card is hidden and all of Player's cards are visible. At the end of the hand all cards are shown, and you may want to show each hand's total value. Write a function for each of these scenarios.
```
def show_some(player,dealer):
for card in player.cards:
print(card)
for card in dealer.cards[1]:
print(card)
def show_all(player,dealer):
for card in player.cards:
print(card)
for card in dealer.cards:
print(card)
```
**Step 10: Write functions to handle end of game scenarios**<br>
Remember to pass player's hand, dealer's hand and chips as needed.
```
def player_busts():
pass
def player_wins():
pass
def dealer_busts():
pass
def dealer_wins():
pass
def push():
pass
```
### And now on to the game!!
```
while True:
# Print an opening statement
# Create & shuffle the deck, deal two cards to each player
# Set up the Player's chips
# Prompt the Player for their bet
# Show cards (but keep one dealer card hidden)
while playing: # recall this variable from our hit_or_stand function
# Prompt for Player to Hit or Stand
# Show cards (but keep one dealer card hidden)
# If player's hand exceeds 21, run player_busts() and break out of loop
break
# If Player hasn't busted, play Dealer's hand until Dealer reaches 17
# Show all cards
# Run different winning scenarios
# Inform Player of their chips total
# Ask to play again
break
```
And that's it! Remember, these steps may differ significantly from your own solution. That's OK! Keep working on different sections of your program until you get the desired results. It takes a lot of time and patience! As always, feel free to post questions and comments to the QA Forums.
# Good job!
| github_jupyter |
# Training a Hierarchical DivNoising network for Convallaria data which is intrinsically noisy
This notebook contains an example on how to train a Hierarchical DivNoising Ladder VAE for an intrinsically noisy data. This requires having a noise model (model of the imaging noise) which can be either measured from calibration data or bootstrapped from raw noisy images themselves. If you haven't done so, please first run '1-CreateNoiseModel.ipynb', which will download the data and create a noise model.
```
import warnings
warnings.filterwarnings('ignore')
# We import all our dependencies.
import numpy as np
import torch
import sys
sys.path.append('../../../')
from models.lvae import LadderVAE
from lib.gaussianMixtureNoiseModel import GaussianMixtureNoiseModel
from boilerplate import boilerplate
import lib.utils as utils
import training
from tifffile import imread
from matplotlib import pyplot as plt
from tqdm import tqdm
use_cuda = torch.cuda.is_available()
device = torch.device("cuda" if use_cuda else "cpu")
```
### Specify ```path``` to load training data
Your data should be stored in the directory indicated by ```path```.
```
path="./data/Convallaria_diaphragm/"
observation= imread(path+'20190520_tl_25um_50msec_05pc_488_130EM_Conv.tif')
```
# Training Data Preparation
For training we need to follow some preprocessing steps first which will prepare the data for training purposes.
We first divide the data into training and validation sets with 85% images allocated to training set and rest to validation set. Then we augment the training data 8-fold by 90 degree rotations and flips.
```
train_data = observation[:int(0.85*observation.shape[0])]
val_data= observation[int(0.85*observation.shape[0]):]
print("Shape of training images:", train_data.shape, "Shape of validation images:", val_data.shape)
train_data = utils.augment_data(train_data) ### Data augmentation disabled for fast training, but can be enabled
### We extract overlapping patches of size ```patch_size x patch_size``` from training and validation images.
### Usually 64x64 patches work well for most microscopy datasets
patch_size = 64
img_width = observation.shape[2]
img_height = observation.shape[1]
num_patches = int(float(img_width*img_height)/float(patch_size**2)*1)
train_images = utils.extract_patches(train_data, patch_size, num_patches)
val_images = utils.extract_patches(val_data, patch_size, num_patches)
val_images = val_images[:1000] # We limit validation patches to 1000 to speed up training but it is not necessary
test_images = val_images[:100]
img_shape = (train_images.shape[1], train_images.shape[2])
print("Shape of training images:", train_images.shape, "Shape of validation images:", val_images.shape)
```
# Configure Hierarchical DivNoising model
<code>model_name</code> specifies the name of the model with which the weights will be saved and wil be loaded later for prediction.<br>
<code>directory_path</code> specifies the directory where the model weights and the intermediate denoising and generation results will be saved. <br>
<code>gaussian_noise_std</code> is only applicable if dataset is synthetically corrupted with Gaussian noise of known std. For real datasets, it should be set to ```None```.<br>
<code>noiseModel</code> specifies a noise model for training. If noisy data is generated synthetically using Gaussian noise, set it to None. Else set it to the GMM based noise model (.npz file) generated from '1-CreateNoiseModel.ipynb'.<br>
<code>batch_size</code> specifies the batch size used for training. The default batch size of $64$ works well for most microscopy datasets.<br>
<code>virtual_batch</code> specifies the virtual batch size used for training. It divides the <code>batch_size</code> into smaller mini-batches of size <code>virtual_batch</code>. Decrease this if batches do not fit in memory.<br>
<code>test_batch_size</code> specifies the batch size used for testing every $1000$ training steps. Decrease this if test batches do not fit in memory, it does not have any consequence on training. It is just for intermediate visual debugging.<br>
<code>lr</code> specifies the learning rate.<br>
<code>max_epochs</code> specifies the total number of training epochs. Around $150-200$ epochs work well generally.<br>
<code>steps_per_epoch</code> specifies how many steps to take per epoch of training. Around $400-500$ steps work well for most datasets.<br>
<code>num_latents</code> specifies the number of stochastic layers. The default setting of $6$ works well for most datasets but quite good results can also be obtained with as less as $4$ layers. However, more stochastic layers may improve performance for some datasets at the cost of increased training time.<br>
<code>z_dims</code> specifies the number of bottleneck dimensions (latent space dimensions) at each stochastic layer per pixel. The default setting of $32$ works well for most datasets.<br>
<code>blocks_per_layer</code> specifies how many residual blocks to use per stochastic layer. Usually, setting it to be $4$ or more works well. However, more residual blocks improve performance at the cost of increased training time.<br>
<code>batchnorm</code> specifies if batch normalization is used or not. Turning it to True is recommended.<br>
<code>free_bits</code> specifies the threshold below which KL loss is not optimized for. This prevents the [KL-collapse problem](https://arxiv.org/pdf/1511.06349.pdf%3Futm_campaign%3DRevue%2520newsletter%26utm_medium%3DNewsletter%26utm_source%3Drevue). The default setting of $1.0$ works well for most datasets.<br>
**__Note:__** With these settings, training will take approximately $24$ hours on Tesla P100/Titan Xp GPU needing about 6 GB GPU memory. We optimized the code to run on less GPU memory. For faster training, consider increasing ```virtual_batch_size``` but since we have not tested with different settings of ```virtual_batch_size```, we do not yet know how this affects results. To reduce traing time, also consider reducing either ```num_latents``` or ```blocks_per_layer``` to $4$. These settings will bring down the training time to around $12-15$ hours while still giving good results.
```
model_name = "convallaria"
directory_path = "./Trained_model/"
# Data-specific
gaussian_noise_std = None
noise_model_params= np.load("./data/Convallaria_diaphragm/GMMNoiseModel_convallaria_3_2_calibration.npz")
noiseModel = GaussianMixtureNoiseModel(params = noise_model_params, device = device)
# Training-specific
batch_size=64
virtual_batch = 8
lr=3e-4
max_epochs = 500
steps_per_epoch=400
test_batch_size=100
# Model-specific
num_latents = 6
z_dims = [32]*int(num_latents)
blocks_per_layer = 5
batchnorm = True
free_bits = 1.0
```
# Train network
```
train_loader, val_loader, test_loader, data_mean, data_std = boilerplate._make_datamanager(train_images,val_images,
test_images,batch_size,
test_batch_size)
model = LadderVAE(z_dims=z_dims,blocks_per_layer=blocks_per_layer,data_mean=data_mean,data_std=data_std,noiseModel=noiseModel,
device=device,batchnorm=batchnorm,free_bits=free_bits,img_shape=img_shape).cuda()
model.train() # Model set in training mode
training.train_network(model=model,lr=lr,max_epochs=max_epochs,steps_per_epoch=steps_per_epoch,directory_path=directory_path,
train_loader=train_loader,val_loader=val_loader,test_loader=test_loader,
virtual_batch=virtual_batch,gaussian_noise_std=gaussian_noise_std,
model_name=model_name,val_loss_patience=30)
```
# Plotting losses
```
trainHist=np.load(directory_path+"model/train_loss.npy")
reconHist=np.load(directory_path+"model/train_reco_loss.npy")
klHist=np.load(directory_path+"model/train_kl_loss.npy")
valHist=np.load(directory_path+"model/val_loss.npy")
plt.figure(figsize=(18, 3))
plt.subplot(1,3,1)
plt.plot(trainHist,label='training')
plt.plot(valHist,label='validation')
plt.xlabel("epochs")
plt.ylabel("loss")
plt.legend()
plt.subplot(1,3,2)
plt.plot(reconHist,label='training')
plt.xlabel("epochs")
plt.ylabel("reconstruction loss")
plt.legend()
plt.subplot(1,3,3)
plt.plot(klHist,label='training')
plt.xlabel("epochs")
plt.ylabel("KL loss")
plt.legend()
plt.show()
```
| github_jupyter |
# Multi-Class Classification with iris flowers dataset
## Problem Description
In this tutorial we will use the standard machine learning problem called the [iris flowers dataset](http://archive.ics.uci.edu/ml/datasets/Iris).
This dataset is well studied and is a good problem for practicing on neural networks because all of the 4 input variables are numeric and have the same scale in centimeters. Each instance describes the properties of an observed flower measurements and the output variable is specific iris species.
This is a multi-class classification problem, meaning that there are more than two classes to be predicted, in fact there are three flower species. This is an important type of problem on which to practice with neural networks because the three class values require specialized handling.
The iris flower dataset is a well studied problem and as such we can expect to achieve a model accuracy in the range of 95% to 97%. This provides a good target to aim for when developing our models.
You can [download the iris flowers dataset](http://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data) from the UCI Machine Learning repository and place it in your current working directory with the filename iris.csv
### Attribute Information:
1. sepal length in cm
2. sepal width in cm
3. petal length in cm
4. petal width in cm
5. class:
* Iris Setosa
* Iris Versicolour
* Iris Virginica
## Import Classes and Functions
We can begin by importing all of the classes and functions we will need in this tutorial.
This includes both the functionality we require from Keras, but also data loading from [pandas](http://pandas.pydata.org/) as well as data preparation and model evaluation from [scikit-learn](http://scikit-learn.org/).
```
import numpy as np
import pandas as pd
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
from sklearn.model_selection import cross_val_score, KFold
from keras.models import Sequential
from keras.layers import Dense, Activation
from keras.wrappers.scikit_learn import KerasClassifier
from livelossplot import PlotLossesKeras
```
## Initialize Random Number Generator
Next we need to initialize the random number generator to a constant value (7).
This is important to ensure that the results we achieve from this model can be achieved again precisely. It ensures that the stochastic process of training a neural network model can be reproduced.
```
# fix random seed for reproducibility
seed = 7
np.random.seed(seed)
```
## Load The Dataset
The dataset can be loaded directly. Because the output variable contains strings, it is easiest to load the data using pandas. We can then split the attributes (columns) into input variables (X) and output variables (Y).
```
# load dataset
dataframe = pd.read_csv("../datasets/iris.csv", header=None)
dataframe.head()
dataset = dataframe.values
X_data = dataset[:,0:4].astype(float)
y_data = dataset[:,4]
print("X matrix shape", X_data.shape)
print("y matrix shape", y_data.shape)
```
# Encode The Output Variable
The output variable contains three different string values.
When modeling multi-class classification problems using neural networks, it is good practice to reshape the output attribute from a vector that contains values for each class value to be a matrix with a boolean for each class value and whether or not a given instance has that class value or not.
This is called [one hot encoding](https://en.wikipedia.org/wiki/One-hot) or creating dummy variables from a categorical variable.
For example, in this problem three class values are Iris-setosa, Iris-versicolor and Iris-virginica. If we had the observations:
* Iris-setosa
* Iris-versicolor
* Iris-virginica
We can turn this into a one-hot encoded binary matrix for each data instance that would look as follows:
<pre>
| Iris-setosa | Iris-versicolor | Iris-virginica |
|-------------|-----------------|----------------|
| 1 | 0 | 0 |
|-------------|-----------------|----------------|
| 0 | 1 | 0 |
|-------------|-----------------|----------------|
| 0 | 0 | 1 |
</pre>
We can do this by first encoding the strings consistently to integers using the scikit-learn class LabelEncoder, followed by one-hot encoding with the class OneHotEncoder.
```
label_encoder = LabelEncoder()
y_data = label_encoder.fit_transform(y_data).reshape(-1,1)
one_hot_encoder = OneHotEncoder()
y_data = one_hot_encoder.fit_transform(y_data).toarray()
```
## Define The Neural Network Model
The Keras library provides wrapper classes to allow you to use neural network models developed with Keras in scikit-learn.
There is a KerasClassifier class in Keras that can be used as an Estimator in scikit-learn, the base type of model in the library. The KerasClassifier takes the name of a function as an argument. This function must return the constructed neural network model, ready for training.
Below is a function that will create a baseline neural network for the iris classification problem. It creates a simple fully connected network with one hidden layer that contains 4 neurons, the same number of inputs (it could be any number of neurons).
The hidden layer uses a rectifier activation function which is a good practice. Because we used one-hot encoding for our iris dataset, the output layer must create 3 output values, one for each class. The output value with the largest value will be taken as the class predicted by the model.
The network topology of this simple one-layer neural network can be summarized as:
* inputs -> [4 hidden nodes] -> 3 outputs
Note, that we use a sigmoid activation function in the output layer. This is to ensure the output values are in the range of 0 and 1 and may be used as predicted probabilities.
Finally, the network uses the efficient ADAM gradient descent optimization algorithm with a logarithmic loss function, which is called categorical_crossentropy in Keras.
```
def baseline_model():
model = Sequential()
model.add(Dense(4, input_dim=4, kernel_initializer='normal'))
model.add(Activation('sigmoid'))
model.add(Dense(3, kernel_initializer='normal'))
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
return model
```
## Create the Model
```
model = baseline_model()
```
## Define training parameters
```
batch_size = 5
epochs = 200
```
## Train the model
```
model.fit(X_data, y_data, batch_size=batch_size, epochs=epochs, verbose=1, callbacks=[PlotLossesKeras()])
```
## Evaluate the model with k-Fold Cross Validation
We can now create our KerasClassifier for use in scikit-learn.
We can also pass arguments in the construction of the KerasClassifier class that will be passed on to the fit() function internally used to train the neural network. Here, we pass the number of epochs as 200 and batch size as 5 to use when training the model.
Debugging is also turned off when training by setting verbose to 0.
```
estimator = KerasClassifier(build_fn=baseline_model, epochs=epochs, batch_size=batch_size, verbose=0)
```
We can now evaluate the neural network model on our training data.
The scikit-learn has excellent capability to evaluate models using a suite of techniques.
The gold standard for evaluating machine learning models is k-fold cross validation.
First we can define the model evaluation procedure.
Here, we set the number of folds to be 10 (an excellent default) and to shuffle the data before partitioning it.
```
kfold = KFold(n_splits=10, shuffle=True, random_state=None)
```
Now we can evaluate our model (estimator) on our dataset (X_data and Y_data) using a 10-fold cross validation procedure (kfold).
Evaluating the model only takes approximately 10 seconds and returns an object that describes the evaluation of the 10 constructed models for each of the splits of the dataset.
```
results = cross_val_score(estimator, X_data, y_data, cv=kfold)
print("Accuracy: %.2f%% (%.2f%%)" % (results.mean()*100, results.std()*100))
```
The results are summarized as both the mean and standard deviation of the model accuracy on the dataset.
This is a reasonable estimation of the performance of the model on unseen data.
It is also within the realm of known top results for this problem.
| github_jupyter |
# A first look at the data
## Introduction
In this section we will set the paths and load the data. We will then explain the data structure and do a few sanity checks. Finally follow some examples for how to perform simples plots of the raw data.
Import the required modules:
```
import os
import numpy as np
import mne
```
## Loading the data
Loading the data. The MEGIN data are stored in the FIF format which is a binary format with embedded labels.
As a first step, we set the path to the data. Note that this will dependent on where you have stored the dataset. Afterwards, we set the file names.
```
data_path = r'C:\Users\JensenO\Dropbox\FLUX\Development\dataRaw'
# data_path = r'THE PATH TO DATA ON YOUR LOCAL SYSTEM'
file_name = ['training_raw-1.fif','training_raw-2.fif']
```
There is a limit to the file size of the FIF files. If the MEG recordings exceed this limit, the MEGIN acquisition system automatically split the data in two or more FIF files. In those cases, by reading the first FIF file MNE-Python you will automatically read all the linked split files. In our case, the sample dataset was broken into 2 sub-files by the operator and we need to read them one by one.
Start by reading the first file:
```
path_data = os.path.join(data_path,file_name[0])
data1 = mne.io.read_raw_fif(path_data)
```
To get some basic information from the FIF file write:
```
print(data1)
```
The 5.4 MB refer to the size of the data set. The 1457.0 s is the recording time. As the sampling rate was 1000 Hz, this results in 1457000 samples recorded in 343 channels. These channels include the MEG, EOG, triggers channels etc.
To get some additional information:
```
print(data1.info)
```
This set of information show important parameters of the dataset. For instance, we can see that the gantry was positioned at 68 degress (*gantry_angle*) and that the sampling frequency was 1000 Hz (*sfreq*). The data were lowpass filtered at 330.0 Hz (*lowpass*) and highpass filtered at 0.1 Hz (*highpass*) prior to the digital sampling.
**Question 1:** What is the purpose of the 330 Hz lowpas-filter (hint: see Analog Filters for Data Conversion, Chapter 3; Smith et al., 1999)
**Question 2:** What is the purpose of 0.1 Hz highpass-filter?
The data object (*data1*) allows for a simple inspection of the data by plotting the spectra:
```
%matplotlib inline
data1.plot_psd(fmax=60);
```
Note the 10 Hz alpha activity on the gradiometers as well as the 50 Hz line noise. The frequency of the line noise is 50 Hz in European countries including Russia whereas it is 60 in South Korea as well as South and North America. In Japan it can be either 50 or 60 Hz depending on the region.
**Question 3:** Why is the line-noise 50 Hz larger for the magnetometers than the gradiometers?
To show the raw data use:
```
%matplotlib inline
data1.plot(duration=10,title='Raw');
```
To enable the interactive functions of the plotting tool:
```
%matplotlib qt
data1.plot(duration=10,title='Raw');
```
This will open the plot in a new window. Use the arrow keys to move over channels and time. Click on the help button for more information.
**Question 3:** Scroll through the MEG data and identify the following artifacts (include figures in the response):
- Eye-blink
- A muscle contraction (characterized by high-frequency activity
- The cardiac artifact
## Preregistration and publication
Preregistration: report the sampling frequency and the properties of the anti-aliasing lowpass filter. Also report the peripheral data to be recorded (e.g. the EOG and ECG)
Publication, example:
"The ongoing MEG data were recorded using the TRIUX system from MEGIN. This system has 102 magnetometers and 204 planer gradiometers. These are placed at 102 locations each having one magnetometer and a set of two orthogonal gradiometers. The horizontal and vertical EOG data as well as the ECG were acquired together with the MEG data. The data were sampled at 1000 Hz and stored for offline analysis. Prior to sampling, a lowpass filter at ~330 Hz was applied. To record the horizontal EOG, a pair of electrodes were attached approximately 2.5 cm away from the outer canthus of each eye. To record the vertical EOG, a pair of electrodes were placed above and below the right eye in line with the pupil. The ECG was recorded from a pair of electrodes placed on the left and right collarbone. Four head position indicator coils (HPIs) were placed behind the left and right ear as well as on the left and right forehead just below the hairline. The positions of the HPIs, the nasion, the left and right preauricular points, as well as the surface points of the scalp, were digitized using a PolhemusTM device. "
## References
Smith, W.S. (1997) The Scientist and Engineer's Guide to Digital Signal Processing. California Technical Publishing. ISBN 0-9660176-3-3 [Online version](http://www.dspguide.com/)
| github_jupyter |
# The Method of Lagrangian Descriptors
## Introduction and Theory
One of the biggest challenges of dynamical systems theory or nonlinear dynamics is the development of mathematical techniques that provide us with the capability of exploring the geometrical template of structures that governs transport in phase space. Since the early 1900, the idea of pursuing a qualitative description of the solutions of differential equations, which emerged from the pioneering work carried out by Henri Poincaré on the three body problem of celestial mechanics \cite{hp1890}, has had a profound impact on our understanding of the nonlinear character of natural phenomena. Indeed, this powerful approach has now been widely embraced by the scientific community and its essence was nicely captured by Vladimir Arnold's statement that a complete description of classical mechanics boils down to the geometrical analysis of phase space.
In this section we present the details of a mathematical tool whose potential brings us one step closer to fulfilling the long sought after dream envisioned by Poincaré. The method is known in the literature as Lagrangian descriptors (LDs) and has the capability of revealing the geometrical template of phase space structures that characterizes trajectories with qualitatively distinct dynamical behavior. As we will see, this method provides us with a systematic way of exploring phase space by means of looking at its dynamical skeleton using low-dimensional slices. This procedure allows for a complete reconstruction, that is, a *phase space tomography*, of the intricate geometry of underlying invariant manifolds that characterize transport mechanisms.
Consider a general dynamical system:
\begin{equation}
\dfrac{d\mathbf{x}}{dt} = \mathbf{v}(\mathbf{x},t) \;,\quad \mathbf{x} \in \mathbb{R}^{n} \;,\; t \in \mathbb{R} \;,
\label{eq:gtp_dynSys}
\end{equation}
where the vector field satifies $\mathbf{v}(\mathbf{x},t) \in C^{r} (r \geq 1)$ in $\mathbf{x}$ and continuous in time. If the vector field does not depend on time, the system is called *autonomous*, and it is *non-autonomous* otherwise. For any initial condition $\mathbf{x}(t_0) = \mathbf{x}_0$ this system of differential equations has a unique solution with the same regularity as that of the vector field, which also depends continuously on the initial data \cite{coddington1984}. The vector field can be specified as an analytical model, or it could also have been retrieved from numerical simulations as a discrete spatio-temporal dataset.
The method of Lagrangian Descriptors was first introduced to analyze Lagrangian transport and mixing in geophysical flows \cite{madrid2009,mendoza2010}. Initially, LDs relied on the arclength of the trajectories as they evolve from their initial conditions forward and backward in time \cite{mendoza2010,mancho2013lagrangian}. Since its proposal as a nonlinear dynamics tool, it was used to plan a transoceanic autonomous underwater vehicle missions \cite{ramos2018}, manage marine oil spills \cite{gg2016}, analyze the Stratospheric Polar Vortex \cite{alvaro1,alvaro2,curbelo2019a,curbelo2019b}, and recently in chemical reaction dynamics \cite{craven2015lagrangian,craven2016deconstructing,craven2017lagrangian,revuelta2019unveiling}.
Our goal in this section is to give an introduction to the method of Lagrangian descriptors, which can reveal the geometry of phase space structures that determine transport in dynamical systems, such as in Eq. \eqref{eq:gtp_dynSys}. The geometry of phase space structures is encoded in the trajectories of the system, which can be extracted using LDs from their initial conditions. The simple idea behind LDs is to seed a given phase space region with initial conditions and integrate a bounded and positive quantity (an intrinsic geometrical and/or physical property of the dynamical system under study) along trajectories for a finite time. This approach reveals the invariant phase space structures that make up the dynamical skeleton governing reaction dynamics. This is similar to the visualization techniques developed in fluid mechanics experiments in the laboratory to uncover patterns of flow structures by studying the evolution of particles in a moving fluid \cite{chien1986}. A powerful analogy can be found in iron filings that align with the magnetic field lines of a magnet.
One of the biggest challenges that one is faced with when exploring the high-dimensional phase space of a dynamical system, such as those that occur in Hamiltonian systems, is that of the qualitative description of the behavior of ensembles of initial conditions, and recovering from their trajectory evolution the underlying template of geometrical phase space structures that governs the dynamical transport mechanisms of the flow. The problem that naturally arises in this context is that the trajectories of ensembles of initial conditions that start nearby might get ''lost'' with respect to each other very quickly, making the use of classical nonlinear dynamics techniques (Small Alignment Index, Lyapunov Exponents, etc...) that rely on tracking the location of neighboring trajectories computationally expensive and difficult to interpret. Furthermore, when Poincaré maps are applied for the dynamical analysis of high-dimensional systems, one might encounter the issue of trajectories not coming back to the selected surface of section, which would yield no relevant information whatsoever.
The method of Lagrangian descriptors provides tremendous advantages in comparison to other methodologies to overcome these issues. For instance, it is a computationally inexpensive and straightforward to implement tool to explore nonlinear dynamics. But probably, the key and revolutionary idea behind the success of this technique is that it focuses on integrating a positive scalar function along trajectories of initial conditions of the system instead of tracking their phase space location. In this way, by emphasizing initial conditions, it directly targets the building blocks where the dynamical structure of phase space is encoded. The methodology offered by LDs has thus the capability of producing a complete and detailed geometrical *phase space tomography* in high dimensions by means of using low-dimensional phase space probes to extract the intersections of the phase space invariant manifolds with these slices \cite{demian2017,naik2019a,naik2019b,GG2019}. Any phase space slice can be selected and sampled with a high-resolution grid of initial conditions, and no information regarding the dynamical skeleton af invariant manifolds at the given slice is lost as the trajectories evolve in time. Moreover, this analysis does not rely on trajectories coming back to the chosen slice, as is required for Ponicaré maps to work. In this respect, there is also another key point that needs to be highlighted which demonstrates the real potential of LDs with respect to other classical nonlinear dynamics techniques. Using LDs we can obtain *all* the invariant manifolds of the dynamical system *simultaneously*, the hyperbolic stable and unstable manifolds coming from *any* NHIM in phase space, and also the KAM tori. This provides an edge over the classical approach of computing stable and unstable manifolds that relies on locating first the NHIMs in phase space individually, and for every NHIM globalize the manifolds separately, for which a knowledge of the eigendirections is crucial. Consequently, the application of LDs offers the capability of recovering *all* the relevant phase space structures in one *shot* without having to study the local dynamics about equilibrium points of the dynamical system.
### Formulations for Lagrangian Descriptors
The Lagrangian descriptor is a scalar valued non-negative function $M$, that is integrated forward and backward for a fixed intergation time $\tau$. Originally $M$ was defined using the arclength:
\begin{equation}
M(\mathbf{x_0},t_0,\tau) = \int_{t_0-\tau}^{t_0+\tau} ||\mathbf{v}(\mathbf{x}(t;\mathbf{x_0},t)|| \; dt \;,
\label{eq:M_function}
\end{equation}
where $||\cdot||$ is the Euclidean distance. $M$ can be naturally broken down into a forward ($M^f$) and backward ($M^b$) integral:
\begin{equation}
M(\mathbf{x}_{0},t_0,\tau) = M^b(\mathbf{x}_{0},t_0,\tau) + M^f(\mathbf{x}_{0},t_0,\tau) \;,
\end{equation}
where we have that:
\begin{equation}
M^f(\mathbf{x}_{0},t_0,\tau) = \int^{t_0+\tau}_{t_0} ||\mathbf{v}(\mathbf{x}(t;\mathbf{x}_0),t)|| \; dt \;\;,\;\;
M^b(\mathbf{x}_{0},t_0,\tau) = \int^{t_0}_{t_0-\tau} ||\mathbf{v}(\mathbf{x}(t;\mathbf{x}_0),t)|| \; dt \;.
\end{equation}
The advantage of this splitting is that $M^f$ highlights the stable manifolds of the dynamical system, and $M^b$ recovers the unstable manfifolds, while $M$ shows all the invariant manifolds simultaneously.
The intuitive idea why this tool works is that the influence of phase space structures on trajectories will result in differences (abrupt change) in arclength of nearby trajectories in the neighborhood of a phase space structure.
The method captures this distinct dynamical behavior separated by invariant phase space structures that results in abrupt changes of values of $M$. This detection of invariant manifolds has been mathematically quantified in terms of ''singular structures'' \cite{mancho2013lagrangian,lopesino2017}, where $M$ is non-differentiable. Once the manifolds are known, one can compute the NHIM at their intersection by means of a root search algorithm. An alternative method to recover the manifolds and their associated NHIM is by minimizing the function $M$ using a search optimization algorithm. This second procedure and some interesting variations are described in \cite{feldmaier2019}.
We remark that there is no general ''golden rule'' for selecting the value of $\tau$ for exploring phase space. The *appropriate* (usually chosen by trial and error) value of $\tau$ will unveil the relevant geometrical template of phase space structures. A very low value of $\tau$ will not reveal any structures and very high value may lead to obscurity due to differences in magnitude of LD values. This means that $\tau$ is intimately related to the time scales of the dynamical phenomena that occur in the system under consideration. One needs to bear in mind the compromise that exists between the complexity of the structures revealed by the method to explain a certain dynamical phenomenon.
An alternative definition of LDs relies on the $p$-norm of the vector field of the dynamical system, where $p \in (0,1]$ and was first introduced in \cite{lopesino2017} as:
\begin{equation}
M_p(\mathbf{x}_{0},t_0,\tau) = \sum_{k=1}^{n} \int^{t_0+\tau}_{t_0-\tau} |v_{k}(\mathbf{x}(t;\mathbf{x}_0),t)|^p \; dt \;,
\label{eq:Mp_function}
\end{equation}
Although this alternative definition of LDs does not have such an intuitive physical interpretation as that of arclength, it has been shown allow for a rigorous analysis of the notion of ''singular structures'' and to establish the mathematical connection of this notion to invariant stable and unstable manifolds in phase space. Furthermore, forward integration will reveal stable manifolds and backward evolution unveils unstable manifolds, as in the arclegth version of LDs. Another important aspect of the $p$-norm of LDs is that, since in the definition all the vector field components contribute separately, one can naturally decompose the LD in a way that allows to isolate distinct dynamical effects such as hyperbolic and elliptic behavior. This was used in \cite {demian2017,naik2019a} to show that the method can be used to successfully detect NHIMs and their stable and unstable manifolds in Hamiltonian systems. Furthermore, it is important to remark that with this definition of LDs one can mathematically prove that these phase space structures are detected as singularities of the $M_p$ scalar field, that is, at points where the function is non-differentiable and therefore its gradient takes very large values \cite{lopesino2017,demian2017,naik2019a}. Moreover, in this context it has also been shown that:
\begin{equation}
\mathcal{W}^u(\mathbf{x}_{0},t_0) = \textrm{argmin } M_p^{(b)}(\mathbf{x}_{0},t_0,\tau) \quad,\quad \mathcal{W}^s(\mathbf{x}_{0},t_0) = \textrm{argmin } M_p^{(f)}(\mathbf{x}_{0},t_0,\tau) \;,
\label{eq:min_LD_manifolds}
\end{equation}
where $\mathcal{W}^u$ and $\mathcal{W}^s$ are, respectively, the unstable and stable manifolds calculated at time $t_0$ and $\textrm{argmin}(\cdot)$ denotes the phase space coordinates $\mathbf{x}_0$ that minimize the function $M_p$. In addition, NHIMs at time $t_0$ can be calculated as the intersection of the stable and unstable manifolds:
\begin{equation}
\mathcal{N}(\mathbf{x}_{0},t_0) = \mathcal{W}^u(\mathbf{x}_{0},t_0) \cap \mathcal{W}^s(\mathbf{x}_{0},t_0) = \textrm{argmin} M_p(\mathbf{x}_{0},t_0,\tau)
\label{eq:min_NHIM_LD}
\end{equation}
At this point, we would like to discuss the issues that arise from the definitions of LDs provided in Eqs. \eqref{eq:M_function} and \eqref{eq:Mp_function} when they are applied to analyze dynamics in open Hamiltonian systems. Notice that in both definitions all the initial conditions are integrated for the same time $\tau$. Recent studies have revealed \cite{junginger2017chemical,naik2019b,GG2019} that computing fixed-time LDs, that is, integrating all initial conditions chosen on a phase space surface for the same integration time $\tau$, could give rise to issues related to the fact that some of the trajectories that escape from the potential energy surface can go to infinity in finite time or at an increasing rate. The trajectories that show this behavior will give NaN values in the LD scalar field, hiding some regions of the phase space, and therefore obscuring the detection of invariant manifolds. In order to circumvent this problem we explain here the approach that has been recently adopted in the literature \cite{junginger2017chemical,naik2019b,GG2019} known as variable integration time Lagrangian descriptors. In this methodology, LDs are calculated, at any initial condition, for a fixed initial integration time $\tau_0$ or until the trajectory of that initial condition leaves a certain phase space region $\mathcal{R}$ that we call the {\em interaction region}. Therefore, the total integration time in this strategy depends on the initial conditions themselves, that is $\tau(\mathbf{x}_0)$. In this variable-time formulation, given a fixed integration time $\tau_0 > 0$, the $p$-norm definition of LDs with $p \in (0,1]$ has the form:
\begin{equation}
M_p(\mathbf{x}_{0},t_0,\tau) = \sum_{k=1}^{n} \int^{t_0 + \tau^{+}_{\mathbf{x}_0}}_{t_0 - \tau^{-}_{\mathbf{x}_0}} |v_{k}(\mathbf{x}(t;\mathbf{x}_0),t)|^p \; dt \;,
\label{eq:Mp_vt}
\end{equation}
and the total integration time is defined as:
\begin{equation}
\tau^{\pm}_{\mathbf{x}_{0}} = \min \left\lbrace \tau_0 \, , \, |t^{\pm}|_{\big| \mathbf{x}\left(t^{\pm}; \, \mathbf{x}_{0}\right) \notin \mathcal{R}} \right\rbrace \; ,
\end{equation}
where $t^{+}$ and $t^{-}$ are the times for which the trajectory leaves the interaction region $\mathcal{R}$ in forward and backward time, respectively. It is important to highlighting that we select a large enough interaction region, the variable integration time LD definition given above in Eq. \eqref{eq:Mp_vt} will approach the fixed-time LD definition in Eq. \eqref{eq:Mp_function}. Thus, NHIMs and their stable and unstable manifolds will be captured by the phase space points for which the LD is non-differentiable and local minimum behavior given in Eqs. \eqref{eq:min_LD_manifolds} and \eqref{eq:min_NHIM_LD} is recovered. Moreover, KAM tori are also detected by contour values of the time-averaged LD. Therefore, the variable integration time LDs provides us with a suitable methodology to study the phase space structures that characterize escaping dynamics in open Hamiltonians, since it avoids the issue of trajectories escaping to infinity very fast.
## Revealing the Phase Space Structure
### Linear System
We describe this result for the two degrees-of-freedom system given by the linear quadratic Hamiltonian
associated to a index-1 saddle at the origin. This Hamiltonian and the equations of motion are
given by the expressions:
\begin{equation}
H(x,y,p_x,p_y) = \dfrac{\lambda}{2}\left(p_x^2 - x^2\right) + \dfrac{\omega}{2} \left(p_y^2 + y^2 \right) \quad,\quad \begin{cases}
\dot{x} = \lambda \, p_x \\
\dot{p}_{x} = \lambda \, x \\
\dot{y} = \omega \, p_y \\
\dot{p}_{y} = -\omega \, y
\end{cases}
\label{eq:index1_Ham}
\end{equation}
Given the initial condition $\mathbf{x}_0 = \mathbf{x}(t_0) = \left(x_0,y_0,p_x^0,p_y^0\right) \in \mathbb{R}^4$, the general solution is:
\begin{equation}
\begin{split}
x(t) = \frac{1}{2} \left[A e^{\lambda t} + B e^{-\lambda t}\right] \;\;&,\;\;
p_x(t) = \frac{1}{2} \left[A e^{\lambda t} - B e^{-\lambda t}\right] \\[.1cm]
y(t) = y_0 \cos(\omega t) + p^0_y \sin(\omega t) \;\;&,\;\;
p_y(t) = p^0_y \cos(\omega t) - y_0 \sin(\omega t)
\end{split}
\label{eq:gen_sol_linham}
\end{equation}
where $A = x_0 + p_x^0$ and $B = x_0 - p_x^0$. Since the system is autonomous, we can choose without loss of generality the initial time as $t_0 = 0$. Decomposing the $p$-norm LDs into the hyperbolic and elliptic components of the system we get:
\begin{equation}
M_{p}(\mathbf{x}_{0},\tau) = M^{h}_{p}(\mathbf{x}_{0},\tau) + M^{e}_{p}(\mathbf{x}_{0},\tau) \;,
\label{eq:hyp_comp_M}
\end{equation}
where the hyperbolic ($M^{h}$) and elliptic ($M^{e}$) parts are:
\begin{equation}
M^{h}_{p}(\mathbf{x}_{0},\tau) = \int^{\tau}_{-\tau} |\dot{x}(t;\mathbf{x}_0)|^p + |\dot{p}_{x}(t;\mathbf{x}_0)|^p \; dt \;\;,\;\; M^{e}_{p}(\mathbf{x}_{0},\tau) = \int^{\tau}_{-\tau} |\dot{y}(t;\mathbf{x}_0)|^p + |\dot{p}_{y}(t;\mathbf{x}_0)|^p \; dt
\label{eq:ell_comp_M}
\end{equation}
By means of an asympotic analysis when the integration time is sufficiently large, i.e. $\tau \gg 1$, it is straightforward to show \cite{lopesino2017,demian2017,naik2019a} that the asymptotic behavior of the hyperbolic component is:
\begin{equation}
M_{p}^{h}\left(\mathbf{x}_0,\tau\right) \sim \left(\dfrac{\lambda}{2}\right)^{p-1} \dfrac{|A|^{p} + |B|^{p}}{p} \, e^{p \lambda \tau}
\label{eq:M_hyp_asymp}
\end{equation}
Therefore, this result shows that $M_{\gamma}^{h}$ grows exponentially with $\tau$ and also that the leading order singularities in $M_{p}^{h}$ occur when $|A| = 0$, that is, when $p^0_x = - x_0$, which corresponds to initial conditions on the stable manifold of the NHIM, or in the case where $|B| = 0$, that is, $p^0_x = x_0$, corresponding to initial conditions on the unstable manifold of the NHIM. Moreover, $M_{p}^{h}$ is non-differentiable at the NHIM, since it is given by the intersection of the stable and unstable manifolds.
\begin{equation}
\lim_{\tau \to \infty} \dfrac{1}{2\tau} M^{e}_{p}(\mathbf{x}_{0},\tau) = \dfrac{2}{\pi} \left(\omega R\right)^{p} B\left(\dfrac{p+1}{2},\dfrac{1}{2}\right)
\label{eq:lim_m_gamma}
\end{equation}
where $R$ is the radius of the circular periodic orbit described by the $y$ DoF in the center space $y-p_y$ . So the time average of the elliptic part of LDs converges to a value that depends on the energy of the $y$ DoF, and therefore in the limit its value is constant for all points of the periodic orbit. This result is of interest because it allows us to use LDs as a tool to recover phase space KAM tori, given that the dynamical system under study satisfies the conditions of the Ergodic Partition Theorem \cite{mezic1999}. Therefore, if one analyzes the time average of LDs on a specific initial condition in phase space $\mathbf{x}_0$, and its value converges, then this initial copndition would lie in an invariant phase space set cosisting of points that share the same time average value. Therefore, the contours of the time average of LDs, when it converges, identify invariant phase space sets. It is also important to highlight here that the limit value to which $M^{e}$ converges is directly related to the limit value to which the classical arclength definition of LDs tends when just applied to the elliptic component of the system. Indeed:
\begin{equation}
\displaystyle{\lim_{\tau \to \infty} \dfrac{1}{2\tau} M^{e}_{p}} = \dfrac{2}{\pi} \left(\lim_{\tau \to \infty} \dfrac{1}{2\tau} M^{e}\right)^{p} B\left(\dfrac{p+1}{2},\dfrac{1}{2}\right)
\end{equation}
where $M^e$ represents the arclength LD in Eq. \eqref{eq:M_function} applied only to the $y$ DoF.
To conclude this theoretical discussion we will show how LDs attains a local minimum at the phase space points corresponding to the stable and unstable manifolds of the NHIM and a global minimum at the NHIM. Given an energy of the system, $H = H_0$, above that of the origin, we know from the Lyapunov Subcenter Manifold Theorem \cite{weinstein1973,moser1976,rabinowitz1982} that a family of NHIM parametrized by the energy bifurcates from the index-1 saddle. The phase space points that lie on the stable (or unstable) manifold, that is $x_0 = -p_x^0$ ($x_0 = p_x^0$), contribute to the hyperbolic component of LDs described in Eq. \eqref{eq:M_hyp_asymp} with $|A| = 0$ ($|B| = 0$) so that the Lagrangian descriptor has a local minimum at the manifold. Moreover, if the initial condition is on the NHIM, that is $x_0 = p_x^0 = 0$, then $|A| = |B| = 0$ and consequently $M^{h}_{p}(\mathbf{x}_{0},\tau) = 0$. Furthermore, all the energy of the system for these points concentrates on the $y$ DoF which evolves periodically, implying that $M^{e}_{p}(\mathbf{x}_{0},\tau) > 0$. As a result, LDs attain a global minimum at those points.
In order to support the theretical argument presented above, we provide also a numerical computation of the $p$-norm LDs in the saddle space $x-p_x$ of the linear Hamiltonian given in Eq. \eqref{eq:index1_Ham} using $p = 1/2$ and for an integration time $\tau = 10$. In Fig. \ref{fig:index1_lds} we illustrate how the method detects the stable (red) and unstable (blue) manifolds of the unstable periodic orbit at the origin, and these manifolds can be directly extracted from the ridges of the scalar field $||\nabla M_p||$. Moreover, we show in Fig. \ref{fig:index1_lds} C) that the LD scalar field is non-differentiable at the manifolds and also attains a local minimum on them, and we do so by looking at the values taken by $M_p$ along the line $p_x = 1/2$.
<img src="figures/fig3.png" width="100%">
\label{fig:index1_lds}
\caption{Phase portrait in the saddle space of the linear Hamiltonian given in Eq. \eqref{eq:index1_Ham}. A) Applicaton of the $p$-norm definition of LDs in Eq. \eqref{eq:Mp_function} using $p = 1/2$ with $\tau = 10$; B) Invariant stable (blue) and unstable (red) manifolds of the unstable periodic orbit at the origin extracted from the gradient of the $M_p$ function; C) Value of LDs along the line $p_x = 0.5$ depicted in panel A) to illustrate how the method detects the stable and unstable manifolds at points where the scalar field is singular or non-differentiable and attains a local minimum.}
### Time-dependent double well potential
In the following example, we illustrate how the arclength definition of LDs captures the stable and unstable manifolds that determine the phase portrait of the time-dependent double well potential, commonly known in dynamical systems as the forced Duffing oscillator. The Duffing equation arises when studying the motion of a particle on a line, i.e. a one degree of freedom system, subjected to the influence of a symmetric double well potential and an external forcing. The second order ODE that describes this oscillator is given by:
\begin{equation}
\ddot{x} + x^3 - x = \varepsilon f(t) \quad \Leftrightarrow \quad
\begin{cases}
\dot{x} = y \\
\dot{y} = x - x^3 + \varepsilon f(t) \\
\end{cases}
\end{equation}
where $\varepsilon$ measures the strength of the forcing term $f(t)$, and we choose for this example a sinusoidal force $f(t) = \sin(t)$. In the autonomous case, i.e. $\varepsilon = 0$, the system has three equilibrium points: a saddle located at the origin and two diametrally opoosed centers at the points $(\pm 1,0)$. The stable and unstable manifolds that emerge from the saddle point form two homoclininc orbits in the form of a figure eight around the two center equilibria:
\begin{equation}
\mathcal{W}^{s} = \mathcal{W}^{u} = \left\{(x,y) \in \mathbb{R}^2 \; \Big| \; 2y^2 + x^4 - 2x^2 = 0 \right\}
\label{eq:duff_homocMani}
\end{equation}
We begin by computing LDs for the unforced Duffing system using $\tau = 2$. For this small integration time, the method highlights the saddle and center fixed points, since the arclength at those points is always zero. Moreover, in this case the phase portrait still looks blurry as shown in Fig. \ref{fig:duffing1_lds} A), and this is a consequence of trajectories not having sufficient time to evolve in order to make distinct dynamical behaviors ditinguishable. If we now increase the integration time to $\tau = 10$, we can see in Fig. \ref{fig:duffing1_lds} B) that the homoclinic connection formed by the stable and unstable manifolds of the saddle point at the origin becomes clearly visible. Moreover, observe that the manifolds are located at points where the scalar values taken by LDs change abruptly. This property is demonstrated in Fig. \ref{fig:duffing1_lds} C), where we have depicted the value of function $M$ along the line $y = 0.5$. Notice that sharp changes in the scalar field of LDs at the manifolds are also related to local minima.
<img src="figures/fig1.png" width="100%">
\label{fig:duffing1_lds}
\caption{Phase portrait of the autonomous and undamped Duffing oscillator obtained by applying the arclength definition of LDs in Eq. \eqref{eq:M_function}. A) LDs with $\tau = 2$; B) LDs with $\tau = 10$; C) Value of LDs along the line $y = 0.5$ depicted in panel B) illustrating how the method detects the stable and unstable manifolds at points where the scalar field changes abruptly.}
We move on to compute LDs for the forced Duffing oscillator. In this situation, the vector field is time-dependent and thus the dynamical system is nonautonomous. The consequence is that the homoclinic connection breaks up and the stable and unstable manifolds intersect, forming an intricate tangle that gives rise to chaos. We illustrate this phenomenon by computing LDs with $\tau = 10$ to reconstruct the phase portrait at the initial time $t_0 = 0$. This result is shown in Fig. \ref{fig:duffing2_lds} C), and we also depict the forward $(M^f)$ and backward $(M^b)$ contributions of LDs in Fig. \ref{fig:duffing2_lds} A) and B) respectively, demonstrating that the method can be used to recover the stable and unstable manifolds separately. Furthermore, by taking the value of LDs along the line $y = 0.5$, the location of the invariant manifolds are highlighted at points corresponding to sharp changes (and local minima) in the scalar field values of LDs.
<img src="figures/fig2.png" width="80%">
\label{fig:duffing2_lds}
\caption{Phase portrait of the nonautonomous and undamped Duffing oscillator obtained at time $t = 0$ by applying the arclength definition of LDs in Eq. \eqref{eq:M_function} with an integration time $\tau = 10$. A) Forward LDs detect stable manifolds; B) Backward LDs highlight unstable manifolds of the system; C) Total LDs (forward $+$ backward) showing that all invariant manifolds are recovered simultaneously. D) Value taken by LDs along the line $y = 0.5$ in panel C) to illustrate how the method detects the stable and unstable manifolds at points where the scalar field changes abruptly.}
### Cubic potential
In order to illustrate the issues encountered by the fixed integration time LDs and how the variable integration approach resolves them, we apply the method to a basic one degree-of-freedom Hamiltonian known as the ''fish potential'', which is given by the formula:
\begin{equation}
H = \dfrac{1}{2} p_x^2 + \dfrac{1}{2} x^2 + \dfrac{1}{3} x^3 \quad \Leftrightarrow \quad
\begin{cases}
\dot{x} = p_x \\
\dot{p}_{x} = - x - x^2
\end{cases} \;.
\label{eq:fish_Ham}
\end{equation}
This dynamical system has a saddle point at the point $(-1,0)$ from which a homoclinic orbit emerges, which surrounds the elliptic point located at the origin. By applying the $p$-norm LD with $p = 1/2$ and integrating all initial conditions for the same time $\tau = 3$ we clearly observe in Fig. \ref{fig:fish_lds} A) the problems that appear in the detection of phase space structures due to trajectories escaping to infinity in finite time. If we increase $\tau$ further, very large and NaN values of LDs completely obscure the phase portrait of the system. On the other hand, if now we use the variable integration time LDs with $\tau = 8$ and we select for the interaction region a circle of radius $r = 15$ centered at the origin, the homoclininc orbit and the equilibrium points are nicely captured. Moreover, we can extract the stable and unstable manifolds of the system from the sharp ridges in the gradient of the scalar field, due to the fact that the method is non-differentiable at the location of the manifolds.
<img src="figures/fig4.png" width="100%">
\label{fig:fish_lds}
\caption{Phase portrait of the ''fish potential'' Hamiltonian in Eq. \eqref{eq:fish_Ham} revealed by the $p$-norm LDs with $p = 1/2$. A) Fixed-time integration LDs in Eq. \eqref{eq:Mp_function} with $\tau = 3$; B) Variable-time integration definition of LDs in Eq. \eqref{eq:Mp_vt} with $\tau = 8$; C) Invariant stable (blue) and unstable (red) manifolds of the saddle fixed point extracted from the gradient of the variable time $M_p$ function.}
### Hénon-Heiles Hamiltonian
We conclude this tutorial on how to apply the method of Lagrangian descriptors to unveil the dynamical skeleton of a four-dimensional phase space by applying this tool to a hallmark Hamiltonian system of nonlinear dynamics, the H\'enon-Heiles Hamiltonian introduced in 1964 to study the motion of stars in galaxies \cite{henon1964}. This system is described by:
\begin{equation}
H = \dfrac{1}{2} \left(p_x^2 + p_y^2\right) + \dfrac{1}{2}\left(x^2 + y^2\right) + x^2y - \dfrac{1}{3} y^3 \quad \Leftrightarrow \quad
\begin{cases}
\dot{x} = p_x \\
\dot{p}_{x} = - x - 2xy \\
\dot{y} = p_y \\
\dot{p}_{y} = - y - x^2 + y^2
\end{cases} \;.
\label{eq:henon_system}
\end{equation}
which has four equilibrium points: one minimum located at the origin and three saddle-center points at $(0,1)$ and $(\pm \sqrt{3}/2,-1/2)$. The potential energy surface is $V(x,y) = x^2/2 + y^2/2 + x^2y - y^3/3$ which has a $\pi/3$ rotational symmetry and is characterized by a central scattering region about the origin and three escape channels, see Fig. \ref{fig:henonHeiles_pes} below for details.
<img src="figures/fig5.png" width="100%">
\label{fig:henonHeiles_pes}
\caption{Potential energy surface for the H\'enon-Heiles system.}
In order to analyze the phase space of the H\'enon-Heiles Hamiltonian by means of the variable integration time LDs, we fix an energy $H = H_0$ of the system and choose an interaction region $\mathcal{R}$ defined in configuration space by a circle of radius $15$ centered at the origin. For our analysis we consider the following phase space slices:
\begin{eqnarray}
\mathcal{U}^{+}_{y,p_y} & = \left\{(x,y,p_x,p_y) \in \mathbb{R}^4 \;|\; H = H_0 \;,\; x = 0 \;,\; p_x > 0\right\} \\[.1cm]
\mathcal{V}^{+}_{x,p_x} &= \left\{(x,y,p_x,p_y) \in \mathbb{R}^4 \;|\; H = H_0 \;,\; y = 0 \;,\; p_y > 0\right\}
\label{eq:psos}
\end{eqnarray}
Once we have fixed the surfaces of section (SOS) where we want to compute LDs, we select a grid of initial conditions and, after discarding those that are energetically unfeasible, we integrate the remaining conditions both forward and backward in time, and compute LDs using the definition in Eq. \eqref{eq:Mp_vt} with $p = 1/2$ along the trajectory for the whole fixed integration time or until the initial condition leaves the interaction region $\mathcal{R}$, what happens first. The result obtained when the LDs values are plotted will reveal the stable and unstable manifolds and also de KAM tori in the surface of section under consideration. Since the stable and unstable manifolds are detected at points where the LD scalar function is non-differentiable, we can directly extract them from the gradient, that is, using $||\nabla \mathcal{M}_p||$. We begin by looking at the phase space structures on the SOS $\mathcal{U}^{+}_{y,p_y}$. To do so, we fix an energy for the system $H = 1/12$, which is below that of the saddle-center equilibrium points. For that energy level, the exit channels of the PES are closed, and therefore, all trajectories are trapped in the scattering region of the central minimum at the origin. We can clearly see in Fig. \ref{fig:henonHeiles_lds} A)-B) that the computation of the $p$-norm variable integration time LDs with $p = 1/2$ using $\tau = 50$ reveals that the motion of the system is completely regular. The method nicely captures the UPO present in the central region of the PES and also its stable and unstable manifolds which form a homoclininc cnnection. In order to demonstrate how the intricate details of chaotic motion are captured by LDs, we increase the energy of the system to $H = 1/3$. This energy level is now above that of the index-1 saddles of the PES, and consequently, phase space bottlenecks open in the energy manifold allowing trajectories of the system to escape to infinity from the scattering region. When we apply LDs using $\tau = 10$ on the SOSs defined in Eq. \eqref{eq:psos}, we observe in Figs. \ref{fig:henonHeiles_lds} C)-F) that we can detect with high-fidelity the intricate homoclinic tangle formed by the stable and unstable manifolds of the UPO associated to the upper index-1 saddle of the PES. Moreover, observe that despite the issue of trajectories escaping to infinity in finite time, LDs succeed in revealing the template of geometrical phase space structures that governs transport and escape dynamics from the PES of the H\'enon-Heiles Hamiltonian system.
<img src="figures/fig6.png" width="100%">
\label{fig:henonHeiles_lds}
\caption{Phase space structures of the H\'enon-Heiles Hamiltonian as revealed by the $p$-norm variable integration time LDs with $p = 1/2$. A) LDs computed for $\tau = 50$ in the SOS $\mathcal{U}^{+}_{y,p_y}$ with energy $H = 1/12$; C) LDs for $\tau = 10$ in the SOS $\mathcal{U}^{+}_{y,p_y}$ with energy $H = 1/3$; E) LDs for $\tau = 10$ in the SOS $\mathcal{V}^{+}_{x,p_x}$ with energy $H = 1/3$;. In the right panels we have extracted the invariant stable (blue) and unstable (red) manifolds from the gradient of LDs.}
To finish this chapter we would like to mention that the method of Lagrangian descriptors has also been adapted to explore the template of geometrical structures present in the phase space of stochastic dynamical systems \cite{balibrea2016lagrangian}. This achievenment clearly evidences the versatility that this mathematical technique brings to the nonlinear dynamics community. We are confident that the analysis of stochastic processes that play a crucial role in chemical reaction dynamics by means of LDs will surely help to shed light and provide new and interesting insights in the development of Chemistry. Definitely, Lagrangian descriptors has become the ultimate tool to reveal the strong bond that exists between Chemistry and Mathematics in phase space.
## Implications for Reaction Dynamics
## References
\bibliography{LDs}
| github_jupyter |
Convolutional Dictionary Learning
=================================
This example demonstrates the use of [dictlrn.cbpdndl.ConvBPDNDictLearn](http://sporco.rtfd.org/en/latest/modules/sporco.dictlrn.cbpdndl.html#sporco.dictlrn.cbpdndl.ConvBPDNDictLearn) for learning a convolutional dictionary from a set of training images. The dictionary learning algorithm is based on the ADMM consensus dictionary update [[1]](http://sporco.rtfd.org/en/latest/zreferences.html#id44) [[26]](http://sporco.rtfd.org/en/latest/zreferences.html#id25).
```
from __future__ import print_function
from builtins import input
import pyfftw # See https://github.com/pyFFTW/pyFFTW/issues/40
import numpy as np
from sporco.dictlrn import cbpdndl
from sporco import util
from sporco import signal
from sporco import plot
plot.config_notebook_plotting()
```
Load training images.
```
exim = util.ExampleImages(scaled=True, zoom=0.25, gray=True)
S1 = exim.image('barbara.png', idxexp=np.s_[10:522, 100:612])
S2 = exim.image('kodim23.png', idxexp=np.s_[:, 60:572])
S3 = exim.image('monarch.png', idxexp=np.s_[:, 160:672])
S4 = exim.image('sail.png', idxexp=np.s_[:, 210:722])
S5 = exim.image('tulips.png', idxexp=np.s_[:, 30:542])
S = np.dstack((S1, S2, S3, S4, S5))
```
Highpass filter training images.
```
npd = 16
fltlmbd = 5
sl, sh = signal.tikhonov_filter(S, fltlmbd, npd)
```
Construct initial dictionary.
```
np.random.seed(12345)
D0 = np.random.randn(8, 8, 64)
```
Set regularization parameter and options for dictionary learning solver.
```
lmbda = 0.2
opt = cbpdndl.ConvBPDNDictLearn.Options({'Verbose': True, 'MaxMainIter': 200,
'CBPDN': {'rho': 50.0*lmbda + 0.5},
'CCMOD': {'rho': 10.0, 'ZeroMean': True}},
dmethod='cns')
```
Create solver object and solve.
```
d = cbpdndl.ConvBPDNDictLearn(D0, sh, lmbda, opt, dmethod='cns')
D1 = d.solve()
print("ConvBPDNDictLearn solve time: %.2fs" % d.timer.elapsed('solve'))
```
Display initial and final dictionaries.
```
D1 = D1.squeeze()
fig = plot.figure(figsize=(14, 7))
plot.subplot(1, 2, 1)
plot.imview(util.tiledict(D0), title='D0', fig=fig)
plot.subplot(1, 2, 2)
plot.imview(util.tiledict(D1), title='D1', fig=fig)
fig.show()
```
Get iterations statistics from solver object and plot functional value, ADMM primary and dual residuals, and automatically adjusted ADMM penalty parameter against the iteration number.
```
its = d.getitstat()
fig = plot.figure(figsize=(20, 5))
plot.subplot(1, 3, 1)
plot.plot(its.ObjFun, xlbl='Iterations', ylbl='Functional', fig=fig)
plot.subplot(1, 3, 2)
plot.plot(np.vstack((its.XPrRsdl, its.XDlRsdl, its.DPrRsdl,
its.DDlRsdl)).T, ptyp='semilogy', xlbl='Iterations',
ylbl='Residual', lgnd=['X Primal', 'X Dual', 'D Primal', 'D Dual'],
fig=fig)
plot.subplot(1, 3, 3)
plot.plot(np.vstack((its.XRho, its.DRho)).T, xlbl='Iterations',
ylbl='Penalty Parameter', ptyp='semilogy',
lgnd=['$\\rho_X$', '$\\rho_D$'], fig=fig)
fig.show()
```
| github_jupyter |
The code for plotting warming stripes is adapted from [Sebastian Beyer's github repository](https://github.com/sebastianbeyer/warmingstripes)
```
import pandas as pd
import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
%matplotlib inline
obs_file = '../data/raw/obs_temp_data/combined_temps_Jan_2019.csv'
models_file = '../data/raw/model_temp_data/Model timeseries.xlsx'
obs_df = pd.read_csv(obs_file)
obs_ds = obs_df.to_xarray().swap_dims({'index': 'year'}).groupby('year').mean()
obs_ds['mean'] = (obs_ds['hadcrut4'] + obs_ds['gistemp'] + obs_ds['noaa'] + obs_ds['berkeley'] + obs_ds['cowtan_way'])/5.
models_df = pd.read_excel(models_file, sheet_name = 'Individual papers')
models_df.columns = map(str.lower, models_df.columns)
models_ds = models_df.to_xarray().swap_dims({'index': 'year'})
vmin = -1.0
vmax = 1.0
timeslice = slice(1960,2017)
meanslice = slice(1978,1998)
plt.figure(figsize=(10,4.5))
plt.subplot(2,1,1)
temp_to_stack = (
obs_ds['mean'].sel(year=timeslice) -
obs_ds['mean'].sel(year=meanslice).mean(dim='year')
).values
stacked_temps = np.stack([temp_to_stack, temp_to_stack],axis=0)
img = plt.imshow(stacked_temps, cmap='RdBu_r', aspect=6, vmin=vmin, vmax=vmax)
plt.gca().set_axis_off()
plt.subplots_adjust(top = 1, bottom = 0, right = 1, left = 0, hspace = 0, wspace = 0)
plt.margins(0,0)
plt.gca().xaxis.set_major_locator(plt.NullLocator())
plt.gca().yaxis.set_major_locator(plt.NullLocator())
plt.plot([1988-1960,1988-1960], [-0.5,1.5],'--', color = 'grey')
plt.annotate(s='Temperature observations', xy=(0.01, 0.05),xycoords='axes fraction', fontsize = 14, color = 'black')
plt.subplot(2,1,2)
temp_to_stack = (
models_ds['hansen_1988_b_t'].sel(year=timeslice) -
models_ds['hansen_1988_b_t'].sel(year=meanslice).mean(dim='year')
).values
stacked_temps = np.stack([temp_to_stack, temp_to_stack],axis=0)
img = plt.imshow(stacked_temps, cmap='RdBu_r', aspect=6, vmin=vmin, vmax=vmax)
plt.gca().set_axis_off()
plt.subplots_adjust(top = 1, bottom = 0, right = 1, left = 0, hspace = 0, wspace = 0)
plt.margins(0,0)
plt.gca().xaxis.set_major_locator(plt.NullLocator())
plt.gca().yaxis.set_major_locator(plt.NullLocator())
plt.annotate(s='Hansen et al. (1988)\nEmissions Scenario B', xy=(0.01, 0.05),xycoords='axes fraction', fontsize = 14, color = 'black')
plt.plot([1988-1960,1988-1960], [-0.5,1.5],'--', color = 'grey')
plt.annotate(s='Publication date (1988)', xy=(1988-1960-1.2, 1.4),xycoords='data', fontsize = 12, color = 'black', rotation=90, alpha=0.7)
plt.savefig("extra_figures/projectionstripes.png", bbox_inches = 'tight', pad_inches = 0, dpi=400)
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#PyMOTW" data-toc-modified-id="PyMOTW-1"><span class="toc-item-num">1 </span>PyMOTW</a></span><ul class="toc-item"><li><span><a href="#Finding-Patterns-in-Text" data-toc-modified-id="Finding-Patterns-in-Text-1.1"><span class="toc-item-num">1.1 </span>Finding Patterns in Text</a></span></li><li><span><a href="#Compiling-Expressions" data-toc-modified-id="Compiling-Expressions-1.2"><span class="toc-item-num">1.2 </span>Compiling Expressions</a></span></li><li><span><a href="#Multiple-Matches" data-toc-modified-id="Multiple-Matches-1.3"><span class="toc-item-num">1.3 </span>Multiple Matches</a></span></li><li><span><a href="#Pattern-Syntax" data-toc-modified-id="Pattern-Syntax-1.4"><span class="toc-item-num">1.4 </span>Pattern Syntax</a></span><ul class="toc-item"><li><span><a href="#Repetition" data-toc-modified-id="Repetition-1.4.1"><span class="toc-item-num">1.4.1 </span>Repetition</a></span></li><li><span><a href="#Character-Sets" data-toc-modified-id="Character-Sets-1.4.2"><span class="toc-item-num">1.4.2 </span>Character Sets</a></span></li><li><span><a href="#Anchoring" data-toc-modified-id="Anchoring-1.4.3"><span class="toc-item-num">1.4.3 </span>Anchoring</a></span></li></ul></li><li><span><a href="#Constraining-the-Search" data-toc-modified-id="Constraining-the-Search-1.5"><span class="toc-item-num">1.5 </span>Constraining the Search</a></span></li><li><span><a href="#Dissecting-Matches-with-Groups" data-toc-modified-id="Dissecting-Matches-with-Groups-1.6"><span class="toc-item-num">1.6 </span>Dissecting Matches with Groups</a></span></li><li><span><a href="#Search-Options" data-toc-modified-id="Search-Options-1.7"><span class="toc-item-num">1.7 </span>Search Options</a></span><ul class="toc-item"><li><span><a href="#Case-insensitive-Matching" data-toc-modified-id="Case-insensitive-Matching-1.7.1"><span class="toc-item-num">1.7.1 </span>Case-insensitive Matching</a></span></li><li><span><a href="#Input-with-Multiple-Lines" data-toc-modified-id="Input-with-Multiple-Lines-1.7.2"><span class="toc-item-num">1.7.2 </span>Input with Multiple Lines</a></span></li><li><span><a href="#Unicode" data-toc-modified-id="Unicode-1.7.3"><span class="toc-item-num">1.7.3 </span>Unicode</a></span></li><li><span><a href="#Verbose-Expression-Syntax" data-toc-modified-id="Verbose-Expression-Syntax-1.7.4"><span class="toc-item-num">1.7.4 </span>Verbose Expression Syntax</a></span></li><li><span><a href="#Embedding-Flags-in-Patterns" data-toc-modified-id="Embedding-Flags-in-Patterns-1.7.5"><span class="toc-item-num">1.7.5 </span>Embedding Flags in Patterns</a></span></li></ul></li><li><span><a href="#Looking-Ahead,-or-behind" data-toc-modified-id="Looking-Ahead,-or-behind-1.8"><span class="toc-item-num">1.8 </span>Looking Ahead, or behind</a></span></li><li><span><a href="#Self-referencing-Expressions" data-toc-modified-id="Self-referencing-Expressions-1.9"><span class="toc-item-num">1.9 </span>Self-referencing Expressions</a></span></li><li><span><a href="#Modifying-Strings-with-Patterns" data-toc-modified-id="Modifying-Strings-with-Patterns-1.10"><span class="toc-item-num">1.10 </span>Modifying Strings with Patterns</a></span></li><li><span><a href="#Splitting-with-Patterns" data-toc-modified-id="Splitting-with-Patterns-1.11"><span class="toc-item-num">1.11 </span>Splitting with Patterns</a></span></li></ul></li><li><span><a href="#笔记" data-toc-modified-id="笔记-2"><span class="toc-item-num">2 </span>笔记</a></span><ul class="toc-item"><li><span><a href="#RE" data-toc-modified-id="RE-2.1"><span class="toc-item-num">2.1 </span><code>RE</code></a></span></li><li><span><a href="#正则要义" data-toc-modified-id="正则要义-2.2"><span class="toc-item-num">2.2 </span>正则要义</a></span></li></ul></li></ul></div>
# PyMOTW
[re – Regular Expressions - Python Module of the Week](https://pymotw.com/2/re/)
## Finding Patterns in Text
```
import re
patterns = [ 'this', 'that' ]
text = 'Does this text match the pattern?'
for pattern in patterns:
print ('Looking for "%s" in "%s" ->' % (pattern, text)),
if re.search(pattern, text):
print ('found a match!')
else:
print ('no match')
import re
pattern = 'this'
text = 'Does this text match the pattern?'
match = re.search(pattern, text)
s = match.start()
e = match.end()
print ('Found "%s" in "%s" from %d to %d ("%s")' % \
(match.re.pattern, match.string, s, e, text[s:e]))
```
## Compiling Expressions
```
import re
# Pre-compile the patterns
regexes = [ re.compile(p) for p in [ 'this',
'that',
]
]
text = 'Does this text match the pattern?'
for regex in regexes:
print ('Looking for "%s" in "%s" ->' % (regex.pattern, text)),
if regex.search(text):
print ('found a match!')
else:
print ('no match')
```
## Multiple Matches
```
import re
text = 'abbaaabbbbaaaaa'
pattern = 'ab'
for match in re.findall(pattern, text):
print ('Found "%s"' % match)
import re
text = 'abbaaabbbbaaaaa'
pattern = 'ab'
for match in re.finditer(pattern, text):
s = match.start()
e = match.end()
print ('Found "%s" at %d:%d' % (text[s:e], s, e))
```
## Pattern Syntax
```
import re
def test_patterns(text, patterns=[]):
"""Given source text and a list of patterns, look for
matches for each pattern within the text and print
them to stdout.
"""
# Show the character positions and input text
print
print (''.join(str(i/10 or ' ') for i in range(len(text))))
print (''.join(str(i%10) for i in range(len(text))))
print (text)
# Look for each pattern in the text and print the results
for pattern in patterns:
print
print ('Matching "%s"' % pattern)
for match in re.finditer(pattern, text):
s = match.start()
e = match.end()
print (' %2d : %2d = "%s"' % \
(s, e-1, text[s:e]))
return
if __name__ == '__main__':
test_patterns('abbaaabbbbaaaaa', ['ab'])
```
### Repetition
```
test_patterns('abbaaabbbbaaaaa',
[ 'ab*', # a followed by zero or more b
'ab+', # a followed by one or more b
'ab?', # a followed by zero or one b
'ab{3}', # a followed by three b
'ab{2,3}', # a followed by two to three b
])
test_patterns('abbaaabbbbaaaaa',
[ 'ab*?', # a followed by zero or more b
'ab+?', # a followed by one or more b
'ab??', # a followed by zero or one b
'ab{3}?', # a followed by three b
'ab{2,3}?', # a followed by two to three b
])
```
### Character Sets
```
test_patterns('abbaaabbbbaaaaa',
[ '[ab]', # either a or b
'a[ab]+', # a followed by one or more a or b
'a[ab]+?', # a followed by one or more a or b, not greedy
])
test_patterns('This is some text -- with punctuation.',
[ '[^-. ]+', # sequences without -, ., or space
])
test_patterns('This is some text -- with punctuation.',
[ '[a-z]+', # sequences of lower case letters
'[A-Z]+', # sequences of upper case letters
'[a-zA-Z]+', # sequences of lower or upper case letters
'[A-Z][a-z]+', # one upper case letter followed by lower case letters
])
test_patterns('abbaaabbbbaaaaa',
[ 'a.', # a followed by any one character
'b.', # b followed by any one character
'a.*b', # a followed by anything, ending in b
'a.*?b', # a followed by anything, ending in b
])
test_patterns('This is a prime #1 example!',
[ r'\d+', # sequence of digits
r'\D+', # sequence of non-digits
r'\s+', # sequence of whitespace
r'\S+', # sequence of non-whitespace
r'\w+', # alphanumeric characters
r'\W+', # non-alphanumeric
])
test_patterns(r'\d+ \D+ \s+ \S+ \w+ \W+',
[ r'\\d\+',
r'\\D\+',
r'\\s\+',
r'\\S\+',
r'\\w\+',
r'\\W\+',
])
```
### Anchoring
```
test_patterns('This is some text -- with punctuation.',
[ r'^\w+', # word at start of string
r'\A\w+', # word at start of string
r'\w+\S*$', # word at end of string, with optional punctuation
r'\w+\S*\Z', # word at end of string, with optional punctuation
r'\w*t\w*', # word containing 't'
r'\bt\w+', # 't' at start of word
r'\w+t\b', # 't' at end of word
r'\Bt\B', # 't', not start or end of word
])
```
## Constraining the Search
```
text = 'This is some text -- with punctuation.'
pattern = 'is'
print ('Text :', text)
print ('Pattern:', pattern)
m = re.match(pattern, text)
print ('Match :', m)
s = re.search(pattern, text)
print ('Search :', s)
text = 'This is some text -- with punctuation.'
pattern = re.compile(r'\b\w*is\w*\b')
print ('Text:', text)
print
pos = 0
while True:
match = pattern.search(text, pos)
if not match:
break
s = match.start()
e = match.end()
print (' %2d : %2d = "%s"' % \
(s, e-1, text[s:e]))
# Move forward in text for the next search
pos = e
```
## Dissecting Matches with Groups
```
test_patterns('abbaaabbbbaaaaa',
[ 'a(ab)', # 'a' followed by literal 'ab'
'a(a*b*)', # 'a' followed by 0-n 'a' and 0-n 'b'
'a(ab)*', # 'a' followed by 0-n 'ab'
'a(ab)+', # 'a' followed by 1-n 'ab'
])
text = 'This is some text -- with punctuation.'
print (text)
print
for pattern in [ r'^(\w+)', # word at start of string
r'(\w+)\S*$', # word at end of string, with optional punctuation
r'(\bt\w+)\W+(\w+)', # word starting with 't' then another word
r'(\w+t)\b', # word ending with 't'
]:
regex = re.compile(pattern)
match = regex.search(text)
print ('Matching "%s"' % pattern)
print (' ', match.groups())
print
text = 'This is some text -- with punctuation.'
print ('Input text :', text)
# word starting with 't' then another word
regex = re.compile(r'(\bt\w+)\W+(\w+)')
print ('Pattern :', regex.pattern)
match = regex.search(text)
print ('Entire match :', match.group(0))
print ('Word starting with "t":', match.group(1))
print ('Word after "t" word :', match.group(2))
print (match.groups())
text = 'This is some text -- with punctuation.'
print (text)
print
for pattern in [ r'^(?P<first_word>\w+)',
r'(?P<last_word>\w+)\S*$',
r'(?P<t_word>\bt\w+)\W+(?P<other_word>\w+)',
r'(?P<ends_with_t>\w+t)\b',
]:
regex = re.compile(pattern)
match = regex.search(text)
print ('Matching "%s"' % pattern)
print (' ', match.groups())
print (' ', match.groupdict())
print
def test_patterns(text, patterns=[]):
"""Given source text and a list of patterns, look for
matches for each pattern within the text and print
them to stdout.
"""
# Show the character positions and input text
print
print (''.join(str(i/10 or ' ') for i in range(len(text))))
print (''.join(str(i%10) for i in range(len(text))))
print (text)
# Look for each pattern in the text and print the results
for pattern in patterns:
print
print ('Matching "%s"' % pattern)
for match in re.finditer(pattern, text):
s = match.start()
e = match.end()
print (' %2d : %2d = "%s"' % \
(s, e-1, text[s:e]))
print (' Groups:', match.groups())
if match.groupdict():
print (' Named groups:', match.groupdict())
print
return
test_patterns('abbaaabbbbaaaaa',
[r'a((a*)(b*))', # 'a' followed by 0-n 'a' and 0-n 'b'
])
re.findall(r'a((a*)(b*))',t)
re.findall(r'b(a)',t)
test_patterns('abbaaabbbbaaaaa',
[r'a((a+)|(b+))', # 'a' followed by a sequence of 'a' or sequence of 'b'
r'a((a|b)+)', # 'a' followed by a sequence of 'a' or 'b'
])
test_patterns('abbaaabbbbaaaaa',
[r'a((a+)|(b+))', # capturing form
r'a((?:a+)|(?:b+))', # non-capturing
])
```
## Search Options
### Case-insensitive Matching
```
text = 'This is some text -- with punctuation.'
pattern = r'\bT\w+'
with_case = re.compile(pattern)
without_case = re.compile(pattern, re.IGNORECASE)
print ('Text :', text)
print ('Pattern :', pattern)
print ('Case-sensitive :', with_case.findall(text))
print ('Case-insensitive:', without_case.findall(text))
```
### Input with Multiple Lines
```
text = 'This is some text -- with punctuation.\nAnd a second line.'
pattern = r'(^\w+)|(\w+\S*$)'
single_line = re.compile(pattern)
multiline = re.compile(pattern, re.MULTILINE)
print ('Text :', repr(text))
print ('Pattern :', pattern)
print ('Single Line :', single_line.findall(text))
print ('Multline :', multiline.findall(text))
text = 'This is some text -- with punctuation.\nAnd a second line.'
pattern = r'.+'
no_newlines = re.compile(pattern)
dotall = re.compile(pattern, re.DOTALL)
# print 'Text :', repr(text)
# print 'Pattern :', pattern
print ('No newlines :', no_newlines.findall(text))
print ('Dotall :', dotall.findall(text))
```
### Unicode
Python3 不需要,因为 Python3 默认对所有的 string 使用 Unicode 编码
```
import re
import codecs
import sys
# Python2
# Set standard output encoding to UTF-8.
sys.stdout = codecs.getwriter('UTF-8')(sys.stdout)
text = u'Français złoty Österreich'
pattern = ur'\w+'
ascii_pattern = re.compile(pattern)
unicode_pattern = re.compile(pattern, re.UNICODE)
print ('Text :', text)
print ('Pattern :', pattern)
print ('ASCII :', u', '.join(ascii_pattern.findall(text)))
print ('Unicode :', u', '.join(unicode_pattern.findall(text)))
```
### Verbose Expression Syntax
```
import re
address = re.compile('[\w\d.+-]+@([\w\d.]+\.)+(com|org|edu)', re.UNICODE)
candidates = [
u'first.last@example.com',
u'first.last+category@gmail.com',
u'valid-address@mail.example.com',
u'not-valid@example.foo',
]
for candidate in candidates:
print
print ('Candidate:', candidate)
match = address.search(candidate)
if match:
print (' Matches')
else:
print (' No match')
# 方括号里的就是元素本身,不代表数量
address = re.compile(
'''
[\w\d.+-]+ # username
@
([\w\d.]+\.)+ # domain name prefix
(com|org|edu) # we should support more top-level domains
''',
re.UNICODE | re.VERBOSE)
candidates = [
u'first.last@example.com',
u'first.last+category@gmail.com',
u'valid-address@mail.example.com',
u'not-valid@example.foo',
]
for candidate in candidates:
print
print ('Candidate:', candidate)
match = address.search(candidate)
if match:
print (' Matches')
else:
print (' No match')
address = re.compile(
'''
# A name is made up of letters, and may include "." for title
# abbreviations and middle initials.
((?P<name>
([\w.,]+\s+)*[\w.,]+)
\s*
# Email addresses are wrapped in angle brackets: < >
# but we only want one if we found a name, so keep
# the start bracket in this group.
<
)? # the entire name is optional
# The address itself: username@domain.tld
(?P<email>
[\w\d.+-]+ # username
@
([\w\d.]+\.)+ # domain name prefix
(com|org|edu) # limit the allowed top-level domains
)
>? # optional closing angle bracket
''',
re.UNICODE | re.VERBOSE)
candidates = [
u'first.last@example.com',
u'first.last+category@gmail.com',
u'valid-address@mail.example.com',
u'not-valid@example.foo',
u'First Last <first.last@example.com>',
u'No Brackets first.last@example.com',
u'First Last',
u'First Middle Last <first.last@example.com>',
u'First M. Last <first.last@example.com>',
u'<first.last@example.com>',
]
for candidate in candidates:
print
print ('Candidate:', candidate)
match = address.search(candidate)
if match:
print (' Match name :', match.groupdict()['name'])
print (' Match email:', match.groupdict()['email'])
else:
print (' No match')
```
### Embedding Flags in Patterns
```
text = 'This is some text -- with punctuation.'
pattern = r'(?i)\bT\w+'
regex = re.compile(pattern)
print ('Text :', text)
print ('Pattern :', pattern)
print ('Matches :', regex.findall(text))
text = 'This is some text -- with punctuation.'
pattern = r'(?i)\bT\w+'
regex = re.compile(pattern)
print ('Text :', text)
print ('Pattern :', pattern)
print ('Matches :', regex.findall(text))
```
## Looking Ahead, or behind
```
address = re.compile(
'''
# A name is made up of letters, and may include "." for title
# abbreviations and middle initials.
((?P<name>
([\w.,]+\s+)*[\w.,]+
)
\s+
) # name is no longer optional
# LOOKAHEAD
# Email addresses are wrapped in angle brackets, but we only want
# the brackets if they are both there, or neither are.
(?= (<.*>$) # remainder wrapped in angle brackets
|
([^<].*[^>]$) # remainder *not* wrapped in angle brackets
)
<? # optional opening angle bracket
# The address itself: username@domain.tld
(?P<email>
[\w\d.+-]+ # username
@
([\w\d.]+\.)+ # domain name prefix
(com|org|edu) # limit the allowed top-level domains
)
>? # optional closing angle bracket
''',
re.UNICODE | re.VERBOSE)
candidates = [
u'First Last <first.last@example.com>',
u'No Brackets first.last@example.com',
u'Open Bracket <first.last@example.com',
u'Close Bracket first.last@example.com>',
]
for candidate in candidates:
print
print ('Candidate:', candidate)
match = address.search(candidate)
if match:
print (' Match name :', match.groupdict()['name'])
print (' Match email:', match.groupdict()['email'])
else:
print (' No match')
address = re.compile(
'''
^
# An address: username@domain.tld
# Ignore noreply addresses
(?!noreply@.*$)
[\w\d.+-]+ # username
@
([\w\d.]+\.)+ # domain name prefix
(com|org|edu) # limit the allowed top-level domains
$
''',
re.UNICODE | re.VERBOSE)
candidates = [
u'first.last@example.com',
u'noreply@example.com',
]
for candidate in candidates:
print
print ('Candidate:', candidate)
match = address.search(candidate)
if match:
print (' Match:', candidate[match.start():match.end()])
else:
print (' No match')
address = re.compile(
'''
^
# An address: username@domain.tld
[\w\d.+-]+ # username
# Ignore noreply addresses
(?<!noreply)
@
([\w\d.]+\.)+ # domain name prefix
(com|org|edu) # limit the allowed top-level domains
$
''',
re.UNICODE | re.VERBOSE)
candidates = [
u'first.last@example.com',
u'noreply@example.com',
]
for candidate in candidates:
print
print ('Candidate:', candidate)
match = address.search(candidate)
if match:
print (' Match:', candidate[match.start():match.end()])
else:
print (' No match')
import re
twitter = re.compile(
'''
# A twitter handle: @username
(?<=@)
([\w\d_]+) # username
''',
re.UNICODE | re.VERBOSE)
text = '''This text includes two Twitter handles.
One for @ThePSF, and one for the author, @doughellmann.
'''
print (text)
for match in twitter.findall(text):
print ('Handle:', match)
import re
twitter = re.compile(
'''
# A twitter handle: @username
# 注意,改成「=」不对
(?=@)
([\w\d_]+) # username
''',
re.UNICODE | re.VERBOSE)
text = '''This text includes two Twitter handles.
One for @ThePSF, and one for the author, @doughellmann.
'''
print (text)
for match in twitter.findall(text):
print ('Handle:', match)
```
## Self-referencing Expressions
```
address = re.compile(
r'''
# The regular name
(\w+) # first name
\s+
(([\w.]+)\s+)? # optional middle name or initial
(\w+) # last name
\s+
<
# The address: first_name.last_name@domain.tld
(?P<email>
\1 # first name
\.
\4 # last name
@
([\w\d.]+\.)+ # domain name prefix
(com|org|edu) # limit the allowed top-level domains
)
>
''',
re.UNICODE | re.VERBOSE | re.IGNORECASE)
candidates = [
u'First Last <first.last@example.com>',
u'Different Name <first.last@example.com>',
u'First Middle Last <first.last@example.com>',
u'First M. Last <first.last@example.com>',
]
for candidate in candidates:
print
print ('Candidate:', candidate)
match = address.search(candidate)
if match:
print (' Match name :', match.group(1), match.group(4))
print (' Match email:', match.group(5))
else:
print (' No match')
address = re.compile(
'''
# The regular name
(?P<first_name>\w+)
\s+
(([\w.]+)\s+)? # optional middle name or initial
(?P<last_name>\w+)
\s+
<
# The address: first_name.last_name@domain.tld
(?P<email>
(?P=first_name)
\.
(?P=last_name)
@
([\w\d.]+\.)+ # domain name prefix
(com|org|edu) # limit the allowed top-level domains
)
>
''',
re.UNICODE | re.VERBOSE | re.IGNORECASE)
candidates = [
u'First Last <first.last@example.com>',
u'Different Name <first.last@example.com>',
u'First Middle Last <first.last@example.com>',
u'First M. Last <first.last@example.com>',
]
for candidate in candidates:
print
print ('Candidate:', candidate)
match = address.search(candidate)
if match:
print (' Match name :', match.groupdict()['first_name'], match.groupdict()['last_name'])
print (' Match email:', match.groupdict()['email'])
else:
print (' No match')
address = re.compile(
'''
^
# A name is made up of letters, and may include "." for title
# abbreviations and middle initials.
(?P<name>
([\w.]+\s+)*[\w.]+
)?
\s*
# Email addresses are wrapped in angle brackets, but we only want
# the brackets if we found a name.
(?(name)
# remainder wrapped in angle brackets because we have a name
(?P<brackets>(?=(<.*>$)))
|
# remainder does not include angle brackets without name
(?=([^<].*[^>]$))
)
# Only look for a bracket if our look ahead assertion found both
# of them.
(?(brackets)<|\s*)
# The address itself: username@domain.tld
(?P<email>
[\w\d.+-]+ # username
@
([\w\d.]+\.)+ # domain name prefix
(com|org|edu) # limit the allowed top-level domains
)
# Only look for a bracket if our look ahead assertion found both
# of them.
(?(brackets)>|\s*)
$
''',
re.UNICODE | re.VERBOSE)
candidates = [
u'First Last <first.last@example.com>',
u'No Brackets first.last@example.com',
u'Open Bracket <first.last@example.com',
u'Close Bracket first.last@example.com>',
u'no.brackets@example.com',
]
for candidate in candidates:
print
print ('Candidate:', candidate)
match = address.search(candidate)
if match:
print (' Match name :', match.groupdict()['name'])
print (' Match email:', match.groupdict()['email'])
else:
print (' No match')
address = re.compile(
'''
# 注意,目前这种情无法匹配到 u'First Last <first.last@example.com>',因为 '<'
^(?P<name>\w+\s\w+\s)
(?(name)(?=(\w+))|(?=([^<].*[^>]$)))
''',re.UNICODE | re.VERBOSE)
candidates = [
u'First Last <first.last@example.com>',
u'No Brackets first.last@example.com',
u'Open Bracket <first.last@example.com',
u'Close Bracket first.last@example.com>',
u'fdf no.brackets@example.com',
]
for candidate in candidates:
print
print ('Candidate:', candidate)
match = address.search(candidate)
if match:
print (' Match name :', match.groupdict()['name'])
else:
print (' No match')
```
## Modifying Strings with Patterns
```
# ?表示非贪婪匹配
bold = re.compile(r'\*{2}(.*?)\*{2}', re.UNICODE)
text = 'Make this **bold**. This **too**.'
print ('Text:', text)
print ('Bold:', bold.sub(r'<b>\1</b>', text))
# 可以用名字
bold = re.compile(r'\*{2}(?P<bold_text>.*?)\*{2}', re.UNICODE)
text = 'Make this **bold**. This **too**.'
print ('Text:', text)
print ('Bold:', bold.sub(r'<b>\g<bold_text></b>', text))
bold = re.compile(r'\*{2}(.*?)\*{2}', re.UNICODE)
text = 'Make this **bold**. This **too**.'
print ('Text:', text)
print ('Bold:', bold.subn(r'<b>\1</b>', text))
```
## Splitting with Patterns
```
text = 'Paragraph one\non two lines.\n\nParagraph two.\n\n\nParagraph three.'
for num, para in enumerate(re.findall(r'(.+?)\n{2,}', text, flags=re.DOTALL)):
print (num, repr(para))
print
text = 'Paragraph one\non two lines.\n\nParagraph two.\n\n\nParagraph three.'
print ('With findall:')
for num, para in enumerate(re.findall(r'(.+?)(\n{2,}|$)', text, flags=re.DOTALL)):
print (num, repr(para))
print
print
print ('With split:')
for num, para in enumerate(re.split(r'\n{2,}', text)):
print (num, repr(para))
print
text = 'Paragraph one\non two lines.\n\nParagraph two.\n\n\nParagraph three.'
print
print ('With split:')
for num, para in enumerate(re.split(r'(\n{2,})', text)):
print (num, repr(para))
print
```
# 笔记
说明:例子可以直接从 1.8 开始看
## `RE`
- `re.compile()`
- compile 常用的表达式更高效
- 使用编译的表达式能避免高速缓存查找开销,将编译工作转移到应用程序启动时,而不是程序响应用户操作
- `re.compile(pattern, re.IGNORECASE)`: IGNORECASE 忽略大小写
- `re.compile(pattern, re.MULTILINE)`: 是否按行匹配,带参数按行,不带参数整个字符串为整体(不考虑换行符)
- `re.compile(pattern, re.DOTALL)`: 默认不匹配换行符,带参数匹配换行符
- `re.compile(pattern, re.Unicode | re.VERBOSE)`: 让 pattern 看起来更易读
- `< and >`: angle brackets,`(?P<pattern_name>pattern)`,给 pattern 命名
- 嵌入标志(编译表达式无法添加标志时,可以在表达式字符串本身内嵌入标志)
- 打开不区分大小写匹配项: `(?i)` 添加到 pattern 的最前面就好
- `i: IGNORECASE; m: MULTILINE; s: DOTALL; u: UNICODE; x: VERBOSE`
- `?:...`: 无捕获组
- `(?(id)yes-expression|no-expression)`, id 是 group name or number
- `yes-expression` is the pattern to use if the group has a value,如果 group 有返回值时执行
- `no-expression` is the pattern to use otherwise.
- `(?pattern)`
- `(?!pattern)`: 不匹配 pattern
- `(?<!noreply)` 放在匹配到的 username 下和 `(?!noreply@.*$)` 放在最前面匹配是一样的
- `(?<=@)`: 等于 @
- `re.search()`
- 使用 pattern 扫描文本
- 如果找到则返回 Match 对象,否则返回 `None`
- `match.start(), match.end()` 返回匹配到对象的起止位置
- **`pattern.search()`**
- 返回子串,而不是 `re.search()` 的整个字符串
- `re.match()`
- 只有 pattern 在文本的最开始才返回结果,否则返回 `None`
- `re.findall()`
- `finditer` 返回一个迭代器,可以使用 `match.start(), match.end()` 返回起止位置
- `groups()`
- **返回匹配到的字符序列**,不需要再用起止位置确定返回的字符串
- `group(index)`: 返回单独的一个 group(匹配到的结果)
- `(?P<name>pattern)`: 给一个 group 命名,`match.groupdict()` 返回 {名字:值} 的字典
- 用「行数」引用或用 `(?P=pattern_name)` 引用
- `re.findall(r'a((a*)(b*))',t)`: 类似这种,第一个 a 是不返回到结果的,只返回括号里面的
- `re.sub()`
- `\g<name>` 语法也适用于编号引用,并且使用它可以消除组号和周围文字数字之间的任何歧义。
- `bold = re.compile(r'\*{2}(?P<bold_text>.*?)\*{2}', re.UNICODE)`,替换时,中间那块不变
- `bold.sub(r'<b>\g<bold_text></b>', text)`: `<bold_text>` 是表达式名字
- `bold = re.compile(r'\*{2}(.*?)\*{2}', re.UNICODE)`
- `bold.sub(r'<b>\1</b>', text)`: `bold` 是正则表达式,`<b>\1</b>` 表示要替换的正则内容,`\1` 表示匹配时的中间部分**不变**
- `subn()` 返回修改后的字符串和替换的数量
- `re.split()`
- `re.split(r'(\n{2,})', text)`: 返回分隔符和分割后的结果
- `re.split(r'\n{2,}', text)`: 返回分割后的结果
## 正则要义
- 数量
- {} 多少次
- \* 等于 {0,},0 次或任意正次数
- \+ 等于 {1,},1 次或任意正次数
- ? 等于 {0,1},0 次或 1 次
- pattern 后面加「?」(英文的问号)就是非贪婪匹配,默认是贪婪匹配(匹配尽可能多的字符)
- 也就是匹配数量少的次数(也就是 {} 左边的数字次数)
- 转义
- 范围
- `[ab]`: a or b,`[\w\d.+-]` 里面的「.+-」就是代表自身
- `a[ab]+`: aa... or ab...
- `a[ab]+?`: aa or ab
- `[^-. ]+`: 不含 `-, ., 空格` 的
- `[A-Z][a-z]+`: 一个大写字母后面跟着小写字母
- 小数点
- `.` 表示任何单独的字符,除了换行符(\n)
- `a.*b`: a 后面任意字符,b 结尾
- `a.*?b`: a 后面任意字符,b 结尾,非贪婪
- 反斜杠
- `\d`: a digit
- `\D`: a non-digit
- `\s`: whitespace(tab, space, newline, etc.)
- `\S`: non-whitespace
- `\w`: alphanumeric 字母数字
- `\W`: non-alphanumeric 非字母数字(标点,符号如#等)
- `\`: 转义
- 位置
- `^`: string or line 开始
- `$`: string or line 结尾
- `\A`: string 开始
- `\Z`: string 结尾
- `\b`: empty string at the beginning or end of a word
- `\B`: empty string not at the beginning or end of a word
- 例子
- `r'^\w+', # word at start of string`
- `r'\A\w+', # word at start of string`
- `r'\w+\S*$', # word at end of string, with optional punctuation`
- `r'\w+\S*\Z', # word at end of string, with optional punctuation`
- `r'\w*t\w*', # word containing 't'`
- `r'\bt\w+', # 't' at start of word`
- `r'\w+t\b', # 't' at end of word`
- `r'\Bt\B', # 't', not start or end of word`
- 逻辑
- `()`: 分组,视为整体
- `|`: 管道符,要么用左边的字符串匹配,要么用右边的匹配
| github_jupyter |
# A Cantera Simulation Using RMG-Py
```
from IPython.display import display, Image
from rmgpy.chemkin import load_chemkin_file
from rmgpy.tools.canteraModel import Cantera, get_rmg_species_from_user_species
from rmgpy.species import Species
```
Load the species and reaction from the RMG-generated chemkin file `chem_annotated.inp` and `species_dictionary.txt` file found in your `chemkin` folder after running a job.
```
species_list, reaction_list = load_chemkin_file('data/ethane_model/chem_annotated.inp',
'data/ethane_model/species_dictionary.txt',
'data/ethane_model/tran.dat')
```
Set a few conditions for how to react the system
```
# Find the species: ethane
user_ethane=Species().from_smiles('CC')
species_dict = get_rmg_species_from_user_species([user_ethane], species_list)
ethane = species_dict[user_ethane]
reactor_type_list = ['IdealGasReactor']
mol_frac_list=[{ethane: 1}]
Tlist = ([1300, 1500, 2000], 'K')
Plist = ([1], 'bar')
reaction_time_list = ([0.5], 'ms')
# Create cantera object, loading in the species and reactions
job = Cantera(species_list=species_list, reaction_list=reaction_list, output_directory='temp')
# The cantera file must be created from an associated chemkin file
# We can either load the Model from the initialized set of rmg species and reactions
job.load_model()
# Or load it from a chemkin file by uncommenting the following line:
#job.load_chemkin_model('data/ethane_model/chem_annotated.inp',transport_file='data/ethane_model/tran.dat')
# Generate the conditions based on the settings we declared earlier
job.generate_conditions(reactor_type_list, reaction_time_list, mol_frac_list, Tlist, Plist)
# Simulate and plot
alldata = job.simulate()
job.plot(alldata)
# Show the plots in the ipython notebook
for i, condition in enumerate(job.conditions):
print('Condition {0}'.format(i+1))
display(Image(filename="temp/{0}_mole_fractions.png".format(i+1)))
# We can get the cantera model Solution's species and reactions
ct_species = job.model.species()
ct_reactions = job.model.reactions()
# We can view a cantera species or reaction object from this
ct_ethane = ct_species[4]
ct_rxn = ct_reactions[0]
print(ct_ethane)
print(ct_rxn)
# We can also do things like modifying the cantera species thermo and reaction kinetics through modifying the
# RMG objects first, then using the `modifyReactionKinetics` or `modifySpeciesThermo` functions
# Alter the RMG objects in place, lets pick ethane and the first reaction
rmg_ethane = species_dict[user_ethane]
rmg_ethane.thermo.change_base_enthalpy(2*4184) # Change base enthalpy by 2 kcal/mol
rmg_rxn = reaction_list[0]
rmg_rxn.kinetics.change_rate(4) # Change A factor by multiplying by a factor of 4
# Take a look at the state of the cantera model before and after
print('Cantera Model: Before')
ct_species = job.model.species()
ct_reactions = job.model.reactions()
print('Ethane Thermo = {} kcal/mol'.format(ct_species[4].thermo.h(300)/1000/4184))
print('Reaction 1 Kinetics = {}'.format(ct_reactions[0].rate))
# Now use the altered RMG objects to modify the kinetics and thermo
job.modify_reaction_kinetics(0, rmg_rxn)
job.modify_species_thermo(4, rmg_ethane, use_chemkin_identifier = True)
# If we modify thermo, the cantera model must be refreshed. If only kinetics are modified, this does not need to be done.
job.refresh_model()
print('')
print('Cantera Model: After')
ct_species = job.model.species()
ct_reactions = job.model.reactions()
print('Ethane Thermo = {} kcal/mol'.format(ct_species[4].thermo.h(300)/1000/4184))
print('Reaction 1 Kinetics = {}'.format(ct_reactions[0].rate))
# Simulate and plot
alldata = job.simulate()
job.plot(alldata)
# Show the plots in the ipython notebook
for i, condition in enumerate(job.conditions):
print('Condition {0}'.format(i+1))
display(Image(filename="temp/{0}_mole_fractions.png".format(i+1)))
```
| github_jupyter |
<div align="right"><i>Peter Norvig<br>April 2015<br>Python 3: Feb 2019</i></div>
# When is Cheryl's Birthday?
[This puzzle](https://www.google.com/webhp?#q=cheryl%27s%20birthday) has been making the rounds:
> 1. Albert and Bernard became friends with Cheryl, and want to know when her birthday is. Cheryl gave them a list of 10 possible dates:
May 15 May 16 May 19
June 17 June 18
July 14 July 16
August 14 August 15 August 17
> 2. Cheryl then tells Albert and Bernard separately the month and the day of the birthday respectively.
> 3. **Albert**: "I don't know when Cheryl's birthday is, and I know that Bernard does not know."
> 4. **Bernard**: "At first I don't know when Cheryl's birthday is, but I know now."
> 5. **Albert**: "Then I also know when Cheryl's birthday is."
> 6. So when is Cheryl's birthday?
Let's work through this puzzle statement by statement.
## 1. Cheryl gave them a list of 10 possible dates:
```
dates = ['May 15', 'May 16', 'May 19',
'June 17', 'June 18',
'July 14', 'July 16',
'August 14', 'August 15', 'August 17']
```
We'll define accessor functions for the month and day of a date:
```
def Month(date): return date.split()[0]
def Day(date): return date.split()[1]
Month('May 15')
Day('May 15')
```
## 2. Cheryl then tells Albert and Bernard separately the month and the day of the birthday respectively.
Now we have to think about what we're doing. We'll use a *set of dates* to represent a *belief set*: a person who has the belief set `{'August 15', 'May 15'}` *believes* that Cheryl's birthday is one of those two days. A person *knows* the birthdate when they get down to a belief set with only one possibility.
We can define the idea of Cheryl **telling** someone a component of her birthdate, and while we're at it, the idea of **knowing** a birthdate:
```
BeliefSet = set
def tell(part, dates=dates) -> BeliefSet:
"Cheryl tells a part of her birthdate to someone; return a set of possible dates."
return {date for date in dates if part in date}
def know(beliefs) -> bool:
"A person `knows` the answer if their belief set has only one possibility."
return len(beliefs) == 1
```
For example: If Cheryl tells Albert that her birthday is in May, he would know there is a set of three possible birthdates:
```
tell('May')
```
And if she tells Bernard that her birthday is on the 15th, he would know there are two possibilities:
```
tell('15')
```
With two possibilities, Bernard does not know the birthdate:
```
know(tell('15'))
```
## Overall Strategy
If Cheryl tells Albert `'May'` then *he* knows there are three possibilities, but *we* (the puzzle solvers) don't know that, because we don't know what Cheryl said.
So what can we do? We can consider *all* of the possible dates, one at a time. For example, first consider `'May 15'`. Cheryl tells Albert `'May'` and Bernard `'15'`, giving them the lists of possible birthdates shown above. We can then check whether statements 3 through 5 are true in this scenario. If they are, then `'May 15'` is a solution to the puzzle. Repeat the process for each of the other possible dates. If all goes well, there should be exactly one date for which all the statements are true.
Here is the main function, `cheryls_birthday`, which takes a set of possible dates, and returns the subset of dates that satisfy statements 3 through 5. The function `satisfy` is similar to the builtin function `filter`: `satisfy` takes a collection of items (here a set of dates) and returns the subset that satisfies all the predicates:
```
def cheryls_birthday(dates=dates) -> BeliefSet:
"Return a subset of the dates for which all three statements are true."
return satisfy(dates, statement3, statement4, statement5)
def satisfy(items, *predicates):
"Return the subset of items that satisfy all the predicates."
return {item for item in items
if all(pred(item) for pred in predicates)}
## TO DO: define statement3, statement4, statement5
```
## 3. Albert: I don't know when Cheryl's birthday is, and I know that Bernard does not know.
The function `statement3` corresponds to the third statement in the problem. It takes as input a single possible birthdate (not a set) and returns `True` if Albert's statement is true for that birthdate. How do we go from Albert's English statement to a Python function? Let's paraphrase it in a form that uses the concepts we have defined:
> **Albert**: After Cheryl told me the month of her birthdate, I didn't know her birthday. But for *any* of the possible dates, if Bernard was told the day of that date, he would not know Cheryl's birthday.
That I can translate directly into code:
```
def statement3(date) -> bool:
"Albert: I don't know when Cheryl's birthday is, but I know that Bernard does not know too."
dates = tell(Month(date))
return (not know(dates)
and all(not know(tell(Day(d))) for d in dates))
```
We haven't solved the puzzle yet, but let's take a peek and see which dates satisfy statement 3:
```
satisfy(dates, statement3)
```
## 4. Bernard: At first I don't know when Cheryl's birthday is, but I know now.
Again, a paraphrase:
> **Bernard:** At first Cheryl told me the day, and I didn't know. Then, out of the possible dates, I considered just the dates for which Albert's statement 3 is true, and now I know.
```
def statement4(date):
"Bernard: At first I don't know when Cheryl's birthday is, but I know now."
dates = tell(Day(date))
return (not know(dates) and know(satisfy(dates, statement3)))
```
Let's see which dates satisfy both statement 3 and statement 4:
```
satisfy(dates, statement3, statement4)
```
Wait a minute—I thought that Bernard **knew**?! Why are there three possible dates? Bernard does indeed know; it is just that we, the puzzle solvers, don't know. That's because Bernard knows something we don't know: the day. If Bernard was told `'15'` then he would know `'August 15'`; if he was told `'17'` he would know `'August 17'`, and if he was told `'16'` he would know `'July 16'`. *We* don't know because we don't know which of these is the case.
## 5. Albert: Then I also know when Cheryl's birthday is.
Albert is saying that after hearing the month and Bernard's statement 4, he now knows Cheryl's birthday:
```
def statement5(date):
"Albert: Then I also know when Cheryl's birthday is."
return know(satisfy(tell(Month(date)), statement4))
```
## 6. So when is Cheryl's birthday?
```
cheryls_birthday()
```
**Success!** We have deduced that Cheryl's birthday is **July 16**. It is now `True` that we know Cheryl's birthday:
```
know(cheryls_birthday())
```
| github_jupyter |
# MisGAN: Learning from Incomplete Data with GANs
[Generative adversarial networks](https://arxiv.org/abs/1406.2661) (GANs)
provide a powerful modeling framework for learning complex
high-dimensional distributions.
Training GANs normally requires access to a large collection of
fully-observed data. However, it is not always possible to obtain a
large amount of fully-observed data. Missing data is well-known to be
prevalent in many real-world application domains where different data
cases might have different missing entries. This arbitrary missingness
poses a significant challenge to many existing machine learning models.
In this notebook, we present a quick introduction to
[MisGAN](https://openreview.net/forum?id=S1lDV3RcKm¬eId=S1lDV3RcKm),
a GAN-based framework for learning from incomplete data.
We demonstrate how to implement MisGAN in [PyTorch](https://pytorch.org/)
and run it on a modified MNIST dataset where the images are partially-observed.
## Missing data
To understand the design of MisGAN, we first talk about how to formally
model missing data.
The generative process for incompletely observed
data can be described below where $\mathbf{x}\in\mathbb{R}^n$
is a complete data vector and $\mathbf{m}\in\{0,1\}^n$
is a binary mask that determines which entries in $\mathbf{x}$ to reveal:
$$
\begin{split}
\mathbf{x}&\sim p_\theta(\mathbf{x}), \\
\quad\mathbf{m}&\sim p_\phi(\mathbf{m}|\mathbf{x}).
\end{split}
$$
We can represent an incomplete data instance as a pair of
a partially-observed data vector $\mathbf{x}\in\mathbb{R}^n$ and
a corresponding mask $\mathbf{m}\in\{0,1\}^n$
that indicates which entries in $\mathbf{x}$ are observed:
$x_d$ is observed if $m_d=1$ otherwise $x_d$ is missing and might
contain an arbitrary value that we should ignore.
With this representation, an incomplete dataset is in the form of
$\mathcal{D}=\{(\mathbf{x}_i,\mathbf{m}_i)\}_{i=1,\dots,N}$
where both $\mathbf{x}_i$ and $\mathbf{m}_i$ are fixed-length vectors.
Before introducing MisGAN, we first create an incomplete dataset for the rest
of the experiments.
We start with configuring the notebook and importing required modules.
```
# !pip install --upgrade torch torchvision
%reload_ext autoreload
%autoreload 2
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import torch
import torch.nn as nn
import torch.optim as optim
from torch.autograd import grad
from torch.utils.data import DataLoader
from torch.utils.data import Dataset
from torchvision import datasets, transforms
import numpy as np
from matplotlib.patches import Rectangle
import pylab as plt
use_cuda = torch.cuda.is_available()
device = torch.device('cuda' if use_cuda else 'cpu')
```
## Incomplete MNIST dataset
In this notebook, all the experiments will be run on the
[MNIST](http://yann.lecun.com/exdb/mnist/) dataset,
which contains 60000 handwritten digits images of size 28x28.
For the missing data distribution,
we choose the "square observation" pattern: all pixels are missing
except for a square occurring at a random location on the image.
For simplicity, we assume that there is no dependency between the mask and
the content of the image.
This is also known as missing completely at random (MCAR).
`BlockMaskedMNIST` creates an incomplete MNIST dataset by turning each image
in MNIST into a pair of a partially-observed image (with type `FloatTensor`)
and a mask (with type `CharTensor`), which both have size (1, 28, 28).
Note that the range of pixel values of each image is rescaled to \[0,1\].
```
class BlockMaskedMNIST(Dataset):
def __init__(self, block_len, data_dir='src/mnist-data', random_seed=0):
self.block_len = block_len
self.rnd = np.random.RandomState(random_seed)
data = datasets.MNIST(data_dir, train=True, download=True,
transform=transforms.ToTensor())
self.data_size = len(data)
self.generate_incomplete_data(data)
def __getitem__(self, index):
# return index so we can retrieve the mask location from self.mask_loc
return self.image[index], self.mask[index], index
def __len__(self):
return self.data_size
def generate_incomplete_data(self, data):
n_masks = self.data_size
self.image = [None] * n_masks
self.mask = [None] * n_masks
self.mask_loc = [None] * n_masks
for i in range(n_masks):
d0 = self.rnd.randint(0, 28 - self.block_len + 1)
d1 = self.rnd.randint(0, 28 - self.block_len + 1)
mask = torch.zeros((28, 28), dtype=torch.uint8)
mask[d0:(d0 + self.block_len), d1:(d1 + self.block_len)] = 1
self.mask[i] = mask.unsqueeze(0) # add an axis for channel
self.mask_loc[i] = d0, d1, self.block_len, self.block_len
# Mask out missing pixels by zero
self.image[i] = data[i][0] * mask.float()
```
We now create a modified MNIST dataset using `BlockMaskedMNIST` with a
random 12x12 observed block on each image, which accounts for
81.6% missing rate.
```
data = BlockMaskedMNIST(block_len=12)
batch_size = 64
data_loader = DataLoader(data, batch_size=batch_size, shuffle=True,
drop_last=True)
```
Below we implement `plot_grid()` for plotting input images on a grid
of `nrow` rows and `ncol` columns.
An optional argument `bbox` can be provided as a list of (x, y, width, height)
to draw a red rectangular frame with that coordinate on each image.
```
def plot_grid(ax, image, bbox=None, gap=1, gap_value=1, nrow=4, ncol=8,
title=None):
image = image.cpu().numpy().squeeze(1)
LEN = 28
grid = np.empty((nrow * (LEN + gap) - gap, ncol * (LEN + gap) - gap))
grid.fill(gap_value)
for i, x in enumerate(image):
if i >= nrow * ncol:
break
p0 = (i // ncol) * (LEN + gap)
p1 = (i % ncol) * (LEN + gap)
grid[p0:(p0 + LEN), p1:(p1 + LEN)] = x
ax.set_axis_off()
ax.imshow(grid, cmap='binary_r', interpolation='none', aspect='equal')
if bbox:
nplot = min(len(image), nrow * ncol)
for i in range(nplot):
d0, d1, d0_len, d1_len = bbox[i]
p0 = (i // ncol) * (LEN + gap)
p1 = (i % ncol) * (LEN + gap)
offset = np.array([p1 + d1, p0 + d0]) - .5
ax.add_patch(Rectangle(
offset, d1_len, d0_len, lw=1.5, edgecolor='red', fill=False))
if title:
ax.set_title(title)
```
### Masking operator
Here we implement the masking operator
$f_\tau(\mathbf{x}, \mathbf{m}) = \mathbf{x} \odot \mathbf{m} + \tau\bar{\mathbf{m}}$.
As we mentioned before, an incomplete data instance can be represented as
a pair of fixed-length vectors $(\mathbf{x}, \mathbf{m})$.
The masking operator transforms an incomplete data instance into a vector of
the same size with all missing entries in $\mathbf{x}$ replaced by a constant
value $\tau$.
This plays an important role in MisGAN that we will describe later.
Before that, we will use it to visualize the incomplete MNIST dataset
we just prepared.
```
def mask_data(data, mask, tau=0):
return mask * data + (1 - mask) * tau
```
### Visualization of incomplete MNIST images
We plot a random subset of images from the incomplete MNIST dataset below.
Gray pixels represent the missing entries in each image.
```
data_samples, mask_samples, _ = next(iter(data_loader))
fig, ax = plt.subplots(figsize=(12, 3))
plot_grid(ax, mask_data(data_samples, mask_samples.float(), .5),
nrow=4, ncol=16)
```
## MisGAN
MisGAN is a GAN-based framework for learning distributions in the presence
of incomplete observations. The overall structure is illustrated below:
<img src="img/misgan.png" width="500" style="display: block; margin: 2em auto" />
MisGAN consists of a data generator $G_x$ that generates complete data.
In addition, it also has a mask generator $G_m$ to explicitly model the
missing data process.
Note that the input $(\mathbf{x}, \mathbf{m})$ is the incomplete data that
follows the representation mentioned earlier.
MisGAN mimics the generation of incomplete data (under the MCAR assumption)
by treating the generated complete data $\tilde{\mathbf{x}}$ together with
the generated mask $\tilde{\mathbf{m}}$ as an incomplete data instance,
with the mask specifies which entries in $\tilde{\mathbf{x}}$ are considered
missing.
We then complete both the real incomplete data $(\mathbf{x}, \mathbf{m})$
and the generated ones $(\tilde{\mathbf{x}}, \tilde{\mathbf{m}})$
using the same masking operator $f_{\tau}$.
We train the data generator $G_x$ by making the masked generated data
$f_{\tau}(\tilde{\mathbf{x}}, \tilde{\mathbf{m}})$ indistinguishable from
the masked real incomplete data $f_{\tau}(\mathbf{x}, \mathbf{m})$ using
a data discriminator $D_x$.
On the other hand, since both generated masks $\tilde{\mathbf{m}}$ and
real masks $\mathbf{m}$ are fully-observed, we can train the mask generator
$G_m$ with a mask discriminator $D_m$ as in a standard GAN.
### Generator
For each generator, we use a linear layer followed by three deconvolution
layers with ReLUs in between.
`ConvDataGenerator` implements the data generator $G_x$.
Since the pixel values are in \[0, 1\], we apply the sigmoid activation to
the real-valued output at the end.
`ConvMaskGenerator` implements the mask generator $G_m$.
Note that the masks are binary-valued.
Since discrete data generating processes have zero gradient almost everywhere,
to carry out gradient-based training for GANs, we relax the output
of the mask generator $G_m$ from $\{0,1\}^n$ to $[0, 1]^n$.
We use the sigmoid activated output $\sigma(z / \lambda)$
with a low temperature $\lambda = 0.66$ to encourage saturation
and make the output closer to zero or one.
```
# Must sub-class ConvGenerator to provide transform()
class ConvGenerator(nn.Module):
def __init__(self, latent_size=128):
super().__init__()
self.DIM = 64
self.latent_size = latent_size
self.preprocess = nn.Sequential(
nn.Linear(latent_size, 4 * 4 * 4 * self.DIM),
nn.ReLU(True),
)
self.block1 = nn.Sequential(
nn.ConvTranspose2d(4 * self.DIM, 2 * self.DIM, 5),
nn.ReLU(True),
)
self.block2 = nn.Sequential(
nn.ConvTranspose2d(2 * self.DIM, self.DIM, 5),
nn.ReLU(True),
)
self.deconv_out = nn.ConvTranspose2d(self.DIM, 1, 8, stride=2)
def forward(self, input):
net = self.preprocess(input)
net = net.view(-1, 4 * self.DIM, 4, 4)
net = self.block1(net)
net = net[:, :, :7, :7]
net = self.block2(net)
net = self.deconv_out(net)
return self.transform(net).view(-1, 1, 28, 28)
class ConvDataGenerator(ConvGenerator):
def __init__(self, latent_size=128):
super().__init__(latent_size=latent_size)
self.transform = lambda x: torch.sigmoid(x)
class ConvMaskGenerator(ConvGenerator):
def __init__(self, latent_size=128, temperature=.66):
super().__init__(latent_size=latent_size)
self.transform = lambda x: torch.sigmoid(x / temperature)
```
### Discriminator
We implement the discriminator (or referred to as the critic in
Wasserstein GANs) in `ConvCritic` with three convolutional layers followed by
a linear layer for both $D_x$ and $D_m$.
```
class ConvCritic(nn.Module):
def __init__(self):
super().__init__()
self.DIM = 64
main = nn.Sequential(
nn.Conv2d(1, self.DIM, 5, stride=2, padding=2),
nn.ReLU(True),
nn.Conv2d(self.DIM, 2 * self.DIM, 5, stride=2, padding=2),
nn.ReLU(True),
nn.Conv2d(2 * self.DIM, 4 * self.DIM, 5, stride=2, padding=2),
nn.ReLU(True),
)
self.main = main
self.output = nn.Linear(4 * 4 * 4 * self.DIM, 1)
def forward(self, input):
input = input.view(-1, 1, 28, 28)
net = self.main(input)
net = net.view(-1, 4 * 4 * 4 * self.DIM)
net = self.output(net)
return net.view(-1)
```
### Training Wasserstein GAN with gradient penalty
MisGAN is compatible with many GAN variations, and here we use
[Wasserstein GAN](https://arxiv.org/abs/1701.07875) to train MisGAN.
`CriticUpdater` computes the loss of the discriminator and updates
its parameters accordingly.
We follow the [WGAN-GP](https://arxiv.org/abs/1704.00028) procedure to
train discriminators with the gradient penalty.
Specifically,
given the data distribution $p_r$ and the model distribution $p_g$,
the loss for the discriminator $D$ is given by
$$
\mathcal{L} =
\mathbb{E}_{\tilde{\mathbf{x}}\sim p_g}\big[D(\tilde{\mathbf{x}})\big] -
\mathbb{E}_{\mathbf{x}\sim p_r}\big[D(\mathbf{x})\big] +
\lambda \mathbb{E}_{\mathbf{y}}
\big[\left(\|\nabla_{\mathbf{y}} D(\mathbf{y})\|_2 - 1\right)^2\big]
$$
where $\mathbf{y}$ is sampled according to
$\mathbf{y}=\xi\mathbf{x} + (1-\xi)\tilde{\mathbf{x}}$ with
$\tilde{\mathbf{x}}\sim p_g$, $\mathbf{x}\sim p_r$,
and $\xi\sim\operatorname{uniform}(0, 1)$.
The gradient penalty term is for enforcing the (soft) 1-Lipschtiz
constraint required by the Wasserstein GAN.
Samples drawn from $p_r$ and $p_g$ are provided as `real` and `fake`
respectively when calling `CriticUpdater`.
```
class CriticUpdater:
def __init__(self, critic, critic_optimizer, batch_size=64, gp_lambda=10):
self.critic = critic
self.critic_optimizer = critic_optimizer
self.gp_lambda = gp_lambda
# Interpolation coefficient
self.eps = torch.empty(batch_size, 1, 1, 1, device=device)
# For computing the gradient penalty
self.ones = torch.ones(batch_size).to(device)
def __call__(self, real, fake):
real = real.detach()
fake = fake.detach()
self.critic.zero_grad()
self.eps.uniform_(0, 1)
interp = (self.eps * real + (1 - self.eps) * fake).requires_grad_()
grad_d = grad(self.critic(interp), interp, grad_outputs=self.ones,
create_graph=True)[0]
grad_d = grad_d.view(real.shape[0], -1)
grad_penalty = ((grad_d.norm(dim=1) - 1)**2).mean() * self.gp_lambda
w_dist = self.critic(fake).mean() - self.critic(real).mean()
loss = w_dist + grad_penalty
loss.backward()
self.critic_optimizer.step()
```
Now, we instantiate all the building blocks for MisGAN: the data/mask
generators and their corresponding discriminators.
We use the [Adam optimizer](https://arxiv.org/abs/1412.6980) to train them.
```
nz = 128 # dimensionality of the latent code
n_critic = 5
alpha = .2
data_gen = ConvDataGenerator().to(device)
mask_gen = ConvMaskGenerator().to(device)
data_critic = ConvCritic().to(device)
mask_critic = ConvCritic().to(device)
data_noise = torch.empty(batch_size, nz, device=device)
mask_noise = torch.empty(batch_size, nz, device=device)
lrate = 1e-4
data_gen_optimizer = optim.Adam(
data_gen.parameters(), lr=lrate, betas=(.5, .9))
mask_gen_optimizer = optim.Adam(
mask_gen.parameters(), lr=lrate, betas=(.5, .9))
data_critic_optimizer = optim.Adam(
data_critic.parameters(), lr=lrate, betas=(.5, .9))
mask_critic_optimizer = optim.Adam(
mask_critic.parameters(), lr=lrate, betas=(.5, .9))
update_data_critic = CriticUpdater(
data_critic, data_critic_optimizer, batch_size)
update_mask_critic = CriticUpdater(
mask_critic, mask_critic_optimizer, batch_size)
```
### Training MisGAN
To describe the training procedure of MisGAN,
we first define the following loss functions,
one for the masks and the other for the data:
$$
\begin{split}
\mathcal{L}_m(D_m, G_m) &=
\mathbb{E}_{(\mathbf{x},\mathbf{m})\sim p_\mathcal{D}}\left[D_m(\mathbf{m})\right] -
\mathbb{E}_{\boldsymbol{\varepsilon}\sim p_\varepsilon}\left[D_m(G_m(\boldsymbol{\varepsilon}))\right],
\\
\mathcal{L}_x(D_x, G_x, G_m) &=
\mathbb{E}_{(\mathbf{x},\mathbf{m})\sim p_\mathcal{D}}\left[D_x(f_\tau(\mathbf{x},\mathbf{m}))\right] -
\mathbb{E}_{\boldsymbol{\varepsilon}\sim p_\varepsilon, \mathbf{z}\sim p_z}\left[
D_x\left(f_\tau\left(G_x(\mathbf{z}),G_m(\boldsymbol{\varepsilon})\right)\right)\right].
\end{split}
$$
We optimize the generators and the discriminators
according to the following objectives:
$$
\begin{split}
&\min_{G_x}\max_{D_x\in\mathcal{F}_x} \mathcal{L}_x(D_x, G_x, G_m),
\\
&\min_{G_m}\max_{D_m\in\mathcal{F}_m} \mathcal{L}_m(D_m, G_m) + \alpha \mathcal{L}_x(D_x, G_x, G_m),
\end{split}
$$
where $\mathcal{F}_x, \mathcal{F}_m$ are defined such that $D_x, D_m$ are
both 1-Lipschitz in Wasserstein GANs.
We alternate between `n_critic` steps of optimizing the discriminators
and one step of optimizing the generators.
The update of the discriminator is implemented in `CriticUpdater` described
earlier, which handles the Lipschitz constraint.
We use $\alpha=0.2$ to encourage the generated masks to match the distribution
of the real masks and the masked generated complete samples
to match masked real data.
In the example,
we use the standard Gaussian $\mathcal{N}(\mathbf{0}, \mathbf{I})$
for both noise distributions $p_z$ and $p_\varepsilon$.
During training, we draw a bunch of generated complete data and mask samples
every `plot_interval` epochs to assess the training progress qualitatively.
```
plot_interval = 50
critic_updates = 0
for epoch in range(300):
for real_data, real_mask, _ in data_loader:
real_data = real_data.to(device)
real_mask = real_mask.to(device).float()
# Update discriminators' parameters
data_noise.normal_()
mask_noise.normal_()
fake_data = data_gen(data_noise)
fake_mask = mask_gen(mask_noise)
masked_fake_data = mask_data(fake_data, fake_mask)
masked_real_data = mask_data(real_data, real_mask)
update_data_critic(masked_real_data, masked_fake_data)
update_mask_critic(real_mask, fake_mask)
critic_updates += 1
if critic_updates == n_critic:
critic_updates = 0
# Update generators' parameters
for p in data_critic.parameters():
p.requires_grad_(False)
for p in mask_critic.parameters():
p.requires_grad_(False)
data_gen.zero_grad()
mask_gen.zero_grad()
data_noise.normal_()
mask_noise.normal_()
fake_data = data_gen(data_noise)
fake_mask = mask_gen(mask_noise)
masked_fake_data = mask_data(fake_data, fake_mask)
data_loss = -data_critic(masked_fake_data).mean()
data_loss.backward(retain_graph=True)
data_gen_optimizer.step()
mask_loss = -mask_critic(fake_mask).mean()
(mask_loss + data_loss * alpha).backward()
mask_gen_optimizer.step()
for p in data_critic.parameters():
p.requires_grad_(True)
for p in mask_critic.parameters():
p.requires_grad_(True)
if plot_interval > 0 and (epoch + 1) % plot_interval == 0:
# Although it makes no difference setting eval() in this example,
# you will need those if you are going to use modules such as
# batch normalization or dropout in the generators.
data_gen.eval()
mask_gen.eval()
with torch.no_grad():
print('Epoch:', epoch)
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(10, 2.5))
data_noise.normal_()
data_samples = data_gen(data_noise)
plot_grid(ax1, data_samples, title='generated complete data')
mask_noise.normal_()
mask_samples = mask_gen(mask_noise)
plot_grid(ax2, mask_samples, title='generated masks')
plt.show()
plt.close(fig)
data_gen.train()
mask_gen.train()
```
## Missing data imputation
Now, we introduce an extension of MisGAN for missing data imputation.
The goal of missing data imputation is to complete the
missing data according to $p(\mathbf{x}_\text{mis}|\mathbf{x}_\text{obs})$.
To do so, we augment MisGAN with an imputer $G_i$ accompanied by
a corresponding discriminator $D_i$ as illustrated below.
<img src="img/misgan-impute.png" width="625" style="display: block; margin: 2em auto" />
The imputer $G_m$ is a function of the incomplete example
$(\mathbf{x},\mathbf{m})$ and a random vector $\boldsymbol{\omega}$
drawn from a noise distribution $p_\omega$.
The noise $\boldsymbol{\omega}$ is for modeling the uncertainty when sampling
from $p(\mathbf{x}_\text{mis}|\mathbf{x}_\text{obs})$.
The imputer is trained by making the completed data $\hat{\mathbf{x}}$
indistinguishable from the generated complete data $\tilde{\mathbf{x}}$.
### Imputer
We construct the imputer $G_i(\mathbf{x},\mathbf{m},\boldsymbol{\omega})$
as the follows:
$$
G_i(\mathbf{x},\mathbf{m},\boldsymbol{\omega})
=\mathbf{x}\odot\mathbf{m}+\hat{G}_i(\mathbf{x}\odot\mathbf{m} + \boldsymbol{\omega}\odot\bar{\mathbf{m}})\odot\bar{\mathbf{m}}.
$$
where $\hat{G}_i$ is a imputer network that generates the imputation result.
The masking on the input of the imputer network,
$\mathbf{x}\odot\mathbf{m} + \boldsymbol{\omega}\odot\bar{\mathbf{m}}$,
ensures that the amount of noise injected to $\hat{G}_i$
is complementary to the size of the observed features.
This is intuitive in the sense that when a data case is almost fully-observed,
we expect less variety in $p(\mathbf{x}_\text{mis}|\mathbf{x}_\text{obs})$
and vice versa.
Note that the noise $\boldsymbol{\omega}$ needs to have the same dimensionality
as $\mathbf{x}$.
The final masking (outside of $\hat{G}_i$)
ensures that the observed entries of $\mathbf{x}$ are kept
intact in the output of the imputer $G_i$.
We implement $G_i$ in `Imputer` as a three-layer fully-connected network
with ReLUs in between.
```
class Imputer(nn.Module):
def __init__(self, arch=(512, 512)):
super().__init__()
self.fc = nn.Sequential(
nn.Linear(784, arch[0]),
nn.ReLU(),
nn.Linear(arch[0], arch[1]),
nn.ReLU(),
nn.Linear(arch[1], arch[0]),
nn.ReLU(),
nn.Linear(arch[0], 784),
)
def forward(self, data, mask, noise):
net = data * mask + noise * (1 - mask)
net = net.view(data.shape[0], -1)
net = self.fc(net)
net = torch.sigmoid(net).view(data.shape)
return data * mask + net * (1 - mask)
```
Now we instantiate the imputer and the corresponding discriminator.
We also use the Adam optimizer to train them.
Note that for MisGAN imputation, we will re-use most of the components
created earlier for MisGAN including the data/mask generators and
the discriminators.
```
imputer = Imputer().to(device)
impu_critic = ConvCritic().to(device)
impu_noise = torch.empty(batch_size, 1, 28, 28, device=device)
imputer_lrate = 2e-4
imputer_optimizer = optim.Adam(
imputer.parameters(), lr=imputer_lrate, betas=(.5, .9))
impu_critic_optimizer = optim.Adam(
impu_critic.parameters(), lr=imputer_lrate, betas=(.5, .9))
update_impu_critic = CriticUpdater(
impu_critic, impu_critic_optimizer, batch_size)
```
### Training MisGAN imputer
To train the imputer-equipped MisGAN,
we define the following losses (MisGAN uses the first two):
$$
\begin{split}
\mathcal{L}_m(D_m, G_m) &=
\mathbb{E}_{(\mathbf{x},\mathbf{m})\sim p_\mathcal{D}}\left[D_m(\mathbf{m})\right] -
\mathbb{E}_{\boldsymbol{\varepsilon}\sim p_\varepsilon}\left[D_m(G_m(\boldsymbol{\varepsilon}))\right],
\\
\mathcal{L}_x(D_x, G_x, G_m) &=
\mathbb{E}_{(\mathbf{x},\mathbf{m})\sim p_\mathcal{D}}\left[D_x(f_\tau(\mathbf{x},\mathbf{m}))\right] -
\mathbb{E}_{\boldsymbol{\varepsilon}\sim p_\varepsilon, \mathbf{z}\sim p_z}\left[
D_x\left(f_\tau\left(G_x(\mathbf{z}),G_m(\boldsymbol{\varepsilon})\right)\right)\right],
\\
\mathcal{L}_i(D_i, G_i, G_x) &=
\mathbb{E}_{\mathbf{z}\sim p_z}\left[D_i(G_x(\mathbf{z}))\right] -
\mathbb{E}_{(\mathbf{x},\mathbf{m})\sim p_\mathcal{D}, \boldsymbol{\omega}\sim p_\omega}
\left[D_i(G_i(\mathbf{x},\mathbf{m},\boldsymbol{\omega}))\right].
\end{split}
$$
We jointly learn the data generating process and
the imputer according to the following objectives:
$$
\begin{split}
\min_{G_i}\max_{D_i\in\mathcal{F}_i}\ &\mathcal{L}_i(D_i, G_i, G_x),
\label{eq:objimputer}\\
\min_{G_x}\max_{D_x\in\mathcal{F}_x}\ &\mathcal{L}_x(D_x, G_x, G_m) + \beta \mathcal{L}_i(D_i, G_i, G_x),
\\
\min_{G_m}\max_{D_m\in\mathcal{F}_m}\ &\mathcal{L}_m(D_m, G_m) + \alpha \mathcal{L}_x(D_x, G_x, G_m),
\end{split}
$$
where we use $\beta=0.1$ to encourage the generated complete data to match
the distribution of the imputed real data in addition to
having the masked generated data to match the masked real data.
In the MNIST example, the noise for the imputer $\boldsymbol{\omega}$
(`impu_noise`) is drawn from $p_\omega=\operatorname{uniform}(0, 1)$
to match the range of the pixel value \[0, 1\].
We plot the imputation results every `plot_interval` epochs.
In each of the plots,
the region inside of each red box are the observed pixels;
the pixels outside of the box are generated by the imputer.
```
beta = .1
plot_interval = 100
critic_updates = 0
for epoch in range(600):
for real_data, real_mask, index in data_loader:
real_data = real_data.to(device)
real_mask = real_mask.to(device).float()
masked_real_data = mask_data(real_data, real_mask)
# Update discriminators' parameters
data_noise.normal_()
fake_data = data_gen(data_noise)
mask_noise.normal_()
fake_mask = mask_gen(mask_noise)
masked_fake_data = mask_data(fake_data, fake_mask)
impu_noise.uniform_()
imputed_data = imputer(real_data, real_mask, impu_noise)
update_data_critic(masked_real_data, masked_fake_data)
update_mask_critic(real_mask, fake_mask)
update_impu_critic(fake_data, imputed_data)
critic_updates += 1
if critic_updates == n_critic:
critic_updates = 0
# Update generators' parameters
for p in data_critic.parameters():
p.requires_grad_(False)
for p in mask_critic.parameters():
p.requires_grad_(False)
for p in impu_critic.parameters():
p.requires_grad_(False)
data_noise.normal_()
fake_data = data_gen(data_noise)
mask_noise.normal_()
fake_mask = mask_gen(mask_noise)
masked_fake_data = mask_data(fake_data, fake_mask)
impu_noise.uniform_()
imputed_data = imputer(real_data, real_mask, impu_noise)
data_loss = -data_critic(masked_fake_data).mean()
mask_loss = -mask_critic(fake_mask).mean()
impu_loss = -impu_critic(imputed_data).mean()
mask_gen.zero_grad()
(mask_loss + data_loss * alpha).backward(retain_graph=True)
mask_gen_optimizer.step()
data_gen.zero_grad()
(data_loss + impu_loss * beta).backward(retain_graph=True)
data_gen_optimizer.step()
imputer.zero_grad()
impu_loss.backward()
imputer_optimizer.step()
for p in data_critic.parameters():
p.requires_grad_(True)
for p in mask_critic.parameters():
p.requires_grad_(True)
for p in impu_critic.parameters():
p.requires_grad_(True)
if plot_interval > 0 and (epoch + 1) % plot_interval == 0:
with torch.no_grad():
imputer.eval()
# Plot imputation results
impu_noise.uniform_()
imputed_data = imputer(real_data, real_mask, impu_noise)
bbox = [data.mask_loc[idx] for idx in index]
print('Epoch:', epoch)
fig, ax = plt.subplots(figsize=(6, 3))
plot_grid(ax, imputed_data, bbox, gap=2)
plt.show()
plt.close(fig)
imputer.train()
```
| github_jupyter |
```
from jupyterthemes import jtplot
jtplot.style(theme='onedork', context='talk', fscale=1.4, spines=False, gridlines='--', ticks=True, grid=False, figsize=(6, 4.5))
from os.path import join
import pandas as pd
import numpy as np
import seaborn as sns
current_palette = sns.color_palette()
%matplotlib inline
import matplotlib.pyplot as plt
from matplotlib_venn import venn2
from matplotlib import rcParams
from matplotlib.ticker import FuncFormatter
from scipy.stats import fisher_exact
from ipywidgets import interact, IntSlider, FloatSlider
```
### Simulation of Association Rule Metrics
```
total_widget = IntSlider(min=10, max=1000, step=10, value=500)
antecedent_widget = IntSlider(min=5, max=1000, step=5, value=100)
consequent_widget = IntSlider(min=5, max=1000, step=5, value=100)
joint_widget = FloatSlider(min=.01, max=1.0, value=.5)
def plot_metrics(antecedent, consequent, joint_percent, total):
"""Interactive Venn Diagram of joint transactions and plot of support, confidence, and lift
Slider Inputs:
- total: total transactions for all itemsets
- antecedent, consequent: all transactions involving either itemset
- joint_percent: percentage of (smaller of) antecedent/consequent involving both itemsets
Venn Diagram Calculations:
- joint = joint_percent * min(antecedent, consequent)
- antecedent, consequent: original values - joint transactions
Metric Calculations:
- Support Antecedent: antecedent/total
- Support Consequent: Consequent/total
- Support Joint Transactions: joint/total
- Rule Confidence: Support Joint Transactions / total
- Rule Lift: Support Joint Transactions / (Support Antecedent * Support Consequent)
"""
fig = plt.figure(figsize=(15, 8))
ax1 = plt.subplot2grid((2, 2), (0, 0))
ax2 = plt.subplot2grid((2, 2), (0, 1))
ax3 = plt.subplot2grid((2, 2), (1, 0))
ax4 = plt.subplot2grid((2, 2), (1, 1))
joint = int(joint_percent * min(antecedent, consequent))
contingency_table = [[joint, consequent - joint], [antecedent - joint, max(total - antecedent - consequent + joint, 0)]]
contingency_df = pd.DataFrame(contingency_table, columns=['Consequent', 'Not Consequent'], index=['Antecedent', 'Not Antecedent']).astype(int)
sns.heatmap(contingency_df, ax=ax1, annot=True, cmap='Blues', square=True, vmin=0, vmax=total, fmt='.0f')
ax1.set_title('Contingency Table')
v = venn2(subsets=(antecedent - joint, consequent - joint, joint),
set_labels=['Antecedent', 'Consequent'],
set_colors=current_palette[:2],
ax=ax2)
ax2.set_title("{} Transactions".format(total))
support_antecedent = antecedent / total
support_consequent = consequent / total
support = pd.Series({'Antecedent': support_antecedent,
'Consequent': support_consequent})
support.plot(kind='bar', ax=ax3,
color=current_palette[:2], title='Support', ylim=(0, 1), rot=0)
ax3.yaxis.set_major_formatter(
FuncFormatter(lambda y, _: '{:.0%}'.format(y)))
support_joint = joint / total
confidence = support_joint / support_antecedent
lift = support_joint / (support_antecedent * support_consequent)
_, pvalue = fisher_exact(contingency_table, alternative='greater')
metrics = pd.Series(
{'Confidence': confidence, 'Lift': lift, 'p-Value': pvalue})
metrics.plot(kind='bar', ax=ax4,
color=current_palette[2:5], rot=0, ylim=(0, 2))
ax3.yaxis.set_major_formatter(
FuncFormatter(lambda y, _: '{:.0%}'.format(y)))
for ax, series in {ax3: support, ax4: metrics}.items():
rects = ax.patches
labels = ['{:.0%}'.format(x) for x in series.tolist()]
for rect, label in zip(rects, labels):
height = min(rect.get_height() + .01, 2.05)
ax.text(rect.get_x() + rect.get_width() / 2,
height, label, ha='center', va='bottom')
plt.suptitle('Assocation Rule Analysis {Antecedent => Consequent}')
plt.tight_layout()
plt.subplots_adjust(top=0.9)
plt.show()
interact(plot_metrics,
antecedent=antecedent_widget,
consequent=consequent_widget,
joint_percent=joint_widget,
total=total_widget);
```
| github_jupyter |
```
import os
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from sklearn.ensemble import RandomForestRegressor
from treeinterpreter import treeinterpreter as ti
from sklearn.metrics import mean_squared_error, mean_absolute_error, mean_absolute_percentage_error
from math import sqrt
from sklearn.svm import SVR
from sklearn.preprocessing import MinMaxScaler
# master_df = pd.DataFrame()
# set columns script, "", "", train_p, "", test_p, "", "", mse, rmse, mae, mape, "", cross_correlation
master_df = pd.DataFrame(columns=['script', '', '', 'train_p', '', 'test_p', '', '', 'mse', 'rmse', 'mae', 'mape', '', 'cross_correlation'])
all_title = 'score'
```
### Cross Correlation function for future use
```
def crosscorr(datax, datay, lag=0, method="pearson"):
""" Lag-N cross correlation.
Parameters
—------—
lag : int, default 0
datax, datay : pandas.Series objects of equal length
Returns
—------—
crosscorr : float
"""
return datax.corr(datay.shift(lag), method=method)
# Read data from csv files
# TODO: change the path to your own path depending on the file that wants to be processed.
# df_read = pd.read_csv(f'{os.getcwd()}/dataset_prediction-models/final_price-and-compund.csv', index_col="date")
df_read = pd.read_csv(f'{os.getcwd()}/dataset_prediction-models/final_price-and-score.csv', index_col="date")
df_read.rename(columns={'close':'Price'}, inplace=True)
# reorder columns compund, price
df_read = df_read[['score', 'Price']]
# df_read = df_read[['compund', 'Price']]
# Scale the data to be between 0 and 1 using MinMaxScaler.fit_transform().
# This is done to make the data easier to work with becasue the data is between 0 and 1.
df_values = df_read.values
sc = MinMaxScaler()
data_scaled = sc.fit_transform(df_values)
# save the numpy array back in a dataframe
df_scaled = pd.DataFrame(data_scaled, columns=df_read.columns, index=df_read.index)
print(df_scaled.head())
```
### Random Forest regression model
```
# Predicting stock prices using Random Forest Regression model
def RandomForestModel(data_scaled):
"""
This function takes in a dataframe and predicts the stock price using
Random Forest Regression model. The function returns the predicted stock
price and the mean absolute error. It also plots the predicted stock price
and the actual stock price.
"""
# split the data into train and test.
train_index = int(0.75 * len(data_scaled))
train = data_scaled[:train_index]
test = data_scaled[train_index:]
print("train,test,val",train.shape, test.shape)
# split the data into features and target.
xtrain, ytrain = train[:,:2], train[:,1]
xtest, ytest = test[:,:2], test[:,1]
X_train = np.zeros((len(xtrain), 2))
y_train = np.zeros((len(ytrain), 1))
X_test = np.zeros((len(xtest), 2))
y_test = np.zeros((len(ytest), 1))
# fill the numpy array with the train data
for i in range(len(xtrain)):
X_train[i] = xtrain[i]
y_train[i] = ytrain[i]
print("x_train", X_train.shape)
print("y_train", y_train.shape)
# fill the numpy array with the test data
for i in range(len(xtest)):
X_test[i] = xtest[i]
y_test[i] = ytest[i]
print("x_test", X_test.shape)
print("y_test", y_test.shape)
###############################################################################
# Create a Random Forest Regressor model.
# n_estimators is the number of trees in the forest. It is the number of trees that are grown.
# random_state is the seed used by the random number generator.
# It is set to 0, because we want the results to be reproducible.
# The random number generator is used to select the features for splitting the data.
# max_depth is the maximum depth of the tree. It is the maximum number of splits that a node can have.
rf = RandomForestRegressor(n_estimators=50, random_state=0, max_depth=5)
rf.fit(X_train, y_train)
# Predict the stock price using the Random Forest Regressor model using a tree interpreter.
# The tree interpreter is used to interpret the tree and get the predicted stock price.
y_pred, bias, contributions = ti.predict(rf, X_test)
mse = mean_squared_error(y_test, y_pred)
rmse = sqrt(mean_squared_error(y_test, y_pred))
mae = mean_absolute_error(y_test, y_pred)
mape = mean_absolute_percentage_error(y_test, y_pred)
# Accuracy of the model
# print("Accuracy of the model on training data:", rf.score(X_train, y_train) * 100, "%")
print("\nAccuracy of the model on testing data: ", round(rf.score(X_test, y_test), 2) * 100, "%")
print('Mean Squared Error:', round(mse, 4))
print('Root mean squared error:', round(rmse, 4))
print('Mean absolute error:', round(mae, 4))
print('Mean absolute percentage error:', round(mape, 4) * 100, '%')
plt.figure(figsize=(18, 8))
plt.plot(y_test, '-', color='blue', label='Actual Price')
plt.plot(y_pred, '.-', color='orange', label='Predicted Price')
plt.title('Random Forest Regression on BTC dataset (Close price and engagement score)')
plt.xlabel('Hours (units of time)')
plt.ylabel('Price (normalised)')
plt.legend()
# plt.gcf().autofmt_xdate()
plt.show()
###############################################################################
# plot the predicted stock price and the actual stock price on the same graph.
fig, ax = plt.subplots(figsize=(18, 8))
ax.plot(y_test, '-', color='blue', label='Actual Price', linewidth=1.2)
ax.plot(y_pred, '.-', color='orange', label='Predicted Price', linewidth=1.6)
# add a background to the plot
ax.axvspan(0, 50, alpha=0.15, color='grey')
# add a title to the plot
ax.set_title('Random Forest Regression on BTC dataset (Close price and engagement score)')
# add a label to the x-axis
ax.set_xlabel('Hours (units of time)')
# add a label to the y-axis
ax.set_ylabel('Price (normalised)')
# add a legend
ax.legend()
# add a grid
ax.grid()
plt.show()
return y_test, y_pred, mse, rmse, mae, mape
y_test, y_pred, mse, rmse, mae, mape = RandomForestModel(data_scaled)
```
### Cross Correlation - Random Forest Regression
```
actual_price_df = pd.DataFrame(y_test, columns=['price'])
predicted_price_df = pd.DataFrame(y_pred, columns=['price'])
# Calculate the Pearson Cross Correlation for lag 0
curr_corr = crosscorr(predicted_price_df['price'], actual_price_df['price'],
method="pearson")
print("Pearson Correlation for lag 0:", curr_corr)
master_df = master_df.append({'script': f'RF_price-{all_title}',
'': '',
'': '',
'': '',
'train_p': 60,
'': '',
'test_p': 40,
'': '',
'mse': round(mse, 4),
'rmse': round(rmse, 4),
'mae': round(mae, 4),
'mape': round(mape, 4),
'': '',
'cross_correlation': curr_corr},
ignore_index=True)
```
### SVR prediction
```
def SVRModel(data_scaled):
"""
This function takes in a dataframe and predicts the stock price using
Support Vector Regression model. The function returns the predicted stock
price and the mean absolute error. It also plots the predicted stock price
and the actual stock price.
"""
# split the data into train and test.
train_index = int(0.6 * len(data_scaled))
train = data_scaled[:train_index]
test = data_scaled[train_index:]
print("train,test,val",train.shape, test.shape)
# split the data into features and target.
xtrain, ytrain = train[:,:2], train[:,1]
xtest, ytest = test[:,:2], test[:,1]
X_train = np.zeros((len(xtrain), 2))
y_train = np.zeros((len(ytrain), 1))
X_test = np.zeros((len(xtest), 2))
y_test = np.zeros((len(ytest), 1))
# fill the numpy array with the train data
for i in range(len(xtrain)):
X_train[i] = xtrain[i]
y_train[i] = ytrain[i]
print("x_train", X_train.shape)
print("y_train", y_train.shape)
# fill the numpy array with the test data
for i in range(len(xtest)):
X_test[i] = xtest[i]
y_test[i] = ytest[i]
print("x_test", X_test.shape)
print("y_test", y_test.shape)
# ###############################################################################
# Create the Support Vector Regression model.
regressor = SVR(kernel='rbf')#, C=1, gamma='auto')
regressor.fit(X_train, y_train)
y_pred = regressor.predict(X_test)
rmse = sqrt(mean_squared_error(y_test, y_pred))
mae = mean_absolute_error(y_test, y_pred)
mape = mean_absolute_percentage_error(y_test, y_pred)
# print('Accuracy of the model on training data:', round(regressor.score(X_train, y_train), 2) * 100, '%')
print('Accuracy of the model on testing data:', round(regressor.score(X_test, y_test), 2) * 100, '%')
print('Root mean squared error:', round(rmse, 4))
print('Mean absolute error:', round(mae, 4))
print('Mean absolute percentage error:', round(mape, 4) * 100, '%')
# Plot the actual stock price and the predicted stock price.
plt.figure(figsize=(18, 8))
plt.plot(y_test, '-', color='blue', label='Actual Price')
plt.plot(y_pred, '.-', color='orange', label='Predicted Price')
plt.title('Support Vector Regression on BTC dataset (Close price and engagement score)')
plt.xlabel('Hours (units of time)')
plt.ylabel('Price (normalised)')
plt.legend()
# plt.gcf().autofmt_xdate()
plt.show()
###############################################################################
# plot the predicted stock price and the actual stock price on the same graph.
fig, ax = plt.subplots(figsize=(18, 8))
ax.plot(y_test, '-', color='blue', label='Actual Price')
ax.plot(y_pred, '.-', color='orange', label='Predicted Price')
# add a background to the plot
ax.axvspan(0, 80, alpha=0.15, color='grey')
# add a title to the plot
ax.set_title('Support Vector Regression on BTC dataset (Close price and engagement score)')
# add a label to the x-axis
ax.set_xlabel('Hours (units of time)')
# add a label to the y-axis
ax.set_ylabel('Price (normalised)')
# add a legend
ax.legend()
# add a grid
ax.grid()
plt.show()
return y_test, y_pred, mse, rmse, mae, mape
y_test, y_pred, mse, rmse, mae, mape = SVRModel(data_scaled)
```
### Cross Correlation on SVM
```
actual_price_df = pd.DataFrame(y_test, columns=['price'])
predicted_price_df = pd.DataFrame(y_pred, columns=['price'])
# Calculate the Pearson Cross Correlation for lag 0
curr_corr = crosscorr(predicted_price_df['price'], actual_price_df['price'],
method="pearson")
print("Pearson Correlation for lag 0:", curr_corr)
master_df = master_df.append({'script': f'SV_price-and-{all_title}',
'': '',
'': '',
'': '',
'train_p': 60,
'': '',
'test_p': 40,
'': '',
'mse': round(mse, 4),
'rmse': round(rmse, 4),
'mae': round(mae, 4),
'mape': round(mape, 4),
'': '',
'cross_correlation': curr_corr},
ignore_index=True)
```
### Percentage of tweets (sentiment analysis)
```
# Create a pie chart to show the percentage of positive and negative sentiment
# positive_sentiment = df_read['positive'].sum()
# negative_sentiment = df_read['negative'].sum()
# neutral_sentiment = df_read['neutral'].sum()
# # Percentage of positive sentiment
# positive_sentiment_percentage = positive_sentiment / (positive_sentiment + negative_sentiment + neutral_sentiment) * 100
# negative_sentiment_percentage = negative_sentiment / (positive_sentiment + negative_sentiment + neutral_sentiment) * 100
# neutral_sentiment_percentage = neutral_sentiment / (positive_sentiment + negative_sentiment + neutral_sentiment) * 100
# # Print the percentage of positive and negative sentiment
# print("\n\n")
# print("Percentage of positive sentiment:", positive_sentiment_percentage)
# print("Percentage of negative sentiment:", negative_sentiment_percentage)
# print("Percentage of neutral sentiment:", neutral_sentiment_percentage)
# # Plot the pie chart
# labels = 'Positive', 'Negative', 'Neutral'
# sizes = [positive_sentiment_percentage, negative_sentiment_percentage, neutral_sentiment_percentage]
# colors = ['gold', 'yellowgreen', 'lightcoral']
# explode = (0.1, 0, 0)
# plt.pie(sizes, explode=explode, labels=labels, colors=colors, autopct='%1.1f%%', shadow=True, startangle=140)
# plt.plot()
if os.path.exists('metrics.csv'):
master_df.to_csv('metrics.csv', mode='a', header=False, index=False)
else:
master_df.to_csv('metrics.csv', index=False)
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import pathlib
import urllib.request
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.transforms
import scipy.ndimage.measurements
import scipy.interpolate
import scipy.optimize
import imageio
import pymedphys._mocks.profiles
import pymedphys._wlutz.findfield
import pymedphys._wlutz.createaxis
import pymedphys._wlutz.interppoints
image_name = '00000B94.png'
image_path = pathlib.Path('.').resolve().parent.joinpath('data', 'images', 'lossless', image_name)
if not image_path.exists():
url = f'https://zenodo.org/record/3520266/files/{image_name}?download=1'
urllib.request.urlretrieve(url, image_path)
img = imageio.imread(image_path)
img = img[:, 1:-1]
assert img.dtype == np.dtype('uint16')
assert np.shape(img) == (1024, 1022)
shape = np.shape(img)
x_axis = np.arange(-shape[1]/2, shape[1]/2)/4
y_axis = np.arange(-shape[0]/2, shape[0]/2)/4
np.shape(x_axis)
plt.imshow(img)
plt.xlim([400, 600])
plt.ylim([600, 400])
plt.colorbar()
scale_and_flip = 1 - img[::-1,:] / 2**16
plt.contourf(x_axis, y_axis, scale_and_flip, 30)
plt.axis('equal')
plt.xlim([-25, 25])
plt.ylim([-25, 25])
plt.colorbar()
interpolation = scipy.interpolate.RectBivariateSpline(x_axis, y_axis, scale_and_flip.T, kx=1, ky=1)
x_axis[:, None], y_axis[None, :]
interpolated = np.swapaxes(interpolation(x_axis, y_axis), 0, 1)
np.shape(interpolated)
np.allclose(interpolated, scale_and_flip)
plt.contourf(x_axis, y_axis, interpolated, 30)
plt.axis('equal')
plt.xlim([-25, 25])
plt.ylim([-25, 25])
plt.colorbar()
xx, yy = np.meshgrid(x_axis, y_axis)
np.shape(y_axis[:, None])
%timeit np.swapaxes(interpolation(x_axis, y_axis), 0, 1)
%timeit interpolation.ev(np.ravel(xx), np.ravel(yy))
def create_interpolated_field(x, y, img):
interpolation = scipy.interpolate.RectBivariateSpline(x, y, img.T, kx=1, ky=1)
def field(x, y):
if np.shape(x) != np.shape(y):
raise ValueError("x and y required to be the same shape")
result = interpolation.ev(np.ravel(x), np.ravel(y))
result.shape = x.shape
return result
return field
# field = create_interpolated_field(x, y, img)
interpolated = interpolation.ev(np.ravel(xx), np.ravel(yy))
interpolated.shape = xx.shape
plt.contourf(x_axis, y_axis, interpolated, 30)
plt.axis('equal')
plt.xlim([-25, 25])
plt.ylim([-25, 25])
plt.colorbar()
field = pymedphys._mocks.profiles.create_rectangular_field_function([0,0], [10,10], 1, 0)
np.shape(field(x_axis[:, None], y_axis[None, :]))
x_centre=-16.45827370949386
y_centre=0.0
x_edge=16.73970349084719
y_edge=15.002190875079133
penumbra=1.1355210572766254
actual_rotation=-88.07256317492568
actual_rotation = actual_rotation
edge_lengths = [x_edge, y_edge]
# penumbra = 0.5
actual_centre = [x_centre, y_centre]
field = pymedphys._mocks.profiles.create_rectangular_field_function(actual_centre, edge_lengths, penumbra, actual_rotation)
(180 - 90) % 90
x = np.arange(-50, 50, 0.1)
y = np.arange(-50, 50, 0.1)
xx, yy = np.meshgrid(x, y)
zz = field(xx, yy)
plt.pcolormesh(xx, yy, zz)
plt.axis('equal')
initial_centre = pymedphys._wlutz.findfield._initial_centre(x, y, zz)
centre, rotation = pymedphys._wlutz.findfield.field_finding_loop(field, edge_lengths, penumbra, initial_centre)
centre, rotation
def draw_by_diff(dx, dy, transform):
draw_x = np.cumsum(dx)
draw_y = np.cumsum(dy)
draw_x, draw_y = pymedphys._wlutz.interppoints.apply_transform(draw_x, draw_y, transform)
return draw_x, draw_y
transform = matplotlib.transforms.Affine2D()
transform.rotate_deg(-rotation)
transform.translate(*centre)
rotation_x_points = np.linspace(-edge_lengths[0]/2, edge_lengths[0]/2, 51)
rotation_y_points = np.linspace(-edge_lengths[1]/2, edge_lengths[1]/2, 61)
rot_xx_points, rot_yy_points = np.meshgrid(rotation_x_points, rotation_y_points)
rot_xx_points, rot_yy_points = pymedphys._wlutz.interppoints.apply_transform(rot_xx_points, rot_yy_points, transform)
rect_dx = [-edge_lengths[0]/2, 0, edge_lengths[0], 0, -edge_lengths[0]]
rect_dy = [-edge_lengths[1]/2, edge_lengths[1], 0, -edge_lengths[1], 0]
rect_crosshair_dx = [-edge_lengths[0]/2, edge_lengths[0], -edge_lengths[0], edge_lengths[0]]
rect_crosshair_dy = [-edge_lengths[1]/2, edge_lengths[1], 0, -edge_lengths[1]]
plt.figure(figsize=(10,10))
plt.pcolormesh(xx, yy, zz)
plt.plot(*draw_by_diff(rect_dx, rect_dy, transform), 'k', lw=2)
plt.plot(*draw_by_diff(rect_crosshair_dx, rect_crosshair_dy, transform), 'k', lw=0.5)
# plt.plot(rot_xx_points, rot_yy_points, '.')
plt.scatter(centre[0], centre[1], c='r', s=1)
plt.axis('equal')
np.allclose(actual_centre, centre)
np.allclose(actual_rotation, rotation)
```
| github_jupyter |
# RadarCOVID-Report
## Data Extraction
```
import datetime
import json
import logging
import os
import shutil
import tempfile
import textwrap
import uuid
import matplotlib.pyplot as plt
import matplotlib.ticker
import numpy as np
import pandas as pd
import pycountry
import retry
import seaborn as sns
%matplotlib inline
current_working_directory = os.environ.get("PWD")
if current_working_directory:
os.chdir(current_working_directory)
sns.set()
matplotlib.rcParams["figure.figsize"] = (15, 6)
extraction_datetime = datetime.datetime.utcnow()
extraction_date = extraction_datetime.strftime("%Y-%m-%d")
extraction_previous_datetime = extraction_datetime - datetime.timedelta(days=1)
extraction_previous_date = extraction_previous_datetime.strftime("%Y-%m-%d")
extraction_date_with_hour = datetime.datetime.utcnow().strftime("%Y-%m-%d@%H")
current_hour = datetime.datetime.utcnow().hour
are_today_results_partial = current_hour != 23
```
### Constants
```
from Modules.ExposureNotification import exposure_notification_io
spain_region_country_code = "ES"
germany_region_country_code = "DE"
default_backend_identifier = spain_region_country_code
backend_generation_days = 7 * 2
daily_summary_days = 7 * 4 * 3
daily_plot_days = 7 * 4
tek_dumps_load_limit = daily_summary_days + 1
```
### Parameters
```
environment_backend_identifier = os.environ.get("RADARCOVID_REPORT__BACKEND_IDENTIFIER")
if environment_backend_identifier:
report_backend_identifier = environment_backend_identifier
else:
report_backend_identifier = default_backend_identifier
report_backend_identifier
environment_enable_multi_backend_download = \
os.environ.get("RADARCOVID_REPORT__ENABLE_MULTI_BACKEND_DOWNLOAD")
if environment_enable_multi_backend_download:
report_backend_identifiers = None
else:
report_backend_identifiers = [report_backend_identifier]
report_backend_identifiers
environment_invalid_shared_diagnoses_dates = \
os.environ.get("RADARCOVID_REPORT__INVALID_SHARED_DIAGNOSES_DATES")
if environment_invalid_shared_diagnoses_dates:
invalid_shared_diagnoses_dates = environment_invalid_shared_diagnoses_dates.split(",")
else:
invalid_shared_diagnoses_dates = []
invalid_shared_diagnoses_dates
```
### COVID-19 Cases
```
report_backend_client = \
exposure_notification_io.get_backend_client_with_identifier(
backend_identifier=report_backend_identifier)
@retry.retry(tries=10, delay=10, backoff=1.1, jitter=(0, 10))
def download_cases_dataframe():
return pd.read_csv("https://raw.githubusercontent.com/owid/covid-19-data/master/public/data/owid-covid-data.csv")
confirmed_df_ = download_cases_dataframe()
confirmed_df_.iloc[0]
confirmed_df = confirmed_df_.copy()
confirmed_df = confirmed_df[["date", "new_cases", "iso_code"]]
confirmed_df.rename(
columns={
"date": "sample_date",
"iso_code": "country_code",
},
inplace=True)
def convert_iso_alpha_3_to_alpha_2(x):
try:
return pycountry.countries.get(alpha_3=x).alpha_2
except Exception as e:
logging.info(f"Error converting country ISO Alpha 3 code '{x}': {repr(e)}")
return None
confirmed_df["country_code"] = confirmed_df.country_code.apply(convert_iso_alpha_3_to_alpha_2)
confirmed_df.dropna(inplace=True)
confirmed_df["sample_date"] = pd.to_datetime(confirmed_df.sample_date, dayfirst=True)
confirmed_df["sample_date"] = confirmed_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_df.sort_values("sample_date", inplace=True)
confirmed_df.tail()
confirmed_days = pd.date_range(
start=confirmed_df.iloc[0].sample_date,
end=extraction_datetime)
confirmed_days_df = pd.DataFrame(data=confirmed_days, columns=["sample_date"])
confirmed_days_df["sample_date_string"] = \
confirmed_days_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_days_df.tail()
def sort_source_regions_for_display(source_regions: list) -> list:
if report_backend_identifier in source_regions:
source_regions = [report_backend_identifier] + \
list(sorted(set(source_regions).difference([report_backend_identifier])))
else:
source_regions = list(sorted(source_regions))
return source_regions
report_source_regions = report_backend_client.source_regions_for_date(
date=extraction_datetime.date())
report_source_regions = sort_source_regions_for_display(
source_regions=report_source_regions)
report_source_regions
def get_cases_dataframe(source_regions_for_date_function, columns_suffix=None):
source_regions_at_date_df = confirmed_days_df.copy()
source_regions_at_date_df["source_regions_at_date"] = \
source_regions_at_date_df.sample_date.apply(
lambda x: source_regions_for_date_function(date=x))
source_regions_at_date_df.sort_values("sample_date", inplace=True)
source_regions_at_date_df["_source_regions_group"] = source_regions_at_date_df. \
source_regions_at_date.apply(lambda x: ",".join(sort_source_regions_for_display(x)))
source_regions_at_date_df.tail()
#%%
source_regions_for_summary_df_ = \
source_regions_at_date_df[["sample_date", "_source_regions_group"]].copy()
source_regions_for_summary_df_.rename(columns={"_source_regions_group": "source_regions"}, inplace=True)
source_regions_for_summary_df_.tail()
#%%
confirmed_output_columns = ["sample_date", "new_cases", "covid_cases"]
confirmed_output_df = pd.DataFrame(columns=confirmed_output_columns)
for source_regions_group, source_regions_group_series in \
source_regions_at_date_df.groupby("_source_regions_group"):
source_regions_set = set(source_regions_group.split(","))
confirmed_source_regions_set_df = \
confirmed_df[confirmed_df.country_code.isin(source_regions_set)].copy()
confirmed_source_regions_group_df = \
confirmed_source_regions_set_df.groupby("sample_date").new_cases.sum() \
.reset_index().sort_values("sample_date")
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df.merge(
confirmed_days_df[["sample_date_string"]].rename(
columns={"sample_date_string": "sample_date"}),
how="right")
confirmed_source_regions_group_df["new_cases"] = \
confirmed_source_regions_group_df["new_cases"].clip(lower=0)
confirmed_source_regions_group_df["covid_cases"] = \
confirmed_source_regions_group_df.new_cases.rolling(7, min_periods=0).mean().round()
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df[confirmed_output_columns]
confirmed_source_regions_group_df = confirmed_source_regions_group_df.replace(0, np.nan)
confirmed_source_regions_group_df.fillna(method="ffill", inplace=True)
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df[
confirmed_source_regions_group_df.sample_date.isin(
source_regions_group_series.sample_date_string)]
confirmed_output_df = confirmed_output_df.append(confirmed_source_regions_group_df)
result_df = confirmed_output_df.copy()
result_df.tail()
#%%
result_df.rename(columns={"sample_date": "sample_date_string"}, inplace=True)
result_df = confirmed_days_df[["sample_date_string"]].merge(result_df, how="left")
result_df.sort_values("sample_date_string", inplace=True)
result_df.fillna(method="ffill", inplace=True)
result_df.tail()
#%%
result_df[["new_cases", "covid_cases"]].plot()
if columns_suffix:
result_df.rename(
columns={
"new_cases": "new_cases_" + columns_suffix,
"covid_cases": "covid_cases_" + columns_suffix},
inplace=True)
return result_df, source_regions_for_summary_df_
confirmed_eu_df, source_regions_for_summary_df = get_cases_dataframe(
report_backend_client.source_regions_for_date)
confirmed_es_df, _ = get_cases_dataframe(
lambda date: [spain_region_country_code],
columns_suffix=spain_region_country_code.lower())
```
### Extract API TEKs
```
raw_zip_path_prefix = "Data/TEKs/Raw/"
base_backend_identifiers = [report_backend_identifier]
multi_backend_exposure_keys_df = \
exposure_notification_io.download_exposure_keys_from_backends(
backend_identifiers=report_backend_identifiers,
generation_days=backend_generation_days,
fail_on_error_backend_identifiers=base_backend_identifiers,
save_raw_zip_path_prefix=raw_zip_path_prefix)
multi_backend_exposure_keys_df["region"] = multi_backend_exposure_keys_df["backend_identifier"]
multi_backend_exposure_keys_df.rename(
columns={
"generation_datetime": "sample_datetime",
"generation_date_string": "sample_date_string",
},
inplace=True)
multi_backend_exposure_keys_df.head()
early_teks_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.rolling_period < 144].copy()
early_teks_df["rolling_period_in_hours"] = early_teks_df.rolling_period / 6
early_teks_df[early_teks_df.sample_date_string != extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
early_teks_df[early_teks_df.sample_date_string == extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
multi_backend_exposure_keys_df = multi_backend_exposure_keys_df[[
"sample_date_string", "region", "key_data"]]
multi_backend_exposure_keys_df.head()
active_regions = \
multi_backend_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
active_regions
multi_backend_summary_df = multi_backend_exposure_keys_df.groupby(
["sample_date_string", "region"]).key_data.nunique().reset_index() \
.pivot(index="sample_date_string", columns="region") \
.sort_index(ascending=False)
multi_backend_summary_df.rename(
columns={"key_data": "shared_teks_by_generation_date"},
inplace=True)
multi_backend_summary_df.rename_axis("sample_date", inplace=True)
multi_backend_summary_df = multi_backend_summary_df.fillna(0).astype(int)
multi_backend_summary_df = multi_backend_summary_df.head(backend_generation_days)
multi_backend_summary_df.head()
def compute_keys_cross_sharing(x):
teks_x = x.key_data_x.item()
common_teks = set(teks_x).intersection(x.key_data_y.item())
common_teks_fraction = len(common_teks) / len(teks_x)
return pd.Series(dict(
common_teks=common_teks,
common_teks_fraction=common_teks_fraction,
))
multi_backend_exposure_keys_by_region_df = \
multi_backend_exposure_keys_df.groupby("region").key_data.unique().reset_index()
multi_backend_exposure_keys_by_region_df["_merge"] = True
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_df.merge(
multi_backend_exposure_keys_by_region_df, on="_merge")
multi_backend_exposure_keys_by_region_combination_df.drop(
columns=["_merge"], inplace=True)
if multi_backend_exposure_keys_by_region_combination_df.region_x.nunique() > 1:
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_combination_df[
multi_backend_exposure_keys_by_region_combination_df.region_x !=
multi_backend_exposure_keys_by_region_combination_df.region_y]
multi_backend_exposure_keys_cross_sharing_df = \
multi_backend_exposure_keys_by_region_combination_df \
.groupby(["region_x", "region_y"]) \
.apply(compute_keys_cross_sharing) \
.reset_index()
multi_backend_cross_sharing_summary_df = \
multi_backend_exposure_keys_cross_sharing_df.pivot_table(
values=["common_teks_fraction"],
columns="region_x",
index="region_y",
aggfunc=lambda x: x.item())
multi_backend_cross_sharing_summary_df
multi_backend_without_active_region_exposure_keys_df = \
multi_backend_exposure_keys_df[multi_backend_exposure_keys_df.region != report_backend_identifier]
multi_backend_without_active_region = \
multi_backend_without_active_region_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
multi_backend_without_active_region
exposure_keys_summary_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.region == report_backend_identifier]
exposure_keys_summary_df.drop(columns=["region"], inplace=True)
exposure_keys_summary_df = \
exposure_keys_summary_df.groupby(["sample_date_string"]).key_data.nunique().to_frame()
exposure_keys_summary_df = \
exposure_keys_summary_df.reset_index().set_index("sample_date_string")
exposure_keys_summary_df.sort_index(ascending=False, inplace=True)
exposure_keys_summary_df.rename(columns={"key_data": "shared_teks_by_generation_date"}, inplace=True)
exposure_keys_summary_df.head()
```
### Dump API TEKs
```
tek_list_df = multi_backend_exposure_keys_df[
["sample_date_string", "region", "key_data"]].copy()
tek_list_df["key_data"] = tek_list_df["key_data"].apply(str)
tek_list_df.rename(columns={
"sample_date_string": "sample_date",
"key_data": "tek_list"}, inplace=True)
tek_list_df = tek_list_df.groupby(
["sample_date", "region"]).tek_list.unique().reset_index()
tek_list_df["extraction_date"] = extraction_date
tek_list_df["extraction_date_with_hour"] = extraction_date_with_hour
tek_list_path_prefix = "Data/TEKs/"
tek_list_current_path = tek_list_path_prefix + f"/Current/RadarCOVID-TEKs.json"
tek_list_daily_path = tek_list_path_prefix + f"Daily/RadarCOVID-TEKs-{extraction_date}.json"
tek_list_hourly_path = tek_list_path_prefix + f"Hourly/RadarCOVID-TEKs-{extraction_date_with_hour}.json"
for path in [tek_list_current_path, tek_list_daily_path, tek_list_hourly_path]:
os.makedirs(os.path.dirname(path), exist_ok=True)
tek_list_base_df = tek_list_df[tek_list_df.region == report_backend_identifier]
tek_list_base_df.drop(columns=["extraction_date", "extraction_date_with_hour"]).to_json(
tek_list_current_path,
lines=True, orient="records")
tek_list_base_df.drop(columns=["extraction_date_with_hour"]).to_json(
tek_list_daily_path,
lines=True, orient="records")
tek_list_base_df.to_json(
tek_list_hourly_path,
lines=True, orient="records")
tek_list_base_df.head()
```
### Load TEK Dumps
```
import glob
def load_extracted_teks(mode, region=None, limit=None) -> pd.DataFrame:
extracted_teks_df = pd.DataFrame(columns=["region"])
file_paths = list(reversed(sorted(glob.glob(tek_list_path_prefix + mode + "/RadarCOVID-TEKs-*.json"))))
if limit:
file_paths = file_paths[:limit]
for file_path in file_paths:
logging.info(f"Loading TEKs from '{file_path}'...")
iteration_extracted_teks_df = pd.read_json(file_path, lines=True)
extracted_teks_df = extracted_teks_df.append(
iteration_extracted_teks_df, sort=False)
extracted_teks_df["region"] = \
extracted_teks_df.region.fillna(spain_region_country_code).copy()
if region:
extracted_teks_df = \
extracted_teks_df[extracted_teks_df.region == region]
return extracted_teks_df
daily_extracted_teks_df = load_extracted_teks(
mode="Daily",
region=report_backend_identifier,
limit=tek_dumps_load_limit)
daily_extracted_teks_df.head()
exposure_keys_summary_df_ = daily_extracted_teks_df \
.sort_values("extraction_date", ascending=False) \
.groupby("sample_date").tek_list.first() \
.to_frame()
exposure_keys_summary_df_.index.name = "sample_date_string"
exposure_keys_summary_df_["tek_list"] = \
exposure_keys_summary_df_.tek_list.apply(len)
exposure_keys_summary_df_ = exposure_keys_summary_df_ \
.rename(columns={"tek_list": "shared_teks_by_generation_date"}) \
.sort_index(ascending=False)
exposure_keys_summary_df = exposure_keys_summary_df_
exposure_keys_summary_df.head()
```
### Daily New TEKs
```
tek_list_df = daily_extracted_teks_df.groupby("extraction_date").tek_list.apply(
lambda x: set(sum(x, []))).reset_index()
tek_list_df = tek_list_df.set_index("extraction_date").sort_index(ascending=True)
tek_list_df.head()
def compute_teks_by_generation_and_upload_date(date):
day_new_teks_set_df = tek_list_df.copy().diff()
try:
day_new_teks_set = day_new_teks_set_df[
day_new_teks_set_df.index == date].tek_list.item()
except ValueError:
day_new_teks_set = None
if pd.isna(day_new_teks_set):
day_new_teks_set = set()
day_new_teks_df = daily_extracted_teks_df[
daily_extracted_teks_df.extraction_date == date].copy()
day_new_teks_df["shared_teks"] = \
day_new_teks_df.tek_list.apply(lambda x: set(x).intersection(day_new_teks_set))
day_new_teks_df["shared_teks"] = \
day_new_teks_df.shared_teks.apply(len)
day_new_teks_df["upload_date"] = date
day_new_teks_df.rename(columns={"sample_date": "generation_date"}, inplace=True)
day_new_teks_df = day_new_teks_df[
["upload_date", "generation_date", "shared_teks"]]
day_new_teks_df["generation_to_upload_days"] = \
(pd.to_datetime(day_new_teks_df.upload_date) -
pd.to_datetime(day_new_teks_df.generation_date)).dt.days
day_new_teks_df = day_new_teks_df[day_new_teks_df.shared_teks > 0]
return day_new_teks_df
shared_teks_generation_to_upload_df = pd.DataFrame()
for upload_date in daily_extracted_teks_df.extraction_date.unique():
shared_teks_generation_to_upload_df = \
shared_teks_generation_to_upload_df.append(
compute_teks_by_generation_and_upload_date(date=upload_date))
shared_teks_generation_to_upload_df \
.sort_values(["upload_date", "generation_date"], ascending=False, inplace=True)
shared_teks_generation_to_upload_df.tail()
today_new_teks_df = \
shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.upload_date == extraction_date].copy()
today_new_teks_df.tail()
if not today_new_teks_df.empty:
today_new_teks_df.set_index("generation_to_upload_days") \
.sort_index().shared_teks.plot.bar()
generation_to_upload_period_pivot_df = \
shared_teks_generation_to_upload_df[
["upload_date", "generation_to_upload_days", "shared_teks"]] \
.pivot(index="upload_date", columns="generation_to_upload_days") \
.sort_index(ascending=False).fillna(0).astype(int) \
.droplevel(level=0, axis=1)
generation_to_upload_period_pivot_df.head()
new_tek_df = tek_list_df.diff().tek_list.apply(
lambda x: len(x) if not pd.isna(x) else None).to_frame().reset_index()
new_tek_df.rename(columns={
"tek_list": "shared_teks_by_upload_date",
"extraction_date": "sample_date_string",}, inplace=True)
new_tek_df.tail()
shared_teks_uploaded_on_generation_date_df = shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.generation_to_upload_days == 0] \
[["upload_date", "shared_teks"]].rename(
columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_teks_uploaded_on_generation_date",
})
shared_teks_uploaded_on_generation_date_df.head()
estimated_shared_diagnoses_df = shared_teks_generation_to_upload_df \
.groupby(["upload_date"]).shared_teks.max().reset_index() \
.sort_values(["upload_date"], ascending=False) \
.rename(columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_diagnoses",
})
invalid_shared_diagnoses_dates_mask = \
estimated_shared_diagnoses_df.sample_date_string.isin(invalid_shared_diagnoses_dates)
estimated_shared_diagnoses_df[invalid_shared_diagnoses_dates_mask] = 0
estimated_shared_diagnoses_df.head()
```
### Hourly New TEKs
```
hourly_extracted_teks_df = load_extracted_teks(
mode="Hourly", region=report_backend_identifier, limit=25)
hourly_extracted_teks_df.head()
hourly_new_tek_count_df = hourly_extracted_teks_df \
.groupby("extraction_date_with_hour").tek_list. \
apply(lambda x: set(sum(x, []))).reset_index().copy()
hourly_new_tek_count_df = hourly_new_tek_count_df.set_index("extraction_date_with_hour") \
.sort_index(ascending=True)
hourly_new_tek_count_df["new_tek_list"] = hourly_new_tek_count_df.tek_list.diff()
hourly_new_tek_count_df["new_tek_count"] = hourly_new_tek_count_df.new_tek_list.apply(
lambda x: len(x) if not pd.isna(x) else 0)
hourly_new_tek_count_df.rename(columns={
"new_tek_count": "shared_teks_by_upload_date"}, inplace=True)
hourly_new_tek_count_df = hourly_new_tek_count_df.reset_index()[[
"extraction_date_with_hour", "shared_teks_by_upload_date"]]
hourly_new_tek_count_df.head()
hourly_summary_df = hourly_new_tek_count_df.copy()
hourly_summary_df.set_index("extraction_date_with_hour", inplace=True)
hourly_summary_df = hourly_summary_df.fillna(0).astype(int).reset_index()
hourly_summary_df["datetime_utc"] = pd.to_datetime(
hourly_summary_df.extraction_date_with_hour, format="%Y-%m-%d@%H")
hourly_summary_df.set_index("datetime_utc", inplace=True)
hourly_summary_df = hourly_summary_df.tail(-1)
hourly_summary_df.head()
```
### Official Statistics
```
import requests
import pandas.io.json
official_stats_response = requests.get("https://radarcovid.covid19.gob.es/kpi/statistics/basics")
official_stats_response.raise_for_status()
official_stats_df_ = pandas.io.json.json_normalize(official_stats_response.json())
official_stats_df = official_stats_df_.copy()
official_stats_df["date"] = pd.to_datetime(official_stats_df["date"], dayfirst=True)
official_stats_df.head()
official_stats_column_map = {
"date": "sample_date",
"applicationsDownloads.totalAcummulated": "app_downloads_es_accumulated",
"communicatedContagions.totalAcummulated": "shared_diagnoses_es_accumulated",
}
accumulated_suffix = "_accumulated"
accumulated_values_columns = \
list(filter(lambda x: x.endswith(accumulated_suffix), official_stats_column_map.values()))
interpolated_values_columns = \
list(map(lambda x: x[:-len(accumulated_suffix)], accumulated_values_columns))
official_stats_df = \
official_stats_df[official_stats_column_map.keys()] \
.rename(columns=official_stats_column_map)
official_stats_df["extraction_date"] = extraction_date
official_stats_df.head()
official_stats_path = "Data/Statistics/Current/RadarCOVID-Statistics.json"
previous_official_stats_df = pd.read_json(official_stats_path, orient="records", lines=True)
previous_official_stats_df["sample_date"] = pd.to_datetime(previous_official_stats_df["sample_date"], dayfirst=True)
official_stats_df = official_stats_df.append(previous_official_stats_df)
official_stats_df.head()
official_stats_df = official_stats_df[~(official_stats_df.shared_diagnoses_es_accumulated == 0)]
official_stats_df.sort_values("extraction_date", ascending=False, inplace=True)
official_stats_df.drop_duplicates(subset=["sample_date"], keep="first", inplace=True)
official_stats_df.head()
official_stats_stored_df = official_stats_df.copy()
official_stats_stored_df["sample_date"] = official_stats_stored_df.sample_date.dt.strftime("%Y-%m-%d")
official_stats_stored_df.to_json(official_stats_path, orient="records", lines=True)
official_stats_df.drop(columns=["extraction_date"], inplace=True)
official_stats_df = confirmed_days_df.merge(official_stats_df, how="left")
official_stats_df.sort_values("sample_date", ascending=False, inplace=True)
official_stats_df.head()
official_stats_df[accumulated_values_columns] = \
official_stats_df[accumulated_values_columns] \
.astype(float).interpolate(limit_area="inside")
official_stats_df[interpolated_values_columns] = \
official_stats_df[accumulated_values_columns].diff(periods=-1)
official_stats_df.drop(columns="sample_date", inplace=True)
official_stats_df.head()
```
### Data Merge
```
result_summary_df = exposure_keys_summary_df.merge(
new_tek_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
shared_teks_uploaded_on_generation_date_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
estimated_shared_diagnoses_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
official_stats_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = confirmed_eu_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df = confirmed_es_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df["sample_date"] = pd.to_datetime(result_summary_df.sample_date_string)
result_summary_df = result_summary_df.merge(source_regions_for_summary_df, how="left")
result_summary_df.set_index(["sample_date", "source_regions"], inplace=True)
result_summary_df.drop(columns=["sample_date_string"], inplace=True)
result_summary_df.sort_index(ascending=False, inplace=True)
result_summary_df.head()
with pd.option_context("mode.use_inf_as_na", True):
result_summary_df = result_summary_df.fillna(0).astype(int)
result_summary_df["teks_per_shared_diagnosis"] = \
(result_summary_df.shared_teks_by_upload_date / result_summary_df.shared_diagnoses).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case"] = \
(result_summary_df.shared_diagnoses / result_summary_df.covid_cases).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case_es"] = \
(result_summary_df.shared_diagnoses_es / result_summary_df.covid_cases_es).fillna(0)
result_summary_df.head(daily_plot_days)
def compute_aggregated_results_summary(days) -> pd.DataFrame:
aggregated_result_summary_df = result_summary_df.copy()
aggregated_result_summary_df["covid_cases_for_ratio"] = \
aggregated_result_summary_df.covid_cases.mask(
aggregated_result_summary_df.shared_diagnoses == 0, 0)
aggregated_result_summary_df["covid_cases_for_ratio_es"] = \
aggregated_result_summary_df.covid_cases_es.mask(
aggregated_result_summary_df.shared_diagnoses_es == 0, 0)
aggregated_result_summary_df = aggregated_result_summary_df \
.sort_index(ascending=True).fillna(0).rolling(days).agg({
"covid_cases": "sum",
"covid_cases_es": "sum",
"covid_cases_for_ratio": "sum",
"covid_cases_for_ratio_es": "sum",
"shared_teks_by_generation_date": "sum",
"shared_teks_by_upload_date": "sum",
"shared_diagnoses": "sum",
"shared_diagnoses_es": "sum",
}).sort_index(ascending=False)
with pd.option_context("mode.use_inf_as_na", True):
aggregated_result_summary_df = aggregated_result_summary_df.fillna(0).astype(int)
aggregated_result_summary_df["teks_per_shared_diagnosis"] = \
(aggregated_result_summary_df.shared_teks_by_upload_date /
aggregated_result_summary_df.covid_cases_for_ratio).fillna(0)
aggregated_result_summary_df["shared_diagnoses_per_covid_case"] = \
(aggregated_result_summary_df.shared_diagnoses /
aggregated_result_summary_df.covid_cases_for_ratio).fillna(0)
aggregated_result_summary_df["shared_diagnoses_per_covid_case_es"] = \
(aggregated_result_summary_df.shared_diagnoses_es /
aggregated_result_summary_df.covid_cases_for_ratio_es).fillna(0)
return aggregated_result_summary_df
aggregated_result_with_7_days_window_summary_df = compute_aggregated_results_summary(days=7)
aggregated_result_with_7_days_window_summary_df.head()
last_7_days_summary = aggregated_result_with_7_days_window_summary_df.to_dict(orient="records")[1]
last_7_days_summary
aggregated_result_with_14_days_window_summary_df = compute_aggregated_results_summary(days=13)
last_14_days_summary = aggregated_result_with_14_days_window_summary_df.to_dict(orient="records")[1]
last_14_days_summary
```
## Report Results
```
display_column_name_mapping = {
"sample_date": "Sample\u00A0Date\u00A0(UTC)",
"source_regions": "Source Countries",
"datetime_utc": "Timestamp (UTC)",
"upload_date": "Upload Date (UTC)",
"generation_to_upload_days": "Generation to Upload Period in Days",
"region": "Backend",
"region_x": "Backend\u00A0(A)",
"region_y": "Backend\u00A0(B)",
"common_teks": "Common TEKs Shared Between Backends",
"common_teks_fraction": "Fraction of TEKs in Backend (A) Available in Backend (B)",
"covid_cases": "COVID-19 Cases (Source Countries)",
"shared_teks_by_generation_date": "Shared TEKs by Generation Date (Source Countries)",
"shared_teks_by_upload_date": "Shared TEKs by Upload Date (Source Countries)",
"shared_teks_uploaded_on_generation_date": "Shared TEKs Uploaded on Generation Date (Source Countries)",
"shared_diagnoses": "Shared Diagnoses (Source Countries – Estimation)",
"teks_per_shared_diagnosis": "TEKs Uploaded per Shared Diagnosis (Source Countries)",
"shared_diagnoses_per_covid_case": "Usage Ratio (Source Countries)",
"covid_cases_es": "COVID-19 Cases (Spain)",
"app_downloads_es": "App Downloads (Spain – Official)",
"shared_diagnoses_es": "Shared Diagnoses (Spain – Official)",
"shared_diagnoses_per_covid_case_es": "Usage Ratio (Spain)",
}
summary_columns = [
"covid_cases",
"shared_teks_by_generation_date",
"shared_teks_by_upload_date",
"shared_teks_uploaded_on_generation_date",
"shared_diagnoses",
"teks_per_shared_diagnosis",
"shared_diagnoses_per_covid_case",
"covid_cases_es",
"app_downloads_es",
"shared_diagnoses_es",
"shared_diagnoses_per_covid_case_es",
]
summary_percentage_columns= [
"shared_diagnoses_per_covid_case_es",
"shared_diagnoses_per_covid_case",
]
```
### Daily Summary Table
```
result_summary_df_ = result_summary_df.copy()
result_summary_df = result_summary_df[summary_columns]
result_summary_with_display_names_df = result_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
result_summary_with_display_names_df
```
### Daily Summary Plots
```
result_plot_summary_df = result_summary_df.head(daily_plot_days)[summary_columns] \
.droplevel(level=["source_regions"]) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
summary_ax_list = result_plot_summary_df.sort_index(ascending=True).plot.bar(
title=f"Daily Summary",
rot=45, subplots=True, figsize=(15, 30), legend=False)
ax_ = summary_ax_list[0]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.95)
_ = ax_.set_xticklabels(sorted(result_plot_summary_df.index.strftime("%Y-%m-%d").tolist()))
for percentage_column in summary_percentage_columns:
percentage_column_index = summary_columns.index(percentage_column)
summary_ax_list[percentage_column_index].yaxis \
.set_major_formatter(matplotlib.ticker.PercentFormatter(1.0))
```
### Daily Generation to Upload Period Table
```
display_generation_to_upload_period_pivot_df = \
generation_to_upload_period_pivot_df \
.head(backend_generation_days)
display_generation_to_upload_period_pivot_df \
.head(backend_generation_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping)
fig, generation_to_upload_period_pivot_table_ax = plt.subplots(
figsize=(12, 1 + 0.6 * len(display_generation_to_upload_period_pivot_df)))
generation_to_upload_period_pivot_table_ax.set_title(
"Shared TEKs Generation to Upload Period Table")
sns.heatmap(
data=display_generation_to_upload_period_pivot_df
.rename_axis(columns=display_column_name_mapping)
.rename_axis(index=display_column_name_mapping),
fmt=".0f",
annot=True,
ax=generation_to_upload_period_pivot_table_ax)
generation_to_upload_period_pivot_table_ax.get_figure().tight_layout()
```
### Hourly Summary Plots
```
hourly_summary_ax_list = hourly_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.plot.bar(
title=f"Last 24h Summary",
rot=45, subplots=True, legend=False)
ax_ = hourly_summary_ax_list[-1]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.9)
_ = ax_.set_xticklabels(sorted(hourly_summary_df.index.strftime("%Y-%m-%d@%H").tolist()))
```
### Publish Results
```
github_repository = os.environ.get("GITHUB_REPOSITORY")
if github_repository is None:
github_repository = "pvieito/Radar-STATS"
github_project_base_url = "https://github.com/" + github_repository
display_formatters = {
display_column_name_mapping["teks_per_shared_diagnosis"]: lambda x: f"{x:.2f}" if x != 0 else "",
display_column_name_mapping["shared_diagnoses_per_covid_case"]: lambda x: f"{x:.2%}" if x != 0 else "",
display_column_name_mapping["shared_diagnoses_per_covid_case_es"]: lambda x: f"{x:.2%}" if x != 0 else "",
}
general_columns = \
list(filter(lambda x: x not in display_formatters, display_column_name_mapping.values()))
general_formatter = lambda x: f"{x}" if x != 0 else ""
display_formatters.update(dict(map(lambda x: (x, general_formatter), general_columns)))
daily_summary_table_html = result_summary_with_display_names_df \
.head(daily_plot_days) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.to_html(formatters=display_formatters)
multi_backend_summary_table_html = multi_backend_summary_df \
.head(daily_plot_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(formatters=display_formatters)
def format_multi_backend_cross_sharing_fraction(x):
if pd.isna(x):
return "-"
elif round(x * 100, 1) == 0:
return ""
else:
return f"{x:.1%}"
multi_backend_cross_sharing_summary_table_html = multi_backend_cross_sharing_summary_df \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(
classes="table-center",
formatters=display_formatters,
float_format=format_multi_backend_cross_sharing_fraction)
multi_backend_cross_sharing_summary_table_html = \
multi_backend_cross_sharing_summary_table_html \
.replace("<tr>","<tr style=\"text-align: center;\">")
extraction_date_result_summary_df = \
result_summary_df[result_summary_df.index.get_level_values("sample_date") == extraction_date]
extraction_date_result_hourly_summary_df = \
hourly_summary_df[hourly_summary_df.extraction_date_with_hour == extraction_date_with_hour]
covid_cases = \
extraction_date_result_summary_df.covid_cases.item()
shared_teks_by_generation_date = \
extraction_date_result_summary_df.shared_teks_by_generation_date.item()
shared_teks_by_upload_date = \
extraction_date_result_summary_df.shared_teks_by_upload_date.item()
shared_diagnoses = \
extraction_date_result_summary_df.shared_diagnoses.item()
teks_per_shared_diagnosis = \
extraction_date_result_summary_df.teks_per_shared_diagnosis.item()
shared_diagnoses_per_covid_case = \
extraction_date_result_summary_df.shared_diagnoses_per_covid_case.item()
shared_teks_by_upload_date_last_hour = \
extraction_date_result_hourly_summary_df.shared_teks_by_upload_date.sum().astype(int)
display_source_regions = ", ".join(report_source_regions)
if len(report_source_regions) == 1:
display_brief_source_regions = report_source_regions[0]
else:
display_brief_source_regions = f"{len(report_source_regions)} 🇪🇺"
def get_temporary_image_path() -> str:
return os.path.join(tempfile.gettempdir(), str(uuid.uuid4()) + ".png")
def save_temporary_plot_image(ax):
if isinstance(ax, np.ndarray):
ax = ax[0]
media_path = get_temporary_image_path()
ax.get_figure().savefig(media_path)
return media_path
def save_temporary_dataframe_image(df):
import dataframe_image as dfi
df = df.copy()
df_styler = df.style.format(display_formatters)
media_path = get_temporary_image_path()
dfi.export(df_styler, media_path)
return media_path
summary_plots_image_path = save_temporary_plot_image(
ax=summary_ax_list)
summary_table_image_path = save_temporary_dataframe_image(
df=result_summary_with_display_names_df)
hourly_summary_plots_image_path = save_temporary_plot_image(
ax=hourly_summary_ax_list)
multi_backend_summary_table_image_path = save_temporary_dataframe_image(
df=multi_backend_summary_df)
generation_to_upload_period_pivot_table_image_path = save_temporary_plot_image(
ax=generation_to_upload_period_pivot_table_ax)
```
### Save Results
```
report_resources_path_prefix = "Data/Resources/Current/RadarCOVID-Report-"
result_summary_df.to_csv(
report_resources_path_prefix + "Summary-Table.csv")
result_summary_df.to_html(
report_resources_path_prefix + "Summary-Table.html")
hourly_summary_df.to_csv(
report_resources_path_prefix + "Hourly-Summary-Table.csv")
multi_backend_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Summary-Table.csv")
multi_backend_cross_sharing_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Cross-Sharing-Summary-Table.csv")
generation_to_upload_period_pivot_df.to_csv(
report_resources_path_prefix + "Generation-Upload-Period-Table.csv")
_ = shutil.copyfile(
summary_plots_image_path,
report_resources_path_prefix + "Summary-Plots.png")
_ = shutil.copyfile(
summary_table_image_path,
report_resources_path_prefix + "Summary-Table.png")
_ = shutil.copyfile(
hourly_summary_plots_image_path,
report_resources_path_prefix + "Hourly-Summary-Plots.png")
_ = shutil.copyfile(
multi_backend_summary_table_image_path,
report_resources_path_prefix + "Multi-Backend-Summary-Table.png")
_ = shutil.copyfile(
generation_to_upload_period_pivot_table_image_path,
report_resources_path_prefix + "Generation-Upload-Period-Table.png")
```
### Publish Results as JSON
```
def generate_summary_api_results(df: pd.DataFrame) -> list:
api_df = df.reset_index().copy()
api_df["sample_date_string"] = \
api_df["sample_date"].dt.strftime("%Y-%m-%d")
api_df["source_regions"] = \
api_df["source_regions"].apply(lambda x: x.split(","))
return api_df.to_dict(orient="records")
summary_api_results = \
generate_summary_api_results(df=result_summary_df)
today_summary_api_results = \
generate_summary_api_results(df=extraction_date_result_summary_df)[0]
summary_results = dict(
backend_identifier=report_backend_identifier,
source_regions=report_source_regions,
extraction_datetime=extraction_datetime,
extraction_date=extraction_date,
extraction_date_with_hour=extraction_date_with_hour,
last_hour=dict(
shared_teks_by_upload_date=shared_teks_by_upload_date_last_hour,
shared_diagnoses=0,
),
today=today_summary_api_results,
last_7_days=last_7_days_summary,
last_14_days=last_14_days_summary,
daily_results=summary_api_results)
summary_results = \
json.loads(pd.Series([summary_results]).to_json(orient="records"))[0]
with open(report_resources_path_prefix + "Summary-Results.json", "w") as f:
json.dump(summary_results, f, indent=4)
```
### Publish on README
```
with open("Data/Templates/README.md", "r") as f:
readme_contents = f.read()
readme_contents = readme_contents.format(
extraction_date_with_hour=extraction_date_with_hour,
github_project_base_url=github_project_base_url,
daily_summary_table_html=daily_summary_table_html,
multi_backend_summary_table_html=multi_backend_summary_table_html,
multi_backend_cross_sharing_summary_table_html=multi_backend_cross_sharing_summary_table_html,
display_source_regions=display_source_regions)
with open("README.md", "w") as f:
f.write(readme_contents)
```
### Publish on Twitter
```
enable_share_to_twitter = os.environ.get("RADARCOVID_REPORT__ENABLE_PUBLISH_ON_TWITTER")
github_event_name = os.environ.get("GITHUB_EVENT_NAME")
if enable_share_to_twitter and github_event_name == "schedule" and \
(shared_teks_by_upload_date_last_hour or not are_today_results_partial):
import tweepy
twitter_api_auth_keys = os.environ["RADARCOVID_REPORT__TWITTER_API_AUTH_KEYS"]
twitter_api_auth_keys = twitter_api_auth_keys.split(":")
auth = tweepy.OAuthHandler(twitter_api_auth_keys[0], twitter_api_auth_keys[1])
auth.set_access_token(twitter_api_auth_keys[2], twitter_api_auth_keys[3])
api = tweepy.API(auth)
summary_plots_media = api.media_upload(summary_plots_image_path)
summary_table_media = api.media_upload(summary_table_image_path)
generation_to_upload_period_pivot_table_image_media = api.media_upload(generation_to_upload_period_pivot_table_image_path)
media_ids = [
summary_plots_media.media_id,
summary_table_media.media_id,
generation_to_upload_period_pivot_table_image_media.media_id,
]
if are_today_results_partial:
today_addendum = " (Partial)"
else:
today_addendum = ""
def format_shared_diagnoses_per_covid_case(value) -> str:
if value == 0:
return "–"
return f"≤{value:.2%}"
display_shared_diagnoses_per_covid_case = \
format_shared_diagnoses_per_covid_case(value=shared_diagnoses_per_covid_case)
display_last_14_days_shared_diagnoses_per_covid_case = \
format_shared_diagnoses_per_covid_case(value=last_14_days_summary["shared_diagnoses_per_covid_case"])
display_last_14_days_shared_diagnoses_per_covid_case_es = \
format_shared_diagnoses_per_covid_case(value=last_14_days_summary["shared_diagnoses_per_covid_case_es"])
status = textwrap.dedent(f"""
#RadarCOVID – {extraction_date_with_hour}
Today{today_addendum}:
- Uploaded TEKs: {shared_teks_by_upload_date:.0f} ({shared_teks_by_upload_date_last_hour:+d} last hour)
- Shared Diagnoses: ≤{shared_diagnoses:.0f}
- Usage Ratio: {display_shared_diagnoses_per_covid_case}
Last 14 Days:
- Usage Ratio (Estimation): {display_last_14_days_shared_diagnoses_per_covid_case}
- Usage Ratio (Official): {display_last_14_days_shared_diagnoses_per_covid_case_es}
Info: {github_project_base_url}#documentation
""")
status = status.encode(encoding="utf-8")
api.update_status(status=status, media_ids=media_ids)
```
| github_jupyter |
# TABLE OF CONTENTS:<a id='toc'></a>
These tips are focused on general Python tips I think that are good to know.
Please go through official documentation if you want more thorough examples.
<b>Topics:</b>
- <b>[Additional Operators](#op)</b>
- <b>[Global](#global)</b>
- <b>[Comparisons](#compare)</b>
- <b>[Enumerate](#enum)</b>
- <b>[Comprehension](#comp)</b>
- [List](#list)
- [Set](#set)
- [Dict](#dict)
```
# # Uncomment if you want to use inline pythontutor
# from IPython.display import IFrame
# IFrame('http://www.pythontutor.com/visualize.html#mode=display', height=1500, width=750)
```
# Additional Operators<a id="op"></a>
[Return to table of contents](#toc)
Operators besides you typical `+`, `-`, `/`, etc.
`~` Inversion, is the bitwise complement operator in python which essentially calculates `-x - 1`
`=` Assign value of right side of expression to left side operand `x = y + z`
`+=` Add AND: Add right side operand with left side operand and then assign to left operand `a+=b` `a=a+b`
`-=` Subtract AND: Subtract right operand from left operand and then assign to left operand `a-=b` `a=a-b`
`*=` Multiply AND: Multiply right operand with left operand and then assign to left operand `a*=b` `a=a*b`
`/=` Divide AND: Divide left operand with right operand and then assign to left operand `a/=b` `a=a/b`
`%=` Modulus AND: Takes modulus using left and right operands and assign result to left operand `a%=b` `a=a%b`
`//=` Divide(floor) AND: Divide left operand with right operand and then assign the value(floor) to left operand `a//=b` `a=a//b`
`**=` Exponent AND: Calculate exponent(raise power) value using operands and assign value to left operand `a**=b` `a=a**b`
`&=` Performs Bitwise AND on operands and assign value to left operand `a&=b` `a=a&b`
`|=` Performs Bitwise OR on operands and assign value to left operand `a|=b` `a=a|b`
`^=` Performs Bitwise xOR on operands and assign value to left operand `a^=b` `a=a^b`
`>>=` Performs Bitwise right shift on operands and assign value to left operand `a>>=b` `a=a>>b`
`<<=` Performs Bitwise left shift on operands and assign value to left operand `a <<= b` `a= a << b`
<b>Or assignment</b>
Assigning a variable based on another variable's assignment.
```
var = None
b = None or var
print(b)
var = 5
b = None or var
print(b)
```
# Global<a id="global"></a>
[Return to table of contents](#toc)
Global lets you access global variables. In this example the global variable `c` is outside of the functions scope.
```
# c = 1
# def add():
# c = c + 2
# print(c)
# add()
# # UnboundLocalError: local variable 'c' referenced before assignment
c = 1
def add():
global c
c = c + 2
print(c)
add()
```
# Comparisons<a id="compare"></a>
[Return to table of contents](#toc)
<b>Max<b>
```
# Find the max of an array/variables/values etc.
print(max(5, 2))
print(max([10,11,12,13]))
```
<b>Min<b>
```
# Find the min of an array/variables/values etc.
print(min(5, 2))
print(min([10,11,12,13]))
```
<b>float("inf")/float("-inf")<b>
```
# Setting a value to infinity or -infinity lets you have an easy comparion
print(float("inf") > 309840)
print(float("-inf") < -930984)
```
# Enumerate<a id="enum"></a>
[Return to table of contents](#toc)
This lets you take the element's index and use it as a variable.
```
pies = ["apple", "blueberry", "lemon"]
for num, i in enumerate(pies):
print(num, ":", i)
# For dictionaris this is what it would look like.
pies = {"pie1":"apple", "pie2":"blueberry", "pie3":"lemon"}
for key, value in pies.items():
print(key, ":", value)
```
# Comprehensions<a id="comp"></a>
[Return to table of contents](#toc)
Are a quicker way to create lists, dicts and sets, they act like for loops and can take conditions as well as if else statements.
<b>List comprehension</b><a id='list'></a>
[Return to table of contents](#toc)
```
# Manual way to make a list
list_1 = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10]
list_2 = [11, 12, 13, 14, 15, 16, 17, 18, 19, 20]
print(list_1)
print(list_2)
# Using list()
list_1_with_list = list(range(1, 11))
list_2_with_list = list(range(11, 21))
print(list_1_with_list)
print(list_2_with_list)
# List comprehension autogenerates the list, they function closely to for loops.
list_1_with_comp = [x for x in range(1, 11)]
list_2_with_comp = [x for x in range(11, 21)]
print(list_1_with_comp)
print(list_2_with_comp)
# Works with functions.
def addition(x):
return x + x
[addition(x) for x in range(0, 3)]
# Also works with conditions
# The % is modulus which gives you the remainder after division.
# If an number is even add 1 if it's odd add 3
[x + 1 if x%2 ==0 else x + 3 for x in range(1, 11)]
# Without an else statement if goes at the end of the statement.
[x + 1 for x in range(1, 11) if x%2 ==0]
# Nested loop example to show how they function like for loops.
for a in range(0, 3):
for b in range(0, 5):
print(a, b)
# List comp also works as a nested loop.
[[a, b] for a in range(0, 3) for b in range(0, 5)]
```
<b>Set comprehension</b><a id='set'></a>
[Return to table of contents](#toc)
```
# Set comprehension is same format as list comprehension but uses curly brackets.
set_1_with_comp = {x for x in range(1, 11)}
set_2_with_comp = {x for x in range(11, 21)}
print(set_1_with_comp)
print(set_2_with_comp)
```
<b>Dict comprehension</b><a id='dict'></a>
[Return to table of contents](#toc)
Examples from: http://cmdlinetips.com/2018/01/5-examples-using-dict-comprehension/ (More samples there as well)
```
# dict comprehension to create dict with numbers as values
{str(i):i for i in [1,2,3,4,5]}
# create list of fruits
fruits = ["apple", "mango", "banana", "cherry"]
# dict comprehension to create dict with fruit name as keys
{f:len(f) for f in fruits}
```
| github_jupyter |
# DEBUG: Inference performance comparison
# TFLite - Performance CPU only vs. Heterogenous execution
In this example notebook, we compare ***TFlite*** inference performance, of a pre-trained Classification model running on CPU (Cortex A**) vs. the same model running in an heterogenous approach (Cortex A** + TIDL offload)
- The user can choose the model (see section titled *Choosing a Pre-Compiled Model*)
- The models used in this example were trained on the ***ImageNet*** dataset because it is a widely used dataset developed for training and benchmarking image classification AI models.
- We perform inference on one sample image.
## Choosing a Pre-Compiled Model
We provide a set of precompiled artifacts to use with this notebook that will appear as a drop-down list once the first code cell is executed.
<img src=docs/images/drop_down.PNG width="400">
**Note**:Users can run this notebook as-is, only action required is to select a model.
```
import os
import cv2
import numpy as np
import ipywidgets as widgets
from scripts.utils import get_eval_configs
last_artifacts_id = selected_model_id.value if "selected_model_id" in locals() else None
prebuilt_configs, selected_model_id = get_eval_configs('classification','tflitert', num_quant_bits = 8, last_artifacts_id = last_artifacts_id)
display(selected_model_id)
print(f'Selected Model: {selected_model_id.label}')
config = prebuilt_configs[selected_model_id.value]
config['session'].set_param('model_id', selected_model_id.value)
config['session'].start()
```
## Define utility function to preprocess input images
Below, we define a utility function to preprocess images for the model. This function takes a path as input, loads the image and preprocesses the images as required by the model. The steps below are shown as a reference (no user action required):
1. Load image
2. Convert BGR image to RGB
3. Scale image
4. Apply per-channel pixel scaling and mean subtraction
5. Convert RGB Image to BGR.
6. Convert the image to NCHW format
- The input arguments of this utility function is selected automatically by this notebook based on the model selected in the drop-down
```
def preprocess(image_path, size, mean, scale, layout, reverse_channels):
# Step 1
img = cv2.imread(image_path)
# Step 2
img = img[:,:,::-1]
# Step 3
img = cv2.resize(img, (size, size), interpolation=cv2.INTER_CUBIC)
# Step 4
img = img.astype('float32')
for mean, scale, ch in zip(mean, scale, range(img.shape[2])):
img[:,:,ch] = ((img.astype('float32')[:,:,ch] - mean) * scale)
# Step 5
if reverse_channels:
img = img[:,:,::-1]
# Step 6
if layout == 'NCHW':
img = np.expand_dims(np.transpose(img, (2,0,1)),axis=0)
else:
img = np.expand_dims(img,axis=0)
return img
from scripts.utils import get_preproc_props
size, mean, scale, layout, reverse_channels = get_preproc_props(config)
print(f'Image size: {size}')
```
## Load and Run a model on ARM (Cortex A**) only
Next cell executes ***TF Lite*** model on Cortex A, and collect benchmark data.
```
import tflite_runtime.interpreter as tflite
import matplotlib.pyplot as plt
from scripts.utils import imagenet_class_to_name
tflite_model_path = config['session'].get_param('model_file')
artifacts_dir = config['session'].get_param('artifacts_folder')
interpreter = tflite.Interpreter(model_path=tflite_model_path)
interpreter.allocate_tensors()
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
img_in = preprocess('sample-images/elephant.bmp' , size, mean, scale, layout, reverse_channels)
if not input_details[0]['dtype'] == np.float32:
img_in = np.uint8(img_in)
interpreter.set_tensor(input_details[0]['index'], img_in)
#Running inference several times to get an stable performance output
for i in range(5):
interpreter.invoke()
res = interpreter.get_tensor(output_details[0]['index'])
print(f'\nTop three results:')
for idx, cls in enumerate(res[0].squeeze()[1:].argsort()[-3:][::-1]):
print('[%d] %s' % (idx, '/'.join(imagenet_class_to_name(cls))))
from scripts.utils import plot_TI_performance_data, plot_TI_DDRBW_data, get_benchmark_output
print(f'\nPerformance CPU EP')
stats = interpreter.get_TI_benchmark_data()
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(10,5))
plot_TI_performance_data(stats, axis=ax)
plt.show()
tt, st, rb, wb = get_benchmark_output(stats)
print(f'Statistics : \n Inferences Per Second : {1000.0/tt :7.2f} fps')
print(f' Inferece Time Per Image : {tt :7.2f} ms \n DDR BW Per Image : {rb+ wb : 7.2f} MB')
```
## Load and Run a model on Heterogenous mode
Next cell executes ***TF Lite*** model in heterogenous mode. Model runs on Cortex A** with graphs offload to TIDL using ***`libtidl_tfl_delegate`*** delegate library. Benchmark data is shown at the end.
```
tflite_model_path = config['session'].get_param('model_file')
artifacts_dir = config['session'].get_param('artifacts_folder')
tidl_delegate = [tflite.load_delegate('libtidl_tfl_delegate.so', {'artifacts_folder': artifacts_dir})]
interpreter = tflite.Interpreter(model_path=tflite_model_path, experimental_delegates=tidl_delegate)
interpreter.allocate_tensors()
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
img_in = preprocess('sample-images/elephant.bmp' , size, mean, scale, layout, reverse_channels)
if not input_details[0]['dtype'] == np.float32:
img_in = np.uint8(img_in)
interpreter.set_tensor(input_details[0]['index'], img_in)
#Running inference several times to get an stable performance output
for i in range(5):
interpreter.invoke()
res = interpreter.get_tensor(output_details[0]['index'])
print(f'\nTop three results:')
for idx, cls in enumerate(res[0].squeeze()[1:].argsort()[-3:][::-1]):
print('[%d] %s' % (idx, '/'.join(imagenet_class_to_name(cls))))
from scripts.utils import plot_TI_performance_data, plot_TI_DDRBW_data, get_benchmark_output
print(f'\nPerformance TFLite + TIDL delegates')
stats = interpreter.get_TI_benchmark_data()
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(10,5))
plot_TI_performance_data(stats, axis=ax)
plt.show()
tt, st, rb, wb = get_benchmark_output(stats)
print(f'Statistics : \n Inferences Per Second : {1000.0/tt :7.2f} fps')
print(f' Inferece Time Per Image : {tt :7.2f} ms \n DDR BW Per Image : {rb+ wb : 7.2f} MB')
```
## Final notes
- With this notebook, user's can quickly compare FPS when running their models only on Cortex A** vs. running their models in heterogenous mode.
- If in Heterogenous mode a model's accuracy, or output, is wrong, a quick sanity check is to run the same model only on Cortex A**
- Accuracy can be improved by modifying TIDL compilation options. For additional tips you can check "run and compare a model compiled with different compilation option" inside debug_tips notebook
| github_jupyter |
# Applying PCA to Interpolated Data
---
I followed the Python Data Science Handbook [In Depth: Principal Component Analysis](https://jakevdp.github.io/PythonDataScienceHandbook/05.09-principal-component-analysis.html) notebook to apply PCA to the interpolated shallow profiler data.
The motivation for this is that time-series data from the oceans can have high dimensionality and strong multicollinearity, and PCA can extract features that (hopefully) maintain the signals we're interested in while providing us with a lower-dimension dataframe to use in modeling. This allows analysis to be scaled up to include several years of measurements for several variables and locations without requiring truly excessive computational resources.
Keep in mind two assumptions of PCA:
1. **Linearity:** PCA detects and controls for linear relationships, so we assume that the data does not hold nonlinear relationships (or that we don't care about these nonlinear relationships).
2. **Large variances define importance:** If data is spread in a direction, that direction is important! If there is little spread in a direction, that direction is not very important.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns; sns.set()
from sklearn.decomposition import PCA
%matplotlib inline
```
---
### Read in the data
This profiler data has been interpolated to a regular 2D grid where the index is time and the columns are different seawater pressures, so that each row is a new observation and each column is a different location in the water column. I used [scipy's `griddata()`](https://docs.scipy.org/doc/scipy/reference/generated/scipy.interpolate.griddata.html) to do this in the [06 interpolation notebook](https://github.com/dgumustel/coastal-upwelling/blob/main/notebooks/06_0_interpolation_griddata.ipynb).
```
df = pd.read_csv('../../coastal_upwelling_output/interpolated.csv')
df.rename({'Unnamed: 0':'time'},inplace=True, axis=1)
df.set_index('time', inplace=True)
df.drop(columns=['CUTI', 'upwelling'], inplace=True)
df
```
The goal is to reduce these 192 columns to a significantly smaller number while retaining most of the original signals in the data.
```
df.shape[0] * df.shape[1]
```
Imagine how many more values this dataframe would have if we added a few more years of observations, increased the depth resolution, or appended more variables, like salinity or dissolved oxygen. Applying PCA will significantly reduce the number of calculations required in modeling.
### Choosing the number of components
```
pca = PCA().fit(df)
plt.plot(np.cumsum(pca.explained_variance_ratio_), '.')
plt.xlim(-1, 50)
plt.xlabel('Number of components')
plt.ylabel('Cumulative explained variance');
```
I'll include the first 4 components to explain roughly 95% of the variance.
### Applying PCA
```
pca = PCA(n_components=4).fit(df)
components = pca.transform(df)
filtered = pca.inverse_transform(components)
fig, [ax1, ax2] = plt.subplots(2, 1, figsize=(20, 12))
# Make sure these plot on the same color scales
vmin = 6
vmax = 19.5
levels = np.linspace(vmin, vmax, 11)
cf1 = ax1.contourf(df.T, levels=levels)
ax1.set_ylabel('Pressure', size=16)
ax1.set_xlabel('Time', size=16)
ax1.set_title('Original Interpolation', size=16)
cb1 = plt.colorbar(cf1, ax=ax1)
cb1.set_label('Temperature (deg C)', size=16)
ax1.invert_yaxis()
ax1.tick_params(axis='both', which='major', labelsize=14)
cb1.ax.tick_params(labelsize=14)
cf2 = ax2.contourf(filtered.T, levels=levels)
ax2.set_ylabel('Pressure', size=16)
ax2.set_xlabel('Time', size=16)
ax2.set_title('Reconstructed with First 4 Components', size=16)
cb2 = plt.colorbar(cf2, ax=ax2)
cb2.set_label('Temperature (deg C)', size=16)
ax2.invert_yaxis()
ax2.tick_params(axis='both', which='major', labelsize=14)
cb2.ax.tick_params(labelsize=14)
plt.tight_layout()
```
We can see how well this process did at reconstructing the interpolated data using the first 4 components. Note that the axes ticks are indices, not actual pressure or time measurements. The true y-axis range is `5.12` to `196.12` dbars at intervals of 1 dbar (approximately 1 meter) and the true x-axis range is `2018-07-17 17:18:50` to `2018-12-25 23:18:50` at intervals of 1 hour and 40 minutes.
| github_jupyter |
# Scientific modules and IPython
Nikolay Koldunov
koldunovn@gmail.com
This is part of [**Python for Geosciences**](https://github.com/koldunovn/python_for_geosciences) notes.
```
%matplotlib inline
import matplotlib.pylab as plt
```
## Core scientific packages
When people say that they do their scientific computations in Python it's only half true. Python is a construction set, similar to MITgcm or other models. Without packages it's only a core, that although very powerful, does not seems to be able to do much by itself.
There is a set of packages, that almost every scientist would need:
<img height="200" src="files/core.png">
We are going to talk about all exept Sympy
## Installation
Installation instructions can be found in the README.md file of this repository. Better to use [rendered version from GitHub](https://github.com/koldunovn/python_for_geosciences/blob/master/README.md).
## IPython
In order to be productive you need comfortable environment, and this is what IPython provides. It was started as enhanced python interactive shell, but with time become architecture for interactive computing.
## Jupyter notebook
Since the 0.12 release, IPython provides a new rich text web interface - IPython notebook. Here you can combine:
#### Code execution
```
print('I love Python')
```
#### Text (Markdown)
IPython [website](http://ipython.org/).
List:
* [Python on Codeacademy](http://www.codecademy.com/tracks/python)
* [Google's Python Class](https://developers.google.com/edu/python/)
Code:
print('hello world')
#### $\LaTeX$ equations
$$\int_0^\infty e^{-x^2} dx=\frac{\sqrt{\pi}}{2}$$
$$
F(x,y)=0 ~~\mbox{and}~~
\left| \begin{array}{ccc}
F''_{xx} & F''_{xy} & F'_x \\
F''_{yx} & F''_{yy} & F'_y \\
F'_x & F'_y & 0
\end{array}\right| = 0
$$
#### Plots
```
x = [1,2,3,4,5]
plt.plot(x);
```
#### Rich media
```
from IPython.display import YouTubeVideo
YouTubeVideo('F4rFuIb1Ie4')
```
* [IPython website](http://ipython.org/)
* [Notebook gallery](https://github.com/ipython/ipython/wiki/A-gallery-of-interesting-IPython-Notebooks)
## Run notebook
In order to start Jupyter notebook you have to type:
jupyter notebook
### You can download and run this lectures:
Web version can be accesed from the [github repository](https://github.com/koldunovn/python_for_geosciences).
## Main IPython features
### Getting help
You can use question mark in order to get help. To execute cell you have to press *Shift+Enter*
```
?
```
Question mark after a function will open pager with documentation. Double question mark will show you source code of the function.
```
plt.plot??
```
Press SHIFT+TAB after opening bracket in order to get help for the function (list of arguments, doc string).
```
sum()
```
### Accessing the underlying operating system
You can access system functions by typing exclamation mark.
```
!pwd
```
If you already have some netCDF file in the directory and *ncdump* is installed, you can for example look at its header.
```
!ncdump -h test_netcdf.nc
```
## Magic functions
The magic function system provides a series of functions which allow you to
control the behavior of IPython itself, plus a lot of system-type
features.
Let's create some set of numbers using [range](http://docs.python.org/2/library/functions.html#range) command:
```
list(range(10))
```
And find out how long does it take to run it with *%timeit* magic function:
```
%timeit list(range(10))
```
Print all interactive variables (similar to Matlab function):
```
%whos
```
### Cell-oriented magic
Receive as argument both the current line where they are declared and the whole body of the cell.
```
%%timeit
range(10)
range(100)
```
Thre are several cell-oriented magic functions that allow you to run code in other languages:
```
%%bash
echo "My shell is:" $SHELL
%%perl
$variable = 1;
print "The variable has the value of $variable\n";
```
You can write content of the cell to a file with *%%writefile* (or *%%file* for ipython < 1.0):
```
%%writefile hello.py
#if you use ipython < 1.0, use %%file comand
#%%file
a = 'hello world!'
print(a)
```
And then run it:
```
%run hello.py
```
The *%run* magic will run your python script and load all variables into your interactive namespace for further use.
```
%whos
```
In order to get information about all magic functions type:
```
%magic
```
### Links:
[The cell magics in IPython](http://nbviewer.ipython.org/urls/raw.github.com/ipython/ipython/1.x/examples/notebooks/Cell%20Magics.ipynb)
| github_jupyter |
# Model Interpretation Methods
Welcome to the final assignment of course 3! In this assignment we will focus on the interpretation of machine learning and deep learning models. Using the techniques we've learned this week we'll revisit some of the models we've built throughout the course and try to understand a little more about what they're doing.
In this assignment you'll use various methods to interpret different types of machine learning models. In particular, you'll learn about the following topics:
- Interpreting Deep Learning Models
- Understanding output using GradCAMs
- Feature Importance in Machine Learning
- Permutation Method
- SHAP Values
Let's get started.
### This assignment covers the folowing topics:
- [1. Interpreting Deep Learning Models](#1)
- [1.1 GradCAM](#1-1)
- [1.1.1 Getting Intermediate Layers](#1-1-1)
- [1.1.2 Getting Gradients](#1-1-2)
- [1.1.3 Implementing GradCAM](#1-1-3)
- [Exercise 1](#ex-01)
- [1.1.4 Using GradCAM to Visualize Multiple Labels](#1-1-4)
- [Exercise 2](#ex-02)
- [2. Feature Importance in Machine Learning](#2)
- [2.1 Permuation Method for Feature Importance](#2-1)
- [2.1.1 Implementing Permutation](#2-1-1)
- [Exercise 3](#ex-03)
- [2.1.2 Implementing Importance](#2-1-2)
- [Exercise 4](#ex-04)
- [2.1.3 Computing our Feature Importance](#2-1-3)
- [2.2 Shapley Values for Random Forests](#2-2)
- [2.2.1 Visualizing Feature Importance on Specific Individuals](#2-2-1)
- [2.2.2 Visualizing Feature Importance on Aggregate](#2-2-2)
- [2.2.3 Visualizing Interactions between Features](#2-2-3)
## Packages
We'll first import the necessary packages for this assignment.
- `keras`: we'll use this framework to interact with our deep learning model
- `matplotlib`: standard plotting library
- `pandas`: we'll use this to manipulate data
- `numpy`: standard python library for numerical operations
- `cv2`: library that contains convenience functions for image processing
- `sklearn`: standard machine learning library
- `lifelines`: we'll use their implementation of the c-index
- `shap`: library for interpreting and visualizing machine learning models using shapley values
```
import keras
from keras import backend as K
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import cv2
import sklearn
import lifelines
import shap
from util import *
# This sets a common size for all the figures we will draw.
plt.rcParams['figure.figsize'] = [10, 7]
```
<a name="1"></a>
## 1 Interpreting Deep Learning Models
To start, let's try understanding our X-ray diagnostic model from Course 1 Week 1. Run the next cell to load in the model (it should take a few seconds to complete).
```
model = load_C3M3_model()
```
Let's load in an X-ray image to develop on. Run the next cell to load and show the image.
```
IMAGE_DIR = 'nih_new/images-small/'
df = pd.read_csv("nih_new/train-small.csv")
im_path = IMAGE_DIR + '00016650_000.png'
x = load_image(im_path, df, preprocess=False)
plt.imshow(x, cmap = 'gray')
plt.show()
```
Next, let's get our predictions. Before we plug the image into our model, we have to normalize it. Run the next cell to compute the mean and standard deviation of the images in our training set.
```
mean, std = get_mean_std_per_batch(df)
```
Now we are ready to normalize and run the image through our model to get predictions.
```
labels = ['Cardiomegaly', 'Emphysema', 'Effusion', 'Hernia', 'Infiltration', 'Mass', 'Nodule', 'Atelectasis',
'Pneumothorax', 'Pleural_Thickening', 'Pneumonia', 'Fibrosis', 'Edema', 'Consolidation']
processed_image = load_image_normalize(im_path, mean, std)
preds = model.predict(processed_image)
pred_df = pd.DataFrame(preds, columns = labels)
pred_df.loc[0, :].plot.bar()
plt.title("Predictions")
plt.show()
```
We see, for example, that the model predicts Cardiomegaly (enlarged heart) with high probability. Indeed, this patient was diagnosed with cardiomegaly. However, we don't know where the model is looking when it's making its own diagnosis. To gain more insight into what the model is looking at, we can use GradCAMs.
<a name="1-1"></a>
### 1.1 GradCAM
GradCAM is a technique to visualize the impact of each region of an image on a specific output for a Convolutional Neural Network model. Through GradCAM, we can generate a heatmap by computing gradients of the specific class scores we are interested in visualizing.
<a name="1-1-1"></a>
#### 1.1.1 Getting Intermediate Layers
Perhaps the most complicated part of computing GradCAM is accessing intermediate activations in our deep learning model and computing gradients with respect to the class output. Now we'll go over one pattern to accomplish this, which you can use when implementing GradCAM.
In order to understand how to access intermediate layers in a computation, first let's see the layers that our model is composed of. This can be done by calling Keras convenience function `model.summary()`. Do this in the cell below.
```
model.summary()
```
There are a lot of layers, but typically we'll only be extracting one of the last few. Remember that the last few layers usually have more abstract information. To access a layer, we can use `model.get_layer(layer).output`, which takes in the name of the layer in question. Let's try getting the `conv5_block16_concat` layer, the raw output of the last convolutional layer.
```
spatial_maps = model.get_layer('conv5_block16_concat').output
print(spatial_maps)
```
Now, this tensor is just a placeholder, it doesn't contain the actual activations for a particular image. To get this we will use [Keras.backend.function](https://www.tensorflow.org/api_docs/python/tf/keras/backend/function) to return intermediate computations while the model is processing a particular input. This method takes in an input and output placeholders and returns a function. This function will compute the intermediate output (until it reaches the given placeholder) evaluated given the input. For example, if you want the layer that you just retrieved (conv5_block16_concat), you could write the following:
```
get_spatial_maps = K.function([model.input], [spatial_maps])
print(get_spatial_maps)
```
We see that we now have a `Function` object. Now, to get the actual intermediate output evaluated with a particular input, we just plug in an image to this function:
```
# get an image
x = load_image_normalize(im_path, mean, std)
print(f"x is of type {type(x)}")
print(f"x is of shape {x.shape}")
# get the spatial maps layer activations (a list of numpy arrays)
spatial_maps_x_l = get_spatial_maps([x])
print(f"spatial_maps_x_l is of type {type(spatial_maps_x_l)}")
print(f"spatial_maps_x_l is has length {len(spatial_maps_x_l)}")
# get the 0th item in the list
spatial_maps_x = spatial_maps_x_l[0]
print(f"spatial_maps_x is of type {type(spatial_maps_x)}")
print(f"spatial_maps_x is of shape {spatial_maps_x.shape}")
```
Notice that the shape is (1, 10, 10, 1024). The 0th dimension of size 1 is the batch dimension. Remove the batch dimension for later calculations by taking the 0th index of spatial_maps_x.
```
# Get rid of the batch dimension
spatial_maps_x = spatial_maps_x[0] # equivalent to spatial_maps_x[0,:]
print(f"spatial_maps_x without the batch dimension has shape {spatial_maps_x.shape}")
print("Output some of the content:")
print(spatial_maps_x[0])
```
We now have the activations for that particular image, and we can use it for interpretation. The function that is returned by calling `K.function([model.input], [spatial_maps])` (saved here in the variable `get_spatial_maps`) is sometimes referred to as a "hook", letting you peek into the intermediate computations in the model.
<a name="1-1-2"></a>
#### 1.1.2 Getting Gradients
The other major step in computing GradCAMs is getting gradients with respect to the output for a particular class. Luckily, Keras makes getting gradients simple. We can use the [Keras.backend.gradients](https://www.tensorflow.org/api_docs/python/tf/keras/backend/gradients) function. The first parameter is the value you are taking the gradient of, and the second is the parameter you are taking that gradient with respect to. We illustrate below:
```
# get the output of the model
output_with_batch_dim = model.output
print(f"Model output includes batch dimension, has shape {output_with_batch_dim.shape}")
```
To get the output without the batch dimension, you can take the 0th index of the tensor. Note that because the batch dimension is 'None', you could actually enter any integer index, but let's just use 0.
```
# Get the output without the batch dimension
output_all_categories = output_with_batch_dim[0]
print(f"The output for all 14 categories of disease has shape {output_all_categories.shape}")
```
The output has 14 categories, one for each disease category, indexed from 0 to 13. Cardiomegaly is the disease category at index 0.
```
# Get the first category's output (Cardiomegaly) at index 0
y_category_0 = output_all_categories[0]
print(f"The Cardiomegaly output is at index 0, and has shape {y_category_0.shape}")
# Get gradient of y_category_0 with respect to spatial_maps
gradient_l = K.gradients(y_category_0, spatial_maps)
print(f"gradient_l is of type {type(gradient_l)} and has length {len(gradient_l)}")
# gradient_l is a list of size 1. Get the gradient at index 0
gradient = gradient_l[0]
print(gradient)
```
Again, this is just a placeholder. Just like for intermediate layers, we can use `K.function` to compute the value of the gradient for a particular input.
The K.function() takes in
- a list of inputs: in this case, one input, 'model.input'
- a list of tensors: in this case, one output tensor 'gradient'
It returns a function that calculates the activations of the list of tensors.
- This returned function returns a list of the activations, one for each tensor that was passed into K.function().
```
# Create the function that gets the gradient
get_gradient = K.function([model.input], [gradient])
type(get_gradient)
# get an input x-ray image
x = load_image_normalize(im_path, mean, std)
print(f"X-ray image has shape {x.shape}")
```
The `get_gradient` function takes in a list of inputs, and returns a list of the gradients, one for each image.
```
# use the get_gradient function to get the gradient (pass in the input image inside a list)
grad_x_l = get_gradient([x])
print(f"grad_x_l is of type {type(grad_x_l)} and length {len(grad_x_l)}")
# get the gradient at index 0 of the list.
grad_x_with_batch_dim = grad_x_l[0]
print(f"grad_x_with_batch_dim is type {type(grad_x_with_batch_dim)} and shape {grad_x_with_batch_dim.shape}")
# To remove the batch dimension, take the value at index 0 of the batch dimension
grad_x = grad_x_with_batch_dim[0]
print(f"grad_x is type {type(grad_x)} and shape {grad_x.shape}")
print("Gradient grad_x (show some of its content:")
print(grad_x[0])
```
Just like we had a hook into the penultimate layer, we now have a hook into the gradient! This allows us to easily compute pretty much anything relevant to our model output.
We can also combine the two to have one function call which gives us both the gradient and the last layer (this might come in handy when implementing GradCAM in the next section).
```
# Use K.function to generate a single function
# Notice that a list of two tensors, is passed in as the second argument of K.function()
get_spatial_maps_and_gradient = K.function([model.input], [spatial_maps, gradient])
print(type(get_spatial_maps_and_gradient))
# The returned function returns a list of the evaluated tensors
tensor_eval_l = get_spatial_maps_and_gradient([x])
print(f"tensor_eval_l is type {type(tensor_eval_l)} and length {len(tensor_eval_l)}")
# store the two numpy arrays from index 0 and 1 into their own variables
spatial_maps_x_with_batch_dim, grad_x_with_batch_dim = tensor_eval_l
print(f"spatial_maps_x_with_batch_dim has shape {spatial_maps_x_with_batch_dim.shape}")
print(f"grad_x_with_batch_dim has shape {grad_x_with_batch_dim.shape}")
# Note: you could also do this directly from the function call:
spatial_maps_x_with_batch_dim, grad_x_with_batch_dim = get_spatial_maps_and_gradient([x])
print(f"spatial_maps_x_with_batch_dim has shape {spatial_maps_x_with_batch_dim.shape}")
print(f"grad_x_with_batch_dim has shape {grad_x_with_batch_dim.shape}")
# Remove the batch dimension by taking the 0th index at the batch dimension
spatial_maps_x = spatial_maps_x_with_batch_dim[0]
grad_x = grad_x_with_batch_dim[0]
print(f"spatial_maps_x shape {spatial_maps_x.shape}")
print(f"grad_x shape {grad_x.shape}")
print("\nSpatial maps (print some content):")
print(spatial_maps_x[0])
print("\nGradient (print some content:")
print(grad_x[0])
```
<a name="1-1-3"></a>
#### 1.1.3 Implementing GradCAM
<a name='ex-01'></a>
### Exercise 1
In the next cell, fill in the `grad_cam` method to produce GradCAM visualizations for an input model and image. This is fairly complicated, so it might help to break it down into these steps:
1. Hook into model output and last layer activations.
2. Get gradients of last layer activations with respect to output.
3. Compute value of last layer and gradients for input image.
4. Compute weights from gradients by global average pooling.
5. Compute the dot product between the last layer and weights to get the score for each pixel.
6. Resize, take ReLU, and return cam.
<details>
<summary>
<font size="3" color="darkgreen"><b>Hints</b></font>
</summary>
The following hints follow the order of the sections described above.
1. Remember that the output shape of our model will be [1, class_amount].
1. The input in this case will always have batch_size = 1
2. See [K.gradients](https://www.tensorflow.org/api_docs/python/tf/keras/backend/gradients)
3. Follow the procedure we used in the previous two sections.
4. Check the axis; make sure weights have shape (C)!
5. See [np.dot](https://docs.scipy.org/doc/numpy/reference/generated/numpy.dot.html)
</details>
To test, you will compare your output on an image to the output from a correct implementation of GradCAM. You will receive full credit if the pixel-wise mean squared error is less than 0.05.
```
# UNQ_C1 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def grad_cam(input_model, image, category_index, layer_name):
"""
GradCAM method for visualizing input saliency.
Args:
input_model (Keras.model): model to compute cam for
image (tensor): input to model, shape (1, H, W, 3)
cls (int): class to compute cam with respect to
layer_name (str): relevant layer in model
H (int): input height
W (int): input width
Return:
cam ()
"""
cam = None
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# 1. Get placeholders for class output and last layer
# Get the model's output
output_with_batch_dim = input_model.output
# Remove the batch dimension
output_all_categories = output_with_batch_dim[0]
# Retrieve only the disease category at the given category index
y_c = output_all_categories[category_index]
# Get the input model's layer specified by layer_name, and retrive the layer's output tensor
spatial_map_layer = input_model.get_layer(layer_name).output
# 2. Get gradients of last layer with respect to output
# get the gradients of y_c with respect to the spatial map layer (it's a list of length 1)
grads_l = K.gradients(y_c,spatial_map_layer)
# Get the gradient at index 0 of the list
grads = grads_l[0]
# 3. Get hook for the selected layer and its gradient, based on given model's input
# Hint: Use the variables produced by the previous two lines of code
spatial_map_and_gradient_function = K.function([input_model.input],[spatial_map_layer, grads])
# Put in the image to calculate the values of the spatial_maps (selected layer) and values of the gradients
spatial_map_all_dims, grads_val_all_dims = spatial_map_and_gradient_function([image])
# Reshape activations and gradient to remove the batch dimension
# Shape goes from (B, H, W, C) to (H, W, C)
# B: Batch. H: Height. W: Width. C: Channel
# Reshape spatial map output to remove the batch dimension
spatial_map_val = spatial_map_all_dims[0]
# Reshape gradients to remove the batch dimension
grads_val = grads_val_all_dims[0]
# print(spatial_map_val.shape,type(spatial_map_val),"---",grads_val.shape,type(grads_val))
# 4. Compute weights using global average pooling on gradient
# grads_val has shape (Height, Width, Channels) (H,W,C)
# Take the mean across the height and also width, for each channel
# Make sure weights have shape (C)
weights = np.mean(grads_val,axis=(0,1))
# 5. Compute dot product of spatial map values with the weights
cam = np.dot(spatial_map_val, weights)
### END CODE HERE ###
# We'll take care of the postprocessing.
H, W = image.shape[1], image.shape[2]
cam = np.maximum(cam, 0) # ReLU so we only get positive importance
cam = cv2.resize(cam, (W, H), cv2.INTER_NEAREST)
cam = cam / cam.max()
return cam
```
Below we generate the CAM for the image and compute the error (pixel-wise mean squared difference) from the expected values according to our reference.
```
im = load_image_normalize(im_path, mean, std)
cam = grad_cam(model, im, 5, 'conv5_block16_concat') # Mass is class 5
# Loads reference CAM to compare our implementation with.
reference = np.load("reference_cam.npy")
error = np.mean((cam-reference)**2)
print(f"Error from reference: {error:.4f}, should be less than 0.05")
```
Run the next cell to visualize the CAM and the original image.
```
plt.imshow(load_image(im_path, df, preprocess=False), cmap='gray')
plt.title("Original")
plt.axis('off')
plt.show()
plt.imshow(load_image(im_path, df, preprocess=False), cmap='gray')
plt.imshow(cam, cmap='magma', alpha=0.5)
plt.title("GradCAM")
plt.axis('off')
plt.show()
```
We can see that it focuses on the large white area in the middle of the chest cavity. Indeed this is a clear case of cardiomegaly, that is, an enlarged heart.
<a name="1-1-4"></a>
#### 1.1.4 Using GradCAM to Visualize Multiple Labels
<a name='ex-02'></a>
### Exercise 2
We can use GradCAMs for multiple labels on the same image. Let's do it for the labels with best AUC for our model, Cardiomegaly, Mass, and Edema.
```
# UNQ_C2 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def compute_gradcam(model, img, mean, std, data_dir, df,
labels, selected_labels, layer_name='conv5_block16_concat'):
"""
Compute GradCAM for many specified labels for an image.
This method will use the `grad_cam` function.
Args:
model (Keras.model): Model to compute GradCAM for
img (string): Image name we want to compute GradCAM for.
mean (float): Mean to normalize to image.
std (float): Standard deviation to normalize the image.
data_dir (str): Path of the directory to load the images from.
df(pd.Dataframe): Dataframe with the image features.
labels ([str]): All output labels for the model.
selected_labels ([str]): All output labels we want to compute the GradCAM for.
layer_name: Intermediate layer from the model we want to compute the GradCAM for.
"""
img_path = data_dir + img
preprocessed_input = load_image_normalize(img_path, mean, std)
predictions = model.predict(preprocessed_input)
print("Ground Truth: ", ", ".join(np.take(labels, np.nonzero(df[df["Image"] == img][labels].values[0]))[0]))
plt.figure(figsize=(15, 10))
plt.subplot(151)
plt.title("Original")
plt.axis('off')
plt.imshow(load_image(img_path, df, preprocess=False), cmap='gray')
j = 1
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# Loop through all labels
for i in range(len(labels)): # complete this line
# Compute CAM and show plots for each selected label.
# Check if the label is one of the selected labels
if labels[i] in selected_labels: # complete this line
# Use the grad_cam function to calculate gradcam
gradcam = grad_cam(model,preprocessed_input,i,layer_name)
### END CODE HERE ###
print("Generating gradcam for class %s (p=%2.2f)" % (labels[i], round(predictions[0][i], 3)))
plt.subplot(151 + j)
plt.title(labels[i] + ": " + str(round(predictions[0][i], 3)))
plt.axis('off')
plt.imshow(load_image(img_path, df, preprocess=False), cmap='gray')
plt.imshow(gradcam, cmap='magma', alpha=min(0.5, predictions[0][i]))
j +=1
```
Run the following cells to print the ground truth diagnosis for a given case and show the original x-ray as well as GradCAMs for Cardiomegaly, Mass, and Edema.
```
df = pd.read_csv("nih_new/train-small.csv")
image_filename = '00016650_000.png'
labels_to_show = ['Cardiomegaly', 'Mass', 'Edema']
compute_gradcam(model, image_filename, mean, std, IMAGE_DIR, df, labels, labels_to_show)
```
The model correctly predicts absence of mass or edema. The probability for mass is higher, and we can see that it may be influenced by the shapes in the middle of the chest cavity, as well as around the shoulder. We'll run it for two more images.
```
image_filename = '00005410_000.png'
compute_gradcam(model, image_filename, mean, std, IMAGE_DIR, df, labels, labels_to_show)
```
In the example above, the model correctly focuses on the mass near the center of the chest cavity.
```
image_name = '00004090_002.png'
compute_gradcam(model, image_name, mean, std, IMAGE_DIR, df, labels, labels_to_show)
```
Here the model correctly picks up the signs of edema near the bottom of the chest cavity. We can also notice that Cardiomegaly has a high score for this image, though the ground truth doesn't include it. This visualization might be helpful for error analysis; for example, we can notice that the model is indeed looking at the expected area to make the prediction.
This concludes the section on GradCAMs. We hope you've gained an appreciation for the importance of interpretation when it comes to deep learning models in medicine. Interpretation tools like this one can be helpful for discovery of markers, error analysis, and even in deployment.
<a name="2"></a>
## 2 Feature Importance in Machine Learning
When developing predictive models and risk measures, it's often helpful to know which features are making the most difference. This is easy to determine in simpler models such as linear models and decision trees. However as we move to more complex models to achieve high performance, we usually sacrifice some interpretability. In this assignment we'll try to regain some of that interpretability using Shapley values, a technique which has gained popularity in recent years, but which is based on classic results in cooperative game theory.
We'll revisit our random forest model from course 2 module 2 and try to analyze it more closely using Shapley values. Run the next cell to load in the data and model from that assignment and recalculate the test set c-index.
```
rf = pickle.load(open('nhanes_rf.sav', 'rb')) # Loading the model
test_df = pd.read_csv('nhanest_test.csv')
test_df = test_df.drop(test_df.columns[0], axis=1)
X_test = test_df.drop('y', axis=1)
y_test = test_df.loc[:, 'y']
cindex_test = cindex(y_test, rf.predict_proba(X_test)[:, 1])
print("Model C-index on test: {}".format(cindex_test))
```
Run the next cell to print out the riskiest individuals according to our model.
```
X_test_risky = X_test.copy(deep=True)
X_test_risky.loc[:, 'risk'] = rf.predict_proba(X_test)[:, 1] # Predicting our risk.
X_test_risky = X_test_risky.sort_values(by='risk', ascending=False) # Sorting by risk value.
X_test_risky.head()
```
<a name="2-1"></a>
### 2.1 Permuation Method for Feature Importance
First we'll try to determine feature importance using the permutation method. In the permutation method, the importance of feature $i$ would be the regular performance of the model minus the performance with the values for feature $i$ permuted in the dataset. This way we can assess how well a model without that feature would do without having to train a new model for each feature.
<a name="2-1-1"></a>
#### 2.1.1 Implementing Permutation
<a name='ex-03'></a>
### Exercise 3
Complete the implementation of the function below, which given a feature name returns a dataset with those feature values randomly permuted.
<details>
<summary>
<font size="3" color="darkgreen"><b>Hints</b></font>
</summary>
<ul>
<li>
See <a href=https://numpy.org/devdocs/reference/random/generated/numpy.random.permutation.html> np.random.permutation</a>
</li>
</ul>
</details>
```
# UNQ_C3 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def permute_feature(df, feature):
"""
Given dataset, returns version with the values of
the given feature randomly permuted.
Args:
df (dataframe): The dataset, shape (num subjects, num features)
feature (string): Name of feature to permute
Returns:
permuted_df (dataframe): Exactly the same as df except the values
of the given feature are randomly permuted.
"""
permuted_df = df.copy(deep=True) # Make copy so we don't change original df
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# Permute the values of the column 'feature'
permuted_features = np.random.permutation(permuted_df[feature])
# Set the column 'feature' to its permuted values.
permuted_df[feature] = permuted_features
### END CODE HERE ###
return permuted_df
print("Test Case")
example_df = pd.DataFrame({'col1': [0, 1, 2], 'col2':['A', 'B', 'C']})
print("Original dataframe:")
print(example_df)
print("\n")
print("col1 permuted:")
print(permute_feature(example_df, 'col1'))
print("\n")
print("Compute average values over 1000 runs to get expected values:")
col1_values = np.zeros((3, 1000))
np.random.seed(0) # Adding a constant seed so we can always expect the same values and evaluate correctly.
for i in range(1000):
col1_values[:, i] = permute_feature(example_df, 'col1')['col1'].values
print("Average of col1: {}, expected value: [0.976, 1.03, 0.994]".format(np.mean(col1_values, axis=1)))
```
<a name="2-1-2"></a>
#### 2.1.2 Implementing Importance
<a name='ex-04'></a>
### Exercise 4
Now we will use the function we just created to compute feature importances (according to the permutation method) in the function below.
<details>
<summary>
<font size="3" color="darkgreen"><b>Hints</b></font>
</summary>
\begin{align}
I_x = \left\lvert perf - perf_x \right\rvert
\end{align}
where $I_x$ is the importance of feature $x$ and
\begin{align}
perf_x = \frac{1}{n}\cdot \sum_{i=1}^{n} perf_i^{sx}
\end{align}
where $perf_i^{sx}$ is the performance with the feature $x$ shuffled in the $i$th permutation.
</details>
```
# UNQ_C4 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def permutation_importance(X, y, model, metric, num_samples = 100):
"""
Compute permutation importance for each feature.
Args:
X (dataframe): Dataframe for test data, shape (num subject, num features)
y (np.array): Labels for each row of X, shape (num subjects,)
model (object): Model to compute importances for, guaranteed to have
a 'predict_proba' method to compute probabilistic
predictions given input
metric (function): Metric to be used for feature importance. Takes in ground
truth and predictions as the only two arguments
num_samples (int): Number of samples to average over when computing change in
performance for each feature
Returns:
importances (dataframe): Dataframe containing feature importance for each
column of df with shape (1, num_features)
"""
importances = pd.DataFrame(index = ['importance'], columns = X.columns)
# Get baseline performance (note, you'll use this metric function again later)
baseline_performance = metric(y, model.predict_proba(X)[:, 1])
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# Iterate over features (the columns in the importances dataframe)
for feature in importances.columns: # complete this line
# Compute 'num_sample' performances by permutating that feature
# You'll see how the model performs when the feature is permuted
# You'll do this num_samples number of times, and save the performance each time
# To store the feature performance,
# create a numpy array of size num_samples, initialized to all zeros
feature_performance_arr = np.zeros(num_samples)
# Loop through each sample
for i in range(num_samples): # complete this line
# permute the column of dataframe X
perm_X = permute_feature(X,feature)
# calculate the performance with the permuted data
# Use the same metric function that was used earlier
feature_performance_arr[i] = metric(y, model.predict_proba(perm_X)[:, 1])
# Compute importance: absolute difference between
# the baseline performance and the average across the
importances[feature]['importance'] = np.abs(baseline_performance - np.mean(feature_performance_arr))
### END CODE HERE ###
return importances
```
**Test Case**
```
print("Test Case")
print("\n")
print("We check our answers on a Logistic Regression on a dataset")
print("where y is given by a sigmoid applied to the important feature.")
print("The unimportant feature is random noise.")
print("\n")
example_df = pd.DataFrame({'important': np.random.normal(size=(1000)), 'unimportant':np.random.normal(size=(1000))})
example_y = np.round(1 / (1 + np.exp(-example_df.important)))
example_model = sklearn.linear_model.LogisticRegression(fit_intercept=False).fit(example_df, example_y)
example_importances = permutation_importance(example_df, example_y, example_model, cindex, num_samples=100)
print("Computed importances:")
print(example_importances)
print("\n")
print("Expected importances (approximate values):")
print(pd.DataFrame({"important": 0.50, "unimportant": 0.00}, index=['importance']))
print("If you round the actual values, they will be similar to the expected values")
```
<a name="2-1-3"></a>
#### 2.1.3 Computing our Feature Importance
Next, we compute importances on our dataset. Since we are computing the permutation importance for all the features, it might take a few minutes to run.
```
importances = permutation_importance(X_test, y_test, rf, cindex, num_samples=100)
importances
```
Let's plot these in a bar chart for easier comparison.
```
importances.T.plot.bar()
plt.ylabel("Importance")
l = plt.legend()
l.remove()
plt.show()
```
You should see age as by far the best prediction of near term mortality, as one might expect. Next is sex, followed by diastolic blood pressure. Interestingly, the poverty index also has a large impact, despite the fact that it is not directly related to an individual's health. This alludes to the importance of social determinants of health in our model.
<a name="2-2"></a>
### 2.2 Shapley Values for Random Forests
We'll contrast the permutation method with a more recent technique known as Shapley values (actually, Shapley values date back to the mid 20th century, but have only been applied to machine learning very recently).
<a name="2-2-1"></a>
#### 2.2.1 Visualizing Feature Importance on Specific Individuals
We can use Shapley values to try and understand the model output on specific individuals. In general Shapley values take exponential time to compute, but luckily there are faster approximations for forests in particular that run in polynomial time. Run the next cell to display a 'force plot' showing how each feature influences the output for the first person in our dataset. If you want more information about 'force plots' and other decision plots, please take a look at [this notebook](https://github.com/slundberg/shap/blob/master/notebooks/plots/decision_plot.ipynb) by the `shap` library creators.
```
explainer = shap.TreeExplainer(rf)
i = 0 # Picking an individual
shap_value = explainer.shap_values(X_test.loc[X_test_risky.index[i], :])[1]
shap.force_plot(explainer.expected_value[1], shap_value, feature_names=X_test.columns, matplotlib=True)
```
For this individual, their age, pulse pressure, and sex were the biggest contributors to their high risk prediction. Note how shapley values give us greater granularity in our interpretations.
Feel free to change the `i` value above to explore the feature influences for different individuals.
<a name="2-2-2"></a>
#### 2.2.2 Visualizing Feature Importance on Aggregate
Just like with the permutation method, we might also want to understand model output in aggregate. Shapley values allow us to do this as well. Run the next cell to initialize the shapley values for each example in the test set (this may also take a few minutes).
```
shap_values = shap.TreeExplainer(rf).shap_values(X_test)[1]
```
You can ignore the `setting feature_perturbation` message.
Run the next cell to see a summary plot of the shapley values for each feature on each of the test examples. The colors indicate the value of the feature. The features are listed in terms of decreasing absolute average shapley value over all the individuals in the dataset.
```
shap.summary_plot(shap_values, X_test)
```
In the above plot, you might be able to notice a high concentration of points on specific SHAP value ranges. This means that a high proportion of our test set lies on those ranges.
As with the permutation method, age, sex, poverty index, and diastolic BP seem to be the most important features. Being older has a negative impact on mortality, and being a woman (sex=2.0) has a positive effect.
<a name="2-2-3"></a>
#### 2.2.3 Visualizing Interactions between Features
The `shap` library also lets you visualize interactions between features using dependence plots. These plot the Shapley value for a given feature for each data point, and color the points in using the value for another feature. This lets us begin to explain the variation in shapley value for a single value of the main feature.
Run the next cell to see the interaction between Age and Sex.
```
shap.dependence_plot('Age', shap_values, X_test, interaction_index = 'Sex')
```
We see that while Age > 50 is generally bad (positive Shapley value), being a woman (red points) generally reduces the impact of age. This makes sense since we know that women generally live longer than men.
Run the next cell to see the interaction between Poverty index and Age
```
shap.dependence_plot('Poverty index', shap_values, X_test, interaction_index='Age')
```
We see that the impact of poverty index drops off quickly, and for higher income individuals age begins to explain much of variation in the impact of poverty index. We encourage you to try some other pairs and see what other interesting relationships you can find!
Congratulations! You've completed the final assignment of course 3, well done!
| github_jupyter |
# Parameterizations
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import pyqg
year = 24*60*60*360.
```
## Define parameterization
In this example, we'll implement the physical parameterization learned by symbolic regression in [Zanna and Bolton 2020](https://doi.org/10.1029/2020GL088376):
```
def zb2020_parameterization(m, factor=-46761284):
"""Implements the parameterization from Equation 6 of
https://doi.org/10.1029/2020GL088376"""
ik = 1j * m.k
il = 1j * m.l
# Compute relative velocity derivatives in spectral space
uh = m.fft(m.u)
vh = m.fft(m.v)
vx = m.ifft(vh * ik)
vy = m.ifft(vh * il)
uy = m.ifft(uh * il)
ux = m.ifft(uh * ik)
# Compute ZB2020 basis functions
rel_vort = vx - uy
shearing = vx + uy
stretching = ux - vy
# Combine them in real space and take their FFT
rv_stretch = m.fft(rel_vort * stretching)
rv_shear = m.fft(rel_vort * shearing)
sum_sqs = m.fft(rel_vort**2 + shearing**2 + stretching**2) / 2.0
# Take spectral-space derivatives and multiply by the scaling factor
Su = factor * m.ifft(ik*(sum_sqs - rv_shear) + il*rv_stretch)
Sv = factor * m.ifft(il*(sum_sqs + rv_shear) + ik*rv_stretch)
return Su, Sv
```
This function parameterizes subgrid forcing in terms of changes that should be applied to the real-space velocity tendencies. To use it in a `pyqg.Model`, we can pass it during initialization as `uv_parameterization`, and then its outputs will be added to the spectral-space potential vorticity tendency $\partial_t \hat{q}$ (after suitable transformations, and at each timestep). Note that we can also define parameterizions in terms of PV directly by initializing the model with a `q_parameterization` function (which should return a single array rather than a tuple of two).
## Run models
To illustrate the effect of the parameterization, we'll run three variants of models:
* one without parameterization at `nx=64` resolution (where $\Delta x$ is larger than the deformation radius $r_d$, preventing the model from fully resolving eddies),
* one with parameterization at `nx=64` resolution,
* one at a higher `nx=256` resolution (where $\Delta x$ is ~4x finer than the deformation radius, so eddies can be almost fully resolved).
```
%%time
m_unparam = pyqg.QGModel(nx=64, dt=3600., tmax=10*year, tavestart=5*year, twrite=10000)
m_unparam.run()
%%time
m_param = pyqg.QGModel(nx=64, dt=3600., tmax=10*year, tavestart=5*year, twrite=10000,
uv_parameterization=zb2020_parameterization)
m_param.run()
%%time
m_hires = pyqg.QGModel(nx=256, dt=3600., tmax=10*year, tavestart=5*year, twrite=10000)
m_hires.run()
```
Note that parameterization does slow down the simulation, but it's still significantly faster than running a high-resolution simulation.
## Plot spectra
Now let's examine how adding the parameterization to our simulation changed its energetic properties:
```
from pyqg import diagnostic_tools as tools
from scipy.stats import linregress
plt.figure(figsize=(10,6))
plt.title("Comparison of kinetic energy spectra", fontsize=16)
for m,label in zip([m_unparam, m_param, m_hires], ['Lo-res','Lo-res + parameterization','Hi-res']):
# Calculate the spectrum
k, spectrum = tools.calc_ispec(m, m.get_diagnostic('KEspec').sum(axis=0))
# Do a power law fit on data in the intertial range
kmin = 5e-5
kmax = 1e-4
i = np.argmin(np.abs(np.log(k) - np.log(kmin)))
j = np.argmin(np.abs(np.log(k) - np.log(kmax)))
lr = linregress(np.log(k[i:j]), np.log(spectrum[i:j]))
# Plot the spectrum with the linear fit
loglog_plot = plt.loglog(k, spectrum, lw=5, alpha=0.9,
label=(label + "\n($\propto k^{" + str(lr.slope.round(2)) + "}$)"))
plt.ylim(1e-12, 1e-8)
plt.xlim(2e-5, 2e-4)
plt.xlabel("Isotropic wavenumber")
plt.ylabel("KEspec")
plt.legend(loc='best', fontsize=14)
plt.show()
```
Although the overall energy is lower, the parameterization ends up with an inverse cascade whose decay rate (log-log slope over intertial lengthscales) is much closer to that of the high-resolution model.
| github_jupyter |
<b>Define environment variables</b>
To be used in future training steps. Note that the BUCKET_NAME defined below must exist in the GCP project.
```
%env BUCKET_NAME=<ADD DETAILS HERE>
%env JOB_NAME=<ADD DETAILS HERE>
%env TRAINING_PACKAGE_PATH=./trainer/
%env MAIN_TRAINER_MODULE=trainer.rf_trainer
%env REGION=<ADD DETAILS HERE>
%env RUNTIME_VERSION=<ADD DETAILS HERE>
%env PYTHON_VERSION=<ADD DETAILS HERE>
%env SCALE_TIER=<ADD DETAILS HERE>
%env MODEL_NAME=<ADD DETAILS HERE>
%env PROJECT_ID=<ADD DETAILS HERE>
%env DATASET_ID=b<ADD DETAILS HERE>
%env VERSION_NAME=<ADD DETAILS HERE>
%env FRAMEWORK=<ADD DETAILS HERE>
# Training and testing files must be in a cloud storage bucket before training runs.
!gsutil mb gs://${BUCKET_NAME}
!gsutil cp train.csv gs://${BUCKET_NAME}
!gsutil cp test.csv gs://${BUCKET_NAME}
# Remove output from previous runs, if any.
!rm input.json
```
<b>Perform training locally with default parameters</b>
[Using AI Platform for Local Training](https://cloud.google.com/sdk/gcloud/reference/ai-platform/local/train)
```
# Give the service account for this project an "Editor" role in IAM for all users of this environment
# to have Bigquery access. The first time this cell is run, set create-data and hp-tune to True. This
# creates input files and the results of hyperparameter tuning available. You can set them to false for
# subsequent runs.
# Fill the incomplete details to train the model locally
!gcloud ai-platform local train \
--<ADD DETAILS HERE> \
--<ADD DETAILS HERE> \
-- \
--<ADD DETAILS HERE> \
--<ADD DETAILS HERE> \
--create-data=True \
--hp-tune=True \
--num-hp-iterations=3
```
<b>Perform training on AI Platform</b>
The training job can also be run on AI Platform.
Important: A single training job (either locally or using AI Platform) must complete with the --create-data and --hp-tune flags set to True for the remainig functionality to complete.
Note that we've updated the compute allocated to the master machine for this job to allow for more muscle.
```
# The first time this cell is run, set create-data and hp-tune to True. This
# creates input files and the results of hyperparameter tuning available. You can set them to false for
# subsequent runs.
now = !date +"%Y%m%d_%H%M%S"
%env JOB_NAME=black_friday_job_$now.s
!gcloud ai-platform jobs submit training $JOB_NAME \
--job-dir gs://${BUCKET_NAME}/rf-job-dir \
--<ADD DETAILS HERE> \
--<ADD DETAILS HERE> \
--<ADD DETAILS HERE> \
--<ADD DETAILS HERE> \
--<ADD DETAILS HERE> \
--<ADD DETAILS HERE> \
--master-machine-type n1-highcpu-16 \
-- \
--<ADD DETAILS HERE> \
--<ADD DETAILS HERE> \
--<ADD DETAILS HERE> \
--dataset-id $DATASET_ID \
--<ADD DETAILS HERE> \
--<ADD DETAILS HERE> \
--<ADD DETAILS HERE>
# Stream logs so that training is done before subsequent cells are run.
# Remove '> /dev/null' to see step-by-step output of the model build steps.
!gcloud ai-platform jobs stream-logs $JOB_NAME > /dev/null
# Model should exit with status "SUCCEEDED"
!gcloud ai-platform jobs describe $JOB_NAME --format="value(state)"
```
<b>Host the trained model on AI Platform</b>
Because our raw prediction output from the model is a numpy array that needs to be converted into a product category, we'll need to implement a custom prediction module.
First, execute the setup script to create a distribution tarball
```
!python setup.py sdist --formats=gztar
```
Next copy the tarball over to Cloud Storage
```
!gsutil cp dist/trainer-0.1.tar.gz gs://${BUCKET_NAME}/staging-dir/trainer-0.1.tar.gz
```
Create a new model on AI Platform. Note that this needs to be done just once, and future iterations are saved as "versions" of the model.
```
# write the command to create a ML MODEL
<ADD DETAILS HERE>
```
Next we create new version using our trained model
```
!gcloud beta ai-platform versions create $VERSION_NAME \
--model $MODEL_NAME \
--origin gs://${BUCKET_NAME}/black_friday_${JOB_NAME}/ \
--runtime-version=1.14 \
--python-version=3.5 \
--package-uris gs://${BUCKET_NAME}/staging-dir/trainer-0.1.tar.gz \
--prediction-class predictor.MyPredictor
```
<b>Prepare a sample for inference</b>
```
!python generate_sample.py \
--project-id $PROJECT_ID \
--bucket-name ${BUCKET_NAME}
```
<b>Make an inference on a new sample.</b>
Pass the sample object to the model hosted in AI Platform to return a prediction.
```
# make an online prediction
!gcloud ai-platform predict --model $MODEL_NAME --version \
$VERSION_NAME --json-instances input.json
```
| github_jupyter |
```
#@title Copyright 2020 Google LLC. Double-click here for license information.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Linear Regression with a Real Dataset
This Colab uses a real dataset to predict the prices of houses in California.
## Learning Objectives:
After doing this Colab, you'll know how to do the following:
* Read a .csv file into a [pandas](https://developers.google.com/machine-learning/glossary/#pandas) DataFrame.
* Examine a [dataset](https://developers.google.com/machine-learning/glossary/#data_set).
* Experiment with different [features](https://developers.google.com/machine-learning/glossary/#feature) in building a model.
* Tune the model's [hyperparameters](https://developers.google.com/machine-learning/glossary/#hyperparameter).
## The Dataset
The [dataset for this exercise](https://developers.google.com/machine-learning/crash-course/california-housing-data-description) is based on 1990 census data from California. The dataset is old but still provides a great opportunity to learn about machine learning programming.
## Use the right version of TensorFlow
The following hidden code cell ensures that the Colab will run on TensorFlow 2.X.
```
#@title Run on TensorFlow 2.x
%tensorflow_version 2.x
```
## Import relevant modules
The following hidden code cell imports the necessary code to run the code in the rest of this Colaboratory.
```
#@title Import relevant modules
import pandas as pd
import tensorflow as tf
from matplotlib import pyplot as plt
# The following lines adjust the granularity of reporting.
pd.options.display.max_rows = 10
pd.options.display.float_format = "{:.1f}".format
```
## The dataset
Datasets are often stored on disk or at a URL in [.csv format](https://wikipedia.org/wiki/Comma-separated_values).
A well-formed .csv file contains column names in the first row, followed by many rows of data. A comma divides each value in each row. For example, here are the first five rows of the .csv file file holding the California Housing Dataset:
```
"longitude","latitude","housing_median_age","total_rooms","total_bedrooms","population","households","median_income","median_house_value"
-114.310000,34.190000,15.000000,5612.000000,1283.000000,1015.000000,472.000000,1.493600,66900.000000
-114.470000,34.400000,19.000000,7650.000000,1901.000000,1129.000000,463.000000,1.820000,80100.000000
-114.560000,33.690000,17.000000,720.000000,174.000000,333.000000,117.000000,1.650900,85700.000000
-114.570000,33.640000,14.000000,1501.000000,337.000000,515.000000,226.000000,3.191700,73400.000000
```
### Load the .csv file into a pandas DataFrame
This Colab, like many machine learning programs, gathers the .csv file and stores the data in memory as a pandas Dataframe. pandas is an open source Python library. The primary datatype in pandas is a DataFrame. You can imagine a pandas DataFrame as a spreadsheet in which each row is identified by a number and each column by a name. pandas is itself built on another open source Python library called NumPy. If you aren't familiar with these technologies, please view these two quick tutorials:
* [NumPy](https://colab.research.google.com/github/google/eng-edu/blob/master/ml/cc/exercises/numpy_ultraquick_tutorial.ipynb?utm_source=linearregressionreal-colab&utm_medium=colab&utm_campaign=colab-external&utm_content=numpy_tf2-colab&hl=en)
* [Pandas DataFrames](https://colab.research.google.com/github/google/eng-edu/blob/master/ml/cc/exercises/pandas_dataframe_ultraquick_tutorial.ipynb?utm_source=linearregressionreal-colab&utm_medium=colab&utm_campaign=colab-external&utm_content=pandas_tf2-colab&hl=en)
The following code cell imports the .csv file into a pandas DataFrame and scales the values in the label (`median_house_value`):
```
# Import the dataset.
training_df = pd.read_csv(filepath_or_buffer="https://download.mlcc.google.com/mledu-datasets/california_housing_train.csv")
# Scale the label.
training_df["median_house_value"] /= 1000.0
# Print the first rows of the pandas DataFrame.
training_df.head()
```
Scaling `median_house_value` puts the value of each house in units of thousands. Scaling will keep loss values and learning rates in a friendlier range.
Although scaling a label is usually *not* essential, scaling features in a multi-feature model usually *is* essential.
## Examine the dataset
A large part of most machine learning projects is getting to know your data. The pandas API provides a `describe` function that outputs the following statistics about every column in the DataFrame:
* `count`, which is the number of rows in that column. Ideally, `count` contains the same value for every column.
* `mean` and `std`, which contain the mean and standard deviation of the values in each column.
* `min` and `max`, which contain the lowest and highest values in each column.
* `25%`, `50%`, `75%`, which contain various [quantiles](https://developers.google.com/machine-learning/glossary/#quantile).
```
# Get statistics on the dataset.
training_df.describe()
```
### Task 1: Identify anomalies in the dataset
Do you see any anomalies (strange values) in the data?
```
#@title Double-click to view a possible answer.
# The maximum value (max) of several columns seems very
# high compared to the other quantiles. For example,
# example the total_rooms column. Given the quantile
# values (25%, 50%, and 75%), you might expect the
# max value of total_rooms to be approximately
# 5,000 or possibly 10,000. However, the max value
# is actually 37,937.
# When you see anomalies in a column, become more careful
# about using that column as a feature. That said,
# anomalies in potential features sometimes mirror
# anomalies in the label, which could make the column
# be (or seem to be) a powerful feature.
# Also, as you will see later in the course, you
# might be able to represent (pre-process) raw data
# in order to make columns into useful features.
```
## Define functions that build and train a model
The following code defines two functions:
* `build_model(my_learning_rate)`, which builds a randomly-initialized model.
* `train_model(model, feature, label, epochs)`, which trains the model from the examples (feature and label) you pass.
Since you don't need to understand model building code right now, we've hidden this code cell. You may optionally double-click the following headline to see the code that builds and trains a model.
```
#@title Define the functions that build and train a model
def build_model(my_learning_rate):
"""Create and compile a simple linear regression model."""
# Most simple tf.keras models are sequential.
model = tf.keras.models.Sequential()
# Describe the topography of the model.
# The topography of a simple linear regression model
# is a single node in a single layer.
model.add(tf.keras.layers.Dense(units=1,
input_shape=(1,)))
# Compile the model topography into code that TensorFlow can efficiently
# execute. Configure training to minimize the model's mean squared error.
model.compile(optimizer=tf.keras.optimizers.RMSprop(lr=my_learning_rate),
loss="mean_squared_error",
metrics=[tf.keras.metrics.RootMeanSquaredError()])
return model
def train_model(model, df, feature, label, epochs, batch_size):
"""Train the model by feeding it data."""
# Feed the model the feature and the label.
# The model will train for the specified number of epochs.
history = model.fit(x=df[feature],
y=df[label],
batch_size=batch_size,
epochs=epochs)
# Gather the trained model's weight and bias.
trained_weight = model.get_weights()[0]
trained_bias = model.get_weights()[1]
# The list of epochs is stored separately from the rest of history.
epochs = history.epoch
# Isolate the error for each epoch.
hist = pd.DataFrame(history.history)
# To track the progression of training, we're going to take a snapshot
# of the model's root mean squared error at each epoch.
rmse = hist["root_mean_squared_error"]
return trained_weight, trained_bias, epochs, rmse
print("Defined the create_model and traing_model functions.")
```
## Define plotting functions
The following [matplotlib](https://developers.google.com/machine-learning/glossary/#matplotlib) functions create the following plots:
* a scatter plot of the feature vs. the label, and a line showing the output of the trained model
* a loss curve
You may optionally double-click the headline to see the matplotlib code, but note that writing matplotlib code is not an important part of learning ML programming.
```
#@title Define the plotting functions
def plot_the_model(trained_weight, trained_bias, feature, label):
"""Plot the trained model against 200 random training examples."""
# Label the axes.
plt.xlabel(feature)
plt.ylabel(label)
# Create a scatter plot from 200 random points of the dataset.
random_examples = training_df.sample(n=200)
plt.scatter(random_examples[feature], random_examples[label])
# Create a red line representing the model. The red line starts
# at coordinates (x0, y0) and ends at coordinates (x1, y1).
x0 = 0
y0 = trained_bias
x1 = 10000
y1 = trained_bias + (trained_weight * x1)
plt.plot([x0, x1], [y0, y1], c='r')
# Render the scatter plot and the red line.
plt.show()
def plot_the_loss_curve(epochs, rmse):
"""Plot a curve of loss vs. epoch."""
plt.figure()
plt.xlabel("Epoch")
plt.ylabel("Root Mean Squared Error")
plt.plot(epochs, rmse, label="Loss")
plt.legend()
plt.ylim([rmse.min()*0.97, rmse.max()])
plt.show()
print("Defined the plot_the_model and plot_the_loss_curve functions.")
```
## Call the model functions
An important part of machine learning is determining which [features](https://developers.google.com/machine-learning/glossary/#feature) correlate with the [label](https://developers.google.com/machine-learning/glossary/#label). For example, real-life home-value prediction models typically rely on hundreds of features and synthetic features. However, this model relies on only one feature. For now, you'll arbitrarily use `total_rooms` as that feature.
```
# The following variables are the hyperparameters.
learning_rate = 0.01
epochs = 30
batch_size = 30
# Specify the feature and the label.
my_feature = "total_rooms" # the total number of rooms on a specific city block.
my_label="median_house_value" # the median value of a house on a specific city block.
# That is, you're going to create a model that predicts house value based
# solely on total_rooms.
# Discard any pre-existing version of the model.
my_model = None
# Invoke the functions.
my_model = build_model(learning_rate)
weight, bias, epochs, rmse = train_model(my_model, training_df,
my_feature, my_label,
epochs, batch_size)
print("\nThe learned weight for your model is %.4f" % weight)
print("The learned bias for your model is %.4f\n" % bias )
plot_the_model(weight, bias, my_feature, my_label)
plot_the_loss_curve(epochs, rmse)
```
A certain amount of randomness plays into training a model. Consequently, you'll get different results each time you train the model. That said, given the dataset and the hyperparameters, the trained model will generally do a poor job describing the feature's relation to the label.
## Use the model to make predictions
You can use the trained model to make predictions. In practice, [you should make predictions on examples that are not used in training](https://developers.google.com/machine-learning/crash-course/training-and-test-sets/splitting-data). However, for this exercise, you'll just work with a subset of the same training dataset. A later Colab exercise will explore ways to make predictions on examples not used in training.
First, run the following code to define the house prediction function:
```
def predict_house_values(n, feature, label):
"""Predict house values based on a feature."""
batch = training_df[feature][10000:10000 + n]
predicted_values = my_model.predict_on_batch(x=batch)
print("feature label predicted")
print(" value value value")
print(" in thousand$ in thousand$")
print("--------------------------------------")
for i in range(n):
print ("%5.0f %6.0f %15.0f" % (training_df[feature][10000 + i],
training_df[label][10000 + i],
predicted_values[i][0] ))
```
Now, invoke the house prediction function on 10 examples:
```
predict_house_values(10, my_feature, my_label)
```
### Task 2: Judge the predictive power of the model
Look at the preceding table. How close is the predicted value to the label value? In other words, does your model accurately predict house values?
```
#@title Double-click to view the answer.
# Most of the predicted values differ significantly
# from the label value, so the trained model probably
# doesn't have much predictive power. However, the
# first 10 examples might not be representative of
# the rest of the examples.
```
## Task 3: Try a different feature
The `total_rooms` feature had only a little predictive power. Would a different feature have greater predictive power? Try using `population` as the feature instead of `total_rooms`.
Note: When you change features, you might also need to change the hyperparameters.
```
my_feature = "?" # Replace the ? with population or possibly
# a different column name.
# Experiment with the hyperparameters.
learning_rate = 2
epochs = 3
batch_size = 120
# Don't change anything below this line.
my_model = build_model(learning_rate)
weight, bias, epochs, rmse = train_model(my_model, training_df,
my_feature, my_label,
epochs, batch_size)
plot_the_model(weight, bias, my_feature, my_label)
plot_the_loss_curve(epochs, rmse)
predict_house_values(15, my_feature, my_label)
#@title Double-click to view a possible solution.
my_feature = "population" # Pick a feature other than "total_rooms"
# Possibly, experiment with the hyperparameters.
learning_rate = 0.05
epochs = 18
batch_size = 3
# Don't change anything below.
my_model = build_model(learning_rate)
weight, bias, epochs, rmse = train_model(my_model, training_df,
my_feature, my_label,
epochs, batch_size)
plot_the_model(weight, bias, my_feature, my_label)
plot_the_loss_curve(epochs, rmse)
predict_house_values(10, my_feature, my_label)
```
Did `population` produce better predictions than `total_rooms`?
```
#@title Double-click to view the answer.
# Training is not entirely deterministic, but population
# typically converges at a slightly higher RMSE than
# total_rooms. So, population appears to be about
# the same or slightly worse at making predictions
# than total_rooms.
```
## Task 4: Define a synthetic feature
You have determined that `total_rooms` and `population` were not useful features. That is, neither the total number of rooms in a neighborhood nor the neighborhood's population successfully predicted the median house price of that neighborhood. Perhaps though, the *ratio* of `total_rooms` to `population` might have some predictive power. That is, perhaps block density relates to median house value.
To explore this hypothesis, do the following:
1. Create a [synthetic feature](https://developers.google.com/machine-learning/glossary/#synthetic_feature) that's a ratio of `total_rooms` to `population`. (If you are new to pandas DataFrames, please study the [Pandas DataFrame Ultraquick Tutorial](https://colab.research.google.com/github/google/eng-edu/blob/master/ml/cc/exercises/pandas_dataframe_ultraquick_tutorial.ipynb?utm_source=linearregressionreal-colab&utm_medium=colab&utm_campaign=colab-external&utm_content=pandas_tf2-colab&hl=en).)
2. Tune the three hyperparameters.
3. Determine whether this synthetic feature produces
a lower loss value than any of the single features you
tried earlier in this exercise.
```
# Define a synthetic feature named rooms_per_person
training_df["rooms_per_person"] = ? # write your code here.
# Don't change the next line.
my_feature = "rooms_per_person"
# Assign values to these three hyperparameters.
learning_rate = ?
epochs = ?
batch_size = ?
# Don't change anything below this line.
my_model = build_model(learning_rate)
weight, bias, epochs, rmse = train_model(my_model, training_df,
my_feature, my_label,
epochs, batch_size)
plot_the_loss_curve(epochs, rmse)
predict_house_values(15, my_feature, my_label)
#@title Double-click to view a possible solution to Task 4.
# Define a synthetic feature
training_df["rooms_per_person"] = training_df["total_rooms"] / training_df["population"]
my_feature = "rooms_per_person"
# Tune the hyperparameters.
learning_rate = 0.06
epochs = 24
batch_size = 30
# Don't change anything below this line.
my_model = build_model(learning_rate)
weight, bias, epochs, mae = train_model(my_model, training_df,
my_feature, my_label,
epochs, batch_size)
plot_the_loss_curve(epochs, mae)
predict_house_values(15, my_feature, my_label)
```
Based on the loss values, this synthetic feature produces a better model than the individual features you tried in Task 2 and Task 3. However, the model still isn't creating great predictions.
## Task 5. Find feature(s) whose raw values correlate with the label
So far, we've relied on trial-and-error to identify possible features for the model. Let's rely on statistics instead.
A **correlation matrix** indicates how each attribute's raw values relate to the other attributes' raw values. Correlation values have the following meanings:
* `1.0`: perfect positive correlation; that is, when one attribute rises, the other attribute rises.
* `-1.0`: perfect negative correlation; that is, when one attribute rises, the other attribute falls.
* `0.0`: no correlation; the two column's [are not linearly related](https://en.wikipedia.org/wiki/Correlation_and_dependence#/media/File:Correlation_examples2.svg).
In general, the higher the absolute value of a correlation value, the greater its predictive power. For example, a correlation value of -0.8 implies far more predictive power than a correlation of -0.2.
The following code cell generates the correlation matrix for attributes of the California Housing Dataset:
```
# Generate a correlation matrix.
training_df.corr()
```
The correlation matrix shows nine potential features (including a synthetic
feature) and one label (`median_house_value`). A strong negative correlation or strong positive correlation with the label suggests a potentially good feature.
**Your Task:** Determine which of the nine potential features appears to be the best candidate for a feature?
```
#@title Double-click here for the solution to Task 5
# The `median_income` correlates 0.7 with the label
# (median_house_value), so median_income` might be a
# good feature. The other seven potential features
# all have a correlation relatively close to 0.
# If time permits, try median_income as the feature
# and see whether the model improves.
```
Correlation matrices don't tell the entire story. In later exercises, you'll find additional ways to unlock predictive power from potential features.
**Note:** Using `median_income` as a feature may raise some ethical and fairness
issues. Towards the end of the course, we'll explore ethical and fairness issues.
| github_jupyter |
<a href="https://colab.research.google.com/github/Shailesh0209/x_tools_in_ds_dipoma-iitm/blob/main/x_get_data_w2_TDS.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#L2.2: Get the data-Nominatim Open Street Maps
## Scraping using Geocoding API of Open Street Maps(OSM)
We would be using the Nominatim API to scrape geocoding imformation of any open ended address text using Python
```
# no need to install these if using Google Colab
!pip install geopandas
!pip install geopy
# import nominatim api
from geopy.geocoders import Nominatim
# activate nominatim geocoder
locator = Nominatim (user_agent="sx81")
# type any address text
location = locator.geocode("Chennai")
# print latitude and longitude of the address
print("Latitude={}, Longitude={}".format(location.latitude, location.longitude))
# the API output has multiple other details as a json like altitude, latitude
# longitude, correct raw address, etc.
# printing all the information
location.raw, location.point, location.longitude, location.latitude, location.altitude, location.address
# typing another address
location2 = locator.geocode("IIT Madras")
location2.raw, location2.point, location2.longitude, location2.latitude, location2.altitude, location2.address
```
# L2.3 Get the data BBC Weather location service
## A tutorial to scrape the location ID of any city in BBC Weather
Thsi code snippet takes city name as input and it hits the BBC Weather API with a request for location ID. This location ID is used as input in the next part of the code to scrape weather forecast for the city using this location ID.
Web scraping might not be legal always. It is a good idea to check the terms of the website you plan to scrape before proceeding. Also, if your code requests a url from a server multiple times, it is a good practice to either cache your requests, o insert a timed delay between consecutive requests.
```
import os
import requests # to get the webpage
import json # to convert API to json format
from urllib.parse import urlencode
import numpy as np
import pandas as pd
import re # regular expressio operators
test_city = "New York"
location_url = 'https://locator-service.api.bcci.co.uk/locations?' + urlencode({
'api_key': 'AGbFAKx58hyjQScCXIYrxuEwJh2W2cmv',
's': test_city,
'stack': 'aws',
'locale': 'en',
'filter': 'international',
'place-types': 'settlement,airport, district',
'order': 'importance',
'a': 'true',
'format': 'json'
})
location_url
result = requests.get(location_url, verify=False).text
result
# Print locationid
result['response']['results']['results'][0]['id']
```
###Creating a function to output location id by taking any city name as input.
```
def getlocid(city):
city = city.lower() # convert city name to lowercase to standardize format
# Convert into an API call using URL encoding
location_url = 'https://locator-services.api.bcci.co.uk/locations?' + urlencode({
'api_key': 'AGbFAKx58hyjQScCXIYrxuEwJh2W2cmv',
's': city,
'stack': 'aws',
'locale': 'en',
'filter': 'international',
'place-types': 'settlement, airport,district',
'order': 'true',
'format': 'json'
})
result = requests.get(location_url).json()
locid = result['response']['results']['results'][0]['id']
return locid
getlocid('Toronto')
```
#L2.4 Get the data-Scraping with Excel
```
```
#L2.5 Get the data-Scraping with Python
## WEB Scrapping IMDB
In this exercise we'll look at scraping data from IMDB. Our goal is to convert the top 250 list of movies in IMDB intoa tabular form using Python. This data can then be used for further analysis.
1: Import Necessary Libraries
```
from bs4 import BeautifulSoup as bs
import requests # to access website
import pandas as pd
```
2: Load the webpage
```
r = requests.get("https://www.imdb.com/chart/top/")
# Convert to a beautiful soup object
soup = bs(r.content)
# Print out HTML
Contents = soup.prettify()
```
3: Creating empty list
```
movie_title = []
movie_year = []
movie_rating = []
```
4: Extract HTML tag contents
```
imdb_table = soup.find(class_="chart full-width")
movie_titlecolumn= imdb_table.find_all(class_="titleColumn")
movie_ratingscolumn = imdb_table.find_all(class_="ratingColumn imdbRating")
for row in movie_titlecolumn:
title = row.a.text # tag content extraction
movie_title.append(title)
movie_title
for row in movie_titlecolumn:
year = row.span.text # tag content extraction # gives text contain inside span tag
movie_year.append(year)
movie_year
for row in movie_ratingscolumn:
rating = row.strong.text # tag content extraction
movie_rating.append(rating)
movie_rating
```
5: Create DataFrame
```
movie_df = pd.DataFrame({'Movie Title': movie_title, 'Year of Release': movie_year, 'IMDB Rating': movie_rating})
movie_df
movie_df.head()
```
#L2.6: Get the data-Wikimedia
This is a self-explanaory short tutorial on using the wikipedia library to extract information from wikipedia.
```
!pip install wikipedia
import wikipedia as wk
print(wk.search("Isaac Newton"))
print(wk.search("IIT Madras", results=2))
print(wk.summary("Isaac Newton"))
print(wk.summary("IIT Madras", sentences=2))
full_page = wk.page("IIT Madras")
print(full_page.content)
print(full_page.url)
print(full_page.references)
print(full_page.images)
print(full_page.images[0])
# extract html code of wikipedia page based on any search text
html = wk.page("IIT Madras").html().encode("UTF-8")
import pandas as pd
df = pd.read_html(html)[6]
df
```
# L2.7:Get the data-Scrape BBC weather with Python
## A tutorial to scrape the web.
This example scrapes the BBC weather for any specific city, and collects weather forecast for the next 14 days and saves it as a csv file.
Web scraping might not be legal always. It is a good idea to check the terms of the website you plan to scrape before proceeding. Also, if your code requests a url from a server ultiple times, it is a good practice to either cache your requests, or insert a timed delay between consecutive requests.
```
import json # to convert API to json format
from urllib.parse import urlencode
import requests # to get the webpage
from bs4 import BeautifulSoup # to parse the webpage
import pandas as pd
import re # regular expression operators
from datetime import datetime
```
We now GET the webpage of interest, from the server
```
required_city = "Mumbai"
location_url = 'https://loator-service.api.bbci.co.uk/locations?' + urlencode({
'api_key': 'AGbFAKx58hyjQScCXIYrxuEwJh2W2cmv',
's': required_city,
'stack': 'aws',
'locale': 'en',
'filter': 'international',
'pace-types': 'settlement,airport,district',
'order': 'importance',
'a': 'true',
'format': 'json'
})
location_url
result = requests.get(location_url, verify=False).json()
result
""" url = 'https//www.bbc.com/weather/1275339' # url to BBC weather,
corresponding to a specific city (Mumbai, in this example"""
url = 'https://www.bbc.com/weather/'+result['response']['results']['results'][0]['id']
response = requests.get(url, verify=False)
```
Next, we initiate an instance fo BeautifulSoup.
```
soup = BeautifulSoup(response.content, 'html.parser')
```
The information we want (daily high and low temp., and daily weather summary), are in specific blocks on the webpage. We need to find the block type, type of identifier, and the identifier name (all these can be figured out by right clicking on the webpage and selecting 'Inspect' on the Chrome browser; similar modus operand for the browsers)
```
daily_high_values = soup.find_all('span',
attrs={'class':
'wr-day-temperature_high-value'})
# block-type: span; identifier type: class; and
#class name: wr-day-temperature_high-value
daily_high_values
daily_low_values = soup.find_all('span',
attrs={'class': 'wr-day-temperature_low-value'})
daily_low-values
daily_summary = soup.find('div',
attrs={'class': 'wr-day-summary'})
daily_sumary
daily_summary.text
```
`General book keeping`:
With the code snippet in the cell above, we get forecast data for 14 days, including today. We will now post process the data to first extract the required information/text and discard all the html wrapper code, then combine all variables into one common list, and finally convert it into a pandas dataFrame.
```
daily_high_values[0].text.strip()
daily_high_values[5].text.strip()
daily_high_values[0].text.strip().split()[0]
daily_high_values_list = [daily_high_values[i].text.strip().split()[0] for i in range(len(daily_high_values))]
daily_high_values_list
daily_low_values_list = [daily_low_values[i].text.strip().split()[0] for i in range(len(daily_low_values))]
daily_low_values_list
daily_summary.text
daily_summary_list = re.fidall('[a-zA-Z][^A-Z]*', daily_summary.text) # split the string on uppercase
daily_summary_list
datelist = pd.date_range(datetime.today(), periods=len(daily_high_values)).tolist()
datelist
datelist = [datelist[i].date().strftime('%y-%m-%d') for i in range(len(datelist))]
datelist
zipped = zip(datelist, daily_high_values_list, daily_low_values_list, daily_summary_list)
df = pd.DataFrame(list(zipped), columns=['Date', 'High', 'Low', 'Summary'])
display(df)
# remove the 'degree' character
df.High = df.High.replace('\°','',regex=True).astype(float)
df.Low = df.Low.replace('\°','',regex=True).astype(float)
display(df)
```
Extract the name of the city for which data is gathered
```
# location = soup.find('div', attrs={'class':'wr-c-location'})
location = soup.find('h1', attrs={'id':'wr-location-name-id'})
location.text.split()
# create a recording
filename_csv = location.text.split()[0]+'.csv'
df.to_csv(filename_csv,index = None)
filename_xlsx = location.text.split()[0]+'.xlsx'
df.to_exvel(filename_xlsx)
```
#L2.8: Get the data-Scraping PDFs
```
```
# GA2
####2.
```
import pandas as pd
data = pd.read_excel("https://rbidocs.rbi.org.in/rdocs/Content/DOCs/IFCB2009_85.xlsx")
data.head()
data.shape
1 + 2
b = print(1)
type(b)
data.groupby("CITY 1").CHENNAI.count()
data
data.
data['CITY 1'].value_counts()
```
####5.
```
!pip install geopandas
!pip install geopy
# import nominatim api
from geopy.geocoders import Nominatim
locator = Nominatim(user_agent='sx7987')
location = locator.geocode("Chennai")
print(type("Latitude={}, Longitude={}".format(location.latitude, location.longitude)))
location.raw, location.point, location.longitude, location.latitude, location.altitude, location.address
```
####7.
```
import requests
from bs4 import BeautifulSoup
from urllib.parse import urljoin
import os
url = 'https://www.premierleague.com/publications'
folder_location = r'/content/drive/MyDrive/Colab Notebooks/premier_league'
if not os.path.exists(folder_location):
os.mkdir(folder_location)
response = requests.get(url)
soup = BeautifulSoup(response.text, "html.parser")
for link in soup.select("a[href$='.pdf']"):
filename = os.path.join(folder_location, link['href'].split('/')[-1])
with open(filename, 'wb') as f:
f.write(requests.get(urljoin(url, link['href'])).content)
import requests
from bs4 import BeautifulSoup
from urllib.parse import urljoin
import os
url = 'https://www.premierleague.com/publications'
folder_location = r'/content/drive/MyDrive/Colab Notebooks/premier_league'
if not os.path.exists(folder_location):
os.mkdir(folder_location)
response = requests.get(url)
soup = BeautifulSoup(response.text, "html.parser")
for link in soup.select("a[href$='.pdf']"):
filename = os.path.join(folder_location, link['href'].split('/')[-1])
with open(filename, 'wb') as f:
f.write(requests.get(urljoin(url,link['href'])).content)
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><ul class="toc-item"><li><span><a href="#Import-the-libraries" data-toc-modified-id="Import-the-libraries-0.1"><span class="toc-item-num">0.1 </span>Import the libraries</a></span></li><li><span><a href="#Example" data-toc-modified-id="Example-0.2"><span class="toc-item-num">0.2 </span>Example</a></span></li></ul></li><li><span><a href="#Status" data-toc-modified-id="Status-1"><span class="toc-item-num">1 </span>Status</a></span></li></ul></div>
<a href="https://colab.research.google.com/github/butchland/fastai_xla_extensions/blob/master/explore_nbs/Explore_gpu_training_comparison.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# A comparison of the GPU vs TPU training
This is the GPU version of the index notebook
of the fastai xla extensions index page
The only difference between the two is the
use of a GPU runtime
```
#hide
!pip install nbdev --upgrade > /dev/null
#hide
#colab
!curl -s https://course.fast.ai/setup/colab | bash
```
Install fastai2 and the fastai_xla_extensions packages
```
#hide_output
!pip install fastai2 > /dev/null
```
### Import the libraries
Import the fastai2 libraries
```
from fastai2.vision.all import *
```
### Example
Build a Pets classifier -- adapted from fastai course [Lesson 5 notebook](https://github.com/fastai/course-v4/blob/master/nbs/05_pet_breeds.ipynb)
```
#hide
path = Path('.')
```
Load Oxford-IIT Pets dataset
```
#colab
path = untar_data(URLs.PETS)/'images'
Path.BASE_PATH = path
pat = r'(.+)_\d+.jpg$'
```
Create Fastai DataBlock
_Note that batch transforms are currently
set to none as they seem to slow the training
on the TPU (for investigation)._
```
datablock = DataBlock(
blocks=(ImageBlock,CategoryBlock),
get_items=get_image_files,
splitter=RandomSplitter(seed=42),
get_y=using_attr(RegexLabeller(pat),'name'),
item_tfms=Resize(224),
batch_tfms=[]
)
```
Get a TPU device
Set the dataloaders to use GPU
```
#colab
dls = datablock.dataloaders(path)
#colab
dls.show_batch()
```
Use the normal wrapper
```
opt_func = Adam
```
Create a Fastai CNN Learner
```
#colab
learner = cnn_learner(dls, resnet34, metrics=accuracy, opt_func=opt_func)
```
Using the `lr_find` works correctly.
```
#colab
learner.lr_find()
```
Fine tune model
_Calling `learner.unfreeze` causes the model to overfit for the TPU so to make a fair comparison with the GPU version we are training using the frozen model only._
```
#colab
learner.freeze()
```
We are using the `lr` values same as the TPU (although these are not the ideal ones as shown by the LR Finder)
```
#colab
learner.fit_one_cycle(1,slice(7e-4),pct_start=0.99)
```
The results show the GPU is running faster but running 1 epoch shows lower accuracy and higher loss values.
```
#colab
learner.save('stage-1')
#colab
learner.load('stage-1')
#hide
#colab
list(learner.model.parameters())[0].device
```
The device used is a GPU ('cuda') not a TPU.
```
#colab
learner.freeze()
#colab
learner.fit_one_cycle(4,lr_max=slice(1e-6,1e-4))
```
It still reflects the general trend of the initial call to `fit_one_cycle` in that it runs 1 epoch per minute but at lower accuracy and higher losses.
Plot loss seems to be working fine.
```
#colab
learner.recorder.plot_loss()
```
## Status
The fastai XLA extensions library is still in very early development phase (not even alpha) which means there's still a lot of things not working.
Use it at your own risk.
If you wish to contribute to the project, fork it and make pull request.
This project uses [nbdev](https://nbdev.fast.ai/) -- a jupyter notebook first development environment and is being developed on [Colab](https://colab.research.google.com).
```
#hide
#colab
!nbdev_clean_nbs
!nbdev_build_docs
```
| github_jupyter |
# An overview of rigid body dynamics
$$
% vector
\newcommand{\v}[1]{\mathbf{\vec{#1}}}
% unit vector
\newcommand{\u}[1]{\mathbf{\hat{#1}}}
% dot product
\newcommand{\dp}[2]{#1 \cdot #2}
% cross product
\newcommand{\cp}[2]{#1 \times #2}
% rotation matrix
\newcommand{\R}[2]{{}^{#1} R ^{#2}}
% vector derivative
\newcommand{\d}[2]{\frac{{}^#2d#1}{dt}}
% angular velocity
\newcommand{\av}[2]{{}^{#2} \v{\omega} {}^{#1}}
% angular acceleration
\newcommand{\aa}[2]{{}^{#2} \v{\alpha} {}^{#1}}
% position
\newcommand{\pos}[2]{\v{r} {}^{#2/#1}}
% velocity
\newcommand{\vel}[2]{{}^#2 \v{v} {}^{#1}}
% acceleration
\newcommand{\acc}[2]{{}^#2 \v{a} {}^{#1}}
$$
Rigid body dynamics is concerned with describing the motion of systems composed of solid bodies; such as vehicles, skeletons, robots [1-4]:

This document borrows heavily from [5, 6].
# Newton's Second Law
For all these systems, our goal is to determine how each body's position changes in time. Newton told us that the acceleration $\v{a}$ of a system is proportional to the force $\v{F}$ applied to it:
$$\v{F} = m\v{a}$$
Newton gave us the bodies' acceleration, but we would really prefer to obtain their position. Thus, Newton gave us a second order ordinary differential equation for the quantity we seek, i.e. $\v{a}=\frac{d^2}{dt^2}\v{x}$. This equation is actually far too simple for the systems we want to study, but it reveals that there are three topics to consider in describing a rigid body system: its kinematics ($\v{a}$), its mass properties ($m$), and the external forces applied to the system ($\v{F}$). In this notebook, we present the tools necessary to mathematically describe a rigid body system. Once equipped with a mathematical description of a system, we can generate equations that describe its motion. Regardless, we always end up with second-order ordinary differential equations in time.
# Vectors
Newton's second law is a vector equation. A vector is a quantity that has a **magnitude** and a **direction**. For example, "5 miles East" is a vector quantity with magnitude 5 miles and direction East. We draw them as arrows:

We represent the magnitude of a vector $\v{v}$ as $|\v{v}|$. We represent the direction of a vector $\v{v}$ using a unit vector $\u{u}$ (magnitude of 1) that has the same direction as $\v{v}$:
$$\u{u} = \frac{\v{v}}{|\v{v}|}$$
We will work with the following vector quantities: positions, velocities, accelerations, forces, angular velocities, and torques/moments. Don't think about these vectors as linear algebra vectors. Our vectors always have a physical interpretation (and thus are always 2 or 3 dimensinonal), while linear algebra vectors are often more abstract.
## Addition
When we add vector $\v{b}$ to vector $\v{a}$, the result is a vector that starts at the tail of $\v{a}$ and ends at the tip of $\v{b}$:

```
from __future__ import print_function, division
from sympy import init_printing
init_printing(use_latex='mathjax', pretty_print=False)
```
Physics vectors in SymPy are created by first specifying a reference frame and using the associated unit vectors to construct vectors of arbritrary magnitude and direction. Reference frames will be discussed later on and for now it is only important that `N.x`, `N.y`, and `N.z` are three mutally orthoganl vectors of unit length.
```
from sympy.physics.vector import ReferenceFrame
N = ReferenceFrame('N')
```
Simple scalar variables can be import from SymPy with:
```
from sympy.abc import c, d, e, f, g, h
```
Finally, the unit vectors and scalars can be combined to create vectors.
```
a = c * N.x + d * N.y + e * N.z
a
a.to_matrix(N)
b = f * N.x + g * N.y + h * N.z
b
```
The magnitude of a vector can be found with::
```
a.magnitude()
a + b
```
## Scaling
Multiplying a vector by a scalar changes its magnitude, but not its direction. Scaling by a negative number changes a vector's magnitude and reverses its sense (rotates it by 180 degrees).

```
b = 2 * a
b
c = -a
c
```
### Exercise
Create three vectors that lie in the $XY$ plane where each vector is:
1. of length $l$ that is at an angle of $\frac{\pi}{4}$ degrees from the $X$ axis
2. of length $10$ and is in the $-Y$ direction
3. of length $l$ and is $\theta$ radians from the $Y$ axis
Finally, add vectors 1 and 2 and substract $5$ times the third vector.
*Hint: SymPy has variables and trigonometic functions, for example `from sympy import tan, pi`*
```
%load exercise_solutions/n01_vector_addition_scaling.py
```
## Dot product (scalar product)
The dot product, which yields a scalar quantity, is defined as:
$$\v{a} \cdot \v{b} = |\v{a}||\v{b}| \cos{\theta}$$
where $\theta$ is the angle between the two vectors. It is used to determine:
* the angle between two vectors: $\theta = \cos^{-1}[\v{a} \cdot \v{b} / (|\v{a}||\v{b}|)]$
* a vector's magnitude: $ |\v{v}| = \sqrt{\v{v} \cdot \v{v}} $
* the length of a vector along a direction/unit vector $\u{u}$ (called the projection): $ \mbox{proj}_{\u{u}}\v{v} = \v{v} \cdot \u{u}$
* if two vectors are perpendicular: $ \v{a} \cdot \v{b} = 0 \mbox{ if } \v{a} \perp \v{b} $
* compute power: $ P = \dp{\v{F}}{\v{v}}$
Also, dot products are used to convert a vector equation into a scalar equation (by "dotting" an entire equation with a vector).

```
from sympy.abc import c, d, e, f, g, h
from sympy.physics.vector import ReferenceFrame, dot
N = ReferenceFrame('N')
a = c * N.x + d * N.y + e * N.z
b = f * N.x + g * N.y + h * N.z
dot(a, b)
```
### Exercise
Given the vectors $\v{v}_1 = a \hat{\mathbf{n}}_x + b\hat{\mathbf{n}}_y + a \hat{\mathbf{n}}_z$ and $\v{v}_2=b \hat{\mathbf{n}}_x + a\hat{\mathbf{n}}_y + b \hat{\mathbf{n}}_z$ find the angle between the two vectors using the dot product.
```
%load exercise_solutions/n01_vector_dot_product.py
```
## Cross product (vector product)
The cross product, which yields a vector quantity, is defined as:
$$ \cp{\v{a}}{\v{b}} = |\v{a}||\v{b}| \sin\theta \u{u}$$
where $\theta$ is the angle between the two vectors, and $\u{u}$ is the unit vector perpendicular to both $\v{a}$ and $\v{b}$ whose sense is given by the right-hand rule. It is used to:
* obtain a vector/direction perpendicular to two other vectors
* determine if two vectors are parallel: $\cp{\v{a}}{\v{b}} = \v{0} \mbox{ if } \v{a} \parallel \v{b}$
* compute moments: $ \cp{\v{r}}{\v{F}}$
* compute the area of a triangle

```
from sympy.abc import c, d, e, f, g, h
from sympy.physics.vector import ReferenceFrame, cross
N = ReferenceFrame('N')
a = c * N.x + d * N.y + e * N.z
b = f * N.x + g * N.y + h * N.z
cross(a, b)
```
### Exercise
Given three points located in reference frame $N$ by:
$$
\v{p}_1 = 23 \u{n}_x - 12 \u{n}_y \\
\v{p}_2 = 16 \u{n}_x + 2 \u{n}_y - 4 \u{n}_z \\
\v{p}_3 = \u{n}_x + 14 \u{n}_z
$$
Find the area of the triangle bounded by these three points using the cross product.
*Hint: Search online for the relationship of the cross product to triangle area.*
```
%load exercise_solutions/n01_vector_cross_product.py
```
## Some vector properties
* The order in which you add them does not matter: $\v{a} + \v{b} = \v{b} + \v{a}$
* You can distrubute a scalar among vectors: $ s (\v{a} + \v{b}) = s\v{a} + s\v{b} $
**Dot product**
* You can pull out scalars: $ c \v{a} \cdot d \v{b} = cd (\v{a} \cdot \v{b})$
* Order does not matter: $\dp{\v{a}}{\v{b}} = \dp{\v{b}}{\v{a}}$
* You can distribute: $\dp{\v{a}}{(\v{b} + \v{c})} = \dp{\v{a}}{\v{b}} + \dp{\v{a}}{\v{c}}$
**Cross product**
* Crossing a vector with itself "cancels" it: $\cp{\v{a}}{\v{b}} = \vec{0}$
* You can pull out scalars: $ c \v{a} \times d \v{b} = cd (\v{a} \times \v{b})$
* Order DOES matter (because of the right-hand rule): $\cp{\v{a}}{\v{b}} = -\cp{\v{b}}{\v{a}}$
* You can distribute: $\cp{\v{a}}{(\v{b} + \v{c})} = \cp{\v{a}}{\v{b}} + \cp{\v{a}}{\v{c}}$
* They are NOT associative: $\cp{\v{a}}{(\cp{\v{b}}{\v{c}})} \neq \cp{(\cp{\v{a}}{\v{b}})}{\v{c}}$
# Reference frames
A reference frame (or simply, frame) is a rigid 3D object. We always attach a reference frame to rigid bodies in order to describe their motion. We may also use "empty" reference frames to make a system easier to model.
A reference frame has some *location* in space, but it does *not* have a position. Reference frames contain points, and those *points* have positions.
A reference frame also has an *orientation* in space. To specify its orientation, we choose a vector basis whose orientation is fixed with respect to the reference frame (but there are infinitely many vector bases we *could* label on the frame). In general, we are only interested in the vector bases we attach to reference frames; from here on, we will instead refer to reference frames in the places where we referred vector bases. That is, we express vectors in a reference frame instead of in a vector basis.
A reference frame's location and orientation vary in time. Two important attributes of a reference frame are its **angular velocity** $\v{\omega}$ and its **angular acceleration** $\v{\alpha}$; we'll describe these shortly.
A **Newtonian reference frame** is one in which Newton's second law holds.

## Expressing vectors with a vector basis
We have shown you what a vector $\v{v}$ looks like, but have yet to express an actual vector mathematically. To do so, we first choose three unit vectors $\u{a}_x$, $\u{a}_y$, and $\u{a}_z$ whose directions we accept as given. Consider the human jumper from above; we choose:
* $\u{a}_x$ to point forward,
* $\u{a}_y$ to point upwards,
* $\u{a}_z$ to point out of the plane (to the subject's right).

These three unit vectors are mutually perpendicular. For pratical reasons, we will always make sure that's the case. If so, the three vectors define a vector basis. We can express the position of the subject's hand from its toes in terms of these three vectors:
$$ \v{r} = d_x \u{a}_x + d_y \u{a}_y + 0 \u{a}_z$$
We call $d_x$ the **measure** of $\v{r}$ along $\u{a}_x$, and it is equal to $\v{r} \cdot \u{a}_x$. Note that a vector basis does not have an origin.
We could have chosen a different vector basis, such as $\u{b}_x$, $\u{b}_y$, $\u{b}_z$. Then, we would express $\v{r}$ as:
$$ \v{r} = f_x \u{b}_x + f_y \u{b}_y + 0 \u{b}_z$$
Using this alternative vector basis does not change the fact that $\v{r}$ is the position of the hand from the toes; it simply changes how we *express* this quantity. It is possible to express a single vector in infinitely many ways, since we can choose to use any valid vector basis. In the next section, we will learn how to relate different vector bases to each other.
#### Operating on vectors expressed in a basis
Once we express a vector in a vector basis, it is easy to perform operations on it with vectors expressed in the same basis. Consider the two vectors:
* $\v{a} = a_x \u{n}_x + a_y \u{n}_y + a_z \u{n}_z$
* $\v{b} = b_x \u{n}_x + b_y \u{n}_y + b_z \u{n}_z$
Here are the addition, dot, and cross operations between these two vectors:
$$
\v{a} + \v{b} = (a_x + b_x) \u{n}_x + (a_y + b_y) \u{n}_y + (a_z + b_z) \u{n}_z \\
\dp{\v{a}}{\v{b}} = a_x b_x + a_y b_y + a_z b_z\\
\cp{\v{a}}{\v{b}} = \det{
\begin{bmatrix} \u{n}_x & \u{n}_y & \u{n}_z \\
a_x & a_y & a_z \\
b_x & b_y & b_z
\end{bmatrix}}
$$
#### We must specify a vector basis
When a vector is expressed in typical linear algebra notation, information is lost. For example, we don't know the basis in which the following vector is expressed:
$$
\v{v} = \begin{bmatrix} v_x \\ v_y \\ v_z \end{bmatrix}
$$
If we don't know the basis in which $v_x$, $v_y$, and $v_z$ are its measures, we cannot add $\v{v}$ to another vector, etc. To express a vector in matrix form, we must carry along the basis in which it is expressed. One option for doing so is the following:
$$
[\v{v}]_{n} = \begin{bmatrix} v_x \\ v_y \\ v_z \end{bmatrix}_{n} $$
The notation $[\v{v}]_{n}$ specifies that $\v{v}$ is expressed in the vector basis $\u{n}_x$, $\u{n}_y$, $\u{n}_z$.
```
from sympy.abc import c, d, e, f, g, h, theta
from sympy.physics.vector import ReferenceFrame, dot, cross
A = ReferenceFrame('A')
B = A.orientnew('B', 'Axis', (theta, A.z))
a = c * A.x + d * A.y + e * A.z
b = f * B.x + g * B.y + h * B.z
a + b
dot(a, b)
cross(a, b)
(a+b).express(A)
```
# Rotation matrices (direction cosine matrices)
In almost every problem, we make use of multiple vector bases. The reason is that there is usually a particular basis in which a vector is most conveniently expressed. And, that convenient basis is usually not the same for all vectors we'll deal with. A side effect is that we will often want to change the basis in which a vector is expressed. To do so, we use a rotation matrix (also called a direction cosine matrix). The rotation matrix ${}^a R^b$ allows us to take a vector $\v{v}$ expressed in $\u{b}_x$, $\u{b}_y$, $\u{b}_z$ and re-express it in $\u{a}_x$, $\u{a}_y$, $\u{a}_z$:
$$
[\v{v}]_{a} = {}^a R^b ~ [\v{v}]_{b}
$$
The rotation matrix is given by dot products across the two the vector bases:
$$
\R{a}{b} =
\begin{bmatrix}
\dp{\u{a}_x}{\u{b}_x} & \dp{\u{a}_x}{\u{b}_y} & \dp{\u{a}_x}{\u{b}_z} \\
\dp{\u{a}_y}{\u{b}_x} & \dp{\u{a}_y}{\u{b}_y} & \dp{\u{a}_y}{\u{b}_z} \\
\dp{\u{a}_z}{\u{b}_x} & \dp{\u{a}_z}{\u{b}_y} & \dp{\u{a}_z}{\u{b}_z} \\
\end{bmatrix}
$$
Because of the nature of vector bases, this matrix is symmetric and orthogonal. If we instead have a vector in basis $a$ and want to express it in $b$, we can simply use the inverse of $\R{a}{b}$. Since the matrix is orthogonal, its inverse is the same as its transpose.
$$
\R{b}{a} = (\R{a}{b})^{-1} = (\R{a}{b})^T \\
[\v{v}]_{b} = {}^b R^a ~ [\v{v}]_{a} \\
[\v{v}]_{b} = ({}^a R^b)^T ~ [\v{v}]_{a}
$$
The columns of $\R{a}{b}$ are the unit vectors $\u{b}_x$, $\u{b}_y$, $\u{b}_z$ expressed in $a$:
$$
\R{a}{b} = \begin{bmatrix} [\u{b}_x]_a & [\u{b}_y]_a & [\u{b}_z]_a \end{bmatrix}
$$
#### Successive rotations
We'll usually need to re-express a vector multiple times. Luckily, we can do so by multiplying rotation matrices together:
$$
\R{d}{a} = (\R{d}{c} )(\R{c}{b}) (\R{b}{a}) \\
[\v{v}]_{d} = \R{d}{a} [\v{v}]_{a} \\
[\v{v}]_{d} = (\R{d}{c} )(\R{c}{b}) (\R{b}{a})[\v{v}]_{a}
$$
#### A point of confusion: rotating vs. re-expressing
Sometimes, rotation matrices are used to rotate vectors; that is, cause the vector to point somewhere different. That is NOT how we are using rotation matrices here. Rotating a vector changes the vector itself, while we are only changing how the *same* vector is expressed.
```
B.dcm(A)
```
### Exercise
Create two reference frames, the first should be attached to your laptop keyboard surface. For the first frame, the $Z$ axis should be directed from the Q key to the P key. The $Y$ unit vector should be directed from the shift key to the tab key. Now on the screen, attach a reference frame where the $Z$ axis is directed from the right side of the screen to the left and lies in the plane of the screen. The $Y$ axis should be directed from the top of the screen to the hinge.
The angle between the laptop and screen is $\theta$ such that $\theta=0$ corresponds to the laptop being closed and $0 < \theta < \pi$ is the laptop being open. With this create a vector that starts at the bottom left hand corner of the wrist rests and ends at the top right corner of the screen. Use $w$ for the width and $l$ for the length of the laptop.
Print the vector expressed in the keyboard frame.
*Hint: You may need to create more than two frames and a simple sketch will help.*
```
%load exercise_solutions/n01_vector_rotation.py
```
# Derivatives of vectors
Consider the vector $\u{a}_x$ in the figure above. To an observer sitting on $A$, $\u{a}_x$ never changes; it is fixed rigidly to $A$. Therefore, the observer would say the time derivative of $\u{a}_x$ is $\v{0}$. However, an observer on $N$ would indeed observe that $\u{a}_x$ changes in time. For this reason, when we take the time derivative of a vector, we must specify the frame in which we take the derivative. The derivative of a generic vector $\v{p}$ in frame $N$ is denoted as:
$$\d{\v{p}}{N}$$
Consider a vector $\v{p}$ expressed in $A$:
$$\v{p} = p_x \u{a}_x + p_y \u{a}_y + p_z \u{a}_z$$
Its time derivative in frame $A$ is:
$$\d{\v{p}}{A} = \dot{p}_x \u{a}_x + \dot{p}_y \u{a}_y + \dot{p}_z \u{a}_z$$
Here, we have benefitted from the fact that $\u{a}_x$, $\u{a}_y$, and $\u{a}_z$ are constant in $A$. We are not so fortunate when taking the derivative in $N$, since these basis vectors are not constant in $N$:
$$\d{\v{p}}{N} = \dot{p}_x \u{a}_x + p_x \d{\u{a}_x}{N} + \dot{p}_y \u{a}_y + p_y \d{\u{a}_y}{A} + \dot{p}_z \u{a}_z + p_z \d{\u{a}_z}{N}$$
This formula for the derivative in $N$ of a vector expressed in $A$ is not so great to use. Once we introduce angular velocity, we will have a much better way to compute such quantities.
```
a
a.diff(c, A)
```
# Angular velocity and angular acceleration
A reference frame's angular velocity describes the rate of change of the frame's orientation. Consider frame $A$. Since angular velocity is a vector quantity, we must specify the frame from which we observe the change in $A$'s orientation.
<div class="text-info" style="margin: 10px">
$\av{A}{N}$: the angular velocity of frame $A$ as observed from frame $N$
</div>
There are some complicated formulas for $\av{A}{N}$, but you usually don't need them. Typically, you know $\av{A}{N}$ by inspection. Take the linkage below:

In this linkage, the only way that frame/body $B$ can move with respect to $A$ is by rotating about $B_o$ by the angle $q_1$. Thus, by inspection:
$$\av{B}{A} = \dot{q}_1 \u{b}_z$$
$C$ is attached to $B$ similarly:
$$\av{C}{B} = \dot{q}_2 \u{c}_z$$
#### Angular velocity addition theorem
We can add angular velocities together, similar to how we multiplied reference frames:
$$\av{C}{A} = \av{B}{A} + \av{C}{B}$$
#### Derivative theorem
For any vector $\v{p}$, the following equation relates the derivative of $\v{p}$ in two different reference frames via the angular velocity between these two frames:
$$\d{\v{p}}{A} = \d{\v{p}}{B} + \av{B}{A} \times \v{p}$$
Again, this works for *any* vector, not just position vectors.
This theorem is really important, and is the primary way that we compute derivatives of vectors in other frames.
#### Angular acceleration
The equations of rigid body dynamics will also require angular accelerations $\aa{B}{A}$ of the rigid bodies in the system, but this can usually be computed automatically from $\av{B}{A}$.
<div class="text-info" style="margin: 10px">
$\aa{A}{N}$: the angular acceleration of frame $A$ as observed from frame $N$
</div>
```
B.ang_vel_in(A)
from sympy import Function
from sympy.abc import t
theta = Function('theta')(t)
theta
theta.diff(t)
B = A.orientnew('B', 'Axis', (theta, A.z))
B.ang_vel_in(A)
```
# Position, velocity, and acceleration
#### Position
Position vectors have the special property that two points must be specified. For example, if I want to obtain the position of point $P$ in the figure above, I must specify the point from which I want that position.
<div class="text-info" style="margin: 10px">
$\pos{Q}{P}$: the position of point $Q$ with respect to point $P$.
</div>
In modeling, we often must write down various position vectors via inspection.
#### Velocity
The velocity of a point is the derivative of its position, and must have associated with it the frame in which the derivative is taken.
<div class="text-info" style="margin: 10px">
$\vel{Q}{N}$: the velocity of point $Q$ in frame $N$
</div>
Previously, we used the symbol $\v{v}$ to denote a generic vector. Henceforth, $\v{v}$ refers to a velocity. If $N_o$ is a point fixed in $N$, then:
$$\vel{Q}{N}=\d{\pos{Q}{N_o}}{N}$$
When using PyDy, we rarely need to use inspection to determine the velocity of points of interest. Instead, we are usually in the situation that we want the velocity (in $N$) of point $Q$ fixed on body $B$, and we already know the velocity of another point $P$ fixed on $B$. In this case, we use the following formula to obtain $\vel{Q}{N}$ (`v2pt_theory` in PyDy):
$$\vel{Q}{N} = \vel{P}{N} + \av{B}{N} \times \pos{Q}{P}$$
#### Acceleration
An acceleration of a point is the derivative of its velocity, and must have associated with it the frame in which the derivative is taken.
<div class="text-info" style="margin: 10px">
$\acc{Q}{N}$: the acceleration of point $Q$ in frame $N$
</div>
Henceforth, $\v{a}$ refers to an acceleration. If the velocity of $Q$ is given, then the acceleration is:
$$\acc{Q}{N}=\d{\vel{Q}{N}}{N}$$
Similarly to velocity, we rarely need to use inspection to determine the acceleration of points of interest. Instead, we are usually in the situation that we want the acceleration (in $N$) of point $Q$ fixed on body $B$, and we already know the velocity of another point $P$ fixed on $B$. In this case, we use the following formula to obtain $\acc{Q}{N}$ (`a2pt_theory` in PyDy):
$$\acc{Q}{N} = \acc{P}{N} + \aa{B}{N} \times \pos{Q}{P} + \av{B}{N} \times (\av{B}{N} \times \pos{Q}{P})$$
# Inertial properties
Each particle or rigid body has interial properties. We will assume that these properties are constant with respect to time. Each particle in a system has a scalar mass and each rigid body has a scalar mass located at it's center of mass and an inertia dyadic (or tensor) that represents how that mass is distributed in space, which is typically defined with respect to the center of mass.
Just as we do with vectors above, we will use a basis dependent expression of tensors. The inertia of a 3D rigid body is typically expressed as a tensor (symmetric 3 x 3 matrix).
$$
I = \begin{bmatrix}
I_{xx} & I_{xy} & I_{xz} \\
I_{xy} & I_{yy} & I_{yz} \\
I_{xz} & I_{yz} & I_{zz}
\end{bmatrix}_N
$$
The three terms on the diagnol are the moments of inertia and represent the resistance to angular acceleration about the respective axis in the subscript. The off diagonal terms are the products of inertia and represent the coupled resistance to angular acceleration from one axis to another. The $N$ subscript denotes that this tensor is expressed in the $N$ reference frame.
We can write this tensor as a dyadic to allow for easy combinations of inertia tensors expressed in different frames, just like we combine vectors expressed in different frames above. This basis dependent tensor takes the form:
$$I =
I_{xx} \begin{bmatrix}
1 & 0 & 0 \\
0 & 0 & 0 \\
0 & 0 & 0
\end{bmatrix}_N +
I_{xy} \begin{bmatrix}
0 & 1 & 0 \\
0 & 0 & 0 \\
0 & 0 & 0
\end{bmatrix}_N +
I_{xz} \begin{bmatrix}
0 & 0 & 1 \\
0 & 0 & 0 \\
0 & 0 & 0
\end{bmatrix}_N +
I_{yx} \begin{bmatrix}
0 & 0 & 0 \\
1 & 0 & 0 \\
0 & 0 & 0
\end{bmatrix}_N +
I_{yy} \begin{bmatrix}
0 & 0 & 0 \\
0 & 1 & 0 \\
0 & 0 & 0
\end{bmatrix}_N +
I_{yz} \begin{bmatrix}
0 & 0 & 0 \\
0 & 0 & 1 \\
0 & 0 & 0
\end{bmatrix}_N + \\
I_{zx} \begin{bmatrix}
0 & 0 & 0 \\
0 & 0 & 0 \\
1 & 0 & 0
\end{bmatrix}_N +
I_{zy} \begin{bmatrix}
0 & 0 & 0 \\
0 & 0 & 0 \\
0 & 1 & 0
\end{bmatrix}_N +
I_{zz} \begin{bmatrix}
0 & 0 & 0 \\
0 & 0 & 0 \\
0 & 0 & 1
\end{bmatrix}_N
$$
These "unit" tensors are simply the outer product of the associated unit vectors and can be written as such:
$$ I = I_{xx} \u{n}_x \otimes \u{n}_x +
I_{xy} \u{n}_x \otimes \u{n}_y +
I_{xz} \u{n}_x \otimes \u{n}_z +
I_{yx} \u{n}_y \otimes \u{n}_x +
I_{yy} \u{n}_y \otimes \u{n}_y +
I_{yz} \u{n}_y \otimes \u{n}_z +
I_{zx} \u{n}_z \otimes \u{n}_x +
I_{zy} \u{n}_z \otimes \u{n}_y +
I_{zz} \u{n}_z \otimes \u{n}_z $$
Inertia dyadics and tensors can be created in the following way:
```
from sympy import symbols
from sympy.physics.mechanics import ReferenceFrame, inertia
ixx, iyy, izz, ixy, iyz, ixz = symbols('I_xx I_yy I_zz I_xy I_yz I_xz')
N = ReferenceFrame('N')
I = inertia(N, ixx, iyy, izz, ixy, iyz, ixz)
I
I.to_matrix(N)
```
# Forces and moments/torques
Forces are vectors which are applied to specific points (bound vectors) and moments/torques are vectors than describe rotational load applied to a body. Both can simply be described as vectors but either a point or reference frame must be associated with each, respectively.
**Equal and Opposite**
Don't forget Newton's third law of motion. If there is a force or torque, there is always an equal and opposit force or torque acting on the opposing point or reference frame.
```
from sympy.abc import a, b, c
from sympy.physics.vector import ReferenceFrame, Point
A = ReferenceFrame('A')
P = Point('P')
f = a * A.x + b * A.y + b * A.z
# We will typically denote a force as tuple of a vector and a point.
force = (f, P)
f
```
# Equations of motion
Once all of the important forces acting on a system, the accelerations of all particles and bodies, and the inertial properties of the system are found, the equations of motion can be formed. For planar dyanmics the equations take on this form:
$$\sum \v{F} = m \v{a} \\ \sum \v{T} = I \v{\alpha}$$
The force equation (Newton's second law) and the torque equation (Euler's equation) make up a set of second ordinary differential equations in time. We typically want these equations in first order form:
$$\dot{x} = f(x, u, t)$$
And to do that kinematical differential equations are introduced, which simply define the relationships between the positional and angular states and their derivatives. For example, we could introduce $\omega$ as the time derivative of an angle $\theta$:
$$\omega = \dot{\theta}$$
The states $x$ are typically positions, angles, and their rates.
In general, the equations of motion are non-linear ordinary differential equations and anayltical solutions do not exist. To find the resulting state trajectories we turn to numerical integration methods.
$$x = \int_{t_0}^{t_f} f(x, u, t) dt$$
# References
[1] Moore, J. K. (2012). Human Control of a Bicycle. University of California, Davis.
[2] Ashby, B. M., & Delp, S. L. (2006). Optimal control simulations reveal mechanisms by which arm movement improves standing long jump performance. Journal of biomechanics, 39(9), 1726–34. doi:10.1016/j.jbiomech.2005.04.017
[3] Gong, X., Bai, Y., Hou, Z., Zhao, C., Tian, Y., & Sun, Q. (2012). Backstepping sliding mode tracking control of quad-rotor under input saturation. International Journal of Intelligent Computing and Cybernetics, 5(4), 515–532. doi:10.1108/17563781211282268
[4] Wu, J. Z., An, K.-N., Cutlip, R. G., Krajnak, K., Welcome, D., & Dong, R. G. (2008). Analysis of musculoskeletal loading in an index finger during tapping. Journal of biomechanics, 41(3), 668–76. doi:10.1016/j.jbiomech.2007.09.025
[5] Mitiguy, P. Advanced Dynamics & Motion Simulation. 2013.
[6] http://docs.sympy.org/latest/modules/physics/mechanics/index.html
| github_jupyter |
<img src="https://microsoft.github.io/Accera/assets/logos/Accera_darktext.png" alt="Accera logo" width="600"/>
# Accera Quickstart Example
In this example, we will:
* Implement matrix multiplication with a ReLU activation (matmul + ReLU), commonly used in in machine learning algorithms
* Generate two implementations: a naive algorithm and one with loop transformations
* Compare the timings of both implementations
### Setup
First, we'll install Accera using `pip`.
#### Optional: if running this notebook locally
* Linux/macOS: install gcc using `apt install gcc`.
* Windows: install Microsoft Visual Studio and run `vcvars64.bat` to setup Visual Studio tools in your `PATH` before starting the Jupyter environment.
```
!pip install accera
```
### Build
Run the code below to implement `ReLU(C + A @ B)` on arrays `A`, `B`, and `C`.
We'll build a package called `"hello_accera"` that will export both versions as C functions.
```
import accera as acc
# define placeholder inputs/output
A = acc.Array(role=acc.Array.Role.INPUT, shape=(512, 512))
B = acc.Array(role=acc.Array.Role.INPUT, shape=(512, 512))
C = acc.Array(role=acc.Array.Role.INPUT_OUTPUT, shape=(512, 512))
# implement the logic for matmul and relu
matmul = acc.Nest(shape=(512, 512, 512))
i1, j1, k1 = matmul.get_indices()
@matmul.iteration_logic
def _():
C[i1, j1] += A[i1, k1] * B[k1, j1]
relu = acc.Nest(shape=(512, 512))
i2, j2 = relu.get_indices()
@relu.iteration_logic
def _():
C[i2, j2] = acc.max(C[i2, j2], 0.0)
package = acc.Package()
# fuse the i and j indices of matmul and relu, add to the package
schedule = acc.fuse(matmul.create_schedule(), relu.create_schedule(), partial=2)
package.add(schedule, args=(A, B, C), base_name="matmul_relu_fusion_naive")
# transform the schedule, add to the package
# here we will focus only on the j index. For a more complete example, see:
# https://microsoft.github.io/Accera/Tutorials/Optimized_MatMul/
tile_size_j = 256
target = acc.Target(category=acc.Target.Category.CPU)
f, i, j, k = schedule.get_indices()
jj = schedule.split(j, tile_size_j)
jjj = schedule.split(jj, (target.vector_bytes // 4) * 2) # there are 2 vfma execution units, each holding (target.vector_bytes // 4) 32-bit float elements
jjjj = schedule.split(jjj, target.vector_bytes // 4) # each SIMD register holds (target.vector_bytes // 4) 32-bit float elements
schedule.reorder(j, f, k, i, jj, jjj, jjjj) # reorder the loops
plan = schedule.create_plan(target)
plan.kernelize(unroll_indices=(jjj,), vectorize_indices=jjjj) # unroll and vectorize
package.add(plan, args=(A, B, C), base_name="matmul_relu_fusion_transformed")
# build a dynamically-linked package (a .dll or .so) that exports both functions
print(package.build(name="hello_accera", format=acc.Package.Format.HAT_DYNAMIC))
```
### Benchmark
In the previous section, we built a binary (`.so`) and a header file (`.hat`).
Next, we will load the package and compare the timings of both implementations.
```
import hatlib as hat
import numpy as np
# load the package
_, functions = hat.load("hello_accera.hat")
# call one of the functions with test inputs
A_test = np.random.rand(512, 512).astype(np.float32)
B_test = np.random.rand(512, 512).astype(np.float32)
C_test = np.zeros((512, 512)).astype(np.float32)
C_numpy = np.maximum(C_test + A_test @ B_test, 0.0)
matmul_relu = functions["matmul_relu_fusion_transformed"]
matmul_relu(A_test, B_test, C_test)
# check correctness
np.testing.assert_allclose(C_test, C_numpy, atol=1e-3)
# benchmark all functions
hat.run_benchmark("hello_accera.hat", batch_size=5, min_time_in_sec=5)
```
### Next Steps
The [Manual](https://microsoft.github.io/Accera/Manual/00%20Introduction/) is a good place to start for an introduction to the Accera Python programming model.
In particular, the [schedule transformations](https://microsoft.github.io/Accera/Manual/03%20Schedules/#schedule-transformations) describe how you can experiment with different loop transformations with just a few lines of Python.
Finally, the `.hat` format is just a C header file containing metadata. Learn more about the [HAT format](https://github.com/microsoft/hat) and [benchmarking](https://github.com/microsoft/hat/tree/main/tools).
## How it works
In a nutshell, Accera takes the Python code that defines the loop schedule and algorithm and converts it into [MLIR](https://mlir.llvm.org/) intermediate representation (IR). Accera's compiler then takes this IR through a series of MLIR pipelines to perform transformations. The result is a binary library with a C header file. The library implements the algorithms that are defined in Python, and is compatible with the target.
To peek into the stages of IR transformation that Accera does, try replacing `format=acc.Package.Format.HAT_DYNAMIC` with `format=acc.Package.Format.MLIR_DYNAMIC` above, re-run the build, and search the `_tmp` subfolder for the intermediate `*.mlir` files. We plan to document these IR constructs in the future.
## Documentation
Get to know Accera by reading the [Documentation](https://microsoft.github.io/Accera/).
You can find more step-by-step examples in the [Tutorials](https://microsoft.github.io/Accera/Tutorials).
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams['font.sans-serif']=['SimHei']
plt.rcParams['axes.unicode_minus']=False
# CITY = 'beijing'
CITY = 'shanghai'
host_df = pd.read_csv('data/{}/host.csv'.format(CITY))
room_df = pd.read_csv('data/{}/room.csv'.format(CITY))
best_room_df = pd.read_csv('data/{}/best_room.csv'.format(CITY))
def plot_translate(raw_room_df: pd.DataFrame):
room_df = raw_room_df.copy(deep=True)
trans_dict = {
'Entire home/apt': '套房',
'Private room':'单间',
'Shared room':'共享'
}
room_df['instant_bookable']=room_df['instant_bookable'].map(lambda x: '不允许' if x=='f' else '允许')
room_df['host_identity_verified'] = room_df['host_identity_verified'].map(lambda x: '未认证' if x=='f' else '已认证')
room_df['room_type'] = room_df['room_type'].map(lambda x: trans_dict[x])
room_df['is_single'] = room_df['is_single'].map(lambda x: '企业房东' if x=='f' else '个人房东')
return room_df
def get_trans_df(room_df: pd.DataFrame, best_room_df:pd.DataFrame,attr:str):
# transform_rate = (valid_count / best_count) / (total_count / room_count)
valid_count = best_room_df[attr].value_counts() # count of attr in best rooms
best_count = len(best_room_df) # count of best rooms
total_count = room_df[attr].value_counts() # count of attr in all rooms
room_count = len(room_df) # count of all rooms
trans_df = valid_count * room_count / (best_count * total_count)
trans_df = trans_df.sort_values(ascending=False).dropna(axis=0,how='any')
return trans_df
def get_plot_elem(trans_df:pd.DataFrame, attr:str):
x= ['{}'.format(num) for num in trans_df.index]
y1 = [] # count of est rooms
for name in trans_df.index:
y1.append(len(best_room_df[best_room_df[attr]==name]))
y2 = trans_df.values.tolist()
return x, y1, y2
def plot_transfer(x, y1, y2, var_name_list: list, xlabel: str, ylabel_list:list, ylim1:list=[0,110], ylim2:list=[0, 3.5],x_rotation:int=45):
fig = plt.figure()
ax1 = fig.add_subplot(111)
ax1.bar(x, y1,label=var_name_list[0],color='c')
ax1.set_ylim(ylim1)
ax1.set_ylabel(ylabel_list[0])
plt.legend(loc=2)
plt.xlabel(xlabel)
plt.xticks(rotation=x_rotation)
ax2 = ax1.twinx()
ax2.plot(x, y2, '-p', label=var_name_list[1])
ax2.set_ylim(ylim2)
ax2.set_ylabel(ylabel_list[1])
y3 = [1] * len(x)
ax2.plot(x, y3,'--')
plt.legend(loc=1)
def analyse_trans(room_df: pd.DataFrame, best_room_df: pd.DataFrame, attr: str, ylim1:list=[0,110], ylim2:list=[0, 3.5], rotation:int=45):
attr_dict = {
'neighbourhood': '行政区划',
'accommodates': '房间容量',
'room_type': '房间类型',
'bedrooms': '卧室数量',
'beds': '床数量',
'instant_bookable': '即时预订',
'is_single': '房东类型',
'host_identity_verified': '房东认证情况',
'host_response_rate': '房东评论回复率'
}
trans_df = get_trans_df(room_df, best_room_df, attr)
x, y1, y2 = get_plot_elem(trans_df, attr)
plot_transfer(x, y1, y2, ['房源数','转化率'],attr_dict.get(attr,attr),['优质房源数(个)','优质房源转化系数'],ylim1, ylim2,x_rotation=rotation)
best_room_df
room_df = pd.merge(room_df, host_df, on='host_id')
room_df = plot_translate(room_df)
best_room_df = pd.merge(best_room_df, host_df, on='host_id')
best_room_df = plot_translate(best_room_df)
analyse_trans(room_df, best_room_df, 'beds',ylim1=[0,120],ylim2=[0,7], rotation=0)
analyse_trans(room_df,best_room_df, 'instant_bookable',ylim2=[0,1.5], rotation=0)
analyse_trans(room_df, best_room_df, 'neighbourhood',ylim2=[0,2])
analyse_trans(room_df,best_room_df,'accommodates',ylim2=[0,3.5],rotation=0)
analyse_trans(room_df,best_room_df,'room_type', ylim2=[0,2],rotation=0)
analyse_trans(room_df, best_room_df, 'bedrooms',ylim1=[0,150],ylim2=[0,2], rotation=0)
analyse_trans(room_df, best_room_df, 'is_single',ylim1=[0,170],ylim2=[0,2],rotation=0)
analyse_trans(room_df, best_room_df, 'host_response_rate',ylim1=[0,150],ylim2=[0,3.5],rotation=0)
analyse_trans(room_df, best_room_df,'host_identity_verified',ylim1=[0, 200],ylim2=[0,5],rotation=0)
```
| github_jupyter |
# ATLAS $H \rightarrow ZZ \rightarrow \ell \ell \ell \ell$ Public Outreach Example
ATLAS has released its higgs discovery dataset as public data:
* Use `ServiceX` to stream the 4 lepton data
* Use `coffea` and `awkward` to produce the final $m_{4\ell}$ plots.
* This is **only** aboput 0.5 GB worth of data!
Outline
1. Use `ServiceX` for general quality and object selection
1. Use `coffea` and `awkward` to do multi-object event wide selection and plots
1. Produce the plot for running on a single MC file
1. Run on all the MC and Data files
```
from func_adl_servicex import ServiceXSourceUpROOT
from servicex.servicex import ServiceXDataset
from coffea.processor.servicex import Analysis, DataSource, LocalExecutor
from func_adl import ObjectStream
from hist import Hist
import asyncio
from typing import List
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (20, 10)
```
## Selecting events and clean objects
The ATLAS analysis ...
First we create the representative data source, and apply the initial trigger requirement:
```
ds = ServiceXSourceUpROOT('cernopendata://dummy', "mimi", backend='open_uproot')
ds.return_qastle = True # Magic
good_events = (ds
.Where(lambda e: e.trigE or e.trigM)
)
```
Next, basic lepton selection:
* Turn the columnar representation into object so we can make cuts
* Apply the common base cuts for electrons and muons
* We also need the event weights, which are baked into the ntuple
```
all_leptons = (good_events
.Select(lambda e: (
Zip({
'lep_pt': e.lep_pt,
'lep_eta': e.lep_eta,
'lep_phi': e.lep_phi,
'lep_energy': e.lep_E,
'lep_charge': e.lep_charge,
'lep_ptcone30': e.lep_ptcone30,
'lep_etcone20': e.lep_etcone20,
'lep_typeid': e.lep_type,
'lep_trackd0pvunbiased': e.lep_trackd0pvunbiased,
'lep_tracksigd0pvunbiased': e.lep_tracksigd0pvunbiased,
'lep_z0': e.lep_z0,
}),
e.mcWeight,
e.scaleFactor_ELE*e.scaleFactor_MUON*e.scaleFactor_LepTRIGGER*e.scaleFactor_PILEUP,
))
)
good_leptons = (all_leptons
.Select(lambda e: {
'ele': e[0]
.Where(lambda lep: lep.lep_typeid == 11)
.Where(lambda lep: (lep.lep_pt > 7000)
and (abs(lep.lep_eta) < 2.47)
and (lep.lep_etcone20/lep.lep_pt < 0.3)
and (lep.lep_ptcone30/lep.lep_pt < 0.3)
and (abs(lep.lep_trackd0pvunbiased) / lep.lep_tracksigd0pvunbiased < 5)
),
'mu': e[0]
.Where(lambda lep: lep.lep_typeid == 13)
.Where(lambda lep: (lep.lep_pt > 5000)
and (abs(lep.lep_eta) < 2.5)
and (lep.lep_etcone20/lep.lep_pt < 0.3)
and (lep.lep_ptcone30/lep.lep_pt < 0.3)
and (abs(lep.lep_trackd0pvunbiased) / lep.lep_tracksigd0pvunbiased < 3)
),
'mcWeight': e[1],
'scaleFactor': e[2],
})
)
```
And finally, we have to turn this into a form that coffea can currently understand.
* We only need to feed columns we will use downstream out of `ServiceX`, reducing the dataload.
```
atlas_selection = (good_leptons
.Select(lambda e: {
'electrons_pt': e.ele.lep_pt,
'electrons_eta': e.ele.lep_eta,
'electrons_phi': e.ele.lep_phi,
'electrons_energy': e.ele.lep_energy,
'electrons_charge': e.ele.lep_charge,
'electrons_z0': e.ele.lep_z0,
'muons_pt': e.mu.lep_pt,
'muons_eta': e.mu.lep_eta,
'muons_phi': e.mu.lep_phi,
'muons_energy': e.mu.lep_energy,
'muons_charge': e.mu.lep_charge,
'muons_z0': e.mu.lep_z0,
'mcWeight': e.mcWeight,
'scaleFactor': e.scaleFactor,
})
.AsParquetFiles('junk.parquet')
)
```
## Performing the analysis
The data from `ServiceX` is now analyzed by `awkward` and `coffea`.
* `coffea` automatically builds _event_ layout for electorns and muons, queuing off the prefix in the names from `ServiceX`
* `coffea` recognizes that $p_T$, $\eta$, $\phi$, and $E$ are availible, and builds a 4-vector that has a `theta` property.
* `awkward` properly translates the `np.sin` to work on an `awkward` array.
```
class ATLAS_Higgs_4L(Analysis):
'''Run the 4 Lepton analysis on ATLAS educational ntuples
'''
@staticmethod
def process(events):
from collections import defaultdict
import numpy as np
import awkward as ak
sumw = defaultdict(float)
mass_hist = (Hist.new
.Reg(60, 60, 180, name='mass', label='$m_{4\ell}$ [GeV]')
.StrCat([], name='dataset', label='Cut Type', growth=True)
.StrCat([], name='channel', label='Channel', growth=True)
.Int64()
)
dataset = events.metadata['dataset']
electrons = events.electrons
muons = events.muons
weight = ak.Array(np.ones(len(events.scaleFactor))) if events.metadata['is_data'] \
else events.scaleFactor*events.mcWeight
# We didn't have the 4-vector in `ServiceX`, so we couldn't do the final good-object cut.
# Good electon selection
electrons_mask = (abs(electrons.z0*np.sin(electrons.theta)) < 0.5)
electrons_good = electrons[electrons_mask]
# Good muon selection
muons_mask = (abs(muons.z0*np.sin(muons.theta)) < 0.5)
muons_good = muons[muons_mask]
# Next are event level cuts
# In order to cut in sorted lepton pt, we have to rebuild a lepton array here
leptons_good = ak.concatenate((electrons_good, muons_good), axis=1)
leptons_good_index = ak.argsort(leptons_good.pt, ascending=False)
leptons_good_sorted = leptons_good[leptons_good_index]
# Event level cuts now that we know the good leptons
# - We need to look at 4 good lepton events only
# - We need same flavor, so check for even numbers of each flavor
# - all charges must be balenced
event_mask = (
(ak.num(leptons_good_sorted) == 4)
& ((ak.num(electrons_good) == 0) | (ak.num(electrons_good) == 2) | (ak.num(electrons_good) == 4))
& ((ak.num(muons_good) == 0) | (ak.num(muons_good) == 2) | (ak.num(muons_good) == 4))
& (ak.sum(electrons_good.charge, axis=1) == 0)
& (ak.sum(muons_good.charge, axis=1) == 0)
)
# Next, we need to cut on the pT for the leading, sub-leading, and sub-sub-leading lepton
leptons_good_preselection = leptons_good[event_mask]
event_good_lepton_mask = (
(leptons_good_preselection[:,0].pt > 25000.0)
& (leptons_good_preselection[:,1].pt > 15000.0)
& (leptons_good_preselection[:,2].pt > 10000.0)
)
# Now, we need to rebuild the good muon and electron lists with those selections
muons_analysis = muons_good[event_mask][event_good_lepton_mask]
electrons_analysis = electrons_good[event_mask][event_good_lepton_mask]
# Lets do eemumu events - as there are no permutations there.abs
# At this point if there are two muons, there must be two electrons
eemumu_mask = (ak.num(muons_analysis) == 2)
muon_eemumu = muons_analysis[eemumu_mask]
electrons_eemumu = electrons_analysis[eemumu_mask]
z1_eemumu = muon_eemumu[:,0] + muon_eemumu[:,1]
z2_eemumu = electrons_eemumu[:,0] + electrons_eemumu[:,1]
h_eemumu = z1_eemumu + z2_eemumu
sumw[dataset] += len(h_eemumu)
mass_hist.fill(
channel=r'$ee\mu\mu$',
mass=h_eemumu.mass/1000.0,
dataset=dataset,
# weight=weight[eemumu_mask]
)
# Next, eeee. For this we have to build permutations and select the best one
def four_leptons_one_flavor(same_flavor_leptons, event_weights, channel: str):
fl_positive = same_flavor_leptons[same_flavor_leptons.charge > 0]
fl_negative = same_flavor_leptons[same_flavor_leptons.charge < 0]
fl_pairs = ak.cartesian((fl_positive, fl_negative))
zs = fl_pairs["0"] + fl_pairs["1"]
delta = abs((91.18*1000.0) - zs.mass[:])
closest_masses = np.min(delta, axis=-1)
the_closest = (delta == closest_masses)
the_furthest = the_closest[:,::-1]
h_eeee = zs[the_closest] + zs[the_furthest]
sumw[dataset] += len(h_eeee)
mass_hist.fill(
channel=channel,
mass=ak.flatten(h_eeee.mass/1000.0),
dataset=dataset,
# weight=event_weights,
)
four_leptons_one_flavor(electrons_analysis[(ak.num(electrons_analysis) == 4)],
weight[(ak.num(electrons_analysis) == 4)],
'$eeee$')
four_leptons_one_flavor(muons_analysis[(ak.num(muons_analysis) == 4)],
weight[(ak.num(muons_analysis) == 4)],
'$\\mu\\mu\\mu\\mu$')
return {
"sumw": sumw,
"mass": mass_hist,
}
```
## Run on a MC file
Define a convience function to load MC files - there are a lot of them.
* Note the `is_data` metadata - which was used above in an if statement
```
def make_ds(name: str, query: ObjectStream):
'''Create a ServiceX Datasource for a particular ATLAS Open data file
'''
from utils import files
is_data = name == 'data'
datasets = [ServiceXDataset(files[name]['files'], backend_type='open_uproot', image='sslhep/servicex_func_adl_uproot_transformer:pr_fix_awk_bug')]
return DataSource(query=query, metadata={'dataset': name, 'is_data': is_data}, datasets=datasets)
from utils import files
all_datasets = list(files.keys())
', '.join(all_datasets)
```
And an easy routine that will run a single file
* Async so we can run multiple queires at once
* Contains a lot of boiler plate
* Runs on multiple datasets at once (which we will need)
```
async def run_analysis(names: List[str]):
'Generate base plot for a multiple datafiles'
executor = LocalExecutor(datatype='parquet')
datasources = [make_ds(ds_name, atlas_selection) for ds_name in names]
# Create the analysis and we can run from there.
analysis = ATLAS_Higgs_4L()
async def run_updates_stream(accumulator_stream, name):
'''Run to get the last item in the stream, with a useful error message'''
coffea_info = None
try:
async for coffea_info in accumulator_stream:
pass
except Exception as e:
raise Exception(f'Failure while processing {name}') from e
return coffea_info
# Run on all items and wait till they are done!
all_plots = await asyncio.gather(*[run_updates_stream(executor.execute(analysis, source), source.metadata['dataset']) for source in datasources])
all_plots_mass = [p['mass'] for p in all_plots]
mass = all_plots_mass[0]
for p in all_plots_mass[1:]:
mass += p
return mass
mc_mass_plot = await run_analysis(['ggH125_ZZ4lep'])
artists = mc_mass_plot.project('mass','channel').plot(stack=True)
ax = artists[0].stairs.axes # get the axis
ax.legend(loc="best");
mc_mass_plot.project('mass').plot()
```
## Running on all MC and Data Files
Here we will repeat the above, but unleash it on all our datasets. These will all be put into a single histogram, with the `dataset` bin marking what sample they are from.
```
mass_plot = await run_analysis(all_datasets)
```
Lets plot everything - not that this is interesting from a physics point of view, but it does make sure everything was added to the histogram in the end!
```
artists = mass_plot.project('mass', 'dataset').plot(stack=True)
ax = artists[0].stairs.axes # get the axis
ax.legend(loc="best");
```
First, lets look at the components. We'd like signal, data, and the sum of everything else (MC prediction).
Data:
```
mass_plot[:, 'data', :].project('mass').plot();
```
Signal:
```
signal_ds = list(d for d in all_datasets if 'H125' in d)
artists = mass_plot[:,signal_ds,:].project('mass', 'dataset').plot(stack=True)
ax = artists[0].stairs.axes # get the axis
ax.legend(loc="best");
```
Sum of the backgrounds:
```
mc_ds = list(str(i) for i in (set(list(mass_plot.axes[1])) - set(signal_ds) - set(['data'])))
mc_ds.sort()
artists = mass_plot[:,mc_ds,:].project('mass', 'dataset').plot(stack=True)
ax = artists[0].stairs.axes # get the axis
ax.legend(loc="best");
```
| github_jupyter |
```
#!/usr/bin/env python3
############## ROS Import ###############
import rospy
import std_msgs
from sensor_msgs.msg import Image
import ros_numpy
import numpy as np
import random
import time
import itertools
import os
import cv2
from cv_bridge import CvBridge, CvBridgeError
%matplotlib notebook
from matplotlib import pyplot as plt
from IPython.display import display, HTML, clear_output
from IPython.display import Image as IPython_Image
import tensorflow as tf
path_raw_video = '/home/kimbring2/Desktop/raw_video.avi'
path_seg_video = '/home/kimbring2/Desktop/seg_video.avi'
path_gan_video = '/home/kimbring2/Desktop/gan_video.avi'
fps = 5
size = (512,512)
raw_video_out = cv2.VideoWriter(path_raw_video, cv2.VideoWriter_fourcc(*'DIVX'), fps, (640,360))
seg_video_out = cv2.VideoWriter(path_seg_video, cv2.VideoWriter_fourcc(*'DIVX'), fps, (640,360))
gan_video_out = cv2.VideoWriter(path_gan_video, cv2.VideoWriter_fourcc(*'DIVX'), fps, (256,256))
imported_rl = tf.saved_model.load("/home/kimbring2/Desktop/rl_model")
imported_seg = tf.saved_model.load("/home/kimbring2/Desktop/seg_model")
imported_gan = tf.saved_model.load("/home/kimbring2/Desktop/gan_model")
f_rl = imported_rl.signatures["serving_default"]
f_seg = imported_seg.signatures["serving_default"]
f_gan = imported_gan.signatures["serving_default"]
rl_test_input = np.zeros([1,128,128,5])
seg_test_input = np.zeros([1,256,256,3])
gan_test_input = np.zeros([1,256,256,3])
rl_test_tensor = tf.convert_to_tensor(rl_test_input, dtype=tf.float32)
seg_test_tensor = tf.convert_to_tensor(seg_test_input, dtype=tf.float32)
gan_test_tensor = tf.convert_to_tensor(gan_test_input, dtype=tf.float32)
memory_state = tf.zeros([1,128], dtype=np.float32)
carry_state = tf.zeros([1,128], dtype=np.float32)
f_rl = imported_rl.signatures["serving_default"]
rl_test_input = np.zeros([1,128,128,5])
print(f_rl(input_1=rl_test_tensor, input_2=memory_state, input_3=carry_state)['core_lstm'].numpy()[0])
print(f_rl(input_1=rl_test_tensor, input_2=memory_state, input_3=carry_state)['core_lstm_1'].numpy()[0])
print(f_rl(input_1=rl_test_tensor, input_2=memory_state, input_3=carry_state)['dense_1'].numpy()[0])
time.sleep(1)
print(f_seg(seg_test_tensor)['conv2d_transpose_4'].numpy()[0])
time.sleep(1)
print(f_gan(gan_test_tensor)['conv2d_transpose_7'].numpy()[0])
bridge = CvBridge()
raw_video_frame = np.zeros([640,360,3])
seg_video_frame = np.zeros([640,360,3])
gan_video_frame = np.zeros([256,256,3])
camera_frame = np.zeros([128,128,3])
step = 0
def arrayShow(imageArray):
image = bytes(cv2.imencode('.jpg', imageArray)[1])
return IPython_Image(data=image)
def image_callback(msg):
global camera_frame
global raw_video_frame
global seg_video_frame
global gan_video_frame
global step
cv_image = bridge.imgmsg_to_cv2(msg, "passthrough")
cv_image = cv2.resize(cv_image, (640, 360), interpolation=cv2.INTER_AREA)
frame = cv_image
raw_video_frame = cv_image
#raw_video_out.write(cv_image)
cv_image_shape = cv_image.shape
width = cv_image_shape[1]
height = cv_image_shape[0]
cv_image = cv2.resize(cv_image, (256, 256), interpolation=cv2.INTER_AREA)
cv_image = cv2.normalize(cv_image, None, 0, 1, cv2.NORM_MINMAX, cv2.CV_32F)
#cv_image = cv2.cvtColor(cv_image, cv2.COLOR_RGB2RGBA).astype(np.float32)
resized = np.array([cv_image])
input_tensor = tf.convert_to_tensor(resized, dtype=tf.float32)
#output = f_cyclegan(input_tensor)['conv2d_transpose_7'].numpy()[0]
pred_mask = f_seg(input_tensor)['conv2d_transpose_4']
pred_mask = tf.argmax(pred_mask, axis=-1)
pred_mask = pred_mask[..., tf.newaxis]
pred_mask = pred_mask[0]
pred_mask = tf.keras.preprocessing.image.array_to_img(pred_mask)
pred_mask = np.array(pred_mask)
ret, thresh = cv2.threshold(pred_mask, 126, 255, cv2.THRESH_BINARY)
kernel = np.ones((5, 5), np.uint8)
erudition_image = cv2.erode(thresh, kernel, iterations=2) #// make dilation image
dilation_image = cv2.dilate(erudition_image, kernel, iterations=2) #// make dilation image
dilation_image = cv2.resize(np.float32(dilation_image), dsize=(640,360), interpolation=cv2.INTER_AREA)
dilation_image = dilation_image != 255.0
# converting from BGR to HSV color space
hsv_frame = cv2.cvtColor(frame, cv2.COLOR_BGR2HSV)
# Red color
low_red = np.array([120, 155, 84])
high_red = np.array([179, 255, 255])
red_mask = cv2.inRange(hsv_frame, low_red, high_red)
red = cv2.bitwise_and(frame, frame, mask=red_mask)
# Blue color
low_blue = np.array([110, 130, 2])
high_blue = np.array([126, 255, 255])
blue_mask = cv2.inRange(hsv_frame, low_blue, high_blue)
kernel = np.ones((10, 10), np.uint8)
blue_mask = cv2.dilate(blue_mask, kernel, iterations=1) #// make dilation image
blue = cv2.bitwise_and(frame, frame, mask=blue_mask)
# Green color
low_green = np.array([25, 52, 72])
high_green = np.array([60, 255, 255])
green_mask = cv2.inRange(hsv_frame, low_green, high_green)
kernel = np.ones((5, 5), np.uint8)
green_mask = cv2.dilate(green_mask, kernel, iterations=1) #// make dilation image
green = cv2.bitwise_and(frame, frame, mask=green_mask)
mask = green_mask + blue_mask + dilation_image
result = cv2.bitwise_and(frame, frame, mask=mask)
result_mean = np.mean(result)
indy, indx, indz = np.where((result==0))
result[indy, indx, indz] = result_mean
#cv2.imwrite("/home/kimbring2/Desktop/output_seg" + "_" + str(step)+ "_.jpg", result)
#seg_video_out.write(np.uint8(result))
seg_video_frame = result
test_image = result
test_image = (test_image / 127.5) - 1
test_tensor = tf.convert_to_tensor(test_image, dtype=tf.float32)
test_tensor = tf.image.resize(test_tensor, [256, 256], method=tf.image.ResizeMethod.NEAREST_NEIGHBOR)
test_tensor = tf.reshape(test_tensor, [1,256,256,3], name=None)
prediction = f_gan(test_tensor)['conv2d_transpose_7'].numpy()
gan_result = prediction[0]* 0.5 + 0.5
#cv2.imwrite("/home/kimbring2/Desktop/output_gan" + "_" + str(step)+ "_.jpg", gan_result * 255.0)
#gan_video_out.write(np.uint8(gan_result * 255.0))
#gan_result = gan_result * 255
gan_video_frame = np.uint8(gan_result * 255.0)
camera_frame = cv2.resize(gan_result, (128, 128), interpolation=cv2.INTER_AREA)
step += 1
lidar_value = 0
def lidar_callback(msg):
global lidar_value
lidar_value = msg.data
#print("lidar: " + str(msg))
infrared_value = 'False'
def infrared_callback(msg):
global infrared_value
infrared_value = msg.data
#print("infrared: " + str(msg))
############## ROS Part ###############
rospy.init_node('deepsoccer')
wheel1 = rospy.Publisher('/deepsoccer_motors/cmd_str_wheel1', std_msgs.msg.String, queue_size=1)
wheel2 = rospy.Publisher('/deepsoccer_motors/cmd_str_wheel2', std_msgs.msg.String, queue_size=1)
wheel3 = rospy.Publisher('/deepsoccer_motors/cmd_str_wheel3', std_msgs.msg.String, queue_size=1)
wheel4 = rospy.Publisher('/deepsoccer_motors/cmd_str_wheel4', std_msgs.msg.String, queue_size=1)
solenoid = rospy.Publisher('/deepsoccer_solenoid/cmd_str', std_msgs.msg.String, queue_size=5)
roller = rospy.Publisher('/deepsoccer_roller/cmd_str', std_msgs.msg.String, queue_size=5)
rospy.Subscriber("/deepsoccer_camera/raw", Image, image_callback)
rospy.Subscriber("/deepsoccer_lidar", std_msgs.msg.String, lidar_callback)
rospy.Subscriber("/deepsoccer_infrared", std_msgs.msg.String, infrared_callback)
rate = rospy.Rate(5000)
stop_action = [0, 0, 0, 0, 'stop', 'none']
forward_action = [50, 1074, 1074, 50, 'in', 'none']
left_action = [1074, 1074, 1074, 1074, 'in', 'none']
right_action = [50, 50, 50, 50, 'in', 'out']
bacward_action = [1074, 50, 50, 1074, 'in', 'none']
hold_action = [0, 0, 0, 0, 'in', 'none']
kick_action = [0, 0, 0, 0, 'stop', 'out']
run_action = [100, 1124, 1124, 100, 'stop', 'none']
robot_action_list = [stop_action, forward_action, left_action, right_action, bacward_action, hold_action, kick_action, run_action]
%matplotlib notebook
############## ROS + Deep Learning Part ###############
memory_state = np.zeros([1,128], dtype=np.float32)
carry_state = np.zeros([1,128], dtype=np.float32)
while not rospy.is_shutdown():
#print("start")
action_index = 0
#print("lidar_value: " + str(lidar_value))
lidar_ = int(lidar_value) / 1200
#print("lidar_: " + str(lidar_))
#print("infrared_value: " + str(infrared_value))
infrared_ = int(infrared_value == 'True')
#print("infrared_: " + str(infrared_))
#print("action: " + str(action))
#print("")
clear_output(wait=True)
#print("camera_frame.shape: ", camera_frame.shape)
imgs_array = np.zeros((360*3,640,3))
imgs_array[0:360,:,:] = raw_video_frame
imgs_array[360:720,:,:] = seg_video_frame
imgs_array[720:720+256,0:256,:] = gan_video_frame
img = arrayShow(imgs_array)
display(img)
frame_state_channel = camera_frame
lidar_state_channel = (np.ones(shape=(128,128,1), dtype=np.float32)) * lidar_
infrared_state_channel = (np.ones(shape=(128,128,1), dtype=np.float32)) * infrared_ / 2.0
state_channel1 = np.concatenate((frame_state_channel, lidar_state_channel), axis=2)
state_channel2 = np.concatenate((state_channel1, infrared_state_channel), axis=2)
state_channel2 = np.array([state_channel2])
#print("state_channel2.shape: " + str(state_channel2.shape))
state_channel_tensor = tf.convert_to_tensor(state_channel2, dtype=tf.float32)
memory_state = tf.convert_to_tensor(memory_state, dtype=tf.float32)
carry_state = tf.convert_to_tensor(carry_state, dtype=tf.float32)
#prediction = f_rl(input_1=state_channel_tensor, input_2=memory_state, input_3=carry_state)
#action_logit = prediction['dense_1'].numpy()[0]
#memory_state = prediction['core_lstm'].numpy()
#carry_state = prediction['core_lstm_1'].numpy()
#predict_value = f_rl(state_channel_tensor)['dueling_model'].numpy()[0].numpy()[0]
#print("action_logit: " + str(action_logit))
#print("memory_state.shape: " + str(memory_state.shape))
#print("carry_state.shape: " + str(carry_state.shape))
#action_index = np.argmax(action_logit, axis=0)
#print("action_index: " + str(action_index))
action_index = 0
#print("")
action = robot_action_list[action_index]
wheel1_action = action[0]
wheel2_action = action[1]
wheel3_action = action[2]
wheel4_action = action[3]
roller_action = action[4]
solenoid_action = action[5]
wheel1.publish(str(wheel1_action))
wheel2.publish(str(wheel2_action))
wheel3.publish(str(wheel3_action))
wheel4.publish(str(wheel4_action))
roller.publish(roller_action)
solenoid.publish(solenoid_action)
#time.sleep(0.1)
rate.sleep()
```
| github_jupyter |
<div class="alert alert-block alert-success">
<b><center>SOFTMAX CLASSIFICATION</center></b>
<b><center>Multiclass Classification에 댸한 기본을 이해하기 위한 코드</center></b>
</div>
# Configure Learning Environment
```
# !pip install git+https://github.com/nockchun/rspy --force
import rspy as rsp
rsp.setSystemWarning(off=True)
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
import seaborn as sns
from ipywidgets import interact
np.set_printoptions(linewidth=200, precision=2)
tf.__version__
```
# Understanding The Concept
```
eduConf = rsp.EduPlotConf(font="NanumGothicCoding", figScale=1)
plotExp = rsp.EduPlot2D(eduConf)
x = np.linspace(-10, 10, 200)
plotExp.addXYData(x, np.exp(x), name="exponential")
plotExp.addText([[1, 3.5]], ["${e}^{x}$"], name="exponential")
plotExp.addXYData(x, 2**x, color="#FF9900", name="exponential2")
plotExp.addText([[2.2, 3]], ["${2}^{x}$"], color="#FF9900", name="exponential2")
plotExp.addXYData(x, 10**x, color="#FF0000", name="exponential10")
plotExp.addText([[0, 3]], ["${10}^{x}$"], color="#FF0000", name="exponential10")
plotExp.genSpace(4)
plt.show()
plt.close()
```
## Optimize
$$ \underset { W }{ minimize } \quad cost(W,b) = W -\alpha \frac { \partial }{ \partial W } cost(W) $$
# Logistic Classification
## Prepare Datas
```
data = [
[1, 9, 8],
[2, 1, 9],
[3, 2, 1],
[4, 3, 2],
[5, 4, 3],
[6, 5, 4],
[7, 6, 5],
[8, 7, 6]
]
label = [
[0, 0, 1],
[0, 0, 1],
[0, 0, 1],
[0, 1, 0],
[0, 1, 0],
[0, 1, 0],
[1, 0, 0],
[1, 0, 0]
]
data = np.array(data, dtype=float)
label = np.array(label, dtype=float)
data.shape, label.shape
```
## Generate Model
```
X = tf.placeholder(tf.float32, shape=[None,3], name="data")
Y = tf.placeholder(tf.float32, shape=[None,3], name="label")
W = tf.Variable(tf.random_normal([3, 3]), name="weight")
b = tf.Variable(tf.random_normal([3]), name="bias")
hypothesis = tf.nn.softmax(tf.matmul(X, W) + b)
loss = - tf.reduce_mean(Y * tf.log(hypothesis) + (1-Y) * tf.log(1-hypothesis))
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.01).minimize(loss)
```
## Training
```
sess = tf.Session()
init = tf.global_variables_initializer()
sess.run(init)
for step in range(20001):
loss_val, hy_val, _ = sess.run([loss, hypothesis, optimizer], feed_dict= {X:data, Y:label})
if step % 10000 == 0:
print (loss_val, hy_val)
```
## Predict
```
predicted = tf.cast(hypothesis>0.5, dtype=tf.float32)
accuracy = tf.reduce_mean(tf.cast(tf.equal(predicted, Y), dtype=tf.float32))
sess.run([hypothesis, predicted, accuracy], feed_dict={X:data, Y:label})
sess.close()
```
| github_jupyter |
```
# Import helper module from local folder
import sys
import os
sys.path.append(os.getcwd())
from resources import helper
# Numerical and plotting tools
import numpy as np
import matplotlib.pyplot as plt
# Import SI unit conversion factors
from resources.helper import GHz, MHz, kHz, us, ns
# Importing standard Qiskit libraries
from qiskit import IBMQ
from qiskit.tools.jupyter import *
IBMQ.save_account('YOUR-IBM-TOKEN')
print(str(IBMQ.stored_account()))
# Loading your IBM Quantum account
IBMQ.load_account()
IBMQ.providers() # see a list of providers you have access to
from qiskit.pulse import DriveChannel, Gaussian
# Get the special provider assigned to you using information from the output above
hub_name = 'iqc2021-1' # e.g. 'iqc2021-1'
group_name = 'challenge-69' # e.g. 'challenge-1'
project_name = 'ex4' # Your project name should be 'ex4'
provider = IBMQ.get_provider(hub=hub_name, group=group_name, project=project_name)
# Get `ibmq_jakarta` backend from the provider
backend_name = 'ibmq_jakarta'
backend = provider.get_backend(backend_name)
backend # See details of the `ibmq_jakarta` quantum system
from qiskit import pulse
from qiskit.pulse import Play, Schedule, DriveChannel
# Please use qubit 0 throughout the notebook
qubit = 0
backend_config = backend.configuration()
exc_chans = helper.get_exc_chans(globals())
dt = backend_config.dt
print(f"Sampling time: {dt*1e9} ns")
backend_defaults = backend.defaults()
center_frequency = backend_defaults.qubit_freq_est
inst_sched_map = backend_defaults.instruction_schedule_map
inst_sched_map.instructions
# Retrieve calibrated measurement pulse from backend
meas = inst_sched_map.get('measure', qubits=[qubit])
meas.exclude(channels=exc_chans).draw(time_range=[0,1000])
from qiskit.pulse import DriveChannel, Gaussian
# The same spec pulse for both 01 and 12 spec
drive_amp = 0.25
drive_duration = inst_sched_map.get('x', qubits=[qubit]).duration
# Calibrated backend pulse use advanced DRAG pulse to reduce leakage to the |2> state.
# Here we will use simple Gaussian pulse
drive_sigma = drive_duration // 4 # DRAG pulses typically 4*sigma long.
spec_pulse = Gaussian(duration=drive_duration, amp=drive_amp,
sigma=drive_sigma, name=f"Spec drive amplitude = {drive_amp}")
# Construct an np array of the frequencies for our experiment
spec_freqs_GHz = helper.get_spec01_freqs(center_frequency, qubit)
# Create the base schedule
# Start with drive pulse acting on the drive channel
spec01_scheds = []
for freq in spec_freqs_GHz:
with pulse.build(name="Spec Pulse at %.3f GHz" % freq) as spec01_sched:
with pulse.align_sequential():
# Pay close attention to this part to solve the problem at the end
pulse.set_frequency(freq*GHz, DriveChannel(qubit))
pulse.play(spec_pulse, DriveChannel(qubit))
pulse.call(meas)
spec01_scheds.append(spec01_sched)
# Draw spec01 schedule
spec01_scheds[-1].exclude(channels=exc_chans).draw(time_range=[0,1000])
from qiskit.tools.monitor import job_monitor
# Run the job on a real backend
spec01_job = backend.run(spec01_scheds, job_name="Spec 01", **helper.job_params)
print(spec01_job.job_id())
job_monitor(spec01_job)
from resources.helper import SpecFitter
amp_guess = 5e6
f01_guess = 10
B = 1.2
C = -0.1
fit_guess = [amp_guess, f01_guess, B, C]
fit = SpecFitter(spec01_job.result(), spec_freqs_GHz, qubits=[qubit], fit_p0=fit_guess)
fit.plot(0, series='z')
f01 = fit.spec_freq(0, series='z')
print("Spec01 frequency is %.6f GHz" % f01)
max_rabi_amp = 0.75
rabi_amps = helper.get_rabi_amps(max_rabi_amp)
rabi_scheds = []
for ridx, amp in enumerate(rabi_amps):
with pulse.build(name="rabisched_%d_0" % ridx) as sched: # '0' corresponds to Rabi
with pulse.align_sequential():
pulse.set_frequency(f01*GHz, DriveChannel(qubit))
rabi_pulse = Gaussian(duration=drive_duration, amp=amp, \
sigma=drive_sigma, name=f"Rabi drive amplitude = {amp}")
pulse.play(rabi_pulse, DriveChannel(qubit))
pulse.call(meas)
rabi_scheds.append(sched)
# Draw rabi schedule
rabi_scheds[-1].exclude(channels=exc_chans).draw(time_range=[0,1000])
# Run the job on a real device
rabi_job = backend.run(rabi_scheds, job_name="Rabi", **helper.job_params)
print(rabi_job.job_id())
job_monitor(rabi_job)
# If the queuing time is too long, you can save the job id
# And retrieve the job after it's done
# Replace 'JOB_ID' with the the your job id and uncomment to line below
#rabi_job = backend.retrieve_job('JOB_ID')
from qiskit.ignis.characterization.calibrations.fitters import RabiFitter
amp_guess = 5e7
fRabi_guess = 2
phi_guess = 0.5
c_guess = 0
fit_guess = [amp_guess, fRabi_guess, phi_guess, c_guess]
fit = RabiFitter(rabi_job.result(), rabi_amps, qubits=[qubit], fit_p0=fit_guess)
fit.plot(qind=0, series='0')
x180_amp = fit.pi_amplitude()
print("Pi amplitude is %.3f" % x180_amp)
```
<div id='problem'></div>
<div class="alert alert-block alert-success">
## Step 3 (Problem): Find $|1\rangle \rightarrow |2\rangle$ transition frequency
In order to observe the transition between the $|1\rangle$ and $|2\rangle$ states of the transmon, you need to:
1. Apply an $X_\pi$ pulse to transition the qubit from $|0\rangle$ to $|1\rangle$.
1. Apply a second pulse with varying frequency to find the $|1\rangle \rightarrow |2\rangle$ transition.
</div>
<div class="alert alert-block alert-danger">
The cell below is the only one you need to modify in the entire notebook.
</div>
```
backend_defaults = backend.defaults()
center_frequency = backend_defaults.qubit_freq_est
inst_sched_map = backend_defaults.instruction_schedule_map
inst_sched_map.instructions
# Define pi pulse
drive_duration = inst_sched_map.get('x', qubits=[qubit]).duration
x_pulse = Gaussian(duration=drive_duration,
amp=x180_amp,
sigma=drive_sigma,
name='x_pulse')
def build_spec12_pulse_schedule(freq, anharm_guess_GHz):
with pulse.build(name="Spec Pulse at %.3f GHz" % (freq+anharm_guess_GHz)) as spec12_schedule:
with pulse.align_sequential():
# WRITE YOUR CODE BETWEEN THESE LINES - START
pulse.set_frequency(f01*GHz, DriveChannel(qubit))
pulse.play(x_pulse, DriveChannel(qubit))
pulse.set_frequency((freq+anharm_guess_GHz)*GHz, DriveChannel(qubit))
pulse.play(spec_pulse, DriveChannel(qubit))
pulse.call(meas)
# WRITE YOUR CODE BETWEEN THESE LINES - END
return spec12_schedule
```
The anharmonicity of our transmon qubits is typically around $-300$ MHz, so we will sweep around that value.
```
anharmonicity_guess_GHz = -0.3 # your anharmonicity guess
freqs_GHz = helper.get_spec12_freqs(f01, qubit)
# Now vary the sideband frequency for each spec pulse
spec12_scheds = []
for freq in freqs_GHz:
spec12_scheds.append(build_spec12_pulse_schedule(freq, anharmonicity_guess_GHz))
# Draw spec12 schedule
spec12_scheds[-1].exclude(channels=exc_chans).draw(time_range=[0,1000])
# Run the job on a real device
spec12_job = backend.run(spec12_scheds, job_name="Spec 12", **helper.job_params)
print(spec12_job.job_id())
job_monitor(spec12_job)
# If the queuing time is too long, you can save the job id
# And retrieve the job after it's done
# Replace 'JOB_ID' with the the your job id and uncomment to line below
#spec12_job = backend.retrieve_job('JOB_ID')
```
### Fit the Spectroscopy Data
<div id='fit-f12'></div>
We will again fit the spectroscopy signal to a Lorentzian function of the form
$$ \frac{AB}{\pi[(f-f_{12})^2 + B^2]} + C $$
to find the frequency of the $|1\rangle \to |2\rangle$ transition $f_{12}$ with these fitting parameters:
Parameter | Corresponds to
--- | ---
$A$ | amplitude
$f_{12}$ | 12 frequency guess (GHz)
$B$ | scale
$C$ | offset
<div class="alert alert-block alert-danger">
**Note:** You may need to modify the fitting parameters below to get a good fit.
</div>
```
amp_guess = 5e7
f12_guess = f01 - 0.3
B = .1
C = 0
fit_guess = [amp_guess, f12_guess, B, C]
fit = SpecFitter(spec12_job.result(), freqs_GHz+anharmonicity_guess_GHz, qubits=[qubit], fit_p0=fit_guess)
fit.plot(0, series='z')
f12 = fit.spec_freq(0, series='z')
print("Spec12 frequency is %.6f GHz" % f12)
# Check your answer using following code
from qc_grader import grade_ex4
grade_ex4(f12,qubit,backend_name)
```
## Calculating $E_J/E_c$
Modifying the equations in the introduction section, we can calculate $E_c$ and $E_J$ using $f_{01}$ and $f_{12}$ obtained from the pulse experiments:
$$
E_c = -\delta = f_{01} - f_{12} \qquad E_J = \frac{(2f_{01}-f_{12})^2}{8(f_{01}-f_{12})}
$$
```
Ec = f01 - f12
Ej = (2*f01-f12)**2/(8*(f01-f12))
print(f"Ej/Ec: {Ej/Ec:.2f}") # This value is typically ~ 30
```
| github_jupyter |
# ES Module 3
Welcome to Module 3!
Last time, we went over:
1. Strings and Intergers
2. Arrays
3. Tables
Today we will continue working with tables, and introduce a new procedure called filtering. Before you start, run the following cell.
```
# Loading our libraries, i.e. tool box for our module
import numpy as np
from datascience import *
```
### Paired Programming
Today we want to introduce a new system of work called paired programming. Wikipedia defines paired programming in the following way:
Pair programming is an agile software development technique in which two programmers work together at one workstation. One, the driver, writes code while the other, the observer or navigator, reviews each line of code as it is typed in. The two programmers switch roles frequently.
This methodolgy is quite known in the computer science realm, and we want to try and see how well it would work in our little class room. Hopefully we would all benefit from this, by closing the gap between more experienced coders and less so we could move forward to more advanced topics! Additionally, there is always the benefit of having a friend when all hell breaks loose (or the code just would not work..)
So after this brief introduction, please team up with a class-mate, hopefully someone you did not know from before that is at a slightly different level of programming experience.
Please start now, as one takes the controls and the other is reviewing the code.
## 0. Comments
Comments are ways of making your code more human readable. It's good practice to add comments to your code so someone else reading your code can get an idea of what's going on.
You can add a comment to your code by preceeding it with a `#` symbol. When the computer sees any line preceeded by a `#` symbol, it'll ignore it. Here's an example below:
```
# Calculating the total number of pets in my house.
num_cats = 4
num_dogs = 10
total = num_cats + num_dogs
total
```
Now, write a comment in the cell below explaining what it is doing, then run the cell to see if you're correct.
```
animals = make_array('Cat', 'Dog', 'Bird', 'Spider')
num_legs = make_array(4, 4, 2, 8)
my_table = Table().with_columns('Animal', animals,
'Number of Legs', num_legs)
my_table
```
## 1. Tables (Continued)
It is time to practice tables again. We want to load the table files you have uploaded last module. This time, you do it by yourself. Load the table "inmates_by_year.csv" and "correctional_population.csv" and assign it to a variable. Remember, to load a table we use `Table.read_table()` and pass the name of the table as an argument to the function.
```
inmates_by_year = Table.read_table('inmates_by_year.csv')
correctional_population = Table.read_table('correctional_population.csv')
```
Good job! Now we have all the tables loaded.
It is time to extract some information from these tables!
In the next several cells, we will guide you through a quick manipulation that will allow us to extract information about the entire correctional population using both tables we have loaded above.
In the correctional_population table, we are given data about the number of supervised per 100,000 U.S. adult residents. That means that if we want to have the approximated number of the entire population under supervision we need to multiply by 100,000.
```
# First, extract the column name "Number supervised per 100,000 U.S. adult residents/c" from
# the correctional_population table and assign it to the variable provided.
c_p = ...
c_p
```
#### filtering
When you run the cell above, you may notice that the values in our array are actually strings (you can tell because each value has quotation marks around it). However, we can't do mathematical operations on strings, so we'll have to convert this array first so it has integers instead of strings. This is called filtering, or cleaning the data, so we can actually do some work on it. In the following cells, when you see the `# filtering` sign, know that we have yet to cover this topic.
Run the following cell to do clean the table. We'll go over how to do this in a later section of this module. If you have any questions about how it works, feel free to ask any of us!
```
# filtering
def string_to_int(val):
return int(val.replace(',', ''))
c_p = correctional_population.apply(string_to_int, 'Number supervised per 100,000 U.S. adult residents/c')
```
Now, let's continue finding the real value of c_p.
```
# In this cell, multiply the correctional population column name "Number supervised per 100,000 U.S. adult residents/c"
# by 100000 and assign it to a new variable (c_p stands for correctional population)
real_c_p = ...
real_c_p
```
Next we want to assign the Total column from inmates_by_year to a variable in order to be able to operate on it.
```
total_inmates = ...
total_inmates
```
Again, run the following line to convert the values in `total_inmates` to ints.
```
# filtering
total_inmates = inmates_by_year.apply(string_to_int, 'Total')
total_inmates
```
#### Switch position, the navigator now takes the wheel.
Now that we have the variables holding all the information we want to manipulate, we can start digging into it.
We want to come up with a scheme that will allow us to see the precentage of people that are incarcerated, from the total supervised population, by year.
Before we do that, though, examine your two variables, `total_inmates` and `real_c_p` and their corresponding tables. Do you foresee any issues with directly comparing these two tables?
The `correctional_population` table has a row corresponding to 2000, which `inmates_by_year` does not have. This not only means that the data from our two tables doesn't match up, but also that our arrays are two different lengths. Recall that we cannot do operations on arrays with different lengths.
To fix this, run the following cell, in which we get rid of the value corresponding to the year 2000 from `real_c_p`. Again, if you have questions about how this works, feel free to ask us!
```
# filtering
real_c_p = real_c_p.take(np.arange(1, real_c_p.size))
real_c_p
```
Now our arrays both correspond to data from the same years and we can do operations with both of them!
```
# Write a short code that stores the precentage of people incarcerated from the supervised population
# (rel stands for relative, c_p stands from correctional population)
inmates_rel_c_p = ...
inmates_rel_c_p
```
Now, this actually gives us useful information!
Why not write it down? Please write down what this information tells you about the judicial infrastructure - we are looking for more mathy/dry explanation (rather than observation of how poorly it is).
```
# A simple sentence will suffice, we want to see intuitive understanding. Please call a teacher when done to check!
extract_information_shows = "YOUR ANSWER HERE"
extract_information_shows
```
For a final touch, please sort inmates_rel_c_p by descending order in the next cell. We won't tell you how to sort, this time please check the last lab module on how to sort a table. It is an important quality of a programmer to be able to reuse code you already have.
Hint: Remember that you can only use `sort` on tables. How might you manipulate your array so that you can sort it?
```
# Please sort inmates_rel_c_p in descending order and display the output
inmates_rel_c_p = ...
...
```
#### Before starting, please switch positions
## Filtering
Right now, we can't really get much extra information from our tables other than by sorting them. In this section, we'll learn how to filter our data so we can get more useful insights from it. This is especially useful when dealing with larger data sets!
For example, say we wanted insights about the total number of inmates after 2012. We can find this out using the `where` function. Check out the cell below for an example of how to use this.
```
inmates_by_year.where('Year', are.above(2012))
```
Notice that `where` takes in two arguments: the name of the column, and the condition we are filtering by.
Now, try it for yourself! In the cell below, filter `correctional_population` so it only includes years after 2008.
If you run the following cell, you'll find a complete description of all such conditions (which we'll call predicates) that you can pass into where. This information can also be found [here](https://www.inferentialthinking.com/chapters/05/2/selecting-rows.html).
```
functions = make_array('are.equal_to(Z)', 'are.above(x)', 'are.above_or_equal_to(x)', 'are.below(x)',
'are.below_or_equal_to(x)', 'are.between(x, y)', 'are.strictly_between(x, y)',
'are.between_or_equal_to(x, y)', 'are.containing(S)')
descriptions = make_array('Equal to Z', 'Greater than x', 'Greater than or equal to x', 'Below x',
'Less than or equal to x', 'Greater than or equal to x, and less than y',
'Greater than x and less than y', 'Greater than or equal to x, and less than or equal to y',
'Contains the string S')
predicates = Table().with_columns('Predicate', functions,
'Description', descriptions)
predicates
```
Now, we'll be using filtering to gain more insights about our two tables. Before we start, be sure to run the following cell so we can ensure every column we're working with is numerical.
```
inmates_by_year = inmates_by_year.drop('Total').with_column('Total', total_inmates).select('Year', 'Total', 'Standard error/a')
correctional_population = correctional_population.drop('Number supervised per 100,000 U.S. adult residents/c').with_column('Number supervised per 100,000 U.S. adult residents/c', c_p).select('Year', 'Number supervised per 100,000 U.S. adult residents/c', 'U.S. adult residents under correctional supervision ').relabel('U.S. adult residents under correctional supervision ', 'U.S. adult residents under correctional supervision')
```
First, find the mean of the total number of inmates. Hint: You can use the `np.mean()` function on arrays to calculate this.
```
avg_inmates = ...
avg_inmates
```
Now, filter `inmates_by_year` to find data for the years in which the number of total inmates was under the average.
```
filtered_inmates = ...
filtered_inmates
```
What does this tell you about the total inmate population? Write your answer in the cell below.
```
answer = "YOUR TEXT HERE"
```
#### Before continuing, please switch positions.
Now, similarly, find the average number of adults under correctional supervision, and filter the table to find the years in which the number of adults under correctional supervision was under the average.
```
avg = ...
filtered_c_p = ...
filtered_c_p
```
Do the years match up? Does this make sense based on the proportions you calculated above in `inmates_rel_c_p`?
```
answer = "YOUR TEXT HERE"
```
Now, from `correctional_population`, filter the table so the value of U.S. adult residents under correctional supervision is 1 in 31. Remember, the values in this column are strings.
```
c_p_1_in_34 = ...
c_p_1_in_34
```
Now, we have one last challenge exercise. Before doing this, finish the challenge exercises from last module. We highly encourage you to work with your partner on this one.
In the following cell, find the year with the max number of supervised adults for which the proportion of US adult residents under correctional supervision was 1 in 32.
The lines given below are from the staff solution. Feel free to solve this problem in as many or as few lines as you need.
```
one_in_32 = ...
one_in_32_sorted = ...
year = ...
year
```
Congratulations, you're done with this module! Before you leave, please fill out this [link](https://docs.google.com/a/berkeley.edu/forms/d/1KQHzw-rh_E--lnQ7ItLrOcH7WJUTexDiKMLcuPwClzo/edit?usp=drive_web) to give us feedback on how we can make these modules more useful for you.
| github_jupyter |
```
from cyllene import *
%initialize 22_activity_5_01
```
# Activity 5 - Exponential and Logarithmic Functions
## Part 1 - Review of one-to-one functions and inverse functions
### Introduction
Recall what is meant by a one-to-one function. Given a function, we say that it is one-to-one if every element in the range of the function corresponds to only one element in the domain of a function. That is, given a function $f(x)$,
> if $f(x_1) = f(x_2)$, then $x_1 = x_2$.
(If the output was the same, then the input must have been the same.)
Consider the following three functions, two of the functions are one-to-one functions; which one is NOT a one-to-one function?
> 1. $f(x) = x + 2$ with domain $x \in (-\infty, \infty )$
> 2. $g(x) = x^2$ with domain $x \in (-\infty, \infty )$
> 3. $h(x) = x^2$ with domain $x \in [0, \infty)$
In the past, you may have sketched the functions and employed the "horizontal line test" to determine whether or not the function was one-to-one. That is, if a horizontal line intersected the graph in more then one point, then the function was not one-to-one.
Upon sketching the first function, $y=f(x)$, we see it is a line with slope $m=1$.
Any horizontal line will cross this sketch in exactly one spot.
In fact, given any value in the range of the function, for example $y=10$, we can calculate the corresponding $x$ value that gets mapped to it; that is, if $x+2=10$, then $x = 10-2 = 8$. [Note $f(8) = 10$]
```
plot(f, (x,-12,12));
```
On first glance, it may appear the the next two functions are the same; however, it is important to remember that associated with any functional definition is the domain of the function. Even though $g(x)=x^2$ and $h(x)=x^2$, the domains are different, thus they are really two different functions. The function $g(x)$ defined on all real numbers is NOT a one-to-one function.
A counter example would be $g(-2) = (-2)^2 = 4$ and $g(2) = 2^2 = 4$. Given an output, $y=4$, we have two inputs from the domain, $x_1= -2$ and $x_2=2$ that get mapped to $y=4$. Sketch the parabola $y=x^2$ from $x=-3$ to $x=3$ and verify that the horizontal line, $y=4$, intersects the graph at two points, $(-2, 4)$ and $(2, 4)$. Thus the function $g(x)$ as defined is NOT a one-to-one function.
```
G_g.show();
```
However, if you restrict your domain to just $x \ge 0$, the "sketch" of the function will only be the right half of the parabola you sketched. The horizontal line $y=4$ will only cross the graph in one place. In fact, any horizontal line $y=k$ where $k>0$ will pass through the right half of parabola exactly once. The third function is a one-to-one function.
```
G_h.show();
```
### The consequence of a one-to-one determination: The existence of an inverse function!
If a function is one-to-one, then an inverse *function* exists. For $f(x) = x + 2$ with domain $x \in (-\infty, \infty )$, we may write out the inverse function by writing $y = x + 2$ and solving for $x$, that is, $x = y-2$, which represents the inverse function, written $f^{-1}(y) = y-2$. Employing the forward function, $f(8) = 8+2 = 10$. Then, the inverse, $f^{-1}(10) = 10-2 = 8$, takes us back!
For the function $h(x) = x^2$ with domain $x \in [0, \infty)$, we can do a similar process to determine the inverse function. We let $y = x^2$ and solve for $x$ to get $x = \sqrt{y}$ and $x = -\sqrt{y}$. However, since we restricted the domain of $h(x)$ to be $x \ge 0$, we need only consider the positive square root and $h^{-1}(y) = \sqrt{y}$.
Notice of the second function from the Introduction, $g(x)$, since the domain was not resricted, the inverse process is not a "functional" process; for example, if $y=4$, going in reverse, there are two $x$-values associated with it, $x=-\sqrt{4}=-2$ and $x=\sqrt{4}=2$ (the horizontal line test).
In short, given $y = f(x)$, then $x = f^{-1}(y)$. Also note that the domain of the forward function $f(x)$ is the range of the inverse function, and the range of $f(x)$ is the domain of the inverse function.
### A note on notation.
Given $f(x) = x+2$, in lieu of writing $f^{-1}(y) = y-2$, since the variable $y$ represents the input and $y-2$ is the rule (what to do with the input), often the inverse is also written with the dummy variable $x$, that is, $f^{-1}(x) = x-2$. However, the "$x$" in $f(x)$ and the "$x$" in $f^{-1}(x)$ represent two different quantities, most often with different units.
### Explore the outputs of a basic exponential function
Consider the function $f(x) = 2^x$ defined on $(-\infty, \infty)$
## Exercise
```
%problem 1
%%answer Problem 1
(1): 1/4
(2): 1/2
(3): 1
(4): 2
(5): 4
```
## Exercise
Below is a plot of the function $f(x) = 2^x$. Use the horizontal line test to determine if the function is one-to-one.
```
plot(f_exp(x), (x,-3,3));
%problem 2
%%answer Problem 2
```
Since the function $f(x) = 2^x$ is one-to-one, we know that a unique inverse function exists. We can't just write $y = 2^x$ and "solve for $x$", we need new notation to depict this new function.
> If $f(x) = 2^x$, then $f^{-1}(y) = \log_2(y)$.
In the graph above, consider the horizontal line through $y=4$ on the vertical axis and a vertical line through $x=2$ on the horizontal axis and note that it intersects the curve at the point $(2, 4)$. This may be interpreted as $2^2=4$ or as $\log_2(4) = 2$
Using this idea, confirm the following values of $\log_2(y)$:
```
%problem 3
%%answer Problem 3
(1):
(2):
(3):
```
### Closing Considerations
>1. If $y = b^x$, where wher the base $b>0$ and $b \neq 1$, then $x = \log_b (y)$; that is, given $y = f(x)$, then $x = f^{-1}(y)$.
>2. Note that the domain (valid input values) of $f(x) = 2^x$ is $(-\infty, \infty)$, but the range (outputs) of $f(x) = 2^x > 0$.
This implies the domain of $f^{-1}(x) = \log_2(x)$ is $x>0$ (See the note on notation above).
In general, the only valid input value into $\log_b(x)$ is a value $x>0$ (e.g. the domain of the function $\log_b(x)$ is $x>0$)
>3. Since $2^0 = 1$, and in general $b^0 = 1$, we have $\log_b(1) = 0$ regardless of base $b$.
| github_jupyter |
<center><img alt="" src="images/Cover_NLPTM.jpg"/></center>
## <center><font color="blue">Representasi Dokumen</font></center>
<b><center>(C) Taufik Sutanto - 2020</center>
<center>tau-data Indonesia ~ https://tau-data.id ~ taufik@tau-data.id</center>
```
# Installing Modules for Google Colab
import nltk
!wget https://raw.githubusercontent.com/taudata-indonesia/eLearning/master/taudataNlpTm.py
!mkdir data
!wget -P data/ https://raw.githubusercontent.com/taudata-indonesia/eLearning/master/data/slang.txt
!wget -P data/ https://raw.githubusercontent.com/taudata-indonesia/eLearning/master/data/stopwords_id.txt
!wget -P data/ https://raw.githubusercontent.com/taudata-indonesia/eLearning/master/data/stopwords_en.txt
!wget -P data/ https://raw.githubusercontent.com/taudata-indonesia/eLearning/master/data/wn-ind-def.tab
!wget -P data/ https://raw.githubusercontent.com/taudata-indonesia/eLearning/master/data/wn-msa-all.tab
!wget -P data/ https://raw.githubusercontent.com/taudata-indonesia/eLearning/master/data/all_indo_man_tag_corpus_model.crf.tagger
!pip install spacy python-crfsuite unidecode textblob sastrawi tweepy twython
!python -m spacy download en
!python -m spacy download xx
!python -m spacy download en_core_web_sm
nltk.download('popular')
import taudataNlpTm as tau, seaborn as sns; sns.set()
import tweepy, json, nltk, urllib.request
from textblob import TextBlob
from nltk.tokenize import TweetTokenizer
from twython import TwythonStreamer
from tqdm import tqdm_notebook as tqdm
```
<h1 id="Vector-Space-Model---VSM">Vector Space Model - VSM</h1>
<p><img alt="" src="images/vsm.png" style="width: 300px; height: 213px;" /></p>
## <font color="blue">Outline Representasi Dokumen :</font>
* Representasi Sparse (VSM): Binary, tf &-/ idf, Custom tf-idf, BM25
* Frequency filtering, n-grams, vocabulary based
* Representasi Dense: Word Embedding (Word2Vec dan FastText)
* Tensor to Matrix representation untuk model Machine Learning di Text Mining
<ul>
<li>Data yang biasanya kita ketahui berbentuk <strong>tabular </strong>(tabel/kolom-baris/matriks/<em>array</em>/larik), data seperti ini disebut data terstruktur (<strong><em>structured data</em></strong>).</li>
<li>Data terstruktur dapat disimpan dengan baik di <em>spreadsheet</em> (misal: <em>Excel/CSV</em>) atau basis data (<em>database</em>) relasional dan secara umum dapat digunakan langsung oleh berbagai model/<em>tools</em> statistik/data mining konvensional.</li>
<li>Sebagian data yang lain memiliki “<em>tags</em>” yang menjelaskan elemen semantik yang berbeda di dalamnya dan cenderung tidak memiliki skema (struktur) yang statis.</li>
<li>Data seperti ini disebut data<em> <strong>semi-structured</strong></em>, contohnya data dalam bentuk <strong><a href="http://www.w3.org/XML/" target="_blank">XML</a></strong>.</li>
<li>Apa bedanya? Apa maksudnya tidak memiliki skema yang statis? Penjelasan mudahnya bayangkan sebuah data terstruktur (tabular), namun dalam setiap baris (<em>record/instance</em>)-nya tidak memiliki jumlah variabel (peubah) yang sama.</li>
<li>Tentu saja data seperti ini tidak sesuai jika disimpan dan diolah dengan <em>tools/software</em> yang mengasumsikan struktur yang statis pada setiap barisnya (misal: Excel dan SPSS).</li>
</ul>
<p><img alt="" src="images/3_tipeData.png" style="height: 400px ; width: 430px" /></p>
<ul>
<li>Data multimedia seperti teks, gambar atau video <strong>tidak dapat</strong> <strong>secara langsung</strong> dianalisa dengan model statistik/data mining.</li>
<li>Sebuah proses awal <em>(pre-process)</em> harus dilakukan terlebih dahulu untuk merubah data-data tidak (semi) terstruktur tersebut menjadi bentuk yang dapat digunakan oleh model statistik/data mining konvensional.</li>
<li>Terdapat berbagai macam cara mengubah data-data tidak terstruktur tersebut ke dalam bentuk yang lebih sederhana, dan ini adalah suatu bidang ilmu tersendiri yang cukup dalam. Sebagai contoh saja sebuah teks biasanya dirubah dalam bentuk vektor/<em>topics</em> terlebih dahulu sebelum diolah.</li>
<li>Vektor data teks sendiri bermacam-macam jenisnya: ada yang berdasarkan eksistensi (<strong><em>binary</em></strong>), frekuensi dokumen (<strong>tf</strong>), frekuensi dan invers jumlah dokumennya dalam corpus (<strong><a href="https://en.wikipedia.org/wiki/Tf%E2%80%93idf" target="_blank">tf-idf</a></strong>), <strong>tensor</strong>, dan sebagainya.</li>
<li> Proses perubahan ini sendiri biasanya tidak <em>lossless</em>, artinya terdapat cukup banyak informasi yang hilang. Maksudnya bagaimana? Sebagai contoh ketika teks direpresentasikan dalam vektor (sering disebut sebagai model <strong>bag-of-words</strong>) maka informasi urutan antar kata menghilang. </li>
</ul>
<p><img alt="" src="images/3_structureData.png" style="height:270px; width:578px" /></p>
<p><strong>Contoh bentuk umum representasi dokumen:</strong></p>
<p><img alt="" src="images/3_Bentuk umum representasi dokumen.JPG" style="height: 294px ; width: 620px" /></p>
<p>Pada Model <em>n-grams</em> kolom bisa juga berupa frase.</p>
<h2 id="Document-Term-Matrix-:-Vector-Space-Model---VSM">Document-Term Matrix : Vector Space Model - VSM</h2>
<p><img alt="" src="images/vsm_matrix.png" style="width: 500px; height: 283px;" /></p>
<p><img alt="" src="images/3_rumus tfidf.png" style="height:370px; width:367px" /></p>
<p><img alt="" src="images/3_tfidf logic.jpg" style="height:359px; width:638px" /></p>
<p><img alt="" src="images/3_variant tfidf.png" style="height:334px; width:955px" /></p>
K = |d|
# pertama-tama mari kita Load Data twitter dari pertemuan sebelumnya
* Silahkan gunakan data baru (crawl lagi) jika diinginkan
```
def loadTweets(file='Tweets.json'):
f=open(file,encoding='utf-8', errors ='ignore', mode='r')
T=f.readlines();f.close()
for i,t in enumerate(T):
T[i] = json.loads(t.strip())
return T
# karena ToS data json ini dikirimkan terpisah hanya untuk kalangan terbatas.
import json
T2 = loadTweets(file='data/tweets_sma-01.json')
print('Total data = {}'.format(len(data)))
print('tweet pertama oleh "{}" : "{}"'.format(T2[0]['user']['screen_name'],T2[0]['full_text']))
# Contoh mengambil hanya data tweet
data = [t['full_text'] for t in T2]
data[:5] # 5 tweet pertama
```
# PreProcessing Data Text-nya
```
# pre processing
import taudataNlpTm as tau
from tqdm import tqdm_notebook as tqdm
# cleanText(T, fix={}, lemma=None, stops = set(), symbols_remove = True, min_charLen = 2, fixTag= True)
stops, lemmatizer = tau.LoadStopWords(lang='id')
stops.add('rt')
stops.add('..')
for i,d in tqdm(enumerate(data)):
data[i] = tau.cleanText(d, lemma=lemmatizer, stops = stops, symbols_remove = True, min_charLen = 2)
print(data[0])
# Menggunakan modul SciKit untuk merubah data tidak terstruktur ke VSM
# Scikit implementation http://scikit-learn.org/stable/modules/feature_extraction.html
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
# http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html#sklearn.feature_extraction.text.CountVectorizer
# http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.TfidfVectorizer.html#sklearn.feature_extraction.text.TfidfVectorizer
# VSM - "binari"
binary_vectorizer = CountVectorizer(binary = True)
binari = binary_vectorizer.fit_transform(data)
binari.shape # ukuran VSM
# Sparse vectors/matrix
binari[0]
# Mengakses Datanya
print(binari[0].data)
print(binari[0].indices)
# Kolom dan term
print(str(binary_vectorizer.vocabulary_)[:93])
# VSM term Frekuensi : "tf"
tf_vectorizer = CountVectorizer(binary = False)
tf = tf_vectorizer.fit_transform(data)
print(tf.shape) # Sama
print(tf[0].data) # Hanya data ini yg berubah
print(tf[0].indices) # Letak kolomnya tetap sama
d = tf_vectorizer.vocabulary_
kata_kolom = {k:v for v,k in d.items()}
kata_kolom[99]
# VSM term Frekuensi : "tf-idf"
tfidf_vectorizer = TfidfVectorizer()
tfidf = tfidf_vectorizer.fit_transform(data)
print(tfidf.shape) # Sama
print(tfidf[0].data) # Hanya data ini yg berubah
print(tfidf[0].indices) # Letak kolomnya berbeda, namun jumlah kolom dan elemennya tetap sama
```
## Custom tf-idf:
* Menurut http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.TfidfVectorizer.html
* default formula tf-idf yang digunakan sk-learn adalah:
* $tfidf = tf * log(\frac{N}{df+1})$ ==> Smooth IDF
* namun kita merubahnya menjadi:
* $tfidf = tf * log(\frac{N}{df})$ ==> Non Smooth IDF
* $tfidf = tf * log(\frac{N}{df+1})$ ==> linear_tf, Smooth IDF
* $tfidf = (1+log(tf)) * log(\frac{N}{df})$ ==> sublinear_tf, Non Smooth IDF
```
# VSM term Frekuensi : "tf-idf"
tfidf_vectorizer = TfidfVectorizer(smooth_idf= False, sublinear_tf=True)
tfidf = tfidf_vectorizer.fit_transform(data)
print(tfidf.shape) # Sama
print(tfidf[0].data) # Hanya data ini yg berubah
print(tfidf[0].indices) # Letak kolomnya = tfidf
```
### Alasan melakukan filtering berdasarkan frekuensi:
* Intuitively filter noise
* Curse of Dimensionality (akan dibahas kemudian)
* Computational Complexity
* Improving accuracy
```
# Frequency Filtering di VSM
tfidf_vectorizer = TfidfVectorizer()
tfidf_1 = tfidf_vectorizer.fit_transform(data)
tfidf_vectorizer = TfidfVectorizer(max_df=0.75, min_df=5)
tfidf_2 = tfidf_vectorizer.fit_transform(data)
print(tfidf_1.shape)
print(tfidf_2.shape)
tfidf_vectorizer = TfidfVectorizer(lowercase=True, smooth_idf= True, sublinear_tf=True,
ngram_range=(1, 2), max_df=0.90, min_df=2)
tfidf_3 = tfidf_vectorizer.fit_transform(data)
print(tfidf_3.shape)
```
<h2 id="Best-Match-Formula-:-BM25">Best-Match Formula : BM25</h2>
<p><img alt="" src="images/3_bm25_simple.png" style="height: 123px; width: 300px;" /></p>
<ol>
<li>di IR nilai b dan k yang optimal adalah : <strong> <em>b</em> = 0.75 dan k = [1.2 - 2.0] </strong><br />
ref: <em>Christopher, D. M., Prabhakar, R., & Hinrich, S. C. H. Ü. T. Z. E. (2008). Introduction to information retrieval. An Introduction To Information Retrieval, 151, 177.</em></li>
<li>Tapi kalau untuk TextMining (clustering) nilai <strong>k optimal adalah 20, nilai b = sembarang (boleh = 0.75)</strong><br />
ref: <em>Whissell, J. S., & Clarke, C. L. (2011). Improving document clustering using Okapi BM25 feature weighting. Information retrieval, 14(5), 466-487.</em></li>
<li><strong>avgDL </strong>adalah rata-rata panjang dokumen di seluruh dataset dan <strong>DL </strong>adalah panjang dokumen D.<br />
hati-hati, ini berbeda dengan tf-idf MySQL diatas.</li>
</ol>
```
# Variasi pembentukan matriks VSM:
d1 = '@udin76, Minum kopi pagi-pagi sambil makan pisang goreng is the best'
d2 = 'Belajar NLP dan Text Mining ternyata seru banget sadiezz'
d3 = 'Sudah lumayan lama bingits tukang Bakso belum lewat'
d4 = 'Aduh ga banget makan Mie Ayam p4k4i kesyap, please deh'
D = [d1, d2, d3, d4]
# Jika kita menggunakan cara biasa:
tfidf_vectorizer = TfidfVectorizer()
vsm = tfidf_vectorizer.fit_transform(D)
print(tfidf_vectorizer.vocabulary_)
# N-Grams VSM
# Bermanfaat untuk menangkap frase kata, misal: "ga banget", "pisang goreng", dsb
tfidf_vectorizer = TfidfVectorizer(ngram_range=(1, 2))
vsm = tfidf_vectorizer.fit_transform(D)
print(tfidf_vectorizer.vocabulary_)
# Vocabulary based VSM
# Bermanfaat untuk menghasilkan hasil analisa yang "bersih"
# variasi 2
d1 = '@udin76, Minum kopi pagi-pagi sambil makan pisang goreng is the best'
d2 = 'Belajar NLP dan Text Mining ternyata seru banget sadiezz'
d3 = 'Sudah lumayan lama bingits tukang Bakso belum lewat seru'
d4 = 'Aduh ga banget makan Mie Ayam p4k4i kesyap, please deh'
D = [d1,d2,d3,d4]
Vocab = {'seru banget':0, 'seru':1, 'the best':2, 'lama':3, 'text mining':4, 'nlp':5, 'ayam':6}
tf_vectorizer = CountVectorizer(binary = False, vocabulary=Vocab)
tf = tf_vectorizer.fit_transform(D)
print(tf.toarray())
tf_vectorizer.vocabulary_
Vocab = {'seru banget':0, 'the best':1, 'lama':2, 'text mining':3, 'nlp':4, 'ayam':5}
tfidf_vectorizer = TfidfVectorizer(max_df=1.0, min_df=1, lowercase=True, vocabulary=Vocab)
vsm = tfidf_vectorizer.fit_transform(D)
print(tfidf_vectorizer.vocabulary_)
# VSM terurut sesuai definisi dan terkesan lebih "bersih"
# Perusahaan besar biasanya memiliki menggunakan teknik ini dengan vocabulary yang comprehensif
# Sangat cocok untuk Sentiment Analysis
```
<h2><strong>Word Embeddings</strong></h2>
<h2><img alt="" src="images/3_word_embeddings.png" style="height: 296px ; width: 602px" /></h2>
<p><img alt="" src="images/3_word2vec_example.png" style="height:400px; width:667px" /></p>
<h3>Word2Vec</h3>
<p><img alt="" src="images/3_word2Vec.png" style="height:400px; width:636px" /><br />
Dikembangkan oleh Tomas Mikolov - Google :</p>
<p>Goldberg, Yoav; Levy, Omer. "word2vec Explained: Deriving Mikolov et al.'s Negative-Sampling Word-Embedding Method". <a href="https://en.wikipedia.org/wiki/ArXiv">arXiv</a>:<a href="https://arxiv.org/abs/1402.3722">1402.3722</a> </p>
<p><img alt="" src="images/BoW_VS_WordEmbedding.png" style="width: 248px; height: 372px;" /></p>
```
data[:3]
# Rubah bentuk data seperti yang dibutuhkan genSim
# Bisa juga dilakukan dengan memodifikasi fungsi "cleanText" (agar lebih efisien)
data_we = []
for doc in data:
Tokens = [str(w) for w in TextBlob(doc).words]
data_we.append(Tokens)
print(data_we[:3])
# https://radimrehurek.com/gensim/models/word2vec.html
# train word2vec dengan data di atas
from gensim.models import Word2Vec
L = 300 # Jumlah neurons = ukuran vektor = jumlah kolom
model_wv = Word2Vec(data_we, min_count=2, size=L, window = 5, workers= -2)
# min_count adalah jumlah kata minimal yang muncul di corpus
# "size" adalah Dimensionality of the word vectors
# (menurut beberapa literature untuk text disarankan 300-500)
# "window" adalah jarak maximum urutan kata yang di pertimbangkan
# workers = jumlah prosesor yang digunakan untuk menjalankan word2vec
print('Done!...')
# di data yang sebenarnya (i.e. besar) Gensim sering membutuhkan waktu cukup lama
# Untungnya kita bisa menyimpan dan me-load kembali hasil perhitungan model word2vec, misal
model_wv.save('data/model_w2v')
model_wv = Word2Vec.load('data/model_w2v')
print('Done!...')
```
### Hati-hati, Word2vec menggunakan Matriks Dense
<p>Penggunaan memory oleh Gensim kurang lebih sebagai berikut:</p>
<p>Jumlah kata x "size" x 12 bytes</p>
<p>Misal terdapat 100 000 kata unik dan menggunakan 200 layers, maka penggunaan memory = </p>
<p>100,000x200x12 bytes = ~229MB</p>
<p>Jika jumlah size semakin banyak, maka jumlah training data yang diperlukan juga semakin banyak, namun model akan semakin akurat.</p>
```
# Melihat vector suatu kata
vektor = model_wv.wv.__getitem__(['psbb'])
print(len(vektor[0])) # Panjang vektor keseluruhan = jumlah neuron yang digunakan
print(vektor[0][:5]) # 5 elemen pertama dari vektornya
# Mencari kata terdekat menurut data training dan Word2Vec
model_wv.wv.most_similar('psbb')
# Melihat similarity antar kata
print(model_wv.wv.similarity('psbb', 'corona'))
print(model_wv.wv.similarity('psbb', 'jakarta'))
print(model_wv.wv.similarity('psbb', 'psbb'))
```
<p><img alt="" src="images/3_cosine.png" style="height:400px; width:683px" /></p>
## Hati-hati Cosine adalah similarity bukan distance
Hal ini akan mempengaruhi interpretasi
```
# error jika kata tidak ada di training data
# beckman bukan beckmans ==> hence di Word Embedding PreProcessing harus thourough
kata = 'copid'
try:
print(model_wv.wv.most_similar(kata))
except:
print('error! kata "',kata,'" tidak ada di training data')
# ini salah satu kelemahan Word2Vec
```
## Tips:
<p>Hati-hati GenSim tidak menggunakan seluruh kata di training data!.</p>
<p>Perintah berikut akan menghasilkan kata-kata yang terdapat di vocabulary GenSim</p>
```
Vocabulary = model_wv.wv.vocab
print(str(Vocabulary.keys())[:250])
# Gunakan vocabulary ini (rubah ke "set") untuk membuat program menjadi lebih robust
```
## Hati-hati menginterpretasikan hasil Word2Vec
<h3 id=" FastText-(Facebook-2016)"> FastText (Facebook-2016)</h3>
<ul>
<li>Menggunakan Sub-words: app, ppl, ple - apple</li>
<li>Paper: https://arxiv.org/abs/1607.04606 </li>
<li>Website: https://fasttext.cc/</li>
<li>Source: https://github.com/facebookresearch/fastText </li>
</ul>
```
# Caution penggunaan memory besar, bila timbul "Memory Error" kecilkan nilai L
from gensim.models import FastText
L = 100 # Jumlah neurons = ukuran vektor = jumlah kolom
model_FT = FastText(data_we, size=L, window=5, min_count=2, workers=-2)
'Done'
# Mencari kata terdekat menurut data training dan Word2Vec
model_FT.wv.most_similar('psbb')
# Melihat similarity antar kata
print(model_FT.wv.similarity('psbb', 'corona'))
print(model_FT.wv.similarity('psbb', 'jakarta'))
print(model_FT.wv.similarity('psbb', 'psbb'))
# Word2Vec VS FastText
try:
print(model_wv.wv.most_similar('coro'))
except:
print('Word2Vec error!')
try:
print(model_FT.wv.most_similar('coro'))
except:
print('FastText error!')
```
# Diskusi:
<ul>
<li>Apakah kelebihan dan kekurangan WE secara umum?</li>
<li>Apakah kira-kira aplikasi WE?</li>
<li>Apakah bisa dijadikan representasi dokumen? Bagaimana caranya?</li>
<li>Bergantung pada apa sajakah performa model WE?</li>
</ul>
* Preprocessing apa yang sebaiknya dilakukan pada model Word Embedding?
* Apakah Pos Tag bermanfaat disini? Jika iya bagaimana menggunakannya?
<h1>End of Module UDA-06</h1>
<hr />
<p><img alt="" src="images/2_Studying_Linguistic.png" style="height:500px; width:667px" /></p>
| github_jupyter |
# Fussing with Healpix
```
#
import os
import numpy as np
import healpy as hp
import pandas
from matplotlib import pyplot as plt
from scipy import interpolate
import iris
%matplotlib notebook
noaa_path = '/home/xavier/Projects/Oceanography/data/SST/NOAA-OI-SST-V2/'
def set_fontsize(ax,fsz):
'''
Parameters
----------
ax : Matplotlib ax class
fsz : float
Font size
'''
for item in ([ax.title, ax.xaxis.label, ax.yaxis.label] +
ax.get_xticklabels() + ax.get_yticklabels()):
item.set_fontsize(fsz)
```
# Load one
```
file_1990 = os.path.join(noaa_path, 'sst.day.mean.1990.nc')
cubes_1990 = iris.load(file_1990)
sst_1990 = cubes_1990[0]
sst_1990
# Realize
_ = sst_1990.data
```
## Grab a day, any day
```
jan1_1990 = sst_1990.data[0,:,:]
jan1_1990.shape
```
## Coords
```
lat_coord = sst_1990.coord('latitude').points
lon_coord = sst_1990.coord('longitude').points
lat_coord[0:5]
```
# Init Healpix
## nside
```
npix = np.product(jan1_1990.shape)
npix
nside_approx = np.sqrt(npix/12.)
nside_approx
nside = 294
npix_hp = hp.nside2npix(nside)
npix_hp
print(
"Approximate resolution at NSIDE {} is {:.2} deg".format(
nside, hp.nside2resol(nside, arcmin=True) / 60
)
)
```
## Init
```
m = np.arange(npix_hp)
hp.mollview(m, title='Mollview image Ring')
hp.graticule()
```
## Angles (deg)
```
theta, phi = np.degrees(hp.pix2ang(nside=nside, ipix=np.arange(npix_hp))) #[0, 1, 2, 3, 4]))
theta.size
np.max(theta)
```
# Interpolate
## Mesh me
```
lat_mesh, lon_mesh = np.meshgrid(lat_coord, lon_coord, indexing='ij')
lat_mesh.shape
jan1_1990.shape
jan1_1990.fill_value = -10.
jan1_1990
```
## Interpolate
```
func = interpolate.RectBivariateSpline(lat_coord, lon_coord, jan1_1990.filled(), kx=2, ky=2)#, epsilon=2)
new_SST = func.ev(90-theta, phi)
new_SST.shape
```
## Plot
```
#m = np.arange(npix_hp)
hp.mollview(new_SST, title='Mollview image Ring', max=30., min=0.)
hp.graticule()
```
# Harmonics
## Mask
```
goodpix = new_SST > -5.
np.mean(new_SST[goodpix])
```
## Remove the monopole
```
diff_SST = new_SST - np.mean(new_SST[goodpix])
diff_SST[~goodpix] = 0.
```
## Calculate
```
cls = hp.sphtfunc.anafast(diff_SST)
cls.size
```
## Plot
```
ls = np.arange(cls.size)
plt.clf()
ax = plt.gca()
ax.plot(ls, ls*(ls+1)*cls, drawstyle='steps-mid')
#
ax.set_xscale('log')
ax.set_xlim(1.,1000)
ax.set_xlabel('Multipole moment l')
ax.set_ylabel(r'$l(l+1)c_l$')
set_fontsize(ax,17)
plt.show()
```
# Save
```
# Write
t = Table()
t['flux'] = DM_tot # the data array
t.meta['ORDERING'] = 'RING'
t.meta['COORDSYS'] = 'G'
t.meta['NSIDE'] = 1024
t.meta['INDXSCHM'] = 'IMPLICIT'
t.write(hp_file, overwrite=True)
```
| github_jupyter |
# DS1000E Rigol Waveform Examples
**Scott Prahl**
**March 2020**
```
import sys
import numpy as np
import matplotlib.pyplot as plt
try:
import RigolWFM.wfm as rigol
except:
print("***** You need to install the module to read Rigol files first *****")
print("***** Execute the following line in a new cell, then retry *****")
print()
print("!{sys.executable} -m pip install RigolWFM")
```
## Introduction
This notebook illustrates shows how to extract signals from a `.wfm` file created by a the Rigol DS1000E scope. It also validates that the process works by comparing with `.csv` and screenshots.
Two different `.wfm` files are examined one for the DS1052E scope and one for the DS1102E scope. The accompanying `.csv` files seem to have t=0 in the zero in the center of the waveform.
The list of Rigol scopes that should produce the same file format are:
```
print(rigol.DS1000E_scopes[:])
```
## DS1052E
We will start with a `.wfm` file from a Rigol DS1052E scope. This test file accompanies [wfm_view.exe](http://meteleskublesku.cz/wfm_view/) a freeware program from <http://www.hakasoft.com.au>.
The waveform looks like
<img src="https://github.com/scottprahl/RigolWFM/raw/master/wfm/DS1052E.png" width="50%">
Now let's look at plot of the data from the corresponding `.csv` file created by [wfm_view.exe](http://meteleskublesku.cz/wfm_view/)
```
csv_filename_52 = "https://raw.githubusercontent.com/scottprahl/RigolWFM/master/wfm/DS1052E.csv"
csv_data = np.genfromtxt(csv_filename_52, delimiter=',', skip_header=19, skip_footer=1, encoding='latin1').T
center_time = csv_data[0][-1]*1e6/2
plt.subplot(211)
plt.plot(csv_data[0]*1e6,csv_data[1], color='green')
plt.title("DS1052E from .csv file")
plt.ylabel("Volts (V)")
plt.xlim(center_time-0.6,center_time+0.6)
plt.xticks([])
plt.subplot(212)
plt.plot(csv_data[0]*1e6,csv_data[2], color='red')
plt.xlabel("Time (µs)")
plt.ylabel("Volts (V)")
plt.xlim(center_time-0.6,center_time+0.6)
plt.show()
```
### Now for the `.wfm` data
First a textual description.
```
help(rigol.Wfm.from_file)
# raw=true is needed because this is a binary file
#wfm_url = "https://github.com/scottprahl/RigolWFM/raw/master/wfm/DS1052E.wfm" + "?raw=true"
#w = rigol.Wfm.from_url(wfm_url, '1000E')
wfm_file = "/Users/prahl/Documents/Code/git/RigolWFM/wfm/DS1102E-D.wfm"
w = rigol.Wfm.from_file(wfm_file, 'E')
description = w.describe()
print(description)
ch = w.channels[0]
plt.subplot(211)
plt.plot(ch.times*1e3, ch.volts, color='green')
plt.title("DS1052E from .wfm file")
plt.ylabel("Volts (V)")
plt.xlim(-0.6,0.6)
plt.xticks([])
ch = w.channels[1]
plt.subplot(212)
plt.plot(ch.times*1e3, ch.volts, color='red')
plt.xlabel("Time (ms)")
plt.ylabel("Volts (V)")
plt.xlim(-0.6,0.6)
plt.show()
```
## DS1102E-B
### First the `.csv` data
This file only has one active channel. Let's look at what the accompanying `.csv` data looks like.
```
csv_filename = "https://github.com/scottprahl/RigolWFM/raw/master/wfm/DS1102E-B.csv"
my_data = np.genfromtxt(csv_filename, delimiter=',', skip_header=2).T
plt.plot(my_data[0]*1e6, my_data[1])
plt.xlabel("Time (µs)")
plt.ylabel("Volts (V)")
plt.title("DS1102E-B with a single trace")
plt.show()
```
### Now for the `wfm` data
First let's have look at the description of the internal file structure. We see that only channel 1 has been enabled.
```
# raw=true is needed because this is a binary file
wfm_url = "https://github.com/scottprahl/RigolWFM/raw/master/wfm/DS1102E-B.wfm" + "?raw=true"
w = rigol.Wfm.from_url(wfm_url, 'DS1102E')
description = w.describe()
print(description)
w.plot()
plt.xlim(-6,6)
plt.show()
```
## DS1102E-E
[Contributed by @Stapelberg](https://github.com/scottprahl/RigolWFM/issues/11#issue-718562669)
This file uses a 10X probe. First let's have look at the description of the internal file structure. We see that only channel 1 has been enabled and it has a 10X probe.
```
# raw=true is needed because this is a binary file
wfm_url = "https://github.com/scottprahl/RigolWFM/raw/master/wfm/DS1102E-E.wfm" + "?raw=true"
w = rigol.Wfm.from_url(wfm_url, 'DS1102E')
description = w.describe()
print(description)
w.plot()
#plt.xlim(-6,6)
plt.show()
```
| github_jupyter |
# "[DL] Build Resnet from scratch using Pytorch"
> "Understanding the architecture of Resnet easily by implementing it with Pytorch"
- toc:false
- branch: master
- badges: false
- comments: true
- author: Peiyi Hung
- categories: [category, learning]
- image: "images/skip conncetion.png"
# Introduction
Resnet is the most used model architecture in computer vision introduced by Kaiming He et al. in the article ["Deep Residual Learning for Image Recognition"](https://arxiv.org/abs/1512.03385). In order to understand this important model, I read the paper and several Deep Learning books about it. However, I have difficulty understanding the architecture thoroughly by just reading these materials. Therefore I decide to implement it by myself with Pytorch.
The core problem Resnet intends to solve is that **a deeper model can perform worse than a shallow one not because of overfitting, but just becuase of it being deeper**. The Resnet paper found that a deeper model gets worse results on both training set and test set than a shallow model. Since the deeper performs badly in both training set and test set, this is obviously not due to overfitting. The paper indicates that not all systems are similarly easy to optimize. A deeper model might be more powerful but harder to train. To solve this problem, the paper proposed a special architecture, **residual block**, to solve this problem.
The main characteristic of resicual block is **the shortcut connections**. It enables the neural network to skip several layers easily by setting some parameters to zero. In addition, the paper designed **bottleneck connections** which allows us to train a deeper model with similar time complexity as a network with residual blocks by adopting $1 \times 1$ convolutional layers. These two types of networks architectures would be explained and implemented by Pytorch in later sections.
In order to see if our model work, I train the models we build to conduct an image classification task: distinguishing between cats and dogs using [the Oxford-IIIT pet dataset](https://www.robots.ox.ac.uk/~vgg/data/pets/).
This post would first gather the pets data and then build models. Two types of model would be built: one type with residual blocks (Resnet18 and Resnet34), and the other with bottleneck blocks (Resnet50, Resnet101, and Resnet152). Specifically, I would build Resnet34 and Resnet50, and other models are simply generalization of those models. All the model would be created with Pytorch and trained with fastai.
Let's get started.
# Gather the Data
Since I would like train models to distinguish between cats and dogs, I need a datasets containing images of cats and dogs. I use fastai's `ImageDataLoaders` to gather the data.
```
from fastai.vision.all import *
img_path = untar_data(URLs.PETS)
images_fnames = get_image_files(img_path, folders="images")
is_cat = lambda x: x.name[0].isupper()
dls = ImageDataLoaders.from_path_func(img_path, images_fnames,
label_func=is_cat,
item_tfms=Resize(224),
batch_tfms=Normalize)
```
Here's how `ImageDataLoaders` gather the images:
* `img_path` tells `ImageDataLoaders` where our images are stored. `untar_data(URLs.PETS)` would download the data and store it on your local machine.
* `get_image_files` find all the images files. Since all images are stored in the "images" folder, we pass `folders="images"` into the function.
* All file names of cats images start with a capital letter (names of dogs images do not), so we make a function `is_cat` to tell `ImageDataLoaders` the label of each image.
* I resize all the images into size of $224 \times 224$ and normalize them using `item_tfms=Resize(224)` and `batch_tfms=Normalize`.
Let's take a look at our training data:
```
dls.show_batch()
```
Once our data is prepared, we can start to build our model.
# ResNet34
The overall structure of a Resnet is `stem` + `multiple Residual Blocks` + `global average pooling` + `classifier`. (See the struture in Pytorch code in the function `get_resnet`)
Here's an overview of how each part of Resnet works:
* `stem` is a convolutional layer with large kernel size (7 in Resnet) to downsize the image size immediately from the beginning.
* Mutiple residual block with different sizes of channels would be connected after `stem`. The detail of residual blocks would be discussed in the next part.
* `global average pooling` downsize a $k \times k$ feature map of each channel into a $1 \times 1$ feature map.
* These $1 \times 1$ feature maps would be flattened as the features for a fully connected layer that serves as a `classifier`.
Next, let's discuss the residual block.
## Residual Block

Let's say `x` is our input of a residual block. The shape of `x` would usually be `[N, C, H, W]` where `N` is the batch size, `C` is the channel size or the feature size, and `H x W` would be the size of feature map. There are two path `x` would go through: one contains two convolutional layers and the other is a identiy path. The convitional path is composed of two $3 \times 3$ convolutional layers and the identical path is just returning what it takes in. After `x` went through these path, they would be add together and passed into a ReLU function.
There is one potential problem: if we want to change size of channels or downsize the feature maps, we can not add the output from these two path together. Here's how we solvee this problem:
* We change the size of channel by $1 \times 1$ convolutional layers.
* We downsize the images by setting the stride of the first convolutional layer as 2.
Here's how a residual block would be implemented in Pytorch code:
```
class ResidualBlock(nn.Module):
def __init__(self, ni: int, nf: int):
super().__init__()
# shorcut
if ni < nf:
# change channel size
self.shortcut = nn.Sequential(
nn.Conv2d(ni, nf, kernel_size=1, stride=2),
nn.BatchNorm2d(nf))
# downsize the feature map
first_stride = 2
else:
self.shortcut = lambda x: x
first_stride = 1
# convnet
self.conv = nn.Sequential(
nn.Conv2d(ni, nf, kernel_size=3, stride=first_stride, padding=1),
nn.BatchNorm2d(nf),
nn.ReLU(True),
nn.Conv2d(nf, nf, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(nf)
)
def forward(self, x):
return F.relu(self.conv(x) + self.shortcut(x))
```
We can clearly see the two path: `self.conv(x)` and `self.shortcut(x)`. `self.conv` is composition of two convolutional layer. The stride of first convolutional layer would change when we want to downsize the feature mape. `self.shortcut` would return `x` itself when we use the same channel size, whereas it would become a $1 \times 1$ convolutional layers when we want to increase the feature size.
Let's see the structure of the whole Resnet34 in Pytorch code:
```
# we should train the model with GPU if available.
device = "cuda" if torch.cuda.is_available() else "cpu"
def get_resnet():
return nn.Sequential(
nn.Conv2d(3, 64, 7, stride=2, padding=3), # stem
nn.BatchNorm2d(64),
nn.ReLU(),
nn.MaxPool2d(3, stride=2, padding=1),
*[ResidualBlock(64, 64)] * 3, # residual blocks
ResidualBlock(64, 128),
*[ResidualBlock(128, 128)] * 3,
ResidualBlock(128, 256),
*[ResidualBlock(256, 256)] * 5,
ResidualBlock(256, 512),
*[ResidualBlock(512, 512)] * 2,
nn.AdaptiveAvgPool2d(1), # global average pooling
nn.Flatten(),
nn.Linear(512, 2) # classifier
).to(device)
resnet34_model = get_resnet()
```
To see if we make some mistakes while building the model, let's train the model for 10 epoch.
```
learn = Learner(dls, resnet34_model, loss_func=nn.CrossEntropyLoss(), metrics=accuracy)
learn.fit(10, 1e-4)
```
We succeed to train the model and get no error while training.
**Get Resnet18**
The difference between Resnet18 and Resnet34 is just the layers of `ResidualBlock`. Therefore, we can obtain Resnet18 by simply modifying the number of `ResidualBlock`.
```
def get_resnet(n_layers=[3, 3, 5, 2]):
'''
n_layer: list of numbers of residual blocks.
the default [3, 3, 5, 2] is for Resnet34
'''
return nn.Sequential(
nn.Conv2d(3, 64, kernel_size=7, stride=2, padding=3),
nn.BatchNorm2d(64),
nn.ReLU(),
nn.MaxPool2d(3, stride=2, padding=1),
*[ResidualBlock(64, 64)] * n_layers[0],
ResidualBlock(64, 128),
*[ResidualBlock(128, 128)] * n_layers[1],
ResidualBlock(128, 256),
*[ResidualBlock(256, 256)] * n_layers[2],
ResidualBlock(256, 512),
*[ResidualBlock(512, 512)] * n_layers[2],
nn.AdaptiveAvgPool2d(1),
nn.Flatten(),
nn.Linear(512, 2)
).to(device)
```
By passing `[2, 1, 1, 1]` into the function, we can get Resnet18.
```
resent18_model = get_resnet([2, 1, 1, 1])
```
# Resnet50
The overall structure of Resnet50 is similar to the one of Resnet34. To obtain a Resnet50 model, we can simply replace the residual block by bottleneck block.
## Bottleneck block

Like residual block, a bottleneck block has a convolutional path and a shortcut path. However, the structure of convolutional path is different in bottleneck block. It consists of three convolutional layer: two $1 \times 1$ conv layers and one $3 \times 3$ convolutional layer in the middle. The size of channels is also different from a residual block as shown in the graph above.
This bottleneck can allow our model to use more nonlinear activation to learn the underelying patterens from the data and maintain almost the same time complexity as a residual block. Resnet50 contains same amount of blocks as Reesnet34. Since Resnet uses bottleneck layers, it has more learnable layers (increasing from 34 to 50).
Here's how a bottleneck block works in Pytorch code:
```
class BottleneckBlock(nn.Module):
def __init__(self, ni: int, nf: int):
super().__init__()
# shortcut
if ni < nf:
stride = 1 if ni == 64 else 2
self.shortcut = nn.Sequential(
nn.Conv2d(ni, nf, kernel_size=1, stride=stride),
nn.BatchNorm2d(nf)
)
else:
self.shortcut = lambda x: x
stride = 1
# convnet
self.conv = nn.Sequential(
nn.Conv2d(ni, nf//4, kernel_size=1, stride=1),
nn.BatchNorm2d(nf//4),
nn.ReLU(True),
nn.Conv2d(nf//4, nf//4, kernel_size=3, stride=stride, padding=1),
nn.BatchNorm2d(nf//4),
nn.ReLU(True),
nn.Conv2d(nf//4, nf, kernel_size=1, stride=1),
nn.BatchNorm2d(nf)
)
def forward(self, x):
return F.relu(self.conv(x) + self.shortcut(x))
```
As in the previous section, we connect all the element to constrcut a Resnet50 and train it to see if it can really work.
```
resnet50_model = nn.Sequential(
nn.Conv2d(3, 64, kernel_size=7, stride=2, padding=3),
nn.BatchNorm2d(64),
nn.ReLU(),
nn.MaxPool2d(3, stride=2, padding=1),
BottleneckBlock(64, 256),
*[BottleneckBlock(256, 256)] * 2,
BottleneckBlock(256, 512),
*[BottleneckBlock(512, 512)] * 3,
BottleneckBlock(512, 1024),
*[BottleneckBlock(1024, 1024)] * 5,
BottleneckBlock(1024, 2048),
*[BottleneckBlock(2048, 2048)] * 2,
nn.AdaptiveAvgPool2d(1),
nn.Flatten(),
nn.Linear(2048, 2)
).to(device)
learn = Learner(dls, resnet50_model, loss_func=nn.CrossEntropyLoss(), metrics=accuracy)
learn.fit(10, 1e-4)
```
There are two deeper variant of Resnet: Resnet101 and Resnt152. These two variants can be created by simply increasing the number of bottleneck blocks as followed:
```
def get_resnet_bottleneck(n_layer=[2, 3, 5, 2], n_classes=2):
return nn.Sequential(
nn.Conv2d(3, 64, kernel_size=7, stride=2, padding=3),
nn.BatchNorm2d(64),
nn.ReLU(),
nn.MaxPool2d(3, stride=2, padding=1),
BottleneckBlock(64, 256),
*[BottleneckBlock(256, 256)] * n_layer[0],
BottleneckBlock(256, 512),
*[BottleneckBlock(512, 512)] * n_layer[1],
BottleneckBlock(512, 1024),
*[BottleneckBlock(1024, 1024)] * n_layer[2],
BottleneckBlock(1024, 2048),
*[BottleneckBlock(2048, 2048)] * n_layer[3],
nn.AdaptiveAvgPool2d(1),
nn.Flatten(),
nn.Linear(2048, n_classes)
)
resnet101_model = get_resnet_bottleneck([2, 3, 22, 2])
resnet152_model = get_resnet_bottleneck([2, 7, 35, 2])
```
# Summary
| github_jupyter |
# Accessing geoprocessing tools
In this page we will observe how you can search for geoprocessing tools and access them in your code. Thus, we will observe:
- [Searching for geoprocessing tools](#searching-for-geoprocessing-tools)
- [Pythonic representation of tools and toolboxes](#pythonic-representation-of-tools-and-toolboxes)
- [Importing toolboxes](#importing-toolboxes)
- [Importing toolbox from an `Item`](#importing-toolbox-from-an-item)
- [Importing toolbox from a geoprocessing service URL](#importing-toolbox-from-a-geoprocessing-service-url)
- [Tool signature, parameters and documentation](#tool-signature-parameters-and-documentation)
<a id="searching-for-geoprocessing-tools"></a>
## Searching for geoprocessing tools
Geoprocessing tools can be considered as web tools that can shared with others. Users organize their tools into toolboxes and share with on their GIS. You can search for geoprocessing tools just like you search for any other item.
To search your GIS for geoprocessing toolboxes, specify `Geoprocessing Toolbox` as the item type.
# Geoprocessing
```
import arcgis
from arcgis.gis import GIS
from IPython.display import display
gis = GIS('https://www.arcgis.com', 'arcgis_python', 'P@ssword123')
toolboxes = gis.content.search('travel', 'Geoprocessing Toolbox',
outside_org=True, max_items=3)
for toolbox in toolboxes:
display(toolbox)
```
<a id="pythonic-representation-of-tools-and-toolboxes"></a>
## Pythonic representation of tools and toolboxes
In ArcGIS API for Python, geoprocessing toolboxes are represented as Python modules and the individual tools as Python functions. These tools, represented as Python functions, take in a set of input parameters and return one or more output values.
To use custom geoprocessing tools, users simply import that toolbox as a module in their programs and call functions within the module.
<a id="importing-toolboxes"></a>
## Importing toolboxes
The `import_toolbox()` function in the `arcgis.geoprocessing` module imports geoprocessing toolboxes as native Python modules. It accepts a toolbox location which could be a Geoprocessing Toolbox item in your GIS, or a URL to a Geoprocessing Service.
Developers can then call the functions available in the imported module to invoke these tools. Let us see how to import toolboxes from these two sources.
<a id="importing-toolbox-from-an-item"></a>
### Importing toolbox from an `Item`
The code snippet below shows how the Ocean Currents toolbox above can be imported as a module:
```
from arcgis.geoprocessing import import_toolbox
ocean_currents_toolbox = toolboxes[1]
ocean_currents_toolbox
ocean_currents = import_toolbox(ocean_currents_toolbox)
```
The `import_toolbox()` function inspects the geoprocessing toolbox and dynamically generates a Python module containing a function for each tool within the toolbox. Invoking the function invokes the corresponding geoprocessing tool.
The code snippet below uses Python's `inspect` module to list the public functions in the imported module. Developers will typically use their IDE's intellisense to discover the functions in the module.
```
ocean_currents.message_in_a_bottle?
import inspect
# list the public functions in the imported module
[ f[0] for f in inspect.getmembers(ocean_currents, inspect.isfunction)
if not f[0].startswith('_')]
```
<a id="importing-toolbox-from-a-geoprocessing-service-url"></a>
### Importing toolbox from a geoprocessing service URL
```
zion_toolbox_url = 'http://gis.ices.dk/gis/rest/services/Tools/ExtractZionData/GPServer'
zion = import_toolbox(zion_toolbox_url)
```
<a id="tool-signature-parameters-and-documentation"></a>
## Tool signature, parameters and documentation
The function for invoking the geoprocessing tool includes documentation about that tool. This doc shows up using your IDE's intellisense and can also be accessed using Python's help function:
```
help(zion.extract_zion_data)
result = zion.extract_zion_data()
type(result)
result.download()
```
As shown in the example above, tool functions are annotated using [type hints](https://www.python.org/dev/peps/pep-0484/) to help indicate the input they accept and the output they produce. The function signature includes default values for the input parameters, so the caller doesn't have to specify them unless required. Parameter documentation includes a description of each parameter, it's expected type, and whether it is required or optional. If the parameter accepts from a list of input values, that list is included with the documentation as a 'Choice List'. The documentation includes the type and description of the functions return value.
Next, head over to the topic [Using geoprocessing tools](/python/guide/using-geoprocessing-tools/) to see how these tools can be used in Python scripts.
| github_jupyter |
```
%matplotlib notebook
import numpy as np
import scipy.io.wavfile as wavfile
import matplotlib
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = [12, 8]
plt.rcParams['savefig.dpi'] = 80
plt.rcParams['figure.dpi'] = 80
#32-bit floating-point [-1.0,+1.0] np.float32
#from scipy.io.wavfile import write
#write('test.wav', int(fs), np.real(chirp).astype(np.float32))
#https://howthingsfly.si.edu/ask-an-explainer/how-does-speed-sound-air-vary-temperature
T_F=61
target_distance_feet=20
T_C=(T_F-32)*5/9;
speed_of_sound_in_air_m_per_sec=331+0.6*T_C
meter_to_feet=3.281;
target_distance_m=target_distance_feet/meter_to_feet;
round_trip_distance_m=2*target_distance_m;
#Tc is the max depth we are interested in
Tc_sec=1
f_start_Hz=3e3
f_stop_Hz=20e3
S_Hz_per_sec = (f_stop_Hz - f_start_Hz) / Tc_sec
original_chirp_filename='../data/audio_chirp_1sec_3KHz_20KHz.wav'
(original_chirps_fs, original_chirp_data)=wavfile.read(original_chirp_filename)
original_chirp_data.shape
collect_filename='../data/house_bose_front_20ft.wav'
(collect_fs, collect_data)=wavfile.read(collect_filename)
collect_data.shape
chirp_corr=np.correlate(original_chirp_data, collect_data[:,0])
chirp_corr.shape[0]
#For normalized cross correlation compute the energy a each position
segment_length=original_chirp_data.shape[0]
energy_level=np.zeros((collect_data.shape[0]-segment_length,1))
for ii in np.arange(0,energy_level.shape[0]):
energy_level[ii,0]=np.sum(np.absolute(collect_data[ii:(ii+segment_length-1),0]))
t_sec=np.arange(0,chirp_corr.shape[0],1)/collect_fs
d_m=speed_of_sound_in_air_m_per_sec*t_sec;
chirp_corr=np.correlate(original_chirp_data, collect_data[:,0])
print(repr(np.squeeze(chirp_corr[0:1607901]).shape))
print(repr(np.squeeze(energy_level[0:1607901]).shape))
fig = plt.figure(figsize=(10,7))
#subject_data = su.get_subject_data(root_directory, subject=1, trials=trials)
plt.plot(d_m[0:-1],np.log10(np.abs(np.squeeze(chirp_corr[0:1607901]))),label=collect_filename)
plt.xlabel('distance (m)')
plt.ylabel('Corr value')
plt.legend()
plt.title(f'Correlation between chirp and collect with round trip target distance of {round_trip_distance_m} m')
fig = plt.figure(figsize=(7,6))
#subject_data = su.get_subject_data(root_directory, subject=1, trials=trials)
plt.plot(d_m[0:-1],(np.abs((np.divide(np.squeeze(chirp_corr[0:1607901]),np.squeeze(energy_level[0:1607901]))))),label=collect_filename)
plt.xlabel('distance (m)')
plt.ylabel('pseudo crosscorrelation value')
plt.title(f'Correlation between chirp and collect \nwith round trip target distance of {round_trip_distance_m:.1f} m')
fig = plt.figure(figsize=(10,7))
#subject_data = su.get_subject_data(root_directory, subject=1, trials=trials)
plt.plot(d_m[0:-1],np.log10(np.abs((np.divide(np.squeeze(chirp_corr[0:1607901]),np.squeeze(energy_level[0:1607901]))))),label=collect_filename)
plt.xlabel('distance (m)')
plt.ylabel('Energy normalized corr value (dB)')
plt.title(f'Correlation between chirp and collect with round trip target distance of {round_trip_distance_m:.1f} m')
fig = plt.figure(figsize=(10,7))
#subject_data = su.get_subject_data(root_directory, subject=1, trials=trials)
plt.plot(d_m,chirp_corr,label=collect_filename)
plt.xlabel('distance (m)')
plt.ylabel('Corr value')
plt.legend()
plt.title(f'Correlation between chirp and collect with round trip target distance of {round_trip_distance_m} m')
NFFT = 512 # the length of the windowing segments
Fs = collect_fs # the sampling frequency
t_sec=np.arange(0,collect_data.shape[0],1)/collect_fs
d_m=speed_of_sound_in_air_m_per_sec*t_sec;
fig, (ax1, ax2) = plt.subplots(nrows=2)
ax1.plot(t_sec, collect_data[:,0])
ax1.set_xlabel('time (sec)')
ax1.set_ylabel('amplitude (sample)')
Pxx, freqs, bins, im = ax2.specgram(collect_data[:,0], NFFT=NFFT, Fs=Fs, noverlap=500)
#fig.colorbar(im)
im.set_clim([-40,5])
ax2.set_xlabel('time (sec)')
ax2.set_ylabel('frequency (Hz)')
plt.show()
NFFT = 512*2 # the length of the windowing segments
Fs = collect_fs # the sampling frequency
t_sec=np.arange(0,collect_data.shape[0],1)/collect_fs
d_m=speed_of_sound_in_air_m_per_sec*t_sec;
fig, (ax1) = plt.subplots(nrows=1)
Pxx, freqs, bins, im = ax1.specgram(collect_data[:,0], NFFT=NFFT, Fs=Fs, noverlap=int(NFFT*0.9))
#fig.colorbar(im)
im.set_clim([-10,20])
ax1.set_xlabel('time (sec)')
ax1.set_ylabel('frequency (Hz)')
plt.show()
NFFT = 512 # the length of the windowing segments
Fs = collect_fs # the sampling frequency
t_sec=np.arange(0,collect_data.shape[0],1)/collect_fs
fig, (ax2) = plt.subplots(nrows=1)
Pxx, freqs, bins, im = ax2.specgram(collect_data[:,0], NFFT=NFFT, Fs=Fs, noverlap=500)
#fig.colorbar(im)
im.set_clim([-40,5])
ax2.set_xlabel('time (sec)')
ax2.set_ylabel('frequency (Hz)')
plt.show()
NFFT = 512 # the length of the windowing segments
Fs = original_chirps_fs # the sampling frequency
t_sec=np.arange(0,original_chirp_data.shape[0],1)/Fs
fig, (ax2) = plt.subplots(nrows=1)
Pxx, freqs, bins, im = ax2.specgram(original_chirp_data, NFFT=NFFT, Fs=Fs, noverlap=500)
#fig.colorbar(im)
im.set_clim([-40,5])
ax2.set_xlabel('time (sec)')
ax2.set_ylabel('frequency (Hz)')
plt.show()
true_target_distance_round_trip_m = 12.2
true_target_time_sec = true_target_distance_round_trip_m/speed_of_sound_in_air_m_per_sec
true_target_freq_Hz = true_target_time_sec*S_Hz_per_sec
print(f'true target_freq_Hz = {true_target_freq_Hz} Hz and target_time_sec = {true_target_time_sec} sec' )
measured_target_time_sec = 15.0467-15.0128
measured_target_freq_Hz = measured_target_time_sec*S_Hz_per_sec
print(f'measured target_freq_Hz = {measured_target_freq_Hz} Hz and target_time_sec = {measured_target_time_sec} sec' )
measured_target_distance_rount_trip_m = speed_of_sound_in_air_m_per_sec*measured_target_time_sec
print(f'measured_target_distance_rount_trip_m = {measured_target_distance_rount_trip_m} m' )
speed_of_sound_in_air_m_per_sec/(17e3*2)*600
speed_of_sound_in_air_m_per_sec*0.032
```
| github_jupyter |
<table width="100%"> <tr>
<td style="background-color:#ffffff;">
<a href="http://qworld.lu.lv" target="_blank"><img src="../images/qworld.jpg" width="35%" align="left"> </a></td>
<td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
prepared by Abuzer Yakaryilmaz (<a href="http://qworld.lu.lv/index.php/qlatvia/" target="_blank">QLatvia</a>)
<br>
updated by Melis Pahalı | December 5, 2019
<br>
updated by Özlem Salehi | September 17, 2020
</td>
</tr></table>
<table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
$ \newcommand{\bra}[1]{\langle #1|} $
$ \newcommand{\ket}[1]{|#1\rangle} $
$ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
$ \newcommand{\dot}[2]{ #1 \cdot #2} $
$ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
$ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
$ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
$ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
$ \newcommand{\mypar}[1]{\left( #1 \right)} $
$ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
$ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
$ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
$ \newcommand{\onehalf}{\frac{1}{2}} $
$ \newcommand{\donehalf}{\dfrac{1}{2}} $
$ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
$ \newcommand{\vzero}{\myvector{1\\0}} $
$ \newcommand{\vone}{\myvector{0\\1}} $
$ \newcommand{\vhadamardzero}{\myvector{ \sqrttwo \\ \sqrttwo } } $
$ \newcommand{\vhadamardone}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
$ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
$ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
$ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
$ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
$ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
$ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
<h2>Quantum Teleportation</h2>
Asja wants to send a qubit to Balvis by using only classical communication.
Let $ \ket{v} = \myvector{a\\ b} \in \mathbb{R}^2 $ be the quantum state of Asja's qubit.
If Asja has many copies of this qubit, then she can collect the statistics based on these qubits and obtain an approximation of $ a $ and $ b $, say $ \tilde{a} $ and $\tilde{b}$, respectively. After this, Asja can send $ \tilde{a} $ and $\tilde{b}$ by using many classical bits, the number of which depends on the precision of the amplitudes.
On the other hand, If Asja and Balvis share the entangled qubits in state $ \sqrttwo\ket{00} + \sqrttwo\ket{11} $ in advance, then it is possible for Balvis to create $ \ket{v} $ in his qubit after receiving two bits of information from Asja.
<h3>What is quantum teleportation?</h3>
It is the process of transmission of quantum information, that is the state of a qubit, using classical communication and previously entangled qubits.
The state of a qubit is transfered onto another qubit, while the state of the source qubit is destroyed.
Note that we never obtain multiple copies of the same qubit - "No Cloning Theorem".
<a href = "https://www.nature.com/news/quantum-teleportation-is-even-weirder-than-you-think-1.22321">Read more </a>
<h3> Protocol </h3>
The protocol uses three qubits as specified below:
<img src='../images/quantum_teleportation_qubits.png' width="25%" align="left">
Asja has two qubits and Balvis has one qubit.
Asja wants to send her first qubit which is in state $ \ket{v} = \myvector{a\\b} = a\ket{0} + b\ket{1} $.
Asja's second qubit and Balvis' qubit are entangled. The quantum state of Asja's second qubit and Balvis' qubit is $ \sqrttwo\ket{00} + \sqrttwo\ket{11} $.
So, the state of the three qubits is
$$ \mypar{a\ket{0} + b\ket{1}}\mypar{\sqrttwo\ket{00} + \sqrttwo\ket{11}}
= \sqrttwo \big( a\ket{000} + a \ket{011} + b\ket{100} + b \ket{111} \big). $$
<h4> CNOT operator by Asja </h4>
Asja applies CNOT gate to her qubits where her first qubit is the control qubit and her second qubit is the target qubit.
<h3>Task 1</h3>
Calculate the new quantum state after this CNOT operator.
<a href="B54_Quantum_Teleportation_Solutions.ipynb#task1">click for our solution</a>
<h3>Hadamard operator by Asja</h3>
Asja applies Hadamard gate to her first qubit.
<h3>Task 2</h3>
Calculate the new quantum state after this Hadamard operator.
Verify that the resulting quantum state can be written as follows:
$$
\frac{1}{2} \ket{00} \big( a\ket{0}+b\ket{1} \big) +
\frac{1}{2} \ket{01} \big( a\ket{1}+b\ket{0} \big) +
\frac{1}{2} \ket{10} \big( a\ket{0}-b\ket{1} \big) +
\frac{1}{2} \ket{11} \big( a\ket{1}-b\ket{0} \big) .
$$
<a href="B54_Quantum_Teleportation_Solutions.ipynb#task2">click for our solution</a>
<h3> Measurement by Asja </h3>
Asja measures her qubits. With probability $ \frac{1}{4} $, she can observe one of the basis states.
Depeding on the measurement outcomes, Balvis' qubit is in the following states:
<ol>
<li> "00": $ \ket{v_{00}} = a\ket{0} + b \ket{1} $ </li>
<li> "01": $ \ket{v_{01}} = a\ket{1} + b \ket{0} $ </li>
<li> "10": $ \ket{v_{10}} = a\ket{0} - b \ket{1} $ </li>
<li> "11": $ \ket{v_{11}} = a\ket{1} - b \ket{0} $ </li>
</ol>
As can be observed, the amplitudes $ a $ and $ b $ are "transferred" to Balvis' qubit in any case.
If Asja sends the measurement outcomes, then Balvis can construct $ \ket{v} $ exactly.
<h3>Task 3</h3>
Asja sends the measurement outcomes to Balvis by using two classical bits: $ x $ and $ y $.
For each $ (x,y) $ pair, determine the quantum operator(s) that Balvis can apply to obtain $ \ket{v} = a\ket{0}+b\ket{1} $ exactly.
<a href="B54_Quantum_Teleportation_Solutions.ipynb#task3">click for our solution</a>
<h3> Task 4 </h3>
Create a quantum circuit with three qubits and two classical bits.
Assume that Asja has the first two qubits and Balvis has the third qubit.
Implement the protocol given above until Balvis makes the measurement.
<ul>
<li>Create entanglement between Asja's second qubit and Balvis' qubit.</li>
<li>The state of Asja's first qubit can be initialized to a randomly picked angle.</li>
<li>Asja applies CNOT and Hadamard operators to her qubits.</li>
<li>Asja measures her own qubits and the results are stored in the classical registers. </li>
</ul>
At this point, read the state vector of the circuit by using "statevector_simulator".
<i> When a circuit having measurement is simulated by "statevector_simulator", the simulator picks one of the outcomes, and so we see one of the states after the measurement.</i>
Verify that the state of Balvis' qubit is in one of these: $ \ket{v_{00}}$, $ \ket{v_{01}}$, $ \ket{v_{10}}$, and $ \ket{v_{11}}$.
<i> Follow the Qiskit order. That is, let qreg[2] be Asja's first qubit, qreg[1] be Asja's second qubit and let qreg[0] be Balvis' qubit.</i>
```
#
# your code is here
#
```
<a href="B54_Quantum_Teleportation_Solutions.ipynb#task4">click for our solution</a>
To implement the measurement by Balvis, we need <i>classically controlled</i> operations.
<h3> Task 5 </h3>
Implement the protocol above by including the post-processing part done by Balvis, i.e., the measurement results by Asja are sent to Balvis and then he may apply $ X $ or $ Z $ gates depending on the measurement results.
We use the classically controlled quantum operators.
Since we do not make measurement on $ q[2] $, we define only 2 classical bits, each of which can also be defined separated.
```python
q = QuantumRegister(3)
c2 = ClassicalRegister(1,'c2')
c1 = ClassicalRegister(1,'c1')
qc = QuantumCircuit(q,c1,c2)
...
qc.measure(q[1],c1)
...
qc.x(q[0]).c_if(c1,1) # x-gate is applied to q[0] if the classical bit c1 is equal to 1
```
Read the state vector and verify that Balvis' state is $ \myvector{a \\ b} $ after the post-processing.
```
#
# your code is here
#
```
<a href="B54_Quantum_Teleportation_Solutions.ipynb#task5">click for our solution</a>
| github_jupyter |
# Creating a Project Contributor/Collaborator/Author list
It's standard practice in Zooniverse projects to acknowledge the contributions of the volunteers in any publication, report, or other kind of description of the results of the project. This can come in many forms, e.g. in the authors section (usually as a footnote) or acknowledgments section of a journal article. There is usually a relatively brief acknowledgment that directs readers to a full list of contributors. This notebook shows you how to use `make_author_list.py` to generate that list. The list is generated in markdown format.
*Note: different projects use different terms to refer to their volunteers, depending on how they interact with them. Commonly used terms (in addition to "volunteer") are "collaborator", "author", and "contributor". Many avoid the term "user" as it can feel sterile and doesn't really describe the nature of the engagement between project team members (it gets used more often when describing people who make use of software features, for example).*
```
import sys, os
import numpy as np
import pandas as pd
from make_author_list import make_author_list, make_author_list_help
print("Python version: %d.%d.%d, numpy version: %s, pandas version: %s." %(sys.version_info[0],
sys.version_info[1],
sys.version_info[2],
np.__version__,
pd.__version__))
print("Originally developed using Py 2.7.11, np v1.11.0, pd v0.19.2")
print("If these versions don't match and stuff breaks, that's probably why.")
```
#### Options for inputs
In order to generate a list, you just need to specify the name of an input file to read in and the name of an output file to write to. However, there are many different options. To see them all:
```
make_author_list_help()
```
***Note:** you will need to have the panoptes python client installed.
If you have used `basic_classification_stats.py`, e.g. by running the previous Jupyter notebook to this one, you will have generated a list of your project's users along with their classification counts, in csv format.
To start, let's assume that's the case and generate a team contributor list in preformatted markdown. If we can, we want to use the `credited_name` field, which is an optional field that a new Zooniverse participant can specify they want used when they're given credit for their contributions. The `user_name` and `credited_name` are both public in the Zooniverse, so it's ok that we're using them in this public example notebook.
Sometimes people use their emails as their usernames, and even though it's clearly stated that Zooniverse usernames are public, we should maybe avoid publishing them on a bunch of team web pages where they might be picked up by spam bots. The code includes a search for email addresses and can sanitize them if we wish.
```
project_name = "my-project"
# This is the file that's output by basic_classification_processing()
# we generated it in the previous Notebook.
vol_file = project_name + "-classifications_nclass_byuser_loggedin_ranked.csv"
outfile = project_name + "-contributors.md"
# we have to use the "user_name" column in this example because the example file
# doesn't have real Zooniverse IDs. Real exports will, and looking up by user id is faster
# so don't specify author_col unless you need to
authorlist = make_author_list(vol_file, outfile, clean_emails=True, preformat=True, author_col="user_name")
# this is... not that fast (as in, for low thousands of users, go get a coffee)
```
#### Warning messages
There are various reasons the user search might not return anything for a given user ID or name. The most common are:
1. The user account has been deleted
2. There is an issue with the API request (timeout, etc)
If the user account has been deleted, their classifications remain in the system but you can no longer attach that username/ID to any other name information. In that case, either the user name or ID will appear in the author list instead.
API errors sometimes happen if there is high server load, so if you suspect that's what's happening, you can try again later or email the contact@zooniverse account to find out more information.
At the moment, both these reasons for a no-result lookup return the same error, so we can't distinguish between the two in the script. But you should always get back either a list of usernames or IDs that didn't pull a result, so you can use that to help you figure out what's happening (if it's just a few names, it's probably that people decided to delete their account).
#### Checking that it worked
```
!head my-project-contributors.md
authorlist
```
If you want to do something else with this author list, you can use the dataframe to do it, or manipulate the markdown file directly.
*Note: the names will show up in the same order they do in the csv file; in this case they're ranked by number of classifications, but you may want to order it differently.*
### Generating an author list directly from a classification export
If you haven't run `basic_classification_processing.py` you might not already have a list of users. But the classification exports provide this information, so you can use that instead to generate a user list.
```
from make_author_list import make_userfile_from_classfile
classification_file = project_name + "-classifications.csv"
userfile = make_userfile_from_classfile(classification_file)
authorlist_fromclassfile = make_author_list(userfile, outfile, clean_emails=True, preformat=True, author_col="user_name")
```
### Why are the two author lists different?
In the first example, we used a list of contributors that was generated from running `basic_classification_processing` on the classification example file *and only extracting classifications from one workflow and version*. The quick-and-dirty version used immediately above doesn't do that. It doesn't remove non-live classifications, or duplicates, or separate classifications by workflow. So because it considers all classifications in the example file instead of just the ones in a specific workflow, it's a longer list.
This gives you the flexibility to choose how you generate these lists, depending on how you intend to use them.
### How should I use this list?
As a bare minimum, it's a good idea to generate a contributor list for all your volunteers who participated while your project was live, and paste it into your **Team** page on your project with an acknowledgment note as a header to the list. [Here's an example of that.](https://www.zooniverse.org/projects/vrooje/planetary-response-network-and-rescue-global-caribbean-storms-2017/about/team)
(It's no accident that the list there overlaps by quite a bit with the one here; the example classifications were taken from that project.)
#### Running `make_author_list` from the command line
At the command line, type:
`%> python make_author_list.py`
without any inputs to see what the CLI syntax is.
| github_jupyter |
# CUSP UCSL 2016
## utility NB for darts gaussian distribution plot (darts)
## FBB JUNE 2016
```
__author__ = "__fbbianco__"
from __future__ import print_function
from __future__ import division
import os
import pylab as pl
import numpy as np
%pylab inline
N = 10000
def circlepoint(r, c, theta):
return()
def circle(r, c):
cx, cy = np.zeros(N), np.zeros(N)
for i, x in enumerate(np.linspace(c[0] - r, c[0] + r, N)):
cx[i] = x
cy[i] = np.sqrt(r ** 2 - (x - c[0]) ** 2) + c[1]
return cx, cy
def wedges(r, c, nw, v, mod):
cx, cy = np.zeros(N), np.zeros(N)
ii = 0
for i, x in enumerate(np.linspace(c[0] - r, c[0] + r, N)):
y = np.sqrt(r ** 2 - (x - c[0]) ** 2) + c[1]
theta = np.arctan(y / x)
if int((theta * 180 / np.pi / 360. + v) * nw) % 2 == mod:
#print "hrtr", theta, int(theta * 180 / np.pi / 360. * nw)%2
cx[ii] = x
cy[ii] = y
ii += 1
return cx, cy
np.random.seed(100)
fig = pl.figure(figsize = (10, 10))
mycircle = circle(12, [0, 0])
pl.fill_between(mycircle[0], mycircle[1] * ( - 1), mycircle[1], \
where = mycircle[1] > mycircle[1] * ( - 1.0), color = 'black')
mycircleout = circle(10, [0, 0])
pl.plot(mycircleout[0], - mycircleout[1], '.', color = 'green')
pl.plot(mycircleout[0], mycircleout[1], '.', color = 'red')
mycirclein = circle(5, [0, 0])
pl.plot(mycirclein[0], - mycirclein[1], '.', color = 'green')
pl.plot(mycirclein[0], mycirclein[1], '.', color = 'red')
mywedges = wedges(5, [0, 0], 20, 360. / 27, 0)
pl.plot(mywedges[0], mywedges[1], '.', color = 'green')
mywedges = wedges(5, [0, 0], 20, - 360. / 27, 1)
pl.plot(mywedges[0], - mywedges[1], '.', color = 'red')
mywedges = wedges(10, [0, 0], 20, 360. / 27, 0)
pl.plot(mywedges[0], mywedges[1], '.', color = 'green')
mywedges = wedges(10, [0, 0], 20, - 360. / 27, 1)
pl.plot(mywedges[0], - mywedges[1], '.', color = 'red')
mycircleinner = circle(0.3, [0, 0])
pl.plot(mycircleinner[0], - mycircleinner[1], '.', color = 'green')
pl.plot(mycircleinner[0], mycircleinner[1], '.', color = 'green')
rs = 2 * np.random.randn(100) - 0.5
print (rs)
for i, f in enumerate(range(100)):
theta = (np.random.rand() * np.pi)
pl.scatter(rs[i] * np.cos(theta), rs[i] * np.sin(theta), color = 'gray')
#pl.savefig(os.getenv('PUI15PLOTS') + ' / board_%02d.png'%f, transparent = True)
mygauss = lambda x: np.exp( - (x) ** 2)
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FixedLocator, FormatStrFormatter
import matplotlib.pyplot as plt
import numpy as np
fig = plt.figure()
ax = fig.gca(projection = '3d')
X = np.arange( - 3, 3, 0.05)
Y = np.arange( - 3, 3, 0.05)
X, Y = np.meshgrid(X, Y)
R = np.sqrt(mygauss(X) ** 2 + mygauss(Y) ** 2)
Z = 1.0 / (2.0 * pi) * mygauss(X) * mygauss(Y)
surf = ax.plot_surface(X, Y, Z, rstride = 1, cstride = 1, cmap = cm.jet,
linewidth = 0, antialiased = False)
ax.set_zlim3d( - .01, .2)
ax.w_zaxis.set_major_locator(LinearLocator(10))
ax.w_zaxis.set_major_formatter(FormatStrFormatter('%.03f'))
fig.colorbar(surf, shrink = 0.5, aspect = 2)
#plt.savefig('figsurface.png')
```
| github_jupyter |
```
import requests
import pandas as pd
import numpy as np
import json
import urllib
```
# We create a function to access Foursquare API and store the results in a dataframe and a CSV
```
def places_search(term,street, zipcode):
dict = {"near": zipcode,
"address": street,
"query" :term,
#"intent":"match",
"zip":zipcode,
"limit":1,
"client_id" : "",
"client_secret" : "",
"v" : "20181122"
}
params =urllib.parse.urlencode(dict)
url = "https://api.foursquare.com/v2/venues/search?"+params
resp = requests.get(url)
#grabbing the JSON result
data = json.loads(resp.text)
return data
df_restaurant = pd.read_csv("restaurants_raw_data_foursquare.csv")
df_restaurant = df_restaurant[df_restaurant["dba"].notnull()]
count =1
#df_restaurant["searched_foursquare"] = False
df_restaurant = df_restaurant.sort_values(by=['searched_foursquare',"dba"],ascending = True)
print(len(df_restaurant))
print(len(df_restaurant[df_restaurant["searched_foursquare"] == True ]))
for restau in df_restaurant.index[0:2000]:
if df_restaurant.loc[restau,"searched_foursquare"] == False:
jsonData = places_search(str(df_restaurant.loc[restau,"dba"]),str(df_restaurant.loc[restau,"street"]),str(df_restaurant.loc[restau,"zipcode"].astype(int)))
try:
df_restaurant.loc[restau,"foursquare_id"] = jsonData["response"]["venues"][0]["id"]
df_restaurant.loc[restau,"foursquare_name"] = jsonData["response"]["venues"][0]["name"]
df_restaurant.loc[restau,"foursquare_address"] = jsonData["response"]["venues"][0]["location"]["address"]
df_restaurant.loc[restau,"foursquare_lat"] = jsonData["response"]["venues"][0]["location"]["lat"]
df_restaurant.loc[restau,"foursquare_lng"] = jsonData["response"]["venues"][0]["location"]["lng"]
df_restaurant.loc[restau,"foursquare_category_id"] = jsonData["response"]["venues"][0]["categories"][0]["id"]
df_restaurant.loc[restau,"foursquare_category_name"] = jsonData["response"]["venues"][0]["categories"][0]["name"]
print(count,str(df_restaurant.loc[restau,"dba"]),jsonData["response"]["venues"][0]["name"])
except:
print("Foursquare couldn't find!")
df_restaurant.loc[restau,"searched_foursquare"] = True
count = count +1
df_restaurant.to_csv("restaurants_raw_data_foursquare.csv",index=False)
```
| github_jupyter |
```
import torch
import torch.nn as nn
import numpy as np
from sklearn import datasets
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import torchvision
import torchvision.transforms as transforms
import matplotlib.pyplot as plt
from torch.utils.data import Dataset, DataLoader
from scipy.special import softmax
# General
from os import path
from random import randrange
from sklearn.model_selection import train_test_split, GridSearchCV #cross validation
from sklearn.metrics import confusion_matrix, plot_confusion_matrix, make_scorer
from sklearn.metrics import accuracy_score, roc_auc_score, balanced_accuracy_score
from sklearn.preprocessing import LabelEncoder
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import xgboost as xgb
import pickle
import joblib
trainDataFull = pd.read_csv("trainData.csv")
trainDataFull.head(3)
trainData = trainDataFull.loc[:,'v1':'v99']
trainData.head(3)
trainLabels = trainDataFull.loc[:,'target']
trainLabels.unique()
# encode string class values as integers
label_encoder = LabelEncoder()
label_encoder = label_encoder.fit(trainLabels)
label_encoded_y = label_encoder.transform(trainLabels)
label_encoded_y
X_train, X_test, y_train, y_test = train_test_split(trainData.values,
label_encoded_y,
test_size = 0.05,
random_state = 33,
shuffle = True,
stratify = label_encoded_y)
# scale
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
X_train = torch.from_numpy(X_train.astype(np.float32))
X_test = torch.from_numpy(X_test.astype(np.float32))
# y_train = torch.from_numpy(y_train.astype(np.float32))
# y_test = torch.from_numpy(y_test.astype(np.float32))
# y_train = y_train.view(y_train.shape[0],1) # from columns vector to row vector
# y_test = y_test.view(y_test.shape[0],1)
y_train
len(X_train)
class TrainDataset(Dataset):
def __init__(self, data, labels):
self.paths = data
self.labels = labels
def __len__(self):
return self.paths.shape[0]
def __getitem__(self, i):
image = self.paths[i]
label = torch.tensor(self.labels[i])
return image, label
class ValidDataset(Dataset):
def __init__(self, data, labels):
self.paths = data
self.labels = labels
def __len__(self):
return self.paths.shape[0]
def __getitem__(self, i):
image = self.paths[i]
label = torch.tensor(self.labels[i])
return image, label
# hyper parameters
input_size = 103
num_classes = 9
num_epochs = 1000
batch_size = 100
learning_rate = 0.00001
device = 'cuda:0'
train_dataset = TrainDataset(X_train, y_train)
trainloader = DataLoader(train_dataset, shuffle=True, batch_size = batch_size, num_workers = 2)
valid_dataset = ValidDataset(X_test, y_test)
validloader = DataLoader(valid_dataset, shuffle=False, batch_size = batch_size, num_workers = 2)
class NeuralNet(nn.Module):
def __init__(self, input_size, num_classes):
super(NeuralNet, self).__init__()
self.l1 = nn.Linear(input_size, 2000,bias = True)
self.relu1 = nn.ReLU()
self.l2 = nn.Linear(2000,1000,bias = True)
self.relu2 = nn.ReLU()
self.drop1 = nn.Dropout(p=0.3)
self.l3 = nn.Linear(1000, num_classes,bias = True)
def forward(self, x):
out = self.l1(x)
out = self.relu1(out)
out = self.l2(out)
out = self.relu2(out)
out = self.drop1(out)
out = self.l3(out)
# It is no need Softmax. Because CrossEntropyLoss includes it
return out
model = NeuralNet(input_size, num_classes)
model.cuda()
# loss and optimizer
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters(),lr=learning_rate)
# training loop
n_total_steps = len(trainloader)
for epoch in range(num_epochs):
for i, (images, labels) in enumerate(trainloader):
# images.size = 100, 1, 28, 28
# labels.size = 100, 784
images = images.to(device)
labels = labels.to(device)
model.train()
# forward
outputs = model(images)
loss = criterion(outputs, labels)
optimizer.zero_grad()
# backwards
loss.backward()
optimizer.step()
if (i+1) % 100 == 0:
print(f'epoch {epoch+1}/{num_epochs}, step {i+1}/{n_total_steps}, loss = {loss.item():.4f}')
# Valid score
with torch.no_grad():
n_correct = 0
n_samples = 0
for images, labels in validloader:
images = images.to(device)
labels = labels.to(device)
outputs = model(images)
# value, index
_, predictions = torch.max(outputs,1)
n_samples += labels.shape[0]
n_correct += (predictions == labels).sum().item()
acc = 100.0 * n_correct / n_samples
print(f'accuracy = {acc}')
# Valid score
with torch.no_grad():
n_correct = 0
n_samples = 0
for images, labels in validloader:
images = images.to(device)
labels = labels.to(device)
outputs = model(images)
# value, index
_, predictions = torch.max(outputs,1)
n_samples += labels.shape[0]
n_correct += (predictions == labels).sum().item()
acc = 100.0 * n_correct / n_samples
print(f'accuracy = {acc}')
class TestDataset(Dataset):
def __init__(self, data, labels):
self.paths = data
self.labels = labels
def __len__(self):
return self.paths.shape[0]
def __getitem__(self, i):
image = self.paths[i]
label = torch.tensor(self.labels[i])
return image, label
testData = pd.read_csv("testData.csv")
testData
test_normalized_standart = sc.transform(testData.values)
test_normalized_standart
testNN = torch.from_numpy(test_normalized_standart.astype(np.float32))
testNN[0]
preds_pair = []
for i in range(len(testNN)):
print(i)
outputs = model(testNN[i].to(device))
predictions_normalize = softmax(outputs.cpu().detach().numpy(),0)
preds_pair.append(list(predictions_normalize))
result = pd.DataFrame(preds_pair, columns=['c1','c2','c3','c4','c5','c6','c7','c8','c9'])
result
result.to_csv('./results/test-submission-model-nn', index = False)
sum(predictions_normalize)
preds_pair = []
for i in range(len(X_test)):
print(i)
outputs = model(X_test[i].to(device))
predictions_normalize = softmax(outputs.cpu().detach().numpy(),0)
preds_pair.append(list(predictions_normalize))
```
| github_jupyter |
# Softmax_Regression_with_MNIST
## 1. Import Packages
```
import input_data
import numpy as np
import matplotlib.pylab as plt
import tensorflow as tf
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
```
## 2. Explore MNIST Data
```
mnist_images = input_data.read_data_sets("./mnist_data", one_hot=False)
# Example of a picture
pic,real_values = mnist_images.train.next_batch(25)
index = 11 # changeable with 0 ~ 24 integer
image = pic[index,:]
image = np.reshape(image,[28,28])
plt.imshow(image)
plt.show()
# Explore MNIST data
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
image = np.reshape(pic[i,:] , [28,28])
plt.imshow(image)
plt.xlabel(real_values[i])
plt.show()
```
## 3. Make Dataset
```
# Download Data : http://yann.lecun.com/exdb/mnist/
# Data input script : https://github.com/tensorflow/tensorflow/tree/master/tensorflow/examples/tutorials/mnist
mnist = input_data.read_data_sets("./samples/MNIST_data/", one_hot=True)
print("the number of train examples :" , mnist.train.num_examples)
print("the number of test examples :" , mnist.test.num_examples)
```
## 4. Building a neural network with tensorflow
```
# Create placeholders
x = tf.placeholder(tf.float32, [None, 784])
# Initialize parameters
W = tf.Variable(tf.zeros([784, 10]))
b = tf.Variable(tf.zeros([10]))
# Forward Propagation
y = tf.nn.softmax(tf.matmul(x, W) + b)
y_ = tf.placeholder(tf.float32, [None, 10])
# Compute cost
cross_entropy = - tf.reduce_sum(y_*tf.log(y))
# Backward Propagation
learning_rate = 0.01
train_step = tf.train.GradientDescentOptimizer(learning_rate).minimize(cross_entropy)
# Initialize all the variables
init = tf.global_variables_initializer()
# Start the session to compute the tensorflow graph
sess = tf.Session()
sess.run(init)
# Do the training loop - Stochastic training
batch_size = 100
epoch_cost = 0
costs = []
for i in range(1000):
batch_xs, batch_ys = mnist.train.next_batch(batch_size) # 100 random data
_ , minibatch_cost = sess.run([train_step, cross_entropy], feed_dict={x: batch_xs, y_: batch_ys})
epoch_cost = minibatch_cost / batch_size
# Print the cost every epoch
if i % 100 == 0:
print ("Cost after epoch %i: %f" % (i, epoch_cost))
if i % 10 == 0:
costs.append(epoch_cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per 10)')
plt.title("Learning rate =" + str(0.01))
plt.show()
```
## 5. Calculate Accuracy
```
# Validation
# Calculate the correct predictions
correct_prediction = tf.equal(tf.argmax(y,1), tf.argmax(y_,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
# Accuracy
print("Train Accuracy : " , sess.run(accuracy, feed_dict={x: mnist.train.images, y_: mnist.train.labels}))
print("Test Accuracy : " , sess.run(accuracy, feed_dict={x: mnist.test.images, y_: mnist.test.labels}))
```
## 6. Check wrong prediction
```
w = []
for r in range(1000):
if sess.run(tf.argmax(mnist.test.labels[r:r+1], 1)) != sess.run (tf.argmax(y, 1), feed_dict={x: mnist.test.images[r:r+1]}):
w.append(r)
print("wrong label : " ,w)
wrong_pred = []
for i in range(len(w)) :
wrong_pred.append(sess.run (tf.argmax(y, 1), feed_dict={x: mnist.test.images[w[i]:w[i]+1]}))
wrong_label = []
for i in range(len(w)) :
wrong_label.append(sess.run(tf.argmax(mnist.test.labels[w[i]:w[i]+1], 1)))
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(mnist.test.images[w[i]].reshape(28, 28), cmap='Greys', interpolation='nearest')
plt.xlabel("Label" + str(wrong_label[i]) + " ,Pred" +str(wrong_pred[i]))
plt.show()
```
| github_jupyter |
# 3. Bidirectional Recurrent Neural Networks (BRNN)
## Limitation of RNN
In the last lecture, we learned about **Recurrent Neural Networks**. What distinguishes itself from feedforward layers is that it can process sequential data. Although it sounds super cool enough, there's also limitation to it.
Let's say we want to build a model that predicts which word would fall into the blank, `[ ]`. What would be the answer in the following sentence?
- He said, "Teddy `[ ]`"
Yes, it's pretty hard to predict the next word since we only have the past data, which are words before the blank. And how would you know if "Teddy" means a Teddy bear, Teddy Roosevelt, Teddy Sears, or the new Indian action thriller [film](https://en.wikipedia.org/wiki/Teddy_(film)) secretly released on 2021? It seems it might be more helpful to know which words come after the blank. The problem of RNNs is that it misses out future information which might hold important context.
## BRNN is here
Here comes Bidirectional Recurrent Neural Networks. As the name suggests, it not only holds a memory of inputs in one order that RNNs does, but the other way around. This way, the output layer can receive more rich information from both directions, which makes it outstanding especially when the context of the input should be considered.
Let's get back to the previous example to intuitively understand why it's awesome. We have the same task to predict the word in the blank, but this time, we are also given next words.
- He said, "Teddy `[ ]` are cuter than you."
Now, it makes sense. The word in the blank should be "Roosevelt."
You spotted it's a joke for sure because you also looked into what comes after the blank. BRNN does the same.
Be careful about when to use BRRN, though. As you might have guessed, we are not always guaranteed to look into future. That half of its structure depends on the future data means that it can perform half poorly when it has no access to one.
Still, it makes perfect sense to use Bidirectional RNN in various areas where we're likely to have complete inputs such as
- Speech Recognition
- Translation
- Handwrrting Recognition
- Protein Structure Prediction
- Part-of-speech tagging
## BRNN's Structure

<center><i>BRNN Image from DIVE INTO DEEP LEARNING</i></center>
BRNN is largely composed of two vertically stacked RNNs.
$X_t \in \mathbb{R}^{n \times d}$ (where $n$ is the number of sequences and $d$ is the number of input features) is an input for each time step. And let $\phi$ be an activation function for hidden layer.
In the forward layer, $\overrightarrow{\text{H}_t}$ passes its output to both its next time step and the output.
$$\overrightarrow{\text{H}_t} = \phi(X_t\overrightarrow{\text{W}}_{xh} + \overrightarrow{\text{H}}_{t-1}\overrightarrow{\text{W}}_{hh} + \overrightarrow{b_h})$$
The layer above $\overleftarrow{\text{H}_t}$ does it backward.
$$\overleftarrow{\text{H}_t} = \phi(X_t\overleftarrow{\text{W}}_{xh} + \overleftarrow{\text{H}}_{t+1}\overleftarrow{\text{W}}_{hh} + \overleftarrow{b_h})$$
We can obtain the output $O_t$ from the concatenation of the two outputs of $\overrightarrow{\text{H}}_t$ and $\overleftarrow{\text{H}}_t$.
$$O_t = H_{t}W_{ho} + b_o$$
where $W_{ho} \in \mathbb{R}^{2h \times o} $ denotes the weights between the hidden layers and the output layer.
## Keras implementation
After covering the theory, let's now look how BRNNs can be implemented using Keras.
```
import tensorflow as tf
model = tf.keras.models.Sequential([
tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(64, return_sequences = True, input_shape = (5, 10))),
tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(32)),
tf.keras.layers.Dense(10)
])
```
As you can see, the `Bidirectional()` layer function handles the structuring of our neural network and the only thing we are required to do is specifying the type of recurrent network we want to use for our Bidirectional layer. In the example case, we used LSTM, but you could use RNN or GRU (we will be covering it in the future tutorials).
| github_jupyter |
# RandomForestRegressor with StandardScaler
This Code template is for regression analysis using a RandomForestRegressor based on the Ensemble Learning technique using feature scaling via StandardScaler
### Required Packages
```
import warnings
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as se
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import r2_score, mean_absolute_error, mean_squared_error
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training .
```
#x_values
features=[]
```
Target variable for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X = df[features]
Y = df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=NullClearner(Y)
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size = 0.2, random_state = 123)
```
### Data Rescaling
Performing StandardScaler data rescaling operation on dataset. The StandardScaler standardize features by removing the mean and scaling to unit variance.
We will fit an object of StandardScaler to **train data** then transform the same data via <Code>fit_transform(X_train)</Code> method, following which we will transform **test data** via <Code>transform(X_test)</Code> method.
```
standard_scaler = StandardScaler()
X_train = standard_scaler.fit_transform(X_train)
X_test = standard_scaler.transform(X_test)
```
### Model
A random forest is a meta estimator that fits a number of classifying decision trees on various sub-samples of the dataset and uses averaging to improve the predictive accuracy and control over-fitting. The sub-sample size is controlled with the <code>max_samples</code> parameter if <code>bootstrap=True</code> (default), otherwise the whole dataset is used to build each tree.
#### Model Tuning Parameters
1. n_estimators : int, default=100
> The number of trees in the forest.
2. criterion : {“mae”, “mse”}, default=”mse”
> The function to measure the quality of a split. Supported criteria are “mse” for the mean squared error, which is equal to variance reduction as feature selection criterion, and “mae” for the mean absolute error.
3. max_depth : int, default=None
> The maximum depth of the tree.
4. max_features : {“auto”, “sqrt”, “log2”}, int or float, default=”auto”
> The number of features to consider when looking for the best split:
5. bootstrap : bool, default=True
> Whether bootstrap samples are used when building trees. If False, the whole dataset is used to build each tree.
6. oob_score : bool, default=False
> Whether to use out-of-bag samples to estimate the generalization accuracy.
7. n_jobs : int, default=None
> The number of jobs to run in parallel. fit, predict, decision_path and apply are all parallelized over the trees. <code>None</code> means 1 unless in a joblib.parallel_backend context. <code>-1</code> means using all processors. See Glossary for more details.
8. random_state : int, RandomState instance or None, default=None
> Controls both the randomness of the bootstrapping of the samples used when building trees (if <code>bootstrap=True</code>) and the sampling of the features to consider when looking for the best split at each node (if <code>max_features < n_features</code>).
9. verbose : int, default=0
> Controls the verbosity when fitting and predicting.
```
model = RandomForestRegressor(n_jobs = -1,random_state = 123)
model.fit(X_train, y_train)
```
#### Model Accuracy
We will use the trained model to make a prediction on the test set.Then use the predicted value for measuring the accuracy of our model.
> **score**: The **score** function returns the coefficient of determination <code>R<sup>2</sup></code> of the prediction.
```
print("Accuracy score {:.2f} %\n".format(model.score(X_test,y_test)*100))
```
> **r2_score**: The **r2_score** function computes the percentage variablility explained by our model, either the fraction or the count of correct predictions.
> **mae**: The **mean abosolute error** function calculates the amount of total error(absolute average distance between the real data and the predicted data) by our model.
> **mse**: The **mean squared error** function squares the error(penalizes the model for large errors) by our model.
```
y_pred=model.predict(X_test)
print("R2 Score: {:.2f} %".format(r2_score(y_test,y_pred)*100))
print("Mean Absolute Error {:.2f}".format(mean_absolute_error(y_test,y_pred)))
print("Mean Squared Error {:.2f}".format(mean_squared_error(y_test,y_pred)))
```
#### Feature Importances
The Feature importance refers to techniques that assign a score to features based on how useful they are for making the prediction.
```
plt.figure(figsize=(8,6))
n_features = len(X.columns)
plt.barh(range(n_features), model.feature_importances_, align='center')
plt.yticks(np.arange(n_features), X.columns)
plt.xlabel("Feature importance")
plt.ylabel("Feature")
plt.ylim(-1, n_features)
```
#### Prediction Plot
First, we make use of a plot to plot the actual observations, with x_train on the x-axis and y_train on the y-axis.
For the regression line, we will use x_train on the x-axis and then the predictions of the x_train observations on the y-axis.
```
plt.figure(figsize=(14,10))
plt.plot(range(20),y_test[0:20], color = "green")
plt.plot(range(20),model.predict(X_test[0:20]), color = "red")
plt.legend(["Actual","prediction"])
plt.title("Predicted vs True Value")
plt.xlabel("Record number")
plt.ylabel(target)
plt.show()
```
#### Creator: Viraj Jayant, Github: [Profile](https://github.com/Viraj-Jayant)
| github_jupyter |
# Predicting The Shift
* savant leaderboard https://baseballsavant.mlb.com/visuals/team-positioning
## Goals for Modeling
* Can we predict the shift?
* Which features are most predictive?
* Use season to date stats (woba, avg launch angle/speed, etc.), lagged outcomes (whether they shifted this batter's last N PAs, whether it worked), batter id, handedness, situation (base state and run diff), avg. launch angle and exit velo, defense-team/team-year-interaction to predict whether an AB will be shifted
### Data Notes
* Throwing out batters with <100 PA may help accuracy, but it's also unrealistic to pretend those don't exist
```
from pybaseball import statcast
from pybaseball.lahman import master
from pybaseball import playerid_reverse_lookup
import pandas as pd
import numpy as np
import warnings
warnings.filterwarnings('ignore')
#df = statcast('2018-03-25','2018-08-17')
#df.to_csv('statcast_0817.csv')
```
### Use Statcast data 2015 - present. Only consider regular-season games.
```
#df = pd.read_csv('statcast_0817.csv')
# using a csv of statcast events April 2015 - Aug. 2018
df = pd.read_csv('/Users/jledoux/Documents/projects/Saber/baseball-data/statcast_with_shifts.csv')
df['game_date'] = pd.to_datetime(df['game_date'])
# R == 'regular season'
df.game_type.value_counts()
# only consider regular season games
df = df.loc[df['game_type'] == 'R',]
df.columns
```
### Bring in batter names because savant only gives pitcher or batter name, not both. Use bbref ids to connect w/ Lahman to get batter handedness.
```
batters = df.batter
names = playerid_reverse_lookup(batters)
names['batter_name'] = names['name_first'] + ' ' + names['name_last']
master = master()
master = master[['bbrefID', 'bats']]
names = names.merge(master, left_on='key_bbref', right_on = 'bbrefID', suffixes=('_chadwick', '_lahman'))
names = pd.DataFrame(names[['key_mlbam', 'batter_name', 'bats']])
df.batter = df.batter.astype(int)
df = df.merge(names, left_on='batter', right_on = 'key_mlbam', suffixes=('_old', ''))
df.if_fielding_alignment.value_counts()
df.of_fielding_alignment.value_counts()
df.head()
```
### Create a temporary shift column indicating whether the IF is shifted at a given pitch. Sum over this for each atbat. Atbats where a shift happened at least once will reveive a positive flag for the variable is_shift. Otherwise, it equals zero.
Two potential flaws with this: first, what if there was a pinch hitter partway through the atbat? In this case, there's no event for the first batter, so that batter-PA pair will be thrown out. Not a problem.
Second, this doesn't take into account the possibility of changing shift states partway through an AB. If they shift at any point, I'm calling it a shifted-atbat. I don't expect that to have a major impact on the results.
```
df['atbat_pk'] = df['game_pk'].astype(str) + df['at_bat_number'].astype(str)
df['is_shift'] = np.where(df['if_fielding_alignment'] == 'Infield shift', 1, 0)
df.is_shift.value_counts()
shifts = pd.DataFrame(df.groupby('atbat_pk')['is_shift'].sum()).reset_index()
shifts.loc[shifts.is_shift > 0, 'is_shift'] = 1
df = df.merge(shifts, on='atbat_pk', suffixes=('_old', ''))
```
### Create features for whether a player was on 1b, 2b, 3b at beginning of PA. Group by atbat pk, take the first, and use that as the column's value.
This is only an approximation of what's happening in reality, as steals, passed balls, etc. can all cause this to change throughout the life of an atbat, but it's a simple feature that impacts beginning-of-PA shift strategy.
```
df['man_on_first'] = np.where(df['on_1b'] > 0 , 1, 0)
df['man_on_second'] = np.where(df['on_2b'] > 0 , 1, 0)
df['man_on_third'] = np.where(df['on_3b'] > 0 , 1, 0)
df['men_on_base'] = df['man_on_first'] + df['man_on_second'] + df['man_on_third']
df[['on_1b', 'on_2b', 'on_3b', 'man_on_first', 'man_on_second', 'man_on_third', 'men_on_base']].head()
df['score_differential'] = df['fld_score'] - df['bat_score']
# drop rows where our value of interest is unknown
# if you drop the original column you'll lose partial atbats. sometimes it's null for only some pitches. dropping it loses event data.
print(df.shape)
df = df[pd.notnull(df['is_shift'])]
print(df.shape)
df.is_shift.value_counts()
```
### Some quick sanity checks to make sure these woba and shift numbers match up with Savant and FanGraphs.
My shift rates are higher than Savant's, but Savant also has lower PA-numbers than me. My PA numbers match FanGraphs', so I think my numbers are correct and Savant is doing some type of filtering that they aren't explaining on their leaderboard page.
Differences aside, my shift leaderboard ranks similarly to Savant's and my wOBA numbers are also reaonably close to FG's.
wOBA leaders:
```
# I want to use woba as a feature in this model. let's make sure these values are close to reality
# looks good https://baseballsavant.mlb.com/expected_statistics
woba_leaderboard = df.groupby(['batter_name', 'batter'])['woba_value'].agg(['mean', 'count'])
woba_leaderboard.loc[woba_leaderboard['count']>100,].sort_values('mean', ascending=False).head()
```
Shift leaders:
A lefty-heavy leaderboard. No righties in sight.
```
shifted_leaderboard = df.groupby(['batter_name', 'batter', 'bats'])['is_shift'].agg(['mean', 'count'])
shifted_leaderboard.loc[shifted_leaderboard['count']>1500,].sort_values('mean', ascending=False).head(15)
```
### Next: get this data to plate appearance level rather than pitch level
```
# for game-state variables we care about the beginning of the PA, since that mostly determines strategy
plate_appearances = df.sort_values(['game_date', 'at_bat_number'], ascending=True).groupby(['atbat_pk']).first().reset_index()
# get final event of the PA, as this will say single/hr/k/walk/etc
events = df.sort_values(['game_date', 'at_bat_number'], ascending=True).groupby(['atbat_pk']).last().reset_index()
events = events[['atbat_pk', 'events', 'woba_value']]
plate_appearances = plate_appearances.merge(events, on='atbat_pk', suffixes=['_ignore', ''])
# remove PAs with no event. these are probably PAs with pinch hitters. (< 1% of observations)
plate_appearances = plate_appearances.loc[plate_appearances.events.isnull()==False,]
plate_appearances.loc[plate_appearances.batter_name=='javier baez',].tail(10)
```
Shift Leaderboard (count = PAs, sum = number of shifts against, mean = % of PAs shifted against)
```
shifted_leaderboard = plate_appearances.groupby(['game_year','batter_name', 'batter', 'bats'])['is_shift'].agg(['mean', 'count', 'sum'])
shifted_leaderboard.loc[shifted_leaderboard['count']>150,].sort_values('mean', ascending=False).head(15)
```
### Make features numeric / model-ready
* drop unwanted columns
* create dummies of categoricals / text columns
* create averages from ones that make more sense as season-to-date (eg woba). Can I do moving averages to have this accurate at all points in time w/ no date leakage?
```
drops = ['Unnamed: 0', 'index', 'pitch_type', 'release_pos_x', 'release_pos_z', 'player_name',
'spin_dir', 'spin_rate_deprecated', 'break_angle_deprecated', 'break_length_deprecated', 'zone',
'des', 'type', 'pfx_x', 'pfx_z', 'plate_x', 'plate_z', 'on_3b', 'on_2b', 'on_1b', 'hc_x', 'hc_y',
'tfs_deprecated', 'tfs_zulu_deprecated', 'pos2_person_id', 'sv_id', 'vx0', 'vy0', 'vz0', 'ax', 'ay', 'az',
'sz_top', 'sz_bot', 'release_spin_rate', 'release_extension', 'pos1_person_id', 'pos2_person_id.1',
'pos3_person_id', 'pos4_person_id', 'pos5_person_id', 'pos6_person_id', 'pos7_person_id', 'pos8_person_id',
'pos9_person_id', 'release_pos_y', 'home_score', 'away_score', 'post_away_score',
'post_home_score', 'post_bat_score', 'post_fld_score', 'if_fielding_alignment', 'of_fielding_alignment',
'release_speed', 'events_ignore','description', 'estimated_ba_using_speedangle', 'woba_value_ignore',
'fld_score', 'is_shift_old', 'game_type', 'bats', 'balls', 'strikes', 'umpire']
plate_appearances = plate_appearances.drop(drops, 1)
plate_appearances.columns
# create lagged features to represent a player's past n PAs
```
### get average-to-date for each PA / player to avoid info leakage
The grouped features will share an index with the original dataframe, so reset the index and join on this + batter id
Do for:
* woba (expanding mean)
* babip
* plate appearances (rolling sum)
* launch angle (expanding mean)
* exit velo (expanding mean)
* hit_distance_sc (expanding mean)
Small sample properties mean these expanding means will behave erratically at first. For this reason I may end up ignoring observations with rolling average PA < N (n = 100 maybe).
```
#get average-to-date for each PA / player to avoid info leakage
woba_to_date = plate_appearances.sort_values(['batter','game_date','at_bat_number'], ascending=True).groupby('batter')['woba_value'].expanding(min_periods=1).mean()
pd.DataFrame(pd.DataFrame(woba_to_date).sort_index()).shape
plate_appearances.shape
plate_appearances = plate_appearances.sort_values(['batter', 'game_date', 'at_bat_number'], ascending=True)
woba_to_date = pd.DataFrame(woba_to_date).reset_index()
woba_to_date.columns = ['batter', 'index', 'woba']
plate_appearances = plate_appearances.reset_index()
plate_appearances = pd.merge(left=plate_appearances, right=woba_to_date, left_on=['batter','index'],
right_on=['batter', 'index'], suffixes=['old',''])
backup = plate_appearances
plate_appearances = backup
# for babip:
# note: this is just averaging the babip_value. It's much lower than actual babip bc doesn't throw out PAs not included in babip.
plate_appearances = plate_appearances.sort_values(['batter', 'game_date', 'at_bat_number'], ascending=True)
babip_to_date = plate_appearances.sort_values(['batter','game_date','at_bat_number'], ascending=True).groupby('batter')['babip_value'].expanding(min_periods=1).mean()
babip_to_date = pd.DataFrame(babip_to_date).reset_index()
babip_to_date.columns = ['batter', 'index', 'babip']
plate_appearances = plate_appearances.drop('index',1)
plate_appearances = plate_appearances.reset_index()
plate_appearances = pd.merge(left=plate_appearances, right=babip_to_date, left_on=['batter','index'],
right_on=['batter', 'index'], suffixes=['old',''])
# for launch angle
plate_appearances = plate_appearances.sort_values(['batter', 'game_date', 'at_bat_number'], ascending=True)
launch_angle_to_date = plate_appearances.sort_values(['batter','game_date','at_bat_number'], ascending=True).groupby('batter')['launch_angle'].expanding(min_periods=1).mean()
launch_angle_to_date = pd.DataFrame(launch_angle_to_date).reset_index()
launch_angle_to_date.columns = ['batter', 'index', 'launch_angle']
plate_appearances = plate_appearances.drop('index',1)
plate_appearances = plate_appearances.reset_index()
plate_appearances = pd.merge(left=plate_appearances, right=launch_angle_to_date, left_on=['batter','index'],
right_on=['batter', 'index'], suffixes=['old',''])
# for exit velo:
plate_appearances = plate_appearances.sort_values(['batter', 'game_date', 'at_bat_number'], ascending=True)
launch_speed_to_date = plate_appearances.sort_values(['batter','game_date','at_bat_number'], ascending=True).groupby('batter')['launch_speed'].expanding(min_periods=1).mean()
launch_speed_to_date = pd.DataFrame(launch_speed_to_date).reset_index()
launch_speed_to_date.columns = ['batter', 'index', 'launch_speed']
plate_appearances = plate_appearances.drop('index',1)
plate_appearances = plate_appearances.reset_index()
plate_appearances = pd.merge(left=plate_appearances, right=launch_speed_to_date, left_on=['batter','index'],
right_on=['batter', 'index'], suffixes=['old',''])
# for hit distance
plate_appearances = plate_appearances.sort_values(['batter', 'game_date', 'at_bat_number'], ascending=True)
hit_distance_sc = plate_appearances.sort_values(['batter','game_date','at_bat_number'], ascending=True).groupby('batter')['hit_distance_sc'].expanding(min_periods=1).mean()
hit_distance_sc = pd.DataFrame(hit_distance_sc).reset_index()
hit_distance_sc.columns = ['batter', 'index', 'hit_distance_sc']
plate_appearances = plate_appearances.drop('index',1)
plate_appearances = plate_appearances.reset_index()
plate_appearances = pd.merge(left=plate_appearances, right=hit_distance_sc, left_on=['batter','index'],
right_on=['batter', 'index'], suffixes=['old',''])
# for is_shift (shift percentage to date on all dates)
plate_appearances = plate_appearances.sort_values(['batter', 'game_date', 'at_bat_number'], ascending=True)
shifts_to_date = plate_appearances.sort_values(['batter','game_date','at_bat_number'], ascending=True).groupby('batter')['is_shift'].expanding(min_periods=1).mean()
shifts_to_date = pd.DataFrame(shifts_to_date).reset_index()
shifts_to_date.columns = ['batter', 'index', 'avg_shifted_against']
plate_appearances = plate_appearances.drop('index',1)
plate_appearances = plate_appearances.reset_index()
plate_appearances = pd.merge(left=plate_appearances, right=shifts_to_date, left_on=['batter','index'],
right_on=['batter', 'index'], suffixes=['old',''])
plate_appearances[['batter', 'atbat_pk', 'is_shift', 'avg_shifted_against']].tail()
# away_team, home_team
# we don't have column for which team is pitching, but we know the home team pitches the top and away pitches bottom
plate_appearances['team_pitching'] = np.where(plate_appearances['inning_topbot']=='Top', plate_appearances['home_team'],
plate_appearances['away_team'])
plate_appearances['team_batting'] = np.where(plate_appearances['inning_topbot']=='Top', plate_appearances['away_team'],
plate_appearances['home_team'])
# for teams (shift percentage to date for each team)
plate_appearances = plate_appearances.sort_values(['team_pitching', 'game_date'], ascending=True)
shifts_to_date = plate_appearances.sort_values(['team_pitching', 'game_date'], ascending=True).groupby('team_pitching')['is_shift'].expanding(min_periods=1).mean()
shifts_to_date = pd.DataFrame(shifts_to_date).reset_index()
shifts_to_date.columns = ['team_pitching', 'index', 'def_shift_pct']
plate_appearances = plate_appearances.drop('index',1)
plate_appearances = plate_appearances.reset_index()
plate_appearances = pd.merge(left=plate_appearances, right=shifts_to_date, left_on=['team_pitching','index'],
right_on=['team_pitching', 'index'], suffixes=['old',''])
plate_appearances.groupby('team_pitching')['def_shift_pct'].last().sort_values(ascending=False)
# rolling count for plate appearances
plate_appearances = plate_appearances.sort_values(['batter', 'game_date', 'at_bat_number'], ascending=True)
pas = plate_appearances.sort_values(['batter','game_date','at_bat_number'], ascending=True).groupby('batter')['index'].expanding(min_periods=1).count()
pas = pd.DataFrame(pas).reset_index()
pas.columns = ['batter', 'index', 'pas']
plate_appearances = plate_appearances.drop('index',1)
plate_appearances = plate_appearances.reset_index()
plate_appearances = pd.merge(left=plate_appearances, right=pas, left_on=['batter','index'],
right_on=['batter', 'index'], suffixes=['old',''])
plate_appearances.loc[plate_appearances.batter_name=='mike trout',].head()
plate_appearances.is_shift.mean()
backup = plate_appearances
plate_appearances = backup
```
### Create dummies
```
#plate_appearances.pitch_name.value_counts()
# pitch_name. does it somehow matter what type of pitch was hit in the past PA?
# removing for now. it's only useful if lagged and sounds unlikely to be helpful.
# throw out low-prevalence pitches to keep dimensionality down.
#dummies = pd.get_dummies(plate_appearances['pitch_name']).rename(columns=lambda x: 'pitch_' + str(x))
#plate_appearances = pd.concat([plate_appearances, dummies], axis=1)
#plate_appearances.drop(['pitch_name', 'pitch_Fastball', 'pitch_Unknown',
# 'pitch_Pitch Out', 'pitch_Screwball', 'pitch_Forkball',
# 'pitch_Eephus'], inplace=True, axis=1)
dummies = pd.get_dummies(plate_appearances['team_pitching']).rename(columns=lambda x: 'defense_' + str(x))
plate_appearances = pd.concat([plate_appearances, dummies], axis=1)
dummies = pd.get_dummies(plate_appearances['team_batting']).rename(columns=lambda x: 'atbat_' + str(x))
plate_appearances = pd.concat([plate_appearances, dummies], axis=1)
plate_appearances.drop(['team_pitching', 'team_batting', 'home_team', 'away_team',
'inning_topbot'], inplace=True, axis=1)
# todo: group by batter, take max of pa count, select if count < 100. drop if batter in that list of group indexes.
# batter
# note: avoid dropping this id and the pitcher id until required. need these intact to get names back post-modeling.
dummies = pd.get_dummies(plate_appearances['batter']).rename(columns=lambda x: 'batterid_' + str(x))
plate_appearances = pd.concat([plate_appearances, dummies], axis=1)
# pitcher
dummies = pd.get_dummies(plate_appearances['pitcher']).rename(columns=lambda x: 'pitcherid_' + str(x))
plate_appearances = pd.concat([plate_appearances, dummies], axis=1)
# pitcher_throws
plate_appearances['pitcher_throws_left'] = np.where(plate_appearances['p_throws'] == 'L', 1, 0)
# stand (batter_bats)
plate_appearances['left_handed_batter'] = np.where(plate_appearances['stand'] == 'L', 1, 0)
# bb type
dummies = pd.get_dummies(plate_appearances['bb_type']).rename(columns=lambda x: 'bb_type_' + str(x))
plate_appearances = pd.concat([plate_appearances, dummies], axis=1)
plate_appearances.drop(['bb_type'], inplace=True, axis=1)
# month dummies. maybe this changes throughout the season.
# don't drop the original columns in case a continuous relatioship ends up being more meaningful.
plate_appearances['Month'] = plate_appearances['game_date'].dt.month
dummies = pd.get_dummies(plate_appearances['Month']).rename(columns=lambda x: 'Month_' + str(x))
plate_appearances = pd.concat([plate_appearances, dummies], axis=1)
# let's create year dummies while we're at it
dummies = pd.get_dummies(plate_appearances['game_year']).rename(columns=lambda x: 'Year_' + str(x))
plate_appearances = pd.concat([plate_appearances, dummies], axis=1)
# events
dummies = pd.get_dummies(plate_appearances['events']).rename(columns=lambda x: 'event_' + str(x))
plate_appearances = pd.concat([plate_appearances, dummies], axis=1)
# omitting triple because there's never been one. python throws errors.
plate_appearances['onbase'] = plate_appearances.event_single + plate_appearances.event_single + plate_appearances.event_double + plate_appearances.event_triple
plate_appearances['hit'] = plate_appearances.event_single + plate_appearances.event_double \
+ plate_appearances.event_triple + plate_appearances.event_home_run
plate_appearances['successful_outcome_defense'] = plate_appearances.event_field_out + plate_appearances.event_strikeout + plate_appearances.event_grounded_into_double_play \
+ plate_appearances.event_double_play + plate_appearances.event_fielders_choice_out + plate_appearances.event_other_out \
+ plate_appearances.event_triple_play
plate_appearances['successful_shift'] = plate_appearances['is_shift'] * plate_appearances['successful_outcome_defense']
```
### impute missing values
```
drops = ['launch_angleold', 'launch_speedold', 'hit_distance_scold']
plate_appearances = plate_appearances.drop(drops, 1)
#for c in plate_appearances.columns:
# print(plate_appearances[c].head())
# simple imputations: hit location, hit_distance_sc, launch_speed, launch_angle, effective_speed,
# estimated_woba_using_speedangle, babip_value, iso_value
plate_appearances.loc[pd.isna(plate_appearances.hit_location), 'hit_location'] = 0
plate_appearances.loc[pd.isna(plate_appearances.hit_distance_sc), 'hit_distance_sc'] = 0
plate_appearances.loc[pd.isna(plate_appearances.launch_speed), 'launch_speed'] = 0
plate_appearances.loc[pd.isna(plate_appearances.launch_angle), 'launch_angle'] = 0
plate_appearances.loc[pd.isna(plate_appearances.effective_speed), 'effective_speed'] = 0
plate_appearances.loc[pd.isna(plate_appearances.estimated_woba_using_speedangle), 'estimated_woba_using_speedangle'] = 0
plate_appearances.loc[pd.isna(plate_appearances.babip_value), 'babip_value'] = 0
plate_appearances.loc[pd.isna(plate_appearances.iso_value), 'iso_value'] = 0
plate_appearances.loc[pd.isna(plate_appearances.woba_denom), 'woba_denom'] = 1
plate_appearances.loc[pd.isna(plate_appearances.launch_speed_angle), 'launch_speed_angle'] = 0
plate_appearances.shape
backup = plate_appearances
```
### Create lag variables to capture recent outcomes (did the shift last time? did it work?)
```
backup = plate_appearances
plate_appearances = backup
plate_appearances = plate_appearances.sort_values(['batter', 'game_date', 'at_bat_number'], ascending=True)
cols_to_lag = ['is_shift', 'onbase', 'hit', 'successful_outcome_defense', 'successful_shift',
'woba_value', 'launch_speed', 'launch_angle', 'hit_distance_sc', 'bb_type_popup', 'bb_type_line_drive',
'bb_type_ground_ball', 'bb_type_fly_ball']
# how many PAs back to we want to consider?
lag_time = 5
for col in cols_to_lag:
for time in range(lag_time):
feature_name = col + '_lag' + '_{}'.format(time+1)
plate_appearances[feature_name] = plate_appearances.groupby('batter')[col].shift(time+1)
#plate_appearances['shift_lag_1'] = plate_appearances.groupby('batter')['is_shift'].shift(1)
#plate_appearances['shift_lag_2'] = plate_appearances.groupby('batter')['is_shift'].shift(2)
#plate_appearances.loc[plate_appearances.batter_name=='ryan howard',['is_shift','hit','successful_shift',
# 'shift_lag_1', 'shift_lag_2']].head(20)
plate_appearances.loc[plate_appearances.batter_name=='ryan howard',].head()
plate_appearances.shape
#[print(c) for c in plate_appearances.columns]
#[print(plate_appearances[c].head()) for c in plate_appearances.columns]
#now drop everythign that can't go into the final model. leave pitcher/batter/game_pk in for now so we don't lose that info.
# make sure to drop everything that includes info we don't know yet at beginnign of atbat (eg events, launch angle)
drops = ['index', 'atbat_pk', 'game_date', 'stand', 'p_throws', 'hit_location', 'effective_speed', 'estimated_woba_using_speedangle',
'woba_denom', 'babip_value', 'iso_value', 'launch_speed_angle', 'pitch_number', 'key_mlbam', 'batter_name',
'events', 'woba_value', 'successful_shift', 'hit', 'onbase', 'event_walk', 'event_triple_play', 'event_triple',
'event_strikeout_double_play', 'event_strikeout', 'event_single', 'event_sac_fly_double_play', 'event_sac_fly',
'event_sac_bunt_double_play', 'event_sac_bunt', 'event_run', 'event_pickoff_caught_stealing_home',
'event_pickoff_caught_stealing_3b', 'event_pickoff_caught_stealing_2b', 'event_pickoff_3b', 'event_pickoff_2b',
'event_pickoff_1b', 'event_other_out', 'event_intent_walk', 'event_home_run', 'event_hit_by_pitch', 'event_grounded_into_double_play',
'event_force_out', 'event_fielders_choice_out', 'event_fielders_choice', 'event_field_out', 'event_field_error',
'event_double_play', 'event_double', 'event_caught_stealing_home', 'event_caught_stealing_3b', 'event_caught_stealing_2b',
'event_catcher_interf', 'event_batter_interference', 'bb_type_popup', 'bb_type_line_drive', 'bb_type_ground_ball',
'bb_type_fly_ball', 'pitch_name']
plate_appearances = plate_appearances.drop(drops, 1)
plate_appearances.shape
# check for nans I've missed
# just the lags
#for c in plate_appearances.columns:
# print(c)
# print(plate_appearances[c].isnull().sum())
# drop nulls created by lag variables (5 nulls per batter. amounts to rougbly 5k observations
# in other words, 0.8% of rows lost to creating 5 PA worth of lag columns
print(plate_appearances.shape)
plate_appearances = plate_appearances.dropna()
print(plate_appearances.shape)
```
# Modeling
Logistic regression benchmark vs. a simple random forest model. Evaluate using accuracy and AUC score.
I'm starting from these sklearn docs: http://scikit-learn.org/stable/auto_examples/ensemble/plot_feature_transformation.html#sphx-glr-auto-examples-ensemble-plot-feature-transformation-py
From there I will improve this to use cross validation to tune some parameters (number of estimators, tree depth), and maybe experiment with some ensembling (RT + logit + nn may be interesting)
```
np.random.seed(10)
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegressionCV
from sklearn.preprocessing import OneHotEncoder
from sklearn.model_selection import train_test_split
from sklearn.metrics import roc_curve
from sklearn.pipeline import make_pipeline
from sklearn.metrics import roc_auc_score
import matplotlib.pyplot as plt
%matplotlib inline
train_percent = .7
train_samples = int((plate_appearances.shape[0] * .9) * train_percent)
test_samples = int((plate_appearances.shape[0] * .9) * (1 - train_percent))
holdout = int((plate_appearances.shape[0] * .9) * .1)
batters = plate_appearances['batter']
pitchers = plate_appearances['pitcher']
y = plate_appearances['is_shift']
X = plate_appearances.drop(['is_shift', 'batter', 'pitcher'], 1)
X_train = X[:train_samples]
X_test = X[train_samples:train_samples+test_samples]
X_holdout = X[train_samples+test_samples:]
y_train = y[:train_samples]
y_test = y[train_samples:train_samples+test_samples]
y_holdout = y[train_samples+test_samples:]
print('X train, test, holdout shapes: ')
print(X_train.shape)
print(X_test.shape)
print(X_holdout.shape)
print('y train, test, holdout shapes: ')
print(y_train.shape)
print(y_test.shape)
print(y_holdout.shape)
n_estimator = 100
rf = RandomForestClassifier(max_depth=3, n_estimators=n_estimator, n_jobs=3, verbose=2)
#rf_enc = OneHotEncoder()
rf.fit(X_train, y_train)
#rf_enc.fit(rf.apply(X_train))
# a simple logistic benchmark. only knows how often defense shifts and how often batter is shifted against.
lr = LogisticRegressionCV(n_jobs=3)
lr.fit(X_train[['avg_shifted_against', 'def_shift_pct']], y_train)
y_pred_rf = rf.predict_proba(X_test)[:, 1]
fpr_rf, tpr_rf, _ = roc_curve(y_test, y_pred_rf)
y_pred_lr = lr.predict_proba(X_test[['avg_shifted_against', 'def_shift_pct']])[:, 1]
fpr_lr, tpr_lr, _ = roc_curve(y_test, y_pred_lr)
plt.xlim(0, 1)
plt.ylim(0, 1)
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr_rf, tpr_rf, label='RF')
plt.plot(fpr_lr, tpr_lr, label='Logit')
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.title('ROC curve')
plt.legend(loc='best')
plt.show()
# pretty good!
print("Random Forest AUC Score:")
roc_auc_score(y_test, y_pred_rf)
print("Logit AUC Score:")
roc_auc_score(y_test, y_pred_lr)
# let's see what was most important
# https://stackoverflow.com/questions/44101458/random-forest-feature-importance-chart-using-python
#features = X_test.columns
#importances = rf.feature_importances_
#indices = np.argsort(importances)
feats = {} # a dict to hold feature_name: feature_importance
for feature, importance in zip(X_test.columns, rf.feature_importances_):
feats[feature] = importance #add the name/value pair
importances = pd.DataFrame.from_dict(feats, orient='index').rename(columns={0: 'Gini-importance'})
importances.sort_values(by='Gini-importance', ascending=False)[0:50]
score = pd.DataFrame(y_test)
score['pred_prob_rf'] = y_pred_rf
score['pred_prob_lr'] = y_pred_lr
score['pred_rf'] = np.where(score['pred_prob_rf']>=.25, 1, 0)
score['pred_lr'] = np.where(score['pred_prob_lr']>=.25, 1, 0)
score['pred_rf'].sum()
from sklearn.metrics import accuracy_score
# a threshold of .25 works well
# some helpful discussion on thresholds: https://stats.stackexchange.com/questions/312119/classification-probability-threshold
print('for random forest:')
for threshold in [0.05, .1, .15, .2, .25, .3, .35, .4, .45, .5]:
print(accuracy_score(score['is_shift'], np.where(score['pred_prob_rf']>=threshold, 1, 0)))
print('for logistic regression: ')
for threshold in [0.05, .1, .15, .2, .25, .3, .35, .4, .45, .5]:
print(accuracy_score(score['is_shift'], np.where(score['pred_prob_lr']>=threshold, 1, 0)))
print("RF Accuracy: ")
print(accuracy_score(score['is_shift'], np.where(score['pred_prob_rf']>=.25, 1, 0)))
print('Logit Accuracy:')
print(accuracy_score(score['is_shift'], np.where(score['pred_prob_lr']>=.4, 1, 0)))
score['is_false_positive'] = np.where((score['pred_rf']==1) & (score['is_shift']==0), 1, 0)
score['is_false_negative'] = np.where((score['pred_rf']==0) & (score['is_shift']==1), 1, 0)
score['is_true_positive'] = np.where((score['pred_rf']==1) & (score['is_shift']==1), 1, 0)
score['is_true_negative'] = np.where((score['pred_rf']==0) & (score['is_shift']==0), 1, 0)
print('Accuracy Metrics for Random Forest')
score[['is_false_positive', 'is_false_negative', 'is_true_positive', 'is_true_negative']].mean()
score['is_false_positive'] = np.where((score['pred_lr']==1) & (score['is_shift']==0), 1, 0)
score['is_false_negative'] = np.where((score['pred_lr']==0) & (score['is_shift']==1), 1, 0)
score['is_true_positive'] = np.where((score['pred_lr']==1) & (score['is_shift']==1), 1, 0)
score['is_true_negative'] = np.where((score['pred_lr']==0) & (score['is_shift']==0), 1, 0)
print('Accuracy Metrics for Logistic Regression')
score[['is_false_positive', 'is_false_negative', 'is_true_positive', 'is_true_negative']].mean()
```
| github_jupyter |
<a href="https://qworld.net" target="_blank" align="left"><img src="../qworld/images/header.jpg" align="left"></a>
$$
\newcommand{\set}[1]{\left\{#1\right\}}
\newcommand{\abs}[1]{\left\lvert#1\right\rvert}
\newcommand{\norm}[1]{\left\lVert#1\right\rVert}
\newcommand{\inner}[2]{\left\langle#1,#2\right\rangle}
\newcommand{\bra}[1]{\left\langle#1\right|}
\newcommand{\ket}[1]{\left|#1\right\rangle}
\newcommand{\braket}[2]{\left\langle#1|#2\right\rangle}
\newcommand{\ketbra}[2]{\left|#1\right\rangle\left\langle#2\right|}
\newcommand{\angleset}[1]{\left\langle#1\right\rangle}
$$
# Spectral Theory
_prepared by Israel Gelover_
### <a name="remark_2_17">Remark 2.17</a> Spectral Theory
Given $A:\mathcal{H} \to \mathcal{H}$ a linear operator, we can consider $A_\lambda = A - \lambda I$ with $\lambda \in \mathbb{C}$. The study of the distribution of the values of $\lambda$ for which $A_\lambda$ whether has or does not have an inverse is called _Spectral Theory_ of the operator.
In quantum computing we will only work with finite-dimensional operators and this type of operators only have one type of spectrum, which are the eigenvalues of the operator.
Let us recall that to obtain the eigenvalues of a matrix $A$ we calculate $\det(A - \lambda I) = 0$. That is, we look for the values of $\lambda$ for which the matrix $A - \lambda I$ has no inverse. Meaning that the eigenvalues are a particular case of the spectrum of an operator.
### <a name="definition_2_18">Definition 2.18</a> Eigenvalues and Eigenvectors
Let $A:\mathcal{H} \to \mathcal{H}$ be a linear operator
1. The set $P(A) = \set{\lambda \in \mathbb{C} \mid A\ket{f} = \lambda\ket{f} \text{ for some } \ket{f} \neq 0}$ is called the point spectrum of $A$, and its elements are known as _Eigenvalues_ of $A$. **Remark:** If $\mathcal{H}$ is of finite dimension, then $P(A) = \set{\text{solutions of } \det(A - \lambda I) = 0}$
2. Let $\lambda_0 \in P(A)$, if $A\ket{f} = \lambda_0 \ket{f}$ has non-trivial solutions, that is, $\lambda_0$ is an eigenvalue, we will denote by$\mathcal{L}(\lambda_0)$ the subspace of $\mathcal{H}$ generated by the solutions
of $A\ket{f} = \lambda_0 \ket{f}$.
3. $\dim \set{\mathcal{L}(\lambda_0)}$ is called the multiplicity of $\lambda_0$. Let us recall that when finding the non-trivial solutions of $\det(A - \lambda I) = 0$ we can find that the monomial associated with an eigenvalue appears more than once in the characteristic polynomial of $A$, this number of repetitions is what we know as the multiplicity of the eigenvalue.
Intuitively we can interpret that finding the eigenvalues of an operator has the geometric equivalence of finding subspaces that remain invariant under the application of the operator. For example, if we have a linear operator $A:\mathbb{R}^2 \to \mathbb{R}^2$ and we find the eigenvalues of $A$, this would be equivalent to finding a line that is fixed with respect to the application of the operator $A$.
### Task 1.
(On paper) Let $A:\mathcal{H} \to \mathcal{H}$ be a self-adjoint operator, (i.e $A=A^\dagger$).
If $A\ket{f} = \lambda\ket{f}$, use <a href="./HilbertSpace.ipynb#definition_2_2">Definition 2.2</a> and <a href="./LinearOperators.ipynb#definition_2_12">Definition 2.12</a> to conclude that the eigenvalues of an autoadjoint operator are real numbers.
<a href="./SpectralTheory_Solutions.ipynb#task_1">Click here for solution</a>
### Task 2.
(On paper) Let $U:\mathcal{H} \to \mathcal{H}$ be a unitary operator.
If $U\ket{f} = \lambda\ket{f}$, calculate $\braket{f}{f}$ to conclude that the eigenvalues of a unit operator are complex numbers with norm one.
<a href="./SpectralTheory_Solutions.ipynb#task_2">Click here for solution</a>
### <a name="proposition_2_19">Proposition 2.19</a>
Let $A:\mathcal{H} \to \mathcal{H}$ be a self-adjoint operator, then the eigenvectors corresponding to different eigenvalues are orthogonal.
**Remark:** From this proposition the _Spectral Theorem_ can be derived, which states that there exists an orthonormal basis of the space consisting of eigenvectors of the self-adjoint operator.
**Proof:**
Let $\lambda_1, \lambda_2$ be different eigenvalues, that is $\lambda_1 \neq \lambda_2$, such that
\begin{equation*}
A\ket{f_1} = \lambda_1\ket{f_1} \enspace \text{ and } \enspace A\ket{f_2} = \lambda_2\ket{f_2}
\end{equation*}
So, on the one hand we have
\begin{equation*}
\ket{f_1}\cdot (A\ket{f_2}) = \ket{f_1} \cdot (\lambda_2\ket{f_2}) = \lambda_2(\ket{f_1} \cdot \ket{f_2})
\end{equation*}
On the other hand
\begin{equation*}
(A\ket{f_1}) \cdot \ket{f_2} = (\lambda_1\ket{f_1}) \cdot \ket{f_2} = \lambda_1^*(\ket{f_1} \cdot \ket{f_2})
\end{equation*}
Since $A$ is a self-adjoint operator, we know that $\ket{f_1}\cdot (A\ket{f_2}) = (A\ket{f_1}) \cdot \ket{f_2}$ and also $\lambda_1, \lambda_2 \in \mathbb{R}$, then $\lambda_1\braket{f_1}{f_2} = \lambda_2\braket{f_1}{f_2}$.
Therefore $\braket{f_1}{f_2} = 0$, since $\lambda_1 \neq \lambda_2$.
### <a name="definition_2_19">Definition 2.19</a> Outer Product
Let $\ket{f}, \ket{g} \in \mathcal{H}$ Hilbert space, the _Outer Product_ of $\ket{f}$ and $\ket{g}$ is a linear operator such that
\begin{equation*}
\begin{split}
\ketbra{f}{g}:\mathcal{H} &\to \mathcal{H} \\
\ket{x} &\to \ket{f}\braket{g}{x}
\end{split}
\end{equation*}
That is, applying the outer product of $\ket{f}$ and $\ket{g}$ to an arbitrary vector $\ket{x}$, is equivalent to calculate the inner product of $\ket{g}$ with $\ket{x}$ and multiplying the resulting scalar by $\ket{f}$.
### <a name="examples">Examples</a>
1. Let $\ket{\psi} = \frac{1}{\sqrt{2}}(\ket{0} + \ket{1})$ and $\ket{\phi} = \ket{0}$, then
\begin{equation*}
\begin{split}
\ketbra{\psi}{\phi} &= \frac{1}{\sqrt{2}}(\ket{0}+\ket{1})\bra{0} = \frac{1}{\sqrt{2}}(\ketbra{0}{0}+\ketbra{1}{0}) \\
&= \frac{1}{\sqrt{2}}\left[\begin{pmatrix} 1 \\ 0 \end{pmatrix} \begin{pmatrix} 1 & 0 \end{pmatrix} + \begin{pmatrix} 0 \\ 1 \end{pmatrix} \begin{pmatrix} 1 & 0 \end{pmatrix}\right] \\
&= \frac{1}{\sqrt{2}}\left[\begin{pmatrix} 1 & 0 \\ 0 & 0 \end{pmatrix} + \begin{pmatrix} 0 & 0 \\ 1 & 0 \end{pmatrix}\right] = \frac{1}{\sqrt{2}}\begin{pmatrix} 1 & 0 \\ 1 & 0 \end{pmatrix}
\end{split}
\end{equation*}
Let us recall that **Definition 2.20** tells us that the outer product of two vectors is a linear operator, in this case we have the outer product of two qubits and therefore we obtain a two-dimensional operator, that is, we obtain a $2 \times 2$ matrix. Note that from the expression of the outer product using Dirac notation we can deduce how it looks like in matrix notation, as follows:
\begin{equation}\label{dirac_matriz}
\ketbra{\psi}{\phi} = \frac{1}{\sqrt{2}}(\overbrace{\ket{0}}^\text{row} \underbrace{\bra{0}}_\text{column} + \overbrace{\ket{1}}^\text{row} \underbrace{\bra{0}}_\text{column})
\end{equation}
This tells us that the associated matrix will have $\frac{1}{\sqrt{2}}$ in row $0$ - column $0$ and in row $1$ - column $0$ and zeros in the rest of the positions, the same as we obtained when calculating using vector notation:
\begin{equation*}
\ketbra{\psi}{\phi} = \frac{1}{\sqrt{2}}(\ketbra{0}+\ketbra{1}{0}) = \frac{1}{\sqrt{2}}\begin{pmatrix} 1 & 0 \\ 1 & 0 \end{pmatrix}
\end{equation*}
2. Let $\ket{\psi}, \ket{\phi}$ be as in the previous example, then
\begin{equation*}
\begin{split}
(\ketbra{\psi}{\phi})\ket{1} = \ket{\psi}\braket{\phi}{1} &= \frac{1}{\sqrt{2}}(\ket{0} + \ket{1})\braket{0}{1} = 0 \\
&= \frac{1}{\sqrt{2}}\begin{pmatrix} 1 & 0 \\ 1 & 0 \end{pmatrix}\begin{pmatrix} 0 \\ 1 \end{pmatrix} = \frac{1}{\sqrt{2}}\begin{pmatrix} 0 \\ 0 \end{pmatrix} = \begin{pmatrix} 0 \\ 0 \end{pmatrix}
\end{split}
\end{equation*}
In both cases we obtain vector **0**, but clearly, Dirac notation is much more practical and therefore we will prefer it in the rest of the notebooks.
### <a name="definition_2_21">Definition 2.21</a> Completeness Relation
Let $\set{\ket{u_i}}_{i=1}^{n}$ be an orthonormal basis of a space $\mathcal{H} \implies \sum_{i=1}^{n}\ketbra{u_i}{u_i} = I$
Let us recall that the outer product of two vectors is a linear operator, and since the sum of linear operators is again a linear operator, in this case we obtain the identity operator.
**Proof:**
Let $\ket{\psi} \in \mathcal{H}$ be a vector, we can express it in terms of the basis as $\ket{\psi} = \sum_{j=1}^{n}c_j\ket{u_j}$, with $c_j \in \mathbb{F}$ elements of the field. Thus
\begin{equation*}
\begin{split}
\left(\sum_{i=1}^{n}\ketbra{u_i}{u_i}\right)\ket{\psi} &= \sum_{i=1}^{n}\ket{u_i}\braket{u_i}{\psi} \\
&= \sum_{i=1}^{n}\ket{u_i} \sum_{j=1}^{n}c_j \braket{u_i}{u_j} = \sum_{i=1}^{n}c_i\ket{u_i} = \ket{\psi}
\end{split}
\end{equation*}
since $\braket{u_i}{u_j}$ takes value $1$ when $i=j$ and $0$ when $i \neq j$, as they are elements of an orthonormal basis.
| github_jupyter |
# ORF307 Homework 7
Due: Friday, April 9, 2021 9:00 pm ET
- Please export your code with output as pdf.
- If there is any additional answers, please combine them as **ONE** pdf file before submitting to the Gradescope.
# Q1 Parametrized primal and dual LPs {-}
Consider the parameterized primal and dual LPs
$$
\begin{array}{ll}
\mbox{minimize} &(c +\epsilon d)^Tx\\
\text{subject to} & Ax \leq b\\
&\\
\end{array}\qquad
\begin{array}{ll}
\mbox{maximize} &-b^Ty\\
\text{subject to} & A^Ty + c + \epsilon d = 0\\
& y \ge 0
\end{array}
$$
where
$$
A = \begin{bmatrix}
1 & 3\\
1 & 1
\end{bmatrix},
\quad\quad
b = \begin{bmatrix}
4\\
4
\end{bmatrix},
\quad\quad
c = \begin{bmatrix}
-1\\
-2
\end{bmatrix},
\quad\quad
d = \begin{bmatrix}
2 \\
1
\end{bmatrix}
$$
(a) Prove that $x^\star= (4, 0)$ and $y^\star = (0.5, 0.5)$ are optimal when $\epsilon = 0$.
(b) How does $p^\star(\epsilon)$ vary as a function of $\epsilon$ around $\epsilon=0$? Give an explicit expression for $p^\star(\epsilon)$.
(c) Also give an explicit expression for the primal and dual optimal solutions for values of
$\epsilon$ around $\epsilon=0$.
*Hint:* The dual looks very similar to a primal LP in standard form.
# Q2 Chocolate manufacturing company {-}
Consider a chocolate manufacturing company that produces three types of chocolate – A, B, and C. All three chocolates require cocoa beans, sugar, milk. To manufacture each unit of A, B, and C, the following quantities are required:
- Each unit of A requires 8 unit of cocoa beans, 4 unit of sugar, 2 unit of milk.
- Each unit of B requires 6 unit of cocoa beans, 2 unit of sugar, 1.5 unit of milk.
- Each unit of C requires 1 unit of cocoa beans, 1.5 unit of sugar, 0.5 unit of milk.
The company has a total of 48 units of cocoa beans, 20 units of sugar, and 8 unit of milk. On each sale, the company makes a profit of
- 60 per unit A sold.
- 30 per unit B sold.
- 20 pre unit C sold.
Now the company wants to increase the supply of cocoa beans, sugar, and milk by $u = (t, t, 5t)$ units, which corresponds to $td$ where $d = (1, 1, 5)$. The company wished to maximize its profit after the supply increase.
(a) Define problem as LP.
(b) Solve the problem for various values of $t$ using cvxpy. You can set $t$ as a parameter as follows:
```python
x = cp.Variable(n)
t = cp.Parameter()
# Formulate problem ...
# problem = cp.Problem(....)
# Solve for all values of t
t_values = np.linspace(-1, 1, 100)
x_vector = []
objective_vector = []
for t_val in t_values:
t.value = t_val
problem.solve()
x_vector.append(x.value)
objective_vector.append(problem.objective.value)
```
(c) Plot the optimal value $p^\star(td)$ and the lower bound defined around $p^\star(0)$
# Q3 Network flow problem {-}
Consider the uncapacitated (i.e., no capacity constraints) network flow problem shown below. The label next to each arc is the cost of sending one unit of flow along that arc, and the label in each node is the number of units of supply available at each node (where less than zero corresponds to a demand). Your goal is to transport the goods from the suppliers to the consumers at minimum cost.

(a) formulate the uncapacitated network flow problem as network flow and give $A$ matrix.
(b) Solve it using cvxpy.
| github_jupyter |
```
import time
#time.sleep(60*60*3)
import os
os.environ["TF_GPU_THREAD_MODE"] = "gpu_private"
import os
# --- Uncomment to use only CPU (e.g. GPU memory is too small)
# os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID" # see issue #152
# os.environ["CUDA_VISIBLE_DEVICES"] = "-1"
import tensorflow as tf
physical_devices = tf.config.list_physical_devices('GPU')
gpu_devices = tf.config.experimental.list_physical_devices("GPU")
tf.config.experimental.set_memory_growth(physical_devices[0], True)
for device in gpu_devices:
tf.config.experimental.set_memory_growth(device, True)
from tensorflow.compat.v1 import ConfigProto
from tensorflow.compat.v1 import InteractiveSession
config = ConfigProto()
config.gpu_options.allow_growth = True
session = InteractiveSession(config=config)
from tensorflow import keras
from tensorflow.python.data import AUTOTUNE
tf.test.is_gpu_available(cuda_only=True)
# reate logger - nicer formatting
import logging
logger = logging.getLogger()
logging._warn_preinit_stderr = 0
logger.setLevel(logging.INFO)
formatter = logging.Formatter('%(asctime)s: %(message)s')
ch = logging.StreamHandler()
ch.setFormatter(formatter)
logger.handlers = [ch]
logger.info("Logger started.")
# Coppied from pickle2sqlite.ipynb
import itertools
import math
import os
import pickle
import numpy as np
def angle_to_sin_cos(angle):
return [math.sin(angle), math.cos(angle)]
def sin_cos_to_angle(sin, cos):
return math.atan2(sin, cos)
def angle_to_sin_and_back(angle):
return math.atan2(*angle_to_sin_cos(angle))
def to_abs_with_sin_cos(arr):
result = list()
for box in arr:
angles = [angle_to_sin_cos(angle) for angle in box[3:]]
result.append(list(box[:3]) + list(itertools.chain(*angles)))
return np.asarray(result)
def process_abs_sample(sample):
""" Changes sample in form of single 6 elements array to houdini compatible vec3 pos and vec4 orient """
orient = [sin_cos_to_angle(sample[3+2*x], sample[4+2*x]) for x in range(3)] + [1]
return sample[:3], orient
class DataProcessor(object):
""" Code responsible for processing of data gathered into a class for readability.
Trying few different approaches to data representation to test what works best:
For every frame n following representations are calculated:
- x_delta: array (4, 6) of delta between frames n and n-1: (Δx, Δy, Δz, Δα , Δβ, Δγ)
- x_abs_n_minus_1: array (2, 4, 9) of absolute values in frame n-1
- x_abs_n: x_abs in in frame n
Structure of x_abs_*: [[x, y, z, sin(α), cos(α), sin(β), cos(β), sin(γ), cos(γ)], [...]]
- x_force: array of forces in frame n: (power, sin(Φ), cos(Φ)
- y_delta: array (4, 6) of delta between frames n and n+1
- y_abs: array (4, 9) of absolute values in frame n+1
Data is passed to neural networks in forms:
- x: (x_delta, x_force) y: y_delta
- x: (x_delta, x_abs_n, x_force) y: y_delta
- x: (x_delta, x_abs_n, x_force) y: y_abs
- x: (x_abs_n_minus_1, x_abs_n, x_force) y: y_abs
- x: (x_abs_n_minus_1, x_abs_n, x_force) y: y_delta
"""
def __init__(self, paths):
# self.x_delta = np.empty((1, 4, 6))
# self.x_abs_n_minus_1 = np.empty((1, 4, 9))
# self.x_abs_n = np.empty((1, 4, 9))
# self.x_force = np.empty((1, 3))
#
# self.y_delta = np.empty((1, 4, 6))
# self.y_abs = np.empty((1, 4, 9))
self.x_delta = list()
self.x_abs_n_minus_1 = list()
self.x_abs_n = list()
self.x_force = list()
self.y_delta = list()
self.y_abs = list()
for num, file_path in enumerate(paths):
print("{} / {}".format(num, len(file_paths)), end="\r")
with open(file_path, "rb") as f:
frames = pickle.load(f)
filename = os.path.basename(file_path)
force, force_angle = [float(part) for part in filename.split("_")]
last_frame = None
for num, (frame, next_frame) in enumerate(zip(frames[:-1], frames[1:])):
# print(num, end="\r")
if num <=2:
frame_force = 0
else:
frame_force = force
if last_frame is not None:
x_force = np.asarray(angle_to_sin_cos(force_angle) + [frame_force])
n_minus_1_abs = np.asarray([f[0] + f[1][:-1] for f in last_frame])
n_abs = np.asarray([f[0] + f[1][:-1] for f in frame])
n_plus_1_abs = np.asarray([f[0] + f[1][:-1] for f in next_frame])
x_delta = n_abs - n_minus_1_abs
y_delta = n_plus_1_abs - n_abs
n_minus_1_abs = to_abs_with_sin_cos(n_minus_1_abs)
n_abs = to_abs_with_sin_cos(n_abs)
y_abs = to_abs_with_sin_cos(n_plus_1_abs)
self.x_delta.append(x_delta)
self.x_abs_n_minus_1.append(n_minus_1_abs)
self.x_abs_n.append(n_abs)
self.x_force.append(x_force)
self.y_delta.append(y_delta)
self.y_abs.append(y_abs)
last_frame = frame
self.x_delta = np.asarray(self.x_delta)
self.x_abs_n_minus_1 = np.asarray(self.x_abs_n_minus_1)
self.x_abs_n = np.asarray(self.x_abs_n)
self.x_force = np.asarray(self.x_force)
self.y_delta = np.asarray(self.y_delta)
self.y_abs = np.asarray(self.y_abs)
from tensorflow.data import Dataset
data_dir = os.path.normpath("samplesGeneration/data")
file_names = os.listdir(data_dir)
file_paths = [os.path.join(data_dir, file_name) for file_name in file_names]
dp = DataProcessor(file_paths)
from tensorflow.keras.callbacks import EarlyStopping, TensorBoard, ModelCheckpoint
early_stopping = EarlyStopping(monitor='val_loss', patience=4)
models_dir = os.path.join(os.getcwd(), "samplesGeneration", "logs")
from tensorflow.keras import Input, Model
from tensorflow.keras.layers import Dense, Dropout, Flatten, Reshape, Concatenate
from tensorflow.keras.layers import LSTM, GRU
import time
import tensorflow.keras.backend as K
def special_loss(y_true, y_pred):
custom_loss=tf.add(tf.square(y_true-y_pred)/tf.add(tf.square(y_pred), 0.1), tf.k.losses.MSE(y_true, y_pred))
return custom_loss
def loss(y_true,y_pred):
return K.sum(K.square(y_pred - y_true) * K.square(100 * y_true) + K.square(y_pred - y_true))
# physical_devices = tf.config.experimental.list_physical_devices('GPU')
# tf.config.experimental.set_memory_growth(physical_devices[0], True)
def create_datasets(dp, y, batch_size=32):
number_of_samples = len(dp.x_abs_n)
indexes = np.arange(number_of_samples)
np.random.shuffle(indexes)
split = int(number_of_samples * 0.9)
val_length = number_of_samples - split
train_indexes = indexes[:split]
val_indexes = indexes[split:]
ds = tf.data.Dataset.from_tensor_slices(({"x_abs_n_minus_1": dp.x_abs_n_minus_1[train_indexes].reshape(split, 1, 4, -1),
"x_abs_n": dp.x_abs_n[train_indexes].reshape(split, 1, 4, -1),
"x_delta": dp.x_delta[train_indexes].reshape(split, 1, 4, -1),
"x_force": dp.x_force[train_indexes].reshape(split, 1, -1),
}, y[train_indexes].reshape(split, 1, 4, -1)))
ds.cache()
ds.shuffle(buffer_size=1000)
ds.batch(batch_size, drop_remainder=True)
ds.prefetch(buffer_size=AUTOTUNE)
val_ds = tf.data.Dataset.from_tensor_slices(({"x_abs_n_minus_1": dp.x_abs_n_minus_1[val_indexes].reshape(val_length, 1, 4, -1),
"x_abs_n": dp.x_abs_n[val_indexes].reshape(val_length, 1, 4, -1),
"x_delta": dp.x_delta[val_indexes].reshape(val_length, 1, 4, -1),
"x_force": dp.x_force[val_indexes].reshape(val_length, 1, -1),
}, y[val_indexes].reshape(val_length, 1, 4, -1)))
val_ds.cache()
val_ds.shuffle(buffer_size=1000)
val_ds.batch(batch_size, drop_remainder=True)
val_ds.prefetch(buffer_size=AUTOTUNE)
return ds, val_ds
def create_and_train_network(dp, y, model_id=0, epohs=10, batch_size=32, model_generator=None):
y_shape = y[0].shape
log_dir = os.path.join(os.getcwd(), "samplesGeneration", "logs_2", "model_{}_{}_{}_{}".format(y.shape, y_shape, model_id, time.time()))
if not os.path.isdir(log_dir):
os.makedirs(log_dir)
tensorboard_callback = TensorBoard(log_dir=log_dir, histogram_freq=1, profile_batch = '500,520')
best_model_file = "{}_{}_{}_model.h5".format(y.shape, y_shape, model_id)
best_model = ModelCheckpoint(best_model_file, monitor='val_loss', mode='min',verbose=1, save_best_only=True)
class CustomSaveCallback(keras.callbacks.Callback):
def on_test_begin(self, logs=None):
self.model.save("model\\model_{}_{}_{}_{}".format(y.shape, y_shape, model_id, time.time()), save_format='tf')
my_cb = CustomSaveCallback()
train_dataset, validation_dataset = create_datasets(dp, y, batch_size)
def create_model0():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_delta_flat = Flatten()(input_delta)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
x = Concatenate()([input_delta_flat, input_abs_flat, input_force_flat])
x = Dense(64, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(128, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(128, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(np.prod(y_shape))(x)
output_1 = Reshape(y_shape)(x)
model = Model(inputs=[input_delta, input_abs, input_force], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
def create_model1():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_delta_flat = Flatten()(input_delta)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
x = Dense(64, activation='relu')(input_force_flat)
x = Dropout(0.2)(x)
x = Concatenate()([x, input_delta_flat])
x = Dense(128, activation='relu')(x)
x = Dropout(0.2)(x)
x = Concatenate()([x, input_abs_flat])
x = Dense(128, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(np.prod(y_shape))(x)
output_1 = Reshape(y_shape)(x)
model = Model(inputs=[input_delta, input_abs, input_force], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
def create_model2():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_delta_flat = Flatten()(input_delta)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
x = Concatenate()([input_delta_flat, input_abs_flat, input_force_flat])
x = Dense(128, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(128, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(64, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(64, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(np.prod(y_shape))(x)
output_1 = Reshape(y_shape)(x)
model = Model(inputs=[input_delta, input_abs, input_force], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
def create_model3():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs_0 = Input(shape=(4, 9), name='x_abs_n_minus_1')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_abs_0_flat = Flatten()(input_abs_0)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
x = Concatenate()([input_abs_0_flat, input_abs_flat])
x = Reshape((2, 4*9))(x)
x = LSTM(np.prod(y_shape), return_sequences=False, dropout=0.2,
activation="tanh", recurrent_activation="sigmoid", recurrent_dropout=0, unroll=False, use_bias=True
)(x)
x = Dense(np.prod(y_shape))(x)
output_1 = Reshape(y_shape)(x)
model = Model(inputs=[input_abs_0, input_abs, input_force], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
def create_model4():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs_0 = Input(shape=(4, 9), name='x_abs_n_minus_1')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_abs_0_flat = Flatten()(input_abs_0)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
x = Concatenate()([input_abs_0_flat, input_abs_flat])
x = Reshape((2, 4*9))(x)
x = GRU(np.prod(y_shape), return_sequences=False,
activation="tanh", recurrent_activation="sigmoid", recurrent_dropout=0, unroll=False, use_bias=True, reset_after=True
)(x)
x = Flatten()(x)
x = Concatenate()([x, input_force_flat])
x = Dropout(0.2)(x)
x = Dense(np.prod(y_shape))(x)
output_1 = Reshape(y_shape)(x)
model = Model(inputs=[input_abs_0, input_abs, input_force], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
logger.info("Create model...")
if model_generator is not None:
model = model_generator()
elif model_id == 0:
model = create_model0()
elif model_id == 1:
model = create_model1()
elif model_id == 2:
model = create_model2()
elif model_id == 3:
model = create_model3()
elif model_id == 4:
model = create_model4()
model.summary()
logger.info("Fit model..")
network = model.fit(train_dataset, validation_data=validation_dataset, verbose=1,
epochs=epohs, callbacks=[my_cb, early_stopping, best_model, tensorboard_callback])
logger.info("Summary")
model.summary()
models_dir = os.path.join(os.getcwd(), "samplesGeneration", "logs", "model_{}".format(time.time()))
model.save(os.path.join(models_dir, "model"))
return network
logger.info("start")
#training_generator = tf.data.Dataset.from_generator(lambda: training_generator,
# output_types=(tf.float32, tf.float32))
#validation_generator = tf.data.Dataset.from_generator(lambda: training_generator,
# output_types=(tf.float32, tf.float32))
# create_and_train_network(dp, dp.y_delta, 0)
# create_and_train_network(dp, dp.y_delta, 1)
# create_and_train_network(dp, dp.y_delta, 2)
# create_and_train_network(dp, dp.y_delta, 1)
#create_and_train_network(dp, dp.y_abs, 0)
#create_and_train_network(dp, dp.y_abs, 1)
#create_and_train_network(dp, dp.y_abs, 4)
#create_and_train_network(dp, dp.y_delta, 4)
def create_model2_improved():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_delta_flat = Flatten()(input_delta)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
x = Concatenate()([input_delta_flat, input_abs_flat, input_force_flat])
x = Dense(128, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(256, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(512, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(2048, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(512, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(256, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(64, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(np.prod(dp.y_delta[1].shape))(x)
output_1 = Reshape(dp.y_delta[1].shape)(x)
model = Model(inputs=[input_delta, input_abs, input_force], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
# 0.0091
# create_and_train_network(dp, dp.y_delta, 10, model_generator=create_model2_improved)
def create_model2_improved():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_delta_flat = Flatten()(input_delta)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
x = Concatenate()([input_delta_flat, input_abs_flat, input_force_flat])
x = Dense(128, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(256, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(512, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(512, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(512, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(256, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(64, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(np.prod(dp.y_abs[1].shape))(x)
output_1 = Reshape(dp.y_abs[1].shape)(x)
model = Model(inputs=[input_delta, input_abs, input_force], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
#create_and_train_network(dp, dp.y_abs, 5, model_generator=create_model2_improved)
def create_model4_improved():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs_0 = Input(shape=(4, 9), name='x_abs_n_minus_1')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_abs_0_flat = Flatten()(input_abs_0)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
x = Concatenate()([input_abs_0_flat, input_abs_flat])
x = Reshape((2, 4*9))(x)
x = GRU(np.prod(dp.y_delta[0].shape) * 2, return_sequences=False,
activation="tanh", recurrent_activation="sigmoid", recurrent_dropout=0, unroll=False, use_bias=True, reset_after=True
)(x)
x = Flatten()(x)
x = Concatenate()([x, input_force_flat])
x = Dropout(0.2)(x)
x = Dense(np.prod(dp.y_delta[0].shape * 2), activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(np.prod(dp.y_delta[0].shape * 3), activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(np.prod(dp.y_delta[0].shape * 2), activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(np.prod(dp.y_delta[0].shape))(x)
output_1 = Reshape(dp.y_delta[0].shape)(x)
model = Model(inputs=[input_abs_0, input_abs, input_force], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
# 0.0092
# create_and_train_network(dp, dp.y_delta, 11, model_generator=create_model4_improved)
def create_model4_improved():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs_0 = Input(shape=(4, 9), name='x_abs_n_minus_1')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_abs_0_flat = Flatten()(input_abs_0)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
x = Concatenate()([input_abs_0_flat, input_abs_flat])
x = Reshape((2, 4*9))(x)
x = GRU(np.prod(dp.y_abs[0].shape * 2), return_sequences=False,
activation="tanh", recurrent_activation="sigmoid", recurrent_dropout=0, unroll=False, use_bias=True, reset_after=True
)(x)
x = Flatten()(x)
x = Concatenate()([x, input_force_flat])
x = Dropout(0.2)(x)
x = Dense(np.prod(dp.y_abs[0].shape * 2))(x)
x = Dropout(0.2)(x)
x = Dense(np.prod(dp.y_abs[0].shape * 2))(x)
x = Dropout(0.2)(x)
x = Dense(np.prod(dp.y_abs[0].shape))(x)
output_1 = Reshape(dp.y_abs[0].shape)(x)
model = Model(inputs=[input_abs_0, input_abs, input_force], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
#create_and_train_network(dp, dp.y_abs, 12, model_generator=create_model4_improved)
from tensorflow.keras.layers import BatchNormalization
def create_model2_simple():
input_delta = Input(shape=(4, 6), name='x_delta')
input_force = Input(shape=(3, 1), name='x_force')
input_delta_flat = Flatten()(input_delta)
input_force_flat = Flatten()(input_force)
x = Concatenate()([input_delta_flat, input_force_flat])
x = Dense(128, activation='relu')(x)
x = Dropout(0.2)(x)
x = BatchNormalization()(x)
x = Dense(256, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(512, activation='relu')(x)
x = Dropout(0.2)(x)
x = BatchNormalization()(x)
x = Dense(2048, activation='relu')(x)
x = Dropout(0.2)(x)
x = BatchNormalization()(x)
x = Dense(512, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(256, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(64, activation='relu')(x)
x = Dropout(0.2)(x)
x = BatchNormalization()(x)
x = Dense(np.prod(dp.y_delta[1].shape))(x)
output_1 = Reshape(dp.y_delta[1].shape)(x)
model = Model(inputs=[input_delta, input_force], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
create_and_train_network(dp, dp.y_delta, 15, model_generator=create_model2_simple)
def create_model4_improved():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs_0 = Input(shape=(4, 9), name='x_abs_n_minus_1')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_abs_0_flat = Flatten()(input_abs_0)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
input_delta_flat = Flatten()(input_delta)
x = Concatenate()([input_abs_0, input_abs])
#x = Reshape((2, 36))(x)
x = GRU(np.prod(dp.y_abs[0].shape), return_sequences=True,
activation="tanh", recurrent_activation="sigmoid", recurrent_dropout=0, unroll=False, use_bias=True, reset_after=True
)(x)
x = GRU(np.prod(dp.y_abs[0].shape * 2), return_sequences=False,
activation="tanh", recurrent_activation="sigmoid", recurrent_dropout=0, unroll=False, use_bias=True, reset_after=True
)(x)
x = BatchNormalization()(x)
x = Flatten()(x)
x = Concatenate()([x, input_force_flat, input_delta_flat])
x = Dropout(0.2)(x)
x = Dense(np.prod(dp.y_delta[0].shape * 2))(x)
x = Dropout(0.2)(x)
x = Dense(np.prod(dp.y_delta[0].shape * 2))(x)
x = Dropout(0.2)(x)
x = Dense(np.prod(dp.y_delta[0].shape))(x)
output_1 = Reshape(dp.y_delta[0].shape)(x)
model = Model(inputs=[input_abs_0, input_abs, input_force, input_delta], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
create_and_train_network(dp, dp.y_delta, 12, model_generator=create_model4_improved)
from tensorflow.keras.layers import Add
def create_model4_improved():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs_0 = Input(shape=(4, 9), name='x_abs_n_minus_1')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_abs_0_flat = Flatten()(input_abs_0)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
input_delta_flat = Flatten()(input_delta)
delta_force = Concatenate()([input_force_flat, input_delta_flat])
delta_force = Dense(np.prod(dp.y_delta[0].shape))(delta_force)
delta_force = Reshape(dp.y_delta[0].shape)(delta_force)
x = Concatenate()([input_abs_0, input_abs])
#x = Reshape((2, 36))(x)
x = GRU(np.prod(dp.y_delta[0].shape), return_sequences=False,
activation="tanh", recurrent_activation="sigmoid", recurrent_dropout=0, unroll=False, use_bias=True, reset_after=True
)(x)
x = Reshape(dp.y_delta[0].shape)(x)
x = Add()([x, delta_force])
x = Dense(np.prod(dp.y_delta[0].shape * 2))(x)
x = Dropout(0.2)(x)
x = Dense(dp.y_delta[0].shape[-1])(x)
# print(x.shape, np.prod(dp.y_delta[0].shape), dp.y_delta[0].shape, "!!!!!!")
output_1 = Reshape(dp.y_delta[0].shape)(x)
model = Model(inputs=[input_abs_0, input_abs, input_force, input_delta], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
create_and_train_network(dp, dp.y_delta, 12, model_generator=create_model4_improved)
from tensorflow.keras.layers import Add
def create_model4_improved():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs_0 = Input(shape=(4, 9), name='x_abs_n_minus_1')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_abs_0_flat = Flatten()(input_abs_0)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
input_delta_flat = Flatten()(input_delta)
delta_force = Concatenate()([input_force_flat, input_delta_flat])
delta_force = Dense(np.prod(dp.y_delta[0].shape))(delta_force)
delta_force = Reshape(dp.y_delta[0].shape)(delta_force)
x = Concatenate()([input_abs_0, input_abs])
#x = Reshape((2, 36))(x)
x = GRU(np.prod(dp.y_delta[0].shape), return_sequences=False,
activation="tanh", recurrent_activation="sigmoid", recurrent_dropout=0, unroll=False, use_bias=True, reset_after=True
)(x)
x = Reshape(dp.y_delta[0].shape)(x)
x = Add()([x, delta_force])
x = Dense(np.prod(dp.y_delta[0].shape * 2))(x)
x = Dropout(0.2)(x)
x = Dense(dp.y_abs[0].shape[-1])(x)
# print(x.shape, np.prod(dp.y_delta[0].shape), dp.y_delta[0].shape, "!!!!!!")
output_1 = Reshape(dp.y_abs[0].shape)(x)
model = Model(inputs=[input_abs_0, input_abs, input_force, input_delta], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
create_and_train_network(dp, dp.y_abs, 12, model_generator=create_model4_improved)
from tensorflow.keras.layers import Add
def create_model4_improved():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs_0 = Input(shape=(4, 9), name='x_abs_n_minus_1')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_abs_0_flat = Flatten()(input_abs_0)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
input_delta_flat = Flatten()(input_delta)
delta_force = Concatenate()([input_force_flat, input_delta_flat])
delta_force = Dense(np.prod(dp.y_delta[0].shape))(delta_force)
delta_force = Reshape(dp.y_delta[0].shape)(delta_force)
x = Dense(np.prod(dp.y_delta[0].shape[-1]))(input_abs)
x = Add()([x, delta_force])
x = Dense(np.prod(128))(x)
x = Dropout(0.2)(x)
x = Dense(dp.y_delta[0].shape[-1])(x)
# print(x.shape, np.prod(dp.y_delta[0].shape), dp.y_delta[0].shape, "!!!!!!")
output_1 = Reshape(dp.y_delta[0].shape)(x)
model = Model(inputs=[input_abs_0, input_abs, input_force, input_delta], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
create_and_train_network(dp, dp.y_delta, 12, model_generator=create_model4_improved)
from tensorflow.keras.layers import Add
def create_model4_improved():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs_0 = Input(shape=(4, 9), name='x_abs_n_minus_1')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_abs_0_flat = Flatten()(input_abs_0)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
input_delta_flat = Flatten()(input_delta)
delta_force = Concatenate()([input_force_flat, input_delta_flat])
delta_force = Dense(np.prod(dp.y_delta[0].shape))(delta_force)
delta_force = Reshape(dp.y_delta[0].shape)(delta_force)
x = Dense(np.prod(dp.y_delta[0].shape[-1]))(input_abs)
x = Reshape(dp.y_delta[0].shape)(x)
x = Add()([x, delta_force])
x = Dense(np.prod(dp.y_delta[0].shape * 2))(x)
x = Dropout(0.2)(x)
x = Dense(dp.y_abs[0].shape[-1])(x)
# print(x.shape, np.prod(dp.y_delta[0].shape), dp.y_delta[0].shape, "!!!!!!")
output_1 = Reshape(dp.y_abs[0].shape)(x)
model = Model(inputs=[input_abs_0, input_abs, input_force, input_delta], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
create_and_train_network(dp, dp.y_abs, 12, model_generator=create_model4_improved)
from tensorflow.keras.layers import Add
def create_model4_improved():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs_0 = Input(shape=(4, 9), name='x_abs_n_minus_1')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_abs_0_flat = Flatten()(input_abs_0)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
input_delta_flat = Flatten()(input_delta)
delta_force = Concatenate()([input_force_flat, input_delta_flat])
delta_force = Dense(np.prod(dp.y_delta[0].shape))(delta_force)
delta_force = Reshape(dp.y_delta[0].shape)(delta_force)
x = Concatenate()([input_abs_0, input_abs])
#x = Reshape((2, 36))(x)
x = GRU(np.prod(dp.y_delta[0].shape), return_sequences=False,
activation="tanh", recurrent_activation="sigmoid", recurrent_dropout=0, unroll=False, use_bias=True, reset_after=True
)(x)
x = Reshape(dp.y_delta[0].shape)(x)
x = Add()([x, delta_force])
x = Dense(dp.y_abs[0].shape[-1] * 2)(x)
x = Dropout(0.2)(x)
x = Dense(dp.y_abs[0].shape[-1] * 4)(x)
x = Dropout(0.2)(x)
x = Dense(dp.y_abs[0].shape[-1] * 2)(x)
x = Dropout(0.2)(x)
x = Dense(dp.y_abs[0].shape[-1])(x)
# print(x.shape, np.prod(dp.y_delta[0].shape), dp.y_delta[0].shape, "!!!!!!")
output_1 = Reshape(dp.y_abs[0].shape)(x)
model = Model(inputs=[input_abs_0, input_abs, input_force, input_delta], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
create_and_train_network(dp, dp.y_abs, 12, model_generator=create_model4_improved)
from tensorflow.keras.layers import Add
def create_model4_improved():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs_0 = Input(shape=(4, 9), name='x_abs_n_minus_1')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_abs_0_flat = Flatten()(input_abs_0)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
input_delta_flat = Flatten()(input_delta)
input_force_part = Concatenate()([input_abs_flat, input_force_flat])
input_force_part = Dense(np.prod(dp.y_delta[0].shape))(input_force_part)
x = Add()([input_delta_flat, input_force_part])
x = Reshape(dp.y_delta[0].shape)(x)
x = Dense(256)(x)
x = Dense(512)(x)
x = Dense(512)(x)
x = Dense(256)(x)
x = Dense(dp.y_delta[0].shape[-1])(x)
# print(x.shape, np.prod(dp.y_delta[0].shape), dp.y_delta[0].shape, "!!!!!!")
output_1 = Reshape(dp.y_delta[0].shape)(x)
model = Model(inputs=[input_abs_0, input_abs, input_force, input_delta], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
create_and_train_network(dp, dp.y_delta, 12, model_generator=create_model4_improved)
from tensorflow.keras.layers import Add
def create_model4_improved():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs_0 = Input(shape=(4, 9), name='x_abs_n_minus_1')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_abs_0_flat = Flatten()(input_abs_0)
input_abs_flat = Flatten()(input_abs)
input_force_flat = Flatten()(input_force)
input_delta_flat = Flatten()(input_delta)
input_force_part = Concatenate()([input_abs_flat, input_force_flat])
input_force_part = Dense(np.prod(dp.y_delta[0].shape))(input_force_part)
x = Add()([input_delta_flat, input_force_part])
x = Dense(256)(x)
x = Dense(512)(x)
x = Dense(512)(x)
x = Dense(256)(x)
x = Dense(np.prod(dp.y_abs[0].shape))(x)
# print(x.shape, np.prod(dp.y_delta[0].shape), dp.y_delta[0].shape, "!!!!!!")
output_1 = Reshape(dp.y_abs[0].shape)(x)
model = Model(inputs=[input_abs_0, input_abs, input_force, input_delta], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
create_and_train_network(dp, dp.y_abs, 12, model_generator=create_model4_improved)
from tensorflow.keras.layers import Add
def create_model4_improved():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs_0 = Input(shape=(4, 9), name='x_abs_n_minus_1')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_abs_0_flat = Flatten()(input_abs_0)
input_abs_flat = Flatten()(input_abs)
input_abs_rs = Dense(16)(input_abs)
input_abs_rs = Dense(32)(input_abs_rs)
input_abs_rs = Dense(8)(input_abs_rs)
input_abs_rs = Dense(6)(input_abs_rs)
input_force_flat = Flatten()(input_force)
input_delta_flat = Flatten()(input_delta)
input_force_part = Concatenate()([input_abs_flat, input_force_flat])
input_force_part = Dense(np.prod(dp.y_delta[0].shape))(input_force_part)
x = Add()([input_delta_flat, input_force_part])
x = Reshape(dp.y_delta[0].shape)(x)
x = Add()([x, input_abs_rs])
x = Dense(256)(x)
x = Dense(512)(x)
x = Dense(512)(x)
x = Dense(256)(x)
x = Dense(dp.y_abs[0].shape[-1])(x)
# print(x.shape, np.prod(dp.y_delta[0].shape), dp.y_delta[0].shape, "!!!!!!")
output_1 = Reshape(dp.y_abs[0].shape)(x)
model = Model(inputs=[input_abs_0, input_abs, input_force, input_delta], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
create_and_train_network(dp, dp.y_abs, 12, model_generator=create_model4_improved)
from tensorflow.keras.layers import Add
def create_model4_improved():
input_delta = Input(shape=(4, 6), name='x_delta')
input_abs_0 = Input(shape=(4, 9), name='x_abs_n_minus_1')
input_abs = Input(shape=(4, 9), name='x_abs_n')
input_force = Input(shape=(3, 1), name='x_force')
input_abs_0_flat = Flatten()(input_abs_0)
input_abs_flat = Flatten()(input_abs)
input_abs_rs = Dense(16)(input_abs)
input_abs_rs = Dense(32)(input_abs_rs)
input_abs_rs = Dense(8)(input_abs_rs)
input_abs_rs = Dense(6)(input_abs_rs)
input_force_flat = Flatten()(input_force)
input_delta_flat = Flatten()(input_delta)
input_force_part = Concatenate()([input_abs_flat, input_force_flat])
input_force_part = Dense(512)(input_force_part)
input_force_part = Dense(np.prod(dp.y_delta[0].shape))(input_force_part)
x = Add()([input_delta_flat, input_force_part])
x = Reshape(dp.y_delta[0].shape)(x)
x = Add()([x, input_abs_rs])
x = Dense(256)(x)
x = Dense(512)(x)
x = Dense(512)(x)
x = Dense(256)(x)
x = Dense(dp.y_abs[0].shape[-1])(x)
# print(x.shape, np.prod(dp.y_delta[0].shape), dp.y_delta[0].shape, "!!!!!!")
output_1 = Reshape(dp.y_abs[0].shape)(x)
model = Model(inputs=[input_abs_0, input_abs, input_force, input_delta], outputs=[output_1])
model.compile(loss="mse", optimizer='rmsprop')
return model
create_and_train_network(dp, dp.y_abs, 12, model_generator=create_model4_improved)
```
| github_jupyter |
<a href='https://www.learntocodeonline.com/'><img src='https://github.com/ProsperousHeart/TrainingUsingJupyter/blob/master/IMGs/learn-to-code-online.png?raw=true'></a>
_Need to go back to Week 1?_ Click **[here](../Week_1)**!
# Variable Type: [List](http://www.tutorialspoint.com/python/python_lists.htm)
This is the most versatile variable type - also known as an object.
It is written as a list of comma-separated values between square brackets (similar to a [Java array](https://docs.oracle.com/javase/tutorial/java/nutsandbolts/arrays.html)).
`new_list = [1, "string", ...]` where `...` is simply indicative of additional elements.
Some reasons why this is so versatile are:
- the items within a list do NOT need to be of the same data type
- you can have as many items as you require (and have capability to hold)
- there are multiple ways to leverage this data structure
Please be advised that [PEP484](https://www.python.org/dev/peps/pep-0484) highly encourages that the items in a list are of the same type, however it is not enforced. It's just a styling guideline.
<div class="alert alert-success">
<b>Try this!</b>
```python
new_list = ['abcd', 786, 2.34, "John", 70.2]
tiny_list = [123, 'john']
print(new_list)
print(tiny_list)
```
</div>
# Basic List Operations
Basic operations for lists are:
- length: `len(new_list)`
- concatentation
- repetition (multiplication)
- membership
- iteration
## Indexing
Similar to string indices, list indices start at 0 & can be sliced, concatenated, etc.
When you using indexing to return an element of a list, it returns that element - it does not return a list.
<div class="alert alert-success">
<b>Try this!</b>
```python
print(new_list[0])
```
</div>
<div class="alert alert-success">
<b>... then this:</b>
```python
print(tiny_list[-1])
```
</div>
## Slicing
Another thing to keep in mind is that with **slicing** is that when you slice a string, it returns a new string.
When you slice a list? It returns a new list.
<div class="alert alert-success">
<b>Try this!</b>
```python
print(new_list[3:])
```
</div>
<div class="alert alert-success">
<b>... then this:</b>
```python
new_tiny_list = tiny_list[:] # makes a new copy - does not refer to same memory location
print(new_tiny_list)
print("new_tiny_list == tiny_list: {}".format(new_tiny_list == tiny_list))
print("new_tiny_list is tiny_list: {}".format(new_tiny_list is tiny_list))
```
</div>
This is the same kind of indexing & slicing mentioned in [Basics 8 - Variable Type: STRING](Python_Basics_08_-_Variable_Type_STRING.ipynb) - as outlined in the video below.
```
# https://stackoverflow.com/a/59712486
from IPython.display import IFrame
# Youtube or Facebook embed
IFrame(src="https://www.youtube.com/embed/snXCYlOa6D4", width="560", height="315")
```
## Multiplication
When you multiply lists, similar to strings it makes a copy and adds to it.
<div class="alert alert-success">
<b>Try this!</b>
```python
print(tiny_list, "\n")
print(tiny_list * 2)
```
</div>
However, unless you reassign the value back into the list, then it will remain unchanged.
<div class="alert alert-success">
<b>Try this!</b>
```python
print(tiny_list)
tiny_list = tiny_list * 2
print(tiny_list)
```
</div>
## Concatenation
Just like in a string, you can use concatenation to add to a list.
Keep in mind that unless you reassign the list returned from the concatenation, then nothing is changed in place.
<div class="alert alert-success">
<b>Try this!</b>
```python
print("tiny_list = {}".format(tiny_list))
print("new_list = {}\n".format(new_list))
print("CONCATENATION: {}\n".format(tiny_list + new_list))
print("tiny_list = {}".format(tiny_list))
print("new_list = {}".format(new_list))
```
</div>
## Updating A List
The biggest difference between a list and a string is that whiel a string is immutable (can't change it)?
You can replace (update) elements in a list very easily. You just need to assign a new value to the element at index `n` such as:
`list_var[n] = some_data`
<div class="alert alert-success">
<b>Try this!</b>
```python
print(new_list)
new_list[1] = 678
print(new_list)
```
</div>
### Adding To A List
There are two ways to add to a list:
- append
- extend
#### List Append
This built-in method for lists will only take a single input, otherwise it will error out.
`list_var.append(something)`
This allows you to add a single element to the end of a list.
<div class="alert alert-success">
<b>Try this!</b>
```python
print(new_tiny_list)
new_tiny_list.append(["This is a list", 2])
print(new_tiny_list)
```
</div>
<div class="alert alert-success">
<b>... then this:</b>
```python
tiny_list.append(1, 2)
```
</div>
<div class="alert alert-warning">
<i>What happens? Why?</i> You'll learn more about this in <a href="https://github.com/ProsperousHeart/Basics-Boot-Camp/tree/main/Week_3">Week 3</a>.
For now, focus on what this is telling you.
How would you go about solving an error from this?
</div>
#### List Extend
This allows you to add every element of what you are trying to add as a new element in the list.
`listVar.extend(new_list)`
This also only takes in a single element, so you must use a list or tuple.
<div class="alert alert-success">
<b>Try this!</b>
```python
print(new_tiny_list)
```
</div>
<div class="alert alert-success">
<b>... then this:</b>
```python
new_tiny_list.extend([1, 2, 3, "fun"])
print(new_tiny_list)
```
</div>
<div class="alert alert-success">
<b>... then this:</b>
```python
new_tiny_list.extend(1, 2, 3, "fun")
```
</div>
<div class="alert alert-warning">
Learn to recognize these errors. Knowing what possibilities to expect will improve your forward testing capabilities.
In other words ... You can make your code stronger and less susceptible to breaches or "broken" code.
</div>
### Removing From A List
You have several options to remove items from a list.
#### Pop
With our **append** we also have a [pop]() feature with lists, however you may want to consider [dequeue](https://github.com/ProsperousHeart/TrainingUsingJupyter/blob/master/Python/Recipes/Data%20Structures%20And%20Algorithms%2000%20-%20Unpacking%20And%20Deque.ipynb).
`list.pop([index])`
The notation above indicates that a single input of an index is optional, so you could do something like:
<div class="alert alert-success">
```python
temp_list = [1, 2, 3, 4, 5]
print('temp_list: {}'.format(temp_list))
temp_var = temp_list.pop(2)
print('temp_var: {}'.format(temp_var))
print('temp_list: {}'.format(temp_list))
temp_var1 = temp_list.pop()
print('temp_var1: {}'.format(temp_var1))
print('temp_list: {}'.format(temp_list))
```
</div>
Why do we have **append** and **pop** but no **push**?
[Here](https://stackoverflow.com/a/1569007/10474024) is a great response.
#### Deletion of An Element
`del new_list[n]`
You can delete an element of a list by simply providing the position.
<div class="alert alert-success">
<b>Try this!</b>
```python
print(temp_list)
del temp_list[1]
print(temp_list)
```
</div>
## Membership
See [prior section](Python_Basics_03_-_Operators.ipynb#Membership-Operators) on this.
<div class="alert alert-success">
<b>Try this!</b>
```python
verdict = False # innocent until proven otherwise, right? ;)
while not(verdict):
print("Is 2 in temp_list?\n{}".format(temp_list))
if 2 in temp_list:
print("Winner winner, chicken dinner.")
verdict = True
else:
print("Nyet.")
temp_list.append(2) # new stuff entered
print("The end!")
```
</div>
## Iteration
An **iterable** is anything you can loop over.
**Sequences** are iterables that have a specific set of features.
Lots of built-in objects in python are iterables, but not all iterables are sequences. (e.g.: sets, dictionaries, generators, files, etc) - Therefore sequences are a *type* of [iterable](https://docs.python.org/3/library/functions.html#iter).
<div class="alert alert-success">
<b>Try this!</b> <i>But remember - there's no one-way to do anything in programming.</i>
```python
tmp_itr = iter(temp_list)
for itm in tmp_itr:
print(itm)
```
</div>
# Additional Resources
http://effbot.org/zone/python-list.htm
http://www.learnpython.org/en/List_Comprehensions
| github_jupyter |
```
import re
import urllib
import urllib3
import csv
import requests
from bs4 import BeautifulSoup
from collections import OrderedDict
headers = {'User-Agent':'Mozilla/5.0'}
urllib3.disable_warnings()
data1=[]
import random
import csv
from time import sleep
li='https://www.floridabids.us/florida-contractors/location-Altamonte%20Springs.2.htm'
page = requests.get(li, headers=headers, verify=False)
data = page.content
soup = BeautifulSoup(data, "html.parser")
cities_link=soup.find_all(id="filter-city")[0].find_all('a')
len(cities_link)
for g in range(0,len(cities_link)):
link_city0=cities_link[g].get('href')
linkto_city='https://www.floridabids.us'+link_city0
sleep(1)
page = requests.get(linkto_city, headers=headers, verify=False)
data = page.content
soup_companies = BeautifulSoup(data, "html.parser")
allcomp=soup_companies.find_all('div',class_="lr-title lr-mar")
for i in range(len(allcomp)):
data0=[]
link0=allcomp[i].find('a').get('href')
linkto_comp='https://www.floridabids.us'+link0
# print (linkto_comp)
data0.append(linkto_comp)
data1.append(data0)
print(len(data1))
if len(allcomp)>0:
total_page=(int((soup_companies.find(class_="list-total").find_all('span')[-1].text).replace(',',''))/25)+1
print 'Total_pages:',total_page
for q in range(2,total_page+1):
url=linkto_city.replace('.htm','.'+str(q)+'.htm')
sleep(1)
page = requests.get(url, headers=headers, verify=False)
data = page.content
soup = BeautifulSoup(data, "html.parser")
allcomp=soup.find_all('div',class_="lr-title lr-mar")
for i in range(len(allcomp)):
data0=[]
link0=allcomp[i].find('a').get('href')
linkto_comp='https://www.floridabids.us'+link0
# print (linkto_comp)
data0.append(linkto_comp)
data1.append(data0)
csvWriter = csv.writer(open('output_0.csv', 'a'))
csvWriter.writerow(data0)
print(len(data1))
78351/25
len(allcomp)
data1
# 11833
https://www.floridabids.us/florida-contractors/contractor-5821354-STRATIGENT-INC.htm
# 20567
https://www.floridabids.us/florida-contractors/contractor-5610736-L-J-DIESEL-SERVICE-INC.htm
# 30978
https://www.floridabids.us/florida-contractors/contractor-5548974-LUNA-SOURCE.htm
# 39187
https://www.floridabids.us/florida-contractors/contractor-5215930-AIRCRAFT-ENGINEERIN.htm
# 41904
https://www.floridabids.us/florida-contractors/contractor-5815006-MAPPING-SUSTAINABILIT.htm
# 57603
https://www.floridabids.us/florida-contractors/contractor-5101540-Venatore-LLC.htm
checklink=[]
link='https://www.floridabids.us/florida-contractors/search.htm?keyword=&city=Tampa%2C+FL&naics='
# link='https://www.floridabids.us/florida-contractors/search.htm?keyword=&city=Palm%2C+FL&naics='
# link='https://www.floridabids.us/florida-contractors/search.htm?keyword=&city=Orlando%2C+FL&naics='
# link='https://www.floridabids.us/florida-contractors/search.htm?keyword=&city=Miami%2C+FL&naics='
# link='https://www.floridabids.us/florida-contractors/search.htm?keyword=&city=Jacksonville%2C+FL&naics='
# link='https://www.floridabids.us/florida-contractors/search.htm?keyword=&city=Fort+Lauderdale%2C+FL&naics='
page = requests.get(link, headers=headers, verify=False)
data = page.content
soup = BeautifulSoup(data, "html.parser")
total_page=(int((soup.find(class_="list-total").find_all('span')[-1].text).replace(',',''))/25)+1
total_page
filters=soup.find_all(class_="filter-box")
linkscate=filters[3].find_all('a')
for k in range(len(linkscate)):
checklink.append('https://www.floridabids.us'+linkscate[k].get('href'))
print (len(checklink))
checklink[0]
'https://www.floridabids.us/florida-contractors/search.htm?city=Fort+Lauderdale%2C+FL&naics=11'
'https://www.floridabids.us/florida-contractors/search.htm?city=Fort+Lauderdale%2C+FL&naics=11&page=2'
checklink[5]
for g in range(5,len(checklink)):
sleep(1)
page = requests.get(checklink[g], headers=headers, verify=False)
data = page.content
soup_companies = BeautifulSoup(data, "html.parser")
allcomp=soup_companies.find_all('div',class_="lr-title lr-mar")
for i in range(len(allcomp)):
data0=[]
link0=allcomp[i].find('a').get('href')
linkto_comp='https://www.floridabids.us'+link0
# print (linkto_comp)
data0.append(linkto_comp)
csvWriter = csv.writer(open('output_rem.csv', 'a'))
csvWriter.writerow(data0)
data1.append(data0)
print(len(data1))
if len(allcomp)>0:
total_page=(int((soup_companies.find(class_="list-total").find_all('span')[-1].text).replace(',',''))/25)+1
print 'Total_pages:',total_page
for q in range(2,total_page+1):
url=checklink[g]+'&page='+str(q)
sleep(1)
page = requests.get(url, headers=headers, verify=False)
data = page.content
soup = BeautifulSoup(data, "html.parser")
allcomp=soup.find_all('div',class_="lr-title lr-mar")
for i in range(len(allcomp)):
data0=[]
link0=allcomp[i].find('a').get('href')
linkto_comp='https://www.floridabids.us'+link0
# print (linkto_comp)
data0.append(linkto_comp)
data1.append(data0)
csvWriter = csv.writer(open('output_rem.csv', 'a'))
csvWriter.writerow(data0)
print(len(data1))
```
| github_jupyter |
```
import os
from datetime import datetime
from collections import defaultdict
import numpy as np
import yaml
import sys
from easydict import EasyDict as edict
from IPython.core.debugger import set_trace
import torch
from torch import nn
import torch.optim as optim
from torch.utils.data import Dataset, DataLoader
from tensorboardX import SummaryWriter
from networks import define_G, define_D, GANLoss, get_scheduler, update_learning_rate
from utils import vis_batch, save_batch, collate_fn, load_config
from dataset import FashionEdgesDataset
import matplotlib.pyplot as plt
from skimage.transform import resize
import cv2
from tensorboardX import SummaryWriter
def fig_to_array(fig):
fig.canvas.draw()
fig_image = np.array(fig.canvas.renderer._renderer)
return fig_image
device = 'cuda:0'
from skimage.feature import canny
from skimage.morphology import dilation, disk, square
from skimage.color import rgb2gray
from skimage.filters import gaussian
from skimage.segmentation import flood
def image2edges(image,
low_thresh=0.01,
high_thresh=0.2,
sigma=0.5,
selem=True,
d = 2,
randomize_disc=True):
'''
image - np.array
'''
image_gray_rescaled = rgb2gray(image)
edges = canny(image_gray_rescaled,
sigma = sigma,
low_threshold=low_thresh,
high_threshold=high_thresh)
if selem:
if randomize_disc:
d = np.random.choice()
selem = disk(d)
edges = dilation(edges, selem)
low_thresh=0.01
high_thresh=0.2
sigma=0.5
selem=True
d = 2
randomize_disc=True
image = cv2.imread('./agata/Sweater/13783346_17197090_1000.png')
image_gray_rescaled = rgb2gray(image)
edges = canny(image_gray_rescaled,
sigma = 6,
low_threshold=low_thresh,
high_threshold=high_thresh)
if selem:
if randomize_disc:
d = 2 #np.random.choice([0,1,2,5], 1)
selem = disk(d)
edges = dilation(edges, selem)
print(d)
plt.imshow(edges)
edges.astype(float)
module = nn.Conv2d(3,3,3)
optimizerG = optim.Adam(module.parameters(), lr=0.1, betas=(0.1, 0.999))
optimizerG.state_dict()['param_groups'][0]['lr']
writer = SummaryWriter('./logs/tmp')
img = cv2.imread('./feidegger/images/8550.jpg')
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
img = resize(img, (512,356), anti_aliasing=True)
fig = plt.figure()
plt.imshow(img)
arr = fig_to_array(fig)
n_iters_total = 10
writer.add_image(f"{n_iters_total}", arr[...,:3].transpose(2,0,1), global_step=n_iters_total)
netG = define_G(config.input_nc, config.output_nc, config.ngf, 'batch', False, 'normal', 0.02, gpu_id=device)
netD = define_D(config.input_nc + config.output_nc, config.ndf, 'basic', gpu_id=device)
s_total = 0
for param in netG.parameters():
s_total+=param.numel()
print ('Params in Generator (M):', round(s_total/(10**6),2))
s_total = 0
for param in netD.parameters():
s_total+=param.numel()
print ('Params in Discriminator (M):', round(s_total/(10**6),2))
real_a, real_b = torch.randn(config.batch_size,1,512,356).cuda(), torch.randn(config.batch_size,3,512,356).cuda()
fake_b = netG(real_a)
fake_ab = torch.cat((real_a, fake_b), 1)
pred_fake = netD.forward(fake_ab)
```
| github_jupyter |
### This notebook chronicles my major learnings from the Uber traffic prediction competition hosted on Zindi.
# Initial Preprocessing
In this competition, the target value was not explicitly given in the training set so we needed to an initial preprocessing to obtain it. Special thanks to the community for providing us with the initial processing code. The code is replicated below.
```
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pylab as plt
uber = pd.read_csv('train_revised.csv', low_memory=False)
test = pd.read_csv('test_questions.csv', low_memory=False)
uber.head()
ride_id_dict = {}
for ride_id in uber["ride_id"]:
if not ride_id in ride_id_dict:
ride_id_dict[ride_id] = 1
else:
ride_id_dict[ride_id] += 1
uber = uber.drop(['seat_number', 'payment_method', 'payment_receipt', 'travel_to'], axis=1)
uber.drop_duplicates(inplace=True)
uber.reset_index(drop= True, inplace=True)
uber["number_of_tickets"]= np.zeros(len(uber))
for i in range(len(uber)):
ride_id = uber.loc[i]["ride_id"]
uber.at[i,"number_of_tickets"] = ride_id_dict[ride_id]
uber.head()
```
# EXPLORATORY DATA ANALYSIS
In this step, we aim to understand the provided data to see if there are patterns, structures, etc. that could be useful in our modelling phase.
N.B: There is another EDA provided through the community. You can also check it out in the Uber Discussion.
```
uber.info()
test.info()
```
From the above codes, we can see that there are no null values meaning this dataset is 'super-clean'. This is not the usual scenerio. Data-cleaning is usually an important step developing a good model.
We can also see that our travel dates is treated as an object. We should reconsider changing it to datetype.
```
uber['travel_date'] = pd.to_datetime(uber['travel_date'])
test['travel_date'] = pd.to_datetime(test['travel_date'])
print(uber['travel_date'].dtypes)
print(test['travel_date'].dtypes)
```
Let us start our exploration proper.
```
sns.barplot(x='car_type',y='max_capacity',data=uber)
sns.countplot(x='max_capacity',data=uber,hue='car_type')
```
From the data description, we already know that all buses and shuttles have the same maximum capacity. That was easily confirmed using the seaborn barplot.
Another thing we explored was the numbers of buses and shuttles in the dataset using seaborn's countplot. We can see that they are rougly the same.
We can confirmed if this holds true for the test set too
```
sns.barplot(x='car_type',y='max_capacity',data=test)
sns.countplot(x='max_capacity',data=test,hue='car_type')
```
It can be seen that the training is fairly representative of the test set. This is important as we want our unseen scenerios to be similar to the seen scenerios to make our models as accurate as possible
Another thing that is worth exploring is the travel times. Are they horly in nature? Are they just morning journeys? etc. To do this, we have to convert the travel times to meaning numerical data. One way will be to just extract the hour term. Another way is to convert is to minutes from midnight (another valuable insight provided by the community).
```
#Extracting the hour term
uber['hour_booked'] = pd.to_numeric(uber['travel_time'].str.extract(r'(^\d*)').loc[:,0])
test['hour_booked'] = pd.to_numeric(test['travel_time'].str.extract(r'(^\d*)').loc[:,0])
uber['hour_booked'].value_counts().plot.bar()
test['hour_booked'].value_counts().plot.bar()
#express travel time in minutes from midnight
test["travel_time"] = test["travel_time"].str.split(':').apply(lambda x: int(x[0]) * 60 + int(x[1]))
uber["travel_time"] = uber["travel_time"].str.split(':').apply(lambda x: int(x[0]) * 60 + int(x[1]))
(uber["travel_time"]/60).plot.hist(bins=100)
(uber["travel_time"]/60).plot.hist(bins=100)
(test["travel_time"]/60).plot.hist(bins=100)
```
Both methods gave us almost the same information about the data. The most frequent travel time is around 7am and most of the journeys take place before noon with some journeys at 7pm and 11pm.
Another column to explore is travel from i.e where do most of our customers come from?
```
uber['travel_from'].value_counts().plot.bar()
test['travel_from'].value_counts().plot.bar()
```
So many of our customers are actually coming from Kisii. The training set is fairly representative of the test set too.
We can also explore to see if people are likely to travel on a particular day of the week more than the rest.
```
uber["travel_day"] = uber["travel_date"].dt.day_name()
test["travel_day"] = test["travel_date"].dt.day_name()
uber["travel_yr"] = uber["travel_date"].dt.year
test["travel_yr"] = test["travel_date"].dt.year
#Calculating the number of weeks in the dataset we have data for
a=uber[uber["travel_yr"]==2018]["travel_date"].dt.week.nunique() + uber[uber["travel_yr"]==2017]["travel_date"].dt.week.nunique()
b=test[test["travel_yr"]==2018]["travel_date"].dt.week.nunique() + test[test["travel_yr"]==2017]["travel_date"].dt.week.nunique()
(uber[uber['car_type']=='shuttle']["travel_day"].value_counts()/a).plot.bar()
(test[test['car_type']=='shuttle']["travel_day"].value_counts()/b).plot.bar()
(uber[uber['car_type']=='Bus']["travel_day"].value_counts()/a).plot.bar()
(test[test['car_type']=='Bus']["travel_day"].value_counts()/b).plot.bar()
(uber["travel_day"].value_counts()/a).plot.bar()
(test["travel_day"].value_counts()/b).plot.bar()
```
From the above analysis, there seem to be no clear indication that the day of the week matters as the average people that travel on a particular day seems to change with context.
```
uber[uber["travel_yr"]==2017]["travel_date"].dt.month.value_counts().sort_index().plot.bar()
uber[uber["travel_yr"]==2018]["travel_date"].dt.month.value_counts().sort_index().plot.bar()
try:
test[test["travel_yr"]==2017]["travel_date"].dt.month.value_counts().sort_index().plot.bar()
except:
print('No data point to plot.')
test[test["travel_yr"]==2018]["travel_date"].dt.month.value_counts().sort_index().plot.bar()
```
It is clear that from the exploration above that there is something inconsistent about the travel date in the training set. There shouldn't be data for anytime earlier time earlier than Oct 2017 and later than April 2018. This make the date an unreliable indicator. Thus, any date-related feature is unnecessary for modelling.
Lastly we can check to see the distribution for ticket sales.
```
((uber[uber['car_type']=='Bus']['number_of_tickets'])).plot.density()
((uber[uber['car_type']=='shuttle']['number_of_tickets'])).plot.density()
```
It can be seen that for buses, the bus is usually almost empty while for the shuttles, they are almost always full or empty.
```
uber = uber[['travel_time','travel_from','car_type','number_of_tickets','hour_booked']]
uber.head()
```
# Feature Engineering
```
#Trying to linearize the travel time feature for better prediction
uber['travel_time_log']=np.log(uber['travel_time'])
test['travel_time_log']=np.log(test['travel_time'])
```
We proceed to create two features: late night and early morning based on our EDA.
```
uber['early_morning']=uber['hour_booked']<8
test['early_morning']=test['hour_booked']<8
uber['late_night']=uber['hour_booked']>18
test['late_night']=test['hour_booked']>18
```
We use the uber .corr function and seaborn's heatmap to see if there is any linear relationships between our features and targets
```
sns.heatmap(abs(uber.corr()))
```
There seem to be no strong relationship between all of our features and target.
We now try to incoporate an external data - distance from town to Nairobi.
```
distance={'Migori':370.9,'Keroka':279.8,'Kisii':305.5,'Homa Bay':305.5,'Keumbu':294.0,
'Rongo':330.3,'Kijauri':276.6,'Oyugis':331.1,'Awendo':349.5,
'Sirare':391.9,'Nyachenge':322.8,'Kehancha':377.5,
'Kendu Bay':367.5,'Sori':392,'Rodi':349.1,'Mbita':399.4,
'Ndhiwa':369.6}
uber['distance']=uber['travel_from'].map({k:v for k,v in distance.items()})
test['distance']=test['travel_from'].map({k:v for k,v in distance.items()})
test=pd.get_dummies(test,prefix=['car_type','travel_from'],columns=['car_type','travel_from'])
uber=pd.get_dummies(uber,prefix=['car_type','travel_from'],columns=['car_type','travel_from'])
```
# MODELLING
```
print("Original features:\n", (list(uber.columns)), "\n")
feature_cols=['travel_time', 'hour_booked', 'travel_time_log', 'early_morning', 'late_night', 'distance',
'car_type_Bus', 'car_type_shuttle', 'travel_from_Awendo', 'travel_from_Homa Bay', 'travel_from_Kehancha',
'travel_from_Kendu Bay', 'travel_from_Keroka', 'travel_from_Keumbu', 'travel_from_Kijauri',
'travel_from_Kisii', 'travel_from_Mbita', 'travel_from_Migori', 'travel_from_Ndhiwa', 'travel_from_Nyachenge',
'travel_from_Oyugis', 'travel_from_Rodi', 'travel_from_Rongo', 'travel_from_Sirare', 'travel_from_Sori']
predicted_col=['number_of_tickets']
X_train=uber[feature_cols].values
Y_train=uber[predicted_col].values
#Reshaping target column to avoid Sklearb throwing in a warning
Y_train=Y_train.ravel()
split_test_size=0.30
from sklearn.model_selection import train_test_split
Xtrain, Xtest, Ytrain, Ytest= train_test_split(X_train,Y_train, test_size=split_test_size, random_state=260)
from sklearn.metrics import mean_squared_error,mean_absolute_error
from sklearn.model_selection import cross_val_score,KFold,StratifiedKFold
kfold=KFold(n_splits=5)
from sklearn.preprocessing import PolynomialFeatures,MinMaxScaler,StandardScaler
poly=PolynomialFeatures(degree=1).fit(Xtrain)
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.linear_model import LinearRegression
Utrain=(poly.transform(Xtrain))
Utest=(poly.transform(Xtest))
scaler=StandardScaler().fit(Utrain)
Utrain=scaler.transform(Utrain)
Utest=scaler.transform(Utest)
```
# First Model
```
gbrt = GradientBoostingRegressor(criterion='mse',random_state=10,n_estimators=100).fit(Utrain,Ytrain)
cv = cross_val_score (gbrt,Utrain,Ytrain,cv=5)
print(" Average CV is: ", cv.mean())
Ypred=gbrt.predict(Utest)
MAE=mean_absolute_error(Ytest,Ypred)
MSE=mean_squared_error(Ytest,Ypred)
print("GBR MAE:", MAE)
print("GBR Training set score: {:.5f}".format(gbrt.score(Utrain,Ytrain)))
print("GBR Test set score: {:.5f}".format(gbrt.score(Utest,Ytest)))
b=list(gbrt.feature_importances_[1:])
pd.DataFrame(index=feature_cols,data=b).plot.bar()
```
We can clearly see that some features are far more important than some others. While we can just minually remove them. It is better we use the sklearn.feature_selection recursive feature selection with or without cross validation tool (it is better with cv).
```
from sklearn.feature_selection import SelectFromModel
from sklearn.feature_selection import RFE,RFECV
select = RFECV(gbrt,cv=5)
select.fit(Utrain,Ytrain)
select.n_features_
```
We see that the feature selection tool reduced the features from about 25 to 13
```
cv = cross_val_score (select,Utrain,Ytrain,cv=5)
print(" Average CV is: ", cv.mean())
Ypred=select.predict(Utest)
MAE=mean_absolute_error(Ytest,Ypred)
print("GBR MAE:", MAE)
print("GBR Training set score: {:.5f}".format(select.score(Utrain,Ytrain)))
print("GBR Test set score: {:.5f}".format(select.score(Utest,Ytest)))
```
It can be seen that the metrics are almost the same for both sets of features but we prefer the select model because according to Ockham razor principle, you always want to the simplest model that performs best.
# Other implementation
While I tried out other implementations like xgboost, Adaboost, Light GBM, Decision trees, Extra trees and Random Forest, I didnt pay much attention to them as I started the challenge late and didnt have the time to tune every model. I also tried out tricks like PCA but the results were not better off than using just select.I focused only on Gradient Boosting. In hindsight, that may not have been the best decision.
I will however be sharing one other implementation I tried out with alongside gradient boosting (after heavy parameter tuning using Grid Search). Because of the time it took to grid-search, I will just be implementing the best model I obtained in my first grid-search range. To learn more about Grid Search ()
```
gbr = GradientBoostingRegressor(learning_rate=.3,random_state=100,n_estimators=220,subsample=0.75,
loss='lad').fit(Utrain,Ytrain)
select2 = RFECV(gbr,cv=5)
select2.fit(Utrain,Ytrain)
cv = cross_val_score (select2,Utrain,Ytrain,cv=5)
print(" Average CV is: ", cv.mean())
Ypred=select2.predict(Utest)
MAE=mean_absolute_error(Ytest,Ypred)
print("GBR MAE:", MAE)
print("GBR Training set score: {:.5f}".format(select2.score(Utrain,Ytrain)))
print("GBR Test set score: {:.5f}".format(select2.score(Utest,Ytest)))
import mlxtend
#You will need to install mlxtend
from mlxtend.regressor import StackingCVRegressor
```
Since we cant use the stackingCVRegressor with the RFECV select model, we need to redefine our inputs such that the only the most informative features are used. (There is a slight increase from 13 - 19) because I choose to include all the categories of the travel_from.
```
feature_cols=['travel_time_log', 'travel_from_Awendo', 'distance','car_type_shuttle',
'travel_from_Homa Bay', 'travel_from_Kehancha', 'travel_from_Kendu Bay', 'travel_from_Keroka',
'travel_from_Keumbu', 'travel_from_Kijauri', 'travel_from_Kisii', 'travel_from_Mbita', 'travel_from_Migori',
'travel_from_Ndhiwa', 'travel_from_Nyachenge', 'travel_from_Rodi',
'travel_from_Rongo', 'travel_from_Sirare', 'travel_from_Sori']
predicted_col=['number_of_tickets']
X_train=uber[feature_cols].values
Y_train=uber[predicted_col].values
Y_train=Y_train.ravel()
split_test_size=0.3
from sklearn.model_selection import train_test_split
Xtrain, Xtest, Ytrain, Ytest= train_test_split(X_train,Y_train, test_size=split_test_size, random_state=260)
```
Stacking is a type of ensembling that combines the results of two or more estimators using another estimator. Please note that my implementation may not be the best. Stacking is supposed to be used when you are trying to merge the results of three very good estimators. I didn't optimize the decision tree and random forest models.
However, it can be seen that stacked model is not so far off from my best model (imagine the potential if I had used it on many highly tuned models).
```
lr=LinearRegression()
dt = DecisionTreeRegressor(criterion='mae',random_state=100)
rf = RandomForestRegressor(random_state=10,n_estimators=100)
gb = GradientBoostingRegressor(learning_rate=.3,random_state=100,n_estimators=220,subsample=0.75,
loss='lad')
stack = StackingCVRegressor(regressors=(gb, dt, rf),
meta_regressor=lr,cv=5)
stack.fit(Utrain,Ytrain)
```
Below I will be slightly changing my implementation. I am using the mean_absolute_error as the scorer so I can easily see whether my model is generalizing well since mae is the objective metric is the challenge.
```
from sklearn.metrics import make_scorer
cv = cross_val_score (stack,Utrain,Ytrain,cv=5,scoring=make_scorer(mean_absolute_error))
print("Average CV is:", round(cv.mean(),3),cv.std())
Ypred=stack.predict(Utest)
Ypred_t=stack.predict(Utrain)
MAE=mean_absolute_error(Ytest,Ypred)
MAE_t=mean_absolute_error(Ytrain,Ypred_t)
print("GBR Training set score: {:.3f}".format(MAE_t))
print("GBR Test set score: {:.3f}".format(MAE))
```
After another set of gridsearch (ran for about five hours), my winning I came up with my best solution which ended out in the top 25% of all the submitted entries. My model ended up about 0.5 MAE behind winning model in the public leaderboard. I ran out of time to try out other grid search parameters unfortunately.
Below I will contrasting my model to a model of a friend that ended in the top 5% of all submitted entries (about 0.2 MAE behind the winning model).
```
gb=GradientBoostingRegressor(learning_rate=.5,random_state=100,n_estimators=250,subsample=0.75,loss='lad',
max_depth=4).fit(Xtrain,Ytrain)
cv = cross_val_score (gb,Utrain,Ytrain,cv=5,scoring=make_scorer(mean_absolute_error))
print("Average CV is:", round(cv.mean(),3),cv.std())
Ypred=gb.predict(Xtest)
Ypred_t=gb.predict(Xtrain)
MAE=mean_absolute_error(Ytest,Ypred)
MAE_t=mean_absolute_error(Ytrain,Ypred_t)
print("GBR Training set score: {:.3f}".format(MAE_t))
print("GBR Test set score: {:.3f}".format(MAE))
```
The major difference between the two implementation is the range of Grid Search. While I constrained myself, his grid search was more extensive but it took about 3 days for these parameters to be obtained. His implementation is available in the folder.
Finally, I will be sharing something I learnt from a friend after I shared this concern with him after the competition ended: Randomized Search. Randomized Search is similar to Grid Search. The only difference is not available permutations are tested. It randomly picks a specified amount of permutations as defined by you. This could save you a lot of time and helps you choose a more extensive range of search parameters.
```
from sklearn.model_selection import RandomizedSearchCV
#There are about 300,000 different combinations in the grid defined below. We would be using random search to pick just ten
#and see how our model fairs (we do tgis using n_iter)
estimator = GradientBoostingRegressor(random_state=12)
param = {'learning_rate':[0.001, 0.003,.01,0.03,0.05,0.1,0.3,0.5,1,3,5 ],
'n_estimators':[i for i in range(50,550,10)],
'subsample':[i/100 for i in range(50,100,5)],
'loss':['lad','ls','huber'],
'max_depth':[i for i in range(1,20)]}
rs=RandomizedSearchCV(estimator, param_distributions = param,
n_iter=10, n_jobs=-1, random_state=81,cv=3,
return_train_score=True)
rs.fit(Xtrain,Ytrain)
a=pd.DataFrame(rs.cv_results_)
a.sort_values('rank_test_score').head().transpose()
cv = cross_val_score (rs,Utrain,Ytrain,cv=5,scoring=make_scorer(mean_absolute_error))
print("Average CV is:", round(cv.mean(),3),cv.std())
Ypred=rs.predict(Xtest)
Ypred_t=rs.predict(Xtrain)
MAE=mean_absolute_error(Ytest,Ypred)
MAE_t=mean_absolute_error(Ytrain,Ypred_t)
print("GBR Training set score: {:.3f}".format(MAE_t))
print("GBR Test set score: {:.3f}".format(MAE))
```
It can be seen that while the implementation above is somewhat simple and time-efficient. The metrics are not particularly bad. One can rerun the search a few number of times and see whether there is a trend in the chosen parameter and then grid search based on the smaller range. One could also random search for a larger number of times.
# Conclusion
I heard that a top 5 model used Genetic Algorithm for its parameter tuning. Apparently, the person had a lot of time for the tuning as GA is very time-consuming like Grid Search. This may not be possible in a short hackathons or even in some real life scenerios where time is severly limited.
This notebook have taken us through some very important concepts in data science (EDA, Feature Selection, Stacking, HypperParameter Tuning) using the uber data from Zindi. It is specifically desined for those who are just starting their data science journey especially as regards working with real-life data. I hope you find it informative.
P.S: Any other top solution could just clone this notebook and mention one or two two things that made their model stand out if they dont have the time to share an extensive notebook like this.
| github_jupyter |
# A Transfer Learning and Optimized CNN Based Intrusion Detection System for Internet of Vehicles
This is the code for the paper entitled "**A Transfer Learning and Optimized CNN Based Intrusion Detection System for Internet of Vehicles**" accepted in IEEE International Conference on Communications (IEEE ICC).
Authors: Li Yang (lyang339@uwo.ca) and Abdallah Shami (Abdallah.Shami@uwo.ca)
Organization: The Optimized Computing and Communications (OC2) Lab, ECE Department, Western University
**Notebook 1: Data pre-processing**
Procedures:
1): Read the dataset
2): Transform the tabular data into images
3): Display the transformed images
4): Split the training and test set
## Import libraries
```
import numpy as np
import pandas as pd
import os
import cv2
import math
import random
import matplotlib.pyplot as plt
import shutil
from sklearn.preprocessing import QuantileTransformer
from PIL import Image
import warnings
warnings.filterwarnings("ignore")
```
## Read the Car-Hacking/CAN-Intrusion dataset
The complete Car-Hacking dataset is publicly available at: https://ocslab.hksecurity.net/Datasets/CAN-intrusion-dataset
In this repository, due to the file size limit of GitHub, we use the 5% subset.
```
#Read dataset
df=pd.read_csv('data/Car_Hacking_5%.csv')
df
# The labels of the dataset. "R" indicates normal patterns, and there are four types of attack (DoS, fuzzy. gear spoofing, and RPM spoofing zttacks)
df.Label.value_counts()
```
## Data Transformation
Convert tabular data to images
Procedures:
1. Use quantile transform to transform the original data samples into the scale of [0,255], representing pixel values
2. Generate images for each category (Normal, DoS, Fuzzy, Gear, RPM), each image consists of 27 data samples with 9 features. Thus, the size of each image is 9*9*3, length 9, width 9, and 3 color channels (RGB).
```
# Transform all features into the scale of [0,1]
numeric_features = df.dtypes[df.dtypes != 'object'].index
scaler = QuantileTransformer()
df[numeric_features] = scaler.fit_transform(df[numeric_features])
# Multiply the feature values by 255 to transform them into the scale of [0,255]
df[numeric_features] = df[numeric_features].apply(
lambda x: (x*255))
df.describe()
```
All features are in the same scale of [0,255]
### Generate images for each class
```
df0=df[df['Label']=='R'].drop(['Label'],axis=1)
df1=df[df['Label']=='RPM'].drop(['Label'],axis=1)
df2=df[df['Label']=='gear'].drop(['Label'],axis=1)
df3=df[df['Label']=='DoS'].drop(['Label'],axis=1)
df4=df[df['Label']=='Fuzzy'].drop(['Label'],axis=1)
# Generate 9*9 color images for class 0 (Normal)
count=0
ims = []
image_path = "train/0/"
os.makedirs(image_path)
for i in range(0, len(df0)):
count=count+1
if count<=27:
im=df0.iloc[i].values
ims=np.append(ims,im)
else:
ims=np.array(ims).reshape(9,9,3)
array = np.array(ims, dtype=np.uint8)
new_image = Image.fromarray(array)
new_image.save(image_path+str(i)+'.png')
count=0
ims = []
# Generate 9*9 color images for class 1 (RPM spoofing)
count=0
ims = []
image_path = "train/1/"
os.makedirs(image_path)
for i in range(0, len(df1)):
count=count+1
if count<=27:
im=df1.iloc[i].values
ims=np.append(ims,im)
else:
ims=np.array(ims).reshape(9,9,3)
array = np.array(ims, dtype=np.uint8)
new_image = Image.fromarray(array)
new_image.save(image_path+str(i)+'.png')
count=0
ims = []
# Generate 9*9 color images for class 2 (Gear spoofing)
count=0
ims = []
image_path = "train/2/"
os.makedirs(image_path)
for i in range(0, len(df2)):
count=count+1
if count<=27:
im=df2.iloc[i].values
ims=np.append(ims,im)
else:
ims=np.array(ims).reshape(9,9,3)
array = np.array(ims, dtype=np.uint8)
new_image = Image.fromarray(array)
new_image.save(image_path+str(i)+'.png')
count=0
ims = []
# Generate 9*9 color images for class 3 (DoS attack)
count=0
ims = []
image_path = "train/3/"
os.makedirs(image_path)
for i in range(0, len(df3)):
count=count+1
if count<=27:
im=df3.iloc[i].values
ims=np.append(ims,im)
else:
ims=np.array(ims).reshape(9,9,3)
array = np.array(ims, dtype=np.uint8)
new_image = Image.fromarray(array)
new_image.save(image_path+str(i)+'.png')
count=0
ims = []
# Generate 9*9 color images for class 4 (Fuzzy attack)
count=0
ims = []
image_path = "train/4/"
os.makedirs(image_path)
for i in range(0, len(df4)):
count=count+1
if count<=27:
im=df4.iloc[i].values
ims=np.append(ims,im)
else:
ims=np.array(ims).reshape(9,9,3)
array = np.array(ims, dtype=np.uint8)
new_image = Image.fromarray(array)
new_image.save(image_path+str(i)+'.png')
count=0
ims = []
```
### Display samples for each category
```
# Read the images for each category, the file name may vary (27.png, 83.png...)
img1 = Image.open('./train_224/0/27.png')
img2 = Image.open('./train_224/1/83.png')
img3 = Image.open('./train_224/2/27.png')
img4 = Image.open('./train_224/3/27.png')
img5 = Image.open('./train_224/4/27.png')
plt.figure(figsize=(10, 10))
plt.subplot(1,5,1)
plt.imshow(img1)
plt.title("Normal")
plt.subplot(1,5,2)
plt.imshow(img2)
plt.title("RPM Spoofing")
plt.subplot(1,5,3)
plt.imshow(img3)
plt.title("Gear Spoofing")
plt.subplot(1,5,4)
plt.imshow(img4)
plt.title("DoS Attack")
plt.subplot(1,5,5)
plt.imshow(img5)
plt.title("Fuzzy Attack")
plt.show() # display it
```
## Split the training and test set
```
# Create folders to store images
Train_Dir='./train/'
Val_Dir='./test/'
allimgs=[]
for subdir in os.listdir(Train_Dir):
for filename in os.listdir(os.path.join(Train_Dir,subdir)):
filepath=os.path.join(Train_Dir,subdir,filename)
allimgs.append(filepath)
print(len(allimgs)) # Print the total number of images
#split a test set from the dataset, train/test size = 80%/20%
Numbers=len(allimgs)//5 #size of test set (20%)
def mymovefile(srcfile,dstfile):
if not os.path.isfile(srcfile):
print ("%s not exist!"%(srcfile))
else:
fpath,fname=os.path.split(dstfile)
if not os.path.exists(fpath):
os.makedirs(fpath)
shutil.move(srcfile,dstfile)
#print ("move %s -> %s"%(srcfile,dstfile))
# The size of test set
Numbers
# Create the test set
val_imgs=random.sample(allimgs,Numbers)
for img in val_imgs:
dest_path=img.replace(Train_Dir,Val_Dir)
mymovefile(img,dest_path)
print('Finish creating test set')
#resize the images 224*224 for better CNN training
def get_224(folder,dstdir):
imgfilepaths=[]
for root,dirs,imgs in os.walk(folder):
for thisimg in imgs:
thisimg_path=os.path.join(root,thisimg)
imgfilepaths.append(thisimg_path)
for thisimg_path in imgfilepaths:
dir_name,filename=os.path.split(thisimg_path)
dir_name=dir_name.replace(folder,dstdir)
new_file_path=os.path.join(dir_name,filename)
if not os.path.exists(dir_name):
os.makedirs(dir_name)
img=cv2.imread(thisimg_path)
img=cv2.resize(img,(224,224))
cv2.imwrite(new_file_path,img)
print('Finish resizing'.format(folder=folder))
DATA_DIR_224='./train_224/'
get_224(folder='./train/',dstdir=DATA_DIR_224)
DATA_DIR2_224='./test_224/'
get_224(folder='./test/',dstdir=DATA_DIR2_224)
```
| github_jupyter |
# Linear Regression with Gradient Descent Algorithm
This notebook demonstrates the implementation of linear regression with gradient descent algorithm.
Consider the following implementation of the gradient descent loop with NumPy arrays based upon [1]:
```
%pylab inline
def gradient_descent_numpy(X, Y, theta, alpha, num_iters):
m = Y.shape[0]
theta_x = 0.0
theta_y = 0.0
for i in range(num_iters):
predict = theta_x + theta_y * X
err_x = (predict - Y)
err_y = (predict - Y) * X
theta_x = theta_x - alpha * (1.0 / m) * err_x.sum()
theta_y = theta_y - alpha * (1.0 / m) * err_y.sum()
theta[0] = theta_x
theta[1] = theta_y
```
To speedup this implementation with Numba, we need to add the `@jit` decorator to annotate the function signature. Then, we need to expand the NumPy array expressions into a loop. The resulting code is shown below:
```
from numba import jit, f8, int32, void
@jit(void(f8[:], f8[:], f8[:], f8, int32))
def gradient_descent_numba(X, Y, theta, alpha, num_iters):
m = Y.shape[0]
theta_x = 0.0
theta_y = 0.0
for i in range(num_iters):
err_acc_x = 0.0
err_acc_y = 0.0
for j in range(X.shape[0]):
predict = theta_x + theta_y * X[j]
err_acc_x += predict - Y[j]
err_acc_y += (predict - Y[j]) * X[j]
theta_x = theta_x - alpha * (1.0 / m) * err_acc_x
theta_y = theta_y - alpha * (1.0 / m) * err_acc_y
theta[0] = theta_x
theta[1] = theta_y
```
The rest of the code generates some artificial data to test our linear regression algorithm.
```
import numpy as np
import pylab
from timeit import default_timer as timer
def populate_data(N, slope, intercept, stdev=10.0):
noise = stdev*np.random.randn(N)
X = np.arange(N, dtype=np.float64)
Y = noise + (slope * X + intercept)
return X, Y
def run(gradient_descent, X, Y, iterations=10000, alpha=1e-6):
theta = np.empty(2, dtype=X.dtype)
ts = timer()
gradient_descent(X, Y, theta, alpha, iterations)
te = timer()
timing = te - ts
print("x-offset = {} slope = {}".format(*theta))
print("time elapsed: {} s".format(timing))
return theta, timing
def plot(X, theta, c='r'):
result = theta[0] + theta[1] * X
pylab.plot(X, result, c=c, linewidth=2)
```
We will a benchmark with 50 elements to compare the pure python version against the numba version.
```
N = 10
X, Y = populate_data(N, 3, 10)
pylab.scatter(X, Y, marker='o', c='b')
pylab.title('Linear Regression')
print('NumPy'.center(30, '-'))
theta_python, time_python = run(gradient_descent_numpy, X, Y)
print('Numba'.center(30, '-'))
theta_numba, time_numba = run(gradient_descent_numba, X, Y)
# make sure all method yields the same result
assert np.allclose(theta_python, theta_numba)
print('Summary'.center(30, '='))
print('Numba speedup %.1fx' % (time_python / time_numba))
plot(X, theta_numba, c='r')
```
## References
[1] http://aimotion.blogspot.com/2011/10/machine-learning-with-python-linear.html
| github_jupyter |
|<img src="http://pierreproulx.espaceweb.usherbrooke.ca/images/usherb_transp.gif"> |Pierre Proulx, ing, professeur|
|:---|:---|
|Département de génie chimique et de génie biotechnologique |** GCH200-Phénomènes d'échanges I **|
### Section 2.3, écoulement d'un film de fluide Newtonien dans une conduite
> Le développement fait dans Transport Phenomena sera répété ici en développant les solutions avec le calculateur formel sympy et en traçant la solution avec sympy.plot. Nous n'irons pas ici autant dans le détail que dans la section 2.2, Les coordonnées cylindriques sont utilisées, évidemment. Remarquez bien comment la courbure de la géométrie a un effet sur le bilan. <img src='http://pierreproulx.espaceweb.usherbrooke.ca/images/Chap-2-Section-2-3.png'>
>Vous pouvez suivre le développement détaillé à l'aide du livre et voir comment les équations de bilans initiales deviennent les profils de vitesse, de force, débit, etc...
```
#
# Pierre Proulx
#
#
# Préparation de l'affichage et des outils de calcul symbolique
#
import sympy as sp
from IPython.display import *
sp.init_printing(use_latex=True)
%matplotlib inline
# Paramètres, variables et fonctions
#
r,delta_r,L,rho,g,mu,R,P_0,P_L=sp.symbols('r,delta_r,L,rho,g,mu,R,P_0,P_L')
C1=sp.symbols('C1')
phi_rz,phi_zz=sp.symbols('phi_rz,phi_zz')
#
# Équation du bilan des forces
#
eq=2*sp.pi*r*L*phi_rz(r) -2*sp.pi*(r+delta_r)*L*phi_rz(r+delta_r)
eq=eq+2*sp.pi*delta_r*r*phi_zz(0)-2*sp.pi*r*delta_r*phi_zz(L)
eq=eq+2*sp.pi*r*delta_r*L*rho*g
#eq=r*phi_rz(r)/delta_r-(r+delta_r)*phi_rz(r+delta_r)/delta_r+r*P_0/L-r*P_L/L
#
# pose de la limite quand dr tends vers 0
#
eq=eq/(2*sp.pi*L*delta_r)
display(eq)
eq1=sp.limit(eq,delta_r,0).doit()
display(eq1)
eq1=eq1.subs(phi_zz(0),P_0) # les termes de convection s'annulent comme en 2.2
eq1=eq1.subs(phi_zz(L),P_L) # mais il reste le terme de pression
display(eq1)
# Loi de Newton, le terme phi ne contient que le cisaillement, pas de vitesse radiale.
tau=sp.symbols('tau_rz')
eq2=sp.Eq(eq1.subs(phi_rz(r),tau(r)))
display(eq2)
eq2=sp.dsolve(eq2)
display(eq2)
vz=sp.Function('v_z')(r)
newton=-mu*sp.Derivative(vz,r)
eq3=eq2.subs(tau(r),newton)
eq3=eq3.subs(C1,0) # C1 doit être nulle sinon on a une force infinie en r=0
display(eq3)
eq4=sp.dsolve(eq3,vz) # rhs pour avoir la droite de l'équation vz=...
display(eq4)
constante=sp.solve(sp.Eq(eq4.rhs.subs(r,R),0),'C1',dict=True) # Condition de non-glissement à la paroi
display(constante)
constante=constante[0]
vz=eq4.subs(constante)
print('vitesse en fonction de r')
display(vz.simplify())
print('Force')
#
# collect et simplify simplifient l'expression, c'est purement cosmétique.
#
display(-mu*vz.lhs.diff(r)*2*sp.pi*r*L)
display('évalué en r=R devient')
display(-mu*vz.rhs.diff(r).subs(r,R)*2*sp.pi*R*L)
print( 'débit et vitesse moyenne')
vz=vz.rhs # On enlève le terme de gauche et à partir d'ici on travaille seulement avec la partie droite
debit=sp.integrate(2*sp.pi*vz*r,(r,0,R))
display(debit)
vmax=vz.subs(r,0)
display(debit/(sp.pi*R**2)/vmax) # ce calcul donne quoi?
# Maintenant traçons le profil en donnant des valeurs de paramètres réalistes
#
dico={'rho':1000, 'mu':0.001, 'R':0.05, 'L':100,'P_0':200,'P_L':0,'g':9.81}
vz=vz.subs(dico)
vzmax=vz.subs(r,0)
display(vz,vzmax)
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize']=10,8
sp.plot(vz.subs(dico)/vzmax,(r,0,R.subs(dico)),title='Profil de vitesse parabolique',
ylabel='V/Vmax',xlabel='r/R');
```
| github_jupyter |
```
%pylab inline
%config InlineBackend.figure_format = 'svg'
import astropy.cosmology as cosmo
from astropy.cosmology import Planck15
import astropy.units as u
import h5py
import lal
import lalsimulation as ls
import multiprocessing as multi
from scipy.integrate import cumtrapz
from scipy.interpolate import interp1d
from scipy.optimize import linprog
import seaborn as sns
from tqdm import tqdm_notebook
sns.set_style('ticks')
sns.set_context('notebook')
sns.set_palette('colorblind')
```
The mass distribution is
$$
p\left( m_1 \right) \propto \frac{1}{m_1}
$$
(flat in log primary mass) with
$$
p\left( m_2 \mid m_1 \right) \propto \mathrm{const}
$$
(flat in mass ratio at fixed $m_1$) with $5 \, M_\odot \leq m_2 \leq m_1 \leq 50 \, M_\odot$.
The precise mass distribution doesn't matter much for the selection function on $\chi_\mathrm{eff}$; as long as there aren't very sharp features in the $q$ distribution the selection function comes out pretty much the same.
```
MMin = 5
MMax = 50
def draw_m1m2(size=1):
u1s = np.random.uniform(size=size)
u2s = np.random.uniform(size=size)
m1s = exp(u1s*log(MMax) + (1-u1s)*log(MMin))
m2s = m1s*u2s + MMin*(1-u2s)
return (m1s, m2s)
def p_m1m2(m1, m2):
return 1/m1/(log(MMax/MMin))/(m1 - MMin)
```
We draw mergers that follow the low-redshift star-formation rate. $z_\mathrm{max}$ is close to the correct value for design sensitivity; it can be reduced considerably for early sensitivity.
```
gamma = 2.7
zMax = 2.1
zs = expm1(linspace(log(1), log(1+zMax), 1024))
dNdz = (1+zs)**(gamma-1)*4*pi*Planck15.differential_comoving_volume(zs).to(u.Gpc**3/u.sr).value
Nz = cumtrapz(dNdz, zs, initial=0)
icdf_z = interp1d(Nz/Nz[-1], zs)
p_z = interp1d(zs, dNdz/Nz[-1])
def draw_z(size=1):
return icdf_z(np.random.uniform(size=size))
```
Uniform on the sky:
```
def draw_ra_dec(size=1):
ra = np.random.uniform(low=0, high=2*pi, size=size)
sin_dec = np.random.uniform(low=-1, high=1, size=size)
return ra, arcsin(sin_dec)
```
Drawn flat in $-1 < \chi_\mathrm{eff} < 1$.
```
def draw_chieff(size=1):
return np.random.uniform(low=-1, high=1, size=size)
```
This is the code that actually generates the mergers and down-selects them according to an SNR threshold. Notable bits:
* If you change the `psd` argument in `compute_detector_snrs`, you can operate at "early" sensitivity (i.e. O1+O2 sensitivity).
* The `approx_det_cut` function is intended to reject hopeless signals by drawing a line in the $m_1$-$z$ plane that can quickly eliminate signals with masses too small to be detectable (so we don't have to generate their waveforms; this is particularly helpful because low-mass waveforms from $\sim 10 \, \mathrm{Hz}$ are very long, so computationally expensive to generate). **It is not curretly properly tuned, however, so will reject detectable signals.** (This is likely not a major issue with $\chi_\mathrm{eff}$ selection effects, but we should check that.
```
fmin = 9.0
fref = 40.0
psdstart = 10.0
snr_thresh = 8*sqrt(2) # 8 in at least two detectors
def next_pow_two(x):
x2 = 1
while x2 < x:
x2 = x2 << 1
return x2
def generate_waveform(m1, m2, chi_eff, z):
dL = cosmo.Planck15.luminosity_distance(z).to(u.Gpc).value
tmax = ls.SimInspiralChirpTimeBound(fmin, m1*(1+z)*lal.MSUN_SI, m2*(1+z)*lal.MSUN_SI, 0.0, 0.0) + 2
df = 1.0/next_pow_two(tmax)
fmax = 2048.0 # Hz --- based on max freq of 5-5 inspiral
cos_i = np.random.uniform(low=-1, high=1)
phi_ref = np.random.uniform(low=0, high=2*pi)
hp, hc = ls.SimInspiralChooseFDWaveform((1+z)*m1*lal.MSUN_SI, (1+z)*m2*lal.MSUN_SI, 0.0, 0.0, chi_eff, 0.0, 0.0, chi_eff, dL*1e9*lal.PC_SI, arccos(cos_i), phi_ref, 0.0, 0.0, 0.0, df, fmin, fmax, fref, None, ls.IMRPhenomPv2)
return hp, hc
def compute_detector_snrs(hp, hc, ra, dec, psd='design'):
psi = np.random.uniform(low=0, high=2*pi)
gmst = np.random.uniform(low=0, high=2*pi)
snrs = []
for det in ['H1', 'L1', 'V1']:
Fp, Fc = lal.ComputeDetAMResponse(lal.cached_detector_by_prefix[det].response, ra, dec, psi, gmst)
h = lal.CreateCOMPLEX16FrequencySeries("h", lal.LIGOTimeGPS(0), 0.0, hp.deltaF, hp.sampleUnits, hp.data.length)
h.data.data = Fp*hp.data.data + Fc*hc.data.data
fmax = hp.deltaF*(hp.data.length - 1)
df = hp.deltaF
Nf = hp.data.length
fs = linspace(0, fmax, Nf)
sel = fs > psdstart
sffs = lal.CreateREAL8FrequencySeries("psds", 0, 0.0, df, lal.DimensionlessUnit, fs.shape[0])
if det == 'H1' or det == 'L1':
if psd == 'early':
ls.SimNoisePSDaLIGOEarlyHighSensitivityP1200087(sffs, psdstart)
elif psd == 'design':
ls.SimNoisePSDaLIGOaLIGODesignSensitivityT1800044(sffs, psdstart)
else:
raise ValueError('psd must be one of early, design')
elif det == 'V1':
if psd == 'early':
ls.SimNoisePSDAdVEarlyHighSensitivityP1200087(sffs, psdstart)
elif psd == 'design':
ls.SimNoisePSDAdVDesignSensitivityP1200087(sffs, psdstart)
else:
raise ValueError('psd must be one of early, design')
else:
raise ValueError('detector must be one of H1, L1, V1')
rho = ls.MeasureSNRFD(h, sffs, psdstart, -1.0)
n = randn()
snrs.append(sqrt((rho+n)*(rho+n)))
return array(snrs)
# def approx_det_cut(z):
# z0 = 0.1
# m0 = 5
# z1 = 0.8
# m1 = 35
# return m1*(z-z0)/(z1-z0) + m0*(z-z1)/(z0-z1)
def approx_det_cut(z):
m, b = [20.80266868, -0.9]
return m*z + b
def draw_snr(m1, m2, chi_eff, z, ra, dec):
if m1 < approx_det_cut(z):
return (m1, m2, chi_eff, z, ra, dec, 0.0)
else:
hp, hc = generate_waveform(m1, m2, chi_eff, z)
snrs = compute_detector_snrs(hp, hc, ra, dec)
snr = np.sqrt(np.sum(snrs*snrs))
return (m1,m2,chi_eff, z,ra,dec,snr)
def draw_snr_onearg(x):
return draw_snr(*x)
def downselect(m1s, m2s, chi_effs, zs, ras, decs, processes=None):
m1s_sel = []
m2s_sel = []
chi_effs_sel = []
zs_sel = []
ras_sel = []
decs_sel = []
snrs_sel = []
#s = m1s > approx_mdet_threshold(zs)
p = multi.Pool(processes=processes)
try:
for m1, m2, chi_eff, z, ra, dec, snr in tqdm_notebook(p.imap(draw_snr_onearg, zip(m1s, m2s, chi_effs, zs, ras, decs), chunksize=1024), total=len(zs)):
if snr > snr_thresh:
m1s_sel.append(m1)
m2s_sel.append(m2)
chi_effs_sel.append(chi_eff)
zs_sel.append(z)
ras_sel.append(ra)
decs_sel.append(dec)
snrs_sel.append(snr)
finally:
p.close()
return array(m1s_sel), array(m2s_sel), array(chi_effs_sel), array(zs_sel), array(ras_sel), array(decs_sel), array(snrs_sel)
```
To accurately fit a population of size $N$, we need to ensure we can compute the Monte-Carlo selection integral with a fractional uncertainty that is $1/2\sqrt{N}$. This will require *at least* $4*N$ Monte-Carlo samples (and actually more, because the population we are fitting will not match the draw population, so each M-C sample will not "count fully" and we will have fewer "effective" samples than detected draws). Conservatively, I chose the number here so that the number of detected samples would be $\mathcal{O}(10 \times 5000)$ so that we can probably analyze a full five years' of design sensitivity (i.e. 5k BBH mergers), though I'm not sure we would want to.
```
N = 1 << 23
m1s, m2s = draw_m1m2(size=N)
zs = draw_z(size=N)
chi_effs = draw_chieff(size=N)
ras, decs = draw_ra_dec(size=N)
```
The Monte-Carlo selection function estimate takes about three hours to run on my laptop (four cores):
```
m1s_sel, m2s_sel, chi_effs_sel, zs_sel, ras_sel, decs_sel, snrs_sel = downselect(m1s, m2s, chi_effs, zs, ras, decs)
```
Checking that the chosen $z_\mathrm{max}$ doesn't artificially truncate the redshifts of observed sources (it sorta does, but not in a major way).
```
sns.distplot(zs_sel)
axvline(zMax)
```
Checking the approximate detection threshold; since some sources run up against this line, it needs to be tuned further (or just dropped, but then the runtime of the Monte-Carlo explodes by a factor of 10 or more!).
```
zs = linspace(0, zMax, 1000)
scatter(zs_sel[::100], m1s_sel[::100])
plot(zs, approx_det_cut(zs))
```
We have about 50k sources.
```
len(zs_sel)
```
The code below solves a constrained optimization problem: maximize the area under the approximate selection cut line, subject to the constraint that every observed system lies above it. **Do not run it on more than ~1k detections, as the computation is expensive.**
```
c = -array([0.5*zMax*zMax, zMax])
A_ub = column_stack((zs_sel, ones_like(zs_sel)))
b_ub = m1s_sel
sol = linprog(c, A_ub, b_ub, bounds=[(0, None), (None, None)])
sol
0.9*(sol.x)
```
Here we finally come to the selection function: $p\left( \chi_eff \mid \mathrm{detected} \right)$:
```
sns.distplot(chi_effs_sel)
```
And now we save the set of selected samples (note that I've also run this notebook in a different configuration, and also selected samples appropriate to early---that is, GWTC-1---sensitivity).
```
with h5py.File('design-selected.h5', 'w') as f:
def cd(n, d):
f.create_dataset(n, data=d, compression='gzip', shuffle=True)
f.attrs['N_draw'] = N
cd('m1s', m1s_sel)
cd('m2s', m2s_sel)
cd('chi_effs', chi_effs_sel)
cd('zs', zs_sel)
cd('ras', ras_sel)
cd('decs', decs_sel)
cd('snrs', snrs_sel)
```
| github_jupyter |

### ODPi Egeria Hands-On Lab
# Welcome to the Understanding an Asset Lab
## Introduction
ODPi Egeria is an open source project that provides open standards and implementation libraries to connect tools, catalogues and platforms together so they can share information about data and technology (called metadata).
In this hands-on lab you will get a chance to explore the types of properties (metadata) that can be stored about an asset.
## The scenario
Callie Quartile is a data scientist at Coco Pharmaceuticals. She is responsible for analysing the data that is generated during a clinical trial.

Callie's userId is `calliequartile`.
```
calliesUserId = "calliequartile"
```
Clinical trial data is stored in Coco Pharmaceutical's data lake as **data sets**. A data set is a collection of related data. For example, a data set may be the names, addresses of all patients in a clinical trail. Or it may be the measurements from a particular test,
of the notes from a physician as a result of a patient consultation. There is no upper or lower limit on the
size of a data set. It just has to contain a useful collection of data.
In this hands-on lab Callie wants to create clusters of patients involved in the trial based on their characteristics.
This will help her look for patterns in the measurement data they receive through the clinical trial.
# Setting up
Coco Pharmaceuticals make widespread use of ODPi Egeria for tracking and managing their data and related assets.
Figure 1 below shows the metadata servers and the platforms that are hosting them.

> **Figure 1:** Coco Pharmaceuticals' OMAG Server Platforms
```
import os
corePlatformURL = os.environ.get('corePlatformURL','http://localhost:8080')
dataLakePlatformURL = os.environ.get('dataLakePlatformURL','http://localhost:8081')
devPlatformURL = os.environ.get('devPlatformURL','http://localhost:8082')
```
Callie is using the research team's metadata server called `cocoMDS3`. This server is hosted on the Core OMAG Server Platform.
```
server = "cocoMDS3"
serverPlatformURL = corePlatformURL
```
The following request checks that this server is running.
```
import requests
import pprint
import json
adminUserId = "garygeeke"
isServerActiveURL = serverPlatformURL + "/open-metadata/platform-services/users/" + adminUserId + "/server-platform/servers/" + server + "/status"
print (" ")
print ("GET " + isServerActiveURL)
print (" ")
response = requests.get(isServerActiveURL)
print ("Returns:")
prettyResponse = json.dumps(response.json(), indent=4)
print (prettyResponse)
print (" ")
serverStatus = response.json().get('active')
if serverStatus == True:
print("Server " + server + " is active - ready to begin")
else:
print("Server " + server + " is down - start it before proceeding")
```
----
The next set of code sets up the asset - it is subject to change.
```
assetOwnerURL = serverPlatformURL + '/servers/' + server + '/open-metadata/access-services/asset-owner/users/peterprofile'
createAssetURL = assetOwnerURL + '/assets/data-files/csv'
print (createAssetURL)
jsonHeader = {'content-type':'application/json'}
body = {
"class" : "NewCSVFileAssetRequestBody",
"displayName" : "Drop Foot Clinical Trial Patients",
"description" : "List of patients registered for the drop foot clinical trial.",
"fullPath" : "file://secured/research/clinical-trials/drop-foot/Patients.csv"
}
response=requests.post(createAssetURL, json=body, headers=jsonHeader)
response.json()
getAssetsURL = serverAssetOwnerURL + '/assets/by-name?startFrom=0&pageSize=50'
searchString="*Patient*"
print (" ")
print ("GET " + getAssetsURL)
print ("{ " + searchString + " }")
print (" ")
response=requests.post(getAssetsURL, data=searchString)
print ("Returns:")
prettyResponse = json.dumps(response.json(), indent=4)
print (prettyResponse)
print (" ")
if response.json().get('assets'):
if len(response.json().get('assets')) == 1:
print ("1 asset found")
else:
print (str(len(response.json().get('assets'))) + " assets found")
else:
print ("No assets found")
```
| github_jupyter |
```
import pickle
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import re
import string
from keras.models import Model, load_model
from keras.layers import Dense, Embedding, Input, GRU
from keras.layers import LSTM, Bidirectional, GlobalMaxPool1D, Dropout,GlobalAveragePooling1D,Conv1D
from keras.preprocessing import text, sequence
from keras.callbacks import EarlyStopping, ModelCheckpoint
from models_def import Attention
# prepare other feat
fl = [
'../features/other_feat.pkl',
'../features/lgb1_feat.pkl',
'../features/rf1_feat.pkl',
'../features/gbrt1_feat.pkl',
'../features/lr_feat1.pkl',
'../features/lr_feat2.pkl',
'../features/ridge_feat1.pkl',
'../features/ridge_feat2.pkl',
'../features/mnb_feat1.pkl',
'../features/mnb_feat2.pkl',
'../features/wordbatch_feat.pkl',
'../features/tilli_lr_feat.pkl',
]
def get_feat(f):
with open(f,'rb') as fin:
a,b = pickle.load(fin)
return a,b
# load feats
train_x,test_x = [],[]
for feat in fl:
print('file path',feat)
a,b = pickle.load(open(feat,'rb'))
print(a.shape,b.shape)
train_x.append(a)
test_x.append(b)
train_x = np.nan_to_num(np.hstack(train_x))
test_x = np.nan_to_num(np.hstack(test_x))
print(train_x.shape)
print(train_x[0])
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
train_x = scaler.fit_transform(train_x)
test_x = scaler.transform(test_x)
print(train_x[0])
max_features = 160000
maxlen = 250
# Contraction replacement patterns
cont_patterns = [
(b'(W|w)on\'t', b'will not'),
(b'(C|c)an\'t', b'can not'),
(b'(I|i)\'m', b'i am'),
(b'(A|a)in\'t', b'is not'),
(b'(\w+)\'ll', b'\g<1> will'),
(b'(\w+)n\'t', b'\g<1> not'),
(b'(\w+)\'ve', b'\g<1> have'),
(b'(\w+)\'s', b'\g<1> is'),
(b'(\w+)\'re', b'\g<1> are'),
(b'(\w+)\'d', b'\g<1> would'),
(b'<3', b' heart '),
(b':d', b' smile '),
(b':dd', b' smile '),
(b':p', b' smile '),
(b'8\)', b' smile '),
(b':-\)', b' smile '),
(b':\)', b' smile '),
(b';\)', b' smile '),
(b'\(-:', b' smile '),
(b'\(:', b' smile '),
(b'yay!', b' good '),
(b'yay', b' good '),
(b'yaay', b' good '),
(b':/', b' worry '),
(b':>', b' angry '),
(b":'\)", b' sad '),
(b':-\(', b' sad '),
(b':\(', b' sad '),
(b':s', b' sad '),
(b':-s', b' sad '),
(b'\d{1,3}.\d{1,3}.\d{1,3}.\d{1,3}', b' '),
(b'(\[[\s\S]*\])', b' '),
(b'[\s]*?(www.[\S]*)', b' ')
]
patterns = [(re.compile(regex), repl) for (regex, repl) in cont_patterns]
def new_clean(text):
""" Simple text clean up process"""
# 1. Go to lower case (only good for english)
# Go to bytes_strings as I had issues removing all \n in r""
clean = bytes(text.lower(), encoding="utf-8")
# replace words like hhhhhhhhhhhhhhi with hi
for ch in string.ascii_lowercase:
pattern = bytes(ch+'{3,}', encoding="utf-8")
clean = re.sub(pattern, bytes(ch, encoding="utf-8"), clean)
# 2. Drop \n and \t
clean = clean.replace(b"\n", b" ")
clean = clean.replace(b"\t", b" ")
clean = clean.replace(b"\b", b" ")
clean = clean.replace(b"\r", b" ")
# 3. Replace english contractions
for (pattern, repl) in patterns:
clean = re.sub(pattern, repl, clean)
# 4. Drop puntuation
# I could have used regex package with regex.sub(b"\p{P}", " ")
exclude = re.compile(b'[%s]' % re.escape(bytes(string.punctuation, encoding='utf-8')))
clean = b" ".join([exclude.sub(b'', token) for token in clean.split()])
# 5. Drop numbers - as a scientist I don't think numbers are toxic ;-)
clean = re.sub(b"\d+", b" ", clean)
# 6. Remove extra spaces - At the end of previous operations we multiplied space accurences
clean = re.sub(b'\s+', b' ', clean)
# Remove ending space if any
clean = re.sub(b'\s+$', b'', clean)
# 7. Now replace words by words surrounded by # signs
# e.g. my name is bond would become #my# #name# #is# #bond#
# clean = re.sub(b"([a-z]+)", b"#\g<1>#", clean)
clean = re.sub(b" ", b"# #", clean) # Replace space
clean = b"#" + clean + b"#" # add leading and trailing #
return str(clean, 'utf-8')
def clean_text( text ):
text = text.lower().split()
text = " ".join(text)
text = re.sub(r"[^A-Za-z0-9^,!.\/'+\-=]", " ", text)
text = re.sub(r"what's", "what is ", text)
text = re.sub(r"\'s", " ", text)
text = re.sub(r"\'ve", " have ", text)
text = re.sub(r"can't", "cannot ", text)
text = re.sub(r"n't", " not ", text)
text = re.sub(r"i'm", "i am ", text)
text = re.sub(r"\'re", " are ", text)
text = re.sub(r"\'d", " would ", text)
text = re.sub(r"\'ll", " will ", text)
text = re.sub(r",", " ", text)
text = re.sub(r"\.", " ", text)
text = re.sub(r"!", " ! ", text)
text = re.sub(r"\/", " ", text)
text = re.sub(r"\^", " ^ ", text)
text = re.sub(r"\+", " + ", text)
text = re.sub(r"\-", " - ", text)
text = re.sub(r"\=", " = ", text)
text = re.sub(r"'", " ", text)
text = re.sub(r"(\d+)(k)", r"\g<1>000", text)
text = re.sub(r":", " : ", text)
text = re.sub(r" e g ", " eg ", text)
text = re.sub(r" b g ", " bg ", text)
text = re.sub(r" u s ", " american ", text)
text = re.sub(r"\0s", "0", text)
text = re.sub(r" 9 11 ", "911", text)
text = re.sub(r"e - mail", "email", text)
text = re.sub(r"j k", "jk", text)
text = re.sub(r"\s{2,}", " ", text)
return new_clean(text)
train = pd.read_csv("../input/train.csv")
test = pd.read_csv("../input/test.csv")
list_sentences_train = train["comment_text"].fillna("CVxTz").apply(clean_text).values
list_classes = ["toxic", "severe_toxic", "obscene", "threat", "insult", "identity_hate"]
y = train[list_classes].values
list_sentences_test = test["comment_text"].fillna("CVxTz").apply(clean_text).values
print(y.shape)
tokenizer = text.Tokenizer(num_words=max_features)
tokenizer.fit_on_texts(list(list_sentences_train))
list_tokenized_train = tokenizer.texts_to_sequences(list_sentences_train)
list_tokenized_test = tokenizer.texts_to_sequences(list_sentences_test)
X_train = sequence.pad_sequences(list_tokenized_train, maxlen=maxlen)
X_test = sequence.pad_sequences(list_tokenized_test, maxlen=maxlen)
print(X_train.shape,X_test.shape)
# check word_index
tmp_cnt = 0
for k in tokenizer.word_index:
print(k,tokenizer.word_index[k])
tmp_cnt += 1
if tmp_cnt >5:
break
word_idx = tokenizer.word_index
# read word2vec
#
word_vec_dict = {}
with open('../crawl-300d-2M.vec') as f:
first_line_flag = True
for line in f:
if first_line_flag:
first_line_flag= False
continue
v_list = line.rstrip().split(' ')
k = str(v_list[0])
v = np.array([float(x) for x in v_list[1:]])
word_vec_dict[k] = v
print(len(word_vec_dict))
print('Preparing embedding matrix')
EMBEDDING_DIM = 300
nb_words = min(max_features,len(word_idx))
embedding_matrix = np.zeros((nb_words, EMBEDDING_DIM))
for word,i in word_idx.items():
if i >= max_features:
continue
else:
if word in word_vec_dict:
embedding_matrix[i] = word_vec_dict[word]
print('Null word embeddings: %d' % np.sum(np.sum(embedding_matrix, axis=1) == 0))
del word_vec_dict
from sklearn.metrics import log_loss,accuracy_score
from keras.models import Model
from keras.layers import Input, Dense, Embedding, SpatialDropout1D, concatenate
from keras.layers import GRU, Bidirectional, GlobalAveragePooling1D, GlobalMaxPooling1D, CuDNNLSTM
from keras.preprocessing import text, sequence
from keras.callbacks import Callback
def eval_val(y,train_x):
res = 0
acc_res = 0
for i in range(6):
curr_loss = log_loss(y[:,i],train_x[:,i])
acc = accuracy_score(y[:,i],train_x[:,i].round())
print(i,curr_loss,acc)
res += curr_loss
acc_res += acc
print('final',res/6, acc_res/6)
def get_model(comp):
inp = Input(shape=(maxlen, ))
inp_2 = Input(shape=[train_x.shape[1]], name="other")
emb = Embedding(nb_words, EMBEDDING_DIM, weights=[embedding_matrix],trainable=False)(inp)
emb = SpatialDropout1D(0.4)(emb)
x = Bidirectional(CuDNNLSTM(128, return_sequences=True))(emb)
avg_pool = GlobalAveragePooling1D()(x)
max_pool = GlobalMaxPooling1D()(x)
att = Attention(maxlen)(x)
conc = concatenate([att, avg_pool, max_pool, inp_2])
conc = Dense(256, activation="relu")(conc)
outp = Dense(6, activation="sigmoid")(conc)
model = Model(inputs=[inp,inp_2], outputs=outp)
if comp:
model.compile(loss='binary_crossentropy',
optimizer='nadam',
metrics=['accuracy'])
return model
print('def model done')
from sklearn.model_selection import KFold
import gc
from keras import backend as K
def kf_train(fold_cnt=3,rnd=1):
kf = KFold(n_splits=fold_cnt, shuffle=False, random_state=233*rnd)
train_pred, test_pred = np.zeros((159571,6)),np.zeros((153164,6))
for train_index, test_index in kf.split(X_train):
# x,y
curr_x,curr_y = X_train[train_index],y[train_index]
curr_other_x = train_x[train_index]
hold_out_x,hold_out_y = X_train[test_index],y[test_index]
hold_out_other_x = train_x[test_index]
# model
model = get_model(True)
batch_size = 64
epochs = 6
file_path="weights_base.best.h5"
checkpoint = ModelCheckpoint(file_path, monitor='val_loss', verbose=1, save_best_only=True, mode='min')
callbacks_list = [checkpoint]
# train and pred
model.fit([curr_x,curr_other_x], curr_y,
batch_size=batch_size, epochs=epochs,
validation_data=([hold_out_x,hold_out_other_x],hold_out_y),
callbacks=callbacks_list)
model.load_weights(file_path)
y_test = model.predict([X_test,test_x])
test_pred += y_test
hold_out_pred = model.predict([hold_out_x,hold_out_other_x])
train_pred[test_index] = hold_out_pred
# clear
del model
gc.collect()
K.clear_session()
test_pred = test_pred / fold_cnt
print('-------------------------------')
print('all eval',eval_val(y,train_pred))
return train_pred, test_pred
print('def done')
import pickle
sample_submission = pd.read_csv("../input/sample_submission.csv")
train_pred,test_pred = kf_train(fold_cnt=5,rnd=42)
print(train_pred.shape,test_pred.shape)
# 40000,150,lstm + global max_pool
# final 0.0407274256871 0.984048897774
# 100000,150 lstm + attention, glove embedding
# final 0.0404159162853 0.984188856371, pub 9849
# 3996, 4093
# 100000,150 lstm + attention, use spacial dropout,spacial 0.2, last dropout 0.5, fasttext embedding
# 1st epo 4016, 2nd epo 4117, not better compare to glove res
# 100000,150,test arch
# x = Embedding(nb_words, EMBEDDING_DIM, weights=[embedding_matrix],trainable=False)(inp)
# x = Dropout(0.2)(x)
# x = Bidirectional(LSTM(64, return_sequences=True))(x)
# x = Attention(maxlen)(x)
# x = Dense(6, activation="sigmoid")(x)
# 1st epo 4116, not good
# 100000,150,test arch
# x = Embedding(nb_words, EMBEDDING_DIM, weights=[embedding_matrix],trainable=False)(inp)
# x = Dropout(0.2)(x)
# x = Bidirectional(CuDNNLSTM(64, return_sequences=True))(x)
# att = Attention(maxlen)(x)
# avg_pool = GlobalAveragePooling1D()(x)
# max_pool = GlobalMaxPooling1D()(x)
# conc = concatenate([att,avg_pool, max_pool])
# x = Dense(256, activation="relu")(conc)
# x = Dense(6, activation="sigmoid")(x)
# 1st epo , old LSTM 3945
# to save time ,change to CuDNNLSTM
# 1st epo , 3928, 4 fold: final 0.0393455938053 0.984445795289
# 10 fold: final 0.0391567844913 0.984588887287 PUB 9857
# new adj
# 5 fold: final 0.037041229455724294 0.985068297706559 PUB 9862
sample_submission[list_classes] = test_pred
sample_submission.to_csv("../results/lstm_attention_fasttext_sample_5.gz", index=False, compression='gzip')
with open('../features/lstm_attention_fasttext_5_feat.pkl','wb') as fout:
pickle.dump([train_pred,test_pred],fout)
print(sample_submission.head())
print('===================================')
train_pred,test_pred = kf_train(fold_cnt=10,rnd=42)
print(train_pred.shape,test_pred.shape)
sample_submission[list_classes] = test_pred
sample_submission.to_csv("../results/lstm_attention_fasttext_sample_10.gz", index=False, compression='gzip')
with open('../features/lstm_attention_fasttext_10_feat.pkl','wb') as fout:
pickle.dump([train_pred,test_pred],fout)
print(sample_submission.head())
print('===================================')
# final 0.03672656107307943 0.9850442749622426 PUB 9863
```
| github_jupyter |
# Deep Reinforcement Learning for Optimal Execution of Portfolio Transactions
```
import utils
# Get the default financial and AC Model parameters
financial_params, ac_params = utils.get_env_param()
financial_params
ac_params
import numpy as np
import syntheticChrissAlmgren as sca
from ddpg_agent import Agent
from collections import deque
# Create simulation environment
env = sca.MarketEnvironment()
# Initialize Feed-forward DNNs for Actor and Critic models.
agent1 = Agent(state_size=env.observation_space_dimension(), action_size=env.action_space_dimension(),random_seed = 1225)
agent2 = Agent(state_size=env.observation_space_dimension(), action_size=env.action_space_dimension(),random_seed = 108)
# Set the liquidation time
lqt = 60
# Set the number of trades
n_trades = 60
# Set trader's risk aversion
tr1 = 1e-6
tr2 = 1e-6
# Set the number of episodes to run the simulation
episodes = 1300
shortfall_list = []
shortfall_hist1 = np.array([])
shortfall_hist2 = np.array([])
shortfall_deque1 = deque(maxlen=100)
shortfall_deque2 = deque(maxlen=100)
for episode in range(episodes):
# Reset the enviroment
cur_state = env.reset(seed = episode, liquid_time = lqt, num_trades = n_trades, lamb1 = tr1,lamb2 = tr2)
# set the environment to make transactions
env.start_transactions()
for i in range(n_trades + 1):
# Predict the best action for the current state.
cur_state1 = np.delete(cur_state,8)
cur_state2 = np.delete(cur_state,7)
#print(cur_state[5:])
action1 = agent1.act(cur_state1, add_noise = True)
action2 = agent2.act(cur_state2, add_noise = True)
#print(action1,action2)
# Action is performed and new state, reward, info are received.
new_state, reward1, reward2, done1, done2, info = env.step(action1,action2)
# current state, action, reward, new state are stored in the experience replay
new_state1 = np.delete(new_state,8)
new_state2 = np.delete(new_state,7)
agent1.step(cur_state1, action1, reward1, new_state1, done1)
agent2.step(cur_state2, action2, reward2, new_state2, done2)
# roll over new state
cur_state = new_state
if info.done1 and info.done2:
shortfall_hist1 = np.append(shortfall_hist1, info.implementation_shortfall1)
shortfall_deque1.append(info.implementation_shortfall1)
shortfall_hist2 = np.append(shortfall_hist2, info.implementation_shortfall2)
shortfall_deque2.append(info.implementation_shortfall2)
break
if (episode + 1) % 100 == 0: # print average shortfall over last 100 episodes
print('\rEpisode [{}/{}]\tAverage Shortfall for Agent1: ${:,.2f}'.format(episode + 1, episodes, np.mean(shortfall_deque1)))
print('\rEpisode [{}/{}]\tAverage Shortfall for Agent2: ${:,.2f}'.format(episode + 1, episodes, np.mean(shortfall_deque2)))
shortfall_list.append([np.mean(shortfall_deque1),np.mean(shortfall_deque2)])
print('\nAverage Implementation Shortfall for Agent1: ${:,.2f} \n'.format(np.mean(shortfall_hist1)))
print('\nAverage Implementation Shortfall for Agent2: ${:,.2f} \n'.format(np.mean(shortfall_hist2)))
shortfall = np.array(shortfall_list)
np.save('1e-6_1e-6_cooporation_shorfall_list.npy',shortfall)
print(tr1,tr2)
cur_state = env.reset(seed = episode, liquid_time = lqt, num_trades = n_trades, lamb1 = tr1,lamb2 = tr2)
# set the environment to make transactions
env.start_transactions()
trajectory = np.zeros([n_trades+1,2])
for i in range(n_trades + 1):
trajectory[i] = cur_state[7:]
print(cur_state[7:])
# Predict the best action for the current state.
cur_state1 = np.delete(cur_state,8)
cur_state2 = np.delete(cur_state,7)
#print(cur_state[5:])
action1 = agent1.act(cur_state1, add_noise = True)
action2 = agent2.act(cur_state2, add_noise = True)
#print(action1,action2)
# Action is performed and new state, reward, info are received.
new_state, reward1, reward2, done1, done2, info = env.step(action1,action2)
# current state, action, reward, new state are stored in the experience replay
new_state1 = np.delete(new_state,8)
new_state2 = np.delete(new_state,7)
agent1.step(cur_state1, action1, reward1, new_state1, done1)
agent2.step(cur_state2, action2, reward2, new_state2, done2)
# roll over new state
cur_state = new_state
if info.done1 and info.done2:
shortfall_hist1 = np.append(shortfall_hist1, info.implementation_shortfall1)
shortfall_deque1.append(info.implementation_shortfall1)
shortfall_hist2 = np.append(shortfall_hist2, info.implementation_shortfall2)
shortfall_deque2.append(info.implementation_shortfall2)
break
if (episode + 1) % 100 == 0: # print average shortfall over last 100 episodes
print('\rEpisode [{}/{}]\tAverage Shortfall for Agent1: ${:,.2f}'.format(episode + 1, episodes, np.mean(shortfall_deque1)))
print('\rEpisode [{}/{}]\tAverage Shortfall for Agent2: ${:,.2f}'.format(episode + 1, episodes, np.mean(shortfall_deque2)))
np.save('1e-6_1e-6_competition_trajectory_1500.npy',trajectory)
%matplotlib inline
import matplotlib.pyplot as plt
import utils
# We set the default figure size
plt.rcParams['figure.figsize'] = [17.0, 7.0]
# Set the number of days to sell all shares (i.e. the liquidation time)
l_time = 60
# Set the number of trades
n_trades = 60
# Set the trader's risk aversion
t_risk = 1e-6
# Plot the trading list and trading trajectory. If show_trl = True, the data frame containing the values of the
# trading list and trading trajectory is printed
utils.plot_trade_list(lq_time = l_time, nm_trades = n_trades, tr_risk = t_risk, show_trl = True)
```
| github_jupyter |
```
# Erasmus+ ICCT project (2018-1-SI01-KA203-047081)
# Toggle cell visibility
from IPython.display import HTML
tag = HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide()
} else {
$('div.input').show()
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
Toggle cell visibility <a href="javascript:code_toggle()">here</a>.''')
display(tag)
# Hide the code completely
# from IPython.display import HTML
# tag = HTML('''<style>
# div.input {
# display:none;
# }
# </style>''')
# display(tag)
continuous_update=False
%matplotlib notebook
import matplotlib.pyplot as plt
import numpy as np
from scipy.integrate import odeint
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
```
## Equazioni differenziali - sistemi lineari e non lineari
Questo notebook copre vari sistemi definiti dalle seguenti equazioni differenziali:
- vibrazioni non lineari: $\ddot{x}+0.25\dot{x}+5\sin{x}=F$,
- vibrazioni lineari: $\ddot{x}+0.25\dot{x}+5x=F$,
- sistema non lineare: $\ddot{x}+0.003\dot{x}x=F$,
- sistema lineare dipendente dal tempo: $\ddot{x}-0.2{x}t^{\frac{1}{4}}=F$,
- sistema non lineare dipendente dal tempo: $\ddot{x}+0.0001x^2t=F$, <br>
dove $F$ denota l'input (funzione gradino), $t$ il tempo e $x$ la variabile dipendente.
La figura a sinistra mostra le risposte temporali per diverse condizioni iniziali e diversi input. La curva blu spessa rappresenta la risposta temporale in base ai valori impostati tramite i cursori. Le altre curve rappresentano risposte temporali ottenute da multipli di 1) un valore della prima condizione iniziale (spostamento) $x(0)$, 2) un valore della seconda condizione iniziale (velocità) $\dot x(0)$, o 3) dell'input del sistema (a seconda della selezione *Cosa variare*). I multipli sono valori interi compresi tra -5 e +5.
<br>
*Esempio*:
<br>
$\text{input}=0.7$, $x(0)=2.0$ e $\dot x(0)=5.5$, opzione *input* selezionata.
Output: la curva spessa si basa sui valori impostati sopra. Le altre curve si basano sugli stessi valori $x(0)$ e $\dot x(0)$, ma con le seguenti ampiezze del gradino:
$0.7\begin{pmatrix}-5& -4& -3& -2& -1& 0& 1& 2& 3& 4& 5\end{pmatrix}=\begin{pmatrix}-3.5& -2.8& -2.1& -1.4& -0.7& 0& 0.7& 1.4& 2.1& 2.8& 3.5\end{pmatrix}$.
La figura a destra mostra i valori di tutte le risposte temporali mostrate nella figura a sinistra nell'istante di tempo selezionato. Le etichette di graduazione dell'asse x indicano i valori interi compresi tra -5 e +5. Per un sistema lineare questo grafico è sempre una linea.
---
### Come usare questo notebook?
Sposta i cursori per modificare le risposte temporali in funzione dell'ampiezza del gradino selezionato (*input*) o delle condizioni iniziali dei sistemi predefiniti introdotti sopra. È possibile passare da un sistema all'altro premendo il pulsante corrispondente.
#### Legenda:
- *input*: valore dell'ampiezza dell'ingresso (gradino).
- *delay*: ritardo di inizio integrazione (per sistemi dipendenti dal tempo).
- *istante di tempo*: istante in cui vengono mostrate le risposte nella figura a destra.
- $x(0)$: valore della prima condizione iniziale (spostamento).
- $\dot x(0)$: valore della seconda condizione iniziale (velocità).
<br>
```
# diferential equations:
def nonlinear_vibr(y,t, x): # y začetni pogoj, t čas, x krmilni vhod
'''nonlinear vibration'''
th, om = y
b=0.25
c=5.
dydt = (om, x - b*om - c*np.sin(th))
return dydt
def linear_vibr(y,t,x):
'''linear vibration'''
th, om = y
b=0.25
c=5.
dydt = (om, x - b*om - c*th)
return dydt
def nonlinear(y,t,x):
'''nonlinear d.e.'''
c, v = y
dydt = (v, x - 0.003*c*v)
return dydt
def time_dependent(y,t,x):
'''time dependent d.e.'''
c, v = y
dydt = (v, x + .2*c*t**(1/4))
return dydt
def first_order (y, t, x):
'''first order nonlinear time dependent d.e.'''
c, v = y
dydt = (v, x - .0001*c*c*t)
return dydt
fig = plt.figure(figsize=(9.8, 4),num="Equazioni differenziali (ED)")
response = fig.add_subplot(1, 2, 1)
pnts_of_resp = fig.add_subplot(1, 2, 2)
y0 = np.array([0.,0.]) # init. conditions
t_end = 10
x=1
t_0=1
t_calc=1
y0_0=1
y0_1=1
def func():
global y0, x, t_0, t_calc, y0_0, y0_1
y0[0] = y0_0
y0[1] = y0_1
t = np.linspace(0,t_end,t_end*10+1) #time points, where response (solution) is calculated
linearity = [] # point to look, if d.e. is linear on not
t = t+ t_0
plus_minus = 5.
input_amp = np.round(np.linspace(-plus_minus,plus_minus,2*int(plus_minus)+1))
response.clear()
pnts_of_resp.clear()
current_button = buttons.index
for amp in input_amp:
if radio_buttons.index == 0:
i=amp*x
input_pack = [i, y0]
if radio_buttons.index == 1:
i=amp*float(y0[0])
input_pack = [x, [i, y0[1]]]
if radio_buttons.index == 2:
i=amp*float(y0[1])
input_pack = [x, [y0[0], i]]
if current_button == 0:
solution = odeint(nonlinear_vibr, input_pack[1], t, args=(input_pack[0],))
if current_button == 1:
solution = odeint(linear_vibr, input_pack[1], t, args=(input_pack[0],))
if current_button == 2:
solution = odeint(nonlinear, input_pack[1], t, args=(input_pack[0],))
if current_button == 3:
solution = odeint(time_dependent, input_pack[1], t, args=(input_pack[0],))
if current_button == 4:
solution = odeint(first_order, input_pack[1], t, args=(input_pack[0],))
solution = solution[:,0]
linearity.append(solution[int(t_calc*10)])
if amp == 1:
response.plot(t, solution, 'b', lw=2, label='risposta')
else:
response.plot(t, solution, lw=.7)
response.grid()
response.set_xlabel('$t$ [s]')
response.set_ylabel('risposta')
response.set_title('Soluzioni della ED')
response.legend()
pnts_of_resp.plot(input_amp, linearity, label='linearità')
pnts_of_resp.grid()
pnts_of_resp.legend()
pnts_of_resp.set_xlabel('Input o condizioni iniziali')
pnts_of_resp.set_ylabel('risposta nell\'istante %.1d' %t_calc)
pnts_of_resp.set_title('Linearità della ED')
style = {'description_width': 'initial'}
x_widget = widgets.FloatSlider(value=0.5,min=0,max=1,step=.1,description='input',
disabled=False,continuous_update=True,orientation='horizontal',readout=True,readout_format='.1f')
t_0_widget = widgets.IntSlider(value=0,min=0,max=10,step=1,description='delay [s]',
disabled=False,continuous_update=True,orientation='horizontal',readout=True,readout_format='.1d')
t_calc_widget = widgets.IntSlider(value=1,min=1,max=10,step=1,description='istante di tempo [s]',
disabled=False,continuous_update=True,orientation='horizontal',readout=True,readout_format='.1d',style=style)
y0_0_widget = widgets.FloatSlider(value=0.,min=0,max=10,step=.1,description='$x(0)$',
disabled=False,continuous_update=True,orientation='horizontal',readout=True,readout_format='.1f')
y0_1_widget = widgets.FloatSlider(value=0.,min=0,max=10,step=.1,description='$\dot x(0)$',
disabled=False,continuous_update=True,orientation='horizontal',readout=True,readout_format='.1f')
style = {'description_width': 'initial','button_width':'180px'}
buttons = widgets.ToggleButtons(options=['vibrazioni non lineari', 'vibrazioni lineari', 'non lineare', 'lineare tempo variante', 'non lineare tempo variante'],
description='Seleziona un sistema:',value = 'vibrazioni non lineari',disabled=False,style=style)
display(buttons)
i=0
def buttons_clicked(event):
global i
i+=1
if i%5==0: # for more efficient program
i=0
func()
buttons.observe(buttons_clicked)
radio_buttons = widgets.RadioButtons(options=['input', 'x(0)', u'\u1E8B''(0)'],
value='input',description='Cosa variare:',disabled=False,style=style)
j=0
def radio_buttons_clicked(event):
global j
j+=1
if j%5==0:
if radio_buttons.index==0:
x_widget.min=.1
y0_0_widget.min=0
y0_1_widget.min=0
if radio_buttons.index==1:
x_widget.min=0
y0_0_widget.min=.1
y0_1_widget.min=0
if radio_buttons.index==2:
x_widget.min=0
y0_0_widget.min=0
y0_1_widget.min=.1
j=0
func()
radio_buttons.observe(radio_buttons_clicked)
vbox = widgets.VBox([x_widget, t_0_widget, t_calc_widget, y0_0_widget, y0_1_widget])
hbox = widgets.HBox([vbox, radio_buttons])
#buttons.observe(buttons_clicked)
display(hbox)
k=0
def set_values(event):
global k, x, t_0, t_calc, y0_0, y0_1
k+=1
if k%3==0: # every three times is because .observe sends 3 events
k=0
x=x_widget.value
t_0=t_0_widget.value
t_calc=t_calc_widget.value
y0_0=y0_0_widget.value
y0_1=y0_1_widget.value
func()
x_widget.observe(set_values)
t_0_widget.observe(set_values)
t_calc_widget.observe(set_values)
y0_0_widget.observe(set_values)
y0_1_widget.observe(set_values)
#initialize:
def initialize():
set_values(0)
set_values(0)
set_values(0)
x_widget.min = .1
initialize()
```
| github_jupyter |
# Genomics experiment details.
We demonstrate the infinitesimal jackknife on a
publicly available data set of mice gene expression in Shoemaker et al. [2015].
Mice were infected with influenza virus, and gene expression was assessed several times after infection, so the observed data consists of expression levels $y_{gt}$ for genes $g = 1, ..., n_g$ and time points
$t = 1, ..., n_t$, where in this case $n_g = 1000$ and $n_t = 42$.
This notebook contains the first of three steps in the analysis. In this notebook, we will first load the data and define a basis with a hyperparameter we wish to select with cross validation. We then describe the two stages of our analysis: a regression stage and a clustering stage. We then save the data for further analysis by the notebooks ``load_and_refit`` and ``calculate_prediction_error``.
This notebook assumes you have already followed the instructions in ``README.md`` to install the necessary packages and create the dataset.
# Step 1: Initial fit.
```
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
import inspect
import os
import sys
import time
np.random.seed(3452453) # nothing special about this seed (we hope)!
from aistats2019_ij_paper import regression_mixture_lib as rm_lib
from aistats2019_ij_paper import regression_lib as reg_lib
from aistats2019_ij_paper import sensitivity_lib as sens_lib
from aistats2019_ij_paper import spline_bases_lib
from aistats2019_ij_paper import transform_regression_lib as trans_reg_lib
from aistats2019_ij_paper import loading_data_utils
from aistats2019_ij_paper import saving_gmm_utils
from aistats2019_ij_paper import mse_utils
import plot_utils_lib
```
## The first stage: Regression
### Load data
```
# Set bnp_data_repo to be the location of a clone of the repo
# https://github.com/NelleV/genomic_time_series_bnp
bnp_data_repo = '../../genomic_time_series_bnp'
y_train, y_test, train_indx, timepoints = loading_data_utils.load_genomics_data(
bnp_data_repo,
split_test_train = True,
train_indx_file = '../fits/train_indx.npy')
n_train = np.shape(y_train)[0]
print('number of genes in training set: \n', n_train)
n_test = np.shape(y_test)[0]
print('number of genes in test set: \n', n_test)
n_genes = n_train + n_test
test_indx = np.setdiff1d(np.arange(n_genes), train_indx)
gene_indx = np.concatenate((train_indx, test_indx))
```
Each gene $y_g$ has 42 observations. Observations are made at 14 timepoints, with 3 replicates at each timepoints.
```
n_t = len(timepoints)
n_t_unique = len(np.unique(timepoints))
print('timepoints: \n ', timepoints, '\n')
print('Distinct timepoints: \n', np.sort(np.unique(timepoints)), '\n')
print('Number of distinct timepoints:', n_t_unique)
```
Here is the raw data for a few randomly chosen genes.
```
f, axarr = plt.subplots(2, 3, figsize=(15,8))
gene_indx = np.sort(np.random.choice(n_train, 6))
for i in range(6):
n = gene_indx[i]
this_plot = axarr[int(np.floor(i / 3)), i % 3]
this_plot.plot(timepoints, y_train[n, :].T, '+', color = 'blue');
this_plot.set_ylabel('gene expression')
this_plot.set_xlabel('time')
this_plot.set_title('gene number {}'.format(n))
f.tight_layout()
```
### Define regressors
We model the time course using cubic B-splines. Let $\alpha$ be the degrees of freedom of the B-splines, and this is the parameter we seek to choose using cross-validation.
For a given degrees of freedom, the B-spline basis is given by an $n_{t}\times n_{x}$
matrix $X_{df}$, where the each column of $X_{df}$ is a B-spline
basis vector evaluated at the $n_{t}$ timepoints. Note that $n_{x}$
increases with increasing degrees of freedom.
Note that we only use B-splines to smooth the first 11 timepoints. For the last three timepoints, $t = 72, 120, 168$, we use indicator functions on each timepoint as three extra basis vectors. In other words, we append to the regressor matrix three columns, where each column is 1 if $t = 72, 120$, or $168$, respectively, and 0 otherwise. We do this to avoid numerical issues in the matrix $X^T X$. Because the later timepoints are more spread out, the B-spline basis are close to zero at the later timepoints, leading to matrices close to being singular.
```
# Simulate passing arguments in on the command line so that the notebook
# looks more like those in ``cluster_scripts``.
class Args():
def __init__(self):
pass
args = Args()
args.df = 7
args.degree = 3
args.num_components = 10
regressors = spline_bases_lib.get_genomics_spline_basis(
timepoints, df=args.df, degree=3)
regs = reg_lib.Regressions(y_train, regressors)
```
We plot the B-spline matrix for several degrees of freedom below:
```
f, axarr = plt.subplots(2, 3, figsize=(15,8))
i = 0
for df in [4, 5, 6, 7, 8, 9]:
_regressors = spline_bases_lib.get_genomics_spline_basis(
timepoints, exclude_num=3, df=df)
this_plot = axarr[int(np.floor(i / 3)), i % 3]
this_plot.plot(timepoints, _regressors);
this_plot.set_xlabel('time')
this_plot.set_ylabel('B-spline value')
this_plot.set_title('B-spliine basis when df = {}'.format(df))
i += 1
f.tight_layout()
```
We display the regressor matrix below.
```
plt.matshow(regs.x.T)
plt.ylabel('basis')
plt.xlabel('timepoint and replicate')
plt.title('The (transposed) regressor matrix when df = {}\n'.format(args.df));
```
With the regressor $X$ defined above, for each gene $g$
we model $P\left(y_{g}\vert\beta_{g},\sigma_{g}^{2}\right) = \mathcal{N}\left(y_{g}\vert X\beta_{g},\sigma_{g}^{2}\right)$. In the second stage, we will want to cluster $\beta_{g}$ taking into account its uncertainty on each gene. To do this, we wish to estimate the posterior mean $\mathbb{E}[\beta_g | y_g]$ and covariance $\mathrm{Cov}(\beta_{g} | y_g)$ with flat priors for both $\beta_g$ and $\sigma^2_g$.
For each gene, we estimate the posterior with a mean field variational Bayes (MFVB) approximation
$q\left(\sigma_{g}^{2},\beta_{g};\hat{\eta}_{g}\right)$ to the posterior
$P\left(\beta_{g},\sigma_{g}^{2}\vert y_{g}\right)$.
In particular, we take $q\left(\sigma_{g}^{2},\beta_{g};\hat{\eta}_{g}\right) = q^*\left(\sigma_{g}^{2}\right)q^*\left(\beta_{g}\right)$, where
$q^*\left(\sigma_{g}^{2}\right)$ is a dirac delta function, and we optimize over its a location parameter; $q^*\left(\beta_{g}\right)$ is a Gaussian density and we optimize over its mean and covariance.
The optimal variational approximation has a closed form that is formally identical to the standard frequentist
mean and covariance estimate for linear regression. Explicitly, the optimal variational distribution is,
\begin{align*}
q^*(\beta_g) &= \mathcal{N}\Big(\beta_g \; \Big| \; (X^TX)^{-1}X^T y_g, \; \hat\tau_g(X^TX)^{-1}\Big)\\
q^*(\sigma^2_g) &= \delta\{\sigma^2_g = \hat\tau_g\}
\end{align*}
where $\hat\tau_g = \frac{1}{n_t - n_x}\|y_g -X(X^TX)^{-1}X^T y_g\|_2^2$.
The advantage of the MVFB construction is that $\hat{\eta}_{g}$
for $g=1,...,n_{g}$ satisfies set of $n_{g}$ independent M-estimation objectives, allowing us to apply our infinitesimal jackknife results.
Specifically, defining $\theta_{reg}:=\left(\eta_{1},...,\eta_{n_{g}}\right)$,
we wish to minimize
\begin{align*}
F_{reg}\left(\theta_{reg},\alpha\right) & =\sum_{g=1}^{n_{g}}KL\left(q\left(\sigma_{g}^{2},\beta_{g};\eta_{g}\right)||P\left(\beta_{g},\sigma_{g}^{2}\vert y_{g}\right)\right)\\
& =-\sum_{g=1}^{n_{g}}\mathbb{E}_{q}\left[\log P\left(\beta_{g},\sigma_{g}^{2}\vert y_{g}\right)\right]+\mathbb{E}_{q}\left[\log q\left(\beta_{g},\sigma_{g}^{2}\vert\eta_{g}\right)\right]\\
& :=\sum_{g=1}^{n_{g}}F_{reg,g}\left(\eta_{g},\alpha\right).
\end{align*}
Our M-estimator, then, is
\begin{align*}
\frac{\partial F_{reg}\left(\theta_{reg},\alpha\right)}{\partial\theta_{reg}} & =0.
\end{align*}
The class $\texttt{regs}$ can calculate the optimal variational parameters for each gene. In particular, the variational parameters $\eta_g$ consist of a variational mean and covariance for $\beta_g$, as well as a location estimate for $\sigma^2_g$.
```
reg_time = time.time()
opt_reg_params = regs.get_optimal_regression_params()
reg_time = time.time() - reg_time
print('Regression time: {} seconds'.format(reg_time))
```
Here are what some of the fits look like. Each regression produces a prediction $\hat{y}_{g}:=X\mathbb{E}_{q}\left[\beta_g\right]$, plotted with the heavy red line above. The light red are predictions when $\beta_g$ is drawn from $q^*(\beta_g)$; the spread of the light red is intended to give a sense of the covariance of $\beta_g$.
```
f, axarr = plt.subplots(2, 3, figsize=(15,8))
for i in range(6):
n = gene_indx[i]
this_plot = axarr[int(np.floor(i / 3)), i % 3]
plot_utils_lib.PlotRegressionLine(
timepoints, regs, opt_reg_params, n, this_plot=this_plot)
f.tight_layout()
```
We also define and save data for the test regressions, which we will use later to evaluate out-of-sample performance. The training regressions will be saved below with the rest of the fit.
```
regs_test = reg_lib.Regressions(y_test, regressors)
test_regression_outfile = '../fits/test_regressions.json'
with open(test_regression_outfile, 'w') as outfile:
outfile.write(regs_test.to_json())
```
## The second stage: fit a mixture model.
### Transform the parameters before clustering
We are interested in the pattern of gene expression, not the absolute
level, so we wish to cluster $\hat{y}_{g}-\bar{\hat{y}}_{g}$, where
$\bar{\hat{y}}_{g}$ is the average over time points. Noting that
the $n_{t}\times n_{t}$ matrix $\text{Cov}_{q}\left(\hat{y}_{g}-\bar{\hat{y}}_{g}\right)$
is rank-deficient because we have subtracted the mean, the final step
is to rotate $\hat{y}_{g}-\bar{\hat{y}}_{g}$ into a basis where the
zero eigenvector is a principle axis and then drop that component.
Call these transformed regression coefficients $\gamma_{g}$ and observe that
$\text{Cov}_{q}\left(\gamma_{g}\right)$ has a closed form and is full-rank. It is these $\gamma_{g}$s that we will cluster in the second stage.
We briefly note that the re-centering operation could have been equivalently achieved by making a constant one of the regressors. We chose this implementation because it also allows the user to cluster more complex, non-linear transformations of the regression coefficients, though we leave this extension for future work.
We note that the transformations described in this section are done automatically in the ``GMM`` class. We are only calculating these transformations here for exposition.
```
# Get the matrix that does the transformation.
transform_mat, unrotate_transform_mat = \
trans_reg_lib.get_reversible_predict_and_demean_matrix(regs.x)
trans_obs_dim = transform_mat.shape[0]
```
If $T$ is the matrix that effects the transformation, then
\begin{align*}
\mathbb{E}_q[\gamma_g] &= T \mathbb{E}_q[\beta_g]\\
\text{Cov}_q(\gamma_q) &= T \text{Cov}_q(\beta_g) T^T
\end{align*}
The transformed parameters are also regression parameters, just in a different space.
```
# Apply the transformation
transformed_reg_params = \
trans_reg_lib.multiply_regression_by_matrix(
opt_reg_params, transform_mat)
```
We now visualize the transformed coefficients and their uncertainty.
```
f, axarr = plt.subplots(2, 3, figsize=(15,8))
transformed_beta = transformed_reg_params['beta_mean']
transformed_beta_info = transformed_reg_params['beta_info']
for i in range(6):
n = gene_indx[i]
this_plot = axarr[int(np.floor(i / 3)), i % 3]
this_plot.plot(transformed_beta[n, :], color = 'red');
this_plot.set_ylabel('transformed coefficient')
this_plot.set_xlabel('index')
this_plot.set_title('gene number {}'.format(n))
# draw from the variational distribution, to plot uncertainties
for j in range(30):
transformed_beta_draw = np.random.multivariate_normal(
transformed_beta[n, :], \
np.linalg.inv(transformed_beta_info[n]))
axarr[int(np.floor(i / 3)), i % 3].plot(transformed_beta_draw,
color = 'red', alpha = 0.08);
f.tight_layout()
```
The heavy red lines are the means of the transformed regression coefficients; shaded lines are draws from the variational distribution.
It is these transformed coefficients, $\gamma_g$, that we cluster in the second stage.
### Estimate an optimal clustering.
We now define a clustering problem for the $\gamma_{g}$. Let $n_{k}$
be the number of clusters, and $\mu_{1},...,\mu_{n_{k}}$ be the
cluster centers. Also let $z_{gk}$be the binary indicator for the
$g$th gene belonging to cluster $k$. We then define the following
generative model
\begin{align*}
\\
P\left(\pi\right) & =Dirichlet\left(\omega\right)\\
P\left(\mu_{k}\right) & =\mathcal{N}\left(\mu_{k}\vert0,\Sigma_{0}\right) \quad \text{for } \quad k = 1, ..., n_k\\
P\left(z_{gk}=1\vert\pi_{k}\right) & =\pi_{k} \quad \text{for } \quad k = 1, ..., n_k; \; n = 1, ..., n_g\\
P\left(\gamma_{g}\vert z_{gk}=1,\mu_{k},\eta_{g}\right) & =\mathcal{N}\left(\gamma_{g}\vert\mu_{k},\text{Cov}_{q}\left(\gamma_{g}\right)+\epsilon I_{n_{t}-1}\right)
\quad \text{for } \quad k = 1, ..., n_k; \; n = 1, ..., n_g.
\end{align*}
where $\epsilon$ is a small regularization parameter, which helped our optimization produce more stable results.
We will estimate the clustering using the maximum
a posteriori (MAP) estimator of $\theta_{clust}:=\left(\mu,\pi\right)$.
This defines an optimization objective that we seek to minimize:
\begin{align*}
F_{clust}\left(\theta_{clust},\theta_{reg}\right) & = - \sum_{g=1}^{n_{g}}E_{q^*_{z}}\Big\{\log P\left(\gamma_{g}\vert\eta_{g},\mu,\pi, z_g \right)-\log P(z_g | \pi)\Big\} - \log P\left(\mu\right)-\log P\left(\pi\right)
\end{align*}
which, for every value of $\theta_{reg}$, we expect to satisfy
\begin{align*}
\frac{\partial F_{clust}\left(\theta_{clust},\theta_{reg}\right)}{\partial\theta_{clust}} & =0.
\end{align*}
Note that $\theta_{clust}$ involves only the ''global'' parameters $\mu$ and $\pi$. We did take a variational distribution for the $z_{gk}$s, represented by independent Bernoulli distribution, but the optimal $q^*_z$ can be written as a function of $\mu$ and $\pi$. Hence, our optimization objective only involves these global parameters.
```
# Define prior parameters.
num_components = args.num_components
epsilon = 0.1
loc_prior_info_scalar = 1e-5
trans_obs_dim = regs.x.shape[1] - 1
prior_params = \
rm_lib.get_base_prior_params(trans_obs_dim, num_components)
prior_params['probs_alpha'][:] = 1
prior_params['centroid_prior_info'] = loc_prior_info_scalar * np.eye(trans_obs_dim)
gmm = rm_lib.GMM(args.num_components,
prior_params, regs, opt_reg_params,
inflate_coef_cov=None,
cov_regularization=epsilon)
```
In our experiment, the number of clusters $n_k$ was chosen to be {{args.num_components}}.
We set $\omega$ to be the ones vector of length $n_{k}$. The prior info for the cluster centers $\Sigma_{0}$ is {{loc_prior_info_scalar}}$\times I$. $\epsilon$ was set to be {{epsilon}}.
Let us examine the optimization objective. First, we'll inspect the likelihood terms. What follows is the likelihood given that gene $g$ belongs to cluster $k$.
```
print(inspect.getsource(rm_lib.get_log_lik_nk))
```
We can then optimize for $q^*_z$, which can be parametrized by its mean $\mathbb{E}_{q^*_z}[z]$. We note that this update has a closed form given $\theta_{clust}$, so there is no need to solve an optimization problem to find $q^*_z(z)$. We additionally note that we do not use the EM algorithm, which we found to have exhibit extremely poor convergence rates. Rather, we set $q^*_z(z)$ to its optimal value given $\theta_{clust}$ and return the objective as a function of $\theta_{clust}$ alone, allowing the use of more general and higher-quality optimization routines.
```
print(inspect.getsource(rm_lib.get_e_z))
```
With the optimal parameters for $z_{nk}$, we combine the likelihood term with the prior and entropy terms.
```
print(inspect.getsource(rm_lib.wrap_get_loglik_terms))
print(inspect.getsource(rm_lib.wrap_get_kl))
```
This objective function is wrapped in the ``GMM`` class method ``get_params_kl``.
```
print(inspect.getsource(gmm.get_params_kl))
```
### Optimization
For optimization we make extensive use of the [autograd](https://github.com/HIPS/autograd) library for automatic differentiation and the [paragami](https://github.com/rgiordan/paragami) library for parameter packing and sparse Hessians. These packages' details are beyond the scope of the current notebook.
First, we do a k-means initialization.
```
print('Running k-means init.')
init_gmm_params = \
rm_lib.kmeans_init(gmm.transformed_reg_params,
gmm.num_components, 50)
print('Done.')
init_x = gmm.gmm_params_pattern.flatten(init_gmm_params, free=True)
```
We note that
the match between "exact" cross-validation (removing time points and
re-optimizing) and the IJ was considerably improved by
using a high-quality second-order optimization method. In particular, for these
experiments, we employed the Newton conjugate-gradient trust region method
(Chapter 7.1 of Wright et al [1999]) as implemented by the method
``trust-ncg`` in ``scipy.optimize``, preconditioned by the Cholesky
decomposition of an inverse Hessian calculated at an initial approximate
optimum.
We found that first-order or quasi-Newton
methods (such as BFGS) often got stuck or terminated at points with fairly large
gradients. At such points our method does not apply in theory nor, we found,
very well in practice.
The inverse Hessian used for the preconditioner was with respect to
the clustering parameters only and so could be calculated quickly, in contrast
to the $H_1$ matrix used for the IJ, which includes the
regression parameters as well.
First, run with a low tolerance to get a point at which to evaluate an initial preconditioner.
```
gmm.conditioned_obj.reset() # Reset the logging and iteration count.
gmm.conditioned_obj.set_print_every(1)
opt_time = time.time()
gmm_opt, init_x2 = gmm.optimize(init_x, gtol=1e-2)
opt_time = time.time() - opt_time
```
Next, set the preconditioner using the square root inverse Hessian at the point ``init_x2``.
```
tic = time.time()
h_cond = gmm.update_preconditioner(init_x2)
opt_time += time.time() - tic
```
The method ``optimize_fully`` repeats this process of optimizing and re-calculating the preconditioner until the optimal point does not change.
```
gmm.conditioned_obj.reset()
tic = time.time()
gmm_opt, gmm_opt_x = gmm.optimize_fully(
init_x2, verbose=True)
opt_time += time.time() - tic
print('Optimization time: {} seconds'.format(opt_time))
```
``paragami`` patterns allow conversion between unconstrained vectors and dictionaries of parameter values. After "folding" the optimal ``gmm_opt_x``, ``opt_gmm_params`` contains a dictionary of optimal cluster centroids and cluster probabilities.
```
opt_gmm_params = gmm.gmm_params_pattern.fold(gmm_opt_x, free=True)
print(opt_gmm_params.keys())
print(np.sort(opt_gmm_params['probs']))
```
Each gene's regression line has an inferred cluster membership given by $\mathbb{E}_{q^*_z}[z_g]$, and an expected posterior centroid given by $\sum_k \mathbb{E}_{q^*_z}[z_{gk}] \mu_k$. This expected posterior centroid can be un-transformed to give a prediction for the observation.
It is the difference between this prediction line --- which is a function of the clustering --- and the actual data that we consider to be the "error" of the model.
```
gmm_pred = mse_utils.get_predictions(gmm, opt_gmm_params, opt_reg_params)
f, axarr = plt.subplots(2, 3, figsize=(15,8))
for i in range(6):
n = gene_indx[i]
this_plot = axarr[int(np.floor(i / 3)), i % 3]
plot_utils_lib.PlotRegressionLine(
timepoints, regs, opt_reg_params, n, this_plot=this_plot)
plot_utils_lib.PlotPredictionLine(
timepoints, regs, gmm_pred, n, this_plot=this_plot)
f.tight_layout()
```
### Calculating $H_1$ for the IJ
We seek to choose the degrees of freedom $\alpha$ for the B-splines using cross-validation. We leave out one or more timepoints, and fit using only the remaining timepoints. We then estimate the test error by predicting the value of the genes at the held out timepoints.
To do this, we define time weights $w_{t}$ by observing that, for each $g$,
the term $\mathbb{E}_{q}\left[\log P\left(\beta_{g},\sigma_{g}^{2}\vert y_{g}\right)\right]$
decomposes into a sum over time points:
\begin{align*}
F_{reg,g}\left(\eta_{g},\alpha,w\right) & :=-\sum_{t=1}^{n_{t}}w_{t}\left(-\frac{1}{2}\sigma_{g}^{-2}\left(y_{g,t}-\left(X\beta_{g}\right)_{t}\right)^{2}-\frac{1}{2}\log\sigma_{g}^{2}\right)+\mathbb{E}_{q}\left[\log q\left(\beta_{g},\sigma_{g}^{2}\vert\eta_{g}\right)\right].
\end{align*}
We naturally define $F_{reg}\left(\theta_{reg},\alpha,w\right):=\sum_{g=1}^{n_{g}}F_{reg,g}\left(\eta_{g},\alpha,w\right).$
By defining $\theta=\left(\theta_{clust},\theta_{reg}\right)$, we
then have an M-estimator
\begin{align*}
G\left(\theta,w,\alpha\right) & :=\left(\begin{array}{c}
\frac{\partial F_{reg}\left(\theta_{reg},w,\alpha\right)}{\partial\theta_{reg}}\\
\frac{\partial F_{clust}\left(\theta_{clust},\theta_{reg}\right)}{\partial\theta_{clust}}
\end{array}\right)=0.
\end{align*}
We can then apply the IJ to approximate the leaving out of various timepoints.
Note that what we call the "Hessian" for this two-step procedure is not really a Hessian, as it is not symmetric. More precisely, it is the Jacobian of $G$, or what we defined as $H_1$ in the text.
Calculating $H_1$ is the most time-consuming part of the infinitesimal jackknife, since the $H_1$ matrix is quite large (though sparse). However, once $H_1$ is computed, calculating each $\theta_{IJ}(w)$ is extremely fast.
$H_1$ can be computed in blocks:
\begin{align*}
H_1 = \begin{pmatrix}
\nabla^2_{\theta_{reg}} F_{reg} & 0 \\ \nabla_{\theta_{reg}} \nabla_{\theta_{clust}} F_{clust} & \nabla^2_{\theta_{clust}} F_{clust} \end{pmatrix}
\end{align*}
The code refers to $\nabla^2_{\theta_{clust}} F_{clust}$ as the "GMM Hessian". It refers to $\nabla_{\theta_{reg}} \nabla_{\theta_{clust}} F_{clust}$ as the "cross Hessian". And it refers to $\nabla^2_{\theta_{reg}} F_{reg}$ as the "regression Hessian", which itself is block diagonal, with each block an observation. Due to details of the implementation of block sparse Hessians using forward mode automatic differnetiation in the class ``vittles.SparseBlockHessian``, the code below confusingly refers to each regression parameter as a "block".
When the ``FitDerivatives`` class is initialized, it calculates these blocks separately and stacks them into the attribute ``full_hess``, which is a sparse matrix representing $H_1$.
```
# Even though $H_1$ is not a Hessian, by force of habit we call the time to
# compute it ``hess_time``.
hess_time = time.time()
fit_derivs = sens_lib.FitDerivatives(
opt_gmm_params, opt_reg_params,
gmm.gmm_params_pattern, regs.reg_params_pattern,
gmm=gmm, regs=regs,
print_every=10)
hess_time = time.time() - hess_time
print('Total hessian time: {} seconds'.format(hess_time))
```
### Save results as a compressed file.
The results, including $H_1$, are now saved. To calculate the exact CV, these results (including the preconditioner) will be loaded and the model will be refit with timepoints left out. To calculate the IJ, the same results will be loaded and $H_1$ will be used to calculate the IJ.
```
extra_metadata = dict()
extra_metadata['opt_time'] = opt_time
extra_metadata['reg_time'] = reg_time
extra_metadata['hess_time'] = hess_time
extra_metadata['df'] = args.df
extra_metadata['degree'] = args.degree
npz_outfile = '../fits/initial_fit.npz'
saving_gmm_utils.save_initial_optimum(
npz_outfile,
gmm=gmm,
regs=regs,
timepoints=timepoints,
fit_derivs=fit_derivs,
extra_metadata=extra_metadata)
```
### Bibliography
J. E. Shoemaker, S. Fukuyama, A. J. Eisfeld, D. Zhao, E. Kawakami, S. Sakabe, T. Maemura,
T. Gorai, H. Katsura, Y. Muramoto, S. Watanabe, T. Watanabe, K. Fuji, Y. Matsuoka, H. Kitano,
and Y. Kawaoka. An Ultrasensitive Mechanism Regulates Influenza Virus-Induced Inflammation.
PLoS Pathogens, 11(6):1–25, 2015
S. Wright and J. Nocedal. Numerical optimization. Springer Science, 35(67-68):7, 1999.
| github_jupyter |
# PyTorchDIA Tutorial
This tutorial demonstrates how to use the code and provides some tips for improving performance.
For a comprehensive outline of these features, check out the paper https://arxiv.org/abs/2104.13715
```
# imports
import PyTorchDIA
from MakeFakeImage import MakeFake
import torch
import numpy as np
from scipy.stats import norm
from astropy.io import fits
import matplotlib.pyplot as plt
%matplotlib inline
# for reproducibility, use deterministic CUDA convolution computations only
torch.backends.cudnn.deterministic = True
```
## 1) Basics
First, let's generate some artifical CCD data to work with. The data -- with per-pixel indices $ij$ -- are independently drawn from $\mathcal{N}(M, \sigma^2)$, where $M_{ij}$ is our image model. For the pixel variances, we adopt the 'standard' CCD noise model,
$\sigma^2 = \frac{\sigma_{0}^{2}}{F_{ij}^{2}} + \frac{M_{ij}}{G\;F_{ij}}$,
where $\sigma_0$ is the read noise (ADU), $G$ is the detector gain ($\textrm{e}^- / \textrm{ADU}$), and $F$ is the master flat field.
```
## Generate an artifical reference and target image pair
# set random seed
np.random.seed(42)
# generate noiseless reference 'ref'
print('Generating reference...')
size = 142
log_density = np.random.uniform(0, 3, 1)[0]
star_density = 10**log_density # stars per 100x100 pixels
n_sources = np.int(star_density * (size**2/100**2))
phi_r = np.random.uniform(0.5, 2.5, 1)[0] # in pixels, this is ~ [1 - 6] fwhm
sky = np.random.uniform(10, 1000, 1)[0] # ADU
# positions
positions_x = np.random.uniform(0, size, (n_sources,1))
positions_y = np.random.uniform(0, size, (n_sources,1))
positions = np.hstack((positions_x, positions_y))
# fluxes
F = np.random.uniform(10**(-9), 10**(-4.5), n_sources)
fluxes = F**(-2./3.)
# Generate the noiseless reference image
ref_noiseless, F_frac = MakeFake(N=1, size=size, n_sources=n_sources,
psf_sigma=phi_r, sky=sky,
positions=positions, fluxes=fluxes, shifts=[0, 0])
print('Reference properties')
print('Reference size:', size)
print('Number of sources:', n_sources)
print('PSF standard deviation:', phi_r)
print('Sky level:', sky)
print('F_max/F_total:', F_frac) # flux ratio of brightest star that of all stars
print('Target kernel properties:')
phi_k = np.random.uniform(0.5, 2.5, 1)[0]
kernel_size = 19
print('Kernel standard deviation:', phi_k)
print('Kernel size:', kernel_size)
# Generate the noiseless (and shifted) target image
phi_i = np.sqrt(phi_k**2 + phi_r**2)
# positions
shift_x = np.random.uniform(-0.5, 0.5, 1)
shift_y = np.random.uniform(-0.5, 0.5, 1)
#shift_x, shift_y = 0, 0
print('shift_x, shift_y:', shift_x, shift_y)
positions_x_shifted = positions_x + shift_x
positions_y_shifted = positions_y + shift_y
positions_shifted = np.hstack((positions_x_shifted, positions_y_shifted))
imag_noiseless, F_frac = MakeFake(N=1, size=size, n_sources=n_sources,
psf_sigma=phi_i, sky=sky,
positions=positions_shifted, fluxes=fluxes,
shifts = [shift_x[0], shift_y[0]])
imag_noiseless_copy = np.copy(imag_noiseless)
ref_noiseless_copy = np.copy(ref_noiseless)
# read noise [ADU]
sigma_0 = 5.
# add gaussian noise to image under the standard CCD noise model
# N.B. Gain and flat-field are equal to 1, so we only include
# the readout noise [ADU] and the photon shot noise (in the gaussian limit)
def add_noise_to_image(image, read_noise):
noise_map = np.random.normal(0, 1, size=image.shape)
sigma_imag = np.sqrt(read_noise**2 + image)
image += noise_map*sigma_imag
return image, sigma_imag
# adds 10 times **less** variance than add_noise_to_image
def add_less_noise_to_image(image, read_noise):
noise_map = np.random.normal(0, 1, size=image.shape)
sigma_imag = 10**(-0.5) * np.sqrt(read_noise**2 + image)
image += noise_map*sigma_imag
return image, sigma_imag
# add noise to the reference image
ref, sigma_ref = add_less_noise_to_image(image=ref_noiseless, read_noise=sigma_0)
ref_copy = np.copy(ref)
# add noise to the target image i.e.
imag, sigma_imag = add_noise_to_image(image=imag_noiseless, read_noise=sigma_0)
print('Reference and target image shapes:')
print(ref.shape, imag.shape)
# plot
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(10, 5))
ax[0].imshow(ref)
ax[1].imshow(imag);
```
**Tip**: It is almost always a good idea to subtract some estimate of the sky from the reference image. This helps to break the strong anticorrelation between the parameters* in the DIA image model.
*Specifically, the photometric scale factor (sum of kernel pixels) and the differential background.
```
### sky subtract the reference image
ref -= sky
```
The user must provide the *scalar objective* function to optimise. For this CCD data, with Gaussian noise, we should use the following (negative) log-likelihood
$-\textrm{ln}\;\textrm{p}(I_{ij} | \boldsymbol{\theta}) = \frac{1}{2}\chi^2\ + \sum_{ij} \ln\;\sigma_{ij} + \frac{N_\mathrm{data}}{2} \textrm{ln}(2\mathrm{\pi}) \;,$
where $\sigma_{ij}$ are the pixel uncertainties, $N_{\mathrm{data}}$ is the number of pixels in the target image, and the $\chi^2$ is equal to
$\chi^2 = \sum_{ij}\left(\frac{I_{ij} - M_{ij}}{\sigma_{ij}} \right)^2 \;.$
$\boldsymbol{\theta}$ are the model parameters; the kernel pixels and differential background term(s). Let's write this in code (ignoring the irrelevant normalisation constant).
The only requirement for loss functions fed to PyTorchDIA is that the **first argument is the model**, and the **second agrument is the data**. If a flat field is available, this can be passed as an optional third agrument (seen example further down). In this artifically simple case, the flat field is assummed to be perfect, so we can just ignore it. We do have to pass some keyword agruments for the readout noise and detector gain though.
```
# gaussian negative log-likelihood
def gaussian_loss_fn(model, data, rdnoise=5, G=1):
# guard against negative pixel-variances should they arise during the optimisation
var = (torch.clamp(model, min=0.) / G) + rdnoise**2
chi2 = torch.sum((model - data) ** 2 / var)
ln_sigma = torch.sum(torch.log(var))
nll = 0.5 * (chi2 + ln_sigma)
return nll
```
OK, we're now ready to fit a model image to our data image. The simplest DIA model -- with just a scalar differential background term -- has the form
$M_{ij} = [R \otimes K]_{ij} + B_0$,
and can be fit as simply as follows...
```
kernel_size = 19 # specify kernel object single axis size (must be odd)
res = PyTorchDIA.DIA(ref, # reference image (np.ndarray)
imag, # data image (np.ndarray)
gaussian_loss_fn, # loss function
ks = kernel_size, # (square) kernel size: ks x ks
show_convergence_plots = True) # plot loss vs iterations
```
The code outputs some diagnostic information about the optimisation e.g. number of steepest descent (SD) steps, L-BFGS steps and a plot of (log) loss vs iterations. And returns the kernel, the differential background term(s) and the model. Let's plot the image (and histogram) of normalised residuals to check all is well.
```
# these are returned as numpy.ndarrays (if multi-dimensional... B0 is a scalar)
kernel, B0, model = res
```
**Tip**: Convolutions are undefined within half a kernel's width of the data image's border, so we need to trim this appropriately to match the size of the model.
```
# trim I such that target image pixels correspond to only those with valid convolution computations
hwidth = np.int((kernel_size - 1) / 2)
nx, ny = imag.shape
trimmed_imag = imag[hwidth:nx-hwidth, hwidth:nx-hwidth]
residuals = trimmed_imag - model
sigma = np.sqrt(model + sigma_0**2)
normalised_residuals = residuals / sigma
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(12,5))
x = np.linspace(-5, 5, 100)
ax[0].imshow(normalised_residuals)
ax[1].hist(normalised_residuals.flatten(), bins=50, density=True)
ax[1].plot(x, norm.pdf(x, 0, 1))
ax[1].set_xlabel('Normalised residuals', fontsize=20)
ax[1].set_ylabel('Probability', fontsize=20);
```
**N.B.** The kernel returned by the code is orientated such that it can be passed to scipy.signal.convolve2d. Beware: torch.nn.functional.conv2d expects a *different* orientation.
## 2) Modelling a spatially varying background
Let's repeat the above, but add a spatially varying sky background to our data image.
```
np.random.seed(42)
nx, ny = imag_noiseless_copy.shape
x = np.linspace(-0.5, 0.5, nx)
y = np.linspace(-0.5, 0.5, ny)
X, Y = np.meshgrid(x, y, copy=False)
## polynomial coefficients - drop back in the sky level as the 0th degree component
coeffs = [sky, 50, 20]
def spatially_varying_sky(X, Y, c):
return c[0] + X*c[1] + Y*c[2]
# subtract off the scalar sky we added above
imag_noiseless = imag_noiseless_copy + spatially_varying_sky(X, Y, coeffs) - sky
# add noise to the target image i.e.
imag_var_bkg, sigma_imag = add_noise_to_image(image=imag_noiseless, read_noise=sigma_0)
# plot background
plt.imshow(spatially_varying_sky(X, Y, coeffs), origin='lower')
plt.colorbar();
## remember to subtract some estimate of the sky level of the reference
ref = ref_copy - sky
kernel_size = 19
res = PyTorchDIA.DIA(ref, # reference image (np.ndarray)
imag_var_bkg, # data image (np.ndarray)
gaussian_loss_fn, # loss function
ks = kernel_size, # (square) kernel size: ks x ks
poly_degree = 1, # polynomial degree for spatially varying background
show_convergence_plots = True) # plot loss vs iterations
kernel, B, model = res
# trim I such that target image pixels correspond to only those with valid convolution computations
hwidth = np.int((kernel_size - 1) / 2)
nx, ny = imag_var_bkg.shape
trimmed_imag = imag_var_bkg[hwidth:nx-hwidth, hwidth:nx-hwidth]
residuals = trimmed_imag - model
sigma = np.sqrt(model + sigma_0**2)
normalised_residuals = residuals / sigma
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(12,5))
x = np.linspace(-5, 5, 100)
ax[0].imshow(normalised_residuals)
ax[1].hist(normalised_residuals.flatten(), bins=50, density=True)
ax[1].plot(x, norm.pdf(x, 0, 1))
ax[1].set_xlabel('Normalised residuals', fontsize=20)
ax[1].set_ylabel('Probability', fontsize=20);
```
## 3) Going beyond Gaussian noise models
Real images have outliers; those pixels not captured by our simple Gaussian noise model. Indeed, these are usually associated with the things we're interested in performing measurements on further downstream. To ensure the fit is not badly influenced by these objects (or any other outlier e.g. cosmics, 'bad' pixels), we can use *robust* loss functions. In the PyTorchDIA paper, we derive a useful robust alternative for data which is otherwise Gaussian distributed; we'll leave the mathetmatics from here, so check out Section 2.4 of the paper (https://arxiv.org/abs/2104.13715) if you're interested.
Here, we'll implement this robust loss on some real EMCCD data of a microlensed target.
```
## load the reference, data and master flat images
ref_emccd = fits.getdata('coll_LOB190560Z_Llr_2019-05-14_00129.fits')
data_emccd = fits.getdata('coll_LOB190560Z_Llr_2019-05-10_00107.fits')
flat_emccd = fits.getdata('master_flat.fits')
# as c -> infinity, this loss function is identical to the Gaussian
# loss we used previously. c = 1.345 is generally recommended.
def robust_loss_fn(model, data, flat, f=25.8, G_EM=300, sigma_EM=60, N=1200, c=1.345):
# Total gain, G, readout noise, and EMCCD excess noise factor, E
G = f / G_EM
sigma0 = N * (sigma_EM / f)
E = 2
# EMCCD noise model
var = E * (torch.clamp(model, min=0.)/(G*flat) + (sigma0 / flat)**2)
sigma = torch.sqrt(var)
ln_sigma = torch.sum(torch.log(sigma))
# gaussian when (model - targ)/NM <= c
# absolute deviation when (model - targ)/NM > c
cond1 = torch.abs((model - data)/sigma) <= c
cond2 = torch.abs((model - data)/sigma) > c
inliers = ((model - data)/sigma)[cond1]
outliers = ((model - data)/sigma)[cond2]
l2 = 0.5*torch.sum(torch.pow(inliers, 2))
l1 = (c * torch.sum(torch.abs(outliers)) - (0.5 * c**2))
nll = l2 + l1 + ln_sigma
return nll
## again, remember to subtract some estimate of the sky from the background
ref_emccd -= np.median(ref_emccd)
kernel_size = 35
res = PyTorchDIA.DIA(ref_emccd, # reference image (np.ndarray)
data_emccd, # data image (np.ndarray)
robust_loss_fn, # loss function
flat_emccd, # flat (np.ndarray)
ks = kernel_size, # (square) kernel size: ks x ks
show_convergence_plots = True) # plot loss vs iterations
kernel, B, model = res
# trim I such that target image pixels correspond to only those with valid convolution computations
hwidth = np.int((kernel_size - 1) / 2)
nx, ny = data_emccd.shape
trimmed_imag = data_emccd[hwidth:nx-hwidth, hwidth:nx-hwidth]
trimmed_flat = flat_emccd[hwidth:nx-hwidth, hwidth:nx-hwidth]
residuals = trimmed_imag - model
# Total gain, G, and EMCCD excess noise factor, E
G = 25.8 / 300
E = 2
shot_noise = model/(G*trimmed_flat)
sigma = np.sqrt(E*shot_noise)
# normalised residuals
normalised_residuals = residuals / sigma
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(12,5))
x = np.linspace(-5, 5, 100)
ax[0].imshow(normalised_residuals)
ax[1].hist(normalised_residuals.flatten(), bins=200, density=True)
ax[1].plot(x, norm.pdf(x, 0, 1))
ax[1].set_xlabel('Normalised residuals', fontsize=20)
ax[1].set_ylabel('Probability', fontsize=20)
ax[1].set_xlim(-5, 5);
```
**Tip:** The normalised residuals could suggest that there are unmodelled noise contributions. These probably arise from the reference image. It is almost always advantageous to try stacking several 'best' images together in practice and average down the reference image noise if you see this sort of thing. Indeed, for the artifical images generated in this notebook, I add 10 times less noise variance to the reference images than the data image to deliberately guard against this.
## 4) Penalised MLE optimisation: Placing a Laplacian prior on the kernel pixels
**Tip:** If the data is at low signal-to-noise, MLE kernel estimates are known to badly overfit by absorbing noise into the kernel solution. If this is an issue, it can be useful to introduce penalty terms (or priors) on the kernel pixels which favour kernels with smooth surfaces. One useful prior to enforce this property is a Laplacian prior on the kernel, which represents the *connectivity graph* between the pixels, and so favours kernels where neighbouring pixels have similar values. Because construction of the Laplacian is somewhat involved, this has been built into PyTorchDIA.
In order to add this penalty term to any user-specified loss function, you just need to pass a non-zero (positive) value to the 'alpha' keyword agrument, where alpha is the hyperparameter specifying the strength of the penalisation. Good choices are usually somewhere in the range 0.1 - 1. In fact, the data image from Example 1 is quite low signal-to-noise, so we may benefit there. Let's compare the MLE and penalised MLE solutions from that example below.
```
kernel_size = 19
res_MLE = PyTorchDIA.DIA(ref, # reference image (np.ndarray)
imag, # data image (np.ndarray)
gaussian_loss_fn, # loss function
ks = kernel_size, # (square) kernel size: ks x ks
show_convergence_plots = True) # plot loss vs iterations
res_PMLE = PyTorchDIA.DIA(ref, # reference image (np.ndarray)
imag, # data image (np.ndarray)
gaussian_loss_fn, # loss function
ks = kernel_size, # (square) kernel size: ks x ks
alpha = 0.1, # the hyperparameter controlling the strenght of the Laplacian penalty
show_convergence_plots = True) # plot loss vs iterations
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(10, 5))
kernel_MLE, kernel_MAP = res_MLE[0], res_PMLE[0]
ax[0].imshow(kernel_MLE)
ax[0].set_title('MLE kernel')
ax[1].imshow(kernel_MAP)
ax[1].set_title('Penalised MLE kernel');
```
| github_jupyter |
```
import numpy as np
import gym
import tensorflow as tf
env = gym.make('CartPole-v1')
obs = env.reset()
# initilize W and b
n_input = 4
n_hidden = 16
n_output = env.action_space.n
W1 = np.random.randn(n_input, n_hidden)
b1 = np.ones([n_hidden])
W2 = np.random.randn(n_hidden,n_output)
b2 = np.zeros([n_output])
def training_step(obs):
# Forward pass
# two layered network with relu activation
N, D = obs.shape
h = obs.dot(W1) + b1
h[h<0] = 0
out_linear = h.dot(W2) + b2
exp_scores = np.exp(out_linear)
probs = exp_scores / np.sum(exp_scores, axis=1, keepdims=True)
# We dont always want to take the best posible action
# To explore better options we will choose the action
# randomly with predicted probablities
action = np.random.choice([x for x in range(n_output)], 1, p=probs[-1])
# backward pass
grads={}
dscores = probs.copy()
dscores[range(N), list(action)] -= 1
dscores /= N
grads['W2'] = h.T.dot(dscores)
grads['b2'] = np.sum(dscores, axis = 0)
dh = dscores.dot( W2.T)
dh_ReLu = (h > 0) * dh
grads['W1'] = obs.T.dot(dh_ReLu)
grads['b1'] = np.sum(dh_ReLu, axis = 0)
return grads, action
def get_action(obs):
# get action when rendering (testing)
# Forward pass
obs = obs.reshape([1,n_input])
h = obs.dot(W1) + b1
h[h<0] = 0
out_linear = h.dot(W2) + b2
exp_scores = np.exp(out_linear)
probs = exp_scores / np.sum(exp_scores, axis=1, keepdims=True)
return np.random.choice([x for x in range(n_output)], 1, p=probs[-1])[0]
def discount(r, gamma = 0.7):
# We dont know which actions cause the rewards
# so we need to make couple previous actions also responsible
discounted = np.zeros_like(r)
running_add = 0
for t in reversed(range(0, r.size)):
running_add = running_add*gamma + r[t]
discounted[t] = running_add
return discounted
n_iter = 100 # we have early stop
callback = 195 # consider the game is solved
n_game_per_iter = 100
n_action_per_game = 200
learning_rate = 1e-1
for itern in range(n_iter):
update_grads = {}
update_grads['W1'] = np.zeros_like(W1)
update_grads['b1'] = np.zeros_like(b1)
update_grads['W2'] = np.zeros_like(W2)
update_grads['b2'] = np.zeros_like(b2)
mean_reward = 0
all_gradients = []
for game in range(n_game_per_iter):
obs = env.reset()
current_rewards = []
current_gradients = []
total_reward = 0
for step in range(n_action_per_game):
obs = obs.reshape([1,n_input])
grads, action = training_step(obs)
obs, reward, done, info = env.step(action[0])
current_rewards.append(reward)
current_gradients.append(grads)
total_reward+=reward
if done:
break
mean_reward+=total_reward
current_rewards = np.array(current_rewards)
current_gradients = np.array(current_gradients)
# normalize the rewards
discounted_rewards = discount(current_rewards)
discounted_rewards -= np.mean(discounted_rewards)
discounted_rewards /= np.std(discounted_rewards)
# mulitply them with gradients
for i in range(current_gradients.shape[0]):
for _, n in enumerate(current_gradients[i]):
current_gradients[i][n] = current_gradients[i][n]*discounted_rewards[i]
all_gradients.append(current_gradients[i])
# take the mean gradient and make the update
for i in range(len(all_gradients)):
for _, n in enumerate(all_gradients[i]):
update_grads[n]+=all_gradients[i][n]
for _, n in enumerate(update_grads):
update_grads[n]/= len(all_gradients)
W1 -= learning_rate*update_grads['W1']
b1 -= learning_rate*update_grads['b1']
W2 -= learning_rate*update_grads['W2']
b2 -= learning_rate*update_grads['b2']
print('\riteration %d / %d: Mean Score %f'% (itern, n_iter, mean_reward/n_game_per_iter), end = "")
# if the mean score of 100 games is higher than 195
# consider solved
if mean_reward/n_game_per_iter > callback:
print("Training done!")
break
# Render the env
n_test = 10
for i in range(n_test):
obs = env.reset()
total_reward = 0
while True:
env.render()
action = get_action(obs)
obs, reward, done, info = env.step(action)
total_reward+=reward
if done :
break
print("Game %d, Total Reward %f"%(i+1, total_reward))
```
| github_jupyter |
## Getting the ida pro comments out of the files
## Authors:
Aaron Gonzales, Andres Ruiz
```
import os, sys, re
import asm_utils
import pymongo as pm
import multiprocessing
from joblib import delayed, Parallel
import src.utils.utils as utils
samples = asm_utils.get_collection()
```
Aaron and Andres,
The four common call convention keywords that are used in the IDA .asm
files are:
__stdcall
__cdecl
__fastcall
__thiscall
the general format of the function comments looks like:
; <TYPE> <CONVENTION> <NAME>(<ARG0>, <ARG1>, ...)
as an example:
; void __stdcall memcpy(char *dst, char *src, int n)
Another possible thing that might be helpful is the DLL files that are
imported. These are listed on lines that look like the following:
;
; Imports from <NAME>.dll
;
as an example:
;
; Imports from KERNEL32.dll
;
these would give a list of dll's that are used by the samples. It might
be as simple as using any line that contains ".dll" or ".DLL".
```
calls = {'__stdcall', '__cdecl', '__fastcall', '__thiscall'}
test_comments = samples.find_one({'class' : '5'})['ida_comments']
dlls = [line for line in test_comments if '.dll' in line.lower()]
dlls = [line.split()[-1] for line in dlls]
dlls
def get_dlls(comments):
""" Returns a lowercase set of DLLs accessed by the file does not preserve order.
Remove set command to preserve redundency"""
dlls = [line for line in comments if '.dll' in line.lower()]
return list(set(line.split()[-1].lower() for line in dlls))
get_dlls(test_comments)
fcalls = [line for line in test_comments for word in line.split() if word in calls]
def get_calls(comments):
call_dict = {}
calls = {'__stdcall', '__cdecl', '__fastcall', '__thiscall'}
fcalls = [line for line in comments for word in line.split() if word in calls]
call_dict['calls'] = fcalls
call_dict['total_calls'] = len(fcalls)
call_dict['stdcall'] = [line for line in fcalls if '__stdcall' in line.split()]
call_dict['cdecl'] = [line for line in fcalls if '__cdecl' in line.split()]
call_dict['fastcall'] = [line for line in fcalls if '__fastcall' in line.split()]
call_dict['thiscall'] = [line for line in fcalls if '__thiscall' in line.split()]
call_dict['stdcall_count'] = len(call_dict['stdcall'])
call_dict['cdecl_count'] = len(call_dict['cdecl'])
call_dict['fastcall_count'] = len(call_dict['fastcall'])
call_dict['thiscall_count'] = len(call_dict['thiscall'])
return call_dict
get_calls(test_comments)
def expert_comment_maker(document):
document['dlls'] = get_dlls(document['ida_comments'])
document['calls'] = get_calls(document['ida_comments'])
test_doc = samples.find_one({'class' : '5'})
test_doc.keys()
expert_comment_maker(test_doc)
test_doc.keys()
expert_needed = samples.find({'dlls':[]})
expert_needed.count()
def _expert_comment_maker(doc):
print('extracting expert comments from %s' % doc['id'])
expert_comment_maker(doc)
samples.save(doc)
samples = asm_utils.get_collection()
expert_needed = samples.find({'dlls':[]})
num_cores = multiprocessing.cpu_count()
print('Running code on %d processors' % num_cores)
Parallel(n_jobs=num_cores)(
delayed(_expert_comment_maker)(doc) for doc in expert_needed)
col = asm_utils.get_collection(collection='test_samples')
nodll = col.find({'dlls': []})
nodll.count()
nodll
def main():
samples = asm_utils.get_collection(collection='test_samples')
expert_needed = samples.find()
print(expert_needed.count())
print(samples)
num_cores = multiprocessing.cpu_count()
print('Running code on %d processors' % num_cores)
Parallel(n_jobs=num_cores)(
delayed(_expert_comment_maker)(doc) for doc in expert_needed)
```
| github_jupyter |
# m3 document collection workspace
This is a very simple notebook to:
1. write versions of the calibration products from the PDS3 archive
2. move other document collection items to their correct locations
and check matches between lids and filenames
```
import datetime as dt
import os
from collections import namedtuple
import pandas as pd
import sh
from m3_bulk import crude_time_log
from converter import place_label, enforce_name_match
from converter_utils import name_root, eqloc
#don't bother doing this multiple times if you're running this concurrently
# sh.s3fs(
# 'mc-al-khwarizmi',
# './remote/m3_input'
# )
# sh.s3fs(
# 'mc-al-khwarizmi-m3-output',
# './remote/m3_output'
# )
# mappings for all document collection files
from src.m3_conversion import M3FlatFieldWriter, M3PipelineLogWriter, \
M3BDEWriter, M3SSCWriter
doc_path_df = pd.read_csv('./directories/m3/m3_document_mappings.csv')
# all our 'oddball' one-off label files
label_files = []
for root, dirs, files in os.walk('./labels/m3/'):
for file in files:
label_files.append(os.path.join(root, file))
label_df = pd.DataFrame([
{'local_path':file,'filename':name_root(file)}
for file in label_files
])
# some of them have locally-converted or edited versions (PDF compliance, etc.)
# note: EARTH_VIEW_IMAGE isn't included in the github repo because it's large,
# but it's just a simple conversion using rasterio and converter_utils.fitsify()
oddball_files = []
for root, dirs, files in os.walk('./converted_oddballs/m3/'):
for file in files:
oddball_files.append(os.path.join(root, file))
oddball_df = pd.DataFrame([
{
'local_path':file,
'filename':name_root(file)
}
for file in oddball_files
])
# where are our bundles mounted?
input_dir = '/home/ubuntu/m3_input'
output_dir = '/home/ubuntu/m3_output'
# sanity check: do we have an individual label for each thing we think needs one?
missing_labels = [
file for file in eqloc(doc_path_df, 'label_type', 'label')['root'].values
if file not in list(map(name_root, label_files))
]
assert len(missing_labels) == 0
for product in doc_path_df.itertuples():
if product.root == 'earth_view_image':
continue
doc_start_time = dt.datetime.now()
print(product.root, product.path, product.Index)
if product.use_local != 'False':
# enforce match between putative filename root and local file
try:
product_file = eqloc(
oddball_df, "filename", product.root
)["local_path"].values[0]
except IndexError:
raise ValueError("missing local version for " + product.root)
else:
product_file = input_dir + product.path
sh.mkdir('-p', output_dir + product.newpath)
if product.label_type == 'label':
writer = place_label(product, label_df, product_file, output_dir)
# validate every oddball; these labels were all manually
# generated
validate_results = sh.validate("-t", writer['label'])
with open("validate_dump.txt", "a") as file:
file.write(validate_results.stdout.decode())
print("validated successfully")
elif product.label_type == 'template':
if 'reduction_pipeline' in product.newpath:
writer = M3PipelineLogWriter(product_file)
writer.write_pds4(
output_dir + product.newpath +"/",
write_product_files=True
)
elif 'flat_field' in product.newpath:
# note: a handful of these are missing envi headers,
# which made rasterio choke;
# as all their headers (within global/target)
# are identical, we just make copies for them
writer = M3FlatFieldWriter(product_file)
writer.write_pds4(
output_dir + product.newpath + "/",
write_product_files=True
)
elif 'bad_detector_element' in product.newpath:
writer = M3BDEWriter(product_file)
writer.write_pds4(
output_dir + product.newpath + "/",
write_product_files=True
)
elif 'smooth_shape_curve' in product.newpath:
writer = M3SSCWriter(product_file)
writer.write_pds4(
output_dir + product.newpath + "/",
write_product_files=True
)
# enforce match between LID, filename, and putative product
enforce_name_match("".join(writer.PDS4_LABEL), writer.pds4_root)
# validate only one in 10 of these
if product.Index % 10 == 0:
print("0-mod-10th templated document: running Validate Tool")
validate_results = sh.validate("-t", writer.pds4_label_file)
with open("validate_dump.txt", "a") as file:
file.write(validate_results.stdout.decode())
print("validated successfully")
# this is just a stupid bandaid for logging
if isinstance(product.root, str):
root = product.root
else:
root = writer.pds4_root
crude_time_log(
"m3_document_conversion_log",
namedtuple('stupid_bandaid','pds4_label_file')(product.newpath+'/'+root+'.xml'),
str((dt.datetime.now() - doc_start_time).total_seconds())
)
print(
"done with this document; total seconds "
+ str((dt.datetime.now() - doc_start_time).total_seconds())
)
```
| github_jupyter |
# SLU02 - Data Structures
### Start by importing the following packages
```
#used for evaluation
import hashlib
import json
import random
```
In this notebook the following is tested:
- Tuples
- Lists
- Dictionaries
## Exercise 1: Tuples <a name="1"></a>
This exercise covers topics learned regarding tuples.
### 1.1) Create a tuple
Create a tuple of __floats__ named `this_tuple` with size 5.
```
# this_tuple = ...
### BEGIN SOLUTION
this_tuple = (1., 2., 3., 4., 5.)
print(type(this_tuple))
print(len(this_tuple))
print(type(this_tuple[3]))
### END SOLUTION
assert isinstance(this_tuple, tuple), "Are you sure this_tuple is a tuple?"
assert len(this_tuple) == 5, "The length is not quite right."
assert isinstance(this_tuple[3], float), "Did you write floats?"
```
### 1.2) Index a tuple
Considering the following tuple:
```
color = ("red", "blue", "green", "yellow", "black", "white")
```
Using __negative__ indexing, assign the index of the element `"green"` to the variable `green_index`.
```
color = ("red", "blue", "green", "yellow", "black", "white")
#hint: the solution should be a number
#green_index = ...
### BEGIN SOLUTION
green_index = -4
print(color[green_index])
print(
'green_index hashed:',
hashlib.sha256(
json.dumps(green_index).encode()
).hexdigest()
)
### END SOLUTION
assert hashlib.sha256(json.dumps(green_index).encode()).hexdigest() == 'e5e0093f285a4fb94c3fcc2ad7fd04edd10d429ccda87a9aa5e4718efadf182e', "The index is not correct. Are you using negative indexing?"
```
### 1.3) Slice a tuple
Extract `("green", "blue", "red")` from tuple `color`.
Assign the results to a variable called `rgb`.
```
color = ("red", "blue", "green", "yellow", "black", "white")
#Hint: use backwards slicing
#rgb = ...
### BEGIN SOLUTION
rgb = color[-4:-7:-1]
print(rgb)
print(
'rgb hashed:',
hashlib.sha256(
json.dumps(rgb).encode()
).hexdigest()
)
### END SOLUTION
assert isinstance(rgb, tuple), "Is your result a tuple?"
assert len(rgb) == 3, "You aren't selecting the correct number of elements."
assert rgb[0] == "green", "Check which elements you are selecting and their order."
assert hashlib.sha256(json.dumps(rgb).encode()).hexdigest() == '43da8949efc00c51f7d96130a25a6b902f6cfd6157817015ca6ee524e3085374'
```
### 1.4) Index a tuple of tuples
Considering the following tuple of tuples:
`random_numbers = ((1, 2, 3),(4, 5, 6),(7, 8, 9),(10, 11, 12))`
What is the right way to extract the number 8 from `random_numbers`?
a) `random_numbers[3][2]`
b) `random_numbers[-1][-1]`
c) `random_numbers[-2][1]`
d) `random_numbers[-1][1]`
```
#uncomment the right answer
#answer = "a"
#answer = "b"
#answer = "c"
#answer = "d"
### BEGIN SOLUTION
random_numbers = ( (1, 2, 3), (4, 5, 6), (7, 8, 9), (10, 11, 12) )
answer = "c"
print(answer)
print(random_numbers[-2][1])
print(
'rgb hashed:',
hashlib.sha256(
json.dumps(answer).encode()
).hexdigest()
)
### END SOLUTION
assert hashlib.sha256(json.dumps(answer).encode()).hexdigest() == '879923da020d1533f4d8e921ea7bac61e8ba41d3c89d17a4d14e3a89c6780d5d', "Wrong answer."
```
### 1.5) Tuple of size one
How can we create a tuple of size 1 __without__ using the function __tuple__?
a) (5)
b) 5
c) [5]
d) 5,
```
#uncomment the right answer
#answer = "a"
#answer = "b"
#answer = "c"
#answer = "d"
### BEGIN SOLUTION
answer = "d"
print(answer)
this_tuple = 5,
print(type(this_tuple))
print(
'rgb hashed:',
hashlib.sha256(
json.dumps(answer).encode()
).hexdigest()
)
### END SOLUTION
assert hashlib.sha256(json.dumps(answer).encode()).hexdigest() == '3fa5834dc920d385ca9b099c9fe55dcca163a6b256a261f8f147291b0e7cf633', "Wrong answer."
```
### 1.6) Replace values in a tuple
Can a tuple be modified after its creation?
```
#uncomment the right answer
#answer = "yes"
#answer = "no"
### BEGIN SOLUTION
answer = "no"
print(answer)
print(
'rgb hashed:',
hashlib.sha256(
json.dumps(answer).encode()
).hexdigest()
)
### END SOLUTION
assert hashlib.sha256(json.dumps(answer).encode()).hexdigest() == '04a06452677210a3cdaec376fd5ebbca1714cb7af9e62bf5cce1644310a9086a', "Wrong answer."
```
### 1.7) Merge two tuples
Considering the following tuples:
```
left = (1,11,22215,7,14,1,11,9,1,6,2,5)
right = (1,24,50,45,2,45,1,1,2,1,2,1,88,9,9,9,44,5,2)
```
Create a tuple by merging tuples above and assign it to a third tuple called `this_tuple`. Elements in tuple __`left`__ should come before the ones on the __`right`__ tuple.
```
left = (1,11,22215,7,14,1,11,9,1,6,2,5)
right = (1,24,50,45,2,45,1,1,2,1,2,1,88,9,9,9,44,5,2)
#hint: Using operations between tuples.
#this_tuple = ...
### BEGIN SOLUTION
this_tuple = left + right
print(this_tuple)
print(type(this_tuple))
print(len(this_tuple))
print(this_tuple[-3])
print(
'rgb hashed:',
hashlib.sha256(
json.dumps(this_tuple).encode()
).hexdigest()
)
### END SOLUTION
assert isinstance(this_tuple, tuple), "The result should be a tuple."
assert len(this_tuple) == 31, "The merging is not right."
assert this_tuple[-3] == 44, "Re-check the order of the tuples."
assert hashlib.sha256(json.dumps(this_tuple).encode()).hexdigest() == '0c1afd35431992aba438b9382b08332f4d4fed7d4380e538c047e42a223a9dd5'
```
## Exercise 2: Lists <a name="2"></a>
This exercise covers topics learned regarding lists.
### 2.1) List Creation
Write a list named `this_list` with length five and with the string `"bananas"` on negative index -4.
```
#bananas = ...
#this_list = ...
### BEGIN SOLUTION
bananas = "bananas"
this_list = ["kiwi", bananas, "melon", "lemon", "kiwi"]
print(this_list)
print(type(this_list))
print(len(this_list))
print(this_list[-4])
### END SOLUTION
assert isinstance(this_list, list), "The result should be a list."
assert len(this_list) == 5, "The length is not quite right."
assert this_list[-4] == "bananas", "There is no \"bananas\" on position -4."
```
### 2.2) Delete and append values in a list
Giving the list `ice_cream`, delete `"chocolate"` and append `"dulce de leche"`.
```
ice_cream = ["lemon", "stracciatella", "pistacchio", "chocolate", "vanilla"]
### BEGIN SOLUTION
print(len(ice_cream))
del ice_cream[-2]
ice_cream.append("dulce de leche")
print("chocolate" in ice_cream)
print(ice_cream[-1])
### END SOLUTION
assert len(ice_cream) == 5, "The operations were not performed as asked."
assert "chocolate" not in ice_cream, "There is still \"chocolate\" on the ice_cream."
assert ice_cream[-1] == "dulce de leche", "Did you append the exact ingredient?"
```
### 2.3) Delete the last value in a list
Considering the following list, `ice_cream = ["lemon", "vanilla", "stracciatella", "pistacchio", "chocolate", "vanilla"]`, what is the right answer if we want to delete `"vanilla"` in the last position?
a) `del ice_cream[1]`
b) `del ice_cream[-1]`
c) `ice_cream.remove("vanilla")`
d) `ice_cream[-1] = None`
```
#uncomment the right answer
#answer = "a"
#answer = "b"
#answer = "c"
#answer = "d"
### BEGIN SOLUTION
answer = "b"
ice_cream = ["lemon", "vanilla", "stracciatella", "pistacchio", "chocolate", "vanilla"]
del ice_cream[1]
print(ice_cream)
ice_cream = ["lemon", "vanilla", "stracciatella", "pistacchio", "chocolate", "vanilla"]
del ice_cream[-1]
print(ice_cream)
ice_cream = ["lemon", "vanilla", "stracciatella", "pistacchio", "chocolate", "vanilla"]
ice_cream.remove("vanilla")
print(ice_cream)
ice_cream = ["lemon", "vanilla", "stracciatella", "pistacchio", "chocolate", "vanilla"]
ice_cream[-1] = None
print(ice_cream)
print(
'rgb hashed:',
hashlib.sha256(
json.dumps(answer).encode()
).hexdigest()
)
### END SOLUTION
assert hashlib.sha256(json.dumps(answer).encode()).hexdigest() == 'c100f95c1913f9c72fc1f4ef0847e1e723ffe0bde0b36e5f36c13f81fe8c26ed', "Wrong answer."
```
### 2.4) List Operations
Using list operations, create a list called `hello_world` of size 10000. It should have just two unique values, `"hello"` and `"world"`. Starting with `"hello"` then `"world"`, then `"hello"` and so on.
```
#Hint: multiply a list by a int
#hello_world = ...
### BEGIN SOLUTION
hello_world = ["hello", "world"] * 5000
print(len(hello_world))
print(hello_world[0])
print(hello_world[3])
print(hello_world[4532])
print(
'rgb hashed:',
hashlib.sha256(
json.dumps(hello_world).encode()
).hexdigest()
)
### END SOLUTION
assert len(hello_world) == 10000, "The size is not right."
assert hello_world[0] == "hello", "I guess you started on the wrong foot."
assert hello_world[3] == "world", "Are the words alternating?"
assert hello_world[4532]== "hello", "Are the words alternating?"
```
### 2.5) Replace values and sort a list
Replace `"pistacchio"` in the list `ice_cream` by `"cream"`. Add the sub-list `others` to the end of the list `ice_cream`.
After these operations, sort the elements in the list and convert it to a __tuple__.
```
ice_cream = ["lemon", "stracciatella", "pistacchio", "chocolate", "vanilla"]
others = ["dulce de leche", "caramel", "cookies", "peanut butter"] * 100
### BEGIN SOLUTION
ice_cream[2] = "cream"
ice_cream = ice_cream + others
ice_cream.sort()
ice_cream = tuple(ice_cream)
print(len(ice_cream))
print(
'rgb hashed:',
hashlib.sha256(
json.dumps(ice_cream).encode()
).hexdigest()
)
### END SOLUTION
assert len(ice_cream) == 405, "The lists weren't merged right."
assert ice_cream.index("cream") == 201, "Did you replaced \"pistacchio\"?"
assert isinstance(ice_cream, tuple), "the result is not a tuple."
assert hashlib.sha256(json.dumps(ice_cream).encode()).hexdigest() == 'bbe23485294172ad0fab1f37eee62d4f518a296aad06ea944d6d7a31e6ef58a6'
```
## Exercise 3: Dictionaries <a name="3"></a>
This exercise covers topics learned regarding dictionaries.
### 3.1) Create a dictionary
Create a dictionary called `this_dict` with 5 key-value pairs where the keys are strings and values are lists.
```
#this_dict = ...
### BEGIN SOLUTION
this_dict = {"a":[],"b":[],"c":[],"d":[],"e":[]}
print(len(this_dict))
print(isinstance(list(this_dict.keys())[0], str))
print(isinstance(list(this_dict.values())[0], list))
print(isinstance(list(this_dict.keys())[2], str))
### END SOLUTION
assert isinstance(this_dict, dict), "The result is not a dictionary."
assert len(this_dict) == 5, "The dictionary doesn't have 5 key-value pairs."
assert isinstance(list(this_dict.keys())[0], str), "The dictionary keys are not strings."
assert isinstance(list(this_dict.values())[0], list), "The dictionary values are not lists."
```
### 3.2) Extract a value from a dictionary
Considering the following dictionary named `groceries`:
```
groceries = {
"bread": {"type": "grains", "price_per_unit": 2, "quantity_purchased": 1},
"onions": {"type": "vegetables", "price_per_unit": 0.5, "quantity_purchased": 2},
"spinages": {"type": "vegetables" , "price_per_unit": 1.5, "quantity_purchased": 1}
}
```
What is the notation that we should use in order to extract "grains"?
a) `groceries["type"]`
b) `groceries["bread"]`
c) `groceries[0][0]`
d) `groceries["bread"]["type"]`
```
#uncomment the right answer
#answer = "a"
#answer = "b"
#answer = "c"
#answer = "d"
### BEGIN SOLUTION
answer = "d"
print(answer)
groceries = {
"bread": {"type": "grains", "price_per_unit": 2, "quantity_purchased": 1},
"onions": {"type": "vegetables", "price_per_unit": 0.5, "quantity_purchased": 2},
"spinages": {"type": "vegetables" , "price_per_unit": 1.5, "quantity_purchased": 1}
}
print(groceries["bread"]["type"])
print(
'rgb hashed:',
hashlib.sha256(
json.dumps(answer).encode()
).hexdigest()
)
### END SOLUTION
assert hashlib.sha256(json.dumps(answer).encode()).hexdigest() == '3fa5834dc920d385ca9b099c9fe55dcca163a6b256a261f8f147291b0e7cf633', "Wrong answer."
```
### 3.3) Replace, Append and Delete operations on dictionaries
Considering the following dictionary named groceries:
```
groceries = {
"bread": {"type": "grains", "price_per_unit": 2, "quantity_purchased": 1},
"onions": {"type": "vegetables", "price_per_unit": 0.5, "quantity_purchased": 2},
"spinages": {"type": "vegetables" , "price_per_unit": 1.5, "quantity_purchased": 1}
}
```
Complete the following questions regarding Replace, Append and Delete of key-value pairs in a dictionary.
#### 3.3.1) Replace a value on a key-value pair
Considering the dictionary `groceries`, update the `price_per_unit` for bread from 2 to 3.
```
### BEGIN SOLUTION
groceries["bread"]["price_per_unit"] = 3
print(groceries["bread"]["price_per_unit"])
### END SOLUTION
assert groceries["bread"]["price_per_unit"]==3
```
#### 3.3.2) Add a key-value pair
Add rice to our groceries. It should be of type grains, with a price per unit of 1 and in quantity 2.
```
### BEGIN SOLUTION
groceries["rice"]={"type": "grains", "price_per_unit": 1, "quantity_purchased": 2}
print(groceries["rice"])
### END SOLUTION
assert groceries["rice"]["type"] == "grains", "The type of grocery is wrong."
assert groceries["rice"]["price_per_unit"] == 1, "The price per unit is wrong."
assert groceries["rice"]["quantity_purchased"] == 2, "The quantity purchased is wrong."
```
#### 3.3.3) Delete a key-value pair
Delete onions from our `groceries` dictionary.
```
### BEGIN SOLUTION
del groceries["onions"]
"onions" not in groceries.keys()
### END SOLUTION
assert "onions" not in groceries.keys(), "There's still \"onions\" on the groceries."
```
### 3.4) Extract keys and values manually
Considering the following `people_age` dictionary, write down a list with all the keys in the dictionary and another one with all the values. Even if a value is duplicated, write it as many times as it appears. Return the lists __sorted__.
`people_age = {"joao":"senior", "bernardo":"adult", "joao":"child", "alberto":"adult", "amilcar":"child" }`
Create the lists manually (without using methods) by just looking at the dictionary above and writing down the values on lists. Also, try to not print the output of the dictionary `people_age`. The idea of this exercise is to guess the output of the dictionary in terms of keys and values.
```
#keys = ...
#values = ...
### BEGIN SOLUTION
keys = ["joao", "bernardo", "alberto", "amilcar"]
values = ["child", "adult", "adult", "child"]
keys.sort()
values.sort()
print(len(keys))
print(len(values))
print(keys)
print(values)
print(
'rgb hashed:',
hashlib.sha256(
json.dumps(keys).encode()
).hexdigest()
)
print(
'rgb hashed:',
hashlib.sha256(
json.dumps(values).encode()
).hexdigest()
)
### END SOLUTION
assert hashlib.sha256(json.dumps(keys).encode()).hexdigest() == '73a6b69681d6d29dc146cfba3fbeed0cf41d233e0ef7ed0cdda84bb0e7c64924', "The keys list is incorrect."
assert hashlib.sha256(json.dumps(values).encode()).hexdigest() == '39fb994d286387448ffae1e44256baace4d8329d6bd59875aeee5d6b7d17b291', "The values list is incorrect."
```
### 3.5) Extract keys and values using methods
Considering the following dictionary called `people_age`, extract keys and values of the dictionary to a variable called `names` and `ages`, respectively. Convert the variable `ages` to a list called `list_ages` and calculate how many people are senior and assign the value to `n_seniors`.
Contrary to exercise 3.4, on this one you should use methods to extract keys and values.
```
#Hint: find out what is the type of the variable age, you might need to convert this variable to another type.
#Hint: you might need to use your list knowledge.
people_age = {"joao":"senior", "bernardo":"adult", "gabriel":"child", "antonio":"senior", "maria":"senior", "joel":"adult", "ines":"adult", "alberto":"adult", "amilcar":"senior", "emilia":"adult", "ana":"adult", "margarida":"adult" }
#names = ...
#ages = ...
#list_ages = ..
#n_seniors = ...
### BEGIN SOLUTION
names = people_age.keys()
print(names)
print(len(names))
ages = people_age.values()
print(ages)
print(len(ages))
list_ages = list(ages)
n_seniors = list_ages.count("senior")
print(
'rgb hashed:',
hashlib.sha256(
json.dumps(list(names)).encode()
).hexdigest()
)
print(
'rgb hashed:',
hashlib.sha256(
json.dumps(list(ages)).encode()
).hexdigest()
)
print(
'rgb hashed:',
hashlib.sha256(
json.dumps(n_seniors).encode()
).hexdigest()
)
### END SOLUTION
assert len(names) == 12, "The number of names is incorrect."
assert hashlib.sha256(json.dumps(list(names)).encode()).hexdigest() == '9a2a381eef2e8212d8c8b48ab6531a62c064e9fbc39091015c8131138174eaf1', "The variable names is incorrect."
assert len(ages) == 12, "The number of ages is incorrect."
assert hashlib.sha256(json.dumps(list(ages)).encode()).hexdigest() == '7ec678f76d3dec3621b7f64343d89d6d9a0b8faaea6b62c529b937439e3b93f9', "The variable ages is incorrect."
assert isinstance(list_ages, list), "list_ages should be a list."
assert hashlib.sha256(json.dumps(n_seniors).encode()).hexdigest() == '4b227777d4dd1fc61c6f884f48641d02b4d121d3fd328cb08b5531fcacdabf8a', "The number of seniors is incorrect."
```
| github_jupyter |
<a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_01_3_python_collections.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# T81-558: Applications of Deep Neural Networks
**Module 1: Python Preliminaries**
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# Module 1 Material
* Part 1.1: Course Overview [[Video]](https://www.youtube.com/watch?v=Rqq-UnVXtMg&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_01_1_overview.ipynb)
* Part 1.2: Introduction to Python [[Video]](https://www.youtube.com/watch?v=czq5d53vKvo&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_01_2_intro_python.ipynb)
* **Part 1.3: Python Lists, Dictionaries, Sets and JSON** [[Video]](https://www.youtube.com/watch?v=kcGx2I5akSs&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_01_3_python_collections.ipynb)
* Part 1.4: File Handling [[Video]](https://www.youtube.com/watch?v=FSuSLCMgCZc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_01_4_python_files.ipynb)
* Part 1.5: Functions, Lambdas, and Map/Reduce [[Video]](https://www.youtube.com/watch?v=jQH1ZCSj6Ng&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_01_5_python_functional.ipynb)
# Google CoLab Instructions
The following code ensures that Google CoLab is running the correct version of TensorFlow.
```
try:
from google.colab import drive
%tensorflow_version 2.x
COLAB = True
print("Note: using Google CoLab")
except:
print("Note: not using Google CoLab")
COLAB = False
```
# Part 1.3: Python Lists, Dictionaries, Sets and JSON
Like most modern programming languages, Python includes Lists, Sets, Dictionaries, and other data structures as built-in types. The syntax appearance of both of these is similar to JSON. Python and JSON compatibility is discussed later in this module. This course will focus primarily on Lists, Sets, and Dictionaries. It is essential to understand the differences between these three fundamental collection types.
* **Dictionary** - A dictionary is a mutable unordered collection that Python indexes with name and value pairs.
* **List** - A list is a mutable ordered collection that allows duplicate elements.
* **Set** - A set is a mutable unordered collection with no duplicate elements.
* **Tuple** - A tuple is an immutable ordered collection that allows duplicate elements.
Most Python collections are mutable, which means that the program can add and remove elements after definition. An immutable collection cannot add or remove items after definition. It is also essential to understand that an ordered collection means that items maintain their order as the program adds them to a collection. This order might not be any specific ordering, such as alphabetic or numeric.
Lists and tuples are very similar in Python and are often confused. The significant difference is that a list is mutable, but a tuple isn’t. So, we include a list when we want to contain similar items, and include a tuple when we know what information goes into it ahead of time.
Many programming languages contain a data collection called an array. The array type is noticeably absent in Python. Generally, the programmer will use a list in place of an array in Python. Arrays in most programming languages were fixed-length, requiring the program to know the maximum number of elements needed ahead of time. This restriction leads to the infamous array-overrun bugs and security issues. The Python list is much more flexible in that the program can dynamically change the size of a list.
The next sections will look at each of these collection types in more detail.
### Lists and Tuples
For a Python program, lists and tuples are very similar. It is possible to get by as a programmer using only lists and ignoring tuples. Both lists and tuples hold an ordered collection of items.
The primary difference that you will see syntactically is that a list is enclosed by square braces [] and a tuple is enclosed by parenthesis (). The following code defines both list and tuple.
```
l = ['a', 'b', 'c', 'd']
t = ('a', 'b', 'c', 'd')
print(l)
print(t)
```
The primary difference that you will see programmatically is that a list is mutable, which means the program can change it. A tuple is immutable, which means the program cannot change it. The following code demonstrates that the program can change a list. This code also illustrates that Python indexes lists starting at element 0. Accessing element one modifies the second element in the collection. One advantage of tuples over lists is that tuples are generally slightly faster to iterate over than lists.
```
l[1] = 'changed'
#t[1] = 'changed' # This would result in an error
print(l)
```
Like many languages, Python has a for-each statement. This statement allows you to loop over every element in a collection, such as a list or a tuple.
```
# Iterate over a collection.
for s in l:
print(s)
```
The **enumerate** function is useful for enumerating over a collection and having access to the index of the element that we are currently on.
```
# Iterate over a collection, and know where your index. (Python is zero-based!)
for i,l in enumerate(l):
print(f"{i}:{l}")
```
A **list** can have multiple objects added to it, such as strings. Duplicate values are allowed. **Tuples** do not allow the program to add additional objects after definition.
```
# Manually add items, lists allow duplicates
c = []
c.append('a')
c.append('b')
c.append('c')
c.append('c')
print(c)
```
Ordered collections, such as lists and tuples, allow you to access an element by its index number, such as is done in the following code. Unordered collections, such as dictionaries and sets, do not allow the program to access them in this way.
```
print(c[1])
```
A **list** can have multiple objects added to it, such as strings. Duplicate values are allowed. Tuples do not allow the program to add additional objects after definition. For the insert function, an index, the programmer must specify an index. These operations are not allowed for tuples because they would result in a change.
```
# Insert
c = ['a', 'b', 'c']
c.insert(0, 'a0')
print(c)
# Remove
c.remove('b')
print(c)
# Remove at index
del c[0]
print(c)
```
### Sets
A Python **set** holds an unordered collection of objects, but sets do *not* allow duplicates. If a program adds a duplicate item to a set, only one copy of each item remains in the collection. Adding a duplicate item to a set does not result in an error. Any of the following techniques will define a set.
```
s = set()
s = { 'a', 'b', 'c'}
s = set(['a', 'b', 'c'])
print(s)
```
A **list** is always enclosed in square braces [], a **tuple** in parenthesis (), and now we see that the programmer encloses a **set** in curly braces. Programs can add items to a **set** as they run. Programs can dynamically add items to a **set** with the **add** function. It is important to note that the **append** function adds items to lists and tuples, whereas the **add** function adds items to a **set**.
```
# Manually add items, sets do not allow duplicates
# Sets add, lists append. I find this annoying.
c = set()
c.add('a')
c.add('b')
c.add('c')
c.add('c')
print(c)
```
## Maps/Dictionaries/Hash Tables
Many programming languages include the concept of a map, dictionary, or hash table. These are all very related concepts. Python provides a dictionary, that is essentially a collection of name-value pairs. Programs define dictionaries using curly-braces, as seen here.
```
d = {'name': "Jeff", 'address':"123 Main"}
print(d)
print(d['name'])
if 'name' in d:
print("Name is defined")
if 'age' in d:
print("age defined")
else:
print("age undefined")
```
Be careful that you do not attempt to access an undefined key, as this will result in an error. You can check to see if a key is defined, as demonstrated above. You can also access the directory and provide a default value, as the following code demonstrates.
```
d.get('unknown_key', 'default')
```
You can also access the individual keys and values of a dictionary.
```
d = {'name': "Jeff", 'address':"123 Main"}
# All of the keys
print(f"Key: {d.keys()}")
# All of the values
print(f"Values: {d.values()}")
```
Dictionaries and lists can be combined. This syntax is closely related to [JSON](https://en.wikipedia.org/wiki/JSON). Dictionaries and lists together are a good way to build very complex data structures. While Python allows quotes (") and apostrophe (') for strings, JSON only allows double-quotes ("). We will cover JSON in much greater detail later in this module.
The following code shows a hybrid usage of dictionaries and lists.
```
# Python list & map structures
customers = [
{"name": "Jeff & Tracy Heaton", "pets": ["Wynton", "Cricket",
"Hickory"]},
{"name": "John Smith", "pets": ["rover"]},
{"name": "Jane Doe"}
]
print(customers)
for customer in customers:
print(f"{customer['name']}:{customer.get('pets', 'no pets')}")
```
The variable **customers** is a list that holds three dictionaries that represent customers. You can think of these dictionaries as records in a table. The fields in these individual records are the keys of the dictionary. Here the keys **name** and **pets** are fields. However, the field **pets** holds a list of pet names. There is no limit to how deep you might choose to nest lists and maps. It is also possible to nest a map inside of a map or a list inside of another list.
## More Advanced Lists
Several advanced features are available for lists that this section introduces. One such function is **zip**. Two lists can be combined into a single list by the **zip** command. The following code demonstrates the **zip** command.
```
a = [1,2,3,4,5]
b = [5,4,3,2,1]
print(zip(a,b))
```
To see the results of the **zip** function, we convert the returned zip object into a list. As you can see, the **zip** function returns a list of tuples. Each tuple represents a pair of items that the function zipped together. The order in the two lists was maintained.
```
a = [1,2,3,4,5]
b = [5,4,3,2,1]
print(list(zip(a,b)))
```
The usual method for using the zip command is inside of a for-loop. The following code shows how a for-loop can assign a variable to each collection that the program is iterating.
```
a = [1,2,3,4,5]
b = [5,4,3,2,1]
for x,y in zip(a,b):
print(f'{x} - {y}')
```
Usually, both collections will be of the same length when passed to the zip command. It is not an error to have collections of different lengths. As the following code illustrates, the zip command will only process elements up to the length of the smaller collection.
```
a = [1,2,3,4,5]
b = [5,4,3]
print(list(zip(a,b)))
```
Sometimes you may wish to know the current numeric index when a for-loop is iterating through an ordered collection. Use the **enumerate** command to track the index location for a collection element. Because the **enumerate** command deals with numeric indexes of the collection, the zip command will assign arbitrary indexes to elements from unordered collections.
Consider how you might construct a Python program to change every element greater than 5 to the value of 5. The following program performs this transformation. The enumerate command allows the loop to know which element index it is currently on, thus allowing the program to be able to change the value of the current element of the collection.
```
a = [2, 10, 3, 11, 10, 3, 2, 1]
for i, x in enumerate(a):
if x>5:
a[i] = 5
print(a)
```
The comprehension command can dynamically build up a list. The comprehension below counts from 0 to 9 and adds each value (multiplied by 10) to a list.
```
lst = [x*10 for x in range(10)]
print(lst)
```
A dictionary can also be a comprehension. The general format for this is:
```
dict_variable = {key:value for (key,value) in dictonary.items()}
```
A common use for this is to build up an index to symbolic column names.
```
text = ['col-zero','col-one', 'col-two', 'col-three']
lookup = {key:value for (value,key) in enumerate(text)}
print(lookup)
```
This can be used to easily find the index of a column by name.
```
print(f'The index of "col-two" is {lookup["col-two"]}')
```
### An Introduction to JSON
Data stored in a CSV file must be flat; that is, it must fit into rows and columns. Most people refer to this type of data as structured or tabular. This data is tabular because the number of columns is the same for every row. Individual rows may be missing a value for a column; however, these rows still have the same columns.
This sort of data is convenient for machine learning because most models, such as neural networks, also expect incoming data to be of fixed dimensions. Real-world information is not always so tabular. Consider if the rows represent customers. These people might have multiple phone numbers and addresses. How would you describe such data using a fixed number of columns? It would be useful to have a list of these courses in each row that can be of a variable length for each row, or student.
JavaScript Object Notation (JSON) is a standard file format that stores data in a hierarchical format similar to eXtensible Markup Language (XML). JSON is nothing more than a hierarchy of lists and dictionaries. Programmers refer to this sort of data as semi-structured data or hierarchical data. The following is a sample JSON file.
```
{
"firstName": "John",
"lastName": "Smith",
"isAlive": true,
"age": 27,
"address": {
"streetAddress": "21 2nd Street",
"city": "New York",
"state": "NY",
"postalCode": "10021-3100"
},
"phoneNumbers": [
{
"type": "home",
"number": "212 555-1234"
},
{
"type": "office",
"number": "646 555-4567"
},
{
"type": "mobile",
"number": "123 456-7890"
}
],
"children": [],
"spouse": null
}
```
The above file may look somewhat like Python code. You can see curly braces that define dictionaries and square brackets that define lists. JSON does require there to be a single root element. A list or dictionary can fulfill this role. JSON requires double-quotes to enclose strings and names. Single quotes are not allowed in JSON.
JSON files are always legal JavaScript syntax. JSON is also generally valid as Python code, as demonstrated by the following Python program.
```
jsonHardCoded = {
"firstName": "John",
"lastName": "Smith",
"isAlive": True,
"age": 27,
"address": {
"streetAddress": "21 2nd Street",
"city": "New York",
"state": "NY",
"postalCode": "10021-3100"
},
"phoneNumbers": [
{
"type": "home",
"number": "212 555-1234"
},
{
"type": "office",
"number": "646 555-4567"
},
{
"type": "mobile",
"number": "123 456-7890"
}
],
"children": [],
"spouse": None
}
```
Generally, it is better to read JSON from files, strings, or the Internet than hard coding, as demonstrated here. However, for internal data structures, sometimes such hard-coding can be useful.
Python contains support for JSON. When a Python program loads a JSON the root list or dictionary is returned, as demonstrated by the following code.
```
import json
json_string = '{"first":"Jeff","last":"Heaton"}'
obj = json.loads(json_string)
print(f"First name: {obj['first']}")
print(f"Last name: {obj['last']}")
```
Python programs can also load JSON from a file or URL.
```
import requests
r = requests.get("https://raw.githubusercontent.com/jeffheaton/"
+"t81_558_deep_learning/master/person.json")
print(r.json())
```
Python programs can easily generate JSON strings from Python objects of dictionaries and lists.
```
python_obj = {"first":"Jeff","last":"Heaton"}
print(json.dumps(python_obj))
```
A data scientist will generally encounter JSON when they access web services to get their data. A data scientist might use the techniques presented in this section to convert the semi-structured JSON data into tabular data for the program to use with a model such as a neural network.
| github_jupyter |
```
import os
#os.environ["CUDA_VISIBLE_DEVICES"]="2"
stage = "stage_2"
experiment_name = 'NONAMED'
import fastai
from fastai.vision import *
from fastai.callbacks import *
from fastai.distributed import *
from fastai.utils.mem import *
import re
import pydicom
import pdb
import pickle
import torch
!ls models
arch = 'resnet18'
fold=4
lr = 1e-3
model_fn = None
SZ = 512
n_folds=5
n_epochs = 15
n_tta = 10
#model_fn = 'resnet34_sz512_cv0.0821_weighted_loss_fold1_of_5'
if model_fn is not None:
model_fn_fold = int(model_fn[-6])-1
assert model_fn_fold == fold
data_dir = Path('data/unzip')
train_dir = data_dir / f'{stage}_train_images'
test_dir = data_dir / f'{stage}_test_images'
fn_to_study_ix = pickle.load(open(f'data/{stage}_fn_to_study_ix.pickle', 'rb'))
study_ix_to_fn = pickle.load(open(f'data/{stage}_study_ix_to_fn.pickle', 'rb'))
fn_to_labels = pickle.load( open(f'data/{stage}_train_fn_to_labels.pickle', 'rb'))
study_to_data = pickle.load (open(f'data/{stage}_study_to_data.pickle', 'rb'))
# gets a pathlib.Path, returns labels
labels = [
'any0', 'epidural0', 'intraparenchymal0', 'intraventricular0', 'subarachnoid0', 'subdural0',
'any1', 'epidural1', 'intraparenchymal1', 'intraventricular1', 'subarachnoid1', 'subdural1',
'any2', 'epidural2', 'intraparenchymal2', 'intraventricular2', 'subarachnoid2', 'subdural2',
]
def get_labels(center_p):
study, center_ix = fn_to_study_ix[center_p.stem]
total = len(study_ix_to_fn[study])
# Only 1 study has 2 different RescaleIntercept values (0.0 and 1.0) in the same series, so assume they're all the
# same and do everything in the big tensor
pixels = {}
img_labels = []
ixs = [ max(0, min(ix, total-1)) for ix in range(center_ix-1, center_ix+2) ]
for i, ix in enumerate(ixs):
fn = study_ix_to_fn[study][ix]
img_labels += [ x + str(i) for x in fn_to_labels[fn] ]
return img_labels
get_labels(train_dir / 'ID_ac7c8fe8b.dcm')
sample_submission = pd.read_csv(f'data/unzip/{stage}_sample_submission.csv')
sample_submission['fn'] = sample_submission.ID.apply(lambda x: '_'.join(x.split('_')[:2]) + '.dcm')
sample_submission.head()
test_fns = sample_submission.fn.unique()
wc = 50
ww = 100
def open_dicom(p_or_fn): # when called by .from_folder it's a Path; when called from .add_test it's a string
#pdb.set_trace()
center_p = Path(p_or_fn)
study, center_ix = fn_to_study_ix[center_p.stem]
total = len(study_ix_to_fn[study])
# Only 1 study has 2 different RescaleIntercept values (0.0 and 1.0) in the same series, so assume they're all the
# same and do everything in the big tensor
pixels = {}
ixs = [ max(0, min(ix, total-1)) for ix in range(center_ix-1, center_ix+2) ]
for ix in ixs:
if ix not in pixels:
p = center_p.parent / f'{study_ix_to_fn[study][ix]}.dcm'
dcm = pydicom.dcmread(str(p))
if ix == center_ix:
rescale_slope, rescale_intercept = float(dcm.RescaleSlope), float(dcm.RescaleIntercept)
pixels[ix] = torch.FloatTensor(dcm.pixel_array.astype(np.float))
t = torch.stack([pixels[ix] for ix in ixs], dim=0) # stack chans together
if (t.shape[1:] != (SZ,SZ)):
t = torch.nn.functional.interpolate(
t.unsqueeze_(0), size=SZ, mode='bilinear', align_corners=True).squeeze_(0) # resize
t = t * rescale_slope + rescale_intercept # rescale
t = torch.clamp(t, wc-ww/2, wc+ww/2) # window
t = (t - (wc-ww/2)) / ww # normalize
return Image(t)
class DicomList(ImageList):
def open(self, fn): return open_dicom(fn)
my_stats = ([0.45, 0.45, 0.45], [0.225, 0.225, 0.225])
class OverSamplingCallback(LearnerCallback):
def __init__(self,learn:Learner):
super().__init__(learn)
self.labels = self.learn.data.train_dl.dataset.y.items
_, counts = np.unique(self.labels,return_counts=True)
self.weights = torch.DoubleTensor((1/counts)[self.labels])
self.label_counts = np.bincount([self.learn.data.train_dl.dataset.y[i].data for i in range(len(self.learn.data.train_dl.dataset))])
self.total_len_oversample = int(self.learn.data.c*np.max(self.label_counts))
def on_train_begin(self, **kwargs):
self.learn.data.train_dl.dl.batch_sampler = BatchSampler(WeightedRandomSampler(weights,self.total_len_oversample), self.learn.data.train_dl.batch_size,False)
# From: https://forums.fast.ai/t/is-there-any-built-in-method-to-oversample-minority-classes/46838/5
class ImbalancedDatasetSampler(torch.utils.data.sampler.Sampler):
def __init__(self, dataset, indices=None, num_samples=None):
# if indices is not provided,
# all elements in the dataset will be considered
self.indices = list(range(len(dataset))) if indices is None else indices
# if num_samples is not provided,
# draw `len(indices)` samples in each iteration
self.num_samples = len(self.indices) if num_samples is None else num_samples
# distribution of classes in the dataset
label_to_count = defaultdict(int)
for idx in self.indices:
label = self._get_label(dataset, idx)
label_to_count[label] += 1
# weight for each sample
weights = [1.0 / label_to_count[self._get_label(dataset, idx)] for idx in self.indices]
self.weights = torch.DoubleTensor(weights)
def _get_label(self, dataset, idx):
return 'any1' in dataset.y[idx].obj
def __iter__(self):
return (self.indices[i] for i in torch.multinomial(
self.weights, self.num_samples, replacement=True))
def __len__(self):
return self.num_samples
folds = np.array_split(list({v['fold'] for v in study_to_data.values()}), n_folds)
folds
df = pd.read_csv(f"data/{stage}_train_dicom_diags_norm.csv")
# split train/val
np.random.seed(666)
studies_in_fold = [k for k,v in study_to_data.items() if np.isin(v['fold'],folds[fold])]
study_id_val_set = set(df[ df['SeriesInstanceUID'].isin(studies_in_fold)]['SOPInstanceUID'] + '.dcm')
def is_val(p_or_fn):
p = Path(p_or_fn)
return p.name in study_id_val_set
tfms = (
[ flip_lr(p=0.5), rotate(degrees=(-180,180), p=1.) ], # train
[] # val
)
data = (DicomList.from_folder(f'data/unzip/{stage}_train_images/', extensions=['.dcm'], presort=True)
.split_by_valid_func(is_val)
.label_from_func(get_labels, classes=labels)
.transform(tfms, mode='nearest')
.add_test(f'data/unzip/{stage}_test_images/' + test_fns))
db = data.databunch().normalize()
balsampler = ImbalancedDatasetSampler(db.train_ds)
db.train_dl.batch_sampler = balsampler
db.train_dl.batch_sampler
db
db.show_batch() # AttributeError: 'list' object has no attribute 'pixel' <-- because using torch datasets or whatever
# yuval reina's loss
yuval_weights = FloatTensor([ 2, 1, 1, 1, 1, 1, 2, 1, 1, 1, 1, 1, 2, 1, 1, 1, 1, 1 ]).cuda()
def yuval_loss(y_pred,y_true):
return F.binary_cross_entropy_with_logits(y_pred,
y_true,
yuval_weights.repeat(y_pred.shape[0],1))
real_lb_weights = FloatTensor([ 2, 1, 1, 1, 1, 1 ])
def real_lb_loss(pred:Tensor, targ:Tensor)->Rank0Tensor:
pred,targ = flatten_check(pred,targ)
tp = pred.view(-1,18)[:,6:12]
tt = targ.view(-1,18)[:,6:12]
return F.binary_cross_entropy_with_logits(tp, tt, real_lb_weights.to(device=pred.device))
w_loss = 0.1
weighted_lb_weights = FloatTensor([ 2.*w_loss, 1.*w_loss, 1.*w_loss, 1.*w_loss, 1.*w_loss, 1.*w_loss,
2., 1., 1., 1., 1., 1.,
2.*w_loss, 1.*w_loss, 1.*w_loss, 1.*w_loss, 1.*w_loss, 1.*w_loss, ])
def weighted_loss(pred:Tensor,targ:Tensor)->Rank0Tensor:
return F.binary_cross_entropy_with_logits(pred,targ,weighted_lb_weights.to(device=pred.device))
lb_weights = FloatTensor([ 2, 1, 1, 1, 1, 1 ]).cuda()
def lb_loss(pred:Tensor, targ:Tensor)->Rank0Tensor:
pred,targ = flatten_check(pred,targ)
tp = pred.view(-1,6)
tt = targ.view(-1,6)
return torch.nn.functional.binary_cross_entropy_with_logits(tp, tt, lb_weights.to(device=pred.device))
learn = cnn_learner(
db, getattr(models, arch), loss_func=weighted_loss, metrics=[real_lb_loss],
pretrained=True, lin_ftrs=[],ps=0.,
model_dir = Path('./models/').resolve())
try:
learn.load(model_fn,strict=False)
print(f"Loaded {model_fn}")
except Exception as e:
print(e)
learn = learn.to_parallel().to_fp16()
arch_to_batch_factor = {
'resnet18' : 4.3,
'resnet34' : 3.4,
'resnet50' : 1.5,
'resnet101' : 1.0,
'resnext50_32x4d': 1.2,
'densenet121': 1.1,
'squeezenet1_0': 2.6,
'vgg16': 1.5
}
bs = arch_to_batch_factor[arch] * 512*512 * gpu_mem_get()[0] * 1e-3 / (SZ*SZ)
bs *= int(int(any([isinstance(cb, MixedPrecision) for cb in learn.callbacks])))+1 # 2x if fp16
if any([isinstance(cb, fastai.distributed.ParallelTrainer) for cb in learn.callbacks]):
bs *= torch.cuda.device_count()
bs = (bs // torch.cuda.device_count()) * torch.cuda.device_count()
bs = int(bs)
learn.data.batch_size = bs
bs
learn
learn.unfreeze()
learn.summary()
try:
if lr is None:
!rm {learn.model_dir}/tmp.pth
learn.lr_find()
learn.recorder.plot(suggestion=True)
except Exception as e:
print(e)
# set lr
learn.fit_one_cycle(n_epochs, lr)
```
# Save
```
v = learn.validate()
cv = float(v[-1])
model_fn = f'{arch}_sz{SZ}_cv{cv:0.4f}_{learn.loss_func.__name__}_fold{fold+1}_of_{n_folds}'
learn.save(model_fn)
model_fn
```
# Predictions
```
learn.data.test_ds.tfms = learn.data.train_ds.tfms
test_preds = []
# Fastai WTF: it figures out to run outputs through sigmoid if a standard loss error is used
# (see loss_func_name2activ and related stuff) but on custom loss funcs sigmoid must be executed explicitly:
# https://forums.fast.ai/t/shouldnt-we-able-to-pass-an-activ-function-to-learner-get-preds/50492
for _ in progress_bar(range(n_tta)):
preds, _ = learn.get_preds(DatasetType.Test, activ=torch.sigmoid)
test_preds.append(preds)
tta_test_preds = torch.cat([p.unsqueeze(0) for p in test_preds],dim=0)
learn.data.valid_ds.tfms = learn.data.train_ds.tfms
valid_preds = []
# Fastai WTF: it figures out to run outputs through sigmoid if a standard loss error is used
# (see loss_func_name2activ and related stuff) but on custom loss funcs sigmoid must be executed explicitly:
# https://forums.fast.ai/t/shouldnt-we-able-to-pass-an-activ-function-to-learner-get-preds/50492
for _ in progress_bar(range(n_tta)):
preds, _ = learn.get_preds(DatasetType.Valid, activ=torch.sigmoid)
valid_preds.append(preds)
tta_valid_preds = torch.cat([p.unsqueeze(0) for p in valid_preds],dim=0)
PREDS_DIR = 'data/predictions'
!mkdir -p {PREDS_DIR}
torch.save(tta_test_preds, f'{PREDS_DIR}/{model_fn}_test.pth')
torch.save(tta_valid_preds, f'{PREDS_DIR}/{model_fn}_valid.pth')
pd.DataFrame({'fn': [p.stem for p in learn.data.valid_ds.x.items]}).to_csv(
f'{PREDS_DIR}/{model_fn}_valid_fns.csv',index=False)
pd.DataFrame({'fn': [Path(p).stem for p in learn.data.test_ds.x.items]}).to_csv(
f'{PREDS_DIR}/{model_fn}_test_fns.csv',index=False)
```
# Submit to Kaggle
```
tta_test_preds_mean = tta_test_preds.mean(dim=0)
tta_test_preds_geomean = torch.expm1(torch.log1p(tta_test_preds).mean(dim=0))
ids = []
labels = []
for fn, pred in zip(test_fns, tta_test_preds_mean):
for i, label in enumerate(db.train_ds.classes):
if label.endswith('1'):
ids.append(f"{fn.split('.')[0]}_{label.strip('1')}")
predicted_probability = '{0:1.10f}'.format(pred[i].item())
labels.append(predicted_probability)
mkdir -p data/submissions
sub_name = experiment_name + "_" + model_fn
sub_path = f'data/submissions/{sub_name}.csv.zip'
sub_name, sub_path
pd.DataFrame({'ID': ids, 'Label': labels}).to_csv(sub_path, compression='zip', index=False)
!ls data/submissions/
!echo {sub_path}
!kaggle competitions submit -c rsna-intracranial-hemorrhage-detection -f {sub_path} -m {model_fn}
```
| github_jupyter |
# Importing the Libraries
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import pickle
%matplotlib inline
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.model_selection import train_test_split,cross_validate,GridSearchCV
data = pd.read_csv('diabetes.csv')
data
```
# Feature Engineering and Exploratory Data Analysis
```
data['diabetes'] = data['diabetes'].apply(lambda x:1 if x==True else 0)
data.info()
data.shape
sns.pairplot(data, hue='diabetes',vars = data.columns[1:-1])
corrmat = data.corr()
top_corr_features = corrmat.index
plt.figure(figsize=(20,20))
#plot heat map
g=sns.heatmap(data[top_corr_features].corr(),annot=True,cmap="RdYlGn")
sns.countplot(data['diabetes'])
data.corr()
print("total number of rows : {0}".format(len(data)))
print("number of rows missing glucose_conc: {0}".format(len(data.loc[data['glucose_conc'] == 0])))
print("number of rows missing diastolic_bp: {0}".format(len(data.loc[data['diastolic_bp'] == 0])))
print("number of rows missing insulin: {0}".format(len(data.loc[data['insulin'] == 0])))
print("number of rows missing bmi: {0}".format(len(data.loc[data['bmi'] == 0])))
print("number of rows missing diab_pred: {0}".format(len(data.loc[data['diab_pred'] == 0])))
print("number of rows missing age: {0}".format(len(data.loc[data['age'] == 0])))
print("number of rows missing skin: {0}".format(len(data.loc[data['skin'] == 0])))
na_columns = ['glucose_conc','thickness','insulin','bmi','diastolic_bp']
data[na_columns] = data[na_columns].replace(0,np.nan)
df = data.copy()
for column in ['glucose_conc','thickness','insulin']:
median_0 = data[column][data['diabetes']==0].median()
median_1 = data[column][data['diabetes']==1].median()
df[column][df['diabetes']==0] = data[column][df['diabetes']==0].fillna(median_0)
df[column][df['diabetes']==1] = data[column][df['diabetes']==1].fillna(median_1)
df
df.diastolic_bp.fillna(df.diastolic_bp.median(),inplace=True)
df.bmi.fillna(df.bmi.median(),inplace=True)
X = df.drop('diabetes',axis=1)
X
y = df.diabetes
y
data.hist(column='bmi',bins=50,by='diabetes',figsize=(12,8))
data.hist(column='age',bins=50,by='diabetes',figsize=(12,8))
data.hist(column='glucose_conc',bins=50,by='diabetes',figsize=(12,8))
data.hist(column='insulin',bins=50,by='diabetes',figsize=(12,8))
```
# Creating metrics for evaluations
```
from sklearn import metrics
f1 = metrics.make_scorer(metrics.f1_score)
accuracy = metrics.make_scorer(metrics.accuracy_score)
precision = metrics.make_scorer(metrics.precision_score)
recall = metrics.make_scorer(metrics.recall_score)
auc = metrics.make_scorer(metrics.roc_auc_score)
scoring = {
"accuracy":accuracy,
"precision":precision,
"recall": recall,
"f1":f1,
}
def printResults(cv):
print("Accuracy {:.3f} ({:.3f})".format(cv["test_accuracy"].mean(), cv["test_accuracy"].std()))
print("Precision {:.3f} ({:.3f})".format(cv["test_precision"].mean(), cv["test_precision"].std()))
print("Recall {:.3f} ({:.3f})".format(cv["test_recall"].mean(), cv["test_recall"].std()))
print("F1 {:.3f} ({:.3f})".format(cv["test_f1"].mean(), cv["test_f1"].std()))
```
# Creating our ML Model
```
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25)
gbc = GradientBoostingClassifier()
gbc.fit(X_train,y_train)
gbc.fit(X_train,y_train)
y_pred = gbc.predict(X_test)
print(metrics.classification_report(y_test, y_pred))
cm = metrics.confusion_matrix(y_test, y_pred)
sns.heatmap(cm, annot=True, cmap="Blues");
cv_gbc = cross_validate(gbc, X, y, scoring=scoring, cv=5)
printResults(cv_gbc)
```
# Tuning our ML Model
```
params = {
'loss': ['deviance','exponential'],
'learning_rate': [0.1,0.2,0.3,0.4,0.5,0.6,0.7,0.8,0.9,1.0],
'n_estimators': [100,200,300,400,500,600,700,800,800,1000],
}
gs = GridSearchCV(estimator = gbc,param_grid=params,cv=5)
gs.fit(X,y)
gs.best_score_
gs.best_params_
gbc_best = GradientBoostingClassifier(learning_rate=0.1,loss='deviance',n_estimators=400)
gbc_best.fit(X_train,y_train)
y_pred = gbc_best.predict(X_test)
print(metrics.classification_report(y_test, y_pred))
cm = metrics.confusion_matrix(y_test, y_pred)
sns.heatmap(cm, annot=True, cmap="Blues");
cv_gbc_best = cross_validate(gbc_best, X, y, cv=5, scoring=scoring)
printResults(cv_gbc_best)
from sklearn.model_selection import cross_val_score
accuracies = cross_val_score(gbc_best, X, y,cv = 10)
#cv is the number of folds you want your training set to split in
import pickle
filename = 'diabetes-model.pkl'
pickle.dump(gbc_best,open(filename,'wb'))
```
| github_jupyter |
# pMEC1138
This vector expresses eight genes and was assembled from the eight single gene expression cassettes below:
Gene | Enzyme | Acronym | Cassette
-------------------------------------------------- |-------------------|---|-----|
[SsXYL1](http://www.ncbi.nlm.nih.gov/gene/4839234) |D-xylose reductase |XR | [pYPK0_TEF1_PsXYL1_N272D_TDH3](pYPK0_TEF1_PsXYL1_N272D_TDH3.ipynb)
[SsXYL2](http://www.ncbi.nlm.nih.gov/gene/4852013) |xylitol dehydrogenase |XDH | [pYPK0_TDH3_PsXYL2_PGI1](pYPK0_TDH3_PsXYL2_PGI1.ipynb)
[ScXKS1](http://www.yeastgenome.org/locus/S000003426/overview) |Xylulokinase |XK | [pYPK0_PGI1_ScXKS1_FBA1](pYPK0_PGI1_ScXKS1_FBA1.ipynb)
[ScTAL1](http://www.yeastgenome.org/locus/S000004346/overview) |Transaldolase |tal1p | [pYPK0_FBA1_ScTAL1_PDC1](pYPK0_FBA1_ScTAL1_PDC1.ipynb)
[ScTKL1](http://www.yeastgenome.org/locus/S000006278/overview) |Transketolase |TKL | [pYPK0_PDC1_ScTKL1_RPS19b](pYPK0_PDC1_ScTKL1_RPS19b.ipynb)
[ScRPE1](http://www.yeastgenome.org/locus/S000003657/overview) |xylitol dehydrogenase |RPE | [pYPK0_RPS19b_ScRPE1_RPS19a](pYPK0_RPS19b_ScRPE1_RPS19a.ipynb)
[ScRKI1](http://www.yeastgenome.org/locus/S000005621/overview) |Xylulokinase |RKI | [pYPK0_RPS19a_ScRKI1_TPI1](pYPK0_RPS19a_ScRKI1_TPI1.ipynb)
[CiGXF1](http://mic.sgmjournals.org/content/154/6/1646.full) |Transaldolase |GXF | [pYPK0_TPI1_CiGXF1_ENO2](pYPK0_TPI1_CiGXF1_ENO2.ipynb)
TEF1_XR_N272D_TDH3
TDH3_XDH_PGI1
PGI1_XK_FBA1
FBA1_TAL1_PDC1
PDC1_ScTKL1_RPS19b
RPS19b_ScRPE1_RPS19a
RPS19a_ScRKI1_TPI1
TPI1_CiGXF1_ENO2
[pMEC1135](pMEC1135.ipynb)
[pMEC1137](pMEC1137.ipynb)
```
from pydna.all import *
import ipynb
from ipynb.fs.full.pMEC1135 import cas1, cas2, cas3, p4, p409
from ipynb.fs.full.pMEC1137 import p1 as p5, cas2 as cas6, cas3 as cas7, cas4 as cas8, p625, p647, p413
cas4 = pcr(p409, p625, p4)
cas5 = pcr(p413, p647, p5)
pYPK0 = read("pYPK0.gb")
pYPK0.cseguid()
from Bio.Restriction import ZraI, AjiI, EcoRV
pYPK0_E_Z, stuffer = pYPK0.cut((EcoRV, ZraI))
pYPK0_E_Z, stuffer
pYPK0_E_Z, cas1, cas2, cas3, cas4, cas5, cas6, cas7, cas8
asm =Assembly( (pYPK0_E_Z, cas1, cas2, cas3, cas4, cas5, cas6, cas7, cas8) , limit = 61)
asm
candidate = asm.assemble_circular()[0]
candidate.figure()
pw = candidate.synced(pYPK0)
len(pw)
pw.cseguid()
import textwrap
pw.name = "pMEC1138"
pw.description="pYPK0_TEF1_XR_N272D_TPI1_XDH_PDC1_XK_FBA1_TAL1_PDC1_ScTKL1_RPS19b_ScRPE1_RPS19a_ScRKI1_TPI1_CiGXF1_ENO2 (alternative name pYPK0_8)"
pw.stamp()
pw.description = "\n".join( textwrap.wrap( pw.description, 40) )
pw.write("pMEC1138.gb")
```
| github_jupyter |
# 环境搭建
# 换源
在容器内/root下创建**sources.list**文件, 复制下列内容到文件中
```bash
deb http://mirrors.aliyun.com/ubuntu/ trusty main restricted universe multiverse
deb http://mirrors.aliyun.com/ubuntu/ trusty-security main restricted universe multiverse
deb http://mirrors.aliyun.com/ubuntu/ trusty-updates main restricted universe multiverse
deb http://mirrors.aliyun.com/ubuntu/ trusty-proposed main restricted universe multiverse
deb http://mirrors.aliyun.com/ubuntu/ trusty-backports main restricted universe multiverse
deb-src http://mirrors.aliyun.com/ubuntu/ trusty main restricted universe multiverse
deb-src http://mirrors.aliyun.com/ubuntu/ trusty-security main restricted universe multiverse
deb-src http://mirrors.aliyun.com/ubuntu/ trusty-updates main restricted universe multiverse
deb-src http://mirrors.aliyun.com/ubuntu/ trusty-proposed main restricted universe multiverse
deb-src http://mirrors.aliyun.com/ubuntu/ trusty-backports main restricted universe multiverse
```
替换默认源为阿里源
```bash
cp source.txt /etc/apt/sources.list
```
## 安装解压工具
```bash
sudo apt install unzip
```
## Ubuntu16.04 升级 python3.5到 python3.6
1. 在Ubuntu中安装python3.6
```bash
sudo apt-get install software-properties-common
sudo add-apt-repository ppa:jonathonf/python-3.6
sudo apt-get update
sudo apt-get install python3.6
```
2. 这个时候使用pip -V查询,会发现pip还是python3.5的pip,如何指向python3.6呢,首先是删除pip
```bash
sudo apt-get remove python3-pip
sudo apt-get autoremove
```
3. 然后再安装pip
```bash
sudo apt-get install python3-pip
```
4. 切换python版本
```bash
sudo update-alternatives --install /usr/bin/python3 python3 /usr/bin/python3.5 1
sudo update-alternatives --install /usr/bin/python3 python3 /usr/bin/python3.6 2
sudo update-alternatives --config python3
```
4. 发现pip还是指向 python3.5的,这个时候再用python3.6指定升级一下pip:
```bash
python3.6 -m pip install --upgrade pip
```
5. 验证pip版本
```bash
python3 -V
pip -V
```
安装openssh-server并启动
apt-get update
apt-get install openssh-server
# 启动之前需手动创建/var/run/sshd,不然启动sshd的时候会报错
mkdir -p /var/run/sshd
# sshd以守护进程运行
/usr/sbin/sshd -D &
# 安装netstat,查看sshd是否监听22端口
apt-get install net-tools
netstat -apn | grep ssh
如果已经监听22端口,说明sshd服务启动成功
4.ssh登陆
# 生成ssh key
ssh-keygen -t rsa
# 修改sshd-config允许root登陆
sed -i 's+PermitRootLogin prohibit-password+PermitRootLogin yes' /etc/ssh/sshd-config
修改完sshd-config之后需要重启sshd服务
// 找到pid
ps -aux | grep ssh
kill -9 pid
/usr/sbin/sshd -D &
查看容器ip
ifconfig
在主机上进行登陆
ssh root@ip
## 环境检查
本项目使用python3.6编写,推荐使用虚拟环境执行,以模块化方式加载
```bash
python3.6 -m venv venv
source venv/bin/activate
```
## 安装项目依赖
```bash
sudo pip install -r requirements.txt -i https://pypi.douban.com/simple/
```
* bleach==1.5.0
* certifi==2017.7.27.1
* chardet==3.0.4
* cycler==0.10.0
* decorator==4.1.2
* html5lib==0.9999999
* idna==2.6
* Keras==2.0.8
* Markdown==2.6.9
* matplotlib==2.1.0
* networkx==2.0
* numpy==1.13.3
* olefile==0.44
* Pillow==4.3.0
* protobuf==3.4.0
* pyparsing==2.2.0
* python-dateutil==2.6.1
* python-resize-image==1.1.11
* pytz==2017.2
* PyWavelets==0.5.2
* PyYAML==3.12
* requests==2.18.4
* scikit-image==0.13.1
* scipy==0.19.1
* six==1.11.0
* tensorflow==1.8.0
* tensorflow-tensorboard==0.1.8
* urllib3==1.22
* Werkzeug==0.12.2
apt install python-pil
pip install python-resize-image
## 安装tree显示文件目录树
安装工具命令
```bash
sudo apt install tree
```
显示文件目录树请执行
```bash
tree ~/imagenet
```
```
~/imagenet
├── fall11_urls.txt
├── imagenet1000_clsid_to_human.pkl
├── inception_resnet_v2_2016_08_30.ckpt
├── original
├── resized
└── tfrecords
```
## Python下 "No module named _tkinter" 问题解决过程总结
如果在某些机型产生找不到模块_tkinter错误,请按照以下方法解决
错误提示最后一行如下
ImportError: No module named _tkinter
请执行如下命令安装依赖
```bash
sudo apt install tk-dev (Ubuntu/Debian)
yum install tk-devel (CentOS)
```
在安装之后,重新执行程序,错误仍然存在。
```bash
sudo apt install python3-tk (Ubuntu)
yum install python3-tk (Centos)
```
## 下载数据集
以imageNet数据集为例,但实际上我们使用的是ins的数据集,分为50个class,图片质量更高
```bash
python3 -m dataset.download -c 100 -s fall1_url.txt -o ~/imagenet/original/ --skip 1000
```
检查目录下文件数量(下载进度)
```bash
ls -l| grep "^" | wc -l
```
## 调整数据集的大小resize(299,299)
```bash
python3 -m dataset.resize -s ~/imagenet/original -o ~/imagenet/resized -v 1000 > ./log/resize.log
```
## 使用预训练模型写入数据
```bash
python3 -m dataset.lab_batch -c ~/imagenet/inception_resnet_v2_2016_08_30.ckpt
```
## 训练与评测模型
划分训练集和验证集
训练脚本将训练所有的训练图像,在每一个epoch结束的时候检查权值,并从测试集预测一些彩色图像并保存。
```bash
python3.6 -m colorization.train
```
加载预训练模型,图像必须先被序列化。
```bash
python3.6 -m colorization.evaluate
```
## 参考
[环境安装教程](https://blog.csdn.net/silence1772/article/details/78118549)
[使用tf-slim的inception_resnet_v2预训练模型进行图像分类](https://blog.csdn.net/Wayne2019/article/details/78109357)
[Tensorflow源码编译,解决tf提示未使用SSE4.1 SSE4.2 AVX警告](https://blog.csdn.net/qq_36810544/article/details/78799037)
| github_jupyter |

[](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/Certification_Trainings/Healthcare/Spark%20v2.7.6%20Notebooks/7.Clinical_NER_Chunk_Merger.ipynb)
# 7. Clinical NER Chunk Merger
```
import json
from google.colab import files
license_keys = files.upload()
with open(list(license_keys.keys())[0]) as f:
license_keys = json.load(f)
%%capture
for k,v in license_keys.items():
%set_env $k=$v
!wget https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/jsl_colab_setup.sh
!bash jsl_colab_setup.sh -p 2.4.4
! pip install spark-nlp-display
import json
import os
from pyspark.ml import Pipeline,PipelineModel
from pyspark.sql import SparkSession
from sparknlp.annotator import *
from sparknlp_jsl.annotator import *
from sparknlp.base import *
import sparknlp_jsl
import sparknlp
params = {"spark.driver.memory":"16G",
"spark.kryoserializer.buffer.max":"2000M",
"spark.driver.maxResultSize":"2000M"}
spark = sparknlp_jsl.start(license_keys['SECRET'],params=params)
print (sparknlp.version())
print (sparknlp_jsl.version())
# if you want to start the session with custom params as in start function above
def start(secret):
builder = SparkSession.builder \
.appName("Spark NLP Licensed") \
.master("local[*]") \
.config("spark.driver.memory", "16G") \
.config("spark.serializer", "org.apache.spark.serializer.KryoSerializer") \
.config("spark.kryoserializer.buffer.max", "2000M") \
.config("spark.jars.packages", "com.johnsnowlabs.nlp:spark-nlp_2.11:"+version) \
.config("spark.jars", "https://pypi.johnsnowlabs.com/"+secret+"/spark-nlp-jsl-"+jsl_version+".jar")
return builder.getOrCreate()
#spark = start(secret)
spark
# Sample data
data_chunk_merge = spark.createDataFrame([
(1,"""A 63-year-old man presents to the hospital with a history of recurrent infections that include cellulitis, pneumonias, and upper respiratory tract infections. He reports subjective fevers at home along with unintentional weight loss and occasional night sweats. The patient has a remote history of arthritis, which was diagnosed approximately 20 years ago and treated intermittently with methotrexate (MTX) and prednisone. On physical exam, he is found to be febrile at 102°F, rather cachectic, pale, and have hepatosplenomegaly. Several swollen joints that are tender to palpation and have decreased range of motion are also present. His laboratory values show pancytopenia with the most severe deficiency in neutrophils.
""")]).toDF("id","text")
data_chunk_merge.show(truncate=50)
# Annotator that transforms a text column from dataframe into an Annotation ready for NLP
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
# Sentence Detector annotator, processes various sentences per line
sentenceDetector = SentenceDetector()\
.setInputCols(["document"])\
.setOutputCol("sentence")
# Tokenizer splits words in a relevant format for NLP
tokenizer = Tokenizer()\
.setInputCols(["sentence"])\
.setOutputCol("token")
# Clinical word embeddings trained on PubMED dataset
word_embeddings = WordEmbeddingsModel.pretrained("embeddings_clinical", "en", "clinical/models")\
.setInputCols(["sentence", "token"])\
.setOutputCol("embeddings")
# NER model trained on i2b2 (sampled from MIMIC) dataset
clinical_ner = NerDLModel.pretrained("ner_deid_large", "en", "clinical/models") \
.setInputCols(["sentence", "token", "embeddings"]) \
.setOutputCol("clinical_ner")
clinical_ner_converter = NerConverter() \
.setInputCols(["sentence", "token", "clinical_ner"]) \
.setOutputCol("clinical_ner_chunk")
# Cancer Genetics NER
bionlp_ner = NerDLModel.pretrained("ner_bionlp", "en", "clinical/models") \
.setInputCols(["sentence", "token", "embeddings"]) \
.setOutputCol("bionlp_ner")
bionlp_ner_converter = NerConverter() \
.setInputCols(["sentence", "token", "bionlp_ner"]) \
.setOutputCol("bionlp_ner_chunk")
# merge ner_chunks by prioritizing the overlapping indices (chunks with longer lengths and highest information will be kept from each ner model)
chunk_merger_1 = ChunkMergeApproach()\
.setInputCols('clinical_ner_chunk', "bionlp_ner_chunk")\
.setOutputCol('clinical_bionlp_ner_chunk')
# internal clinical NER (general terms)
jsl_ner = NerDLModel.pretrained("ner_jsl", "en", "clinical/models") \
.setInputCols(["sentence", "token", "embeddings"]) \
.setOutputCol("jsl_ner")
jsl_ner_converter = NerConverter() \
.setInputCols(["sentence", "token", "jsl_ner"]) \
.setOutputCol("jsl_ner_chunk")
# merge ner_chunks by prioritizing the overlapping indices (chunks with longer lengths and highest information will be kept from each ner model)
chunk_merger_2 = ChunkMergeApproach()\
.setInputCols('clinical_bionlp_ner_chunk', "jsl_ner_chunk")\
.setOutputCol('final_ner_chunk')
# merge ner_chunks regardess of overlapping indices
# only works with 2.7 and later
chunk_merger_NonOverlapped = ChunkMergeApproach()\
.setInputCols('clinical_bionlp_ner_chunk', "jsl_ner_chunk")\
.setOutputCol('nonOverlapped_ner_chunk')\
.setMergeOverlapping(False)
nlpPipeline = Pipeline(stages=[
documentAssembler,
sentenceDetector,
tokenizer,
word_embeddings,
clinical_ner,
clinical_ner_converter,
bionlp_ner,
bionlp_ner_converter,
chunk_merger_1,
jsl_ner,
jsl_ner_converter,
chunk_merger_2,
chunk_merger_NonOverlapped])
empty_data = spark.createDataFrame([[""]]).toDF("text")
model = nlpPipeline.fit(empty_data)
merged_data = model.transform(data_chunk_merge).cache()
from pyspark.sql import functions as F
result_df = merged_data.select('id',F.explode('final_ner_chunk').alias("cols")) \
.select('id',F.expr("cols.begin").alias("begin"),
F.expr("cols.end").alias("end"),
F.expr("cols.result").alias("chunk"),
F.expr("cols.metadata.entity").alias("entity"))
result_df.show(50, truncate=100)
```
## NonOverlapped Chunk
all the entities form each ner model will be returned one by one
```
from pyspark.sql import functions as F
result_df2 = merged_data.select('id',F.explode('nonOverlapped_ner_chunk').alias("cols")) \
.select('id',F.expr("cols.begin").alias("begin"),
F.expr("cols.end").alias("end"),
F.expr("cols.result").alias("chunk"),
F.expr("cols.metadata.entity").alias("entity"))
result_df2.show(50, truncate=100)
```
| github_jupyter |
# Prediction over Iris Dataset with XGB classifer :
In this project we are going perform a simple prediction using very popular boosting model XGBoostClassifier.
```
data_path = 'iris.csv'
# Important Libraries
import pandas as pd
from matplotlib import pyplot as plt
import seaborn as sns
iris_data = pd.read_csv(data_path)
# Visualize the head of the dataframe
iris_data.head()
print('shape of the data :',iris_data.shape)
# Overview of the data
iris_data.describe()
# Visualizing the target density
counts = iris_data['species'].value_counts()
plt.pie(counts.values,labels=counts.index)
plt.show()
# Performimg Min-Max scaling :
def minmaxscaling(data):
for feature in data.columns:
if data[feature].dtype != 'object':
min_value = min(data[feature])
max_value = max(data[feature])
data[feature] = (data[feature]-min_value) / (max_value-min_value)
return data
scaled_df = minmaxscaling(iris_data)
scaled_df.head()
# Creating train and test data :
# In this part we are going to split the train and test data as such
# that both the train and test have equality in target feature.
train = pd.DataFrame(columns=scaled_df.columns)
test = pd.DataFrame(columns=scaled_df.columns)
for sp_type in scaled_df['species'].unique():
temp_df = scaled_df[scaled_df['species'] == sp_type ]
temp_df = temp_df.sample(frac = 1) # shuffling the data
length = len(temp_df)//5 # test_size defined
train = pd.concat([train,temp_df[length:]] , axis = 0)
test = pd.concat([test,temp_df[:length]] , axis = 0)
print(train.shape,test.shape)
# splitting featureset and target value
X_train = train.drop('species',1)
y_train=train['species']
X_test = test.drop('species',1)
y_test = test['species']
y_train.unique()
# Numerating the targets :
#-------------------------
# sentosa : 0
# versicolor : 1
# virginica : 2
labels = ['setosa', 'versicolor', 'virginica']
for i in range(3):
y_train.replace(labels[i] , i , inplace = True)
y_test.replace(labels[i] , i , inplace = True )
print('X_train --------> ',X_train.shape)
print('y_train --------> ',y_train.shape)
print('X_train --------> ',X_test.shape)
print('X_train --------> ',y_test.shape)
# Importing training model
from xgboost import XGBClassifier as xgb
model = xgb(verbosity = 1)
print(model)
# Model training on train data
model.fit(X_train,y_train)
print('Train Data Accuracy :',model.score(X_train,y_train)*100,'%')
print('Test Data Accuracy :',model.score(X_test,y_test)*100,'%')
# for confusion matrix plotting
from mlxtend.plotting import plot_confusion_matrix
from sklearn.metrics import confusion_matrix
def plot_conf_matrix( featureset , target , model ):
prediction = model.predict( featureset ).tolist()
mat = confusion_matrix( target , prediction )
plot_confusion_matrix( mat , figsize=(5,5) , colorbar = True)
plot_conf_matrix(X_train,y_train,model)
plot_conf_matrix(X_test,y_test,model)
```
## HURRAH !!!
We've completed the project.
| github_jupyter |
# Sentiment Analysis with an RNN
In this notebook, you'll implement a recurrent neural network that performs sentiment analysis.
>Using an RNN rather than a strictly feedforward network is more accurate since we can include information about the *sequence* of words.
Here we'll use a dataset of movie reviews, accompanied by sentiment labels: positive or negative.
<img src="assets/reviews_ex.png" width=40%>
### Network Architecture
The architecture for this network is shown below.
<img src="assets/network_diagram.png" width=40%>
>**First, we'll pass in words to an embedding layer.** We need an embedding layer because we have tens of thousands of words, so we'll need a more efficient representation for our input data than one-hot encoded vectors. You should have seen this before from the Word2Vec lesson. You can actually train an embedding with the Skip-gram Word2Vec model and use those embeddings as input, here. However, it's good enough to just have an embedding layer and let the network learn a different embedding table on its own. *In this case, the embedding layer is for dimensionality reduction, rather than for learning semantic representations.*
>**After input words are passed to an embedding layer, the new embeddings will be passed to LSTM cells.** The LSTM cells will add *recurrent* connections to the network and give us the ability to include information about the *sequence* of words in the movie review data.
>**Finally, the LSTM outputs will go to a sigmoid output layer.** We're using a sigmoid function because positive and negative = 1 and 0, respectively, and a sigmoid will output predicted, sentiment values between 0-1.
We don't care about the sigmoid outputs except for the **very last one**; we can ignore the rest. We'll calculate the loss by comparing the output at the last time step and the training label (pos or neg).
---
### Load in and visualize the data
```
import numpy as np
# read data from text files
with open('data/reviews.txt', 'r') as f:
reviews = f.read()
with open('data/labels.txt', 'r') as f:
labels = f.read()
print(reviews[:2000])
print()
print(labels[:20])
```
## Data pre-processing
The first step when building a neural network model is getting your data into the proper form to feed into the network. Since we're using embedding layers, we'll need to encode each word with an integer. We'll also want to clean it up a bit.
You can see an example of the reviews data above. Here are the processing steps, we'll want to take:
>* We'll want to get rid of periods and extraneous punctuation.
* Also, you might notice that the reviews are delimited with newline characters `\n`. To deal with those, I'm going to split the text into each review using `\n` as the delimiter.
* Then I can combined all the reviews back together into one big string.
First, let's remove all punctuation. Then get all the text without the newlines and split it into individual words.
```
from string import punctuation
print(punctuation)
# get rid of punctuation
reviews = reviews.lower() # lowercase, standardize
all_text = ''.join([c for c in reviews if c not in punctuation])
type(reviews)
# split by new lines and spaces
reviews_split = all_text.split('\n')
all_text = ' '.join(reviews_split)
words = all_text.split()
words[:30]
```
### Encoding the words
The embedding lookup requires that we pass in integers to our network. The easiest way to do this is to create dictionaries that map the words in the vocabulary to integers. Then we can convert each of our reviews into integers so they can be passed into the network.
> **Exercise:** Now you're going to encode the words with integers. Build a dictionary that maps words to integers. Later we're going to pad our input vectors with zeros, so make sure the integers **start at 1, not 0**.
> Also, convert the reviews to integers and store the reviews in a new list called `reviews_ints`.
```
# feel free to use this import
from collections import Counter
## Build a dictionary that maps words to integers
counts = Counter(words)
# sort counts dict by appearing frequency
vocab = sorted(counts, key=counts.get, reverse=True)
# create an index for each word from most common
vocab_to_int = {word: ii for ii, word in enumerate(vocab, 1)}
## use the dict to tokenize each review in reviews_split
## store the tokenized reviews in reviews_ints
reviews_ints = []
for review in reviews_split:
reviews_ints.append([vocab_to_int[word] for word in review.split()])
```
**Test your code**
As a text that you've implemented the dictionary correctly, print out the number of unique words in your vocabulary and the contents of the first, tokenized review.
```
# stats about vocabulary
print('Unique words: ', len((vocab_to_int))) # should ~ 74000+
print()
# print tokens in first review
print('Tokenized review: \n', reviews_ints[:1])
```
### Encoding the labels
Our labels are "positive" or "negative". To use these labels in our network, we need to convert them to 0 and 1.
> **Exercise:** Convert labels from `positive` and `negative` to 1 and 0, respectively, and place those in a new list, `encoded_labels`.
```
# 1=positive, 0=negative label conversion
labels_split = labels.split('\n')
encoded_labels = np.array([1 if label == 'positive' else 0 for label in labels_split])
```
### Removing Outliers
As an additional pre-processing step, we want to make sure that our reviews are in good shape for standard processing. That is, our network will expect a standard input text size, and so, we'll want to shape our reviews into a specific length. We'll approach this task in two main steps:
1. Getting rid of extremely long or short reviews; the outliers
2. Padding/truncating the remaining data so that we have reviews of the same length.
<img src="assets/outliers_padding_ex.png" width=40%>
Before we pad our review text, we should check for reviews of extremely short or long lengths; outliers that may mess with our training.
```
# outlier review stats
review_lens = Counter([len(x) for x in reviews_ints])
print("Zero-length reviews: {}".format(review_lens[0]))
print("Maximum review length: {}".format(max(review_lens)))
```
Okay, a couple issues here. We seem to have one review with zero length. And, the maximum review length is way too many steps for our RNN. We'll have to remove any super short reviews and truncate super long reviews. This removes outliers and should allow our model to train more efficiently.
> **Exercise:** First, remove *any* reviews with zero length from the `reviews_ints` list and their corresponding label in `encoded_labels`.
```
print('Number of reviews before removing outliers: ', len(reviews_ints))
## remove any reviews/labels with zero length from the reviews_ints list.
# checking for reviews that are not 0 length
non_zero_idx = [i for i, review in enumerate(reviews_ints) if len(review) != 0]
reviews_ints = [reviews_ints[i] for i in non_zero_idx]
encoded_labels = np.array([encoded_labels[i] for i in non_zero_idx])
print('Number of reviews after removing outliers: ', len(reviews_ints))
```
---
## Padding sequences
To deal with both short and very long reviews, we'll pad or truncate all our reviews to a specific length. For reviews shorter than some `seq_length`, we'll pad with 0s. For reviews longer than `seq_length`, we can truncate them to the first `seq_length` words. A good `seq_length`, in this case, is 200.
> **Exercise:** Define a function that returns an array `features` that contains the padded data, of a standard size, that we'll pass to the network.
* The data should come from `review_ints`, since we want to feed integers to the network.
* Each row should be `seq_length` elements long.
* For reviews shorter than `seq_length` words, **left pad** with 0s. That is, if the review is `['best', 'movie', 'ever']`, `[117, 18, 128]` as integers, the row will look like `[0, 0, 0, ..., 0, 117, 18, 128]`.
* For reviews longer than `seq_length`, use only the first `seq_length` words as the feature vector.
As a small example, if the `seq_length=10` and an input review is:
```
[117, 18, 128]
```
The resultant, padded sequence should be:
```
[0, 0, 0, 0, 0, 0, 0, 117, 18, 128]
```
**Your final `features` array should be a 2D array, with as many rows as there are reviews, and as many columns as the specified `seq_length`.**
This isn't trivial and there are a bunch of ways to do this. But, if you're going to be building your own deep learning networks, you're going to have to get used to preparing your data.
```
def pad_features(reviews_ints, seq_length):
''' Return features of review_ints, where each review is padded with 0's
or truncated to the input seq_length.
'''
## implement function
# getting the correct rows x cols shape
features = np.zeros((len(reviews_ints), seq_length), dtype=int)
# for each review, I grab that review and
for i, row in enumerate(reviews_ints):
features[i, -len(row):] = np.array(row)[:seq_length]
return features
# Test your implementation!
seq_length = 200
features = pad_features(reviews_ints, seq_length=seq_length)
## test statements - do not change - ##
assert len(features)==len(reviews_ints), "Your features should have as many rows as reviews."
assert len(features[0])==seq_length, "Each feature row should contain seq_length values."
# print first 10 values of the first 30 batches
print(features[:30,:10])
```
## Training, Validation, Test
With our data in nice shape, we'll split it into training, validation, and test sets.
> **Exercise:** Create the training, validation, and test sets.
* You'll need to create sets for the features and the labels, `train_x` and `train_y`, for example.
* Define a split fraction, `split_frac` as the fraction of data to **keep** in the training set. Usually this is set to 0.8 or 0.9.
* Whatever data is left will be split in half to create the validation and *testing* data.
```
split_frac = 0.8
## split data into training, validation, and test data (features and labels, x and y)
train_idx = int(len(features)*split_frac)
train_x, remaining_x = features[:train_idx], features[train_idx:]
train_y, remaining_y = encoded_labels[:train_idx], encoded_labels[train_idx:]
valid_test_idx = int(len(remaining_x)*0.5)
val_x, test_x = remaining_x[:valid_test_idx], remaining_x[:valid_test_idx]
val_y, test_y = remaining_y[:valid_test_idx], remaining_y[:valid_test_idx]
## print out the shapes of your resultant feature data
print("\t\t\tFeatures Shapes:",
"\n Train set: \t\t{}".format(train_x.shape),
"\n Valid set: \t\t{}".format(val_x.shape),
"\n Test set: \t\t{}".format(test_x.shape))
```
**Check your work**
With train, validation, and test fractions equal to 0.8, 0.1, 0.1, respectively, the final, feature data shapes should look like:
```
Feature Shapes:
Train set: (20000, 200)
Validation set: (2500, 200)
Test set: (2500, 200)
```
---
## DataLoaders and Batching
After creating training, test, and validation data, we can create DataLoaders for this data by following two steps:
1. Create a known format for accessing our data, using [TensorDataset](https://pytorch.org/docs/stable/data.html#) which takes in an input set of data and a target set of data with the same first dimension, and creates a dataset.
2. Create DataLoaders and batch our training, validation, and test Tensor datasets.
```
train_data = TensorDataset(torch.from_numpy(train_x), torch.from_numpy(train_y))
train_loader = DataLoader(train_data, batch_size=batch_size)
```
This is an alternative to creating a generator function for batching our data into full batches.
```
import torch
from torch.utils.data import TensorDataset, DataLoader
# create Tensor datasets
train_data = TensorDataset(torch.from_numpy(train_x), torch.from_numpy(train_y))
valid_data = TensorDataset(torch.from_numpy(val_x), torch.from_numpy(val_y))
test_data = TensorDataset(torch.from_numpy(test_x), torch.from_numpy(test_y))
# dataloaders
batch_size = 50
# make sure to SHUFFLE your data
train_loader = DataLoader(train_data, shuffle=True, batch_size=batch_size)
valid_loader = DataLoader(valid_data, shuffle=True, batch_size=batch_size)
test_loader = DataLoader(test_data, shuffle=True, batch_size=batch_size)
# obtain one batch of training data
dataiter = iter(train_loader)
sample_x, sample_y = dataiter.next()
print('Sample input size: ', sample_x.size()) # batch_size, seq_length
print('Sample input: \n', sample_x)
print()
print('Sample label size: ', sample_y.size()) # batch_size
print('Sample label: \n', sample_y)
```
---
# Sentiment Network with PyTorch
Below is where you'll define the network.
<img src="assets/network_diagram.png" width=40%>
The layers are as follows:
1. An [embedding layer](https://pytorch.org/docs/stable/nn.html#embedding) that converts our word tokens (integers) into embeddings of a specific size.
2. An [LSTM layer](https://pytorch.org/docs/stable/nn.html#lstm) defined by a hidden_state size and number of layers
3. A fully-connected output layer that maps the LSTM layer outputs to a desired output_size
4. A sigmoid activation layer which turns all outputs into a value 0-1; return **only the last sigmoid output** as the output of this network.
### The Embedding Layer
We need to add an [embedding layer](https://pytorch.org/docs/stable/nn.html#embedding) because there are 74000+ words in our vocabulary. It is massively inefficient to one-hot encode that many classes. So, instead of one-hot encoding, we can have an embedding layer and use that layer as a lookup table. You could train an embedding layer using Word2Vec, then load it here. But, it's fine to just make a new layer, using it for only dimensionality reduction, and let the network learn the weights.
### The LSTM Layer(s)
We'll create an [LSTM](https://pytorch.org/docs/stable/nn.html#lstm) to use in our recurrent network, which takes in an input_size, a hidden_dim, a number of layers, a dropout probability (for dropout between multiple layers), and a batch_first parameter.
Most of the time, you're network will have better performance with more layers; between 2-3. Adding more layers allows the network to learn really complex relationships.
> **Exercise:** Complete the `__init__`, `forward`, and `init_hidden` functions for the SentimentRNN model class.
Note: `init_hidden` should initialize the hidden and cell state of an lstm layer to all zeros, and move those state to GPU, if available.
```
# First checking if GPU is available
train_on_gpu=torch.cuda.is_available()
if(train_on_gpu):
print('Training on GPU.')
else:
print('No GPU available, training on CPU.')
import torch.nn as nn
class SentimentRNN(nn.Module):
"""
The RNN model that will be used to perform Sentiment analysis.
"""
def __init__(self, vocab_size, output_size, embedding_dim, hidden_dim, n_layers, drop_prob=0.5):
"""
Initialize the model by setting up the layers.
"""
super(SentimentRNN, self).__init__()
self.output_size = output_size
self.n_layers = n_layers
self.hidden_dim = hidden_dim
# define all layers
self.in_embed = nn.Embedding(vocab_size, embedding_dim)
self.lstm = nn.LSTM(embedding_dim, hidden_dim, n_layers, dropout=drop_prob, batch_first=True)
self.dropout = nn.Dropout(p=drop_prob)
self.fc = nn.Linear(hidden_dim, output_size)
def forward(self, x, hidden):
"""
Perform a forward pass of our model on some input and hidden state.
"""
batch_size = x.size(0)
# return last sigmoid output and hidden state
embeds = self.embedding(x)
lstm_output, hidden = self.lstm(embeds, hidden)
output = self.dropout(lstm_output)
# stacking up LSTM layers
output = output.continguous.view(-1, self.hidden_dim)
# Dropout and linear layer
output = self.dropout(output)
sig_out = F.sigmoid(self.fc(output, output_size))
sig_out = sig_out.view(batch_size, -1)
sig_out = sig_out[:, -1] # get last batch of labels
return sig_out, hidden
def init_hidden(self, batch_size):
''' Initializes hidden state '''
# Create two new tensors with sizes n_layers x batch_size x hidden_dim,
# initialized to zero, for hidden state and cell state of LSTM
return hidden
```
## Instantiate the network
Here, we'll instantiate the network. First up, defining the hyperparameters.
* `vocab_size`: Size of our vocabulary or the range of values for our input, word tokens.
* `output_size`: Size of our desired output; the number of class scores we want to output (pos/neg).
* `embedding_dim`: Number of columns in the embedding lookup table; size of our embeddings.
* `hidden_dim`: Number of units in the hidden layers of our LSTM cells. Usually larger is better performance wise. Common values are 128, 256, 512, etc.
* `n_layers`: Number of LSTM layers in the network. Typically between 1-3
> **Exercise:** Define the model hyperparameters.
```
# Instantiate the model w/ hyperparams
vocab_size = len(vocab_to_int)+1
output_size = 1
embedding_dim = 400
hidden_dim = 512
n_layers = 2
net = SentimentRNN(vocab_size, output_size, embedding_dim, hidden_dim, n_layers)
print(net)
```
---
## Training
Below is the typical training code. If you want to do this yourself, feel free to delete all this code and implement it yourself. You can also add code to save a model by name.
>We'll also be using a new kind of cross entropy loss, which is designed to work with a single Sigmoid output. [BCELoss](https://pytorch.org/docs/stable/nn.html#bceloss), or **Binary Cross Entropy Loss**, applies cross entropy loss to a single value between 0 and 1.
We also have some data and training hyparameters:
* `lr`: Learning rate for our optimizer.
* `epochs`: Number of times to iterate through the training dataset.
* `clip`: The maximum gradient value to clip at (to prevent exploding gradients).
```
# loss and optimization functions
lr=0.001
criterion = nn.BCELoss()
optimizer = torch.optim.Adam(net.parameters(), lr=lr)
# training params
epochs = 4 # 3-4 is approx where I noticed the validation loss stop decreasing
counter = 0
print_every = 100
clip=5 # gradient clipping
# move model to GPU, if available
if(train_on_gpu):
net.cuda()
net.train()
# train for some number of epochs
for e in range(epochs):
# initialize hidden state
h = net.init_hidden(batch_size)
# batch loop
for inputs, labels in train_loader:
counter += 1
if(train_on_gpu):
inputs, labels = inputs.cuda(), labels.cuda()
# Creating new variables for the hidden state, otherwise
# we'd backprop through the entire training history
h = tuple([each.data for each in h])
# zero accumulated gradients
net.zero_grad()
# get the output from the model
output, h = net(inputs, h)
# calculate the loss and perform backprop
loss = criterion(output.squeeze(), labels.float())
loss.backward()
# `clip_grad_norm` helps prevent the exploding gradient problem in RNNs / LSTMs.
nn.utils.clip_grad_norm_(net.parameters(), clip)
optimizer.step()
# loss stats
if counter % print_every == 0:
# Get validation loss
val_h = net.init_hidden(batch_size)
val_losses = []
net.eval()
for inputs, labels in valid_loader:
# Creating new variables for the hidden state, otherwise
# we'd backprop through the entire training history
val_h = tuple([each.data for each in val_h])
if(train_on_gpu):
inputs, labels = inputs.cuda(), labels.cuda()
output, val_h = net(inputs, val_h)
val_loss = criterion(output.squeeze(), labels.float())
val_losses.append(val_loss.item())
net.train()
print("Epoch: {}/{}...".format(e+1, epochs),
"Step: {}...".format(counter),
"Loss: {:.6f}...".format(loss.item()),
"Val Loss: {:.6f}".format(np.mean(val_losses)))
```
---
## Testing
There are a few ways to test your network.
* **Test data performance:** First, we'll see how our trained model performs on all of our defined test_data, above. We'll calculate the average loss and accuracy over the test data.
* **Inference on user-generated data:** Second, we'll see if we can input just one example review at a time (without a label), and see what the trained model predicts. Looking at new, user input data like this, and predicting an output label, is called **inference**.
```
# Get test data loss and accuracy
test_losses = [] # track loss
num_correct = 0
# init hidden state
h = net.init_hidden(batch_size)
net.eval()
# iterate over test data
for inputs, labels in test_loader:
# Creating new variables for the hidden state, otherwise
# we'd backprop through the entire training history
h = tuple([each.data for each in h])
if(train_on_gpu):
inputs, labels = inputs.cuda(), labels.cuda()
# get predicted outputs
output, h = net(inputs, h)
# calculate loss
test_loss = criterion(output.squeeze(), labels.float())
test_losses.append(test_loss.item())
# convert output probabilities to predicted class (0 or 1)
pred = torch.round(output.squeeze()) # rounds to the nearest integer
# compare predictions to true label
correct_tensor = pred.eq(labels.float().view_as(pred))
correct = np.squeeze(correct_tensor.numpy()) if not train_on_gpu else np.squeeze(correct_tensor.cpu().numpy())
num_correct += np.sum(correct)
# -- stats! -- ##
# avg test loss
print("Test loss: {:.3f}".format(np.mean(test_losses)))
# accuracy over all test data
test_acc = num_correct/len(test_loader.dataset)
print("Test accuracy: {:.3f}".format(test_acc))
```
### Inference on a test review
You can change this test_review to any text that you want. Read it and think: is it pos or neg? Then see if your model predicts correctly!
> **Exercise:** Write a `predict` function that takes in a trained net, a plain text_review, and a sequence length, and prints out a custom statement for a positive or negative review!
* You can use any functions that you've already defined or define any helper functions you want to complete `predict`, but it should just take in a trained net, a text review, and a sequence length.
```
# negative test review
test_review_neg = 'The worst movie I have seen; acting was terrible and I want my money back. This movie had bad acting and the dialogue was slow.'
# Preprocess the test review
from string import punctuation
test_review_neg
def tokenize_review(test_review):
# lowercase
test_review = test_review.lower()
# remove punctuation
test_text = ''.join([c for c in test_review if c not in punctuation])
# split test_text to rows of words alone
test_text_words = test_text.split()
# tokenize the words
test_text_tokenized = []
for review in test_text_words:
test_text_tokenized.append([vocab_to_int[word] for word in test_text_words])
return test_text_tokenized
test_text_tokenized = tokenize_review(test_review_neg)
print(test_text_tokenized)
# pad the features to avoid dealing with reviews that are not in the same length
seq_length = 200
features = pad_features(test_text_tokenized, seq_length)
# Conversion to tensor
feature_tensor = torch.from_numpy(features)
def predict(net, test_review, sequence_length=200):
''' Prints out whether a give review is predicted to be
positive or negative in sentiment, using a trained model.
params:
net - A trained net
test_review - a review made of normal text and punctuation
sequence_length - the padded length of a review
'''
# turn on evaluation mode
net.eval()
# generate tokens from text
test_text_tokenized = tokenize_review(test_review)
# pad features
features = pad_features(test_text_tokenized, seq_length = sequence_length)
# convert to tensor
features_tensor = torch.from_numpy(features)
batch_size = feature_tensor.size(0)
# initiate hidden state
h = net.init_hidden(batch_size)
if(train_on_gpu):
feature_tensor = feature_tensor.cuda()
# get the output from the model
output, h = net(feature_tensor, h)
# convert output probabilities to predicted class (0 or 1)
pred = torch.round(output.squeeze())
# printing output value, before rounding
print('Prediction value, pre-rounding: {:.6f}'.format(output.item()))
# print custom response
if(pred.item()==1):
print("Positive review detected!")
else:
print("Negative review detected.")
# positive test review
test_review_pos = 'This movie had the best acting and the dialogue was so good. I loved it.'
# call function
# try negative and positive reviews!
seq_length=200
predict(net, test_review_neg, seq_length)
```
### Try out test_reviews of your own!
Now that you have a trained model and a predict function, you can pass in _any_ kind of text and this model will predict whether the text has a positive or negative sentiment. Push this model to its limits and try to find what words it associates with positive or negative.
Later, you'll learn how to deploy a model like this to a production environment so that it can respond to any kind of user data put into a web app!
| github_jupyter |
# UNet/FPN Fastai
* The entire operation into fastai lib
```
import os
from pathlib import Path
HOME = Path(os.environ["HOME"])
DATA = HOME/'ucsi'
BS = 7
EPOCHS = 5
MODEL_NAME = "efficientnet-b5"
REMARK = "_"
THRESHOLD = 0.5
# cut to 400*4?
CUT_HORI = True
DICE_LOSS = True
FP16 = True
MODEL_PTH = "ssdd_%s_b%s%s.pth"%(MODEL_NAME,BS,REMARK)
print(MODEL_PTH)
```
Choices of model names
### VGG
vgg11, vgg13, vgg16, vgg19, vgg11bn, vgg13bn, vgg16bn, vgg19bn,
### Densenet
densenet121, densenet169, densenet201, densenet161, dpn68, dpn98, dpn131,
### Resnet
inceptionresnetv2,
resnet18, resnet34, resnet50, resnet101, resnet152,
resnext50_32x4d, resnext101_32x8d,
### SeNet
se_resnet50, se_resnet101, se_resnet152,
se_resnext50_32x4d, se_resnext101_32x4d,
senet154,
#### EfficientNet
efficientnet-b0, efficientnet-b1, efficientnet-b2, efficientnet-b3, efficientnet-b4, efficientnet-b5, efficientnet-b6, efficientnet-b7
```
import segmentation_models_pytorch as smp
import os
import cv2
import pdb
import time
#import warnings
import random
import numpy as np
import pandas as pd
from tqdm import tqdm_notebook as tqdm
from torch.optim.lr_scheduler import ReduceLROnPlateau
from sklearn.model_selection import train_test_split
import torch
import torch.nn as nn
from torch.nn import functional as F
import torch.optim as optim
import torch.backends.cudnn as cudnn
from torch.utils.data import DataLoader, Dataset, sampler
from matplotlib import pyplot as plt
from albumentations import (HorizontalFlip, RGBShift, ElasticTransform, GridDistortion,RandomBrightness ,ShiftScaleRotate, Normalize, Resize, Compose, OpticalDistortion,GaussNoise)
from albumentations.pytorch import ToTensor
#from onecyclelr import OneCycleLR
from sklearn.model_selection import train_test_split
from sklearn import model_selection
from fastai.vision import *
from ranger import Ranger
seed = 42
random.seed(seed)
os.environ["PYTHONHASHSEED"] = str(seed)
np.random.seed(seed)
torch.cuda.manual_seed(seed)
torch.backends.cudnn.deterministic = True
```
### RLE-Mask utility functions
```
def mask2rle(img):
'''
Numpy image to run length encoding
img: numpy array, 1 -> mask, 0 -> background
Returns run length as string formated
'''
pixels= img.T.flatten()
pixels = np.concatenate([[0], pixels, [0]])
runs = np.where(pixels[1:] != pixels[:-1])[0] + 1
runs[1::2] -= runs[::2]
return ' '.join(str(x) for x in runs)
def make_mask(row_id, df):
fname = df.iloc[row_id].name
labels = df.iloc[row_id][:4]
masks = np.zeros((1400, 2100, 4), dtype=np.float32) # float32 is V.Imp
# 4:class 1~4 (ch:0~3)
for idx, label in enumerate(labels.values):
if label is not np.nan:
label = label.split(" ")
positions = map(int, label[0::2])
length = map(int, label[1::2])
mask = np.zeros(1400 * 2100, dtype=np.uint8)
for pos, le in zip(positions, length):
mask[pos:(pos + le)] = 1
masks[:, :, idx] = mask.reshape(1400, 2100, order='F')
return fname, masks
```
### DataLoader
```
import fastai
#def cutHorizontal(x):
# return torch.cat(list(x[...,i*400:(i+1)*400] for i in range(4)), dim=0)
#def to416(x):
# size = list(x.size())
# size[-1]=416
# new = torch.zeros(size)
# new[...,8:-8] = x
# return new
class CloudDataset(Dataset):
def __init__(self, df, data_folder, mean, std, phase):
self.df = df
self.root = data_folder
self.mean = mean
self.std = std
self.phase = phase
self.aug_trans,self.x_trans, self.y_trans = get_transforms(phase, mean, std)
self.fnames = self.df.index.tolist()
def __getitem__(self, idx):
image_id, mask = make_mask(idx, self.df)
image_path = os.path.join(self.root, "train_images", image_id)
img = cv2.imread(image_path)
auged = self.aug_trans(image=img,mask = mask)
img = auged["image"]
mask = auged["mask"]
img = self.x_trans(image = img)
mask = self.y_trans(image = mask)
img = img["image"]
mask = mask["image"]
# mask = mask.permute(2, 0, 1)
return img, mask
def __len__(self):
return len(self.fnames)
def get_transforms(phase, mean, std):
list_transforms = []
if phase == "train":
auglist = [
HorizontalFlip(p=0.5),
ShiftScaleRotate(scale_limit=0.5, rotate_limit=0, shift_limit=0.1, p=0.5, border_mode=0),
GridDistortion(p=0.5),
OpticalDistortion(p=0.5, distort_limit=2, shift_limit=0.5),
]
else:
auglist = []
auglist.extend([
Normalize(mean=mean, std=std, p=1),
])
x_transforms=[Resize(640, 1280),ToTensor(),]
y_transforms=[Resize(320, 640),ToTensor(),]
return Compose(auglist),Compose(x_transforms),Compose(y_transforms)
return Compose(x_transforms), Compose(y_transforms)
def provider(
data_folder,
df_path,
mean=None,
std=None,
batch_size=BS,
num_workers=8,
):
'''Returns dataloader for the model training'''
df = pd.read_csv(df_path)
# https://www.kaggle.com/amanooo/defect-detection-starter-u-net
df['ImageId'], df['ClassId'] = zip(*df['Image_Label'].str.split('_'))
#df['ClassId'] = df['ClassId'].astype(int)
df = df.pivot(index='ImageId',columns='ClassId',values='EncodedPixels')
df['defects'] = df.count(axis=1)
train_df, val_df = train_test_split(df, test_size=0.02, stratify=df["defects"], random_state=69)
# df = train_df if phase == "train" else val_df
train_ds = CloudDataset(train_df, data_folder, mean, std, "train")
val_ds =CloudDataset(val_df, data_folder, mean,std, "val")
databunch = DataBunch.create(train_ds,val_ds,path = data_folder,bs=BS,num_workers=num_workers)
return databunch
data = provider(
data_folder=DATA,
df_path=DATA/"train.csv",
mean=(0.485, 0.456, 0.406),
std=(0.229, 0.224, 0.225),
batch_size=BS,
num_workers=4,
)
x,y = data.one_batch()
x.size(),y.size()
plt.imshow(x[1,0],)
plt.imshow(y[0,0])
```
### Utility function
#### Dice and IoU metric implementations, metric logger for training and validation.
```
def metricSeg(probability, truth, threshold=THRESHOLD, reduction='none'):
'''Calculates dice of positive and negative images seperately'''
'''probability and truth must be torch tensors'''
# probability = torch.sigmoid(probability)
batch_size = len(truth)
with torch.no_grad():
probability = probability.view(batch_size, -1)
truth = truth.view(batch_size, -1)
assert(probability.shape == truth.shape)
p = (probability > threshold).float()
t = (truth > threshold).float()
t_sum = t.sum(-1)
p_sum = p.sum(-1)
neg_index = torch.nonzero(t_sum == 0)
pos_index = torch.nonzero(t_sum >= 1)
dice_neg = (p_sum == 0).float()
dice_pos = 2 * (p*t).sum(-1)/((p+t).sum(-1))
dice_neg = dice_neg[neg_index]
dice_pos = dice_pos[pos_index]
dice = torch.cat([dice_pos, dice_neg])
dice_neg = np.nan_to_num(dice_neg.mean().item(), 0)
dice_pos = np.nan_to_num(dice_pos.mean().item(), 0)
dice = dice.mean().item()
num_neg = len(neg_index)
num_pos = len(pos_index)
return dice, dice_neg, dice_pos, num_neg, num_pos
class dices(LearnerCallback):
_order = -20 # Needs to run before the recorder, very CRITICAL step
def __init__(self,learn):
super().__init__(learn)
def on_train_begin(self,**kwargs):
self.learn.recorder.add_metric_names(['dice','dice_neg','dice_pos','num_neg','num_pos'])
def on_epoch_begin(self,**kwargs):
self.ttl_dice = []
self.ttl_dice_neg = []
self.ttl_dice_pos = []
self.ttl_num_neg = []
self.ttl_num_pos = []
self.ttl_lists = [self.ttl_dice,self.ttl_dice_neg,self.ttl_dice_pos,self.ttl_num_neg,self.ttl_num_pos]
def on_batch_end(self,last_output,last_target,**kwargs):
dice, dice_neg, dice_pos, num_neg, num_pos = metricSeg(last_output,last_target)
self.ttl_dice.append(dice)
self.ttl_dice_neg.append(dice_neg)
self.ttl_dice_pos.append(dice_pos)
self.ttl_num_neg.append(num_neg)
self.ttl_num_pos.append(num_pos)
def on_epoch_end(self,last_metrics,**kwargs):
extras = [sum(i)/float(len(i)) for i in self.ttl_lists]
return add_metrics(last_metrics, extras)
```
### Define Model
```
seg_model = smp.FPN(MODEL_NAME, encoder_weights="imagenet", classes=4, activation="sigmoid")
class majorModel(nn.Module):
def __init__(self, seg_model):
super().__init__()
self.seq = nn.Sequential(*[
nn.Conv2d(3,12,kernel_size=(3,3), padding=1, stride=1, ),
nn.ReLU(),
nn.Conv2d(12,3,kernel_size=(3,3), padding=1, stride=2),
nn.ReLU(),
seg_model,])
def forward(self,x):
return self.seq(x)
model = majorModel(seg_model).cuda()
optar = partial(Ranger)
```
### Fastai Learner
```
learn = Learner(data, model, metrics = [], opt_func=optar,
loss_func = smp.utils.losses.BCEDiceLoss(eps=1.) if DICE_LOSS else nn.BCEWithLogitsLoss(),
callback_fns = dices).to_fp16()
learn.path = Path("./fastai")
#learn.lr_find()
#learn.recorder.plot()
from fastai.callbacks import SaveModelCallback
lr = 1e-3
learn.unfreeze()
# learn.fit_one_cycle(5,lr, callbacks=[SaveModelCallback(learn, every='epoch', monitor='loss')])
#learn.fit_one_cycle(6, max_lr=slice(1e-6, 1e-4))
learn.fit_one_cycle(EPOCHS,(lr/2),wd=1e-3, callbacks=[
# dices(learn),
# cutHorizonCallback(),
SaveModelCallback(learn, every='epoch', monitor='loss')])
```
### Predict
```
import gc
torch.cuda.empty_cache()
gc.collect()
test_dataset = CloudDataset(df=sub, datatype='test', img_ids=test_ids, transforms = get_validation_augmentation(), preprocessing=get_preprocessing(preprocessing_fn))
test_loader = DataLoader(test_dataset, batch_size=8, shuffle=False, num_workers=0)
loaders = {"test": test_loader}
encoded_pixels = []
image_id = 0
for i, test_batch in enumerate(tqdm.tqdm(loaders['test'])):
runner_out = runner.predict_batch({"features": test_batch[0].cuda()})['logits']
for i, batch in enumerate(runner_out):
for probability in batch:
probability = probability.cpu().detach().numpy()
if probability.shape != (350, 525):
probability = cv2.resize(probability, dsize=(525, 350), interpolation=cv2.INTER_LINEAR)
predict, num_predict = post_process(sigmoid(probability), class_params[image_id % 4][0], class_params[image_id % 4][1])
if num_predict == 0:
encoded_pixels.append('')
else:
r = mask2rle(predict)
encoded_pixels.append(r)
image_id += 1
sub['EncodedPixels'] = encoded_pixels
sub.to_csv('submission.csv', columns=['Image_Label', 'EncodedPixels'], index=False)
```
| github_jupyter |
### Notes:
* python script stores result of the simulation in a predetermined location
* jupyter notebook shows visualizations of the results from solidity implementation and cadCAD model and error metrics
```
from pathlib import Path
import os
import csv
import sys
import datetime as dt
import json
from enum import Enum
import pandas as pd
import numpy as np
%matplotlib inline
sys.path.append('./models')
path = Path().resolve()
root_path = str(path).split('notebooks')[0]
os.chdir(root_path)
with open('./tests/data/historic_market_prices.csv', newline='') as f:
reader = csv.reader(f)
market_prices = list(reader)[0]
market_prices = [float(price) for price in market_prices]
#python stores a file containing Kp, Ki and a time series of market prices
#(an array of tuples, or two arrays, containing a timestamp and the price,
#or the time elapsed and the price change - we can go with whatever is easier to implement on the JS side)
#instead of file name being hardcoded, we could have it stored in an environment
#variable that both python and JS read from
Kp = 5e-7
Ki = -1e-7
per_second_leak = 1000000000000000000000000000
control_period = 3600
length = len(market_prices)
delta_t = [control_period] * length
SIMULATION_TIMESTEPS = length - 1
save_dir = "../truffle/test/saved_sims/pi_second/raw/custom-config-sim.txt"
config_file = {
"Kp": str(int(Kp * 1e18)),
"Ki": str(int(Ki * 1e18 / control_period)),
"noise_barrier": "1000000000000000000",
"per_second_leak": str(int(per_second_leak)),
"oracle_initial_price": "2000000000000000000",
"initial_redemption_price": "2000000000000000000000000000",
"delta_t": [str(i) for i in delta_t],
"market_prices": [str(int(i*1e18)) for i in market_prices],
"save_dir": save_dir
}
with open('./cross-model/truffle/test/config/pi_second_raw.json', 'w') as fp:
json.dump(config_file, fp)
config_file = {
"alpha": per_second_leak,
"kp": Kp,
"ki": lambda control_period=control_period: Ki/control_period,
"delta_t": delta_t,
"market_prices": market_prices,
}
market_prices[0:5]
# Compile, test, and deploy contracts to Ganache
# !npm run pi-raw-second-imported-config --prefix './cross-model/truffle'
os.chdir('./cross-model/truffle')
!npm run install-deps
!npm run pi-raw-second-imported-config
os.chdir('../../')
```
[redemptionRate](https://github.com/reflexer-labs/geb-rrfm-truffle-sims/blob/master/contracts/mock/MockOracleRelayer.sol#L14) = $1+d_t$ (units: per second)
$p_{t+\Delta{t}} = p_{t} (1+d_t)^{\Delta{t}}$ where $\Delta{t}$ is measured in seconds
Meaning:
* $d_t > 0$ implies $redemptionRate > 1$
* $d_t < 0$ implies $redemptionRate < 1$
In the results dataset this is the PerSecondRedemptionRate column
```
from models.system_model_v1.model.state_variables.init import state_variables
def pre_process_initial_conditions(genesis_dict):
base_genesis_states = genesis_dict.copy()
for k in base_genesis_states.keys():
if k in config_file:
try:
base_genesis_states[k] = config_file[k][0]
except:
base_genesis_states[k] = config_file[k]
return base_genesis_states
test_specific_initial_conditions = pre_process_initial_conditions(state_variables)
from models.system_model_v1.model.params.init import params
def pre_process_params_dict(params_dict):
base_params_dict = params_dict.copy()
for k in config_file.keys():
if type(config_file[k]) == list:
base_params_dict[k] = [config_file[k]]
else:
try:
base_params_dict[k] = [config_file[k]]
except OverflowError:
print(config_file[k])
raise
return base_params_dict
test_specific_params = pre_process_params_dict(params)
def test_specific_resolve_time_passed(params, substep, state_history, state):
index = state['timestep']+1
value = params['delta_t'][index]
return {'seconds_passed': value}
def test_specific_update_market_price(params, substep, state_history, state, policy_input):
key = 'market_price'
index = state['timestep']
value = params['market_prices'][index]
return key, value
from models.system_model_v1.model.parts.markets import resolve_time_passed, update_market_price
from models.system_model_v1.model.partial_state_update_blocks import partial_state_update_blocks
for psub in partial_state_update_blocks:
for psub_part in psub.values():
if type(psub_part)==dict:
for k,v in psub_part.items():
if v == resolve_time_passed:
psub_part[k] = test_specific_resolve_time_passed
if v == update_market_price:
psub_part[k] = test_specific_update_market_price
from cadCAD.configuration.utils import config_sim
from cadCAD.configuration import Experiment
from cadCAD import configs
del configs[:]
exp = Experiment()
sim_config_dict = {
'T': range(SIMULATION_TIMESTEPS),
'N': 1,
'M': test_specific_params
}
c = config_sim(sim_config_dict)
exp.append_configs(
initial_state=test_specific_initial_conditions, #dict containing variable names and initial values
partial_state_update_blocks=partial_state_update_blocks, #dict containing state update functions
sim_configs=c #preprocessed dictionaries containing simulation parameters
)
from cadCAD.engine import ExecutionMode, ExecutionContext, Executor
exec_mode = ExecutionMode()
local_mode_ctx = ExecutionContext(exec_mode.local_mode)
executor = Executor(local_mode_ctx, configs) # Pass the configuration object inside an array
raw_result, tensor, sessions = executor.execute() # The `execute()` method returns a tuple; its first elements contains the raw results
full = pd.DataFrame(raw_result)
max_substep = max(full.substep)
df = full.copy()
df = df[(df.substep==max_substep) | (df.substep==0)]
save_dir = "./cross-model/truffle/test/saved_sims/pi_second/raw/custom-config-sim.txt"
data = pd.read_csv(save_dir, sep=" ", header=None, skiprows=1)
data.columns = [x.replace(' ','') for x in pd.read_csv(save_dir, sep="|", nrows=1).columns]
for c in data.columns:
data[c]=data[c].apply(float)
#define normalizing constants
WAD = 10**18
RAY = 10**27
RAD = 10**45
#normalize data to "USD" dimension
data['MarketPrice(USD)'] = data['MarketPrice(WAD)']/WAD
data['RedemptionPrice(USD)'] = data['RedemptionPrice(RAY)']/RAY
data['PerSecondRedemptionRate(float)'] = data['PerSecondRedemptionRate(RAY)']/RAY - 1
data['time'] = data['DelaySinceLastUpdate'].cumsum()
data['timestamp'] = state_variables['timestamp'] + data['time'].apply(lambda x: dt.timedelta(seconds=x))
df
data
combined = df.merge(data, how='inner', on=['timestamp'])
combined = combined.iloc[25:]
combined
import matplotlib.pyplot as plt
fig, axs = plt.subplots(2, 2, figsize=(15,10))
i = 0
axs.flat[i].plot(combined['timestamp'], combined['market_price'], label='Market')
axs.flat[i].plot(combined['timestamp'], combined['target_price'], label='Target')
axs.flat[i].set_ylabel('Python')
axs.flat[i].set_title('Market and Target Price')
axs.flat[i].legend()
i += 1
axs.flat[i].plot(combined['timestamp'], combined['target_rate'])
axs.flat[i].set_title('Per Second Redemption Rate')
i += 1
axs.flat[i].plot(combined['timestamp'], combined['MarketPrice(USD)'], label='Market')
axs.flat[i].plot(combined['timestamp'], combined['RedemptionPrice(USD)'], label='Target')
axs.flat[i].legend()
axs.flat[i].set_ylabel('Solidity')
i += 1
axs.flat[i].plot(combined['timestamp'], combined['PerSecondRedemptionRate(float)'])
fig.tight_layout()
plt.show()
fig.savefig('plots/solidity-cadcad/solidity-cadcad-market.png')
combined['market_price_error'] = combined['market_price'] - combined['MarketPrice(USD)']
combined['market_price_error'].plot(legend=True)
combined['market_price_cumulative_error'] = combined['market_price_error'].cumsum()
combined['market_price_cumulative_error'].plot(legend=True)
combined['target_price_error'] = combined['target_price'] - combined['RedemptionPrice(USD)']
combined['target_price_error'].plot(legend=True)
combined['target_price_cumulative_error'] = combined['target_price_error'].cumsum()
combined['target_price_cumulative_error'].plot(legend=True)
plt.savefig('plots/solidity-cadcad/solidity-cadcad-error.png')
print(f'''
Max error: {combined['target_price_error'].max()}
Std. error: {combined['target_price_error'].std()}
Mean error: {combined['target_price_error'].mean()}
Abs. mean error: {combined['target_price_error'].abs().mean()}
''')
combined['target_price_cumulative_error'].describe()
```
| github_jupyter |
```
import numpy as np
import scipy.sparse as sp
import scipy.io as spio
import matplotlib.pyplot as plt
import pandas as pd
from scipy.stats import pearsonr
#df = pd.read_csv('../simple/processed_data/noacut/final/simple_noacut_final.csv', sep=',')
#cuts = spio.loadmat('../simple/processed_data/noacut/final/simple_noacut_final_cuts')['cuts']
#keep_index = np.nonzero((df.total_count >= 20) & ((df.seq.str.slice(50+20, 56+20) == 'AATAAA') | (df.seq.str.slice(50+20, 56+20) == 'ATTAAA')))[0]
df = pd.read_csv('../doubledope/processed_data/score_60/final/doubledope_score_60_final.csv', sep=',')
cuts = spio.loadmat('../doubledope/processed_data/score_60/final/doubledope_score_60_final_cuts')['cuts']
keep_index = np.nonzero(df.total_count >= 10)[0]
df = df.iloc[keep_index].copy()
cuts = cuts[keep_index, :]
print(len(df))
proximal_usage = (np.sum(cuts[:, 57+20:87+20], axis=1) + 0.1) / (np.sum(cuts, axis=1) + 2. * 0.1)
cuts = None
df['proximal_usage'] = proximal_usage
df['seq'] = df['seq'].str.slice(20, 186+20)
#Generate feature matrix
mer4_dict = {}
mer4_list = []
mer6_dict = {}
mer6_list = []
bases = list('ACGT')
#Build dictionary of 6-mer -> index
i6 = 0
i4 = 0
for b1 in bases :
for b2 in bases :
for b3 in bases :
for b4 in bases :
mer4_dict[b1 + b2 + b3 + b4] = i4
mer4_list.append(b1 + b2 + b3 + b4)
i4 += 1
for b5 in bases :
for b6 in bases :
mer6_dict[b1 + b2 + b3 + b4 + b5 + b6] = i6
mer6_list.append(b1 + b2 + b3 + b4 + b5 + b6)
i6 += 1
#Loop over dataframe, fill matrix X with 6-mer counts
X = sp.lil_matrix((len(df), 256 * 50 + 4096 + 256 * 40 + 4096))
use_regions = list(df.seq.str.slice(0, 50 + 4 - 1).values)
pas_regions = list(df.seq.str.slice(50, 56).values)
dse_regions = list(df.seq.str.slice(56, 96 + 4 - 1).values)
fdse_regions = list(df.seq.str.slice(96, 186).values)
i = 0
for _, row in df.iterrows() :
if i % 50000 == 0 :
print('Extracting features from sequence ' + str(i))
use_region = use_regions[i]
pas_region = pas_regions[i]
dse_region = dse_regions[i]
fdse_region = fdse_regions[i]
#USE
for j in range(0, len(use_region) - 4 + 1) :
if use_region[j:j+4] in mer4_dict :
X[i, j * 256 + mer4_dict[use_region[j:j+4]]] = 1.
#PAS
for j in range(0, len(pas_region) - 6 + 1) :
if pas_region[j:j+6] in mer6_dict :
X[i, 50 * 256 + mer6_dict[pas_region[j:j+6]]] = 1.
#DSE
for j in range(0, len(dse_region) - 4 + 1) :
if dse_region[j:j+4] in mer4_dict :
X[i, 50 * 256 + 4096 + j * 256 + mer4_dict[dse_region[j:j+4]]] = 1.
#FDSE
for j in range(0, len(fdse_region) - 6 + 1) :
if fdse_region[j:j+6] in mer6_dict :
X[i, 50 * 256 + 4096 + 40 * 256 + mer6_dict[fdse_region[j:j+6]]] += 1.
i += 1
X = sp.csr_matrix(X)
y = np.ravel(df['proximal_usage'].values)
print('Shape of X = ' + str(X.shape))
print('Shape of y = ' + str(y.shape))
test_size = 20000
X_train = X[:X.shape[0] - test_size, :]
y_train = y[:y.shape[0] - test_size]
X_test = X[-test_size:, :]
y_test = y[-test_size:]
print(X_train.shape)
print(X_test.shape)
#Helper function for computing log(x / y) in a safe way (whenever x or y is 0).
def safe_kl_log(num, denom) :
log_vec = np.zeros(num.shape)
log_vec[(num > 0) & (denom > 0)] = np.log(num[(num > 0) & (denom > 0)] / denom[(num > 0) & (denom > 0)])
return log_vec
def get_y_pred(X, w, w_0) :
score = X.dot(w) + w_0
return 1. / (1. + np.exp(-score))
def get_kl_div_loss(X, w, w_0, y_true, alpha=0.0) :
y_pred = get_y_pred(X, w, w_0)
kl = y_true * safe_kl_log(y_true, y_pred) + (1. - y_true) * safe_kl_log((1. - y_true), (1. - y_pred)) + (1./2.) * alpha * np.dot(w, w)
return np.sum(kl)
def get_kl_div_gradients(X, w, w_0, y_true, alpha=0.0) :
N = float(X.shape[0])
y_pred = get_y_pred(X, w, w_0)
kl_grad_w = (1. / N) * X.T.dot(y_pred - y_true) + alpha * w
kl_grad_w_0 = (1. / N) * np.sum(y_pred - y_true)
return kl_grad_w, kl_grad_w_0
def sgd(X, y, X_test, y_test, w, w_0, step_size=0.1, batch_size=64, alpha=0.0, n_epochs=10) :
N = float(X.shape[0])
N_test = float(X_test.shape[0])
n_batches = int(X.shape[0] / batch_size)
mean_train_losses = [get_kl_div_loss(X, w, w_0, y, alpha) / N]
mean_test_losses = [get_kl_div_loss(X_test, w, w_0, y_test, alpha) / N_test]
for epoch in range(n_epochs) :
if epoch % 10 == 0 :
print('Training epoch = ' + str(epoch))
print('Training set KL-div = ' + str(round(mean_train_losses[-1], 4)))
print('Test set KL-div = ' + str(round(mean_test_losses[-1], 4)))
total_loss = 0
for batch in range(n_batches) :
X_batch = X[batch * batch_size:(batch+1) * batch_size, :]
y_batch = y[batch * batch_size:(batch+1) * batch_size]
#Compute gradients
grad_w, grad_w_0 = get_kl_div_gradients(X_batch, w, w_0, y_batch, alpha)
#Update weights with small step in opposite direction of Loss gradient
w, w_0 = w - step_size * grad_w, w_0 - step_size * grad_w_0
mean_train_losses.append(get_kl_div_loss(X, w, w_0, y, alpha) / N)
mean_test_losses.append(get_kl_div_loss(X_test, w, w_0, y_test, alpha) / N_test)
if len(mean_train_losses) >= 2 :
if mean_train_losses[-2] / mean_train_losses[-1] <= 1.00001 :
break
print('Gradient descent completed.')
print('Final training set KL-div = ' + str(round(mean_train_losses[-1], 4)))
print('Final test set KL-div = ' + str(round(mean_test_losses[-1], 4)))
return w, w_0, mean_train_losses, mean_test_losses
w, w_0 = np.zeros(X.shape[1]), 0
#Train model with SGD
n_epochs = 20
alpha = 0.0#0.0001
step_size = 0.1
batch_size = 256#X_train.shape[0]
w, w_0, train_losses, test_losses = sgd(X_train, y_train, X_test, y_test, w, w_0, step_size=step_size, batch_size=batch_size, alpha=alpha, n_epochs=n_epochs)
f = plt.figure(figsize=(8, 6))
l1, = plt.plot(np.arange(len(train_losses)), train_losses, linewidth=3, color='red', label='Train loss')
l2, = plt.plot(np.arange(len(train_losses)), test_losses, linewidth=3, color='green', linestyle='--', label='Test loss')
plt.xlabel('Training epoch', fontsize=16)
plt.ylabel('Mean KL-div', fontsize=16)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.xlim(0, len(train_losses))
plt.legend(handles=[l1, l2], fontsize=16)
plt.tight_layout()
plt.show()
#TODO: Scatter plot of true vs. pred SD1 usage on test set, and print R^2 coefficient.
y_pred_test = get_y_pred(X_test, w, w_0)
r_val, _ = pearsonr(y_pred_test, y_test)
f = plt.figure(figsize=(6, 6))
plt.scatter(y_pred_test, y_test, s=5, alpha=0.25, c='black')
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.xlabel('Predicted SD1 Usage', fontsize=16)
plt.ylabel('Observed SD1 Usage', fontsize=16)
plt.title('R^2 = ' + str(round(r_val * r_val, 2)))
plt.tight_layout()
plt.show()
#Store trained weights
#np.save('simple_position_nmer_weights', w)
np.save('doubledope_position_nmer_weights', w)
```
| github_jupyter |
```
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import style
from matplotlib import cm
import seaborn as sns
sns.set(style="ticks")
from scipy.optimize import curve_fit
import os
import numpy as np
import glob
import scipy as sc
plt.rcParams['font.sans-serif'] = "Arial"
plt.rcParams['font.family'] = "sans-serif"
from scipy import stats
def linearRegression(df):
x = df[df.DiscoveryTime.notnull()].trialNum.values
y = df[df.DiscoveryTime.notnull()].DiscoveryTime.values
slope, intercept, r_value, p_value, std_err = sc.stats.linregress(x, y)
return(slope, intercept, r_value, p_value, std_err)
# Define exponential function func,
# where a = alpha, b = Tau, c = y intercept
def fit_to_exponential(x, a, b, c):
return a * np.exp(-(x / b)) + c
```
# Individual floral shape
```
direc = r'./dataFolders/Output/Step5_v4'
outpath = r'./dataFolders/Output/Step6_v5'
outpath_fig = r'./dataFolders/Output/Step6_v5/Figure'
```
Save the all shape specific files as a single file
```
Lightlevel = ['L0.1', 'L50']
for Lightlevel in Lightlevel:
Light_specific_file = glob.glob(direc + "\\" + Lightlevel + '*Raw*.csv')
# get the data together and save as single dataset
df1=[]
df2=[]
df3=[]
df4=[]
df5=[]
df6=[]
ctr = 0
for fpath in Light_specific_file:
df = pd.read_csv(fpath)
DiscoveryTime = df['DiscoveryTime']
trialNum= df.index
# fname = os.path.basename(fpath)[:-30]
a,b, c, _ = os.path.basename(fpath).split("_")
fname = a + "_" + b + "_" + c
names = [fname]*len(trialNum)
mothIn = df['In_Frame']
mothOut = df['Out_Frame']
Proboscis = df['ProboscisDetect']
df1.extend(DiscoveryTime)
df2.extend(trialNum)
df3.extend(names)
df4.extend(mothIn)
df5.extend(mothOut)
df6.extend(Proboscis)
ctr+=1
print(ctr)
new_df = (pd.DataFrame({'In_Frame' : df4, 'Out_Frame' : df5, 'Proboscis' : df6,
'DiscoveryTime': df1, 'trialNum': df2, 'name' : df3}))
print(Lightlevel + '\t' + "has a total of" + '\t'+ str(ctr) + '\t' + "moths")
new_df.to_csv(outpath + "\\" + Lightlevel + "Allmoths.csv")
dFrame = pd.DataFrame()
file = glob.glob(outpath + "\\" + '*Allmoths.csv')
for f in file:
print(f)
df = pd.read_csv(f)
dFrame = dFrame.append(df)
dFrame.to_csv(outpath + "\\" + "AllLight_EveryMoth.csv")
```
Fit to curve and plot
```
f, ax = plt.subplots(1,2, figsize = (20,10), sharex = True, sharey = True)
for idx, f in enumerate(file):
df = pd.read_csv(f)
names = df.name.unique()
n = len(names)
color = cm.tab20b(np.linspace(0,1,n))
for name, c in zip(names, color):
x = dFrame[(dFrame.name == name) & (dFrame.DiscoveryTime.notnull())].trialNum
y = dFrame[(dFrame.name == name) & (dFrame.DiscoveryTime.notnull())].DiscoveryTime
ax[idx].plot(x, y/100, 'o-', markersize = 5, label = name, color = c)
ax[idx].set_ylabel('Discovery Time')
ax[idx].set_xlabel('Visit Number')
ax[0].set_title('L50')
ax[1].set_title('L0.1')
ax[0].legend()
ax[1].legend()
plt.savefig(outpath_fig + 'individualMoth_learningCurve.png')
# do the fit for three different y0 - computed across different lengths of the saturation - curve
Lightlevel = ['L0.1','L50']
fig1, ax0 = plt.subplots(figsize = (10,6))
fig2, ax1 = plt.subplots(figsize = (10,6))
for Lightlevel in Lightlevel:
file = glob.glob(outpath + "\\" + Lightlevel + 'Allmoths.csv')
print(file)
new_df = pd.read_csv(file[0])
xdata = new_df.trialNum.values
ydata = new_df.DiscoveryTime.values
xdata_notnan = new_df[new_df.DiscoveryTime.notnull()].trialNum.values
ydata_notnan = new_df[new_df.DiscoveryTime.notnull()].DiscoveryTime.values
numMoth = len(new_df.name.unique())
length_to_estimate_y0 = [2,4,8]
# initialize all the variables you want to save
slope = []
std_err = []
p_value = []
r_value = []
y_intercept = []
y_intercept_std_dev = []
average_firstTrial = []
alpha = []
tau= []
covariance = []
for length in length_to_estimate_y0:
totalTrial = max(new_df.trialNum)
part_of_df = new_df[new_df.trialNum >= (totalTrial * (1-1/length))]
x = part_of_df[part_of_df.DiscoveryTime.notnull()].trialNum.values
y = part_of_df[part_of_df.DiscoveryTime.notnull()].DiscoveryTime.values
m, y0, r_val, p_val, se = linearRegression(part_of_df)
mean_y0 = part_of_df.DiscoveryTime.mean()
std_y0 = part_of_df.DiscoveryTime.std()
mean_a0andy0 = new_df[new_df.trialNum==0].DiscoveryTime.mean()
a0 = mean_a0andy0 - mean_y0
# plot the regression line
x = part_of_df.trialNum
y = part_of_df.DiscoveryTime
ax0.plot(x,y*1/100, 'ob', markersize=5, alpha=.5, label = 'Raw data')
test_x = range(min(x)-5, max(x)+5)
ax0.plot(test_x, (m*test_x + y0)*1/100, label = 'Fitted line')
ax0.fill_between(test_x, ((m-se)*test_x + y0)*1/100, ((m+se)*test_x + y0)*1/100, alpha=0.2)
ax0.set_title(Lightlevel + '_1/' + str(length) + '_lengthData')
ax0.legend()
fig1.savefig(outpath_fig + "\\" + Lightlevel + '_1-' + str(length) + '_LinearRegression.png')
ax0.cla()
slope.append(m/100)
std_err.append(se/100)
p_value.append(p_val)
r_value.append(r_val)
y_intercept.append(mean_y0/100)
y_intercept_std_dev.append(std_y0/100)
average_firstTrial.append(mean_a0andy0/100)
alpha.append(a0/100)
popt,pcov = curve_fit(lambda x, b: fit_to_exponential(x, a0, b, mean_y0), xdata_notnan, ydata_notnan)
tau.append(popt[0]/100)
covariance.append(pcov.flatten()[0])
# plot the final curve fit
max_trial=max(xdata)
ax1.plot(xdata, ydata*1/100, 'or', markersize=7, alpha=.5, label = 'Raw data')
ax1.plot(range(0,max_trial), fit_to_exponential(range(0,max_trial), a0, *popt, mean_y0)*1/100,
'g--', linewidth=2,
label = 'fit: alpha= %5.3f , Tau= %5.3f, y0= %5.3f' %(a0/100, popt/100, mean_y0/100))
ax1.set_title(Lightlevel + '_1-' + str(length) + '_length', y=1.2)
ax1.set_xlabel('Trial number')
ax1.set_ylabel('Exploration Time (second)')
leg = plt.legend()
leg.get_frame().set_linewidth(1.5)
ax1.legend(bbox_to_anchor=(0., 1.05, 1., .102), loc=3, mode="expand", borderaxespad=0.)
ax1.set_xlim([-5,50])
ax1.set_ylim([-5,70])
ax1.text(60, 40, 'N= ' + str(numMoth))
fig2.tight_layout()
fig2.savefig(outpath_fig + "\\" + 'LearningCurve_' + Lightlevel + '_1-' + str(length) + '_length.png')
ax1.cla()
all_param_df = pd.DataFrame({'slope':slope, 'std_err':std_err,
'p_value':p_value, 'r_value':r_value,
'y_intercept':y_intercept,
'y_intercept_std' : y_intercept_std_dev,
'average_firstTrial':average_firstTrial,
'alpha':alpha,'tau':tau,
'covariance':covariance},
index = ['one-half', 'last fourth', 'last eight'])
all_param_df.to_csv(outpath + "\\" + Lightlevel + '_FittedParameters.csv')
```
## Without Outliers
```
L01Data = pd.read_csv(outpath + "\\L0.1Allmoths.csv")
L01Data = L01Data[L01Data.DiscoveryTime.notnull()]
z = np.abs(stats.zscore(L01Data.DiscoveryTime.values))
outlier_ID = z > 4
plt.plot(z, 'o')
L01Data['zscore'] = z
L01Data['outlier_ID'] = outlier_ID
L01Data['condition'] = ['Low']*len(z)
notoutliers_01 = L01Data[L01Data.outlier_ID == False]
notoutliers_01.to_csv(outpath + "\\L0.1Allmoths_notoutliers.csv")
L50Data = pd.read_csv(outpath + "\\L50Allmoths.csv")
L50Data = L50Data[L50Data.DiscoveryTime.notnull()]
z = np.abs(stats.zscore(L50Data.DiscoveryTime.values))
outlier_ID = z > 4
plt.plot(z, 'o')
L50Data['zscore'] = z
L50Data['outlier_ID'] = outlier_ID
L50Data['condition'] = ['High']*len(z)
notoutliers_50 = L50Data[L50Data.outlier_ID == False]
notoutliers_50.to_csv(outpath + "\\L50Allmoths_notoutliers.csv")
plt.plot(L01Data.DiscoveryTime[L01Data.outlier_ID == True], 'o', color = 'red')
plt.plot(L01Data.DiscoveryTime[L01Data.outlier_ID == False], 'o', color = 'blue')
notoutliers = L01Data[L01Data.outlier_ID == False]
plt.plot(L50Data.DiscoveryTime[L50Data.outlier_ID == True], 'o', color = 'red')
plt.plot(L50Data.DiscoveryTime[L50Data.outlier_ID == False], 'o', color = 'blue')
notoutliers = L50Data[L50Data.outlier_ID == False]
dFrame = pd.DataFrame()
file = glob.glob(outpath + "\\" + '*Allmoths_notOutliers.csv')
for f in file:
print(f)
df = pd.read_csv(f)
dFrame = dFrame.append(df)
dFrame.to_csv(outpath + "\\" + "AllLight_EveryMoth_notOutliers.csv")
# do the fit for three different y0 - computed across different lengths of the saturation - curve
Lightlevel = ['L0.1','L50']
fig1, ax0 = plt.subplots(figsize = (10,6))
fig2, ax1 = plt.subplots(figsize = (10,6))
for Lightlevel in Lightlevel:
file = glob.glob(outpath + "\\" + Lightlevel + 'Allmoths_notoutliers.csv')
print(file)
new_df = pd.read_csv(file[0])
xdata = new_df.trialNum.values
ydata = new_df.DiscoveryTime.values
xdata_notnan = new_df[new_df.DiscoveryTime.notnull()].trialNum.values
ydata_notnan = new_df[new_df.DiscoveryTime.notnull()].DiscoveryTime.values
numMoth = len(new_df.name.unique())
length_to_estimate_y0 = [2,4,8]
# initialize all the variables you want to save
slope = []
std_err = []
p_value = []
r_value = []
y_intercept = []
y_intercept_std_dev = []
average_firstTrial = []
alpha = []
tau= []
covariance = []
for length in length_to_estimate_y0:
totalTrial = max(new_df.trialNum)
part_of_df = new_df[new_df.trialNum >= (totalTrial * (1-1/length))]
x = part_of_df[part_of_df.DiscoveryTime.notnull()].trialNum.values
y = part_of_df[part_of_df.DiscoveryTime.notnull()].DiscoveryTime.values
m, y0, r_val, p_val, se = linearRegression(part_of_df)
mean_y0 = part_of_df.DiscoveryTime.mean()
std_y0 = part_of_df.DiscoveryTime.std()
mean_a0andy0 = new_df[new_df.trialNum==0].DiscoveryTime.mean()
a0 = mean_a0andy0 - mean_y0
# plot the regression line
x = part_of_df.trialNum
y = part_of_df.DiscoveryTime
ax0.plot(x,y*1/100, 'ob', markersize=5, alpha=.5, label = 'Raw data')
test_x = range(min(x)-5, max(x)+5)
ax0.plot(test_x, (m*test_x + y0)*1/100, label = 'Fitted line')
ax0.fill_between(test_x, ((m-se)*test_x + y0)*1/100, ((m+se)*test_x + y0)*1/100, alpha=0.2)
ax0.set_title(Lightlevel + '_1/' + str(length) + '_lengthData')
ax0.legend()
fig1.savefig(outpath_fig + "//"+ Lightlevel + '_1-' + str(length) + '_LinearRegression_NoOutlier.png')
ax0.cla()
slope.append(m/100)
std_err.append(se/100)
p_value.append(p_val)
r_value.append(r_val)
y_intercept.append(mean_y0/100)
y_intercept_std_dev.append(std_y0/100)
average_firstTrial.append(mean_a0andy0/100)
alpha.append(a0/100)
popt,pcov = curve_fit(lambda x, b: fit_to_exponential(x, a0, b, mean_y0), xdata_notnan, ydata_notnan)
tau.append(popt[0]/100)
covariance.append(pcov.flatten()[0])
# plot the final curve fit
max_trial=max(xdata)
ax1.plot(xdata, ydata*1/100, 'or', markersize=7, alpha=.5, label = 'Raw data')
ax1.plot(range(0,max_trial), fit_to_exponential(range(0,max_trial), a0, *popt, mean_y0)*1/100,
'g--', linewidth=2,
label = 'fit: alpha= %5.3f , Tau= %5.3f, y0= %5.3f' %(a0/100, popt/100, mean_y0/100))
ax1.set_title(Lightlevel + '_1-' + str(length) + '_length', y=1.2)
ax1.set_xlabel('Trial number')
ax1.set_ylabel('Exploration Time (second)')
leg = plt.legend()
leg.get_frame().set_linewidth(1.5)
ax1.legend(bbox_to_anchor=(0., 1.05, 1., .102), loc=3, mode="expand", borderaxespad=0.)
ax1.set_xlim([-5,90])
ax1.set_ylim([-5,55])
ax1.text(60, 40, 'N= ' + str(numMoth))
fig2.tight_layout()
fig2.savefig(outpath_fig + "\\" + 'LearningCurve_' + Lightlevel + '_1-' + str(length) + '_length_NotOutlier.png')
ax1.cla()
all_param_df = pd.DataFrame({'slope':slope, 'std_err':std_err,
'p_value':p_value, 'r_value':r_value,
'y_intercept':y_intercept,
'y_intercept_std' : y_intercept_std_dev,
'average_firstTrial':average_firstTrial,
'alpha':alpha,'tau':tau,
'covariance':covariance},
index = ['one-half', 'last fourth', 'last eight'])
all_param_df.to_csv(outpath + "\\" + Lightlevel + '_FittedParameters_notOutliers.csv')
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import io
import requests
```
## 1) 載入資料集
```
url = 'https://github.com/1010code/iris-dnn-tensorflow/raw/master/data/Iris.csv'
s=requests.get(url).content
df_train=pd.read_csv(io.StringIO(s.decode('utf-8')))
df_train = df_train.drop(labels=['Id'],axis=1) # 移除Id
df_train
```
## 2) 手動編碼
處理名目資料 (Nominal variables) - 資料前處理
依據特徵資料的特性,可以選擇手動編碼或自動編碼。
### 使用編碼時機?
進行深度學習時,神經網路只能處理數值資料。因此我們需要將所有非數字型態的特徵進行轉換。
ex:
| Iris-setosa | Iris-versicolor | Iris-virginica |
|:---:|:---:|:---:|
| 1 | 2 | 3 |
```
label_map = {'Iris-setosa': 0, 'Iris-versicolor': 1, 'Iris-virginica': 2}
#將編碼後的label map存至df_train['Species']中。
df_train['Class'] = df_train['Species'].map(label_map)
df_train
```
## 3) 檢查缺失值
使用 numpy 所提供的函式來檢查是否有 NA 缺失值,假設有缺失值使用dropna()來移除。使用的時機在於當只有少量的缺失值適用,若遇到有大量缺失值的情況,或是本身的資料量就很少的情況下建議可以透過機器學習的方法補值來預測缺失值。
```python
# 移除缺失值
train=train.dropna()
```
```
X = df_train.drop(labels=['Species','Class'],axis=1).values # 移除Species (因為字母不參與訓練)
# checked missing data
print("checked missing data(NAN mount):",len(np.where(np.isnan(X))[0]))
```
## 4) 切割訓練集與測試集
```
from sklearn.model_selection import train_test_split
X=df_train.drop(labels=['Class','Species'],axis=1)
y=df_train['Class']
X_train , X_test , y_train , y_test = train_test_split(X,y , test_size=.3 , random_state=42)
print('Training data shape:',X_train.shape)
print('Testing data shape:',X_test.shape)
```
## PCA
Parameters:
- n_components: 指定PCA降維後的特徵維度數目。
- whiten: 是否進行白化True/False。白化意指,對降維後的數據的每個特徵進行正規化,即讓方差都為1、平均值為0。默認值為False。
- random_state: 亂數種子,設定常數能夠保證每次PCA結果都一樣。
Attributes:
- explained_variance_: array類型。降維後的各主成分的方差值,主成分方差值越大,則說明這個主成分越重要
- explained_variance_ratio_: array類型。降維後的各主成分的方差值佔總方差值的比例,主成分所佔比例越大,則說明這個主成分越重要。
- n_components_: int類型。返回保留的特徵個數。
Methods:
- fit(X,y):把數據放入模型中訓練模型。
- fit_transform(X,[,y])all:訓練模型同時返回降維後的數據。
- transform(X):對於訓練好的數據降維。
```
from sklearn.decomposition import PCA
pca = PCA(n_components=2, iterated_power=1)
train_reduced = pca.fit_transform(X_train)
print('PCA方差比: ',pca.explained_variance_ratio_)
print('PCA方差值:',pca.explained_variance_)
plt.figure(figsize=(8,6))
plt.scatter(train_reduced[:, 0], train_reduced[:, 1], c=y_train, alpha=0.5,
cmap=plt.cm.get_cmap('nipy_spectral', 10))
plt.colorbar()
plt.show()
test_reduced = pca.transform(X_test)
plt.figure(figsize=(8,6))
plt.scatter(test_reduced[:, 0], test_reduced[:, 1], c=y_test, alpha=0.5,
cmap=plt.cm.get_cmap('nipy_spectral', 10))
plt.colorbar()
plt.show()
```
### KernelPCA
KernelPCA模型類似於非線性支持向量機,使用核技巧處理非線性數據的降維,主要是選擇合適的核函數。
### IncrementalPCA
IncrementalPCA模型主要是為了解決計算機內存限制問題。工業上樣本量和維度都是非常大的,如果直接擬合數據,機器性能一般都無法支撐。IncrementalPCA則會將數據分成多個batch,然後對每個batch依次遞增調用partial_fit函數對樣本降維。
### SparsePCA
SparsePCA模型相較於普通的PCA區別在於使用了L1正則化,即對非主成分的影響降為0,避免了噪聲對降維的影響。
### MiniBatchSparsePCA
MiniBatchSparsePCA模型類似於SparsePCA,不同之處在於MiniBatchSparsePCA模型通過使用一部分樣本特徵和給定的迭代次數進行降維,以此來解決特徵分解過慢的問題。
## t-SNE
t-SNE使用了更複雜的公式來表達高維與低維之間的關係。且能夠允許非線性的轉換。
Parameters:
- n_components: 指定t-SNE降維後的特徵維度數目。
- n_iter: 設定迭代次數。
- random_state: 亂數種子,設定常數能夠保證每次t-SNE結果都一樣。
Attributes:
- explained_variance_: array類型。降維後的各主成分的方差值,主成分方差值越大,則說明這個主成分越重要
- explained_variance_ratio_: array類型。降維後的各主成分的方差值佔總方差值的比例,主成分所佔比例越大,則說明這個主成分越重要。
- n_components_: int類型。返回保留的特徵個數。
Methods:
- fit(X,y):把數據放入模型中訓練模型。
- fit_transform(X):訓練模型同時返回降維後的數據。
- transform(X):對於訓練好的數據降維。
```
from sklearn.manifold import TSNE
tsneModel = TSNE(n_components=2, random_state=42,n_iter=1000)
train_reduced = tsneModel.fit_transform(X_train)
plt.figure(figsize=(8,6))
plt.scatter(train_reduced[:, 0], train_reduced[:, 1], c=y_train, alpha=0.5,
cmap=plt.cm.get_cmap('nipy_spectral', 10))
plt.colorbar()
plt.show()
```
### t-SNE 不適用於新資料
PCA 降維可以適用新資料,可呼叫transform() 函式即可。而 t-SNE 則不行。因為演算法的關係在 scikit-learn 套件中的 t-SNE 演算法並沒有transform() 函式可以呼叫。
| github_jupyter |
## 4712 errors when extracting #bedrooms
```
def extractBd(x):
val = x['facts and features']
max_idx = val.find(' bd')
if max_idx < 0:
max_idx = len(val)
s = val[:max_idx]
# find comma before
split_idx = s.rfind(',')
if split_idx < 0:
split_idx = 0
else:
split_idx += 2
r = s[split_idx:]
return int(r)
# case I: Studio apartments
extractBd({'facts and features' : 'Studio , 1 ba , -- sqft'})
# or extractBd({'facts and features' : 'Studio , 1 ba , 550 sqft'})
# case II: multiple apartments listed
extractBd({'facts and features' : '3 $2,800+'})
# case III: invalid data
extractBd({'facts and features' : 'Price/sqft: -- , -- bds , -- ba , -- sqft'})
```
Issues:
- forgot about Studio apartments
- ill-formatted data with missing info
## Single error row when extracting zipcode
```
f = lambda x: '%05d' % int(x['postal_code'])
f(None)
```
=> Issue: Forgot to check for NULL values!
## 167 errors when extracting bathrooms
```
def extractBa(x):
val = x['facts and features']
max_idx = val.find(' ba')
if max_idx < 0:
max_idx = len(val)
s = val[:max_idx]
# find comma before
split_idx = s.rfind(',')
if split_idx < 0:
split_idx = 0
else:
split_idx += 2
r = s[split_idx:]
return int(r)
# Row 92 (OperatorID=100007, ecCode=ValueError):
# ('Townhouse For Rent','7 Conant Rd APT 58','WINCHESTER','MA',1890.00000,'$2,850/mo','2 bds , 2.5 ba , 1,409 sqft','https://www.zillow.com/homedetails/7-Conant-Rd-APT-58-Winchester-MA-01890/56541131_zpid/')
# Row 93 (OperatorID=100007, ecCode=ValueError):
# ('Townhouse For Rent','48 Spruce St','Winchester','MA',1890.00000,'$4,250/mo','4 bds , 3.5 ba , 2,000 sqft','https://www.zillow.com/homedetails/48-Spruce-St-Winchester-MA-01890/165961280_zpid/')
# Row 94 (OperatorID=100007, ecCode=ValueError):
# ('Townhouse For Rent','44 Spruce St # 44','Winchester','MA',1890.00000,'$4,500/mo','4 bds , 2.5 ba , 2,500 sqft','https://www.zillow.com/homedetails/44-Spruce-St-44-Winchester-MA-01890/2102250563_zpid/')
extractBa({'facts and features' : '2 bds , 2.5 ba , 1,409 sqft'})
```
Issue:
- forgot about half rooms, need to clean them explicitly!
## 170 errors when extracting Sqft
```
def extractSqft(x):
val = x['facts and features']
max_idx = val.find(' sqft')
if max_idx < 0:
max_idx = len(val)
s = val[:max_idx]
split_idx = s.rfind('ba ,')
if split_idx < 0:
split_idx = 0
else:
split_idx += 5
r = s[split_idx:]
r = r.replace(',', '')
return int(r)
# example: ('House For Rent','232 Washington St','WINCHESTER','MA',1890.00000,'$1,700/mo','2 bds , 1 ba , -- sqft','https://www.zillow.com/homedetails/232-Washington-St-Winchester-MA-01890/56541255_zpid/')
extractSqft({'facts and features' : '2 bds , 1 ba , -- sqft'})
```
+------------+------+
| ValueError | 4712 |
+------------+------+
|- filter
|- withColumn
|- filter
|- withColumn
+-----------+---+
| TypeError | 1 |
+-----------+---+
|- mapColumn
|- withColumn
+------------+---+
| ValueError | 3 |
+------------+---+
|- withColumn
+------------+-----+
| ValueError | 222 |
+------------+-----+
```
# number of exception rows
num_except_rows = 4712 + 1 + 3 + 222
total_rows = 38570 # -1 for header
num_except_rows / total_rows * 100
```
| github_jupyter |
# 05 - Logistic Regression
by [Alejandro Correa Bahnsen](albahnsen.com/)
version 0.1, Feb 2016
## Part of the class [Practical Machine Learning](https://github.com/albahnsen/PracticalMachineLearningClass)
This notebook is licensed under a [Creative Commons Attribution-ShareAlike 3.0 Unported License](http://creativecommons.org/licenses/by-sa/3.0/deed.en_US). Special thanks goes to [Kevin Markham](https://github.com/justmarkham)
# Review: Predicting a Continuous Response
```
# glass identification dataset
import pandas as pd
url = 'http://archive.ics.uci.edu/ml/machine-learning-databases/glass/glass.data'
col_names = ['id','ri','na','mg','al','si','k','ca','ba','fe','glass_type']
glass = pd.read_csv(url, names=col_names, index_col='id')
glass.sort_values('al', inplace=True)
glass.head()
```
**Question:** Pretend that we want to predict **ri**, and our only feature is **al**. How could we do it using machine learning?
**Answer:** We could frame it as a regression problem, and use a linear regression model with **al** as the only feature and **ri** as the response.
**Question:** How would we **visualize** this model?
**Answer:** Create a scatter plot with **al** on the x-axis and **ri** on the y-axis, and draw the line of best fit.
```
%matplotlib inline
import matplotlib.pyplot as plt
# scatter plot using Pandas
glass.plot(kind='scatter', x='al', y='ri')
# equivalent scatter plot using Matplotlib
plt.scatter(glass.al, glass.ri)
plt.xlabel('al')
plt.ylabel('ri')
# fit a linear regression model
from sklearn.linear_model import LinearRegression
linreg = LinearRegression()
feature_cols = ['al']
X = glass[feature_cols]
y = glass.ri
linreg.fit(X, y)
# make predictions for all values of X
glass['ri_pred'] = linreg.predict(X)
glass.head()
# plot those predictions connected by a line
plt.plot(glass.al, glass.ri_pred, color='red')
plt.xlabel('al')
plt.ylabel('Predicted ri')
# put the plots together
plt.scatter(glass.al, glass.ri)
plt.plot(glass.al, glass.ri_pred, color='red')
plt.xlabel('al')
plt.ylabel('ri')
```
### Refresher: interpreting linear regression coefficients
Linear regression equation: $y = \beta_0 + \beta_1x$
```
# compute prediction for al=2 using the equation
linreg.intercept_ + linreg.coef_ * 2
# compute prediction for al=2 using the predict method
linreg.predict(2)
# examine coefficient for al
print(feature_cols, linreg.coef_)
```
**Interpretation:** A 1 unit increase in 'al' is associated with a 0.0025 unit decrease in 'ri'.
```
# increasing al by 1 (so that al=3) decreases ri by 0.0025
1.51699012 - 0.0024776063874696243
# compute prediction for al=3 using the predict method
linreg.predict(3)
```
# Predicting a Categorical Response
```
# examine glass_type
glass.glass_type.value_counts().sort_index()
# types 1, 2, 3 are window glass
# types 5, 6, 7 are household glass
glass['household'] = glass.glass_type.map({1:0, 2:0, 3:0, 5:1, 6:1, 7:1})
glass.head()
```
Let's change our task, so that we're predicting **household** using **al**. Let's visualize the relationship to figure out how to do this:
```
plt.scatter(glass.al, glass.household)
plt.xlabel('al')
plt.ylabel('household')
```
Let's draw a **regression line**, like we did before:
```
# fit a linear regression model and store the predictions
feature_cols = ['al']
X = glass[feature_cols]
y = glass.household
linreg.fit(X, y)
glass['household_pred'] = linreg.predict(X)
# scatter plot that includes the regression line
plt.scatter(glass.al, glass.household)
plt.plot(glass.al, glass.household_pred, color='red')
plt.xlabel('al')
plt.ylabel('household')
```
If **al=3**, what class do we predict for household? **1**
If **al=1.5**, what class do we predict for household? **0**
We predict the 0 class for **lower** values of al, and the 1 class for **higher** values of al. What's our cutoff value? Around **al=2**, because that's where the linear regression line crosses the midpoint between predicting class 0 and class 1.
Therefore, we'll say that if **household_pred >= 0.5**, we predict a class of **1**, else we predict a class of **0**.
## $$h_\beta(x) = \beta_0 + \beta_1x_1 + \beta_2x_2 + ... + \beta_nx_n$$
- $h_\beta(x)$ is the response
- $\beta_0$ is the intercept
- $\beta_1$ is the coefficient for $x_1$ (the first feature)
- $\beta_n$ is the coefficient for $x_n$ (the nth feature)
### if $h_\beta(x)\le 0.5$ then $\hat y = 0$
### if $h_\beta(x)> 0.5$ then $\hat y = 1$
```
# understanding np.where
import numpy as np
nums = np.array([5, 15, 8])
# np.where returns the first value if the condition is True, and the second value if the condition is False
np.where(nums > 10, 'big', 'small')
# transform household_pred to 1 or 0
glass['household_pred_class'] = np.where(glass.household_pred >= 0.5, 1, 0)
glass.head()
# plot the class predictions
plt.scatter(glass.al, glass.household)
plt.plot(glass.al, glass.household_pred_class, color='red')
plt.xlabel('al')
plt.ylabel('household')
```
$h_\beta(x)$ can be lower 0 or higher than 1, which is countra intuitive
## Using Logistic Regression Instead
Logistic regression can do what we just did:
```
# fit a logistic regression model and store the class predictions
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression(C=1e9)
feature_cols = ['al']
X = glass[feature_cols]
y = glass.household
logreg.fit(X, y)
glass['household_pred_class'] = logreg.predict(X)
# plot the class predictions
plt.scatter(glass.al, glass.household)
plt.plot(glass.al, glass.household_pred_class, color='red')
plt.xlabel('al')
plt.ylabel('household')
```
What if we wanted the **predicted probabilities** instead of just the **class predictions**, to understand how confident we are in a given prediction?
```
# store the predicted probabilites of class 1
glass['household_pred_prob'] = logreg.predict_proba(X)[:, 1]
# plot the predicted probabilities
plt.scatter(glass.al, glass.household)
plt.plot(glass.al, glass.household_pred_prob, color='red')
plt.xlabel('al')
plt.ylabel('household')
# examine some example predictions
print(logreg.predict_proba(1))
print(logreg.predict_proba(2))
print(logreg.predict_proba(3))
```
The first column indicates the predicted probability of **class 0**, and the second column indicates the predicted probability of **class 1**.
## Probability, odds, e, log, log-odds
$$probability = \frac {one\ outcome} {all\ outcomes}$$
$$odds = \frac {one\ outcome} {all\ other\ outcomes}$$
Examples:
- Dice roll of 1: probability = 1/6, odds = 1/5
- Even dice roll: probability = 3/6, odds = 3/3 = 1
- Dice roll less than 5: probability = 4/6, odds = 4/2 = 2
$$odds = \frac {probability} {1 - probability}$$
$$probability = \frac {odds} {1 + odds}$$
```
# create a table of probability versus odds
table = pd.DataFrame({'probability':[0.1, 0.2, 0.25, 0.5, 0.6, 0.8, 0.9]})
table['odds'] = table.probability/(1 - table.probability)
table
```
What is **e**? It is the base rate of growth shared by all continually growing processes:
```
# exponential function: e^1
np.exp(1)
```
What is a **(natural) log**? It gives you the time needed to reach a certain level of growth:
```
# time needed to grow 1 unit to 2.718 units
np.log(2.718)
```
It is also the **inverse** of the exponential function:
```
np.log(np.exp(5))
# add log-odds to the table
table['logodds'] = np.log(table.odds)
table
```
## What is Logistic Regression?
**Linear regression:** continuous response is modeled as a linear combination of the features:
$$y = \beta_0 + \beta_1x$$
**Logistic regression:** log-odds of a categorical response being "true" (1) is modeled as a linear combination of the features:
$$\log \left({p\over 1-p}\right) = \beta_0 + \beta_1x$$
This is called the **logit function**.
Probability is sometimes written as pi:
$$\log \left({\pi\over 1-\pi}\right) = \beta_0 + \beta_1x$$
The equation can be rearranged into the **logistic function**:
$$\pi = \frac{e^{\beta_0 + \beta_1x}} {1 + e^{\beta_0 + \beta_1x}}$$
In other words:
- Logistic regression outputs the **probabilities of a specific class**
- Those probabilities can be converted into **class predictions**
The **logistic function** has some nice properties:
- Takes on an "s" shape
- Output is bounded by 0 and 1
We have covered how this works for **binary classification problems** (two response classes). But what about **multi-class classification problems** (more than two response classes)?
- Most common solution for classification models is **"one-vs-all"** (also known as **"one-vs-rest"**): decompose the problem into multiple binary classification problems
- **Multinomial logistic regression** can solve this as a single problem
## Part 6: Interpreting Logistic Regression Coefficients
```
# plot the predicted probabilities again
plt.scatter(glass.al, glass.household)
plt.plot(glass.al, glass.household_pred_prob, color='red')
plt.xlabel('al')
plt.ylabel('household')
# compute predicted log-odds for al=2 using the equation
logodds = logreg.intercept_ + logreg.coef_[0] * 2
logodds
# convert log-odds to odds
odds = np.exp(logodds)
odds
# convert odds to probability
prob = odds/(1 + odds)
prob
# compute predicted probability for al=2 using the predict_proba method
logreg.predict_proba(2)[:, 1]
# examine the coefficient for al
feature_cols, logreg.coef_[0]
```
**Interpretation:** A 1 unit increase in 'al' is associated with a 4.18 unit increase in the log-odds of 'household'.
```
# increasing al by 1 (so that al=3) increases the log-odds by 4.18
logodds = 0.64722323 + 4.1804038614510901
odds = np.exp(logodds)
prob = odds/(1 + odds)
prob
# compute predicted probability for al=3 using the predict_proba method
logreg.predict_proba(3)[:, 1]
```
**Bottom line:** Positive coefficients increase the log-odds of the response (and thus increase the probability), and negative coefficients decrease the log-odds of the response (and thus decrease the probability).
```
# examine the intercept
logreg.intercept_
```
**Interpretation:** For an 'al' value of 0, the log-odds of 'household' is -7.71.
```
# convert log-odds to probability
logodds = logreg.intercept_
odds = np.exp(logodds)
prob = odds/(1 + odds)
prob
```
That makes sense from the plot above, because the probability of household=1 should be very low for such a low 'al' value.
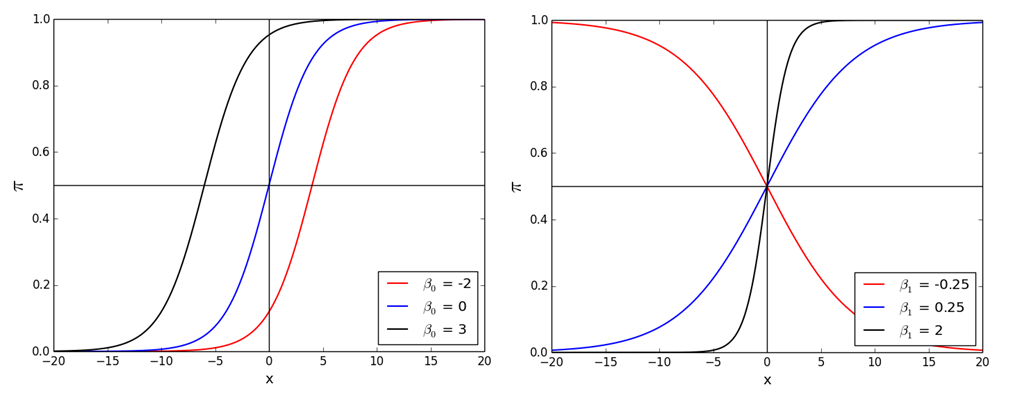
Changing the $\beta_0$ value shifts the curve **horizontally**, whereas changing the $\beta_1$ value changes the **slope** of the curve.
## Comparing Logistic Regression with Other Models
Advantages of logistic regression:
- Highly interpretable (if you remember how)
- Model training and prediction are fast
- No tuning is required (excluding regularization)
- Features don't need scaling
- Can perform well with a small number of observations
- Outputs well-calibrated predicted probabilities
Disadvantages of logistic regression:
- Presumes a linear relationship between the features and the log-odds of the response
- Performance is (generally) not competitive with the best supervised learning methods
- Can't automatically learn feature interactions
| github_jupyter |
# ISAF (إسعاف)
## Integrated Security Assessments for your dev Flow
```
EeeiiiiiEEiiiii.....
\|/
n______ .....iiiiiEEiiiieeEE
:~; : \|/
-----;``~' + ;------------ ______n --------------------------------
`-@-----@-= : :~:
=========================== ; + '~``; =============================
=-@-----@-'
jgs------------------------------------------------------------------
DEVSECOPS IN A PYTHON NUTSHELL
```
# Purpose
This project is a "simple" python implementation of the DevSecOps Methodology, boiled down to the following picture:
[](https://www.gartner.com/doc/1896617/devopssec-creating-agile-triangle)
and an implementation attempt:
[](https://medium.com/@H.A.T/how-to-implement-webs-hospital-b0d8b85389ce)
# Stack Deployement
You can quickly start this PoC (based on OpenFaaS) on Docker Swarm online using the community-run Docker playground: play-with-docker.com (PWD) by clicking the button below:
[](http://labs.play-with-docker.com/?stack=https://gist.githubusercontent.com/h-a-t/eafbb19d7ce46c4ee4a541df018a5f37/raw/d1aa99f6fec38620f09ecd3a9bbfb79207cf3dde/docker-compose.yml&stack_name=func)
Or use the docker-compose.yml file.
# Prerequisite
- Get your Jupyter token:
```bash
$ docker service logs func_jupyter 2>&1 | grep token
func_jupyter.1.xam77gaqxi5s@node1 | [I 18:42:54.562 LabApp] The Jupyter Notebook is running at: http://[all ip addresses on your system]:8888/?token=5afdefee1c98acac0bbf29ae9972b7ccd23c50c115e74e49
func_jupyter.1.xam77gaqxi5s@node1 | to login with a token:
func_jupyter.1.xam77gaqxi5s@node1 | http://localhost:8888/?token=5afdefee1c98acac0bbf29ae9972b7ccd23c50c115e74e49
```
- Install git, unzip, curl and faas-cli in the Jupyter container:
```bash
docker exec --user root -ti func_jupyter.1.shw9s15u6co3cuzp5sjft697t bash
root@b9300915e6ad:~# apt-get update && apt-get -y install unzip git curl
root@b9300915e6ad:~# curl -sSL https://cli.openfaas.com | sh # Not cool :/
```
- Change permissions of docker.sock for the sack of this PoC. :warning: Do not do this in a production environment :bomb:
```bash
chmod 777 /var/run/docker.sock
```
- Upload ISAF.iynb to your Jupyter instance, and press play! \o/
# PoC Stack built upon
| Name | Link | License |
|------------|--------------------------------------------|---------------------------|
| Clair | https://github.com/coreos/clair | Apache License 2.0 |
| JupyterLab | https://github.com/jupyterlab/jupyterlab | BSD 3-Clause |
| Klar | https://github.com/optiopay/klar | MIT |
| Nmap | https://github.com/nmap/nmap | GNU General Public |
| OpenFaaS | https://github.com/openfaas/faas | MIT |
| Sonarqube | https://github.com/SonarSource/sonarqube/ | GNU Lesser General Public |
| WhatWeb | https://github.com/urbanadventurer/WhatWeb | GPLv2 |
| WPScan | https://github.com/wpscanteam/wpscan | Dual-Licensed |
# Build
### What App?
A tiny vulnerable application written in PHP.
[](https://github.com/h-a-t/RedPill/blob/master/src/php/index.php)
These 2 lines are vulnerable to SQL injections and to XSS attacks.
```php
$user_id = $_GET['id'];
$sql = mysql_query("SELECT username, nom, prenom, email FROM users WHERE user_id = $user_id") or die(mysql_error());
```
### Let's deploy it!
```
!cd ~ && git clone https://github.com/h-a-t/RedPill
!pip install docker
import docker
import io
import tarfile
import os
import time
cli = docker.from_env()
cli.containers.list()
## Build app image and pull dependencies
## May take a fair amount of time
os.chdir(os.path.expanduser('~/RedPill/'))
cli.images.build(path='./src/php', tag='hat/app')
#zap_img = cli.images.pull('owasp/zap2docker-weekly:latest')
db_img = cli.images.pull('mariadb:latest')
alpine = cli.images.pull('alpine:latest')
cli.images.list()
## Create a dedicated network
cli.networks.create("app_net", driver="overlay")
for x in cli.networks.list():
print("%s %s" % (x.id,x.name))
## Create Volumes
cli.volumes.create(name='db_data', driver='local')
cli.volumes.create(name='db_init', driver='local')
cli.volumes.create(name='app_data', driver='local')
## Database provisionning
os.chdir(os.path.expanduser('~/RedPill/src/sql'))
tarstream = io.BytesIO()
tar = tarfile.TarFile(fileobj=tarstream, mode='w')
tar.add('staging.sql')
tar.close()
## https://gist.github.com/zbyte64/6800eae10ce082bb78f0b7a2cca5cbc2
tmp=cli.containers.create(
image='alpine',
volumes={'db_init':{'bind': '/data/', 'mode' : 'rw'}})
tarstream.seek(0)
tmp.put_archive(
path='/data/',
data=tarstream
)
## Database run
db_cont = cli.services.create(
image='mariadb:latest',
mounts=[
"db_init:/docker-entrypoint-initdb.d/:rw",
"db_data:/var/lib/mysql/:rw"
],
networks= ['app_net'],
name='db',
env=['MYSQL_RANDOM_ROOT_PASSWORD=yes','MYSQL_USER=user',
'MYSQL_PASSWORD=password','MYSQL_DATABASE=sqli']
)
db_cont?
## Webserver provisionning
os.chdir(os.path.expanduser('~/RedPill/src/php'))
tarstream = io.BytesIO()
tar = tarfile.TarFile(fileobj=tarstream, mode='w')
tar.add('.')
tar.close()
tmp=cli.containers.create(
image='alpine',
volumes={'app_data':{'bind': '/data/', 'mode' : 'rw'}})
tarstream.seek(0)
tmp.put_archive(
path='/data/',
data=tarstream
)
## webserver run
app_cont = cli.services.create(
image='hat/app',
mounts=[
"app_data:/var/www/html:rw",
],
networks= ['app_net','func_functions'],
name='app_web',
endpoint_spec={
'Mode': 'vip',
"Ports": [
{
"Protocol": "tcp",
"TargetPort": 80,
"PublishedPort": 80
}]
},
env=['DB_ENV_MYSQL_USER=user','DB_ENV_MYSQL_PASSWORD=password','BUILD_STAGE=Python'],
)
app_cont?
```
# Security Assessment in a Synchronous Execution Flow
## Static Code Analysis
### Push code to SonarQube for code analysis
```
## Download Sonarqube scanner
os.chdir(os.path.expanduser('~/RedPill'))
!wget https://sonarsource.bintray.com/Distribution/sonar-scanner-cli/sonar-scanner-cli-3.0.3.778-linux.zip
!unzip sonar-scanner-cli-3.0.3.778-linux.zip
!./sonar-scanner-3.0.3.778-linux/bin/sonar-scanner -Dsonar.host.url=http://sonarqube:9000 -Dsonar.projectKey=Redpill:latest -Dsonar.sources=./src/php -Dsonar.language=php
```
### Push to Clair for a container layer scan
TODO: Texte à broder
```
!wget -O clair_conf.yml https://raw.githubusercontent.com/coreos/clair/master/config.yaml.sample
# Database connection string
!sed -i "s/host=.*/postgresql:\/\/postgres:password@db_clair:5432?sslmode=disable/" clair_conf.yml
## Database run
db_clair = cli.services.create(
image='postgres:latest',
networks= ['app_net'],
name='db_clair',
env=['POSTGRES_PASSWORD=password']
)
db_clair?
## Config Provisionning to Clair
tarstream = io.BytesIO()
tar = tarfile.TarFile(fileobj=tarstream, mode='w')
tar.add('clair_conf.yml')
tar.close()
## https://gist.github.com/zbyte64/6800eae10ce082bb78f0b7a2cca5cbc2
tmp=cli.containers.create(
image='alpine',
volumes={'clair_init':{'bind': '/data/', 'mode' : 'rw'}})
tarstream.seek(0)
tmp.put_archive(
path='/data/',
data=tarstream
)
## Clair run
app_clair = cli.services.create(
image='quay.io/coreos/clair:latest',
mounts=[
"clair_init:/config/:rw",
],
networks= ['app_net','func_functions'],
name='app_clair',
endpoint_spec={
'Mode': 'vip',
"Ports": [
{
"Protocol": "tcp",
"TargetPort": 6060,
"PublishedPort": 6060
},
{
"Protocol": "tcp",
"TargetPort": 6061,
"PublishedPort": 6061
}
]
},
command=['/clair','-config','/config/clair_conf.yml']
)
app_clair?
!wget https://github.com/optiopay/klar/releases/download/v1.5-RC2/klar-1.5-RC2-linux-amd64 -O klar
!chmod +x ./klar
!CLAIR_ADDR=http://app_clair:6060 ./klar postgres
!CLAIR_ADDR=http://app_clair:6060 ./klar wpscanteam/vulnerablewordpress
```
## Dynamic Runtime Analysis
### Using the Web-GUI of your favorite Pentesting tool from OWASP: ZAP
- Let L33t do the testing by running a GUI instance of ZAP, just browse localhost:8666/?anonym=true&app=ZAP to start :)
- Warning, image size: 1,52 Go
- Cf. https://github.com/zaproxy/zaproxy/wiki/WebSwing
```
scan_cont = cli.services.create(
image='owasp/zap2docker-stable:latest',
name='app_scan',
networks= ['app_net','func_functions'],
endpoint_spec={
'Mode': 'vip',
"Ports": [
{
"Protocol": "tcp",
"TargetPort": 8080,
"PublishedPort": 8666
},
{
"Protocol": "tcp",
"TargetPort": 8090,
"PublishedPort": 8777
}
]
},
command=['sh','-c','zap-webswing.sh']
)
scan_cont?
```
### Using the flexibility of your favorite cutting-edge technology: OpenFaas
Dockerfile:
```
FROM alexellis2/faas-alpinefunction:latest
RUN apk update && apk add nmap
ENV fprocess="xargs nmap"
CMD ["fwatchdog"]
```
nmap_stack:
```yaml
provider:
name: faas
gateway: http://gateway:8080
functions:
nmap:
lang: Dockerfile
handler: ./Dockerfile
image: hat/nmap
```
```
import tarfile
import time
from io import BytesIO
Dockerfile ='''
FROM alexellis2/faas-alpinefunction:latest
RUN apk update && apk add nmap
ENV fprocess="xargs nmap"
CMD ["fwatchdog"]
'''
with open("Dockerfile", "w") as stack:
stack.write("%s" % Dockerfile)
#write the Dockerfile to a tarred archive
pw_tarstream = BytesIO()
pw_tar = tarfile.TarFile(fileobj=pw_tarstream, mode='w')
file_data = Dockerfile.encode('utf8')
tarinfo = tarfile.TarInfo(name='Dockerfile')
tarinfo.size = len(file_data)
tarinfo.mtime = time.time()
#tarinfo.mode = 0600
pw_tar.addfile(tarinfo, BytesIO(file_data))
pw_tar.close()
pw_tarstream.seek(0)
nmap = cli.images.build(
fileobj=pw_tarstream,
custom_context=True,
tag='hat/nmap'
)
nmap?
## nmap Stack
func_stack='''
provider:
name: faas
gateway: http://gateway:8080
functions:
nmap:
lang: Dockerfile
handler: ./Dockerfile
image: hat/nmap
'''
with open("nmap_func.yml", "w") as stack:
stack.write("%s" % func_stack)
!faas-cli build -f nmap_func.yml
!faas-cli deploy -f nmap_func.yml
## Testing
!curl -v http://gateway:8080/system/functions
## Executing nmap \o/
!curl -v --data "-T4 app_web" --max-time 900 http://gateway:8080/function/nmap
!curl -v http://gateway:8080/system/functions
nmap_serv=cli.services.get('nmap')
# BORDEL COMMENT MARCHE nmap_serv.logs() !!
## Need screenshot? \o/
app_screen = cli.services.create(
image='scrapinghub/splash',
networks= ['app_net','func_functions'],
name='app_screen',
endpoint_spec={
'Mode': 'vip',
"Ports": [
{
"Protocol": "tcp",
"TargetPort": 8050,
"PublishedPort": 8050
}]
}
)
app_screen?
!curl "http://app_screen:8050/render.png?wait=10&render_all=1&url=http://sonarqube:9000/project/issues?id=Redpill%3Alatest&resolved=false&types=BUG" > bug.png
## THE PYTHON WAY \o/
```
Async IO with docker
-> faas in python
https://pypi.python.org/pypi/aiodocker/0.8.2
http://aiodocker.readthedocs.io/en/latest/
https://curio.readthedocs.io/en/latest/tutorial.html
```
import asyncio
from aiodocker.docker import Docker
from aiodocker.exceptions import DockerError
async def demo(docker):
try:
await docker.images.get('alpine:latest')
except DockerError as e:
if e.status == 404:
await docker.pull('alpine:latest')
else:
print('Error retrieving alpine:latest image.')
return
config = {
# "Cmd": ["/bin/ash", "-c", "sleep 1; echo a; sleep 1; echo a; sleep 1; echo a; sleep 1; echo x"],
"Cmd": ["/bin/ash"],
"Image": "alpine:latest",
"AttachStdin": True,
"AttachStdout": True,
"AttachStderr": True,
"Tty": False,
"OpenStdin": True,
"StdinOnce": True,
}
container = await docker.containers.create_or_replace(
config=config, name='aiodocker-example')
print("created and started container {}".format(container._id[:12]))
try:
ws = await container.websocket(stdin=True, stdout=True, stderr=True, stream=True)
await container.start()
async def _send():
await asyncio.sleep(0.5)
await ws.send_bytes(b'echo "hello world"\n')
print("sent a shell command")
asyncio.ensure_future(_send())
resp = await ws.receive()
print("received: {}".format(resp))
await ws.close()
output = await container.log(stdout=True)
print("log output: {}".format(output))
finally:
print("removing container")
await container.delete(force=True)
if __name__ == '__main__':
loop = asyncio.get_event_loop()
docker = Docker()
try:
loop.run_until_complete(demo(docker))
finally:
loop.run_until_complete(docker.close())
loop.close()
import asyncio
from aiodocker.docker import Docker
async def demo(docker):
print('--------------------------------')
print('- Check Docker Version Information')
data_version = await docker.version()
for key, value in data_version.items():
print(key, ':', value)
print('--------------------------------')
print('- Check Docker Image List')
images = await docker.images.list()
for image in images:
for key, value in image.items():
if key == 'RepoTags':
print(key, ':', value)
print('--------------------------------')
print('- Check Docker Container List')
containers = await docker.containers.list()
for container in containers:
container_show = await container.show()
for key, value in container_show.items():
if key == 'Id':
print('Id', ':', value[:12])
print('--------------------------------')
if __name__ == '__main__':
loop = asyncio.get_event_loop()
docker = Docker()
try:
loop.run_until_complete(demo(docker))
finally:
loop.run_until_complete(docker.close())
loop.close()
import asyncio
from aiodocker.docker import Docker
from aiodocker.exceptions import DockerError
async def demo(docker):
try:
await docker.images.get('alpine:latest')
except DockerError as e:
if e.status == 404:
await docker.pull('alpine:latest')
else:
print('Error retrieving alpine:latest image.')
return
subscriber = docker.events.subscribe()
config = {
"Cmd": ["tail", "-f", "/var/log/dmesg"],
"Image":"alpine:latest",
"AttachStdin": False,
"AttachStdout": True,
"AttachStderr": True,
"Tty": False,
"OpenStdin": False,
"StdinOnce": False,
}
container = await docker.containers.create_or_replace(
config=config, name='testing')
await container.start()
print("=> created and started container {}".format(container._id[:12]))
while True:
event = await subscriber.get()
if event is None:
break
for key, value in event.items():
print(key,':', value)
# Demonstrate simple event-driven container mgmt.
if event['Actor']['ID'] == container._id:
if event['Action'] == 'start':
await container.stop()
print("=> killed {}".format(container._id[:12]))
elif event['Action'] == 'stop':
await container.delete(force=True)
print("=> deleted {}".format(container._id[:12]))
elif event['Action'] == 'destroy':
print('=> done with this container!')
break
if __name__ == '__main__':
loop = asyncio.get_event_loop()
docker = Docker()
try:
# do our stuffs.
loop.run_until_complete(demo(docker))
finally:
loop.run_until_complete(docker.close())
loop.close()
import asyncio
import aiodocker
async def list_things():
docker = aiodocker.Docker()
print('== Images ==')
for image in (await docker.images.list()):
tags = image['RepoTags'][0] if image['RepoTags'] else ''
print(image['Id'], tags)
print('== Containers ==')
for container in (await docker.containers.list()):
print(f" {container._id}")
await docker.close()
async def run_container():
docker = aiodocker.Docker()
print('== Running a hello-world container ==')
container = await docker.containers.create_or_replace(
config={
'Cmd': ['/bin/ash', '-c', 'echo "hello world"'],
'Image': 'alpine:latest',
},
name='testing',
)
await container.start()
logs = await container.log(stdout=True)
print(''.join(logs))
await container.delete(force=True)
await docker.close()
if __name__ == '__main__':
loop2 = asyncio.get_event_loop()
loop2.run_until_complete(list_things())
loop2.run_until_complete(run_container())
loop2.close()
```
| github_jupyter |
### The company of this exercise is a social network. They decided to add a feature called: Recommended Friends, i.e. they suggest people you may know.
### A data scientist has built a model to suggest 5 people to each user. These potential friends will be shown on the user newsfeed. At first, the model is tested just on a random subset of users to see how it performs compared to the newsfeed without the new feature.
### The test has been running for some time and your boss asks you to check the results. You are asked to check, for each user, the number of pages visited during their first session since the test started. If this number increased, the test is a success. Specifically, your boss wants to know:
#### (1) Is the test winning? That is, should 100\% of the users see the Recommended Friends feature?
#### (2) Is the test performing similarly for all user segments or are there differences among different segments?
#### (3) If you identified segments that responded differently to the test, can you guess the reason? Would this change your point 1 conclusions?
### Load the package would be used
```
import pandas as pd
pd.set_option("display.max_columns", 10)
pd.set_option("display.width", 350)
from scipy import stats
import matplotlib.pyplot as plt
from matplotlib import rcParams
rcParams.update({"figure.autolayout": True})
import seaborn as sns
sns.set(style = "white")
sns.set(style = "whitegrid", color_codes = True)
user = pd.read_csv("../Datasets/Engagement_Test/user_table.csv")
test = pd.read_csv("../Datasets/Engagement_Test/test_table.csv")
```
### Look into data
```
print(user.shape)
print(test.shape)
print(user.head)
print(test.head)
print(user.info)
print(test.info)
print(len(user["user_id"]) == len(pd.unique(user["user_id"])))
print(len(test["user_id"]) == len(pd.unique(test["user_id"])))
```
### Data processing
```
dat = user.merge(test, on = "user_id", how = "inner")
dat["signup_date"] = pd.to_datetime(dat["signup_date"])
dat["date"] = pd.to_datetime(dat["date"])
dat.head()
```
#### (1) Is the test winning? That is, should 100% of the users see the Recommended Friends feature?
#### Overall
##### Define the function
```
def overall_ttest_mean(dat, variable, test):
overall = stats.ttest_ind(dat[dat[test] == 1][variable], dat[dat[test] == 0][variable], equal_var = False)
test_group = dat[dat[test] == 1][variable].mean()
control_group = dat[dat[test] == 0][variable].mean()
pvalue = overall.pvalue
overall_result = {"test_group": test_group, "control_group": control_group, "pvalue": pvalue}
return overall_result
overall_ttest_mean(dat = dat, variable = "pages_visited", test = "test")
```
#### (2) Is the test performing similarly for all user segments or are there differences among different segments?
#### Stratified test
##### Define the function
```
def stratified_ttest_mean(dat, stratified, variable, test):
stratified_result = dat.groupby(stratified)[variable].agg({
"test_group": lambda x: x[dat[test] == 1].mean(),
"control_group": lambda x: x[dat[test] == 0].mean(),
"p_value": lambda x: stats.ttest_ind(x[dat[test] == 1], x[dat[test] == 0], equal_var = False).pvalue
}).reindex(["test_group", "control_group", "p_value"], axis = 1)
return stratified_result.sort_values(by = "p_value")
```
##### Stratified by browser
```
stratified_ttest_mean(dat = dat, stratified = "browser", variable = "pages_visited", test = "test").reset_index()
```
##### Stratified by date
```
stratified_ttest_mean(dat = dat, stratified = "date", variable = "pages_visited", test = "test").reset_index().sort_values(by = "date")
```
#### (3) If you identified segments that responded differently to the test, can you guess the reason? Would this change your point 1 conclusions?
```
overall_ttest_mean(dat = dat[dat["browser"] != "Opera"], variable = "pages_visited", test = "test")
```
Still rush to make the decision to change. Need to consider the novelty effect.
| github_jupyter |
# Demonstration of OVRO-LWA Stage 3 M&C
## Python APIs
1. ARX (Larry, Rick)
2. F-engine (Jack)
3. X-engine (Jack)
4. Data capture (Jayce)
## Setup
Set up scripts often run on lxdlwacr. lxdlwagpu03 is also ok.
Jack recommends:
1. Start F-engine (tested on lxdlwacr and lxdlwagpu03)
```
from lwa_f import snap2_fengine
f = snap2_fengine.Snap2Fengine('snap01')
f.cold_start_from_config('/home/ubuntu/proj/caltech-lwa/control_sw/config/lwa_corr_config.yaml')
```
2. Start X-engines
```
from lwa352_pipeline_control import Lwa352CorrelatorControl
p = Lwa352CorrelatorControl(['lxdlwagpu0%d' % i for i in range(1,3)]) #or however many pipelines are being used.
p.stop_pipelines()
p.start_pipelines()
import time; time.sleep(15) # Wait for pipelines to come up
p.configure_corr()
```
Older instructions:
1. Program and initialize SNAPs with `lwa_snap_feng_init.py -e -s -m -i -p -o ~/proj/lwa-shell/caltech-lwa/control_sw/config/lwa_corr_config.yaml snap01`. Adding `-t` will send test data.
2. Fire up the GPU pipelines with `lwa352-start-pipeline.sh 0 1` on gpu01 and gpu02 (possibly killing existing pipelines first). One server can run up to 4 pipelines (192 channels x ~24 kHz each). All servers needed to run 32 pipelines with 73MHz bandwidth.
3. Watch X-engine log files until things are running with names `<hostname>.<pipeline_id>.log`
4. Trigger correlator output with `lwa352_arm_correlator.py <server>`, where server can be "lxdlwagpu01".
5. Currently, success means 10s integrations sent to lxdlwagpu03 for capture by Jayce's bifrost pipeline. Good logging:
`2021-07-08 20:33:17 [INFO ] CORR OUTPUT >> Sending complete for time_tag 318652171392122880 in 2.12 seconds (381714432 Bytes; 1.44 Gb/s)`
Note:
* See also video recording of demo at https://youtu.be/H4ihj0EBHXE.
## Breakdown
1. lwa352-stop-pipeline.sh # essentially just a `killall lwa352-pipeline.py`
## Set up
```
%matplotlib inline
from bokeh.layouts import column, row
from bokeh.models import ColumnDataSource, DataRange1d, Select
from bokeh.palettes import Blues4
from bokeh.plotting import figure
from bokeh.io import show, output_notebook
import pandas as pd
output_notebook()
from lwa_f import snap2_fengine, snap2_feng_etcd_client
cd ~/ovro_data_recorder/
import matplotlib.pyplot as plt
import numpy as np
from common import ETCD_HOST, ETCD_PORT # works in ovro_data_recorder directory
```
## etcd level M&C (low level)
```
import etcd3
le = etcd3.client(ETCD_HOST, ETCD_PORT)
# e.g., print a key:
# -- "le.get('/cmd/arx/2')"
# -- "le.get('/mon/snap/1')"
```
## ARX M&C
* Repo forthcoming "lwa-pyutils"
* This will be wrapped to refer to antenna, rather than ARX.
* Note that ARX come up with max attenuation (31.5)
```
from lwautils import lwa_arx
ma = lwa_arx.ARX()
adrs = [17,21,27,31] # currently installed
for adr in adrs:
ma.load_cfg(adr, 1) # optimal preset values
# ma.raw(adr, 'SETSC387') # reasonable default
# setting custom config with dictionary
#ARX_CHAN_CFG = {}
#ARX_CHAN_CFG["dc_on"] = True
#ARX_CHAN_CFG["sig_on"] = True
#ARX_CHAN_CFG["narrow_lpf"] = False
#ARX_CHAN_CFG["narrow_hpf"] = False
#ARX_CHAN_CFG["first_atten"] = 7.5
#ARX_CHAN_CFG["second_atten"] = 15.0
#for adr in adrs:
# ma.set_all_chan_cfg(adr, ARX_CHAN_CFG) # this returns KeyError atm
```
## F-engine M&C
* Repo https://github.com/realtimeradio/caltech-lwa
* python install "control_sw"
```
lwa_f = snap2_fengine.Snap2Fengine('snap01')
lwa_fe = snap2_feng_etcd_client.Snap2FengineEtcdClient('snap01', 1)
print(lwa_f.blocks.keys())
print(lwa_f.is_connected(), le.is_polling())
st, fl = lwa_f.eqtvg.get_status()
print(st)
st0, st1 = lwa_f.input.get_status()
pows = np.array([v for (k,v) in st0.items() if 'pow' in k])
means = np.array([v for (k,v) in st0.items() if 'mean' in k])
TOOLTIPS = [("SNAP2 input", "$index")]
plot = figure(plot_width=950, plot_height=500, title='F stats', tools='hover,wheel_zoom,reset',
tooltips=TOOLTIPS)
plot.xaxis.axis_label = 'Powers'
plot.yaxis.axis_label = 'Means'
_ = plot.scatter(x=pows, y=means)
show(plot)
hist = lwa_f.input.plot_histogram(50)
specs = lwa_f.autocorr.get_new_spectra() # before 4-bit scaling/quantization
len(specs), len(specs[0])
autospecs = []
for i in range(64):
spec = lwa_f.corr.get_new_corr(i, i).real # 8-channel average, normalized by accumulated time/chans
autospecs.append(spec)
fig, (ax0) = plt.subplots(1, 1, figsize=(12,12))
#for autospec in autospecs:
ax0.imshow(np.vstack(autospecs), origin='bottom')
#ax0.imshow(specs, origin='bottom')
np.shape(autospecs)
from pyuvdata import UVData
from lwa_antpos import stations
uvd = UVData()
uvd.Nants_data = np.shape(autospecs)[0]
uvd.Nants_telescope = np.shape(autospecs)[0] # Need to update
uvd.Nbls = uvd.Nblts = np.shape(autospecs)[0]
uvd.Nfreqs = np.shape(autospecs)[1]
uvd.Npols = 1
uvd.Nspws = 1
uvd.Ntimes = 1
uvd.ant_1_array = uvd.ant_2_array = np.arange(uvd.Nblts)
uvd.antenna_names = np.arange(uvd.Nblts).astype('str') # Get actual antenna numbers
uvd.antenna_numbers = np.arange(uvd.Nblts) # Get actual antenna numbers
uvd.antenna_positions = None # Need antenna positions
uvd.baseline_array = 2048*(uvd.ant_1_array+1)+(uvd.ant_1_array+1)+2^16
uvd.channel_width = None # Need channel width
uvd.freq_array = None # Need frequency
uvd.instrument = uvd.telescope_name ='LWA'
uvd.integration_time = None # Need integration time
uvd.phase_type = 'drift'
uvd.vis_units = 'uncalib'
uvd.data_array = np.array(autospecs)[:, np.newaxis, :, np.newaxis]
uvd.check()
#dd = {'x': list(range(512))}
#
#for i in range(64):
# dd[str(i)] = autospecs[i]
#
#source = ColumnDataSource(dd)
TOOLTIPS = [
# ("Frequency channel", "$index"),
# ("Autocorrelation value", "$y"),
("SNAP2 autocorr", "$name"),
]
plot = figure(plot_width=950, plot_height=500, title='autocorr spectra', tooltips=TOOLTIPS, tools='hover,wheel_zoom,reset')
plot.xaxis.axis_label = 'Channel'
plot.yaxis.axis_label = 'Power'
for i in range(64):
_ = plot.line(x=list(range(512)), y=autospecs[i], color='grey', name=str(i), hover_color='firebrick')
show(plot)
```
## X-engine M&C
* Repo https://github.com/realtimeradio/caltech-lwa
* python install "control_sw"
"/mon/corr/xeng/<hostname>/pipeline/<pid>/corr" or "beamform"
```
from lwa352_pipeline_control import Lwa352PipelineControl
lwa_x = Lwa352PipelineControl(pipeline_id=0, etcdhost=ETCD_HOST, host='lxdlwagpu01')
lwa_x.corr.get_bifrost_status()
from astropy import time
time.Time.now().unix
```
## Data capture M&C
```
# gpu03 runs capture server
import mcs # works in ovro_data_recorder directory
from common import LWATime # works in ovro_data_recorder directory
from datetime import timedelta, datetime
c = mcs.Client()
mcs_id = 'drvs19' # data recorder visibilities slow server 19 (end of x-engine IP address)
# start
t_now = LWATime(datetime.utcnow() + timedelta(seconds=15), format='datetime', scale='utc')
mjd_now = int(t_now.mjd)
mpm_now = int((t_now.mjd - mjd_now)*86400.0*1000.0)
r = c.send_command(mcs_id, 'start', start_mjd=mjd_now, start_mpm=mpm_now)
# stop
r = c.send_command(mcs_id, 'stop', stop_mjd='now', stop_mpm=0)
print(le.get(f'/mon/{mcs_id}/bifrost/rx_rate'), le.get(f'/mon/{mcs_id}/bifrost/max_process'))
le.get(f'/mon/{mcs_id}/statistics/avg')
```
| github_jupyter |
# Tutorial 2
**CS3481 Fundamentals of Data Science**
*Semester B 2019/20*
___
**Instructions:**
- same as [Tutorial 1](http://bit.ly/CS3481T1).
___
## Exercise 1 (submit via uReply)
Complete the tutorial exercises of [[Witten11]](https://ebookcentral.proquest.com/lib/cityuhk/reader.action?docID=634862&ppg=595) from **Exercise 17.1.3** to **17.1.7**, and read up to and including the subsection [**The Visualize Panel**](https://ebookcentral.proquest.com/lib/cityuhk/reader.action?docID=634862&ppg=597). Submit your answers of 17.1.3-5 through [uReply](https://cityu.ed2.mobi/student/mobile_index.php) section number **LM715**.
[*Hint: See the [documentation](https://ebookcentral.proquest.com/lib/cityuhk/reader.action?ppg=438&docID=634862&tm=1547446912037) of WEKA for more details.*]
___
**Answers to 17.1.3:**
**Answers to 17.1.4:**
**Answers to 17.1.5:**
___
## Exercise 2 (no submission required)
Use a text editor of your choice to create an ARFF file for the AND gate $Y=X_1\cdot X_2$, and then load the file into WEKA to ensure it is correct.
[*See the [documentation](https://waikato.github.io/weka-wiki/formats_and_processing/arff_stable/) for the ARFF file format, or take a look at some of the ARFF files in the WEKA data folder as examples.*]
___
**Answer**: Modify the following to create the desired ARFF.
```
text = '''@RELATION AND
@ATTRIBUTE X1 {0, 1}
@ATTRIBUTE X2 {_, _}
@ATTRIBUTE Y {_, _}
@DATA
0, _, _
0, _, _
0, _, _
0, _, _
'''
with open('AND.arff','w') as file:
file.write(text)
```
Load the ARFF file into a dataframe to check if the ARFF file is correct.
```
from scipy.io import arff
import pandas as pd
data = arff.loadarff('AND.arff')
df = pd.DataFrame(data[0])
df.head()
```
## Exercise 3 (no submission required)
[[Han11]](https://www.sciencedirect.com/science/article/pii/B9780123814791000022#s0185) **Question 2.5**: Briefly outline how to compute the dissimilarity between objects described by the following:
(a) Nominal attributes
___
Answer:
___
(b) Asymmetric binary attributes
___
Answer:
___
(c) Numeric attributes
___
Answer:
___
(d) Term-frequency vectors
___
Answer:
___
## Exercise 4 (Optional)
The following illustrates some methods of loading datasets into CoLab. You can execute the code by `shift+enter`.
### (a) Load sample datasets from scikit-learn.
Import the `sklearn.datasets` package, which contains the desired iris dataset. Then, load the iris data sets and print its content.
```
from sklearn import datasets # see https://scikit-learn.org/stable/datasets/index.html
iris = datasets.load_iris()
iris # to print out the content
```
The field `DESCR` (description) contains some background information of the dataset. We can pretty-print only the description as follows.
```
print(iris.DESCR)
```
Convert the dataset to a Pandas dataframe.
```
import pandas as pd
import numpy as np #
iris_pd = pd.DataFrame(data = np.c_[iris['data'], iris['target']], columns = iris['feature_names']+['target'])
iris_pd
```
The function `np.c_` above concatenates the columns `iris['data']` of input features and the column `iris['target']` of class values together. For more details of the function, use the `help` function.
Note however that the datatype of the target becomes floating point, which is not desired. It is also unclear what the number means. The following add the target separately as a vector of strings.
```
iris_pd = pd.DataFrame(data = iris['data'], columns = iris['feature_names'])
iris_pd.insert(len(iris_pd.columns),'target',[iris.target_names[i] for i in iris['target']])
iris_pd
```
### (b) Download and load datasets from openML
Download the `weather.nominal` dataset from [openml.org](https://www.openml.org/d/41521).
[*See the [documentation](https://scikit-learn.org/stable/datasets/index.html#downloading-datasets-from-the-openml-org-repository) for more details.*]
```
from sklearn.datasets import fetch_openml
weather = fetch_openml(data_id=41521)
weather
```
Pretty-print with text and wrap it to 100 characters per line.
```
import textwrap
print(textwrap.fill(weather.DESCR,100))
```
Conversion to dataframe.
```
weather_pd = pd.DataFrame(data=np.c_[weather.data,weather.target],columns=weather.feature_names+['target'])
weather_pd
weather_pd["temperature"] = pd.to_numeric(weather_pd["temperature"])
weather_pd["humidity"] = pd.to_numeric(weather_pd["humidity"])
weather_pd
```
**Exercise:** Modify the dataframe so that the columns for `outlook` and `windy` use their respective category labels instead of indexes.
```
```
### (c) Download a CSV file from UCI Machine Learning repository and read it into a dataframe directly
```
import urllib.request
import io
import pandas as pd
url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data'
ftpstream = urllib.request.urlopen(url)
iris = pd.read_csv(io.StringIO(ftpstream.read().decode('utf-8')))
iris
```
**Exercise:** There is something wrong with the above dataframe. Use the additional options `names` and `index_col` of `read_csv` to read the CSV file correctly.
```
```
| github_jupyter |
# Measuring intensity on label borders
In some applications it is reasonable to measure the intensity on label borders. For example, to measure the signal intensity in an image showing the nuclear envelope, one can segment nuclei, identify their borders and then measure the intensity there.
```
import numpy as np
from skimage.io import imread, imshow
import pyclesperanto_prototype as cle
from cellpose import models, io
from skimage import measure
import matplotlib.pyplot as plt
```
## The example dataset
In this example we load an image showing a zebrafish eye, courtesy of Mauricio Rocha Martins, Norden lab, MPI CBG Dresden.
```
multichannel_image = imread("../../data/zfish_eye.tif")
multichannel_image.shape
cropped_image = multichannel_image[200:600, 500:900]
nuclei_channel = cropped_image[:,:,0]
cle.imshow(nuclei_channel)
```
## Image segmentation
First, we use cellpose to segment the cells
```
# load cellpose model
model = models.Cellpose(gpu=False, model_type='nuclei')
# apply model
channels = [0,0] # This means we are processing single channel greyscale images.
label_image, flows, styles, diams = model.eval(nuclei_channel, diameter=None, channels=channels)
# show result
cle.imshow(label_image, labels=True)
```
## Labeling pixels on label borders
Next, we will extract the outline of the segmented nuclei.
```
binary_borders = cle.detect_label_edges(label_image)
labeled_borders = binary_borders * label_image
cle.imshow(label_image, labels=True)
cle.imshow(binary_borders)
cle.imshow(labeled_borders, labels=True)
```
## Dilating outlines
We extend the outlines a bit to have a more robust measurement.
```
extended_outlines = cle.dilate_labels(labeled_borders, radius=2)
cle.imshow(extended_outlines, labels=True)
```
## Overlay visualization
Using this label image of nuclei outlines, we can measure the intensity in the nuclear envelope.
```
nuclear_envelope_channel = cropped_image[:,:,2]
cle.imshow(nuclear_envelope_channel)
cle.imshow(nuclear_envelope_channel, alpha=0.5, continue_drawing=True)
cle.imshow(extended_outlines, alpha=0.5, labels=True)
```
## Label intensity statistics
Measuring the intensity in the image works using the right intensty and label images.
```
stats = cle.statistics_of_labelled_pixels(nuclear_envelope_channel, extended_outlines)
stats["mean_intensity"]
```
## Parametric maps
These measurements can also be visualized using parametric maps
```
intensity_map = cle.mean_intensity_map(nuclear_envelope_channel, extended_outlines)
cle.imshow(intensity_map, min_display_intensity=3000, colorbar=True, colormap="jet")
```
## Exercise
Measure and visualizae the intensity at the label borders in the nuclei channel.
| github_jupyter |
## Work
1. 請比較使用不同層數以及不同 Dropout rate 對訓練的效果
2. 將 optimizer 改成使用 Adam 並加上適當的 dropout rate 檢視結果
```
import os
import keras
import itertools
from keras.datasets import cifar10
from keras.models import Model
from keras.layers import Input, Dense, Dropout
from keras.optimizers import adam
# Disable GPU
os.environ["CUDA_VISIBLE_DEVICES"] = ""
train, test = cifar10.load_data()
## 資料前處理
def preproc_x(x, flatten=True):
x = x / 255.
if flatten:
x = x.reshape((len(x), -1))
return x
def preproc_y(y, num_classes=10):
if y.shape[-1] == 1:
y = keras.utils.to_categorical(y, num_classes)
return y
x_train, y_train = train
x_test, y_test = test
# Preproc the inputs
x_train = preproc_x(x_train)
x_test = preproc_x(x_test)
input_shape = x_train.shape[1:]
# Preprc the outputs
y_train = preproc_y(y_train)
y_test = preproc_y(y_test)
def build_mlp(input_shape, hidden_neuron_units, dropout_rate, output_shape):
input_layer = Input(shape=input_shape)
for ind, num in enumerate(hidden_neuron_units):
if ind == 0:
x = Dense(units=num, activation='relu')(input_layer)
else:
x = Dense(units=num, activation='relu')(x)
x = Dropout(dropout_rate)(x)
output_layer = Dense(output_shape, activation='softmax')(x)
model = Model(inputs=[input_layer], outputs=[output_layer])
return model
model = build_mlp(input_shape, hidden_neuron_units=[256, 128], dropout_rate=0.3, output_shape=10)
model.summary()
## 超參數設定
EPOCHS = 10
LEARNING_RATE = 1e-3
BATCH_SIZE = 256
DROPOUT_RATE = [.3, .4, .5]
HIDDEN_NEURON_UNITS = [256, 128]
OUTPUT_SHAPE = 10
results = {}
for rate in DROPOUT_RATE:
keras.backend.clear_session()
model = build_mlp(input_shape=input_shape,
hidden_neuron_units=HIDDEN_NEURON_UNITS,
dropout_rate=rate, output_shape=OUTPUT_SHAPE)
model.compile(optimizer=adam(lr=LEARNING_RATE),
loss='categorical_crossentropy',
metrics=['accuracy'])
model.fit(x_train, y_train, epochs=EPOCHS,
batch_size=BATCH_SIZE,
validation_data=(x_test, y_test),shuffle=True)
# Collect results
train_loss = model.history.history["loss"]
valid_loss = model.history.history["val_loss"]
train_acc = model.history.history["acc"]
valid_acc = model.history.history["val_acc"]
experiment = 'DropRate:{}'.format(rate)
results[experiment] = {
'train_loss': train_loss,
'valid_loss': valid_loss,
'train_acc': train_acc,
'valid_acc': valid_acc
}
import matplotlib.pyplot as plt
%matplotlib inline
color_bar = ["r", "g", "b", "y", "m", "k"]
plt.figure(figsize=(8,6))
for i, cond in enumerate(results.keys()):
plt.plot(range(len(results[cond]['train_loss'])),results[cond]['train_loss'], '-', label=cond, color=color_bar[i])
plt.plot(range(len(results[cond]['valid_loss'])),results[cond]['valid_loss'], '--', label=cond, color=color_bar[i])
plt.title("Loss")
plt.ylim([0, 5])
plt.legend()
plt.show()
plt.figure(figsize=(8,6))
for i, cond in enumerate(results.keys()):
plt.plot(range(len(results[cond]['train_acc'])),results[cond]['train_acc'], '-', label=cond, color=color_bar[i])
plt.plot(range(len(results[cond]['valid_acc'])),results[cond]['valid_acc'], '--', label=cond, color=color_bar[i])
plt.title("Accuracy")
plt.legend()
plt.show()
```
| github_jupyter |
```
# hide
import sys
sys.path.append("..")
# default_exp sweep_ensemble_utils
```
# Ensemble utils for WandB Sweeps
Per run, wandb stores pred_test and pred_valid pickle files containing:
* preds,targs,inp,losses,preds_denorm,targs_denorm,inp_denorm,pred_date
Per sweep, per run need to access these predictions to be ensembled. WandB API:
* https://docs.wandb.com/library/reference/wandb_api
* https://docs.wandb.com/library/api
* https://docs.wandb.ai/library/public-api-guide
* https://docs.wandb.ai/ref/public-api
Need to ensemble these predictions calculate mean/median and variance as an uncertainty on the model prediction.
```
# export
import pandas as pd
import numpy as np
import datetime as dt
import matplotlib as mpl
import matplotlib.pyplot as plt
import pickle
import pathlib
import wandb
from pathlib import Path
from sklearn.metrics import mean_squared_error, mean_absolute_error
from scipy.stats import pearsonr
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import GradientBoostingRegressor
from lib.read_data import *
from lib.stats_utils import cErrorMetrics, cStationary
# careful, they change things a lot...
#!pip3 install --upgrade wandb
!wandb --version
#!wandb login <add your key here>
```
## Data from WandB API
```
# export
def fdownload_wandb_files(api, entity, project, sweep_ids, datadirs, wandbfname="preds_test_fname"):
"""
download files with fname for all runs associated with list of sweep_ids into datadirs
output:
data[run_id] = [path, run.config]
"""
data = {}
for i in range(len(sweep_ids)):
sweep = api.sweep("{}/{}/{}".format(entity, project, sweep_ids[i]))
print("Sweep: ", sweep.config["name"])
datadir = pathlib.Path(datadirs[i])
datadir.mkdir(parents=True, exist_ok=True)
for run in sweep.runs:
run_id = run.id
# fname on wandb api same for each run, need to rename with run_id
fname_todownload = run.config[wandbfname]#.split("/")[1]
fname_downloaded = "{}_{}.{}".format(fname_todownload.split(".")[0], run_id, fname_todownload.split(".")[1])
# download (if doesn't already exist) and rename
if not (datadir/fname_downloaded).is_file():
file = run.file(fname_todownload)
print("{} downloading {}...".format(run_id, fname_todownload))
file.download(replace=False, root=datadir)
(datadir/fname_todownload).rename(datadir/fname_downloaded)
else:
print("{} already downloaded.".format(fname_downloaded))
# store location of file and run info
data[run_id] = [datadir/fname_downloaded, run.config]
return data
# export
def fget_pickled(sweeps, datadir, entity, project, pklname="models/preds_test.pickle",
download=True, sweep_type="txt"):
"""
download pickle files associated with runs in sweeps
read data into dictionary first broken down by horizon, then indiviual runs
sweep_type: "api" or "txt" [for manual .txt files with sweep run ids]
sweeps == list of sweep ids or list of paths/filenames containing run ids
"""
# path to download files to
datadir = Path(datadir)
datadir.mkdir(parents=True, exist_ok=True)
api = wandb.Api()
data = {}
for sweep_i in sweeps:
# case of wandb api working correctly:
if sweep_type == "api":
sweep = api.sweep("{}/{}/{}".format(entity, project, sweep_i))
runs = sweep.runs
lrun = len(runs)
# case of txt files containing list of run_ids:
elif sweep_type == "txt":
df = pd.read_csv(sweep_i, delimiter=", ")
run_ids = df["ID"].to_numpy().flatten()
horizon = df["H"].to_numpy().flatten()
lrun = len(run_ids)
print(sweep_i, lrun)
for i in range(lrun):
if sweep_type == "api":
run = runs[i]
run_id = run.id
h = run.config["horizon"]
elif sweep_type == "txt":
run_id = run_ids[i]
# need access to run.config to get horizon (this is slow...)
#run = api.run("{}/{}/{}".format(entity, project, run_id))
h = horizon[i]
# need to change name to include run_id or will overwrite
fname_todownload = pklname
fname_downloaded = "{}_{}.{}".format(fname_todownload.split(".")[0], run_id, fname_todownload.split(".")[1])
# download pickle file (if doesn't already exist)
if download:
if not (datadir/fname_downloaded).is_file():
file = run.file(fname_todownload)
print("{} downloading {}...".format(run_id, fname_todownload))
file.download(replace=False, root=datadir)
(datadir/fname_todownload).rename(datadir/fname_downloaded)
else:
print("{} already downloaded.".format(fname_downloaded))
# read in pickle
with open(datadir/fname_downloaded, 'rb') as handle:
d = pickle.load(handle)
# store by horizon, run_id
try:
data[h][run_id] = d
except KeyError:
data[h] = {}
data[h][run_id] = d
return data
entity = "stardust-r"
project = "deep-learning-space-weather-forecasting"
# wandb sweep ids
#sweeps = ["inztzkl4", "607ppbji", "9ihcleuh", "2ajetr8i"]
# path to manual id text files
sweeps = ["./wandb_ensemble/ensemble/ensembleH3.txt", "./wandb_ensemble/ensemble/ensembleH5H7.txt",
"./wandb_ensemble/ensemble/ensembleH10H14.txt", "./wandb_ensemble/ensemble/ensembleH21H27.txt"]
# path to downloaded sweep run files
datadir = "../data/wandb_ensemble/ensemble"
%%time
data_valid = fget_pickled(sweeps, datadir, entity, project, pklname="models/preds_test.pickle",
download=False, sweep_type="txt")
%%time
# for now this is not logged with wandb: must be manually generated by downloading model file and applying to training data
data_train = fget_pickled(sweeps, datadir, entity, project, pklname="models/preds_train.pickle",
download=False, sweep_type="txt")
data_valid[3]["t8r3v9or"]
horizons = list(sorted(data_valid.keys()))
horizons
```
## Ensemble
```
# export
def fget_preds(subdict, limit_date=False, lcompdate=None, ucompdate=None):
"""
stack predictions over ensemble runs
get predictions and targets from runs
shape: (90, 20729, 3) (n_ensemble_runs, n_windows, H)
limit_date to crop for comparison with cls, esa etc.
"""
# shorter lookbacks have more predictions
# need to ensure all predictions are of same length and are combining the right set of predictions
common_idx = []
for run_id in subdict.keys():
common_idx.append(len(subdict[run_id]["pred_date"]))
idx = min(common_idx)
# get common targets and prediction dates
keys = list(subdict.keys())
targs = subdict[keys[0]]["targs_denorm"][-idx:]
epoch = subdict[keys[0]]["pred_date"][-idx:]
if limit_date:
try:
larg = np.argwhere(epoch==lcompdate)[0][0]
except IndexError:
larg = 0
try:
uarg = np.argwhere(epoch==ucompdate)[0][0] + 1
except IndexError:
uarg = -1
targs = targs[larg:uarg]
epoch = epoch[larg:uarg]
# get predictions for each run
preds = []
for run_id in subdict.keys():
if limit_date:
preds.append(subdict[run_id]["preds_denorm"][-idx:][larg:uarg])
else:
preds.append(subdict[run_id]["preds_denorm"][-idx:])
print(np.array(preds).shape, epoch[0], epoch[-1])
return np.array(epoch), np.array(preds), np.array(targs)
# export
def fensemble(preds_valid, mode="mean"):
"""
simple mean/median ensemble, std over ensemble
"""
# reshape
stack = np.dstack(preds_valid)
if mode == "mean":
ensemble_preds = np.mean(stack, axis=2)
elif mode == "median":
ensemble_preds = np.median(stack, axis=2)
# variance in ensemble
ensemble_std = np.std(stack, axis=2)
#print(ensemble_std.shape)
return ensemble_preds, ensemble_std
def fensemble_linear_regression(preds_valid, targs_valid, preds_train, targs_train):
"""
learn linear regression coefficients from training data (one for each day over horizon, H)
apply to combine validation data
X = np.array([[1, 1], [1, 2], [2, 2], [2, 3]])
# y = 1 * x_0 + 2 * x_1 + 3
y = np.dot(X, np.array([1, 2])) + 3
reg = LinearRegression().fit(X, y)
reg.coef_ = array([1., 2.])
reg.predict(np.array([[3, 5]])) = array([16.])
"""
ensemble_preds = []
H = preds_valid.shape[2]
for h in range(H):
# should be 90 (n_ensemble_runs) coefficients
X = preds_train[:,:,h].T
y = targs_train[:,h]
reg = LinearRegression(fit_intercept=False).fit(X, y)
X_valid = preds_valid[:,:,h].T
y_valid = targs_valid[:,h]
#print(reg.predict(X_valid), np.dot(X_valid, reg.coef_) + reg.intercept_, y_valid) # np.sum(np.multiply(X[-1], reg.coef_)) + intercept
print(h, reg.score(X, y), reg.score(X_valid, y_valid))
ensemble_preds.append(np.dot(X_valid, reg.coef_) + reg.intercept_)
# not yet implemented with uncertainty
return np.stack(ensemble_preds).T
# export
def fensemble_boosting_regressor(preds_valid, targs_valid, preds_train, targs_train, alpha=0.9):
"""
Learn combination of ensemble members from training data using Gradient Boosting Regression
Also provides prediction intervals (using quantile regression)
alpha = % prediction interval
https://scikit-learn.org/stable/auto_examples/ensemble/plot_gradient_boosting_quantile.html
https://towardsdatascience.com/how-to-generate-prediction-intervals-with-scikit-learn-and-python-ab3899f992ed
"""
ensemble_preds = []
ensemble_lower = []
ensemble_upper = []
H = preds_valid.shape[2]
# run for each day over horizon
for h in range(H):
X_train = preds_train[:,:,h].T
y_train = targs_train[:,h]
X_test = preds_valid[:,:,h].T
y_test = targs_valid[:,h]
upper_model = GradientBoostingRegressor(loss="quantile", alpha=alpha)
mid_model = GradientBoostingRegressor(loss="ls")
lower_model = GradientBoostingRegressor(loss="quantile", alpha=(1.0-alpha))
# fit models
lower_model.fit(X_train, y_train)
mid_model.fit(X_train, y_train)
upper_model.fit(X_train, y_train)
# store predictions
ensemble_preds.append(mid_model.predict(X_test))
ensemble_lower.append(lower_model.predict(X_test))
ensemble_upper.append(upper_model.predict(X_test))
return np.stack(ensemble_preds).T, np.stack(ensemble_lower).T, np.stack(ensemble_upper).T
limit_date = False
lcompdate = None
ucompdate = None
predictions = {}
for horizon in horizons:
print("Horizon: {}".format(horizon))
# get training (if learning weights) and validation predictions and targets
epoch_train, preds_train, targs_train = fget_preds(data_train[horizon])
epoch_valid, preds_valid, targs_valid = fget_preds(data_valid[horizon], limit_date, lcompdate, ucompdate)
ensemble_preds, ensemble_std = fensemble(preds_valid, mode="mean")
print(ensemble_preds.shape, ensemble_std.shape)
# ensemble and store predictions and uncertainty
predictions[horizon] = [epoch_valid, targs_valid, ensemble_preds, ensemble_std]
%%time
limit_date = False
lcompdate = None
ucompdate = None
mode = "mean" # "median", "boosting_regressor"
# read from pickled file (need for "boosting_regressor" as slow to train)
use_pickle = False
fname = "./wandb_ensemble/ensemble/ensemble_preds_{}_{}-{}.pickle".format(mode, lcompdate, ucompdate)
if use_pickle:
try:
with open(fname, 'rb') as handle:
predictions = pickle.load(handle)
except FileNotFoundError:
print("Pickle file does not exist.")
use_pickle = False
if not use_pickle:
predictions = {}
for horizon in horizons:
print("Horizon: {}".format(horizon))
# get training (if learning weights) and validation predictions and targets
epoch_train, preds_train, targs_train = fget_preds(data_train[horizon])
epoch_valid, preds_valid, targs_valid = fget_preds(data_valid[horizon], limit_date, lcompdate, ucompdate)
# ensemble and store predictions and uncertainty or prediction interval
if mode == "boosting_regressor"
ensemble_preds, ensemble_lower, ensemble_upper = fensemble_boosting_regressor(preds_valid, targs_valid,
preds_train, targs_train,
alpha=0.95)
predictions[horizon] = [epoch_valid, targs_valid, ensemble_preds, ensemble_lower, ensemble_upper]
else:
ensemble_preds, ensemble_std = fensemble(preds_valid, mode=mode)
predictions[horizon] = [epoch_valid, targs_valid, ensemble_preds, ensemble_std]
# if pickle does not already exist, generate
if not Path(fname).is_file():
with open(fname, 'wb') as handle:
pickle.dump(predictions, handle, protocol=pickle.HIGHEST_PROTOCOL)
predictions
```
## Metrics
```
# export
def fmse(y_true, y_pred):
"""
Mean square error.
(from sklearn.metrics import mean_squared_error)
"""
# return np.sum( (np.array(y_true) - np.array(y_pred))**2 ) / len(y_true)
return mean_squared_error(y_true, y_pred)
def fmape(y_true, y_pred):
"""
mean_absolute_percentage_error.
(can cause division-by-zero errors)
"""
y_true, y_pred = np.array(y_true), np.array(y_pred)
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
def fcc_pearsonr(y_true, y_pred):
"""
pearson correlation coefficient
"""
try:
return pearsonr(y_true, y_pred)
except TypeError:
return pearsonr(y_true.flatten(), y_pred.flatten())
def frmse(y_true, y_pred):
"""
Root mean square error.
(from sklearn.metrics import mean_squared_error)
"""
return np.sqrt(mean_squared_error(y_true, y_pred))
def fmae(y_true, y_pred):
"""
mean absolute error
"""
#return np.sum( np.abs(np.array(y_true) - np.array(y_pred)) ) / len(y_true)
return mean_absolute_error(y_true, y_pred)
def fme(y_true, y_pred):
"""
bias
see Liemohn 2018 (model - observed)
"""
return np.sum( np.array(y_pred) - np.array(y_true) ) / len(y_true)
# export
def fmse_std(y_true, y_pred):
"""
Mean square error.
(from sklearn.metrics import mean_squared_error)
There mse stds are massive: would need to plot using rmse
"""
se = (np.array(y_true) - np.array(y_pred))**2
mse = np.mean(se)#, axis=0
sdse = np.std(se)#, axis=0
return sdse
def fmape_std(y_true, y_pred):
"""
mean_absolute_percentage_error.
(can cause division-by-zero errors)
"""
y_true, y_pred = np.array(y_true), np.array(y_pred)
ape = np.abs((y_true - y_pred) / y_true)
mape = np.mean(ape) * 100
sdape = np.std(ape) * 100
return sdape
def fcc_pearsonr_std(y_true, y_pred):
"""
pearson correlation coefficient
"""
return np.nan
def frmse_std(y_true, y_pred):
"""
Root mean square error.
(from sklearn.metrics import mean_squared_error)
"""
se = (np.array(y_true) - np.array(y_pred))**2
mse = np.mean(se)#, axis=0
sdse = np.std(se)#, axis=0
return np.sqrt(sdse)
def fmae_std(y_true, y_pred):
"""
mean absolute error
"""
ae = np.abs(np.array(y_true) - np.array(y_pred))
mae = np.mean(ae)#, axis=0
sdae = np.std(ae)#, axis=0
return sdae
def fme_std(y_true, y_pred):
"""
bias
see Liemohn 2018 (model - observed)
"""
e = np.array(y_pred) - np.array(y_true)
me = np.mean(e)#, axis=0
sde = np.std(e)#, axis=0
return sde
# export
def fget_metrics(y_pred, y_true):
return [fmse(y_true, y_pred), fmape(y_true, y_pred), fcc_pearsonr(y_true, y_pred)[0],
frmse(y_true, y_pred), fmae(y_true, y_pred), fme(y_true, y_pred)]
def fget_metrics_std(y_pred, y_true):
return [fmse_std(y_true, y_pred), fmape_std(y_true, y_pred), fcc_pearsonr_std(y_true, y_pred),
frmse_std(y_true, y_pred), fmae_std(y_true, y_pred), fme_std(y_true, y_pred)]
```
## Compare to external data
```
# export
def fget_external_forecasts(config):
"""
generate dataframe containing forecasts and "truths" for external sources
get persistence
test:
#config = AttrDict()
#config.update(user_config)
#fget_external_forecasts(config)
"""
# ESA
if "esa" in config.data_comp:
dataobj_esa = cESA_SWE()
# read in archive data
df = dataobj_esa.fget_data(filenames=config.esa_archive_fname)[config.esa_archive_key]
df.set_index('ds', inplace=True)
udate_a = (dt.datetime.strptime(config.date_ulim, "%Y-%m-%d") + dt.timedelta(days=27)).strftime("%Y-%m-%d")
df_lim = df[config.date_llim:udate_a]
#dfa_esa = dataobj_esa.finterpolate(df_lim, config.interp_freq)
df_daily = dataobj_esa.fget_daily(df_lim, config.get_daily_method)
dfa_esa = dataobj_esa.fmissing_data(df_daily, config.missing_data_method)
# read in esa forecast data
dff_esa = dataobj_esa.fget_data(filenames=config.esa_forecast_fname)[config.esa_forecast_key]
# add "truth" from archive to forecast
dff_comp_esa = dataobj_esa.fget_forecast_comp(dff_esa, dfa_esa, cname="y")
# rename columns
#dfa_esa.columns = ['gendate' , 'ds', 'y_esa_true']
dff_comp_esa.columns = ['gendate' , 'ds', 'y_esa', 'y_esa_true']
# Calculate persistence
#dff_comp_esa = dataobj_esa.fget_persistence(dfa_esa, 'y_esa_true', "persistence_esa")
dff_comp_esa = dataobj_esa.fget_persistence(dff_comp_esa, 'y_esa_true', "persistence_esa")
# CLS-CNES
if "cls" in config.data_comp:
dataobj_cls = cCLS_CNES()
# read in archive data and restrict to ds and key variable
# need to ensure upper date for archive is 30 days ahead of upper date of forecast gendate
udate_a = (dt.datetime.strptime(config.cls_forecast_udate, "%Y-%m-%d") + dt.timedelta(days=30)).strftime("%Y-%m-%d")
dfa_cls = dataobj_cls.fget_archive_data(config.cls_datadir, config.cls_forecast_ldate, udate_a)
dfa_cls = dfa_cls[['ds',config.cls_key]]
dfa_cls = dfa_cls.set_index("ds")
# read in forecast data and restrict to key variable
dff_cls = dataobj_cls.fget_forecast_data(config.cls_datadir, config.cls_forecast_ldate, config.cls_forecast_udate)
dff_cls = dff_cls[['gendate', 'ds', "{}_c".format(config.cls_key)]]
# add "truth" from archive to forecast
dff_comp_cls = dataobj_cls.fget_forecast_comp(dff_cls, dfa_cls, cname=config.cls_key)
# rename columns
dff_comp_cls.columns = ['gendate' , 'ds', 'y_cls', 'y_cls_true']
# Calculate persistence
dff_comp_cls = dataobj_cls.fget_persistence(dff_comp_cls, 'y_cls_true', "persistence_cls")
# botch
# include esa persistence in cls only table
if "esa" not in config.data_comp:
dataobj_esa = cESA_SWE()
# read in archive data
df = dataobj_esa.fget_data(filenames=config.esa_archive_fname)[config.esa_archive_key]
df.set_index('ds', inplace=True)
ldate = (dt.datetime.strptime(config.cls_forecast_ldate, "%Y-%m-%d") - dt.timedelta(days=1)).strftime("%Y-%m-%d")
df_lim = df[ldate:udate_a] # hard code as takes 1 day later for some reason
#dfa_esa = dataobj_esa.finterpolate(df_lim, config.interp_freq)
df_daily = dataobj_esa.fget_daily(df_lim, config.get_daily_method)
dfa_esa = dataobj_esa.fmissing_data(df_daily, config.missing_data_method)
# add "truth" from archive to forecast
dff_comp_cls1 = dataobj_cls.fget_forecast_comp(dff_cls, dfa_esa, cname="y_cls")
dff_comp_cls1.columns = ['gendate' , 'ds', 'y_cls', 'y_esa_true']
# drop gendates that don't give full forcast (depends on horizon)
dff_comp_cls1 = dff_comp_cls1[dff_comp_cls1.groupby('gendate').gendate.transform('count')>=27].copy()
# Calculate persistence
dff_comp_cls1 = dataobj_cls.fget_persistence(dff_comp_cls1, 'y_esa_true', "persistence_esa")
dff_comp_cls = pd.merge(dff_comp_cls, dff_comp_cls1, on=['gendate', 'ds', 'y_cls'])
# COMBINE AND RETURN
if ("esa" in config.data_comp) and ("cls" in config.data_comp):
df_comp = pd.merge(dff_comp_esa, dff_comp_cls, on=['gendate', 'ds'])
return df_comp
elif "esa" in config.data_comp:
return dff_comp_esa
elif "cls" in config.data_comp:
return dff_comp_cls
# export
def fget_external_metrics(dff, config):
metrics = {}
metrics_std = {}
if "esa" in config.data_comp:
metrics["ESA"] = fget_metrics(dff.y_esa, dff.y_esa_true)
metrics_std["ESA"] = fget_metrics_std(dff.y_esa, dff.y_esa_true)
if "cls" in config.data_comp:
metrics["CLS"] = fget_metrics(dff.y_cls, dff.y_cls_true)
metrics_std["CLS"] = fget_metrics_std(dff.y_cls, dff.y_cls_true)
metrics["PERSISTENCE"] = fget_metrics(dff.persistence_esa, dff.y_esa_true)
metrics_std["PERSISTENCE"] = fget_metrics_std(dff.persistence_esa, dff.y_esa_true)
#metrics["PERSISTENCE"] = fget_metrics(dff.persistence_cls, dff.y_cls_true)
return metrics, metrics_std
```
## Export
```
from nbdev.export import *
notebook2script()
```
| github_jupyter |
Importing the Dependencies
```
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
```
Data Collection and Processing
```
# loading the csv data to a Pandas DataFrame
heart_data = pd.read_csv('heart.csv')
# print first 5 rows of the dataset
heart_data.head()
# print last 5 rows of the dataset
heart_data.tail()
# number of rows and columns in the dataset
heart_data.shape
# getting some info about the data
heart_data.info()
# checking for missing values
heart_data.isnull().sum()
# statistical measures about the data
heart_data.describe()
# checking the distribution of Target Variable
heart_data['target'].value_counts()
```
1 --> Defective Heart
0 --> Healthy Heart
Splitting the Features and Target
```
X = heart_data.drop(columns='target', axis=1)
Y = heart_data['target']
print(X)
print(Y)
```
Splitting the Data into Training data & Test Data
```
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.2, stratify=Y, random_state=2)
#X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.2, random_state=2)
print(X.shape, X_train.shape, X_test.shape)
```
Model Training
Logistic Regression
```
model = LogisticRegression()
# training the LogisticRegression model with Training data
model.fit(X_train, Y_train)
```
Model Evaluation
Accuracy Score
```
# accuracy on training data
X_train_prediction = model.predict(X_train)
training_data_accuracy = accuracy_score(X_train_prediction, Y_train)
print('Accuracy on Training data : ', training_data_accuracy)
# accuracy on test data
X_test_prediction = model.predict(X_test)
test_data_accuracy = accuracy_score(X_test_prediction, Y_test)
print('Accuracy on Test data : ', test_data_accuracy)
```
Building a Predictive System
```
input_data = (62,0,0,140,268,0,0,160,0,3.6,0,2,2)
# change the input data to a numpy array
input_data_as_numpy_array= np.asarray(input_data)
# reshape the numpy array as we are predicting for only on instance
input_data_reshaped = input_data_as_numpy_array.reshape(1,-1)
prediction = model.predict(input_data_reshaped)
print(prediction)
if (prediction[0]== 0):
print('The Person does not have a Heart Disease')
else:
print('The Person has Heart Disease')
```
saving the model
```
import pickle
with open('heart_disease_model','wb') as file:
pickle.dump(model,file)
#reading the model
with open('heart_disease_model','rb') as file:
model = pickle.load(file)
model.predict([[1,2,3,4,5,6,7,8,9,10,11,12,13]])
```
| github_jupyter |
# Ex1: Ring Slot
The Vector Fitting feature is demonstrated using the *ring slot* example network from the scikit-rf `data` folder. Additional explanations and background information can be found in the [Vector Fitting tutorial](../../tutorials/VectorFitting.ipynb).
```
import skrf
import numpy as np
import matplotlib.pyplot as mplt
```
To create a `VectorFitting` instance, a `Network` containing the frequency responses of the N-port is passed. In this example the *ring slot* is used, which can be loaded directly as a `Network` from scikit-rf:
```
nw = skrf.data.ring_slot
vf = skrf.VectorFitting(nw)
```
Now, the vector fit can be performed. The number of poles has to be specified, which depends on the *behaviour* of the responses. A smooth response would only require very few poles (2-5). In this case, 3 real poles are sufficient:
```
vf.vector_fit(n_poles_real=3, n_poles_cmplx=0)
```
As printed in the logging output (not shown), the pole relocation process converged quickly after just 5 iteration steps. This can also be checked with the convergence plot:
```
vf.plot_convergence()
```
The fitted model parameters are now stored in the class variables `poles`, `zeros`, `proportional_coeff` and `constant_coeff` for further use. To verify the result, the model response can be compared to the original network response. As the model will return a response at any given frequency, it makes sense to also check its response outside the frequency range of the original samples:
```
freqs1 = np.linspace(0, 200e9, 201)
fig, ax = mplt.subplots(2, 2)
fig.set_size_inches(12, 8)
vf.plot_s_mag(0, 0, freqs1, ax=ax[0][0]) # plot s11
vf.plot_s_mag(1, 0, freqs1, ax=ax[1][0]) # plot s21
vf.plot_s_mag(0, 1, freqs1, ax=ax[0][1]) # plot s12
vf.plot_s_mag(1, 1, freqs1, ax=ax[1][1]) # plot s22
fig.tight_layout()
mplt.show()
```
To use the model in a circuit simulation, an equivalent circuit can be created based on the fitting parameters. This is currently only implemented for SPICE, but the structure of the equivalent circuit can be adopted to any kind of circuit simulator.
`vf1.write_spice_subcircuit_s('/home/vinc/Desktop/ring_slot.sp')`
For a quick test, the subcircuit is included in a schematic in [QUCS-S](https://ra3xdh.github.io/) for AC simulation and S-parameter calculation based on the port voltages and currents (see the equations):
<img src="./ngspice_ringslot_schematic.svg" />
The simulation outputs from [ngspice](http://ngspice.sourceforge.net/) compare well to the plots above:
<img src="./ngspice_ringslot_sp_mag.svg" />
<img src="./ngspice_ringslot_sp_smith.svg" />
| github_jupyter |
# One-class SVM
### Credit Card Fraud Detection
#### A One-Class Support Vector Machine is an unsupervised learning algorithm that is trained only on the ‘normal’ data, It learns the boundaries of these points and is therefore able to classify any points that lie outside the boundary as outliers.
```
import numpy as np
import pandas as pd
from sklearn import svm
df=pd.read_csv("creditcard.csv")
df.head()
df.info()
#Renaming the class labels as labels
df=df.rename(columns={'Class':'Label'})
#visualizing the data
import seaborn as sns
import matplotlib.pyplot as plt
plt.style.use("ggplot")
sns.FacetGrid(df, hue="Label", size = 6).map(plt.scatter, "Time", "Amount", edgecolor="k").add_legend()
plt.show()
fig,(ax1,ax2)=plt.subplots(ncols=2, figsize=(12,6))
s=sns.boxplot(ax=ax1,x="Label",y="Amount",hue="Label",data=df,palette="PRGn",showfliers=True)
s=sns.boxplot(ax=ax2,x="Label",y="Amount",hue="Label",data=df,palette="PRGn",showfliers=False)
plt.show()
normal=df.loc[df.Label==0]
anamalous=df.loc[df.Label==1]
#In one class SVM, The model is trained with only one class,here model is trained only with observations of normal transactions only.
#remaing will be considered in test set
train_df=normal.loc[0:200000,:]
train_df=train_df.drop('Label',1)
y1=normal.loc[200000:,'Label'] #labels with normal transaction
y2=anamalous['Label']
x_test1=normal.loc[200000:,:].drop('Label',1)
x_test2=anamalous.drop('Label',1)
x_test=x_test1.append(x_test2) #test set of features
oneclass = svm.OneClassSVM(kernel='linear', gamma=0.001, nu=0.95)
y_test1=normal.loc[200000:,'Label']
y_test2=anamalous['Label']
y_test=y_test1.append(y_test2)
oneclass.fit(train_df)
fraud_prediction=oneclass.predict(x_test)
#number of outliers predicted by the model
unique,counts=np.unique(fraud_prediction,return_counts=True)
print(np.asarray((unique, counts)).T)
y_test=y_test.to_frame()
y_test=y_test.reset_index()
fraud_prediction=pd.DataFrame(fraud_prediction)
fraud_prediction=fraud_prediction.rename(columns={0:'prediction'})
#TP: True Positive
#TN: True Negative
#FP: False Positve
#FN: False Negative
TP=FP=TN=FN=0
for i in range(len(y_test)):
if y_test['Label'][i]== 0 and fraud_prediction['prediction'][i] == 1:
TP = TP+1
elif y_test['Label'][i]== 0 and fraud_prediction['prediction'][i] == -1:
FN = FN+1
elif y_test['Label'][i]== 1 and fraud_prediction['prediction'][i] == 1:
FP = FP+1
else:
TN = TN +1
print (TP,FN,FP,TN)
accuracy=(TP+TN)/(TP+FN+FP+TN)
sensitivity=TP/(TP+FN)
specificity=TN/(TN+FP)
print("Accuracy : ",accuracy)
print("Sensitivity: ",sensitivity)
print("Specificity: ",specificity)
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.