text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Convolutional Neural Networks: Step by Step
Welcome to Course 4's first assignment! In this assignment, you will implement convolutional (CONV) and pooling (POOL) layers in numpy, including both forward propagation and (optionally) backward propagation.
**Notation**:
- Superscript $[l]$ denotes an object of the $l^{th}$ layer.
- Example: $a^{[4]}$ is the $4^{th}$ layer activation. $W^{[5]}$ and $b^{[5]}$ are the $5^{th}$ layer parameters.
- Superscript $(i)$ denotes an object from the $i^{th}$ example.
- Example: $x^{(i)}$ is the $i^{th}$ training example input.
- Subscript $i$ denotes the $i^{th}$ entry of a vector.
- Example: $a^{[l]}_i$ denotes the $i^{th}$ entry of the activations in layer $l$, assuming this is a fully connected (FC) layer.
- $n_H$, $n_W$ and $n_C$ denote respectively the height, width and number of channels of a given layer. If you want to reference a specific layer $l$, you can also write $n_H^{[l]}$, $n_W^{[l]}$, $n_C^{[l]}$.
- $n_{H_{prev}}$, $n_{W_{prev}}$ and $n_{C_{prev}}$ denote respectively the height, width and number of channels of the previous layer. If referencing a specific layer $l$, this could also be denoted $n_H^{[l-1]}$, $n_W^{[l-1]}$, $n_C^{[l-1]}$.
We assume that you are already familiar with `numpy` and/or have completed the previous courses of the specialization. Let's get started!
## <font color='darkblue'>Updates</font>
#### If you were working on the notebook before this update...
* The current notebook is version "v2a".
* You can find your original work saved in the notebook with the previous version name ("v2")
* To view the file directory, go to the menu "File->Open", and this will open a new tab that shows the file directory.
#### List of updates
* clarified example used for padding function. Updated starter code for padding function.
* `conv_forward` has additional hints to help students if they're stuck.
* `conv_forward` places code for `vert_start` and `vert_end` within the `for h in range(...)` loop; to avoid redundant calculations. Similarly updated `horiz_start` and `horiz_end`. **Thanks to our mentor Kevin Brown for pointing this out.**
* `conv_forward` breaks down the `Z[i, h, w, c]` single line calculation into 3 lines, for clarity.
* `conv_forward` test case checks that students don't accidentally use n_H_prev instead of n_H, use n_W_prev instead of n_W, and don't accidentally swap n_H with n_W
* `pool_forward` properly nests calculations of `vert_start`, `vert_end`, `horiz_start`, and `horiz_end` to avoid redundant calculations.
* `pool_forward' has two new test cases that check for a correct implementation of stride (the height and width of the previous layer's activations should be large enough relative to the filter dimensions so that a stride can take place).
* `conv_backward`: initialize `Z` and `cache` variables within unit test, to make it independent of unit testing that occurs in the `conv_forward` section of the assignment.
* **Many thanks to our course mentor, Paul Mielke, for proposing these test cases.**
## 1 - Packages
Let's first import all the packages that you will need during this assignment.
- [numpy](www.numpy.org) is the fundamental package for scientific computing with Python.
- [matplotlib](http://matplotlib.org) is a library to plot graphs in Python.
- np.random.seed(1) is used to keep all the random function calls consistent. It will help us grade your work.
```
import numpy as np
import h5py
import matplotlib.pyplot as plt
%matplotlib inline
plt.rcParams['figure.figsize'] = (5.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
%load_ext autoreload
%autoreload 2
np.random.seed(1)
```
## 2 - Outline of the Assignment
You will be implementing the building blocks of a convolutional neural network! Each function you will implement will have detailed instructions that will walk you through the steps needed:
- Convolution functions, including:
- Zero Padding
- Convolve window
- Convolution forward
- Convolution backward (optional)
- Pooling functions, including:
- Pooling forward
- Create mask
- Distribute value
- Pooling backward (optional)
This notebook will ask you to implement these functions from scratch in `numpy`. In the next notebook, you will use the TensorFlow equivalents of these functions to build the following model:
<img src="images/model.png" style="width:800px;height:300px;">
**Note** that for every forward function, there is its corresponding backward equivalent. Hence, at every step of your forward module you will store some parameters in a cache. These parameters are used to compute gradients during backpropagation.
## 3 - Convolutional Neural Networks
Although programming frameworks make convolutions easy to use, they remain one of the hardest concepts to understand in Deep Learning. A convolution layer transforms an input volume into an output volume of different size, as shown below.
<img src="images/conv_nn.png" style="width:350px;height:200px;">
In this part, you will build every step of the convolution layer. You will first implement two helper functions: one for zero padding and the other for computing the convolution function itself.
### 3.1 - Zero-Padding
Zero-padding adds zeros around the border of an image:
<img src="images/PAD.png" style="width:600px;height:400px;">
<caption><center> <u> <font color='purple'> **Figure 1** </u><font color='purple'> : **Zero-Padding**<br> Image (3 channels, RGB) with a padding of 2. </center></caption>
The main benefits of padding are the following:
- It allows you to use a CONV layer without necessarily shrinking the height and width of the volumes. This is important for building deeper networks, since otherwise the height/width would shrink as you go to deeper layers. An important special case is the "same" convolution, in which the height/width is exactly preserved after one layer.
- It helps us keep more of the information at the border of an image. Without padding, very few values at the next layer would be affected by pixels as the edges of an image.
**Exercise**: Implement the following function, which pads all the images of a batch of examples X with zeros. [Use np.pad](https://docs.scipy.org/doc/numpy/reference/generated/numpy.pad.html). Note if you want to pad the array "a" of shape $(5,5,5,5,5)$ with `pad = 1` for the 2nd dimension, `pad = 3` for the 4th dimension and `pad = 0` for the rest, you would do:
```python
a = np.pad(a, ((0,0), (1,1), (0,0), (3,3), (0,0)), mode='constant', constant_values = (0,0))
```
```
# GRADED FUNCTION: zero_pad
def zero_pad(X, pad):
"""
Pad with zeros all images of the dataset X. The padding is applied to the height and width of an image,
as illustrated in Figure 1.
Argument:
X -- python numpy array of shape (m, n_H, n_W, n_C) representing a batch of m images
pad -- integer, amount of padding around each image on vertical and horizontal dimensions
Returns:
X_pad -- padded image of shape (m, n_H + 2*pad, n_W + 2*pad, n_C)
"""
### START CODE HERE ### (≈ 1 line)
X_pad = None
### END CODE HERE ###
return X_pad
np.random.seed(1)
x = np.random.randn(4, 3, 3, 2)
x_pad = zero_pad(x, 2)
print ("x.shape =\n", x.shape)
print ("x_pad.shape =\n", x_pad.shape)
print ("x[1,1] =\n", x[1,1])
print ("x_pad[1,1] =\n", x_pad[1,1])
fig, axarr = plt.subplots(1, 2)
axarr[0].set_title('x')
axarr[0].imshow(x[0,:,:,0])
axarr[1].set_title('x_pad')
axarr[1].imshow(x_pad[0,:,:,0])
```
**Expected Output**:
```
x.shape =
(4, 3, 3, 2)
x_pad.shape =
(4, 7, 7, 2)
x[1,1] =
[[ 0.90085595 -0.68372786]
[-0.12289023 -0.93576943]
[-0.26788808 0.53035547]]
x_pad[1,1] =
[[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]]
```
### 3.2 - Single step of convolution
In this part, implement a single step of convolution, in which you apply the filter to a single position of the input. This will be used to build a convolutional unit, which:
- Takes an input volume
- Applies a filter at every position of the input
- Outputs another volume (usually of different size)
<img src="images/Convolution_schematic.gif" style="width:500px;height:300px;">
<caption><center> <u> <font color='purple'> **Figure 2** </u><font color='purple'> : **Convolution operation**<br> with a filter of 3x3 and a stride of 1 (stride = amount you move the window each time you slide) </center></caption>
In a computer vision application, each value in the matrix on the left corresponds to a single pixel value, and we convolve a 3x3 filter with the image by multiplying its values element-wise with the original matrix, then summing them up and adding a bias. In this first step of the exercise, you will implement a single step of convolution, corresponding to applying a filter to just one of the positions to get a single real-valued output.
Later in this notebook, you'll apply this function to multiple positions of the input to implement the full convolutional operation.
**Exercise**: Implement conv_single_step(). [Hint](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.sum.html).
**Note**: The variable b will be passed in as a numpy array. If we add a scalar (a float or integer) to a numpy array, the result is a numpy array. In the special case when a numpy array contains a single value, we can cast it as a float to convert it to a scalar.
```
# GRADED FUNCTION: conv_single_step
def conv_single_step(a_slice_prev, W, b):
"""
Apply one filter defined by parameters W on a single slice (a_slice_prev) of the output activation
of the previous layer.
Arguments:
a_slice_prev -- slice of input data of shape (f, f, n_C_prev)
W -- Weight parameters contained in a window - matrix of shape (f, f, n_C_prev)
b -- Bias parameters contained in a window - matrix of shape (1, 1, 1)
Returns:
Z -- a scalar value, the result of convolving the sliding window (W, b) on a slice x of the input data
"""
### START CODE HERE ### (≈ 2 lines of code)
# Element-wise product between a_slice_prev and W. Do not add the bias yet.
s = None
# Sum over all entries of the volume s.
Z = None
# Add bias b to Z. Cast b to a float() so that Z results in a scalar value.
Z = None
### END CODE HERE ###
return Z
np.random.seed(1)
a_slice_prev = np.random.randn(4, 4, 3)
W = np.random.randn(4, 4, 3)
b = np.random.randn(1, 1, 1)
Z = conv_single_step(a_slice_prev, W, b)
print("Z =", Z)
```
**Expected Output**:
<table>
<tr>
<td>
**Z**
</td>
<td>
-6.99908945068
</td>
</tr>
</table>
### 3.3 - Convolutional Neural Networks - Forward pass
In the forward pass, you will take many filters and convolve them on the input. Each 'convolution' gives you a 2D matrix output. You will then stack these outputs to get a 3D volume:
<center>
<video width="620" height="440" src="images/conv_kiank.mp4" type="video/mp4" controls>
</video>
</center>
**Exercise**:
Implement the function below to convolve the filters `W` on an input activation `A_prev`.
This function takes the following inputs:
* `A_prev`, the activations output by the previous layer (for a batch of m inputs);
* Weights are denoted by `W`. The filter window size is `f` by `f`.
* The bias vector is `b`, where each filter has its own (single) bias.
Finally you also have access to the hyperparameters dictionary which contains the stride and the padding.
**Hint**:
1. To select a 2x2 slice at the upper left corner of a matrix "a_prev" (shape (5,5,3)), you would do:
```python
a_slice_prev = a_prev[0:2,0:2,:]
```
Notice how this gives a 3D slice that has height 2, width 2, and depth 3. Depth is the number of channels.
This will be useful when you will define `a_slice_prev` below, using the `start/end` indexes you will define.
2. To define a_slice you will need to first define its corners `vert_start`, `vert_end`, `horiz_start` and `horiz_end`. This figure may be helpful for you to find out how each of the corner can be defined using h, w, f and s in the code below.
<img src="images/vert_horiz_kiank.png" style="width:400px;height:300px;">
<caption><center> <u> <font color='purple'> **Figure 3** </u><font color='purple'> : **Definition of a slice using vertical and horizontal start/end (with a 2x2 filter)** <br> This figure shows only a single channel. </center></caption>
**Reminder**:
The formulas relating the output shape of the convolution to the input shape is:
$$ n_H = \lfloor \frac{n_{H_{prev}} - f + 2 \times pad}{stride} \rfloor +1 $$
$$ n_W = \lfloor \frac{n_{W_{prev}} - f + 2 \times pad}{stride} \rfloor +1 $$
$$ n_C = \text{number of filters used in the convolution}$$
For this exercise, we won't worry about vectorization, and will just implement everything with for-loops.
#### Additional Hints if you're stuck
* You will want to use array slicing (e.g.`varname[0:1,:,3:5]`) for the following variables:
`a_prev_pad` ,`W`, `b`
Copy the starter code of the function and run it outside of the defined function, in separate cells.
Check that the subset of each array is the size and dimension that you're expecting.
* To decide how to get the vert_start, vert_end; horiz_start, horiz_end, remember that these are indices of the previous layer.
Draw an example of a previous padded layer (8 x 8, for instance), and the current (output layer) (2 x 2, for instance).
The output layer's indices are denoted by `h` and `w`.
* Make sure that `a_slice_prev` has a height, width and depth.
* Remember that `a_prev_pad` is a subset of `A_prev_pad`.
Think about which one should be used within the for loops.
```
# GRADED FUNCTION: conv_forward
def conv_forward(A_prev, W, b, hparameters):
"""
Implements the forward propagation for a convolution function
Arguments:
A_prev -- output activations of the previous layer,
numpy array of shape (m, n_H_prev, n_W_prev, n_C_prev)
W -- Weights, numpy array of shape (f, f, n_C_prev, n_C)
b -- Biases, numpy array of shape (1, 1, 1, n_C)
hparameters -- python dictionary containing "stride" and "pad"
Returns:
Z -- conv output, numpy array of shape (m, n_H, n_W, n_C)
cache -- cache of values needed for the conv_backward() function
"""
### START CODE HERE ###
# Retrieve dimensions from A_prev's shape (≈1 line)
(m, n_H_prev, n_W_prev, n_C_prev) = None
# Retrieve dimensions from W's shape (≈1 line)
(f, f, n_C_prev, n_C) = None
# Retrieve information from "hparameters" (≈2 lines)
stride = None
pad = None
# Compute the dimensions of the CONV output volume using the formula given above.
# Hint: use int() to apply the 'floor' operation. (≈2 lines)
n_H = None
n_W = None
# Initialize the output volume Z with zeros. (≈1 line)
Z = None
# Create A_prev_pad by padding A_prev
A_prev_pad = None
for i in range(None): # loop over the batch of training examples
a_prev_pad = None # Select ith training example's padded activation
for h in range(None): # loop over vertical axis of the output volume
# Find the vertical start and end of the current "slice" (≈2 lines)
vert_start = None
vert_end = None
for w in range(None): # loop over horizontal axis of the output volume
# Find the horizontal start and end of the current "slice" (≈2 lines)
horiz_start = None
horiz_end = None
for c in range(None): # loop over channels (= #filters) of the output volume
# Use the corners to define the (3D) slice of a_prev_pad (See Hint above the cell). (≈1 line)
a_slice_prev = None
# Convolve the (3D) slice with the correct filter W and bias b, to get back one output neuron. (≈3 line)
weights = None
biases = None
Z[i, h, w, c] = None
### END CODE HERE ###
# Making sure your output shape is correct
assert(Z.shape == (m, n_H, n_W, n_C))
# Save information in "cache" for the backprop
cache = (A_prev, W, b, hparameters)
return Z, cache
np.random.seed(1)
A_prev = np.random.randn(10,5,7,4)
W = np.random.randn(3,3,4,8)
b = np.random.randn(1,1,1,8)
hparameters = {"pad" : 1,
"stride": 2}
Z, cache_conv = conv_forward(A_prev, W, b, hparameters)
print("Z's mean =\n", np.mean(Z))
print("Z[3,2,1] =\n", Z[3,2,1])
print("cache_conv[0][1][2][3] =\n", cache_conv[0][1][2][3])
```
**Expected Output**:
```
Z's mean =
0.692360880758
Z[3,2,1] =
[ -1.28912231 2.27650251 6.61941931 0.95527176 8.25132576
2.31329639 13.00689405 2.34576051]
cache_conv[0][1][2][3] = [-1.1191154 1.9560789 -0.3264995 -1.34267579]
```
Finally, CONV layer should also contain an activation, in which case we would add the following line of code:
```python
# Convolve the window to get back one output neuron
Z[i, h, w, c] = ...
# Apply activation
A[i, h, w, c] = activation(Z[i, h, w, c])
```
You don't need to do it here.
## 4 - Pooling layer
The pooling (POOL) layer reduces the height and width of the input. It helps reduce computation, as well as helps make feature detectors more invariant to its position in the input. The two types of pooling layers are:
- Max-pooling layer: slides an ($f, f$) window over the input and stores the max value of the window in the output.
- Average-pooling layer: slides an ($f, f$) window over the input and stores the average value of the window in the output.
<table>
<td>
<img src="images/max_pool1.png" style="width:500px;height:300px;">
<td>
<td>
<img src="images/a_pool.png" style="width:500px;height:300px;">
<td>
</table>
These pooling layers have no parameters for backpropagation to train. However, they have hyperparameters such as the window size $f$. This specifies the height and width of the $f \times f$ window you would compute a *max* or *average* over.
### 4.1 - Forward Pooling
Now, you are going to implement MAX-POOL and AVG-POOL, in the same function.
**Exercise**: Implement the forward pass of the pooling layer. Follow the hints in the comments below.
**Reminder**:
As there's no padding, the formulas binding the output shape of the pooling to the input shape is:
$$ n_H = \lfloor \frac{n_{H_{prev}} - f}{stride} \rfloor +1 $$
$$ n_W = \lfloor \frac{n_{W_{prev}} - f}{stride} \rfloor +1 $$
$$ n_C = n_{C_{prev}}$$
```
# GRADED FUNCTION: pool_forward
def pool_forward(A_prev, hparameters, mode = "max"):
"""
Implements the forward pass of the pooling layer
Arguments:
A_prev -- Input data, numpy array of shape (m, n_H_prev, n_W_prev, n_C_prev)
hparameters -- python dictionary containing "f" and "stride"
mode -- the pooling mode you would like to use, defined as a string ("max" or "average")
Returns:
A -- output of the pool layer, a numpy array of shape (m, n_H, n_W, n_C)
cache -- cache used in the backward pass of the pooling layer, contains the input and hparameters
"""
# Retrieve dimensions from the input shape
(m, n_H_prev, n_W_prev, n_C_prev) = A_prev.shape
# Retrieve hyperparameters from "hparameters"
f = hparameters["f"]
stride = hparameters["stride"]
# Define the dimensions of the output
n_H = int(1 + (n_H_prev - f) / stride)
n_W = int(1 + (n_W_prev - f) / stride)
n_C = n_C_prev
# Initialize output matrix A
A = np.zeros((m, n_H, n_W, n_C))
### START CODE HERE ###
for i in range(None): # loop over the training examples
for h in range(None): # loop on the vertical axis of the output volume
# Find the vertical start and end of the current "slice" (≈2 lines)
vert_start = None
vert_end = None
for w in range(None): # loop on the horizontal axis of the output volume
# Find the vertical start and end of the current "slice" (≈2 lines)
horiz_start = None
horiz_end = None
for c in range (None): # loop over the channels of the output volume
# Use the corners to define the current slice on the ith training example of A_prev, channel c. (≈1 line)
a_prev_slice = None
# Compute the pooling operation on the slice.
# Use an if statement to differentiate the modes.
# Use np.max and np.mean.
if mode == "max":
A[i, h, w, c] = None
elif mode == "average":
A[i, h, w, c] = None
### END CODE HERE ###
# Store the input and hparameters in "cache" for pool_backward()
cache = (A_prev, hparameters)
# Making sure your output shape is correct
assert(A.shape == (m, n_H, n_W, n_C))
return A, cache
# Case 1: stride of 1
np.random.seed(1)
A_prev = np.random.randn(2, 5, 5, 3)
hparameters = {"stride" : 1, "f": 3}
A, cache = pool_forward(A_prev, hparameters)
print("mode = max")
print("A.shape = " + str(A.shape))
print("A =\n", A)
print()
A, cache = pool_forward(A_prev, hparameters, mode = "average")
print("mode = average")
print("A.shape = " + str(A.shape))
print("A =\n", A)
```
** Expected Output**
```
mode = max
A.shape = (2, 3, 3, 3)
A =
[[[[ 1.74481176 0.90159072 1.65980218]
[ 1.74481176 1.46210794 1.65980218]
[ 1.74481176 1.6924546 1.65980218]]
[[ 1.14472371 0.90159072 2.10025514]
[ 1.14472371 0.90159072 1.65980218]
[ 1.14472371 1.6924546 1.65980218]]
[[ 1.13162939 1.51981682 2.18557541]
[ 1.13162939 1.51981682 2.18557541]
[ 1.13162939 1.6924546 2.18557541]]]
[[[ 1.19891788 0.84616065 0.82797464]
[ 0.69803203 0.84616065 1.2245077 ]
[ 0.69803203 1.12141771 1.2245077 ]]
[[ 1.96710175 0.84616065 1.27375593]
[ 1.96710175 0.84616065 1.23616403]
[ 1.62765075 1.12141771 1.2245077 ]]
[[ 1.96710175 0.86888616 1.27375593]
[ 1.96710175 0.86888616 1.23616403]
[ 1.62765075 1.12141771 0.79280687]]]]
mode = average
A.shape = (2, 3, 3, 3)
A =
[[[[ -3.01046719e-02 -3.24021315e-03 -3.36298859e-01]
[ 1.43310483e-01 1.93146751e-01 -4.44905196e-01]
[ 1.28934436e-01 2.22428468e-01 1.25067597e-01]]
[[ -3.81801899e-01 1.59993515e-02 1.70562706e-01]
[ 4.73707165e-02 2.59244658e-02 9.20338402e-02]
[ 3.97048605e-02 1.57189094e-01 3.45302489e-01]]
[[ -3.82680519e-01 2.32579951e-01 6.25997903e-01]
[ -2.47157416e-01 -3.48524998e-04 3.50539717e-01]
[ -9.52551510e-02 2.68511000e-01 4.66056368e-01]]]
[[[ -1.73134159e-01 3.23771981e-01 -3.43175716e-01]
[ 3.80634669e-02 7.26706274e-02 -2.30268958e-01]
[ 2.03009393e-02 1.41414785e-01 -1.23158476e-02]]
[[ 4.44976963e-01 -2.61694592e-03 -3.10403073e-01]
[ 5.08114737e-01 -2.34937338e-01 -2.39611830e-01]
[ 1.18726772e-01 1.72552294e-01 -2.21121966e-01]]
[[ 4.29449255e-01 8.44699612e-02 -2.72909051e-01]
[ 6.76351685e-01 -1.20138225e-01 -2.44076712e-01]
[ 1.50774518e-01 2.89111751e-01 1.23238536e-03]]]]
```
```
# Case 2: stride of 2
np.random.seed(1)
A_prev = np.random.randn(2, 5, 5, 3)
hparameters = {"stride" : 2, "f": 3}
A, cache = pool_forward(A_prev, hparameters)
print("mode = max")
print("A.shape = " + str(A.shape))
print("A =\n", A)
print()
A, cache = pool_forward(A_prev, hparameters, mode = "average")
print("mode = average")
print("A.shape = " + str(A.shape))
print("A =\n", A)
```
**Expected Output:**
```
mode = max
A.shape = (2, 2, 2, 3)
A =
[[[[ 1.74481176 0.90159072 1.65980218]
[ 1.74481176 1.6924546 1.65980218]]
[[ 1.13162939 1.51981682 2.18557541]
[ 1.13162939 1.6924546 2.18557541]]]
[[[ 1.19891788 0.84616065 0.82797464]
[ 0.69803203 1.12141771 1.2245077 ]]
[[ 1.96710175 0.86888616 1.27375593]
[ 1.62765075 1.12141771 0.79280687]]]]
mode = average
A.shape = (2, 2, 2, 3)
A =
[[[[-0.03010467 -0.00324021 -0.33629886]
[ 0.12893444 0.22242847 0.1250676 ]]
[[-0.38268052 0.23257995 0.6259979 ]
[-0.09525515 0.268511 0.46605637]]]
[[[-0.17313416 0.32377198 -0.34317572]
[ 0.02030094 0.14141479 -0.01231585]]
[[ 0.42944926 0.08446996 -0.27290905]
[ 0.15077452 0.28911175 0.00123239]]]]
```
Congratulations! You have now implemented the forward passes of all the layers of a convolutional network.
The remainder of this notebook is optional, and will not be graded.
## 5 - Backpropagation in convolutional neural networks (OPTIONAL / UNGRADED)
In modern deep learning frameworks, you only have to implement the forward pass, and the framework takes care of the backward pass, so most deep learning engineers don't need to bother with the details of the backward pass. The backward pass for convolutional networks is complicated. If you wish, you can work through this optional portion of the notebook to get a sense of what backprop in a convolutional network looks like.
When in an earlier course you implemented a simple (fully connected) neural network, you used backpropagation to compute the derivatives with respect to the cost to update the parameters. Similarly, in convolutional neural networks you can calculate the derivatives with respect to the cost in order to update the parameters. The backprop equations are not trivial and we did not derive them in lecture, but we will briefly present them below.
### 5.1 - Convolutional layer backward pass
Let's start by implementing the backward pass for a CONV layer.
#### 5.1.1 - Computing dA:
This is the formula for computing $dA$ with respect to the cost for a certain filter $W_c$ and a given training example:
$$ dA += \sum _{h=0} ^{n_H} \sum_{w=0} ^{n_W} W_c \times dZ_{hw} \tag{1}$$
Where $W_c$ is a filter and $dZ_{hw}$ is a scalar corresponding to the gradient of the cost with respect to the output of the conv layer Z at the hth row and wth column (corresponding to the dot product taken at the ith stride left and jth stride down). Note that at each time, we multiply the the same filter $W_c$ by a different dZ when updating dA. We do so mainly because when computing the forward propagation, each filter is dotted and summed by a different a_slice. Therefore when computing the backprop for dA, we are just adding the gradients of all the a_slices.
In code, inside the appropriate for-loops, this formula translates into:
```python
da_prev_pad[vert_start:vert_end, horiz_start:horiz_end, :] += W[:,:,:,c] * dZ[i, h, w, c]
```
#### 5.1.2 - Computing dW:
This is the formula for computing $dW_c$ ($dW_c$ is the derivative of one filter) with respect to the loss:
$$ dW_c += \sum _{h=0} ^{n_H} \sum_{w=0} ^ {n_W} a_{slice} \times dZ_{hw} \tag{2}$$
Where $a_{slice}$ corresponds to the slice which was used to generate the activation $Z_{ij}$. Hence, this ends up giving us the gradient for $W$ with respect to that slice. Since it is the same $W$, we will just add up all such gradients to get $dW$.
In code, inside the appropriate for-loops, this formula translates into:
```python
dW[:,:,:,c] += a_slice * dZ[i, h, w, c]
```
#### 5.1.3 - Computing db:
This is the formula for computing $db$ with respect to the cost for a certain filter $W_c$:
$$ db = \sum_h \sum_w dZ_{hw} \tag{3}$$
As you have previously seen in basic neural networks, db is computed by summing $dZ$. In this case, you are just summing over all the gradients of the conv output (Z) with respect to the cost.
In code, inside the appropriate for-loops, this formula translates into:
```python
db[:,:,:,c] += dZ[i, h, w, c]
```
**Exercise**: Implement the `conv_backward` function below. You should sum over all the training examples, filters, heights, and widths. You should then compute the derivatives using formulas 1, 2 and 3 above.
```
def conv_backward(dZ, cache):
"""
Implement the backward propagation for a convolution function
Arguments:
dZ -- gradient of the cost with respect to the output of the conv layer (Z), numpy array of shape (m, n_H, n_W, n_C)
cache -- cache of values needed for the conv_backward(), output of conv_forward()
Returns:
dA_prev -- gradient of the cost with respect to the input of the conv layer (A_prev),
numpy array of shape (m, n_H_prev, n_W_prev, n_C_prev)
dW -- gradient of the cost with respect to the weights of the conv layer (W)
numpy array of shape (f, f, n_C_prev, n_C)
db -- gradient of the cost with respect to the biases of the conv layer (b)
numpy array of shape (1, 1, 1, n_C)
"""
### START CODE HERE ###
# Retrieve information from "cache"
(A_prev, W, b, hparameters) = None
# Retrieve dimensions from A_prev's shape
(m, n_H_prev, n_W_prev, n_C_prev) = None
# Retrieve dimensions from W's shape
(f, f, n_C_prev, n_C) = None
# Retrieve information from "hparameters"
stride = None
pad = None
# Retrieve dimensions from dZ's shape
(m, n_H, n_W, n_C) = None
# Initialize dA_prev, dW, db with the correct shapes
dA_prev = None
dW = None
db = None
# Pad A_prev and dA_prev
A_prev_pad = None
dA_prev_pad = None
for i in range(None): # loop over the training examples
# select ith training example from A_prev_pad and dA_prev_pad
a_prev_pad = None
da_prev_pad = None
for h in range(None): # loop over vertical axis of the output volume
for w in range(None): # loop over horizontal axis of the output volume
for c in range(None): # loop over the channels of the output volume
# Find the corners of the current "slice"
vert_start = None
vert_end = None
horiz_start = None
horiz_end = None
# Use the corners to define the slice from a_prev_pad
a_slice = None
# Update gradients for the window and the filter's parameters using the code formulas given above
da_prev_pad[vert_start:vert_end, horiz_start:horiz_end, :] += None
dW[:,:,:,c] += None
db[:,:,:,c] += None
# Set the ith training example's dA_prev to the unpadded da_prev_pad (Hint: use X[pad:-pad, pad:-pad, :])
dA_prev[i, :, :, :] = None
### END CODE HERE ###
# Making sure your output shape is correct
assert(dA_prev.shape == (m, n_H_prev, n_W_prev, n_C_prev))
return dA_prev, dW, db
# We'll run conv_forward to initialize the 'Z' and 'cache_conv",
# which we'll use to test the conv_backward function
np.random.seed(1)
A_prev = np.random.randn(10,4,4,3)
W = np.random.randn(2,2,3,8)
b = np.random.randn(1,1,1,8)
hparameters = {"pad" : 2,
"stride": 2}
Z, cache_conv = conv_forward(A_prev, W, b, hparameters)
# Test conv_backward
dA, dW, db = conv_backward(Z, cache_conv)
print("dA_mean =", np.mean(dA))
print("dW_mean =", np.mean(dW))
print("db_mean =", np.mean(db))
```
** Expected Output: **
<table>
<tr>
<td>
**dA_mean**
</td>
<td>
1.45243777754
</td>
</tr>
<tr>
<td>
**dW_mean**
</td>
<td>
1.72699145831
</td>
</tr>
<tr>
<td>
**db_mean**
</td>
<td>
7.83923256462
</td>
</tr>
</table>
## 5.2 Pooling layer - backward pass
Next, let's implement the backward pass for the pooling layer, starting with the MAX-POOL layer. Even though a pooling layer has no parameters for backprop to update, you still need to backpropagation the gradient through the pooling layer in order to compute gradients for layers that came before the pooling layer.
### 5.2.1 Max pooling - backward pass
Before jumping into the backpropagation of the pooling layer, you are going to build a helper function called `create_mask_from_window()` which does the following:
$$ X = \begin{bmatrix}
1 && 3 \\
4 && 2
\end{bmatrix} \quad \rightarrow \quad M =\begin{bmatrix}
0 && 0 \\
1 && 0
\end{bmatrix}\tag{4}$$
As you can see, this function creates a "mask" matrix which keeps track of where the maximum of the matrix is. True (1) indicates the position of the maximum in X, the other entries are False (0). You'll see later that the backward pass for average pooling will be similar to this but using a different mask.
**Exercise**: Implement `create_mask_from_window()`. This function will be helpful for pooling backward.
Hints:
- [np.max()]() may be helpful. It computes the maximum of an array.
- If you have a matrix X and a scalar x: `A = (X == x)` will return a matrix A of the same size as X such that:
```
A[i,j] = True if X[i,j] = x
A[i,j] = False if X[i,j] != x
```
- Here, you don't need to consider cases where there are several maxima in a matrix.
```
def create_mask_from_window(x):
"""
Creates a mask from an input matrix x, to identify the max entry of x.
Arguments:
x -- Array of shape (f, f)
Returns:
mask -- Array of the same shape as window, contains a True at the position corresponding to the max entry of x.
"""
### START CODE HERE ### (≈1 line)
mask = None
### END CODE HERE ###
return mask
np.random.seed(1)
x = np.random.randn(2,3)
mask = create_mask_from_window(x)
print('x = ', x)
print("mask = ", mask)
```
**Expected Output:**
<table>
<tr>
<td>
**x =**
</td>
<td>
[[ 1.62434536 -0.61175641 -0.52817175] <br>
[-1.07296862 0.86540763 -2.3015387 ]]
</td>
</tr>
<tr>
<td>
**mask =**
</td>
<td>
[[ True False False] <br>
[False False False]]
</td>
</tr>
</table>
Why do we keep track of the position of the max? It's because this is the input value that ultimately influenced the output, and therefore the cost. Backprop is computing gradients with respect to the cost, so anything that influences the ultimate cost should have a non-zero gradient. So, backprop will "propagate" the gradient back to this particular input value that had influenced the cost.
### 5.2.2 - Average pooling - backward pass
In max pooling, for each input window, all the "influence" on the output came from a single input value--the max. In average pooling, every element of the input window has equal influence on the output. So to implement backprop, you will now implement a helper function that reflects this.
For example if we did average pooling in the forward pass using a 2x2 filter, then the mask you'll use for the backward pass will look like:
$$ dZ = 1 \quad \rightarrow \quad dZ =\begin{bmatrix}
1/4 && 1/4 \\
1/4 && 1/4
\end{bmatrix}\tag{5}$$
This implies that each position in the $dZ$ matrix contributes equally to output because in the forward pass, we took an average.
**Exercise**: Implement the function below to equally distribute a value dz through a matrix of dimension shape. [Hint](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ones.html)
```
def distribute_value(dz, shape):
"""
Distributes the input value in the matrix of dimension shape
Arguments:
dz -- input scalar
shape -- the shape (n_H, n_W) of the output matrix for which we want to distribute the value of dz
Returns:
a -- Array of size (n_H, n_W) for which we distributed the value of dz
"""
### START CODE HERE ###
# Retrieve dimensions from shape (≈1 line)
(n_H, n_W) = None
# Compute the value to distribute on the matrix (≈1 line)
average = None
# Create a matrix where every entry is the "average" value (≈1 line)
a = None
### END CODE HERE ###
return a
a = distribute_value(2, (2,2))
print('distributed value =', a)
```
**Expected Output**:
<table>
<tr>
<td>
distributed_value =
</td>
<td>
[[ 0.5 0.5]
<br\>
[ 0.5 0.5]]
</td>
</tr>
</table>
### 5.2.3 Putting it together: Pooling backward
You now have everything you need to compute backward propagation on a pooling layer.
**Exercise**: Implement the `pool_backward` function in both modes (`"max"` and `"average"`). You will once again use 4 for-loops (iterating over training examples, height, width, and channels). You should use an `if/elif` statement to see if the mode is equal to `'max'` or `'average'`. If it is equal to 'average' you should use the `distribute_value()` function you implemented above to create a matrix of the same shape as `a_slice`. Otherwise, the mode is equal to '`max`', and you will create a mask with `create_mask_from_window()` and multiply it by the corresponding value of dA.
```
def pool_backward(dA, cache, mode = "max"):
"""
Implements the backward pass of the pooling layer
Arguments:
dA -- gradient of cost with respect to the output of the pooling layer, same shape as A
cache -- cache output from the forward pass of the pooling layer, contains the layer's input and hparameters
mode -- the pooling mode you would like to use, defined as a string ("max" or "average")
Returns:
dA_prev -- gradient of cost with respect to the input of the pooling layer, same shape as A_prev
"""
### START CODE HERE ###
# Retrieve information from cache (≈1 line)
(A_prev, hparameters) = None
# Retrieve hyperparameters from "hparameters" (≈2 lines)
stride = None
f = None
# Retrieve dimensions from A_prev's shape and dA's shape (≈2 lines)
m, n_H_prev, n_W_prev, n_C_prev = None
m, n_H, n_W, n_C = None
# Initialize dA_prev with zeros (≈1 line)
dA_prev = None
for i in range(None): # loop over the training examples
# select training example from A_prev (≈1 line)
a_prev = None
for h in range(None): # loop on the vertical axis
for w in range(None): # loop on the horizontal axis
for c in range(None): # loop over the channels (depth)
# Find the corners of the current "slice" (≈4 lines)
vert_start = None
vert_end = None
horiz_start = None
horiz_end = None
# Compute the backward propagation in both modes.
if mode == "max":
# Use the corners and "c" to define the current slice from a_prev (≈1 line)
a_prev_slice = None
# Create the mask from a_prev_slice (≈1 line)
mask = None
# Set dA_prev to be dA_prev + (the mask multiplied by the correct entry of dA) (≈1 line)
dA_prev[i, vert_start: vert_end, horiz_start: horiz_end, c] += None
elif mode == "average":
# Get the value a from dA (≈1 line)
da = None
# Define the shape of the filter as fxf (≈1 line)
shape = None
# Distribute it to get the correct slice of dA_prev. i.e. Add the distributed value of da. (≈1 line)
dA_prev[i, vert_start: vert_end, horiz_start: horiz_end, c] += None
### END CODE ###
# Making sure your output shape is correct
assert(dA_prev.shape == A_prev.shape)
return dA_prev
np.random.seed(1)
A_prev = np.random.randn(5, 5, 3, 2)
hparameters = {"stride" : 1, "f": 2}
A, cache = pool_forward(A_prev, hparameters)
dA = np.random.randn(5, 4, 2, 2)
dA_prev = pool_backward(dA, cache, mode = "max")
print("mode = max")
print('mean of dA = ', np.mean(dA))
print('dA_prev[1,1] = ', dA_prev[1,1])
print()
dA_prev = pool_backward(dA, cache, mode = "average")
print("mode = average")
print('mean of dA = ', np.mean(dA))
print('dA_prev[1,1] = ', dA_prev[1,1])
```
**Expected Output**:
mode = max:
<table>
<tr>
<td>
**mean of dA =**
</td>
<td>
0.145713902729
</td>
</tr>
<tr>
<td>
**dA_prev[1,1] =**
</td>
<td>
[[ 0. 0. ] <br>
[ 5.05844394 -1.68282702] <br>
[ 0. 0. ]]
</td>
</tr>
</table>
mode = average
<table>
<tr>
<td>
**mean of dA =**
</td>
<td>
0.145713902729
</td>
</tr>
<tr>
<td>
**dA_prev[1,1] =**
</td>
<td>
[[ 0.08485462 0.2787552 ] <br>
[ 1.26461098 -0.25749373] <br>
[ 1.17975636 -0.53624893]]
</td>
</tr>
</table>
### Congratulations !
Congratulations on completing this assignment. You now understand how convolutional neural networks work. You have implemented all the building blocks of a neural network. In the next assignment you will implement a ConvNet using TensorFlow.
| github_jupyter |
# Lecture 21: Covariance, Correlation, Variance of a sum, Variance of Binomial & Hypergeometric
## Stat 110, Prof. Joe Blitzstein, Harvard University
----
### Definition
Covariance of any 2 random variables $X, Y$ is defined as
\begin{align}
\operatorname{Cov}(X, Y) &= \mathbb{E}\left( (X - \mathbb{E}(X)) (Y - \mathbb{E}(Y)) \right) \\
&= \mathbb{E}(XY) - \mathbb{E}(X) \, \mathbb{E}(Y) & \quad \text{similar to definition of variance}
\end{align}
Covariance is a measure of how $X, Y$ might vary _in tandem_.
If the product of $(X - \mathbb{E}(X))$ and $(Y - \mathbb{E}(Y))$ is _positive_ then that means that both values are either _positive_ ($X,Y$ tend to be greater than their respective means); or they are both _negative_ ($X,Y$ tend to be less than their means).
Correlation is defined in terms of covariance, as you will see in a bit.
### Properties
\begin{align}
& \text{[1]} & \operatorname{Cov}(X,X) &= \operatorname{Var}(X) \\
\\
& \text{[2]} & \operatorname{Cov}(X,Y) &= \operatorname{Cov}(Y,X) \\
\\
& \text{[3]} & \operatorname{Cov}(X, c) &= 0 & \quad \text{for some constant }c \\
\\
& \text{[4]} & \operatorname{Cov}(cX, Y) &= c \, \operatorname{Cov}(X,Y) & \quad \text{bilinearity} \\
\\
& \text{[5]} & \operatorname{Cov}(X, Y+Z) &= \operatorname{Cov}(X,Y) + \operatorname{Cov}(X,Z) & \quad \text{bilinearity} \\
\\
& \text{[6]} & \operatorname{Cov}(X+Y, Z+W) &= \operatorname{Cov}(X,Z) + \operatorname{Cov}(X,W) + \operatorname{Cov}(Y,Z) + \operatorname{Cov}(Y,W) & \quad \text{applying (5)} \\
\\
& \Rightarrow & \operatorname{Cov} \left( \sum_{i=1}^{m} a_{i} \, X_{i}, \sum_{j=1}^{n} b_{j} \, Y_{j} \right) &= \sum_{i,j} a_{i} \, b_{j} \, Cov \left( X_{i}, Y_{j} \right) \\
\\
& \text{[7]} & \operatorname{Var}(X_1 + X_2) &= \operatorname{Cov}(X_1+X_2, X_1+X_2) \\
& & &= \operatorname{Cov}X_1, X_1) + \operatorname{Cov}(X_1, X_2) + \operatorname{Cov}(X_2, X_1) + \operatorname{Cov}(X_2, X_2) \\
& & &= \operatorname{Var}(X_1) + \operatorname{Var}(X_2) + 2 \, \operatorname{Cov}(X_1, X_2) \\
\\
& \Rightarrow & \operatorname{Var}(X_1 + X_2) &= \operatorname{Var}(X_1) + \operatorname{Var}(X_2) & \quad \text{if }X_1, X_2 \text{ are uncorrelated} \\
\\
& \Rightarrow & \operatorname{Var}(X_1 + \dots + X_n) &= \operatorname{Var}(X_1) + \dots + \operatorname{Var}(X_n) + 2 \, \sum_{i \lt j} Cov(X_i, X_j) \\
\end{align}
#### Bilinearity
If you imagine treating one variable as fixed and then start working with the other, it sort of looks like _linearity_. It also sort of looks like the distributive property, too.
Bilinearity lets us
\begin{align}
\operatorname{Cov}(\lambda X, Y) &= \lambda \, \operatorname{Cov}(X, Y) = \operatorname{Cov}(X, \lambda Y) & \quad \text{freely move scaling factors} \\
\\
\operatorname{Cov}(X, Y_1 + Y_2) &= \operatorname{Cov}(X, Y_1) + \operatorname{Cov}(X, Y_2) \\
\operatorname{Cov}(X_1 + X_2, Y) &= \operatorname{Cov}(X_1, Y) + \operatorname{Cov}(X_2, Y) & \quad \text{distribution in }\operatorname{Cov} \text{ operator} \\
\end{align}
### Theorem If $X,Y$ are independent, then they are also uncorrelated, i.e., $\operatorname{Cov}(X,Y) = 0$
The converse is not true, however: $\operatorname{Cov}(X,Y) = 0$ does not necessarily mean that $X,Y$ are independent.
Consider $Z \sim \mathcal{N}(0,1)$, and $X \sim Z, Y \sim Z^2$.
\begin{align}
\operatorname{Cov}(X,Y) &= \mathbb{E}(XY) - \mathbb{E}(X)\,\mathbb{E}(Y) \\
&= \mathbb{E}(Z^3) - \mathbb{E}(Z) \, \mathbb{E}(Z^2) \\
&= 0 - 0 & \quad \text{odd moments of }Z \text{ are 0} \\
&= 0
\end{align}
But given $X \sim Z$, we know *exactly* everything about $Y$. And knowing $Y$, gives us everything about $X$, _except for the sign_.
So $X,Y$ are dependent, yet their covariance (and hence correlation) is 0.
## Definition
Correlation is defined in terms of covariance.
\begin{align}
\operatorname{Corr}(X,Y) &= \frac{\operatorname{Cov}(X,Y)}{\sigma_{X} \, \sigma_{Y}} = \frac{\operatorname{Cov}(X,Y)}{\sqrt{\operatorname{Var}(X)} \, \sqrt{\operatorname{Var}(Y)}} \\
\\
&= \operatorname{Cov} \left( \frac{X-\mathbb{E}(X)}{\sigma_{X} }, \frac{Y-\mathbb{E}(Y)}{\sigma_{Y}} \right) & \quad \text{standardize first, then find covariance} \\
\end{align}
Correlation is dimensionless, and like standard deviation, is unit-less.
Correlation also ranges from -1 to 1. This is what happens when we divide by $\sqrt{\operatorname{Var}(X)} \, \sqrt{\operatorname{Var}(Y)}$
### Theorem $-1 \le \operatorname{Corr}(X,Y) \le 1$
Without loss of generality, assume that $X,Y$ are already standarized (mean 0, variance 1).
\begin{align}
\operatorname{Va}r(X + Y) &=\operatorname{Var}(X) + \operatorname{Var}(Y) + 2\, \operatorname{Corr}(X,Y) \\
&= 1 + 1 + 2 \, \rho & \quad \text{ where } \rho = \operatorname{Corr}(X,Y) \\
&= 2 + 2 \, \rho \\
0 &\le 2 + 2 \, \rho & \quad \text{since }Var \ge 0 \\
0 &\le 1 + \rho & \Rightarrow \rho \text{ has a floor of }-1 \\
\\
\operatorname{Var}(X - Y) &= \operatorname{Var}(X) + \operatorname{Var}(-Y) - 2\, \operatorname{Corr}(X,Y) \\
&= 1 + 1 - 2 \, \rho & \quad \text{ where } \rho = \operatorname{Corr}(X,Y) \\
&= 2 - 2 \, \rho \\
0 &\le 2 - 2 \, \rho & \quad \text{since }Var \ge 0 \\
0 &\le 1 - \rho & \Rightarrow \rho \text{ has a ceiling of }1 \\
\\
&\therefore -1 \le \operatorname{Corr}(X,Y) \le 1 &\quad \blacksquare
\end{align}
## $\operatorname{Cov}$ in a Multinomial
Given $(X_1, \dots , X_k) \sim \operatorname{Mult}(n, \vec{p})$, find $\operatorname{Cov}(X_i, X_j)$ for all $i,j$.
\begin{align}
\text{case 1, where } i=j \text{ ...} \\
\\
\operatorname{Cov}(X_i, X_j) &= \operatorname{Var}(X_i) \\
&= n \, p_i \, (1 - p_i) \\
\\
\\
\text{case 2, where } i \ne j \text{ ...} \\
\\
\operatorname{Var}(X_i + X_j) &= \operatorname{Var}(X_i) + \operatorname{Var}(X_j) + 2 \, \operatorname{Cov}(X_i, X_j) & \quad \text{property [7] above} \\
n \, (p_i + p_j) \, (1 - (p_i + p_j)) &= n \, p_i \, (1 - p_i) + n \, p_j \, (1 - p_j) + 2 \, \operatorname{Cov}(X_i, X_j) & \quad \text{lumping property} \\
2 \, \operatorname{Cov}(X_i, X_j) &= n \, (p_i + p_j) \, (1 - (p_i + p_j)) - n \, p_i \, (1 - p_i) - n \, p_j \, (1 - p_j) \\
&= n \left( (p_i + p_j) - (p_i + p_j)^2 - p_i + p_i^2 - p_j + p_j^2 \right) \\
&= n ( -2 \, p_i \, p_j ) \\
\operatorname{Cov}(X_i, X_j) &= - n \, p_i \, p_j
\end{align}
Notice how case 2 where $i \ne j$ yields $\operatorname{Cov}(X_i, X_j) = - n \, p_i \, p_j$, which is negative. This should agree with your intuition that for $(X_1, \dots , X_k) \sim \operatorname{Mult}(n, \vec{p})$, categories $i,j$ are competing with another, and so they cannot be positively correlated nor independent of each other.
## Variance of a Binomial
Applying what we now know of covariance, we can obtain the variance of $X \sim \operatorname{Bin}(n, p)$.
We can describe $X = X_1 + \dots + X_n$ where $X_i$ are i.i.d. $\operatorname{Bern}(p)$.
Now consider _indicator random variables_. Let $I_A$ be the indicator of event $A$.
\begin{align}
I_A &\in \{0, 1\} \\
\\
\Rightarrow I_A^2 &= I_A \\
I_A^3 &= I_A \\
& \text{...} \\
\\
\Rightarrow I_A \, I_B &= I_{A \cap B} & \quad \text{for another event } B \\
\\
Var(X_i) &= \mathbb{E}X_i^2 - ( \mathbb{E}X_i)^2 \\
&= p_i - p_i^2 \\
&= p_i \, (1 - p_i) \\
&= p \, q & \quad \text{variance of Bernoulli} \\
\\
\Rightarrow Var(X) &= Var(X_i + \dots + X_n) \\
&= \operatorname{Var}(X_i) + \dots + \operatorname{Var}(X_n) + 2 \, \left( \sum_{i \ne j} Cov(X_i, X_j) \right) \\
&= n \, p \, q + 2 \, (0) & \quad \operatorname{Cov}(X_i,X_j) = 0 \text{ since } X_i \text{ are i.i.d.} \\
&= n \, p \, q
\end{align}
## Variance of Hypergeometric
Given $X \sim \operatorname{HGeom}(w, b, n)$ where
\begin{align}
X &= X_1 + \dots + X_n \\
\\
X_j &=
\begin{cases}
1, &\text{ if }j^{th} \text{ ball is White} \\
0, &\text{ otherwise } \\
\end{cases} \\
\end{align}
Recall that the balls are sampled _without replacement_, so the draws are not independent.
However, for any draw, it is not the case that any particular ball would prefer to be drawn for that particular iteration. So there is some symmetry we can leverage!
\begin{align}
\operatorname{Var}(X) &= n \, \operatorname{Var}(X_1) + 2 \, \binom{n}{2} \operatorname{Cov}(X_1, X_2) & \quad \text{symmetry!} \\
\\
\text{now } \operatorname{Cov}(X_1, X_2) &= \mathbb{E}(X_1 X_2) - \mathbb{E}(X_1) \, \mathbb{E}(X_2) \\
&= \frac{w}{w+b} \, \frac{w-1}{w+b-1} - \left( \frac{w}{w+b} \right)^2 & \quad \text{recall } I_A \, I_B = I_{A \cap B} \text{; and symmetry} \\
\end{align}
Prof. Blitzstein runs out of time, but the rest is just algebra. _To be concluded in the next lecture._
----
# Appendix: Variance, Covariance and Linear Algebra
Let's relate what we know about variance, covariance and correlation with the concept of the [dot product](https://www.mathsisfun.com/algebra/vectors-dot-product.html) in linear algebra:
\begin{align}
\operatorname{Cov}(X, Y) &= X \cdot Y \\
&= x_1 \, y_1 + \dots + x_n \, y_n \\
\\
\operatorname{Var}(X) &= \operatorname{Cov}(X, X) \\
&= X \cdot X \\
&= x_1 \, x_1 + \dots + x_n \, x_n \\
&= |X|^2 \\
\\
\Rightarrow |X| &= \sigma_{X} \\
&= \sqrt{\operatorname{Var}(X)} \\
&= \sqrt{x_1 \, x_1 + \dots + x_n \, x_n} & \quad \text{the size of }\vec{X} \text{ is its std. deviation} \\
\end{align}
Now let's take this a bit further...
\begin{align}
X \cdot Y &= |X| \, |Y| \, cos \theta \\
\\
\Rightarrow cos \theta &= \frac{X \cdot Y}{|X| \, |Y|} \\
&= \frac{\operatorname{Cov}(X, Y)}{\sigma_{X} \, \sigma_{Y}} \\
&= \rho & \quad \text{... otherwise known as }\operatorname{Corr}(X,Y) \\
\end{align}
Since $\rho_{X,Y} = cos_{X,Y} \theta$, we can see that the correlation of $X,Y$ is also the factor that projects vector $X$ onto $Y$, or vice versa. Now you can see why we can use cosine similarity as a metric to measure how close 2 vectors $X,Y$ are, since it is $\operatorname{Corr}(X,Y)$.
----
View [Lecture 21: Covariance and Correlation | Statistics 110](http://bit.ly/2MXxn89) on YouTube.
| github_jupyter |
```
#cell-width control
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:80% !important; }</style>"))
```
# Imports
```
#packages
import numpy
import tensorflow as tf
from tensorflow.core.example import example_pb2
#utils
import os
import random
import pickle
import struct
import time
from generators import *
#keras
import keras
from keras.preprocessing import text, sequence
from keras.preprocessing.text import Tokenizer
from keras.models import Model, Sequential
from keras.models import load_model
from keras.layers import Dense, Dropout, Activation, Concatenate, Dot, Embedding, LSTM, Conv1D, MaxPooling1D, Input, Lambda
#callbacks
from keras.callbacks import TensorBoard, ModelCheckpoint, Callback
```
# Seed
```
sd = 7
from numpy.random import seed
seed(sd)
from tensorflow import set_random_seed
set_random_seed(sd)
```
# CPU usage
```
#os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID" # see issue #152
#os.environ["CUDA_VISIBLE_DEVICES"] = ""
```
# Global parameters
```
# Embedding
max_features = 400000
maxlen_text = 400
maxlen_summ = 80
embedding_size = 100 #128
# Convolution
kernel_size = 5
filters = 64
pool_size = 4
# LSTM
lstm_output_size = 70
# Training
batch_size = 32
epochs = 20
```
# Load data
```
data_dir = '/mnt/disks/500gb/experimental-data-mini/experimental-data-mini/generator-dist-1to1/1to1/'
processing_dir = '/mnt/disks/500gb/stats-and-meta-data/400000/'
with open(data_dir+'partition.pickle', 'rb') as handle: partition = pickle.load(handle)
with open(data_dir+'labels.pickle', 'rb') as handle: labels = pickle.load(handle)
with open(processing_dir+'tokenizer.pickle', 'rb') as handle: tokenizer = pickle.load(handle)
embedding_matrix = numpy.load(processing_dir+'embedding_matrix.npy')
#the p_n constant
c = 80000
#stats
maxi = numpy.load(processing_dir+'training-stats-all/maxi.npy')
mini = numpy.load(processing_dir+'training-stats-all/mini.npy')
sample_info = (numpy.random.uniform, mini,maxi)
```
# Model
```
#2way input
text_input = Input(shape=(maxlen_text,embedding_size), dtype='float32')
summ_input = Input(shape=(maxlen_summ,embedding_size), dtype='float32')
#1way dropout
#text_route = Dropout(0.25)(text_input)
summ_route = Dropout(0.25)(summ_input)
#1way conv
#text_route = Conv1D(filters,
#kernel_size,
#padding='valid',
#activation='relu',
#strides=1)(text_route)
summ_route = Conv1D(filters,
kernel_size,
padding='valid',
activation='relu',
strides=1)(summ_route)
#1way max pool
#text_route = MaxPooling1D(pool_size=pool_size)(text_route)
summ_route = MaxPooling1D(pool_size=pool_size)(summ_route)
#1way lstm
#text_route = LSTM(lstm_output_size)(text_route)
summ_route = LSTM(lstm_output_size)(summ_route)
#negate results
#merged = Lambda(lambda x: -1*x)(merged)
#add p_n constant
#merged = Lambda(lambda x: x + c)(merged)
#output
output = Dense(1, activation='sigmoid')(summ_route)
#define model
model = Model(inputs=[text_input, summ_input], outputs=[output])
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
print(model.summary())
```
# Train model
```
#callbacks
class BatchHistory(keras.callbacks.Callback):
def on_train_begin(self, logs={}):
self.losses = []
self.accs = []
def on_batch_end(self, batch, logs={}):
self.losses.append(logs.get('loss'))
self.accs.append(logs.get('acc'))
history = BatchHistory()
tensorboard = TensorBoard(log_dir='./logs', histogram_freq=0, batch_size=batch_size, write_graph=True, write_grads=True)
modelcheckpoint = ModelCheckpoint('best.h5', monitor='val_loss', verbose=0, save_best_only=True, mode='min', period=1)
#batch generator parameters
params = {'dim': [(maxlen_text,embedding_size),(maxlen_summ,embedding_size)],
'batch_size': batch_size,
'shuffle': True,
'tokenizer':tokenizer,
'embedding_matrix':embedding_matrix,
'maxlen_text':maxlen_text,
'maxlen_summ':maxlen_summ,
'data_dir':data_dir,
'sample_info':sample_info}
#generators
training_generator = ContAllGenerator(partition['train'], labels, **params)
validation_generator = ContAllGenerator(partition['validation'], labels, **params)
# Train model on dataset
model.fit_generator(generator=training_generator,
validation_data=validation_generator,
use_multiprocessing=True,
workers=5,
epochs=epochs,
callbacks=[tensorboard, modelcheckpoint, history])
with open('losses.pickle', 'wb') as handle: pickle.dump(history.losses, handle, protocol=pickle.HIGHEST_PROTOCOL)
with open('accs.pickle', 'wb') as handle: pickle.dump(history.accs, handle, protocol=pickle.HIGHEST_PROTOCOL)
```
| github_jupyter |
# 100 numpy exercises
This is a collection of exercises that have been collected in the numpy mailing list, on stack overflow
and in the numpy documentation. The goal of this collection is to offer a quick reference for both old
and new users but also to provide a set of exercises for those who teach.
If you find an error or think you've a better way to solve some of them, feel
free to open an issue at <https://github.com/rougier/numpy-100>.
File automatically generated. See the documentation to update questions/answers/hints programmatically.
Run the `initialize.py` module, then for each question you can query the
answer or an hint with `hint(n)` or `answer(n)` for `n` question number.
```
import matplotlib.pyplot as plt
%matplotlib inline
```
#### 1. Import the numpy package under the name `np` (★☆☆)
```
import numpy as np
```
#### 2. Print the numpy version and the configuration (★☆☆)
```
np.__version__
```
#### 3. Create a null vector of size 10 (★☆☆)
```
# 尺寸参数 要么是标量 要么是元组
arr = np.zeros(10)
arr
```
#### 4. How to find the memory size of any array (★☆☆)
```
arr.size
```
#### 5. How to get the documentation of the numpy add function from the command line? (★☆☆)
```
# 就是显示文档用的
np.info(np.sum)
```
#### 6. Create a null vector of size 10 but the fifth value which is 1 (★☆☆)
```
vec = np.zeros(10)
vec[4] = 1
vec
```
#### 7. Create a vector with values ranging from 10 to 49 (★☆☆)
```
np.arange(10, 50)
```
#### 8. Reverse a vector (first element becomes last) (★☆☆)
```
arr = np.arange(10)
arr[::-1]
```
#### 9. Create a 3x3 matrix with values ranging from 0 to 8 (★☆☆)
```
np.arange(9).reshape(3, 3)
```
#### 10. Find indices of non-zero elements from [1,2,0,0,4,0] (★☆☆)
```
arr = np.array([1,2,0,0,4,0])
arr.nonzero()
```
#### 11. Create a 3x3 identity matrix (★☆☆)
```
# 单位矩阵
np.eye(3, 3)
```
#### 12. Create a 3x3x3 array with random values (★☆☆)
```
# rand 是0-1均匀分布
# randn 是正态分布
# randint 是指定上下界的随机整数 均匀分布
np.random.rand(3, 3, 3)
#x = np.arange(1000)
yu = np.random.rand(10000) * 6
yn = np.random.randn(10000)
plt.hist(yu, bins=100, label="u")
plt.hist(yn, bins=100, label="n")
plt.legend()
plt.show()
```
#### 13. Create a 10x10 array with random values and find the minimum and maximum values (★☆☆)
```
vec = np.random.rand(10, 10)
print(vec.max(), vec.min())
```
#### 14. Create a random vector of size 30 and find the mean value (★☆☆)
```
vec = np.random.rand(30)
vec.mean()
```
#### 15. Create a 2d array with 1 on the border and 0 inside (★☆☆)
```
vec = np.ones((4, 4))
vec[1:-1, 1:-1] = 0
vec
```
#### 16. How to add a border (filled with 0's) around an existing array? (★☆☆)
```
# S1 可以用多维度索引选出区间
vec = np.random.randint(1, 10, (5, 5))
mask = np.zeros((5, 5), dtype=np.int64)
mask[1:-1, 1:-1] = 1
vec *= mask
vec
# pad 填充限于边界元素 可以指定填充宽度和填充值
vec = np.random.randint(1, 10, (5, 5))
np.pad(vec, 1, mode="constant", constant_values=0)
```
#### 17. What is the result of the following expression? (★☆☆)
```python
0 * np.nan
np.nan == np.nan
np.inf > np.nan
np.nan - np.nan
np.nan in set([np.nan])
0.3 == 3 * 0.1
```
```
# NAN 和其他值运算的结果都是nan 和任何数比较的bool值都是nan 但是可以作为一个元素放到set中
print("0 * np.nan", 0 * np.nan)
print("np.nan == np.nan", np.nan == np.nan)
print("np.inf > np.nan", np.inf > np.nan)
print("np.nan - np.nan", np.nan - np.nan)
print("np.nan in set([np.nan])", np.nan in set([np.nan]))
print("0.3 == 3 * 0.1", 0.3 == 3 * 0.1)
```
#### 18. Create a 5x5 matrix with values 1,2,3,4 just below the diagonal (★☆☆)
```
vec = np.eye(5, 5)
vec *= np.arange(5).reshape(-1, 1)
vec
# 如果diag传入的是1D array 那么就是将1D array 作为对角线元素生成矩阵
np.diag(np.arange(5))
# 如果diag传入的是2D array 那么取出了对角线的元素
vec2 = np.random.randint(1, 10, (5, 5))
np.diag(vec2)
```
#### 19. Create a 8x8 matrix and fill it with a checkerboard pattern (★☆☆)
```
# check board 是啥
Z = np.zeros((8,8),dtype=int)
Z[1::2,::2] = 1
Z[::2,1::2] = 1
print(Z)
```
#### 20. Consider a (6,7,8) shape array, what is the index (x,y,z) of the 100th element?
```
vec = np.random.randint(1, 100, (6, 7, 8))
# 第一种方式是将多为数组转变为1维 然后找出对应位置的元素
vec.reshape(-1)[99]
# 第二种是使用 unravel_index 将一个一维的索引转化到对应形状的索引中 返回转化后的多维索引
# 这里的多维索引 默认使用的是 C style的方式 也就是行优先, 按照z字形来增加索引
print("index={}".format(np.unravel_index(99,(6,7,8))))
vec[np.unravel_index(99,(6,7,8))]
```
#### 21. Create a checkerboard 8x8 matrix using the tile function (★☆☆)
```
# np.tile 是按照对应维度顺序进行复制的
brick = np.array([[0, 1],[1, 0]])
np.tile(brick, (4, 4))
```
#### 22. Normalize a 5x5 random matrix (★☆☆)
```
# mean 计算均值
# std 计算标准差
random_vec = np.random.randint(1, 100, (5, 5))
(random_vec - random_vec.mean()) / random_vec.std()
```
#### 23. Create a custom dtype that describes a color as four unsigned bytes (RGBA) (★☆☆)
```
# 创建了一种自定义的类型 可以看做这个类型就是一个小号的字典 由key和value组成 字符串对应于key 后面的类型就是value的类型
rgb_dtype = np.dtype([("r", np.uint8), ("g", np.uint8), ("b", np.uint8), ("a", np.uint8)])
rgb_data = [
(1, 2, 3, 4), (2, 2, 3, 4), (3, 2, 3, 4), (4, 2, 3, 4)
]
arr = np.array(rgb_data, dtype=rgb_dtype)
# 使用数字索引获得了第一个元素 然后用["r"] 获取了这个元素中对应的名称为"r"的成员
arr[0]["r"]
```
#### 24. Multiply a 5x3 matrix by a 3x2 matrix (real matrix product) (★☆☆)
```
mat1 = np.arange(15).reshape(5, 3)
mat2 = np.arange(6).reshape(3, 2)
mat1.dot(mat2)
# 这个@运算符还是第一次见
Z = np.ones((5,3)) @ np.ones((3,2))
Z
```
#### 25. Given a 1D array, negate all elements which are between 3 and 8, in place. (★☆☆)
#### 26. What is the output of the following script? (★☆☆)
```python
# Author: Jake VanderPlas
print(sum(range(5),-1))
from numpy import *
print(sum(range(5),-1))
```
#### 27. Consider an integer vector Z, which of these expressions are legal? (★☆☆)
```python
Z**Z
2 << Z >> 2
Z <- Z
1j*Z
Z/1/1
Z<Z>Z
```
#### 28. What are the result of the following expressions?
```python
np.array(0) / np.array(0)
np.array(0) // np.array(0)
np.array([np.nan]).astype(int).astype(float)
```
#### 29. How to round away from zero a float array ? (★☆☆)
#### 30. How to find common values between two arrays? (★☆☆)
#### 31. How to ignore all numpy warnings (not recommended)? (★☆☆)
#### 32. Is the following expressions true? (★☆☆)
```python
np.sqrt(-1) == np.emath.sqrt(-1)
```
#### 33. How to get the dates of yesterday, today and tomorrow? (★☆☆)
#### 34. How to get all the dates corresponding to the month of July 2016? (★★☆)
#### 35. How to compute ((A+B)*(-A/2)) in place (without copy)? (★★☆)
#### 36. Extract the integer part of a random array using 5 different methods (★★☆)
#### 37. Create a 5x5 matrix with row values ranging from 0 to 4 (★★☆)
#### 38. Consider a generator function that generates 10 integers and use it to build an array (★☆☆)
#### 39. Create a vector of size 10 with values ranging from 0 to 1, both excluded (★★☆)
#### 40. Create a random vector of size 10 and sort it (★★☆)
#### 41. How to sum a small array faster than np.sum? (★★☆)
#### 42. Consider two random array A and B, check if they are equal (★★☆)
#### 43. Make an array immutable (read-only) (★★☆)
#### 44. Consider a random 10x2 matrix representing cartesian coordinates, convert them to polar coordinates (★★☆)
#### 45. Create random vector of size 10 and replace the maximum value by 0 (★★☆)
#### 46. Create a structured array with `x` and `y` coordinates covering the [0,1]x[0,1] area (★★☆)
#### 47. Given two arrays, X and Y, construct the Cauchy matrix C (Cij =1/(xi - yj))
#### 48. Print the minimum and maximum representable value for each numpy scalar type (★★☆)
#### 49. How to print all the values of an array? (★★☆)
#### 50. How to find the closest value (to a given scalar) in a vector? (★★☆)
#### 51. Create a structured array representing a position (x,y) and a color (r,g,b) (★★☆)
#### 52. Consider a random vector with shape (100,2) representing coordinates, find point by point distances (★★☆)
#### 53. How to convert a float (32 bits) array into an integer (32 bits) in place?
#### 54. How to read the following file? (★★☆)
```
1, 2, 3, 4, 5
6, , , 7, 8
, , 9,10,11
```
#### 55. What is the equivalent of enumerate for numpy arrays? (★★☆)
#### 56. Generate a generic 2D Gaussian-like array (★★☆)
#### 57. How to randomly place p elements in a 2D array? (★★☆)
#### 58. Subtract the mean of each row of a matrix (★★☆)
#### 59. How to sort an array by the nth column? (★★☆)
#### 60. How to tell if a given 2D array has null columns? (★★☆)
#### 61. Find the nearest value from a given value in an array (★★☆)
#### 62. Considering two arrays with shape (1,3) and (3,1), how to compute their sum using an iterator? (★★☆)
#### 63. Create an array class that has a name attribute (★★☆)
#### 64. Consider a given vector, how to add 1 to each element indexed by a second vector (be careful with repeated indices)? (★★★)
#### 65. How to accumulate elements of a vector (X) to an array (F) based on an index list (I)? (★★★)
#### 66. Considering a (w,h,3) image of (dtype=ubyte), compute the number of unique colors (★★★)
#### 67. Considering a four dimensions array, how to get sum over the last two axis at once? (★★★)
#### 68. Considering a one-dimensional vector D, how to compute means of subsets of D using a vector S of same size describing subset indices? (★★★)
#### 69. How to get the diagonal of a dot product? (★★★)
#### 70. Consider the vector [1, 2, 3, 4, 5], how to build a new vector with 3 consecutive zeros interleaved between each value? (★★★)
#### 71. Consider an array of dimension (5,5,3), how to mulitply it by an array with dimensions (5,5)? (★★★)
#### 72. How to swap two rows of an array? (★★★)
#### 73. Consider a set of 10 triplets describing 10 triangles (with shared vertices), find the set of unique line segments composing all the triangles (★★★)
#### 74. Given an array C that is a bincount, how to produce an array A such that np.bincount(A) == C? (★★★)
#### 75. How to compute averages using a sliding window over an array? (★★★)
#### 76. Consider a one-dimensional array Z, build a two-dimensional array whose first row is (Z[0],Z[1],Z[2]) and each subsequent row is shifted by 1 (last row should be (Z[-3],Z[-2],Z[-1]) (★★★)
#### 77. How to negate a boolean, or to change the sign of a float inplace? (★★★)
#### 78. Consider 2 sets of points P0,P1 describing lines (2d) and a point p, how to compute distance from p to each line i (P0[i],P1[i])? (★★★)
#### 79. Consider 2 sets of points P0,P1 describing lines (2d) and a set of points P, how to compute distance from each point j (P[j]) to each line i (P0[i],P1[i])? (★★★)
#### 80. Consider an arbitrary array, write a function that extract a subpart with a fixed shape and centered on a given element (pad with a `fill` value when necessary) (★★★)
#### 81. Consider an array Z = [1,2,3,4,5,6,7,8,9,10,11,12,13,14], how to generate an array R = [[1,2,3,4], [2,3,4,5], [3,4,5,6], ..., [11,12,13,14]]? (★★★)
#### 82. Compute a matrix rank (★★★)
#### 83. How to find the most frequent value in an array?
#### 84. Extract all the contiguous 3x3 blocks from a random 10x10 matrix (★★★)
#### 85. Create a 2D array subclass such that Z[i,j] == Z[j,i] (★★★)
#### 86. Consider a set of p matrices wich shape (n,n) and a set of p vectors with shape (n,1). How to compute the sum of of the p matrix products at once? (result has shape (n,1)) (★★★)
#### 87. Consider a 16x16 array, how to get the block-sum (block size is 4x4)? (★★★)
#### 88. How to implement the Game of Life using numpy arrays? (★★★)
#### 89. How to get the n largest values of an array (★★★)
#### 90. Given an arbitrary number of vectors, build the cartesian product (every combinations of every item) (★★★)
#### 91. How to create a record array from a regular array? (★★★)
#### 92. Consider a large vector Z, compute Z to the power of 3 using 3 different methods (★★★)
#### 93. Consider two arrays A and B of shape (8,3) and (2,2). How to find rows of A that contain elements of each row of B regardless of the order of the elements in B? (★★★)
#### 94. Considering a 10x3 matrix, extract rows with unequal values (e.g. [2,2,3]) (★★★)
#### 95. Convert a vector of ints into a matrix binary representation (★★★)
#### 96. Given a two dimensional array, how to extract unique rows? (★★★)
#### 97. Considering 2 vectors A & B, write the einsum equivalent of inner, outer, sum, and mul function (★★★)
#### 98. Considering a path described by two vectors (X,Y), how to sample it using equidistant samples (★★★)?
#### 99. Given an integer n and a 2D array X, select from X the rows which can be interpreted as draws from a multinomial distribution with n degrees, i.e., the rows which only contain integers and which sum to n. (★★★)
#### 100. Compute bootstrapped 95% confidence intervals for the mean of a 1D array X (i.e., resample the elements of an array with replacement N times, compute the mean of each sample, and then compute percentiles over the means). (★★★)
| github_jupyter |
# Exploring the Lorenz System of Differential Equations
In this Notebook we explore the Lorenz system of differential equations:
$$
\begin{aligned}
\dot{x} & = \sigma(y-x) \\
\dot{y} & = \rho x - y - xz \\
\dot{z} & = -\beta z + xy
\end{aligned}
$$
This is one of the classic systems in non-linear differential equations. It exhibits a range of different behaviors as the parameters (\\(\sigma\\), \\(\beta\\), \\(\rho\\)) are varied.
## Imports
First, we import the needed things from IPython, NumPy, Matplotlib and SciPy.
```
%matplotlib inline
from ipywidgets import interact, interactive
from IPython.display import clear_output, display, HTML
import numpy as np
from scipy import integrate
from matplotlib import pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib.colors import cnames
from matplotlib import animation
```
## Computing the trajectories and plotting the result
We define a function that can integrate the differential equations numerically and then plot the solutions. This function has arguments that control the parameters of the differential equation (\\(\sigma\\), \\(\beta\\), \\(\rho\\)), the numerical integration (`N`, `max_time`) and the visualization (`angle`).
```
def solve_lorenz(N=10, angle=0.0, max_time=4.0, sigma=10.0, beta=8./3, rho=28.0):
fig = plt.figure()
ax = fig.add_axes([0, 0, 1, 1], projection='3d')
ax.axis('off')
# prepare the axes limits
ax.set_xlim((-25, 25))
ax.set_ylim((-35, 35))
ax.set_zlim((5, 55))
def lorenz_deriv(x_y_z, t0, sigma=sigma, beta=beta, rho=rho):
"""Compute the time-derivative of a Lorenz system."""
x, y, z = x_y_z
return [sigma * (y - x), x * (rho - z) - y, x * y - beta * z]
# Choose random starting points, uniformly distributed from -15 to 15
np.random.seed(1)
x0 = -15 + 30 * np.random.random((N, 3))
# Solve for the trajectories
t = np.linspace(0, max_time, int(250*max_time))
x_t = np.asarray([integrate.odeint(lorenz_deriv, x0i, t)
for x0i in x0])
# choose a different color for each trajectory
colors = plt.cm.viridis(np.linspace(0, 1, N))
for i in range(N):
x, y, z = x_t[i,:,:].T
lines = ax.plot(x, y, z, '-', c=colors[i])
plt.setp(lines, linewidth=2)
ax.view_init(30, angle)
plt.show()
return t, x_t
```
Let's call the function once to view the solutions. For this set of parameters, we see the trajectories swirling around two points, called attractors.
```
t, x_t = solve_lorenz(angle=0, N=10)
```
Using IPython's `interactive` function, we can explore how the trajectories behave as we change the various parameters.
```
w = interactive(solve_lorenz, angle=(0.,360.), max_time=(0.1, 4.0),
N=(0,50), sigma=(0.0,50.0), rho=(0.0,50.0))
display(w)
```
The object returned by `interactive` is a `Widget` object and it has attributes that contain the current result and arguments:
```
t, x_t = w.result
w.kwargs
```
After interacting with the system, we can take the result and perform further computations. In this case, we compute the average positions in \\(x\\), \\(y\\) and \\(z\\).
```
xyz_avg = x_t.mean(axis=1)
xyz_avg.shape
```
Creating histograms of the average positions (across different trajectories) show that on average the trajectories swirl about the attractors.
```
plt.hist(xyz_avg[:,0])
plt.title('Average $x(t)$');
plt.hist(xyz_avg[:,1])
plt.title('Average $y(t)$');
```
| github_jupyter |
# Custom Estimator with Keras
**Learning Objectives**
- Learn how to create custom estimator using tf.keras
## Introduction
Up until now we've been limited in our model architectures to premade estimators. But what if we want more control over the model? We can use the popular Keras API to create a custom model. Keras is a high-level API to build and train deep learning models. It is user-friendly, modular and makes writing custom building blocks of Tensorflow code much easier.
Once we've build a Keras model we then it to an estimator using `tf.keras.estimator.model_to_estimator()`This gives us access to all the flexibility of Keras for creating deep learning models, but also the production readiness of the estimator framework!
```
# Ensure that we have Tensorflow 1.13 installed.
!pip3 freeze | grep tensorflow==1.13.1 || pip3 install tensorflow==1.13.1
import tensorflow as tf
import numpy as np
import shutil
print(tf.__version__)
```
## Train and Evaluate input functions
For the most part, we can use the same train and evaluation input functions that we had in previous labs. Note the function `create_feature_keras_input` below. We will use this to create the first layer of the model. This function is called in turn during the `train_input_fn` and `eval_input_fn` as well.
```
CSV_COLUMN_NAMES = ["fare_amount","dayofweek","hourofday","pickuplon","pickuplat","dropofflon","dropofflat"]
CSV_DEFAULTS = [[0.0],[1],[0],[-74.0], [40.0], [-74.0], [40.7]]
def read_dataset(csv_path):
def parse_row(row):
# Decode the CSV row into list of TF tensors
fields = tf.decode_csv(records = row, record_defaults = CSV_DEFAULTS)
# Pack the result into a dictionary
features = dict(zip(CSV_COLUMN_NAMES, fields))
# NEW: Add engineered features
features = add_engineered_features(features)
# Separate the label from the features
label = features.pop("fare_amount") # remove label from features and store
return features, label
# Create a dataset containing the text lines.
dataset = tf.data.Dataset.list_files(file_pattern = csv_path) # (i.e. data_file_*.csv)
dataset = dataset.flat_map(map_func = lambda filename: tf.data.TextLineDataset(filenames = filename).skip(count = 1))
# Parse each CSV row into correct (features,label) format for Estimator API
dataset = dataset.map(map_func = parse_row)
return dataset
def create_feature_keras_input(features, label):
features = tf.feature_column.input_layer(features = features, feature_columns = create_feature_columns())
return features, label
def train_input_fn(csv_path, batch_size = 128):
#1. Convert CSV into tf.data.Dataset with (features, label) format
dataset = read_dataset(csv_path)
#2. Shuffle, repeat, and batch the examples.
dataset = dataset.shuffle(buffer_size = 1000).repeat(count = None).batch(batch_size = batch_size)
#3. Create single feature tensor for input to Keras Model
dataset = dataset.map(map_func = create_feature_keras_input)
return dataset
def eval_input_fn(csv_path, batch_size = 128):
#1. Convert CSV into tf.data.Dataset with (features, label) format
dataset = read_dataset(csv_path)
#2.Batch the examples.
dataset = dataset.batch(batch_size = batch_size)
#3. Create single feature tensor for input to Keras Model
dataset = dataset.map(map_func = create_feature_keras_input)
return dataset
```
## Feature Engineering
We'll use the same engineered features that we had in previous labs.
```
def add_engineered_features(features):
features["dayofweek"] = features["dayofweek"] - 1 # subtract one since our days of week are 1-7 instead of 0-6
features["latdiff"] = features["pickuplat"] - features["dropofflat"] # East/West
features["londiff"] = features["pickuplon"] - features["dropofflon"] # North/South
features["euclidean_dist"] = tf.sqrt(x = features["latdiff"]**2 + features["londiff"]**2)
return features
def create_feature_columns():
# One hot encode dayofweek and hourofday
fc_dayofweek = tf.feature_column.categorical_column_with_identity(key = "dayofweek", num_buckets = 7)
fc_hourofday = tf.feature_column.categorical_column_with_identity(key = "hourofday", num_buckets = 24)
# Cross features to get combination of day and hour
fc_day_hr = tf.feature_column.crossed_column(keys = [fc_dayofweek, fc_hourofday], hash_bucket_size = 24 * 7)
# Bucketize latitudes and longitudes
NBUCKETS = 16
latbuckets = np.linspace(start = 38.0, stop = 42.0, num = NBUCKETS).tolist()
lonbuckets = np.linspace(start = -76.0, stop = -72.0, num = NBUCKETS).tolist()
fc_bucketized_plat = tf.feature_column.bucketized_column(source_column = tf.feature_column.numeric_column(key = "pickuplon"), boundaries = lonbuckets)
fc_bucketized_plon = tf.feature_column.bucketized_column(source_column = tf.feature_column.numeric_column(key = "pickuplat"), boundaries = latbuckets)
fc_bucketized_dlat = tf.feature_column.bucketized_column(source_column = tf.feature_column.numeric_column(key = "dropofflon"), boundaries = lonbuckets)
fc_bucketized_dlon = tf.feature_column.bucketized_column(source_column = tf.feature_column.numeric_column(key = "dropofflat"), boundaries = latbuckets)
feature_columns = [
#1. Engineered using tf.feature_column module
tf.feature_column.indicator_column(categorical_column = fc_day_hr), # 168 columns
fc_bucketized_plat, # 16 + 1 = 17 columns
fc_bucketized_plon, # 16 + 1 = 17 columns
fc_bucketized_dlat, # 16 + 1 = 17 columns
fc_bucketized_dlon, # 16 + 1 = 17 columns
#2. Engineered in input functions
tf.feature_column.numeric_column(key = "latdiff"), # 1 column
tf.feature_column.numeric_column(key = "londiff"), # 1 column
tf.feature_column.numeric_column(key = "euclidean_dist") # 1 column
]
return feature_columns
```
Calculate the number of feature columns that will be input to our Keras model
```
num_feature_columns = 168 + (16 + 1) * 4 + 3
print("num_feature_columns = {}".format(num_feature_columns))
```
## Build Custom Keras Model
Now we can begin building our Keras model. Have a look at [the guide here](https://www.tensorflow.org/guide/keras) to see more explanation.
```
def create_keras_model():
model = tf.keras.Sequential()
model.add(tf.keras.layers.InputLayer(input_shape = (num_feature_columns,), name = "dense_input"))
model.add(tf.keras.layers.Dense(units = 64, activation = "relu", name = "dense0"))
model.add(tf.keras.layers.Dense(units = 64, activation = "relu", name = "dense1"))
model.add(tf.keras.layers.Dense(units = 64, activation = "relu", name = "dense2"))
model.add(tf.keras.layers.Dense(units = 64, activation = "relu", name = "dense3"))
model.add(tf.keras.layers.Dense(units = 8, activation = "relu", name = "dense4"))
model.add(tf.keras.layers.Dense(units = 1, activation = None, name = "logits"))
def rmse(y_true, y_pred): # Root Mean Squared Error
return tf.sqrt(x = tf.reduce_mean(input_tensor = tf.square(x = y_pred - y_true)))
model.compile(
optimizer = tf.train.AdamOptimizer(),
loss = "mean_squared_error",
metrics = [rmse])
return model
```
## Serving input function
Once we've constructed our model in Keras, we next create the serving input function. This is also similar to what we have done in previous labs. Note that we use our `create_feature_keras_input` function again so that we perform our feature engineering during inference.
```
# Create serving input function
def serving_input_fn():
feature_placeholders = {
"dayofweek": tf.placeholder(dtype = tf.int32, shape = [None]),
"hourofday": tf.placeholder(dtype = tf.int32, shape = [None]),
"pickuplon": tf.placeholder(dtype = tf.float32, shape = [None]),
"pickuplat": tf.placeholder(dtype = tf.float32, shape = [None]),
"dropofflon": tf.placeholder(dtype = tf.float32, shape = [None]),
"dropofflat": tf.placeholder(dtype = tf.float32, shape = [None]),
}
features = {key: tensor for key, tensor in feature_placeholders.items()}
# Perform our feature engineering during inference as well
features, _ = create_feature_keras_input((add_engineered_features(features)), None)
return tf.estimator.export.ServingInputReceiver(features = {"dense_input": features}, receiver_tensors = feature_placeholders)
```
## Train and Evaluate
To train our model, we can use `train_and_evaluate` as we have before. Note that we use `tf.keras.estimator.model_to_estimator` to create our estimator. It takes as arguments the compiled keras model, the OUTDIR, and optionally a `tf.estimator.Runconfig`. Have a look at [the documentation for tf.keras.estimator.model_to_estimator](https://www.tensorflow.org/api_docs/python/tf/keras/estimator/model_to_estimator) to make sure you understand how arguments are used.
```
def train_and_evaluate(output_dir):
tf.logging.set_verbosity(v = tf.logging.INFO) # so loss is printed during training
estimator = tf.keras.estimator.model_to_estimator(
keras_model = create_keras_model(),
model_dir = output_dir,
config = tf.estimator.RunConfig(
tf_random_seed = 1, # for reproducibility
save_checkpoints_steps = 100 # checkpoint every N steps
)
)
train_spec = tf.estimator.TrainSpec(
input_fn = lambda: train_input_fn(csv_path = "./taxi-train.csv"),
max_steps = 500)
exporter = tf.estimator.LatestExporter(name = 'exporter', serving_input_receiver_fn = serving_input_fn)
eval_spec = tf.estimator.EvalSpec(
input_fn = lambda: eval_input_fn(csv_path = "./taxi-valid.csv"),
steps = None,
start_delay_secs = 10, # wait at least N seconds before first evaluation (default 120)
throttle_secs = 10, # wait at least N seconds before each subsequent evaluation (default 600)
exporters = exporter) # export SavedModel once at the end of training
tf.estimator.train_and_evaluate(
estimator = estimator,
train_spec = train_spec,
eval_spec = eval_spec)
%%time
OUTDIR = "taxi_trained"
shutil.rmtree(path = OUTDIR, ignore_errors = True) # start fresh each time
tf.summary.FileWriterCache.clear() # ensure filewriter cache is clear for TensorBoard events file
train_and_evaluate(OUTDIR)
```
Copyright 2019 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| github_jupyter |
<sub>© 2021-present Neuralmagic, Inc. // [Neural Magic Legal](https://neuralmagic.com/legal)</sub>
# PyTorch Classification Model Pruning using SparseML
This notebook provides a step-by-step walkthrough for pruning an already trained (dense) model to enable better performance at inference time using the [DeepSparse Engine](https://github.com/neuralmagic/deepsparse). You will:
- Set up the model and dataset
- Define a generic PyTorch training flow
- Integrate the PyTorch flow with SparseML
- Prune the model using the PyTorch+SparseML flow
- Export to [ONNX](https://onnx.ai/)
Reading through this notebook will be reasonably quick to gain an intuition for how to plug SparseML into your PyTorch training flow. Rough time estimates for fully pruning the default model are given. Note that training with the PyTorch CPU implementation will be much slower than a GPU:
- 15 minutes on a GPU
- 45 minutes on a laptop CPU
## Step 1 - Requirements
To run this notebook, you will need the following packages already installed:
* SparseML and SparseZoo
* PyTorch and torchvision
You can install any package that is not already present via `pip`.
```
import sparseml
import sparsezoo
import torch
import torchvision
```
## Step 2 - Setting Up the Model and Dataset
By default, you will prune a [ResNet50](https://arxiv.org/abs/1512.03385) model trained on the [Imagenette dataset](https://github.com/fastai/imagenette). The model's pretrained weights are downloaded from the SparseZoo model repo. The Imagenette dataset is downloaded from its repository via a helper class from SparseML.
If you would like to try out your model for pruning, modify the appropriate lines for your model and dataset, specifically:
- model = ModelRegistry.create(...)
- train_dataset = ImagenetteDataset(...)
- val_dataset = ImagenetteDataset(...)
Take care to keep the variable names the same, as the rest of the notebook is set up according to those and update any parts of the training flow as needed.
```
from sparseml.pytorch.models import ModelRegistry
from sparseml.pytorch.datasets import ImagenetteDataset, ImagenetteSize
#######################################################
# Define your model below
#######################################################
print("loading model...")
model = ModelRegistry.create(
key="resnet50",
pretrained=True,
pretrained_dataset="imagenette",
num_classes=10,
)
input_shape = ModelRegistry.input_shape("resnet50")
input_size = input_shape[-1]
print(model)
#######################################################
# Define your train and validation datasets below
#######################################################
print("\nloading train dataset...")
train_dataset = ImagenetteDataset(
train=True, dataset_size=ImagenetteSize.s320, image_size=input_size
)
print(train_dataset)
print("\nloading val dataset...")
val_dataset = ImagenetteDataset(
train=False, dataset_size=ImagenetteSize.s320, image_size=input_size
)
print(val_dataset)
```
## Step 3 - Set Up a PyTorch Training Loop
SparseML can plug directly into your existing PyTorch training flow by overriding the Optimizer object. To demonstrate this, in the cell below, we define a simple PyTorch training loop adapted from [here](https://pytorch.org/tutorials/beginner/blitz/cifar10_tutorial.html). To prune your existing models using SparseML, you can use your own training flow.
```
from tqdm.auto import tqdm
import math
import torch
def run_model_one_epoch(model, data_loader, criterion, device, train=False, optimizer=None):
if train:
model.train()
else:
model.eval()
running_loss = 0.0
total_correct = 0
total_predictions = 0
for step, (inputs, labels) in tqdm(enumerate(data_loader), total=len(data_loader)):
inputs = inputs.to(device)
labels = labels.to(device)
if train:
optimizer.zero_grad()
outputs, _ = model(inputs) # model returns logits and softmax as a tuple
loss = criterion(outputs, labels)
if train:
loss.backward()
optimizer.step()
running_loss += loss.item()
predictions = outputs.argmax(dim=1)
total_correct += torch.sum(predictions == labels).item()
total_predictions += inputs.size(0)
loss = running_loss / (step + 1.0)
accuracy = total_correct / total_predictions
return loss, accuracy
```
## Step 4 - Set Up PyTorch Training Objects
In this step, you will select a device to train your model with, set up DataLoader objects, a loss function, and optimizer. All of these variables and objects can be replaced to fit your training flow.
```
from torch.utils.data import DataLoader
from torch.nn import CrossEntropyLoss
from torch.optim import SGD
# setup device
device = "cuda" if torch.cuda.is_available() else "cpu"
model.to(device)
print("Using device: {}".format(device))
# setup data loaders
batch_size = 128
train_loader = DataLoader(
train_dataset, batch_size, shuffle=True, pin_memory=True, num_workers=8
)
val_loader = DataLoader(
val_dataset, batch_size, shuffle=False, pin_memory=True, num_workers=8
)
# setup loss function and optimizer, LR will be overriden by sparseml
criterion = CrossEntropyLoss()
optimizer = SGD(model.parameters(), lr=0.001, momentum=0.9)
```
## Step 5 - Apply a SparseML Recipe and Prune Model
To prune a model with SparseML, you will download a recipe from SparseZoo and use it to create a `ScheduledModifierManager` object. This manager will be used to wrap the optimizer object to gradually prune the model using unstructured weight magnitude pruning after each optimizer step.
You can create SparseML recipes to perform various model pruning schedules, quantization aware training, sparse transfer learning, and more. If you are using a different model than the default, you will have to modify the recipe YAML file to target the new model's parameters.
Finally, using the wrapped optimizer object, you will call the training function to prune your model.
If the kernel shuts down during training, this may be an out of memory error, to resolve this, try lowering the `batch_size` in the cell above.
#### Downloading a Recipe from SparseZoo
The [SparseZoo](https://github.com/neuralmagic/sparsezoo) API provides precofigured recipes for its optimized model. In the cell below, you will download a recipe for pruning ResNet50 on the Imagenette dataset and record it's saved path.
```
from sparsezoo import Zoo
recipe = Zoo.search_recipes(
domain="cv",
sub_domain="classification",
architecture="resnet_v1",
sub_architecture="50",
framework="pytorch",
repo="sparseml",
dataset="imagenette",
optim_name="pruned",
)[0] # unwrap search result
recipe.download()
recipe_path = recipe.downloaded_path()
print(f"Recipe downloaded to: {recipe_path}")
from sparseml.pytorch.optim import (
ScheduledModifierManager,
ScheduledOptimizer,
)
# create ScheduledModifierManager and Optimizer wrapper
manager = ScheduledModifierManager.from_yaml(recipe_path)
optimizer = ScheduledOptimizer(
optimizer, model, manager, steps_per_epoch=len(train_loader), loggers=[],
)
# Run model pruning
epoch = manager.min_epochs
while epoch < manager.max_epochs:
# run training loop
epoch_name = "{}/{}".format(epoch + 1, manager.max_epochs)
print("Running Training Epoch {}".format(epoch_name))
train_loss, train_acc = run_model_one_epoch(
model, train_loader, criterion, device, train=True, optimizer=optimizer
)
print(
"Training Epoch: {}\nTraining Loss: {}\nTop 1 Acc: {}\n".format(
epoch_name, train_loss, train_acc
)
)
# run validation loop
print("Running Validation Epoch {}".format(epoch_name))
val_loss, val_acc = run_model_one_epoch(
model, val_loader, criterion, device
)
print(
"Validation Epoch: {}\nVal Loss: {}\nTop 1 Acc: {}\n".format(
epoch_name, val_loss, val_acc
)
)
epoch += 1
```
## Step 6 - View Model Sparsity
To see the effects of the model pruning, in this step, you will print out the sparsities of each Conv and FC layer in your model.
```
from sparseml.pytorch.utils import get_prunable_layers, tensor_sparsity
# print sparsities of each layer
for (name, layer) in get_prunable_layers(model):
print("{}.weight: {:.4f}".format(name, tensor_sparsity(layer.weight).item()))
```
## Step 7 - Exporting to ONNX
Now that the model is fully recalibrated, you need to export it to an ONNX format, which is the format used by the [DeepSparse Engine](https://github.com/neuralmagic/deepsparse). For PyTorch, exporting to ONNX is natively supported. In the cell block below, a convenience class, ModuleExporter(), is used to handle exporting.
Once the model is saved as an ONNX file, it is ready to be used for inference with the DeepSparse Engine. For saving a custom model, you can override the sample batch for ONNX graph freezing and locations to save to.
```
from sparseml.pytorch.utils import ModuleExporter
save_dir = "pytorch_classification"
exporter = ModuleExporter(model, output_dir=save_dir)
exporter.export_pytorch(name="resnet50_imagenette_pruned.pth")
exporter.export_onnx(torch.randn(1, 3, 224, 224), name="resnet50_imagenette_pruned.onnx")
```
## Next Steps
Congratulations, you have pruned a model and exported it to ONNX for inference! Next steps you can pursue include:
* Pruning different models using SparseML
* Trying different pruning and optimization recipes
* Running your model on the [DeepSparse Engine](https://github.com/neuralmagic/deepsparse)
| github_jupyter |
```
# Trying tf probability
from __future__ import print_function
import collections
import matplotlib.pyplot as plt
import numpy as np
import math
import os
import pandas as pd
import scipy
import scipy.stats
import tensorflow.compat.v2 as tf
tf.enable_v2_behavior()
# import tensorflow as tf
# import tensorflow.compat.v1 as tf
#tf.enable_v2_behavior()
import tensorflow_probability as tfp
tfd = tfp.distributions
tfb = tfp.bijectors
# try:
# tf.compat.v1.enable_eager_execution()
# except ValueError:
# pass
from six.moves import urllib
import warnings
tf.enable_eager_execution()
n = tfd.Normal(loc = 0., scale = 1.)
test = n.log_prob(n.sample(10))
# Reshape Manipulation
six_way_multinomial = tfd.Multinomial(total_count = 1000., probs = [.3, .25, .2, .15, .08, .02])
six_way_multinomial
# Now we can tranform the event shape dimension
transformed_multinomial = tfd.TransformedDistribution(
distribution=six_way_multinomial,
bijector=tfb.Reshape(event_shape_out=[2, 3]))
transformed_multinomial
# Now we can evaluate likelihood of the appropriately shaped object
transformed_multinomial.log_prob([[500., 100., 100.], [150., 100., 50.]])
# Now something more complicated: COVARIANCE ESTIMATION
# We're assuming 2-D data with a known true mean of (0, 0)
true_mean = np.zeros([2], dtype=np.float32)
# We'll make the 2 coordinates correlated
true_cor = np.array([[1.0, 0.9], [0.9, 1.0]], dtype=np.float32)
# And we'll give the 2 coordinates different variances
true_var = np.array([4.0, 1.0], dtype=np.float32)
# Combine the variances and correlations into a covariance matrix
true_cov = np.expand_dims(np.sqrt(true_var), axis=1).dot(np.expand_dims(np.sqrt(true_var), axis=1).T) * true_cor
# We'll be working with precision matrices, so we'll go ahead and compute the
# true precision matrix here
true_precision = np.linalg.inv(true_cov)
print(true_cov)
print('eigenvalues:', np.linalg.eigvals(true_cov))
np.random.seed(123)
my_data = np.random.multivariate_normal(mean = true_mean,
cov = true_cov,
size = 100,
check_valid = 'ignore').astype(np.float32)
my_data.shape
# Do a scatter plot
plt.scatter(my_data[:, 0], my_data[:, 1], alpha = 0.75)
print('mean of observations:', np.mean(my_data, axis=0))
print('true mean:', true_mean)
# Implement likelihood function
def log_lik_data_numpy(precision, data):
# np.linalg.inv is a really inefficient way to get the covariance matrix, but
# remember we don't care about speed here
cov = np.linalg.inv(precision)
rv = scipy.stats.multivariate_normal(true_mean, cov)
return np.sum(rv.logpdf(data))
# test case: compute the log likelihood of the data given the true parameters
log_lik_data_numpy(true_precision, my_data)
# Define priors
PRIOR_DF = 3
PRIOR_SCALE = np.eye(2, dtype = np.float32) / PRIOR_DF
def log_lik_prior_numpy(precision):
rv = scipy.stats.wishart(df = PRIOR_DF, scale = PRIOR_SCALE)
return rv.logpdf(precision)
log_lik_prior_numpy(true_precision)
# Getting analytical posteriors
n = my_data.shape[0]
nu_prior = PRIOR_DF
v_prior = PRIOR_SCALE
nu_posterior = nu_prior + n
v_posterior = np.linalg.inv(np.linalg.inv(v_prior) + my_data.T.dot(my_data))
posterior_mean = nu_posterior * v_posterior
v_post_diag = np.expand_dims(np.diag(v_posterior), axis=1)
posterior_sd = np.sqrt(nu_posterior *
(v_posterior ** 2.0 + v_post_diag.dot(v_post_diag.T)))
# Plot
sample_precision = np.linalg.inv(np.cov(my_data, rowvar=False, bias=False))
fig, axes = plt.subplots(2, 2)
fig.set_size_inches(10, 10)
for i in range(2):
for j in range(2):
ax = axes[i, j]
loc = posterior_mean[i, j]
scale = posterior_sd[i, j]
xmin = loc - 3.0 * scale
xmax = loc + 3.0 * scale
x = np.linspace(xmin, xmax, 1000)
y = scipy.stats.norm.pdf(x, loc=loc, scale=scale)
ax.plot(x, y)
ax.axvline(true_precision[i, j], color='red', label='True precision')
ax.axvline(sample_precision[i, j], color='red', linestyle=':', label='Sample precision')
ax.set_title('precision[%d, %d]' % (i, j))
plt.legend()
plt.show()
# NOW USING TF PROBABILITY
# Data log likelihood
VALIDATE_ARGS = True
ALLOW_NAN_STATS = False
def log_lik_data(precisions, replicated_data):
n = tf.shape(precisions)[0] # number of precision matrices
# We're estimating a precision matrix; we have to invert to get log
# probabilities. Cholesky inversion should be relatively efficient,
# but as we'll see later, it's even better if we can avoid doing the Cholesky
# decomposition altogether.
precisions_cholesky = tf.linalg.cholesky(precisions)
covariances = tf.linalg.cholesky_solve(
precisions_cholesky, tf.linalg.eye(2, batch_shape=[n]))
rv_data = tfd.MultivariateNormalFullCovariance(
loc=tf.zeros([n, 2]),
covariance_matrix=covariances,
validate_args=VALIDATE_ARGS,
allow_nan_stats=ALLOW_NAN_STATS)
return tf.reduce_sum(rv_data.log_prob(replicated_data), axis=0)
# Make replicated data
n = 2
replicated_data = np.tile(np.expand_dims(my_data, axis = 1), reps = [1, 2, 1])
print(replicated_data.shape)
# Check against the numpy implemention
precisions = np.stack([np.eye(2, dtype = np.float32), true_precision])
n = precisions.shape[0]
lik_tf = log_lik_data(precisions, replicated_data = replicated_data).numpy()
for i in range(n):
print(i)
print('numpy: ', log_lik_data_numpy(precisions[i], my_data))
print('tensorflow: ', lik_tf[i])
@tf.function(autograph = False)
def log_lik_prior(precisions):
rv_precision = tfd.Wishart(
df = PRIOR_DF,
scale_tril = tf.linalg.cholesky(PRIOR_SCALE),
validate_args = VALIDATE_ARGS,
allow_nan_stats = ALLOW_NAN_STATS)
return rv_precision.log_prob(precisions)
# check against the numpy implementation
precisions = np.stack([np.eye(2, dtype=np.float32), true_precision])
n = precisions.shape[0]
lik_tf = log_lik_prior(precisions).numpy()
for i in range(n):
print(i)
print('numpy:', log_lik_prior_numpy(precisions[i]))
print('tensorflow:', lik_tf[i])
# Get log Likelihood
def get_log_lik(data, n_chains = 1):
# The data argument that is passed in will be available to the inner function
# below so it doesn't have to be passed in as a parameter.
replicated_data = np.tile(np.expand_dims(data, axis = 1), reps = [1, n_chains, 1])
@tf.function(autograph = False)
def _log_lik(precision):
return log_lik_data(precision, replicated_data) + log_lik_prior(precision)
return _log_lik
@tf.function(autograph = False)
def sample():
tf.random.set_seed(123)
init_precision = tf.expand_dims(tf.eye(2), axis = 0)
# Use expand_dims because we want to pass in a tensor of starting values
log_lik_fn = get_log_lik(my_data, n_chains = 1)
# we'll just do a few steps here
num_results = 10
num_burnin_steps = 10
states = tfp.mcmc.sample_chain(
num_results = num_results,
num_burnin_steps = num_burnin_steps,
current_state = [
init_precision,
],
kernel=tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn = log_lik_fn,
step_size = 0.1,
num_leapfrog_steps = 3,
seed = 123),
trace_fn = None,
parallel_iterations = 1)
return states
try:
states = sample()
except Exception as e:
# shorten the giant stack trace
lines = str(e).split('\n')
print('\n'.join(lines[:5]+['...'] + lines[-3:]))
def get_log_lik_verbose(data, n_chains=1):
# The data argument that is passed in will be available to the inner function
# below so it doesn't have to be passed in as a parameter.
replicated_data = np.tile(np.expand_dims(data, axis=1), reps=[1, n_chains, 1])
def _log_lik(precisions):
# An internal method we'll make into a TensorFlow operation via tf.py_func
def _print_precisions(precisions):
print('precisions:\n', precisions)
return False # operations must return something!
# Turn our method into a TensorFlow operation
print_op = tf.compat.v1.py_func(_print_precisions, [precisions], tf.bool)
# Assertions are also operations, and some care needs to be taken to ensure
# that they're executed
assert_op = tf.assert_equal(
precisions, tf.linalg.matrix_transpose(precisions),
message = 'not symmetrical', summarize=4, name='symmetry_check')
# The control_dependencies statement forces its arguments to be executed
# before subsequent operations
with tf.control_dependencies([print_op, assert_op]):
return (log_lik_data(precisions, replicated_data) +
log_lik_prior(precisions))
return _log_lik
@tf.function(autograph=False)
def sample():
tf.random.set_seed(123)
init_precision = tf.eye(2)[tf.newaxis, ...]
log_lik_fn = get_log_lik_verbose(my_data)
# we'll just do a few steps here
num_results = 10
num_burnin_steps = 10
states = tfp.mcmc.sample_chain(
num_results=num_results,
num_burnin_steps=num_burnin_steps,
current_state=[
init_precision,
],
kernel=tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn=log_lik_fn,
step_size=0.1,
num_leapfrog_steps=3,
seed=123),
trace_fn=None,
parallel_iterations=1)
try:
states = sample()
except Exception as e:
# shorten the giant stack trace
lines = str(e).split('\n')
print('\n'.join(lines[:5]+['...']+lines[-3:]))
# Our transform has 3 stages that we chain together via composition:
precision_to_unconstrained = tfb.Chain([
# step 3: flatten the lower triangular portion of the matrix
tfb.Invert(tfb.FillTriangular(validate_args=VALIDATE_ARGS)),
# step 2: take the log of the diagonals
tfb.TransformDiagonal(tfb.Invert(tfb.Exp(validate_args=VALIDATE_ARGS))),
# step 1: decompose the precision matrix into its Cholesky factors
tfb.Invert(tfb.CholeskyOuterProduct(validate_args=VALIDATE_ARGS)),
])
m = tf.constant([[1., 2.], [2., 8.]])
m_fwd = precision_to_unconstrained.forward(m)
m_inv = precision_to_unconstrained.inverse(m_fwd)
# bijectors handle tensors of values, too!
m2 = tf.stack([m, tf.eye(2)])
m2_fwd = precision_to_unconstrained.forward(m2)
m2_inv = precision_to_unconstrained.inverse(m2_fwd)
print('single input:')
print('m:\n', m.numpy())
print('precision_to_unconstrained(m):\n', m_fwd.numpy())
print('inverse(precision_to_unconstrained(m)):\n', m_inv.numpy())
print()
print('tensor of inputs:')
print('m2:\n', m2.numpy())
print('precision_to_unconstrained(m2):\n', m2_fwd.numpy())
print('inverse(precision_to_unconstrained(m2)):\n', m2_inv.numpy())
def log_lik_prior_transformed(transformed_precisions):
rv_precision = tfd.TransformedDistribution(
tfd.Wishart(
df=PRIOR_DF,
scale_tril=tf.linalg.cholesky(PRIOR_SCALE),
validate_args=VALIDATE_ARGS,
allow_nan_stats=ALLOW_NAN_STATS),
bijector=precision_to_unconstrained,
validate_args=VALIDATE_ARGS)
return rv_precision.log_prob(transformed_precisions)
# Check against the numpy implementation. Note that when comparing, we need
# to add in the Jacobian correction.
precisions = np.stack([np.eye(2, dtype=np.float32), true_precision])
transformed_precisions = precision_to_unconstrained.forward(precisions)
lik_tf = log_lik_prior_transformed(transformed_precisions).numpy()
corrections = precision_to_unconstrained.inverse_log_det_jacobian(
transformed_precisions, event_ndims=1).numpy()
n = precisions.shape[0]
for i in range(n):
print(i)
print('numpy:', log_lik_prior_numpy(precisions[i]) + corrections[i])
print('tensorflow:', lik_tf[i])
def log_lik_data_transformed(transformed_precisions, replicated_data):
# We recover the precision matrix by inverting our bijector. This is
# inefficient since we really want the Cholesky decomposition of the
# precision matrix, and the bijector has that in hand during the inversion,
# but we'll worry about efficiency later.
n = tf.shape(transformed_precisions)[0]
precisions = precision_to_unconstrained.inverse(transformed_precisions)
precisions_cholesky = tf.linalg.cholesky(precisions)
covariances = tf.linalg.cholesky_solve(
precisions_cholesky, tf.linalg.eye(2, batch_shape=[n]))
rv_data = tfd.MultivariateNormalFullCovariance(
loc=tf.zeros([n, 2]),
covariance_matrix=covariances,
validate_args=VALIDATE_ARGS,
allow_nan_stats=ALLOW_NAN_STATS)
return tf.reduce_sum(rv_data.log_prob(replicated_data), axis=0)
# sanity check
precisions = np.stack([np.eye(2, dtype=np.float32), true_precision])
transformed_precisions = precision_to_unconstrained.forward(precisions)
lik_tf = log_lik_data_transformed(
transformed_precisions, replicated_data).numpy()
for i in range(precisions.shape[0]):
print(i)
print('numpy:', log_lik_data_numpy(precisions[i], my_data))
print('tensorflow:', lik_tf[i])
def get_log_lik_transformed(data, n_chains=1):
# The data argument that is passed in will be available to the inner function
# below so it doesn't have to be passed in as a parameter.
replicated_data = np.tile(np.expand_dims(data, axis=1), reps=[1, n_chains, 1])
@tf.function(autograph=False)
def _log_lik_transformed(transformed_precisions):
return (log_lik_data_transformed(transformed_precisions, replicated_data) +
log_lik_prior_transformed(transformed_precisions))
return _log_lik_transformed
# make sure everything runs
log_lik_fn = get_log_lik_transformed(my_data)
m = tf.eye(2)[tf.newaxis, ...]
lik = log_lik_fn(precision_to_unconstrained.forward(m)).numpy()
print(lik)
# We'll choose a proper random initial value this time
np.random.seed(123)
initial_value_cholesky = np.array(
[[0.5 + np.random.uniform(), 0.0],
[-0.5 + np.random.uniform(), 0.5 + np.random.uniform()]],
dtype=np.float32)
initial_value = initial_value_cholesky.dot(
initial_value_cholesky.T)[np.newaxis, ...]
# The sampler works with unconstrained values, so we'll transform our initial
# value
initial_value_transformed = precision_to_unconstrained.forward(
initial_value).numpy()
# Sample!
@tf.function(autograph=False)
def sample():
tf.random.set_seed(123)
log_lik_fn = get_log_lik_transformed(my_data, n_chains=1)
num_results = 1000
num_burnin_steps = 1000
states, is_accepted = tfp.mcmc.sample_chain(
num_results=num_results,
num_burnin_steps=num_burnin_steps,
current_state=[
initial_value_transformed,
],
kernel=tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn=log_lik_fn,
step_size=0.1,
num_leapfrog_steps=3,
seed=123),
trace_fn=lambda _, pkr: pkr.is_accepted,
parallel_iterations=1)
# transform samples back to their constrained form
precision_samples = precision_to_unconstrained.inverse(states)
return states, precision_samples, is_accepted
states, precision_samples, is_accepted = sample()
print('True posterior mean:\n', posterior_mean)
print('Sample mean:\n', np.mean(np.reshape(precision_samples, [-1, 2, 2]), axis=0))
# NOW ADAPTIVE STEPSIZE
N_CHAINS = 3
np.random.seed(123)
initial_values = []
for i in range(N_CHAINS):
initial_value_cholesky = np.array(
[[0.5 + np.random.uniform(), 0.0],
[-0.5 + np.random.uniform(), 0.5 + np.random.uniform()]],
dtype = np.float32)
initial_values.append(initial_value_cholesky.dot(initial_value_cholesky.T))
initial_values = np.stack(initial_values)
initial_values_transformed = precision_to_unconstrained.forward(
initial_values).numpy()
@tf.function(autograph=False)
def sample():
tf.random.set_seed(123)
log_lik_fn = get_log_lik_transformed(my_data)
# Tuning acceptance rates:
dtype = np.float32
num_burnin_iter = 3000
num_warmup_iter = int(0.8 * num_burnin_iter)
num_chain_iter = 2500
# Set the target average acceptance ratio for the HMC as suggested by
# Beskos et al. (2013):
# https://projecteuclid.org/download/pdfview_1/euclid.bj/1383661192
target_accept_rate = 0.651
# Initialize the HMC sampler.
hmc = tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn = log_lik_fn,
step_size = 0.01,
num_leapfrog_steps = 3)
# Adapt the step size using standard adaptive MCMC procedure. See Section 4.2
# of Andrieu and Thoms (2008):
# http://www4.ncsu.edu/~rsmith/MA797V_S12/Andrieu08_AdaptiveMCMC_Tutorial.pdf
adapted_kernel = tfp.mcmc.SimpleStepSizeAdaptation(
inner_kernel = hmc,
num_adaptation_steps = num_warmup_iter,
target_accept_prob = target_accept_rate)
states, is_accepted = tfp.mcmc.sample_chain(
num_results = num_chain_iter,
num_burnin_steps = num_burnin_iter,
current_state = initial_values_transformed,
kernel = adapted_kernel,
trace_fn = lambda _, pkr: pkr.inner_results.is_accepted,
parallel_iterations = 1)
# transform samples back to their constrained form
precision_samples = precision_to_unconstrained.inverse(states)
return states, precision_samples, is_accepted
states, precision_samples, is_accepted = sample()
print(np.mean(is_accepted))
precision_samples_reshaped = np.reshape(precision_samples, [-1, 2, 2])
print('True posterior mean:\n', posterior_mean)
print('Mean of samples:\n', np.mean(precision_samples_reshaped, axis=0))
print('True posterior standard deviation:\n', posterior_sd)
print('Standard deviation of samples:\n', np.std(precision_samples_reshaped, axis=0))
r_hat = tfp.mcmc.potential_scale_reduction(precision_samples).numpy()
print(r_hat)
fig, axes = plt.subplots(2, 2, sharey=True)
fig.set_size_inches(8, 8)
for i in range(2):
for j in range(2):
ax = axes[i, j]
ax.hist(precision_samples_reshaped[:, i, j])
ax.axvline(true_precision[i, j], color='red',
label='True precision')
ax.axvline(sample_precision[i, j], color='red', linestyle=':',
label='Sample precision')
ax.set_title('precision[%d, %d]' % (i, j))
plt.tight_layout()
plt.legend()
plt.show()
fig, axes = plt.subplots(4, 4)
fig.set_size_inches(12, 12)
for i1 in range(2):
for j1 in range(2):
index1 = 2 * i1 + j1
for i2 in range(2):
for j2 in range(2):
index2 = 2 * i2 + j2
ax = axes[index1, index2]
ax.scatter(precision_samples_reshaped[:, i1, j1],
precision_samples_reshaped[:, i2, j2], alpha=0.1)
ax.axvline(true_precision[i1, j1], color='red')
ax.axhline(true_precision[i2, j2], color='red')
ax.axvline(sample_precision[i1, j1], color='red', linestyle=':')
ax.axhline(sample_precision[i2, j2], color='red', linestyle=':')
ax.set_title('(%d, %d) vs (%d, %d)' % (i1, j1, i2, j2))
plt.tight_layout()
plt.show()
# WE CAN ALSO USE A TRANSFORMED TRANSITION KERNEL
# The bijector we need for the TransformedTransitionKernel is the inverse of
# the one we used above
unconstrained_to_precision = tfb.Chain([
# step 3: take the product of Cholesky factors
tfb.CholeskyOuterProduct(validate_args=VALIDATE_ARGS),
# step 2: exponentiate the diagonals
tfb.TransformDiagonal(tfb.Exp(validate_args=VALIDATE_ARGS)),
# step 1: map a vector to a lower triangular matrix
tfb.FillTriangular(validate_args=VALIDATE_ARGS),
])
@tf.function(autograph=False)
def sample():
tf.random.set_seed(123)
log_lik_fn = get_log_lik(my_data)
# Tuning acceptance rates:
dtype = np.float32
num_burnin_iter = 3000
num_warmup_iter = int(0.8 * num_burnin_iter)
num_chain_iter = 2500
# Set the target average acceptance ratio for the HMC as suggested by
# Beskos et al. (2013):
# https://projecteuclid.org/download/pdfview_1/euclid.bj/1383661192
target_accept_rate = 0.651
# Initialize the HMC sampler.
hmc = tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn=log_lik_fn,
step_size=0.01,
num_leapfrog_steps=3)
ttk = tfp.mcmc.TransformedTransitionKernel(
inner_kernel=hmc, bijector=unconstrained_to_precision)
# Adapt the step size using standard adaptive MCMC procedure. See Section 4.2
# of Andrieu and Thoms (2008):
# http://www4.ncsu.edu/~rsmith/MA797V_S12/Andrieu08_AdaptiveMCMC_Tutorial.pdf
adapted_kernel = tfp.mcmc.SimpleStepSizeAdaptation(
inner_kernel=ttk,
num_adaptation_steps=num_warmup_iter,
target_accept_prob=target_accept_rate)
states = tfp.mcmc.sample_chain(
num_results=num_chain_iter,
num_burnin_steps=num_burnin_iter,
current_state=initial_values,
kernel=adapted_kernel,
trace_fn=None,
parallel_iterations=1)
# transform samples back to their constrained form
return states
precision_samples = sample()
# quick sanity check
m = [[1., 2.], [2., 8.]]
m_inv = unconstrained_to_precision.inverse(m).numpy()
m_fwd = unconstrained_to_precision.forward(m_inv).numpy()
print('m:\n', m)
print('unconstrained_to_precision.inverse(m):\n', m_inv)
print('forward(unconstrained_to_precision.inverse(m)):\n', m_fwd)
precision_samples_reshaped = np.reshape(precision_samples, [-1, 2, 2])
print('True posterior mean:\n', posterior_mean)
print('Mean of samples:\n', np.mean(precision_samples_reshaped, axis=0))
r_hat = tfp.mcmc.potential_scale_reduction(precision_samples).numpy()
print(r_hat)
# 8 Schools Experiment
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import seaborn as sns
import warnings
plt.style.use("ggplot")
warnings.filterwarnings('ignore')
num_schools = 8
treatment_effects = np.array(
[30, 10, -5, 10, -2, 2, 20, 15], dtype = np.float32)
# treatment_effects = np.array(
# [28, 8, -3, 7, -1, 1, 18, 12], dtype = np.float32) # treatment effects
treatment_stddevs = np.array(
[15, 10, 16, 11, 9, 11, 10, 18], dtype = np.float32) # treatment SE
fig, ax = plt.subplots()
plt.bar(range(num_schools), treatment_effects, yerr=treatment_stddevs)
plt.title("8 Schools treatment effects")
plt.xlabel("School")
plt.ylabel("Treatment effect")
fig.set_size_inches(10, 8)
plt.show()
model = tfd.JointDistributionSequential([
tfd.Normal(loc = 0., scale = 10., name = "avg_effect"), # `mu` above
tfd.Normal(loc = 5., scale = 1., name = "avg_stddev"), # `log(tau)` above
tfd.Independent(tfd.Normal(loc = tf.zeros(num_schools),
scale = tf.ones(num_schools),
name = "school_effects_standard"), # `theta_prime`
reinterpreted_batch_ndims = 1),
lambda school_effects_standard, avg_stddev, avg_effect: (
tfd.Independent(tfd.Normal(loc = (avg_effect[..., tf.newaxis] +
tf.exp(avg_stddev[..., tf.newaxis]) *
school_effects_standard), # `theta` above
scale = treatment_stddevs),
name = "treatment_effects", # `y` above
reinterpreted_batch_ndims = 1))
])
def target_log_prob_fn(avg_effect, avg_stddev, school_effects_standard):
"""Unnormalized target density as a function of states."""
# Note: We rely on the fact that treatment effects are specified outside of the function
return model.log_prob((
avg_effect, avg_stddev, school_effects_standard, treatment_effects))
num_results = 5000
num_burnin_steps = 3000
# Improve performance by tracing the sampler using `tf.function`
# and compiling it using XLA.
@tf.function(autograph = False)
def do_sampling():
return tfp.mcmc.sample_chain(
num_results = num_results,
num_burnin_steps = num_burnin_steps,
current_state = [
tf.zeros([], name = 'init_avg_effect'),
tf.zeros([], name = 'init_avg_stddev'),
tf.ones([num_schools], name = 'init_school_effects_standard'),
],
kernel = tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn = target_log_prob_fn,
step_size = 0.4,
num_leapfrog_steps = 3))
states, kernel_results = do_sampling()
avg_effect, avg_stddev, school_effects_standard = states
school_effects_samples = (
avg_effect[:, np.newaxis] +
np.exp(avg_stddev)[:, np.newaxis] * school_effects_standard)
num_accepted = np.sum(kernel_results.is_accepted)
print('Acceptance rate: {}'.format(num_accepted / num_results))
fig, axes = plt.subplots(8, 2, sharex='col', sharey='col')
fig.set_size_inches(12, 10)
for i in range(num_schools):
axes[i][0].plot(school_effects_samples[:,i].numpy())
axes[i][0].title.set_text("School {} treatment effect chain".format(i))
sns.kdeplot(school_effects_samples[:,i].numpy(), ax=axes[i][1], shade=True)
axes[i][1].title.set_text("School {} treatment effect distribution".format(i))
axes[num_schools - 1][0].set_xlabel("Iteration")
axes[num_schools - 1][1].set_xlabel("School effect")
fig.tight_layout()
plt.show()
print("E[avg_effect] = {}".format(np.mean(avg_effect)))
print("E[avg_stddev] = {}".format(np.mean(avg_stddev)))
print("E[school_effects_standard] =")
print(np.mean(school_effects_standard[:, ]))
print("E[school_effects] =")
print(np.mean(school_effects_samples[:, ], axis=0))
# Compute the 95% interval for school_effects
school_effects_low = np.array([
np.percentile(school_effects_samples[:, i], 2.5) for i in range(num_schools)
])
school_effects_med = np.array([
np.percentile(school_effects_samples[:, i], 50) for i in range(num_schools)
])
school_effects_hi = np.array([
np.percentile(school_effects_samples[:, i], 97.5)
for i in range(num_schools)
])
fig, ax = plt.subplots(nrows=1, ncols=1, sharex=True)
ax.scatter(np.array(range(num_schools)), school_effects_med, color='red', s=60)
ax.scatter(
np.array(range(num_schools)) + 0.1, treatment_effects, color='blue', s=60)
plt.plot([-0.2, 7.4], [np.mean(avg_effect),
np.mean(avg_effect)], 'k', linestyle='--')
ax.errorbar(
np.array(range(8)),
school_effects_med,
yerr=[
school_effects_med - school_effects_low,
school_effects_hi - school_effects_med
],
fmt='none')
ax.legend(('avg_effect', 'HMC', 'Observed effect'), fontsize=14)
plt.xlabel('School')
plt.ylabel('Treatment effect')
plt.title('HMC estimated school treatment effects vs. observed data')
fig.set_size_inches(10, 8)
plt.show()
# Bayesian Gaussian Mixture
# We'll use the following directory to store files we download as well as our
# preprocessed dataset.
CACHE_DIR = os.path.join(os.sep, 'tmp', 'radon')
def cache_or_download_file(cache_dir, url_base, filename):
"""Read a cached file or download it."""
filepath = os.path.join(cache_dir, filename)
if tf.io.gfile.exists(filepath):
return filepath
if not tf.io.gfile.exists(cache_dir):
tf.io.gfile.makedirs(cache_dir)
url = os.path.join(url_base, filename)
print("Downloading {url} to {filepath}.".format(url=url, filepath=filepath))
urllib.request.urlretrieve(url, filepath)
return filepath
def download_radon_dataset(cache_dir=CACHE_DIR):
"""Download the radon dataset and read as Pandas dataframe."""
url_base = 'http://www.stat.columbia.edu/~gelman/arm/examples/radon/'
# Alternative source:
# url_base = ('https://raw.githubusercontent.com/pymc-devs/uq_chapter/'
# 'master/reference/data/')
srrs2 = pd.read_csv(cache_or_download_file(cache_dir, url_base, 'srrs2.dat'))
srrs2.rename(columns=str.strip, inplace=True)
cty = pd.read_csv(cache_or_download_file(cache_dir, url_base, 'cty.dat'))
cty.rename(columns=str.strip, inplace=True)
return srrs2, cty
def preprocess_radon_dataset(srrs2, cty, state='MN'):
"""Preprocess radon dataset as done in "Bayesian Data Analysis" book."""
srrs2 = srrs2[srrs2.state==state].copy()
cty = cty[cty.st==state].copy()
# We will now join datasets on Federal Information Processing Standards
# (FIPS) id, ie, codes that link geographic units, counties and county
# equivalents. http://jeffgill.org/Teaching/rpqm_9.pdf
srrs2['fips'] = 1000 * srrs2.stfips + srrs2.cntyfips
cty['fips'] = 1000 * cty.stfips + cty.ctfips
df = srrs2.merge(cty[['fips', 'Uppm']], on='fips')
df = df.drop_duplicates(subset='idnum')
df = df.rename(index=str, columns={'Uppm': 'uranium_ppm'})
# For any missing or invalid activity readings, we'll use a value of `0.1`.
df['radon'] = df.activity.apply(lambda x: x if x > 0. else 0.1)
# Remap categories to start from 0 and end at max(category).
county_name = sorted(df.county.unique())
df['county'] = df.county.astype(
pd.api.types.CategoricalDtype(categories=county_name)).cat.codes
county_name = map(str.strip, county_name)
df['log_radon'] = df['radon'].apply(np.log)
df['log_uranium_ppm'] = df['uranium_ppm'].apply(np.log)
df = df[['log_radon', 'floor', 'county', 'log_uranium_ppm']]
return df, county_name
radon, county_name = preprocess_radon_dataset(*download_radon_dataset())
radon.head()
fig, ax = plt.subplots(figsize=(22, 5));
county_freq = radon['county'].value_counts()
county_freq.plot(kind='bar', color='#436bad');
plt.xlabel('County index')
plt.ylabel('Number of radon readings')
plt.title('Number of radon readings per county', fontsize=16)
county_freq = np.array(zip(county_freq.index, county_freq.values)) # We'll use this later.
fig, ax = plt.subplots(ncols=2, figsize=[10, 4]);
radon['log_radon'].plot(kind='density', ax=ax[0]);
ax[0].set_xlabel('log(radon)')
radon['floor'].value_counts().plot(kind='bar', ax=ax[1]);
ax[1].set_xlabel('Floor');
ax[1].set_ylabel('Count');
fig.subplots_adjust(wspace=0.25)
# Handy snippet to reset the global graph and global session.
tf.reset_default_graph()
with warnings.catch_warnings():
warnings.simplefilter('ignore')
tf.reset_default_graph()
try:
sess.close()
except:
pass
sess = tf.InteractiveSession()
inv_scale_transform = lambda y: np.log(y) # Not using TF here.
fwd_scale_transform = tf.exp
def _make_weights_prior(num_counties, dtype):
"""Returns a `len(log_uranium_ppm)` batch of univariate Normal."""
raw_prior_scale = tf.get_variable(
name='raw_prior_scale',
initializer=np.array(inv_scale_transform(1.), dtype=dtype))
return tfp.distributions.Independent(
tfp.distributions.Normal(
loc=tf.zeros(num_counties, dtype=dtype),
scale=fwd_scale_transform(raw_prior_scale)),
reinterpreted_batch_ndims=1)
make_weights_prior = tf.make_template(
name_='make_weights_prior', func_=_make_weights_prior)
```
| github_jupyter |
___
<a href='http://www.pieriandata.com'> <img src='../../Pierian_Data_Logo.png' /></a>
___
# Matplotlib Overview Lecture
## Introduction
Matplotlib is the "grandfather" library of data visualization with Python. It was created by John Hunter. He created it to try to replicate MatLab's (another programming language) plotting capabilities in Python. So if you happen to be familiar with matlab, matplotlib will feel natural to you.
It is an excellent 2D and 3D graphics library for generating scientific figures.
Some of the major Pros of Matplotlib are:
* Generally easy to get started for simple plots
* Support for custom labels and texts
* Great control of every element in a figure
* High-quality output in many formats
* Very customizable in general
Matplotlib allows you to create reproducible figures programmatically. Let's learn how to use it! Before continuing this lecture, I encourage you just to explore the official Matplotlib web page: http://matplotlib.org/
## Installation
You'll need to install matplotlib first with either:
conda install matplotlib
or
pip install matplotlib
## Importing
Import the `matplotlib.pyplot` module under the name `plt` (the tidy way):
```
import matplotlib.pyplot as plt
```
You'll also need to use this line to see plots in the notebook:
```
%matplotlib inline
```
That line is only for jupyter notebooks, if you are using another editor, you'll use: **plt.show()** at the end of all your plotting commands to have the figure pop up in another window.
# Basic Example
Let's walk through a very simple example using two numpy arrays:
### Example
Let's walk through a very simple example using two numpy arrays. You can also use lists, but most likely you'll be passing numpy arrays or pandas columns (which essentially also behave like arrays).
** The data we want to plot:**
```
import numpy as np
x = np.linspace(0, 5, 11)
y = x ** 2
x
y
```
## Basic Matplotlib Commands
We can create a very simple line plot using the following ( I encourage you to pause and use Shift+Tab along the way to check out the document strings for the functions we are using).
```
plt.plot(x, y, 'r') # 'r' is the color red
plt.xlabel('X Axis Title Here')
plt.ylabel('Y Axis Title Here')
plt.title('String Title Here')
plt.show()
```
## Creating Multiplots on Same Canvas
```
# plt.subplot(nrows, ncols, plot_number)
plt.subplot(1, 2, 1)
plt.plot(x, y, 'r--') # More on color options later
plt.subplot(1, 2, 2)
plt.plot(y, x, 'g*-');
```
___
# Matplotlib Object Oriented Method
Now that we've seen the basics, let's break it all down with a more formal introduction of Matplotlib's Object Oriented API. This means we will instantiate figure objects and then call methods or attributes from that object.
## Introduction to the Object Oriented Method
The main idea in using the more formal Object Oriented method is to create figure objects and then just call methods or attributes off of that object. This approach is nicer when dealing with a canvas that has multiple plots on it.
To begin we create a figure instance. Then we can add axes to that figure:
```
# Create Figure (empty canvas)
fig = plt.figure()
# Add set of axes to figure
axes = fig.add_axes([0.1, 0.1, 0.8, 0.8]) # left, bottom, width, height (range 0 to 1)
# Plot on that set of axes
axes.plot(x, y, 'b')
axes.set_xlabel('Set X Label') # Notice the use of set_ to begin methods
axes.set_ylabel('Set y Label')
axes.set_title('Set Title')
```
Code is a little more complicated, but the advantage is that we now have full control of where the plot axes are placed, and we can easily add more than one axis to the figure:
```
# Creates blank canvas
fig = plt.figure()
axes1 = fig.add_axes([0.1, 0.1, 0.8, 0.8]) # main axes
axes2 = fig.add_axes([0.2, 0.5, 0.4, 0.3]) # inset axes
# Larger Figure Axes 1
axes1.plot(x, y, 'b')
axes1.set_xlabel('X_label_axes2')
axes1.set_ylabel('Y_label_axes2')
axes1.set_title('Axes 2 Title')
# Insert Figure Axes 2
axes2.plot(y, x, 'r')
axes2.set_xlabel('X_label_axes2')
axes2.set_ylabel('Y_label_axes2')
axes2.set_title('Axes 2 Title');
```
## subplots()
The plt.subplots() object will act as a more automatic axis manager.
Basic use cases:
```
# Use similar to plt.figure() except use tuple unpacking to grab fig and axes
fig, axes = plt.subplots()
# Now use the axes object to add stuff to plot
axes.plot(x, y, 'r')
axes.set_xlabel('x')
axes.set_ylabel('y')
axes.set_title('title');
```
Then you can specify the number of rows and columns when creating the subplots() object:
```
# Empty canvas of 1 by 2 subplots
fig, axes = plt.subplots(nrows = 1,
ncols = 2)
# Axes is an array of axes to plot on
axes
```
We can iterate through this array:
```
for ax in axes:
ax.plot(x, y, 'b')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_title('title')
# Display the figure object
fig
```
A common issue with matplolib is overlapping subplots or figures. We ca use **fig.tight_layout()** or **plt.tight_layout()** method, which automatically adjusts the positions of the axes on the figure canvas so that there is no overlapping content:
```
fig, axes = plt.subplots(nrows=1, ncols=2)
for ax in axes:
ax.plot(x, y, 'g')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_title('title')
fig
plt.tight_layout()
```
### Figure size, aspect ratio and DPI
Matplotlib allows the aspect ratio, DPI and figure size to be specified when the Figure object is created. You can use the `figsize` and `dpi` keyword arguments.
* `figsize` is a tuple of the width and height of the figure in inches
* `dpi` is the dots-per-inch (pixel per inch).
For example:
```
fig = plt.figure(figsize = (8, 4),
dpi = 100)
```
The same arguments can also be passed to layout managers, such as the `subplots` function:
```
fig, axes = plt.subplots(figsize=(12,3))
axes.plot(x, y, 'r')
axes.set_xlabel('x')
axes.set_ylabel('y')
axes.set_title('title');
```
## Saving figures
Matplotlib can generate high-quality output in a number formats, including PNG, JPG, EPS, SVG, PGF and PDF.
To save a figure to a file we can use the `savefig` method in the `Figure` class:
```
fig.savefig("filename.png")
```
Here we can also optionally specify the DPI and choose between different output formats:
```
fig.savefig("filename.png", dpi=200)
```
____
## Legends, labels and titles
Now that we have covered the basics of how to create a figure canvas and add axes instances to the canvas, let's look at how decorate a figure with titles, axis labels, and legends.
**Figure titles**
A title can be added to each axis instance in a figure. To set the title, use the `set_title` method in the axes instance:
```
ax.set_title("title");
```
**Axis labels**
Similarly, with the methods `set_xlabel` and `set_ylabel`, we can set the labels of the X and Y axes:
```
ax.set_xlabel("x")
ax.set_ylabel("y");
```
### Legends
You can use the **label="label text"** keyword argument when plots or other objects are added to the figure, and then using the **legend** method without arguments to add the legend to the figure:
```
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
ax.plot(x,
x ** 2,
label = "x**2")
ax.plot(x,
x ** 3,
label= "x**3")
ax.legend()
```
Notice how are legend overlaps some of the actual plot!
The **legend** function takes an optional keyword argument **loc** that can be used to specify where in the figure the legend is to be drawn. The allowed values of **loc** are numerical codes for the various places the legend can be drawn. See the [documentation page](http://matplotlib.org/users/legend_guide.html#legend-location) for details. Some of the most common **loc** values are:
```
# Lots of options....
ax.legend(loc = 1) # upper right corner
ax.legend(loc = 2) # upper left corner
ax.legend(loc = 3) # lower left corner
ax.legend(loc = 4) # lower right corner
# .. many more options are available
# Most common to choose
ax.legend(loc = 0) # let matplotlib decide the optimal location
fig
```
## Setting colors, linewidths, linetypes
Matplotlib gives you *a lot* of options for customizing colors, linewidths, and linetypes.
There is the basic MATLAB like syntax (which I would suggest you avoid using for more clairty sake:
### Colors with MatLab like syntax
With matplotlib, we can define the colors of lines and other graphical elements in a number of ways. First of all, we can use the MATLAB-like syntax where `'b'` means blue, `'g'` means green, etc. The MATLAB API for selecting line styles are also supported: where, for example, 'b.-' means a blue line with dots:
```
# MATLAB style line color and style
fig, ax = plt.subplots()
ax.plot(x,
x ** 2,
'b.-') # blue line with dots
ax.plot(x,
x ** 3,
'g--') # green dashed line
```
### Colors with the color= parameter
We can also define colors by their names or RGB hex codes and optionally provide an alpha value using the `color` and `alpha` keyword arguments. Alpha indicates opacity.
```
fig, ax = plt.subplots()
ax.plot(x, x + 1, color = "blue", alpha=0.5) # half-transparant
ax.plot(x, x + 2, color = "#8B008B") # RGB hex code
ax.plot(x, x + 3, color = "#FF8C00") # RGB hex code
```
### Line and marker styles
To change the line width, we can use the `linewidth` or `lw` keyword argument. The line style can be selected using the `linestyle` or `ls` keyword arguments:
```
fig, ax = plt.subplots(figsize=(12,6))
ax.plot(x, x + 1, color = "red", linewidth = 0.25)
ax.plot(x, x + 2, color = "red", linewidth = 0.50)
ax.plot(x, x + 3, color = "red", linewidth = 1.00)
ax.plot(x, x + 4, color = "red", linewidth = 2.00)
# possible linestype options ‘-‘, ‘–’, ‘-.’, ‘:’, ‘steps’
ax.plot(x, x + 5, color = "green", lw = 3, linestyle = '-')
ax.plot(x, x + 6, color = "green", lw = 3, ls = '-.')
ax.plot(x, x + 7, color = "green", lw = 3, ls = ':')
# custom dash
line, = ax.plot(x, x + 8, color = "black", lw = 1.50)
line.set_dashes([5, 10, 15, 10]) # format: line length, space length, ...
# possible marker symbols: marker = '+', 'o', '*', 's', ',', '.', '1', '2', '3', '4', ...
ax.plot(x, x + 9, color = "blue", lw = 3, ls = '-', marker = '+')
ax.plot(x, x + 10, color = "blue", lw = 3, ls = '--', marker = 'o')
ax.plot(x, x + 11, color = "blue", lw = 3, ls = '-', marker = 's')
ax.plot(x, x + 12, color = "blue", lw = 3, ls = '--', marker = '1')
# marker size and color
ax.plot(x, x + 13, color = "purple", lw = 1, ls = '-', marker = 'o', markersize = 2)
ax.plot(x, x + 14, color = "purple", lw = 1, ls = '-', marker = 'o', markersize = 4)
ax.plot(x, x + 15, color = "purple", lw = 1, ls = '-', marker = 'o', markersize = 8, markerfacecolor = "red")
ax.plot(x, x + 16, color = "purple", lw = 1, ls = '-', marker = 's', markersize = 8,
markerfacecolor = "yellow", markeredgewidth = 3, markeredgecolor = "green");
```
### Control over axis appearance
In this section we will look at controlling axis sizing properties in a matplotlib figure.
## Plot range
We can configure the ranges of the axes using the `set_ylim` and `set_xlim` methods in the axis object, or `axis('tight')` for automatically getting "tightly fitted" axes ranges:
```
fig, axes = plt.subplots(1, 3, figsize = (12, 4))
axes[0].plot(x, x ** 2, x, x ** 3)
axes[0].set_title("default axes ranges")
axes[1].plot(x, x ** 2, x, x ** 3)
axes[1].axis('tight')
axes[1].set_title("tight axes")
axes[2].plot(x, x ** 2, x, x ** 3)
axes[2].set_ylim([0, 60])
axes[2].set_xlim([2, 5])
axes[2].set_title("custom axes range");
```
# Special Plot Types
There are many specialized plots we can create, such as barplots, histograms, scatter plots, and much more. Most of these type of plots we will actually create using pandas. But here are a few examples of these type of plots:
```
plt.scatter(x, y)
from random import sample
data = sample(range(1, 1000), 100)
plt.hist(data)
data = [np.random.normal(0, std, 100) for std in range(1, 4)]
# rectangular box plot
plt.boxplot(data,
vert = True,
patch_artist = True);
```
## Further reading
* http://www.matplotlib.org - The project web page for matplotlib.
* https://github.com/matplotlib/matplotlib - The source code for matplotlib.
* http://matplotlib.org/gallery.html - A large gallery showcaseing various types of plots matplotlib can create. Highly recommended!
* http://www.loria.fr/~rougier/teaching/matplotlib - A good matplotlib tutorial.
* http://scipy-lectures.github.io/matplotlib/matplotlib.html - Another good matplotlib reference.
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(context = 'notebook', #The base context is “notebook”, and the other contexts are “paper”, “talk”, and “poster”
style = 'darkgrid', #dict, None, or one of {darkgrid, whitegrid, dark, white, ticks}
palette = 'deep', # Should be something that color_palette() can process.
font_scale = 1,
color_codes = False,
rc = None)
from IPython.display import display
%matplotlib notebook
np.version.version, pd.__version__
# some functions to load
def head_with_full_columns(pd_in, row_amount = 5):
with pd.option_context('display.max_columns', len(pd_in.iloc[0])):
display(pd_in[:row_amount])
def balanced_sample(df_in, total_size, rand_state):
s0 = df_in[df_in['TARGET']==0].sample(n = total_size//2, random_state = rand_state)
s1 = df_in[df_in['TARGET']==1].sample(n = total_size//2, random_state = rand_state)
new_df = pd.concat([s0,s1])
new_df.sort_index(inplace = True)
return new_df
def which_df(feature):
if feature in application_train_df_f_list:
return feature, 'application_train_df'
if feature in bureau_df_f_list:
return feature, 'bureau_df'
if feature in bureau_balance_df_f_list:
return feature, 'bureau_balance_df'
if feature in credit_card_balance_df_f_list:
return feature, 'credit_card_balance_df'
if feature in installments_payments_df_f_list:
return feature, 'installments_payments_df'
if feature in POS_CASH_balance_df_f_list:
return feature, 'POS_CASH_balance_df'
if feature in previous_application_df_f_list:
return feature, 'previous_application_df'
pwd
cd '/Users/DonBunk/Desktop/Google Drive/data_science/Python_Projects/Home_Credit_Default_Risk/'
path_w = 'raw_loan_data_from_Kaggle/'
path_a = 'aggregation/TEST_aggregation/'
test_SK_ID_index_df = pd.read_csv(path_w + 'application_test.csv', usecols=['SK_ID_CURR'], index_col = 'SK_ID_CURR')
test_SK_ID_index_df.sort_values(by = ['SK_ID_CURR'], inplace=True)
```
# aggregate data for each df and merge dfs
## load bureau csv
```
bureau_df = pd.read_csv(path_w + 'bureau.csv')#, index_col = 'SK_ID_CURR')
# replace NaNs in object entries for now is pretty safe.
f_list = list(bureau_df.select_dtypes('object').columns)
bureau_df[f_list] = bureau_df[f_list].fillna(value = 'NA')
# AMT_ANNUITY appears in multiple files, so rename more specifically.
bureau_df.rename( columns = {"AMT_ANNUITY":"AMT_ANNUITY_from_bureau"} , inplace=True)# index = str,
del f_list
```
### aggregate and define df
```
bureau_df.info(verbose = True, null_counts = True)
len(np.unique(bureau_df['SK_ID_CURR'].values))
```
All the floats are really floats, and all the integers are continous numerical.
```
# group interger features
bureau_df_float_list = list(bureau_df.select_dtypes('float64').columns)
bureau_df_float_grouped = bureau_df[bureau_df_float_list].groupby(bureau_df['SK_ID_CURR'])
# apply functions to these features and name them appropriately
function_list = ['mean', 'median','max','min']
bureau_df_float_grouped_agg = bureau_df_float_grouped.agg(function_list)
new_cols = ['_'.join(col).strip() for col in bureau_df_float_grouped_agg.columns.values]
bureau_df_float_grouped_agg.columns = new_cols
# group float features, remove ID features
bureau_df_int_list = list( bureau_df.select_dtypes('int64').columns )
bureau_df_int_list.remove('SK_ID_CURR')
bureau_df_int_list.remove('SK_ID_BUREAU')
bureau_df_int_grouped = bureau_df[bureau_df_int_list].groupby(bureau_df['SK_ID_CURR'])
# apply functions to these features and name them appropriately
function_list = ['mean', 'median','max','min']
bureau_df_int_grouped_agg = bureau_df_int_grouped.agg(function_list)
new_cols = ['_'.join(col).strip() for col in bureau_df_int_grouped_agg.columns.values]
bureau_df_int_grouped_agg.columns = new_cols
# group string features
bureau_df_string_list = list(bureau_df.select_dtypes('object').columns)
bureau_df_string_grouped = bureau_df[bureau_df_string_list].groupby(bureau_df['SK_ID_CURR'])
# calculate the mode for each aggregated feature
# this takes a VERY long time to run
bureau_df_string_grouped_agg = bureau_df_string_grouped.agg(lambda x:x.value_counts().index[0])
bureau_df_string_grouped_agg.columns = [x + '_mode' for x in bureau_df_string_grouped_agg.columns]
# merge int, float, and string features into new df
bureau_df_float_and_int_grouped = pd.merge(bureau_df_float_grouped_agg, bureau_df_int_grouped_agg, left_index=True, right_index=True)
bureau_df_aggregated_final = pd.merge(bureau_df_float_and_int_grouped, bureau_df_string_grouped_agg, left_index=True, right_index=True)
bureau_df_aggregated_final.info()
# add in all SK_IDS, which will appear as NaNs in new df
bureau_df_aggregated_final = pd.merge(test_SK_ID_index_df, bureau_df_aggregated_final, left_index=True, right_index=True, how = 'left')
bureau_df_aggregated_final.to_csv(path_a + 'bureau_df_aggregated_final.csv', columns = list(bureau_df_aggregated_final.columns))
del bureau_df
del bureau_df_float_list
del bureau_df_float_grouped
del bureau_df_float_grouped_agg
del bureau_df_int_list
del bureau_df_int_grouped
del function_list
del bureau_df_int_grouped_agg
del bureau_df_string_list
del bureau_df_string_grouped
del bureau_df_string_grouped_agg
del bureau_df_float_and_int_grouped
del new_cols
```
## installments_payments_df
```
installments_payments_df = pd.read_csv(path_w + 'installments_payments.csv')#, index_col = 'SK_ID_CURR')
# replace NaNs in object entries for now is pretty safe.
f_list = list(installments_payments_df.select_dtypes('object').columns)
installments_payments_df[f_list] = installments_payments_df[f_list].fillna(value = 'NA')
del f_list
```
### add new column features to full df
```
installments_payments_df.info(null_counts=True)
# add these new features in full DF.
installments_payments_df['DAYS_PAYMENT_LATE'] = installments_payments_df['DAYS_ENTRY_PAYMENT'] - installments_payments_df['DAYS_INSTALMENT']
installments_payments_df['AMT_OVERPAY'] = installments_payments_df['AMT_PAYMENT'] - installments_payments_df['AMT_INSTALMENT']
# organize indices to call later.
installments_payments_df.set_index(['SK_ID_CURR','SK_ID_PREV', 'NUM_INSTALMENT_NUMBER'], inplace = True)
installments_payments_df.sort_index(inplace=True)
# this takes WHILE to run for the full df.
# peel off, group, and find diffs of NUM_INSTALMENT_VERSION to see how often loan payment calender changed.
# NOTE YOU NEED DOUBLE SQUARE BRACKETS BELOW SO THAT YOU DEFINE A DF NOT A SERIES, group by and diff only seem to work on former!
NUM_INSTALMENT_VERSION_df = installments_payments_df[['NUM_INSTALMENT_VERSION']]
NUM_INSTALMENT_VERSION_df_grouped = NUM_INSTALMENT_VERSION_df[['NUM_INSTALMENT_VERSION']].groupby(level = 1)
# THIS TAKES AWHILE!
NUM_INSTALMENT_VERSION_df_grouped_diff = NUM_INSTALMENT_VERSION_df_grouped.diff()
# rename this new feature and join with original full DF.
NUM_INSTALMENT_VERSION_df_grouped_diff['NUM_INSTALMENT_VERSION_diff'] = NUM_INSTALMENT_VERSION_df_grouped_diff['NUM_INSTALMENT_VERSION']
installments_payments_df = installments_payments_df.join( NUM_INSTALMENT_VERSION_df_grouped_diff['NUM_INSTALMENT_VERSION_diff'])
# reset NUM_INSTALMENT_NUMBER and SK_ID_PREV to columns
installments_payments_df.reset_index(level=2, inplace=True)
installments_payments_df.reset_index(level=1, inplace=True)
def count_unique(list_in):
return len(np.unique(list_in))
def count_postive(list_in):
return len([x for x in list_in if x > 0])
def count_negative(list_in):
return len([x for x in list_in if x < 0])
installments_payments_df_grouped = installments_payments_df.groupby('SK_ID_CURR')
# create the new feature and make a new summary feature DF out of it.
num_of_loans = installments_payments_df_grouped['SK_ID_PREV'].agg([('NUM_OF_LOANS', count_unique)])
installments_payments_df_summary = num_of_loans
# demote SK_ID_CURR index to column for aggregation
installments_payments_df.reset_index(level=0, inplace=True)
# group for aggregation
installments_payments_df_grouped = installments_payments_df.groupby('SK_ID_CURR')
# early/ late features.
agg_list = [('DAYS_PAYMENT_LATE_mean', np.mean),
('DAYS_PAYMENT_LATE_median', np.median),
('DAYS_PAYMENT_LATE_sd', np.std),
('NUM_TIMES_LATE', count_postive),
('NUM_TIMES_EARLY', count_negative)]
some_feats = installments_payments_df_grouped['DAYS_PAYMENT_LATE'].agg(agg_list)
installments_payments_df_summary = installments_payments_df_summary.join(some_feats)
# over/under pay features.
agg_list = [('AMT_OVERPAY_MEAN', np.mean),
('AMT_OVERPAY_MEDIAN', np.median),
('AMT_OVERPAY_SD', np.std),
('NUM_TIMES_OVERPAY', count_postive),
('NUM_TIMES_UNDERPAY', count_negative)]
some_feats = installments_payments_df_grouped['NUM_INSTALMENT_VERSION_diff'].agg(agg_list)
installments_payments_df_summary = installments_payments_df_summary.join(some_feats)
# loan terms change features.
agg_list = [('TERMS_CHANGE_TIMES', np.count_nonzero)]
some_feats = installments_payments_df_grouped['NUM_INSTALMENT_VERSION_diff'].agg(agg_list)
installments_payments_df_summary = installments_payments_df_summary.join(some_feats)
installments_payments_df_summary['TERMS_CHANGE_PER_LOAN'] \
= installments_payments_df_summary['TERMS_CHANGE_TIMES']/installments_payments_df_summary['NUM_OF_LOANS']
installments_payments_df_summary.info(verbose = True, null_counts = True)
del NUM_INSTALMENT_VERSION_df
del NUM_INSTALMENT_VERSION_df_grouped
del NUM_INSTALMENT_VERSION_df_grouped_diff
del num_of_loans
del installments_payments_df_grouped
del agg_list
del some_feats
```
### basic aggregate data
```
# have to drop these columns if I have previously ran the section above.
installments_payments_df.drop(columns = ['DAYS_PAYMENT_LATE','AMT_OVERPAY','NUM_INSTALMENT_VERSION_diff'], inplace = True)
installments_payments_df.info(verbose = True, null_counts = True)
```
All the floats are really floats, and all the integers are continous numerical.
No string features.
```
installments_payments_df_float_grouped = installments_payments_df.groupby(installments_payments_df['SK_ID_CURR'])
# group the float features
installments_payments_df_float_list = list(installments_payments_df.select_dtypes('float64').columns)
installments_payments_df_float_grouped = \
installments_payments_df[installments_payments_df_float_list].groupby(installments_payments_df['SK_ID_CURR'])
# apply these functions to the aggregated feaures, and name acoordingly
function_list = ['mean', 'median', 'max', 'min']
installments_payments_df_float_grouped_agg = installments_payments_df_float_grouped.agg(function_list)
new_cols = ['_'.join(col).strip() for col in installments_payments_df_float_grouped_agg.columns.values]
installments_payments_df_float_grouped_agg.columns = new_cols
# group the int features
installments_payments_df_int_list = list( installments_payments_df.select_dtypes('int64').columns )
installments_payments_df_int_list.remove('SK_ID_CURR')
installments_payments_df_int_list.remove('SK_ID_PREV')
installments_payments_df_int_grouped = \
installments_payments_df[installments_payments_df_int_list].groupby(installments_payments_df['SK_ID_CURR'])
# apply these functions to the aggregated feaures, and name acoordingly
function_list = ['mean', 'median','max','min']
installments_payments_df_int_grouped_agg = installments_payments_df_int_grouped.agg(function_list)
new_cols = ['_'.join(col).strip() for col in installments_payments_df_int_grouped_agg.columns.values]
installments_payments_df_int_grouped_agg.columns = new_cols
# merge all the above as new df
installments_payments_df_aggregated_final = \
pd.merge(installments_payments_df_float_grouped_agg, installments_payments_df_int_grouped_agg, left_index=True, right_index=True)
del installments_payments_df_float_list
del installments_payments_df_float_grouped
del function_list
del installments_payments_df_float_grouped_agg
del new_cols
del installments_payments_df_int_list
del installments_payments_df_int_grouped
del installments_payments_df_int_grouped_agg
```
### merge dfs from two sections above
```
# add in all SK_IDS, which will appear as NaNs in new df
installments_payments_df_final = pd.merge(test_SK_ID_index_df, installments_payments_df_summary, left_index=True, right_index=True, how = 'left')
installments_payments_df_final = pd.merge(installments_payments_df_final, installments_payments_df_aggregated_final, left_index=True, right_index=True, how = 'left')
installments_payments_df_final.to_csv(path_a + 'installments_payments_df_final.csv', columns = list(installments_payments_df_final.columns))
installments_payments_df_final.columns
del installments_payments_df_summary
del installments_payments_df
del installments_payments_df_final
del installments_payments_df_aggregated_final
```
## POS_CASH_balance_df
```
POS_CASH_balance_df = pd.read_csv(path_w + 'POS_CASH_balance.csv')
# replace NaNs in object entries for now is pretty safe.
f_list = list(POS_CASH_balance_df.select_dtypes('object').columns)
POS_CASH_balance_df[f_list] = POS_CASH_balance_df[f_list].fillna(value = 'NA')
del f_list
```
### Aggregate data
```
POS_CASH_balance_df.info(verbose = True, null_counts = True)
```
All the floats are really floats, and all the integers are continous numerical.
```
# group all the float features and aggregate
POS_CASH_balance_df_float_list = list(POS_CASH_balance_df.select_dtypes('float64').columns)
POS_CASH_balance_df_float_grouped = POS_CASH_balance_df[POS_CASH_balance_df_float_list].groupby(POS_CASH_balance_df['SK_ID_CURR'])
# apply the functions to the aggregated features and name accordingly
function_list = ['mean', 'median','max','min']
POS_CASH_balance_df_float_grouped_agg = POS_CASH_balance_df_float_grouped.agg(function_list)
new_cols = ['_'.join(col).strip() for col in POS_CASH_balance_df_float_grouped_agg.columns.values]
POS_CASH_balance_df_float_grouped_agg.columns = new_cols
# group all int features and aggregate, drop ID features
POS_CASH_balance_df_int_list = list( POS_CASH_balance_df.select_dtypes('int64').columns )
POS_CASH_balance_df_int_list.remove('SK_ID_CURR')
POS_CASH_balance_df_int_list.remove('SK_ID_PREV')
POS_CASH_balance_df_int_grouped = POS_CASH_balance_df[POS_CASH_balance_df_int_list].groupby(POS_CASH_balance_df['SK_ID_CURR'])
# apply the functions to the aggregated features and name accordingly
function_list = ['mean', 'median','max','min']
POS_CASH_balance_df_int_grouped_agg = POS_CASH_balance_df_int_grouped.agg(function_list)
new_cols = ['_'.join(col).strip() for col in POS_CASH_balance_df_int_grouped_agg.columns.values]
POS_CASH_balance_df_int_grouped_agg.columns = new_cols
# group all object features and aggregate
POS_CASH_balance_df_string_list = list(POS_CASH_balance_df.select_dtypes('object').columns)
POS_CASH_balance_df_string_grouped = POS_CASH_balance_df[POS_CASH_balance_df_string_list].groupby(POS_CASH_balance_df['SK_ID_CURR'])
# calculate the mode for these features and name accordingly
# this takes a long time to run
POS_CASH_balance_df_string_grouped_agg = POS_CASH_balance_df_string_grouped.agg(lambda x:x.value_counts().index[0])
POS_CASH_balance_df_string_grouped_agg.columns = [x + '_mode' for x in POS_CASH_balance_df_string_grouped_agg.columns]
# merge above dfs
POS_CASH_balance_df_float_and_int_grouped = pd.merge(POS_CASH_balance_df_float_grouped_agg, POS_CASH_balance_df_int_grouped_agg, left_index=True, right_index=True)
POS_CASH_balance_df_aggregated_final = pd.merge(POS_CASH_balance_df_float_and_int_grouped, POS_CASH_balance_df_string_grouped_agg, left_index=True, right_index=True)
# add in all SK_IDS, which will appear as NaNs in new df
POS_CASH_balance_df_aggregated_final = pd.merge(test_SK_ID_index_df, POS_CASH_balance_df_aggregated_final, left_index=True, right_index=True, how = 'left')
POS_CASH_balance_df_aggregated_final.to_csv(path_a +'POS_CASH_balance_df_aggregated_final.csv', columns = list(POS_CASH_balance_df_aggregated_final.columns))
del POS_CASH_balance_df
del POS_CASH_balance_df_aggregated_final
del POS_CASH_balance_df_float_list
del POS_CASH_balance_df_float_grouped
del function_list
del POS_CASH_balance_df_float_grouped_agg
del new_cols
del POS_CASH_balance_df_int_list
del POS_CASH_balance_df_int_grouped
del POS_CASH_balance_df_int_grouped_agg
del POS_CASH_balance_df_string_list
del POS_CASH_balance_df_string_grouped
del POS_CASH_balance_df_string_grouped_agg
del POS_CASH_balance_df_float_and_int_grouped
```
## previous_application_df
```
previous_application_df = pd.read_csv(path_w + 'previous_application.csv')
# replace NaNs in object entries for now is pretty safe.
f_list = list(previous_application_df.select_dtypes('object').columns)
previous_application_df[f_list] = previous_application_df[f_list].fillna(value = 'NA')
del f_list
```
### Aggregate data
```
# I have to rename this bc there is an AMT_ANNUITY and NAME_CONTRACT_STATUS in the application data set too, and need to distinguish after merging.
previous_application_df.rename(columns= {'AMT_ANNUITY':'AMT_ANNUITY_PREV_APP', 'NAME_CONTRACT_STATUS':'NAME_CONTRACT_STATUS_PREV_APP'}, inplace=True)
previous_application_df.info(verbose = True, null_counts = True)
```
All the floats are really floats, and all the integers are continous numerical except NFLAG_LAST_APPL_IN_DAY so subtract that from int,
and add to catagorical list.
```
# get float features and group them
previous_application_df_float_list = list(previous_application_df.select_dtypes('float64').columns)
previous_application_df_float_grouped = previous_application_df[previous_application_df_float_list].groupby(previous_application_df['SK_ID_CURR'])
# apply these functions and name new features accordinly
function_list = ['mean', 'median','max','min']
previous_application_df_float_grouped_agg = previous_application_df_float_grouped.agg(function_list)
new_cols = ['_'.join(col).strip() for col in previous_application_df_float_grouped_agg.columns.values]
previous_application_df_float_grouped_agg.columns = new_cols
# get int features and group them, and drop ID features
previous_application_df_int_list = list( previous_application_df.select_dtypes('int64').columns )
previous_application_df_int_list.remove('SK_ID_CURR')
previous_application_df_int_list.remove('SK_ID_PREV')
previous_application_df_int_list.remove('NFLAG_LAST_APPL_IN_DAY')
previous_application_df_int_grouped = previous_application_df[previous_application_df_int_list].groupby(previous_application_df['SK_ID_CURR'])
# apply these functions and name new features accordinly
function_list = ['mean', 'median','max','min']
previous_application_df_int_grouped_agg = previous_application_df_int_grouped.agg(function_list)
new_cols = ['_'.join(col).strip() for col in previous_application_df_int_grouped_agg.columns.values]
previous_application_df_int_grouped_agg.columns = new_cols
previous_application_df_string_list = list(previous_application_df.select_dtypes('object').columns) + ['NFLAG_LAST_APPL_IN_DAY']
previous_application_df_string_grouped \
= previous_application_df[previous_application_df_string_list].groupby(previous_application_df['SK_ID_CURR'])
# calculate the mode for these features and name accordingly
# this takes a long time to run
previous_application_df_string_grouped_agg \
= previous_application_df_string_grouped.agg(lambda x:x.value_counts().index[0])
previous_application_df_string_grouped_agg;
previous_application_df_string_grouped_agg.columns = [x + '_mode' for x in previous_application_df_string_grouped_agg.columns]
# merge all these dfs together
previous_application_df_float_and_int_grouped = \
pd.merge(previous_application_df_float_grouped_agg, previous_application_df_int_grouped_agg, left_index=True, right_index=True)
previous_application_df_aggregated_final = \
pd.merge(previous_application_df_float_and_int_grouped, previous_application_df_string_grouped_agg, left_index=True, right_index=True)
# add in all SK_IDS, which will appear as NaNs in new df
previous_application_df_aggregated_final = pd.merge(test_SK_ID_index_df, previous_application_df_aggregated_final, left_index=True, right_index=True, how = 'left')
previous_application_df_aggregated_final.to_csv(path_a + 'previous_application_df_aggregated_final.csv', columns = list(previous_application_df_aggregated_final.columns))
del previous_application_df
del previous_application_df_aggregated_final
del previous_application_df_float_list
del previous_application_df_float_grouped
del function_list
del previous_application_df_float_grouped_agg
del new_cols
del previous_application_df_int_list
del previous_application_df_int_grouped
del previous_application_df_string_list
del previous_application_df_string_grouped
del previous_application_df_string_grouped_agg
del previous_application_df_float_and_int_grouped
```
| github_jupyter |
# Tutorial on Causal Inference and its Connections to Machine Learning (Using DoWhy+EconML)
This tutorial presents a walk-through on using DoWhy+EconML libraries for causal inference. Along the way, we'll highlight the connections to machine learning---how machine learning helps in building causal effect estimators, and how causal reasoning can be help build more robust machine learning models.
Examples of data science questions that are fundamentally causal inference questions:
* **A/B experiments**: If I change the algorithm, will it lead to a higher success rate?
* **Policy decisions**: If we adopt this treatment/policy, will it lead to a healthier patient/more revenue/etc.?
* **Policy evaluation**: Knowing what I know now, did my policy help or hurt?
* **Credit attribution**: Are people buying because of the recommendation algorithm? Would they have bought anyway?
In this tutorial, you will:
* Learn how causal reasoning is necessary for decision-making, and the difference between a prediction and decision-making task.
<br>
* Get hands-on with estimating causal effects using the four steps of causal inference: **model, identify, estimate and refute**.
<br>
* See how DoWhy+EconML can help you estimate causal effects with **4 lines of code**, using the latest methods from statistics and machine learning to estimate the causal effect and evaluate its robustness to modeling assumptions.
<br>
* Work through **real-world case-studies** with Jupyter notebooks on applying causal reasoning in different scenarios including estimating impact of a customer loyalty program on future transactions, predicting which users will be positively impacted by an intervention (such as an ad), pricing products, and attributing which factors contribute most to an outcome.
<br>
* Learn about the connections between causal inference and the challenges of modern machine learning models.
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Why-causal-inference?" data-toc-modified-id="Why-causal-inference?-1"><span class="toc-item-num">1 </span>Why causal inference?</a></span><ul class="toc-item"><li><span><a href="#Defining-a-causal-effect" data-toc-modified-id="Defining-a-causal-effect-1.1"><span class="toc-item-num">1.1 </span>Defining a causal effect</a></span></li><li><span><a href="#The-difference-between-prediction-and-causal-inference" data-toc-modified-id="The-difference-between-prediction-and-causal-inference-1.2"><span class="toc-item-num">1.2 </span>The difference between prediction and causal inference</a></span></li><li><span><a href="#Two-fundamental-challenges-for-causal-inference" data-toc-modified-id="Two-fundamental-challenges-for-causal-inference-1.3"><span class="toc-item-num">1.3 </span>Two fundamental challenges for causal inference</a></span></li></ul></li><li><span><a href="#The-four-steps-of-causal-inference" data-toc-modified-id="The-four-steps-of-causal-inference-2"><span class="toc-item-num">2 </span>The four steps of causal inference</a></span><ul class="toc-item"><li><span><a href="#The-DoWhy+EconML-solution" data-toc-modified-id="The-DoWhy+EconML-solution-2.1"><span class="toc-item-num">2.1 </span>The DoWhy+EconML solution</a></span></li><li><span><a href="#A-mystery-dataset:-Can-you-find-out-if-if-there-is-a-causal-effect?" data-toc-modified-id="A-mystery-dataset:-Can-you-find-out-if-if-there-is-a-causal-effect?-2.2"><span class="toc-item-num">2.2 </span>A mystery dataset: Can you find out if if there is a causal effect?</a></span><ul class="toc-item"><li><span><a href="#Model-assumptions-about-the-data-generating-process-using-a-causal-graph" data-toc-modified-id="Model-assumptions-about-the-data-generating-process-using-a-causal-graph-2.2.1"><span class="toc-item-num">2.2.1 </span>Model assumptions about the data-generating process using a causal graph</a></span></li><li><span><a href="#Identify-the-correct-estimand-for-the-target-quantity-based-on-the-causal-model" data-toc-modified-id="Identify-the-correct-estimand-for-the-target-quantity-based-on-the-causal-model-2.2.2"><span class="toc-item-num">2.2.2 </span>Identify the correct estimand for the target quantity based on the causal model</a></span></li><li><span><a href="#Estimate-the-target-estimand" data-toc-modified-id="Estimate-the-target-estimand-2.2.3"><span class="toc-item-num">2.2.3 </span>Estimate the target estimand</a></span></li><li><span><a href="#Check-robustness-of-the-estimate-using-refutation-tests" data-toc-modified-id="Check-robustness-of-the-estimate-using-refutation-tests-2.2.4"><span class="toc-item-num">2.2.4 </span>Check robustness of the estimate using refutation tests</a></span></li></ul></li></ul></li><li><span><a href="#Case-studies-using-DoWhy+EconML" data-toc-modified-id="Case-studies-using-DoWhy+EconML-3"><span class="toc-item-num">3 </span>Case-studies using DoWhy+EconML</a></span><ul class="toc-item"><li><span><a href="#Estimating-the-impact-of-a-customer-loyalty-program" data-toc-modified-id="Estimating-the-impact-of-a-customer-loyalty-program-3.1"><span class="toc-item-num">3.1 </span>Estimating the impact of a customer loyalty program</a></span></li><li><span><a href="#Recommendation-A/B-testing-at-an-online-company" data-toc-modified-id="Recommendation-A/B-testing-at-an-online-company-3.2"><span class="toc-item-num">3.2 </span>Recommendation A/B testing at an online company</a></span></li><li><span><a href="#User-segmentation-for-targeting-interventions" data-toc-modified-id="User-segmentation-for-targeting-interventions-3.3"><span class="toc-item-num">3.3 </span>User segmentation for targeting interventions</a></span></li><li><span><a href="#Multi-investment-attribution-at-a-software-company" data-toc-modified-id="Multi-investment-attribution-at-a-software-company-3.4"><span class="toc-item-num">3.4 </span>Multi-investment attribution at a software company</a></span></li></ul></li><li><span><a href="#Connections-to-fundamental-machine-learning-challenges" data-toc-modified-id="Connections-to-fundamental-machine-learning-challenges-4"><span class="toc-item-num">4 </span>Connections to fundamental machine learning challenges</a></span></li><li><span><a href="#Further-resources" data-toc-modified-id="Further-resources-5"><span class="toc-item-num">5 </span>Further resources</a></span><ul class="toc-item"><li><span><a href="#DoWhy+EconML-libraries" data-toc-modified-id="DoWhy+EconML-libraries-5.1"><span class="toc-item-num">5.1 </span>DoWhy+EconML libraries</a></span></li><li><span><a href="#Detailed-KDD-Tutorial-on-Causal-Inference" data-toc-modified-id="Detailed-KDD-Tutorial-on-Causal-Inference-5.2"><span class="toc-item-num">5.2 </span>Detailed KDD Tutorial on Causal Inference</a></span></li><li><span><a href="#Book-chapters-on-causality-and-machine-learning" data-toc-modified-id="Book-chapters-on-causality-and-machine-learning-5.3"><span class="toc-item-num">5.3 </span>Book chapters on causality and machine learning</a></span></li><li><span><a href="#Causality-and-Machine-Learning-group-at-Microsoft" data-toc-modified-id="Causality-and-Machine-Learning-group-at-Microsoft-5.4"><span class="toc-item-num">5.4 </span>Causality and Machine Learning group at Microsoft</a></span></li></ul></li></ul></div>
## Why causal inference?
Many key data science tasks are about decision-making. Data scientists are regularly called upon to support decision-makers at all levels, helping them make the best use of data in support of achieving desired outcomes. For example, an executive making investment and resourcing decisions, a marketer determining discounting policies, a product team prioritizing which features to ship, or a doctor deciding which treatment to administer to a patient.
Each of these decision-makers is asking a what-if question. Data-driven answers to such questions require understanding the *causes* of an event and how to take action to improve future outcomes.
### Defining a causal effect
Suppose that we want to find the causal effect of taking an action A on the outcome Y. To define the causal effect, consider two worlds:
1. World 1 (Real World): Where the action A was taken and Y observed
2. World 2 (*Counterfactual* World): Where the action A was not taken (but everything else is the same)
Causal effect is the difference between Y values attained in the real world versus the counterfactual world.
$${E}[Y_{real, A=1}] - E[Y_{counterfactual, A=0}]$$

In other words, A causes Y iff changing A leads to a change in Y,
*keeping everything else constant*. Changing A while keeping everything else constant is called an **intervention**, and represented by a special notation, $do(A)$.
Formally, causal effect is the magnitude by which Y is changed by a unit *interventional* change in A:
$$E[Y│do(A=1)]−E[Y|do(A=0)]$$
To estimate the effect, the *gold standard* is to conduct a randomized experiment where a randomized subset of units is acted upon ($A=1$) and the other subset is not ($A=0$). These subsets approximate the disjoint real and counterfactual worlds and randomization ensures that there is not systematic difference between the two subsets (*"keeping everything else constant"*).
However, it is not always feasible to a run a randomized experiment. To answer causal questions, we often need to rely on observational or logged data. Such observed data is biased by correlations and unobserved confounding and thus there are systematic differences in which units were acted upon and which units were not. For example, a new marketing campaign may be deployed during the holiday season, a new feature may only have been applied to high-activity users, or the older patients may have been more likely to receive the new drug, and so on. The goal of causal inference methods is to remove such correlations and confounding from the data and estimate the *true* effect of an action, as given by the equation above.
### The difference between prediction and causal inference
<table><tr>
<td> <img src="images/supervised_ml_schematic.png" alt="Drawing" style="width: 400px;"/> </td>
<td> <img src="images/causalinference_schematic.png" alt="Drawing" style="width: 400px;"/> </td>
</tr></table>
### Two fundamental challenges for causal inference
We never observe the counterfactual world
* Cannot directly calculate the causal effect
* Must estimate the counterfactuals
* Challenges in validation
Multiple causal mechanisms can be fit to a single data distribution
* Data alone is not enough for causal inference
* Need domain knowledge and assumptions
## The four steps of causal inference
Since there is no ground-truth test dataset available that an estimate can be compared to, causal inference requires a series of principled steps to achieve a good estimator.
Let us illustrate the four steps through a sample dataset. This tutorial requires you to download two libraries: DoWhy and EconML. Both can be installed by the following command: `pip install dowhy econml`.
```
# Required libraries
import dowhy
from dowhy import CausalModel
import dowhy.datasets
# Avoiding unnecessary log messges and warnings
import logging
logging.getLogger("dowhy").setLevel(logging.WARNING)
import warnings
from sklearn.exceptions import DataConversionWarning
warnings.filterwarnings(action='ignore', category=DataConversionWarning)
# Load some sample data
data = dowhy.datasets.linear_dataset(
beta=10,
num_common_causes=5,
num_instruments=2,
num_samples=10000,
treatment_is_binary=True)
```
**I. Modeling**
The first step is to encode our domain knowledge into a causal model, often represented as a graph. The final outcome of a causal inference analysis depends largely on the input assumptions, so this step is quite important. To estimate the causal effect, most common problems involve specifying two types of variables:
1. **Confounders**: These are variables that cause both the action and the outcome. As a result, any observed correlation between the action and the outcome may simply be due to the confounder variables, and not due to any causal relationship from the action to the outcome.
2. **Instrumental Variables**: These are special variables that cause the action, but do not directly affect the outcome. In addition, they are not affected by any variable that affects the outcome. Instrumental variables can help reduce bias, if used in the correct way.
```
# I. Create a causal model from the data and domain knowledge.
model = CausalModel(
data=data["df"],
treatment=data["treatment_name"],
outcome=data["outcome_name"],
common_causes=data["common_causes_names"],
intrumental_variables=data["instrument_names"])
```
To visualize the graph, we can write,
```
model.view_model(layout="dot")
from IPython.display import Image, display
display(Image(filename="causal_model.png"))
```
In general, you can specify a causal graph that describes the mechanisms of the data-generating process for a given dataset. Each arrow in the graph denotes a causal mechanism: "A->B" implies that the variable A causes variable B.
```
# I. Create a causal model from the data and given graph.
model = CausalModel(
data=data["df"],
treatment=data["treatment_name"][0],
outcome=data["outcome_name"][0],
graph=data["gml_graph"])
model.view_model(layout="dot")
```
**II. Identification**
Both ways of providing domain knowledge (either through named variable sets of confounders and instrumental variables, or through a causal graph) correspond to an underlying causal graph. Given a causal graph and a target quantity (e.g., effect of A on B), the process of identifcation is to check whether the target quantity can be estimated given the observed variables. Importantly, identification only considers the names of variables that are available in the observed data; it does not need access to the data itself. Related to the two kinds of variables above, there are two main identification methods for causal inference.
1. **Backdoor criterion** (or more generally, adjustment sets): If all common causes of the action A and the outcome Y are observed, then the backdoor criterion implies that the causal effect can be identified by conditioning on all the common causes. This is a simplified definition (refer to Chapter 3 of the CausalML book for a formal definition).
$$ E[Y│do(A=a)] = E_W E[Y|A=a, W=w]$$
where $W$ refers to the set of common causes (confounders) of $A$ and $Y$.
2. **Instrumental variable (IV) identification**: If there is an instrumental variable available, then we can estimate effect even when any (or none) of the common causes of action and outcome are unobserved. The IV identification utilizes the fact that the instrument only affects the action directly, so the effect of the instrument on the outcome can be broken up into two sequential parts: the effect of the instrument on the action and the effect of the action on the treatment. It then relies on estimating the effect of the instrument on the action and the outcome to estimate the effect of the action on the outcome. For a binary instrument, the effect estimate is given by,
$$ E[Y│do(A=1)] -E[Y│do(A=0)] =\frac{E[Y│Z=1]- E[Y│Z=0]}{E[A│Z=1]- E[A│Z=0]} $$
```
# II. Identify causal effect and return target estimands
identified_estimand = model.identify_effect()
print(identified_estimand)
```
**III. Estimation**
As the name suggests, the estimation step involves building a statistical estimator that can compute the target estimand identified in the previous step. Many estimators have been proposed for causal inference. DoWhy implements a few of the standard estimators while EconML implements a powerful set of estimators that use machine learning.
We show an example of using Propensity Score Stratification using DoWhy, and a machine learning-based method called Double-ML using EconML.
```
# III. Estimate the target estimand using a statistical method.
propensity_strat_estimate = model.estimate_effect(identified_estimand,
method_name="backdoor.dowhy.propensity_score_stratification")
print(propensity_strat_estimate)
import econml
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LassoCV
from sklearn.ensemble import GradientBoostingRegressor
dml_estimate = model.estimate_effect(identified_estimand,
method_name="backdoor.econml.dml.DMLCateEstimator",
method_params={
'init_params': {'model_y':GradientBoostingRegressor(),
'model_t': GradientBoostingRegressor(),
'model_final':LassoCV(fit_intercept=False), },
'fit_params': {}
})
print(dml_estimate)
```
**IV. Refutation**
Finally, checking robustness of the estimate is probably the most important step of a causal analysis. We obtained an estimate using Steps 1-3, but each step may have made certain assumptions that may not be true. Absent of a proper validation "test" set, this step relies on *refutation* tests that seek to refute the correctness of an obtained estimate using properties of a good estimator. For example, a refutation test (`placebo_treatment_refuter`) checks whether the estimator returns an estimate value of 0 when the action variable is replaced by a random variable, independent of all other variables.
```
# IV. Refute the obtained estimate using multiple robustness checks.
refute_results = model.refute_estimate(identified_estimand, propensity_strat_estimate,
method_name="placebo_treatment_refuter")
print(refute_results)
```
### The DoWhy+EconML solution
We will use the DoWhy+EconML libraries for causal inference. DoWhy provides a general API for the four steps and EconML provides advanced estimators for the Estimation step.
DoWhy allows you to visualize, formalize, and test the assumptions they are making, so that you can better understand the analysis and avoid reaching incorrect conclusions. It does so by focusing on assumptions explicitly and introducing automated checks on validity of assumptions to the extent possible. As you will see, the power of DoWhy is that it provides a formal causal framework to encode domain knowledge and it can run automated robustness checks to validate the causal estimate from any estimator method.
Additionally, as data becomes high-dimensional, we need specialized methods that can handle known confounding. Here we use EconML that implements many of the state-of-the-art causal estimation approaches. This package has a common API for all the techniques, and each technique is implemented as a sequence of machine learning tasks allowing for the use of any existing machine learning software to solve these subtasks, allowing you to plug-in the ML models that you are already familiar with rather than learning a new toolkit. The power of EconML is that you can now implement the state-of-the-art in causal inference just as easily as you can run a linear regression or a random forest.
Together, DoWhy+EconML make answering what if questions a whole lot easier by providing a state-of-the-art, end-to-end framework for causal inference, including the latest causal estimation and automated robustness procedures.
### A mystery dataset: Can you find out if if there is a causal effect?
To walk-through the four steps, let us consider the **Mystery Dataset** problem. Suppose you are given some data with treatment and outcome. Can you determine whether the treatment causes the outcome, or the correlation is purely due to another common cause?
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import math
import dowhy.datasets, dowhy.plotter
```
Below we create a dataset where the true causal effect is decided by random variable. It can be either 0 or 1.
```
rvar = 1 if np.random.uniform() > 0.2 else 0
is_linear = False # A non-linear dataset. Change to True to see results for a linear dataset.
data_dict = dowhy.datasets.xy_dataset(10000, effect=rvar,
num_common_causes=2,
is_linear=is_linear,
sd_error=0.2)
df = data_dict['df']
print(df.head())
dowhy.plotter.plot_treatment_outcome(df[data_dict["treatment_name"]], df[data_dict["outcome_name"]],
df[data_dict["time_val"]])
```
#### Model assumptions about the data-generating process using a causal graph
```
model= CausalModel(
data=df,
treatment=data_dict["treatment_name"],
outcome=data_dict["outcome_name"],
common_causes=data_dict["common_causes_names"],
instruments=data_dict["instrument_names"])
model.view_model(layout="dot")
```
#### Identify the correct estimand for the target quantity based on the causal model
```
identified_estimand = model.identify_effect()
print(identified_estimand)
```
Since this is observed data, the warning asks you if there are any unobserved confounders that are missing in this dataset. If there are, then ignoring them will lead to an incorrect estimate.
If you want to disable the warning, you can use `proceed_when_unidentifiable=True` as an additional parameter to `identify_effect`.
#### Estimate the target estimand
```
estimate = model.estimate_effect(identified_estimand,
method_name="backdoor.linear_regression")
print(estimate)
print("Causal Estimate is " + str(estimate.value))
# Plot Slope of line between action and outcome = causal effect
dowhy.plotter.plot_causal_effect(estimate, df[data_dict["treatment_name"]], df[data_dict["outcome_name"]])
```
As you can see, for a non-linear data-generating process, the linear regression model is unable to distinguish the causal effect from the observed correlation.
If the DGP was linear, however, then simple linear regression would have worked. To see that, try setting `is_linear=True` in cell **10** above.
To model non-linear data (and data with high-dimensional confounders), we need more advanced methods. Below is an example using the double machine learning estimator from EconML. This estimator uses machine learning-based methods like gradient boosting trees to learn the relationship between the outcome and confounders, and the treatment and confounders, and then finally compares the residual variation between the outcome and treatment.
```
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LassoCV
from sklearn.ensemble import GradientBoostingRegressor
dml_estimate = model.estimate_effect(identified_estimand, method_name="backdoor.econml.dml.DMLCateEstimator",
control_value = 0,
treatment_value = 1,
confidence_intervals=False,
method_params={"init_params":{'model_y':GradientBoostingRegressor(),
'model_t': GradientBoostingRegressor(),
"model_final":LassoCV(fit_intercept=False),
'featurizer':PolynomialFeatures(degree=2, include_bias=True)},
"fit_params":{}})
print(dml_estimate)
```
As you can see, the DML method obtains a better estimate, that is closer to the true causal effect of 1.
#### Check robustness of the estimate using refutation tests
```
res_random=model.refute_estimate(identified_estimand, dml_estimate, method_name="random_common_cause")
print(res_random)
res_placebo=model.refute_estimate(identified_estimand, dml_estimate,
method_name="placebo_treatment_refuter", placebo_type="permute",
num_simulations=20)
print(res_placebo)
```
## Case-studies using DoWhy+EconML
In practice, as the data becomes high-dimensional, simple estimators will not estimate the correct causal effect. More advanced supervised machine learning models also do not work and often are worse than simple regression, because they include additional regularization techniques that help in minimizing predictive error, but can have unwanted effects on estimating the causal effect. Therefore, we need methods targeted to estimate the causal effect. At the same time, we also need suitable refutation methods that can check the robustness of the estimate.
Here is an example of using DoWhy+EconML for a high-dimensional dataset.
More details are in this [notebook](https://github.com/microsoft/dowhy/blob/master/docs/source/example_notebooks/dowhy-conditional-treatment-effects.ipynb).
Below we provide links to case studies that illustrate the use of DoWhy+EconML.
### Estimating the impact of a customer loyalty program
[Link to full notebook](https://github.com/microsoft/dowhy/blob/master/docs/source/example_notebooks/dowhy_example_effect_of_memberrewards_program.ipynb)
### Recommendation A/B testing at an online company
[Link to full notebook](https://github.com/microsoft/EconML/blob/master/notebooks/CustomerScenarios/Case%20Study%20-%20Recommendation%20AB%20Testing%20at%20An%20Online%20Travel%20Company%20-%20EconML%20%2B%20DoWhy.ipynb)
### User segmentation for targeting interventions
[Link to full notebook](https://github.com/microsoft/EconML/blob/master/notebooks/CustomerScenarios/Case%20Study%20-%20Customer%20Segmentation%20at%20An%20Online%20Media%20Company%20-%20EconML%20%2B%20DoWhy.ipynb)
### Multi-investment attribution at a software company
[Link to full notebook](https://github.com/microsoft/EconML/blob/master/notebooks/CustomerScenarios/Case%20Study%20-%20Multi-investment%20Attribution%20at%20A%20Software%20Company%20-%20EconML%20%2B%20DoWhy.ipynb)
## Connections to fundamental machine learning challenges
Causality is connected to many fundamental challenges in building machine learning models, including out-of-distribution generalization, fairness, explanability and privacy.

How causality can help in solving many of the challenges above is an active area of research.
## Further resources
### DoWhy+EconML libraries
DoWhy code: https://github.com/microsoft/dowhy
DoWhy notebooks: https://github.com/microsoft/dowhy/tree/master/docs/source/example_notebooks
EconML code: https://github.com/microsoft/econml
EconML notebooks: https://github.com/microsoft/EconML/tree/master/notebooks
### Detailed KDD Tutorial on Causal Inference
https://causalinference.gitlab.io/kdd-tutorial/
### Book chapters on causality and machine learning
http://causalinference.gitlab.io/
### Causality and Machine Learning group at Microsoft
https://www.microsoft.com/en-us/research/group/causal-inference/
| github_jupyter |
```
from tokenizers import Tokenizer
import torch
tokenizer = Tokenizer.from_file("tokenizer_captions.json")
VOCAB_SIZE = tokenizer.get_vocab_size()
def tokenize(texts, context_length = 256, add_start = False, add_end = False, truncate_text = False):
if isinstance(texts, str):
texts = [texts]
sot_tokens = tokenizer.encode("<|startoftext|>").ids if add_start else []
eot_tokens = tokenizer.encode("<|endoftext|>").ids if add_end else []
all_tokens = [sot_tokens + tokenizer.encode(text).ids + eot_tokens for text in texts]
result = torch.zeros(len(all_tokens), context_length, dtype=torch.long)
for i, tokens in enumerate(all_tokens):
if len(tokens) > context_length:
if truncate_text:
tokens = tokens[:context_length]
else:
raise RuntimeError(f"Input {texts[i]} is too long for context length {context_length}")
result[i, :len(tokens)] = torch.tensor(tokens)
return result
!wget --no-clobber <dropbox_url>
!wget --no-clobber https://www.dropbox.com/s/hl5hyzhyal3vfye/dalle_iconic_butterfly_149.pt
%pip install tokenizers
%pip install gpustat
!git clone https://github.com/lucidrains/DALLE-pytorch
%cd ./DALLE-pytorch/
!python3 setup.py install
!sudo apt-get -y install llvm-9-dev cmake
!git clone https://github.com/microsoft/DeepSpeed.git /tmp/Deepspeed
%cd /tmp/Deepspeed
!DS_BUILD_SPARSE_ATTN=1 ./install.sh -r
checkpoint_path = "dalle_iconic_butterfly_149.pt"
import os
import glob
text = "an armchair imitating a pikachu. an armchair in the shape of a pikachu." #@param
!python /content/DALLE-pytorch/generate.py --batch_size=32 --taming --dalle_path=$checkpoint_path --num_images=128 --text="$text"; wait;
text_cleaned = text.replace(" ", "_")
_folder = f"/content/outputs/{text_cleaned}/"
import glob
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
%matplotlib inline
text_cleaned = text.replace(" ", "_")
output_dir = f"/content/outputs/{text_cleaned}/" #@param
images = []
for img_path in glob.glob(f'{output_dir}*.jpg'):
images.append(mpimg.imread(img_path))
plt.figure(figsize=(128,128))
columns = 5
for i, image in enumerate(images):
plt.subplot(len(images) / columns + 1, columns, i + 1)
plt.imshow(image)
%pip install "git+https://github.com/openai/CLIP.git"
import clip
import torch
device = "cuda" if torch.cuda.is_available() else "cpu"
model, preprocess = clip.load("ViT-B/32", device=device)
""" Get rank by CLIP! """
image = F.interpolate(images, size=224)
text = clip.tokenize(["this colorful bird has a yellow breast , with a black crown and a black cheek patch."]).to(device)
with torch.no_grad():
image_features = model.encode_image(image)
text_features = model.encode_text(text)
logits_per_image, logits_per_text = model(image, text)
probs = logits_per_text.softmax(dim=-1).cpu().numpy()
print("Label probs:", probs)
np_images = images.cpu().numpy()
scores = probs[0]
def show_reranking(images, scores, sort=True):
img_shape = images.shape
if sort:
scores_sort = scores.argsort()
scores = scores[scores_sort[::-1]]
images = images[scores_sort[::-1]]
rows = 4
cols = img_shape[0] // 4
img_idx = 0
for col in range(cols):
fig, axs = plt.subplots(1, rows, figsize=(20,20))
plt.subplots_adjust(wspace=0.01)
for row in range(rows):
tran_img = np.transpose(images[img_idx], (1,2,0))
axs[row].imshow(tran_img, interpolation='nearest')
axs[row].set_title("{}%".format(np.around(scores[img_idx]*100, 5)))
axs[row].set_xticks([])
axs[row].set_yticks([])
img_idx += 1
show_reranking(np_images, scores)
from torchvision import transforms
txt = "this bird has wings that are brown with a white belly"
img_path = "images/Yellow_Headed_Blackbird_0013_8362.jpg"
img = Image.open(img_path)
tf = transforms.Compose([
transforms.Lambda(lambda img: img.convert("RGB") if img.mode != "RGB" else img),
transforms.RandomResizedCrop(256, scale=(0.6, 1.0), ratio=(1.0, 1.0)),
transforms.ToTensor(),
])
img = tf(img).cuda()
sot_token = vocab.encode("<|startoftext|>").ids[0]
eot_token = vocab.encode("<|endoftext|>").ids[0]
codes = [0] * dalle_dict['hparams']['text_seq_len']
text_token = vocab.encode(txt).ids
tokens = [sot_token] + text_token + [eot_token]
codes[:len(tokens)] = tokens
caption_token = torch.LongTensor(codes).cuda()
imgs = img.repeat(16,1,1,1)
caps = caption_token.repeat(16,1)
mask = (caps != 0).cuda()
images = dalle.generate_images(
caps,
mask = mask,
img = imgs,
num_init_img_tokens = (100), # you can set the size of the initial crop, defaults to a little less than ~1/2 of the tokens, as done in the paper
filter_thres = 0.9,
temperature = 1.0
)
grid = make_grid(images, nrow=4, normalize=False, range=(-1, 1)).cpu()
show(grid)
# import wandb
# run = wandb.init()
# artifact = run.use_artifact('afiaka87/dalle_train_transformer/trained-dalle:v14', type='model')
# artifact_dir = artifact.download()
```
| github_jupyter |
## Host Genome
### Extract short fragment
```
from Bio import SeqIO
sp34 = "/home/junyuchen/Lab/Phage-SOP/Result/Assembly-2020-06-14/SP34_FDSW202399941-1r/scaffolds.fasta"
cd1382 = "/home/junyuchen/Lab/Phage-SOP/Result/Assembly-2020-06-14/CD1382_FDSW202399938-1r/scaffolds.fasta"
OutDir = "/home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract"
ShortS = []
for seq in SeqIO.parse(sp34, "fasta"):
if len(seq) < 10000:
ShortS.append(seq)
#print(ShortS)
SeqIO.write(ShortS, OutDir + "/sp34.fasta", "fasta")
ShortS = []
for seq in SeqIO.parse(cd1382, "fasta"):
if len(seq) < 10000:
ShortS.append(seq)
#print(ShortS)
SeqIO.write(ShortS, OutDir + "/cd1382.fasta", "fasta")
cd1382 = "/home/junyuchen/Lab/Phage-SOP/Result/Meta-spades-test/CD1382_FDSW202399938-1r/scaffolds.fasta"
sp34 = "/home/junyuchen/Lab/Phage-SOP/Result/Meta-spades-test/SP34_FDSW202399941-1r/scaffolds.fasta"
OutDir = "/home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract"
ShortS = []
for seq in SeqIO.parse(sp34, "fasta"):
if len(seq) < 10000:
ShortS.append(seq)
#print(ShortS)
SeqIO.write(ShortS, OutDir + "/sp34-1.fasta", "fasta")
ShortS = []
for seq in SeqIO.parse(cd1382, "fasta"):
if len(seq) < 10000:
ShortS.append(seq)
#print(ShortS)
SeqIO.write(ShortS, OutDir + "/cd1382-1.fasta", "fasta")
```
### Use blastp?
#### translate seq first
```
transeq /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/sp34.fasta /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/sp34_tran.fasta -frame 6 -table=11 -sformat pearson
transeq /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/cd1382.fasta /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/cd1382_tran.fasta -frame 6 -table=11 -sformat pearson
transeq /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/sp34-1.fasta /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/sp34_tran-1.fasta -frame 6 -table=11 -sformat pearson
transeq /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/cd1382-1.fasta /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/cd1382_tran-1.fasta -frame 6 -table=11 -sformat pearson
```
```shell
diamond blastx --db /home/malab/databases_of_malab/nr/nr --query /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/sp34_tran.fasta --out /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/sp34_blastp.out --evalue 1e-05 --outfmt 6 --max-target-seqs 1 --threads 10
```
```shell
diamond blastx --db /home/malab/databases_of_malab/nr/nr --query /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/cd1382_tran.fasta --out /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/cd1382_blastp.out --evalue 1e-05 --outfmt 6 --max-target-seqs 1 --threads 10
```
```shell
diamond blastp --db /home/malab/databases_of_malab/nr/nr --query /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/sp34_tran.fasta --out /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/sp34_blastp.out --evalue 1e-05 --outfmt 6 --max-target-seqs 1 --threads 10
```
```shell
diamond blastp --db /home/malab/databases_of_malab/nr/nr --query /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/cd1382_tran.fasta --out /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/cd1382_blastp.out --evalue 1e-05 --outfmt 6 --max-target-seqs 1 --threads 10
```
```shell
diamond blastp --db /home/malab/databases_of_malab/nr/nr --query /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/sp34_tran-1.fasta --out /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/sp34_blastp-1.out --evalue 1e-05 --outfmt 6 --max-target-seqs 1 --threads 10
```
```shell
diamond blastp --db /home/malab/databases_of_malab/nr/nr --query /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/cd1382_tran.fasta --out /home/junyuchen/Lab/Phage-SOP/Result/ShortReadsExtract/cd1382_blastp.out --evalue 1e-05 --outfmt 6 --max-target-seqs 1 --threads 10
```
| github_jupyter |
# Machine Learning Engineer Nanodegree
## Reinforcement Learning
## Project: Train a Smartcab to Drive
Welcome to the fourth project of the Machine Learning Engineer Nanodegree! In this notebook, template code has already been provided for you to aid in your analysis of the *Smartcab* and your implemented learning algorithm. You will not need to modify the included code beyond what is requested. There will be questions that you must answer which relate to the project and the visualizations provided in the notebook. Each section where you will answer a question is preceded by a **'Question X'** header. Carefully read each question and provide thorough answers in the following text boxes that begin with **'Answer:'**. Your project submission will be evaluated based on your answers to each of the questions and the implementation you provide in `agent.py`.
>**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. In addition, Markdown cells can be edited by typically double-clicking the cell to enter edit mode.
-----
## Getting Started
In this project, you will work towards constructing an optimized Q-Learning driving agent that will navigate a *Smartcab* through its environment towards a goal. Since the *Smartcab* is expected to drive passengers from one location to another, the driving agent will be evaluated on two very important metrics: **Safety** and **Reliability**. A driving agent that gets the *Smartcab* to its destination while running red lights or narrowly avoiding accidents would be considered **unsafe**. Similarly, a driving agent that frequently fails to reach the destination in time would be considered **unreliable**. Maximizing the driving agent's **safety** and **reliability** would ensure that *Smartcabs* have a permanent place in the transportation industry.
**Safety** and **Reliability** are measured using a letter-grade system as follows:
| Grade | Safety | Reliability |
|:-----: |:------: |:-----------: |
| A+ | Agent commits no traffic violations,<br/>and always chooses the correct action. | Agent reaches the destination in time<br />for 100% of trips. |
| A | Agent commits few minor traffic violations,<br/>such as failing to move on a green light. | Agent reaches the destination on time<br />for at least 90% of trips. |
| B | Agent commits frequent minor traffic violations,<br/>such as failing to move on a green light. | Agent reaches the destination on time<br />for at least 80% of trips. |
| C | Agent commits at least one major traffic violation,<br/> such as driving through a red light. | Agent reaches the destination on time<br />for at least 70% of trips. |
| D | Agent causes at least one minor accident,<br/> such as turning left on green with oncoming traffic. | Agent reaches the destination on time<br />for at least 60% of trips. |
| F | Agent causes at least one major accident,<br />such as driving through a red light with cross-traffic. | Agent fails to reach the destination on time<br />for at least 60% of trips. |
To assist evaluating these important metrics, you will need to load visualization code that will be used later on in the project. Run the code cell below to import this code which is required for your analysis.
```
# Import the visualization code
import visuals as vs
# Pretty display for notebooks
%matplotlib inline
```
### Understand the World
Before starting to work on implementing your driving agent, it's necessary to first understand the world (environment) which the *Smartcab* and driving agent work in. One of the major components to building a self-learning agent is understanding the characteristics about the agent, which includes how the agent operates. To begin, simply run the `agent.py` agent code exactly how it is -- no need to make any additions whatsoever. Let the resulting simulation run for some time to see the various working components. Note that in the visual simulation (if enabled), the **white vehicle** is the *Smartcab*.
### Question 1
In a few sentences, describe what you observe during the simulation when running the default `agent.py` agent code. Some things you could consider:
- *Does the Smartcab move at all during the simulation?*
- *What kind of rewards is the driving agent receiving?*
- *How does the light changing color affect the rewards?*
**Hint:** From the `/smartcab/` top-level directory (where this notebook is located), run the command
```bash
'python smartcab/agent.py'
```
**Answer:**Smartcab does not move at all during the simulation. The reward depends on the light. If green and no oncoming traffic and smartcab is idle, driver gets negative rewards(for example: -4.20). If the light is red and smartcab is idle all the time, driver gets positive rewards(for example: 1.20). Because our smartcab does not move at all, If the light is red driver always gets positive rewards otherwise light is green driver gets negative rewards.
### Understand the Code
In addition to understanding the world, it is also necessary to understand the code itself that governs how the world, simulation, and so on operate. Attempting to create a driving agent would be difficult without having at least explored the *"hidden"* devices that make everything work. In the `/smartcab/` top-level directory, there are two folders: `/logs/` (which will be used later) and `/smartcab/`. Open the `/smartcab/` folder and explore each Python file included, then answer the following question.
### Question 2
- *In the *`agent.py`* Python file, choose three flags that can be set and explain how they change the simulation.*
- *In the *`environment.py`* Python file, what Environment class function is called when an agent performs an action?*
- *In the *`simulator.py`* Python file, what is the difference between the *`'render_text()'`* function and the *`'render()'`* function?*
- *In the *`planner.py`* Python file, will the *`'next_waypoint()`* function consider the North-South or East-West direction first?*
**Answer:**
1.Three flags I choose from the agent.py are : learning -If set to True, this flag forces the driving agent to use Q-learning; tolerance - epsilon tolerance before beginning testing, default is 0.05 ; enforce_deadline - If set to True, this flag enforces a deadline metric.
2.act() function is the first function is called when an agent performs an action.
3.render_text() prints the step resuts above the simulator and render() updates the grid and the smartcab's using pygame in the simulator.
4.next_waypoint() function consider the East-West direction first.
-----
## Implement a Basic Driving Agent
The first step to creating an optimized Q-Learning driving agent is getting the agent to actually take valid actions. In this case, a valid action is one of `None`, (do nothing) `'left'` (turn left), `right'` (turn right), or `'forward'` (go forward). For your first implementation, navigate to the `'choose_action()'` agent function and make the driving agent randomly choose one of these actions. Note that you have access to several class variables that will help you write this functionality, such as `'self.learning'` and `'self.valid_actions'`. Once implemented, run the agent file and simulation briefly to confirm that your driving agent is taking a random action each time step.
### Basic Agent Simulation Results
To obtain results from the initial simulation, you will need to adjust following flags:
- `'enforce_deadline'` - Set this to `True` to force the driving agent to capture whether it reaches the destination in time.
- `'update_delay'` - Set this to a small value (such as `0.01`) to reduce the time between steps in each trial.
- `'log_metrics'` - Set this to `True` to log the simluation results as a `.csv` file in `/logs/`.
- `'n_test'` - Set this to `'10'` to perform 10 testing trials.
Optionally, you may disable to the visual simulation (which can make the trials go faster) by setting the `'display'` flag to `False`. Flags that have been set here should be returned to their default setting when debugging. It is important that you understand what each flag does and how it affects the simulation!
Once you have successfully completed the initial simulation (there should have been 20 training trials and 10 testing trials), run the code cell below to visualize the results. Note that log files are overwritten when identical simulations are run, so be careful with what log file is being loaded!
Run the agent.py file after setting the flags from projects/smartcab folder instead of projects/smartcab/smartcab.
```
# Load the 'sim_no-learning' log file from the initial simulation results
vs.plot_trials('sim_no-learning.csv')
```
### Question 3
Using the visualization above that was produced from your initial simulation, provide an analysis and make several observations about the driving agent. Be sure that you are making at least one observation about each panel present in the visualization. Some things you could consider:
- *How frequently is the driving agent making bad decisions? How many of those bad decisions cause accidents?*
- *Given that the agent is driving randomly, does the rate of reliability make sense?*
- *What kind of rewards is the agent receiving for its actions? Do the rewards suggest it has been penalized heavily?*
- *As the number of trials increases, does the outcome of results change significantly?*
- *Would this Smartcab be considered safe and/or reliable for its passengers? Why or why not?*
**Answer:**
1. Totally driving agent makes bad decisions about 40% of times. About 5% of the decisions cause major accidents which means 12.5% of bad decision the agent makes cause major accidents. Minor acciden fluctuates around the same number as major accidents and another 12.5% of bad decision the agent makes cause minor accident. In Sum, 25% of bad decisions cause accidents.
2. Rate of reliability is around 20% which is way much lower than 60% and causes at least one major accident that is why we got reliability rate of F. Given the agent driving randomly, this rate make sense because our driver does not learn anything and does not change anything to get better result.
3. On the average agent receives rewards lower than -4 close to -5. From the above graph, we can see that about 10% of times agent makes minor or major accident.The major accidents return reward of -40 and minor accidents return reward of -5. If the rewards were heavily penalized, rolling average reward per action graph should show a lower average number so it hasn't been heavily penalized.
4. Outcome of results doesn't change significantly because our driving agent makes random choices and does not learn anything.
5. This smartcab is not safe or reliable for its passenger because it cause accidents 10% of the times and it has a rate of reliability of 20%. Looking at the reliabiliy and safety chart those results gave this car a grade of F which is the lowest grade.I think a smartcab should get an A+ to be safe for a passanger because in a real world traffic our smartcab will meet unpredicted possibities of facing accidents, so it is better for us to build our smartcab as safe and reliable as possible.
-----
## Inform the Driving Agent
The second step to creating an optimized Q-learning driving agent is defining a set of states that the agent can occupy in the environment. Depending on the input, sensory data, and additional variables available to the driving agent, a set of states can be defined for the agent so that it can eventually *learn* what action it should take when occupying a state. The condition of `'if state then action'` for each state is called a **policy**, and is ultimately what the driving agent is expected to learn. Without defining states, the driving agent would never understand which action is most optimal -- or even what environmental variables and conditions it cares about!
### Identify States
Inspecting the `'build_state()'` agent function shows that the driving agent is given the following data from the environment:
- `'waypoint'`, which is the direction the *Smartcab* should drive leading to the destination, relative to the *Smartcab*'s heading.
- `'inputs'`, which is the sensor data from the *Smartcab*. It includes
- `'light'`, the color of the light.
- `'left'`, the intended direction of travel for a vehicle to the *Smartcab*'s left. Returns `None` if no vehicle is present.
- `'right'`, the intended direction of travel for a vehicle to the *Smartcab*'s right. Returns `None` if no vehicle is present.
- `'oncoming'`, the intended direction of travel for a vehicle across the intersection from the *Smartcab*. Returns `None` if no vehicle is present.
- `'deadline'`, which is the number of actions remaining for the *Smartcab* to reach the destination before running out of time.
### Question 4
*Which features available to the agent are most relevant for learning both **safety** and **efficiency**? Why are these features appropriate for modeling the *Smartcab* in the environment? If you did not choose some features, why are those features* not *appropriate? Please note that whatever features you eventually choose for your agent's state, must be argued for here. That is: your code in agent.py should reflect the features chosen in this answer.
*
NOTE: You are not allowed to engineer new features for the smartcab.
**Answer:**
From the input features light and oncoming features are relevant for safety and waypoint features is relevant for learning efficiency. It is important for our smarcab to learn traffic rules of the grid world. light is needed for safety because it determines if our car should move or stay idle. Doing the opposite of what the light direct us will be unsafe. Oncoming is important when our smartcab decide to move direction of the oncoming traffic is important.
While all of the features I mention for safety also important for efficiency, waypoint feature is also important for efficiency because looking at those features we can calculate our agent's reliability. Reliability and safety are both relevant for our agent to learn efficiency.
I didn't choose right and left features from input features because potential cars on either side does not affect the results as oncoming does. Oncoming will determine our left turn and right turn rule and change the reward results we get from the potential accidents and traffic violations.
I eliminate deadline because it does not affect the safety and does not needed to find optimal policy.
I choose to use waypoint, light and oncoming to find optimal policy.
### Define a State Space
When defining a set of states that the agent can occupy, it is necessary to consider the *size* of the state space. That is to say, if you expect the driving agent to learn a **policy** for each state, you would need to have an optimal action for *every* state the agent can occupy. If the number of all possible states is very large, it might be the case that the driving agent never learns what to do in some states, which can lead to uninformed decisions. For example, consider a case where the following features are used to define the state of the *Smartcab*:
`('is_raining', 'is_foggy', 'is_red_light', 'turn_left', 'no_traffic', 'previous_turn_left', 'time_of_day')`.
How frequently would the agent occupy a state like `(False, True, True, True, False, False, '3AM')`? Without a near-infinite amount of time for training, it's doubtful the agent would ever learn the proper action!
### Question 5
*If a state is defined using the features you've selected from **Question 4**, what would be the size of the state space? Given what you know about the environment and how it is simulated, do you think the driving agent could learn a policy for each possible state within a reasonable number of training trials?*
**Hint:** Consider the *combinations* of features to calculate the total number of states!
**Answer:**
There are;
2 possible values for light: green and red,
4 possible values for oncoming: None, left, right, forward,
3 possible values for waypoint: left, right, forward,
2*4*3 = 24
This number of state is reasonable for our driving agent to learn within a reasonable number of trials. At least 24 trials are enough to try all the possible state in each trial.
### Update the Driving Agent State
For your second implementation, navigate to the `'build_state()'` agent function. With the justification you've provided in **Question 4**, you will now set the `'state'` variable to a tuple of all the features necessary for Q-Learning. Confirm your driving agent is updating its state by running the agent file and simulation briefly and note whether the state is displaying. If the visual simulation is used, confirm that the updated state corresponds with what is seen in the simulation.
**Note:** Remember to reset simulation flags to their default setting when making this observation!
-----
## Implement a Q-Learning Driving Agent
The third step to creating an optimized Q-Learning agent is to begin implementing the functionality of Q-Learning itself. The concept of Q-Learning is fairly straightforward: For every state the agent visits, create an entry in the Q-table for all state-action pairs available. Then, when the agent encounters a state and performs an action, update the Q-value associated with that state-action pair based on the reward received and the iterative update rule implemented. Of course, additional benefits come from Q-Learning, such that we can have the agent choose the *best* action for each state based on the Q-values of each state-action pair possible. For this project, you will be implementing a *decaying,* $\epsilon$*-greedy* Q-learning algorithm with *no* discount factor. Follow the implementation instructions under each **TODO** in the agent functions.
Note that the agent attribute `self.Q` is a dictionary: This is how the Q-table will be formed. Each state will be a key of the `self.Q` dictionary, and each value will then be another dictionary that holds the *action* and *Q-value*. Here is an example:
```
{ 'state-1': {
'action-1' : Qvalue-1,
'action-2' : Qvalue-2,
...
},
'state-2': {
'action-1' : Qvalue-1,
...
},
...
}
```
Furthermore, note that you are expected to use a *decaying* $\epsilon$ *(exploration) factor*. Hence, as the number of trials increases, $\epsilon$ should decrease towards 0. This is because the agent is expected to learn from its behavior and begin acting on its learned behavior. Additionally, The agent will be tested on what it has learned after $\epsilon$ has passed a certain threshold (the default threshold is 0.05). For the initial Q-Learning implementation, you will be implementing a linear decaying function for $\epsilon$.
### Q-Learning Simulation Results
To obtain results from the initial Q-Learning implementation, you will need to adjust the following flags and setup:
- `'enforce_deadline'` - Set this to `True` to force the driving agent to capture whether it reaches the destination in time.
- `'update_delay'` - Set this to a small value (such as `0.01`) to reduce the time between steps in each trial.
- `'log_metrics'` - Set this to `True` to log the simluation results as a `.csv` file and the Q-table as a `.txt` file in `/logs/`.
- `'n_test'` - Set this to `'10'` to perform 10 testing trials.
- `'learning'` - Set this to `'True'` to tell the driving agent to use your Q-Learning implementation.
In addition, use the following decay function for $\epsilon$:
$$ \epsilon_{t+1} = \epsilon_{t} - 0.05, \hspace{10px}\textrm{for trial number } t$$
If you have difficulty getting your implementation to work, try setting the `'verbose'` flag to `True` to help debug. Flags that have been set here should be returned to their default setting when debugging. It is important that you understand what each flag does and how it affects the simulation!
Once you have successfully completed the initial Q-Learning simulation, run the code cell below to visualize the results. Note that log files are overwritten when identical simulations are run, so be careful with what log file is being loaded!
```
# Load the 'sim_default-learning' file from the default Q-Learning simulation
vs.plot_trials('sim_default-learning.csv')
```
### Question 6
Using the visualization above that was produced from your default Q-Learning simulation, provide an analysis and make observations about the driving agent like in **Question 3**. Note that the simulation should have also produced the Q-table in a text file which can help you make observations about the agent's learning. Some additional things you could consider:
- *Are there any observations that are similar between the basic driving agent and the default Q-Learning agent?*
- *Approximately how many training trials did the driving agent require before testing? Does that number make sense given the epsilon-tolerance?*
- *Is the decaying function you implemented for $\epsilon$ (the exploration factor) accurately represented in the parameters panel?*
- *As the number of training trials increased, did the number of bad actions decrease? Did the average reward increase?*
- *How does the safety and reliability rating compare to the initial driving agent?*
**Answer:**
Comparing to the first graph with the graph from the question 3:
1. Total bad action rate decreased to 12.5% from about 40%
2. 5% of total major accident dropped to 4% as the minor accidents. This change is not significant so we can consider this is similar result for those two graphs.
3. After 20 trial our smartcab reach 80% reliability rate which result an A+ for the reliability. This number make sense because our smartcab learning and its reliability increased from initial models rate 20% to 80%
4. Our used epsilon function which decrease by 0.05 from 1 to 0 is accurately displayed in the exploration factor - learning factor graph by a straight line going from 1 to 0.
5. As the number of trials increase, number of bad decisions decrease from about 35% to 12.5%. Average reward increase as the trial number increase.
6. Safety rating is the same as the initial diriving agent but the reliability rate is increased significantly looking at the letter grades. Safety stayed getting grade of F because total major accident and minor accident rates are still too close to 5%. Reliability get A+ because it increased from 20% to 80%.
-----
## Improve the Q-Learning Driving Agent
The third step to creating an optimized Q-Learning agent is to perform the optimization! Now that the Q-Learning algorithm is implemented and the driving agent is successfully learning, it's necessary to tune settings and adjust learning paramaters so the driving agent learns both **safety** and **efficiency**. Typically this step will require a lot of trial and error, as some settings will invariably make the learning worse. One thing to keep in mind is the act of learning itself and the time that this takes: In theory, we could allow the agent to learn for an incredibly long amount of time; however, another goal of Q-Learning is to *transition from experimenting with unlearned behavior to acting on learned behavior*. For example, always allowing the agent to perform a random action during training (if $\epsilon = 1$ and never decays) will certainly make it *learn*, but never let it *act*. When improving on your Q-Learning implementation, consider the implications it creates and whether it is logistically sensible to make a particular adjustment.
### Improved Q-Learning Simulation Results
To obtain results from the initial Q-Learning implementation, you will need to adjust the following flags and setup:
- `'enforce_deadline'` - Set this to `True` to force the driving agent to capture whether it reaches the destination in time.
- `'update_delay'` - Set this to a small value (such as `0.01`) to reduce the time between steps in each trial.
- `'log_metrics'` - Set this to `True` to log the simluation results as a `.csv` file and the Q-table as a `.txt` file in `/logs/`.
- `'learning'` - Set this to `'True'` to tell the driving agent to use your Q-Learning implementation.
- `'optimized'` - Set this to `'True'` to tell the driving agent you are performing an optimized version of the Q-Learning implementation.
Additional flags that can be adjusted as part of optimizing the Q-Learning agent:
- `'n_test'` - Set this to some positive number (previously 10) to perform that many testing trials.
- `'alpha'` - Set this to a real number between 0 - 1 to adjust the learning rate of the Q-Learning algorithm.
- `'epsilon'` - Set this to a real number between 0 - 1 to adjust the starting exploration factor of the Q-Learning algorithm.
- `'tolerance'` - set this to some small value larger than 0 (default was 0.05) to set the epsilon threshold for testing.
Furthermore, use a decaying function of your choice for $\epsilon$ (the exploration factor). Note that whichever function you use, it **must decay to **`'tolerance'`** at a reasonable rate**. The Q-Learning agent will not begin testing until this occurs. Some example decaying functions (for $t$, the number of trials):
$$ \epsilon = a^t, \textrm{for } 0 < a < 1 \hspace{50px}\epsilon = \frac{1}{t^2}\hspace{50px}\epsilon = e^{-at}, \textrm{for } 0 < a < 1 \hspace{50px} \epsilon = \cos(at), \textrm{for } 0 < a < 1$$
You may also use a decaying function for $\alpha$ (the learning rate) if you so choose, however this is typically less common. If you do so, be sure that it adheres to the inequality $0 \leq \alpha \leq 1$.
If you have difficulty getting your implementation to work, try setting the `'verbose'` flag to `True` to help debug. Flags that have been set here should be returned to their default setting when debugging. It is important that you understand what each flag does and how it affects the simulation!
Once you have successfully completed the improved Q-Learning simulation, run the code cell below to visualize the results. Note that log files are overwritten when identical simulations are run, so be careful with what log file is being loaded!
```
# Load the 'sim_improved-learning' file from the improved Q-Learning simulation
vs.plot_trials('sim_improved-learning.csv')
```
### Question 7
Using the visualization above that was produced from your improved Q-Learning simulation, provide a final analysis and make observations about the improved driving agent like in **Question 6**. Questions you should answer:
- *What decaying function was used for epsilon (the exploration factor)?*
- *Approximately how many training trials were needed for your agent before begining testing?*
- *What epsilon-tolerance and alpha (learning rate) did you use? Why did you use them?*
- *How much improvement was made with this Q-Learner when compared to the default Q-Learner from the previous section?*
- *Would you say that the Q-Learner results show that your driving agent successfully learned an appropriate policy?*
- *Are you satisfied with the safety and reliability ratings of the *Smartcab*?*
**Answer:**
1. I used decay function for epsilon epsilon = 0.92t .
2. After 40th trial exploration curve converges to 0. After 40 trial we can begin to test so we need 40 trial before begining our test.
3. I used epsilon-tolerance as 0.02 and alpha as 0.7. with that high value of alpha my model did not learn anything and did not even run completely.Then I decided to keep alpha as close as default value and set it to 0.3. I used those values to ensure my model to learn as good as it can. I kept epsilon-tolerance 0.02 to give enough time to learner to benefit the learning factor and alpha 0.3 learner not to be biased.
4. This Q-learner results are far better than the previous results.Major and minor accidents are close to 0% which means 100% of the times our smartcab drive safely whereas the previous section this value was aproximately 87.5%. 12,5% improvement in significant for this model. reliability rate is increased by 20% from the previos section.
5. I would say yes. After 20th trial our smartcab does not get negative reward.Also, both of reliability and safety ratings increase significantly to a satisfactory number.
6.Yes I am satisfied because the smartcab gets A+ for safety and A for reliability. Reliability being A not A+ does not bother me that much because this is a minor issue.
### Define an Optimal Policy
Sometimes, the answer to the important question *"what am I trying to get my agent to learn?"* only has a theoretical answer and cannot be concretely described. Here, however, you can concretely define what it is the agent is trying to learn, and that is the U.S. right-of-way traffic laws. Since these laws are known information, you can further define, for each state the *Smartcab* is occupying, the optimal action for the driving agent based on these laws. In that case, we call the set of optimal state-action pairs an **optimal policy**. Hence, unlike some theoretical answers, it is clear whether the agent is acting "incorrectly" not only by the reward (penalty) it receives, but also by pure observation. If the agent drives through a red light, we both see it receive a negative reward but also know that it is not the correct behavior. This can be used to your advantage for verifying whether the **policy** your driving agent has learned is the correct one, or if it is a **suboptimal policy**.
### Question 8
1. Please summarize what the optimal policy is for the smartcab in the given environment. What would be the best set of instructions possible given what we know about the environment?
_You can explain with words or a table, but you should thoroughly discuss the optimal policy._
2. Next, investigate the `'sim_improved-learning.txt'` text file to see the results of your improved Q-Learning algorithm. _For each state that has been recorded from the simulation, is the **policy** (the action with the highest value) correct for the given state? Are there any states where the policy is different than what would be expected from an optimal policy?_
3. Provide a few examples from your recorded Q-table which demonstrate that your smartcab learned the optimal policy. Explain why these entries demonstrate the optimal policy.
4. Try to find at least one entry where the smartcab did _not_ learn the optimal policy. Discuss why your cab may have not learned the correct policy for the given state.
Be sure to document your `state` dictionary below, it should be easy for the reader to understand what each state represents.
**Answer:**
My state space is formed by (waypoint, light, oncoming) which is a resonable number of states for our algorithm to learn in a reasonable time.The optimal policy for the smartcab can be summarized as follows:
* Red light -- Smartcab should stay idle. Otherwise, it gets a negative reward. Right turn is allowed if there is no car coming from left. Since we can not detect cars coming from left, our smartcab should stay idle on red all the time.
* Green light -- Smartcab can move left, right or forward.
* Oncoming -- If there is an oncoming traffic to right or forward, turning left is not allowed.
The policy displays the following states:
('forward', 'red', 'forward')
-- forward : 0.00
-- right : 0.31
-- None : 1.56
-- left : 0.00
This complies with the red light policy I explain in the summary above. If our smartcab stay idle it will get highest positive reward and small positive reward if it turns right.
('forward', 'green', 'left')
-- forward : 1.93
-- right : 0.63
-- None : 0.00
-- left : 0.52
This complies with the green light rule. Only staying idle result a negative reward for our smartcab.
('left', 'red', None)
-- forward : -16.36
-- right : 0.32
-- None : 2.06
-- left : -18.00
This complies with the red light policy I explain in the summary above. Our car is allowed to turn right and stay idle in red. Therefore, None has the highest positive value and right has a positive value. At this state our car should stay idle.
('right', 'red', None)
-- forward : -2.86
-- right : -4.73
-- None : 0.49
-- left : -4.98
At this state optimal policy is to stay idle and this state give the highest positive reward for staying idle. Turning right is allowed for our summary of optimal policy.
('left', 'green', 'right')
-- forward : 0.00
-- right : 0.92
-- None : -2.37
-- left : 0.00
Given the features I choose, the smartcab did not learn the optimal policy in this state becasue Oncoming car wants to turn right and our smartcab gets 0.00 to left which will cause a collision and a big traffic violation. This state action pair was never explored by the agent,therefore it is our suboptimal policy. If that state occurs in real life we expect forward to get positive reward and left get a negative reward. Turning right should be positive reward and None should be negative , sure , but those two are the only actions our agent discovered and no idea about other actions.
-----
### Optional: Future Rewards - Discount Factor, `'gamma'`
Curiously, as part of the Q-Learning algorithm, you were asked to **not** use the discount factor, `'gamma'` in the implementation. Including future rewards in the algorithm is used to aid in propagating positive rewards backwards from a future state to the current state. Essentially, if the driving agent is given the option to make several actions to arrive at different states, including future rewards will bias the agent towards states that could provide even more rewards. An example of this would be the driving agent moving towards a goal: With all actions and rewards equal, moving towards the goal would theoretically yield better rewards if there is an additional reward for reaching the goal. However, even though in this project, the driving agent is trying to reach a destination in the allotted time, including future rewards will not benefit the agent. In fact, if the agent were given many trials to learn, it could negatively affect Q-values!
### Optional Question 9
*There are two characteristics about the project that invalidate the use of future rewards in the Q-Learning algorithm. One characteristic has to do with the *Smartcab* itself, and the other has to do with the environment. Can you figure out what they are and why future rewards won't work for this project?*
**Answer:** Even if we are given deadline metric, our agent only knows the next waypoint. Deadline changes for every intersectionand is not linked to specific intersection. It does not make sense to talk about the distance between the destination and any specific states,Therefore we can not propagate rewards towards states which are "close to the destination". Because of that feature rewards won't work for this project. Before each trial our values of the environment keeps resetting and prior rewards and results does not mean anything for the current trial which makes it impossible to evaluate future rewards.
Sestination does not stay at the same intersection over the trials, so if we want to propagate reward away from the destination, after a while, we would not know which way to go.
> **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to
**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
| github_jupyter |
## MNIST CNN (Keras + Tensorflow)
### Get started
Assuming you have Keras > 2.2 and Tensorflow > 1.10, you will need the following libraries for comparison
- Install **DeepExplain**
https://github.com/marcoancona/DeepExplain
- Install **DeepLIFT**
https://github.com/kundajelab/deeplift
- Install **SHAP**
https://github.com/slundberg/shap
```
%load_ext autoreload
%autoreload 2
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import tempfile, sys, os, pickle
sys.path.insert(0, os.path.abspath('../..'))
#os.environ["CUDA_VISIBLE_DEVICES"]="-1"
import keras
from keras.datasets import mnist
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential, Model, load_model
from keras.layers import Dense, Dropout, Flatten, Activation, Input, Conv2D, MaxPooling2D
from keras import regularizers
from keras import backend as K
import numpy as np
import tensorflow as tf
import scipy
print ("Using TF:", tf.__version__)
print ("Using Keras:", keras.__version__)
# Import DASP
from dasp import DASP
#Import DeepLift
# Installation instructions: https://github.com/kundajelab/deeplift
import deeplift
from deeplift.layers import NonlinearMxtsMode
from deeplift.conversion import kerasapi_conversion as kc
from deeplift.util import compile_func
# Import Deep Explain (for Grad * Input, Integrated Gradients and Occlusion implementations)
# Installation instructions: https://github.com/marcoancona/DeepExplain
from deepexplain.tensorflow import DeepExplain
# Build and train a network.
SKIP_TRAIN = False # after first run, this can be set to True
saved_model_file = '.model_cnn.h5'
saved_model_weights_file = '.model_weights.h5'
batch_size = 128
num_classes = 10
epochs = 6
# input image dimensions
img_rows, img_cols = 28, 28
# the data, shuffled and split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train = x_train.reshape(-1, img_rows,img_cols,1)
x_test = x_test.reshape(-1, img_rows,img_cols,1)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
x_train = (x_train - 0.5) * 2
x_test = (x_test - 0.5) * 2
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# LeNet-5
model = Sequential()
model.add(Conv2D(6, kernel_size=(5, 5),activation='relu',input_shape=x_train.shape[1:], name='conv_1'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(16, (5, 5), activation='relu', name='conv_2'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(120, activation='relu', name='dense_1'))
model.add(Dense(84, activation='relu', name='dense_2'))
model.add(Dense(num_classes, name='dense_3'))
model.add(Activation('softmax'))
# ^ IMPORTANT: notice that the final softmax must be in its own layer
# if we want to target pre-softmax units
if SKIP_TRAIN is True:
model = load_model(saved_model_file)
else:
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
model.save(saved_model_file)
model.save_weights(saved_model_weights_file)
model.summary()
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
```
### Define a subset of the test set to generate explanations for
```
xs = x_test[0:10]
ys = y_test[0:10]
a_intgrad = np.zeros_like(xs)
a_res = np.zeros_like(xs)
a_rc = np.zeros_like(xs)
a_occlusion = np.zeros_like(xs)
a_dasp = np.zeros_like(xs)
a_sampling = np.zeros_like(xs)
```
### Use Deep Shapley propagation to compute approximate Shapley Values.
Notice that this requires to convert our original model into a propababilistic one. We provide probabilistic layers for this.
Also, this will require O(c*n) evaluations of the probabilistic network, where n is the number of input features and c is the number of coalition sizes to be tested (ideally c = n)
```
# Clone the model to remove last softmax activation
reduced_model = Model(model.inputs, model.layers[-2].output)
# Init DASP
dasp = DASP(reduced_model)
# Get model description (optional)
dasp.model_summary()
# Run DASP with 16 coalition sizes (enough for convergence)
dasp_results = dasp.run(xs, 16)
# Select only the explanation for the target unit corresponding to the correct class
a_dasp = np.array([dasp_results[i, c, :] for i, c in enumerate(np.argmax(ys, 1))])
```
### Use DeepExplain framework to compute attributions using Gradient * Input, Integrated Gradients and Occlusion
Occlusion is performed by replacing one pixel at the time with a zero value and measuring the difference in the target output caused by such occlusion.
```
%%time
with DeepExplain(session=K.get_session()) as de: # <-- init DeepExplain context
# Need to reconstruct the graph in DeepExplain context, using the same weights.
# With Keras this is very easy:
# 1. Get the input tensor to the original model
input_tensor = model.layers[0].input
# 2. We now target the output of the last dense layer (pre-softmax)
# To do so, create a new model sharing the same layers untill the last dense (index -2)
fModel = Model(inputs=input_tensor, outputs = model.layers[-2].output)
target_tensor = fModel(input_tensor)
a_gradin = de.explain('grad*input', target_tensor * ys, input_tensor, xs)
a_intgrad = de.explain('intgrad', target_tensor * ys, input_tensor, xs)
a_occlusion = de.explain('occlusion', target_tensor * ys, input_tensor, xs)
print ("Done")
```
### Use DeepLIFT framework to compute attributions according to both Rescale and RevealCancel methods
```
%%time
# Compute DeepLift attributions
revealcancel_model = kc.convert_model_from_saved_files(
h5_file=saved_model_file,
nonlinear_mxts_mode=NonlinearMxtsMode.RevealCancel)
rescale_model = kc.convert_model_from_saved_files(
h5_file=saved_model_file,
nonlinear_mxts_mode=NonlinearMxtsMode.Rescale)
revealcancel_func = revealcancel_model.get_target_contribs_func(find_scores_layer_idx=0, target_layer_idx=-2)
rescale_func = rescale_model.get_target_contribs_func(find_scores_layer_idx=0, target_layer_idx=-2)
a_rc = np.array([np.array(revealcancel_func(
task_idx=np.argmax(y),
input_data_list=[[x]],
input_references_list=[[np.zeros_like(x)]],
batch_size=100,
progress_update=None)) for x, y in zip(xs,ys)])
a_res = np.array([np.array(rescale_func(
task_idx=np.argmax(y),
input_data_list=[[x]],
input_references_list=[[np.zeros_like(x)]],
batch_size=100,
progress_update=None)) for x, y in zip(xs,ys)])
```
### Use Shapley sampling to approximate the Ground Truth
Notice that this might take a while. Sampling method as described in https://www.sciencedirect.com/science/article/pii/S0305054808000804
```
from utils.shapley_sampling import run_shapley_sampling
a_sampling = run_shapley_sampling(fModel, xs, ys, runs=2**10, feat_dims=[1,2])
print ("Done")
```
## Plot attribution maps for qualitative comparison
```
attributions = [
('Integrated\nGradients', a_intgrad.reshape(xs.shape)),
('DeepLIFT\n(Rescale)', a_res.reshape(xs.shape)),
('DeepLIFT\n(RevCancel)', a_rc.reshape(xs.shape)),
('Occlusion', a_occlusion.reshape(xs.shape)),
('DASP (ours)', a_dasp.reshape(xs.shape)),
('Sampling\n(Ground Truth)', a_sampling.reshape(xs.shape))
]
# Plot attributions
%matplotlib inline
from utils.utils import plot_attribution_maps
# Plot all
plot_attribution_maps("mnist_cnn", xs, [x[1] for x in attributions], [x[0] for x in attributions], idxs=range(8))
```
## Quantitative comparison (MSE and Spearman rank correlation)
```
from utils.utils import plot_mse_comparison, plot_correlation_comparison
plot_mse_comparison("mnist_cnn", [x[1] for x in attributions], [x[0] for x in attributions], gt_idx=-1)
plot_correlation_comparison("mnist_cnn", [x[1] for x in attributions], [x[0] for x in attributions], gt_idx=-1)
```
## Quantitative comparison (pixel perturbation)
Pixels in the original images are ranked by their attribution value and sequentially "removed" by setting their value to zero. We expect good explanations to highlight pixels that are important for the classification. Assuming that the importance of a pixels is positively correlated with its marginal contribution to the output, we expect the target output to drop faster when the most important pixels are removed first. These curve showa the output variations while pixels are removed according to the ranking given by the different methods. Better methods produce the largest variations. See "Evaluating the visualization of what a Deep Neural Network has learned", Samek et al., 2015 for details.
```
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
from utils.accuracy_robustness import run_robustness_test
init_notebook_mode(connected=True)
run_robustness_test(fModel, xs, ys, [x[1] for x in attributions], [x[0] for x in attributions], 'mnist_cnn', 1,
result_path='.', mode='prediction', reduce_dim=None)
```
| github_jupyter |
```
#Make sure dependencies are installed
import cobra
import BOFdat as bd
import pandas as pd
from Bio import SeqIO
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sb
from cobra.util.solver import linear_reaction_coefficients
sb.set_style('whitegrid')
#Write another proteomic file that has protein id instead of locus tags
ecoli_gb = SeqIO.read('Ecoli_K12_MG1655.gbff','genbank')
proteomic_orig = pd.read_csv('proteomic.csv')
ecoli_dict ={}
for i in ecoli_gb.features:
if i.type == 'CDS':
if 'locus_tag' in i.qualifiers and 'protein_id' in i.qualifiers:
ecoli_dict[i.qualifiers['locus_tag'][0]] = i.qualifiers['protein_id'][0]
prot_id = []
for i in proteomic_orig.gene_ID:
prot_id.append(ecoli_dict.get(i))
new_proteomic = pd.DataFrame({'gene_id':prot_id,'abundance':[i for i in proteomic_orig.Mean]},columns=['gene_id','abundance'])
new_proteomic.to_csv('proteomic.csv')
#Give the path to each file as function parameters
#Genome file in BioPython supported format (.faa, .fna) and GenBank file
#also in BioPython supported format (.gb, .gbff)
genome = 'Ecoli_DNA.fna'
genbank = 'Ecoli_K12_MG1655.gbff'
#OMICs data as a 2 column csv file, gene and abundance
transcriptomic = 'transcriptomic.csv'
proteomic = 'proteomic.csv'
#Lipidomic abundances and conversion to model identifier
lipidomic_abundances = 'lipidomic_abundances.csv'
lipidomic_conversion = 'lipidomic_conversion.csv'
#Metabolomic abundances and conversion to model identifier
metabolomic_abundances = 'metabolomic_abundances.csv'
metabolite_conversion = 'metabolites_to_add.csv'
#Growth data on different carbon sources, uptake and secretion rates
maintenance = 'maintenance.csv'
#The model for which the coefficients are generated
model = 'iML1515.json'
ecoli = cobra.io.load_json_model(model)
```
# Figure S2
```
#calculate and plot fold change
import math
old_biomass = {}
for m in ecoli.reactions.Ec_biomass_iML1515_WT_75p37M.reactants:
old_biomass[m.id] = abs(ecoli.reactions.Ec_biomass_iML1515_WT_75p37M.get_coefficient(m))
data_df = pd.DataFrame(index=np.linspace(0.01,0.9,num=100),columns=['dna','rna','protein','lipids','metabolites'])
for ratio in np.linspace(0.01,0.9,num=100):
dna_coeff = bd.dna.generate_coefficients(genome,model,DNA_WEIGHT_FRACTION=ratio)
rna_coeff = bd.rna.generate_coefficients(genbank,model,transcriptomic,RNA_WEIGHT_FRACTION=ratio)
prot_coeff = bd.protein.generate_coefficients(genbank,model,proteomic,PROTEIN_WEIGHT_FRACTION=ratio)
lipid_coeff = bd.lipid.generate_coefficients(lipidomic_abundances,lipidomic_conversion,model,LIPID_WEIGHT_FRACTION=ratio)
metab_coeff = bd.metabolite.generate_coefficients(metabolomic_abundances,metabolite_conversion,model,METAB_WEIGHT_FRACTION=ratio)
new_biomass = [dna_coeff, rna_coeff, prot_coeff, lipid_coeff,metab_coeff]
#Generate a new biomass objective function by adding all the components together
mean_error = []
for i in range(len(new_biomass)):
if i == 4:
#Dealing with metab error
number_of_matches = []
for r,orig_coeff in old_biomass.iteritems():
for m,new_coeff in new_biomass[i].iteritems():
if r[:-2] == m.id[:-2]:
#Get order of magnitude of original coeff
#Verify the order of magnitude
if abs(int(math.log10(new_coeff)))-abs(int(math.log10(orig_coeff))) == 0:
number_of_matches.append(m.id)
mean_error.append(len(number_of_matches))
else:
d = new_biomass[i]
df1 = pd.DataFrame({'new_coefficient':d.values(),'metab':[m.id for m in d.keys()]})
df2 = pd.DataFrame({'original_coefficient':old_biomass.values(),'metab': old_biomass.keys()})
df3 = pd.merge(left=df1,right=df2,how='outer',on='metab')
#Remove data that is absurdely divergent
cut_off = (df3.new_coefficient.median() + df3.original_coefficient.median())/2 + (df3.new_coefficient.std()+df3.original_coefficient.std())
#Get all data below cut-off
df3['difference'] = abs(df3.new_coefficient[df3['new_coefficient']<cut_off]) - abs(df3.original_coefficient[df3['original_coefficient']<cut_off])
df3['percent_error'] = abs(df3['difference']/df3.original_coefficient[df3['original_coefficient']<cut_off])*100
data = df3[df3['percent_error'].isnull() == False].percent_error
mean_error.append(data.mean())
data_df.loc[ratio] = mean_error
data_df
data_df.to_csv('figS2_data.csv')
ploting_df = data_df[data_df.columns[:-1]]
#Plot
color_palette = sb.color_palette('dark',10)
colors = [color_palette[int(i)] for i in np.linspace(0,9, num=9)]
#Change column names
new_cols_dict = {'dna':'DNA','rna':'RNA','protein':'Protein','lipids':'Lipids'}
ploting_df.rename(columns=new_cols_dict,inplace=True)
ax = ploting_df.plot(color=colors)
# zoom-in / limit the view to different portions of the data
ax.set_ylim(0., 100.) # outliers only
plt.ylabel('BOFsc percent difference')
plt.xlabel('Macromolecular category weight fraction')
plt.savefig('/home/jean-christophe/Documents/Maitrise_UCSD/biomass/Paper_figures/supp2a.svg')
plt.show()
DNA_RATIO = 0.031
RNA_RATIO = 0.205
PROTEIN_RATIO = 0.55
LIPID_RATIO = 0.091
METABOLITE_RATIO = 0.1
iML_ratios = [DNA_RATIO,RNA_RATIO,PROTEIN_RATIO,LIPID_RATIO]
sum(iML_ratios)
import matplotlib.
min_error, weight_fraction_min = [],[]
for col in ploting_df.columns:
min_error.append(ploting_df[col].min())
weight_fraction_min.append(ploting_df[col].idxmin())
result_df = pd.DataFrame({'Minimal percent difference':weight_fraction_min,'iML1515':[i for i in iML_ratios]},index=['DNA','RNA','Protein','Lipid'])
color_palette = sb.color_palette('Blues',4)
result_df.plot(kind='bar',color=[color_palette[1],color_palette[3]],fontsize=16,)
plt.xlabel('Macromolecular category',fontsize=18)
plt.ylabel('Weight fraction',fontsize=18)
plt.savefig('/home/jean-christophe/Documents/Maitrise_UCSD/biomass/Paper_figures/supp2b.svg')
plt.show()
result_df.plot.bar?
lipid_coeff = bd.lipid.generate_coefficients(lipidomic_abundances,lipidomic_conversion,model)
lipid_coeff
import cobra
import BOFdat
import pandas as pd
from cobra.util.solver import linear_reaction_coefficients
ecoli = cobra.io.load_json_model('iML1515.json')
[m.id for m in ecoli.reactions.Ec_biomass_iML1515_WT_75p37M.reactants if m.id.startswith('pg') or m.id.startswith('pe')]
biomass = ecoli.reactions.
for m in biomass.reactants:
if m.id.startswith('pe') or m.id.startswith('pg'):
print(m.id)
[i.formula_weight for i in lipid_coeff.keys()]
import BOFdat
lipid_df = BOFdat.lipid.filter_for_model_metab('lipidomic_conversion.csv','iML1515.json')
lipid_df
from cobra.util.solver import linear_reaction_coefficients
biomass = linear_reaction_coefficients(ecoli).keys()[0]
for m in biomass.reactants:
print(m.id)
```
# Figure S3
# Figure S3
| github_jupyter |
<a href="https://colab.research.google.com/github/3778/COVID-19/blob/master/notebooks/%5Bissue_62%5D_Covid_Auto_Arima.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
!git clone https://github.com/3778/COVID-19.git
!pip install pmdarima
import os
os.chdir('COVID-19')
import pandas as pd
from pmdarima import auto_arima
from data.collectors import load_dump_covid_19_data, load_jh_data
load_dump_covid_19_data()
jhu = load_jh_data()
```
# Simular como teria sido acompanhar o histórico que já temos fazendo previsões para os próximos `forecast_n_days`
(ex: Simulação do que teria acontecido se eu tivesse tentado prever de 7 em 7 dias os próximos 7 dias)
```
def simulate(data, forecast_n_days=7, n_history=None):
lag = pd.DateOffset(days=forecast_n_days)
one_day = pd.DateOffset(days=1)
dmin, dmax = data.index.min(), data.index.max()
date_range = (
pd
.date_range(
start=dmin+lag,
end=dmax-lag,
freq=f'{forecast_n_days}D'
)
)
look_back = pd.DateOffset(
days=(dmax-dmin).days if n_history is None else n_history
)
forecasts = []
for date in date_range:
train = data.loc[date-look_back:date]
test = data.loc[date+one_day:date+lag]
model = (
auto_arima(
train,
start_p=1,
d=None,
start_q=1,
max_p=15,
max_d=15,
max_q=15,
seasonal=False,
suppress_warnings=True,
maxiter=500,
stepwise=False
)
)
forecast = model.predict(test.shape[0])
forecasts.append(
pd.Series(forecast, index=test.index)
)
return (
pd
.concat(
[
data,
(
pd
.concat(forecasts)
.rename('forecast')
)
],
axis=1
)
)
```
# Brasil
```
data = (
pd
.read_csv(
'data/csv/covid_19/by_uf/by_uf.csv'
)
.query('date >= "2020-03-01"')
.groupby('date')
['cases']
.sum()
)
data.index = pd.to_datetime(data.index)
sim = simulate(data, forecast_n_days=3, n_history=None)
sim.plot.bar(figsize=(18, 5));
```
# Mundo
```
ser = 'China'
data = (
jhu
.set_index('country')
.loc[ser]
.reset_index(drop=True)
.set_index('date')
['cases']
)
data.index = pd.to_datetime(data.index)
sim = simulate(data, forecast_n_days=3, n_history=None)
sim.plot.bar(figsize=(18, 5));
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import cv2
from datetime import datetime
import time
from tqdm import tqdm
import os
import sys
from skimage import io
from matplotlib import pyplot as plt
%matplotlib inline
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
from unet.maskprocessor import *
from unet.normalization import *
from unet.loss import *
```
The purpose of this notebook is:
To evaluate the model performance on unseen test set. The test set is sourced from DigitalGlobe (WorldView 2) Open Data Program which is lower resolution and noisier than the hi-res MapBox satellite imageries used in training set and validation set.
Pre-requisite:
Directory containing the source of truth MapBox street tiles. These could be obtained via the MapBox API.
Directory containing the inferred mask tiles. These tiles could be generated by roadSegmentationMaskGen.py.
```
index = 2
dir_truth = '/Users/jkwok/Documents/Insight/tools/jTileDownloader 2/digitalglobe/mapbox_custom_street/'
dir_inferred = ['/Users/jkwok/Documents/Insight/tools/jTileDownloader 2/digitalglobe/jackkwok.digitalglobe_harvey_3020132_tif_mask/',
'/Users/jkwok/Documents/Insight/tools/jTileDownloader 2/digitalglobe/141812_post_mask/',
'/Users/jkwok/Documents/Insight/tools/jTileDownloader 2/digitalglobe/214437_post_mask/']
model_file = ['/Users/jkwok/Documents/Insight/models/Unet_Dilated-20170917-223544.hdf5',
'/Users/jkwok/Documents/Insight/models/Unet_Dilated-20170921-141812.hdf5',
'/Users/jkwok/Documents/Insight/models/Unet_Dilated-20170921-214437.hdf5']
def image_file_list(dir_path):
""" limitation: the images files must have an image extension: webp, jpg, png, or jpeg """
result = []
for root, dirs, files in os.walk(dir_path):
for file in files:
if file.endswith('.webp') or file.endswith('.png') or file.endswith('.jpg') or file.endswith('.jpeg'):
result.append(os.path.join(root, file))
return result
inferred_filelist = image_file_list(dir_inferred[index])
truth_filelist = []
for img_file in inferred_filelist:
truth_file = img_file.replace(dir_inferred[index], dir_truth)
truth_file = truth_file.replace('.jpg', '.png')
truth_filelist.append(truth_file)
img_df = pd.DataFrame(
{'infer': inferred_filelist,
'truth': truth_filelist,
})
img_df.head()
infer_jpg_filename = img_df.loc[0, 'infer']
truth_jpg_filename = img_df.loc[0, 'truth']
print(infer_jpg_filename)
print(truth_jpg_filename)
infer_jpg_img = cv2.imread(infer_jpg_filename)
truth_jpg_img = cv2.imread(truth_jpg_filename)
print(infer_jpg_img.shape)
print(truth_jpg_img.shape)
infer_image = cv2.cvtColor(infer_jpg_img, cv2.COLOR_BGR2GRAY)
mask = get_street_mask(truth_jpg_img)
print('binary mask', np.unique(mask))
plt.imshow(mask, cmap=plt.cm.binary)
new_style = {'grid': False}
mask.dtype='uint8'
mask[mask == 1] = 255
print(np.unique(infer_image))
print(np.unique(mask))
dice = np.sum(infer_image[mask==255])*2.0 / (np.sum(infer_image) + np.sum(mask))
print(dice)
# compute dice coef score between source of truth and prediction
for i in tqdm(range(len(inferred_filelist)), miniters=10):
infer_jpg_filename = img_df.loc[i, 'infer']
truth_jpg_filename = img_df.loc[i, 'truth']
infer_jpg_img = cv2.imread(infer_jpg_filename)
truth_jpg_img = cv2.imread(truth_jpg_filename)
infer_image = cv2.cvtColor(infer_jpg_img, cv2.COLOR_BGR2GRAY)
mask = get_street_mask(truth_jpg_img)
mask.dtype='uint8'
mask[mask == 1] = 255
#print(np.unique(infer_image))
#print(np.unique(mask))
dice = np.sum(infer_image[mask==255])*2.0 / (np.sum(infer_image) + np.sum(mask))
img_df.set_value(i, 'dice_coef', dice)
img_df.head()
# 0.270229 (no blur) Unet_Dilated-20170917-223544.hdf5
# 0.305918 (with blur aug during training) Unet_Dilated-20170921-141812.hdf5
# 0.334121 (with blur aug during training large dataset)
x = img_df[img_df['dice_coef']!=0]
avg_dice = x.mean()
print 'average dice score: ', avg_dice
sorted_df = img_df.sort_values('dice_coef', ascending=False)
sorted_df.head(10)
```
| github_jupyter |
<h1><center>Tutorial 3: Probability</center></h1>
<div style="text-align: center">Adnane Ez-zizi, 16 Oct 2019</div>
$\newcommand{\E}{\mathrm{E}}$
$\newcommand{\Var}{\mathrm{Var}}$
$\newcommand{\Cov}{\mathrm{Cov}}$
$\newcommand{\Corr}{\mathrm{Corr}}$
This notebook introduces the elementary concepts from the probability theory that are necessary to understand and build machine learning models. For example, we will cover notions such as a random variable, a probability distribution and independence, as well as useful properties of probability like the chain rule or Bayes theorem. We will also illustrate these notions using Python code.
## 0. Preliminary steps
```
import numpy as np
from scipy import stats
import matplotlib.pyplot as plt
#%precision 3
```
## 1. Definition of probability
Probability, at its core, refers to the likelihood or the chance that a certain event will occur. For instance, when we roll a fair die, we can intuitively report that there is a probability of $\frac{1}{6}$ that one of the six possible outcomes (1, 2, 3, 4, 5 or 6) will occur.
Probability, as a field, is branch of mathematics that provides tools to deal and measure uncertainty. In machine learning, we often have to deal with uncertain or stochastic quantities, whether the system that we try to model is assumed inherently stochastic like the trajectory of a subatomic particle, or cannot be completely observed like a robot with a low quality camera trying to identify objects, or because of incomplete modelling like when have to discard some of the information that we have collected (e.g., a robot that discretises the space when making a prediction about the position of an object).
Before we move to more advanced notions, let's cover some elementary terminology that is commonly used in probability theory:
- **Experiment:** any process, usually random, that results in one of many possible results. For example in our first example, the toss of a die can be considered as an experiment.
- **Trial:** one repetition of the process of an experiment. For example, in the experiment of tossing a die 10 times and recording the sequence of results, each toss is considered as a trial.
- **Outcome:** each distinct possible result of an experiment. For example, in the experiment of tossing a die twice, one possible outcome is (3,5) where 3 is the result from the first trial and 5 is the result from the second trial.
- **Event:** consists of one or more outcomes. For example, in the experiment of tossing on die, drawing an even number can be considered as an event, which has 3 possible outcomes (2, 4 and 6).
- **Sample space** set of all possible outcome. For the die experiment, the sample space is {1,2,3,4,5,6}.
In mathematical notation, the probability of an event E is often denoted as $P(E)$ or $Pr(E)$.
## 2. Random variables
A random variable is a function that maps a possible outcome of a random process to a real number. For example, we can define a random variable that represents the outcome from a die roll, $X \in \{1,2,3,4,5,6\}$, or the sum of the outcomes from rolling two dice, $Y \in \{1,2,\cdots,12\}$.
Once we have defined a random variable, we can talk about its probability of taking a certain value or of taking a value in a given interval. Thus, the probability of observing a 3 when we roll a die is expressed as $P(X = 3)$, while the probability of observing an even outcome can be expressed as $P(X \in \{2,4,6\})$.
There are two types of random variables: discrete and continuous. A random variable is said to be discrete if it takes a finite or countably infinite number of values such as the variable $X$ representing the die roll's outcome. A continous random variable can take real values such as the value of a share in a stock exchange market.
## 3. Discrete variables
### 3.1. Probability mass function
The probability mass function, or distribution of a discrete random variable $X$ maps the possible values of $X$ ($x_1, x_2, \dots$) to their probabilities. We will denote a probability mass function by a lower-cased $p(.)$: $p(x_i) = P(X = x_i)$ for all possible values $x_i$'s of X.
For example, the probability mass function of the random variable $X$ that represents a die roll outcome is defined by:
\begin{equation}
p(x) = P(X = x) = \frac{1}{6} \;\; \text{for} \; x \in \{1, 2, 3, 4, 5, 6\}
\end{equation}
A probability mass function $p$ of a random variable $X$ must satisfy 3 conditions:
1. The domain of $p$ must be the set of all possible outcomes of $X$.
2. for all possible values $x$ of $X$, $0\leq p(x) \leq 1$, where $p(x) = 0$ if $x$ can never occur (impossible outcome) and $p(x) = 1$ if $x$ always occur.
3. $\sum_{x} {p(x)} = 1$, that is, the probabilities of all possible outcomes sum to 1.
### 3.2. Expectation and variance
#### Expectation
The expectation or expected value (or mean) of a discrete random variable $X$ is defined as:
$\E(X) = \sum_{i} {x_ip(x_i)}$. In other words, the expectation is a weighted average of the possible values $x_i$ with weights $p(x_i)$, similar to a center of mass in physics. It gives us an idea about the central value taken by the random variable.
For the die roll example above, the expectation is equal to:
\begin{equation}
\E(X) = 1 \cdot \frac{1}{6} + 2 \cdot \frac{1}{6} + 3 \cdot \frac{1}{6} + 4 \cdot \frac{1}{6} + 5 \cdot \frac{1}{6} + 6 \cdot \frac{1}{6} = \frac{21}{6} = 3.5
\end{equation}
The expectation has a few properties:
- The expectation of a deterministic value is equal to the value itself: $\E(x) = x$, where x is a non-random value.
- The expectation is a linear function, that is: $\E(aX + bY) = a\E(X) + b\E(Y)$.
- $\E(f(X)) = \sum_{i} {f(x_i)p(x_i)}$
#### Variance
The variance provides a measure of how much the values of a random variable $X$ vary as we sample different values of $X$ from its probability distribution. Mathematically, it is defined as: $\Var(X) = \E(\left[X - \E(X)\right]^2) = \E(X^2) - \E(X)^2$. For instance, the variance of $X$ in the die roll example is given by:
\begin{equation}
\Var(X) = \E(X^2) - \E(X)^2 = (1^2 \cdot \frac{1}{6} + 2^2 \cdot \frac{1}{6} + 3^2 \cdot \frac{1}{6} + 4^2 \cdot \frac{1}{6} + 5^2 \cdot \frac{1}{6} + 6^2 \cdot \frac{1}{6}) - \frac{21}{6} = \frac{71}{6}
\end{equation}
The variance has a few properties:
- The variance of a deterministic value is equal to 0: $\Var(x) = 0$, where $x$ is a non-random value.
- The variance preserves the sum: $\Var(X + Y) = \Var(X) + \Var(Y)$.
- The variance is a non linear function for the multiplication by a scalar: $\Var(aX) = a^2\Var(X)$.
- $\Var(f(X)) = \E(f(X)^2) - \E(f(X))^2$
The standard deviation is a related measure defined as the square root of the variance: $SD(X) = \sqrt{\Var(X)}$.
### 3.3. Some popular discrete probability distributions
#### Bernoulli distribution
Suppose that we run a random experiment whose outcome can be classified as either a success or a failure. An example of such an experiment is tossing a fair coin where a head is considered as success and tail as failure. If we let $X = 1$ when the outcome is a success (e.g. head) and $X = 0$ when it is a failure (e.g. tail), then the probability mass function of $X$ is given by:
\begin{aligned}
& p(0) = P(X = 0) = 1 − p\\
& p(1) = P(X = 1) = p
\end{aligned}
where $p$ is the probability that the experiment is a success ($0\leq p\leq 1$).
A random variable $X$ is said to be a Bernoulli random variable if its probability mass function is given by the equations
shown just above for some $p \in (0, 1)$. We also say that $X$ follows a Bernoulli distribution with parameter $p$ and write this mathematically as $X \sim Bernoulli(p)$.
```
# Mass function of a Bernoulli(0.7) distribution
print('p(0) ={p0: 1.1f}'.format(p0 = stats.bernoulli.pmf(0, p = 0.7)))
print('p(1) ={p0: 1.1f}'.format(p0 = stats.bernoulli.pmf(1, p = 0.7)))
```
We can draw samples from a Bernoulli distribution, using the function `stats.bernoulli.rvs()`. For example, if we want to draw 10 values from a $Bernoulli(0.7)$ distribution, we would use:
```
# Draw 10 values from a Bernoulli variable with p = 0.7
stats.bernoulli.rvs(p = 0.7, size = 10)
```
The expection and variance of a random variable $X \sim Bernoulli(p)$ can be computed in a straightforward manner as:
\begin{align}
& \E(X) = 0 \cdot p(0) + 1 \cdot p(1) = p\\
& \Var(X) = E(X^2) - E(X)^2 = (0^2 \cdot p(0) + 1^2 \cdot p(1)) - p^2 = p(1-p)
\end{align}
```
# Expectation and variance of a X ~ Bernoulli(0.7)
print('E(X) ={exp: 1.2f}'.format(exp = stats.bernoulli.mean(p = 0.7)))
print('Var(X) ={var: 1.2f}'.format(var = stats.bernoulli.var(p = 0.7)))
```
#### Binomial distribution
Suppose now that we perform an experiment with $n$ independent trials, each of which results in a success with probability $p$ and in a failure with probability $1 − p$. If $X$ represents the number of successes that occur in the $n$ trials, then $X$ is said to be a binomial random variable with parameters $(n, p)$. A Bernoulli random variable is thus just a binomial random variable with parameters $(1, p)$.
The probability mass function of a binomial random variable, $X$, having parameters $(n, p)$ (we can write $X \sim B(n,p)$), is given by:
\begin{equation}
p(i) = {n \choose i} p^i (1 − p)^{n−i} \quad i = 0, 1,\dots, n
\end{equation}
where ${n \choose i}$ is the number of possible combinations that result from randomly drawing $i$ objects out of $n$ different objects. Mathematically, ${n \choose i} = \frac{n!}{i!\,(n-i)!}$, where $k! = k\times(k-1)\times(k-2)\dots\times1$.
```
# Mass function of a B(6,0.5) distribution
for i in range(7):
print('p({j:d}) = {p0:1.4f}'.format(j = i, p0 = stats.binom.pmf(i, n = 6, p = 0.5)))
# Plor of the mass function
fig, ax = plt.subplots(1, 1)
n, p = 6, 0.5
x = list(range(7))
ax.plot(x, stats.binom.pmf(x, n, p), 'bo', ms=8, label='binom pmf')
ax.vlines(x, 0, stats.binom.pmf(x, n, p), colors='b', lw=5, alpha=0.5)
plt.show()
```
In code, to draw samples from a Binomial distribution, we can call `stats.binom.rvs()`. For example, drawing 3 values from a $B(6, 0.5)$ distribution can be done using the following code:
```
# Draw 3 values from a variable following a B(6, 0.5)
print(stats.binom.rvs(n = 6, p = 0.5, size = 3))
```
This means that we got 4 successes ($X = 1$) in each of the first two draws, and got 2 successes in the last draw.
The expection and variance of a random variable $X \sim B(n,p)$ are given by:
\begin{align}
& \E(X) = np\\
& \Var(X) = p(1-p)
\end{align}
```
# Expectation and variance of a X ~ B(6, 0.5)
print('E(X) ={exp: 1.1f}'.format(exp = stats.binom.mean(n = 6, p = 0.5)))
print('Var(X) ={var: 1.1f}'.format(var = stats.binom.var(n = 6, p = 0.5)))
```
## 4. Continuous variables
### 4.1. Probability density function
When working with a continuous random variable, we no longer want to find the probability that the variable take a specific value such as asking what is the probability that the value of a certain share in a financial market is exactly £100.684, since this probability will virtually be 0. Instead, we are more interested in the probability that the continuous random variable lies in a certain interval. For that, we normally describe the distribution of a random variable using a probability density function instead of a probability mass function. More specifically, a function $f$ is said to be the density of a random variable $X$ if, for any interval $I$, $P(X \in I) = \int_{I}{f(x)dx}$.
In general, a function $f$ must satisfy 3 conditions to be considered as a density function of a random variable $X$:
1. The domain of $f$ must be the set of all possible outcomes of $X$.
2. for all possible values $x$ of $X$, $f(x) \geq 0$.
3. $\int{f(x)dx} = 1$.
### 4.2. Expectation and variance
#### Expectation
The definition of expectation in the case of continuous random variables is very similar to that in the discrete case, where we only have to replace the summation with integration, and the probability mass function with the probability density function. More specifically, if $X\sim f$ (meaning that X follows a distribution whose density function is $f$) and the domain of $X$ is $\mathcal{X}$, then: $\E(X) = \int_{\mathcal{X}} {xf(x)dx}$.
#### Variance
For variance, the definition remains the same: $\Var(X) = \E(\left[X - \E(X)\right]^2) = \E(X^2) - \E(X)^2$.
### 4.3. Cumulative distribution function
Let $X$ be a random variable. The function $F$ defined by:
\begin{equation}
F(x) = P(X \leq x), \quad -\infty \leq x\leq \infty
\end{equation}
is called the cumulative distribution function of $X$. Thus, $F$ specifies, for all real values $x$, the probability that the random variable is less than or equal to $x$.
Similarly to the probability density function, the cumulative distribution function $F$ describes fully the distribution of a random variable, with the additional advantage that it can work with any type of random variables (discrete or continuous). Actually, F can be obtained from $f$, the probability density function, using the formulas: $F(x) = \int{f(t)dt}$
The One interesting property of $F$ is that it is a nondecreasing function; that is, if $a < b$, then $F(a) \leq F(b)$.
### 4.4. Some popular continuous probability distributions
#### Uniform distribution
A random variable $X$ is said to be uniformly distributed over the interval $(a, b)$, noted in short as $X \sim U(a,b)$, if $X$ is equally likely to fall anywhere in this interval. The probability density function of such random variable is given by:
\begin{equation}
f(x) =
\left\{
\begin{array}{ll}
\frac{1}{b-a} & \mbox{if } a\leq x\leq b \\
0 & \mbox{otherwise }
\end{array}
\right.
\end{equation}
```
# density function of a U(0,1) distribution
print('f(-0.2) = {fx:1.1f}'.format(fx = stats.uniform.pdf(-0.2, loc = 0, scale = 1)))
print('f(0) = {fx:1.1f}'.format(fx = stats.uniform.pdf(0, loc = 0, scale = 1)))
print('f(0.3) = {fx:1.1f}'.format(fx = stats.uniform.pdf(0.1, loc = 0, scale = 1)))
print('f(1) = {fx:1.1f}'.format(fx = stats.uniform.pdf(1, loc = 0, scale = 1)))
print('f(4.5) = {fx:1.1f}'.format(fx = stats.uniform.pdf(4.5, loc = 0, scale = 1)))
# Plot of the density function
fig, ax = plt.subplots(1, 1)
a, b = 0, 1
x = np.linspace(-2, 2, 200)
ax.plot(x, stats.uniform.pdf(x, loc = a, scale = b), 'r-', lw=2, label='frozen pdf')
plt.show()
```
In code, to draw samples from a Uniform distribution, we can call `stats.uniform.rvs()`. For example, drawing 10 values from a $U(0, 1)$ distribution can be done using the following code:
```
# Draw 10 values from a variable following a U(0, 1)
stats.uniform.rvs(loc = 0, scale = 1, size = 10)
```
Since $F(x) = \int{f(t)dt}$, it follows from the above equation that the distribution function of a uniform random variable on the interval $(a, b)$ is given by:
\begin{equation}
F(x) =
\left\{
\begin{array}{ll}
0 & \mbox{if } x< a \\
\frac{x-a}{b-a} & \mbox{if } a\leq x\leq b \\
1 & \mbox{otherwise }
\end{array}
\right.
\end{equation}
The expectation and variance of the $U(a,b)$ distribution are given by:
\begin{align}
& \E(X) = \frac{a+b}{2}\\
& \Var(X) = \frac{(b-a)^2}{12}
\end{align}
```
# Expectation and variance of a X ~ U(0, 1)
print('E(X) ={exp: 1.1f}'.format(exp = stats.uniform.mean(loc = 0, scale = 1)))
print('Var(X) ={var: 1.4f}'.format(var = stats.uniform.var(loc = 0, scale = 1)))
```
#### Normal distribution
$X$ is said to be a normal random variable (or normally distributed), with parameters $\mu$ and $\sigma^2$ if the density of $X$ is given by:
\begin{equation}
f(x) = \frac{1}{\sqrt{2\pi \sigma^2}} \exp\left(-\frac{1}{2 \sigma^2} (x-\mu)^2\right)
\end{equation}
In short, we denote a normal distribution with parameters $\mu$ and $\sigma^2$ by $\mathcal{N}(\mu,\sigma^2)$, so we will also write as before $X \sim \mathcal{N}(\mu,\sigma^2)$ to introduce a normal random variable $X$.
```
# density function of a N(0,1) distribution
print('f(-1.96) = {fx:1.2f}'.format(fx = stats.norm.pdf(-1.96, loc = 0, scale = 1)))
print('f(0) = {fx:1.2f}'.format(fx = stats.norm.pdf(0, loc = 0, scale = 1)))
print('f(1.96) = {fx:1.2f}'.format(fx = stats.norm.pdf(1.96, loc = 0, scale = 1)))
print('f(1) = {fx:1.2f}'.format(fx = stats.norm.pdf(1, loc = 0, scale = 1)))
# Plot of the density function
fig, ax = plt.subplots(1, 1)
mu, sigma = 0, 1
x = np.linspace(-5, 5, 100)
ax.plot(x, stats.norm.pdf(x, loc = mu, scale = sigma), 'r-', lw=2, label='frozen pdf')
plt.show()
```
In code, to draw samples from a Uniform distribution, we can call `stats.uniform.rvs()`. For example, drawing 10 values from a $U(0, 1)$ distribution can be done using the following code:
```
# Draw 10 values from a variable following a N(0, 1)
stats.norm.rvs(loc = 0, scale = 1, size = 10)
```
The expectation and variance of a normally distributed random variable, $X \sim \mathcal{N}(\mu,\sigma^2)$, are respectively equal to the paramters of the distribution, that is: $\E(X) = \mu$ and $\Var(X) = \sigma^2$. In other words a normal distribution is completely specified using its expectation and variance.
```
# Expectation and variance of a X ~ U(0, 1)
print('E(X) ={exp: 1.1f}'.format(exp = stats.norm.mean(loc = 0, scale = 1)))
print('Var(X) ={var: 1.1f}'.format(var = stats.norm.var(loc = 0, scale = 1)))
```
## 5. Joint and conditional distributions
### 5.1. Joint probability
Up to now, we have only discussed probability distributions for single random variables. But what if we are interested in the outcomes of two or more random variables? For example, suppose we toss two fair coins and wants to know the probability that the first coin turns up head and the second turns up tail, or in a formal way, $P(X_1 = H, X_2 = T)$, where $X_1$ and $X_2$ respectively represent the outcome of the first and second coin. Let's write the probabilities of all possible joint outcomes in a table:
outcome | probability
---- | -------
HT | 1/4
HH | 1/4
TH | 1/4
TT | 1/4
What we just did is to define a joint probability distribution for the random variables $X_1$ and $X2$. Technically, as before, we can talk either about a joint probability mass function in the case of discrete variables or joint probability density function (or cumulative distribution function) when the variables are continuous.
#### Joint probability mass function (discrete case):
Assume we have two discrete random variables $X$ and $Y$, which can take on values $x_1, x_2, \dots$ and $y_1, y_2, \dots$, respectively. Their joint probability mass function is defined as:
\begin{equation}
p(x_i, y_i) = P(X = x_i, Y = y_i) \; \text{for all possible values $x_i$'s and $y_i$'s of $X$ and $Y$}.
\end{equation}
A joint probability mass function $p$ of a random variable $X$ must satisfy the same 3 conditions seen before for a standard mass function:
1. The domain of $p$ must be the set of all possible joint outcomes of $X$ and $Y$.
2. for all possible values $x$ of $X$ and $y$ of $Y$, $0\leq p(x, y) \leq 1$
3. $\sum_{x,y} {p(x,y)} = 1$, that is, the probabilities of all possible outcomes sum to 1.
One of the widely-used joint distributions is the [multinomial distribution](https://en.wikipedia.org/wiki/Multinomial_distribution), which generalises the binomial distribution when we are interested in the probability of any possible combination of number of success of each of $k$ categories (instead of two) in $n$ independent trials. For example, say we roll a 6-sided die $n$ times independently. The multinomial distribution would then give the probabiliy of counts of each of the 6 sides (here, $k=6$). The parameters of a multinomial are: $n$, the number of independent trials and $p_1, \dots, p_k$ the probabilities of successes of each of the $k$ categories to compute the count of successes for. We thus often denote the multinomial distribution by $Mult(n, p_1, \dots, p_k)$
#### Marginal mass function (discrete case)
If we have a joint distribution of multiple discrete variables, we can extract the mass function of one of the variables by summing over the possible values of the other variables. The obtained distribution of that variable is called marginal distribution. Concretelely, say we have two random variables $X$ and $Y$ that represent the outcomes of coin flips. Their joint mass function is given by:
\begin{equation}
p(i,j) = \frac{1}{4} \;\text{for } i,j \in \{H,T\}
\end{equation}
The marginal mass function of $X$ can be extracted as follows:
\begin{equation}
p(i) = \sum_{j \in \{H,T\}}{p(i,j)} = \frac{1}{4} + \frac{1}{4} = \frac{1}{2}
\end{equation}
#### Joint cumulative and density functions (continuous case):
Joint cumulative distribution function can be easily generalised from the single variable case to the multiple-variable case. If $X$ and $Y$ are two continous random variables, their joint cumulative distribution function is defined as:
\begin{equation}
F(x, y) = P(X \leq x, Y \leq y), \quad -\infty \leq x,y\leq \infty
\end{equation}
We say that $f$ is the joint probabiliy density function of $X$ and $Y$ if, for any interval $I$ and $J$, $P(X \in I, Y \in J) = \int_{I,J}{f(x,y)dxdy}$. The density function $f$ also satisfies the same 3 conditions seen in the case of a single variable, that is:
1. The domain of $f$ must be the set of all possible joint outcomes of $X$ and $Y$.
2. for all possible values $x$ of $X$ and $y$ of $Y$, $f(x, y) \geq 0$.
3. $\int{f(x,y)dxdy} = 1$.
#### Marginal density function (continous case)
As in the discrete case, if we have a joint distribution of two (or multiple) continuous random variables $X$ and $Y$, we can marginalizing over $Y$ to get the marginal distrubution of $X$. This is done by integrating over the values of $Y$:
\begin{equation}
f(x) = \int{f(x,y)dy}
\end{equation}
### 5.2. Conditional probability
#### Definition
We often want to know the probability of one event, given that we know another event is true. For instance, given that a house is located in a posh area, what is the probability that its price is greater than £500,000. This is refered to as a conditional probability, and in our case, we write it as $P(\text{price } > 500000 \text{ | area is posh})$.
If we know the joint distribution of two random variables, then we can get the conditional distribution of one variable given the other variable as follows:
- In the discrete case, the conditional mass function of $X$ given $Y$ is equal to: $p(x|y) = \frac{p(x,y)}{p(y)} = \frac{p(x,y)}{\sum_{x}{p(x,y)}}$
- In the continuous case, the conditional density function of $X$ given $Y$ is equal to: $f(x|y) = \frac{f(x,y)}{f(y)} = \frac{f(x,y)}{\int{f(x,y)dx}}$
The definition can be extended easily to the case of more than two variables by simply replacing one of the variable in the formula with the multiple variables that we want to condition on or compute the conditial distribution for.
#### Chain rule
Any joint probability distribution over many random variables may be decomposed into conditional distributions over only one variable:
\begin{equation}
P(x_1, \cdots , x_n) = P(x_n|x_{n-1}, \cdots, x_1)P(x_{n-1}|x_{n-2}, \cdots, x_1) \cdots P(x_2|x_1) P(x_1)
\end{equation}
Thus, for example, we have:
\begin{equation}
P(x,y,z) = P(x|y,z)P(y|z)P(z)
\end{equation}
#### Bayes theorem
Bayes theorem, named after the reputable Mathematician Thomas Bayes, is a simple yet a powerful formula that allows to flip a conditional probability. It basically states that:
\begin{equation}
p(y|x) = \frac{p(x|y)p(y)}{p(x)} = \frac{p(x|y)p(y)}{\sum_y{p(x|y)p(y)}}
\end{equation}
One major application of Bayes theorem is to compute the probability of a model (or the parameters of the model to be more precise) given some data. Knowing the model, we can determine the probability of the data given a specific set of model parameters. Bayes’ rule allows us to get from the probability of the data given the model $P(data|model)$, to the probability of the model given the data $P(model|data)$. In other words, it allows us to estimate the parameters of a model having collected a set of data.
### 5.3. Independence and conditional independence
#### Independence
Two random variables $X$ and $Y$ are said to be independent (often denoted as $X\perp Y$) if the values taken by one of the variables have no influence in the values taken by the other variable. Formally, this can be expressed using conditional probabilities as: $p(x|y) = p(x)$ in the discrete case, or $f(x|y) = f(x)$ in the continous case.
knowing that $p(x|y) = \frac{p(x,y)}{p(y)}$
then $\frac{p(x,y)}{p(y)} = p(x)$
which means that $p(x,y) = p(x)p(y)$
Actually, in textbooks of probability, you often find that independence is introduced using the last formula instead of using conditional probabilities.
#### Conditional independence
Two random variables $X$ and $Y$ are conditionally independent given a random variable $z$ (denoted as $X\perp Y \;|\; Z$) if the conditional joint mass (or density) function of $X$ and $Y$ is equal to the product of the conditional mass (or density) functions of the two random variables:
\begin{equation}
p(x,y|z) = p(x|z)p(y|z)
\end{equation}
### 5.4. Covariance and correlation
#### Covariance
The covariance between $X$ and $Y$, denoted by $\Cov(X,Y)$, is defined by
\begin{equation}
\Cov(X,Y) = \E\left[(X-\E(X)) (Y-\E(Y))\right]
\end{equation}
The covariance has a few properties:
- The covariance is symmetric: $\Cov(X,Y) = \Cov(Y,X)$
- The covariance is a linear function of each argument, that is: $\Cov(aX,Y) = a\,\Cov(Y,X)$.
- If $X$ and $Y$ are independent, then $\Cov(X,Y) = 0$.
#### Correlation
The correlation of two random variables $X$ and $Y$, denoted by $\Corr(X,Y)$, is defined by:
\begin{equation}
\Corr(X,Y)= \frac{\Cov(X,Y)}{\sqrt{\Var(X)\Var(Y)}}
\end{equation}
where $\Var(X)$ and $\Var(Y)$ are strictly positive. One can show that $-1\leq\Corr(X,Y)\leq 1$
The correlation coefficient is a measure of the degree of linearity between $X$ and $Y$. A value of $\Corr(X,Y)$ near $1$ or $−1$ indicates a high degree of linearity between $X$ and $Y$, whereas a value near $0$ indicates that such linearity is absent. A positive value of $\Corr(X,Y)$ indicates that $Y$ tends to increase when $X$ does, whereas a negative value indicates that $Y$ tends to decrease when $X$ increases. If $\Corr(X,Y) = 0$, then $X$ and $Y$ are said to be uncorrelated.
## 6. Law of large numbers, Central limit theorem
Here we will cover two of the most fundamental theorems in Probability
### 6.1. Law of large numbers
Suppose that we have a collection of random variables $X_1, X_2, \cdots, X_n$ that are independent and identically distributed. The strong law of large numbers states that as the number of random variables increase ($n\to \infty$), the empirical average of the random variables will get closer to the true common expected value of the variables normal distribution:
\begin{equation}
\frac{1}{n}\sum_{i=1}^n{X_i} \xrightarrow[n\to \infty]{} \E(X)
\end{equation}
where the convergence is almost sure, that is:
\begin{equation}
P\left( \lim_{n \to \infty} \frac{1}{n}\sum_{i=1}^n{X_i} = \E(X) \right) = 1
\end{equation}
### 6.2. Central limit theorem
Let's $X_1, X_2, \cdots, X_n$ be a collection of random variables that are independent and identically distributed. The central limit theorem states that if we sum them up, then the larger the collection, the closer the sum will be to a normal distribution. Mathematically, this is translated as:
\begin{equation}
\frac{1}{\sqrt{n}} \sum_{i=1}^n \frac{X_i - \mu}{\sigma} \xrightarrow[n\to \infty]{} \mathcal{N}(0, 1) \quad \left(\text{or equivalently}, \sum_{i=1}^n{X_i} \underset{n\to \infty}{\sim} \mathcal{N}\left(\mu, \frac{\sigma}{n}\right)\right)
\end{equation}
where $\mu$ and $\sigma^2$ are respectively the common expectation and variance of the random variables $X_1, X_2, \cdots, X_n$, and the convergence is in law, that is:
\begin{equation}
\lim_{n \to \infty} P\left( \frac{1}{\sqrt{n}} \sum_{i=1}^n \frac{x_i - \mu}{\sigma} \le z \right) = \int_{-\infty}^z
(1/\sqrt{2 \pi}) \exp(-u^2/2) \, du
\end{equation}
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import theano
import pymc3 as pm
%matplotlib inline
```
The data was downloaded from https://www.kaggle.com/camnugent/sandp500
```
df = pd.read_csv('../data/all_stocks_5yr.csv.zip', compression='zip')
df.head()
stocks = ['MSFT', 'AAPL', 'GOOG', 'NEM', 'ALB']
df_stocks = df.loc[df.Name.isin(stocks), ['Date', 'Close', 'Name']]
df_stocks.shape
df_stocks = df_stocks.pivot(index='Date', columns='Name', values='Close')
df_stocks.head()
df_stocks.plot(figsize=(12, 5))
plt.show()
y_mean = df_stocks.mean()
y_std = df_stocks.std()
df_stocks = (df_stocks - y_mean) / y_std
df_stocks.plot(figsize=(12, 5))
plt.show()
```
## Generating Synthetic data
```
N = 1000
D = 3
Q = np.random.randn(D, D)
Σ = Q.dot(Q.T)
e = np.random.randn(N, D)
e = e.dot(Σ)
y2 = np.cumsum(e, axis=0)
plt.figure(figsize=(12, 5))
plt.plot(y2)
plt.show()
```
## Inference Process
```
def inference(y, t, window=365, n_samples=100):
N, D = y.shape
t_section = np.int64(np.arange(len(y)))//window
k = t_section.max()+1
t_t = theano.shared(np.repeat(t[:,np.newaxis], D, axis=1))
y_t = theano.shared(y)
t_section_t = theano.shared(t_section)
with pm.Model() as model:
sd_α = pm.HalfCauchy.dist(beta=0.5)
packed_L_α = pm.LKJCholeskyCov('packed_L_α', n=D, eta=2., sd_dist=sd_α)
L_α = pm.expand_packed_triangular(D, packed_L_α)
Σ_α = pm.Deterministic('Σ_α', L_α.dot(L_α.T))
μ_β = pm.Bound(pm.Normal, lower=0, upper=1)('μ_β', sd=0.5, shape=D)
sd_β = pm.HalfCauchy.dist(beta=0.5)
packed_L_β = pm.LKJCholeskyCov('packed_L_β', n=D, eta=2., sd_dist=sd_β)
L_β = pm.expand_packed_triangular(D, packed_L_β)
Σ_β = pm.Deterministic('Σ_β', L_β.dot(L_β.T))
α = pm.MvGaussianRandomWalk('alpha', shape=(k+1, D), cov=Σ_α)
β = pm.MvGaussianRandomWalk('beta', shape=(k+1, D), mu=μ_β, cov=Σ_β)
alpha_r = α[t_section_t]
beta_r = β[t_section_t]
regression = alpha_r+beta_r*t_t
sd = pm.Uniform('sd', 0, 1)
likelihood = pm.Normal('y', mu=regression, sd=sd, observed=y_t)
trace = pm.sample(n_samples, njobs=4)
return trace, t_section
t = np.arange(len(df_stocks))
t = (t-t.mean())/t.std()
trace, t_section = inference(df_stocks.values, t, n_samples=500)
a_mean = trace['alpha'][-1000:].mean(axis=0)
b_mean = trace['beta'][-1000:].mean(axis=0)
y_pred = a_mean[t_section] + b_mean[t_section]*t[:,None]
# Un-normalise the data
y_pred = y_pred*y_std[None,:] + y_mean[None,:]
y = df_stocks.values*y_std[None,:] + y_mean[None,:]
dates = np.array([np.datetime64(d) for d in df_stocks.index])
stocks = df_stocks.columns.values
for i, stock in enumerate(stocks):
plt.figure(figsize=(12,5))
plt.plot(dates, y[:, i], label=stock)
plt.plot(dates, y_pred[:, i], label="trend")
plt.legend()
plt.title(stock.capitalize())
plt.show()
plt.imshow(trace['Σ_β'][1000:].mean(axis=0))
plt.colorbar()
plt.show()
plt.imshow(trace['Σ_β'][1000:].std(axis=0))
plt.colorbar()
plt.show()
```
## Correlation
```
stocks
trace_sigma2 = np.zeros_like(trace['Σ_β'][1000:])
for i,t in enumerate(trace['Σ_β'][1000:]):
t_diag = np.sqrt(t.diagonal())
trace_sigma2[i] = (t/t_diag[:,None])/t_diag[None,:]
plt.imshow(trace_sigma2.mean(axis=0))
plt.colorbar()
plt.show()
```
NEM and ALB have a correlation of 0.5 (as expected).
```
plt.imshow(trace_sigma2.std(axis=0))
plt.colorbar()
plt.show()
```
| github_jupyter |
# Building Data Genome Project 2.0
## Exploratory data analysis of metadata
Biam! (pic.biam@gmail.com)
```
# data and numbers
import numpy as np
import pandas as pd
# Visualization
import matplotlib.pyplot as plt
import matplotlib as mpl
import matplotlib.pylab as pylab
%matplotlib inline
import seaborn as sns
sns.set_style("darkgrid")
mpl.style.use('ggplot')
import gc
```
# Dataset
* `building_id`: building code-name with the structure <i>SiteID_[SimplifiedUsage](https://github.com/buds-lab/building-data-genome-project-2/wiki/Simplified-PSU)_UniqueName</i>.
* `site_id`: animal-code-name for the site.
* `building_id_kaggle`: building ID used for the [Kaggle competition](https://www.kaggle.com/c/ashrae-energy-prediction) (numeric).
* `site_id_kaggle`: site ID used for the [Kaggle competition](https://www.kaggle.com/c/ashrae-energy-prediction) (numeric).
* `primaryspaceusage`: Primary space usage of all buildings is mapped using the [energystar scheme building description types](https://www.energystar.gov/buildings/facility-owners-and-managers/existing-buildings/use-portfolio-manager/identify-your-property-type).
* `sub_primaryspaceusage`: [energystar scheme building description types](https://www.energystar.gov/buildings/facility-owners-and-managers/existing-buildings/use-portfolio-manager/identify-your-property-type) subcategory.
* `sqm`: Floor area of building in square meters (m2).
* `lat`: Latitude of building location to city level.
* `lng`: Longitude of building location to city level.
* `timezone`: site's timezone.
* `electricity`: presence of this kind of meter in the building. `Yes` if affirmative, `NaN` if negative.
* `hotwater`: presence of this kind of meter in the building. `Yes` if affirmative, `NaN` if negative.
* `chilledwater`: presence of this kind of meter in the building. `Yes` if affirmative, `NaN` if negative.
* `steam`: presence of this kind of meter in the building. `Yes` if affirmative, `NaN` if negative.
* `water`: presence of this kind of meter in the building. `Yes` if affirmative, `NaN` if negative.
* `irrigation`: presence of this kind of meter in the building. `Yes` if affirmative, `NaN` if negative.
* `solar`: presence of this kind of meter in the building. `Yes` if affirmative, `NaN` if negative.
* `gas`: presence of this kind of meter in the building. `Yes` if affirmative, `NaN` if negative.
* `industry`: Industry type corresponding to building.
* `subindustry`: More detailed breakdown of Industry type corresponding to building.
* `heatingtype`: Type of heating in corresponding building.
* `yearbuilt`: Year corresponding to when building was first constructed, in the format YYYY.
* `date_opened`: Date building was opened for use, in the format D/M/YYYY.
* `numberoffloors`: Number of floors corresponding to building.
* `occupants`: Usual number of occupants in the building.
* `energystarscore`: Rating of building corresponding to building energystar scheme ([Energy Star Score](https://www.energystar.gov/buildings/facility-owners-and-managers/existing-buildings/use-portfolio-manager/understand-metrics/how-1-100)).
* `eui`: [Energy use intensity](https://www.energystar.gov/buildings/facility-owners-and-managers/existing-buildings/use-portfolio-manager/understand-metrics/what-energy) of the building (kWh/year/m2).
* `site_eui`: Energy (Consumed/Purchased) use intensity of the site (kWh/year/m2).
* `source_eui`: Total primary energy consumption normalized by area (Takes into account conversion efficiency of primary energy into secondary energy).
* `leed_level`: LEED rating of the building ([Leadership in Energy and Environmental Design](https://en.wikipedia.org/wiki/Leadership_in_Energy_and_Environmental_Design")), most widely used green building rating system.
* `rating`: Other building energy ratings.
```
path = "..\\data\\metadata\\"
# Buildings data
metadata = pd.read_csv(path + "metadata.csv")
metadata.info()
```
# Exploratory Data Analysis
## Missing values
```
# Percentage of missing values in each feature
round(metadata.isna().sum()/len(metadata)*100,2)
```
## Categories
```
cat = ["site_id","primaryspaceusage","sub_primaryspaceusage","industry","subindustry","timezone"]
col = []
for feature in cat:
col_list = list(metadata[feature].unique())
len_col_list = len(list(metadata[feature].unique()))
col_list.insert(0, len_col_list)
col.append(col_list)
cat_df = pd.DataFrame.from_records(col).T.rename(
columns={
0: "site_id",
1: "primaryspaceusage",
2: "sub_primaryspaceusage",
3: "industry",
4: "subindustry",
5: "timezone",
}
)
cat_df
cat_df.to_csv("..\\temp\\cat_df.csv", index=False)
```
## Sites location
```
import geopandas as gpd
from shapely.geometry import Point, Polygon
# World map
world = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
# Exclude Antartica
world = world[(world.name != "Antarctica") & (world.name != "Fr. S. Antarctic Lands")]
# Coordinate reference system used in this map
world.crs
```
Latitude and longitude are of the site location (all buildings from the same location shares <code>lng</code> and <code>lat</code> values).
```
# All the sites
sites = metadata[["site_id","lat","lng"]].groupby("site_id").median()
# Converts lat and lng to geometry objects
geometry = [Point(xy) for xy in zip (sites["lng"], sites["lat"])]
# Creates geoDataFrame
geo_sites = gpd.GeoDataFrame(sites, crs = world.crs, geometry = geometry)
geo_sites
# Plot
sns.set(font_scale = 1)
fig, ax = plt.subplots(figsize = (15,15))
world.plot(ax = ax, alpha = 0.4, color = "grey")
geo_sites.plot(ax = ax, alpha = 0.8, color = "dodgerblue")
fig.savefig("..\\figures\\map.pdf", bbox_inches='tight')
fig.savefig("..\\figures\\map.png", dpi=72, bbox_inches='tight')
# Zoom Plot
fig, ax = plt.subplots(figsize = (15,15))
world.plot(ax = ax, alpha = 0.4, color = "grey")
geo_sites.plot(ax = ax, color = "dodgerblue")
ax.set_xlim([-125, 25])
ax.set_ylim([20, 60])
```
## Features frequency plots
```
len(np.unique(metadata["building_id"]))
# colors = ["crimson","mediumvioletred","orangered","gold","yellowgreen","lightseagreen","royalblue","rebeccapurple","slategray"]
sns.set(rc={'figure.figsize':(36,21)})
sns.set(font_scale = 2)
f, axes = plt.subplots(3, 3)
axes = axes.flatten()
color = "yellowgreen"
# primary use category countplot in decreasing order
# Temporary dataset
top5 = list(metadata['primaryspaceusage'].value_counts().iloc[:5].index)
temp = metadata[["primaryspaceusage"]].copy()
temp.loc[temp.primaryspaceusage.isin(top5) == False, "primaryspaceusage"] = "Other"
# Plot
ax1 = axes[0]
g1 = sns.countplot(data=temp, y="primaryspaceusage", color= color, orient="h", ax=ax1, order = temp['primaryspaceusage'].value_counts().index)
ax1.title.set_text('Primary use category')
ax1.set(ylabel="", xlabel="", xlim=(0,1600))
# adds percentage
total = float(len(metadata)) # number of buildings
for p in g1.patches:
width = p.get_width()
g1.text(width + 150,
p.get_y() + p.get_height()/1.5,
'{:1.2%}'.format(width/total),
ha="center")
del(top5, temp)
# primary use subcategory countplot in decreasing order
# Temporary dataset
top5 = list(metadata['sub_primaryspaceusage'].value_counts().iloc[:8].index)
temp = metadata[["sub_primaryspaceusage"]].copy()
temp.loc[temp.sub_primaryspaceusage.isin(top5) == False, "sub_primaryspaceusage"] = "Other"
# Plot
ax2 = axes[1]
g2 = sns.countplot(data=temp, y="sub_primaryspaceusage", color= color, orient="h", ax=ax2, order = temp['sub_primaryspaceusage'].value_counts().iloc[:16].index)
ax2.title.set_text('Primary use subcategory')
ax2.set(ylabel="", xlabel="", xlim=(0,1600))
# adds percentage
total = float(len(metadata)) # number of buildings
for p in g2.patches:
width = p.get_width()
g2.text(width + 150,
p.get_y() + p.get_height()/1.5,
'{:1.2%}'.format(width/total),
ha="center")
del(top5, temp)
# industry countplot in decreasing order
ax3 = axes[2]
g3 = sns.countplot(data=metadata, y="industry", color=color, ax=ax3, orient="h", order = metadata['industry'].value_counts().index)
ax3.title.set_text('Industry category (65% missing values)')
ax3.set(ylabel="", xlabel="", xlim=(0,1600))
# adds percentage
total = float(len(metadata)) # number of buildings
for p in g3.patches:
width = p.get_width()
g3.text(width + 150,
p.get_y() + p.get_height()/1.5,
'{:1.2%}'.format(width/total),
ha="center")
# subindustry countplot in decreasing order
# Temporary dataset
top5 = list(metadata['subindustry'].value_counts().iloc[:5].index)
temp = metadata[["subindustry"]].copy()
temp.loc[temp.subindustry.isin(top5) == False, "subindustry"] = "Other"
# Plot
ax4 = axes[3]
g4 = sns.countplot(data=temp, y="subindustry", color=color, ax=ax4, orient="h", order = temp['subindustry'].value_counts().index)
ax4.title.set_text('Subindustry category (65% missing values)')
ax4.set(ylabel="", xlabel="", xlim=(0,1600))
# adds percentage
total = float(len(metadata)) # number of buildings
for p in g4.patches:
width = p.get_width()
g4.text(width + 150,
p.get_y() + p.get_height()/1.5,
'{:1.2%}'.format(width/total),
ha="center")
del(top5, temp)
# timezone countplot in decreasing order
ax5 = axes[4]
g5 = sns.countplot(data=metadata, y="timezone", color=color, ax=ax5, orient="h", order = metadata['timezone'].value_counts().index)
ax5.title.set_text('Timezone')
ax5.set(ylabel="", xlabel="", xlim=(0,1600))
# adds percentage
total = float(len(metadata)) # number of buildings
for p in g5.patches:
width = p.get_width()
g5.text(width + 150,
p.get_y() + p.get_height()/1.5,
'{:1.2%}'.format(width/total),
ha="center")
# Meters type frequency
ax6 = axes[5]
# Temporal datafram
temp = pd.melt(metadata[["building_id","electricity","hotwater","chilledwater","steam","water","irrigation","gas","solar"]],id_vars = "building_id", var_name="meter")
# plot
g6 = sns.countplot(data=temp.loc[temp['value']=="Yes"], y='meter', color= color, ax=ax6, orient="h", order = temp.loc[temp['value']=="Yes"]["meter"].value_counts().index)
g6.title.set_text('Meter type frequency')
g6.set(ylabel="", xlabel="", xlim=(0,1600))
# adds percentage
total = temp.loc[temp['value']=="Yes"]["value"].value_counts()[0] # number of meters
for p in g6.patches:
width = p.get_width()
g6.text(width + 150,
p.get_y() + p.get_height()/1.5,
'{:1.2%}'.format(width/total),
ha="center")
del(temp)
# "sqft" histogram
ax7 = axes[6]
g7 = sns.distplot(metadata["sqm"], ax=ax7, color=color)
g7.set(ylabel="", xlabel="")
ax7.set_title('Building Area (square meters)')
# "yearbuilt" histogram
ax8 = axes[7]
g8 = sns.distplot(metadata["yearbuilt"].dropna(), ax=ax8, color=color)
g8.set(ylabel="", xlabel="")
ax8.set_title('Year built (50% missing values)')
# "occupants" histogram
ax9 = axes[8]
g9 = sns.distplot(metadata["occupants"].dropna(), ax=ax9, color=color)
g9.set(ylabel="", xlabel="")
ax9.set_title('Ocuppants (85% missing values)')
plt.tight_layout()
f.savefig("..\\figures\\metadata_features.pdf", bbox_inches='tight')
f.savefig("..\\figures\\metadata_features.png", bbox_inches='tight')
```
### Number of buildings in each site
```
metadata.groupby("site_id").building_id.count()
```
### Number of meters per site
```
temp = pd.melt(metadata[["site_id","electricity","hotwater","chilledwater","steam","water","irrigation","gas","solar"]],id_vars = "site_id", var_name="meter")
bysite = temp[temp.value == "Yes"].groupby(["site_id","meter"]).count().groupby("site_id").sum()
bysite
bysite.value.sum()
print("Total number of meters: " + str(len(temp.dropna())))
```
| github_jupyter |
```
from __future__ import division, print_function, absolute_import
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import matplotlib.gridspec as gridspec
import yash_bls
def doSearch(time, flux, minPeriod, maxPeriod, ):
SNR, period, epoch, duration, depth, transitModel, period_guesses, \
convolved_bls = yash_bls.do_bls_and_fit(time, flux, minPeriod, maxPeriod)
phase, phasedFlux = yash_bls.getPhase(time, flux, period, epoch)
phaseModel, phasedFluxModel = yash_bls.getPhase(time, transitModel, period, epoch)
secTime, secSNR, secPer, secEpoch, secDur, secModel = yash_bls.findSecondary(time, flux, period, epoch, duration)
if secSNR > 5 and abs(period - secPer) < 0.05:
secPhase, secPhaseModel = yash_bls.getPhase(secTime, secModel, secPer, epoch)
idx = len(secPhase[secPhase < 0])
else:
secPhase, secPhaseModel, idx = [], [], 1
# Odd/Even plot
fitT_odd, fitT_even = yash_bls.computeOddEvenModels(time, flux, period, epoch)
phaseModel_odd, phasedFluxModel_odd = yash_bls.getPhase(time, fitT_odd.transitmodel, period * 2, epoch)
phaseModel_even, phasedFluxModel_even = yash_bls.getPhase(time, fitT_even.transitmodel, period * 2, epoch + period)
depthOdd = fitT_odd.fitresultplanets['pnum0']['rprs'] ** 2
depthEven = fitT_even.fitresultplanets['pnum0']['rprs'] ** 2
phaseOdd, fluxOdd = yash_bls.getPhase(time, flux, period * 2, epoch)
phaseEven, fluxEven = yash_bls.getPhase(time, flux, period * 2, epoch + period)
x1, x2 = -duration, duration
y1, y2 = -3*np.std(fluxOdd), 3*np.std(fluxOdd)
if min(fluxOdd) < y1:
y1 = min(fluxOdd) - np.std(fluxOdd)
# sigma = abs(depth1 - depth2) / sqrt(u1^2 + u2^2)
durOdd = yash_bls.computeTransitDuration(period, fitT_odd.fitresultstellar['rho'], fitT_odd.fitresultplanets['pnum0']['rprs'])
durEven = yash_bls.computeTransitDuration(period, fitT_odd.fitresultstellar['rho'], fitT_even.fitresultplanets['pnum0']['rprs'])
sigma = yash_bls.computePointSigma(time, flux, transitModel, period, epoch, duration)
nOddPoints = np.sum((-durOdd*0.5 < phaseOdd) & (phaseOdd < durOdd * 0.5))
nEvenPoints = np.sum((-durEven*0.5 < phaseEven) & (phaseEven < durEven * 0.5))
uOdd, uEven = sigma / np.sqrt(nOddPoints), sigma / np.sqrt(nEvenPoints)
depthDiffSigma = abs(depthOdd - depthEven) / np.sqrt(uOdd**2 + uEven**2)
return locals()
def plotSearch(outs, secondary=False):
x1 = outs['x1']
x2 = outs['x2']
gs = gridspec.GridSpec(3,2)
ax1 = plt.subplot(gs[0,:])
axOdd = plt.subplot(gs[1,0])
axEven = plt.subplot(gs[1,1])
ax3 = plt.subplot(gs[2,:])
gs.update(wspace = 0, hspace = 0.5)
ax1.plot(outs['time'], outs['flux'], 'k')
y1, y2 = ax1.get_ylim()
ax1.vlines(np.arange(outs['epoch'], outs['time'][-1], outs['period']), y1, y2,
color = 'r', linestyles = 'dashed', linewidth = 0.5)
ax1.axis([outs['time'][0], outs['time'][-1], y1, y2])
ax1.set_title('kplr%s; best period = %8.6g days; SNR = %8.6g' %('K2', outs['period'], outs['SNR']))
ax1.set_xlabel('days')
axOdd.set_ylabel('flux')
axOdd.scatter(outs['phaseOdd'], outs['fluxOdd'], marker = '.', s = 1, color = 'k', alpha = 1)
axOdd.plot(outs['phaseModel_odd'], outs['phasedFluxModel_odd'], 'r')
axOdd.axhline(-outs['depthOdd'], x1, x2)
axOdd.axis([x1,x2,y1,y2])
axOdd.set_title('odd')
axEven.scatter(outs['phaseEven'], outs['fluxEven'], marker = '.', s = 1, color = 'k', alpha = 1)
axEven.plot(outs['phaseModel_even'], outs['phasedFluxModel_even'], 'r')
axEven.axhline(-outs['depthEven'], x1, x2)
axEven.yaxis.tick_right()
axEven.axis([x1,x2,y1,y2])
axEven.set_title('even')
if secondary:
plt.plot(outs['secPhase'][:idx], outs['secPhaseModel'][:idx], 'c')
plt.plot(outs['secPhase'][idx:], outs['secPhaseModel'][idx:], 'c')
ax3.scatter(outs['phase'], outs['phasedFlux'], marker = '.', s = 1, color = 'k')
ax3.plot(outs['phaseModel'], outs['phasedFluxModel'], 'r')
y1, y2 = -3*np.std(outs['phasedFlux']), 3*np.std(outs['phasedFlux'])
if min(outs['phasedFlux']) < y1:
y1 = min(outs['phasedFlux']) - np.std(outs['phasedFlux'])
ax3.axis([outs['phase'][0], outs['phase'][-1], y1, y2])
ax3.set_xlabel('phase [hours]')
ax3.text(0.5, 1.25, 'depth diff sigma = %.3f' %outs['depthDiffSigma'], horizontalalignment = 'center',
verticalalignment = 'center', transform = ax3.transAxes)
def get_qf(time,flux,epoch,period,transitmodel=None):
date1 = (time - epoch) + 0.5*period
phi1 = (((date1 / period) - np.floor(date1/period)) * period) - 0.5*period
q1 = np.sort(phi1)
f1 = (flux[np.argsort(phi1)]) * -1.E6
if transitmodel is not None:
m1 = (transitmodel[np.argsort(phi1)]) * -1.E6
return q1,f1,m1
else:
return q1,f1
filename = 'wasp47.csv'
time, flux = np.genfromtxt(filename, unpack = True, delimiter=',')
mask = np.isfinite(flux) * np.isfinite(time)
time1 = flux[mask] # !!!! I mixed up flux and time in my wasp47.csv file
flux1 = time[mask] # !!!!
time = time1
flux = flux1
flux = (flux / np.median(flux) ) -1.0
time, flux = yash_bls.outlierRemoval(time, flux)
flux = yash_bls.medianDetrend(flux, 26)
# Main transit search
minPeriod = 0.5 # Limitations of BLS Fortran code
maxPeriod = (time[-1] - time[0]) / 3.
outs = doSearch(time, flux, minPeriod, maxPeriod, )
fig, [ax1,ax2,ax3] = plt.subplots(3,1,figsize=[12,9])
ax1.plot(time, flux )
ax1.plot(time, outs['transitModel'] )
#ax2.plot(time, flux )
ax2.plot(outs['period_guesses'], outs['convolved_bls'])
q,f = get_qf(time,flux, outs['epoch'], outs['period'])
ax3.scatter(q,f, s=4)
ax3.set_ylim([np.max(f),np.min(f)])
ax3.set_xlim([np.max(q),np.min(q)])
fluxmin = flux - outs['transitModel']
#time, flux = yash_bls.outlierRemoval(time, fluxmin)
#fluxmin = yash_bls.medianDetrend(fluxmin, 26)
# Main transit search
minPeriod = 0.5 # Limitations of BLS Fortran code
maxPeriod = (time[-1] - time[0]) / 3.
outsmin = doSearch(time, fluxmin, minPeriod, maxPeriod, )
fig, [ax1,ax2,ax3] = plt.subplots(3,1,figsize=[12,9])
ax1.plot(time, fluxmin )
ax1.plot(time, outsmin['transitModel'] )
#ax2.plot(time, flux )
ax2.plot(outsmin['period_guesses'], outsmin['convolved_bls'])
q,f = get_qf(time,fluxmin, outsmin['epoch'], outsmin['period'])
ax3.scatter(q,f, s=4)
ax3.set_ylim([np.max(f),np.min(f)])
ax3.set_xlim([np.max(q),np.min(q)])
fluxmin2 = fluxmin - outsmin['transitModel']
#time, flux = yash_bls.outlierRemoval(time, fluxmin)
#fluxmin = yash_bls.medianDetrend(fluxmin, 26)
# Main transit search
minPeriod = 0.5 # Limitations of BLS Fortran code
maxPeriod = (time[-1] - time[0]) / 3.
outsmin2 = doSearch(time, fluxmin2, minPeriod, maxPeriod, )
fig, [ax1,ax2,ax3] = plt.subplots(3,1,figsize=[12,9])
ax1.plot(time, fluxmin2 )
ax1.plot(time, outsmin2['transitModel'] )
#ax2.plot(time, flux )
ax2.plot(outsmin2['period_guesses'], outsmin2['convolved_bls'])
q,f = get_qf(time,fluxmin2, outsmin2['epoch'], outsmin2['period'])
ax3.scatter(q,f, s=4)
ax3.set_ylim([np.max(f),np.min(f)])
ax3.set_xlim([np.max(q),np.min(q)])
print(outs['period'], outs['SNR'])
print(outsmin['period'], outsmin['SNR'])
print(outsmin2['period'], outsmin2['SNR'])
4.1591708630839328 / 0.78954682307784974
fluxmin3 = fluxmin2 - outsmin2['transitModel']
#time, flux = yash_bls.outlierRemoval(time, fluxmin)
#fluxmin = yash_bls.medianDetrend(fluxmin, 26)
# Main transit search
minPeriod = 0.5 # Limitations of BLS Fortran code
maxPeriod = (time[-1] - time[0]) / 3.
outsmin3 = doSearch(time, fluxmin3, minPeriod, maxPeriod, )
fig, [ax1,ax2,ax3] = plt.subplots(3,1,figsize=[12,9])
ax1.plot(time, fluxmin3 )
ax1.plot(time, outsmin3['transitModel'] )
#ax2.plot(time, flux )
ax2.plot(outsmin3['period_guesses'], outsmin3['convolved_bls'])
q,f = get_qf(time,fluxmin3, outsmin3['epoch'], outsmin3['period'])
ax3.scatter(q,f, s=4)
ax3.set_ylim([np.max(f),np.min(f)])
ax3.set_xlim([np.max(q),np.min(q)])
outs
plotSearch(outs)
fig = plt.gcf()
fig.set_figwidth(12)
fig.set_figheight(9)
```
| github_jupyter |
**[Course Home Page](https://www.kaggle.com/learn/machine-learning-for-insights)**
---
# Intro
One of the most basic questions we might ask of a model is *What features have the biggest impact on predictions?*
This concept is called *feature importance*. I've seen feature importance used effectively many times for every purpose in the list of use cases above.
There are multiple ways to measure feature importance. Some approaches answer subtly different versions of the question above. Other approaches have documented shortcomings.
In this lesson, we'll focus on *permutation importance*. Compared to most other approaches, permutation importance is:
- Fast to calculate
- Widely used and understood
- Consistent with properties we would want a feature importance measure to have
# How it Works
Permutation importance uses models differently than anything you've seen so far, and many people find it confusing at first. So we'll start with an example to make it more concrete.
Consider data with the following format:

We want to predict a person's height when they become 20 years old, using data that is available at age 10.
Our data includes useful features (*height at age 10*), features with little predictive power (*socks owned*), as well as some other features we won't focus on in this explanation.
**Permutation importance is calculated after a model has been fitted.** So we won't change the model or change what predictions we'd get for a given value of height, sock-count, etc.
Instead we will ask the following question: If I randomly shuffle a single column of the validation data, leaving the target and all other columns in place, how would that affect the accuracy of predictions in that now-shuffled data?

Randomly re-ordering a single column should cause less accurate predictions, since the resulting data no longer corresponds to anything observed in the real world. Model accuracy especially suffers if we shuffle a column that the model relied on heavily for predictions. In this case, shuffling `height at age 10` would cause terrible predictions. If we shuffled `socks owned` instead, the resulting predictions wouldn't suffer nearly as much.
With this insight, the process is as follows:
1. Get a trained model
2. Shuffle the values in a single column, make predictions using the resulting dataset. Use these predictions and the true target values to calculate how much the loss function suffered from shuffling. That performance deterioration measures the importance of the variable you just shuffled.
3. Return the data to the original order (undoing the shuffle from step 2.) Now repeat step 2 with the next column in the dataset, until you have calculated the importance of each column.
# Code Example
Our example will use a model that predicts whether a soccer/football team will have the "Man of the Game" winner based on the team's statistics. The "Man of the Game" award is given to the best player in the game. Model-building isn't our current focus, so the cell below loads the data and builds a rudimentary model.
```
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
data = pd.read_csv('../input/fifa-2018-match-statistics/FIFA 2018 Statistics.csv')
y = (data['Man of the Match'] == "Yes") # Convert from string "Yes"/"No" to binary
feature_names = [i for i in data.columns if data[i].dtype in [np.int64]]
X = data[feature_names]
train_X, val_X, train_y, val_y = train_test_split(X, y, random_state=1)
my_model = RandomForestClassifier(random_state=0).fit(train_X, train_y)
```
Here is how to calculate and show importances with the [eli5](https://eli5.readthedocs.io/en/latest/) library:
```
import eli5
from eli5.sklearn import PermutationImportance
perm = PermutationImportance(my_model, random_state=1).fit(val_X, val_y)
eli5.show_weights(perm, feature_names = val_X.columns.tolist())
```
# Interpreting Permutation Importances
The values towards the top are the most important features, and those towards the bottom matter least.
The first number in each row shows how much model performance decreased with a random shuffling (in this case, using "accuracy" as the performance metric).
Like most things in data science, there is some randomness to the exact performance change from a shuffling a column. We measure the amount of randomness in our permutation importance calculation by repeating the process with multiple shuffles. The number after the **±** measures how performance varied from one-reshuffling to the next.
You'll occasionally see negative values for permutation importances. In those cases, the predictions on the shuffled (or noisy) data happened to be more accurate than the real data. This happens when the feature didn't matter (should have had an importance close to 0), but random chance caused the predictions on shuffled data to be more accurate. This is more common with small datasets, like the one in this example, because there is more room for luck/chance.
In our example, the most important feature was **Goals scored**. That seems sensible. Soccer fans may have some intuition about whether the orderings of other variables are surprising or not.
# Your Turn
**[Get started here](https://www.kaggle.com/kernels/fork/1637562)** to flex your new permutation importance knowledge.
---
**[Course Home Page](https://www.kaggle.com/learn/machine-learning-for-insights)**
| github_jupyter |
```
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
import warnings
warnings.filterwarnings('ignore')
plt.rcParams['figure.figsize'] = [10, 5]
def calStability(raw, task_order, method):
values = {k:[] for k in method}
for t in task_order:
rows = raw[raw["task_order"]==t]
offline = rows[rows["method"]=="offline"]
for m in method:
target = rows[rows["method"]==m]
_m = target[target["task_index"]==1][["accuracy", "no_of_test", "no_of_correct_prediction"]]
if m=="offline":
continue
_ideal = offline[offline["task_index"]==1]["accuracy"]
val = float((_m["accuracy"]/float(_ideal)).sum()/len(_m))
values[m].append(val)
return values
def calPlasticity(raw, task_order, method):
values = {k:[] for k in method}
for t in task_order:
rows = raw[raw["task_order"]==t]
offline = rows[rows["method"]=="offline"]
for m in method:
_sum = 0.0
target = rows[rows["method"]==m]
train_session = target["train_session"].unique()
if m=="offline":
continue
_m = target[target["train_session"]==(target["task_index"])][["accuracy", "no_of_test", "no_of_correct_prediction"]]
_ideal = offline["accuracy"]
if len(_m) != len(_ideal):
values[m].append(np.nan)
continue
val = _m["accuracy"].div(_ideal.values, axis=0).sum()/len(_m)
# Ignore when val is much more than 4; due to deternimator is too large
if val > 4:
_m = list(_m["accuracy"])
_ideal = list(_ideal)
cnt = 0
acc = 0.0
for i in range(len(_m)):
if _m[i] / _ideal[i] > 4:
print("\t SKIP", "m=", _m[i] / _ideal[i], "v=", _ideal[i])
continue
cnt +=1
acc += _m[i] / _ideal[i]
val = acc
values[m].append(val)
return values
def calOverallAcc(raw, task_order, method):
values = {k:[] for k in method}
for t in task_order:
rows = raw[raw["task_order"]==t]
offline = rows[rows["method"]=="offline"]
for m in method:
if m=="offline":
continue
_sum = 0.0
target = rows[rows["method"]==m]
task_index = target["task_index"].unique()
train_session = target["train_session"].unique()
_m = target[target["train_session"]==len(task_index)][["accuracy", "no_of_test", "no_of_correct_prediction"]]
_ideal = offline["accuracy"]
if len(_m) != len(_ideal):
# print("SKIP", m, t)
values[m].append(np.nan)
continue
val = np.nansum(_m["accuracy"].div(_ideal.values, axis=0))/len(_m)
# Ignore when val is much more than 4; due to deternimator is too large
if val > 4:
_m = list(_m["accuracy"])
_ideal = list(_ideal)
cnt = 0
acc = 0.0
for i in range(len(_m)):
if _m[i] / _ideal[i] > 4:
print("\t SKIP", "m=", _m[i] / _ideal[i], "v=", _ideal[i])
continue
cnt +=1
acc += _m[i] / _ideal[i]
val = acc
values[m].append(val)
return values
all_values = {}
for d in ["HouseA", "CASAS", "PAMAP", "DSADS"]:
# for d in ["CASAS"]:
dataset = d
folder = "../../Results/"+dataset+"/exp_component_sensitivity/"
raw = pd.read_csv(folder+"results.txt")
raw.columns = [c.strip() for c in raw.columns]
raw["train_session"] = pd.to_numeric(raw["train_session"], errors='coerce')
cmd = raw["cmd"].unique()
task_order = raw["task_order"].unique()
method = raw["method"].unique()
stability = []
plasticity = []
overallAcc = []
for c in cmd:
print(c)
target = raw[raw["cmd"]==c]
m = calStability(target, task_order, method)
stability.append(m)
m = calPlasticity(target, task_order, method)
plasticity.append(m)
m = calOverallAcc(target, task_order, method)
overallAcc.append(m)
all_values[d] = (stability, plasticity, overallAcc)
print(d, "DONE")
```
# Plots
```
from scipy import stats
def plot(values, title, width=0.85, offset_ratio=0, xticks=[], models=None, rotation=-45, filename="plot.pdf"):
plt.rcParams['figure.figsize'] = [10, 7]
plt.rcParams.update({'font.size': 25})
plt.rcParams['axes.titlepad'] = 10
m = []
merr = []
if models is None:
models = ["mp-gan", "mp-wgan", "sg-cgan", "sg-cwgan"]
# values[cmd]["sg-cgan"][task_order]
for model in models:
tmp = []
tmperr = []
# for i, v in enumerate(values):
for i in [0, 1, 2, 3, 4, 6]:
v = values[i]
avg = np.nanmean(v[model])
err = stats.sem(v[model], nan_policy="omit")
tmp.append(avg)
tmperr.append(err)
m.append(tmp)
merr.append(tmperr)
# m[model_index][cmd]
print("Plot values")
print(m[0])
print(merr[0])
ind = np.arange(len(xticks)) # the x locations for the groups
fig, ax = plt.subplots()
patterns = [ "/" , "\\" , "x" , "-" , "+" , "|", "o", "O", ".", "*" ]
for i, model in enumerate(models):
offset = (float(i)/len(models))*width
offset -= (offset_ratio)*width
ax.bar(ind + offset, m[i], width*(1.0/len(models)), yerr=merr[i], label=model, hatch=patterns[i])
X = np.arange(-0.5, len(m[0])+0.5)
Y = [m[i][0] for _ in range(len(X))]
ax.plot(X, Y, linestyle=':')
ax.set_title(title)
ax.set_xticks(ind)
ax.set_xticklabels(xticks, rotation=rotation, rotation_mode="default", fontdict={"fontsize":20}, horizontalalignment='left')
# ax.legend()
fig.tight_layout()
fig.savefig(filename, bbox_inches='tight')
plt.show()
xticks = [
"None",
"Self-verify",
"SMOTE",
"EWC",
"LwF",
# "Instance Noise",
"All"
]
stability, plasticity, overallAcc = all_values["HouseA"]
models = ["sg-cgan"]
plot(stability, "Stability of the model", xticks=xticks, models=models, filename="comp_HouseA_sta.pdf")
plot(plasticity, "Plasticity of the model", xticks=xticks, models=models, filename="comp_HouseA_pla.pdf")
plot(overallAcc, "Overall accuracy of the model", xticks=xticks, models=models, filename="comp_HouseA_ova.pdf")
stability, plasticity, overallAcc = all_values["CASAS"]
models = ["sg-cgan"]
plot(stability, "Stability of the model", xticks=xticks, models=models, filename="comp_CASAS_sta.pdf")
plot(plasticity, "Plasticity of the model", xticks=xticks, models=models, filename="comp_CASAS_pla.pdf")
plot(overallAcc, "Overall accuracy of the model", xticks=xticks, models=models, filename="comp_CASAS_ova.pdf")
stability, plasticity, overallAcc = all_values["PAMAP"]
models = ["sg-cgan"]
plot(stability, "Stability of the model", xticks=xticks, models=models, filename="comp_PAMAP_sta.pdf")
plot(plasticity, "Plasticity of the model", xticks=xticks, models=models, filename="comp_PAMAP_pla.pdf")
plot(overallAcc, "Overall accuracy of the model", xticks=xticks, models=models, filename="comp_PAMAP_ova.pdf")
stability, plasticity, overallAcc = all_values["DSADS"]
models = ["sg-cgan"]
plot(stability, "Stability of the model", xticks=xticks, models=models, filename="comp_DSADS_sta.pdf")
plot(plasticity, "Plasticity of the model", xticks=xticks, models=models, filename="comp_DSADS_pla.pdf")
plot(overallAcc, "Overall accuracy of the model", xticks=xticks, models=models, filename="comp_DSADS_ova.pdf")
```
| github_jupyter |
# HW 03
1. The lowest and highest bins are filled. We expect clipping in the data.
2. Increase sampling frequency till spectra does not change anymore.
3. Here, we want the resolution $\Delta V$ to be 0.1% of the range (full scale: $FS = V_{max} - V_{min}$).
\begin{align}
\frac{\Delta V}{FS} & = \frac{1}{V_{max} - V_{min}} \cdot \frac{V_{max} - V_{min}}{2^{N}} \\
& = \frac{1}{2^{N}} \lt 0.001
\end{align}
This implies $N = 10$ bit, ($2^{10} = 1,024$).
4. The signal is:
\begin{align}
y(t) = 1.1 + 3.9 \sin(20,000 \pi t)
\end{align}
Its amplitude will be between 2.8 and 5 V, and the frequency is $f = 10,000$ Hz. We need to make sure the DAQ system has a high enough sampling rate to prevent aliasing and large enough input rate to prevent clipping. Then we will select the DAQ system with the best resolution (or the smallest quantization error).
(a): sampling rate is too low to satisfy Nyquiest criterion
(b): input range is too small, we would like to have a safety margin of 20-40 %.
(c) \& (d): sampling rate and input range are ok.
```
Q_c = 20/2**(8+1)
Q_d = 20/2**(16+1)
print('Q (c) = ', Q_c, ' V, Q (d) = ', Q_d, ' V')
```
Case (d) is the best design.
5.
\begin{align}
y(t) = 0.03 + 0.05 \cos (1,000 \pi t)
\end{align}
First, we select the gain: Voltage range: $A_{PT} = 0.02 - 0.08$ V. The smallest range of $\pm 0.01$ V is the best and has adequate safety margin to guaranty that there will be not clipping.
Then, we select the sampling rate: The signal is periodic and we will optimize the frequency resolution to make sure we do not have aliazing. The signal frequency is $f = 500$ Hz, to satisfy Nyquiest criterion, we would a sample rate $f_s > 2\cdot f > 1,000$ Hz. An appropriate sampling frequency would be in the range $f_s = [1,024 - 4,000]$ Hz.
6. (a) The ideal input range on the DAQ would be: $\pm 60-80$ mV. We will use an amplifier with an output voltage: $A_{amp}\pm 80$ mV for the analysis here.
The gain should be: $G = A_{amp} / A_{PT} = 80/8 = 10$
(b) Inverting amplifiers are less susceptible to electromagnetic noise, so we should build an amplifier stage based on inverting amplifiers. However, inverting amplifiers are susceptible to impedance loading, so it would be safe to use a buffer on the input of the amplifier stage.
7. (a) The desired bandwidth is $f = 100,000$ Hz. The amplifier gain bandwidth product is $GBP = 1 MHz$. Remembering the definition of $GBP$:
\begin{align*}
GBP & = G_{theoretical} \times f_c
\end{align*}
So for us the $f_c = f = 100,000$ kHz. This forces a theoretical gain per amplifier stage that should not exceed $G_{theoretical} = GBP / f_c = 10^6/10^5 = 10$.
So for a total desired gain of $G=100$, we should use at least two stages of $G_{stage} = G_{theoretical} = 10$ in series.
(b) You have two options to build your circuit:
1- Use two non-inverting amplifiers of gain $G=10$ in series. Non-inverting amplifiers have infinite input impedance and not susceptible to impedance loading.
2- Use two inverting amplifiers of gain $G=-10$ in series. Now you have have to worry about impedance loading, so you should put a buffer first, then the two inverting amplifiers.
8. For a Butterworth high-pass filter of order $n$ the gain is:
\begin{align*}
G = \frac{1}{\sqrt{1+\left( \frac{f_{cutoff}}{f} \right)^{2n} }}
\end{align*}
Here, $f$ is the noise: $f = 60$ Hz, the cutoff frequency is $f_{cutoff} = 300$ Hz.
Gain in decibel, assuming the gain, $G$, is defined as the ratio of amplitudes (like it is the case here):
\begin{align}
G_{dB} = 20 \log_{10} ( G ) = 20 \log_{10} \left( \frac{V_{out}}{V_{in}} \right)
\end{align}
```
import numpy
f = 60 # Hz
f_c = 300 # Hz
G_1 = 1/numpy.sqrt(1+(f_c/f)**(2*1))
G_2 = 1/numpy.sqrt(1+(f_c/f)**(2*2))
G_4 = 1/numpy.sqrt(1+(f_c/f)**(2*4))
print('the gains in percentage are:')
print("order 1: G = %2.4f " % (G_1*100), '%')
print("order 2: G = %2.4f " % (G_2*100), '%')
print("order 4: G = %2.4f " % (G_4*100), '%')
# convert to dB:
G_1dB = 20*numpy.log10(G_1)
G_2dB = 20*numpy.log10(G_2)
G_4dB = 20*numpy.log10(G_4)
print('the gains in dB are: ')
print("order 1: G = %2.4f " % (G_1dB), 'dB')
print("order 2: G = %2.4f " % (G_2dB), 'dB')
print("order 4: G = %2.4f " % (G_4dB), 'dB')
```
9. The signal could be written analytically as:
\begin{align}
y(t) = 1.50 \sin(40 \pi t) + 0.20 \sin(20,000 \pi t)
\end{align}
So the carrier frequency is $f_{carrier} = 20$ Hz, and the noise frequency is $f_{noise} = 10,000$ Hz. We acquire $N=2^{12}$ points. The DAQ has $n=16$ bits.
We also have $f_s = 30,000$ Hz.
(a) The frequency resolution is: $\Delta f = f_s/N$.
The quantization error is: $Q = (V_{max}-V_{min})/2^{n+1}$
```
f_s = 30000 # Hz
N = 2**12 # points
n = 16 # # bits
Q = 20/2**(n+1) # quantization error in V
print("\u0394 f = %4.4f" %(f_s/N), 'Hz')
print("Q = %1.4f" %(Q*1000), 'mV')
```
(b) This takes 3 steps.
_Step 1_ Select the type of filter and cutoff frequency, $f_c$: Here we want to remove high frequency noise, so we will use a low-pass filter. As a rule of thumb, it should be at least $10>f_{carrier}$ and $10<f_{noise}$. Here any values between: $f_c = [200-1,000]$ Hz is acceptable. I would select the lowest to more efficiently filter the noise: $f_c = 200$ Hz.
_Step 2_ Select the gain of the amplifier to make the most of the DAQ system: Ideally, the amplitude of the carrier should be 60-80% of the input range of the DAQ. So the gain should be in the range: $G = [6-8]/1.5 = 4-5.3$. Let's select: $G = 5$.
_Step 3_ Filter the signal: We wish for the noise amplitude to be smaller than the quantization error $Q$ of our DAQ system. Remember that the low pass filter has to be applied to the amplified signal. So after amplification, the noise has amplitude: $A_{noise} = 5\times 0.20 = 1$ V.
```
A_noise = 1 #V
f_c = 200 # Hz
f_noise = 10000 # Hz
# try LPF of order 1
n_LPF = 1
G = 1/numpy.sqrt(1+(f_noise/f_c)**(2*n_LPF))
print('order ',n_LPF,' : A_noise', (G*A_noise), 'V, Q/A_noise = ',Q/(G*A_noise))
# too big
# try LPF of order 2
n_LPF = 2
G = 1/numpy.sqrt(1+(f_noise/f_c)**(2*n_LPF))
print('order ',n_LPF,' : A_noise', (G*A_noise), 'V, Q/A_noise = ',Q/(G*A_noise))
# still too big
# try LPF of order 3
n_LPF = 3
G = 1/numpy.sqrt(1+(f_noise/f_c)**(2*n_LPF))
print('order ',n_LPF,' : A_noise', (G*A_noise), 'V, Q/A_noise = ',Q/(G*A_noise))
print('I need a filter of order 3')
```
| github_jupyter |
[](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/jupyter/transformers/HuggingFace%20in%20Spark%20NLP%20-%20BertForSequenceClassification.ipynb)
## Import BertForSequenceClassification models from HuggingFace 🤗 into Spark NLP 🚀
Let's keep in mind a few things before we start 😊
- This feature is only in `Spark NLP 3.3.2` and after. So please make sure you have upgraded to the latest Spark NLP release
- You can import BERT models trained/fine-tuned for token classification via `BertForSequenceClassification` or `TFBertForSequenceClassification`. These models are usually under `Token Classification` category and have `bert` in their labels
- Reference: [TFBertForSequenceClassification](https://huggingface.co/transformers/model_doc/bert.html#tfbertforsequenceclassification)
- Some [example models](https://huggingface.co/models?filter=bert&pipeline_tag=text-classification)
## Export and Save HuggingFace model
- Let's install `HuggingFace` and `TensorFlow`. You don't need `TensorFlow` to be installed for Spark NLP, however, we need it to load and save models from HuggingFace.
- We lock TensorFlow on `2.4.4` version and Transformers on `4.15.0`. This doesn't mean it won't work with the future releases, but we wanted you to know which versions have been tested successfully.
```
!pip install -q transformers==4.15.0 tensorflow==2.4.4
```
- HuggingFace comes with a native `saved_model` feature inside `save_pretrained` function for TensorFlow based models. We will use that to save it as TF `SavedModel`.
- We'll use [finiteautomata/beto-sentiment-analysis](https://huggingface.co/finiteautomata/beto-sentiment-analysis) model from HuggingFace as an example
- In addition to `TFBertForSequenceClassification` we also need to save the `BertTokenizer`. This is the same for every model, these are assets needed for tokenization inside Spark NLP.
```
from transformers import TFBertForSequenceClassification, BertTokenizer
MODEL_NAME = 'finiteautomata/beto-sentiment-analysis'
tokenizer = BertTokenizer.from_pretrained(MODEL_NAME)
tokenizer.save_pretrained('./{}_tokenizer/'.format(MODEL_NAME))
try:
model = TFBertForSequenceClassification.from_pretrained(MODEL_NAME)
except:
model = TFBertForSequenceClassification.from_pretrained(MODEL_NAME, from_pt=True)
model.save_pretrained("./{}".format(MODEL_NAME), saved_model=True)
```
Let's have a look inside these two directories and see what we are dealing with:
```
!ls -l {MODEL_NAME}
!ls -l {MODEL_NAME}/saved_model/1
!ls -l {MODEL_NAME}_tokenizer
```
- As you can see, we need the SavedModel from `saved_model/1/` path
- We also be needing `vocab.txt` from the tokenizer
- All we need is to just copy the `vocab.txt` to `saved_model/1/assets` which Spark NLP will look for
- In addition to vocabs, we also need `labels` and their `ids` which is saved inside the model's config. We will save this inside `labels.txt`
```
asset_path = '{}/saved_model/1/assets'.format(MODEL_NAME)
!cp {MODEL_NAME}_tokenizer/vocab.txt {asset_path}
# get label2id dictionary
labels = model.config.label2id
# sort the dictionary based on the id
labels = sorted(labels, key=labels.get)
with open(asset_path+'/labels.txt', 'w') as f:
f.write('\n'.join(labels))
```
Voila! We have our `vocab.txt` and `labels.txt` inside assets directory
```
!ls -l {MODEL_NAME}/saved_model/1/assets
```
## Import and Save BertForSequenceClassification in Spark NLP
- Let's install and setup Spark NLP in Google Colab
- This part is pretty easy via our simple script
```
! wget http://setup.johnsnowlabs.com/colab.sh -O - | bash
```
Let's start Spark with Spark NLP included via our simple `start()` function
```
import sparknlp
# let's start Spark with Spark NLP
spark = sparknlp.start()
```
- Let's use `loadSavedModel` functon in `BertForSequenceClassification` which allows us to load TensorFlow model in SavedModel format
- Most params can be set later when you are loading this model in `BertForSequenceClassification` in runtime like `setMaxSentenceLength`, so don't worry what you are setting them now
- `loadSavedModel` accepts two params, first is the path to the TF SavedModel. The second is the SparkSession that is `spark` variable we previously started via `sparknlp.start()`
- NOTE: `loadSavedModel` only accepts local paths and not distributed file systems such as `HDFS`, `S3`, `DBFS`, etc. That is why we use `write.save` so we can use `.load()` from any file systems
```
from sparknlp.annotator import *
from sparknlp.base import *
sequenceClassifier = BertForSequenceClassification.loadSavedModel(
'{}/saved_model/1'.format(MODEL_NAME),
spark
)\
.setInputCols(["document",'token'])\
.setOutputCol("class")\
.setCaseSensitive(True)\
.setMaxSentenceLength(128)
```
- Let's save it on disk so it is easier to be moved around and also be used later via `.load` function
```
sequenceClassifier.write().overwrite().save("./{}_spark_nlp".format(MODEL_NAME))
```
Let's clean up stuff we don't need anymore
```
!rm -rf {MODEL_NAME}_tokenizer {MODEL_NAME}
```
Awesome 😎 !
This is your BertForSequenceClassification model from HuggingFace 🤗 loaded and saved by Spark NLP 🚀
```
! ls -l {MODEL_NAME}_spark_nlp
```
Now let's see how we can use it on other machines, clusters, or any place you wish to use your new and shiny BertForSequenceClassification model 😊
```
sequenceClassifier_loaded = BertForSequenceClassification.load("./{}_spark_nlp".format(MODEL_NAME))\
.setInputCols(["document",'token'])\
.setOutputCol("class")
```
You can see what labels were used to train this model via `getClasses` function:
```
# .getClasses was introduced in spark-nlp==3.4.0
sequenceClassifier_loaded.getClasses()
```
This is how you can use your loaded classifier model in Spark NLP 🚀 pipeline:
```
document_assembler = DocumentAssembler() \
.setInputCol('text') \
.setOutputCol('document')
tokenizer = Tokenizer() \
.setInputCols(['document']) \
.setOutputCol('token')
pipeline = Pipeline(stages=[
document_assembler,
tokenizer,
sequenceClassifier_loaded
])
# couple of simple examples
example = spark.createDataFrame([["Te quiero. Te amo."]]).toDF("text")
result = pipeline.fit(example).transform(example)
# result is a DataFrame
result.select("text", "class.result").show()
```
That's it! You can now go wild and use hundreds of `BertForSequenceClassification` models from HuggingFace 🤗 in Spark NLP 🚀
```
```
| github_jupyter |
# Roku Python
```
import roku
roku.Roku?
cfg = {
"host": "192.168.1.128",
"port": 8060,
"timeout": 10, # E
}
r = roku.Roku(**cfg)
pbs_kids = next(a for a in r.apps if a.name=='PBS KIDS')
pbs_kids.launch()
r.home()
r.play()
r.back()
r.right()
r.select()
r.select()
r.pause=r.play
r.pause()
r.back()
for _ in range(10):
r.left()
r.down()
import time
class PBSKids(roku.Roku):
def __init__(self, **kwargs):
self.roku = roku.Roku(**kwargs)
self.pbs_kids = next(a for a in self.roku.apps if a.name=='PBS KIDS')
self.pbs_kids.launch()
time.sleep(5)
self.zero()
def zero(self):
for _ in range(10):
self.roku.left()
self.roku.down()
p = PBSKids(**cfg)
p.roku.back()
p.zero()
class PBSKids(roku.Roku):
def __init__(self, **kwargs):
self.roku = roku.Roku(**kwargs)
self.pbs_kids = next(a for a in self.roku.apps if a.name=='PBS KIDS')
self.pbs_kids.launch()
time.sleep(5)
self.zero()
def zero(self):
for _ in range(10):
self.roku.left()
self.roku.down()
def home(self):
p.roku.back()
self.zero()
p = PBSKids(**cfg)
self = p
channel_map = {
"live": [0, 1],
"odd_squad": [0,0],
"daniel_tiger": [1, 1],
"wild_kratts": [1, 0],
}
self.home()
self.roku.back()
self.zero()
channel_map = {
"live": [0, 1],
"odd_squad": [0,0],
"daniel_tiger": [1, 1],
"wild_kratts": [1, 0],
}
self.zero()
show = "wild_kratts"
for _ in range(channel_map[show][0]):
self.roku.right()
for _ in range(channel_map[show][1]):
self.roku.up()
self.roku.back()
self.zero()
show = "daniel_tiger"
for _ in range(channel_map[show][0]):
self.roku.right()
for _ in range(channel_map[show][1]):
self.roku.up()
time.sleep(0.1)
self.roku.select()
time.sleep(1)
self.roku.select()
p.roku.back()
class PBSKids(roku.Roku):
def __init__(self, **kwargs):
self.roku = roku.Roku(**kwargs)
self.pbs_kids = next(a for a in self.roku.apps if a.name=='PBS KIDS')
self.pbs_kids.launch()
time.sleep(5)
self.zero()
def zero(self):
for _ in range(10):
self.roku.left()
self.roku.down()
def home(self):
p.roku.back()
self.zero()
def sleep(self, sleep_time=0.1):
time.sleep(sleep_time)
# Practical Pressing
def back(self):
cmd = getattr(self.roku, "back")
cmd()
self.sleep()
def left(self):
self.roku.back()
self.sleep()
def left(self):
self.roku.back()
self.sleep()
def left(self):
self.roku.back()
self.sleep()
def left(self):
self.roku.back()
self.sleep()
p = PBSKids(**cfg)
self.roku.find_remote()
self.roku.input_tuner()
self.roku.select()
channel_map = {
"live": [0, 1],
"odd_squad": [0,0],
"daniel_tiger": [1, 1],
"wild_kratts": [1, 0],
}
class PBSKids(roku.Roku):
def __init__(self, **kwargs):
self.roku = roku.Roku(**kwargs)
self.pbs_kids = next(a for a in self.roku.apps if a.name=='PBS KIDS')
self.pbs_kids.launch()
time.sleep(5)
self.zero()
def zero(self):
for _ in range(10):
self.roku.left()
self.roku.down()
def live_home(self):
p.roku.back()
self.zero()
def home(self):
p.roku.back()
p.roku.back()
self.zero()
def sleep(self, sleep_time=0.1):
time.sleep(sleep_time)
def play_show(self, show):
for _ in range(channel_map[show][0]):
self.roku.right()
for _ in range(channel_map[show][1]):
self.roku.up()
# Practical Pressing
def back(self):
cmd = getattr(self.roku, "back")
cmd()
self.sleep()
def left(self):
self.roku.left()
self.sleep()
def right(self):
self.roku.right()
self.sleep()
def up(self):
self.roku.up()
self.sleep()
def down(self):
self.roku.down()
self.sleep()
def select(self):
self.roku.select()
self.sleep(1)
p = PBSKids(**cfg)
p.play_show("daniel_tiger")
p.roku.select()
p.roku.select()
p.live_home()
p.play_show("daniel_tiger")
p.roku.select()
se
p.roku.select()
```
| github_jupyter |
```
import sys
sys.path.append("../")
from __future__ import division
from pymacrospin import *
from pymacrospin import crystal, demag, energy, normalize
import numpy as np
import pandas as pd
from mpl_toolkits.mplot3d import Axes3D, proj3d
import matplotlib.pyplot as plt
from matplotlib import rcParams
rcParams['font.size'] = 16
```
# Cubic anisotropy #
Author: Colin Jermain
Following the [Crystal orientation](Crystal orientation.ipynb) discussion, the cubic anisotropy can be defined in the context of a given crystal surface. This notebook will illustrate the free energy of the cubic anisotropy in the $(110)$ surface of the yttrium iron garnet (YIG) material. The magnetic parameters for YIG are given in the [Yttrium iron garnet](Yttrium iron garnet.ipynb) notebook.
The origins of cubic anisotropy are in spin-orbit coupling [(O'Handley, 2000)](http://www.wiley.com/WileyCDA/WileyTitle/productCd-0471155667.html).
## Free energy description ##
The cubic anisotropy contributes to the free energy in terms of the square of the magnetization. This can be explained by time-reversal symmetry, under which the direction of magnetization ($\vec{M} = \vec{r} \times \vec{J}$) is expected to reverse [(Landau et al., 1960)](https://www.elsevier.com/books/electrodynamics-of-continuous-media/landau/978-0-7506-2634-7). The free energy should not reverse sign since this would make stable equilibriums into unstable energy maxima. The first two terms of the anisotropy part of the free energy take the form
$$F_c = \frac{K_{c1}}{M_s^4}(M_l^2 M_m^2 + M_m^2 M_n^2 + M_l^2 M_n^2) + \frac{K_{c2}}{M_s^6}(M_l^2 M_m^2 M_n^2)$$
in terms of the magnetization along the Miller indices $[lmn]$ of the crystal, saturation magnetization ($M_s$), and the first and second order energy coefficients ($K_{c1}$ and $K_{c2}$).
```
l,m,n = crystal.l, crystal.m, crystal.n
z = normalize(np.array([1,1,0]))
x = normalize(np.array([1,-1,1]))
y = np.cross(z, x)
T = np.concatenate((x, y, z)).reshape(3,3)
in_plane_indices = crystal.in_plane_indices(z, magnitude=1)
in_plane_normals = normalize(in_plane_indices)
in_plane_angles = crystal.angle_between_directions(x, z,
in_plane_normals, rotation='right')
# Sort the angles and corresponding indices
sorting_mask = in_plane_angles.argsort()
in_plane_angles = in_plane_angles[sorting_mask]
in_plane_indices = in_plane_indices[sorting_mask]
in_plane_normals = in_plane_normals[sorting_mask]
in_plane_labels = [crystal.miller_indices_display(i) for i in in_plane_indices]
angles = np.linspace(0, 360, num=100) # degrees
rotations = rotation_array(Rz, angles)
orientations = l.dot(rotations.dot(T))
plt.plot(angles, orientations.dot(l), label='$l$')
plt.plot(angles, orientations.dot(m), label='$m$')
plt.plot(angles, orientations.dot(n), label='$n$')
plt.xlim(0,360)
plt.ylabel("Magnitude")
plt.xlabel("Angle (degrees)")
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.twiny()
plt.xlabel("Angle $[lmn]$")
plt.xticks(in_plane_angles, in_plane_labels, rotation=45, verticalalignment='bottom')
plt.grid(which='major')
plt.xlim(0, 360)
plt.twinx()
plt.yticks([-1/np.sqrt(3), 1/np.sqrt(3)], alpha=0.0)
plt.ylim(-1, 1)
plt.grid(which='major')
plt.show()
```
| github_jupyter |
```
from pyspark.sql import SparkSession
import pyspark.sql.functions as f
spark = SparkSession.builder \
.master("local[*]")\
.config("spark.driver.memory", "4g") \
.config("spark.executor.memory", "1g") \
.getOrCreate()
import s2sphere
from pyspark.sql.types import StringType, FloatType
def cell_id(level: int, lat: int, lng: int) -> str:
return s2sphere.CellId.from_lat_lng(s2sphere.LatLng.from_degrees(lat, lng)).parent(level).to_token()
cell_id_udf = f.udf(cell_id, StringType())
def sphere_distance(token_from: str, token_to: str) -> float:
r = 6373.0
cell_from = s2sphere.CellId.from_token(token_from)
cell_to = s2sphere.CellId.from_token(token_to)
return cell_from.to_lat_lng().get_distance(cell_to.to_lat_lng()).radians * r
sphere_distance_udf = f.udf(sphere_distance, FloatType())
train_df = spark.read.option('header', 'true').csv('../data/raw/train.csv')\
.select('pickup_datetime', 'pickup_longitude', 'pickup_latitude', 'dropoff_longitude', 'dropoff_latitude')\
.withColumnRenamed('pickup_latitude', 'pickup_lat')\
.withColumn('pickup_lat', f.col('pickup_lat').cast('double'))\
.withColumnRenamed('pickup_longitude', 'pickup_lon')\
.withColumn('pickup_lon', f.col('pickup_lon').cast('double'))\
.withColumnRenamed('dropoff_latitude', 'dropoff_lat')\
.withColumn('dropoff_lat', f.col('dropoff_lat').cast('double'))\
.withColumnRenamed('dropoff_longitude', 'dropoff_lon')\
.withColumn('dropoff_lon', f.col('dropoff_lon').cast('double'))\
.withColumn('pickup_cell', cell_id_udf(f.lit(18), f.col('pickup_lat'), f.col('pickup_lon')))\
.withColumn('dropoff_cell', cell_id_udf(f.lit(18), f.col('dropoff_lat'), f.col('dropoff_lon')))
test_df = spark.read.option('header', 'true').csv('../data/raw/test.csv')\
.select('pickup_datetime', 'pickup_longitude', 'pickup_latitude', 'dropoff_longitude', 'dropoff_latitude')\
.withColumnRenamed('pickup_latitude', 'pickup_lat')\
.withColumn('pickup_lat', f.col('pickup_lat').cast('double'))\
.withColumnRenamed('pickup_longitude', 'pickup_lon')\
.withColumn('pickup_lon', f.col('pickup_lon').cast('double'))\
.withColumnRenamed('dropoff_latitude', 'dropoff_lat')\
.withColumn('dropoff_lat', f.col('dropoff_lat').cast('double'))\
.withColumnRenamed('dropoff_longitude', 'dropoff_lon')\
.withColumn('dropoff_lon', f.col('dropoff_lon').cast('double'))\
.withColumn('pickup_cell', cell_id_udf(f.lit(18), f.col('pickup_lat'), f.col('pickup_lon')))\
.withColumn('dropoff_cell', cell_id_udf(f.lit(18), f.col('dropoff_lat'), f.col('dropoff_lon')))
df = train_df.union(test_df)\
.select('pickup_cell', 'dropoff_cell')\
.dropDuplicates()\
.withColumn('distance', sphere_distance_udf(f.col('pickup_cell'), f.col('dropoff_cell')))
df.write.parquet('../data/processed/distance_matrix', mode='overwrite')
df.show(3)
df.printSchema()
df.count()
```
| github_jupyter |
# Check `GDS` Python stack
This notebook checks all software requirements for the course Geographic Data Science are correctly installed.
A successful run of the notebook implies no errors returned in any cell *and* every cell beyond the first one returning a printout of `True`. This ensures a correct environment installed.
```
! echo "You are running version $GDS_ENV_VERSION of the GDS env"
import black
import bokeh
import boto3
import bottleneck
import cenpy
import clustergram
import contextily
import cython
import dask
import dask_ml
import datashader
import flake8
import geocube
import geopandas
import geopy
import h3
import hdbscan
import pystac_client
import ipyleaflet
import ipympl
import ipywidgets
import legendgram
import lxml
import momepy
import nbdime
import netCDF4
import networkx
import osmnx
import palettable
import pandana
import polyline
import psycopg2
import pyarrow
import pygeos
import pyrosm
import pysal
import pysal.lib
import pysal.explore
import pysal.model
import pysal.viz
import rasterio
import rasterstats
import rio_cogeo
import rioxarray
import skimage
import sklearn
import skfusion
import seaborn
import spatialpandas
import sqlalchemy
import statsmodels
import tabulate
import urbanaccess
import xarray_leaflet
import xrspatial
import xlrd
import xlsxwriter
```
---
`pip` installs:
```
import dask_geopandas
import download
import geoalchemy2
import matplotlib_scalebar
import pygeoda
import pytest_cov
import pytest_tornasync
#Does not support Windows/Python3.9 for now#import simplification
import topojson
# This is broken at point of release because of:#https://github.com/urbangrammarai/graphics/issues/2#import urbangrammar_graphics
```
---
**Legacy checks** (in some ways superseded by those above but in some still useful)
```
import bokeh as bk
float(bk.__version__[:1]) >= 1
import matplotlib as mpl
float(mpl.__version__[:3]) >= 1.5
import seaborn as sns
float(sns.__version__[:3]) >= 0.6
import datashader as ds
float(ds.__version__[:3]) >= 0.6
import palettable as pltt
float(pltt.__version__[:3]) >= 3.1
sns.palplot(pltt.matplotlib.Viridis_10.hex_colors)
```
---
```
import pandas as pd
float(pd.__version__[:3]) >= 1
import dask
float(dask.__version__[:1]) >= 1
import sklearn
float(sklearn.__version__[:4]) >= 0.20
import statsmodels.api as sm
float(sm.__version__[2:4]) >= 10
```
---
```
import fiona
float(fiona.__version__[:3]) >= 1.8
import geopandas as gpd
float(gpd.__version__[:3]) >= 0.4
import pysal as ps
float(ps.__version__[:1]) >= 2
import rasterio as rio
float(rio.__version__[:1]) >= 1
```
# Test
```
from pysal import lib as libps
shp = libps.examples.get_path('columbus.shp')
db = geopandas.read_file(shp)
db.head()
db[['AREA', 'PERIMETER']].to_feather('db.feather')
tst = pd.read_feather('db.feather')
! rm db.feather
db.to_parquet('db.pq')
tst = gpd.read_parquet('db.pq')
! rm db.pq
import matplotlib.pyplot as plt
%matplotlib inline
f, ax = plt.subplots(1)
db.plot(facecolor='yellow', ax=ax)
ax.set_axis_off()
plt.show()
db.crs = 'EPSG:26918'
db_wgs84 = db.to_crs(epsg=4326)
db_wgs84.plot()
plt.show()
from pysal.viz import splot
from splot.mapping import vba_choropleth
f, ax = vba_choropleth(db['INC'], db['HOVAL'], db)
```
---
```
db.plot(column='INC', scheme='fisher_jenks', cmap=plt.matplotlib.cm.Blues)
plt.show()
city = osmnx.geocode_to_gdf('Berkeley, California, US')
osmnx.plot_footprints(osmnx.project_gdf(city));
from pyrosm import get_data
# Download data for the city of Helsinki
fp = get_data("Helsinki", directory="./")
print(fp)
# Get filepath to test PBF dataset
fp = pyrosm.get_data("test_pbf")
print("Filepath to test data:", fp)
# Initialize the OSM object
osm = pyrosm.OSM(fp)
# See the type
print("Type of 'osm' instance: ", type(osm))
from pyrosm import get_data
# Pyrosm comes with a couple of test datasets
# that can be used straight away without
# downloading anything
fp = get_data("test_pbf")
# Initialize the OSM parser object
osm = pyrosm.OSM(fp)
# Read all drivable roads
# =======================
drive_net = osm.get_network(network_type="driving")
drive_net.plot()
! rm Helsinki.osm.pbf
```
---
```
import numpy as np
import contextily as ctx
tl = ctx.providers.CartoDB.Positron
db = geopandas.read_file(ps.lib.examples.get_path('us48.shp'))
db.crs = "EPSG:4326"
dbp = db.to_crs(epsg=3857)
w, s, e, n = dbp.total_bounds
# Download raster
_ = ctx.bounds2raster(w, s, e, n, 'us.tif', source=tl)
# Load up and plot
source = rio.open('us.tif', 'r')
red = source.read(1)
green = source.read(2)
blue = source.read(3)
pix = np.dstack((red, green, blue))
bounds = (source.bounds.left, source.bounds.right, \
source.bounds.bottom, source.bounds.top)
f = plt.figure(figsize=(6, 6))
ax = plt.imshow(pix, extent=bounds)
ax = db.plot()
ctx.add_basemap(ax, crs=db.crs.to_string())
from ipyleaflet import Map, basemaps, basemap_to_tiles, SplitMapControl
m = Map(center=(42.6824, 365.581), zoom=5)
right_layer = basemap_to_tiles(basemaps.NASAGIBS.ModisTerraTrueColorCR, "2017-11-11")
left_layer = basemap_to_tiles(basemaps.NASAGIBS.ModisAquaBands721CR, "2017-11-11")
control = SplitMapControl(left_layer=left_layer, right_layer=right_layer)
m.add_control(control)
m
# Convert us.tiff to COG with rio-cogeo
! rio cogeo create us.tif us_cog.tif
! rio cogeo validate us_cog.tif
! rio info us_cog.tif
! rio warp --dst-crs EPSG:4326 us_cog.tif us_cog_ll.tif
# rioxarray + xarray_leaflet
import xarray
us = xarray.open_rasterio(
"us_cog.tif"
).sel(
band=1,
y=slice(7000000, 5000000),
x=slice(-10000000, -8000000)
)
import xarray_leaflet
from ipyleaflet import Map
m = Map(zoom=1, basemap=basemaps.CartoDB.DarkMatter)
m
l = us.astype(
float
).rio.reproject(
"EPSG:4326"
).leaflet.plot(m)
l.interact(opacity=(0.0,1.0))
! rm us.tif us_cog.tif us_cog_ll.tif
! rm -rf cache/
from IPython.display import GeoJSON
GeoJSON({
"type": "Feature",
"geometry": {
"type": "Point",
"coordinates": [-118.4563712, 34.0163116]
}
})
```
| github_jupyter |
## Tutorial Introduction
This tutorial will go through the Apache Spark basic and high level API focusing on Spark streaming. All the codes were run on Windows 7 OS.
Apache Spark is an open source fast and general-purpose cluster computing system, it was originally developed at the University of California, Berkeley's AMPLab, later it was open sourced on Apache Software Foundation and maintenaced by Apache till now. Spark provides high level interface for clusters programming with implicit and efficient data parallelism and fault-tolerance. _(To make this tutorial easy to review in Juypter notebook, I ran Spark on a multi core CPU machine in pseudo-distributed local mode, this mode is for development and testing purposes)_
Spark is claimed to be 100x faster than Hadoop MapReduce in memory, or 10x faster on disk. After some research, there are three advantages make Spark faster:
1. Unlike MapReduce persists each step's results on disk (e.g. Hadoop File System) and read by next step's computation as input, Spark can directly pass the result of previous step to next step, which saves lots of disk and network I/O. The advantage comes from the cheaper memory nowadays and the TB level memory addressing abilities of 64-bit platform.
2. Apache Spark has an advanced Directed Acyclic Graph (DAG) execution engine that supports cyclic data flow and in-memory computing. It can optimize many operations into one stage, where in MapReduce these operations are scheduled in multiple stages.
3. Apache Spark saves lots of Java Virtual Machine (JVM) setup time by keeping a running executor JVM on each cluster node, where in MapReduce a new JVM is created for each task.
## 0. Include Spark in Jupyter Notebook
Download pre-built Spark package: [Package Link](http://spark.apache.org/downloads.html).
Options selected while writing this tutorial:
- Spark release: 2.0.1 (Oct 03, 2016)
- Package type: Pre-built for Hadoop 2.7 and later
- Download type: direct download
Download `spark-2.0.1-bin-hadoop2.7.tgz` and unzip it to the same folder of this notebook file, then include Spark into the notebook as following:
```
import os
import sys
import random
import time
spark_home = os.getcwd()+'/spark-2.0.1-bin-hadoop2.7'
os.environ['SPARK_HOME'] = spark_home
sys.path.insert(0, os.path.join(spark_home, 'python'))
sys.path.insert(0, os.path.join(spark_home, 'python\lib\py4j-0.10.3-src.zip'))
```
To test if Spark is included into this notebook successfully, please try to build SparkContext. There should be no warning or exception.
```
from pyspark import SparkConf, SparkContext
from pyspark.streaming import StreamingContext
from operator import add
from pyspark.sql import SparkSession
conf = SparkConf()
# In local mode, specify the number of CPU cores Spark can use in bracket, or use * to let Spark to detect
conf.setMaster("local[*]")
conf.setAppName("Spark Tutorial")
# specify the memory Spark can use
conf.set("spark.executor.memory", "1g")
sc = SparkContext(conf = conf)
```
<b>Note1</b> :_ If exception _`[Java gateway process exited before sending the driver its port number]`_ is thrown on Windows OS, modify `spark-2.0.1-bin-hadoop2.7/bin/spark-class2.cmd` line 33, remove the double quotes
<b>Note2</b> :_ While writing the tutorial, I encountered _`[global name 'accumulators' is not defined]`_ exception from `context.py`, I added <code>`print(accumulators)`</code> in <code>`_do_init_`</code> function body before the problematic code, then the exception mysteriously disappeared...
<b>Note3</b> : In the cluster environment, you can pass the cluster URL to <code>conf.setMaster()</code>, like Spark, [Mesos](http://mesos.apache.org/) or [YARN](http://hadoop.apache.org/docs/stable/hadoop-yarn/hadoop-yarn-site/YARN.html) cluster URL.
<b>Note4</b> : Always call sc.stop() before generate another SparkContext.
```
# test if Spark is functioning, count the number words in LICENSE file
spark = SparkSession.builder.appName("Spark Tutorial").getOrCreate()
lines = spark.read.text(spark_home+'LICENSE').rdd.map(lambda r: r[0])
counts = lines.flatMap(lambda x: x.split(' ')).map(lambda x: 1).reduce(add)
print counts
```
The result should be 3453.
## 1. Spark Basic
### 1.1 Create Resilient Distributed Datasets (RDDs)
Spark's cluster programming API is centered on this RDD data structure, RDD is a read-only multiset of data items can be easily distributed amony cluster nodes, also provide an easy way for Spark to use the memory.
```
# create RDD from a list, (parallelizing an existing collection), then find the max value
test_list = [random.randint(1, sys.maxint) for i in range(10000)]
distData = sc.parallelize(test_list)
max_val = distData.reduce(lambda a, b: a if a>b else b)
print max_val == max(test_list)
# create RDD from existing data file
# Count number of appearance of each word in LICENSE file
test_file = sc.textFile(spark_home+'LICENSE') # reads it as a collection of lines
word_counts = test_file.flatMap(lambda x: x.split(' ')).map(lambda x: (x, 1)).reduceByKey(add)
output = word_counts.collect()
for (word, count) in output:
print("%s: %i" % (word, count))
```
First ten word-count pairs:
<pre>
: 1430
all: 3
customary: 1
(org.antlr:ST4:4.0.4: 1
managed: 1
Works,: 2
APPENDIX:: 1
details.: 2
granting: 1
Subcomponents:: 1
</pre>
### 1.2 Persist RDD Object in Memory
RDD object cannot be reused after reduce since RDD object is lazily created, i.e. only execute the RDD create operation when it is needed by reduce, and only the reduced result is returned, if the same RDD object is used later, RDD create operation needs to be executed again. If the generation of RDD object is time consuming, Spark can persist the RDD into memory for future uses
```
# make a larger file by repeating LICENSE 2000 times to make file reading time longer, result file size is around 34.5 MB
filenames = [spark_home+'LICENSE' for i in range(2000)]
with open(spark_home+'LARGE_LICENSE', 'w') as outfile:
for fname in filenames:
with open(fname) as infile:
outfile.write(infile.read())
line_len = sc.textFile(spark_home+'LARGE_LICENSE').map(lambda x: len(x))
line_len.persist() # persist to memory
%%time
# time with persisted line_len
max_len = line_len.reduce(lambda a,b : a if a>b else b)
min_len = line_len.reduce(lambda a,b : a if a<b else b)
total_len = line_len.reduce(add)
print max_len, min_len, total_len
line_len.unpersist() # remove persisted RDD from memory, compare the time for the same operations
%%time
# time without persisted line_len
max_len = line_len.reduce(lambda a,b : a if a>b else b)
min_len = line_len.reduce(lambda a,b : a if a<b else b)
total_len = line_len.reduce(add)
print max_len, min_len, total_len
```
### 1.3 Use Complex Function in Map and Reduce
We can pass a complex function to map and reduce function.
```
def complexMap(s):
'''
Get the individual words of current line
'''
words = s.strip().split(" ")
words_num = len(words)
# only counts line with more than ten words, and the first word starts with an alphabetic character
if words_num>10 and words[0][0].isalpha():
return (words[0], words_len)
else:
return (None, 0)
def complexReduce(a, b):
'''
conditional sum reduce
'''
if a[0] and b[0]:
return ("Total", a[1]+b[1])
if a[0]:
return ("Total", a[1])
if b[0]:
return ("Total", b[1])
return (None, 0)
sc.textFile(spark_home+'LICENSE').map(complexMap).reduce(complexReduce)
```
## 2. Spark Streaming
Spark Streaming is an extension of the core Spark API to process streaming data. Various streaming data sources can be applied like Kafka, Flume, Kinesis, or TCP sockets, Spark Streaming application checks newly arrived/created data with pre-defined time interval. Spark provides many high-level data processing method like <code>map</code>, <code>reduce</code>, <code>join</code>, <code>window</code> and their variants. At the end, processed data can be pushed out to filesystems, databases, and live dashboards.
More conveniently, Spark’s machine learning and graph processing algorithms can also be applied to data streams.
<img src="streaming-arch.png"/>
Spark Streaming provides a high-level abstraction on streaming data representation, called discretized stream (DStream). DStreams can be created either from input data streams from sources such as Kafka, Flume, and Kinesis, or from another DStreams. Internally, a DStream is represented as a sequence of RDDs.
<img src="streaming-dstream.png"/>
### 2.0 Spark Streaming Workflow
1. Spark Streaming context initialization
2. Input DStream creation (streaming data source)
3. Streaming data processing definition, i.e. applying transformation and output operations to DStreams
4. Start the application with <code>streamingContext.start()</code>
5. Wait for the finish of processing with <code>streamingContext.awaitTermination()</code>, the streaming application will be terminated when exception occurs or <code>streamingContext.stop()</code> is called.
### 2.1 Initialize Spark Streaming Context
```
ssc = StreamingContext(sc, 1) # time interval is defined as 1 second
```
<b>Note1: </b>Spark Streaming needs at least one node/thread as data receiver, and needs extra nodes/threads to do the processing job. In pseudo-distributed local mode, use `local` or `local[1]` as master URL will leave Spark no data processing thread since the only one thread acts as data receiver.
<b>Note2</b> : The batch interval should be set based on the latency requirements of the application and available cluster resources to make Spark streaming application stable, i.e. the application should be able to process data as fast as the streaming data being generated and received. More details on how to figure out the batch interal is on [Spark documentation](http://spark.apache.org/docs/latest/streaming-programming-guide.html#setting-the-right-batch-interval).
### 2.2 Create Input DStream
In cluster environment, we can create input DStream from socket, text file and binary file, each has a initialization function respectively. The text file and binary file are required to be hosted on Hadoop-compatible file system, everytime a new file is <b>moved</b> or <b>renamed</b> in the monitored directory, since Spark reads file data at once when new file is found by name, copied file or editing file will be seen as empty file by Spark.
Since the local file system is not Hadoop-compatible, and Sparking streaming is more stable on distributed file system, we use another way to generate the input DStream in this tutorial, <code>queueStream()</code>. <code>queueStream()</code> generates a data stream from a collection of RDDs, then feed the StreamingContext one RDD per time interval.
We use Twitter API to get a random sample of most recent tweets (around 1% of total new tweets), then combine a number tweets into a group, then each group is transformed into a distributed dataset that can be operated on in parallel, a seriers of such dataset will be formed into data stream using <code>queueStream()</code>.
```
# install twitter package into Jupyter notebook
!pip install twitter
import twitter
def connect_twitter():
'''
Connect to Twitter with developer API keys, then call Twitter API on TwitterStream
'''
consumer_key = "bm7qkiTCNPMzsBIkkSnwgnzVU"
consumer_secret = "fkID5ttsNogh4eQyWiKpgRg7P80yXbsglj9nAYA6peN4QGSNlX"
access_token = "794210841440686081-utUrhHReNXcUcD3KligzLb95MpyXv7c"
access_secret = "PDD7XWdAoJNYaMbwEzNHNfc1UueWNfepQIep4ABPoHHpq"
auth = twitter.OAuth(token = access_token, token_secret = access_secret, consumer_key = consumer_key, consumer_secret = consumer_secret)
return twitter.TwitterStream(auth=auth)
twitter_stream = connect_twitter()
def get_tweet(content_generator):
'''
Get valid Twitter content from a generator returned by Twitter sample API
'''
while True:
tweet = content_generator.next()
if 'delete' in tweet:
continue
return tweet['text'].encode('utf-8')
def gen_rdd_queue(twitter_stream, tweets_num=10, queue_len=10):
'''
Generate a RDD list out of the groups of tweets, this list will be transformed into data stream
'''
rddQueue = []
# Get most recent tweets samples
content_generator = twitter_stream.statuses.sample(block=True)
for q in range(queue_len):
contents = []
for i in range(tweets_num):
contents.append(get_tweet(content_generator))
# Generate the distributed dataset from a group of tweets content
rdd = ssc.sparkContext.parallelize(contents, 5)
rddQueue += [rdd]
return rddQueue
rddQueue = gen_rdd_queue(twitter_stream)
```
### 2.3 Stream Data Processing Definition
We will get the word with top occurences from the samples within every time interval, and print the result.
```
def process_tweet(new_values, last_sum):
'''
Word count update function
'''
return sum(new_values) + (last_sum or 0)
def output(time, rdd, top = 10):
'''
Print the words with top occurences
'''
result = []
read = rdd.take(top)
print("Time: %s:" % time)
for record in read:
print(record)
print("")
```
### 2.4 Start and Stop the Streaming application
```
ssc.checkpoint("./checkpoint-tweet")
# Get input DStream
lines = ssc.queueStream(rddQueue)
# Get words by split space of raw tweet text, the count word ocurrence and sort in descending order
counts = lines.flatMap(lambda line: line.split(" "))\
.map(lambda word: (word, 1))\
.updateStateByKey(process_tweet)\
.transform(lambda rdd: rdd.sortBy(lambda x: x[1],False))
# Print result on the result of each time interval
counts.foreachRDD(output)
ssc.start()
print "Spark Streaming started"
# To save time, the streaming application will be terminated manually after 30 seconds
time.sleep(30)
ssc.stop(stopSparkContext=False, stopGraceFully=True)
print "Spark Streaming finished"
```
Sample output:
Spark Streaming started<br/>
Time: 2016-11-03 18:19:05:<br/>
('RT', 8)<br/>
("j'ai", 2)<br/>
('de', 2)<br/>
('en', 2)<br/>
('que', 2)<br/>
('\xd8\x8c', 2)<br/>
('#MourinhoOut', 1)<br/>
(')', 1)<br/>
('somos', 1)<br/>
('m\xc3\xa1s', 1)<br/>
('#Gala9GH17', 1)<br/>
<br/>
Time: 2016-11-03 18:19:06:<br/>
('RT', 13)<br/>
('e', 4)<br/>
('\xd8\xa7\xd9\x84\xd9\x84\xd9\x87', 3)<br/>
('\xd9\x88\xd8\xa7\xd9\x84\xd9\x84\xd9\x87', 2)<br/>
('\xce\xba\xce\xb1\xce\xb9', 2)<br/>
('en', 2)<br/>
('\xd9\x85\xd9\x86', 2)<br/>
('de', 2)<br/>
("j'ai", 2)<br/>
('que', 2)<br/>
('\xd8\xb9\xd9\x86', 2)<br/>
<br/>
Time: 2016-11-03 18:19:07:<br/>
('RT', 18)<br/>
('de', 6)<br/>
('e', 5)<br/>
('to', 4)<br/>
('for', 4)<br/>
('no', 4)<br/>
('\xd8\xa7\xd9\x84\xd9\x84\xd9\x87', 3)<br/>
('it', 3)<br/>
('', 2)<br/>
('dos', 2)<br/>
('win', 2)<br/>
...
## 3. Further Resources
### 3.1 Spark DataFrames and SQL
Spark DataFrame is a distributed collection of data organized into named columns, more powerful lambda functions can be applied on. It is conceptually equivalent to a table in a relational database or a data frame in Python, but with more optimizations under the hood. Spark SQL provides ability to execute SQL queries on Spark data.
Reference Link: <http://spark.apache.org/docs/latest/sql-programming-guide.html>
### 3.2 Spark Machine Learning Library (MLlib)
MLlib is a rich library on common machine learning algorithms and methods, it makes the machine learning application more scalable and easier. These functionalities provided by MLlib can be applied on RDD easily.
Reference Link: <http://spark.apache.org/docs/latest/ml-guide.html>
## 4. References
1. Apache Spark Wikipedia: (https://en.wikipedia.org/wiki/Apache_Spark)
2. Spark Programming Guides: (http://spark.apache.org/docs/latest/programming-guide.html)
3. Why Spark is faster: (https://www.quora.com/What-makes-Spark-faster-than-MapReduce)
| github_jupyter |
Character issues, Unicode
```
s = 'cafe'
len(s)
b = s.encode('utf-8')
b
len(b)
b.decode('utf-8')
cafe = bytes('cafe', encoding='utf-8')
cafe
cafe[0]
cafe[2]
```
bytes: immutable
bytearray: mutable
```
bytes.fromhex('63616665')
b'cafe'.hex()
cafe_arr = bytearray(cafe)
cafe_arr
cafe_arr[-1:]
cafe_arr[1]
import array
numbers = array.array('h', [-2, -1, 0, 1, 2]) #short integers
octets = bytes(numbers)
octets
import struct
fmt = '<3s3sHH' # < little-endian, 3s3s 3 bytes, HH 16bit integers
with open('wizard.gif', 'rb') as fp:
img = memoryview(fp.read())
header = img[:10]
bytes(header)
struct.unpack(fmt, header)
del header, img
```
Basic encoders/decoders and errors include UnicodeEncodeError, UnicodeDecodeError, SyntaxError, UnicodeError
```
for codec in ['latin_1', 'utf_8', 'utf_16']:
print(codec, 'El Niño'.encode(codec), sep='\t')
city = 'São Paulo'
city.encode('utf_8')
city.encode('utf-16')
city.encode('cp437')
city.encode('cp437', errors='ignore')
city.encode('cp437', errors='replace')
octets = b'Montr\xe9al'
octets.decode('cp1252')
octets.decode('utf_8')
octets.decode('utf-8', errors='replace')
```
Handle text files
```
open('cafe.txt', 'w', encoding='utf_8').write('café')
open('cafe.txt').read()
fp = open('cafe.txt', 'w')
fp
fp.close()
fp.encoding
#Encoding defaults: a madhouse
#default_encodings.py
import sys
import locale
expressions = """
locale.getpreferredencoding()
my_file.encoding
sys.stdout.isatty()
sys.stdout.encoding
sys.stdin.isatty()
sys.stdin.encoding
sys.stderr.isatty()
sys.stderr.encoding
sys.getdefaultencoding()
sys.getfilesystemencoding()
"""
for expression in expressions.split():
value = eval(expression)
print(expression.rjust(30), '->', repr(value))
```
Normalizing Unicode
```
s1 = 'café'
s2 = 'cafe\u0301'
s1, s2
len(s1), len(s2)
s1 == s2
from unicodedata import normalize
len(normalize('NFC', s1)), len(normalize('NFC', s2))
len(normalize('NFD', s1)), len(normalize('NFD', s2))
normalize('NFC', s1) == normalize('NFC', s2)
normalize('NFD', s1) == normalize('NFD', s2)
from unicodedata import normalize, name
ohm = '\u2126'
name(ohm)
ohm_c = normalize('NFC', ohm)
name(ohm_c)
ohm == ohm_c
normalize('NFC', ohm) == normalize('NFC', ohm_c)
from unicodedata import normalize
def nfc_equal(str1, str2):
return normalize('NFC', str1) == normalize('NFC', str2)
def fold_equal(str1, str2):
return (normalize('NFC', str1).casefold() ==
normalize('NFC', str2).casefold())
s1 = 'café'
s2 = 'cafe\u0301'
nfc_equal(s1, s2)
#santize.py
import unicodedata
import string
def shave_marks(txt):
"""Remove all diacritic marks"""
norm_txt = unicodedata.normalize('NFD', txt)
shaved = ''.join(c for c in norm_txt if not unicodedata.combining(c))
return unicodedata.normalize('NFC', shaved)
def shave_marks_latin(txt):
"""Remove all diacritic marks from latin base characters"""
norm_txt = unicodedata.normlize('NFD', txt)
latin_base = False
keepers = []
for c in norm_txt:
if unicodedata.combining(c) and latin_base:
continue
keepers.append(c)
if not unicodedata.combining(c):
latin_base = c in string.ascii_lettters
shaved = ''.join(keepers)
return unicodedata.normalize('NFC', shaved)
single_map = str.maketrans("""‚ƒ„†ˆ‹‘’“”•–—˜›""",
"""'f"*^<''""---~>""")
multi_map = str.maketrans({
'€': '<euro>',
'…': '...',
'OE': 'OE',
'™': '(TM)',
'oe': 'oe',
'‰': '<per mille>',
'‡': '**',
})
multi_map.update(single_map)
def dewinize(txt):
"""Replace Win1252 symbols with ASCII chars or sequences"""
return txt.translate(multi_map)
def asciize(txt):
no_marks = shave_marks_latin(dewinize(txt))
no_marks = no_marks.replace('ß', 'ss')
return unicodedata.normalize('NFKC', no_marks)
order = '“Herr Voß: • 1⁄2 cup of ŒtkerTM caffè latte • bowl of açaí.”'
shave_marks(order)
fruits = ['caju', 'atemoia', 'cajá', 'açaí', 'acerola']
sorted(fruits)
```
| github_jupyter |
```
%matplotlib inline
```
GroupLasso for linear regression with dummy variables
=====================================================
A sample script for group lasso with dummy variables
Setup
-----
```
import matplotlib.pyplot as plt
import numpy as np
from sklearn.linear_model import Ridge
from sklearn.metrics import r2_score
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import OneHotEncoder
from group_lasso import GroupLasso
from group_lasso.utils import extract_ohe_groups
np.random.seed(42)
GroupLasso.LOG_LOSSES = True
```
Set dataset parameters
----------------------
```
num_categories = 30
min_options = 2
max_options = 10
num_datapoints = 10000
noise_std = 1
```
Generate data matrix
--------------------
```
X_cat = np.empty((num_datapoints, num_categories))
for i in range(num_categories):
X_cat[:, i] = np.random.randint(min_options, max_options, num_datapoints)
ohe = OneHotEncoder()
X = ohe.fit_transform(X_cat)
groups = extract_ohe_groups(ohe)
group_sizes = [np.sum(groups == g) for g in np.unique(groups)]
active_groups = [np.random.randint(0, 2) for _ in np.unique(groups)]
```
Generate coefficients
---------------------
```
w = np.concatenate(
[
np.random.standard_normal(group_size) * is_active
for group_size, is_active in zip(group_sizes, active_groups)
]
)
w = w.reshape(-1, 1)
true_coefficient_mask = w != 0
intercept = 2
```
Generate regression targets
---------------------------
```
y_true = X @ w + intercept
y = y_true + np.random.randn(*y_true.shape) * noise_std
```
View noisy data and compute maximum R^2
---------------------------------------
```
plt.figure()
plt.plot(y, y_true, ".")
plt.xlabel("Noisy targets")
plt.ylabel("Noise-free targets")
# Use noisy y as true because that is what we would have access
# to in a real-life setting.
R2_best = r2_score(y, y_true)
```
Generate pipeline and train it
------------------------------
```
pipe = pipe = Pipeline(
memory=None,
steps=[
(
"variable_selection",
GroupLasso(
groups=groups,
group_reg=0.1,
l1_reg=0,
scale_reg=None,
supress_warning=True,
n_iter=100000,
frobenius_lipschitz=False,
),
),
("regressor", Ridge(alpha=1)),
],
)
pipe.fit(X, y)
```
Extract results and compute performance metrics
-----------------------------------------------
```
# Extract from pipeline
yhat = pipe.predict(X)
sparsity_mask = pipe["variable_selection"].sparsity_mask_
coef = pipe["regressor"].coef_.T
# Construct full coefficient vector
w_hat = np.zeros_like(w)
w_hat[sparsity_mask] = coef
R2 = r2_score(y, yhat)
# Print performance metrics
print(f"Number variables: {len(sparsity_mask)}")
print(f"Number of chosen variables: {sparsity_mask.sum()}")
print(f"R^2: {R2}, best possible R^2 = {R2_best}")
```
Visualise regression coefficients
---------------------------------
```
for i in range(w.shape[1]):
plt.figure()
plt.plot(w[:, i], ".", label="True weights")
plt.plot(w_hat[:, i], ".", label="Estimated weights")
plt.figure()
plt.plot([w.min(), w.max()], [coef.min(), coef.max()], "gray")
plt.scatter(w, w_hat, s=10)
plt.ylabel("Learned coefficients")
plt.xlabel("True coefficients")
plt.show()
```
| github_jupyter |
## callbacks.misc
Miscellaneous callbacks that don't belong to any specific group are to be found here.
```
from fastai.gen_doc.nbdoc import *
from fastai.callbacks.misc import *
show_doc(StopAfterNBatches)
```
[`StopAfterNBatches`](/callbacks.misc.html#StopAfterNBatches)
There could be various uses for this handy callback.
The initial purpose of it was to be able to quickly check memory requirements for a given set of hyperparamaters like `bs` and `size`.
Since all the required GPU memory is setup during the first batch of the first epoch [see tutorial](https://docs.fast.ai/tutorial.resources.html#gpu-memory-usage-anatomy), it's enough to run just 1-2 batches to measure whether your hyperparameters are right and won't lead to Out-Of-Memory (OOM) errors. So instead of waiting for minutes or hours to just discover that your `bs` or `size` are too large, this callback allows you to do it seconds.
You can deploy it on a specific learner (or fit call) just like with any other callback:
```
from fastai.callbacks.misc import StopAfterNBatches
[...]
learn = cnn_learner([...])
learn.callbacks.append(StopAfterNBatches(n_batches=2))
learn.fit_one_cycle(3, max_lr=1e-2)
```
and it'll either fit into the existing memory or it'll immediately fail with OOM error. You may want to add [ipyexperiments](https://github.com/stas00/ipyexperiments/) to show you the memory usage, including the peak usage.
This is good, but it's cumbersome since you have to change the notebook source code and often you will have multiple learners and fit calls in the same notebook, so here is how to do it globally by placing the following code somewhere on top of your notebook and leaving the rest of your notebook unmodified:
```
from fastai.callbacks.misc import StopAfterNBatches
# True turns the speedup on, False return to normal behavior
tune = True
#tune = False
if tune:
defaults.silent = True # don't print the fit metric headers
defaults.extra_callbacks = [StopAfterNBatches(n_batches=2)]
else:
defaults.silent = False
defaults.extra_callbacks = None
```
When you're done tuning your hyper-parameters, just set `tune` to `False` and re-run the notebook to do true fitting.
The setting `defaults.silent` controls whether [`fit`](/basic_train.html#fit) calls print out any output.
Do note that when you run this callback, each fit call will be interrupted resulting in the red colored output - that's just an indication that the normal fit didn't happen, so you shouldn't expect any qualitative results out of it.
## Undocumented Methods - Methods moved below this line will intentionally be hidden
| github_jupyter |
```
# imports
import sympy as sm
import numpy as np
import math
import matplotlib.pyplot as plt
%matplotlib inline
```
In order to recast the Huber loss as a valid likelihood function to be used in likelihood ratio tests, we need to derive the associated normalisation constant. We can achieve this using sympy, a symbolic computation package.
Using the same notation as in the PyTorchDIA paper, we can start by defining the class of distributions,
$f(x) = Q \times \frac{1}{\sigma} \times \exp({-\rho(x)})$.
$Q$ is the normalisation constant we're interested in deriving. In order to find $Q$, we need to integrate everything to its right over the entire range of $x$. Setting $Q$ equal to the reciprocal of the result of this integration will ensure that when $f(x)$ is integrated over the entire range of $x$ the result will always be $1$ i.e. $f(x)$ is now a valid probability distribution.
We'll start with the Gaussian case, where
$\rho(x) = \frac{1}{2}\frac{x^2}{\sigma^2}\;.$
N.B. Under the above definition for $f(x)$, we already have the $\sigma$ term that turns up in the usual derivation of the normalisation constant for a Gaussian, so integrating this expression should give us $\sqrt{2\pi}$. Let's use sympy to do this for us.
```
# first, test this out for the gaussian case
x, sigma = sm.symbols("x, sigma", real=True, positive=True)
rho = ((x / sigma) ** 2 )/ 2
loss = sm.exp(-rho) / sigma
norm = 2*sm.integrate(loss, (x, 0, sm.oo))
sm.simplify(norm)
```
OK great, that seems to work! Now let's do the same for the Huber loss.
\begin{equation}
\rho_{\text{Huber}, c}(x) =
\begin{cases}
\frac{1}{2}\frac{x^2}{\sigma^2}, & \text{for } |\frac{x}{\sigma}|\leq c \\
c(|\frac{x}{\sigma}| - \frac{1}{2}c), & \text{for } |\frac{x}{\sigma}| > c \;.
\end{cases}
\label{eq:huber_loss}
\end{equation}
I don't expect the result will be quite as tidy as above!
```
# normalisation constant for Huber 'likelihood'
x, c, sigma = sm.symbols("x, c, sigma", real=True, positive=True)
rho = sm.Piecewise(
((x / sigma) ** 2 / 2, (x / sigma) <= c),
(c * (sm.Abs(x / sigma) - c / 2), ((x / sigma) > c))
)
loss = sm.exp(-rho) / sigma
norm = 2 * sm.integrate(loss, (x, 0, sm.oo))
sm.simplify(norm)
#const = (np.sqrt(2 * np.pi) * math.erf(c / np.sqrt(2))) + ((2 / c) * np.exp(-0.5 * c**2))
```
OK, on to the meat of this notebook. I'm not certain that it makes sense to compare a Huber likelihood against a Gaussian, even for instances in which there are no outlying data points. We can probe this question with a toy example; fitting a straight line to data with known Gaussian uncertainties.
It may be that the Huber likelihood (evaluated at the MLE) is strongly dependent on the tuning paramter, $c$. In the case where $c$ tends to infinity, we should expect the same value for the likelihood as for the Gaussian case. In this special case, not only are all residuals from the model treated as 'inliers', but note too that the normalisation constant we found above would also tend to $\sqrt{2\pi}$; the error function in the first term becomes unity and the latter term becomes neglibily small. However, for useful, smaller values of $c$ e.g. 1.345, in which 'outlying' residuals are treated linearly rather than quadratically, should we expect to get roughly the same values for the likelihoods? In other words, does the linear treatment of outliers balance with the change to the normalisation constant, which is itself dependent on $c$. I don't see how it could... but's let's verify this numerically.
```
## Generate some mock data
# The linear model with slope 2 and intercept 1:
true_params = [2.0, 1.0]
# points drawn from true model
np.random.seed(42)
x = np.sort(np.random.uniform(-2, 2, 30))
yerr = 0.4 * np.ones_like(x)
y = true_params[0] * x + true_params[1] + yerr * np.random.randn(len(x))
# true line
x0 = np.linspace(-2.1, 2.1, 200)
y0 = np.dot(np.vander(x0, 2), true_params)
# plot
plt.errorbar(x, y, yerr=yerr, fmt=",k", ms=0, capsize=0, lw=1, zorder=999)
plt.scatter(x, y, marker="s", s=22, c="k", zorder=1000)
plt.plot(x0, y0, c='k')
# analytic solution - MLE for a priori known Gaussian noise
def linear_regression(x, y, yerr):
A = np.vander(x, 2)
result = np.linalg.solve(np.dot(A.T, A / yerr[:, None]**2), np.dot(A.T, y / yerr**2))
return result
res = linear_regression(x, y, yerr)
m, b = res
# optimise Huber loss
import torch
# initialise model parameters (y = m*x + b)
m_robust = torch.nn.Parameter(1e-3*torch.ones(1), requires_grad = True)
b_robust = torch.nn.Parameter(1e-3*torch.ones(1), requires_grad = True)
params_robust = list([m_robust, b_robust])
# negative log-likelihood for Huber likelihood (excluding the irrelevant normalisation constant)
def nll_Huber(model, data, yerr, c):
# ln_sigma is same as above
ln_sigma = torch.log(yerr).sum()
# define inliers and outliers with a threshold, c
resid = torch.abs((model - data)/yerr)
cond1 = resid <= c
cond2 = resid > c
inliers = ((model - data)/yerr)[cond1]
outliers = ((model - data)/yerr)[cond2]
# Huber loss can be thought of as a hybrid of l2 and l1 loss
# apply l2 (i.e. normal) loss to inliers, and l1 to outliers
l2 = 0.5*torch.pow(inliers, 2).sum()
l1 = (c * torch.abs(outliers) - (0.5 * c**2)).sum()
nll = ln_sigma + l2 + l1
return nll
# pass paramterers to optimizer
optimizer_robust = torch.optim.Adam(params_robust, lr = 0.01)
# tuning parameter, c
c = 1.345
xt, yt, yerrt = torch.from_numpy(x), torch.from_numpy(y), torch.from_numpy(yerr)
for epoch in range(2000):
model = m_robust*xt + b_robust
# negative loglikelihood
loss = nll_Huber(model, yt, yerrt, c)
optimizer_robust.zero_grad()
loss.backward()
optimizer_robust.step()
if np.mod(epoch, 100) == 0:
# You can see the alpha+scale parameters moving around.
print('{:<4}: loss={:03f} m={:03f} b={:03f}'.format(
epoch, loss.item(), m_robust.item(), b_robust.item()))
m_r, b_r = m_robust.item(), b_robust.item()
def evaluate_gaussian_log_likelihood(data, model, var):
print('\nGaussian log-likelihood')
chi2 = 0.5 * ((data - model)**2 / var).sum()
lnsigma = np.log(np.sqrt(var)).sum()
norm_constant = len(data.flatten()) * 0.5 * np.log(2 * np.pi)
print('chi2, lnsigma, norm_constant:', chi2, lnsigma, norm_constant)
return -(chi2 + lnsigma + norm_constant)
def evaluate_huber_log_likelihood(data, model, var, c):
print('\nHuber log-likelihood')
## PyTorchDIA - 'Huber' likelihood
sigma = np.sqrt(var)
ln_sigma = np.log(sigma).sum()
# gaussian when (model - targ)/sigma <= c
# absolute deviation when (model - targ)/sigma > c
cond1 = np.abs((model - data)/sigma) <= c
cond2 = np.abs((model - data)/sigma) > c
inliers = ((model - data)/sigma)[cond1]
outliers = ((model - data)/sigma)[cond2]
l2 = 0.5*(np.power(inliers, 2)).sum()
l1 = (c *(np.abs(outliers)) - (0.5 * c**2)).sum()
constant = (np.sqrt(2 * np.pi) * math.erf(c / np.sqrt(2))) + ((2 / c) * np.exp(-0.5 * c**2))
norm_constant = len(data.flatten()) * np.log(constant)
ll = -(l2 + l1 + ln_sigma + norm_constant)
print('l2, l1, ln_sigma, norm_constant:', l2, l1, ln_sigma, norm_constant)
return ll
# MLE models
model = m*x + b # Gaussian
model_r = m_r*x + b_r # Huber
ll_gaussian = evaluate_gaussian_log_likelihood(x, model, yerr**2)
print(ll_gaussian)
ll_huber = evaluate_huber_log_likelihood(x, model_r, yerr**2, c=c)
print(ll_huber)
```
Unless I've blundered, it seems like the Huber log-likelihood is always going to exceed the Gaussian due to how the numerics are treated. If this is indeed correct, then any comparison between the two would be meaningless.
| github_jupyter |
# 2 - Updated Sentiment Analysis
In the previous notebook, we got the fundamentals down for sentiment analysis. In this notebook, we'll actually get decent results.
We will use:
- packed padded sequences
- pre-trained word embeddings
- different RNN architecture
- bidirectional RNN
- multi-layer RNN
- regularization
- a different optimizer
This will allow us to achieve ~84% test accuracy.
## Preparing Data
As before, we'll set the seed, define the `Fields` and get the train/valid/test splits.
We'll be using *packed padded sequences*, which will make our RNN only process the non-padded elements of our sequence, and for any padded element the `output` will be a zero tensor. To use packed padded sequences, we have to tell the RNN how long the actual sequences are. We do this by setting `include_lengths = True` for our `TEXT` field. This will cause `batch.text` to now be a tuple with the first element being our sentence (a numericalized tensor that has been padded) and the second element being the actual lengths of our sentences.
```
import torch
from torchtext import data
from torchtext import datasets
SEED = 1234
torch.manual_seed(SEED)
torch.backends.cudnn.deterministic = True
TEXT = data.Field(tokenize = 'spacy',
tokenizer_language = 'en_core_web_sm',
include_lengths = True)
LABEL = data.LabelField(dtype = torch.float)
```
We then load the IMDb dataset.
```
from torchtext import datasets
train_data, test_data = datasets.IMDB.splits(TEXT, LABEL)
```
Then create the validation set from our training set.
```
import random
train_data, valid_data = train_data.split(random_state = random.seed(SEED))
```
Next is the use of pre-trained word embeddings. Now, instead of having our word embeddings initialized randomly, they are initialized with these pre-trained vectors.
We get these vectors simply by specifying which vectors we want and passing it as an argument to `build_vocab`. `TorchText` handles downloading the vectors and associating them with the correct words in our vocabulary.
Here, we'll be using the `"glove.6B.100d" vectors"`. `glove` is the algorithm used to calculate the vectors, go [here](https://nlp.stanford.edu/projects/glove/) for more. `6B` indicates these vectors were trained on 6 billion tokens and `100d` indicates these vectors are 100-dimensional.
You can see the other available vectors [here](https://github.com/pytorch/text/blob/master/torchtext/vocab.py#L113).
The theory is that these pre-trained vectors already have words with similar semantic meaning close together in vector space, e.g. "terrible", "awful", "dreadful" are nearby. This gives our embedding layer a good initialization as it does not have to learn these relations from scratch.
**Note**: these vectors are about 862MB, so watch out if you have a limited internet connection.
By default, TorchText will initialize words in your vocabulary but not in your pre-trained embeddings to zero. We don't want this, and instead initialize them randomly by setting `unk_init` to `torch.Tensor.normal_`. This will now initialize those words via a Gaussian distribution.
```
MAX_VOCAB_SIZE = 25_000
TEXT.build_vocab(train_data,
max_size = MAX_VOCAB_SIZE,
vectors = "glove.6B.100d",
unk_init = torch.Tensor.normal_)
LABEL.build_vocab(train_data)
```
As before, we create the iterators, placing the tensors on the GPU if one is available.
Another thing for packed padded sequences all of the tensors within a batch need to be sorted by their lengths. This is handled in the iterator by setting `sort_within_batch = True`.
```
BATCH_SIZE = 64
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
train_iterator, valid_iterator, test_iterator = data.BucketIterator.splits(
(train_data, valid_data, test_data),
batch_size = BATCH_SIZE,
sort_within_batch = True,
device = device)
```
## Build the Model
The model features the most drastic changes.
### Different RNN Architecture
We'll be using a different RNN architecture called a Long Short-Term Memory (LSTM). Why is an LSTM better than a standard RNN? Standard RNNs suffer from the [vanishing gradient problem](https://en.wikipedia.org/wiki/Vanishing_gradient_problem). LSTMs overcome this by having an extra recurrent state called a _cell_, $c$ - which can be thought of as the "memory" of the LSTM - and the use use multiple _gates_ which control the flow of information into and out of the memory. For more information, go [here](https://colah.github.io/posts/2015-08-Understanding-LSTMs/). We can simply think of the LSTM as a function of $x_t$, $h_t$ and $c_t$, instead of just $x_t$ and $h_t$.
$$(h_t, c_t) = \text{LSTM}(x_t, h_t, c_t)$$
Thus, the model using an LSTM looks something like (with the embedding layers omitted):

The initial cell state, $c_0$, like the initial hidden state is initialized to a tensor of all zeros. The sentiment prediction is still, however, only made using the final hidden state, not the final cell state, i.e. $\hat{y}=f(h_T)$.
### Bidirectional RNN
The concept behind a bidirectional RNN is simple. As well as having an RNN processing the words in the sentence from the first to the last (a forward RNN), we have a second RNN processing the words in the sentence from the **last to the first** (a backward RNN). At time step $t$, the forward RNN is processing word $x_t$, and the backward RNN is processing word $x_{T-t+1}$.
In PyTorch, the hidden state (and cell state) tensors returned by the forward and backward RNNs are stacked on top of each other in a single tensor.
We make our sentiment prediction using a concatenation of the last hidden state from the forward RNN (obtained from final word of the sentence), $h_T^\rightarrow$, and the last hidden state from the backward RNN (obtained from the first word of the sentence), $h_T^\leftarrow$, i.e. $\hat{y}=f(h_T^\rightarrow, h_T^\leftarrow)$
The image below shows a bi-directional RNN, with the forward RNN in orange, the backward RNN in green and the linear layer in silver.

### Multi-layer RNN
Multi-layer RNNs (also called *deep RNNs*) are another simple concept. The idea is that we add additional RNNs on top of the initial standard RNN, where each RNN added is another *layer*. The hidden state output by the first (bottom) RNN at time-step $t$ will be the input to the RNN above it at time step $t$. The prediction is then made from the final hidden state of the final (highest) layer.
The image below shows a multi-layer unidirectional RNN, where the layer number is given as a superscript. Also note that each layer needs their own initial hidden state, $h_0^L$.

### Regularization
Although we've added improvements to our model, each one adds additional parameters. Without going into overfitting into too much detail, the more parameters you have in in your model, the higher the probability that your model will overfit (memorize the training data, causing a low training error but high validation/testing error, i.e. poor generalization to new, unseen examples). To combat this, we use regularization. More specifically, we use a method of regularization called *dropout*. Dropout works by randomly *dropping out* (setting to 0) neurons in a layer during a forward pass. The probability that each neuron is dropped out is set by a hyperparameter and each neuron with dropout applied is considered indepenently. One theory about why dropout works is that a model with parameters dropped out can be seen as a "weaker" (less parameters) model. The predictions from all these "weaker" models (one for each forward pass) get averaged together withinin the parameters of the model. Thus, your one model can be thought of as an ensemble of weaker models, none of which are over-parameterized and thus should not overfit.
### Implementation Details
Another addition to this model is that we are not going to learn the embedding for the `<pad>` token. This is because we want to explitictly tell our model that padding tokens are irrelevant to determining the sentiment of a sentence. This means the embedding for the pad token will remain at what it is initialized to (we initialize it to all zeros later). We do this by passing the index of our pad token as the `padding_idx` argument to the `nn.Embedding` layer.
To use an LSTM instead of the standard RNN, we use `nn.LSTM` instead of `nn.RNN`. Also, note that the LSTM returns the `output` and a tuple of the final `hidden` state and the final `cell` state, whereas the standard RNN only returned the `output` and final `hidden` state.
As the final hidden state of our LSTM has both a forward and a backward component, which will be concatenated together, the size of the input to the `nn.Linear` layer is twice that of the hidden dimension size.
Implementing bidirectionality and adding additional layers are done by passing values for the `num_layers` and `bidirectional` arguments for the RNN/LSTM.
Dropout is implemented by initializing an `nn.Dropout` layer (the argument is the probability of dropping out each neuron) and using it within the `forward` method after each layer we want to apply dropout to. **Note**: never use dropout on the input or output layers (`text` or `fc` in this case), you only ever want to use dropout on intermediate layers. The LSTM has a `dropout` argument which adds dropout on the connections between hidden states in one layer to hidden states in the next layer.
As we are passing the lengths of our sentences to be able to use packed padded sequences, we have to add a second argument, `text_lengths`, to `forward`.
Before we pass our embeddings to the RNN, we need to pack them, which we do with `nn.utils.rnn.packed_padded_sequence`. This will cause our RNN to only process the non-padded elements of our sequence. The RNN will then return `packed_output` (a packed sequence) as well as the `hidden` and `cell` states (both of which are tensors). Without packed padded sequences, `hidden` and `cell` are tensors from the last element in the sequence, which will most probably be a pad token, however when using packed padded sequences they are both from the last non-padded element in the sequence. Note that the `lengths` argument of `packed_padded_sequence` must be a CPU tensor so we explicitly make it one by using `.to('cpu')`.
We then unpack the output sequence, with `nn.utils.rnn.pad_packed_sequence`, to transform it from a packed sequence to a tensor. The elements of `output` from padding tokens will be zero tensors (tensors where every element is zero). Usually, we only have to unpack output if we are going to use it later on in the model. Although we aren't in this case, we still unpack the sequence just to show how it is done.
The final hidden state, `hidden`, has a shape of _**[num layers * num directions, batch size, hid dim]**_. These are ordered: **[forward_layer_0, backward_layer_0, forward_layer_1, backward_layer 1, ..., forward_layer_n, backward_layer n]**. As we want the final (top) layer forward and backward hidden states, we get the top two hidden layers from the first dimension, `hidden[-2,:,:]` and `hidden[-1,:,:]`, and concatenate them together before passing them to the linear layer (after applying dropout).
```
import torch.nn as nn
class RNN(nn.Module):
def __init__(self, vocab_size, embedding_dim, hidden_dim, output_dim, n_layers,
bidirectional, dropout, pad_idx):
super().__init__()
self.embedding = nn.Embedding(vocab_size, embedding_dim, padding_idx = pad_idx)
self.rnn = nn.LSTM(embedding_dim,
hidden_dim,
num_layers=n_layers,
bidirectional=bidirectional,
dropout=dropout)
self.fc = nn.Linear(hidden_dim * 2, output_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, text, text_lengths):
#text = [sent len, batch size]
embedded = self.dropout(self.embedding(text))
#embedded = [sent len, batch size, emb dim]
#pack sequence
# lengths need to be on CPU!
packed_embedded = nn.utils.rnn.pack_padded_sequence(embedded, text_lengths.to('cpu'))
packed_output, (hidden, cell) = self.rnn(packed_embedded)
#unpack sequence
output, output_lengths = nn.utils.rnn.pad_packed_sequence(packed_output)
#output = [sent len, batch size, hid dim * num directions]
#output over padding tokens are zero tensors
#hidden = [num layers * num directions, batch size, hid dim]
#cell = [num layers * num directions, batch size, hid dim]
#concat the final forward (hidden[-2,:,:]) and backward (hidden[-1,:,:]) hidden layers
#and apply dropout
hidden = self.dropout(torch.cat((hidden[-2,:,:], hidden[-1,:,:]), dim = 1))
#hidden = [batch size, hid dim * num directions]
return self.fc(hidden)
```
Like before, we'll create an instance of our RNN class, with the new parameters and arguments for the number of layers, bidirectionality and dropout probability.
To ensure the pre-trained vectors can be loaded into the model, the `EMBEDDING_DIM` must be equal to that of the pre-trained GloVe vectors loaded earlier.
We get our pad token index from the vocabulary, getting the actual string representing the pad token from the field's `pad_token` attribute, which is `<pad>` by default.
```
INPUT_DIM = len(TEXT.vocab)
EMBEDDING_DIM = 100
HIDDEN_DIM = 256
OUTPUT_DIM = 1
N_LAYERS = 2
BIDIRECTIONAL = True
DROPOUT = 0.5
PAD_IDX = TEXT.vocab.stoi[TEXT.pad_token]
model = RNN(INPUT_DIM,
EMBEDDING_DIM,
HIDDEN_DIM,
OUTPUT_DIM,
N_LAYERS,
BIDIRECTIONAL,
DROPOUT,
PAD_IDX)
```
We'll print out the number of parameters in our model.
Notice how we have almost twice as many parameters as before!
```
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
```
The final addition is copying the pre-trained word embeddings we loaded earlier into the `embedding` layer of our model.
We retrieve the embeddings from the field's vocab, and check they're the correct size, _**[vocab size, embedding dim]**_
```
pretrained_embeddings = TEXT.vocab.vectors
print(pretrained_embeddings.shape)
```
We then replace the initial weights of the `embedding` layer with the pre-trained embeddings.
**Note**: this should always be done on the `weight.data` and not the `weight`!
```
model.embedding.weight.data.copy_(pretrained_embeddings)
```
As our `<unk>` and `<pad>` token aren't in the pre-trained vocabulary they have been initialized using `unk_init` (an $\mathcal{N}(0,1)$ distribution) when building our vocab. It is preferable to initialize them both to all zeros to explicitly tell our model that, initially, they are irrelevant for determining sentiment.
We do this by manually setting their row in the embedding weights matrix to zeros. We get their row by finding the index of the tokens, which we have already done for the padding index.
**Note**: like initializing the embeddings, this should be done on the `weight.data` and not the `weight`!
```
UNK_IDX = TEXT.vocab.stoi[TEXT.unk_token]
model.embedding.weight.data[UNK_IDX] = torch.zeros(EMBEDDING_DIM)
model.embedding.weight.data[PAD_IDX] = torch.zeros(EMBEDDING_DIM)
print(model.embedding.weight.data)
```
We can now see the first two rows of the embedding weights matrix have been set to zeros. As we passed the index of the pad token to the `padding_idx` of the embedding layer it will remain zeros throughout training, however the `<unk>` token embedding will be learned.
## Train the Model
Now to training the model.
The only change we'll make here is changing the optimizer from `SGD` to `Adam`. SGD updates all parameters with the same learning rate and choosing this learning rate can be tricky. `Adam` adapts the learning rate for each parameter, giving parameters that are updated more frequently lower learning rates and parameters that are updated infrequently higher learning rates. More information about `Adam` (and other optimizers) can be found [here](http://ruder.io/optimizing-gradient-descent/index.html).
To change `SGD` to `Adam`, we simply change `optim.SGD` to `optim.Adam`, also note how we do not have to provide an initial learning rate for Adam as PyTorch specifies a sensibile default initial learning rate.
```
import torch.optim as optim
optimizer = optim.Adam(model.parameters())
```
The rest of the steps for training the model are unchanged.
We define the criterion and place the model and criterion on the GPU (if available)...
```
criterion = nn.BCEWithLogitsLoss()
model = model.to(device)
criterion = criterion.to(device)
```
We implement the function to calculate accuracy...
```
def binary_accuracy(preds, y):
"""
Returns accuracy per batch, i.e. if you get 8/10 right, this returns 0.8, NOT 8
"""
#round predictions to the closest integer
rounded_preds = torch.round(torch.sigmoid(preds))
correct = (rounded_preds == y).float() #convert into float for division
acc = correct.sum() / len(correct)
return acc
```
We define a function for training our model.
As we have set `include_lengths = True`, our `batch.text` is now a tuple with the first element being the numericalized tensor and the second element being the actual lengths of each sequence. We separate these into their own variables, `text` and `text_lengths`, before passing them to the model.
**Note**: as we are now using dropout, we must remember to use `model.train()` to ensure the dropout is "turned on" while training.
```
def train(model, iterator, optimizer, criterion):
epoch_loss = 0
epoch_acc = 0
model.train()
for batch in iterator:
optimizer.zero_grad()
text, text_lengths = batch.text
predictions = model(text, text_lengths).squeeze(1)
loss = criterion(predictions, batch.label)
acc = binary_accuracy(predictions, batch.label)
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
```
Then we define a function for testing our model, again remembering to separate `batch.text`.
**Note**: as we are now using dropout, we must remember to use `model.eval()` to ensure the dropout is "turned off" while evaluating.
```
def evaluate(model, iterator, criterion):
epoch_loss = 0
epoch_acc = 0
model.eval()
with torch.no_grad():
for batch in iterator:
text, text_lengths = batch.text
predictions = model(text, text_lengths).squeeze(1)
loss = criterion(predictions, batch.label)
acc = binary_accuracy(predictions, batch.label)
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
```
And also create a nice function to tell us how long our epochs are taking.
```
import time
def epoch_time(start_time, end_time):
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
```
Finally, we train our model...
```
N_EPOCHS = 5
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
start_time = time.time()
train_loss, train_acc = train(model, train_iterator, optimizer, criterion)
valid_loss, valid_acc = evaluate(model, valid_iterator, criterion)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), 'tut2-model.pt')
print(f'Epoch: {epoch+1:02} | Epoch Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train Acc: {train_acc*100:.2f}%')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. Acc: {valid_acc*100:.2f}%')
```
...and get our new and vastly improved test accuracy!
```
model.load_state_dict(torch.load('tut2-model.pt'))
test_loss, test_acc = evaluate(model, test_iterator, criterion)
print(f'Test Loss: {test_loss:.3f} | Test Acc: {test_acc*100:.2f}%')
```
## User Input
We can now use our model to predict the sentiment of any sentence we give it. As it has been trained on movie reviews, the sentences provided should also be movie reviews.
When using a model for inference it should always be in evaluation mode. If this tutorial is followed step-by-step then it should already be in evaluation mode (from doing `evaluate` on the test set), however we explicitly set it to avoid any risk.
Our `predict_sentiment` function does a few things:
- sets the model to evaluation mode
- tokenizes the sentence, i.e. splits it from a raw string into a list of tokens
- indexes the tokens by converting them into their integer representation from our vocabulary
- gets the length of our sequence
- converts the indexes, which are a Python list into a PyTorch tensor
- add a batch dimension by `unsqueeze`ing
- converts the length into a tensor
- squashes the output prediction from a real number between 0 and 1 with the `sigmoid` function
- converts the tensor holding a single value into an integer with the `item()` method
We are expecting reviews with a negative sentiment to return a value close to 0 and positive reviews to return a value close to 1.
```
import spacy
nlp = spacy.load('en_core_web_sm')
def predict_sentiment(model, sentence):
model.eval()
tokenized = [tok.text for tok in nlp.tokenizer(sentence)]
indexed = [TEXT.vocab.stoi[t] for t in tokenized]
length = [len(indexed)]
tensor = torch.LongTensor(indexed).to(device)
tensor = tensor.unsqueeze(1)
length_tensor = torch.LongTensor(length)
prediction = torch.sigmoid(model(tensor, length_tensor))
return prediction.item()
```
An example negative review...
```
predict_sentiment(model, "This film is terrible")
```
An example positive review...
```
predict_sentiment(model, "This film is great")
```
## Next Steps
We've now built a decent sentiment analysis model for movie reviews! In the next notebook we'll implement a model that gets comparable accuracy with far fewer parameters and trains much, much faster.
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# CycleGAN
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/generative/cyclegan"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/generative/cyclegan.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/generative/cyclegan.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/generative/cyclegan.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
This notebook demonstrates unpaired image to image translation using conditional GAN's, as described in [Unpaired Image-to-Image Translation using Cycle-Consistent Adversarial Networks](https://arxiv.org/abs/1703.10593), also known as CycleGAN. The paper proposes a method that can capture the characteristics of one image domain and figure out how these characteristics could be translated into another image domain, all in the absence of any paired training examples.
This notebook assumes you are familiar with Pix2Pix, which you can learn about in the [Pix2Pix tutorial](https://www.tensorflow.org/tutorials/generative/pix2pix). The code for CycleGAN is similar, the main difference is an additional loss function, and the use of unpaired training data.
CycleGAN uses a cycle consistency loss to enable training without the need for paired data. In other words, it can translate from one domain to another without a one-to-one mapping between the source and target domain.
This opens up the possibility to do a lot of interesting tasks like photo-enhancement, image colorization, style transfer, etc. All you need is the source and the target dataset (which is simply a directory of images).


## Set up the input pipeline
Install the [tensorflow_examples](https://github.com/tensorflow/examples) package that enables importing of the generator and the discriminator.
```
!pip install git+https://github.com/tensorflow/examples.git
import tensorflow as tf
import tensorflow_datasets as tfds
from tensorflow_examples.models.pix2pix import pix2pix
import os
import time
import matplotlib.pyplot as plt
from IPython.display import clear_output
AUTOTUNE = tf.data.AUTOTUNE
```
## Input Pipeline
This tutorial trains a model to translate from images of horses, to images of zebras. You can find this dataset and similar ones [here](https://www.tensorflow.org/datasets/datasets#cycle_gan).
As mentioned in the [paper](https://arxiv.org/abs/1703.10593), apply random jittering and mirroring to the training dataset. These are some of the image augmentation techniques that avoids overfitting.
This is similar to what was done in [pix2pix](https://www.tensorflow.org/tutorials/generative/pix2pix#load_the_dataset)
* In random jittering, the image is resized to `286 x 286` and then randomly cropped to `256 x 256`.
* In random mirroring, the image is randomly flipped horizontally i.e left to right.
```
dataset, metadata = tfds.load('cycle_gan/horse2zebra',
with_info=True, as_supervised=True)
train_horses, train_zebras = dataset['trainA'], dataset['trainB']
test_horses, test_zebras = dataset['testA'], dataset['testB']
BUFFER_SIZE = 1000
BATCH_SIZE = 1
IMG_WIDTH = 256
IMG_HEIGHT = 256
def random_crop(image):
cropped_image = tf.image.random_crop(
image, size=[IMG_HEIGHT, IMG_WIDTH, 3])
return cropped_image
# normalizing the images to [-1, 1]
def normalize(image):
image = tf.cast(image, tf.float32)
image = (image / 127.5) - 1
return image
def random_jitter(image):
# resizing to 286 x 286 x 3
image = tf.image.resize(image, [286, 286],
method=tf.image.ResizeMethod.NEAREST_NEIGHBOR)
# randomly cropping to 256 x 256 x 3
image = random_crop(image)
# random mirroring
image = tf.image.random_flip_left_right(image)
return image
def preprocess_image_train(image, label):
image = random_jitter(image)
image = normalize(image)
return image
def preprocess_image_test(image, label):
image = normalize(image)
return image
train_horses = train_horses.map(
preprocess_image_train, num_parallel_calls=AUTOTUNE).cache().shuffle(
BUFFER_SIZE).batch(1)
train_zebras = train_zebras.map(
preprocess_image_train, num_parallel_calls=AUTOTUNE).cache().shuffle(
BUFFER_SIZE).batch(1)
test_horses = test_horses.map(
preprocess_image_test, num_parallel_calls=AUTOTUNE).cache().shuffle(
BUFFER_SIZE).batch(1)
test_zebras = test_zebras.map(
preprocess_image_test, num_parallel_calls=AUTOTUNE).cache().shuffle(
BUFFER_SIZE).batch(1)
sample_horse = next(iter(train_horses))
sample_zebra = next(iter(train_zebras))
plt.subplot(121)
plt.title('Horse')
plt.imshow(sample_horse[0] * 0.5 + 0.5)
plt.subplot(122)
plt.title('Horse with random jitter')
plt.imshow(random_jitter(sample_horse[0]) * 0.5 + 0.5)
plt.subplot(121)
plt.title('Zebra')
plt.imshow(sample_zebra[0] * 0.5 + 0.5)
plt.subplot(122)
plt.title('Zebra with random jitter')
plt.imshow(random_jitter(sample_zebra[0]) * 0.5 + 0.5)
```
## Import and reuse the Pix2Pix models
Import the generator and the discriminator used in [Pix2Pix](https://github.com/tensorflow/examples/blob/master/tensorflow_examples/models/pix2pix/pix2pix.py) via the installed [tensorflow_examples](https://github.com/tensorflow/examples) package.
The model architecture used in this tutorial is very similar to what was used in [pix2pix](https://github.com/tensorflow/examples/blob/master/tensorflow_examples/models/pix2pix/pix2pix.py). Some of the differences are:
* Cyclegan uses [instance normalization](https://arxiv.org/abs/1607.08022) instead of [batch normalization](https://arxiv.org/abs/1502.03167).
* The [CycleGAN paper](https://arxiv.org/abs/1703.10593) uses a modified `resnet` based generator. This tutorial is using a modified `unet` generator for simplicity.
There are 2 generators (G and F) and 2 discriminators (X and Y) being trained here.
* Generator `G` learns to transform image `X` to image `Y`. $(G: X -> Y)$
* Generator `F` learns to transform image `Y` to image `X`. $(F: Y -> X)$
* Discriminator `D_X` learns to differentiate between image `X` and generated image `X` (`F(Y)`).
* Discriminator `D_Y` learns to differentiate between image `Y` and generated image `Y` (`G(X)`).

```
OUTPUT_CHANNELS = 3
generator_g = pix2pix.unet_generator(OUTPUT_CHANNELS, norm_type='instancenorm')
generator_f = pix2pix.unet_generator(OUTPUT_CHANNELS, norm_type='instancenorm')
discriminator_x = pix2pix.discriminator(norm_type='instancenorm', target=False)
discriminator_y = pix2pix.discriminator(norm_type='instancenorm', target=False)
to_zebra = generator_g(sample_horse)
to_horse = generator_f(sample_zebra)
plt.figure(figsize=(8, 8))
contrast = 8
imgs = [sample_horse, to_zebra, sample_zebra, to_horse]
title = ['Horse', 'To Zebra', 'Zebra', 'To Horse']
for i in range(len(imgs)):
plt.subplot(2, 2, i+1)
plt.title(title[i])
if i % 2 == 0:
plt.imshow(imgs[i][0] * 0.5 + 0.5)
else:
plt.imshow(imgs[i][0] * 0.5 * contrast + 0.5)
plt.show()
plt.figure(figsize=(8, 8))
plt.subplot(121)
plt.title('Is a real zebra?')
plt.imshow(discriminator_y(sample_zebra)[0, ..., -1], cmap='RdBu_r')
plt.subplot(122)
plt.title('Is a real horse?')
plt.imshow(discriminator_x(sample_horse)[0, ..., -1], cmap='RdBu_r')
plt.show()
```
## Loss functions
In CycleGAN, there is no paired data to train on, hence there is no guarantee that the input `x` and the target `y` pair are meaningful during training. Thus in order to enforce that the network learns the correct mapping, the authors propose the cycle consistency loss.
The discriminator loss and the generator loss are similar to the ones used in [pix2pix](https://www.tensorflow.org/tutorials/generative/pix2pix#define_the_loss_functions_and_the_optimizer).
```
LAMBDA = 10
loss_obj = tf.keras.losses.BinaryCrossentropy(from_logits=True)
def discriminator_loss(real, generated):
real_loss = loss_obj(tf.ones_like(real), real)
generated_loss = loss_obj(tf.zeros_like(generated), generated)
total_disc_loss = real_loss + generated_loss
return total_disc_loss * 0.5
def generator_loss(generated):
return loss_obj(tf.ones_like(generated), generated)
```
Cycle consistency means the result should be close to the original input. For example, if one translates a sentence from English to French, and then translates it back from French to English, then the resulting sentence should be the same as the original sentence.
In cycle consistency loss,
* Image $X$ is passed via generator $G$ that yields generated image $\hat{Y}$.
* Generated image $\hat{Y}$ is passed via generator $F$ that yields cycled image $\hat{X}$.
* Mean absolute error is calculated between $X$ and $\hat{X}$.
$$forward\ cycle\ consistency\ loss: X -> G(X) -> F(G(X)) \sim \hat{X}$$
$$backward\ cycle\ consistency\ loss: Y -> F(Y) -> G(F(Y)) \sim \hat{Y}$$

```
def calc_cycle_loss(real_image, cycled_image):
loss1 = tf.reduce_mean(tf.abs(real_image - cycled_image))
return LAMBDA * loss1
```
As shown above, generator $G$ is responsible for translating image $X$ to image $Y$. Identity loss says that, if you fed image $Y$ to generator $G$, it should yield the real image $Y$ or something close to image $Y$.
If you run the zebra-to-horse model on a horse or the horse-to-zebra model on a zebra, it should not modify the image much since the image already contains the target class.
$$Identity\ loss = |G(Y) - Y| + |F(X) - X|$$
```
def identity_loss(real_image, same_image):
loss = tf.reduce_mean(tf.abs(real_image - same_image))
return LAMBDA * 0.5 * loss
```
Initialize the optimizers for all the generators and the discriminators.
```
generator_g_optimizer = tf.keras.optimizers.Adam(2e-4, beta_1=0.5)
generator_f_optimizer = tf.keras.optimizers.Adam(2e-4, beta_1=0.5)
discriminator_x_optimizer = tf.keras.optimizers.Adam(2e-4, beta_1=0.5)
discriminator_y_optimizer = tf.keras.optimizers.Adam(2e-4, beta_1=0.5)
```
## Checkpoints
```
checkpoint_path = "./checkpoints/train"
ckpt = tf.train.Checkpoint(generator_g=generator_g,
generator_f=generator_f,
discriminator_x=discriminator_x,
discriminator_y=discriminator_y,
generator_g_optimizer=generator_g_optimizer,
generator_f_optimizer=generator_f_optimizer,
discriminator_x_optimizer=discriminator_x_optimizer,
discriminator_y_optimizer=discriminator_y_optimizer)
ckpt_manager = tf.train.CheckpointManager(ckpt, checkpoint_path, max_to_keep=5)
# if a checkpoint exists, restore the latest checkpoint.
if ckpt_manager.latest_checkpoint:
ckpt.restore(ckpt_manager.latest_checkpoint)
print ('Latest checkpoint restored!!')
```
## Training
Note: This example model is trained for fewer epochs (40) than the paper (200) to keep training time reasonable for this tutorial. Predictions may be less accurate.
```
EPOCHS = 40
def generate_images(model, test_input):
prediction = model(test_input)
plt.figure(figsize=(12, 12))
display_list = [test_input[0], prediction[0]]
title = ['Input Image', 'Predicted Image']
for i in range(2):
plt.subplot(1, 2, i+1)
plt.title(title[i])
# getting the pixel values between [0, 1] to plot it.
plt.imshow(display_list[i] * 0.5 + 0.5)
plt.axis('off')
plt.show()
```
Even though the training loop looks complicated, it consists of four basic steps:
* Get the predictions.
* Calculate the loss.
* Calculate the gradients using backpropagation.
* Apply the gradients to the optimizer.
```
@tf.function
def train_step(real_x, real_y):
# persistent is set to True because the tape is used more than
# once to calculate the gradients.
with tf.GradientTape(persistent=True) as tape:
# Generator G translates X -> Y
# Generator F translates Y -> X.
fake_y = generator_g(real_x, training=True)
cycled_x = generator_f(fake_y, training=True)
fake_x = generator_f(real_y, training=True)
cycled_y = generator_g(fake_x, training=True)
# same_x and same_y are used for identity loss.
same_x = generator_f(real_x, training=True)
same_y = generator_g(real_y, training=True)
disc_real_x = discriminator_x(real_x, training=True)
disc_real_y = discriminator_y(real_y, training=True)
disc_fake_x = discriminator_x(fake_x, training=True)
disc_fake_y = discriminator_y(fake_y, training=True)
# calculate the loss
gen_g_loss = generator_loss(disc_fake_y)
gen_f_loss = generator_loss(disc_fake_x)
total_cycle_loss = calc_cycle_loss(real_x, cycled_x) + calc_cycle_loss(real_y, cycled_y)
# Total generator loss = adversarial loss + cycle loss
total_gen_g_loss = gen_g_loss + total_cycle_loss + identity_loss(real_y, same_y)
total_gen_f_loss = gen_f_loss + total_cycle_loss + identity_loss(real_x, same_x)
disc_x_loss = discriminator_loss(disc_real_x, disc_fake_x)
disc_y_loss = discriminator_loss(disc_real_y, disc_fake_y)
# Calculate the gradients for generator and discriminator
generator_g_gradients = tape.gradient(total_gen_g_loss,
generator_g.trainable_variables)
generator_f_gradients = tape.gradient(total_gen_f_loss,
generator_f.trainable_variables)
discriminator_x_gradients = tape.gradient(disc_x_loss,
discriminator_x.trainable_variables)
discriminator_y_gradients = tape.gradient(disc_y_loss,
discriminator_y.trainable_variables)
# Apply the gradients to the optimizer
generator_g_optimizer.apply_gradients(zip(generator_g_gradients,
generator_g.trainable_variables))
generator_f_optimizer.apply_gradients(zip(generator_f_gradients,
generator_f.trainable_variables))
discriminator_x_optimizer.apply_gradients(zip(discriminator_x_gradients,
discriminator_x.trainable_variables))
discriminator_y_optimizer.apply_gradients(zip(discriminator_y_gradients,
discriminator_y.trainable_variables))
for epoch in range(EPOCHS):
start = time.time()
n = 0
for image_x, image_y in tf.data.Dataset.zip((train_horses, train_zebras)):
train_step(image_x, image_y)
if n % 10 == 0:
print ('.', end='')
n+=1
clear_output(wait=True)
# Using a consistent image (sample_horse) so that the progress of the model
# is clearly visible.
generate_images(generator_g, sample_horse)
if (epoch + 1) % 5 == 0:
ckpt_save_path = ckpt_manager.save()
print ('Saving checkpoint for epoch {} at {}'.format(epoch+1,
ckpt_save_path))
print ('Time taken for epoch {} is {} sec\n'.format(epoch + 1,
time.time()-start))
```
## Generate using test dataset
```
# Run the trained model on the test dataset
for inp in test_horses.take(5):
generate_images(generator_g, inp)
```
## Next steps
This tutorial has shown how to implement CycleGAN starting from the generator and discriminator implemented in the [Pix2Pix](https://www.tensorflow.org/tutorials/generative/pix2pix) tutorial. As a next step, you could try using a different dataset from [TensorFlow Datasets](https://www.tensorflow.org/datasets/datasets#cycle_gan).
You could also train for a larger number of epochs to improve the results, or you could implement the modified ResNet generator used in the [paper](https://arxiv.org/abs/1703.10593) instead of the U-Net generator used here.
| github_jupyter |
```
!pip install tensorflow==2.0.0b1
import tensorflow as tf
print(tf.__version__)
```
The next code block will set up the time series with seasonality, trend and a bit of noise.
```
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
def plot_series(time, series, format="-", start=0, end=None):
plt.plot(time[start:end], series[start:end], format)
plt.xlabel("Time")
plt.ylabel("Value")
plt.grid(True)
def trend(time, slope=0):
return slope * time
def seasonal_pattern(season_time):
"""Just an arbitrary pattern, you can change it if you wish"""
return np.where(season_time < 0.4,
np.cos(season_time * 2 * np.pi),
1 / np.exp(3 * season_time))
def seasonality(time, period, amplitude=1, phase=0):
"""Repeats the same pattern at each period"""
season_time = ((time + phase) % period) / period
return amplitude * seasonal_pattern(season_time)
def noise(time, noise_level=1, seed=None):
rnd = np.random.RandomState(seed)
return rnd.randn(len(time)) * noise_level
time = np.arange(4 * 365 + 1, dtype="float32")
baseline = 10
series = trend(time, 0.1)
baseline = 10
amplitude = 40
slope = 0.05
noise_level = 5
# Create the series
series = baseline + trend(time, slope) + seasonality(time, period=365, amplitude=amplitude)
# Update with noise
series += noise(time, noise_level, seed=42)
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
```
Now that we have the time series, let's split it so we can start forecasting
```
split_time = 1000
time_train = time[:split_time]
x_train = series[:split_time]
time_valid = time[split_time:]
x_valid = series[split_time:]
plt.figure(figsize=(10, 6))
plot_series(time_train, x_train)
plt.show()
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plt.show()
```
# Naive Forecast
```
naive_forecast = series[split_time - 1:-1]
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, naive_forecast)
```
Let's zoom in on the start of the validation period:
```
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid, start=0, end=150)
plot_series(time_valid, naive_forecast, start=1, end=151)
```
You can see that the naive forecast lags 1 step behind the time series.
Now let's compute the mean squared error and the mean absolute error between the forecasts and the predictions in the validation period:
```
print(keras.metrics.mean_squared_error(x_valid, naive_forecast).numpy())
print(keras.metrics.mean_absolute_error(x_valid, naive_forecast).numpy())
```
That's our baseline, now let's try a moving average:
```
def moving_average_forecast(series, window_size):
"""Forecasts the mean of the last few values.
If window_size=1, then this is equivalent to naive forecast"""
forecast = []
for time in range(len(series) - window_size):
forecast.append(series[time:time + window_size].mean())
return np.array(forecast)
moving_avg = moving_average_forecast(series, 30)[split_time - 30:]
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, moving_avg)
print(keras.metrics.mean_squared_error(x_valid, moving_avg).numpy())
print(keras.metrics.mean_absolute_error(x_valid, moving_avg).numpy())
```
That's worse than naive forecast! The moving average does not anticipate trend or seasonality, so let's try to remove them by using differencing. Since the seasonality period is 365 days, we will subtract the value at time *t* – 365 from the value at time *t*.
```
diff_series = (series[365:] - series[:-365])
diff_time = time[365:]
plt.figure(figsize=(10, 6))
plot_series(diff_time, diff_series)
plt.show()
```
Great, the trend and seasonality seem to be gone, so now we can use the moving average:
```
diff_moving_avg = moving_average_forecast(diff_series, 50)[split_time - 365 - 50:]
plt.figure(figsize=(10, 6))
plot_series(time_valid, diff_series[split_time - 365:])
plot_series(time_valid, diff_moving_avg)
plt.show()
```
Now let's bring back the trend and seasonality by adding the past values from t – 365:
```
diff_moving_avg_plus_past = series[split_time - 365:-365] + diff_moving_avg
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, diff_moving_avg_plus_past)
plt.show()
print(keras.metrics.mean_squared_error(x_valid, diff_moving_avg_plus_past).numpy())
print(keras.metrics.mean_absolute_error(x_valid, diff_moving_avg_plus_past).numpy())
```
Better than naive forecast, good. However the forecasts look a bit too random, because we're just adding past values, which were noisy. Let's use a moving averaging on past values to remove some of the noise:
```
diff_moving_avg_plus_smooth_past = moving_average_forecast(series[split_time - 370:-360], 10) + diff_moving_avg
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, diff_moving_avg_plus_smooth_past)
plt.show()
print(keras.metrics.mean_squared_error(x_valid, diff_moving_avg_plus_smooth_past).numpy())
print(keras.metrics.mean_absolute_error(x_valid, diff_moving_avg_plus_smooth_past).numpy())
```
| github_jupyter |
# T1550.003 - Use Alternate Authentication Material: Pass the Ticket
Adversaries may “pass the ticket” using stolen Kerberos tickets to move laterally within an environment, bypassing normal system access controls. Pass the ticket (PtT) is a method of authenticating to a system using Kerberos tickets without having access to an account's password. Kerberos authentication can be used as the first step to lateral movement to a remote system.
In this technique, valid Kerberos tickets for [Valid Accounts](https://attack.mitre.org/techniques/T1078) are captured by [OS Credential Dumping](https://attack.mitre.org/techniques/T1003). A user's service tickets or ticket granting ticket (TGT) may be obtained, depending on the level of access. A service ticket allows for access to a particular resource, whereas a TGT can be used to request service tickets from the Ticket Granting Service (TGS) to access any resource the user has privileges to access.(Citation: ADSecurity AD Kerberos Attacks)(Citation: GentilKiwi Pass the Ticket)
[Silver Ticket](https://attack.mitre.org/techniques/T1558/002) can be obtained for services that use Kerberos as an authentication mechanism and are used to generate tickets to access that particular resource and the system that hosts the resource (e.g., SharePoint).(Citation: ADSecurity AD Kerberos Attacks)
[Golden Ticket](https://attack.mitre.org/techniques/T1558/001) can be obtained for the domain using the Key Distribution Service account KRBTGT account NTLM hash, which enables generation of TGTs for any account in Active Directory.(Citation: Campbell 2014)
## Atomic Tests
```
#Import the Module before running the tests.
# Checkout Jupyter Notebook at https://github.com/cyb3rbuff/TheAtomicPlaybook to run PS scripts.
Import-Module /Users/0x6c/AtomicRedTeam/atomics/invoke-atomicredteam/Invoke-AtomicRedTeam.psd1 - Force
```
### Atomic Test #1 - Mimikatz Kerberos Ticket Attack
Similar to PTH, but attacking Kerberos
**Supported Platforms:** windows
#### Attack Commands: Run with `command_prompt`
```command_prompt
mimikatz # kerberos::ptt Administrator@atomic.local
```
```
Invoke-AtomicTest T1550.003 -TestNumbers 1
```
## Detection
Audit all Kerberos authentication and credential use events and review for discrepancies. Unusual remote authentication events that correlate with other suspicious activity (such as writing and executing binaries) may indicate malicious activity.
Event ID 4769 is generated on the Domain Controller when using a golden ticket after the KRBTGT password has been reset twice, as mentioned in the mitigation section. The status code 0x1F indicates the action has failed due to "Integrity check on decrypted field failed" and indicates misuse by a previously invalidated golden ticket.(Citation: CERT-EU Golden Ticket Protection)
| github_jupyter |
<a href="https://colab.research.google.com/github/piyush5566/Genetic-Mutation-Classification/blob/master/genetic_mutation_classification_part_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import numpy as np
import pandas as pd
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import train_test_split
from nltk.corpus import stopwords
import re
# Loading training_variants. Its a comma seperated file
data_variants = pd.read_csv('Variants_data')
# Loading training_text dataset. This is seperated by ||
data_text =pd.read_csv("Text_data",sep="\|\|",engine="python",names=["ID","TEXT"],skiprows=1)
# We would like to remove all stop words like a, is, an, the, ...
# so we collecting all of them from nltk library
import nltk
nltk.download('stopwords')
stop_words = set(stopwords.words('english'))
def data_text_preprocess(total_text, ind, col):
# Remove int values from text data as that might not be imp
if type(total_text) is not int:
string = ""
# replacing all special char with space
total_text = re.sub('[^a-zA-Z0-9\n]', ' ', str(total_text))
# replacing multiple spaces with single space
total_text = re.sub('\s+',' ', str(total_text))
# bring whole text to same lower-case scale.
total_text = total_text.lower()
for word in total_text.split():
# if the word is a not a stop word then retain that word from text
if not word in stop_words:
string += word + " "
data_text[col][ind] = string
# Below code will take some time because its huge text (took 4 minute on my 16 GB RAM system), so run it and have a cup of coffee :)
for index, row in data_text.iterrows():
if type(row['TEXT']) is str:
data_text_preprocess(row['TEXT'], index, 'TEXT')
#merging both gene_variations and text data based on ID
result = pd.merge(data_variants, data_text,on='ID', how='left')
result.head()
result.loc[result['TEXT'].isnull(),'TEXT'] = result['Gene'] +' '+result['Variation']
y_true = result['Class'].values
result.Gene = result.Gene.str.replace('\s+', '_')
result.Variation = result.Variation.str.replace('\s+', '_')
result=result.drop(columns=['Class','ID'])
# Splitting the data into train and test set
X_train, test_df, y_train, y_test = train_test_split(result, y_true, stratify=y_true, test_size=0.2)
# split the train data now into train validation and cross validation
train_df, cv_df, y_train, y_cv = train_test_split(X_train, y_train, stratify=y_train, test_size=0.2)
print(train_df.shape,y_train.shape)
from sklearn.feature_extraction.text import TfidfVectorizer
# one-hot encoding of Gene feature.
#gene_vectorizer = CountVectorizer()
#gene_feature_onehotCoding = gene_vectorizer.fit_transform(result['Gene'])
#gene_feature_onehotCoding.shape
gene_vectorizer = TfidfVectorizer()
train_gene_feature_TFIDF=gene_vectorizer.fit_transform(train_df['Gene'])
cv_gene_feature_TFIDF=gene_vectorizer.transform(cv_df['Gene'])
test_gene_feature_TFIDF=gene_vectorizer.transform(test_df['Gene'])
# one-hot encoding of variation feature.
#variation_vectorizer = CountVectorizer()
#variation_feature_onehotCoding = variation_vectorizer.fit_transform(result['Variation'])
var_vectorizer = TfidfVectorizer()
train_variation_feature_TFIDF=var_vectorizer.fit_transform(train_df['Variation'])
cv_variation_feature_TFIDF=var_vectorizer.transform(cv_df['Variation'])
test_variation_feature_TFIDF=var_vectorizer.transform(test_df['Variation'])
from sklearn.preprocessing import normalize
text_vectorizer = TfidfVectorizer()
train_text_feature_TFIDF=text_vectorizer.fit_transform(train_df['TEXT'])
train_text_feature_TFIDF=normalize(train_text_feature_TFIDF,axis=0)
cv_text_feature_TFIDF=text_vectorizer.transform(cv_df['TEXT'])
cv_text_feature_TFIDF=normalize(cv_text_feature_TFIDF,axis=0)
test_text_feature_TFIDF=text_vectorizer.transform(test_df['TEXT'])
test_text_feature_TFIDF=normalize(test_text_feature_TFIDF,axis=0)
train_gene_var_TFIDF = np.hstack((train_gene_feature_TFIDF.toarray(),train_variation_feature_TFIDF.toarray()))
train_x_TFIDF = np.hstack((train_gene_var_TFIDF, train_text_feature_TFIDF.toarray()))
cv_gene_var_TFIDF = np.hstack((cv_gene_feature_TFIDF.toarray(),cv_variation_feature_TFIDF.toarray()))
cv_x_TFIDF = np.hstack((cv_gene_var_TFIDF, cv_text_feature_TFIDF.toarray()))
test_gene_var_TFIDF = np.hstack((test_gene_feature_TFIDF.toarray(),test_variation_feature_TFIDF.toarray()))
test_x_TFIDF = np.hstack((test_gene_var_TFIDF, test_text_feature_TFIDF.toarray()))
train_x_TFIDF.shape
```
TFIDF + RANDOM OVERSAMPLER + XGBOOST
```
from imblearn.over_sampling import RandomOverSampler
ros = RandomOverSampler()
train_X_ros, train_y_ros = ros.fit_sample(train_x_TFIDF, y_train)
train_X_ros.shape
df_y=pd.Series(train_y_ros,name='Class')
df_y.value_counts().sort_index()
import xgboost as xgb
#dtrain = xgb.DMatrix(X_train, label=y_train)
#dcv=xgb.DMatrix(X_cv,label=y_cv)
from sklearn.metrics.classification import accuracy_score, log_loss
model = xgb.XGBClassifier()
eval_set = [(cv_x_TFIDF, y_cv)]
model.fit(train_X_ros, train_y_ros, eval_metric="mlogloss", eval_set=eval_set, verbose=True)
# make predictions for test data
predict_y = model.predict_proba(train_x_TFIDF)
print("The train log loss is:",log_loss(y_train, predict_y, labels=model.classes_, eps=1e-15))
predict_y = model.predict_proba(cv_x_TFIDF)
print("The cross validation log loss is:",log_loss(y_cv, predict_y, labels=model.classes_, eps=1e-15))
predict_y = model.predict_proba(test_x_TFIDF)
print("The test log loss is:",log_loss(y_test, predict_y, labels=model.classes_, eps=1e-15))
y_pred=model.predict(test_x_TFIDF)
from sklearn.metrics import classification_report
print(classification_report(y_test,y_pred))
```
TFIDF + SMOTE + XGBOOST
```
from imblearn.over_sampling import SMOTE
smote = SMOTE(ratio='minority')
train_X_ros, train_y_ros = smote.fit_sample(train_x_TFIDF, y_train)
train_X_ros.shape
df_y=pd.Series(train_y_ros,name='Class')
df_y.value_counts().sort_index()
#from sklearn.metrics.classification import accuracy_score, log_loss
model = xgb.XGBClassifier()
eval_set = [(cv_x_TFIDF, y_cv)]
model.fit(train_X_ros, train_y_ros, eval_metric="mlogloss", eval_set=eval_set, verbose=True)
# make predictions for test data
predict_y = model.predict_proba(train_x_TFIDF)
print("The train log loss is:",log_loss(y_train, predict_y, labels=model.classes_, eps=1e-15))
predict_y = model.predict_proba(cv_x_TFIDF)
print("The cross validation log loss is:",log_loss(y_cv, predict_y, labels=model.classes_, eps=1e-15))
predict_y = model.predict_proba(test_x_TFIDF)
print("The test log loss is:",log_loss(y_test, predict_y, labels=model.classes_, eps=1e-15))
y_pred=model.predict(test_x_TFIDF)
from sklearn.metrics import classification_report
print(classification_report(y_test,y_pred))
```
| github_jupyter |
# UNSUPERVISED LEARNING MODELS
```
import numpy as np
from sklearn.cluster import KMeans
from sklearn.datasets import make_blobs
# define number of samples
n_samples = 100
# define random state value to initialize the center
random_state = 20
# define number of feature as 5
X,y = make_blobs(n_samples=n_samples, n_features=5, random_state=None)
# define number of cluster to be formed as 3 and
# in random state and fit features into the model
predict_y = KMeans(n_clusters=3, random_state=random_state).fit_predict(X)
# estimator function
predict_y
```
# REDUCING DIMENSIONS USING PCA
```
from sklearn.decomposition import PCA
from sklearn.datasets import make_blobs
# define sample and random state
n_sample = 20
random_state = 20
# generates the data with 10 features
X,y = make_blobs(n_samples=n_sample, n_features=10, random_state=None)
# view the shape of the dataset
X.shape
# define PCA estimator with the number of reduced components
pca = PCA(n_components=3)
# fit the data into the PCA estimator
pca.fit(X)
pca.explained_variance_ratio_
# first PCA component
first_pca = pca.components_[0]
first_pca
# transform the fitted data to apply dimensionality reduction
pca_reduced = pca.transform(X)
# VIEW THE REDUCED SHAPE
pca_reduced.shape
# output: number of features reduced from 10 to 3
```
# BUILDING A PIPELINE
```
from sklearn.pipeline import Pipeline
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import LogisticRegression
from sklearn.decomposition import PCA
# chain the estimators
estimator = [('dim_reduction', PCA()),('logres_model', LogisticRegression())]
estimator
# put them in a pipeline object
pipeline_estimator = Pipeline(steps=estimator)
# check the chain of estimators
pipeline_estimator
# view first step
pipeline_estimator.steps[0]
# view the second step
pipeline_estimator.steps[1]
# view all the steps in the pipeline
pipeline_estimator.steps
```
# MODEL PERSISTENCE
```
from sklearn.datasets import load_iris
iris_ds = load_iris()
iris_ds.feature_names
iris_ds.target
X_feature = iris_ds.data
Y_target = iris_ds.target
X_new = [[3,3,3,3],[4,4,4,4]]
logreg = LogisticRegression()
logreg.fit(X_feature, Y_target)
logreg.predict(X_new)
# for model persistence
import pickle as pkl
# using dumps method to persist the model
persist_model = pkl.dumps(logreg)
persist_model
# persist the model to file
from sklearn.externals import joblib
joblib.dump(logreg, 'ml_unsupervised_models_1.pkl')
new_logreg_estimator = joblib.load('ml_unsupervised_models_1.pkl')
new_logreg_estimator
new_logreg_estimator.predict(X_new)
# output: the new model predicts same as the old one
```
| github_jupyter |
This notebook was prepared by [Donne Martin](http://donnemartin.com). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# Solution Notebook
## Problem: Remove duplicates from a linked list.
* [Constraints](#Constraints)
* [Test Cases](#Test-Cases)
* [Algorithm: Hash Map Lookup](#Algorithm:-Hash-Map-Lookup)
* [Algorithm: In-Place](#Algorithm:-In-Place)
* [Code](#Code)
* [Unit Test](#Unit-Test)
## Constraints
* Can we assume this is a non-circular, singly linked list?
* Yes
* Can you insert None values in the list?
* No
* Can we assume we already have a linked list class that can be used for this problem?
* Yes
* Can we use additional data structures?
* Implement both solutions
* Can we assume this fits in memory?
* Yes
## Test Cases
* Empty linked list -> []
* One element linked list -> [element]
* General case with no duplicates
* General case with duplicates
## Algorithm: Hash Map Lookup
Loop through each node
* For each node
* If the node's value is in the hash map
* Delete the node
* Else
* Add node's value to the hash map
Complexity:
* Time: O(n)
* Space: O(n)
## Algorithm: In-Place
* For each node
* Compare node with every other node
* Delete nodes that match current node
Complexity:
* Time: O(n^2)
* Space: O(1)
Note:
* We'll need to use a 'runner' to check every other node and compare it to the current node
## Code
```
%run ../linked_list/linked_list.py
class MyLinkedList(LinkedList):
def remove_dupes(self):
if self.head is None:
return
node = self.head
seen_data = set()
while node is not None:
if node.data not in seen_data:
seen_data.add(node.data)
prev = node
node = node.next
else:
prev.next = node.next
node = node.next
def remove_dupes_single_pointer(self):
if self.head is None:
return
node = self.head
seen_data = set({node.data})
while node.next is not None:
if node.next.data in seen_data:
node.next = node.next.next
else:
seen_data.add(node.next.data)
node = node.next
def remove_dupes_in_place(self):
curr = self.head
while curr is not None:
runner = curr
while runner.next is not None:
if runner.next.data == curr.data:
runner.next = runner.next.next
else:
runner = runner.next
curr = curr.next
```
## Unit Test
```
%%writefile test_remove_duplicates.py
import unittest
class TestRemoveDupes(unittest.TestCase):
def test_remove_dupes(self, linked_list):
print('Test: Empty list')
linked_list.remove_dupes()
self.assertEqual(linked_list.get_all_data(), [])
print('Test: One element list')
linked_list.insert_to_front(2)
linked_list.remove_dupes()
self.assertEqual(linked_list.get_all_data(), [2])
print('Test: General case, duplicates')
linked_list.insert_to_front(1)
linked_list.insert_to_front(1)
linked_list.insert_to_front(3)
linked_list.insert_to_front(2)
linked_list.insert_to_front(3)
linked_list.insert_to_front(1)
linked_list.insert_to_front(1)
linked_list.remove_dupes()
self.assertEqual(linked_list.get_all_data(), [1, 3, 2])
print('Test: General case, no duplicates')
linked_list.remove_dupes()
self.assertEqual(linked_list.get_all_data(), [1, 3, 2])
print('Success: test_remove_dupes\n')
def main():
test = TestRemoveDupes()
linked_list = MyLinkedList(None)
test.test_remove_dupes(linked_list)
if __name__ == '__main__':
main()
run -i test_remove_duplicates.py
```
| github_jupyter |
```
import pandas as pd
import spacy
import ast
from sklearn.preprocessing import MultiLabelBinarizer
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import train_test_split
from sklearn.linear_model import SGDClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.svm import LinearSVC
from sklearn.multiclass import OneVsRestClassifier
from spacy.lang.en.stop_words import STOP_WORDS
from nltk.tokenize import RegexpTokenizer
from nltk.corpus import stopwords
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
from sklearn.preprocessing import LabelEncoder
import nltk
nltk.download('wordnet')
from nltk.stem import WordNetLemmatizer
nltk.download('stopwords')
from google.colab import files
from sklearn import model_selection
from tensorflow import keras
from tensorflow.keras import layers, models
import numpy as np
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from keras.layers import Flatten
from keras.layers.embeddings import Embedding
from tensorflow.keras.layers import BatchNormalization
df = pd.read_json('https://raw.githubusercontent.com/triantafillu/Bootcamp-Repository-Language-2/main/data/ne_data.json')
df.head()
```
# Preprocessing
```
df.shape
df = df.dropna()
themes_encoder = MultiLabelBinarizer()
y = themes_encoder.fit_transform(df['themes'])
df['themes_encoded'] = [x for x in y]
df['themes_encoded'][0]
df['full_text'] = df.apply(lambda row: row.title + " " + row.text, axis = 1)
df.head()
df.drop(['title', 'text', 'themes'], axis=1, inplace=True)
df.head()
labelencoder = LabelEncoder()
df['author'] = labelencoder.fit_transform(df['author'])
df.head()
# Decontraction
def full_form(word):
if word == "nt": word = 'not'
if word == "re": word = 'be'
if word == "d": word = 'would'
if word == "m": word = 'am'
if word == "s": word = 'be'
if word == "ve": word = 'have'
return word
def preprocessing(text):
tokenizer = RegexpTokenizer(r'\w+')
text = tokenizer.tokenize(text)
stop_words = set(stopwords.words('english'))
cleaned_text = []
for word in text:
if word not in stop_words:
cleaned_text.append(word)
wnl = WordNetLemmatizer()
text = [wnl.lemmatize(token) for token in cleaned_text]
text = [full_form(w).lower() for w in text]
return text
df['full_text'] = df['full_text'].apply(preprocessing)
df['full_text']
texts_len = df['full_text'].apply(len)
df.drop(df[texts_len<50].index, inplace=True)
df.drop(df[texts_len>150].index, inplace=True)
tokenizer = Tokenizer(num_words=3000)
tokenizer.fit_on_texts(df['full_text'])
# Encode training data sentences into sequences
df['full_text'] = tokenizer.texts_to_sequences(df['full_text'])
df['full_text']
# Get max training sequence length
maxlen = 150 #max([len(x) for x in df['full_text']])
# Pad the training sequences
padded = pad_sequences(df['full_text'], padding='post', truncating='post', maxlen=maxlen)
padded
df['full_text'] = [x for x in padded]
df['full_text']
df.head()
df.drop(df.columns[0], axis=1, inplace = True)
df.head()
def convert_to_decade(x):
dec = x // 10
res = dec * 10
return res
df.year = df.year.apply(convert_to_decade)
df
```
# Model
```
X = np.array(df['full_text'].to_list())
Y = np.array(df['year'].to_list())
X_train, X_test, Y_train, Y_test = model_selection.train_test_split(X, Y, test_size=0.1, random_state=37)
max_features = 10000
max_len = 150
embedding_dim = 16
model = keras.models.Sequential([
keras.layers.Embedding(input_dim=max_features,
output_dim=embedding_dim,
input_length=max_len),
keras.layers.Flatten(),
keras.layers.Dense(64,activation='relu'),
keras.layers.Dense(32,activation='relu'),
keras.layers.Dense(16,activation='relu'),
keras.layers.Dense(8,activation='relu'),
keras.layers.Dense(1, activation='relu')
])
model.build()
model.compile(optimizer='adam',
loss='mean_squared_error',
metrics=['MAE'])
model.summary()
model.fit(X_train, Y_train, epochs=10, batch_size = 64, validation_split=0.1)
score = model.evaluate(np.array(X_test), np.array(Y_test))
print("Test Score:", score2[0])
print("Test Accuracy:", score2[1])
from google.colab import drive
drive.mount('/drive')
model.save('/drive/My Drive/Colab Notebooks/year_prediction.h5')
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Neural machine translation with attention
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/text/nmt_with_attention">
<img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/text/nmt_with_attention.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/text/nmt_with_attention.ipynb">
<img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/text/nmt_with_attention.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
This notebook trains a sequence to sequence (seq2seq) model for Spanish to English translation. This is an advanced example that assumes some knowledge of sequence to sequence models.
After training the model in this notebook, you will be able to input a Spanish sentence, such as *"¿todavia estan en casa?"*, and return the English translation: *"are you still at home?"*
The translation quality is reasonable for a toy example, but the generated attention plot is perhaps more interesting. This shows which parts of the input sentence has the model's attention while translating:
<img src="https://tensorflow.org/images/spanish-english.png" alt="spanish-english attention plot">
Note: This example takes approximately 10 minutes to run on a single P100 GPU.
```
import tensorflow as tf
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
from sklearn.model_selection import train_test_split
import unicodedata
import re
import numpy as np
import os
import io
import time
```
## Download and prepare the dataset
We'll use a language dataset provided by http://www.manythings.org/anki/. This dataset contains language translation pairs in the format:
```
May I borrow this book? ¿Puedo tomar prestado este libro?
```
There are a variety of languages available, but we'll use the English-Spanish dataset. For convenience, we've hosted a copy of this dataset on Google Cloud, but you can also download your own copy. After downloading the dataset, here are the steps we'll take to prepare the data:
1. Add a *start* and *end* token to each sentence.
2. Clean the sentences by removing special characters.
3. Create a word index and reverse word index (dictionaries mapping from word → id and id → word).
4. Pad each sentence to a maximum length.
```
# Download the file
path_to_zip = tf.keras.utils.get_file(
'spa-eng.zip', origin='http://storage.googleapis.com/download.tensorflow.org/data/spa-eng.zip',
extract=True)
path_to_file = os.path.dirname(path_to_zip)+"/spa-eng/spa.txt"
# Converts the unicode file to ascii
def unicode_to_ascii(s):
return ''.join(c for c in unicodedata.normalize('NFD', s)
if unicodedata.category(c) != 'Mn')
def preprocess_sentence(w):
w = unicode_to_ascii(w.lower().strip())
# creating a space between a word and the punctuation following it
# eg: "he is a boy." => "he is a boy ."
# Reference:- https://stackoverflow.com/questions/3645931/python-padding-punctuation-with-white-spaces-keeping-punctuation
w = re.sub(r"([?.!,¿])", r" \1 ", w)
w = re.sub(r'[" "]+', " ", w)
# replacing everything with space except (a-z, A-Z, ".", "?", "!", ",")
w = re.sub(r"[^a-zA-Z?.!,¿]+", " ", w)
w = w.strip()
# adding a start and an end token to the sentence
# so that the model know when to start and stop predicting.
w = '<start> ' + w + ' <end>'
return w
en_sentence = u"May I borrow this book?"
sp_sentence = u"¿Puedo tomar prestado este libro?"
print(preprocess_sentence(en_sentence))
print(preprocess_sentence(sp_sentence).encode('utf-8'))
# 1. Remove the accents
# 2. Clean the sentences
# 3. Return word pairs in the format: [ENGLISH, SPANISH]
def create_dataset(path, num_examples):
lines = io.open(path, encoding='UTF-8').read().strip().split('\n')
word_pairs = [[preprocess_sentence(w) for w in l.split('\t')] for l in lines[:num_examples]]
return zip(*word_pairs)
en, sp = create_dataset(path_to_file, None)
print(en[-1])
print(sp[-1])
def tokenize(lang):
lang_tokenizer = tf.keras.preprocessing.text.Tokenizer(
filters='')
lang_tokenizer.fit_on_texts(lang)
tensor = lang_tokenizer.texts_to_sequences(lang)
tensor = tf.keras.preprocessing.sequence.pad_sequences(tensor,
padding='post')
return tensor, lang_tokenizer
def load_dataset(path, num_examples=None):
# creating cleaned input, output pairs
targ_lang, inp_lang = create_dataset(path, num_examples)
input_tensor, inp_lang_tokenizer = tokenize(inp_lang)
target_tensor, targ_lang_tokenizer = tokenize(targ_lang)
return input_tensor, target_tensor, inp_lang_tokenizer, targ_lang_tokenizer
```
### Limit the size of the dataset to experiment faster (optional)
Training on the complete dataset of >100,000 sentences will take a long time. To train faster, we can limit the size of the dataset to 30,000 sentences (of course, translation quality degrades with less data):
```
# Try experimenting with the size of that dataset
num_examples = 30000
input_tensor, target_tensor, inp_lang, targ_lang = load_dataset(path_to_file, num_examples)
# Calculate max_length of the target tensors
max_length_targ, max_length_inp = target_tensor.shape[1], input_tensor.shape[1]
# Creating training and validation sets using an 80-20 split
input_tensor_train, input_tensor_val, target_tensor_train, target_tensor_val = train_test_split(input_tensor, target_tensor, test_size=0.2)
# Show length
print(len(input_tensor_train), len(target_tensor_train), len(input_tensor_val), len(target_tensor_val))
def convert(lang, tensor):
for t in tensor:
if t!=0:
print ("%d ----> %s" % (t, lang.index_word[t]))
print ("Input Language; index to word mapping")
convert(inp_lang, input_tensor_train[0])
print ()
print ("Target Language; index to word mapping")
convert(targ_lang, target_tensor_train[0])
```
### Create a tf.data dataset
```
BUFFER_SIZE = len(input_tensor_train)
BATCH_SIZE = 64
steps_per_epoch = len(input_tensor_train)//BATCH_SIZE
embedding_dim = 256
units = 1024
vocab_inp_size = len(inp_lang.word_index)+1
vocab_tar_size = len(targ_lang.word_index)+1
dataset = tf.data.Dataset.from_tensor_slices((input_tensor_train, target_tensor_train)).shuffle(BUFFER_SIZE)
dataset = dataset.batch(BATCH_SIZE, drop_remainder=True)
example_input_batch, example_target_batch = next(iter(dataset))
example_input_batch.shape, example_target_batch.shape
```
## Write the encoder and decoder model
Implement an encoder-decoder model with attention which you can read about in the TensorFlow [Neural Machine Translation (seq2seq) tutorial](https://github.com/tensorflow/nmt). This example uses a more recent set of APIs. This notebook implements the [attention equations](https://github.com/tensorflow/nmt#background-on-the-attention-mechanism) from the seq2seq tutorial. The following diagram shows that each input words is assigned a weight by the attention mechanism which is then used by the decoder to predict the next word in the sentence. The below picture and formulas are an example of attention mechanism from [Luong's paper](https://arxiv.org/abs/1508.04025v5).
<img src="https://www.tensorflow.org/images/seq2seq/attention_mechanism.jpg" width="500" alt="attention mechanism">
The input is put through an encoder model which gives us the encoder output of shape *(batch_size, max_length, hidden_size)* and the encoder hidden state of shape *(batch_size, hidden_size)*.
Here are the equations that are implemented:
<img src="https://www.tensorflow.org/images/seq2seq/attention_equation_0.jpg" alt="attention equation 0" width="800">
<img src="https://www.tensorflow.org/images/seq2seq/attention_equation_1.jpg" alt="attention equation 1" width="800">
This tutorial uses [Bahdanau attention](https://arxiv.org/pdf/1409.0473.pdf) for the encoder. Let's decide on notation before writing the simplified form:
* FC = Fully connected (dense) layer
* EO = Encoder output
* H = hidden state
* X = input to the decoder
And the pseudo-code:
* `score = FC(tanh(FC(EO) + FC(H)))`
* `attention weights = softmax(score, axis = 1)`. Softmax by default is applied on the last axis but here we want to apply it on the *1st axis*, since the shape of score is *(batch_size, max_length, hidden_size)*. `Max_length` is the length of our input. Since we are trying to assign a weight to each input, softmax should be applied on that axis.
* `context vector = sum(attention weights * EO, axis = 1)`. Same reason as above for choosing axis as 1.
* `embedding output` = The input to the decoder X is passed through an embedding layer.
* `merged vector = concat(embedding output, context vector)`
* This merged vector is then given to the GRU
The shapes of all the vectors at each step have been specified in the comments in the code:
```
class Encoder(tf.keras.Model):
def __init__(self, vocab_size, embedding_dim, enc_units, batch_sz):
super(Encoder, self).__init__()
self.batch_sz = batch_sz
self.enc_units = enc_units
self.embedding = tf.keras.layers.Embedding(vocab_size, embedding_dim)
self.gru = tf.keras.layers.GRU(self.enc_units,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
def call(self, x, hidden):
x = self.embedding(x)
output, state = self.gru(x, initial_state = hidden)
return output, state
def initialize_hidden_state(self):
return tf.zeros((self.batch_sz, self.enc_units))
encoder = Encoder(vocab_inp_size, embedding_dim, units, BATCH_SIZE)
# sample input
sample_hidden = encoder.initialize_hidden_state()
sample_output, sample_hidden = encoder(example_input_batch, sample_hidden)
print ('Encoder output shape: (batch size, sequence length, units) {}'.format(sample_output.shape))
print ('Encoder Hidden state shape: (batch size, units) {}'.format(sample_hidden.shape))
class BahdanauAttention(tf.keras.layers.Layer):
def __init__(self, units):
super(BahdanauAttention, self).__init__()
self.W1 = tf.keras.layers.Dense(units)
self.W2 = tf.keras.layers.Dense(units)
self.V = tf.keras.layers.Dense(1)
def call(self, query, values):
# query hidden state shape == (batch_size, hidden size)
# query_with_time_axis shape == (batch_size, 1, hidden size)
# values shape == (batch_size, max_len, hidden size)
# we are doing this to broadcast addition along the time axis to calculate the score
query_with_time_axis = tf.expand_dims(query, 1)
# score shape == (batch_size, max_length, 1)
# we get 1 at the last axis because we are applying score to self.V
# the shape of the tensor before applying self.V is (batch_size, max_length, units)
score = self.V(tf.nn.tanh(
self.W1(query_with_time_axis) + self.W2(values)))
# attention_weights shape == (batch_size, max_length, 1)
attention_weights = tf.nn.softmax(score, axis=1)
# context_vector shape after sum == (batch_size, hidden_size)
context_vector = attention_weights * values
context_vector = tf.reduce_sum(context_vector, axis=1)
return context_vector, attention_weights
attention_layer = BahdanauAttention(10)
attention_result, attention_weights = attention_layer(sample_hidden, sample_output)
print("Attention result shape: (batch size, units) {}".format(attention_result.shape))
print("Attention weights shape: (batch_size, sequence_length, 1) {}".format(attention_weights.shape))
class Decoder(tf.keras.Model):
def __init__(self, vocab_size, embedding_dim, dec_units, batch_sz):
super(Decoder, self).__init__()
self.batch_sz = batch_sz
self.dec_units = dec_units
self.embedding = tf.keras.layers.Embedding(vocab_size, embedding_dim)
self.gru = tf.keras.layers.GRU(self.dec_units,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
self.fc = tf.keras.layers.Dense(vocab_size)
# used for attention
self.attention = BahdanauAttention(self.dec_units)
def call(self, x, hidden, enc_output):
# enc_output shape == (batch_size, max_length, hidden_size)
context_vector, attention_weights = self.attention(hidden, enc_output)
# x shape after passing through embedding == (batch_size, 1, embedding_dim)
x = self.embedding(x)
# x shape after concatenation == (batch_size, 1, embedding_dim + hidden_size)
x = tf.concat([tf.expand_dims(context_vector, 1), x], axis=-1)
# passing the concatenated vector to the GRU
output, state = self.gru(x)
# output shape == (batch_size * 1, hidden_size)
output = tf.reshape(output, (-1, output.shape[2]))
# output shape == (batch_size, vocab)
x = self.fc(output)
return x, state, attention_weights
decoder = Decoder(vocab_tar_size, embedding_dim, units, BATCH_SIZE)
sample_decoder_output, _, _ = decoder(tf.random.uniform((BATCH_SIZE, 1)),
sample_hidden, sample_output)
print ('Decoder output shape: (batch_size, vocab size) {}'.format(sample_decoder_output.shape))
```
## Define the optimizer and the loss function
```
optimizer = tf.keras.optimizers.Adam()
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(
from_logits=True, reduction='none')
def loss_function(real, pred):
mask = tf.math.logical_not(tf.math.equal(real, 0))
loss_ = loss_object(real, pred)
mask = tf.cast(mask, dtype=loss_.dtype)
loss_ *= mask
return tf.reduce_mean(loss_)
```
## Checkpoints (Object-based saving)
```
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
checkpoint = tf.train.Checkpoint(optimizer=optimizer,
encoder=encoder,
decoder=decoder)
```
## Training
1. Pass the *input* through the *encoder* which return *encoder output* and the *encoder hidden state*.
2. The encoder output, encoder hidden state and the decoder input (which is the *start token*) is passed to the decoder.
3. The decoder returns the *predictions* and the *decoder hidden state*.
4. The decoder hidden state is then passed back into the model and the predictions are used to calculate the loss.
5. Use *teacher forcing* to decide the next input to the decoder.
6. *Teacher forcing* is the technique where the *target word* is passed as the *next input* to the decoder.
7. The final step is to calculate the gradients and apply it to the optimizer and backpropagate.
```
@tf.function
def train_step(inp, targ, enc_hidden):
loss = 0
with tf.GradientTape() as tape:
enc_output, enc_hidden = encoder(inp, enc_hidden)
dec_hidden = enc_hidden
dec_input = tf.expand_dims([targ_lang.word_index['<start>']] * BATCH_SIZE, 1)
# Teacher forcing - feeding the target as the next input
for t in range(1, targ.shape[1]):
# passing enc_output to the decoder
predictions, dec_hidden, _ = decoder(dec_input, dec_hidden, enc_output)
loss += loss_function(targ[:, t], predictions)
# using teacher forcing
dec_input = tf.expand_dims(targ[:, t], 1)
batch_loss = (loss / int(targ.shape[1]))
variables = encoder.trainable_variables + decoder.trainable_variables
gradients = tape.gradient(loss, variables)
optimizer.apply_gradients(zip(gradients, variables))
return batch_loss
EPOCHS = 10
for epoch in range(EPOCHS):
start = time.time()
enc_hidden = encoder.initialize_hidden_state()
total_loss = 0
for (batch, (inp, targ)) in enumerate(dataset.take(steps_per_epoch)):
batch_loss = train_step(inp, targ, enc_hidden)
total_loss += batch_loss
if batch % 100 == 0:
print('Epoch {} Batch {} Loss {:.4f}'.format(epoch + 1,
batch,
batch_loss.numpy()))
# saving (checkpoint) the model every 2 epochs
if (epoch + 1) % 2 == 0:
checkpoint.save(file_prefix = checkpoint_prefix)
print('Epoch {} Loss {:.4f}'.format(epoch + 1,
total_loss / steps_per_epoch))
print('Time taken for 1 epoch {} sec\n'.format(time.time() - start))
```
## Translate
* The evaluate function is similar to the training loop, except we don't use *teacher forcing* here. The input to the decoder at each time step is its previous predictions along with the hidden state and the encoder output.
* Stop predicting when the model predicts the *end token*.
* And store the *attention weights for every time step*.
Note: The encoder output is calculated only once for one input.
```
def evaluate(sentence):
attention_plot = np.zeros((max_length_targ, max_length_inp))
sentence = preprocess_sentence(sentence)
inputs = [inp_lang.word_index[i] for i in sentence.split(' ')]
inputs = tf.keras.preprocessing.sequence.pad_sequences([inputs],
maxlen=max_length_inp,
padding='post')
inputs = tf.convert_to_tensor(inputs)
result = ''
hidden = [tf.zeros((1, units))]
enc_out, enc_hidden = encoder(inputs, hidden)
dec_hidden = enc_hidden
dec_input = tf.expand_dims([targ_lang.word_index['<start>']], 0)
for t in range(max_length_targ):
predictions, dec_hidden, attention_weights = decoder(dec_input,
dec_hidden,
enc_out)
# storing the attention weights to plot later on
attention_weights = tf.reshape(attention_weights, (-1, ))
attention_plot[t] = attention_weights.numpy()
predicted_id = tf.argmax(predictions[0]).numpy()
result += targ_lang.index_word[predicted_id] + ' '
if targ_lang.index_word[predicted_id] == '<end>':
return result, sentence, attention_plot
# the predicted ID is fed back into the model
dec_input = tf.expand_dims([predicted_id], 0)
return result, sentence, attention_plot
# function for plotting the attention weights
def plot_attention(attention, sentence, predicted_sentence):
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(1, 1, 1)
ax.matshow(attention, cmap='viridis')
fontdict = {'fontsize': 14}
ax.set_xticklabels([''] + sentence, fontdict=fontdict, rotation=90)
ax.set_yticklabels([''] + predicted_sentence, fontdict=fontdict)
ax.xaxis.set_major_locator(ticker.MultipleLocator(1))
ax.yaxis.set_major_locator(ticker.MultipleLocator(1))
plt.show()
def translate(sentence):
result, sentence, attention_plot = evaluate(sentence)
print('Input: %s' % (sentence))
print('Predicted translation: {}'.format(result))
attention_plot = attention_plot[:len(result.split(' ')), :len(sentence.split(' '))]
plot_attention(attention_plot, sentence.split(' '), result.split(' '))
```
## Restore the latest checkpoint and test
```
# restoring the latest checkpoint in checkpoint_dir
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
translate(u'hace mucho frio aqui.')
translate(u'esta es mi vida.')
translate(u'¿todavia estan en casa?')
# wrong translation
translate(u'trata de averiguarlo.')
```
## Next steps
* [Download a different dataset](http://www.manythings.org/anki/) to experiment with translations, for example, English to German, or English to French.
* Experiment with training on a larger dataset, or using more epochs
| github_jupyter |
Web Scraping
============
- HTTP requests
- HTML, XML, DOM, CSS selectors, XPath
- browser automation
- cleanse and export extracted data
Web-based (or browser-based) user interfaces are ubiquitous
- web browser as universal platform to run software (at least, the user interface)
- if a human is able to access information in WWW using a web browser,
also a computer program can access the same information and automatically extract it
- challenges: navigate a web page, execute user interaction (mouse clicks, forms)
- real challenges: login forms, captchas, IP blocking, etc.
- not covered here
- also: ethical considerations whether or not to get around access blocking
- well-defined technology stack
- HTTP
- HTML / XML
- DOM
- CSS
- XPath
- JavaScript
## Web Browser
- render HTML page to make it readable for humans
- basic navigation in the WWW (follow links)
- [text-based browsers](https://en.wikipedia.org/wiki/Text-based_web_browser)
```
lynx https://www.bundestag.de/parlament/fraktionen/cducsu
```
- modern graphical browsers
- interpret JavaScript
- show multi-media content
- run “web applications”
- headless vs. headful browsers
- headful: graphical user interface attached
- [headless](https://en.wikipedia.org/wiki/Headless_browser)
- controlled programmatically or via command-line
- interaction but no mandatory page rendering (saves resources: CPU, RAM)
### Tip: Extract Text and Links Using a Text-Based Browser
Tip: text-based browsers usually have an option to "dump" the text and/or link lists into a file, e.g.
```
lynx -dump https://www.bundestag.de/parlament/fraktionen/cducsu \
>data/bundestag/fraktionen.cducsu.txt
```
### Tip: Explore Web Pages and Web Technologies using the Developer Tool of your Web Browser
Modern web browsers (Firefox, Chromium, IE, etc.) include a set of [web development tools](https://en.wikipedia.org/wiki/Web_development_tools). Originally addressed to web developers to test and debug the code (HTML, CSS, Javascript) used to build a web site, the browser web developer tools are the easiest way to explore and understand the technologies used to build a web site. The initial exploration later helps to scrape data from the web site.
### Browser Automation
- load a page by URL including page dependencies (CSS, Javascript, images, media)
- simulate user interaction (clicks, input, scrolling)
- take screenshots
- access the DOM tree or the HTML modified by executed Javascript and user interactions
from/in the browser to extract data
## Process HTML Pages in Python
- [requests](https://pypi.org/project/requests/) to fetch pages via HTTP
- [beautifulsoup](https://pypi.org/project/beautifulsoup4/) to parse HTML
```
import requests
request_url = 'https://www.bundestag.de/parlament/fraktionen/cducsu'
response = requests.get(request_url)
response
response.headers
response.status_code
!pip install beautifulsoup4
from bs4 import BeautifulSoup
html = BeautifulSoup(response.text)
html.head.title # tree-style path addressing of HTML elements
```
Note: the HTML document can be represented as a tree structure aka. [DOM tree](https://en.wikipedia.org/wiki/Document_Object_Model):
```
html
├── head
│ ├── meta
│ │ └── @charset=utf-8
│ └── title
│ └── ...(text)
└── body
└── ...
```
The tree above is an equivalent representation for the HTML snippet
```html
<!DOCTYPE html>
<html>
<head>
<meta charset="utf-8" />
<title>Deutscher Bundestag - CDU/CSU-Fraktion</title>
</head>
<body>
...
</body>
</html>
```
```
# access the plain text of an HTML element
# (inside the opening and closing tag)
html.head.title.text
# beautifulsoup also allows to select elements by tag name without a tree-like path
html.find('title').text
# or if a tag is expected to appear multiple times:
# select all `a` elements and show the first three
html.findAll('a')[0:3]
# selection by CSS class name
html.find(class_='bt-standard-content')
html.find(class_='bt-standard-content').findAll('a')
# but we are also interested in the function of the members:
html.find(class_='bt-standard-content').findAll('h4')
from urllib.parse import urljoin
for role_node in html.find(class_='bt-standard-content').findAll('h4'):
role = role_node.text.rstrip(':')
for link_node in role_node.next_sibling.findAll('a'):
name = link_node.text
link = urljoin(request_url, link_node.get('href'))
print(role, name, link)
```
Now we put everything together, so that we can run this for all factions of the parliament:
- we use a function to
- fetch the page of the faction and
- extract the members from the page content
- iterate over all factions
- store the list of faction roles and MPs in a data frame and CSV
```
%%script false --no-raise-error
# uncomment the above instruction to run this code
# note: do not run the cell by default
# because sending 6 HTTP requests may take long
import requests
from time import sleep
from urllib.parse import urljoin
from bs4 import BeautifulSoup
import pandas as pd
request_base_url = 'https://www.bundestag.de/parlament/fraktionen/'
factions = 'cducsu spd fdp linke gruene afd'.split()
def get_members_of_faction(faction):
global request_base_url
url = request_base_url + faction
response = requests.get(url)
if not response.ok:
return
result = []
html = BeautifulSoup(response.text)
for role_node in html.find(class_='bt-standard-content').findAll('h4'):
role = role_node.text.strip().rstrip(':')
if not role_node.next_sibling:
continue
for link_node in role_node.next_sibling.findAll('a'):
name = link_node.text
link = urljoin(url, link_node.get('href'))
result.append([name, faction, role, link])
return result
faction_roles = []
for faction in factions:
if faction_roles:
# be polite and wait before the next request
sleep(5)
faction_roles += get_members_of_faction(faction)
df_faction_roles = pd.DataFrame(faction_roles, columns=['name', 'faction', 'role', 'link'])
df_faction_roles.to_csv('data/bundestag/faction_roles.csv')
import pandas as pd
df_faction_roles = pd.read_csv('data/bundestag/faction_roles.csv')
df_faction_roles.value_counts('faction')
# not all members of the parliament have a role in their faction
# and are listed on the landing page of the faction
df_faction_roles.shape
df_faction_roles[df_faction_roles['role'].str.startswith('Fraktionsvorsitz')]
# now let's try whether we can fetch the biography and other information of a single MP
member_url = df_faction_roles.loc[df_faction_roles['name']=='Andreas Jung','link'].values[0]
member_response = requests.get(member_url)
member_html = BeautifulSoup(member_response.text)
# let's try first using the CSS class "bundestag-standard-content"
for node in member_html.findAll(class_='bt-standard-content'):
print(node.text)
```
### Automatic Cleansing of Text
A trivial extraction of all text in the body of web page would include a lot of unwanted content (navigation menus, header, footer, side bars), the "main" content could be even only a small part in the middle of the page. There are heuristics and algorithms for automatic removal of "boilerplate" content:
- Mozilla Readability: the [reader view](https://support.mozilla.org/en-US/kb/firefox-reader-view-clutter-free-web-pages) of the Firefox browser
- originally implemented in JavaScript, see [Readability.js](https://github.com/mozilla/readability/blob/master/Readability.js)
- but there is a Python port - [ReadabiliPy](https://github.com/alan-turing-institute/ReadabiliPy) or [ReadabiliPy on pypi](https://pypi.org/project/readabilipy/)
- [jusText](https://nlp.fi.muni.cz/projects/justext/) or [jusText on pypi](https://pypi.org/project/jusText/)
Here an example usage of ReadabiliPy with the latest fetched page (without any manual selection of elements by CSS class):
```
!pip install readabilipy
from readabilipy import simple_json_from_html_string
article = simple_json_from_html_string(member_response.text, use_readability=True)
for paragraph in article['plain_text']:
print(paragraph['text'])
print()
# but there's also a "readable" and simple HTML snippet
# (shown as rendered HTML in the output)
from IPython.core.display import HTML
HTML(article['plain_content'])
```
## Processing XML
The [Open Data](https://www.bundestag.de/services/opendata) portal of the German parliament offers a zip file "Stammdaten aller Abgeordneten seit 1949 im XML-Format (Stand 12.03.2021)" for free download. Most likely we should get the information about all PMs from this source. But how do we process XML?
Assumed the zip archive has been downloaded, unzipped and the files are all placed in `data/bundestag/`, we can simply read the file and pass it to beautifulsoup which will parse it. But we request a specific parser feature (`lxml-xml`) so that the casing of XML elements is preserved.
```
from bs4 import BeautifulSoup
xml = BeautifulSoup(open('data/bundestag/MDB_STAMMDATEN.XML').read(),
features='lxml-xml')
xml.MDB
len(xml.findAll('MDB'))
from collections import Counter
mp_acad_title = Counter()
mp_with_acad_title, mp_total = 0, 0
for mp in xml.findAll('MDB'):
mp_total += 1
has_academic_title = False
for nn in mp.findAll("NAME"):
if nn.AKAD_TITEL.text:
has_academic_title = True
mp_acad_title[nn.AKAD_TITEL.text] += 1
if has_academic_title:
# count a title only once (in case of multiple names)
mp_with_acad_title += 1
mp_with_acad_title / mp_total
mp_acad_title.most_common()
```
A final note: Reading the XML file describing the members of the German parliament into a tabular data structure will be painful (similar as for JSON data source) because of
- the nested structure
- some list-like data, for example the fact that one MP can have multiple names
Instead of coding the conversion in Python: with [XSLT](https://en.wikipedia.org/wiki/XSLT) there is a dedicated language for transforming XML documents into other document formats.
The [Open Discourse](https://opendiscourse.de/) projects hosts the proceedings of the German parliament and also a list of MPs in data formats easy to consume. See the [Open Discourse data sets](https://dataverse.harvard.edu/dataverse/opendiscourse) page.
## Browser automation with Python
- [Selenium](https://pypi.org/project/selenium/)
- nice example: [impf-botpy](https://github.com/alfonsrv/impf-botpy)
- [Playwright](https://playwright.dev/python/docs/intro)
- [Playwright on pypi](https://pypi.org/project/playwright/) including nice examples (some cited below)
- [Python API docs](https://playwright.dev/python/docs/api/class-playwright)
Note: Playwright does not run in a Jupyter notebook. We'll run the scripts directly in the Python interpreter.
Installation:
```
pip install playwright
playwright install
```
Take a screenshot using two different browsers:
```python
from playwright.sync_api import sync_playwright
with sync_playwright() as p:
for browser_type in [p.chromium, p.firefox]:
browser = browser_type.launch()
page = browser.new_page()
page.goto('http://whatsmyuseragent.org/')
_ = page.screenshot(path=f'figures/example-{browser_type.name}.png')
browser.close()
```
Just run the script [scripts/playwright_whatsmyuseragent_screenshot.py](scripts/playwright_whatsmyuseragent_screenshot.py) in the console / shell:
```
python ./scripts/playwright_whatsmyuseragent_screenshot.py
```
The screenshots are then found in the folder `figures/` for [chromium](./figures/example-chromium.png) and [firefox](./figures/example-firefox.png).
Playwright can record user interactions (mouse clicks, keyboard input) and create Python code to replay the recorded actions:
```
playwright codegen https://www.bundestag.de/abgeordnete/biografien
```
The created Python code is then modified, here to loop over all overlays showing the members of the parliament:
```python
from time import sleep
from playwright.sync_api import sync_playwright
def run(playwright):
browser = playwright.chromium.launch(headless=False)
context = browser.new_context(viewport={'height': 1080, 'width': 1920})
page = context.new_page()
page.goto("https://www.bundestag.de/abgeordnete/biografien")
while True:
try:
sleep(3)
page.click("button:has-text(\"Vor\")")
except Exception:
break
with sync_playwright() as p:
run(p)
```
Again: best run the [replay script](./scripts/playwright_replay.py) in the console:
```
python ./scripts/playwright_replay.py
```
| github_jupyter |
```
import sys, os, os.path
import glob
import scipy as sp
import numpy as np
import matplotlib
import matplotlib.pyplot as pp
import yt
from yt.frontends.boxlib.data_structures import AMReXDataset
%pylab inline
# Try Cori's scratch... otherwise the user will have to manually input the data root
scratch_env_key = "SCRATCH"
if scratch_env_key in os.environ.keys():
data_root = os.environ[scratch_env_key]
else:
data_root = ""
print(RuntimeWarning("`SCRATCH` is not an environment variable => data_root is empty"))
data_dir = "."
data_path = os.path.join(data_root, data_dir)
step = 1
n_fill = 5
prefix = "plt"
file_fmt = prefix + "{:0" + str(n_fill) + "d}"
data_glob = os.path.join(data_path, prefix + "*")
data_files = glob.glob(data_glob)
data_files.sort()
def check_sequential(files_sorted, step):
sequential = True
for i in range(len(files_sorted)):
c_file = file_fmt.format(i*step)
c_path = os.path.join(data_path, c_file)
missing = list()
if c_path not in files_sorted:
missing.append(c_path)
sequential = False
return sequential, missing
check_sequential(data_files, step)
class SoA:
_pref = "particle_"
_pos = "position_"
_vel = "vel"
_mass = "mass"
def __init__(self, data):
str_pos = self._pref+self._pos
self.px = np.array(data[str_pos + "x"])
self.py = np.array(data[str_pos + "y"])
self.pz = np.array(data[str_pos + "z"])
str_vel = self._pref+self._vel
self.vx = np.array(data[str_vel + "x"])
self.vy = np.array(data[str_vel + "y"])
self.vz = np.array(data[str_vel + "z"])
self.mass = np.array(data[self._pref + self._mass])
def __str__(self):
return "{pos:" + str(self.px) + "," + str(self.py) + "," + str(self.pz) + \
"; vel:" + str(self.vx) + "," + str(self.vy) + "," + str(self.vz) + \
"; mass:" + str(self.mass) + "}"
def __repr__(self):
return str(self)
class Particle:
def __init__(self, px, py, pz, vx, vy, vz, mass):
self.pos = np.array([px, py, pz])
self.vel = np.array([vx, vy, vz])
self.mass = mass
def __str__(self):
return "P(" + str(self.pos) + "," + str(self.vel) + "," + str(self.mass) + ")"
def __repr__(self):
return str(self)
class AoS:
def __init__(self, amrex_data):
self.particles = list()
soa = SoA(amrex_data)
data = zip(soa.px, soa.py, soa.pz, soa.vx, soa.vy, soa.vz, soa.mass)
for elt in data:
self.particles.append(Particle(* elt))
ds = AMReXDataset(data_files[-1])
ds.particle_fields_by_type
ad = ds.all_data()
soa = SoA(ad)
aos = AoS(ad)
aos.particles
ad["particle_position_x"]
part_x = np.array(ad["particle_position_x"])
part_x
np.array([2])
ad["particle_position_y"][0]
ad["particle_position_z"][0]
ad["particle_velx"][0]
ad["particle_vely"][0]
ad["particle_velz"][0]
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Load CSV data
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/load_data/csv"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/load_data/csv.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/load_data/csv.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/load_data/csv.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
This tutorial provides an example of how to load CSV data from a file into a `tf.data.Dataset`.
The data used in this tutorial are taken from the Titanic passenger list. The model will predict the likelihood a passenger survived based on characteristics like age, gender, ticket class, and whether the person was traveling alone.
## Setup
```
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
from __future__ import absolute_import, division, print_function, unicode_literals
import functools
import numpy as np
import tensorflow as tf
TRAIN_DATA_URL = "https://storage.googleapis.com/tf-datasets/titanic/train.csv"
TEST_DATA_URL = "https://storage.googleapis.com/tf-datasets/titanic/eval.csv"
train_file_path = tf.keras.utils.get_file("train.csv", TRAIN_DATA_URL)
test_file_path = tf.keras.utils.get_file("eval.csv", TEST_DATA_URL)
# Make numpy values easier to read.
np.set_printoptions(precision=3, suppress=True)
```
## Load data
To start, let's look at the top of the CSV file to see how it is formatted.
```
!head {train_file_path}
```
You can [load this using pandas](pandas.ipynb), and pass the NumPy arrays to TensorFlow. If you need to scale up to a large set of files, or need a loader that integrates with [TensorFlow and tf.data](../../guide/data,ipynb) then use the `tf.data.experimental.make_csv_dataset` function:
The only column you need to identify explicitly is the one with the value that the model is intended to predict.
```
LABEL_COLUMN = 'survived'
LABELS = [0, 1]
```
Now read the CSV data from the file and create a dataset.
(For the full documentation, see `tf.data.experimental.make_csv_dataset`)
```
def get_dataset(file_path, **kwargs):
dataset = tf.data.experimental.make_csv_dataset(
file_path,
batch_size=5, # Artificially small to make examples easier to show.
label_name=LABEL_COLUMN,
na_value="?",
num_epochs=1,
ignore_errors=True,
**kwargs)
return dataset
raw_train_data = get_dataset(train_file_path)
raw_test_data = get_dataset(test_file_path)
def show_batch(dataset):
for batch, label in dataset.take(1):
for key, value in batch.items():
print("{:20s}: {}".format(key,value.numpy()))
```
Each item in the dataset is a batch, represented as a tuple of (*many examples*, *many labels*). The data from the examples is organized in column-based tensors (rather than row-based tensors), each with as many elements as the batch size (5 in this case).
It might help to see this yourself.
```
show_batch(raw_train_data)
```
As you can see, the columns in the CSV are named. The dataset constructor will pick these names up automatically. If the file you are working with does not contain the column names in the first line, pass them in a list of strings to the `column_names` argument in the `make_csv_dataset` function.
```
CSV_COLUMNS = ['survived', 'sex', 'age', 'n_siblings_spouses', 'parch', 'fare', 'class', 'deck', 'embark_town', 'alone']
temp_dataset = get_dataset(train_file_path, column_names=CSV_COLUMNS)
show_batch(temp_dataset)
```
This example is going to use all the available columns. If you need to omit some columns from the dataset, create a list of just the columns you plan to use, and pass it into the (optional) `select_columns` argument of the constructor.
```
SELECT_COLUMNS = ['survived', 'age', 'n_siblings_spouses', 'class', 'deck', 'alone']
temp_dataset = get_dataset(train_file_path, select_columns=SELECT_COLUMNS)
show_batch(temp_dataset)
```
## Data preprocessing
A CSV file can contain a variety of data types. Typically you want to convert from those mixed types to a fixed length vector before feeding the data into your model.
TensorFlow has a built-in system for describing common input conversions: `tf.feature_column`, see [this tutorial](../keras/feature_columns) for details.
You can preprocess your data using any tool you like (like [nltk](https://www.nltk.org/) or [sklearn](https://scikit-learn.org/stable/)), and just pass the processed output to TensorFlow.
The primary advantage of doing the preprocessing inside your model is that when you export the model it includes the preprocessing. This way you can pass the raw data directly to your model.
### Continuous data
If your data is already in an apropriate numeric format, you can pack the data into a vector before passing it off to the model:
```
SELECT_COLUMNS = ['survived', 'age', 'n_siblings_spouses', 'parch', 'fare']
DEFAULTS = [0, 0.0, 0.0, 0.0, 0.0]
temp_dataset = get_dataset(train_file_path,
select_columns=SELECT_COLUMNS,
column_defaults = DEFAULTS)
show_batch(temp_dataset)
example_batch, labels_batch = next(iter(temp_dataset))
```
Here's a simple function that will pack together all the columns:
```
def pack(features, label):
return tf.stack(list(features.values()), axis=-1), label
```
Apply this to each element of the dataset:
```
packed_dataset = temp_dataset.map(pack)
for features, labels in packed_dataset.take(1):
print(features.numpy())
print()
print(labels.numpy())
```
If you have mixed datatypes you may want to separate out these simple-numeric fields. The `tf.feature_column` api can handle them, but this incurs some overhead and should be avoided unless really necessary. Switch back to the mixed dataset:
```
show_batch(raw_train_data)
example_batch, labels_batch = next(iter(temp_dataset))
```
So define a more general preprocessor that selects a list of numeric features and packs them into a single column:
```
class PackNumericFeatures(object):
def __init__(self, names):
self.names = names
def __call__(self, features, labels):
numeric_freatures = [features.pop(name) for name in self.names]
numeric_features = [tf.cast(feat, tf.float32) for feat in numeric_freatures]
numeric_features = tf.stack(numeric_features, axis=-1)
features['numeric'] = numeric_features
return features, labels
NUMERIC_FEATURES = ['age','n_siblings_spouses','parch', 'fare']
packed_train_data = raw_train_data.map(
PackNumericFeatures(NUMERIC_FEATURES))
packed_test_data = raw_test_data.map(
PackNumericFeatures(NUMERIC_FEATURES))
show_batch(packed_train_data)
example_batch, labels_batch = next(iter(packed_train_data))
```
#### Data Normalization
Continuous data should always be normalized.
```
import pandas as pd
desc = pd.read_csv(train_file_path)[NUMERIC_FEATURES].describe()
desc
MEAN = np.array(desc.T['mean'])
STD = np.array(desc.T['std'])
def normalize_numeric_data(data, mean, std):
# Center the data
return (data-mean)/std
```
Now create a numeric column. The `tf.feature_columns.numeric_column` API accepts a `normalizer_fn` argument, which will be run on each batch.
Bind the `MEAN` and `STD` to the normalizer fn using [`functools.partial`](https://docs.python.org/3/library/functools.html#functools.partial).
```
# See what you just created.
normalizer = functools.partial(normalize_numeric_data, mean=MEAN, std=STD)
numeric_column = tf.feature_column.numeric_column('numeric', normalizer_fn=normalizer, shape=[len(NUMERIC_FEATURES)])
numeric_columns = [numeric_column]
numeric_column
```
When you train the model, include this feature column to select and center this block of numeric data:
```
example_batch['numeric']
numeric_layer = tf.keras.layers.DenseFeatures(numeric_columns)
numeric_layer(example_batch).numpy()
```
The mean based normalization used here requires knowing the means of each column ahead of time.
### Categorical data
Some of the columns in the CSV data are categorical columns. That is, the content should be one of a limited set of options.
Use the `tf.feature_column` API to create a collection with a `tf.feature_column.indicator_column` for each categorical column.
```
CATEGORIES = {
'sex': ['male', 'female'],
'class' : ['First', 'Second', 'Third'],
'deck' : ['A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I', 'J'],
'embark_town' : ['Cherbourg', 'Southhampton', 'Queenstown'],
'alone' : ['y', 'n']
}
categorical_columns = []
for feature, vocab in CATEGORIES.items():
cat_col = tf.feature_column.categorical_column_with_vocabulary_list(
key=feature, vocabulary_list=vocab)
categorical_columns.append(tf.feature_column.indicator_column(cat_col))
# See what you just created.
categorical_columns
categorical_layer = tf.keras.layers.DenseFeatures(categorical_columns)
print(categorical_layer(example_batch).numpy()[0])
```
This will be become part of a data processing input later when you build the model.
### Combined preprocessing layer
Add the two feature column collections and pass them to a `tf.keras.layers.DenseFeatures` to create an input layer that will extract and preprocess both input types:
```
preprocessing_layer = tf.keras.layers.DenseFeatures(categorical_columns+numeric_columns)
print(preprocessing_layer(example_batch).numpy()[0])
```
## Build the model
Build a `tf.keras.Sequential`, starting with the `preprocessing_layer`.
```
model = tf.keras.Sequential([
preprocessing_layer,
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid'),
])
model.compile(
loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
```
## Train, evaluate, and predict
Now the model can be instantiated and trained.
```
train_data = packed_train_data.shuffle(500)
test_data = packed_test_data
model.fit(train_data, epochs=20)
```
Once the model is trained, you can check its accuracy on the `test_data` set.
```
test_loss, test_accuracy = model.evaluate(test_data)
print('\n\nTest Loss {}, Test Accuracy {}'.format(test_loss, test_accuracy))
```
Use `tf.keras.Model.predict` to infer labels on a batch or a dataset of batches.
```
predictions = model.predict(test_data)
# Show some results
for prediction, survived in zip(predictions[:10], list(test_data)[0][1][:10]):
print("Predicted survival: {:.2%}".format(prediction[0]),
" | Actual outcome: ",
("SURVIVED" if bool(survived) else "DIED"))
```
| github_jupyter |
# Batch Normalization – Practice
Batch normalization is most useful when building deep neural networks. To demonstrate this, we'll create a convolutional neural network with 20 convolutional layers, followed by a fully connected layer. We'll use it to classify handwritten digits in the MNIST dataset, which should be familiar to you by now.
This is **not** a good network for classfying MNIST digits. You could create a _much_ simpler network and get _better_ results. However, to give you hands-on experience with batch normalization, we had to make an example that was:
1. Complicated enough that training would benefit from batch normalization.
2. Simple enough that it would train quickly, since this is meant to be a short exercise just to give you some practice adding batch normalization.
3. Simple enough that the architecture would be easy to understand without additional resources.
This notebook includes two versions of the network that you can edit. The first uses higher level functions from the `tf.layers` package. The second is the same network, but uses only lower level functions in the `tf.nn` package.
1. [Batch Normalization with `tf.layers.batch_normalization`](#example_1)
2. [Batch Normalization with `tf.nn.batch_normalization`](#example_2)
The following cell loads TensorFlow, downloads the MNIST dataset if necessary, and loads it into an object named `mnist`. You'll need to run this cell before running anything else in the notebook.
```
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True, reshape=False)
```
# Batch Normalization using `tf.layers.batch_normalization`<a id="example_1"></a>
This version of the network uses `tf.layers` for almost everything, and expects you to implement batch normalization using [`tf.layers.batch_normalization`](https://www.tensorflow.org/api_docs/python/tf/layers/batch_normalization)
We'll use the following function to create fully connected layers in our network. We'll create them with the specified number of neurons and a ReLU activation function.
This version of the function does not include batch normalization.
```
"""
DO NOT MODIFY THIS CELL
"""
def fully_connected(prev_layer, num_units):
"""
Create a fully connectd layer with the given layer as input and the given number of neurons.
:param prev_layer: Tensor
The Tensor that acts as input into this layer
:param num_units: int
The size of the layer. That is, the number of units, nodes, or neurons.
:returns Tensor
A new fully connected layer
"""
layer = tf.layers.dense(prev_layer, num_units, activation=tf.nn.relu)
return layer
```
We'll use the following function to create convolutional layers in our network. They are very basic: we're always using a 3x3 kernel, ReLU activation functions, strides of 1x1 on layers with odd depths, and strides of 2x2 on layers with even depths. We aren't bothering with pooling layers at all in this network.
This version of the function does not include batch normalization.
```
"""
DO NOT MODIFY THIS CELL
"""
def conv_layer(prev_layer, layer_depth):
"""
Create a convolutional layer with the given layer as input.
:param prev_layer: Tensor
The Tensor that acts as input into this layer
:param layer_depth: int
We'll set the strides and number of feature maps based on the layer's depth in the network.
This is *not* a good way to make a CNN, but it helps us create this example with very little code.
:returns Tensor
A new convolutional layer
"""
strides = 2 if layer_depth % 3 == 0 else 1
conv_layer = tf.layers.conv2d(prev_layer, layer_depth*4, 3, strides, 'same', activation=tf.nn.relu)
return conv_layer
```
**Run the following cell**, along with the earlier cells (to load the dataset and define the necessary functions).
This cell builds the network **without** batch normalization, then trains it on the MNIST dataset. It displays loss and accuracy data periodically while training.
```
"""
DO NOT MODIFY THIS CELL
"""
def train(num_batches, batch_size, learning_rate):
# Build placeholders for the input samples and labels
inputs = tf.placeholder(tf.float32, [None, 28, 28, 1])
labels = tf.placeholder(tf.float32, [None, 10])
# Feed the inputs into a series of 20 convolutional layers
layer = inputs
for layer_i in range(1, 20):
layer = conv_layer(layer, layer_i)
# Flatten the output from the convolutional layers
orig_shape = layer.get_shape().as_list()
layer = tf.reshape(layer, shape=[-1, orig_shape[1] * orig_shape[2] * orig_shape[3]])
# Add one fully connected layer
layer = fully_connected(layer, 100)
# Create the output layer with 1 node for each
logits = tf.layers.dense(layer, 10)
# Define loss and training operations
model_loss = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=logits, labels=labels))
train_opt = tf.train.AdamOptimizer(learning_rate).minimize(model_loss)
# Create operations to test accuracy
correct_prediction = tf.equal(tf.argmax(logits,1), tf.argmax(labels,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
# Train and test the network
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for batch_i in range(num_batches):
batch_xs, batch_ys = mnist.train.next_batch(batch_size)
# train this batch
sess.run(train_opt, {inputs: batch_xs, labels: batch_ys})
# Periodically check the validation or training loss and accuracy
if batch_i % 100 == 0:
loss, acc = sess.run([model_loss, accuracy], {inputs: mnist.validation.images,
labels: mnist.validation.labels})
print('Batch: {:>2}: Validation loss: {:>3.5f}, Validation accuracy: {:>3.5f}'.format(batch_i, loss, acc))
elif batch_i % 25 == 0:
loss, acc = sess.run([model_loss, accuracy], {inputs: batch_xs, labels: batch_ys})
print('Batch: {:>2}: Training loss: {:>3.5f}, Training accuracy: {:>3.5f}'.format(batch_i, loss, acc))
# At the end, score the final accuracy for both the validation and test sets
acc = sess.run(accuracy, {inputs: mnist.validation.images,
labels: mnist.validation.labels})
print('Final validation accuracy: {:>3.5f}'.format(acc))
acc = sess.run(accuracy, {inputs: mnist.test.images,
labels: mnist.test.labels})
print('Final test accuracy: {:>3.5f}'.format(acc))
# Score the first 100 test images individually. This won't work if batch normalization isn't implemented correctly.
correct = 0
for i in range(100):
correct += sess.run(accuracy,feed_dict={inputs: [mnist.test.images[i]],
labels: [mnist.test.labels[i]]})
print("Accuracy on 100 samples:", correct/100)
num_batches = 800
batch_size = 64
learning_rate = 0.002
tf.reset_default_graph()
with tf.Graph().as_default():
train(num_batches, batch_size, learning_rate)
```
With this many layers, it's going to take a lot of iterations for this network to learn. By the time you're done training these 800 batches, your final test and validation accuracies probably won't be much better than 10%. (It will be different each time, but will most likely be less than 15%.)
Using batch normalization, you'll be able to train this same network to over 90% in that same number of batches.
# Add batch normalization
We've copied the previous three cells to get you started. **Edit these cells** to add batch normalization to the network. For this exercise, you should use [`tf.layers.batch_normalization`](https://www.tensorflow.org/api_docs/python/tf/layers/batch_normalization) to handle most of the math, but you'll need to make a few other changes to your network to integrate batch normalization. You may want to refer back to the lesson notebook to remind yourself of important things, like how your graph operations need to know whether or not you are performing training or inference.
If you get stuck, you can check out the `Batch_Normalization_Solutions` notebook to see how we did things.
**TODO:** Modify `fully_connected` to add batch normalization to the fully connected layers it creates. Feel free to change the function's parameters if it helps.
```
def fully_connected(prev_layer, num_units, training_bool):
"""
Create a fully connectd layer with the given layer as input and the given number of neurons.
:param prev_layer: Tensor
The Tensor that acts as input into this layer
:param num_units: int
The size of the layer. That is, the number of units, nodes, or neurons.
:returns Tensor
A new fully connected layer
"""
layer_temp = tf.layers.dense(prev_layer,num_units,activation = None, use_bias = False)
batch_norm_output = tf.layers.batch_normalization(layer_temp, training = training_bool)
layer = tf.nn.relu(batch_norm_output)
return layer
```
**TODO:** Modify `conv_layer` to add batch normalization to the convolutional layers it creates. Feel free to change the function's parameters if it helps.
```
def conv_layer(prev_layer, layer_depth, training_bool):
"""
Create a convolutional layer with the given layer as input.
:param prev_layer: Tensor
The Tensor that acts as input into this layer
:param layer_depth: int
We'll set the strides and number of feature maps based on the layer's depth in the network.
This is *not* a good way to make a CNN, but it helps us create this example with very little code.
:returns Tensor
A new convolutional layer
"""
strides = 2 if layer_depth % 3 == 0 else 1
conv_layer_temp = tf.layers.conv2d(prev_layer, layer_depth*4, 3, strides, 'same', activation=None, use_bias=False)# See Batch_Normalization_Solutions for alternate implementations, including with batch norm after nonlinear activation function and/or with bias
batch_norm_output = tf.layers.batch_normalization(conv_layer_temp,training=training_bool)
conv_layer = tf.nn.relu(batch_norm_output)
return conv_layer
```
**TODO:** Edit the `train` function to support batch normalization. You'll need to make sure the network knows whether or not it is training, and you'll need to make sure it updates and uses its population statistics correctly.
```
def train(num_batches, batch_size, learning_rate):
# Build placeholders for the input samples and labels
inputs = tf.placeholder(tf.float32, [None, 28, 28, 1])
labels = tf.placeholder(tf.float32, [None, 10])
# Placeholder to indicate whether or not training
training_bool = tf.placeholder(tf.bool)
# Feed the inputs into a series of 20 convolutional layers
layer = inputs
for layer_i in range(1, 20):
layer = conv_layer(layer, layer_i, training_bool)
# Flatten the output from the convolutional layers
orig_shape = layer.get_shape().as_list()
layer = tf.reshape(layer, shape=[-1, orig_shape[1] * orig_shape[2] * orig_shape[3]])
# Add one fully connected layer
layer = fully_connected(layer, 100, training_bool)
# Create the output layer with 1 node for each
logits = tf.layers.dense(layer, 10)
# Define loss and training operations
model_loss = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=logits, labels=labels))
#The with statement tells tensorflow to update population stats while training
with tf.control_dependencies(tf.get_collection(tf.GraphKeys.UPDATE_OPS)):
train_opt = tf.train.AdamOptimizer(learning_rate).minimize(model_loss)
# Create operations to test accuracy
correct_prediction = tf.equal(tf.argmax(logits,1), tf.argmax(labels,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
# Train and test the network
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for batch_i in range(num_batches):
batch_xs, batch_ys = mnist.train.next_batch(batch_size)
# train this batch
sess.run(train_opt, {inputs: batch_xs, labels: batch_ys, training_bool: True})
# Periodically check the validation or training loss and accuracy
if batch_i % 100 == 0:
loss, acc = sess.run([model_loss, accuracy], {inputs: mnist.validation.images,
labels: mnist.validation.labels, training_bool: False})
print('Batch: {:>2}: Validation loss: {:>3.5f}, Validation accuracy: {:>3.5f}'.format(batch_i, loss, acc))
elif batch_i % 25 == 0:
loss, acc = sess.run([model_loss, accuracy], {inputs: batch_xs, labels: batch_ys, training_bool: False})
print('Batch: {:>2}: Training loss: {:>3.5f}, Training accuracy: {:>3.5f}'.format(batch_i, loss, acc))
# At the end, score the final accuracy for both the validation and test sets
acc = sess.run(accuracy, {inputs: mnist.validation.images,
labels: mnist.validation.labels, training_bool: False})
print('Final validation accuracy: {:>3.5f}'.format(acc))
acc = sess.run(accuracy, {inputs: mnist.test.images,
labels: mnist.test.labels, training_bool: False})
print('Final test accuracy: {:>3.5f}'.format(acc))
# Score the first 100 test images individually. This won't work if batch normalization isn't implemented correctly.
correct = 0
for i in range(100):
correct += sess.run(accuracy,feed_dict={inputs: [mnist.test.images[i]],
labels: [mnist.test.labels[i]],
training_bool: False})
print("Accuracy on 100 samples:", correct/100)
num_batches = 800
batch_size = 64
learning_rate = 0.002
tf.reset_default_graph()
with tf.Graph().as_default():
train(num_batches, batch_size, learning_rate)
```
With batch normalization, you should now get an accuracy over 90%. Notice also the last line of the output: `Accuracy on 100 samples`. If this value is low while everything else looks good, that means you did not implement batch normalization correctly. Specifically, it means you either did not calculate the population mean and variance while training, or you are not using those values during inference.
# Batch Normalization using `tf.nn.batch_normalization`<a id="example_2"></a>
Most of the time you will be able to use higher level functions exclusively, but sometimes you may want to work at a lower level. For example, if you ever want to implement a new feature – something new enough that TensorFlow does not already include a high-level implementation of it, like batch normalization in an LSTM – then you may need to know these sorts of things.
This version of the network uses `tf.nn` for almost everything, and expects you to implement batch normalization using [`tf.nn.batch_normalization`](https://www.tensorflow.org/api_docs/python/tf/nn/batch_normalization).
**Optional TODO:** You can run the next three cells before you edit them just to see how the network performs without batch normalization. However, the results should be pretty much the same as you saw with the previous example before you added batch normalization.
**TODO:** Modify `fully_connected` to add batch normalization to the fully connected layers it creates. Feel free to change the function's parameters if it helps.
**Note:** For convenience, we continue to use `tf.layers.dense` for the `fully_connected` layer. By this point in the class, you should have no problem replacing that with matrix operations between the `prev_layer` and explicit weights and biases variables.
```
def fully_connected(prev_layer, num_units, training_bool):
"""
Create a fully connectd layer with the given layer as input and the given number of neurons.
:param prev_layer: Tensor
The Tensor that acts as input into this layer
:param num_units: int
The size of the layer. That is, the number of units, nodes, or neurons.
:returns Tensor
A new fully connected layer
"""
# See the notebook Batch_Normalization_lesson for extensive notes for settig ths up
layer_temp = tf.layers.dense(prev_layer, num_units, activation=None, use_bias = False)
# Batch normalization has trainable variables gamma for scaling and beta for shifting
gamma = tf.Variable(tf.ones([num_units]))
beta = tf. Variable(tf.ones([num_units]))
# Small value for bacth nor to make sure we don't divide by zero
epsilon = 1e-3
# Variables for the mean and variance necessary for tf.nn.batch_normalization. Trainable set to false to tell tensorflow not to modify these variables during back prop
pop_mean = tf.Variable(tf.zeros([num_units]), trainable = False)
pop_variance = tf.Variable(tf.ones([num_units]), trainable = False)
def batch_norm_training():
# Calculating mean and variance coming out layer's linear-combination step
batch_mean, batch_variance = tf.nn.moments(layer_temp, [0])
# Calculating moving average of training data's mean and variance while training to be used during inference
decay = 0.99# has to be less than 1 apparently, it's like the momentum parameter in tf.layers.batch_normalization
train_mean = tf.assign(pop_mean, pop_mean*decay + batch_mean*(1-decay))
train_variance = tf.assign(pop_variance, pop_variance*decay + batch_variance*(1-decay))
with tf.control_dependencies([train_mean, train_variance]):
return tf.nn.batch_normalization(layer_temp, batch_mean, batch_variance, beta, gamma, epsilon)
def batch_norm_inference():
return tf.nn.batch_normalization(layer_temp, pop_mean, pop_variance, beta, gamma, epsilon)
batch_norm_output = tf.cond(training_bool, batch_norm_training, batch_norm_inference)
layer = tf.nn.relu(batch_norm_output)
return layer
```
**TODO:** Modify `conv_layer` to add batch normalization to the fully connected layers it creates. Feel free to change the function's parameters if it helps.
**Note:** Unlike in the previous example that used `tf.layers`, adding batch normalization to these convolutional layers _does_ require some slight differences to what you did in `fully_connected`.
```
def conv_layer(prev_layer, layer_depth, training_bool):
"""
Create a convolutional layer with the given layer as input.
:param prev_layer: Tensor
The Tensor that acts as input into this layer
:param layer_depth: int
We'll set the strides and number of feature maps based on the layer's depth in the network.
This is *not* a good way to make a CNN, but it helps us create this example with very little code.
:returns Tensor
A new convolutional layer
"""
strides = 2 if layer_depth % 3 == 0 else 1
in_channels = prev_layer.get_shape().as_list()[3]
out_channels = layer_depth*4
weights = tf.Variable(
tf.truncated_normal([3, 3, in_channels, out_channels], stddev=0.05))
conv_layer = tf.nn.conv2d(prev_layer, weights, strides=[1,strides, strides, 1], padding='SAME')
gamma = tf.Variable(tf.ones([out_channels]))
beta = tf.Variable(tf.zeros([out_channels]))
epsilon = 1e-3
pop_mean = tf.Variable(tf.zeros([out_channels]), trainable=False)
pop_variance = tf.Variable(tf.ones([out_channels]), trainable=False)
def batch_norm_training():
#Must make sure to use correct dimensions here
batch_mean, batch_variance = tf.nn.moments(conv_layer, [0,1,2], keep_dims=False)
decay = 0.99
train_mean = tf.assign(pop_mean, pop_mean*decay + batch_mean*(1-decay))
train_variance = tf.assign(pop_variance, pop_varianc*decay + batch_variance*(1-decay))
with tf.control_dependencies([train_mean, train_variance]):
return tf.nn.batch_normalization(conv_layer, batch_mean, batch_variance, beta, gamma, epsilon)
def batch_norm_inference():
return tf.nn.batch_normalization(conv_layer, pop_mean, pop_variance, beta, gamma, epsilon)
batch_norm_output = tf.cond(training_bool, batch_norm_training, batch_norm_inference)
conv_layer = tf.nn.relu(conv_layer)
return conv_layer
```
**TODO:** Edit the `train` function to support batch normalization. You'll need to make sure the network knows whether or not it is training.
```
def train(num_batches, batch_size, learning_rate):
# Build placeholders for the input samples and labels
inputs = tf.placeholder(tf.float32, [None, 28, 28, 1])
labels = tf.placeholder(tf.float32, [None, 10])
# Placeholder to determine whether or not training
training_bool = tf.placeholder(tf.bool)
# Feed the inputs into a series of 20 convolutional layers
layer = inputs
for layer_i in range(1, 20):
layer = conv_layer(layer, layer_i, training_bool)
# Flatten the output from the convolutional layers
orig_shape = layer.get_shape().as_list()
layer = tf.reshape(layer, shape=[-1, orig_shape[1] * orig_shape[2] * orig_shape[3]])
# Add one fully connected layer
layer = fully_connected(layer, 100, training_bool)
# Create the output layer with 1 node for each
logits = tf.layers.dense(layer, 10)
# Define loss and training operations
model_loss = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=logits, labels=labels))
train_opt = tf.train.AdamOptimizer(learning_rate).minimize(model_loss)
# Create operations to test accuracy
correct_prediction = tf.equal(tf.argmax(logits,1), tf.argmax(labels,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
# Train and test the network
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for batch_i in range(num_batches):
batch_xs, batch_ys = mnist.train.next_batch(batch_size)
# train this batch
sess.run(train_opt, {inputs: batch_xs, labels: batch_ys, training_bool: True})
# Periodically check the validation or training loss and accuracy
if batch_i % 100 == 0:
loss, acc = sess.run([model_loss, accuracy], {inputs: mnist.validation.images,
labels: mnist.validation.labels, training_bool: False})
print('Batch: {:>2}: Validation loss: {:>3.5f}, Validation accuracy: {:>3.5f}'.format(batch_i, loss, acc))
elif batch_i % 25 == 0:
loss, acc = sess.run([model_loss, accuracy], {inputs: batch_xs, labels: batch_ys, training_bool: False})
print('Batch: {:>2}: Training loss: {:>3.5f}, Training accuracy: {:>3.5f}'.format(batch_i, loss, acc))
# At the end, score the final accuracy for both the validation and test sets
acc = sess.run(accuracy, {inputs: mnist.validation.images,
labels: mnist.validation.labels, training_bool: False})
print('Final validation accuracy: {:>3.5f}'.format(acc))
acc = sess.run(accuracy, {inputs: mnist.test.images,
labels: mnist.test.labels, training_bool: False})
print('Final test accuracy: {:>3.5f}'.format(acc))
# Score the first 100 test images individually. This won't work if batch normalization isn't implemented correctly.
correct = 0
for i in range(100):
correct += sess.run(accuracy,feed_dict={inputs: [mnist.test.images[i]],
labels: [mnist.test.labels[i]], training_bool: False})
print("Accuracy on 100 samples:", correct/100)
num_batches = 800
batch_size = 64
learning_rate = 0.002
tf.reset_default_graph()
with tf.Graph().as_default():
train(num_batches, batch_size, learning_rate)
```
Once again, the model with batch normalization should reach an accuracy over 90%. There are plenty of details that can go wrong when implementing at this low level, so if you got it working - great job! If not, do not worry, just look at the `Batch_Normalization_Solutions` notebook to see what went wrong.
| github_jupyter |
# Enrichment analysis: TORUS and GREGOR
This workflow document has several pipelines written in SoS Workflow language to perform data preparation and analysis for enrichment of GWAS signals in given annotation features.
For enrichment analysis currently it implements the `torus` pipeline, based on some `snakemake` codes originally written by Jean at Xin He Lab, UChicago.
```
%revisions -s -n 10
sos run gwas_enrichment.ipynb -h
```
## Copy this GIT depository
```
git clone https://github.com/cumc/bioworkflows
```
## To run enrichment analysis (overview)
### Step 1 (start from `bed` format annotation)
```
sos run fine-mapping/gwas_enrichment.ipynb range2var_annotation --z-score ... --single-annot ... --multi-annot ...
```
### Step 2 (start from binary annotation for SNPs)
```
sos run fine-mapping/gwas_enrichment.ipynb enrichment --z-score ... --single-annot ... --multi-annot ...
```
## Software intallation
### `SoS`
SoS introduction [webpage](https://vatlab.github.io/sos-docs/)
Installation:
```
pip install sos sos-notebook
```
SoS basic usage [webpage](https://vatlab.github.io/sos-docs/running.html#content)
### `docker`
Some steps in this pipeline uses docker images we prepare. If you do not have docker installed, you can install it via:
- Run commands below:
```
curl -fsSL get.docker.com -o get-docker.sh
sudo sh get-docker.sh
sudo usermod -aG docker $USER
```
- Log out and log back in (no need to reboot computer).
**In this pipeline, `bedops` and `torus` is executed via docker image we prepared and uploaded to dockerhub.com. The pipeline should automatically download the image and run the docker container instance.**
## Reference data preparation
### hg19.fa
- only for `deepsea` and `gregor` steps; if you only use enrichment analysis, you do not need to download it.
- Genome Reference: hg19 (GRCh37) or hg38 (GRCh38).
- Download [link](http://hgdownload.cse.ucsc.edu/goldenPath/hg19/bigZips/hg19.fa.gz)
## Annotation files
### Annotation files download
Alkes Price's lab has some annotation files in `bed` format available for download:
https://data.broadinstitute.org/alkesgroup/LDSCORE/
eg `baselineLD_v2.2_bedfiles.tgz`.
### `bed` format annotation input
**If you use the type of annotation file below, you need to run step 1 and 2.**
Annotation files are in `bed` format (for example: Promoter_UCSC.bed).
chr1 9873 16361
chr1 32610 36610
chr1 67090 71090
...
### Binary annotations of interest
**If you use the type of annotation file below, you only need to run step 2.**
0/1 binary indicated. For example, a few lines in `Coding_UCSC_annotation.torus.gz`.
SNP Coding_UCSC_d
9:140192349:G:C 0
9:140192576:C:T 0
9:140194020:G:A 0
9:140194735:G:T 1
9:140194793:A:G 1
9:140195019:C:A 1
There are many annotations one can use. For this workflow you should prepare two lists:
- For the ones that you'd like to use for one variable `torus`, put their file name prefix in `data/single_annotations.txt`
- For the ones that you'd like to use for multi-variable `torus`, put their file name prefix in `data/multi_annotations.txt`
If you want to remove one annotation just use `#` to comment it out in the list files.
```
%preview ../data/general_annotations.txt -l 10
```
## `torus` workflow data format
### GWAS z-scores specification
Example GWAS data (European)
- [SCZ Sweden data](https://www.med.unc.edu/pgc/results-and-downloads/data-use-agreement-forms/SwedenSCZ_data_download_agreement)
- [PTSD European Ancestry](https://www.med.unc.edu/pgc/results-and-downloads/data-use-agreement-forms/PTSD%20EA_data_download_agreement)
Format `chr:pos:alt:ref ld_label z-score`. 3 columns in total: `chr:pos:alt:ref` (chromosome, position, alternative allele and reference allele), `ld_label` (LD chunk label) and `z-score` (summary statistics).
```
1:10583:A:G 1 0.116319
...
22:51228910:A:G 1703 -0.866894
```
where the 2nd column is the analysis block ID (see below). All GWAS summary statistics have to be converted to this format in order to use.
### LD block files
The last column indicates LD chunk. LD chunk blocks depend on chromosome and position. There are 1703 LD chunks in European human genome.
```
chr1 10583 1892607 1
chr1 1892607 3582736 2
...
chr22 49824534 51243298 1703
```
- LD block reference paper [link](https://academic.oup.com/bioinformatics/article/32/2/283/1743626).
### SNP map
GWAS SNP map. Identical number of SNPs with GWAS.
Format `chrom.pos chr pos`. 3 columns in total: `chrom.pos` (example: `chr1.729679`), `chr` and `pos`.
```
chr1.729679 1 729679
chr1.731718 1 731718
chr1.734349 1 734349
...
```
### Minimal working example for `torus` workflow
- GWAS z-score example [link](https://www.dropbox.com/s/y2q29a71bb2wmr1/SCZGWAS.zscore.gz?dl=0)
- SNP map example [link](https://www.dropbox.com/s/oi4yr0au3oy3hi2/smap.gz?dl=0)
- annotation example: `Promoter_UCSC.bed` download [link](https://www.dropbox.com/s/huo5k78wptxmrpe/Promoter_UCSC.bed?dl=0). 22,436 lines in total.
- analysis block example: [link](https://www.dropbox.com/s/up78cnpkpoodark/ld_chunk.bed?dl=0). Used to identify LD chunk for each GWAS SNP.
## Workflow
**You can edit and change the following 5 directories and file pathes.**
First edit and run the following 5 command to define bash variables.
```
work_dir=~/Documents/GWAS_ATAC
anno_dir=~/Documents/GWAS_ATAC/bed_annotations
z_file=~/Documents/GWAS_ATAC/SCZGWAS_zscore/SCZGWAS.zscore.gz
single=~/GIT/fine-mapping/data/general_annotations.txt
blk=~/Documents/GWAS_ATAC/blocks.txt
snps=~/Documents/GWAS_ATAC/SCZGWAS_zscore/SCZGWAS.snps.gz
```
Then run following commands.
```
sos run fine-mapping/gwas_enrichment.ipynb range2var_annotation --cwd $work_dir --annotation_dir $anno_dir --z-score $z_file --single-annot $single
sos run fine-mapping/gwas_enrichment.ipynb enrichment --cwd $work_dir --annotation_dir $anno_dir --z-score $z_file --single-annot $single --blocks $blk --snps $snps
```
```
[global]
# working directory
parameter: cwd = path()
# hg19 path
parameter: hg19 = path()
# Deepsea model for use with `deepsee_apply`
parameter: deepsea_model = path()
# Path to directory of annotation files
parameter: annotation_dir = path() # 2
# Path to z-score file
parameter: z_score = path() # 3
# Path to list of single annotations to use
parameter: single_annot = path() # 4
# Path to lists of multi-annotations to use
parameter: multi_annot = paths() # 5
def get_bed_name(directory, basename):
filename = path(f"{directory}/{basename}.bed")
if not filename.is_file():
raise ValueError(f"Cannot find file ``{directory}/{basename}.bed``")
return filename
try:
single_anno = [get_bed_name(f"{annotation_dir:a}", x.split()[0]) for x in open(single_annot).readlines() if not x.startswith('#')]
multi_anno = dict([(f'{y:bn}', [get_bed_name(f"{annotation_dir:a}", x.split()[0]) for x in open(y).readlines() if not x.startswith('#')]) for y in multi_annot])
except (FileNotFoundError, IsADirectoryError):
single_anno = []
multi_anno = dict()
out_dir = f'{cwd:a}/{z_score:bn}'.replace('.', '_')
```
## Utility steps
### Convert variants from z-score file to bed format
```
# Auxiliary step to get variant in bed format based on variant ID in z-score file
[zscore2bed_1]
parameter: in_file = path()
input: in_file
output: f'{_input:n}.bed.unsorted'
R: expand = "${ }", container = 'gaow/atac-gwas', workdir = cwd, stdout = f'{_output:n}.stdout'
library(readr)
library(stringr)
library(dplyr)
var_file <- ${_input:r}
out_file <- ${_output:r}
variants <- read_tsv(var_file, col_names=FALSE, col_types='ccd')
colnames(variants) = c('variant', 'ld', 'zscore')
var_info <- str_split(variants$variant, ":")
variants <- mutate(variants, chr = paste0("chr", sapply(var_info, function(x){x[1]})),
pos = sapply(var_info, function(x){x[2]})) %>%
mutate(start = as.numeric(pos), stop=as.numeric(pos) + 1) %>%
select(chr, start, stop, variant)
options(scipen=1000) # So that positions are always fully written out)
write.table(variants, file=out_file, quote=FALSE, col.names=FALSE, row.names=FALSE, sep="\t")
[zscore2bed_2]
output: f'{_input:n}'
bash: expand = True, container = 'gaow/atac-gwas', workdir = cwd
sort-bed {_input} > {_output}
[get_variants: provides = '{data}.bed']
output: f'{data}.bed'
sos_run('zscore2bed', in_file = f'{_output:n}.gz')
```
### Merge annotations for joint torus analysis
```
# Auxiliary step to merge multiple annotations
[merge_annotations]
parameter: out_file=path()
parameter: data_files=paths()
input: data_files
output: out_file
R: expand = '${ }', container = 'gaow/atac-gwas', workdir = cwd, stdout = f'{_output:n}.stdout'
library(readr)
library(dplyr)
library(stringr)
out_name <- ${_output:r}
annots <- c(${_input:r,})
var_annot <- read_tsv(annots[1], col_names=T)
for(i in 2:length(annots)){
var_annot2 <- read_tsv(annots[i], col_names=T)
stopifnot(all(var_annot$SNP == var_annot2$SNP))
var_annot <- cbind(var_annot, var_annot2[,2])
}
write.table(var_annot, file=gzfile(out_name),
row.names=FALSE, quote=FALSE, sep="\t")
```
## Prepare `torus` format binary annotations
```
# Get variants in data that falls in target region
[range2var_annotation_1]
depends: f'{z_score:n}.bed'
input: set(paths(single_anno + list(multi_anno.values()))), group_by = 1
output: f'{out_dir}/{_input:bn}.{z_score:bn}.bed'
bash: expand = True, container = 'gaow/atac-gwas', workdir = cwd, volumes = [f'{_input:d}:{_input:d}', f'{_output:d}:{_output:d}']
bedops -e {z_score:n}.bed {_input} > {_output}
# Make binary annotation file
[range2var_annotation_2]
parameter: discrete = 1
depends: z_score
output: f'{_input:n}.gz'
R: expand = "${ }", container = 'gaow/atac-gwas', workdir = cwd, stdout = f'{_output:n}.stdout', volumes = [f'{_input:d}:{_input:d}', f'{_output:d}:{_output:d}']
library(readr)
library(dplyr)
library(stringr)
variant_tsv <- ${z_score:r}
annotation_var_bed <- ${_input:r}
annot_name <- ${_input:bnr} %>% str_replace(paste0(".",${z_score:bnr}), "")
out_name <- ${_output:r}
vars <- read_tsv(variant_tsv, col_names=FALSE, col_types='ccd')[,1]
annot_vars = read_tsv(annotation_var_bed, col_names=FALSE, col_types='cddc')
names(vars) <- "SNP"
vars <- vars %>%
mutate(annot_d = case_when(SNP %in% annot_vars$X4 ~ 1,
TRUE ~ 0))
names(vars)[2] <- paste0(annot_name, '${"_d" if discrete else "_c"}')
write.table(vars, file=gzfile(out_name),
col.names=TRUE, row.names=FALSE, sep="\t", quote=FALSE)
# Obtain multiple annotations per SNP for enrichment analysis
[range2var_annotation_3]
for k, value in multi_anno.items():
sos_run('merge_annotations', out_file = f'{out_dir}/{k}.{z_score:bn}.gz', data_files = [f'{out_dir}/{v:bn}.{z_score:bn}.gz' for v in paths(value)])
```
## Enrichment analysis via `torus`
```
# Run torus with annotation + z-score file
[enrichment_1 (run torus)]
depends: z_score
parameter: blocks = path()
parameter: snps = path()
input_files = [f'{out_dir}/{value:bn}.{z_score:bn}.gz' for value in paths(single_anno)] + [f'{out_dir}/{k}.{z_score:bn}.gz' for k in multi_anno]
fail_if(len(input_files) == 0, msg = "No annotations to use! Please use ``--single-annot`` and ``--multi-annot`` to pass annotation lists")
input: input_files, group_by = 1
output: f'{_input:n}.torus'
bash: container = 'gaow/atac-gwas', workdir = cwd, expand = True, stdout = f'{_output:n}.stdout', stderr = f'{_output:n}.stderr', volumes = [f'{_input:d}:{_input:d}', f'{_output:d}:{_output:d}']
rm -rf {_output:n}_prior
torus -d {z_score} -smap {snps} -gmap {blocks} -annot {_input} -est --load_zval -dump_prior {_output:n}_prior > {_output}
# Consolidate all torus result into one table
[enrichment_2 (make output table)]
input: group_by = 'all'
output: f'{out_dir}/{z_score:bn}.torus.merged.csv'
python: expand = '${ }', workdir = cwd
# get log2 folds of enrichment
import pandas as pd
def get_torus_res(fn):
import math, os
res = pd.read_table(fn, sep = "\s+", header = None, names = ["annotation", "log2_fold_enrichment", "CI_low", "CI_high"])
res = res.iloc[1:]
res["source"] = [os.path.split(fn)[-1] for x in range(res.shape[0])]
res["annotation"] = res["annotation"].apply(lambda x: x.rsplit('.',1)[0])
res["log2_fold_enrichment"] = [round(math.log2(math.exp(x)),4) for x in res["log2_fold_enrichment"]]
res["CI_low"] = [round(math.log2(math.exp(x)),4) for x in res["CI_low"]]
res["CI_high"] = [round(math.log2(math.exp(x)),4) for x in res["CI_high"]]
return res
res = pd.concat([get_torus_res(x) for x in [${_input:r,}]])
res.to_csv(${_output:r}, index=False)
```
## Apply pre-trained `deepsea` model to variants
Credits to Yanyu Liang. Required inputs are:
- Path to hg19 reference genome
- Path to model HDF5 file
- Path to list of variants, in the format of:
```
chr1 68090 68091 G T
chr1 68090 68091 G A
chr1 68090 68091 G C
chr1 68091 68092 C G
chr1 68091 68092 C T
chr1 68091 68092 C A
chr1 68092 68093 T A
chr1 68092 68093 T C
chr1 68092 68093 T G
chr1 68093 68094 A C
```
To run this workflow:
```
sos run workflow/gwas_enrichment.ipynb deepsea_apply --variants /path/to/variant/list.file
```
```
sos run workflow/gwas_enrichment.ipynb deepsea_apply --variants ~/Documents/GWAS_ATAC/GWAS_data/SCZSweden/scz.swe.var.txt
```
It requires much memory. So we test for first 10K line of variant and it worked.
```
sos run workflow/gwas_enrichment.ipynb deepsea_apply --variants ~/Documents/GWAS_ATAC/GWAS_data/SCZSweden/test.var.txt
sos run workflow/gwas_enrichment.ipynb deepsea_apply --variants ~/Documents/GWAS_ATAC/GWAS_data/SCZSweden/test100K.var.txt
```
Break 9898079 snps into two halves and run deepsea separately.
```
less scz.swe.var.txt | head -4949039 | gzip > half1.var.txt
```
```
sos run workflow/gwas_enrichment.ipynb deepsea_apply --variants ~/Documents/GWAS_ATAC/GWAS_data/SCZSweden/half1.var.txt
sos run workflow/gwas_enrichment.ipynb deepsea_apply --variants ~/Documents/GWAS_ATAC/GWAS_data/SCZSweden/half2.var.txt
```
```
[deepsea_apply_1 (prepare config file)]
parameter: variants = path()
input: variants
output: f'{_input:n}.config'
report: expand = True, output = _output
var_list: {_input:r}
pred_model:
path: '{deepsea_model}'
label:
CN: 1
DN: 2
GA: 3
ips: 4
NSC: 5
reference_genome: '{hg19}'
script_dir: '/opt/deepann'
out_dir: {_output:dr}
out_prefix: '{_output:bn}_deepsea'
##### some default setup #####
#### usually don't change ####
check_allele: False
design: '499-1-500'
[deepsea_apply_2 (apply deepsea weights)]
output: f'{_input:n}.deepsea.list'
bash: expand = True, container = 'gaow/deepann', volumes = [f'{hg19:d}:{hg19:d}', f'{deepsea_model:d}:{deepsea_model:d}', f'{os.path.expanduser("~")}:/home/$USER'], extra_args = "-e HOME=/home/$USER -e USER=$USER"
snakemake --snakefile /opt/deepann/Snakefile --configfile {_input} && ls {_input:n}_deepsea/output__*.gz > {_output}
```
## Enrichment analysis via GREGOR
To properly perform enrichment analysis we want to match the control SNPs with the SNPs of interest -- that is, SNPs inside CS -- in terms of LD, distance to nearest gene and MAF. The [GREGOR](http://csg.sph.umich.edu/GREGOR/index.php/site/download) software can generate list of matched SNPs. I will use SNPs inside CS as input and expect a list of output SNPs matching these inputs.
GREGOR is release under University of Michigan license so I'll not make it into a docker image. So path to GREGOR directory is required. Also we need reference files, prepared by:
```
cat \
GREGOR.AFR.ref.r2.greater.than.0.7.tar.gz.part.00 \
GREGOR.AFR.ref.r2.greater.than.0.7.tar.gz.part.01 \
> GREGOR.AFR.ref.r2.greater.than.0.7.tar.gz
tar zxvf GREGOR.AFR.ref.r2.greater.than.0.7.tar.gz
```
MD5SUM check:
```
AFR.part.0 ( MD5: 9926904128dd58d6bf1ad4f1e90638af )
AFR.part.1 ( MD5: c1d30aff89a584bfa8c1fa1bdc197f21 )
```
Same for EUR data-set.
#### To run GREGOR
`PGC3`
```
sos run workflow/gwas_enrichment.ipynb gregor --gregor-input ~/Documents/GWAS_ATAC/scz2_zscore/gregor.scz2.txt.gz --single-annot data/gregor.txt
```
`
PGC2
`
```
sos run workflow/gwas_enrichment.ipynb gregor --gregor-input ~/Documents/GWAS_ATAC/pgc2_zscore/gregor.pgc2.txt.gz --single-annot data/gregor.txt
```
```
[gregor_1 (make SNP index)]
depends: executable('zcat')
parameter: gregor_input = path()
input: gregor_input
output: f'{_input:nn}.rsid.txt', f'{_input:nn}.annotations.list'
bash: expand = '${ }'
zcat ${_input} | cut -f 2,3 -d "_" | sed 's/_/:/g' > ${_output[0]}
with open(_output[1], 'w') as f:
f.write('\n'.join(single_anno))
[gregor_2 (make configuration file)]
depends: executable('sed')
parameter: gregor_db = path('~/Documents/hg19/GREGOR_DB')
parameter: pop = 'EUR'
output: f'{_input[0]:nn}.gregor.conf'
report: output = f'{_output}', expand = True
##############################################################################
# CHIPSEQ ENRICHMENT CONFIGURATION FILE
# This configuration file contains run-time configuration of
# CHIP_SEQ ENRICHMENT
###############################################################################
## KEY ELEMENTS TO CONFIGURE : NEED TO MODIFY
###############################################################################
INDEX_SNP_FILE = {_input[0]}
BED_FILE_INDEX = {_input[1]}
REF_DIR = {gregor_db}
R2THRESHOLD = 0.7 ## must be greater than 0.7
LDWINDOWSIZE = 10000 ## must be less than 1MB; these two values define LD buddies
OUT_DIR = {_output:nn}_gregor_output
MIN_NEIGHBOR_NUM = 10 ## define the size of neighborhood
BEDFILE_IS_SORTED = true ## false, if the bed files are not sorted
POPULATION = {pop} ## define the population, you can specify EUR, AFR, AMR or ASN
TOPNBEDFILES = 2
JOBNUMBER = 10
###############################################################################
#BATCHTYPE = mosix ## submit jobs on MOSIX
#BATCHOPTS = -E/tmp -i -m2000 -j10,11,12,13,14,15,16,17,18,19,120,122,123,124,125 sh -c
###############################################################################
#BATCHTYPE = slurm ## submit jobs on SLURM
#BATCHOPTS = --partition=broadwl --account=pi-mstephens --time=0:30:0
###############################################################################
BATCHTYPE = local ## run jobs on local machine
bash: expand = True
sed -i '/^$/d' {_output}
```
GREGOR is written in `perl`. Some libraries are required before one can run GREGOR:
```
sudo apt-get install libdbi-perl libswitch-perl libdbd-sqlite3-perl
```
```
[gregor_3 (run gregor)]
depends: executable('perl')
parameter: gregor_path = path('~/Documents/GREGOR')
output: f'{_input:nn}_gregor_output/StatisticSummaryFile.txt'
bash: expand = True
perl {gregor_path}/script/GREGOR.pl --conf {_input} && touch {_output}
[gregor_4 (format output)]
depends: executable('sed')
output: f'{_input:n}.csv'
bash: expand = True
sed 's/\t/,/g' {_input} > {_output}
```
| github_jupyter |
```
# noexport
import os
os.system('export_notebook identify_domain_v3.ipynb')
from tmilib import *
#user = get_test_users()[0]
#print user
#tab_focus_times = get_tab_focus_times_for_user(user)
#print tab_focus_times[0].keys()
#print active_second_to_domain_id.keys()[:10]
#seen_domains_only_real = get_seen_urls_and_domains_real_and_history()['seen_domains_only_real']
#[domain_to_id(x) for x in ['www.mturk.com', 'apps.facebook.com', 'www.facebook.com', 'www.reddit.com', 'www.youtube.com']]
#sumkeys(total_tabbed_back_domains_all, 'www.mturk.com', 'apps.facebook.com', 'www.facebook.com', 'www.reddit.com', 'www.youtube.com')
#728458.0/2382221
#print_counter(total_tabbed_back_domains_first)
'''
total_active_seconds = 0
for user in get_test_users():
total_active_seconds += len(get_active_seconds_for_user(user))
#for span in get_active_seconds_for_user(user)
print total_active_seconds
'''
#for user in get_test_users():
# print user
# print (get_recently_seen_domain_stats_for_user(user)['stats'])
@memoized
def get_recently_seen_domain_stats_for_user_v3(user):
ordered_visits = get_history_ordered_visits_corrected_for_user(user)
ordered_visits = exclude_bad_visits(ordered_visits)
active_domain_at_time = get_active_domain_at_time_for_user(user)
active_seconds_set = set(get_active_insession_seconds_for_user(user))
#active_second_to_domain_id = get_active_second_to_domain_id_for_user(user)
active_second_to_domain_id = {int(k):v for k,v in get_active_second_to_domain_id_for_user(user).viewitems()}
recently_seen_domain_ids = [-1]*100
seen_domain_ids_set = set()
stats = Counter()
nomatch_domains = Counter()
tabbed_back_domains_first = Counter()
tabbed_back_domains_second = Counter()
tabbed_back_domains_all = Counter()
distractors_list = [domain_to_id(x) for x in ['www.mturk.com', 'apps.facebook.com', 'www.facebook.com', 'www.reddit.com', 'www.youtube.com']]
distractors = set(distractors_list)
most_recent_distractor = distractors_list[0]
for idx,visit in enumerate(ordered_visits):
if idx+1 >= len(ordered_visits):
break
next_visit = ordered_visits[idx+1]
cur_domain = url_to_domain(visit['url'])
cur_domain_id = domain_to_id(cur_domain)
if cur_domain_id in distractors:
most_recent_distractor = cur_domain_id
recently_seen_domain_ids.append(cur_domain_id)
#if cur_domain_id != recently_seen_domain_ids[-1]:
# if cur_domain_id in seen_domain_ids_set:
# recently_seen_domain_ids.remove(cur_domain_id)
# seen_domain_ids_set.add(cur_domain_id)
# recently_seen_domain_ids.append(cur_domain_id)
#if cur_domain_id != recently_seen_domain_ids[-1]:
# if cur_domain_id in seen_domain_ids_set:
# recently_seen_domain_ids.remove(cur_domain_id)
# seen_domain_ids_set.add(cur_domain_id)
# recently_seen_domain_ids.append(cur_domain_id)
next_domain = url_to_domain(next_visit['url'])
next_domain_id = domain_to_id(next_domain)
cur_time_sec = int(round(visit['visitTime'] / 1000.0))
next_time_sec = int(round(next_visit['visitTime'] / 1000.0))
if cur_time_sec > next_time_sec:
continue
for time_sec in xrange(cur_time_sec, next_time_sec+1):
if time_sec not in active_seconds_set:
continue
ref_domain_id = active_second_to_domain_id[time_sec]
stats['total'] += 1
if most_recent_distractor == ref_domain_id:
stats['most_recent_distractor_total'] += 1
if cur_domain_id == ref_domain_id:
stats['most_recent_distractor_curdomain'] += 1
elif cur_domain_id == next_domain_id:
stats['most_recent_distractor_nextdomain'] += 1
else:
stats['most_recent_distractor_some_prev_domain'] += 1
if cur_domain_id == ref_domain_id:
if next_domain_id == cur_domain_id:
stats['first and next equal and correct'] += 1
else:
stats['first correct only'] += 1
continue
if next_domain_id == ref_domain_id:
stats['next correct only'] += 1
continue
stats['both incorrect'] += 1
found_match = False
ref_domain = id_to_domain(ref_domain_id)
for i in range(1,101):
if recently_seen_domain_ids[-1-i] == ref_domain_id:
stats['nth previous correct ' + str(abs(i))] += 1
stats['some previous among past 100 correct'] += 1
found_match = True
tabbed_back_domains_all[ref_domain] += 1
if i == 1:
tabbed_back_domains_first[ref_domain] += 1
if i == 2:
tabbed_back_domains_second[ref_domain] += 1
break
if not found_match:
ref_domain = id_to_domain(ref_domain_id)
#if ref_domain == 'newtab':
# stats['newtab'] += 1
# continue
stats['no match found'] += 1
nomatch_domains[id_to_domain(ref_domain_id)] += 1
'''
if cur_domain_id == ref_domain_id:
if next_domain_id == cur_domain_id:
stats['first and next equal and correct'] += 1
continue
else:
stats['first correct only'] += 1
continue
else:
if next_domain_id == cur_domain_id:
stats['both incorrect'] += 1
found_match = False
for i in range(1,101):
if recently_seen_domain_ids[-1-i] == ref_domain_id:
stats['nth previous correct ' + str(abs(i))] += 1
stats['some previous among past 100 correct'] += 1
found_match = True
break
if not found_match:
ref_domain = id_to_domain(ref_domain_id)
if ref_domain == 'newtab':
stats['newtab'] += 1
continue
stats['no match found'] += 1
nomatch_domains[id_to_domain(ref_domain_id)] += 1
continue
if next_domain_id == ref_domain_id:
stats['next correct only'] += 1
continue
'''
return {
'stats': stats,
'nomatch_domains': nomatch_domains,
'tabbed_back_domains_all': tabbed_back_domains_all,
'tabbed_back_domains_first': tabbed_back_domains_first,
'tabbed_back_domains_second': tabbed_back_domains_second,
}
#total_stats = Counter({'total': 544544, 'first and next equal and correct': 351081, 'first correct only': 88522, 'both incorrect': 51663, 'some previous among past 20 correct': 41231, 'nth previous correct 1': 31202, 'next correct only': 23569, 'no match found': 10432, 'nth previous correct 2': 3311, 'nth previous correct 3': 1635, 'nth previous correct 4': 905, 'nth previous correct 5': 862, 'nth previous correct 6': 545, 'nth previous correct 7': 412, 'nth previous correct 8': 357, 'nth previous correct 9': 269, 'nth previous correct 10': 259, 'nth previous correct 13': 234, 'nth previous correct 11': 229, 'nth previous correct 12': 190, 'nth previous correct 15': 183, 'nth previous correct 14': 140, 'nth previous correct 17': 139, 'nth previous correct 20': 95, 'nth previous correct 16': 90, 'nth previous correct 19': 88, 'nth previous correct 18': 86})
total_stats = Counter()
for user in get_test_users():
for k,v in get_recently_seen_domain_stats_for_user_v3(user)['stats'].viewitems():
total_stats[k] += v
#total_stats = Counter({'total': 544544, 'first and next equal and correct': 351081, 'first correct only': 88522, 'both incorrect': 51663, 'some previous among past 20 correct': 41136, 'nth previous correct 2': 31202, 'next correct only': 23569, 'no match found': 10527, 'nth previous correct 3': 3311, 'nth previous correct 4': 1635, 'nth previous correct 5': 905, 'nth previous correct 6': 862, 'nth previous correct 7': 545, 'nth previous correct 8': 412, 'nth previous correct 9': 357, 'nth previous correct 10': 269, 'nth previous correct 11': 259, 'nth previous correct 14': 234, 'nth previous correct 12': 229, 'nth previous correct 13': 190, 'nth previous correct 16': 183, 'nth previous correct 15': 140, 'nth previous correct 18': 139, 'nth previous correct 17': 90, 'nth previous correct 20': 88, 'nth previous correct 19': 86})
print_counter(total_stats)
print_counter(total_stats)
def sumkeys(d, *args):
return sum(d.get(x, 0.0) for x in args)
norm = {k:float(v)/total_stats['total'] for k,v in total_stats.viewitems()}
print 'select prev gets answer correct', sumkeys(norm, 'first and next equal and correct', 'first correct only')
print 'prev or next gets answer correct', sumkeys(norm, 'first and next equal and correct', 'first correct only', 'next correct only')
print 'prev or next or newtab gets answer correct', sumkeys(norm, 'first and next equal and correct', 'first correct only', 'next correct only', 'newtab')
for i in range(1, 101):
sumprev = sum([norm.get('nth previous correct '+str(x),0.0) for x in range(i+1)])
print 'prev or next or past ' + str(i), sumkeys(norm, 'first and next equal and correct', 'first correct only', 'next correct only', 'newtab')+sumprev
norm = {k:float(v)/total_stats['total'] for k,v in total_stats.viewitems()}
print 'select prev gets answer correct', sumkeys(norm, 'first and next equal and correct', 'first correct only')
print 'prev or next gets answer correct', sumkeys(norm, 'first and next equal and correct', 'first correct only', 'next correct only')
print 'prev or next or newtab gets answer correct', sumkeys(norm, 'first and next equal and correct', 'first correct only', 'next correct only', 'newtab')
for i in range(1, 101):
sumprev = sum([norm.get('nth previous correct '+str(x),0.0) for x in range(i+1)])
print 'prev or next or past ' + str(i), sumkeys(norm, 'first and next equal and correct', 'first correct only', 'next correct only', 'newtab')+sumprev
#print norm['most_recent_distractor_total']
#print norm['most_recent_distractor_curdomain']
#print norm['most_recent_distractor_nextdomain']
#print norm['most_recent_distractor_some_prev_domain']
```
| github_jupyter |
# freud.cluster.Cluster and freud.cluster.ClusterProperties
The `freud.cluster` module determines clusters of points and computes cluster quantities like centers of mass, gyration tensors, and radii of gyration. The example below generates random points, and shows that they form clusters. This case is two-dimensional (with $z=0$ for all particles) for simplicity, but the cluster module works for both 2D and 3D simulations.
```
import freud
import matplotlib.pyplot as plt
import numpy as np
```
First, we generate a box and random points to cluster.
```
box = freud.Box.square(L=6)
points = np.empty(shape=(0, 2))
for center_point in [(-1.8, 0), (1.5, 1.5), (-0.8, -2.8), (1.5, 0.5)]:
points = np.concatenate(
(
points,
np.random.multivariate_normal(
mean=center_point, cov=0.08 * np.eye(2), size=(100,)
),
)
)
points = np.hstack((points, np.zeros((points.shape[0], 1))))
points = box.wrap(points)
system = freud.AABBQuery(box, points)
system.plot(ax=plt.gca(), s=10)
plt.title("Raw points before clustering", fontsize=20)
plt.gca().tick_params(axis="both", which="both", labelsize=14, size=8)
plt.show()
```
Now we create a box and a cluster compute object.
```
cl = freud.cluster.Cluster()
```
Next, we use the `computeClusters` method to determine clusters and the `clusterIdx` property to return their identities. Note that we use `freud`'s *method chaining* here, where a compute method returns the compute object.
```
cl.compute(system, neighbors={"r_max": 1.0})
print(cl.cluster_idx)
fig, ax = plt.subplots(1, 1, figsize=(9, 6))
for cluster_id in range(cl.num_clusters):
cluster_system = freud.AABBQuery(
system.box, system.points[cl.cluster_keys[cluster_id]]
)
cluster_system.plot(ax=ax, s=10, label=f"Cluster {cluster_id}")
print(
f"There are {len(cl.cluster_keys[cluster_id])} points in cluster {cluster_id}."
)
ax.set_title("Clusters identified", fontsize=20)
ax.legend(loc="best", fontsize=14)
ax.tick_params(axis="both", which="both", labelsize=14, size=8)
plt.show()
```
We may also compute the clusters' centers of mass and gyration tensor using the `ClusterProperties` class.
```
clp = freud.cluster.ClusterProperties()
clp.compute(system, cl.cluster_idx);
```
Plotting these clusters with their centers of mass, with size proportional to the number of clustered points:
```
fig, ax = plt.subplots(1, 1, figsize=(9, 6))
for i in range(cl.num_clusters):
cluster_system = freud.AABBQuery(system.box, system.points[cl.cluster_keys[i]])
cluster_system.plot(ax=ax, s=10, label=f"Cluster {i}")
for i, c in enumerate(clp.centers):
ax.scatter(c[0], c[1], s=len(cl.cluster_keys[i]), label=f"Cluster {i} Center")
plt.title("Center of mass for each cluster", fontsize=20)
plt.legend(loc="best", fontsize=14)
plt.gca().tick_params(axis="both", which="both", labelsize=14, size=8)
plt.gca().set_aspect("equal")
plt.show()
```
The 3x3 gyration tensors $G$ can also be computed for each cluster. For this two-dimensional case, the $z$ components of the gyration tensor are zero. The gyration tensor can be used to determine the principal axes of the cluster and radius of gyration along each principal axis. Here, we plot the gyration tensor's eigenvectors with length corresponding to the square root of the eigenvalues (the singular values).
```
fig, ax = plt.subplots(1, 1, figsize=(9, 6))
for i in range(cl.num_clusters):
cluster_system = freud.AABBQuery(system.box, system.points[cl.cluster_keys[i]])
cluster_system.plot(ax=ax, s=10, label=f"Cluster {i}")
for i, c in enumerate(clp.centers):
ax.scatter(c[0], c[1], s=len(cl.cluster_keys[i]), label=f"Cluster {i} Center")
for cluster_id in range(cl.num_clusters):
com = clp.centers[cluster_id]
G = clp.gyrations[cluster_id]
evals, evecs = np.linalg.eig(G[:2, :2])
arrows = np.sqrt(evals) * evecs
for arrow in arrows.T:
plt.arrow(com[0], com[1], arrow[0], arrow[1], width=0.05, color="k")
plt.title("Eigenvectors of the gyration tensor for each cluster", fontsize=20)
plt.legend(loc="best", fontsize=14)
ax.tick_params(axis="both", which="both", labelsize=14, size=8)
ax.set_aspect("equal")
plt.show()
```
| github_jupyter |
#### Setting Up The Athena Driver
Jupyter Notebooks, like this one, provide a hybrid experience between interactive analytics and purpose built application development. You can leverage powerful statical analysis libraries like numpy or pandas and feed the results into pyplot or seaborn visualization libraries.
```
import sys
!{sys.executable} -m pip install PyAthena
from pyathena import connect
import pandas as pd
conn = connect(work_group='packt-athena-analytics', region_name='us-east-1', schema_name='packt_serverless_analytics')
athena_results = pd.read_sql("""SELECT year, COUNT(*) as num_rides
FROM chapter_3_nyc_taxi_parquet
GROUP BY year
ORDER BY num_rides DESC""", conn)
athena_results.head(3)
from matplotlib import pyplot as plt
import seaborn as sns
sns.barplot(x="year", y="num_rides", data=athena_results)
from scipy import stats
#surpressing warning related to chained assignment of zscore to existing data frame
pd.options.mode.chained_assignment = None
zscore = stats.zscore(athena_results['num_rides'])
athena_results['zscore']=zscore
print(athena_results)
athena_filtered= athena_results[athena_results['zscore'] > 0]
sns.barplot(x="year", y="num_rides", data=athena_filtered)
athena_results_2 = pd.read_sql("""
SELECT date_trunc('day', date_parse(tpep_pickup_datetime,'%Y-%m-%d %H:%i:%s')) as day,
COUNT(*) as ride_count,
AVG(fare_amount) as avg_fare_amount,
AVG(tip_amount) as avg_tip_amount
FROM chapter_7_nyc_taxi_parquet
GROUP BY date_trunc('day', date_parse(tpep_pickup_datetime,'%Y-%m-%d %H:%i:%s'))
ORDER BY day ASC
""", conn)
zscore2 = stats.zscore(athena_results_2['ride_count'])
athena_results_2['zscore']=zscore2
athena_filtered_2= athena_results_2[athena_results_2['zscore'] > -1]
import matplotlib.dates as mdates
import matplotlib.ticker as ticker
fig, ax = pyplot.subplots(figsize= (16.7, 6.27))
plot = sns.scatterplot(ax=ax, x="day", y="ride_count",size="avg_fare_amount",sizes=(1, 150), hue="avg_tip_amount", data=athena_filtered_2)
plot.xaxis.set_major_locator(ticker.MultipleLocator(125))
plot.xaxis.set_major_formatter(mdates.DateFormatter('%m/%Y'))
plt.show()
athena_results_3=pd.read_sql("""SELECT
max(hour(date_parse(tpep_pickup_datetime,'%Y-%m-%d %H:%i:%s'))) as hour_val,
avg(date_diff('second', date_parse(tpep_pickup_datetime,'%Y-%m-%d %H:%i:%s'),
date_parse(tpep_dropoff_datetime,'%Y-%m-%d %H:%i:%s'))) as duration,
avg(trip_distance) as trip_distance,
avg(fare_amount) as fare_amount,
avg(tip_amount) as tip_amount,
count(*) as cnt
from chapter_7_nyc_taxi_parquet
WHERE year=2018
group by date_trunc('hour', date_parse(tpep_pickup_datetime,'%Y-%m-%d %H:%i:%s')) """, conn)
athena_results_3.corr()
```
| github_jupyter |
This notebook detects IEDs using different parameters.
```
import mne
import scipy.io
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
import math
import pyedflib
import random
from tqdm import tqdm
import pandas as pd
matplotlib.rcParams['figure.figsize'] = (10, 2)
mne.set_log_level('WARNING')
%run DetectSpike_SEEG.py
```
Open the file
## DaKa
```
# file_name = 'Data/TrialPatientBeSa/X~ X_74addeec-ea9c-4b25-8280-cab2db067651.EDF'
# file_name = 'Data/TrialPatientArRa/X~ X_2fc7b4bb-d0f1-45b1-b60d-0ec24a942259.EDF'
file_name = 'Data/TrialPatientDaKa/X~ X_0696a934-8738-44ab-8904-99b1d92c4b39.EDF'
f = pyedflib.EdfReader(file_name)
min_min, max_min = 720, 1200
# Load file containing suitable channels and convert to list
df = pd.read_excel('Data/TrialPatientDaKa/Electrodes.xlsx', header=None)
valid_channels = df[0].tolist()
df = pd.read_excel('Data/TrialPatientDaKa/Electrode_mapping.xlsx')
valid_channels_mapped = []
for valid_channel in valid_channels:
valid_channels_mapped.append("C" + str(int(df[df.Original == valid_channel].Mapped)))
signal_labels = f.getSignalLabels()
freq = 2048
# artifact_chans_idx = [46, 47, 49, 51, 52, 53, 57, 59, 98, 99, 113, 137, 146]
# artifact_chans_names = ['C' + str(idx) for idx in artifact_chans_idx]
# valid_channels_mapped_clean = [chan for chan in valid_channels_mapped if chan not in artifact_chans_names]
# valid_channels_clean = [valid_channels[valid_channels_mapped.index(chan)] for chan in valid_channels_mapped_clean]
sample_size = 50
valid_channels_sample = [valid_channels_mapped[i] for i in sorted(random.sample(range(len(valid_channels_mapped)), sample_size))]
from random import randrange
np.save('eeg_samples/sampled_channels.npy', np.array(valid_channels_sample))
np.array(valid_channels_sample)
bh, ah = butter(2, 0.5 / (freq / 2), 'highpass')
bl, al = butter(4, 70 / (freq / 2), 'lowpass')
from pyedflib import EdfWriter, FILETYPE_EDF, FILETYPE_EDFPLUS
from datetime import datetime
start_date = datetime(2017, 5, 24, 15, 53, 49)
def save_eeg(data, valid_channels_sample, filename, all_annotations=None):
with pyedflib.EdfWriter(filename, len(valid_channels_sample), FILETYPE_EDFPLUS) as writer:
writer.setPatientCode('')
writer.setRecordingAdditional('')
writer.setStartdatetime(start_date)
writer.setSignalHeaders([{
# 'label': valid_channels[valid_channels_mapped.index(valid_channel)],
'label': str(i),
'sample_rate': float(freq),
'dimension': 'uV',
'physical_min': -8711.0,
'physical_max': 8711.0,
'digital_min': -32767,
'digital_max': 32767,
'transducer': '',
'prefilter': ''
} for i, valid_channel in enumerate(valid_channels_sample)])
writer.writeSamples(data)
if all_annotations is not None:
for all_annotation in all_annotations:
for annotation in all_annotation:
# add seconds of index of all_annotation?
writer.writeAnnotation(annotation[0], annotation[1], 'IED sequence')
# writer.close()
spacing = (max_min - min_min) // 100
%run utils.py
combined_data = []
all_annotations = []
for iteration, min_start in enumerate(tqdm(range(min_min, max_min, spacing))):
random.seed(iteration)
random_start = randrange(0, len(valid_channels_mapped) - sample_size + 1)
# valid_channels_sample = valid_channels_mapped[random_start:random_start + sample_size]
valid_channels_sample = valid_channels[random_start:random_start + sample_size]
data = np.zeros((len(valid_channels_sample), freq * 60))
# Populate area of data for the current block across all channels
for i, chan in enumerate(valid_channels_sample):
data[i, :] = f.readSignal(signal_labels.index(chan), \
start = min_start * freq * 60, \
n = freq * 60)
filename = 'eeg_samples/from_' + str(min_start) + '.edf'
save_eeg(data, valid_channels_sample, filename)
# Detect IEDs in current block across all chanels
spike_id, chan_id, _ = DetectSpikes(data, freq, STDCoeff=2)
all_seq_spikes, all_seq_chans = spikes_to_sequences(spike_id, chan_id, 0.015, 0.015)
data = filtfilt(bh, ah, data)
data = filtfilt(bl, al, data)
# seconds 1 to 2
combined_data.append(data[:, 2048:2048*2])
curr_annotations = []
for all_seq_spike in all_seq_spikes:
if (1 <= all_seq_spike[0] / 2048 <= 2) or (1 <= all_seq_spike[-1] / 2048 <= 2):
start = iteration - 1 + all_seq_spike[0] / 2048
end = iteration - 1 + all_seq_spike[-1] / 2048
duration = end - start
curr_annotations.append((start, duration))
print(start, duration)
elif all_seq_spike[-1] > 2:
break
all_annotations.append(curr_annotations)
# print(len(all_seq_spikes))
filename = 'eeg_samples/BeSa.edf'
save_eeg(np.concatenate(combined_data, axis=-1), valid_channels_sample, filename, all_annotations)
spacing = (max_min - min_min) // 20
%run utils.py
combined_data = []
all_annotations = []
for iteration, min_start in enumerate(tqdm(range(min_min, max_min, spacing))):
data = np.zeros((len(valid_channels_sample), freq * 60))
# Populate area of data for the current block across all channels
for i, chan in enumerate(valid_channels_sample):
data[i, :] = f.readSignal(signal_labels.index(chan), \
start = min_start * freq * 60, \
n = freq * 60)
filename = 'eeg_samples/BeSa/from_' + str(min_start) + '.edf'
save_eeg(data, valid_channels_sample, filename)
# Detect IEDs in current block across all chanels
spike_id, chan_id, _ = DetectSpikes(data, freq, STDCoeff=2)
all_seq_spikes, all_seq_chans = spikes_to_sequences(spike_id, chan_id, 0.05, 0.015)
data = filtfilt(bh, ah, data)
data = filtfilt(bl, al, data)
# seconds 1 to 2
combined_data.append(data[:, 2048:2048*2])
curr_annotations = []
for all_seq_spike in all_seq_spikes:
if (1 <= all_seq_spike[0] / 2048 <= 2) or (1 <= all_seq_spike[-1] / 2048 <= 2):
start = iteration - 1 + all_seq_spike[0] / 2048
end = iteration - 1 + all_seq_spike[-1] / 2048
duration = end - start
curr_annotations.append((start, duration))
print(start, duration)
elif all_seq_spike[-1] > 2:
break
all_annotations.append(curr_annotations)
# print(len(all_seq_spikes))
```
## BeSa
```
file_name = 'Data/TrialPatientBeSa/X~ X_74addeec-ea9c-4b25-8280-cab2db067651.EDF'
f = pyedflib.EdfReader(file_name)
min_min, max_min = 720, 2100
# Load file containing suitable channels and convert to list
df = pd.read_excel('Data/TrialPatientBeSa/BeSA/sub-01/Electrodes.xlsx', header=None)
valid_channels = ['EEG ' + name for name in df[0].tolist()]
signal_labels = f.getSignalLabels()
sample_size = 50
valid_channels_sample = [valid_channels[i] for i in sorted(random.sample(range(len(valid_channels)), sample_size))]
np.save('eeg_samples/BeSa/sampled_channels.npy', np.array(valid_channels_sample))
np.array(valid_channels_sample)
from pyedflib import EdfWriter, FILETYPE_EDF
from datetime import datetime
start_date = datetime(2017, 5, 24, 15, 53, 49)
def save_eeg(data, valid_channels_sample, filename, all_annotations=None):
with pyedflib.EdfWriter(filename, len(valid_channels_sample), FILETYPE_EDF) as writer:
writer.setPatientCode('')
writer.setRecordingAdditional('')
writer.setStartdatetime(start_date)
writer.setSignalHeaders([{
'label': valid_channel,
'sample_rate': float(freq),
'dimension': 'uV',
'physical_min': -8711.0,
'physical_max': 8711.0,
'digital_min': -32767,
'digital_max': 32767,
'transducer': '',
'prefilter': ''
} for valid_channel in valid_channels_sample])
writer.writeSamples(data)
# if all_annotations is not None:
# for all_annotation in all_annotations:
# for annotation in all_annotation:
# # add seconds of index of all_annotation?
# writer.writeAnnotation(annotation[0], annotation[1], 'IED sequence')
# writer.close()
spacing = (max_min - min_min) // 20
%run utils.py
combined_data = []
all_annotations = []
for iteration, min_start in enumerate(tqdm(range(min_min, max_min, spacing))):
data = np.zeros((len(valid_channels_sample), freq * 60))
# Populate area of data for the current block across all channels
for i, chan in enumerate(valid_channels_sample):
data[i, :] = f.readSignal(signal_labels.index(chan), \
start = min_start * freq * 60, \
n = freq * 60)
filename = 'eeg_samples/BeSa/from_' + str(min_start) + '.edf'
save_eeg(data, valid_channels_sample, filename)
# Detect IEDs in current block across all chanels
spike_id, chan_id, _ = DetectSpikes(data, freq, STDCoeff=2)
all_seq_spikes, all_seq_chans = spikes_to_sequences(spike_id, chan_id, freq)
data = filtfilt(bh, ah, data)
data = filtfilt(bl, al, data)
# seconds 1 to 2
combined_data.append(data[:, 2048:2048*2])
curr_annotations = []
for all_seq_spike in all_seq_spikes:
if (1 <= all_seq_spike[0] / 2048 <= 2) or (1 <= all_seq_spike[-1] / 2048 <= 2):
curr_annotations.append((all_seq_spike[0] / 2048, (all_seq_spike[-1] - all_seq_spike[0]) / 2048))
print(iteration + all_seq_spike[0] / 2048, iteration + all_seq_spike[-1] / 2048)
elif all_seq_spike[-1] > 2:
break
all_annotations.append(curr_annotations)
# print(len(all_seq_spikes))
filename = 'eeg_samples/BeSa/combined.edf'
save_eeg(np.concatenate(combined_data, axis=-1), valid_channels_sample, filename, all_annotations)
```
Define file specific parameters
```
freq = 2048
```
Get channels and number of channels
```
num_chans = f.signals_in_file # change to valid_channels.shape[0]
signal_labels = f.getSignalLabels()
```
Get list of channels to use from channels.mat
```
# Load file containing suitable channels and convert to list
df = pd.read_excel('Data/TrialPatientDaKa/Electrodes.xlsx', header=None)
valid_channels = df[0].tolist()
df = pd.read_excel('Data/TrialPatientDaKa/Electrode_mapping.xlsx')
valid_channels_mapped = []
for valid_channel in valid_channels:
valid_channels_mapped.append("C" + str(int(df[df.Original == valid_channel].Mapped)))
# Load file containing suitable channels and convert to list
mat = scipy.io.loadmat('Data/TrialPatientArRa/channels.mat')
valid_channels = [channel[0] for channel in mat['channels'][:, 0]]
# valid_channels = signal_labels
import pandas as pd
# Load file containing suitable channels and convert to list
df = pd.read_excel('Data/TrialPatientDaKa/Electrodes.xlsx', header=None)
valid_channels_unmapped = df[0].tolist()
df = pd.read_excel('Data/TrialPatientDaKa/Electrode_mapping.xlsx')
valid_channels = []
for valid_channel in valid_channels_unmapped:
valid_channels.append("C" + str(int(df[df.Original == valid_channel].Mapped)))
valid_channels_unmapped[valid_channels.index('C47')]
```
Process a random minute of data with a range of params and select random 5s segments in random channels to show detections for each set of params.
```
matplotlib.rcParams['figure.figsize'] = (2, 2)
# Process 1 minute at a time
mins_to_process = 1
M = f.getNSamples()[0]# / (freq * 60)
NumSecs = M / freq
Blocks = math.floor(NumSecs / (mins_to_process * 60))
bh, ah = butter(2, 0.5 / (freq / 2), 'highpass')
bl, al = butter(4, 70 / (freq / 2), 'lowpass')
def generate_detections(params_list):
# Randomly choose a minute to select data from
minute = random.randrange(60, Blocks)
# minute = 28
data_block = np.zeros((len(valid_channels_mapped), freq * 60 * mins_to_process))
# reference = f.readSignal(signal_labels.index(ref_chan), \
# start = minute * freq * 60 * mins_to_process, \
# n = freq * 60 * mins_to_process)
# Populate area of data for the current block across all channels
for i, chan in enumerate(valid_channels_mapped):
data_block[i, :] = f.readSignal(signal_labels.index(chan), \
start = minute * freq * 60 * mins_to_process, \
n = freq * 60 * mins_to_process)# - reference
# Index of channel to consider
chan = random.choices(valid_channels)[0]
# Randomly select a 5 second segment within the minute of data to show detections for (note: 12 blocks of 5s)
five_second_segment = random.randrange(12)
print(minute, chan, five_second_segment)
for params in params_list:
# Detect IEDs in current block across all chanels
SpikeIndex, ChanId, _ = DetectSpikes(data_block, freq, **params)
data = filtfilt(bh, ah, data_block, padlen=150)
data_block = filtfilt(bl, al, data_block, padlen=150)
info = mne.create_info([chan], freq, ch_types='seeg')
data = mne.io.RawArray(data = [data_block[valid_channels.index(chan)] \
[five_second_segment * freq * 5 : \
(five_second_segment + 1) * freq * 5]], info = info)
onsets = []
durations = []
descriptions = []
SpikeIds_from_zero = SpikeIndex - five_second_segment * freq * 5
spike_indices = np.logical_and(ChanId == valid_channels.index(chan), SpikeIds_from_zero < 5 * freq)
for spike in SpikeIds_from_zero[spike_indices]:
if spike > 0:
onsets.append(spike / 2048)
durations.append(0.01)
descriptions.append('IED')
my_annot = mne.Annotations(onset=onsets, # in seconds
duration=durations, # in seconds, too
description=descriptions)
data = data.set_annotations(my_annot)
# test = data.plot(start=20, duration=2, n_channels=6, scalings=dict(eeg=5e-4))
fig = data.plot(scalings=dict(seeg=3e3), show_scrollbars=False)
plt.show()
DetThresholds_list = [{'DetThresholds':[7, 7, 200, 10, 10]},
{'DetThresholds':[7, 7, 400, 10, 10]},
{'DetThresholds':[7, 7, 600, 10, 10]},
{'DetThresholds':[7, 7, 800, 10, 10]}]
STDCoeff_list = [2, 3]
SCALE_list = [70]
params1 = {'STDCoeff': 2}
params2 = {'STDCoeff': 3}
# generate_detections(DetThresholds_list)
generate_detections([params1, params2])
```
Det Thresholds: [7, 7, 600, 10, 10]
7 seems fine for the first two arguments. Definitely not 9. (explored 5-9)
STDCoeff: 3 seems okay (explored 2-6). 2 has less false negatives, but more false positives.
scales of 60-90 so far (explored 50-90). 70 seems good.
| github_jupyter |
```
library(dplyr)
library(ggplot2)
mydir = "/nfs/leia/research/stegle/dseaton/hipsci/singlecell_neuroseq/data/data_processed/pool1_17_D52/"
mysuffix = "pool1_17_D52.scanpy.w_metadata.w_celltype.scanpy.obs_df.groupedby.donor_id-pool_id-time_point-treatment.celltype_counts.tsv"
myfilename = paste0(mydir,mysuffix)
df2 = read.table(myfilename, header = T)
df2$celltype <- as.character(df2$celltype)
df2$celltype[df2$celltype == "CHem"] <- "U_Neur1"
df2$celltype[df2$celltype == "unknown"] <- "U_Neur3"
head(df2)
df_tot_d52_mid = df2[df2$celltype %in% c("DA","Sert"),] %>% group_by(donor_id,pool_id) %>%
summarize(total_midbrain_cells = sum(n_cells))
nrow(df_tot_d52_mid)
df_tot_d52 = df2 %>% group_by(donor_id,pool_id) %>% summarize(total_cells = sum(n_cells))
nrow(df_tot_d52)
df = inner_join(df_tot_d52, df_tot_d52_mid, by = c("donor_id","pool_id"))
nrow(df)
head(df)
df$diff_eff = df$total_midbrain_cells/df$total_cells
head(df)
library(lme4)
coldata = read.csv("/hps/nobackup/stegle/users/acuomo/all_scripts/sc_neuroseq/metadata_all_lines.csv", row.names = 1)
head(coldata)
df2 = inner_join(df, as.data.frame(coldata), by = "donor_id")
head(df2)
lmm2 <- lmer(diff_eff ~ (1 | donor_id) + (1 | pool_id) + (1 | Gender) + (1 | Age), data = df2, REML = F)
summary(lmm2)
donor = summary(lmm2)$varcor$donor_id[1,1]
pool = summary(lmm2)$varcor$pool_id[1,1]
sex = summary(lmm2)$varcor$Gender[1,1]
age = summary(lmm2)$varcor$Age[1,1]
residual = summary(lmm2)$sigma **2
#
sum = donor+pool+sex+age+residual
options(repr.plot.width = 5, repr.plot.height = 3.5)
df_plot = data.frame(component = c("Donor/line","Pool","Donor Sex","Donor Age","Residuals"),
variance = c(donor/sum, pool/sum, sex/sum, age/sum, residual/sum))
p = ggplot(df_plot, aes(x = component, y = variance*100, fill = component)) + geom_bar(stat="identity", alpha = 0.8)
p = p + xlab("") + ylab("% variance explained") + theme_classic()
p = p + theme(axis.text.x = element_text(angle = 45, hjust = 1, size = 10))
p + scale_fill_manual(values = c("darkmagenta","steelblue4","turquoise4","goldenrod4","firebrick"))
fig_dir = "/hps/nobackup/stegle/users/acuomo/all_scripts/sc_neuroseq/figures/extended_figures/"
pdf(paste0(fig_dir,"SF_9c.pdf"), width=5, height=3.5)
p + scale_fill_manual(values = c("darkmagenta","steelblue4","turquoise4","goldenrod4","firebrick"))
dev.off()
xci_data = "/nfs/leia/research/stegle/dseaton/hipsci/singlecell_neuroseq/data/metadata/neuroseq_x_chrom_inactivation_data.tsv"
xci = read.csv(xci_data, sep = "\t")
xci$donor_id = xci$cell_line_id
head(xci)
nrow(df2)
df3 = inner_join(xci, df2, by = c("donor_id","Gender"))
nrow(df3)
head(df3,2)
df3$xci[df3$mean_x_chrom_ase > 0 & df3$mean_x_chrom_ase < 0.1] = '0-0.1'
df3$xci[df3$mean_x_chrom_ase > 0.1 & df3$mean_x_chrom_ase < 0.2] = '0.1-0.2'
df3$xci[df3$mean_x_chrom_ase > 0.2 & df3$mean_x_chrom_ase < 0.3] = '0.2-0.3'
df3$xci[df3$mean_x_chrom_ase > 0.3 & df3$mean_x_chrom_ase < 0.4] = '0.3-0.4'
df3$xci[df3$mean_x_chrom_ase > 0.4 & df3$mean_x_chrom_ase < 0.5] = '0.4-0.5'
head(df3,2)
lmm3 <- lmer(diff_eff ~ (1 | donor_id) + (1 | pool_id) + (1 | xci) + (1 | Age), data = df3, REML = F)
summary(lmm3)
donor = summary(lmm3)$varcor$donor_id[1,1]
pool = summary(lmm3)$varcor$pool_id[1,1]
xci = summary(lmm3)$varcor$xci[1,1]
sex = summary(lmm3)$varcor$Gender[1,1]
age = summary(lmm3)$varcor$Age[1,1]
residual = summary(lmm3)$sigma **2
#
sum = donor+pool+xci+age+residual
options(repr.plot.width = 5, repr.plot.height = 3.5)
df_plot = data.frame(component = c("Donor/line","Pool","Line XCI","Donor Age","Residuals"),
variance = c(donor/sum, pool/sum, xci/sum, age/sum, residual/sum))
p = ggplot(df_plot, aes(x = component, y = variance*100, fill = component)) + geom_bar(stat="identity", alpha = 0.8)
p = p + xlab("") + ylab("% variance explained") + theme_classic()
p = p + theme(axis.text.x = element_text(angle = 45, hjust = 1, size = 10))
p + scale_fill_manual(values = c("darkmagenta","turquoise4","palegreen3","goldenrod4","firebrick"))
cor.test(df3$diff_eff, df3$mean_x_chrom_ase)
pdf(paste0(fig_dir,"SF_9c.pdf"), width=5, height=3.5)
p + scale_fill_manual(values = c("darkmagenta","steelblue4","turquoise4","goldenrod4","firebrick"))
dev.off()
```
| github_jupyter |
```
from __future__ import division
import os
import cv2
import numpy as np
import sys
import pickle
from optparse import OptionParser
import time
import tensorflow as tf
from keras import backend as K
from keras.layers import Input
from keras.models import Model
from keras.backend.tensorflow_backend import set_session
from keras_frcnn import roi_helpers
from skimage import data
from skimage.filters import unsharp_mask
import math
with open('config.pickle', 'rb') as f_in:
C = pickle.load(f_in)
import keras_frcnn.vgg as nn
# turn off any data augmentation at test time
C.use_horizontal_flips = False
C.use_vertical_flips = False
C.rot_90 = False
C.num_rois=32
C.anchor_box_scales = [8,16,24,32,40,64,72,92,128,256]
C.anchor_box_ratios = [[1, 1], [1./math.sqrt(2), 2./math.sqrt(2)], [2./math.sqrt(2), 1./math.sqrt(2)],[2./math.sqrt(2), 3./math.sqrt(2)],[1./math.sqrt(2), 3./math.sqrt(2)]]
working_dir=os.getcwd()
img_path = os.path.join(working_dir,"images")
def format_img_size(img, C):
""" formats the image size based on config """
img_min_side = 480
(height,width,_) = img.shape
if width <= height:
ratio = img_min_side/width
new_height = int(ratio * height)
new_width = int(img_min_side)
else:
ratio = img_min_side/height
new_width = int(ratio * width)
new_height = int(img_min_side)
img = cv2.resize(img, (new_width, img_min_side), interpolation=cv2.INTER_CUBIC)
print (img.shape)
return img,ratio
def format_img_channels(img, C):
""" formats the image channels based on config """
img = img[:, :, (2, 1, 0)]
img = img.astype(np.float32)
print (C.img_channel_mean[0])
img[:, :, 0] -= C.img_channel_mean[0]
img[:, :, 1] -= C.img_channel_mean[1]
img[:, :, 2] -= C.img_channel_mean[2]
img /= C.img_scaling_factor
img = np.transpose(img, (2, 0, 1))
img = np.expand_dims(img, axis=0)
return img
def format_img(img, C):
""" formats an image for model prediction based on config """
img, ratio = format_img_size(img, C)
img = format_img_channels(img, C)
return img,ratio
# Method to transform the coordinates of the bounding box to its original size
def get_real_coordinates(ratio, x1, y1, x2, y2):
real_x1 = int(round(x1 // ratio))
real_y1 = int(round(y1 // ratio))
real_x2 = int(round(x2 // ratio))
real_y2 = int(round(y2 // ratio))
return (real_x1, real_y1, real_x2 ,real_y2)
class_mapping = C.class_mapping
if 'bg' not in class_mapping:
class_mapping['bg'] = len(class_mapping)
class_mapping = {v: k for k, v in class_mapping.items()}
print(class_mapping)
class_to_color = {class_mapping[v]: np.random.randint(0, 255, 3) for v in class_mapping}
num_features = 512
input_shape_img = (None, None, 3)
input_shape_features = (None, None, num_features)
img_input = Input(shape=input_shape_img)
roi_input = Input(shape=(C.num_rois, 4))
feature_map_input = Input(shape=input_shape_features)
# define the base network (resnet here, can be VGG, Inception, etc)
shared_layers = nn.nn_base(img_input, trainable=True)
# define the RPN, built on the base layers
num_anchors = len(C.anchor_box_scales) * len(C.anchor_box_ratios)
rpn_layers = nn.rpn(shared_layers, num_anchors)
classifier = nn.classifier(feature_map_input, roi_input, C.num_rois, nb_classes=len(class_mapping), trainable=True)
model_rpn = Model(img_input, rpn_layers)
model_classifier_only = Model([feature_map_input, roi_input], classifier)
model_classifier = Model([feature_map_input, roi_input], classifier)
model_path=os.path.join(working_dir,"model/Faster_RCNN_model.hdf5")
print('Loading weights')
model_rpn.load_weights(model_path, by_name=True)
model_classifier.load_weights(model_path, by_name=True)
model_rpn.compile(optimizer='sgd', loss='mse')
model_classifier.compile(optimizer='sgd', loss='mse')
model_rpn.summary()
model_classifier.summary()
all_imgs = []
classes = {}
bbox_threshold = 0.5
visualise = True
st=0
for idx, img_name in enumerate((os.listdir(img_path))):
if not img_name.lower().endswith(('.bmp', '.jpeg', '.jpg', '.png', '.tif', '.tiff')):
continue
print(img_name)
st = st+1
filepath = os.path.join(img_path,'IMG_20201109_191530-17720.jpg')
img = cv2.imread(filepath)
#below are some image enhancement techniques used
#img = improve_contrast_image_using_clahe(img)
#img = unsharp_mask(img)
X, ratio = format_img(img, C)
if K.common.image_dim_ordering() == 'tf':
X = np.transpose(X, (0, 2, 3, 1))
# get the feature maps and output from the RPN
[Y1, Y2, F] = model_rpn.predict(X)
R = roi_helpers.rpn_to_roi(Y1, Y2, C, K.common.image_dim_ordering(), overlap_thresh=0.9)
# convert from (x1,y1,x2,y2) to (x,y,w,h)
R[:, 2] -= R[:, 0]
R[:, 3] -= R[:, 1]
# apply the spatial pyramid pooling to the proposed regions
bboxes = {}
probs = {}
for jk in range(R.shape[0]//C.num_rois + 1):
ROIs = np.expand_dims(R[C.num_rois*jk:C.num_rois*(jk+1), :], axis=0)
if ROIs.shape[1] == 0:
break
if jk == R.shape[0]//C.num_rois:
#pad R
curr_shape = ROIs.shape
target_shape = (curr_shape[0],C.num_rois,curr_shape[2])
ROIs_padded = np.zeros(target_shape).astype(ROIs.dtype)
ROIs_padded[:, :curr_shape[1], :] = ROIs
ROIs_padded[0, curr_shape[1]:, :] = ROIs[0, 0, :]
ROIs = ROIs_padded
[P_cls, P_regr] = model_classifier_only.predict([F, ROIs])
for ii in range(P_cls.shape[1]):
print (class_mapping[np.argmax(P_cls[0, ii, :])])
if np.max(P_cls[0, ii, :]) < .2 or np.argmax(P_cls[0, ii, :]) == (P_cls.shape[2] - 1):
continue
cls_name = class_mapping[np.argmax(P_cls[0, ii, :])]
if cls_name not in bboxes:
bboxes[cls_name] = []
probs[cls_name] = []
(x, y, w, h) = ROIs[0, ii, :]
cls_num = np.argmax(P_cls[0, ii, :])
try:
(tx, ty, tw, th) = P_regr[0, ii, 4*cls_num:4*(cls_num+1)]
tx /= C.classifier_regr_std[0]
ty /= C.classifier_regr_std[1]
tw /= C.classifier_regr_std[2]
th /= C.classifier_regr_std[3]
x, y, w, h = roi_helpers.apply_regr(x, y, w, h, tx, ty, tw, th)
except:
pass
bboxes[cls_name].append([C.rpn_stride*x, C.rpn_stride*y, C.rpn_stride*(x+w), C.rpn_stride*(y+h)])
probs[cls_name].append(np.max(P_cls[0, ii, :]))
all_dets = []
print (bboxes)
for key in bboxes:
bbox = np.array(bboxes[key])
new_boxes, new_probs = roi_helpers.non_max_suppression_fast(bbox, np.array(probs[key]), overlap_thresh=0.2)
for jk in range(new_boxes.shape[0]):
(x1, y1, x2, y2) = new_boxes[jk,:]
(real_x1, real_y1, real_x2, real_y2) = get_real_coordinates(ratio, x1, y1, x2, y2)
cv2.rectangle(img,(real_x1, real_y1), (real_x2, real_y2), (int(class_to_color[key][0]), int(class_to_color[key][1]), int(class_to_color[key][2])),2)
textLabel = key
all_dets.append((key,100*new_probs[jk]))
(retval,baseLine) = cv2.getTextSize(textLabel,cv2.FONT_HERSHEY_COMPLEX,1,1)
textOrg = (real_x1, real_y1-0)
cv2.rectangle(img, (textOrg[0] - 5, textOrg[1]+baseLine - 5), (textOrg[0]+retval[0] + 5, textOrg[1]-retval[1] - 5), (0, 0, 0), 2)
cv2.rectangle(img, (textOrg[0] - 5,textOrg[1]+baseLine - 5), (textOrg[0]+retval[0] + 5, textOrg[1]-retval[1] - 5), (255, 255, 255), -1)
cv2.putText(img, textLabel, textOrg, cv2.FONT_HERSHEY_DUPLEX, 1, (0, 0, 0), 1)
#print(f'Elapsed time = {time.time() - st)}')
print(all_dets)
if st==1:
break
from matplotlib.pyplot import imshow,subplots
import numpy as np
from PIL import Image
%matplotlib inline
fig, ax = subplots(figsize=(18, 12))
imshow(img,aspect='auto')
```
| github_jupyter |
```
# default_exp meta.xlearner
#hide
%reload_ext autoreload
%autoreload 2
%matplotlib inline
```
# X-Learner
> X-Learner
```
#hide
from nbdev.showdoc import *
#export
# REFERENCE: https://github.com/uber/causalml
# Copyright 2019 Uber Technology, Inc.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
from copy import deepcopy
import logging
import numpy as np
import pandas as pd
from tqdm import tqdm
from scipy.stats import norm
from causalnlp.meta.base import BaseLearner
from causalnlp.meta.utils import check_treatment_vector, check_p_conditions, convert_pd_to_np
from causalnlp.meta.explainer import Explainer
from causalnlp.meta.utils import regression_metrics, classification_metrics
from causalnlp.meta.propensity import compute_propensity_score
logger = logging.getLogger('causalnlp')
class BaseXLearner(BaseLearner):
"""A parent class for X-learner regressor classes.
An X-learner estimates treatment effects with four machine learning models.
Details of X-learner are available at Kunzel et al. (2018) (https://arxiv.org/abs/1706.03461).
"""
def __init__(self,
learner=None,
control_outcome_learner=None,
treatment_outcome_learner=None,
control_effect_learner=None,
treatment_effect_learner=None,
ate_alpha=.05,
control_name=0):
"""Initialize a X-learner.
Args:
learner (optional): a model to estimate outcomes and treatment effects in both the control and treatment
groups
control_outcome_learner (optional): a model to estimate outcomes in the control group
treatment_outcome_learner (optional): a model to estimate outcomes in the treatment group
control_effect_learner (optional): a model to estimate treatment effects in the control group
treatment_effect_learner (optional): a model to estimate treatment effects in the treatment group
ate_alpha (float, optional): the confidence level alpha of the ATE estimate
control_name (str or int, optional): name of control group
"""
assert (learner is not None) or ((control_outcome_learner is not None) and
(treatment_outcome_learner is not None) and
(control_effect_learner is not None) and
(treatment_effect_learner is not None))
if control_outcome_learner is None:
self.model_mu_c = deepcopy(learner)
else:
self.model_mu_c = control_outcome_learner
if treatment_outcome_learner is None:
self.model_mu_t = deepcopy(learner)
else:
self.model_mu_t = treatment_outcome_learner
if control_effect_learner is None:
self.model_tau_c = deepcopy(learner)
else:
self.model_tau_c = control_effect_learner
if treatment_effect_learner is None:
self.model_tau_t = deepcopy(learner)
else:
self.model_tau_t = treatment_effect_learner
self.ate_alpha = ate_alpha
self.control_name = control_name
self.propensity = None
self.propensity_model = None
def __repr__(self):
return ('{}(control_outcome_learner={},\n'
'\ttreatment_outcome_learner={},\n'
'\tcontrol_effect_learner={},\n'
'\ttreatment_effect_learner={})'.format(self.__class__.__name__,
self.model_mu_c.__repr__(),
self.model_mu_t.__repr__(),
self.model_tau_c.__repr__(),
self.model_tau_t.__repr__()))
def fit(self, X, treatment, y, p=None):
"""Fit the inference model.
Args:
X (np.matrix or np.array or pd.Dataframe): a feature matrix
treatment (np.array or pd.Series): a treatment vector
y (np.array or pd.Series): an outcome vector
p (np.ndarray or pd.Series or dict, optional): an array of propensity scores of float (0,1) in the
single-treatment case; or, a dictionary of treatment groups that map to propensity vectors of
float (0,1); if None will run ElasticNetPropensityModel() to generate the propensity scores.
"""
X, treatment, y = convert_pd_to_np(X, treatment, y)
check_treatment_vector(treatment, self.control_name)
self.t_groups = np.unique(treatment[treatment != self.control_name])
self.t_groups.sort()
if p is None:
self._set_propensity_models(X=X, treatment=treatment, y=y)
p = self.propensity
else:
p = self._format_p(p, self.t_groups)
self._classes = {group: i for i, group in enumerate(self.t_groups)}
self.models_mu_c = {group: deepcopy(self.model_mu_c) for group in self.t_groups}
self.models_mu_t = {group: deepcopy(self.model_mu_t) for group in self.t_groups}
self.models_tau_c = {group: deepcopy(self.model_tau_c) for group in self.t_groups}
self.models_tau_t = {group: deepcopy(self.model_tau_t) for group in self.t_groups}
self.vars_c = {}
self.vars_t = {}
for group in self.t_groups:
mask = (treatment == group) | (treatment == self.control_name)
treatment_filt = treatment[mask]
X_filt = X[mask]
y_filt = y[mask]
w = (treatment_filt == group).astype(int)
# Train outcome models
self.models_mu_c[group].fit(X_filt[w == 0], y_filt[w == 0])
self.models_mu_t[group].fit(X_filt[w == 1], y_filt[w == 1])
# Calculate variances and treatment effects
var_c = (y_filt[w == 0] - self.models_mu_c[group].predict(X_filt[w == 0])).var()
self.vars_c[group] = var_c
var_t = (y_filt[w == 1] - self.models_mu_t[group].predict(X_filt[w == 1])).var()
self.vars_t[group] = var_t
# Train treatment models
d_c = self.models_mu_t[group].predict(X_filt[w == 0]) - y_filt[w == 0]
d_t = y_filt[w == 1] - self.models_mu_c[group].predict(X_filt[w == 1])
self.models_tau_c[group].fit(X_filt[w == 0], d_c)
self.models_tau_t[group].fit(X_filt[w == 1], d_t)
def predict(self, X, treatment=None, y=None, p=None, return_components=False,
verbose=True):
"""Predict treatment effects.
Args:
X (np.matrix or np.array or pd.Dataframe): a feature matrix
treatment (np.array or pd.Series, optional): a treatment vector
y (np.array or pd.Series, optional): an outcome vector
p (np.ndarray or pd.Series or dict, optional): an array of propensity scores of float (0,1) in the
single-treatment case; or, a dictionary of treatment groups that map to propensity vectors of
float (0,1); if None will run ElasticNetPropensityModel() to generate the propensity scores.
return_components (bool, optional): whether to return outcome for treatment and control seperately
verbose (bool, optional): whether to output progress logs
Returns:
(numpy.ndarray): Predictions of treatment effects.
"""
X, treatment, y = convert_pd_to_np(X, treatment, y)
if p is None:
logger.info('Generating propensity score')
p = dict()
for group in self.t_groups:
p_model = self.propensity_model[group]
p[group] = p_model.predict(X)
else:
p = self._format_p(p, self.t_groups)
te = np.zeros((X.shape[0], self.t_groups.shape[0]))
dhat_cs = {}
dhat_ts = {}
for i, group in enumerate(self.t_groups):
model_tau_c = self.models_tau_c[group]
model_tau_t = self.models_tau_t[group]
dhat_cs[group] = model_tau_c.predict(X)
dhat_ts[group] = model_tau_t.predict(X)
_te = (p[group] * dhat_cs[group] + (1 - p[group]) * dhat_ts[group]).reshape(-1, 1)
te[:, i] = np.ravel(_te)
if (y is not None) and (treatment is not None) and verbose:
mask = (treatment == group) | (treatment == self.control_name)
treatment_filt = treatment[mask]
X_filt = X[mask]
y_filt = y[mask]
w = (treatment_filt == group).astype(int)
yhat = np.zeros_like(y_filt, dtype=float)
yhat[w == 0] = self.models_mu_c[group].predict(X_filt[w == 0])
yhat[w == 1] = self.models_mu_t[group].predict(X_filt[w == 1])
logger.info('Error metrics for group {}'.format(group))
regression_metrics(y_filt, yhat, w)
if not return_components:
return te
else:
return te, dhat_cs, dhat_ts
def fit_predict(self, X, treatment, y, p=None, return_ci=False, n_bootstraps=1000, bootstrap_size=10000,
return_components=False, verbose=True):
"""Fit the treatment effect and outcome models of the R learner and predict treatment effects.
Args:
X (np.matrix or np.array or pd.Dataframe): a feature matrix
treatment (np.array or pd.Series): a treatment vector
y (np.array or pd.Series): an outcome vector
p (np.ndarray or pd.Series or dict, optional): an array of propensity scores of float (0,1) in the
single-treatment case; or, a dictionary of treatment groups that map to propensity vectors of
float (0,1); if None will run ElasticNetPropensityModel() to generate the propensity scores.
return_ci (bool): whether to return confidence intervals
n_bootstraps (int): number of bootstrap iterations
bootstrap_size (int): number of samples per bootstrap
return_components (bool, optional): whether to return outcome for treatment and control seperately
verbose (str): whether to output progress logs
Returns:
(numpy.ndarray): Predictions of treatment effects. Output dim: [n_samples, n_treatment]
If return_ci, returns CATE [n_samples, n_treatment], LB [n_samples, n_treatment],
UB [n_samples, n_treatment]
"""
X, treatment, y = convert_pd_to_np(X, treatment, y)
self.fit(X, treatment, y, p)
if p is None:
p = self.propensity
else:
p = self._format_p(p, self.t_groups)
te = self.predict(X, treatment=treatment, y=y, p=p, return_components=return_components)
if not return_ci:
return te
else:
t_groups_global = self.t_groups
_classes_global = self._classes
models_mu_c_global = deepcopy(self.models_mu_c)
models_mu_t_global = deepcopy(self.models_mu_t)
models_tau_c_global = deepcopy(self.models_tau_c)
models_tau_t_global = deepcopy(self.models_tau_t)
te_bootstraps = np.zeros(shape=(X.shape[0], self.t_groups.shape[0], n_bootstraps))
logger.info('Bootstrap Confidence Intervals')
for i in tqdm(range(n_bootstraps)):
te_b = self.bootstrap(X, treatment, y, p, size=bootstrap_size)
te_bootstraps[:, :, i] = te_b
te_lower = np.percentile(te_bootstraps, (self.ate_alpha / 2) * 100, axis=2)
te_upper = np.percentile(te_bootstraps, (1 - self.ate_alpha / 2) * 100, axis=2)
# set member variables back to global (currently last bootstrapped outcome)
self.t_groups = t_groups_global
self._classes = _classes_global
self.models_mu_c = deepcopy(models_mu_c_global)
self.models_mu_t = deepcopy(models_mu_t_global)
self.models_tau_c = deepcopy(models_tau_c_global)
self.models_tau_t = deepcopy(models_tau_t_global)
return (te, te_lower, te_upper)
def estimate_ate(self, X, treatment, y, p=None, bootstrap_ci=False, n_bootstraps=1000, bootstrap_size=10000):
"""Estimate the Average Treatment Effect (ATE).
Args:
X (np.matrix or np.array or pd.Dataframe): a feature matrix
treatment (np.array or pd.Series): a treatment vector
y (np.array or pd.Series): an outcome vector
p (np.ndarray or pd.Series or dict, optional): an array of propensity scores of float (0,1) in the
single-treatment case; or, a dictionary of treatment groups that map to propensity vectors of
float (0,1); if None will run ElasticNetPropensityModel() to generate the propensity scores.
bootstrap_ci (bool): whether run bootstrap for confidence intervals
n_bootstraps (int): number of bootstrap iterations
bootstrap_size (int): number of samples per bootstrap
Returns:
The mean and confidence interval (LB, UB) of the ATE estimate.
"""
te, dhat_cs, dhat_ts = self.fit_predict(X, treatment, y, p, return_components=True)
X, treatment, y = convert_pd_to_np(X, treatment, y)
if p is None:
p = self.propensity
else:
p = self._format_p(p, self.t_groups)
ate = np.zeros(self.t_groups.shape[0])
ate_lb = np.zeros(self.t_groups.shape[0])
ate_ub = np.zeros(self.t_groups.shape[0])
for i, group in enumerate(self.t_groups):
_ate = te[:, i].mean()
mask = (treatment == group) | (treatment == self.control_name)
treatment_filt = treatment[mask]
w = (treatment_filt == group).astype(int)
prob_treatment = float(sum(w)) / w.shape[0]
dhat_c = dhat_cs[group][mask]
dhat_t = dhat_ts[group][mask]
p_filt = p[group][mask]
# SE formula is based on the lower bound formula (7) from Imbens, Guido W., and Jeffrey M. Wooldridge. 2009.
# "Recent Developments in the Econometrics of Program Evaluation." Journal of Economic Literature
se = np.sqrt((
self.vars_t[group] / prob_treatment + self.vars_c[group] / (1 - prob_treatment) +
(p_filt * dhat_c + (1 - p_filt) * dhat_t).var()
) / w.shape[0])
_ate_lb = _ate - se * norm.ppf(1 - self.ate_alpha / 2)
_ate_ub = _ate + se * norm.ppf(1 - self.ate_alpha / 2)
ate[i] = _ate
ate_lb[i] = _ate_lb
ate_ub[i] = _ate_ub
if not bootstrap_ci:
return ate, ate_lb, ate_ub
else:
t_groups_global = self.t_groups
_classes_global = self._classes
models_mu_c_global = deepcopy(self.models_mu_c)
models_mu_t_global = deepcopy(self.models_mu_t)
models_tau_c_global = deepcopy(self.models_tau_c)
models_tau_t_global = deepcopy(self.models_tau_t)
logger.info('Bootstrap Confidence Intervals for ATE')
ate_bootstraps = np.zeros(shape=(self.t_groups.shape[0], n_bootstraps))
for n in tqdm(range(n_bootstraps)):
cate_b = self.bootstrap(X, treatment, y, p, size=bootstrap_size)
ate_bootstraps[:, n] = cate_b.mean()
ate_lower = np.percentile(ate_bootstraps, (self.ate_alpha / 2) * 100, axis=1)
ate_upper = np.percentile(ate_bootstraps, (1 - self.ate_alpha / 2) * 100, axis=1)
# set member variables back to global (currently last bootstrapped outcome)
self.t_groups = t_groups_global
self._classes = _classes_global
self.models_mu_c = deepcopy(models_mu_c_global)
self.models_mu_t = deepcopy(models_mu_t_global)
self.models_tau_c = deepcopy(models_tau_c_global)
self.models_tau_t = deepcopy(models_tau_t_global)
return ate, ate_lower, ate_upper
class BaseXRegressor(BaseXLearner):
"""
A parent class for X-learner regressor classes.
"""
def __init__(self,
learner=None,
control_outcome_learner=None,
treatment_outcome_learner=None,
control_effect_learner=None,
treatment_effect_learner=None,
ate_alpha=.05,
control_name=0):
"""Initialize an X-learner regressor.
Args:
learner (optional): a model to estimate outcomes and treatment effects in both the control and treatment
groups
control_outcome_learner (optional): a model to estimate outcomes in the control group
treatment_outcome_learner (optional): a model to estimate outcomes in the treatment group
control_effect_learner (optional): a model to estimate treatment effects in the control group
treatment_effect_learner (optional): a model to estimate treatment effects in the treatment group
ate_alpha (float, optional): the confidence level alpha of the ATE estimate
control_name (str or int, optional): name of control group
"""
super().__init__(
learner=learner,
control_outcome_learner=control_outcome_learner,
treatment_outcome_learner=treatment_outcome_learner,
control_effect_learner=control_effect_learner,
treatment_effect_learner=treatment_effect_learner,
ate_alpha=ate_alpha,
control_name=control_name)
class BaseXClassifier(BaseXLearner):
"""
A parent class for X-learner classifier classes.
"""
def __init__(self,
outcome_learner=None,
effect_learner=None,
control_outcome_learner=None,
treatment_outcome_learner=None,
control_effect_learner=None,
treatment_effect_learner=None,
ate_alpha=.05,
control_name=0):
"""Initialize an X-learner classifier.
Args:
outcome_learner (optional): a model to estimate outcomes in both the control and treatment groups.
Should be a classifier.
effect_learner (optional): a model to estimate treatment effects in both the control and treatment groups.
Should be a regressor.
control_outcome_learner (optional): a model to estimate outcomes in the control group.
Should be a classifier.
treatment_outcome_learner (optional): a model to estimate outcomes in the treatment group.
Should be a classifier.
control_effect_learner (optional): a model to estimate treatment effects in the control group.
Should be a regressor.
treatment_effect_learner (optional): a model to estimate treatment effects in the treatment group
Should be a regressor.
ate_alpha (float, optional): the confidence level alpha of the ATE estimate
control_name (str or int, optional): name of control group
"""
if outcome_learner is not None:
control_outcome_learner = outcome_learner
treatment_outcome_learner = outcome_learner
if effect_learner is not None:
control_effect_learner = effect_learner
treatment_effect_learner = effect_learner
super().__init__(
learner=None,
control_outcome_learner=control_outcome_learner,
treatment_outcome_learner=treatment_outcome_learner,
control_effect_learner=control_effect_learner,
treatment_effect_learner=treatment_effect_learner,
ate_alpha=ate_alpha,
control_name=control_name)
if ((control_outcome_learner is None) or (treatment_outcome_learner is None)) and (
(control_effect_learner is None) or (treatment_effect_learner is None)):
raise ValueError("Either the outcome learner or the effect learner pair must be specified.")
def fit(self, X, treatment, y, p=None):
"""Fit the inference model.
Args:
X (np.matrix or np.array or pd.Dataframe): a feature matrix
treatment (np.array or pd.Series): a treatment vector
y (np.array or pd.Series): an outcome vector
p (np.ndarray or pd.Series or dict, optional): an array of propensity scores of float (0,1) in the
single-treatment case; or, a dictionary of treatment groups that map to propensity vectors of
float (0,1); if None will run ElasticNetPropensityModel() to generate the propensity scores.
"""
X, treatment, y = convert_pd_to_np(X, treatment, y)
check_treatment_vector(treatment, self.control_name)
self.t_groups = np.unique(treatment[treatment != self.control_name])
self.t_groups.sort()
if p is None:
self._set_propensity_models(X=X, treatment=treatment, y=y)
p = self.propensity
else:
p = self._format_p(p, self.t_groups)
self._classes = {group: i for i, group in enumerate(self.t_groups)}
self.models_mu_c = {group: deepcopy(self.model_mu_c) for group in self.t_groups}
self.models_mu_t = {group: deepcopy(self.model_mu_t) for group in self.t_groups}
self.models_tau_c = {group: deepcopy(self.model_tau_c) for group in self.t_groups}
self.models_tau_t = {group: deepcopy(self.model_tau_t) for group in self.t_groups}
self.vars_c = {}
self.vars_t = {}
for group in self.t_groups:
mask = (treatment == group) | (treatment == self.control_name)
treatment_filt = treatment[mask]
X_filt = X[mask]
y_filt = y[mask]
w = (treatment_filt == group).astype(int)
# Train outcome models
self.models_mu_c[group].fit(X_filt[w == 0], y_filt[w == 0])
self.models_mu_t[group].fit(X_filt[w == 1], y_filt[w == 1])
# Calculate variances and treatment effects
var_c = (y_filt[w == 0] - self.models_mu_c[group].predict_proba(X_filt[w == 0])[:, 1]).var()
self.vars_c[group] = var_c
var_t = (y_filt[w == 1] - self.models_mu_t[group].predict_proba(X_filt[w == 1])[:, 1]).var()
self.vars_t[group] = var_t
# Train treatment models
d_c = self.models_mu_t[group].predict_proba(X_filt[w == 0])[:, 1] - y_filt[w == 0]
d_t = y_filt[w == 1] - self.models_mu_c[group].predict_proba(X_filt[w == 1])[:, 1]
self.models_tau_c[group].fit(X_filt[w == 0], d_c)
self.models_tau_t[group].fit(X_filt[w == 1], d_t)
def predict(self, X, treatment=None, y=None, p=None, return_components=False,
verbose=True):
"""Predict treatment effects.
Args:
X (np.matrix or np.array or pd.Dataframe): a feature matrix
treatment (np.array or pd.Series, optional): a treatment vector
y (np.array or pd.Series, optional): an outcome vector
p (np.ndarray or pd.Series or dict, optional): an array of propensity scores of float (0,1) in the
single-treatment case; or, a dictionary of treatment groups that map to propensity vectors of
float (0,1); if None will run ElasticNetPropensityModel() to generate the propensity scores.
return_components (bool, optional): whether to return outcome for treatment and control seperately
return_p_score (bool, optional): whether to return propensity score
verbose (bool, optional): whether to output progress logs
Returns:
(numpy.ndarray): Predictions of treatment effects.
"""
X, treatment, y = convert_pd_to_np(X, treatment, y)
if p is None:
logger.info('Generating propensity score')
p = dict()
for group in self.t_groups:
p_model = self.propensity_model[group]
p[group] = p_model.predict(X)
else:
p = self._format_p(p, self.t_groups)
te = np.zeros((X.shape[0], self.t_groups.shape[0]))
dhat_cs = {}
dhat_ts = {}
for i, group in enumerate(self.t_groups):
model_tau_c = self.models_tau_c[group]
model_tau_t = self.models_tau_t[group]
dhat_cs[group] = model_tau_c.predict(X)
dhat_ts[group] = model_tau_t.predict(X)
_te = (p[group] * dhat_cs[group] + (1 - p[group]) * dhat_ts[group]).reshape(-1, 1)
te[:, i] = np.ravel(_te)
if (y is not None) and (treatment is not None) and verbose:
mask = (treatment == group) | (treatment == self.control_name)
treatment_filt = treatment[mask]
X_filt = X[mask]
y_filt = y[mask]
w = (treatment_filt == group).astype(int)
yhat = np.zeros_like(y_filt, dtype=float)
yhat[w == 0] = self.models_mu_c[group].predict_proba(X_filt[w == 0])[:, 1]
yhat[w == 1] = self.models_mu_t[group].predict_proba(X_filt[w == 1])[:, 1]
logger.info('Error metrics for group {}'.format(group))
classification_metrics(y_filt, yhat, w)
if not return_components:
return te
else:
return te, dhat_cs, dhat_ts
#hide
from nbdev.export import notebook2script; notebook2script()
```
| github_jupyter |
```
%matplotlib inline
```
# Segmenting the picture of greek coins in regions
This example uses `spectral_clustering` on a graph created from
voxel-to-voxel difference on an image to break this image into multiple
partly-homogeneous regions.
This procedure (spectral clustering on an image) is an efficient
approximate solution for finding normalized graph cuts.
There are two options to assign labels:
* with 'kmeans' spectral clustering will cluster samples in the embedding space
using a kmeans algorithm
* whereas 'discrete' will iteratively search for the closest partition
space to the embedding space.
```
print(__doc__)
# Author: Gael Varoquaux <gael.varoquaux@normalesup.org>, Brian Cheung
# License: BSD 3 clause
import time
import numpy as np
from distutils.version import LooseVersion
from scipy.ndimage.filters import gaussian_filter
import matplotlib.pyplot as plt
import skimage
from skimage.data import coins
from skimage.transform import rescale
from sklearn.feature_extraction import image
from sklearn.cluster import spectral_clustering
# these were introduced in skimage-0.14
if LooseVersion(skimage.__version__) >= '0.14':
rescale_params = {'anti_aliasing': False, 'multichannel': False}
else:
rescale_params = {}
# load the coins as a numpy array
orig_coins = coins()
# Resize it to 20% of the original size to speed up the processing
# Applying a Gaussian filter for smoothing prior to down-scaling
# reduces aliasing artifacts.
smoothened_coins = gaussian_filter(orig_coins, sigma=2)
rescaled_coins = rescale(smoothened_coins, 0.2, mode="reflect",
**rescale_params)
# Convert the image into a graph with the value of the gradient on the
# edges.
graph = image.img_to_graph(rescaled_coins)
# Take a decreasing function of the gradient: an exponential
# The smaller beta is, the more independent the segmentation is of the
# actual image. For beta=1, the segmentation is close to a voronoi
beta = 10
eps = 1e-6
graph.data = np.exp(-beta * graph.data / graph.data.std()) + eps
# Apply spectral clustering (this step goes much faster if you have pyamg
# installed)
N_REGIONS = 25
```
Visualize the resulting regions
```
for assign_labels in ('kmeans', 'discretize'):
t0 = time.time()
labels = spectral_clustering(graph, n_clusters=N_REGIONS,
assign_labels=assign_labels, random_state=42)
t1 = time.time()
labels = labels.reshape(rescaled_coins.shape)
plt.figure(figsize=(5, 5))
plt.imshow(rescaled_coins, cmap=plt.cm.gray)
for l in range(N_REGIONS):
plt.contour(labels == l,
colors=[plt.cm.nipy_spectral(l / float(N_REGIONS))])
plt.xticks(())
plt.yticks(())
title = 'Spectral clustering: %s, %.2fs' % (assign_labels, (t1 - t0))
print(title)
plt.title(title)
plt.show()
```
| github_jupyter |
```
from fugue_notebook import setup
setup()
from dask_sql.integrations import fugue
%%fsql
airports =
LOAD CSV "/tmp/airports.csv"
COLUMNS airport_id:long,name:str,city:str,country:str,iata:str,icao:str,lat:double,lng:double,alt:long,timezone:str,dst:str,type:str,source:str
YIELD DATAFRAME
PRINT
airlines =
LOAD CSV "/tmp/airlines.csv"
COLUMNS airline_id:long,name:str,alias:str,iata:str,icao:str,callsign:str,country:str,active:str
YIELD DATAFRAME
PRINT
airports.native
import pandas as pd
from triad import Schema
from typing import List, Iterable, Dict, Any
df = pd.read_parquet("https://s3.amazonaws.com/bsql/data/air_transport/flight_ontime_2020-01.parquet")
schema = Schema(df.iloc[: , :-1])
print(schema)
import pandas as pd
files = [[f"https://s3.amazonaws.com/bsql/data/air_transport/flight_ontime_2020-0{i}.parquet"] for i in [1,2,3,4,5]]
files_df = pd.DataFrame(files, columns=["path"])
files_df
from typing import Dict, Any, List, Iterable
import os
from shutil import rmtree
def download(files:pd.DataFrame,path:str) -> None:
os.makedirs(path,exist_ok=True)
for file in files["path"]:
fn = os.path.basename(file)
npath = os.path.join(path,fn)
print(npath)
pd.read_parquet(file)[schema.names].to_parquet(npath)
download(files_df.head(2), "/tmp/1.parquet")
%%fsql dask
OUTTRANSFORM files_df
EVEN PREPARTITION ROWCOUNT
USING download(path="/tmp/flights.parquet")
%%fsql dask
LOAD "/tmp/flights.parquet"
PRINT ROWCOUNT
SELECT FL_DATE,CRS_DEP_TIME,DEP_TIME,DEP_DELAY LIMIT 50 PERSIST
YIELD DATAFRAME AS test
#schema: *,ts:datetime,day_of_year:int,hour_of_week:int
def generate_time_metrics(df:pd.DataFrame) -> pd.DataFrame:
date = df["FL_DATE"].astype(str) + " "+df["CRS_DEP_TIME"].astype(str)
df["ts"]=pd.to_datetime(date, format="%Y-%m-%d %H%M")
df["day_of_year"]=df["ts"].dt.dayofyear
df["hour_of_week"]=df["ts"].dt.dayofweek*24+df["ts"].dt.hour
return df
generate_time_metrics(test.as_pandas())
%%fsql dask
LOAD "/tmp/flights.parquet"
TRANSFORM USING generate_time_metrics
SELECT
ts,
day_of_year,
hour_of_week,
ORIGIN AS origin,
DEST AS dest,
OP_UNIQUE_CARRIER AS carrier,
DEP_DELAY AS delay
PERSIST
YIELD DATAFRAME AS flights
PRINT ROWCOUNT
import matplotlib.pyplot as plt
def plot(df:pd.DataFrame,x:Any,y:Any,sort:Any,**kwargs) -> None:
df.sort_values(sort).plot(x=x,y=y,**kwargs)
plt.show()
%%fsql dask
SELECT day_of_year, AVG(delay) AS avg_delay FROM flights GROUP BY day_of_year
OUTPUT USING plot(x="day_of_year",y="avg_delay",sort="day_of_year")
SELECT hour_of_week, AVG(delay) AS avg_delay FROM flights GROUP BY hour_of_week
OUTPUT USING plot(x="hour_of_week",y="avg_delay",sort="hour_of_week")
%%fsql dask
info =
SELECT ts
, carrier
, B.name AS carrier_name
, origin
, C.name AS origin_name
, C.country AS origin_country
, C.lat AS origin_lat
, C.lng AS origin_lng
, dest
, D.name AS dest_name
, D.country AS dest_country
, D.lat AS dest_lat
, D.lng AS dest_lng
, delay
FROM flights AS A
LEFT OUTER JOIN airlines AS B
ON A.carrier = B.iata
LEFT OUTER JOIN airports AS C
ON A.origin = C.iata
LEFT OUTER JOIN airports AS D
ON A.dest = D.iata
WHERE C.lat IS NOT NULL AND C.lng IS NOT NULL
AND D.lat IS NOT NULL AND D.lng IS NOT NULL
PERSIST YIELD DATAFRAME
PRINT ROWCOUNT
SELECT * WHERE origin_country = dest_country AND origin_country = 'United States'
PERSIST YIELD DATAFRAME AS info_us
PRINT ROWCOUNT
def plot_bar(df:pd.DataFrame,x:Any,y:Any,sort:Any,**kwargs) -> None:
df.sort_values(sort).plot.bar(x=x,y=y,**kwargs)
plt.show()
%%fsql dask
SELECT origin, AVG(delay) AS delay FROM info_us GROUP BY origin
SELECT * ORDER BY delay DESC LIMIT 10
OUTPUT USING plot_bar(x="origin",y="delay",sort="delay", title="By Origin")
SELECT dest, AVG(delay) AS delay FROM info_us GROUP BY dest
SELECT * ORDER BY delay DESC LIMIT 10
OUTPUT USING plot_bar(x="dest",y="delay",sort="delay", title="By Dest")
top =
SELECT carrier, COUNT(*) AS ct
FROM info_us GROUP BY carrier
ORDER BY ct DESC LIMIT 20
PERSIST YIELD DATAFRAME
info_top =
SELECT info_us.* FROM info_us INNER JOIN top ON info_us.carrier = top.carrier
SELECT carrier_name, AVG(delay) AS delay FROM info_top GROUP BY carrier_name
SELECT * ORDER BY delay DESC LIMIT 10
OUTPUT USING plot_bar(x="carrier_name",y="delay",sort="delay", title="By Top Carriers")
%%fsql dask
LOAD "/tmp/flights.parquet"
TRANSFORM USING generate_time_metrics
flights =
SELECT
ts,
day_of_year,
hour_of_week,
ORIGIN AS origin,
DEST AS dest,
OP_UNIQUE_CARRIER AS carrier,
DEP_DELAY AS delay
PERSIST
SELECT day_of_year, AVG(delay) AS avg_delay FROM flights GROUP BY day_of_year
OUTPUT USING plot(x="day_of_year",y="avg_delay",sort="day_of_year")
SELECT hour_of_week, AVG(delay) AS avg_delay FROM flights GROUP BY hour_of_week
OUTPUT USING plot(x="hour_of_week",y="avg_delay",sort="hour_of_week")
info =
SELECT ts
, carrier
, B.name AS carrier_name
, origin
, C.name AS origin_name
, C.country AS origin_country
, C.lat AS origin_lat
, C.lng AS origin_lng
, dest
, D.name AS dest_name
, D.country AS dest_country
, D.lat AS dest_lat
, D.lng AS dest_lng
, delay
FROM flights AS A
LEFT OUTER JOIN airlines AS B
ON A.carrier = B.iata
LEFT OUTER JOIN airports AS C
ON A.origin = C.iata
LEFT OUTER JOIN airports AS D
ON A.dest = D.iata
WHERE C.lat IS NOT NULL AND C.lng IS NOT NULL
AND D.lat IS NOT NULL AND D.lng IS NOT NULL
info_us =
SELECT * WHERE origin_country = dest_country AND origin_country = 'United States'
PERSIST
SELECT origin, AVG(delay) AS delay FROM info_us GROUP BY origin
SELECT * ORDER BY delay DESC LIMIT 10
OUTPUT USING plot_bar(x="origin",y="delay",sort="delay", title="By Origin")
SELECT dest, AVG(delay) AS delay FROM info_us GROUP BY dest
SELECT * ORDER BY delay DESC LIMIT 10
OUTPUT USING plot_bar(x="dest",y="delay",sort="delay", title="By Dest")
top =
SELECT carrier, COUNT(*) AS ct
FROM info_us GROUP BY carrier
ORDER BY ct DESC LIMIT 20
info_top =
SELECT info_us.* FROM info_us INNER JOIN top ON info_us.carrier = top.carrier
SELECT carrier_name, AVG(delay) AS delay FROM info_top GROUP BY carrier_name
SELECT * ORDER BY delay DESC LIMIT 10
OUTPUT USING plot_bar(x="carrier_name",y="delay",sort="delay", title="By Top Carriers")
```
| github_jupyter |
```
from brian2 import *
prefs.codegen.target = "numpy"
start_scope()
# Parameters
area = 20000*umetre**2
Cm = 1*ufarad*cm**-2 * area
gl = 5e-5*siemens*cm**-2 * area
El = -65*mV
EK = -90*mV
ENa = 50*mV
g_na = 100*msiemens*cm**-2 * area
g_kd = 30*msiemens*cm**-2 * area
VT = -63*mV
# The model
eqs_HH = '''
dv/dt = (gl*(El-v) - g_na*(m*m*m)*h*(v-ENa) - g_kd*(n*n*n*n)*(v-EK) + I)/Cm : volt
dm/dt = 0.32*(mV**-1)*(13.*mV-v+VT)/
(exp((13.*mV-v+VT)/(4.*mV))-1.)/ms*(1-m)-0.28*(mV**-1)*(v-VT-40.*mV)/
(exp((v-VT-40.*mV)/(5.*mV))-1.)/ms*m : 1
dn/dt = 0.032*(mV**-1)*(15.*mV-v+VT)/
(exp((15.*mV-v+VT)/(5.*mV))-1.)/ms*(1.-n)-.5*exp((10.*mV-v+VT)/(40.*mV))/ms*n : 1
dh/dt = 0.128*exp((17.*mV-v+VT)/(18.*mV))/ms*(1.-h)-4./(1+exp((40.*mV-v+VT)/(5.*mV)))/ms*h : 1
I : amp
'''
group = NeuronGroup(1, eqs_HH,
threshold='v > -40*mV',
refractory='v > -40*mV',
method='exponential_euler')
group.v = El
statemon = StateMonitor(group, 'v', record=True)
spikemon = SpikeMonitor(group, variables='v')
figure(figsize=(9, 4))
for l in range(5):
group.I = rand()*50*nA
run(10*ms)
axvline(l*10, ls='--', c='k')
axhline(El/mV, ls='-', c='lightgray', lw=3)
plot(statemon.t/ms, statemon.v[0]/mV, '-b')
plot(spikemon.t/ms, spikemon.v/mV, 'ob')
xlabel('Time (ms)')
ylabel('v (mV)');
start_scope()
N = 10
neuron_spacing = 50*umetre
width = N/4.0*neuron_spacing
eqs_HH_mod = '''
dv/dt = (gl*(El-v) - g_na*(m*m*m)*h*(v-ENa) - g_kd*(n*n*n*n)*(v-EK) + I)/Cm : volt
dm/dt = 0.32*(mV**-1)*(13.*mV-v+VT)/
(exp((13.*mV-v+VT)/(4.*mV))-1.)/ms*(1-m)-0.28*(mV**-1)*(v-VT-40.*mV)/
(exp((v-VT-40.*mV)/(5.*mV))-1.)/ms*m : 1
dn/dt = 0.032*(mV**-1)*(15.*mV-v+VT)/
(exp((15.*mV-v+VT)/(5.*mV))-1.)/ms*(1.-n)-.5*exp((10.*mV-v+VT)/(40.*mV))/ms*n : 1
dh/dt = 0.128*exp((17.*mV-v+VT)/(18.*mV))/ms*(1.-h)-4./(1+exp((40.*mV-v+VT)/(5.*mV)))/ms*h : 1
I : amp
x : metre
'''
# Neuron has one variable x, its position
# G = NeuronGroup(N, '')
G = NeuronGroup(
N,
eqs_HH_mod,
threshold='v > -40*mV',
refractory='v > -40*mV',
method='exponential_euler'
)
G.v = El
G.x = 'i*neuron_spacing'
statemon = StateMonitor(G, 'v', record=True)
spikemon = SpikeMonitor(G, variables='v')
# # All synapses are connected (excluding self-connections)
# S = Synapses(G, G, 'w : 1')
# # S.connect(condition='i!=j')
# S.connect()
# # Weight varies with distance
# S.w = 'exp(-(x_pre-x_post)**2/(2*width**2))'
S = Synapses(G, G, 'w : 1')
S.connect(condition='i!=j')
S.w = '-exp(-(x_pre-x_post)**2/(2*width**2))'
S2 = Synapses(G, G, 'w : 1')
S2.connect(condition='i==j')
S2.w = 'exp(-(x_pre-x_post)**2/(2*width**2))'
scatter(S.x_pre/um, S.x_post/um, S.w*20)
xlabel('Source neuron position (um)')
ylabel('Target neuron position (um)');
run(10*ms)
plot(statemon.t/ms, statemon.v[0]/mV, '-b')
plot(spikemon.t/ms, spikemon.v/mV, 'ob')
xlabel('Time (ms)')
ylabel('v (mV)');
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
from matplotlib import pyplot as plt
import copy
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
# trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
# testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
# foreground_classes = {'plane', 'car', 'bird'}
# background_classes = {'cat', 'deer', 'dog', 'frog', 'horse','ship', 'truck'}
# fg1,fg2,fg3 = 0,1,2
trainloader = torch.utils.data.DataLoader(trainset, batch_size=256,shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=256,shuffle=False)
import torch.nn as nn
import torch.nn.functional as F
class CNN(nn.Module):
def __init__(self):
super(CNN, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=16, kernel_size=3, padding=0)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(in_channels=16, out_channels=32, kernel_size=3, padding=0)
self.conv3 = nn.Conv2d(in_channels=32, out_channels=32, kernel_size=3, padding=0)
self.fc1 = nn.Linear(512, 256)
self.fc2 = nn.Linear(256, 64)
self.fc3 = nn.Linear(64, 10)
# self.fc4 = nn.Linear(40,10)
# self.fc5 = nn.Linear(10,2)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
# print(x.shape)
x = (F.relu(self.conv3(x)))
x = x.view(x.size(0), -1)
# print(x.shape)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = (self.fc3(x))
# x = F.relu(self.fc4(x))
# x = self.fc4(x)
return x
cnn_net = CNN()#.double()
cnn_net = cnn_net.to("cuda")
import torch.optim as optim
criterion_cnn = nn.CrossEntropyLoss()
optimizer_cnn = optim.SGD(cnn_net.parameters(), lr=0.01, momentum=0.9)
acti = []
loss_curi = []
epochs = 300
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
# zero the parameter gradients
optimizer_cnn.zero_grad()
# forward + backward + optimize
outputs = cnn_net(inputs)
loss = criterion_cnn(outputs, labels)
loss.backward()
optimizer_cnn.step()
# print statistics
running_loss += loss.item()
mini_batch = 50
if i % mini_batch == mini_batch-1: # print every 50 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, i + 1, running_loss / mini_batch))
ep_lossi.append(running_loss/mini_batch) # loss per minibatch
running_loss = 0.0
if(np.mean(ep_lossi) <= 0.01):
break;
loss_curi.append(np.mean(ep_lossi)) #loss per epoch
print('Finished Training')
torch.save(cnn_net.state_dict(),"/content/drive/My Drive/Research/Cheating_data/CIFAR10_class_classify/weights/"+"cnn_3layer_16_32_32"+".pt")
correct = 0
total = 0
with torch.no_grad():
for data in trainloader:
images, labels = data
images, labels = images.to("cuda"), labels.to("cuda")
outputs = cnn_net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the %d train images: %d %%' % (total, 100 * correct / total))
print(total,correct)
correct = 0
total = 0
out = []
pred = []
with torch.no_grad():
for data in testloader:
images, labels = data
images, labels = images.to("cuda"),labels.to("cuda")
out.append(labels.cpu().numpy())
outputs= cnn_net(images)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print(total,correct)
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
with torch.no_grad():
for data in testloader:
images, labels = data
images, labels = images.to("cuda"),labels.to("cuda")
outputs = cnn_net(images)
_, predicted = torch.max(outputs, 1)
c = (predicted == labels).squeeze()
for i in range(4):
label = labels[i]
class_correct[label] += c[i].item()
class_total[label] += 1
for i in range(10):
print('Accuracy of %5s : %2d %%' % (
classes[i], 100 * class_correct[i] / class_total[i]))
```
| github_jupyter |
# kneed -- knee detection in Python
For the purposes of the walkthrough, import `DataGenerator` to create simulated datasets.
In practice, the `KneeLocator` class will be used to identify the knee point.
```
%matplotlib inline
from kneed.data_generator import DataGenerator as dg
from kneed.knee_locator import KneeLocator
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
x = [3.07, 3.38, 3.55, 3.68, 3.78, 3.81, 3.85, 3.88, 3.9, 3.93]
y = [0.0, 0.3, 0.47, 0.6, 0.69, 0.78, 0.845, 0.903, 0.95, 1.0]
kl = KneeLocator(x, y, S=1.0, curve='convex', direction='increasing', interp_method='interp1d')
kl.x_normalized
np.diff(kl.x_normalized).mean()
np.diff(x).mean()
from scipy.signal import argrelextrema
argrelextrema(kl.y_difference, np.greater)
argrelextrema(kl.y_difference, np.less)
kl.y_difference_maxima
plt.plot(kl.x_normalized, kl.y_normalized);
plt.plot(kl.x_difference, kl.y_difference);
np.random.seed(23) # only for the walkthrough
x,y = dg.noisy_gaussian(N=1000)
x[:5],y[:5]
```
The knee is located by passing `x` and `y` values to `knee_locator`.
`S` is the sensitivity parameter
`curve`='concave'
```
kneedle = KneeLocator(x, y, S=1.0, curve='concave', direction='increasing', interp_method='polynomial')
kneedle.plot_knee_normalized()
kneedle.plot_knee()
kneedle.knee
```
There are plotting functions to visualize the knee point on the raw data and the normalized data.
## Average Knee for NoisyGaussian from 50 random iterations
```
knees = []
for i in range(50):
x,y = dg.noisy_gaussian(N=1000)
kneedle = KneeLocator(x, y, direction='increasing', curve='concave', interp_method='polynomial')
knees.append(kneedle.knee)
np.mean(knees)
```
# Test all type of functions
```
x = np.arange(0,10)
y_convex_inc = np.array([1,2,3,4,5,10,15,20,40,100])
y_convex_dec = y_convex_inc[::-1]
y_concave_dec = 100 - y_convex_inc
y_concave_inc = 100 - y_convex_dec
kn = KneeLocator(x, y_convex_inc, curve='convex')
knee_yconvinc = kn.knee
kn = KneeLocator(x, y_convex_dec, curve='convex', direction='decreasing')
knee_yconvdec = kn.knee
kn = KneeLocator(x, y_concave_inc, curve='concave')
knee_yconcinc = kn.knee
kn = KneeLocator(x, y_concave_dec, curve='concave', direction='decreasing')
knee_yconcdec = kn.knee
f, axes = plt.subplots(2, 2, figsize=(10,10));
yconvinc = axes[0][0]
yconvdec = axes[0][1]
yconcinc = axes[1][0]
yconcdec = axes[1][1]
sns.lineplot(x, y_convex_inc, ax=axes[0][0])
yconvinc.vlines(x=knee_yconvinc, ymin=0, ymax=100, linestyle='--')
yconvinc.set_title("curve='convex', direction='increasing'")
sns.lineplot(x, y_convex_dec, ax=axes[0][1])
yconvdec.vlines(x=knee_yconvdec, ymin=0, ymax=100, linestyle='--')
yconvdec.set_title("curve='convex', direction='decreasing'")
sns.lineplot(x, y_concave_inc, ax=axes[1][0])
yconcinc.vlines(x=knee_yconcinc, ymin=0, ymax=100, linestyle='--')
yconcinc.set_title("curve='concave', direction='increasing'")
sns.lineplot(x, y_concave_dec, ax=axes[1][1])
yconcdec.vlines(x=knee_yconcdec, ymin=0, ymax=100, linestyle='--')
yconcdec.set_title("curve='concave', direction='decreasing'");
```
# Polynomial line fit
An example of a "bumpy" line where the traditional `interp1d` spline fitting method does not provide the best estimate for the point of maximum curvature.
We demonstrate that setting the parameter `interp_method='polynomial'` will choose the right point by smoothing the line.
```
x = list(range(90))
y = [
7304.99, 6978.98, 6666.61, 6463.2, 6326.53, 6048.79, 6032.79, 5762.01, 5742.77,
5398.22, 5256.84, 5226.98, 5001.72, 4941.98, 4854.24, 4734.61, 4558.75, 4491.1,
4411.61, 4333.01, 4234.63, 4139.1, 4056.8, 4022.49, 3867.96, 3808.27, 3745.27,
3692.34, 3645.55, 3618.28, 3574.26, 3504.31, 3452.44, 3401.2, 3382.37, 3340.67,
3301.08, 3247.59, 3190.27, 3179.99, 3154.24, 3089.54, 3045.62, 2988.99, 2993.61,
2941.35, 2875.6, 2866.33, 2834.12, 2785.15, 2759.65, 2763.2, 2720.14, 2660.14,
2690.22, 2635.71, 2632.92, 2574.63, 2555.97, 2545.72, 2513.38, 2491.57, 2496.05,
2466.45, 2442.72, 2420.53, 2381.54, 2388.09, 2340.61, 2335.03, 2318.93, 2319.05,
2308.23, 2262.23, 2235.78, 2259.27, 2221.05, 2202.69, 2184.29, 2170.07, 2160.05,
2127.68, 2134.73, 2101.96, 2101.44, 2066.4, 2074.25, 2063.68, 2048.12, 2031.87
]
kneedle = KneeLocator(x, y, S=1.0, curve='convex', direction='decreasing')
kneedle.plot_knee_normalized()
kneedle = KneeLocator(x, y, S=1.0, curve='convex', direction='decreasing', interp_method='polynomial')
kneedle.plot_knee_normalized()
```
| github_jupyter |
# Project - Measure Interpolation Impact

## Goal of Project
- The goal of the project is to see how big impact interpolation can have on results.
- The focus is mainly on step 2.
- To see the impact we will make simple model usages.
- The project will not go into details of steps 3 to 5.
## Step 1: Acquire
- Explore problem
- Identify data
- Import data
### Step 1.a: Import libraries
- Execute the cell below (SHIFT + ENTER)
```
import pandas as pd
import numpy as np
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score
import matplotlib.pyplot as plt
%matplotlib inline
```
### Step 1.b: Read the data
- Use ```pd.read_parquet()``` to read the file `files/weather-predict.parquet`
- NOTE: Remember to assign the result to a variable (e.g., ```data```)
- Apply ```.head()``` on the data to see all is as expected
## Step 2: Prepare
- Explore data
- Visualize ideas
- Cleaning data
### Step 2.a: Check the data types
- This step tells you if some numeric column is not represented numeric.
- Get the data types by ```.dtypes```
### Step 2.b: Check the length, null-values, and zero values
- Check the length
- HINT: Use `len()`
- Check the number of null-values
- HINT: Use `.isna().sum()`
- Check the number of zero-values
- HINT: Use `(data == 0).sum()`
### Step 2.c: Baseline
- Check the correlation to have a measure if we did nothing
- HINT: Use `corr()`
### Step 2.d: Prepare data
- We know `Pressure+24` has NaN and 0 values.
- These are not correct values and we cannot use them in our model.
- Create a `dataset` without these rows.
- HINT: Use filters like `data[data['Pressure+24h'] != 0]` and `dropna()`
### Step 2.e: Check the size and zero values
- Check the size of datasets `data` and `datasets`
- Check how many zero-values each dataset has
### Step 2.f: Check the correlation
- For fun check the correlation of `dataset`
- Then do the same after you interpolated 0 values
- HINT: Apply `replace` and `interpolate`
- Does the result surprice you?
- Notice how much interpolation improves the result
### Step 2.g: Linear Regression Function
- Create function `regression_score` to calculate the r-square score
- It should take independent features X and dependent feature y
- Then split that into training and testing sets.
- Fit the training set.
- Predict the test set.
- Return the r-square score
### Step 2.h: Test baseline
- Test the `regression_score` function on `dataset`
### Step 2.i: Test on interploated dataset
- Make a interpolated dataset
- Get the result (from `regression_score`) for interpolated dataset
| github_jupyter |
```
%load_ext rpy2.ipython
%matplotlib inline
import pandas as pd
import numpy as np
from fbprophet import Prophet
import logging
logging.getLogger('fbprophet').setLevel(logging.ERROR)
import warnings
warnings.filterwarnings("ignore")
df = pd.DataFrame({
'ds': pd.date_range(start='2020-01-01', periods=20),
'y': np.arange(20),
})
m = Prophet(weekly_seasonality=False)
m.fit(df)
%%R
library(prophet)
df <- data.frame(
ds=seq(as.Date("2020-01-01"), by = "day", length.out = 20),
y=1:20
)
m <- prophet(df, weekly.seasonality=FALSE)
```
### Saving models
It is possible to save fitted Prophet models so that they can be loaded and used later.
In R, this is done with `saveRDS` and `readRDS`:
```
%%R
saveRDS(m, file="model.RDS") # Save model
m <- readRDS(file="model.RDS") # Load model
```
In Python, models should not be saved with pickle; the Stan backend attached to the model object will not pickle well, and will produce issues under certain versions of Python. Instead, you should use the built-in serialization functions to serialize the model to json:
```
import json
from fbprophet.serialize import model_to_json, model_from_json
with open('serialized_model.json', 'w') as fout:
json.dump(model_to_json(m), fout) # Save model
with open('serialized_model.json', 'r') as fin:
m = model_from_json(json.load(fin)) # Load model
```
The json file will be portable across systems, and deserialization is backwards compatible with older versions of fbprophet.
### Flat trend and custom trends
For time series that exhibit strong seasonality patterns rather than trend changes, it may be useful to force the trend growth rate to be flat. This can be achieved simply by passing `growth=flat` when creating the model:
```
%%R
m <- prophet(df, growth='flat')
m = Prophet(growth='flat')
```
Note that if this is used on a time series that doesn't have a constant trend, any trend will be fit with the noise term and so there will be high predictive uncertainty in the forecast.
To use a trend besides these three built-in trend functions (piecewise linear, piecewise logistic growth, and flat), you can download the source code from github, modify the trend function as desired in a local branch, and then install that local version. This PR provides a good illustration of what must be done to implement a custom trend: https://github.com/facebook/prophet/pull/1466/files.
### Updating fitted models
A common setting for forecasting is fitting models that need to be updated as additional data come in. Prophet models can only be fit once, and a new model must be re-fit when new data become available. In most settings, model fitting is fast enough that there isn't any issue with re-fitting from scratch. However, it is possible to speed things up a little by warm-starting the fit from the model parameters of the earlier model. This code example shows how this can be done in Python:
```
def stan_init(m):
"""Retrieve parameters from a trained model.
Retrieve parameters from a trained model in the format
used to initialize a new Stan model.
Parameters
----------
m: A trained model of the Prophet class.
Returns
-------
A Dictionary containing retrieved parameters of m.
"""
res = {}
for pname in ['k', 'm', 'sigma_obs']:
res[pname] = m.params[pname][0][0]
for pname in ['delta', 'beta']:
res[pname] = m.params[pname][0]
return res
df = pd.read_csv('../examples/example_wp_log_peyton_manning.csv')
df1 = df.loc[df['ds'] < '2016-01-19', :] # All data except the last day
m1 = Prophet().fit(df1) # A model fit to all data except the last day
%timeit m2 = Prophet().fit(df) # Adding the last day, fitting from scratch
%timeit m2 = Prophet().fit(df, init=stan_init(m1)) # Adding the last day, warm-starting from m1
```
As can be seen, the parameters from the previous model are passed in to the fitting for the next with the kwarg `init`. In this case, model fitting was almost 2x faster when using warm starting. The speedup will generally depend on how much the optimal model parameters have changed with the addition of the new data.
There are few caveats that should be kept in mind when considering warm-starting. First, warm-starting may work well for small updates to the data (like the addition of one day in the example above) but can be worse than fitting from scratch if there are large changes to the data (i.e., a lot of days have been added). This is because when a large amount of history is added, the location of the changepoints will be very different between the two models, and so the parameters from the previous model may actually produce a bad trend initialization. Second, as a detail, the number of changepoints need to be consistent from one model to the next or else an error will be raised because the changepoint prior parameter `delta` will be the wrong size.
| github_jupyter |
### PROBLEM 1
Assume s is a string of lower case characters.
Write a program that counts up the number of vowels contained in the string s. Valid vowels are: 'a', 'e', 'i', 'o', and 'u'. For example, if s = 'azcbobobegghakl', your program should print:
Number of vowels: 5
```
def ex_ps1(s):
vowels = ['a', 'e', 'i', 'o', 'u']
list_s = list(s)
vowels_counter = 0
for i in list_s:
for v in vowels:
if i == v:
vowels_counter= vowels_counter+1
print("Number of vowels: "+str(vowels_counter))
ex_ps1("azcbobobegghakl")
```
### PROBLEM 2
Assume s is a string of lower case characters.
Write a program that prints the number of times the string 'bob' occurs in s. For example, if s = 'azcbobobegghakl', then your program should print
Number of times bob occurs is: 2
```
def ex_ps2():
s = 'azcbobobegghakl'
special_code = "bob"
lower_letter = 0
max_letter = 3
word_len = len(s)
bobs_counter = 0
while max_letter <= word_len:
looking_for_bob = s[lower_letter:max_letter]
if looking_for_bob == special_code:
lower_letter = lower_letter+1
max_letter = max_letter+1
bobs_counter = bobs_counter+1
else:
lower_letter = lower_letter+1
max_letter = max_letter+1
print("Number of times bob occurs is: "+str(bobs_counter))
ex_ps2()
```
### PROBLEM 3
Assume s is a string of lower case characters.
Write a program that prints the longest substring of s in which the letters occur in alphabetical order. For example, if s = 'azcbobobegghakl', then your program should print
Longest substring in alphabetical order is: beggh
In the case of ties, print the first substring. For example, if s = 'abcbcd', then your program should print
Longest substring in alphabetical order is: abc
Note: This problem may be challenging. We encourage you to work smart. If you've spent more than a few hours on this problem, we suggest that you move on to a different part of the course. If you have time, come back to this problem after you've had a break and cleared your head.
```
def ex_ps3():
s = 'abcbcd'
first_letter_number = 0
last_letter_number = 1
word_len = len(s)
longest_substring_counter = 0
longest_substring = ""
current_substring_counter = 0
current_substring = ""
while last_letter_number <= word_len:
current_string = s[first_letter_number:last_letter_number]
if current_string == ''.join(sorted(current_string)):
last_letter_number = last_letter_number+1
current_substring_counter = current_substring_counter+1
current_substring = current_string
if current_substring_counter > longest_substring_counter:
longest_substring_counter = current_substring_counter
longest_substring = current_substring
else:
print("Finished abc time to move")
first_letter_number = first_letter_number+1
last_letter_number = first_letter_number+1
current_substring_counter=0
current_substring=""
current_string = s[first_letter_number:last_letter_number]
print("Longest substring in alphabetical order is: "+str(longest_substring))
ex_ps3()
```
| github_jupyter |
```
import numpy as np
import seaborn as sb
import pandas
import sys
import itertools
import matplotlib.pyplot as plt
import nltk
import csv
import datetime
%matplotlib notebook
```
# Recurrent Neural Networks (RNNs)
The following is taken from the excellent tutorial on RNNs and adapted a bit.
http://www.wildml.com/2015/09/recurrent-neural-networks-tutorial-part-2-implementing-a-language-model-rnn-with-python-numpy-and-theano/
Our goal is to learn dependencies between words from a corpus given a recurrent neural network architecture.
## Step 1: Reading in the corpus
In the following, we are going to use a Shakespeare corpus (that only includes his plays, not the sonnets) to train our RNN.
In order to get our network to properly train sentences, we are going to prepend sentence start tokens and append sentence end tokens to each of the corpus's sentences.
We are going to make use of the natural language toolkit to help with some processing.
In order to get this, you need to `pip3 install nltk` and then type from a python console `ntlk.download()` and select the popular packages for downloading.
```
vocabularySize = 8000
unknownToken = "UNK"
sentenceStartToken = "S_START"
sentenceEndToken = "S_END"
# Read the data and append SENTENCE_START and SENTENCE_END tokens
print("Reading shakespeare plays...")
with open('data/shakespeare_plays.txt') as f:
# split the data into sentences using nltk magic
# this simply iterates through each line, lowercases it, and then
# sends it to nltk, which splits the lines into sentences according
# to English punctuation rules (i.e., .!?) and also handles problem cases
sentences = itertools.chain(*[nltk.sent_tokenize(x.lower()) for x in f])
# for each of the sentences we prepend S_START and
# and append S_END tokens for our network to learn
sentences = ["%s %s %s" % (sentenceStartToken, x, sentenceEndToken) for x in sentences]
print("read {:d} sentences".format(len(sentences)))
```
Now we are going to use ntlk to tokenize our sentences - tokenize in this case means to split it into words and sentence markers (such as ,.! and so on). In our case, we would like to get words.
```
# now get words from the sentences - nltk uses much smarter methods
# to do this than our crude regular expressions from the first assignment
# but: if also takes a lot longer...
tokenizedSentences = [nltk.word_tokenize(sent) for sent in sentences]
# now we determine the unigram
unigram = nltk.FreqDist(itertools.chain(*tokenizedSentences))
print ("found {:d} unique words".format(len(unigram.items())))
```
## Step 2: Preparing the data structures
The RNN will not directly work on words, but will process each word as its index in the corresponding unigram of the text.
So, we will use nltk to give us the unigram and the indices, and then create a variable `index2word` and `word2index` that will convert back and forth between the two representations.
The RNN will be trained on the output of `word2index`.
```
# use only the vocabularySize most common words - yes, nltk has a
# helper function for that, too ^^
vocab = unigram.most_common(vocabularySize-1)
# create index2word and word2index indexing vectors that we need later
index2word = [x[0] for x in vocab]
# don't forget our unknown token:
index2word.append(unknownToken)
# and reverse index2word:
word2index = dict([(w,i) for i,w in enumerate(index2word)])
print("We choose a vocabulary size of {:d}".format(vocabularySize))
print("The least frequent word in the vocabulary is '{:s}' and appeared {:d} times.".format(vocab[-1][0], vocab[-1][1]))
print("The most frequent non start/end token word in the vocabulary is '{:s}' and appeared {:d} times.".format(vocab[2][0], vocab[2][1]))
```
Note that we restricted our vocabulary to 8000, so the unigram will contain words that our not in our dictionary.
We have to replace these words by the UNK token in order to deal with them during training and testing.
```
# now we need to go back and replace all words that are NOT
# in the vocabulary with the UNK token!
for i, sent in enumerate(tokenizedSentences):
tokenizedSentences[i] = [w if w in word2index else unknownToken for w in sent]
print("\nExample sentence: '{:s}'".format(sentences[1000]))
print("\nExample sentence after Pre-processing:",tokenizedSentences[1000])
```
## Step 3: Create training data
This is the easiest part. We simply get all the words and put them into an array.
```
# create training data for the network
Xtrain = np.asarray([[word2index[w] for w in sentence[:-1]] for sentence in tokenizedSentences])
ytrain = np.asarray([[word2index[w] for w in sentence[1:]] for sentence in tokenizedSentences])
# show examples sentences for X and y
for i in Xtrain[10]:
print(index2word[i])
for i in ytrain[10]:
print(index2word[i])
```
## Step 4: The RNN
Now for the actual beef of this. In the following, we will create an RNN-class that handles all the training for us.
The reason for doing a class is that this nicely capsulates all the functionality for us. Also, all current python interfaces to neural network tools (such as tensorflow, Theano, etc.) use class-based approaches, so that we can get some feeling here as well.
### Step 4.1: Initialization
Let's first add the constructor to the class:
```
class RNN:
# the constructor of the class - this function gets automatically
# called if you make a new instance of the class
# the first variable is the name under which you refer to the
# current instance itself and is usually named "self"
# wordDim: the number words in our dictionary
# hiddenDim: the number of hidden states or "memory"
# bpttTruncate: the number of time-steps we can go back
def __init__(self, wordDim, hiddenDim=100, bpttTruncate=4):
# assign the instance variables
self.wordDim = wordDim
self.hiddenDim = hiddenDim
self.bpttTruncate = bpttTruncate
# initialize the network weights with small random numbers
# there is a bit of "witchcraft" here in how you initialize these
# for now, let's just say that small, random values are fine, but that
# you can do better with [-1/sqrt(n),1/sqrt(n)]
self.U = np.random.uniform(-np.sqrt(1./wordDim), np.sqrt(1./wordDim), (hiddenDim, wordDim))
self.V = np.random.uniform(-np.sqrt(1./hiddenDim), np.sqrt(1./hiddenDim), (wordDim, hiddenDim))
self.W = np.random.uniform(-np.sqrt(1./hiddenDim), np.sqrt(1./hiddenDim), (hiddenDim, hiddenDim))
```
### Step 4.2: The forward propagation and the loss function
Now let's define one forward pass through the RNN. For this, we need the output of the final layer, which is an array of probabilities for each of the words in our dictionary.
There are many ways to define this, but a popular loss is the softmax-loss, which uses the softmax-function $S$ that takes as input a vector $\vec{x}$ of dimension $d$ and returns as output:
$S(\vec{x}) = \frac{e^{\vec{x}}}{\sum_{k=1}^{D}e^{x_k}}$
One easy view of the effects of this function is that it stretches the contrast of the input, such that high elements of the vector in the input will be even higher in the output. The output is normalized to be between 0 and 1, and that gives us basically a probabilistic output!
Because of numerical issues with the exponential function, large elements will lead to overflow (think $e^{1000}$), so people use the fact that the function $S(\vec{x})=S(C\vec{x})$ for an arbitrary $C$, which we can freely choose. In practice $C=max_k(x_k)$
```
# output loss for the final layer
def softmax(x):
xt = np.exp(x - np.max(x))
return xt / np.sum(xt)
```
Now for the actual forward pass.
```
# one forward pass
def forwardProp(self, x):
# how many time steps we have - this is the number of
# words in the sentence!
T = len(x)
# we save all hidden states in s, because we need them later
# we reserve one additional element for the initial hidden state
s = np.zeros((T + 1, self.hiddenDim))
# the outputs for each time step - each time step will predict
# probabilities for all wordDim words!!!
o = np.zeros((T, self.wordDim))
# now go through each time step
for t in np.arange(T):
# the current hidden state is the activation function of
# the input, which is the previous hidden state multiplied
# by W and the addition of the current variable weighted by U
# in order to multiply U with x[t], we would need to do some
# sort of one-hot encoding of x[t]! This would be done in
# standard NN packages
# but we can also use a trick: we can simply index into
# our weight array U with x[t], which will be the same
# as multiplying U with a one-hot vector
s[t] = np.tanh(self.U[:,x[t]] + self.W.dot(s[t-1]))
o[t] = softmax(self.V.dot(s[t]))
return [o, s]
# we only do this here, since we left the "class" scope and are
# doing an "outside" decleration of that class functionality
RNN.forwardProp=forwardProp
```
### Step 4.3: Predicting something with the RNN
The output of the RNN is a large vector of probabilities, with one element for each of our dictionary words.
So, if we want to predict the "best" word, we simply need to select the index with the highest probability.
```
def predict(self, x):
# do forward propagation and the index of the highest score
o, _ = self.forwardProp(x)
return np.argmax(o, axis=1)
RNN.predict = predict
```
### Testing the forward pass
Let's choose our example sentence and push it through.
```
np.random.seed(1)
# init
model = RNN(vocabularySize)
# do one pass with an example sentence that has 11 words
o, s = model.forwardProp(Xtrain[40])
# so our output will have probabilities for each of the 11 words for
# all entries in our dictionary
print(o.shape)
print(o)
# here are the hidden states [we have one more to model the initial state]
print(s.shape)
```
These probabilities are completely random, of course, since we actually have not trained the model!
### Predicting something with the model
Let's use that example sentence to predict the words that we would get and compare that to the actual words that we wanted to have.
```
predictions = model.predict(Xtrain[40])
print(predictions.shape)
print(predictions)
for idx,i in enumerate(predictions):
print('predicted:',index2word[i],' wanted:',index2word[ytrain[40][idx]])
```
As we can see, we don't have much of a match, which is understandable, since all the weights are currently random.
### Step 4.4 Determine loss
The final layer calculates the output, which consists of the probabilities for each word in the dictionary. Now, to determine the full loss, we need to of course do this **for each** of the sentences and then sum up the log of the probabilities to keep things small and easy.
```
def calculateTotalLoss(self, x, y):
# initialize loss
L = 0
# go through each sentence of the target
for i in np.arange(len(y)):
# do a forward pass and get the output (and the state for which we do not care here)
o, _ = self.forwardProp(x[i])
# now we check the output for those words that are contained
# in the target sentence:
correctWordProbabilities = o[np.arange(len(y[i])), y[i]]
# this tells us how wrong we are, so we add that to the loss
L += -1 * np.sum(np.log(correctWordProbabilities))
return L
# this one is easy, we just some up all the words we have in the
# target set
def calculateLoss(self, x, y):
# total loss divided by the number of training examples
N = np.sum((len(y_i) for y_i in y))
return self.calculateTotalLoss(x,y)/N
RNN.calculateTotalLoss = calculateTotalLoss
RNN.calculateLoss = calculateLoss
```
Let's test this loss. Running this on the full dataset will be very slow, so here we only use a subset to show that our initial loss is close to the random prediction:
```
print("Expected Loss for random predictions: {:f}".format(np.log(vocabularySize)))
# running this on the full data set is prohibitively slow
# so just use the first 100 to "approximate"
print("Actual loss: {:f}".format(model.calculateLoss(Xtrain[:100], ytrain[:100])))
```
### Step 4.5: Backpropagation through time (BPTT)
Here comes the second piece of magic and the core of RNNs: the back-propagation through time.
If you remember our discussion on training standard multi-layer networks, you also remember backpropagation.
Here, the algorithm is exactly the same with the added twist that time gets unfolded - note that we would need to do this for the full length of the training data!
The actual weight updates are calculated in the exact same way as for the standard backprop algorithm.
You initialize with the output of the last layer and then push back through all hidden layers.
Note that a full understanding of this algorithm requires the calculation of a rather lengthy set of derivatives. For a full explanation and derivation, see here:
https://github.com/go2carter/nn-learn/blob/master/grad-deriv-tex/rnn-grad-deriv.pdf
Note also that there are many more methods to implement this, including shortening the loop:
https://gist.github.com/m-liu/36c5e78d98c92fd8fa0c0f1efe2269c4
```
def bptt(self, x, y):
# number of "time steps" - again this is the length of the sentence!
T = len(y)
# push everything through the network once
o, s = self.forwardProp(x)
# these store the gradients
dLdU = np.zeros(self.U.shape)
dLdV = np.zeros(self.V.shape)
dLdW = np.zeros(self.W.shape)
# final layer initialization:
delta_o = o
delta_o[np.arange(len(y)), y] -= 1.
# now we take a look at each output
# and go backwards through each word in the sentence
for t in np.arange(T)[::-1]:
# update of loss with respect to hidden node
dLdV += np.outer(delta_o[t], s[t].T)
# now calculate first delta
delta_t = self.V.T.dot(delta_o[t]) * (1 - (s[t] ** 2))
# now unfold network and go back through time
# we do a maximum of self.bpttTruncate time travel steps
# the max's are there to avoid boundary effects
# the higher this parameter, the better in "theory"
for bpttStep in np.arange(max(0, t-self.bpttTruncate), t+1)[::-1]:
#print("bptt: step t={:d} bptt step={:d} ".format(t, bpttStep))
# update W
dLdW += np.outer(delta_t, s[bpttStep-1])
# update U
dLdU[:,x[bpttStep]] += delta_t
# change delta for next step
delta_t = self.W.T.dot(delta_t) * (1 - s[bpttStep-1] ** 2)
return [dLdU, dLdV, dLdW]
RNN.bptt = bptt
```
### Step 4.6: Make training
In order to demonstrate the BPTT, we implement a function that only does one update step of all weights.
This is a very simple gradient descent update on all three parameters.
```
def gd1step(self,Xtrain,ytrain,learningRate):
# first calculate the gradients
dLdU, dLdV, dLdW = self.bptt(Xtrain, ytrain)
# then change parameters according to gradients and learning rate
self.U -= learningRate * dLdU
self.V -= learningRate * dLdV
self.W -= learningRate * dLdW
RNN.gd1step = gd1step
```
Now for the actual training. Here we put in the gradient descent.
```
# gradient descent training
# - model: The RNN model instance
# - Xtrain: The training data set [words x sentences]
# - ytrain: The target training data [words [shifted by one] x sentences]
# - learningRate: Initial learning rate for SGD
# - nEpoch: Number of times to iterate through the complete dataset
# - evaluateLossEvery: Evaluate the loss every epochs
def train(model, Xtrain, ytrain, learningRate=0.005, nEpoch=100, evaluateLossEvery=5):
# keep track of the losses so we can plot them later
losses = []
numExamplesSeen = 0
# loop through all epochs
for epoch in range(nEpoch):
# do we want to evaluate the loss?
# caution: this can be expensive!
if (epoch % evaluateLossEvery == 0 and epoch>0):
loss = model.calculateLoss(Xtrain, ytrain)
losses.append((numExamplesSeen, loss))
time = datetime.datetime.now().strftime('%Y-%m-%d %H:%M:%S')
print ("{:s}: Loss after numExamplesSeen={:d} epoch={:d}: {:f}".format(time, numExamplesSeen, epoch, loss))
# Adjust the learning rate if loss increases
#if (len(losses) >= 1 and losses[-1][1] >= losses[-2][1]):
# learningRate = learningRate * 0.5
# print("Setting learning rate to {:f}".format(learning_rate))
sys.stdout.flush()
# for each training sentence...
for i in range(len(ytrain)):
# do one gradient step
model.gd1step(Xtrain[i], ytrain[i], learningRate)
numExamplesSeen += 1
```
### Step 5: Testing
Let's test how fast one step is for one sentence.
```
np.random.seed(1)
model = RNN(vocabularySize)
%timeit model.gd1step(Xtrain[40], ytrain[40], 0.005)
```
Ouch. Now imagine that for one whole epoch, where we have 160K sentences, which means roughly one hour to go through our full training set! In order to do good training, we should probably do a few 10s of epochs!!
```
np.random.seed(1)
# Train on a small subset of the data to see what happens
model = RNN(vocabularySize)
losses = train(model, Xtrain[:10], ytrain[:10], nEpoch=1000, evaluateLossEvery=100)
indexToTest=200
predictions = model.predict(Xtrain[indexToTest])
print(predictions.shape)
print(predictions)
for idx,i in enumerate(predictions):
print('predicted:',index2word[i],' wanted:',index2word[ytrain[indexToTest][idx]])
```
Much better :)
```
def generateSentence(model):
# since the model predict indices, we are going to get
# index values into a variable called newSentence
# sentence beginning is defined by the sentenceStartToken
newSentence = [word2index[sentenceStartToken]]
# from this, we predict until we get sentenceEndToken
while not newSentence[-1] == word2index[sentenceEndToken]:
# that's the main part - here we use the model to predict
# the next words, given the current state
nextWordProbabilities,_ = model.forwardProp(newSentence)
# from this, we will sample a random word
sampledWord = word2index[unknownToken]
# iterate through list until we hit word that is not unknown
while sampledWord == word2index[unknownToken]:
samples = np.random.multinomial(1, nextWordProbabilities[-1])
sampledWord = np.argmax(samples)
newSentence.append(sampledWord)
# now convert the index into strings
sentence = [index2word[x] for x in newSentence[1:-1]]
return sentence
np.random.seed(1)
# Train on a small subset of the data to see what happens
modelBig = RNN(vocabularySize)
losses = train(modelBig, Xtrain[:10000], ytrain[:10000], nEpoch=50, evaluateLossEvery=5)
numSentences = 10
sentenceMinLength = 3
for i in range(numSentences):
sentence = []
# We want long sentences, not sentences with one or two words
while len(sentence) <= sentenceMinLength:
sentence = generateSentence(modelBig)
print(" ".join(sentence))
```
## Interpreting the results
Example sentences do not look too promising, even though training proceeded for quite some time.
In fact, this implementation does not suffer from computational resources (i.e., simply training longer would be better), but from a very fundamental issue.
# Reading in characters instead
Our model is based on word prediction. What about predicting the next character instead?
Let's try:
```
# reading Shakespeare as full text
allData = open('data/shakespeare_plays.txt', 'r').read()
# splitting into characters done smartly making the use of sets/lists
characters = list(set(allData.lower()))
# checking the size of the data
dataSize = len(allData)
vocabularySize = len(characters)
print('Shakespeare has {:d} characters in total with {:d} unique characters in the vocabulary.'.format(dataSize, vocabularySize))
# creating dictionaries of indices and converting back
character2Index = { ch:i for i,ch in enumerate(characters) }
index2character = { i:ch for i,ch in enumerate(characters) }
# make the training data
p=0
# this will hold the sentences for training and testing
XtrainChar=list()
ytrainChar=list()
# this is the number of characters we want to investigate
scope = 25
# now go through our data
while p+scope+1<=dataSize:
# grab the current scope
i = [character2Index[ch] for ch in allData[p:p+scope].lower()]
XtrainChar.append(i)
# offset the scope by one
t = [character2Index[ch] for ch in allData[p+1:p+scope+1].lower()]
ytrainChar.append(t)
p=p+scope
np.random.seed(1)
# Train on a small subset of the data to see what happens
modelChar = RNN(vocabularySize)
losses = train(modelChar, XtrainChar[:1000], ytrainChar[:1000], nEpoch=200, evaluateLossEvery=1,learningRate=0.001)
pred=modelChar.predict(XtrainChar[10])
for idx,p in enumerate(pred):
print("p:",index2character[p]," wanted:",index2character[ytrainChar[10][idx]])
modelChar = RNN(vocabularySize)
probs,_=modelChar.forwardProp(XtrainChar[0])
len(probs)
x=[character2Index[ch] for ch in ['']]
string=list()
string.append(index2character[1])
for i in range(100):
probs,_=modelChar.forwardProp(x)
idx = np.random.choice(np.arange(84),1,p=probs[0])
x.append(idx[0])
string.append(index2character[idx[0]])
print(''.join(string))
```
| github_jupyter |
# Studying Stock Returns vs Social Sentiments
Are stock returns and social sentiments related? What is the relationship if we expect one?
And more importantly, could we use social sentiments to predict stock returns?
In this study, we will quickly look at **Social Sentiments** on **Tweets** using the `TextBlob` and `ntlk.vadar` package. And we will compare daily sentiments with daily stock returns.
The **Stock returns** in question actually would be the [**Alpha**](https://corporatefinanceinstitute.com/resources/knowledge/finance/alpha/) of the stock, where
$$ \alpha = {R}-{Rf}- \beta ({Rm} - {Rf}) $$
for our purpose, we are going to assume that ${Rf}$ is 0
### Preliminary
For this study, we will use `pandas_datareader` to get historical returns;
And `vaderSentiment` and `TextBlob` for Sentiment Analysis
```
import pandas as pd
import numpy as np
from functools import reduce
import re
from unidecode import unidecode
# sentiment analysis
from vaderSentiment.vaderSentiment import SentimentIntensityAnalyzer
from textblob import TextBlob
from datetime import timedelta
# pandas data reader
import pandas_datareader.data as web
from datetime import datetime as dt
# Alpha Vantage requirements
av_api_key = 'TGXS2LEUKMHXTKT5'
from alpha_vantage.timeseries import TimeSeries
ts = TimeSeries( key = av_api_key, output_format = 'pandas')
# plotting
%matplotlib inline
import seaborn as sns
import matplotlib.pyplot as plt
import plotly.plotly as py
import plotly.graph_objs as go
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
import cufflinks as cf
# Required for Plot.ly Offline
init_notebook_mode(connected=True)
# Cufflinks bind plotly to pandas dataframe in IPython Notebooks
cf.set_config_file(offline = False, world_readable = True, theme = 'ggplot')
```
## Getting Stock Returns
for our study, we are interested in the [**FANG** stocks](https://www.investopedia.com/terms/f/fang-stocks-fb-amzn.asp): `FB`, `AMZN`, `NFLX`, [`GOOG`](https://investorplace.com/2019/01/goog-google-stock-split/)
The [relevant benchmark would then be NASDAQ](https://www.forbes.com/sites/jaysomaney/2016/12/30/comparing-facebook-amazon-netflix-and-google-aka-fang-performance-in-2016/#57a2bf8952f9), which we'll use `QQQ` as proxy for.
We'll also get `SPY` just to test our calculated **Beta** vs Yahoo! Finance. The calcuation is reference from [here](https://medium.com/python-data/capm-analysis-calculating-stock-beta-as-a-regression-in-python-c82d189db536)
```
l_symbols = ['FB','AMZN', 'NFLX', 'GOOG', 'QQQ', 'SPY']
sdate = dt(2018,1,1)
edate = dt(2019,1,23)
yhoo_data = web.DataReader( l_symbols, 'yahoo', sdate, edate)
yhoo_data.head()
data = yhoo_data.stack()
data.reset_index(inplace = True)
l_col_to_keep = ['Date', 'Symbols', 'Adj Close', 'Volume']
r_dfs = []
# 1
for sym in l_symbols:
r_df = data[ data.Symbols == sym ].loc[:, l_col_to_keep]
r_df = r_df.set_index('Date')
r_df[f'r({sym})'] = r_df['Adj Close']/ r_df['Adj Close'].shift(1) - 1
# or just do df.pct_change()
r_dfs.append( r_df.iloc[1:,:])
# 2
df = reduce( lambda x, y : pd.concat([x,y], axis =1),
[ r_df.iloc[:,-1] for r_df in r_dfs ]
)
df.sort_values(by = 'Date',ascending = False).head(3)
from scipy import stats
def GetBeta( r_sym , r_benchmark):
slope, intercept, r_value, p_value, std_err = stats.linregress( r_sym, r_benchmark)
return slope
def GetAlpha( r_sym , r_benchmark):
beta = GetBeta( r_sym, r_benchmark)
return r_sym - beta * r_benchmark
GetBeta( df['r(FB)'], df['r(QQQ)'])
alpha_fb = GetAlpha( df['r(FB)'], df['r(QQQ)'])
alpha_fb.sort_index(ascending= False).head()
```
### Visualizing **Alpha**, **${Rm}$**, **${R}$**
```
cols = ['alpha','r(FB)', 'r(QQQ)']
l_series = [alpha_fb, df['r(FB)'], df['r(QQQ)']]
plot_df = pd.concat( l_series, axis = 1)
plot_df.columns = cols
#plot_df = pd.DataFrame(data = l_series, columns = cols, index = alpha_fb.index)
plot_df.plot(figsize = (12,5))
plotly_data = [go.Scatter(x = plot_df.index, y = plot_df['alpha'], name = 'alpha'),
go.Scatter(x = plot_df.index, y = plot_df['r(FB)'], name = 'r(FB)'),
go.Scatter(x = plot_df.index, y = plot_df['r(QQQ)'],name = 'r(QQQ)')
]
iplot(plotly_data)
```
## Getting Sentiments
```
t_nflx = pd.read_csv('dataset/twitter/NFLX.csv')
t_fb = pd.read_csv('dataset/twitter/FB.csv')
t_amzn = pd.read_csv('dataset/twitter/AMZN.csv')
t_goog = pd.read_csv('dataset/twitter/GOOG.csv')
def data_processing(data, stockcode):
data = data[['unix_timestamp','content']]
data = data.dropna()
data['stockcode'] = data['content'].copy()
data['stockcode'] = data['stockcode'].apply(lambda x: stockcode)
data['vader'] = data['content'].copy()
data['vader'] = data['vader'].apply(lambda x: str(x).lower())
data['vader'] = data['vader'].apply(lambda x: unidecode(re.sub('[^a-zA-z0-9\s]','',x)))
data['textblob'] = data['vader'].copy()
# Generating Vader Sentiment
analyzer = SentimentIntensityAnalyzer()
data['vader'] = data['vader'].apply(analyzer.polarity_scores)
data['vader'] = data['vader'].apply(lambda x: x['compound'])
# Generating TextBlob Sentiment
data['textblob'] = data['textblob'].apply(TextBlob)
data['textblob'] = data['textblob'].apply(lambda x: x.sentiment[0])
data['unix_timestamp'] = data['unix_timestamp'].apply(lambda x: str(x)[0:10])
data['date'] = data['unix_timestamp'].apply(lambda x: dt.fromtimestamp(int(x)).strftime('%Y-%m-%d'))
data['unix_timestamp'] = data['unix_timestamp'].apply(lambda x: dt.fromtimestamp(int(x)).strftime('%Y-%m-%d %H:%M:%S'))
data['unix_timestamp'] = pd.to_datetime(data['unix_timestamp'])
data['date'] = pd.to_datetime(data['date'])
data.index = data['date']
del data['date']
return data.sort_index()
def daily_sentiment_calculator(data):
data = data[['vader','textblob']].reset_index()
data = data[['vader','textblob']].groupby(data['date']).sum()
return data
test = data_processing( t_nflx, 'NFLX')
test_sent_daily = daily_sentiment_calculator(test)
test_sent_daily.plot(figsize = (12,5))
plotly_data = [go.Scatter(x = test_sent_daily.index, y = test_sent_daily['vader'], name = 'vader'),
go.Scatter(x = test_sent_daily.index, y = test_sent_daily['textblob'], name = 'textblob')
]
iplot(plotly_data)
fb_data = data_processing( t_fb, 'FB')
fb_sent = daily_sentiment_calculator(fb_data)
l_series = [
alpha_fb,
fb_sent['vader'],
fb_sent['textblob']
]
compare_df = pd.concat(l_series, axis =1)
compare_df.columns = ['alpha', 'sent_vader', 'sent_textblob']
compare_df.tail(10)
compare_df.dropna()[['alpha','sent_vader']].plot()
scatterdf = compare_df.dropna()[['alpha','sent_textblob']]
plotly_data = [go.Scatter(x = scatterdf.alpha, y = scatterdf.sent_textblob, mode = 'markers')
]
layout = go.Layout(
title = 'excess return vs sentiments',
xaxis = {'title': 'excess return'},
yaxis = {'title': 'sentiment'}
)
fig = go.Figure( data = plotly_data, layout= layout)
iplot(fig)
```
| github_jupyter |

# Tutorial 1 - Weather Data: Accesing it, understanding it, visualizing it!
This notebook explores a standard type of weather data, the typical meteorological year (TMY), and how to summarize it with Python and [Pandas](https://pandas.pydata.org/).

## Steps:
- [Weather data in PV performance models](#Weather-Data-&-PV)
- Looking at a sample weather data file
- Where to get weather data from?
- Weather data to API
## PV Concepts:
- TMY
- GHI, DNI, DHI
- DryBulb, Wspd
- Irradiance vs. Insolation
## Python Concepts:
- Exploring a Pandas dataframe (`df`): `len()`, `df.head()`, `df.keys()`
- [pvlib input-output tools](https://pvlib-python.readthedocs.io/en/stable/api.html#io-tools)
- Ploting a Pandas dataframe (`df`): `df.plot()`
- Aggregating data in a dataframe (`df`): [`df.resample(freq).sum()`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.resample.html)
- Pandas [`DateOffsets`](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#dateoffset-objects) - shortcuts to set the frequency when resampling
- Getting NREL irradiance data from the web based API using pvlib
## Weather Data & PV
Weather and irradiance data are used as input to PV performance models.

These data are directly measured, derived from measured data, or simulated using a stochastic model.
If you want to learn more about irradiance data, you can visit the [Solar RAdiometry webpage](https://www.pmodwrc.ch/en/world-radiation-center-2/srs/)
## Typical Meteorological Year
TMY datasets are intended to represent the weather for a typical year at a given location.
TMY datasets provide hourly **solar irradiance**, **air temperature**, **wind speed**, and other weather measurements for a hypothetical year that represents more or less a "median year" for solar resource.

TMY datasets are created by selecting individual months out of an extended period of weather measurememts (say, 20 years of data) to construct a single year's worth of data. There are several methods for selecting which months to include, but the general idea is to calculate monthly summary statistics and take the month that lies in the middle of the distribution. For example, no two Januaries will be exactly the same, so summing the total solar irradiance for each January will give a [normal distribution](https://en.wikipedia.org/wiki/Normal_distribution), and the month that falls closest to the [median](https://en.wikipedia.org/wiki/Median) is chosen as the representative month. The same process is followed for February, March, and so on, and all twelve representative months are stitched together into a year-long dataset.
The oldest TMYs were calculated using data from the nearest weather station (airports and such). Today, it's common to use TMYs calculated using simulated weather data from satellite imagery because of the improved spatial resolution.
To get a better feel for TMY data, we'll first explore an example TMY dataset that is bundled with pvlib.
## First Step: Import Libraries
In Python, some functions are builtin like `print()` but others must be imported before they can be used. For this notebook we're going to import three packages:
* [pvlib](https://pvlib-python.readthedocs.io/en/stable/) - library for simulating performance of photovoltaic energy systems.
* [pandas](https://pandas.pydata.org/) - analysis tool for timeseries and tabular data
* [matplotlib](https://matplotlib.org/) - data visualization for Python
Some Python modules are part of the [standard library](https://docs.python.org/3/library/index.html), but are not imported with builtins. We'll use the `pathlib` module which is useful for accessing files and folders.
```
import os # for getting environment variables
import pathlib # for finding the example dataset
import pvlib
import pandas as pd # for data wrangling
import matplotlib.pyplot as plt # for visualization
```
Query which version you are using of pvlib:
```
print(pvlib.__version__)
```
## Reading a TMY dataset with pvlib
First, we'll read the TMY dataset with [`pvlib.iotools.read_tmy3()`](https://pvlib-python.readthedocs.io/en/stable/generated/pvlib.iotools.read_tmy3.html) which returns a [Pandas DataFrame](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.html) of the timeseries weather data and a second output with a Python dictionary of the TMY metadata like longitude, latitude, elevation, etc.
We will use the Python [`pathlib`](https://docs.python.org/3/library/pathlib.html) to get the path to the `'data'` directory which comes with the `pvlib` package. Then we can use the slash operator, `/` to make the full path to the TMY file.
```
help(pvlib.iotools.read_tmy3)
DATA_DIR = pathlib.Path(pvlib.__file__).parent / 'data'
df_tmy, meta_dict = pvlib.iotools.read_tmy3(DATA_DIR / '723170TYA.CSV', coerce_year=1990)
meta_dict # display the dictionary of metadata
```
Let's display the first 4 lines of the dataframe
```
df_tmy.head(4)
```
This dataset follows the standard format of handling timeseries data with pandas -- one row per timestamp, one column per measurement type. Because TMY files represent one year of data (no leap years), that means they'll have 8760 rows. The number of columns can vary depending on the source of the data.
```
print("Number of rows:", len(df_tmy))
print("Number of columns:", len(df_tmy.columns))
```
You can access single rows by pointing to its number location (iloc) or by using the index name it has. In this case, that is a dateTime
```
df_tmy.iloc[0];
df_tmy.loc['1990-01-01 01:00:00-05:00'];
```
You can also print all the column names in the dataframe
```
df_tmy.keys()
```
There are 71 columns, which is quite a lot! For now, let's focus just on the ones that are most important for PV modeling -- the irradiance, temperature, and wind speed columns, and extract them into a new DataFrame.
## Irradiance
Irradiance is an instantaneous measurement of solar power over some area. For practical purposes of measurement and interpretation, irradiance is expressed and separated into different components.

The units of irradiance are watts per square meter.
## Wind
Wind speed is measured with an anemometer. The most common type is a the cup-type anemometer, shown on the right side of the picture below. The number of rotations per time interval is used to calculate the wind speed. The vane on the left is used to measure the direction of the wind. Wind direction is reported as the direction from which the wind is blowing.
<img src="https://pvpmc.sandia.gov/wp-content/uploads/2012/04/anemometer.jpg"></img>
## Air temperature
Also known as dry-bulb temperature, is the temperature of the ambient air when the measurement device is shielded from radiation and moisture. The most common method of air temperature measurement uses a resistive temperature device (RTD) or thermocouple within a radiation shield. The shield blocks sunlight from reaching the sensor (avoiding radiative heating), yet allows natural air flow around the sensor. More accurate temperature measurement devices utilize a shield which forces air across the sensor.
Air temperature is typically measured on the Celsius scale.
Air temperature plays a large role in PV system performance as PV modules and inverters are cooled convectively by the surrounding air.
<img src="https://pvpmc.sandia.gov/wp-content/uploads/2012/04/AmbTemp.jpg" width="400" height="400"> </img>
## Downselect columns
There are a lot more weather data in that file that you can access. To investigate all the column headers, we used `.keys()` above. Always read the [Instruction Manual](https://www.nrel.gov/docs/fy08osti/43156.pdf) for the weather files to get more details on how the data is aggregated, units, etc.
At this point we are interested in <b> GHI, DHI, DNI, DryBulb </b> and <b> Wind Speed </b>. For this NREL TMY3 dataset the units of irradiance are W/m², dry bulb temperature is in °C, and wind speed is m/s.
```
# GHI, DHI, DNI are irradiance measurements
# DryBulb is the "dry-bulb" (ambient) temperature
# Wspd is wind speed
df = df_tmy[['GHI', 'DHI', 'DNI', 'DryBulb', 'Wspd']]
# show the first 15 rows:
df.head(15)
```
## Plotting time series data with pandas and matplotlib
Let's make some plots to get a better idea of what TMY data gives us.
### Irradiance
First, the three irradiance fields:
```
first_week = df.head(24*7) # Plotting 7 days, each one has 24 hours or entries
first_week[['GHI', 'DHI', 'DNI']].plot()
plt.ylabel('Irradiance [W/m$^2$]');
```
Let's control the parameters a bit more
```
birthday = df.loc['1990-11-06':'1990-11-06']
plt.plot(birthday['DNI'], color='r')
plt.plot(birthday['DHI'], color='g', marker='.')
plt.plot(birthday['GHI'], color='b', marker='s')
plt.ylabel('Irradiance [W/m$^2$]');
```
#### Exercise
How does the Irradiance look like in YOUR birthday?
Hint: the next cell is 'Markdown', you need to switch it to 'Code' for it to run
birthday = df.loc[] # Type your birthday here using the 1990 year, i.e. '1990-11-06':'1990-11-06'
plt.plot(birthday['DNI'], color='g')
plt.plot(birthday['DHI'], color='b', marker='.')
plt.plot(birthday['GHI'], color='r', marker='s')
plt.ylabel('Irradiance [W/m$^2$]');
GHI, DHI, and DNI are the three "basic" ways of measuring irradiance, although each of them is measured in units of power per area (watts per square meter):
- GHI: Global Horizontal Irradiance; the total sunlight intensity falling on a horizontal plane
- DHI: Diffuse Horizontal Irradiance; the subset of sunlight falling on a horizontal plane that isn't coming directly from the sun (e.g., the light that makes the sky blue)
- DNI: Direct Normal Irradiance; the subset of sunlight coming directly from the sun



Later tutorials will show how these three values are used in PV modeling. For now, let's just get a qualitative understanding of the differences between them: looking at the above plot, there is a pattern where when DNI is high, DHI is low. The sun puts out a (roughly) constant amount of energy, which means photons either make it through the atmosphere without scattering and are counted as direct irradiance, or they tend to get scattered and become part of the diffuse irradiance, but not both. Looking at DNI makes it easy to pick out which hours are cloud and which are sunny -- most days in January are rather overcast with low irradiance, but the sun does occasionally break through.
In addition to daily variation, there is also seasonal variation in irradiance. Let's compare a winter week with a summer week, using pandas to select out a summertime subset. Notice the increased intensity in summer -- GHI peaks around 900 W/m^2, whereas in winter the maximum was more like 500 W/m^2.
```
summer_week = df.loc['1990-06-01':'1990-06-08']
summer_week[['GHI', 'DHI', 'DNI']].plot()
plt.ylabel('Irradiance [W/m$^2$]');
```
### Temperature
Next up is temperature:
```
first_week['DryBulb'].plot()
plt.ylabel('Ambient Temperature [°C]');
```
### Wind speed
And finally, wind speed:
```
first_week['Wspd'].plot()
plt.ylabel('Wind Speed [m/s]');
```
## Aggregating hourly data to monthly summaries
Pandas makes it easy to roll-up timeseries data into summary values. We can use the [`DataFrame.resample()`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.resample.html) function with [`DateOffsets`](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#dateoffset-objects) like `'M'` for months. For example, we can calculate total monthly GHI as a quick way to visualize the seasonality of solar resource:
```
# summing hourly irradiance (W/m^2) gives insolation (W h/m^2)
monthly_ghi = df['GHI'].resample('M').sum()
monthly_ghi.head(4)
monthly_ghi = monthly_ghi.tz_localize(None) # don't need timezone for monthly data
monthly_ghi.plot.bar()
plt.ylabel('Monthly Global Horizontal Irradiance\n[W h/m$^2$]');
```
We can also take monthly averages instead of monthly sums:
```
fig, ax1 = plt.subplots()
ax2 = ax1.twinx() # add a second y-axis
monthly_average_temp_wind = df[['DryBulb', 'Wspd']].resample('M').mean()
monthly_average_temp_wind['DryBulb'].plot(ax=ax1, c='tab:blue')
monthly_average_temp_wind['Wspd'].plot(ax=ax2, c='tab:orange')
ax1.set_ylabel(r'Ambient Temperature [$\degree$ C]')
ax2.set_ylabel(r'Wind Speed [m/s]')
ax1.legend(loc='lower left')
ax2.legend(loc='lower right');
```
### Exercise
Plot the Average DNI per Day
```
try:
daily_average_DNI = df[['']].resample('').mean() # Add the column name, and resample by day. Month is 'M', day is..
daily_average_DNI.plot()
except:
print("You haven't finished this exercise correctly, try again!")
```
## Where to get _Free_ Solar Irradiance Data?
There are many different sources of solar irradiance data. For your projects, these are some of the most common:
- [TMY3/TMY2](https://nsrdb.nrel.gov/data-sets/archives.html) - like the data we loaded earlier on this example. This database of TMY3 `.csv` files and TMY2 files have been archived by NREL.
- [NSRDB](https://maps.nrel.gov/nsrdb-viewer/) - National Solar Radiation Database. You can access data through the website for many locations accross the world, or you can use their [web API](https://developer.nrel.gov/docs/solar/nsrdb/) to download data programmatically. An "API" is an ["application programming interface"](https://en.wikipedia.org/wiki/API), and a "web API" is a programming interface that allows you to write code to interact with web services like the NSRDB.
- [EPW](https://www.energy.gov/eere/buildings/downloads/energyplus-0) - Energy Plus Weather data is available for many locations accross the world. It's in its own format file ('EPW') so you can't open it easily in a spreadsheet program like Excel, but you can use [`pvlib.iotools.read_epw()`](https://pvlib-python.readthedocs.io/en/stable/generated/pvlib.iotools.read_epw.html) to get it into a dataframe and use it.
- [PVGIS](https://ec.europa.eu/jrc/en/pvgis) - Free global weather data provided by the European Union and derived from many govermental agencies including the NSRDB. PVGIS also provides a web API. You can get PVGIS TMY data using [`pvlib.iotools.get_pvgis_tmy()`](https://pvlib-python.readthedocs.io/en/stable/generated/pvlib.iotools.get_pvgis_tmy.html).
- Perhaps another useful link: https://sam.nrel.gov/weather-data.html
## Where else can you get historical irradiance data?
There are several commercial providers of solar irradiance data. Data is available at different spatial and time resolutions. Each provider offers data under subscription that will provide access to irradiance (and other weather variables) via API to leverage in python.
* [SolarAnywhere](https://www.solaranywhere.com/)
* [SolarGIS](https://solargis.com/)
* [Vaisala](https://www.vaisala.com/en)
* [Meteonorm](https://meteonorm.com/en/)
* [DNV Solar Resource Compass](https://src.dnvgl.com/)

## NREL API Key
At the [NREL Developer Network](https://developer.nrel.gov/), there are [APIs](https://en.wikipedia.org/wiki/API) to a lot of valuable [solar resources](https://developer.nrel.gov/docs/solar/) like [weather data from the NSRDB](https://developer.nrel.gov/docs/solar/nsrdb/), [operational data from PVDAQ](https://developer.nrel.gov/docs/solar/pvdaq-v3/), or indicative calculations using [PVWatts](https://developer.nrel.gov/docs/solar/pvwatts/). In order to use these resources from NREL, you need to [register for a free API key](https://developer.nrel.gov/signup/). You can test out the APIs using the `DEMO_KEY` but it has limited bandwidth compared to the [usage limit for registered users](https://developer.nrel.gov/docs/rate-limits/). NREL has some [API usage instructions](https://developer.nrel.gov/docs/api-key/), but pvlib has a few builtin functions, like [`pvlib.iotools.get_psm3()`](https://pvlib-python.readthedocs.io/en/stable/generated/pvlib.iotools.get_psm3.html), that wrap the NREL API, and call them for you to make it much easier to use. Skip ahead to the next section to learn more. But before you do...
**Please pause now to visit https://developer.nrel.gov/signup/ and get an API key.**
### Application Programming Interface (API)
What exactly is an API? Nowadays, the phrase is used interchangeably with a "web API" but in general an API is just a recipe for how to interface with a application programmatically, _IE_: in code. An API could be as simple as a function signature or its published documentation, _EG_: the API for the `solarposition` function is you give it an ISO8601 formatted date with a timezone, the latitude, longitude, and elevation as numbers, and it returns the zenith and azimuth as numbers.
A web API is the same, except the application is a web service, that you access at its URL using web methods. We won't go into too much more detail here, but the most common web method is `GET` which is pretty self explanatory. Look over the [NREL web usage instructions](https://developer.nrel.gov/docs/api-key/) for some examples, but interacting with a web API can be as easy as entering a URL into a browser. Try the URL below to _get_ the PVWatts energy output for a fixed tilt site in [Broomfield, CO](https://goo.gl/maps/awkEcNGzSur9Has18).
https://developer.nrel.gov/api/pvwatts/v6.json?api_key=DEMO_KEY&lat=40&lon=-105&system_capacity=4&azimuth=180&tilt=40&array_type=1&module_type=1&losses=10
In addition to just using your browser, you can also access web APIs programmatically. The most popular Python package to interact with web APIs is [requests](https://docs.python-requests.org/en/master/). There's also free open source command-line tools like [cURL](https://curl.se/) and [HTTPie](https://httpie.io/), and a popular nagware/freemium GUI application called [Postman](https://www.postman.com/).
**If you have an NREL API key please enter it in the next cell.**
```
NREL_API_KEY = None # <-- please set your NREL API key here
# note you must use "quotes" around your key, for example:
# NREL_API_KEY = 'DEMO_KEY' # single or double both work fine
# during the live tutorial, we've stored a dedicated key on our server
if NREL_API_KEY is None:
try:
NREL_API_KEY = os.environ['NREL_API_KEY'] # get dedicated key for tutorial from servier
except KeyError:
NREL_API_KEY = 'DEMO_KEY' # OK for this demo, but better to get your own key
```
## Fetching TMYs from the NSRDB
The example TMY dataset used here is from an airport in North Carolina, but what if we wanted to model a PV system somewhere else? The NSRDB, one of many sources of weather data intended for PV modeling, is free and easy to access using pvlib. As an example, we'll fetch a TMY dataset for Albuquerque, New Mexico at coordinates [(35.0844, -106.6504)](https://www.google.com/maps/@35.0844,-106.6504,9z). We use [`pvlib.iotools.get_psm3()`](https://pvlib-python.readthedocs.io/en/stable/generated/pvlib.iotools.get_psm3.html) which returns a Python dictionary of metadata and a Pandas dataframe of the timeseries weather data.
```
metadata, df_abq = pvlib.iotools.get_psm3(
latitude=35.0844, longitude=-106.6504,
api_key=NREL_API_KEY,
email='mark.mikofski@dnv.com', # <-- any email works here fine
names='tmy')
metadata
```
TMY datasets from the PSM3 service of the NSRDB are timestamped using the real year that the measurements came from. The [`pvlib.iotools.read_tmy3()`](https://pvlib-python.readthedocs.io/en/stable/generated/pvlib.iotools.read_tmy3.html) function had a `coerce_year` argument to force everything to align to a single dummy year, but `pvlib.iotools.get_psm3()` doesn't have that feature. For convenience let's standardize the data to 1990 and then compare monthly GHI to the North Carolina location:
```
df_abq['Year'] = 1990
df_abq.index = pd.to_datetime(df_abq[['Year', 'Month', 'Day', 'Hour']])
ghi_comparison = pd.DataFrame({
'NC': monthly_ghi, # using the monthly values from earlier
'NM': df_abq['GHI'].resample('M').sum(),
})
ghi_comparison.plot.bar()
plt.ylabel('Monthly GHI [W h/m^2]');
```
It's not too surprising to see that our New Mexico location is significantly sunnier than the one in North Carolina.
[](http://creativecommons.org/licenses/by/4.0/)
This work is licensed under a [Creative Commons Attribution 4.0 International License](http://creativecommons.org/licenses/by/4.0/).
| github_jupyter |
# ORENIST Dynamic Filter Classification
Note: This notebook is desinged to run with Python3 and CPU (no GPU) runtime.
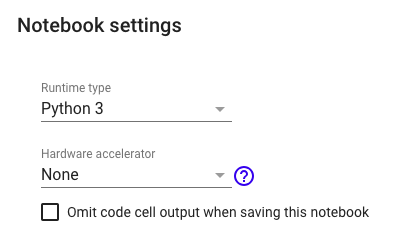
This notebook uses TensorFlow2.x.
```
%tensorflow_version 2.x
```
####[ODF-01]
Import modules and set a random seed.
```
import numpy as np
import matplotlib.pyplot as plt
from pandas import DataFrame
import pickle
import tensorflow as tf
from tensorflow.keras import layers, models, initializers
np.random.seed(20190225)
tf.random.set_seed(20190225)
```
####[ODF-02]
Download the ORENIST dataset and store into NumPy arrays.
```
!curl -LO https://github.com/enakai00/colab_tfbook/raw/master/Chapter04/ORENIST.data
with open('ORENIST.data', 'rb') as file:
images, labels = pickle.load(file)
```
####[ODF-03]
Define a model to classify the ORENIST dataset with the convolutional filters.
The ouputs from filters are converted to a single pixel image with the pooling layer.
```
model = models.Sequential()
model.add(layers.Reshape((28, 28, 1), input_shape=(28*28,), name='reshape'))
model.add(layers.Conv2D(2, (5, 5), padding='same',
kernel_initializer=initializers.TruncatedNormal(),
use_bias=False, name='conv_filter'))
model.add(layers.Lambda(lambda x: abs(x), name='abs'))
model.add(layers.MaxPooling2D((28, 28), name='max_pooling'))
model.add(layers.Flatten(name='flatten'))
model.add(layers.Dense(3, activation='softmax', name='softmax'))
model.summary()
```
####[ODF-04]
Compile the model using the Adam optimizer, and Cross entroy as a loss function.
```
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['acc'])
```
####[ODF-05]
Train the model.
```
history = model.fit(images, labels,
batch_size=len(images), epochs=2000, verbose=0)
```
####[ODF-06]
Plot charts to see the accuracy and loss values. It achieves the 100% accuracy.
```
DataFrame({'acc': history.history['acc']}).plot()
DataFrame({'loss': history.history['loss']}).plot()
```
####[ODF-07]
Define a model to extract outputs from intermediate layers.
```
layer_outputs = [model.get_layer('abs').output,
model.get_layer('max_pooling').output]
model2 = models.Model(inputs=model.input, outputs=layer_outputs)
```
####[ODF-08]
Apply the trained filters to the ORENST dataset.
```
conv_output, pool_output = model2.predict(images[:9])
filter_vals = model.get_layer('conv_filter').get_weights()[0]
```
####[ODF-09]
Show images after applying the convolutional filters.
```
fig = plt.figure(figsize=(10, 3))
v_max = np.max(conv_output)
for i in range(2):
subplot = fig.add_subplot(3, 10, 10*(i+1)+1)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.imshow(filter_vals[:, :, 0, i], cmap=plt.cm.gray_r)
for i in range(9):
subplot = fig.add_subplot(3, 10, i+2)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.set_title('%d' % np.argmax(labels[i]))
subplot.imshow(images[i].reshape((28, 28)),
vmin=0, vmax=1, cmap=plt.cm.gray_r)
subplot = fig.add_subplot(3, 10, 10+i+2)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.imshow(conv_output[i,:,:,0],
vmin=0, vmax=v_max, cmap=plt.cm.gray_r)
subplot = fig.add_subplot(3, 10, 20+i+2)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.imshow(conv_output[i,:,:,1],
vmin=0, vmax=v_max, cmap=plt.cm.gray_r)
```
####[ODF-10]
Show images after applying the pooling layer.
```
fig = plt.figure(figsize=(10, 3))
v_max = np.max(pool_output)
for i in range(2):
subplot = fig.add_subplot(3, 10, 10*(i+1)+1)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.imshow(filter_vals[:, :, 0, i], cmap=plt.cm.gray_r)
for i in range(9):
subplot = fig.add_subplot(3, 10, i+2)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.set_title('%d' % np.argmax(labels[i]))
subplot.imshow(images[i].reshape((28, 28)),
vmin=0, vmax=1, cmap=plt.cm.gray_r)
subplot = fig.add_subplot(3, 10, 10+i+2)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.imshow(pool_output[i, :, :, 0],
vmin=0, vmax=v_max, cmap=plt.cm.gray_r)
subplot = fig.add_subplot(3, 10, 20+i+2)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.imshow(pool_output[i, :, :, 1],
vmin=0, vmax=v_max, cmap=plt.cm.gray_r)
```
####[ODF-11]
Convert outputs from the pooling layer into binary values with a threshold 7.0.
```
fig = plt.figure(figsize=(10, 3))
bin_index = np.sign(pool_output-7.0)
for i in range(2):
subplot = fig.add_subplot(3, 10, 10*(i+1)+1)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.imshow(filter_vals[:, :, 0, i], cmap=plt.cm.gray_r)
for i in range(9):
subplot = fig.add_subplot(3, 10, i+2)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.set_title('%d' % np.argmax(labels[i]))
subplot.imshow(images[i].reshape((28, 28)),
vmin=0, vmax=1, cmap=plt.cm.gray_r)
subplot = fig.add_subplot(3, 10, 10+i+2)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.imshow(bin_index[i, :, :, 0],
vmin=-1, vmax=1, cmap=plt.cm.gray_r)
subplot = fig.add_subplot(3, 10, 20+i+2)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.imshow(bin_index[i, :, :, 1],
vmin=-1, vmax=1, cmap=plt.cm.gray_r)
```
| github_jupyter |
<img src="header.png" align="left"/>
# Exercise: Classification of MNIST (10 points)
Die goal of this exercise is to create a simple image classification network and to work on the improvement of a model and how to debug and check the training data. We start with a simple CNN model for digit classification of the MNIST dataset [1]. This dataset contains 60,000 scans of digits for training and 10,000 scans of digits for validation. A sample consists of 28x28 features with values between 0 and 255, note that the features are inverted. Actually digits are rather dark on a light background. MNIST digits are light on a dark background.
This example is partly based on a tutorial by Jason Brownlee [2].
Please follow the instructions in the notebook.
```
[1] http://yann.lecun.com/exdb/mnist/
[2] https://machinelearningmastery.com/how-to-develop-a-convolutional-neural-network-from-scratch-for-mnist-handwritten-digit-classification/
```
**NOTE**
Document your results by simply adding a markdown cell or a python cell (as comment) and writing your statements into this cell. For some tasks the result cell is already available.
[](https://colab.research.google.com/github/ditomax/mlexercises/blob/master/04%20Exercise%20Classification%20MNIST.ipynb)
```
#
# Prepare colab
#
COLAB=False
try:
%tensorflow_version 2.x
print("running on google colab")
COLAB=True
except:
print("not running on google colab")
#
# Turn off errors and warnings (does not work sometimes)
#
from warnings import simplefilter
# ignore all future warnings
simplefilter(action='ignore', category=FutureWarning)
simplefilter(action='ignore', category=Warning)
#
# Import some modules
#
import tensorflow as tf
from tensorflow.keras.datasets import mnist
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Conv2D
from tensorflow.keras.layers import MaxPooling2D
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Flatten
from tensorflow.keras.layers import BatchNormalization
from tensorflow.keras.layers import Dropout
from tensorflow.keras.optimizers import SGD
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.models import model_from_json
from tensorflow.keras.callbacks import ModelCheckpoint
from tensorflow.keras.callbacks import EarlyStopping
from sklearn.metrics import confusion_matrix
import os
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
#
# Diagram size
#
plt.rcParams['figure.figsize'] = [16, 9]
#
# nasty hack for macos
#
os.environ['KMP_DUPLICATE_LIB_OK']='True'
#
# check version of tensorflow
#
print('starting notebook with tensorflow version {}'.format(tf.version.VERSION))
```
# Load and prepare data
```
#
# Loading of the data (very simplified) with split into train and test data (fixed split)
#
(x_train, y_train), (x_test, y_test) = mnist.load_data()
#
# Check some data rows
#
x_train[0][10]
#
# Print shapes of data
#
print('training data: X=%s, y=%s' % (x_train.shape, y_train.shape))
print('test data: X=%s, y=%s' % (x_test.shape, y_test.shape))
#
# Display some examples of the data
#
for i in range(15):
plt.subplot(4,4,1 + i)
plt.imshow(x_train[i], cmap=plt.get_cmap('gray'))
plt.show()
#
# Display labels of some data
#
for i in range(15):
print('label {}'.format(y_train[i]))
```
<div class="alert alert-block alert-info">
## Task
Plot a histogram of the classes of the training data (1 point).
After plotting, give a short estimation if this distribution is OK for use in a classification situation.
</div>
```
#
# Histogram of class counts (digits)
#
# Task: plot the histogram as array or as plot
#
```
# Prepare data for classification
<div class="alert alert-block alert-info">
## Task
Find out why the unusual shap of the input data is required? Why is (-1,28,28) not sufficien? (1 point)
Give a short description here in the comment.
Hint: check the tensorflow keras documentation about 2D cnn layer.
</div>
```
#
# Change shape of data for model
#
x_train = x_train.reshape((x_train.shape[0], 28, 28, 1))
x_test = x_test.reshape((x_test.shape[0], 28, 28, 1))
#
# your anser here
#
#
# Scale pixel values into range of 0 to 1
#
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train = x_train / 255.0
x_test = x_test / 255.0
# check one transformed sample row
x_train[0][10]
#
# One-hot encoding for classes
#
y_train = to_categorical(y_train, num_classes=10)
y_test = to_categorical(y_test, num_classes=10)
# check the one-hot encoding
y_train
```
# Build the first model
<div class="alert alert-block alert-info">
## Task
Complete the code for a simple convolutional neural network (CNN) with one CNN layer (2 Points).
Hint: look for examples in the internet or in the slides.
</div>
```
model = Sequential()
...
model.add(Dense(10, activation='softmax'))
# compile model
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
# get a short summary of the model
model.summary()
# train model
history = model.fit(x_train, y_train, batch_size=128, epochs=5 )
```
# First prediction with model
<div class="alert alert-block alert-info">
## Task
Describe the meaning of the numbers returned from the prediction. (1 point)
Write your findings here in the comments
Hint: look at the definition of the output layer (last layer) in the model.
</div>
```
model.predict(x_train[:1])
# compare with expexted result
y_train[:1]
#
# Measure the accuracy
#
_, acc = model.evaluate(x_test, y_test, verbose=0)
print('accuracy {:.5f}'.format(acc))
#
# Estimate the number of false classifications in production use
#
print('with {} samples there are about {:.0f} false classifications to expect.'.format( x_test.shape[0], (x_test.shape[0]*(1-acc))))
```
# Print out training progress
```
#
# Plot loss and accuracy
#
def summarize_diagnostics(history,modelname):
plt.subplot(211)
plt.title('Cross Entropy Loss')
plt.plot(history.history['loss'], color='blue', label='train')
plt.subplot(212)
plt.title('Classification Accuracy')
plt.plot(history.history['accuracy'], color='green', label='train')
plt.subplots_adjust(hspace=0.5)
plt.savefig( 'results/' + modelname + '_plot.png')
plt.show()
plt.close()
summarize_diagnostics(history,'03_model1')
```
# Improve the model significantly
<div class="alert alert-block alert-info">
## Task
Your customer requires to have less than 1% of wrong classifications. Start to build a better
model with significantly less than 100 wrong classifications in the 10000 test samples.
Research the internet for the optimal model setup for MNIST classification and try to replicate this model here.
Make sure to document the source where you found the hints for the improvement (links to sources) (2 Points).
</div>
```
#
# Setup new model
#
def create_model_2():
model = Sequential()
...
model.add(Dense(10, activation='softmax'))
return model
#
# instantiate model
#
model2 = create_model_2()
#
# compile
#
model2.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
#
# train with history
#
history = model2.fit(x_train, y_train, batch_size=128, epochs=15 )
#
# Measure the accuracy
#
_, acc = model2.evaluate(x_test, y_test, verbose=0)
print('Accuracy {:.5f}'.format(acc))
#
# Estimate the number of false classifications in production use
#
print('with {} samples there are about {:.0f} false classifications to expect.'.format( x_test.shape[0], (x_test.shape[0]*(1-acc))))
# Result: (describe where you found the hints for improvement and how much it improved)
model2.summary()
summarize_diagnostics(history,'03_model2')
```
# Save the model
```
#
# Save a model for later use
#
prefix = 'results/03_'
modelName = prefix + "model.json"
weightName = prefix + "model.h5"
# set to True if the model should be saved
save_model = True
if save_model:
model_json = model2.to_json()
with open( modelName , "w") as json_file:
json_file.write(model_json)
# serialize weights to HDF5
model2.save_weights( weightName )
print("saved model to disk as {} {}".format(modelName,weightName))
else:
# load model (has to be saved before, model is not part of git)
json_file = open(modelName, 'r')
loaded_model_json = json_file.read()
json_file.close()
model2 = model_from_json(loaded_model_json)
# load weights into new model
model2.load_weights(weightName)
print("loaded model from disk")
```
# Find characteristics in the errors of the model
<div class="alert alert-block alert-info">
## Task
There are still too many false classifications using the model.
Evaluate all test data and plot examples of failed classifications to get a better undestanding what goes wring.
Plot a confusion matrix to get a better insight. (1 Point).
</div>
```
y_test_predictions = model2.predict(x_test)
#
# generate confusion matrix
# Task: find a suitable function for generating a confusion matrix as array
#
confusion = ...
print(confusion)
# make a nice plot of the confusion matrix
import itertools
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
#print(cm)
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j],
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# plot confusion matrix
plot_confusion_matrix(confusion,['0','1','2','3','4','5','6','7','8','9'] )
```
# Improve the training
Beside many other options, there are two streight forward ways to improve your model:
1. Add more data for those classes which are poorely classified
1. Add augmentation for the training data
Implement the augmentation strategy and test if there is an improvement.
## Augmentation
<div class="alert alert-block alert-info">
## Task
Task: Search the internet for the ImageDataGenerator class of the Keras framework
and implement such a generator for the training of the model.
Select suitable augmentation which fits to the use-case.
Document the resulting accuracy. (2 Points)
</div>
```
# Augmentation solution
...
# instantiate model
model3 = create_model_2()
# Training
...
#
# Evaluierung
#
_, acc = model3.evaluate(x_test, y_test, verbose=0)
print('accuracy {:.3f} '.format(acc) )
summarize_diagnostics(history,'03_model3')
y_test_predictions = model3.predict(x_test)
# generate confusion matrix
confusion = confusion_matrix(np.argmax(y_test,axis=1), np.argmax(y_test_predictions,axis=1))
# plot confusion matrix
plot_confusion_matrix(confusion,['0','1','2','3','4','5','6','7','8','9'] )
```
| github_jupyter |
# Text Generation
```
import string
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Embedding, LSTM, Dense, Bidirectional
```
### Helping Functions
```
def create_lyrics_corpus(dataset, field):
# Remove all other punctuation
dataset[field] = dataset[field].str.replace('[{}]'.format(string.punctuation), '')
# Make it lowercase
dataset[field] = dataset[field].str.lower()
# Make it one long string to split by line
lyrics = dataset[field].str.cat()
corpus = lyrics.split('\n')
# Remove any trailing whitespace
for l in range(len(corpus)):
corpus[l] = corpus[l].rstrip()
# Remove any empty lines
corpus = [l for l in corpus if l != '']
return corpus
def tokenize_corpus(corpus, num_words=-1):
# Fit a Tokenizer on the corpus
if num_words > -1:
tokenizer = Tokenizer(num_words=num_words)
else:
tokenizer = Tokenizer()
tokenizer.fit_on_texts(corpus)
return tokenizer
```
## Step 1 : Get the Corpus
```
# Read the dataset from csv - just first 10 songs for now
path = tf.keras.utils.get_file('songdata.csv',
'https://drive.google.com/uc?id=1LiJFZd41ofrWoBtW-pMYsfz1w8Ny0Bj8')
print (path)
dataset = pd.read_csv(path, dtype=str)[:10]
corpus = create_lyrics_corpus(dataset, 'text')
```
## Step 2 : Tokenize the Corpus
```
# Tokenize the corpus
tokenizer = tokenize_corpus(corpus)
total_words = len(tokenizer.word_index) + 19 # why 19?
#print(tokenizer.word_index)
print(total_words)
dataset.head()
```
## Step 3 : Create n-Gram
```
sequences = []
for line in corpus:
token_list = tokenizer.texts_to_sequences([line])[0]
for i in range(1, len(token_list)):
n_gram_sequence = token_list[:i+1]
sequences.append(n_gram_sequence)
```
## Step 4 : Pad sequences
```
# Pad sequences for equal input length
max_sequence_len = max([len(seq) for seq in sequences])
sequences = np.array(pad_sequences(sequences, maxlen=max_sequence_len, padding='pre'))
```
## Step 5 : X and y - Values
```
# Split sequences between the "input" sequence and "output" predicted word
X = sequences[:,:-1]
y_label = sequences[:,-1]
# One-hot encode the labels
y = tf.keras.utils.to_categorical(y_label, num_classes = total_words)
```
### Explore and Trace
```
# Check out how some of our data is being stored
# The Tokenizer has just a single index per word
print(tokenizer.word_index['know'])
print(tokenizer.word_index['feeling'])
# Input sequences will have multiple indexes
print(X[5])
print(X[6])
# And the one hot labels will be as long as the full spread of tokenized words
print(y[5])
print(y[6])
```
## Step 6 : Create Model
```
model = Sequential()
model.add(Embedding(total_words, 64, input_length=max_sequence_len-1))
model.add(Bidirectional(LSTM(20)))
model.add(Dense(total_words, activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
model.summary()
history = model.fit(X, y, epochs=200, verbose=1)
def plot_graphs(history, string):
plt.plot(history.history[string])
plt.xlabel("Epochs")
plt.ylabel(string)
plt.show()
plot_graphs(history, 'accuracy')
```
## Step 7 : Generate Text
```
seed_text = "im feeling chills"
next_words = 100
for _ in range(next_words):
token_list = tokenizer.texts_to_sequences([seed_text])[0]
token_list = pad_sequences([token_list], maxlen=max_sequence_len-1, padding='pre')
predicted = np.argmax(model.predict(token_list), axis=-1)
output_word = ""
for word, index in tokenizer.word_index.items():
if index == predicted:
output_word = word
break
seed_text += " " + output_word
print(seed_text)
```
| github_jupyter |
PyGSLIB
========
PPplot
---------------
```
#general imports
import pygslib
```
Getting the data ready for work
---------
If the data is in GSLIB format you can use the function `pygslib.gslib.read_gslib_file(filename)` to import the data into a Pandas DataFrame.
```
#get the data in gslib format into a pandas Dataframe
mydata= pygslib.gslib.read_gslib_file('../datasets/cluster.dat')
true= pygslib.gslib.read_gslib_file('../datasets/true.dat')
true['Declustering Weight'] = 1
```
## gslib probplot with bokeh
```
parameters_probplt = {
# gslib parameters for histogram calculation
'iwt' : 0, # input boolean (Optional: set True). Use weight variable?
'va' : mydata['Primary'], # input rank-1 array('d') with bounds (nd). Variable
'wt' : mydata['Declustering Weight'], # input rank-1 array('d') with bounds (nd) (Optional, set to array of ones). Declustering weight.
# visual parameters for figure (if a new figure is created)
'figure' : None, # a bokeh figure object (Optional: new figure created if None). Set none or undefined if creating a new figure.
'title' : 'Prob blot', # string (Optional, "Histogram"). Figure title
'xlabel' : 'Primary', # string (Optional, default "Z"). X axis label
'ylabel' : 'P[Z<c]', # string (Optional, default "f(%)"). Y axis label
'xlog' : 1, # boolean (Optional, default True). If true plot X axis in log sale.
'ylog' : 1, # boolean (Optional, default True). If true plot Y axis in log sale.
# visual parameter for the probplt
'style' : 'cross', # string with valid bokeh chart type
'color' : 'blue', # string with valid CSS colour (https://www.w3schools.com/colors/colors_names.asp), or an RGB(A) hex value, or tuple of integers (r,g,b), or tuple of (r,g,b,a) (Optional, default "navy")
'legend': 'Non declustered', # string (Optional, default "NA").
'alpha' : 1, # float [0-1] (Optional, default 0.5). Transparency of the fill colour
'lwidth': 0, # float (Optional, default 1). Line width
# leyend
'legendloc': 'bottom_right'} # float (Optional, default 'top_right'). Any of top_left, top_center, top_right, center_right, bottom_right, bottom_center, bottom_left, center_left or center
parameters_probplt_dcl = parameters_probplt.copy()
parameters_probplt_dcl['iwt']=1
parameters_probplt_dcl['legend']='Declustered'
parameters_probplt_dcl['color'] = 'red'
parameters_probplt_true = parameters_probplt.copy()
parameters_probplt_true['va'] = true['Primary']
parameters_probplt_true['wt'] = true['Declustering Weight']
parameters_probplt_true['iwt']=0
parameters_probplt_true['legend']='True'
parameters_probplt_true['color'] = 'black'
parameters_probplt_true['style'] = 'line'
parameters_probplt_true['lwidth'] = 1
results, fig = pygslib.plothtml.probplt(parameters_probplt)
# add declustered to the plot
parameters_probplt_dcl['figure']= fig
results, fig = pygslib.plothtml.probplt(parameters_probplt_dcl)
# add true CDF to the plot
parameters_probplt_true['figure']=parameters_probplt_dcl['figure']
results, fig = pygslib.plothtml.probplt(parameters_probplt_true)
# show the plot
pygslib.plothtml.show(fig)
```
| github_jupyter |
# Running membership inference attacks on the Nursery data
In this tutorial we will show how to run black-box membership attacks. This will be demonstrated on the Nursery dataset (original dataset can be found here: https://archive.ics.uci.edu/ml/datasets/nursery).
We have already preprocessed the dataset such that all categorical features are one-hot encoded, and the data was scaled using sklearn's StandardScaler.
## Load data
```
import os
import sys
sys.path.insert(0, os.path.abspath('..'))
from art.utils import load_nursery
(x_train, y_train), (x_test, y_test), _, _ = load_nursery(test_set=0.5)
```
## Train random forest model
```
from sklearn.ensemble import RandomForestClassifier
from art.estimators.classification.scikitlearn import ScikitlearnRandomForestClassifier
model = RandomForestClassifier()
model.fit(x_train, y_train)
art_classifier = ScikitlearnRandomForestClassifier(model)
print('Base model accuracy: ', model.score(x_test, y_test))
```
## Attack
### Rule-based attack
The rule-based attack uses the simple rule to determine membership in the training data: if the model's prediction for a sample is correct, then it is a member. Otherwise, it is not a member.
```
import numpy as np
from art.attacks.inference.membership_inference import MembershipInferenceBlackBoxRuleBased
attack = MembershipInferenceBlackBoxRuleBased(art_classifier)
# infer attacked feature
inferred_train = attack.infer(x_train, y_train)
inferred_test = attack.infer(x_test, y_test)
# check accuracy
train_acc = np.sum(inferred_train) / len(inferred_train)
test_acc = 1 - (np.sum(inferred_test) / len(inferred_test))
acc = (train_acc * len(inferred_train) + test_acc * len(inferred_test)) / (len(inferred_train) + len(inferred_test))
print(train_acc)
print(test_acc)
print(acc)
```
This means that for 51% of the data, membership status is inferred correctly.
```
def calc_precision_recall(predicted, actual, positive_value=1):
score = 0 # both predicted and actual are positive
num_positive_predicted = 0 # predicted positive
num_positive_actual = 0 # actual positive
for i in range(len(predicted)):
if predicted[i] == positive_value:
num_positive_predicted += 1
if actual[i] == positive_value:
num_positive_actual += 1
if predicted[i] == actual[i]:
if predicted[i] == positive_value:
score += 1
if num_positive_predicted == 0:
precision = 1
else:
precision = score / num_positive_predicted # the fraction of predicted “Yes” responses that are correct
if num_positive_actual == 0:
recall = 1
else:
recall = score / num_positive_actual # the fraction of “Yes” responses that are predicted correctly
return precision, recall
# rule-based
print(calc_precision_recall(np.concatenate((inferred_train, inferred_test)),
np.concatenate((np.ones(len(inferred_train)), np.zeros(len(inferred_test))))))
```
### Black-box attack
The black-box attack basically trains an additional classifier (called the attack model) to predict the membership status of a sample. It can use as input to the learning process probabilities/logits or losses, depending on the type of model and provided configuration.
#### Train attack model
```
from art.attacks.inference.membership_inference import MembershipInferenceBlackBox
attack_train_ratio = 0.5
attack_train_size = int(len(x_train) * attack_train_ratio)
attack_test_size = int(len(x_test) * attack_train_ratio)
bb_attack = MembershipInferenceBlackBox(art_classifier)
# train attack model
bb_attack.fit(x_train[:attack_train_size], y_train[:attack_train_size],
x_test[:attack_test_size], y_test[:attack_test_size])
```
#### Infer sensitive feature and check accuracy
```
# get inferred values
inferred_train_bb = bb_attack.infer(x_train[attack_train_size:], y_train[attack_train_size:])
inferred_test_bb = bb_attack.infer(x_test[attack_test_size:], y_test[attack_test_size:])
# check accuracy
train_acc = np.sum(inferred_train_bb) / len(inferred_train_bb)
test_acc = 1 - (np.sum(inferred_test_bb) / len(inferred_test_bb))
acc = (train_acc * len(inferred_train_bb) + test_acc * len(inferred_test_bb)) / (len(inferred_train_bb) + len(inferred_test_bb))
print(train_acc)
print(test_acc)
print(acc)
```
Acheives slightly better results than the rule-based attack.
```
# black-box
print(calc_precision_recall(np.concatenate((inferred_train_bb, inferred_test_bb)),
np.concatenate((np.ones(len(inferred_train_bb)), np.zeros(len(inferred_test_bb))))))
```
## Train neural network model
```
import torch
from torch import nn, optim
from torch.utils.data import DataLoader
from torch.utils.data.dataset import Dataset
from art.estimators.classification.pytorch import PyTorchClassifier
class ModelToAttack(nn.Module):
def __init__(self, num_classes, num_features):
super(ModelToAttack, self).__init__()
self.fc1 = nn.Sequential(
nn.Linear(num_features, 1024),
nn.Tanh(), )
self.fc2 = nn.Sequential(
nn.Linear(1024, 512),
nn.Tanh(), )
self.classifier = nn.Linear(512, num_classes)
# self.softmax = nn.Softmax(dim=1)
def forward(self, x):
out = self.fc1(x)
out = self.fc2(out)
return self.classifier(out)
mlp_model = ModelToAttack(4, 24)
mlp_model = torch.nn.DataParallel(mlp_model)
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(mlp_model.parameters(), lr=0.0001)
class NurseryDataset(Dataset):
def __init__(self, x, y=None):
self.x = torch.from_numpy(x.astype(np.float64)).type(torch.FloatTensor)
if y is not None:
self.y = torch.from_numpy(y.astype(np.int8)).type(torch.LongTensor)
else:
self.y = torch.zeros(x.shape[0])
def __len__(self):
return len(self.x)
def __getitem__(self, idx):
if idx >= len(self.x):
raise IndexError("Invalid Index")
return self.x[idx], self.y[idx]
train_set = NurseryDataset(x_train, y_train)
train_loader = DataLoader(train_set, batch_size=100, shuffle=True, num_workers=0)
for epoch in range(20):
for (input, targets) in train_loader:
input, targets = torch.autograd.Variable(input), torch.autograd.Variable(targets)
optimizer.zero_grad()
outputs = mlp_model(input)
loss = criterion(outputs, targets)
loss.backward()
optimizer.step()
mlp_art_model = PyTorchClassifier(model=mlp_model, loss=criterion, optimizer=optimizer, input_shape=(24,), nb_classes=4)
pred = np.array([np.argmax(arr) for arr in mlp_art_model.predict(x_test.astype(np.float32))])
print('Base model accuracy: ', np.sum(pred == y_test) / len(y_test))
```
### Rule-based attack
```
mlp_attack = MembershipInferenceBlackBoxRuleBased(mlp_art_model)
# infer
mlp_inferred_train = mlp_attack.infer(x_train.astype(np.float32), y_train)
mlp_inferred_test = mlp_attack.infer(x_test.astype(np.float32), y_test)
# check accuracy
mlp_train_acc = np.sum(mlp_inferred_train) / len(mlp_inferred_train)
mlp_test_acc = 1 - (np.sum(mlp_inferred_test) / len(mlp_inferred_test))
mlp_acc = (mlp_train_acc * len(mlp_inferred_train) + mlp_test_acc * len(mlp_inferred_test)) / (len(mlp_inferred_train) + len(mlp_inferred_test))
print(mlp_train_acc)
print(mlp_test_acc)
print(mlp_acc)
print(calc_precision_recall(np.concatenate((mlp_inferred_train, mlp_inferred_test)),
np.concatenate((np.ones(len(mlp_inferred_train)), np.zeros(len(mlp_inferred_test))))))
```
### Black-box attack
```
mlp_attack_bb = MembershipInferenceBlackBox(mlp_art_model, attack_model_type='rf')
# train attack model
mlp_attack_bb.fit(x_train[:attack_train_size].astype(np.float32), y_train[:attack_train_size],
x_test[:attack_test_size].astype(np.float32), y_test[:attack_test_size])
# infer
mlp_inferred_train_bb = mlp_attack_bb.infer(x_train.astype(np.float32), y_train)
mlp_inferred_test_bb = mlp_attack_bb.infer(x_test.astype(np.float32), y_test)
# check accuracy
mlp_train_acc_bb = np.sum(mlp_inferred_train_bb) / len(mlp_inferred_train_bb)
mlp_test_acc_bb = 1 - (np.sum(mlp_inferred_test_bb) / len(mlp_inferred_test_bb))
mlp_acc_bb = (mlp_train_acc_bb * len(mlp_inferred_train_bb) + mlp_test_acc_bb * len(mlp_inferred_test_bb)) / (len(mlp_inferred_train_bb) + len(mlp_inferred_test_bb))
print(mlp_train_acc_bb)
print(mlp_test_acc_bb)
print(mlp_acc_bb)
print(calc_precision_recall(np.concatenate((mlp_inferred_train_bb, mlp_inferred_test_bb)),
np.concatenate((np.ones(len(mlp_inferred_train_bb)), np.zeros(len(mlp_inferred_test_bb))))))
```
Using a random forest as the attack model we were able to acheive better performance than the rule-based attack, both in terms of accuracy and precision.
| github_jupyter |
As we have seen in previous sections, hvPlot bakes in interactivity by automatically creating widgets when using ``groupby``. These widgets can be refined using [Panel](https://panel.pyviz.org). Panel allows you to customize the interactivity of your hvPlot output and provides more fine-grained control over the layout.
<div class="alert alert-warning" role="alert">
When viewing on a static website, the widgets will be inoperable. To explore this functionality fully, download the notebook and run it!
</div>
```
import panel as pn
import hvplot.pandas # noqa
from bokeh.sampledata.iris import flowers
```
When ``groupby`` is used, the default widget is selected for the data type of the column. In this case since 'species' is composed of strings, the widget an instance of the class ``pn.widgets.Select``.
```
flowers.hvplot.bivariate(x='sepal_width', y='sepal_length', width=600,
groupby='species')
```
### Customizing Widgets
We can change where the widget is shown using the ``widget_location`` option.
```
flowers.hvplot.bivariate(x='sepal_width', y='sepal_length', width=600,
groupby='species', widget_location='left_top')
```
We can also change what the class of the widget is, using the ``widgets`` dict. For instance if we want to use a slider instead of a selector we can specify that.
```
flowers.hvplot.bivariate(x='sepal_width', y='sepal_length', width=600,
groupby='species', widgets={'species': pn.widgets.DiscreteSlider})
```
### Using widgets as arguments
So far we have only been dealing with widgets that are produced when using the ``groupby`` key word. But panel provides many other ways of expanding the interactivity of hvplot objects. For instance we might want to allow the user to select which fields to plot on the ``x`` and ``y`` axes. Or even what ``kind`` of plot to produce.
```
x = pn.widgets.Select(name='x', options=['sepal_width', 'petal_width'])
y = pn.widgets.Select(name='y', options=['sepal_length', 'petal_length'])
kind = pn.widgets.Select(name='kind', value='scatter', options=['bivariate', 'scatter'])
plot = flowers.hvplot(x=x, y=y, kind=kind, colorbar=False, width=600)
pn.Row(pn.WidgetBox(x, y, kind), plot)
```
### Using functions
In addition to using widgets directly as arguments, we can also use functions that have been decorated with ``pn.depends``
```
x = pn.widgets.Select(name='x', options=['sepal_width', 'petal_width'])
y = pn.widgets.Select(name='y', options=['sepal_length', 'petal_length'])
kind = pn.widgets.Select(name='kind', value='scatter', options=['bivariate', 'scatter'])
by_species = pn.widgets.Checkbox(name='By species')
color = pn.widgets.ColorPicker(value='#ff0000')
@pn.depends(by_species, color)
def by_species_fn(by_species, color):
return 'species' if by_species else color
plot = flowers.hvplot(x=x, y=y, kind=kind, c=by_species_fn, colorbar=False, width=600, legend='top_right')
pn.Row(pn.WidgetBox(x, y, kind, color, by_species), plot)
```
We can keep even add a callback to disable the color options when 'bivariate' is selected. After running the cell below, try changing 'kind' above and notice how the color and 'By species' areas turn grey to indicate that they are disabled.
```
def update(event):
if kind.value == 'bivariate':
color.disabled = True
by_species.disabled = True
else:
color.disabled = False
by_species.disabled = False
kind.param.watch(update, 'value');
```
To learn more about Panel and how to use it with output from hvPlot, see the [Panel docs on the HoloViews pane](http://panel.pyviz.org/reference/panes/HoloViews.html). To learn more about available widgets, see the Widgets' section of the [Panel Reference Gallery](http://panel.pyviz.org/reference/index.html).
| github_jupyter |
```
from IPython.display import Image
```
# CNTK 204: Sequence to Sequence Networks with Text Data
## Introduction and Background
This hands-on tutorial will take you through both the basics of sequence-to-sequence networks, and how to implement them in the Microsoft Cognitive Toolkit. In particular, we will implement a sequence-to-sequence model with attention to perform grapheme to phoneme translation. We will start with some basic theory and then explain the data in more detail, and how you can download it.
Andrej Karpathy has a [nice visualization](http://karpathy.github.io/2015/05/21/rnn-effectiveness/) of five common paradigms of neural network architectures:
```
# Figure 1
Image(url="http://cntk.ai/jup/paradigms.jpg", width=750)
```
In this tutorial, we are going to be talking about the fourth paradigm: many-to-many where the length of the output does not necessarily equal the length of the input, also known as sequence-to-sequence networks. The input is a sequence with a dynamic length, and the output is also a sequence with some dynamic length. It is the logical extension of the many-to-one paradigm in that previously we were predicting some category (which could easily be one of `V` words where `V` is an entire vocabulary) and now we want to predict a whole sequence of those categories.
The applications of sequence-to-sequence networks are nearly limitless. It is a natural fit for machine translation (e.g. English input sequences, French output sequences); automatic text summarization (e.g. full document input sequence, summary output sequence); word to pronunciation models (e.g. character [grapheme] input sequence, pronunciation [phoneme] output sequence); and even parse tree generation (e.g. regular text input, flat parse tree output).
## Basic theory
A sequence-to-sequence model consists of two main pieces: (1) an encoder; and (2) a decoder. Both the encoder and the decoder are recurrent neural network (RNN) layers that can be implemented using a vanilla RNN, an LSTM, or GRU Blocks (here we will use LSTM). In the basic sequence-to-sequence model, the encoder processes the input sequence into a fixed representation that is fed into the decoder as a context. The decoder then uses some mechanism (discussed below) to decode the processed information into an output sequence. The decoder is a language model that is augmented with some "strong context" by the encoder, and so each symbol that it generates is fed back into the decoder for additional context (like a traditional LM). For an English to German translation task, the most basic setup might look something like this:
```
# Figure 2
Image(url="http://cntk.ai/jup/s2s.png", width=700)
```
The basic sequence-to-sequence network passes the information from the encoder to the decoder by initializing the decoder RNN with the final hidden state of the encoder as its initial hidden state. The input is then a "sequence start" tag (`<s>` in the diagram above) which primes the decoder to start generating an output sequence. Then, whatever word (or note or image, etc.) it generates at that step is fed in as the input for the next step. The decoder keeps generating outputs until it hits the special "end sequence" tag (`</s>` above).
A more complex and powerful version of the basic sequence-to-sequence network uses an attention model. While the above setup works well, it can start to break down when the input sequences get long. At each step, the hidden state `h` is getting updated with the most recent information, and therefore `h` might be getting "diluted" in information as it processes each token. Further, even with a relatively short sequence, the last token will always get the last say and therefore the thought vector will be somewhat biased/weighted towards that last word. To deal with this problem, we use an "attention" mechanism that allows the decoder to look not only at all of the hidden states from the input, but it also learns which hidden states, for each step in decoding, to put the most weight on. In this tutorial we will implement a sequence-to-sequence network that can be run either with or without attention enabled.
```
# Figure 3
Image(url="https://cntk.ai/jup/cntk204_s2s2.png", width=700)
```
The `Attention` layer above takes the current value of the hidden state in the Decoder, all of the hidden states in the Encoder, and calculates an augmented version of the hidden state to use. More specifically, the contribution from the Encoder's hidden states will represent a weighted sum of all of its hidden states where the highest weight corresponds both to the biggest contribution to the augmented hidden state and to the hidden state that will be most important for the Decoder to consider when generating the next word.
## Problem: Grapheme-to-Phoneme Conversion
The [grapheme](https://en.wikipedia.org/wiki/Grapheme) to [phoneme](https://en.wikipedia.org/wiki/Phoneme) problem is a translation task that takes the letters of a word as the input sequence (the graphemes are the smallest units of a writing system) and outputs the corresponding phonemes; that is, the units of sound that make up a language. In other words, the system aims to generate an unambigious representation of how to pronounce a given input word.
**Example**
The graphemes or the letters are translated into corresponding phonemes:
> **Grapheme** : **|** T **|** A **|** N **|** G **|** E **|** R **|**
**Phonemes** : **|** ~T **|** ~AE **|** ~NG **|** ~ER **|**
**Model structure overview**
As discussed above, the task we are interested in solving is creating a model that takes some sequence as an input, and generates an output sequence based on the contents of the input. The model's job is to learn the mapping from the input sequence to the output sequence that it will generate. The job of the encoder is to come up with a good representation of the input that the decoder can use to generate a good output. For both the encoder and the decoder, the LSTM does a good job at this.
Note that the LSTM is simply one of a whole set of different types of Blocks that can be used to implement an RNN. This is the code that is run for each step in the recurrence. In the Layers library, there are three built-in recurrent Blocks: the (vanilla) `RNN`, the `GRU`, and the `LSTM`. Each processes its input slightly differently and each has its own benefits and drawbacks for different types of tasks and networks. To get these blocks to run for each of the elements recurrently in a network, we create a `Recurrence` over them. This "unrolls" the network to the number of steps that are in the given input for the RNN layer.
**Importing CNTK and other useful libraries**
CNTK is a Python module that contains several submodules like `io`, `learner`, `graph`, etc. We make extensive use of numpy as well.
```
from __future__ import print_function
import numpy as np
import os
import cntk as C
import cntk.tests.test_utils
cntk.tests.test_utils.set_device_from_pytest_env() # (only needed for our build system)
C.cntk_py.set_fixed_random_seed(1) # fix a random seed for CNTK components
# Check if this is a test environment
def isTest():
return ('TEST_DEVICE' in os.environ)
```
### Downloading the data
In this tutorial we will use a lightly pre-processed version of the CMUDict (version 0.7b) dataset from http://www.speech.cs.cmu.edu/cgi-bin/cmudict. The CMUDict data refers to the Carnegie Mellon University Pronouncing Dictionary and is an open-source machine-readable pronunciation dictionary for North American English. The data is in the CNTKTextFormatReader format. Here is an example sequence pair from the data, where the input sequence (S0) is in the left column, and the output sequence (S1) is on the right:
```
0 |S0 3:1 |# <s> |S1 3:1 |# <s>
0 |S0 4:1 |# A |S1 32:1 |# ~AH
0 |S0 5:1 |# B |S1 36:1 |# ~B
0 |S0 4:1 |# A |S1 31:1 |# ~AE
0 |S0 7:1 |# D |S1 38:1 |# ~D
0 |S0 12:1 |# I |S1 47:1 |# ~IY
0 |S0 1:1 |# </s> |S1 1:1 |# </s>
```
The code below will download the required files (training, testing, the single sequence above for visual validation, and a small vocab file) and put them in a local folder (the training file is ~34 MB, testing is ~4MB, and the validation file and vocab file are both less than 1KB).
```
import requests
def download(url, filename):
""" utility function to download a file """
response = requests.get(url, stream=True)
with open(filename, "wb") as handle:
for data in response.iter_content():
handle.write(data)
MODEL_DIR = "."
DATA_DIR = os.path.join('..', 'Examples', 'SequenceToSequence', 'CMUDict', 'Data')
# If above directory does not exist, just use current.
if not os.path.exists(DATA_DIR):
DATA_DIR = '.'
dataPath = {
'validation': 'tiny.ctf',
'training': 'cmudict-0.7b.train-dev-20-21.ctf',
'testing': 'cmudict-0.7b.test.ctf',
'vocab_file': 'cmudict-0.7b.mapping',
}
for k in sorted(dataPath.keys()):
path = os.path.join(DATA_DIR, dataPath[k])
if os.path.exists(path):
print("Reusing locally cached:", path)
else:
print("Starting download:", dataPath[k])
url = "https://github.com/Microsoft/CNTK/blob/release/2.6/Examples/SequenceToSequence/CMUDict/Data/%s?raw=true"%dataPath[k]
download(url, path)
print("Download completed")
dataPath[k] = path
```
### Data Reader
To efficiently collect our data, randomize it for training, and pass it to the network, we use the CNTKTextFormat reader. We will create a small function that will be called when training (or testing) that defines the names of the streams in our data, and how they are referred to in the raw training data.
```
# Helper function to load the model vocabulary file
def get_vocab(path):
# get the vocab for printing output sequences in plaintext
vocab = [w.strip() for w in open(path).readlines()]
i2w = { i:w for i,w in enumerate(vocab) }
w2i = { w:i for i,w in enumerate(vocab) }
return (vocab, i2w, w2i)
# Read vocabulary data and generate their corresponding indices
vocab, i2w, w2i = get_vocab(dataPath['vocab_file'])
input_vocab_dim = 69
label_vocab_dim = 69
# Print vocab and the correspoding mapping to the phonemes
print("Vocabulary size is", len(vocab))
print("First 15 letters are:")
print(vocab[:15])
print()
print("Print dictionary with the vocabulary mapping:")
print(i2w)
```
We will use the above to create a reader for our training data. Let's create it now:
```
def create_reader(path, is_training):
return C.io.MinibatchSource(C.io.CTFDeserializer(path, C.io.StreamDefs(
features = C.io.StreamDef(field='S0', shape=input_vocab_dim, is_sparse=True),
labels = C.io.StreamDef(field='S1', shape=label_vocab_dim, is_sparse=True)
)), randomize = is_training, max_sweeps = C.io.INFINITELY_REPEAT if is_training else 1)
# Train data reader
train_reader = create_reader(dataPath['training'], True)
# Validation data reader
valid_reader = create_reader(dataPath['validation'], True)
```
**Set our model hyperparameters**
We have a number of settings that control the complexity of our network, the shapes of our inputs, and other options such as whether we will use an embedding (and what size to use), and whether or not we will employ attention. We set them now as they will be made use of when we build the network graph in the following sections.
```
hidden_dim = 512
num_layers = 2
attention_dim = 128
use_attention = True
use_embedding = True
embedding_dim = 200
vocab = ([w.strip() for w in open(dataPath['vocab_file']).readlines()]) # all lines of vocab_file in a list
length_increase = 1.5
```
## Model Creation
We will set two more parameters now: the symbols used to denote the start of a sequence (sometimes called 'BOS') and the end of a sequence (sometimes called 'EOS'). In this case, our sequence-start symbol is the tag $<s>$ and our sequence-end symbol is the end-tag $</s>$.
Sequence start and end tags are important in sequence-to-sequence networks for two reasons. The sequence start tag is a "primer" for the decoder; in other words, because we are generating an output sequence and RNNs require some input, the sequence start token "primes" the decoder to cause it to emit its first generated token. The sequence end token is important because the decoder will learn to output this token when the sequence is finished. Otherwise the network wouldn't know how long of a sequence to generate. For the code below, we setup the sequence start symbol as a `Constant` so that it can later be passed to the Decoder LSTM as its `initial_state`. Further, we get the sequence end symbol's index so that the Decoder can use it to know when to stop generating tokens.
```
sentence_start =C.Constant(np.array([w=='<s>' for w in vocab], dtype=np.float32))
sentence_end_index = vocab.index('</s>')
```
## Step 1: setup the input to the network
### Dynamic axes in CNTK (Key concept)
One of the important concepts in understanding CNTK is the idea of two types of axes:
- **static axes**, which are the traditional axes of a variable's shape, and
- **dynamic axes**, which have dimensions that are unknown until the variable is bound to real data at computation time.
The dynamic axes are particularly important in the world of recurrent neural networks. Instead of having to decide a maximum sequence length ahead of time, padding your sequences to that size, and wasting computation, CNTK's dynamic axes allow for variable sequence lengths that are automatically packed in minibatches to be as efficient as possible.
When setting up sequences, there are *two dynamic axes* that are important to consider. The first is the *batch axis*, which is the axis along which multiple sequences are batched. The second is the dynamic axis particular to that sequence. The latter is specific to a particular input because of variable sequence lengths in your data. For example, in sequence to sequence networks, we have two sequences: the **input sequence**, and the **output (or 'label') sequence**. One of the things that makes this type of network so powerful is that the length of the input sequence and the output sequence do not have to correspond to each other. Therefore, both the input sequence and the output sequence require their own unique dynamic axis.
We first create the `inputAxis` for the input sequence and the `labelAxis` for the output sequence. We then define the inputs to the model by creating sequences over these two unique dynamic axes. Note that `InputSequence` and `LabelSequence` are *type declarations*. This means that the `InputSequence` is a type that consists of a sequence over the `inputAxis` axis.
```
# Source and target inputs to the model
inputAxis = C.Axis('inputAxis')
labelAxis = C.Axis('labelAxis')
InputSequence = C.layers.SequenceOver[inputAxis]
LabelSequence = C.layers.SequenceOver[labelAxis]
```
## Step 2: define the network
As discussed before, the sequence-to-sequence network is, at its most basic, an RNN (LSTM) encoder followed by an RNN (LSTM) decoder, and a dense output layer. We will implement both the Encoder and the Decoder using the CNTK Layers library. Both of these will be created as CNTK Functions. Our `create_model()` Python function creates both the `encode` and `decode` CNTK Functions. The `decode` function directly makes use of the `encode` function and the return value of `create_model()` is the CNTK Function `decode` itself.
We start by passing the input through an embedding (learned as part of the training process). So that this function can be used in the `Sequential` block of the Encoder and the Decoder whether we want an embedding or not, we will use the `identity` function if the `use_embedding` parameter is `False`. We then declare the Encoder layers as follows:
First, we pass the input through our `embed` function and then we stabilize it. This adds an additional scalar parameter to the learning that can help our network converge more quickly during training. Then, for each of the number of LSTM layers that we want in our encoder, except the final one, we set up an LSTM recurrence. The final recurrence will be a `Fold` if we are not using attention because we only pass the final hidden state to the decoder. If we are using attention, however, then we use another normal LSTM `Recurrence` that the Decoder will put its attention over later on.
Below we see a diagram of how the layered version of the sequence-to-sequence network with attention works. As the code shows below, the output of each layer of the Encoder and Decoder is used as the input to the layer just above it. The Attention model focuses on the top layer of the Encoder and informs the first layer of the Decoder.
```
# Figure 4
Image(url="https://cntk.ai/jup/cntk204_s2s3.png", width=900)
```
For the decoder, we first define several sub-layers: the `Stabilizer` for the decoder input, the `Recurrence` blocks for each of the decoder's layers, the `Stabilizer` for the output of the stack of LSTMs, and the final `Dense` output layer. If we are using attention, then we also create an `AttentionModel` function `attention_model` which returns an augmented version of the decoder's hidden state with emphasis placed on the encoder hidden states that should be most used for the given step while generating the next output token.
We then build the CNTK Function `decode`. The decorator `@Function` turns a regular Python function into a proper CNTK Function with the given arguments and return value. The Decoder works differently during training than it does during test time. During training, the history (i.e. input) to the Decoder `Recurrence` consists of the ground-truth labels. This means that while generating $y^{(t=2)}$, for example, the input will be $y^{(t=1)}$. During evaluation, or "test time", however, the input to the Decoder will be the actual output of the model. For a greedy decoder -- which we are implementing here -- that input is therefore the `hardmax` of the final `Dense` layer.
The Decoder Function `decode` takes two arguments: (1) the `input` sequence; and (2) the Decoder `history`. First, it runs the `input` sequence through the Encoder function `encode` that we setup earlier. We then get the `history` and map it to its embedding if necessary. Then the embedded representation is stabilized before running it through the Decoder's `Recurrence`. For each layer of `Recurrence`, we run the embedded `history` (now represented as `r`) through the `Recurrence`'s LSTM. If we are not using attention, we run it through the `Recurrence` with its initial state set to the value of the final hidden state of the encoder (note that since we run the Encoder backwards when not using attention that the "final" hidden state is actually the first hidden state in chronological time). If we are using attention, however, then we calculate the auxiliary input `h_att` using our `attention_model` function and we splice that onto the input `x`. This augmented `x` is then used as input for the Decoder's `Recurrence`.
Finally, we stabilize the output of the Decoder, put it through the final `Dense` layer `proj_out`, and label the output using the `Label` layer which allows for simple access to that layer later on.
```
# create the s2s model
def create_model(): # :: (history*, input*) -> logP(w)*
# Embedding: (input*) --> embedded_input*
embed = C.layers.Embedding(embedding_dim, name='embed') if use_embedding else identity
# Encoder: (input*) --> (h0, c0)
# Create multiple layers of LSTMs by passing the output of the i-th layer
# to the (i+1)th layer as its input
# Note: We go_backwards for the plain model, but forward for the attention model.
with C.layers.default_options(enable_self_stabilization=True, go_backwards=not use_attention):
LastRecurrence = C.layers.Fold if not use_attention else C.layers.Recurrence
encode = C.layers.Sequential([
embed,
C.layers.Stabilizer(),
C.layers.For(range(num_layers-1), lambda:
C.layers.Recurrence(C.layers.LSTM(hidden_dim))),
LastRecurrence(C.layers.LSTM(hidden_dim), return_full_state=True),
(C.layers.Label('encoded_h'), C.layers.Label('encoded_c')),
])
# Decoder: (history*, input*) --> unnormalized_word_logp*
# where history is one of these, delayed by 1 step and <s> prepended:
# - training: labels
# - testing: its own output hardmax(z) (greedy decoder)
with C.layers.default_options(enable_self_stabilization=True):
# sub-layers
stab_in = C.layers.Stabilizer()
rec_blocks = [C.layers.LSTM(hidden_dim) for i in range(num_layers)]
stab_out = C.layers.Stabilizer()
proj_out = C.layers.Dense(label_vocab_dim, name='out_proj')
# attention model
if use_attention: # maps a decoder hidden state and all the encoder states into an augmented state
attention_model = C.layers.AttentionModel(attention_dim,
name='attention_model') # :: (h_enc*, h_dec) -> (h_dec augmented)
# layer function
@C.Function
def decode(history, input):
encoded_input = encode(input)
r = history
r = embed(r)
r = stab_in(r)
for i in range(num_layers):
rec_block = rec_blocks[i] # LSTM(hidden_dim) # :: (dh, dc, x) -> (h, c)
if use_attention:
if i == 0:
@C.Function
def lstm_with_attention(dh, dc, x):
h_att = attention_model(encoded_input.outputs[0], dh)
x = C.splice(x, h_att)
return rec_block(dh, dc, x)
r = C.layers.Recurrence(lstm_with_attention)(r)
else:
r = C.layers.Recurrence(rec_block)(r)
else:
# unlike Recurrence(), the RecurrenceFrom() layer takes the initial hidden state as a data input
r = C.layers.RecurrenceFrom(rec_block)(*(encoded_input.outputs + (r,))) # :: h0, c0, r -> h
r = stab_out(r)
r = proj_out(r)
r = C.layers.Label('out_proj_out')(r)
return r
return decode
```
The network that we defined above can be thought of as an "abstract" model that must first be wrapped to be used. In this case, we will use it first to create a "training" version of the model (where the history for the Decoder will be the ground-truth labels), and then we will use it to create a greedy "decoding" version of the model where the history for the Decoder will be the `hardmax` output of the network. Let's set up these model wrappers next.
## Training
Before starting training, we will define the training wrapper, the greedy decoding wrapper, and the criterion function used for training the model. Let's start with the training wrapper.
```
def create_model_train(s2smodel):
# model used in training (history is known from labels)
# note: the labels must NOT contain the initial <s>
@C.Function
def model_train(input, labels): # (input*, labels*) --> (word_logp*)
# The input to the decoder always starts with the special label sequence start token.
# Then, use the previous value of the label sequence (for training) or the output (for execution).
past_labels = C.layers.Delay(initial_state=sentence_start)(labels)
return s2smodel(past_labels, input)
return model_train
```
Above, we create the CNTK Function `model_train` again using the `@Function` decorator. This function takes the input sequence `input` and the output sequence `labels` as arguments. The `past_labels` are setup as the `history` for the model we created earlier by using the `Delay` layer. This will return the previous time-step value for the input `labels` with an `initial_state` of `sentence_start`. Therefore, if we give the labels `['a', 'b', 'c']`, then `past_labels` will contain `['<s>', 'a', 'b', 'c']` and then return our abstract base model called with the history `past_labels` and the input `input`.
Let's go ahead and create the greedy decoding model wrapper now as well:
```
def create_model_greedy(s2smodel):
# model used in (greedy) decoding (history is decoder's own output)
@C.Function
@C.layers.Signature(InputSequence[C.layers.Tensor[input_vocab_dim]])
def model_greedy(input): # (input*) --> (word_sequence*)
# Decoding is an unfold() operation starting from sentence_start.
# We must transform s2smodel (history*, input* -> word_logp*) into a generator (history* -> output*)
# which holds 'input' in its closure.
unfold = C.layers.UnfoldFrom(lambda history: s2smodel(history, input) >> C.hardmax,
# stop once sentence_end_index was max-scoring output
until_predicate=lambda w: w[...,sentence_end_index],
length_increase=length_increase)
return unfold(initial_state=sentence_start, dynamic_axes_like=input)
return model_greedy
```
Above we create a new CNTK Function `model_greedy` which this time only takes a single argument. This is of course because when using the model at test time we don't have any labels -- it is the model's job to create them for us! In this case, we use the `UnfoldFrom` layer which runs the base model with the current `history` and funnels it into the `hardmax`. The `hardmax`'s output then becomes part of the `history` and we keep unfolding the `Recurrence` until the `sentence_end_index` has been reached. The maximum length of the output sequence (the maximum unfolding of the Decoder) is determined by a multiplier passed to `length_increase`. In this case we set `length_increase` to `1.5` above so the maximum length of each output sequence is 1.5x its input.
The last thing we will do before setting up the training loop is define the function that will create the criterion function for our model.
```
def create_criterion_function(model):
@C.Function
@C.layers.Signature(input=InputSequence[C.layers.Tensor[input_vocab_dim]],
labels=LabelSequence[C.layers.Tensor[label_vocab_dim]])
def criterion(input, labels):
# criterion function must drop the <s> from the labels
postprocessed_labels = C.sequence.slice(labels, 1, 0) # <s> A B C </s> --> A B C </s>
z = model(input, postprocessed_labels)
ce = C.cross_entropy_with_softmax(z, postprocessed_labels)
errs = C.classification_error(z, postprocessed_labels)
return (ce, errs)
return criterion
```
Above, we create the criterion function which drops the sequence-start symbol from our labels for us, runs the model with the given `input` and `labels`, and uses the output to compare to our ground truth. We use the loss function `cross_entropy_with_softmax` and get the `classification_error` which gives us the percent-error per-word of our generation accuracy. The CNTK Function `criterion` returns these values as a tuple and the Python function `create_criterion_function(model)` returns that CNTK Function.
Now let's move on to creating the training loop...
```
def train(train_reader, valid_reader, vocab, i2w, s2smodel, max_epochs, epoch_size):
# create the training wrapper for the s2smodel, as well as the criterion function
model_train = create_model_train(s2smodel)
criterion = create_criterion_function(model_train)
# also wire in a greedy decoder so that we can properly log progress on a validation example
# This is not used for the actual training process.
model_greedy = create_model_greedy(s2smodel)
# Instantiate the trainer object to drive the model training
minibatch_size = 72
lr = 0.001 if use_attention else 0.005
learner = C.fsadagrad(model_train.parameters,
#apply the learning rate as if it is a minibatch of size 1
lr = C.learning_parameter_schedule_per_sample([lr]*2+[lr/2]*3+[lr/4], epoch_size),
momentum = C.momentum_schedule(0.9366416204111472, minibatch_size=minibatch_size),
gradient_clipping_threshold_per_sample=2.3,
gradient_clipping_with_truncation=True)
trainer = C.Trainer(None, criterion, learner)
# Get minibatches of sequences to train with and perform model training
total_samples = 0
mbs = 0
eval_freq = 100
# print out some useful training information
C.logging.log_number_of_parameters(model_train) ; print()
progress_printer = C.logging.ProgressPrinter(freq=30, tag='Training')
# a hack to allow us to print sparse vectors
sparse_to_dense = create_sparse_to_dense(input_vocab_dim)
for epoch in range(max_epochs):
while total_samples < (epoch+1) * epoch_size:
# get next minibatch of training data
mb_train = train_reader.next_minibatch(minibatch_size)
# do the training
trainer.train_minibatch({criterion.arguments[0]: mb_train[train_reader.streams.features],
criterion.arguments[1]: mb_train[train_reader.streams.labels]})
progress_printer.update_with_trainer(trainer, with_metric=True) # log progress
# every N MBs evaluate on a test sequence to visually show how we're doing
if mbs % eval_freq == 0:
mb_valid = valid_reader.next_minibatch(1)
# run an eval on the decoder output model (i.e. don't use the groundtruth)
e = model_greedy(mb_valid[valid_reader.streams.features])
print(format_sequences(sparse_to_dense(mb_valid[valid_reader.streams.features]), i2w))
print("->")
print(format_sequences(e, i2w))
# visualizing attention window
if use_attention:
debug_attention(model_greedy, mb_valid[valid_reader.streams.features])
total_samples += mb_train[train_reader.streams.labels].num_samples
mbs += 1
# log a summary of the stats for the epoch
progress_printer.epoch_summary(with_metric=True)
# done: save the final model
model_path = "model_%d.cmf" % epoch
print("Saving final model to '%s'" % model_path)
s2smodel.save(model_path)
print("%d epochs complete." % max_epochs)
```
In the above function, we created one version of the model for training (plus its associated criterion function) and one version of the model for evaluation. Normally this latter version would not be required but here we have done it so that we can periodically sample from the non-training model to visually understand how our model is converging by seeing the kinds of sequences that it generates as the training progresses.
We then setup some standard variables required for the training loop. We set the `minibatch_size` (which refers to the total number of elements -- NOT sequences -- in a minibatch), the initial learning rate `lr`, we initialize a `learner` using the `adam_sgd` algorithm and a `learning_rate_schedule` that slowly reduces our learning rate. We make use of gradient clipping to help control exploding gradients, and we finally create our `Trainer` object `trainer`.
We make use of CNTK's `ProgressPrinter` class which takes care of calculating average metrics per minibatch/epoch and we set it to update every 30 minibatches. And finally, before starting the training loop, we initialize a function called `sparse_to_dense` which we use to properly print out the input sequence data that we use for validation because it is sparse. That function is defined just below:
```
# dummy for printing the input sequence below. Currently needed because input is sparse.
def create_sparse_to_dense(input_vocab_dim):
I = C.Constant(np.eye(input_vocab_dim))
@C.Function
@C.layers.Signature(InputSequence[C.layers.SparseTensor[input_vocab_dim]])
def no_op(input):
return C.times(input, I)
return no_op
```
Inside the training loop, we proceed much like many other CNTK networks. We request the next bunch of minibatch data, we perform our training, and we print our progress to the screen using the `progress_printer`. Where we diverge from the norm, however, is where we run an evaluation using our `model_greedy` version of the network and run a single sequence, "ABADI" through to see what the network is currently predicting.
Another difference in the training loop is the optional attention window visualization. Calling the function `debug_attention` shows the weight that the Decoder put on each of the Encoder's hidden states for each of the output tokens that it generated. This function, along with the `format_sequences` function required to print the input/output sequences to the screen, are given below.
```
# Given a vocab and tensor, print the output
def format_sequences(sequences, i2w):
return [" ".join([i2w[np.argmax(w)] for w in s]) for s in sequences]
# to help debug the attention window
def debug_attention(model, input):
q = C.combine([model, model.attention_model.attention_weights])
#words, p = q(input) # Python 3
words_p = q(input)
words = words_p[0]
p = words_p[1]
output_seq_len = words[0].shape[0]
p_sq = np.squeeze(p[0][:output_seq_len,:,:]) # (batch, output_len, input_len, 1)
opts = np.get_printoptions()
np.set_printoptions(precision=5)
print(p_sq)
np.set_printoptions(**opts)
```
Let's try training our network for a small part of an epoch. In particular, we'll run through 25,000 tokens (about 3% of one epoch):
```
model = create_model()
train(train_reader, valid_reader, vocab, i2w, model, max_epochs=1, epoch_size=25000)
```
As we can see above, while the loss has come down quite a ways, the output sequence is still quite a ways off from what we expect. Uncomment the code below to run for a full epoch (notice that we switch the `epoch_size` parameter to the actual size of the training data) and by the end of the first epoch you will already see a very good grapheme-to-phoneme translation model running!
```
# Uncomment the line below to train the model for a full epoch
#train(train_reader, valid_reader, vocab, i2w, model, max_epochs=1, epoch_size=908241)
```
## Testing the network
Now that we've trained a sequence-to-sequence network for graphme-to-phoneme translation, there are two important things we should do with it. First, we should test its accuracy on a held-out test set. Then, we should try it out in an interactive environment so that we can put in our own input sequences and see what the model predicts. Let's start by determining the test string error rate.
At the end of training, we saved the model using the line `s2smodel.save(model_path)`. Therefore, to test it, we will need to first `load` that model and then run some test data through it. Let's `load` the model, then create a reader configured to access our testing data. Note that we pass `False` to the `create_reader` function this time to denote that we are in testing mode so we should only pass over the data a single time.
```
# load the model for epoch 0
model_path = "model_0.cmf"
model = C.Function.load(model_path)
# create a reader pointing at our testing data
test_reader = create_reader(dataPath['testing'], False)
```
Now we need to define our testing function. We pass the `reader`, the learned `s2smodel`, and the vocabulary map `i2w` so that we can directly compare the model's predictions to the test set labels. We loop over the test set, evaluate the model on minibatches of size 512 for efficiency, and keep track of the error rate. Note that below we test *per-sequence*. This means that every single token in a generated sequence must match the tokens in the label for that sequence to be considered as correct.
```
# This decodes the test set and counts the string error rate.
def evaluate_decoding(reader, s2smodel, i2w):
model_decoding = create_model_greedy(s2smodel) # wrap the greedy decoder around the model
progress_printer = C.logging.ProgressPrinter(tag='Evaluation')
sparse_to_dense = create_sparse_to_dense(input_vocab_dim)
minibatch_size = 512
num_total = 0
num_wrong = 0
while True:
mb = reader.next_minibatch(minibatch_size)
if not mb: # finish when end of test set reached
break
e = model_decoding(mb[reader.streams.features])
outputs = format_sequences(e, i2w)
labels = format_sequences(sparse_to_dense(mb[reader.streams.labels]), i2w)
# prepend sentence start for comparison
outputs = ["<s> " + output for output in outputs]
num_total += len(outputs)
num_wrong += sum([label != output for output, label in zip(outputs, labels)])
rate = num_wrong / num_total
print("string error rate of {:.1f}% in {} samples".format(100 * rate, num_total))
return rate
```
Now we will evaluate the decoding using the above function. If you use the version of the model we trained above with just a small 50000 sample of the training data, you will get an error rate of 100% because we cannot possibly get every single token correct with such a small amount of training. However, if you uncommented the training line above that trains the network for a full epoch, you should have ended up with a much-improved model that showed approximately the following training statistics:
```
Finished Epoch[1 of 300]: [Training] loss = 0.878420 * 799303, metric = 26.23% * 799303 1755.985s (455.2 samples/s);
```
Now let's evaluate the model's test set performance below.
```
# print the string error rate
evaluate_decoding(test_reader, model, i2w)
```
If you did not run the training for the full first epoch, the output above will be a `1.0` meaning 100% string error rate. If, however, you uncommented the line to perform training for a full epoch, you should get an output of `0.569`. A string error rate of `56.9` is actually not bad for a single pass over the data. Let's now modify the above `evaluate_decoding` function to output the per-phoneme error rate. This means that we are calculating the error at a higher precision and also makes things easier in some sense because with the string error rate we could have every phoneme correct but one in each example and still end up with a 100% error rate. Here is the modified version of that function:
```
# This decodes the test set and counts the string error rate.
def evaluate_decoding(reader, s2smodel, i2w):
model_decoding = create_model_greedy(s2smodel) # wrap the greedy decoder around the model
progress_printer = C.logging.ProgressPrinter(tag='Evaluation')
sparse_to_dense = create_sparse_to_dense(input_vocab_dim)
minibatch_size = 512
num_total = 0
num_wrong = 0
while True:
mb = reader.next_minibatch(minibatch_size)
if not mb: # finish when end of test set reached
break
e = model_decoding(mb[reader.streams.features])
outputs = format_sequences(e, i2w)
labels = format_sequences(sparse_to_dense(mb[reader.streams.labels]), i2w)
# prepend sentence start for comparison
outputs = ["<s> " + output for output in outputs]
for s in range(len(labels)):
for w in range(len(labels[s])):
num_total += 1
if w < len(outputs[s]): # in case the prediction is longer than the label
if outputs[s][w] != labels[s][w]:
num_wrong += 1
rate = num_wrong / num_total
print("{:.1f}".format(100 * rate))
return rate
# print the phoneme error rate
test_reader = create_reader(dataPath['testing'], False)
evaluate_decoding(test_reader, model, i2w)
```
If you're using the model that was trained for one full epoch, then you should get a phoneme error rate of around 10%. Not bad! This means that for each of the 383,294 phonemes in the test set, our model predicted nearly 90% of them correctly (if you used the quickly-trained version of the model then you will get an error rate of around 45%). Now, let's work with an interactive session where we can input our own input sequences and see how the model predicts their pronunciation (i.e. phonemes). Additionally, we will visualize the Decoder's attention for these samples to see which graphemes in the input it deemed to be important for each phoneme that it produces. Note that in the examples below the results will only be good if you use a model that has been trained for at least one epoch.
## Interactive session
Here we will write an interactive function to make it easy to interact with the trained model and try out your own input sequences that do not appear in the test set. Please note that the results will be very poor if you just use the model that was trained for a very short amount of time. The model we used just above that was trained for one epoch does a good job, and if you have the time and patience to train the model for a full 30 epochs, it will perform very nicely.
We will first import some graphics libraries that make the attention visualization possible and then we will define the `translate` function that takes a numpy-based representation of the input and runs our model.
```
# imports required for showing the attention weight heatmap
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
def translate(tokens, model_decoding, vocab, i2w, show_attention=False):
vdict = {v:i for i,v in enumerate(vocab)}
try:
w = [vdict["<s>"]] + [vdict[c] for c in tokens] + [vdict["</s>"]]
except:
print('Input contains an unexpected token.')
return []
# convert to one_hot
query = C.Value.one_hot([w], len(vdict))
pred = model_decoding(query)
pred = pred[0] # first sequence (we only have one) -> [len, vocab size]
if use_attention:
pred = np.squeeze(pred) # attention has extra dimensions
# print out translation and stop at the sequence-end tag
prediction = np.argmax(pred, axis=-1)
translation = [i2w[i] for i in prediction]
# show attention window (requires matplotlib, seaborn, and pandas)
if use_attention and show_attention:
q = C.combine([model_decoding.attention_model.attention_weights])
att_value = q(query)
# get the attention data up to the length of the output (subset of the full window)
att_value = np.squeeze(att_value[0][0:len(prediction),0:len(w)])
# set up the actual words/letters for the heatmap axis labels
columns = [i2w[ww] for ww in prediction]
index = [i2w[ww] for ww in w]
dframe = pd.DataFrame(data=np.fliplr(att_value.T), columns=columns, index=index)
sns.heatmap(dframe)
plt.show()
return translation
```
The `translate` function above takes a list of letters input by the user as `tokens`, the greedy decoding version of our model `model_decoding`, the vocabulary `vocab`, a map of index to vocab `i2w`, and the `show_attention` option which determines if we will visualize the attention vectors or not.
We convert our input into a `one_hot` representation, run it through the model with `model_decoding(query)` and, since each prediction is actually a probability distribution over the entire vocabulary, we take the `argmax` to get the most probable token for each step.
To visualize the attention window, we use `combine` to turn the `attention_weights` into a CNTK Function that takes the inputs that we expect. This way, when we run the function `q`, the output will be the values of the `attention_weights`. We do some data manipulation to get this data into the format that `sns` expects, and we show the visualization.
Finally, we need to write the user-interaction loop which allows a user to enter multiple inputs.
```
def interactive_session(s2smodel, vocab, i2w, show_attention=False):
model_decoding = create_model_greedy(s2smodel) # wrap the greedy decoder around the model
import sys
print('Enter one or more words to see their phonetic transcription.')
while True:
if isTest(): # Testing a prefilled text for routine testing
line = "psychology"
else:
line = input("> ")
if line.lower() == "quit":
break
# tokenize. Our task is letter to sound.
out_line = []
for word in line.split():
in_tokens = [c.upper() for c in word]
out_tokens = translate(in_tokens, model_decoding, vocab, i2w, show_attention=True)
out_line.extend(out_tokens)
out_line = [" " if tok == '</s>' else tok[1:] for tok in out_line]
print("=", " ".join(out_line))
sys.stdout.flush()
if isTest(): #If test environment we will test the translation only once
break
```
The above function simply creates a greedy decoder around our model and then continually asks the user for an input which we pass to our `translate` function. Visualizations of the attention will continue being appended to the notebook until you exit the loop by typing `quit`. Please uncomment the following line to try out the interaction session.
```
interactive_session(model, vocab, i2w, show_attention=True)
```
Notice how the attention weights show how important different parts of the input are for generating different tokens in the output. For tasks like machine translation, where the order of one-to-one words often changes due to grammatical differences between languages, this becomes very interesting as we see the attention window move further away from the diagonal that is mostly displayed in grapheme-to-phoneme translations.
**What's next**
With the above model, you have the basics for training a powerful sequence-to-sequence model with attention in a number of distinct domains. The only major changes required are preparing a dataset with pairs input and output sequences and in general the rest of the building blocks will remain the same. Good luck, and have fun!
| github_jupyter |
# Feature importance
In this notebook, we will detail methods to investigate the importance of
features used by a given model. We will look at:
1. interpreting the coefficients in a linear model;
2. the attribute `feature_importances_` in RandomForest;
3. `permutation feature importance`, which is an inspection technique that
can be used for any fitted model.
## 0. Presentation of the dataset
This dataset is a record of neighborhoods in California district, predicting
the **median house value** (target) given some information about the
neighborhoods, as the average number of rooms, the latitude, the longitude or
the median income of people in the neighborhoods (block).
```
from sklearn.datasets import fetch_california_housing
import pandas as pd
X, y = fetch_california_housing(as_frame=True, return_X_y=True)
```
To speed up the computation, we take the first 10000 samples
```
X = X[:10000]
y = y[:10000]
X.head()
```
The feature reads as follow:
- MedInc: median income in block
- HouseAge: median house age in block
- AveRooms: average number of rooms
- AveBedrms: average number of bedrooms
- Population: block population
- AveOccup: average house occupancy
- Latitude: house block latitude
- Longitude: house block longitude
- MedHouseVal: Median house value in 100k$ *(target)*
To assert the quality of our inspection technique, let's add some random
feature that won't help the prediction (un-informative feature)
```
import numpy as np
# Adding random features
rng = np.random.RandomState(0)
bin_var = pd.Series(rng.randint(0, 1, X.shape[0]), name='rnd_bin')
num_var = pd.Series(np.arange(X.shape[0]), name='rnd_num')
X_with_rnd_feat = pd.concat((X, bin_var, num_var), axis=1)
```
We will split the data into training and testing for the remaining part of
this notebook
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X_with_rnd_feat, y,
random_state=29)
```
Let's quickly inspect some features and the target
```
import seaborn as sns
train_dataset = X_train.copy()
train_dataset.insert(0, "MedHouseVal", y_train)
_ = sns.pairplot(
train_dataset[['MedHouseVal', 'Latitude', 'AveRooms', 'AveBedrms', 'MedInc']],
kind='reg', diag_kind='kde', plot_kws={'scatter_kws': {'alpha': 0.1}})
```
We see in the upper right plot that the median income seems to be positively
correlated to the median house price (the target).
We can also see that the average number of rooms `AveRooms` is very
correlated to the average number of bedrooms `AveBedrms`.
## 1. Linear model inspection
In linear models, the target value is modeled as a linear combination of the
features.
Coefficients represent the relationship between the given feature $X_i$ and
the target $y$, assuming that all the other features remain constant
(conditional dependence). This is different from plotting $X_i$ versus $y$
and fitting a linear relationship: in that case all possible values of the
other features are taken into account in the estimation (marginal
dependence).
```
from sklearn.linear_model import RidgeCV
model = RidgeCV()
model.fit(X_train, y_train)
print(f'model score on training data: {model.score(X_train, y_train)}')
print(f'model score on testing data: {model.score(X_test, y_test)}')
```
Our linear model obtains a $R^2$ score of .60, so it explains a significant
part of the target. Its coefficient should be somehow relevant. Let's look at
the coefficient learnt
```
import matplotlib.pyplot as plt
coefs = pd.DataFrame(
model.coef_,
columns=['Coefficients'], index=X_train.columns
)
coefs.plot(kind='barh', figsize=(9, 7))
plt.title('Ridge model')
plt.axvline(x=0, color='.5')
plt.subplots_adjust(left=.3)
```
### Sign of coefficients
```{admonition} A surprising association?
**Why is the coefficient associated to `AveRooms` negative?** Does the
price of houses decreases with the number of rooms?
```
The coefficients of a linear model are a *conditional* association:
they quantify the variation of a the output (the price) when the given
feature is varied, **keeping all other features constant**. We should
not interpret them as a *marginal* association, characterizing the link
between the two quantities ignoring all the rest.
The coefficient associated to `AveRooms` is negative because the number
of rooms is strongly correlated with the number of bedrooms,
`AveBedrms`. What we are seeing here is that for districts where the houses
have the same number of bedrooms on average, when there are more rooms
(hence non-bedroom rooms), the houses are worth comparatively less.
### Scale of coefficients
The `AveBedrms` have the higher coefficient. However, we can't compare the
magnitude of these coefficients directly, since they are not scaled. Indeed,
`Population` is an integer which can be thousands, while `AveBedrms` is
around 4 and Latitude is in degree.
So the Population coefficient is expressed in "$100k\\$$ / habitant" while the
AveBedrms is expressed in "$100k\\$$ / nb of bedrooms" and the Latitude
coefficient in "$100k\\$$ / degree".
We see that changing population by one does not change the outcome, while as
we go south (latitude increase) the price becomes cheaper. Also, adding a
bedroom (keeping all other feature constant) shall rise the price of the
house by 80k$.
So looking at the coefficient plot to gauge feature importance can be
misleading as some of them vary on a small scale, while others vary a lot
more, several decades.
This becomes visible if we compare the standard deviations of our different
features.
```
X_train.std(axis=0).plot(kind='barh', figsize=(9, 7))
plt.title('Features std. dev.')
plt.subplots_adjust(left=.3)
plt.xlim((0, 100))
```
So before any interpretation, we need to scale each column (removing the mean
and scaling the variance to 1).
```
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
model = make_pipeline(StandardScaler(), RidgeCV())
model.fit(X_train, y_train)
print(f'model score on training data: {model.score(X_train, y_train)}')
print(f'model score on testing data: {model.score(X_test, y_test)}')
coefs = pd.DataFrame(
model[1].coef_,
columns=['Coefficients'], index=X_train.columns
)
coefs.plot(kind='barh', figsize=(9, 7))
plt.title('Ridge model')
plt.axvline(x=0, color='.5')
plt.subplots_adjust(left=.3)
```
Now that the coefficients have been scaled, we can safely compare them.
The median income feature, with longitude and latitude are the three
variables that most influence the model.
The plot above tells us about dependencies between a specific feature and the
target when all other features remain constant, i.e., conditional
dependencies. An increase of the `HouseAge` will induce an increase of the
price when all other features remain constant. On the contrary, an increase
of the average rooms will induce an decrease of the price when all other
features remain constant.
### Checking the variability of the coefficients
We can check the coefficient variability through cross-validation: it is a
form of data perturbation.
If coefficients vary significantly when changing the input dataset their
robustness is not guaranteed, and they should probably be interpreted with
caution.
```
from sklearn.model_selection import cross_validate
from sklearn.model_selection import RepeatedKFold
cv_model = cross_validate(
model, X_with_rnd_feat, y, cv=RepeatedKFold(n_splits=5, n_repeats=5),
return_estimator=True, n_jobs=2
)
coefs = pd.DataFrame(
[model[1].coef_
for model in cv_model['estimator']],
columns=X_with_rnd_feat.columns
)
plt.figure(figsize=(9, 7))
sns.boxplot(data=coefs, orient='h', color='cyan', saturation=0.5)
plt.axvline(x=0, color='.5')
plt.xlabel('Coefficient importance')
plt.title('Coefficient importance and its variability')
plt.subplots_adjust(left=.3)
```
Every coefficient looks pretty stable, which mean that different Ridge model
put almost the same weight to the same feature.
### Linear models with sparse coefficients (Lasso)
In it important to keep in mind that the associations extracted depend
on the model. To illustrate this point we consider a Lasso model, that
performs feature selection with a L1 penalty. Let us fit a Lasso model
with a strong regularization parameters `alpha`
```
from sklearn.linear_model import Lasso
model = make_pipeline(StandardScaler(), Lasso(alpha=.015))
model.fit(X_train, y_train)
print(f'model score on training data: {model.score(X_train, y_train)}')
print(f'model score on testing data: {model.score(X_test, y_test)}')
coefs = pd.DataFrame(
model[1].coef_,
columns=['Coefficients'], index=X_train.columns
)
coefs.plot(kind='barh', figsize=(9, 7))
plt.title('Lasso model, strong regularization')
plt.axvline(x=0, color='.5')
plt.subplots_adjust(left=.3)
```
Here the model score is a bit lower, because of the strong regularization.
However, it has zeroed out 3 coefficients, selecting a small number of
variables to make its prediction.
We can see that out of the two correlated features `AveRooms` and
`AveBedrms`, the model has selected one. Note that this choice is
partly arbitrary: choosing one does not mean that the other is not
important for prediction. **Avoid over-interpreting models, as they are
imperfect**.
As above, we can look at the variability of the coefficients:
```
cv_model = cross_validate(
model, X_with_rnd_feat, y, cv=RepeatedKFold(n_splits=5, n_repeats=5),
return_estimator=True, n_jobs=2
)
coefs = pd.DataFrame(
[model[1].coef_
for model in cv_model['estimator']],
columns=X_with_rnd_feat.columns
)
plt.figure(figsize=(9, 7))
sns.boxplot(data=coefs, orient='h', color='cyan', saturation=0.5)
plt.axvline(x=0, color='.5')
plt.xlabel('Coefficient importance')
plt.title('Coefficient importance and its variability')
plt.subplots_adjust(left=.3)
```
We can see that both the coefficients associated to `AveRooms` and
`AveBedrms` have a strong variability and that they can both be non
zero. Given that they are strongly correlated, the model can pick one
or the other to predict well. This choice is a bit arbitrary, and must
not be over-interpreted.
### Lessons learned
Coefficients must be scaled to the same unit of measure to retrieve feature
importance, or comparing them.
Coefficients in multivariate linear models represent the dependency between a
given feature and the target, conditional on the other features.
Correlated features might induce instabilities in the coefficients of linear
models and their effects cannot be well teased apart.
Inspecting coefficients across the folds of a cross-validation loop gives an
idea of their stability.
## 2. RandomForest `feature_importances_`
On some algorithms, there are some feature importance methods,
inherently built within the model. It is the case in RandomForest models.
Let's investigate the built-in `feature_importances_` attribute.
```
from sklearn.ensemble import RandomForestRegressor
model = RandomForestRegressor()
model.fit(X_train, y_train)
print(f'model score on training data: {model.score(X_train, y_train)}')
print(f'model score on testing data: {model.score(X_test, y_test)}')
```
Contrary to the testing set, the score on the training set is almost perfect,
which means that our model is overfitting here.
```
importances = model.feature_importances_
```
The importance of a feature is basically: how much this feature is used in
each tree of the forest. Formally, it is computed as the (normalized) total
reduction of the criterion brought by that feature.
```
indices = np.argsort(importances)
fig, ax = plt.subplots()
ax.barh(range(len(importances)), importances[indices])
ax.set_yticks(range(len(importances)))
_ = ax.set_yticklabels(np.array(X_train.columns)[indices])
```
Median income is still the most important feature.
It also has a small bias toward high cardinality features, such as the noisy
feature `rnd_num`, which are here predicted having .07 importance, more than
`HouseAge` (which has low cardinality).
## 3. Feature importance by permutation
We introduce here a new technique to evaluate the feature importance of any
given fitted model. It basically shuffles a feature and sees how the model
changes its prediction. Thus, the change in prediction will correspond to
the feature importance.
```
# Any model could be used here
model = RandomForestRegressor()
# model = make_pipeline(StandardScaler(),
# RidgeCV())
model.fit(X_train, y_train)
print(f'model score on training data: {model.score(X_train, y_train)}')
print(f'model score on testing data: {model.score(X_test, y_test)}')
```
As the model gives a good prediction, it has captured well the link
between X and y. Hence, it is reasonable to interpret what it has
captured from the data.
### Feature importance
Lets compute the feature importance for a given feature, say the `MedInc`
feature.
For that, we will shuffle this specific feature, keeping the other feature as
is, and run our same model (already fitted) to predict the outcome. The
decrease of the score shall indicate how the model had used this feature to
predict the target. The permutation feature importance is defined to be the
decrease in a model score when a single feature value is randomly shuffled.
For instance, if the feature is crucial for the model, the outcome would also
be permuted (just as the feature), thus the score would be close to zero.
Afterward, the feature importance is the decrease in score. So in that case,
the feature importance would be close to the score.
On the contrary, if the feature is not used by the model, the score shall
remain the same, thus the feature importance will be close to 0.
```
def get_score_after_permutation(model, X, y, curr_feat):
""" return the score of model when curr_feat is permuted """
X_permuted = X.copy()
col_idx = list(X.columns).index(curr_feat)
# permute one column
X_permuted.iloc[:, col_idx] = np.random.permutation(
X_permuted[curr_feat].values)
permuted_score = model.score(X_permuted, y)
return permuted_score
def get_feature_importance(model, X, y, curr_feat):
""" compare the score when curr_feat is permuted """
baseline_score_train = model.score(X, y)
permuted_score_train = get_score_after_permutation(model, X, y, curr_feat)
# feature importance is the difference between the two scores
feature_importance = baseline_score_train - permuted_score_train
return feature_importance
curr_feat = 'MedInc'
feature_importance = get_feature_importance(model, X_train, y_train, curr_feat)
print(f'feature importance of "{curr_feat}" on train set is '
f'{feature_importance:.3}')
```
Since there is some randomness, it is advisable to run it multiple times and
inspect the mean and the standard deviation of the feature importance.
```
n_repeats = 10
list_feature_importance = []
for n_round in range(n_repeats):
list_feature_importance.append(
get_feature_importance(model, X_train, y_train, curr_feat))
print(
f'feature importance of "{curr_feat}" on train set is '
f'{np.mean(list_feature_importance):.3} '
f'+/- {np.std(list_feature_importance):.3}')
```
0.67 over 0.98 is very relevant (note the $R^2$ score could go below 0). So
we can imagine our model relies heavily on this feature to predict the class.
We can now compute the feature permutation importance for all the features.
```
def permutation_importance(model, X, y, n_repeats=10):
"""Calculate importance score for each feature."""
importances = []
for curr_feat in X.columns:
list_feature_importance = []
for n_round in range(n_repeats):
list_feature_importance.append(
get_feature_importance(model, X, y, curr_feat))
importances.append(list_feature_importance)
return {'importances_mean': np.mean(importances, axis=1),
'importances_std': np.std(importances, axis=1),
'importances': importances}
# This function could directly be access from sklearn
# from sklearn.inspection import permutation_importance
def plot_feature_importances(perm_importance_result, feat_name):
""" bar plot the feature importance """
fig, ax = plt.subplots()
indices = perm_importance_result['importances_mean'].argsort()
plt.barh(range(len(indices)),
perm_importance_result['importances_mean'][indices],
xerr=perm_importance_result['importances_std'][indices])
ax.set_yticks(range(len(indices)))
_ = ax.set_yticklabels(feat_name[indices])
```
Let's compute the feature importance by permutation on the training data.
```
perm_importance_result_train = permutation_importance(
model, X_train, y_train, n_repeats=10)
plot_feature_importances(perm_importance_result_train, X_train.columns)
```
We see again that the feature `MedInc`, `Latitude` and `Longitude` are very
important for the model.
We note that our random variable `rnd_num` is now very less important than
latitude. Indeed, the feature importance built-in in RandomForest has bias
for continuous data, such as `AveOccup` and `rnd_num`.
However, the model still uses these `rnd_num` feature to compute the output.
It is in line with the overfitting we had noticed between the train and test
score.
### discussion
1. For correlated feature, the permutation could give non realistic sample
(e.g. nb of bedrooms higher than the number of rooms)
2. It is unclear whether you should use training or testing data to compute
the feature importance.
3. Note that dropping a column and fitting a new model will not allow to
analyse the feature importance for a specific model, since a *new* model
will be fitted.
# Take Away
* One could directly interpret the coefficient in linear model (if the
feature have been scaled first)
* Model like RandomForest have built-in feature importance
* `permutation_importance` gives feature importance by permutation for any
fitted model
| github_jupyter |
```
import torch
import os
import cv2
import torch.nn as nn
import torch.nn.functional as F
import numpy as np
from torchvision import transforms
transform_data = transforms.Compose(
[
transforms.ToPILImage(),
transforms.RandomVerticalFlip(),
transforms.RandomHorizontalFlip(),
transforms.RandomCrop((112)),
transforms.ToTensor(),
]
)
def load_data(img_size=112):
data = []
labels = {}
index = -1
for label in os.listdir('./data/'):
index += 1
labels[label] = index
print(len(labels))
X = []
y = []
for label in labels:
for file in os.listdir(f'./data/{label}/'):
path = f'./data/{label}/{file}'
img = cv2.imread(path)
img = cv2.resize(img,(img_size,img_size))
data.append([np.array(transform_data(np.array(img))),labels[label]])
X.append(np.array(transform_data(np.array(img))))
y.append(labels[label])
np.random.shuffle(data)
np.random.shuffle(data)
np.random.shuffle(data)
np.random.shuffle(data)
np.random.shuffle(data)
np.save('./data.npy',data)
VAL_SPLIT = 0.25
VAL_SPLIT = len(X)*VAL_SPLIT
VAL_SPLIT = int(VAL_SPLIT)
X_train = X[:-VAL_SPLIT]
y_train = y[:-VAL_SPLIT]
X_test = X[-VAL_SPLIT:]
y_test = y[-VAL_SPLIT:]
X = torch.from_numpy(np.array(X))
y = torch.from_numpy(np.array(y))
X_train = np.array(X_train)
X_test = np.array(X_test)
y_train = np.array(y_train)
y_test = np.array(y_test)
X_train = torch.from_numpy(X_train)
X_test = torch.from_numpy(X_test)
y_train = torch.from_numpy(y_train)
y_test = torch.from_numpy(y_test)
return X,y,X_train,X_test,y_train,y_test
X,y,X_train,X_test,y_train,y_test = load_data()
import torch.nn as nn
import torch.nn.functional as F
class BaseLine(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(3,32,5)
self.conv2 = nn.Conv2d(32,64,5)
self.conv2batchnorm = nn.BatchNorm2d(64)
self.conv3 = nn.Conv2d(64,128,5)
self.fc1 = nn.Linear(128*10*10,256)
self.fc2 = nn.Linear(256,128)
self.fc3 = nn.Linear(128,50)
self.relu = nn.ReLU()
def forward(self,X):
preds = F.max_pool2d(self.relu(self.conv1(X)),(2,2))
preds = F.max_pool2d(self.relu(self.conv2batchnorm(self.conv2(preds))),(2,2))
preds = F.max_pool2d(self.relu(self.conv3(preds)),(2,2))
preds = preds.view(-1,128*10*10)
preds = self.relu(self.fc1(preds))
preds = self.relu(self.fc2(preds))
preds = self.relu(self.fc3(preds))
return preds
device = torch.device('cuda')
from torchvision import models
# model = BaseLine().to(device)
# model = model.to(device)
model = models.resnet18(pretrained=True).to(device)
in_f = model.fc.in_features
model.fc = nn.Linear(in_f,50)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(),lr=0.1)
PROJECT_NAME = 'Car-Brands-Images-Clf'
import wandb
EPOCHS = 100
BATCH_SIZE = 32
from tqdm import tqdm
# wandb.init(project=PROJECT_NAME,name='transfer-learning')
# for _ in tqdm(range(EPOCHS)):
# for i in range(0,len(X_train),BATCH_SIZE):
# X_batch = X_train[i:i+BATCH_SIZE].view(-1,3,112,112).to(device)
# y_batch = y_train[i:i+BATCH_SIZE].to(device)
# model.to(device)
# preds = model(X_batch)
# preds = preds.to(device)
# loss = criterion(preds,y_batch)
# optimizer.zero_grad()
# loss.backward()
# optimizer.step()
# wandb.log({'loss':loss.item()})
# TL vs Custom Model best = TL
def get_loss(criterion,y,model,X):
model.to('cuda')
preds = model(X.view(-1,3,112,112).to('cuda').float())
preds.to('cuda')
loss = criterion(preds,torch.tensor(y,dtype=torch.long).to('cuda'))
loss.backward()
return loss.item()
def test(net,X,y):
device = 'cuda'
net.to(device)
correct = 0
total = 0
net.eval()
with torch.no_grad():
for i in range(len(X)):
real_class = torch.argmax(y[i]).to(device)
net_out = net(X[i].view(-1,3,112,112).to(device).float())
net_out = net_out[0]
predictied_class = torch.argmax(net_out)
if predictied_class == real_class:
correct += 1
total += 1
net.train()
net.to('cuda')
return round(correct/total,3)
import torch
import os
import cv2
import torch.nn as nn
import torch.nn.functional as F
import numpy as np
from torchvision import transforms
transform_data = transforms.Compose(
[
transforms.ToPILImage(),
transforms.RandomVerticalFlip(),
transforms.RandomHorizontalFlip(),
transforms.RandomCrop((112)),
transforms.ToTensor(),
]
)
def load_data(img_size=112):
data = []
labels = {}
index = -1
for label in os.listdir('./data/'):
index += 1
labels[label] = index
print(len(labels))
X = []
y = []
for label in labels:
for file in os.listdir(f'./data/{label}/'):
path = f'./data/{label}/{file}'
img = cv2.imread(path)
img = cv2.resize(img,(img_size,img_size))
data.append([np.array(transform_data(np.array(img))),labels[label]])
X.append(np.array(transform_data(np.array(img))))
y.append(labels[label])
np.random.shuffle(data)
np.random.shuffle(data)
np.random.shuffle(data)
np.random.shuffle(data)
np.random.shuffle(data)
np.save('./data.npy',data)
VAL_SPLIT = 0.25
VAL_SPLIT = len(X)*VAL_SPLIT
VAL_SPLIT = int(VAL_SPLIT)
X_train = X[:-VAL_SPLIT]
y_train = y[:-VAL_SPLIT]
X_test = X[-VAL_SPLIT:]
y_test = y[-VAL_SPLIT:]
X = torch.from_numpy(np.array(X))
y = torch.from_numpy(np.array(y))
X_train = np.array(X_train)
X_test = np.array(X_test)
y_train = np.array(y_train)
y_test = np.array(y_test)
X_train = torch.from_numpy(X_train)
X_test = torch.from_numpy(X_test)
y_train = torch.from_numpy(y_train)
y_test = torch.from_numpy(y_test)
return X,y,X_train,X_test,y_train,y_test
X,y,X_train,X_test,y_train,y_test = load_data()
import torch.nn as nn
import torch.nn.functional as F
class BaseLine(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(3,32,5)
self.conv2 = nn.Conv2d(32,64,5)
self.conv2batchnorm = nn.BatchNorm2d(64)
self.conv3 = nn.Conv2d(64,128,5)
self.fc1 = nn.Linear(128*10*10,256)
self.fc2 = nn.Linear(256,128)
self.fc3 = nn.Linear(128,50)
self.relu = nn.ReLU()
def forward(self,X):
preds = F.max_pool2d(self.relu(self.conv1(X)),(2,2))
preds = F.max_pool2d(self.relu(self.conv2batchnorm(self.conv2(preds))),(2,2))
preds = F.max_pool2d(self.relu(self.conv3(preds)),(2,2))
preds = preds.view(-1,128*10*10)
preds = self.relu(self.fc1(preds))
preds = self.relu(self.fc2(preds))
preds = self.relu(self.fc3(preds))
return preds
device = torch.device('cuda')
from torchvision import models
# model = BaseLine().to(device)
# model = model.to(device)
model = models.resnet18(pretrained=True).to(device)
in_f = model.fc.in_features
model.fc = nn.Linear(in_f,50)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(),lr=0.1)
PROJECT_NAME = 'Car-Brands-Images-Clf'
import wandb
EPOCHS = 100
BATCH_SIZE = 32
from tqdm import tqdm
def get_loss(criterion,y,model,X):
model.to('cuda')
preds = model(X.view(-1,3,112,112).to('cuda').float())
preds.to('cuda')
loss = criterion(preds,torch.tensor(y,dtype=torch.long).to('cuda'))
loss.backward()
return loss.item()
def test(net,X,y):
device = 'cuda'
net.to(device)
correct = 0
total = 0
net.eval()
with torch.no_grad():
for i in range(len(X)):
real_class = torch.argmax(y[i]).to(device)
net_out = net(X[i].view(-1,3,112,112).to(device).float())
net_out = net_out[0]
predictied_class = torch.argmax(net_out)
if predictied_class == real_class:
correct += 1
total += 1
net.train()
net.to('cuda')
return round(correct/total,3)
wandb.init(project=PROJECT_NAME,name='transfer-learning')
for _ in tqdm(range(EPOCHS)):
for i in range(0,len(X_train),BATCH_SIZE):
X_batch = X_train[i:i+BATCH_SIZE].view(-1,3,112,112).to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model.to(device)
preds = model(X_batch)
preds = preds.to(device)
loss = criterion(preds,y_batch)
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item(),'val_loss':get_loss(criterion,y_test,model,X_test),'accuracy':test(model,X_train,y_train),'val_accuracy':test(model,X_test,y_test)})
# wandb.init(project=PROJECT_NAME,name='transfer-learning')
# for _ in tqdm(range(EPOCHS)):
# for i in range(0,len(X_train),BATCH_SIZE):
# X_batch = X_train[i:i+BATCH_SIZE].view(-1,3,112,112).to(device)
# y_batch = y_train[i:i+BATCH_SIZE].to(device)
# model.to(device)
# preds = model(X_batch)
# preds = preds.to(device)
# loss = criterion(preds,y_batch)
# optimizer.zero_grad()
# loss.backward()
# optimizer.step()
# wandb.log({'loss':loss.item(),'val_loss':get_loss(criterion,y_test,model,X_test),'accuracy':test(model,X_train,y_train),'val_accuracy':test(model,X_test,y_test)})
# TL vs Custom Model best = TL
EPOCHS = 100
BATCH_SIZE = 32
model = models.mobilenet_v3_large(pretrained=False, num_classes=50).to(device)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(),lr=0.1)
wandb.init(project=PROJECT_NAME,name=f'models.mobilenet_v3_large')
for _ in tqdm(range(EPOCHS),leave=False):
for i in tqdm(range(0,len(X_train),BATCH_SIZE),leave=False):
X_batch = X_train[i:i+BATCH_SIZE].view(-1,3,112,112).to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model.to(device)
preds = model(X_batch)
preds = preds.to(device)
loss = criterion(preds,y_batch)
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item(),'val_loss':get_loss(criterion,y_test,model,X_test),'accuracy':test(model,X_train,y_train),'val_accuracy':test(model,X_test,y_test)})
```
| github_jupyter |
```
# Console related imports.
from subprocess import Popen, PIPE
import os
from IPython.utils.py3compat import bytes_to_str, string_types
# Widget related imports.
from IPython.html import widgets
from IPython.display import display
```
Define function to run a process without blocking the input.
```
def read_process(process, append_output):
""" Try to read the stdout and stderr of a process and render it using
the append_output method provided
Parameters
----------
process: Popen handle
append_output: method handle
Callback to render output. Signature of
append_output(output, [prefix=])"""
try:
stdout = process.stdout.read()
if stdout is not None and len(stdout) > 0:
append_output(stdout, prefix=' ')
except:
pass
try:
stderr = process.stderr.read()
if stderr is not None and len(stderr) > 0:
append_output(stderr, prefix='ERR ')
except:
pass
def set_pipe_nonblocking(pipe):
"""Set a pipe as non-blocking"""
try:
import fcntl
fl = fcntl.fcntl(pipe, fcntl.F_GETFL)
fcntl.fcntl(pipe, fcntl.F_SETFL, fl | os.O_NONBLOCK)
except:
pass
kernel = get_ipython().kernel
def run_command(command, append_output, has_user_exited=None):
"""Run a command asyncronously
Parameters
----------
command: str
Shell command to launch a process with.
append_output: method handle
Callback to render output. Signature of
append_output(output, [prefix=])
has_user_exited: method handle
Check to see if the user wants to stop the command.
Must return a boolean."""
# Echo input.
append_output(command, prefix='>>> ')
# Create the process. Make sure the pipes are set as non-blocking.
process = Popen(command, shell=True, stdout=PIPE, stderr=PIPE)
set_pipe_nonblocking(process.stdout)
set_pipe_nonblocking(process.stderr)
# Only continue to read from the command
while (has_user_exited is None or not has_user_exited()) and process.poll() is None:
read_process(process, append_output)
kernel.do_one_iteration() # Run IPython iteration. This is the code that
# makes this operation non-blocking. This will
# allow widget messages and callbacks to be
# processed.
# If the process is still running, the user must have exited.
if process.poll() is None:
process.kill()
else:
read_process(process, append_output) # Read remainer
```
Create the console widgets without displaying them.
```
console_container = widgets.VBox(visible=False)
console_container.padding = '10px'
output_box = widgets.Textarea()
output_box.height = '400px'
output_box.font_family = 'monospace'
output_box.color = '#AAAAAA'
output_box.background_color = 'black'
output_box.width = '800px'
input_box = widgets.Text()
input_box.font_family = 'monospace'
input_box.color = '#AAAAAA'
input_box.background_color = 'black'
input_box.width = '800px'
console_container.children = [output_box, input_box]
```
Hook the process execution methods up to our console widgets.
```
def append_output(output, prefix):
if isinstance(output, string_types):
output_str = output
else:
output_str = bytes_to_str(output)
output_lines = output_str.split('\n')
formatted_output = '\n'.join([prefix + line for line in output_lines if len(line) > 0]) + '\n'
output_box.value += formatted_output
output_box.scroll_to_bottom()
def has_user_exited():
return not console_container.visible
def handle_input(sender):
sender.disabled = True
try:
command = sender.value
sender.value = ''
run_command(command, append_output=append_output, has_user_exited=has_user_exited)
finally:
sender.disabled = False
input_box.on_submit(handle_input)
```
Create the button that will be used to display and hide the console. Display both the console container and the new button used to toggle it.
```
toggle_button = widgets.Button(description="Start Console")
def toggle_console(sender):
console_container.visible = not console_container.visible
if console_container.visible:
toggle_button.description="Stop Console"
input_box.disabled = False
else:
toggle_button.description="Start Console"
toggle_button.on_click(toggle_console)
display(toggle_button)
display(console_container)
```
| github_jupyter |
# Q3
In this question, you'll be doing some error handling. There's only one function to finish coding below--the logger function--but there's a LOT of other code that's already written for you! This will start introducing you to working with existing codebases.
### A
In this question, you'll write a logger. This is a critical component of any full-scale application: when something goes wrong, and you start the debugging process, it helps immensely to have logs of activity of your application to see where it started going off the rails.
The log will take the form of a dictionary. The keys of the dictionary will be the name of the method called, and the value will be the integer code of the method's output. OR, in the case of an exception, the integer code of the exception. You can access this code in the `except` block:
```
try:
some_code
except Exception as e:
e.value
```
You'll need to save this `e.value` in your log dictionary for the output of the function call.
Call all three of the functions below:
- unreliable1
- unreliable2
- unreliable3
and record their outputs in the dictionary.
In summary, these are your goals:
1. In logger(), call each of the three methods once: unreliable1(), unreliable2(), unreliable3()
2. When calling each method, wrap them in try / except blocks so no exceptions escape to crash the program.
3. If an exception occurs, record its value to the dictionary for that function.
4. If NO exception occurs, and the function completes, it will return a value. Record that value in the dictionary for that function.
```
"""
The below classes--Error1, Error2, and Error3--are the names of the possible
errors that can be raised, and which you'll have to protect against.
"""
import random
class Error1(Exception):
def __init__(self, value):
self.value = value
def __str__(self):
return repr(self.value)
class Error2(Exception):
def __init__(self, value):
self.value = value
def __str__(self):
return repr(self.value)
class Error3(Exception):
def __init__(self, value):
self.value = value
def __str__(self):
return repr(self.value)
"""
The following three functions do SUPER IMPORTANT THINGS!
However, they are very prone to throwing exceptions, so when
you call them, you'll need to properly protect the calls!
"""
def unreliable1():
for i in range(1, 4):
if random.randint(0, 1) == 1:
raise Error1(i)
return random.randint(10, 1000000)
def unreliable2():
for i in range(4, 7):
if random.randint(0, 1) == 1:
raise Error2(i)
return random.randint(10, 1000000)
def unreliable3():
for i in range(7, 10):
if random.randint(0, 1) == 1:
raise Error3(i)
return random.randint(10, 1000000)
###
# Finish this method!
###
def logger():
logs = {
'unreliable1': 0,
'unreliable2': 0,
'unreliable3': 0,
}
### BEGIN SOLUTION
### END SOLUTION
return logs
random.seed(10)
logs = logger()
assert logs['unreliable1'] == 2
assert logs['unreliable2'] == 4
assert logs['unreliable3'] == 9
random.seed(381734)
logs = logger()
assert logs['unreliable1'] == 1
assert logs['unreliable2'] == 5
assert logs['unreliable3'] == 7
random.seed(895572877675748)
logs = logger()
assert logs['unreliable1'] == 1
assert logs['unreliable2'] == 297436
assert logs['unreliable3'] == 8
```
| github_jupyter |
# SVM with sigmoid kernel
The goal of this notebook is to find the best parameters for polynomial kernel. We also want to check if the parameters depend on stock.
We will use [sklearn.svm](http://scikit-learn.org/stable/modules/generated/sklearn.svm.SVC.html#sklearn.svm.SVC) library to perform calculations. We want to pick the best parameters for **SVC**:
* C (default 1.0)
* gamma (default 1/number_of_features, so 1 in our case)
* coef0 (default 0.0)
Sigmoid kernel function: $(\tanh(\gamma \langle x,x'\rangle + coef0))$
```
%matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.dates as md
from statsmodels.distributions.empirical_distribution import ECDF
import numpy as np
import seaborn as sns
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
from sklearn.metrics import classification_report
from sklearn import svm
import warnings
from lob_data_utils import lob
sns.set_style('whitegrid')
warnings.filterwarnings('ignore')
```
# Data
We use data from 5 stocks (from dates 2013-09-01 - 2013-11-16) for which logistic regression yielded the best results.
We selected 3 subsets for each stock:
* training set (60% of data)
* test set (20% of data)
* cross-validation set (20% of data)
```
stocks = ['11234']
dfs = {}
dfs_cv = {}
dfs_test = {}
for s in stocks:
df, df_cv, df_test = lob.load_prepared_data(s, cv=True, length=6416)
dfs[s] = df
dfs_test[s] = df_cv
dfs_cv[s] = df_test
dfs[stocks[0]].head()
def svm_classification(d, kernel, gamma='auto', C=1.0, degree=3, coef0=0.0):
clf = svm.SVC(kernel=kernel, gamma=gamma, C=C, degree=degree, coef0=coef0)
X = d['queue_imbalance'].values.reshape(-1, 1)
y = d['mid_price_indicator'].values.reshape(-1, 1)
clf.fit(X, y)
return clf
```
# Methodology
We will use at first naive approach to grasp how each of the parameter influences the ROC area score and what values make sense, when the other parameters are set to defaults for parameters C and gamma.
Next we will choose parameter coef0 for SVM with gamma and C set to the best values we got.
### C parameter
The C parameter has influence over margin picked by SVM:
* for large values of **C** SVM will choose a smaller-margin hyperplane, which means that more data points will be classified correctly
* for small values of **C** SVM will choose a bigger-margin hyperplane, so there may be more misclassifications
At first we tried parameters: [0.0001, 0.001, 0.01, 0.1, 1, 10, 1000], but after first calculations it seems that it wasn't enough, so a few more values were introduced or removed.
```
cs = [0.01, 0.09, 0.1, 0.102, 0.105, 0.107, 0.11, 1.5]
df_css = {}
ax = plt.subplot()
ax.set_xscale("log", basex=10)
for s in stocks:
df_cs = pd.DataFrame(index=cs)
df_cs['roc'] = np.zeros(len(df_cs))
for c in cs:
reg_svm = svm_classification(dfs[s], 'sigmoid', C=c, coef0=0)
pred_svm_out_of_sample = reg_svm.predict(dfs_cv[s]['queue_imbalance'].values.reshape(-1, 1))
logit_roc_auc = roc_auc_score(dfs_cv[s]['mid_price_indicator'], pred_svm_out_of_sample)
df_cs.loc[c] = logit_roc_auc
plt.plot(df_cs, linestyle='--', label=s, marker='x', alpha=0.5)
df_css[s] = df_cs
plt.legend()
```
##### Best values of C parameter
Choice of parameter C should be small - less than 0.1
```
for s in stocks:
idx = df_css[s]['roc'].idxmax()
print('For {} the best is {}'.format(s, idx))
```
##### Influence of C parameter
The score difference between SVM with the worst choice of parameter **C** and the best choice one is shown on the output below.
```
for s in stocks:
err_max = df_css[s]['roc'].max()
err_min = df_css[s]['roc'].min()
print('For {} the diff between best and worst {}'.format(s, err_max - err_min))
```
### Gamma
Gamma is a parameter which has influence over decision region - the bigger it is, the bigger influence every single row of data has. When gamma is low the decision region is very broad. When gamma is high it can even create islands of decision-boundaries around data points.
```
gammas = [0.5, 0.85, 0.87, 0.9, 0.92, 0.93, 0.94, 0.95, 0.96, 0.97]
df_gammas = {}
ax = plt.subplot()
ax.set_xscale("log", basex=10)
for s in stocks:
df_gamma = pd.DataFrame(index=gammas)
df_gamma['roc'] = np.zeros(len(df_gamma))
for g in gammas:
reg_svm = svm_classification(dfs[s], 'sigmoid', gamma=g, coef0=0)
pred_svm_out_of_sample = reg_svm.predict(dfs_cv[s]['queue_imbalance'].values.reshape(-1, 1))
logit_roc_auc = roc_auc_score(dfs_cv[s]['mid_price_indicator'], pred_svm_out_of_sample)
df_gamma.loc[g] = logit_roc_auc
plt.plot(df_gamma, linestyle='--', label=s, marker='x', alpha=0.5)
df_gammas[s] = df_gamma
plt.legend()
```
##### Best values of gamma
There is no rule, how to set this parameter.
```
for s in stocks:
idx = df_gammas[s]['roc'].idxmax()
print('For {} the best is {}'.format(s, idx))
```
##### Influence of gamma
The score difference between SVM with the worst choice of **gamma** and the best choice one is shown on the output below. For scoring method we used *roc_area*. For stocks **10795**, **11618** and **4481** the difference is more than 0.1, so it's definitelly worth to experiment more.
```
for s in stocks:
err_max = df_gammas[s]['roc'].max()
err_min = df_gammas[s]['roc'].min()
print('For {} the diff between best and worst {}'.format(s, err_max - err_min))
df_params = {}
for s in stocks:
print(s)
params = []
for c in cs:
for g in gammas:
reg_svm = svm_classification(dfs[s], 'sigmoid', C=c, gamma=g)
prediction = reg_svm.predict(dfs_cv[s]['queue_imbalance'].values.reshape(-1, 1))
score = roc_auc_score(dfs_cv[s]['mid_price_indicator'], prediction)
params.append({'score': score, 'gamma': g, 'c': c})
df_params[s] = pd.DataFrame(params)
for s in stocks:
df_g = df_params[s].pivot(index='c', columns='gamma', values='score')
sns.heatmap(df_g)
plt.title('Best params for ' + s)
plt.figure()
for s in stocks:
print(s, df_params[s].iloc[df_params[s]['score'].idxmax()])
df_params[stocks[0]].sort_values(by='score', ascending=False).head(5)
```
### Coef0
For sigmoid kernel we use function:
$(\tanh(\gamma \langle x,x'\rangle + r))$, where r is specified by coef0.
```
coeffs = [0, 0.0000001, 0.000001, 0.00009, 0.0001, 0.0002]
# 10, 1
df_coefs = {}
ax = plt.subplot()
ax.set_xscale("log", basex=10)
for s in stocks:
df_coef = pd.DataFrame(index=cs)
df_coef['roc'] = np.zeros(len(df_coef))
for c in coeffs:
best_idx = df_params[s]['score'].idxmax()
reg_svm = svm_classification(dfs[s], 'sigmoid', C=df_params[s].iloc[best_idx]['c'],
gamma=df_params[s].iloc[best_idx]['gamma'], coef0=c)
pred_svm_out_of_sample = reg_svm.predict(dfs_cv[s]['queue_imbalance'].values.reshape(-1, 1))
logit_roc_auc = roc_auc_score(dfs_cv[s]['mid_price_indicator'], pred_svm_out_of_sample)
df_coef.loc[c] = logit_roc_auc
plt.plot(df_coef, linestyle='--', label=s, marker='x', alpha=0.5)
df_coefs[s] = df_coef
plt.legend()
```
##### Best values of coef0
The value of coef0 should be rather small - less than 0.5.
```
for s in stocks:
idx = df_coefs[s]['roc'].idxmax()
print('For {} the best is {}'.format(s, idx))
```
##### Influence of coef0
For all stocks choice of coef0 is significant, it can make even 0.5 difference.
```
for s in stocks:
err_max = df_coefs[s]['roc'].max()
err_min = df_coefs[s]['roc'].min()
print('For {} the diff between best and worst {}'.format(s, err_max - err_min))
```
#### Best params so far
```
print(s, '\t', 'C', '\t', 'gamma', '\t', 'coef0')
for s in stocks:
print(s, '\t', df_css[s]['roc'].idxmax(), '\t', df_gammas[s]['roc'].idxmax(),
'\t', df_coefs[s]['roc'].idxmax())
df_params_coef = {}
for s in stocks:
print(s)
params = []
for idx, row in df_params[s].iterrows():
for coef in coeffs:
reg_svm = svm_classification(dfs[s], 'sigmoid', C=row['c'],
gamma=row['gamma'], coef0=coef)
prediction = reg_svm.predict(dfs_cv[s]['queue_imbalance'].values.reshape(-1, 1))
score = roc_auc_score(dfs_cv[s]['mid_price_indicator'], prediction)
params.append({'score': score, 'gamma': row['gamma'], 'c': row['c'], 'coef0': coef})
df_params_coef[s] = pd.DataFrame(params)
print(df_params_coef[s]['score'].idxmax())
df_params_coef[stocks[0]].sort_values(by='score',
ascending=False)[df_params_coef[
stocks[0]]['score']> 0.6]
```
# Results
We compare reults of SVMs with the best choices of parameter against the logistic regression and SVM with defaults.
We will use naive approach - for each stock we will just pick the best values we found in the previous section.
#### Naive approach
We pick the best **C** parameter and the best **gamma** separately from the results of [section above](#Methodology), which were obtained using cross-validation set. The **coef0** parameter "depend" of the choice of the rest of parameters.
For 3 stocks the results are quite good, the rest is very bad.
```
df_results = pd.DataFrame(index=stocks)
df_results['logistic'] = np.zeros(len(stocks))
df_results['sigmoid-tunned'] = np.zeros(len(stocks))
df_results['c-tunned'] = np.zeros(len(stocks))
df_results['coef-tunned'] = np.zeros(len(stocks))
df_results['gamma-tunned'] = np.zeros(len(stocks))
plt.subplot(121)
for s in stocks:
best_idx = df_params_coef[s]['score'].idxmax()
c = df_params_coef[s].iloc[best_idx]['c']
gamma = df_params_coef[s].iloc[best_idx]['gamma']
coef0 = df_params_coef[s].iloc[best_idx]['coef0']
print(c, gamma, coef0)
df_results['c-tunned'][s] = c
df_results['coef-tunned'][s] = coef0
df_results['gamma-tunned'][s] = gamma
reg_svm = svm_classification(
dfs[s], 'sigmoid', C=c, gamma=gamma, coef0=coef0)
roc_score = lob.plot_roc(dfs_test[s], reg_svm, stock=s, title='ROC for test set with the best params')
df_results['sigmoid-tunned'][s] = roc_score
plt.subplot(122)
for s in stocks:
reg_log = lob.logistic_regression(dfs[s], 0, len(dfs[s]))
roc_score = lob.plot_roc(dfs_test[s], reg_log, stock=s, title='ROC for test set with logistic')
df_results['logistic'][s] = roc_score
plt.subplots_adjust(left=0, wspace=0.1, top=1, right=2)
df_results
for s in stocks:
ddd = df_params_coef[s]
print(s)
print(ddd.iloc[ddd['score'].idxmax()])
print()
s = stocks[0]
reg_svm = svm_classification(dfs[s], 'sigmoid', C=0.005,
gamma=10, coef0=1)
prediction = reg_svm.predict(dfs_test[s]['queue_imbalance'].values.reshape(-1, 1))
score = roc_auc_score(dfs_test[s]['mid_price_indicator'], prediction)
score
```
# Conclusions
We didn't use valid grid approach for choosing the best parameters, so there is possibility that these parameters could be improved. For one stock **2051** we have significant improvement (even when we consider other results). For stock **11618** we have small improvement.
### Resources
1. [Queue Imbalance as a One-Tick-Ahead Price Predictor in a Limit Order Book](https://arxiv.org/abs/1512.03492) <a class="anchor-link" href="#1">¶</a>
| github_jupyter |
```
%matplotlib inline
import shapely.geometry, shapely.wkt
from shapely.geometry.point import Point
from shapely.geometry.polygon import LinearRing
from shapely.geometry.linestring import LineString
from shapely.geometry.polygon import Polygon
import shapely as sl
import fiona
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import pylab
from utils.shapely_plot import draw
pylab.rcParams['figure.figsize'] = (17.0, 15.0)
shp_path = r'../data/SurfaceHydrology/SurfaceHydrologyAustralia_major_MurrayDarling.shp'
lengths = []
with fiona.collection(shp_path, "r") as input:
for f in input:
geom = sl.geometry.shape(f['geometry'])
lengths.append(geom.length)
fig = plt.figure(figsize=(17, 17))
ax = plt.axes()
ax.set_aspect('equal', 'datalim')
with fiona.collection(shp_path, "r") as input:
for f in input:
geom = sl.geometry.shape(f['geometry'])
draw(geom, outline='#aaaaff', alpha=0.5)
plt.show()
np.logspace(0, 0.01, 50)
fig = plt.figure(figsize=(17, 2))
ax = plt.axes()
ax.set_xscale('log')
bins = 10 ** np.linspace(np.log10(0.0001), np.log10(0.5), 50)
_ = plt.hist(lengths, bins)
def pair(list):
'''Iterate over pairs in a list -> pair of points '''
for i in range(1, len(list)):
yield list[i-1], list[i]
for seg_start, seg_end in pair(geom.coords):
line_start = Point(seg_start)
line_end = Point(seg_end)
segment = LineString([line_start.coords[0],line_end.coords[0]])
print segment
# get first geom
shp_path = r'../data/SurfaceHydrology/SurfaceHydrologyAustralia_major_MurrayDarling.shp'
g = None
with fiona.collection(shp_path, "r") as input:
for f in input:
g = sl.geometry.shape(f['geometry'])
break
print(g.length)
g
def getDistance(coord1, coord2):
return Point(coord1).distance(Point(coord2))
def explodeLineString(lineString, segmentLength):
length = lineString.length
coords = lineString.coords
# first point
pt1 = coords[0]
ipt2 = 1
currentLength = 0.0
segments = []
segmentId = 0
while currentLength < length and ipt2 < len(coords):
segmentPoints = []
segmentPoints.append(pt1)
currentSegmentLength = 0.0
# print('ipt2:{0}, N:{1} Lc:{2}, L:{3}'
# .format(ipt2, len(coords), currentLength, length))
while currentSegmentLength < segmentLength:
if ipt2 == len(coords):
break
pt2 = coords[ipt2]
distance = getDistance(pt1, pt2)
# print('d:{0}, Lc: {1} Ls: {2} ipt2: {3}'
# .format(distance, currentSegmentLength, segmentLength, ipt2))
if currentSegmentLength + distance > segmentLength: # split
# hypotenuse lengths
h1 = segmentLength - currentSegmentLength # short
h2 = distance # long
# cathetus2
dx2 = pt2[0] - pt1[0]
dy2 = pt2[1] - pt1[1]
# cathetus1
ratio = h1 / h2
x1 = pt1[0] + dx2 * ratio
y1 = pt1[1] + dy2 * ratio
currentSegmentLength += h1
pt1 = [x1, y1]
segmentPoints.append(pt1)
else: # next
currentSegmentLength += distance
segmentPoints.append(pt2)
pt1 = pt2
ipt2 += 1
currentLength += currentSegmentLength
segmentId += 1
segments.append(LineString(segmentPoints))
return segments
print(getDistance(g.coords[0], g.coords[1]))
print(g.length)
segments = explodeLineString(g, 0.02)
len(segments)
shp_path = r'../data/SurfaceHydrology\SurfaceHydrologyAustralia_major_MurrayDarling.shp'
lengths_exploded = []
with fiona.collection(shp_path, "r") as input:
for f in input:
segments = explodeLineString(sl.geometry.shape(f['geometry']), 0.02)
for s in segments:
lengths_exploded.append(s.length)
print('Total length: ' + str(sum(lengths)))
print(len(lengths))
print(len(lengths_exploded))
f, (ax1, ax2) = plt.subplots(1,2, figsize=(20,5))
ax1.set_xlim(0, 0.5)
_ = ax1.hist(lengths, 100)
ax2.set_xlim(0, 0.01)
_ = ax2.hist(lengths_exploded, 100)
plt.show()
f, (ax1, ax2) = plt.subplots(1,2, figsize=(20,5))
bins = 10 ** np.linspace(np.log10(0.0001), np.log10(0.5), 50)
_ = ax1.hist(lengths, bins)
ax1.set_xscale('log')
bins = 10 ** np.linspace(np.log10(0.0001), np.log10(0.5), 50)
_ = ax2.hist(lengths_exploded, bins)
ax2.set_xscale('log')
plt.show()
fig = plt.figure(figsize=(17, 17))
ax = plt.axes()
ax.set_aspect('equal', 'datalim')
# original
# draw(g, outline='#000000')
cmap = matplotlib.colors.ListedColormap (np.random.rand ( 256,3))
# segments
for (i, s) in enumerate(segments):
draw(s, outline=cmap.colors[i])
plt.show()
from shapely.geometry import mapping
# take polylines and split them into segments of a given length
shp_path = r'../data/SurfaceHydrology/SurfaceHydrologyAustralia_major_MurrayDarling.shp'
shp_segments_path = r'../data/SurfaceHydrology/SurfaceHydrologyAustralia_major_MurrayDarling-segments.shp'
step = 0.02 # degrees
with fiona.collection(shp_path, "r") as input:
schema = {'geometry': 'LineString', 'properties': {'segment': 'int', 'aushydro_id': 'int'}}
# schema = input.schema
# schema['properties']['segment'] = 'int'
with fiona.open(shp_segments_path, 'w', 'ESRI Shapefile', schema) as output:
for f in input:
geom = sl.geometry.shape(f['geometry'])
segments = explodeLineString(geom, step)
for (i, s) in enumerate(segments):
properties = {'aushydro_id': f['properties']['aushydro_i']}
properties['segment'] = i
output.write({'geometry': mapping(s), 'properties': properties})
# properties = f['properties']
# properties['segment'] = i
# output.write({'geometry': mapping(s), 'properties': properties})
```
| github_jupyter |
# Stress Analysis in Social Media
Leverage the newly published and labelled reddit dataset for stress analysis to develop and improve supervised learning methods for identifying stress, both neural and traditional, and analyze the complexity and diversity of the data and characteristics of each category.
```
import os
import re
import pandas as pd
import numpy as np
import tensorflow as tf
from tensorflow import keras
import tensorflow_hub as hub
from datetime import datetime
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
print(tf.__version__)
import tensorflow as tf
from tensorflow.python.client import device_lib
print(device_lib.list_local_devices())
import bert
from bert import run_classifier
from bert import optimization
from bert import tokenization
import warnings
warnings.filterwarnings("ignore")
import logging, sys
logging.disable(sys.maxsize)
```
## Data
```
path = '../../data/preprocessed/'
# path = '/content/Insight_Stress_Analysis/data/'
train = pd.read_pickle(path + 'train.pkl')
test = pd.read_pickle(path + 'test.pkl')
DATA_COLUMN = 'text'
LABEL_COLUMN = 'label'
# label_list is the list of labels, i.e. True, False or 0, 1 or 'dog', 'cat'
label_list = [0, 1]
```
#Data Preprocessing
We'll need to transform our data into a format BERT understands. This involves two steps. First, we create `InputExample`'s using the constructor provided in the BERT library.
- `text_a` is the text we want to classify, which in this case, is the `Request` field in our Dataframe.
- `text_b` is used if we're training a model to understand the relationship between sentences (i.e. is `text_b` a translation of `text_a`? Is `text_b` an answer to the question asked by `text_a`?). This doesn't apply to our task, so we can leave `text_b` blank.
- `label` is the label for our example, i.e. True, False
```
# Use the InputExample class from BERT's run_classifier code to create examples from the data
train_InputExamples = train.apply(lambda x: bert.run_classifier.InputExample(guid=None, # Globally unique ID for bookkeeping, unused in this example
text_a = x[DATA_COLUMN],
text_b = None,
label = x[LABEL_COLUMN]), axis = 1)
test_InputExamples = test.apply(lambda x: bert.run_classifier.InputExample(guid=None,
text_a = x[DATA_COLUMN],
text_b = None,
label = x[LABEL_COLUMN]), axis = 1)
```
Next, we need to preprocess our data so that it matches the data BERT was trained on. For this, we'll need to do a couple of things (but don't worry--this is also included in the Python library):
1. Lowercase our text (if we're using a BERT lowercase model)
2. Tokenize it (i.e. "sally says hi" -> ["sally", "says", "hi"])
3. Break words into WordPieces (i.e. "calling" -> ["call", "##ing"])
4. Map our words to indexes using a vocab file that BERT provides
5. Add special "CLS" and "SEP" tokens (see the [readme](https://github.com/google-research/bert))
6. Append "index" and "segment" tokens to each input (see the [BERT paper](https://arxiv.org/pdf/1810.04805.pdf))
Happily, we don't have to worry about most of these details.
```
# Load a vocabulary file and lowercasing information directly from the BERT tf hub module
# This is a path to an uncased (all lowercase) version of BERT
BERT_MODEL_HUB = "https://tfhub.dev/google/bert_uncased_L-12_H-768_A-12/1"
def create_tokenizer_from_hub_module():
"""Get the vocab file and casing info from the Hub module."""
with tf.Graph().as_default():
bert_module = hub.Module(BERT_MODEL_HUB)
tokenization_info = bert_module(signature="tokenization_info", as_dict=True)
with tf.Session() as sess:
vocab_file, do_lower_case = sess.run([tokenization_info["vocab_file"],
tokenization_info["do_lower_case"]])
return bert.tokenization.FullTokenizer(vocab_file=vocab_file,
do_lower_case=do_lower_case)
tokenizer = create_tokenizer_from_hub_module()
# Set the maximum sequence length.
def get_max_len(text):
max_len = 0
for i in range(len(train)):
if len(text.iloc[i]) > max_len:
max_len = len(text.iloc[i])
return max_len
temp = train.text.str.split(' ')
max_len = get_max_len(temp)
MAX_SEQ_LENGTH = max_len
# Convert our train and test features to InputFeatures that BERT understands.
train_features = bert.run_classifier.convert_examples_to_features(train_InputExamples,
label_list,
MAX_SEQ_LENGTH,
tokenizer)
test_features = bert.run_classifier.convert_examples_to_features(test_InputExamples,
label_list,
MAX_SEQ_LENGTH,
tokenizer)
```
## Classification Model
Now that we've prepared our data, let's focus on building a model. `create_model` does just this below. First, it loads the BERT tf hub module again (this time to extract the computation graph). Next, it creates a single new layer that will be trained to adapt BERT to our sentiment task (i.e. classifying whether a movie review is positive or negative). This strategy of using a mostly trained model is called [fine-tuning](http://wiki.fast.ai/index.php/Fine_tuning).
```
def create_model(is_predicting, input_ids, input_mask, segment_ids, labels,
num_labels):
"""Creates a classification model."""
bert_module = hub.Module(BERT_MODEL_HUB, trainable=True)
bert_inputs = dict(input_ids=input_ids, input_mask=input_mask, segment_ids=segment_ids)
bert_outputs = bert_module(inputs=bert_inputs, signature="tokens", as_dict=True)
# Use "pooled_output" for classification tasks on an entire sentence.
# Use "sequence_outputs" for token-level output.
output_layer = bert_outputs["pooled_output"]
hidden_size = output_layer.shape[-1].value
# Create our own layer to tune for politeness data.
output_weights = tf.get_variable(
"output_weights", [num_labels, hidden_size],
initializer=tf.truncated_normal_initializer(stddev=0.02))
output_bias = tf.get_variable("output_bias",
[num_labels],
initializer=tf.zeros_initializer())
with tf.variable_scope("loss"):
# Dropout helps prevent overfitting
output_layer = tf.nn.dropout(output_layer, keep_prob=0.9)
logits = tf.matmul(output_layer, output_weights, transpose_b=True)
logits = tf.nn.bias_add(logits, output_bias)
log_probs = tf.nn.log_softmax(logits, axis=-1)
# Convert labels into one-hot encoding
one_hot_labels = tf.one_hot(labels, depth=num_labels, dtype=tf.float32)
predicted_labels = tf.squeeze(tf.argmax(log_probs, axis=-1, output_type=tf.int32))
# If we're predicting, we want predicted labels and the probabiltiies.
if is_predicting:
return (predicted_labels, log_probs)
# If we're train/eval, compute loss between predicted and actual label
per_example_loss = -tf.reduce_sum(one_hot_labels * log_probs, axis=-1)
loss = tf.reduce_mean(per_example_loss)
return (loss, predicted_labels, log_probs)
```
Next we'll wrap our model function in a `model_fn_builder` function that adapts our model to work for training, evaluation, and prediction.
```
# model_fn_builder actually creates our model function
# using the passed parameters for num_labels, learning_rate, etc.
def model_fn_builder(num_labels, learning_rate, num_train_steps, num_warmup_steps):
"""Returns `model_fn` closure for TPUEstimator."""
def model_fn(features, labels, mode, params): # pylint: disable=unused-argument
"""The `model_fn` for TPUEstimator."""
input_ids = features["input_ids"]
input_mask = features["input_mask"]
segment_ids = features["segment_ids"]
label_ids = features["label_ids"]
is_predicting = (mode == tf.estimator.ModeKeys.PREDICT)
# TRAIN and EVAL
if not is_predicting:
(loss, predicted_labels, log_probs) = create_model(is_predicting,
input_ids,
input_mask,
segment_ids,
label_ids,
num_labels)
train_op = bert.optimization.create_optimizer(loss,
learning_rate,
num_train_steps,
num_warmup_steps,
use_tpu=False)
# Calculate evaluation metrics.
def metric_fn(label_ids, predicted_labels):
accuracy = tf.metrics.accuracy(label_ids, predicted_labels)
f1_score = tf.contrib.metrics.f1_score(label_ids, predicted_labels)
auc = tf.metrics.auc(label_ids, predicted_labels)
recall = tf.metrics.recall(label_ids, predicted_labels)
precision = tf.metrics.precision(label_ids, predicted_labels)
true_pos = tf.metrics.true_positives(label_ids, predicted_labels)
true_neg = tf.metrics.true_negatives(label_ids, predicted_labels)
false_pos = tf.metrics.false_positives(label_ids, predicted_labels)
false_neg = tf.metrics.false_negatives(label_ids, predicted_labels)
return {
"eval_accuracy": accuracy,
"f1_score": f1_score,
"auc": auc,
"precision": precision,
"recall": recall,
"true_positives": true_pos,
"true_negatives": true_neg,
"false_positives": false_pos,
"false_negatives": false_neg
}
eval_metrics = metric_fn(label_ids, predicted_labels)
if mode == tf.estimator.ModeKeys.TRAIN:
return tf.estimator.EstimatorSpec(mode=mode, loss=loss, train_op=train_op)
else:
return tf.estimator.EstimatorSpec(mode=mode, loss=loss, eval_metric_ops=eval_metrics)
else:
(predicted_labels, log_probs) = create_model(is_predicting,
input_ids,
input_mask,
segment_ids,
label_ids,
num_labels)
predictions = {'probabilities': log_probs, 'labels': predicted_labels}
return tf.estimator.EstimatorSpec(mode, predictions=predictions)
# Return the actual model function in the closure
return model_fn
# Compute train and warmup steps from batch size
# These hyperparameters are copied from this colab notebook (https://colab.sandbox.google.com/github/tensorflow/tpu/blob/master/tools/colab/bert_finetuning_with_cloud_tpus.ipynb)
BATCH_SIZE = 8
LEARNING_RATE = 2e-5
NUM_TRAIN_EPOCHS = 3
# Warmup is a period of time where the learning rate
# is small and gradually increases--usually helps training.
WARMUP_PROPORTION = 0.1
# Model configs
SAVE_CHECKPOINTS_STEPS = 500
SAVE_SUMMARY_STEPS = 100
# Compute # train and warmup steps from batch size
num_train_steps = int(len(train_features) / BATCH_SIZE * NUM_TRAIN_EPOCHS)
num_warmup_steps = int(num_train_steps * WARMUP_PROPORTION)
# Specify outpit directory and number of checkpoint steps to save
OUTPUT_DIR = 'output'
run_config = tf.estimator.RunConfig(model_dir=OUTPUT_DIR,
save_summary_steps=SAVE_SUMMARY_STEPS,
save_checkpoints_steps=SAVE_CHECKPOINTS_STEPS)
model_fn = model_fn_builder(num_labels=len(label_list),
learning_rate=LEARNING_RATE,
num_train_steps=num_train_steps,
num_warmup_steps=num_warmup_steps)
estimator = tf.estimator.Estimator(model_fn=model_fn,
config=run_config,
params={"batch_size": BATCH_SIZE})
```
Next we create an input builder function that takes our training feature set (`train_features`) and produces a generator. This is a pretty standard design pattern for working with Tensorflow [Estimators](https://www.tensorflow.org/guide/estimators).
```
# Create an input function for training. drop_remainder = True for using TPUs.
train_input_fn = bert.run_classifier.input_fn_builder(features=train_features,
seq_length=MAX_SEQ_LENGTH,
is_training=True,
drop_remainder=False)
```
Now we train our model! For me, using a Colab notebook running on Google's GPUs, my training time was about 14 minutes.
```
print(f'Beginning Training!')
current_time = datetime.now()
# train the model
estimator.train(input_fn=train_input_fn, max_steps=num_train_steps)
print("Training took time ", datetime.now() - current_time)
# check the test result
test_input_fn = run_classifier.input_fn_builder(features=test_features,
seq_length=MAX_SEQ_LENGTH,
is_training=False,
drop_remainder=False)
estimator.evaluate(input_fn=test_input_fn, steps=None)
def predict(in_sentences):
labels = ["non-stress", "stress"]
labels_idx = [0, 1]
input_examples = [run_classifier.InputExample(guid="", text_a = x, text_b = None, label = 0) for x in in_sentences] # here, "" is just a dummy label
input_features = run_classifier.convert_examples_to_features(input_examples,
labels_idx,
MAX_SEQ_LENGTH,
tokenizer)
predict_input_fn = run_classifier.input_fn_builder(features=input_features,
seq_length=MAX_SEQ_LENGTH,
is_training=False,
drop_remainder=False)
predictions = estimator.predict(predict_input_fn)
return [{"text": sentence, "confidence": list(prediction['probabilities']), "labels": labels[prediction['labels']]}
for sentence, prediction in zip(in_sentences, predictions)]
pred_sentences = ["It's Friday! We wish you a nice start into the weekend!",
"Deep breathing exercises are very relaxing. It can also relieve the symptoms of stress and anxiety.",
"Do you like fruits? I like so much! Be Happy, Keep Smiling!"
]
predictions = predict(pred_sentences)
predictions
```
## Save Ckpts to PB file
```
import os
import tensorflow as tf
trained_checkpoint_prefix = 'Learn2Relax/learn2relax/notebooks/output/model.ckpt-1064'
export_dir = './bert_output/'
graph = tf.Graph()
with tf.compat.v1.Session(graph=graph) as sess:
# Restore from checkpoint
loader = tf.compat.v1.train.import_meta_graph(trained_checkpoint_prefix + '.meta')
loader.restore(sess, trained_checkpoint_prefix)
# Export checkpoint to SavedModel
builder = tf.compat.v1.saved_model.builder.SavedModelBuilder(export_dir)
builder.add_meta_graph_and_variables(sess,
[tf.saved_model.TRAINING, tf.saved_model.SERVING],
strip_default_attrs=True)
builder.save()
```
## BERT fine-tuned model for Tensorflow serving
Reference: https://medium.com/delvify/bert-rest-inference-from-the-fine-tuned-model-499997b32851
```
def serving_input_fn():
label_ids = tf.placeholder(tf.int32, [None], name='label_ids')
input_ids = tf.placeholder(tf.int32, [None, MAX_SEQ_LENGTH], name='input_ids')
input_mask = tf.placeholder(tf.int32, [None, MAX_SEQ_LENGTH], name='input_mask')
segment_ids = tf.placeholder(tf.int32, [None, MAX_SEQ_LENGTH], name='segment_ids')
input_fn = tf.estimator.export.build_raw_serving_input_receiver_fn({
'label_ids': label_ids,
'input_ids': input_ids,
'input_mask': input_mask,
'segment_ids': segment_ids,
})()
return input_fn
estimator._export_to_tpu = False
estimator.export_savedmodel(OUTPUT_DIR, serving_input_fn)
# !saved_model_cli show --dir /home/gillianchiang/Insight_Stress_Analysis/framework/output/1581039388/ --all
# !tensorflow_model_server --port=8500 --rest_api_port=8501 --model_name=bert_model --model_base_path=/home/gillianchiang/Insight_Stress_Analysis/framework/output/
```
| github_jupyter |
# Plotting the DFT
In this notebook we will look at the practical issues associated to plotting the DFT and in particular the DFT of real-world signals. We will examine how to map the DFT coefficients to real-world frequencies and we will investigate the frequency resolution of the DFT and the effects of zero padding.
As a quick reminder, the definition of the DFT for a length-$N$ signal is:
$$
X[k] = \sum_{n=0}^{N-1} x[n]\, e^{-j\frac{2\pi}{N}nk}, \quad k=0, \ldots, N-1
$$
As we have seen, the above formula is just the expression of a change of basis in $\mathbb{C}^N$: we're expressing the information contained in the signal in terms of sinusoidal components rather than in terms of pointwise data. The sinusoidal components have all an integer number of periods over the length of the data signal.
In Python, we will use the `fft` module in Numpy to compute the DFT
```
# first our usual bookkeeping
%matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
plt.rcParams["figure.figsize"] = (14,4)
```
Typically, we will take a vector of data points, compute the DFT and plot the magnitude of the result. For instance, consider the DFT of a linear ramp:
```
x = np.arange(0, 1.02, 0.02) - 0.5
plt.stem(x)
X = np.fft.fft(x);
plt.stem(abs(X));
```
## Positive and negative frequencies
The coefficient number $k$ indicates the contribution (in amplitude and phase) of a sinusoidal component of frequency
$$
\omega_k = \frac{2\pi}{N}k
$$
Because of the rotational symmetry of complex exponentials, a positive frequency $\omega$ between $\pi$ and $2\pi$ is equivalent to a negative frequency of $\omega - 2\pi$; this means that half of the DFT coefficients correspond to negative frequencies and when we concentrate on the physical properties of the DFT it would probably make more sense to plot the coefficients centered around zero with positive frequencies on the right and negative frequencies on the left.
The reason why this is not usuall done are many, including
* convenience
* since we are manipulating finite-length signals, the convention dictates that we start at index zero
* when dealing with real-valued data, the DFT is symmetric in magnitude, so the first half of the coefficients is enough
* if we're looking for maxima in the magnitude, it's just easier to start at zero.
There is also another subtle point that we must take into account when shifting a DFT vector: **we need to differentiate between odd and even length signals**. With $k=0$ as the center point, odd-length vectors will produce symmetric data sets with $(N-1)/2$ points left and right of the oring, whereas even-length vectors will be asymmetric, with one more point on the positive axis; indeed, the highest positive frequency for even-length signals will be equal to $\omega_{N/2} = \pi$. Since the frequencies of $\pi$ and $-\pi$ are identical, we can copy the top frequency data point to the negative axis and obtain a symmetric vector also for even-length signals.
Here is a function that does that:
```
def dft_shift(X):
N = len(X)
if (N % 2 == 0):
# even-length: return N+1 values
return np.concatenate((X[(N/2):], X[:(N/2)+1]))
else:
# odd-length: return N values
return np.concatenate((X[int((N+1)/2):], X[:int((N-1)/2)]))
plt.stem(abs(dft_shift(X)));
```
While the function does shift the vector, the indices are still from zero to $N-1$. Let's modify it so that we returs also the proper values for the indices:
```
def dft_shift(X):
N = len(X)
if (N % 2 == 0):
# even-length: return N+1 values
return np.arange(-int(N/2), int(N/2) + 1), np.concatenate((X[int(N/2):], X[:int(N/2)+1]))
else:
# odd-length: return N values
return np.arange(-int((N-1)/2), int((N-1)/2) + 1), np.concatenate((X[int((N+1)/2):], X[:int((N+1)/2)]))
n, y = dft_shift(X)
plt.stem(n, abs(y));
```
## Mapping the DFT index to real-world frequencies
The next step is to use the DFT to analyze real-world signals. As we have seen in previous examples, what we need to do is set the time interval between samples or, in other words, set the "clock" of the system. For audio, this is equivalent to the sampling rate of the file.
Here for instance is the sound of a piano
```
import IPython
from scipy.io import wavfile
Fs, x = wavfile.read("piano.wav")
IPython.display.Audio(x, rate=Fs)
```
In order to look at the spectrum of the sound file with a DFT we need to map the digital frequency "bins" of the DFT to real-world frequencies.
The $k$-th basis function over $\mathbb{C}^N$ completes $k$ periods over $N$ samples. If the time between samples is $1/F_s$, then the real-world frequency of the $k$-th basis function is periods over time, namely $k(F_s/N).
Let's remap the DFT coefficients using the sampling rate:
```
def dft_map(X, Fs, shift=True):
resolution = float(Fs) / len(X)
if shift:
n, Y = dft_shift(X)
else:
Y = X
n = np.arange(0, len(Y))
f = n * resolution
return f, Y
# let's cut the signal otherwise it's too big
x = x[:32768]
X = np.fft.fft(x);
f, y = dft_map(X, Fs)
plt.plot(f, abs(y));
```
The plot shows what a spectrum analyzer would display. We can see the periodic pattern in the sound, like for all musical tones. If we want to find out the original pitch we need to zoom in in the plot and find the first peak. This is one of the instances in which shifting the DFT does not help, since we'll be looking in the low-frequency range. So let's re-plot withouth the shift, but still mapping the frequencies:
```
X = np.fft.fft(x);
f, y = dft_map(X, Fs, shift=False)
plt.plot(f[:2000], abs(y[:2000]));
```
We can see that the first peak is in the vicinity of 200Hz; to find the exact frequency (to within the resolution afforded by this DFT) let's find the location
```
dft_resolution = float(Fs)/ len(x)
print("DFT resolution is", dft_resolution, "Hz")
# let's search up to 300Hz
max_range = int(300 / dft_resolution)
ix = np.argmax(abs(y[:max_range]))
pitch = f[ix]
print("the note has a pitch of", pitch, "Hz")
```
so the note is a A, half the frequency of concert pitch.
## Zero-padding
Since the resolution of a DFT depends on the length of the data vector, one may erroneously assume that, by *artificially* extending a given data set, the resulting resolution would improve. Note that here we're not talking about *collecting* more data; rather, we have a data set and we append zeros (or any other constant value) to the end of it. This extension is called zero-padding.
The derivation of why zero-padding does not increase the resolution is detailed in the book. Here we will just present a simple example.
Assume we're in $\mathbb{C}^N$ with $N=256$. The resolution of the DFT in this space is
$$
\Delta = 2\pi/256 \approx 0.0245
$$
Let's build a signal with two sinusoids with frequencies more than $\Delta$ apart and let's look at the spectrum:
```
N = 256
Delta = 2*np.pi / N
n = np.arange(0, N)
# main frequency (not a multiple of the fundamental freq for the space)
omega = 2*np.pi / 10
x = np.cos(omega * n) + np.cos((omega + 3*Delta) * n)
plt.plot(abs(np.fft.fft(x))[:100]);
```
we can tell the two frequencies apart and, if you zoom in on the plot, you will see that they are indeed three indices apart. Now let's build a signal with two frequencies that are less than $\Delta$ apart:
```
x = np.cos(omega * n) + np.cos((omega + 0.5*Delta) * n)
plt.plot(abs(np.fft.fft(x))[:100]);
```
The two frequencies cannot be resolved by the DFT. If you try to increase the data vector by zero padding, the plot will still display just one peak:
```
xzp = np.concatenate((x, np.zeros(2000)))
plt.plot(abs(np.fft.fft(xzp))[:500]);
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
# This code creates a virtual display to draw game images on.
# If you are running locally, just ignore it
import os
if type(os.environ.get("DISPLAY")) is not str or len(os.environ.get("DISPLAY")) == 0:
!bash ../xvfb start
%env DISPLAY = : 1
```
### OpenAI Gym
We're gonna spend several next weeks learning algorithms that solve decision processes. We are then in need of some interesting decision problems to test our algorithms.
That's where OpenAI gym comes into play. It's a python library that wraps many classical decision problems including robot control, videogames and board games.
So here's how it works:
```
import gym
env = gym.make("MountainCar-v0")
plt.imshow(env.render('rgb_array'))
print("Observation space:", env.observation_space)
print("Action space:", env.action_space)
```
Note: if you're running this on your local machine, you'll see a window pop up with the image above. Don't close it, just alt-tab away.
### Gym interface
The three main methods of an environment are
* __reset()__ - reset environment to initial state, _return first observation_
* __render()__ - show current environment state (a more colorful version :) )
* __step(a)__ - commit action __a__ and return (new observation, reward, is done, info)
* _new observation_ - an observation right after commiting the action __a__
* _reward_ - a number representing your reward for commiting action __a__
* _is done_ - True if the MDP has just finished, False if still in progress
* _info_ - some auxilary stuff about what just happened. Ignore it ~~for now~~.
```
obs0 = env.reset()
print("initial observation code:", obs0)
# Note: in MountainCar, observation is just two numbers: car position and velocity
print("taking action 2 (right)")
new_obs, reward, is_done, _ = env.step(2)
print("new observation code:", new_obs)
print("reward:", reward)
print("is game over?:", is_done)
# Note: as you can see, the car has moved to the riht slightly (around 0.0005)
```
### Play with it
Below is the code that drives the car to the right.
However, it doesn't reach the flag at the far right due to gravity.
__Your task__ is to fix it. Find a strategy that reaches the flag.
You're not required to build any sophisticated algorithms for now, feel free to hard-code :)
_Hint: your action at each step should depend either on __t__ or on __s__._
```
# create env manually to set time limit. Please don't change this.
TIME_LIMIT = 250
env = gym.wrappers.TimeLimit(gym.envs.classic_control.MountainCarEnv(),
max_episode_steps=TIME_LIMIT + 1)
s = env.reset()
actions = {'left': 0, 'stop': 1, 'right': 2}
# prepare "display"
%matplotlib notebook
fig = plt.figure()
ax = fig.add_subplot(111)
fig.show()
for t in range(TIME_LIMIT):
# change the line below to reach the flag
if s[1]>=0:
s, r, done, _ = env.step(actions['right'])
else:
s, r, done, _ = env.step(actions['left'])
# draw game image on display
ax.clear()
ax.imshow(env.render('rgb_array'))
fig.canvas.draw()
if done:
print("Well done!")
break
else:
print("Time limit exceeded. Try again.")
assert s[0] > 0.47
print("You solved it!")
```
| github_jupyter |
# Opioids in Tennessee
**An Exploration of the Washington Post Opioid Dataset for Tennessee**
**Contact**
* [Qiusheng Wu](https://wetlands.io)
* [Michael Camponovo](https://geography.utk.edu/about-us/faculty/michael-camponovo/)
## Data
* [Tennessee Population by County (2019)](http://worldpopulationreview.com/us-counties/tn/)
* [Tennessee Opioid Data (2006-2012)](https://www.washingtonpost.com/wp-stat/dea-pain-pill-database/summary/arcos-tn-statewide-itemized.tsv.gz) (zip file: 139 MB; uncompressed: 2.4 GB; 5.75 million rows)
## Useful Links
* [StoryMaps of Opioids in Tennessee by Michael Camponovo](https://storymaps.arcgis.com/stories/92678e77f9f9467bbad17c06f68ac67b)
* [Pandas Basics (Reading Data Files, DataFrames, Data Selection](https://data36.com/pandas-tutorial-1-basics-reading-data-files-dataframes-data-selection/)
* [Pandas Tutorial 2: Aggregation and Grouping](https://data36.com/pandas-tutorial-2-aggregation-and-grouping/)
* [ipyleaflet: Interactive maps in the Jupyter notebook](https://ipyleaflet.readthedocs.io/en/latest/index.html)
* https://spatial.utk.edu/maps/opioids-tn.html
## Table of Content
* [Loading Opioid Data](#loading-opioid-data)
* [Selecting Data Columns](#selecting-data-columns)
* [Descriptive Statistics](#descriptive-statistics)
* [Pill Count Summary by Year, Month, and Day of Week](#pill-count-summary-by-date)
* [Top 10 Distributors, Manufacturers, and Buyers](#top-10)
### Loading Opioid Data <a class="anchor" id="loading-opioid-data"></a>
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import calendar
from ipyleaflet import *
import warnings
warnings.filterwarnings('ignore')
# magic function making plot outputs appear within the notebook
%matplotlib inline
# change the default plot output size
plt.rcParams['figure.figsize'] = [18, 8]
```
The Tennesee opioid data (2006-2012) can be downloaded via this [direct link](https://www.washingtonpost.com/wp-stat/dea-pain-pill-database/summary/arcos-tn-statewide-itemized.tsv.gz). Note that the downloaded zip file is only 139 MB. When uncompressed, the tab separated values (TSV) file is 2.4 GB, which contains 5.75 millions rows. If you are interested in opioid data for other U.S. states, check this [DEA opioid database](https://www.washingtonpost.com/graphics/2019/investigations/dea-pain-pill-database/) by the Washington Post.
```
# set file paths of opioid and population data
data_path = "/media/hdd/Data/Opioids/arcos-tn-statewide-itemized.tsv"
pop_path = '/media/hdd/Data/Opioids/tn-county-pop-2019.csv'
# read population data
pop = pd.read_csv(pop_path)
# read opioid data
data = pd.read_csv(data_path, sep='\t')
```
There are 42 columns in this dataset. The **DOSAGE_UNIT** column is the number of pills. More information about each individual column can be found in the [DEA ARCOS's Handbook](https://www.deadiversion.usdoj.gov/arcos/handbook/full.pdf).
Some key columns we will be using include:
* **REPORTER_NAME**: Distributor
* **BUYER_BUS_ACT**: Buyer Type
* **BUYER_NAME**: Pharmacies
* **BUYER_CITY**: Buyer City
* **BUYER_COUNTY**: Buyer County
* **BUYER_STATE**: Buyer State
* **BUYER_ZIP**: Buyer Zip Code
* **DRUG_NAME**: Drug Name
* **TRANSACTION_DATE**: Transaction Date
* **DOSAGE_UNIT**: Dosage Unit
* **Combined_Labeler_Name**: Manufacturers
```
for col in data.columns:
print(col)
print(f"\nTotal number of columns: {len(data.columns)}")
# Take a look at the first five rows
data.head()
# Take a look at the last five rows
data.tail()
```
### Selecting Data Columns <a class="anchor" id="selecting-data-columns"></a>
```
# Select 11 columns for further investigation
sel_cols = ['REPORTER_NAME', 'BUYER_BUS_ACT', 'BUYER_NAME', 'BUYER_CITY', 'BUYER_COUNTY', 'BUYER_STATE',
'BUYER_ZIP', 'DRUG_NAME', 'TRANSACTION_DATE', 'DOSAGE_UNIT', 'Combined_Labeler_Name']
data = data[sel_cols]
for col in data.columns:
print(col)
data.head()
print(f"Total number of rows: {len(data.index):,}")
# Check the transaction date format. Note that it is in the MMDDYYYY format, but we need to pad zeros
data['TRANSACTION_DATE'].head()
# Convert TRANSACTION_DATE from String to DateTime and pad zeros
data['TRANSACTION_DATE'] = pd.to_datetime(data['TRANSACTION_DATE'].astype(str).str.zfill(8), format='%m%d%Y')
# Take a look at the revised TRANSACTION_DATE
data['TRANSACTION_DATE'].head()
# Take a look at the first five rows again with revised TRANSACTION_DATE
data.head()
```
### Descriptive Statistics <a class="anchor" id="descriptive-statistics"></a>
```
total_pills = int(data.DOSAGE_UNIT.sum())
print(f"Total number of pills distributed to Tennessee (2006-2012): {total_pills: ,}")
total_pop = pop['Pop'].sum()
print(f"Total population in Tennessee: {total_pop:,}")
avg_pills = int(total_pills / total_pop)
print(f"Average number of pills per person: {avg_pills}")
# Check TN counties
counties = data['BUYER_COUNTY']
print(sorted(set(counties)))
print(f"\nTotal number of counties: {len(set(counties))}")
# Check TN cities
cities = data['BUYER_CITY']
# print(sorted(set(cities)))
print(f"Total number of cities: {len(set(cities))}")
```
### Pill Count Summary by Year, Month, and Day of Week <a class="anchor" id="pill-count-summary-by-date"></a>
Let's summarize the dataset by year to see how the number of opioids were distributed to Tennessee over time.
```
pills_by_year = data[['TRANSACTION_DATE', 'DOSAGE_UNIT']].groupby(data.TRANSACTION_DATE.dt.year).sum()
pills_by_year
# Plot the trend by year
ax = pills_by_year.plot.bar(title="Pill Count Summary by Year", rot=0)
ax.set_xlabel("Year")
ax.set_ylabel("Pill Count")
vals = ax.get_yticks().astype(int)
ax.set_yticklabels(['{:,}'.format(x) for x in vals])
```
Let's summarize the dataset by month to see how the number of opioids were distributed to Tennessee over time.
```
pills_by_month = data[['TRANSACTION_DATE', 'DOSAGE_UNIT']].groupby(data.TRANSACTION_DATE.dt.month).sum()
pills_by_month['Month'] = pills_by_month.index
pills_by_month.index = pills_by_month['Month'].apply(lambda x: calendar.month_abbr[x])
pills_by_month = pills_by_month[['DOSAGE_UNIT']]
pills_by_month
# Plot the trend by month
ax = pills_by_month.plot.bar(title="Pill Count Summary by Month", rot=0)
ax.set_xlabel("Month")
ax.set_ylabel("Pill Count")
vals = ax.get_yticks().astype(int)
ax.set_yticklabels(['{:,}'.format(x) for x in vals])
```
Let's summarize the dataset by day to see how the number of opioids were distributed to Tennessee over time.
```
pills_by_day = data[['TRANSACTION_DATE', 'DOSAGE_UNIT']].groupby(data.TRANSACTION_DATE.dt.day).sum()
pills_by_day
# Plot the trend by day
ax = pills_by_day.plot(title="Pill Count Summary by Day", rot=0, style='-', marker='o')
ax.set_xlabel("Day")
ax.set_ylabel("Pill Count")
vals = ax.get_yticks().astype(int)
ax.set_yticklabels(['{:,}'.format(x) for x in vals])
```
Let's summarize the dataset by day of week to see how the number of opioids were distributed to Tennessee over time.
```
pills_by_dayofweek = data[['TRANSACTION_DATE', 'DOSAGE_UNIT']].groupby(data.TRANSACTION_DATE.dt.dayofweek).sum()
pills_by_dayofweek['DayofWeek'] = pills_by_dayofweek.index
pills_by_dayofweek.index = pills_by_dayofweek['DayofWeek'].apply(lambda x: calendar.day_abbr[x])
pills_by_dayofweek = pills_by_dayofweek[['DOSAGE_UNIT']]
pills_by_dayofweek
ax = pills_by_dayofweek.plot.bar(title="Pill Count Summary by Day of Week", rot=0)
ax.set_xlabel("Day of Week")
ax.set_ylabel("Pill Count")
vals = ax.get_yticks().astype(int)
ax.set_yticklabels(['{:,}'.format(x) for x in vals])
```
Let's look at the total pills per month by year:
```
pills_by_year_month = data[['TRANSACTION_DATE', 'DOSAGE_UNIT']].groupby([data.TRANSACTION_DATE.dt.month,
data.TRANSACTION_DATE.dt.year]).sum()
# pills_by_year_month.index
# pills_by_month['Month'] = pills_by_month.index
# pills_by_month.index = pills_by_month['Month'].apply(lambda x: calendar.month_abbr[x])
# pills_by_month = pills_by_month[['DOSAGE_UNIT']]
ax = pills_by_year_month['DOSAGE_UNIT'].unstack().plot(style='-', marker='o')
ax.set_xlabel("Month")
ax.set_ylabel("Pill Count")
vals = ax.get_yticks().astype(int)
ax.set_yticklabels(['{:,}'.format(x) for x in vals])
```
### Top 10 Distributors, Manufacturers, and Buyers<a class="anchor" id="top-10"></a>
Let's take a look at the top 10 distributors:
```
distributors = data[['REPORTER_NAME', 'DOSAGE_UNIT']].groupby('REPORTER_NAME').sum()
distributors_top10 = distributors.sort_values('DOSAGE_UNIT', ascending=False).head(10)
distributors_top10
```
The table above shows that the 2nd distributor (**WALGREEN CO**) and 8th distributor (**WALGREEN CO.**) are treated as different distirbutors. We need to remove the extra period from the 8th distributor.
```
data['REPORTER_NAME'] = data['REPORTER_NAME'].str.replace('.', '', regex=False)
distributors = data[['REPORTER_NAME', 'DOSAGE_UNIT']].groupby('REPORTER_NAME').sum()
distributors_top10 = distributors.sort_values('DOSAGE_UNIT', ascending=False).head(10)
distributors_top10
distributors_top10.index = distributors_top10.index.str[:15]
ax = distributors_top10.plot.bar(title="Pill Count Summary by Distributor", rot=0)
ax.set_xlabel("Distributor")
ax.set_ylabel("Pill Count")
vals = ax.get_yticks().astype(int)
ax.set_yticklabels(['{:,}'.format(x) for x in vals])
```
Let's take a look at the top 10 manufacturers:
```
manufacturers = data[['Combined_Labeler_Name', 'DOSAGE_UNIT']].groupby('Combined_Labeler_Name').sum()
manufacturers_top10 = manufacturers.sort_values('DOSAGE_UNIT', ascending=False).head(10)
manufacturers_top10
manufacturers_top10.index = manufacturers_top10.index.str[:15]
ax = manufacturers_top10.plot.bar(title="Pill Count Summary by Manufacturer", rot=0)
ax.set_xlabel("Manufacturer")
ax.set_ylabel("Pill Count")
vals = ax.get_yticks().astype(int)
ax.set_yticklabels(['{:,}'.format(x) for x in vals])
```
Let's take a look at the top 10 buyers:
```
buyers = data[['BUYER_NAME', 'DOSAGE_UNIT']].groupby('BUYER_NAME').sum()
buyers_top10 = buyers.sort_values('DOSAGE_UNIT', ascending=False).head(10)
buyers_top10
buyers_top10.index = buyers_top10.index.str[:15]
ax = buyers_top10.plot.bar(title="Pill Count Summary by Buyer", rot=0)
ax.set_xlabel("Buyer")
ax.set_ylabel("Pill Count")
vals = ax.get_yticks().astype(int)
ax.set_yticklabels(['{:,}'.format(x) for x in vals])
counties = data[['BUYER_COUNTY', 'DOSAGE_UNIT']].groupby('BUYER_COUNTY').sum()
counties_top10 = counties.sort_values('DOSAGE_UNIT', ascending=False).head(10)
counties_top10
ax = counties_top10.plot.bar(title="Pill Count Summary by County", rot=0)
ax.set_xlabel("County")
ax.set_ylabel("Pill Count")
vals = ax.get_yticks().astype(int)
ax.set_yticklabels(['{:,}'.format(x) for x in vals])
cities = data[['BUYER_CITY', 'DOSAGE_UNIT']].groupby('BUYER_CITY').sum()
cities_top10 = cities.sort_values('DOSAGE_UNIT', ascending=False).head(10)
cities_top10
ax = cities_top10.plot.bar(title="Pill Count Summary by City", rot=0)
ax.set_xlabel("City")
ax.set_ylabel("Pill Count")
vals = ax.get_yticks().astype(int)
ax.set_yticklabels(['{:,}'.format(x) for x in vals])
```
| github_jupyter |
# Analyzing the community activity for version control systems
## Exercise
### Background
Technology choices are different. There may be objective reasons for technology at a specific time. But those reasons often change over time. But the developed deep love for an now outdated technology can prevent every progress. Thus objective reasons may become subjective which can create a toxic environment when technology updates are addressed.
### Your task
You are a new team member in a software company. The developers there are using a version control system ("VCS" for short) called CVS (Concurrent Versions System). Some want to migrate to a better VCS. They prefer one that's called SVN (Subversion). You are young but not inexperienced. You heard about newer version control system named "Git". So you propose Git as an alternative to the team. They are very sceptical about your suggestion. Find evidence that shows that the software development community is mainly adopting the Git version control system!
### The dataset
There is a dataset from the online software developer community Stack Overflow in `../datasets/stackoverflow_vcs_data_subset.gz` (with a subset of columns) available with the following data:
* `CreationDate`: the timestamp of the creation date of a Stack Overflow post (= question)
* `TagName`: the tag name for a technology (in our case for only 4 VCSes: "cvs", "svn", "git" and "mercurial")
* `ViewCount`: the numbers of views of a post
## Your solution
### Step 1: Load in the dataset
###### <span style="color:green;">SOLUTION <small>(Click the arrow on the left side if a hint is needed)</small></span>
```
import pandas as pd
vcs_data = pd.read_csv('../datasets/stackoverflow_vcs_data_subset.gz')
vcs_data.head()
```
### Step 2: Explore the dataset by displaying the number of all posts for each VCS
###### <span style="color:green;">SOLUTION <small>(Click the arrow on the left side if a hint is needed)</small></span>
```
vcs_data['TagName'].value_counts()
```
### Step 3: Convert the column with the time stamp
###### <span style="color:green;">SOLUTION <small>(Click the arrow on the left side if a hint is needed)</small></span>
```
vcs_data['CreationDate'] = pd.to_datetime(vcs_data['CreationDate'])
vcs_data.head()
```
### Step 4: Sum up the view counts by the timestamp and the VCSes
###### <span style="color:green;">SOLUTION <small>(Click the arrow on the left side if a hint is needed)</small></span>
```
number_of_posts = vcs_data.groupby(['CreationDate', 'TagName']).sum()
number_of_posts.head()
```
### Step 5: List the number of views by date for each VCS
Hint: You may unstack and fill in the data (and get rid of the hierarchical column)
###### <span style="color:green;">SOLUTION <small>(Click the arrow on the left side if a hint is needed)</small></span>
```
views_per_vcs = number_of_posts.unstack(fill_value=0)['ViewCount']
views_per_vcs.head()
```
### Step 6: Accumulate the number of views for the VCSes for every month over all the years
Hint: First, you have to resample the data and summing it up
###### <span style="color:green;">SOLUTION <small>(Click the arrow on the left side if a hint is needed)</small></span>
```
cumulated_posts = views_per_vcs.resample("1M").sum().cumsum()
cumulated_posts.head()
```
### Step 7: Visualize the number of views over time for all VCSes
###### <span style="color:green;">SOLUTION <small>(Click the arrow on the left side if a hint is needed)</small></span>
```
%matplotlib inline
cumulated_posts.plot(title="accumulated monthly stackoverflow post views");
```
### Step 8: What are your conclusions? Discuss!
| github_jupyter |
[View in Colaboratory](https://colab.research.google.com/github/nishi1612/SC374-Computational-and-Numerical-Methods/blob/master/Set_7.ipynb)
```
from scipy.interpolate import CubicSpline
import numpy as np
import matplotlib.pyplot as plt
def linear_lagrange(a,x):
y = a[1]*(x - a[2])/(a[0] - a[2]) + a[3]*(x - a[0])/(a[2] - a[0])
#plt.plot(x,y,color='blue',label='Linear Lagrange')
#graph_details()
return y
def zero_newton(a,x):
return a[1]
def one_newton(a,x,f):
y = zero_newton(a,x) + ((x-a[0])*(a[1]-a[3])/(a[0]-a[2]))
if(f==1):
plt.plot(x,y,color = 'blue',label='First Order Newton Divided Difference')
plt.legend()
plt.grid(True)
plt.savefig('Graph_' + str(number) + '.png')
files.download('Graph_' + str(number) + '.png')
plt.show()
return y
def func_one(x1,x2,y1,y2):
return (y2 - y1) / (x2 - x1)
def two_newton(a,x,f):
j = a[4] - a[0]
i = func_one(a[2],a[4],a[3],a[5])
k = func_one(a[0],a[2],a[1],a[3])
j = (i - k) / j
y = one_newton(a,x,0) + ((x-a[0])*(x-a[2])*j)
if(f==1):
plt.plot(x,y,color = 'blue',label='Second Order Newton Divided Difference')
plt.legend()
plt.grid(True)
plt.savefig('Graph_' + str(number) + '.png')
files.download('Graph_' + str(number) + '.png')
plt.show()
return y
def func_two(x1,x2,x3,y1,y2,y3):
a = func_one(x1,x2,y1,y2)
b = func_one(x2,x3,y2,y3)
return (b - a) / (x3 - x1)
def three_newton(a,x,f):
j = a[6] - a[0]
k = func_two(a[2],a[4],a[6],a[3],a[5],a[7]) - func_two(a[0],a[2],a[4],a[1],a[3],a[5])
y = two_newton(a,x,0) + ((x-a[0])*(x-a[2])*(x-a[4])*k/j)
if(f==1):
plt.plot(x,y,color = 'blue',label='Third Order Newton Divided Difference')
plt.legend()
plt.grid(True)
plt.savefig('Graph_' + str(number) + '.png')
files.download('Graph_' + str(number) + '.png')
plt.show()
return y
point1 = [1,1]
point2 = [2,1/2]
point3 = [3,1/3]
point4 = [4,1/4]
a = point1 + point2
x1 = np.arange(a[0],a[2],0.00001)
y1 = linear_lagrange(a,x1)
a = point2 + point3
x2 = np.arange(a[0],a[2],0.00001)
y2 = linear_lagrange(a,x2)
a = point3 + point4
x3 = np.arange(a[0],a[2],0.00001)
y3 = linear_lagrange(a,x3)
plt.plot(x1,y1,color='blue',label='Linear Lagrange')
plt.plot(x2,y2,color='blue')
plt.plot(x3,y3,color='blue')
x = [1,2,3,4]
y = [1,1/2,1/3,1/4]
cs = CubicSpline(x,y)
t = np.arange(1,4,0.0001)
plt.plot(t,cs(t),color='green',label='Cubic Spline')
for i in range(len(x)):
plt.plot(x[i],y[i],color='red',marker='o')
plt.legend()
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.grid(True)
plt.title('Comparing Linear Interpolation and Cubic Spline')
plt.show()
plt.plot(t,cs(t),color='green',label='Cubic Spline')
plt.plot(t,1/t,color='blue',label='Actual function')
plt.legend()
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.grid(True)
plt.title('Comparing Actual function and Cubic Spline')
plt.show()
plt.plot(t,cs(t) - 1/t)
plt.xlabel('$x$')
plt.ylabel('Error')
plt.grid(True)
plt.title('Error between Cubic Spline and Actual function')
plt.show()
number = 5
point1 = [0 , 2.5]
point2 = [1 , 0.5]
point3 = [2 , 0.5]
point4 = [2.5 , 1.5]
point5 = [3 , 1.5]
point6 = [3.5 , 1.125]
point7 = [4 , 0]
a = point1 + point2
x1 = np.arange(a[0],a[2],0.00001)
y1 = linear_lagrange(a,x1)
a = point2 + point3
x2 = np.arange(a[0],a[2],0.00001)
y2 = linear_lagrange(a,x2)
a = point3 + point4
x3 = np.arange(a[0],a[2],0.00001)
y3 = linear_lagrange(a,x3)
a = point4 + point5
x4 = np.arange(a[0],a[2],0.00001)
y4 = linear_lagrange(a,x4)
a = point5 + point6
x5 = np.arange(a[0],a[2],0.00001)
y5 = linear_lagrange(a,x5)
a = point6 + point7
x6 = np.arange(a[0],a[2],0.00001)
y6 = linear_lagrange(a,x6)
plt.plot(x1,y1,color='green',label='Linear Lagrange')
plt.plot(x2,y2,color='green')
plt.plot(x3,y3,color='green')
plt.plot(x4,y4,color='green')
plt.plot(x5,y5,color='green')
plt.plot(x6,y6,color='green')
x = [0,1,2,2.5,3,3.5,4]
y = [2.5,0.5,0.5,1.5,1.5,1.125,0]
cs = CubicSpline(x,y)
t = np.arange(x[0],x[len(x)-1],0.0001)
plt.plot(t,cs(t),color='purple',label='Cubic Spline')
for i in range(len(x)):
plt.plot(x[i],y[i],color='blue',marker='o')
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.grid(True)
plt.legend()
plt.title('Comparing Linear Interpolation and Cubic Spline')
plt.show()
point1 = [-0.5 , 0.731531]
point2 = [0 , 1.0]
point3 = [0.25 , 1.268400]
point4 = [1 , 1.718282]
a = point1 + point2
x1 = np.arange(a[0],a[2],0.00001)
y1 = linear_lagrange(a,x1)
a = point2 + point3
x2 = np.arange(a[0],a[2],0.00001)
y2 = linear_lagrange(a,x2)
a = point3 + point4
x3 = np.arange(a[0],a[2],0.00001)
y3 = linear_lagrange(a,x3)
plt.plot(x1,y1,color='green',label='Linear Lagrange')
plt.plot(x2,y2,color='green')
plt.plot(x3,y3,color='green')
x = [-0.5, 0 , 0.25 ,1]
y = [0.731531 ,1.0, 1.268400 , 1.718282]
cs = CubicSpline(x,y)
t = np.arange(x[0],x[len(x)-1],0.0001)
plt.plot(t,cs(t),color='purple',label='Cubic Spline')
for i in range(len(x)):
plt.plot(x[i],y[i],color='blue',marker='o')
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.grid(True)
plt.legend()
plt.title('Comparing Linear Interpolation and Cubic Spline')
plt.show()
plt.plot(t,cs(t),color='purple',label='Cubic Spline')
plt.plot(t,np.exp(t) - t**3,color='green',label='Actual function')
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.grid(True)
plt.legend()
plt.title('Comparing Cubic Spline and Actual function')
plt.show()
plt.plot(cs(t) - np.exp(t) + t**3)
plt.xlabel('$x$')
plt.ylabel('Error')
plt.title('Error between actual function and cubic spline')
plt.show()
point1 = [0, 1]
point2 = [1, 1]
point3 = [2, 5]
a = point1 + point2 + point3
x = np.arange(a[0],a[4],0.0001)
y = two_newton(a,x,0)
plt.plot(x,y,color='green',label='Second Order Newton Divided Difference')
plt.xlabel('$x$')
plt.ylabel('$y$')
# plt.show()
# a = point1 + point2
# x1 = np.arange(a[0],a[2],0.0001)
# y1 = linear_lagrange(a,x1)
# a = point2 + point3
# x2 = np.arange(a[0],a[2],0.0001)
# y2 = linear_lagrange(a,x2)
# plt.plot(x1,y1,color='green',label='Linear Lagrange')
# plt.plot(x2,y2,color='green')
x = [0, 1, 2]
y = [1, 1, 5]
cs = CubicSpline(x,y)
t = np.arange(x[0],x[len(x)-1],0.0001)
plt.plot(t,cs(t),color='purple',label='Cubic Spline')
for i in range(len(x)):
plt.plot(x[i],y[i],color='blue',marker='o')
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.legend()
plt.title('Comparing Second Order Newton Divided Difference and Cubic Spline')
plt.grid(True)
plt.show()
point1 = [1 , 3]
point2 = [2 , 1]
point3 = [3 , 2]
point4 = [4 , 3]
point5 = [5 , 2]
a = point1 + point2
x1 = np.arange(a[0],a[2],0.00001)
y1 = linear_lagrange(a,x1)
a = point2 + point3
x2 = np.arange(a[0],a[2],0.00001)
y2 = linear_lagrange(a,x2)
a = point3 + point4
x3 = np.arange(a[0],a[2],0.00001)
y3 = linear_lagrange(a,x3)
a = point4 + point5
x4 = np.arange(a[0],a[2],0.00001)
y4 = linear_lagrange(a,x4)
plt.plot(x1,y1,color='green',label='Linear Lagrange')
plt.plot(x2,y2,color='green')
plt.plot(x3,y3,color='green')
plt.plot(x4,y4,color='green')
x = [1,2,3,4,5]
y = [3,1,2,3,2]
cs = CubicSpline(x,y)
t = np.arange(x[0],x[len(x)-1],0.0001)
plt.plot(t,cs(t),color='purple',label='Cubic Spline')
for i in range(len(x)):
plt.plot(x[i],y[i],color='blue',marker='o')
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.grid(True)
plt.legend()
plt.title('Comparing Linear Interpolation and Cubic Spline')
plt.show()
point1 = [0 , 0]
point2 = [1/2 , 1/4]
point3 = [1 , 1]
point4 = [2 , -1]
point5 = [3 , -1]
a = point1 + point2
x1 = np.arange(a[0],a[2],0.00001)
y1 = linear_lagrange(a,x1)
a = point2 + point3
x2 = np.arange(a[0],a[2],0.00001)
y2 = linear_lagrange(a,x2)
a = point3 + point4
x3 = np.arange(a[0],a[2],0.00001)
y3 = linear_lagrange(a,x3)
a = point4 + point5
x4 = np.arange(a[0],a[2],0.00001)
y4 = linear_lagrange(a,x4)
plt.plot(x1,y1,color='green',label='Linear Lagrange')
plt.plot(x2,y2,color='green')
plt.plot(x3,y3,color='green')
plt.plot(x4,y4,color='green')
x = [0,1/2,1,2,3]
y = [0,1/4,1,-1,-1]
cs = CubicSpline(x,y)
t = np.arange(x[0],x[len(x)-1],0.0001)
plt.plot(t,cs(t),color='purple',label='Cubic Spline')
for i in range(len(x)):
plt.plot(x[i],y[i],color='blue',marker='o')
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.grid(True)
plt.legend()
plt.title('Comparing Linear Interpolation and Cubic Spline')
plt.show()
point1 = [0 , 1.4]
point2 = [1 , 0.6]
point3 = [2 , 1]
point4 = [2.5 , 0.65]
point5 = [3 , 0.6]
point6 = [4 , 1]
a = point1 + point2
x1 = np.arange(a[0],a[2],0.00001)
y1 = linear_lagrange(a,x1)
a = point2 + point3
x2 = np.arange(a[0],a[2],0.00001)
y2 = linear_lagrange(a,x2)
a = point3 + point4
x3 = np.arange(a[0],a[2],0.00001)
y3 = linear_lagrange(a,x3)
a = point4 + point5
x4 = np.arange(a[0],a[2],0.00001)
y4 = linear_lagrange(a,x4)
a = point5 + point6
x5 = np.arange(a[0],a[2],0.00001)
y5 = linear_lagrange(a,x5)
a = point6 + point7
x6 = np.arange(a[0],a[2],0.00001)
y6 = linear_lagrange(a,x6)
plt.plot(x1,y1,color='green',label='Linear Lagrange')
plt.plot(x2,y2,color='green')
plt.plot(x3,y3,color='green')
plt.plot(x4,y4,color='green')
plt.plot(x5,y5,color='green')
plt.plot(x6,y6,color='green')
x = [0,1,2,2.5,3,4]
y = [1.4,0.6,1,0.65,0.6,1]
cs = CubicSpline(x,y)
t = np.arange(x[0],x[len(x)-1],0.0001)
plt.plot(t,cs(t),color='purple',label='Cubic Spline')
for i in range(len(x)):
plt.plot(x[i],y[i],color='blue',marker='o')
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.grid(True)
plt.legend()
plt.title('Comparing Linear Interpolation and Cubic Spline')
plt.show()
```
| github_jupyter |
```
print(' '*20+'Version: String Encoder v0.0.1.5')
print('.'*150)
print('@copyright Bijoy_Maji')
print('By - Bijoy Maji. Email:- majibijoy00@gmail.com')
print('Note: There have a problem with some key. if you find it , please mail me the key at majibijoy00@gmail.')
print('.'*150)
import random
import math as m
#input stering
in_put=input('Enter your string:')
#genarate a key for encription
output=[in_put[i] for i in range(len(in_put))]
#gggggsjgfdvghggjgsjg
words_list1=[['l', 'z', 'x', 'c', 'v', 'b','u','i','o','p','a','s'],
['d', 'f', 'g', 'h', 'j', 'k', 'q','w','e','r','t','y'],
['n', 'm', '~', '!', '#', '$', '%', '^', '&', '*', '(', ')'],
['_', '+', '@', '-', '=', '{', '}', '|', '[', ']', ':', '"'],
['`', ';', "'", '<', '>', '?', ',', '.', '/', '€', '¤', ' '],
['Z', 'X', 'C', 'V', 'B', 'Y', 'U', 'I', 'O', 'P', 'A', 'S'],
['D', 'F', 'G', 'H', 'J', 'K', 'L','Q', 'W', 'E', 'R', 'T' ],
['N', 'M', '0','5', '6', '7', '8', '9','1', '2', '3', '4']]
lower_words="qwertyuiopasdfghjklzxcvbnm"
upper_words=lower_words.upper()
number="0123456789"
symbols='''~!#$%^&*()_+@-={}|[]:"`;'<>?,./€¤ '''
words=lower_words+symbols+upper_words+number
words_list=[words[i] for i in range(len(words))]
output1=[words_list.index(i) for i in output]
xp,yp,zp=[],[],[]
chack_key=input("Have you a key ? (y/n):")
yes1=['y','Y','yes','Yes']
no1=['n','N','No','no']
key=[]
keyerror=0
try:
if chack_key in no1:
#key work
key=[str(int(random.uniform(0,9))) for i in range (10) ]
elif chack_key in yes1:
key_in=input('Input our key: ')
key_2=['@','$', 'G','A','#','%','X','!','_','+','/','&','z','r','H','Q','W','D','Y','~']
key_1=[key_in[i] for i in range(len(key_in))]
j=int(key_1[10])
key_3=[]
key_4=[]
position=[]
z=[]
for i in range(12,len(key_1),2):
key_3.append(key_1[i])
for k in key_1[:10]:
if k in key_2:
key_4.append(k)
z.append(key_1.index(k))
position.append(key_2.index(k)-j)
y=[]
for l in range (min(position),max(position)+1):
y.append(key_3[position.index(l)])
c=0
for a in z:
key_1[a]=y[c]
c=c+1
key=key_1[:10]
else:
print("Sorry! only 'y' and 'n' are accepted")
keyerror +=1
# mixing word with key
word_with_key=[]
for p in key:
word_with_key.append(8-int(p))
for q in word_with_key:
if q==8:
word_with_key.remove(q)
for i in word_with_key:
if word_with_key.count(i)>1:
word_with_key.remove(i)
key_len_en=len(word_with_key)
for i in range(8):
if i not in word_with_key:
word_with_key.append(i)
#print ('word_with_key: ',word_with_key)
new_words_list=[]
for i in word_with_key:
new_words_list.append(words_list1[i])
for j in output1 :
a1=int((j+1)/12)
b1=int((j+1)%12)
yp.append(b1-1)
if (j+1)%12==0:
xp.append(a1-1)
zp.append(new_words_list[a1-1][b1-1])
else:
xp.append(a1)
zp.append(new_words_list[a1][b1-1])
key_2=['@','$', 'G','A','#','%','X','!','_','+','/','&','z','r','H','Q','W','D','Y','~']
j=int(random.uniform(-1,5))
main_constant=j
x=[]
for i in key:
position=key.index(i)
counter =0
if key.count(i)>1:
x.append(i)
j3=int(random.uniform(0,len(key_2)))
x.append(key_2[j3])
position1=key.index(i)
j+=1
key[key.index(i,position1+1)]=key_2[j]
key.append(str(main_constant))
j2=int(random.uniform(11,len(key_2)))
key.append(key_2[j2])
key=key+x
key_3=key[0]
for key_in_word in range(1,len(key)):
key_3=key_3+key[key_in_word]
if chack_key in no1:
print('Your key is: ',key_3)
#print('Words=',new_words_list)
#print('Your encoded string:',output)
#print('Out put:',output1)
#print(xp,yp,zp)
result=zp[0]
for key_in_word in range(1,len(zp)):
result=result+zp[key_in_word]
if keyerror ==0:
print('Encoded string is :',result)
except:
print ("some error in input key ! Enter right key and try again !")
print(' '*20+'Version: String Decoder v0.0.1.5')
print('.'*150)
print('@copyright Bijoy_Maji')
print('By - Bijoy Maji. Email:- majibijoy00@gmail.com')
print('Note: There have a problem with some key. if you find it , please mail me the key at majibijoy00@gmail.')
print('.'*150)
import math as m
try:
key=input('Input our key: ')
key_2=['@','$', 'G','A','#','%','X','!','_','+','/','&','z','r','H','Q','W','D','Y','~']
key_1=[key[i] for i in range(len(key))]
j=int(key_1[10])
key_3=[]
key_4=[]
position=[]
z=[]
for i in range(12,len(key_1),2):
key_3.append(key_1[i])
for k in key_1[:10]:
if k in key_2:
key_4.append(k)
z.append(key_1.index(k))
position.append(key_2.index(k)-j)
y=[]
for l in range (min(position),max(position)+1):
y.append(key_3[position.index(l)])
c=0
for a in z:
key_1[a]=y[c]
c=c+1
out_come=key_1[:10]
word_with_key=[]
for p in out_come:
word_with_key.append(8-int(p))
for q in word_with_key:
if q==8:
word_with_key.remove(q)
for i in word_with_key:
if word_with_key.count(i)>1:
word_with_key.remove(i)
for i in range(8):
if i not in word_with_key:
word_with_key.append(i)
#print(word_with_key)
in_put=input('Enter your enoded string:')
output=[in_put[i] for i in range(len(in_put))]
words_list1=[['l', 'z', 'x', 'c', 'v', 'b','u','i','o','p','a','s'],
['d', 'f', 'g', 'h', 'j', 'k', 'q','w','e','r','t','y'],
['n', 'm', '~', '!', '#', '$', '%', '^', '&', '*', '(', ')'],
['_', '+', '@', '-', '=', '{', '}', '|', '[', ']', ':', '"'],
['`', ';', "'", '<', '>', '?', ',', '.', '/', '€', '¤', ' '],
['Z', 'X', 'C', 'V', 'B', 'Y', 'U', 'I', 'O', 'P', 'A', 'S'],
['D', 'F', 'G', 'H', 'J', 'K', 'L','Q', 'W', 'E', 'R', 'T' ],
['N', 'M', '0','5', '6', '7', '8', '9','1', '2', '3', '4']]
lower_words="qwertyuiopasdfghjklzxcvbnm"
upper_words=lower_words.upper()
number="0123456789"
symbols='''~!#$%^&*()_+@-={}|[]:"`;'<>?,./€¤ '''
words=lower_words+symbols+upper_words+number
words_list=[words[i] for i in range(len(words))]
output1=[words_list.index(i) for i in output]
new_words_list=[]
for i in word_with_key:
new_words_list.append(words_list1[i])
words_list2=[]
for i in new_words_list:
words_list2=words_list2+i
result=""
try:
for i in in_put:
#print(i,words_list2.index(i))
result+=words[words_list2.index(i)]
print('Your decoded string is: ',result)
except IndexError:
print('Input right values....')
except:
print("This key is wrong. Please input right key")
```
| github_jupyter |
```
import gzip
import json
import re
from typing import Iterator
import nltk
import numpy as np
import pandas as pd
from mizani.breaks import date_breaks
from mizani.formatters import date_format
from nltk.corpus import stopwords
from plotnine import *
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics.pairwise import linear_kernel, pairwise_distances
from sklearn.pipeline import FeatureUnion, Pipeline
from sklearn.preprocessing import FunctionTransformer
pd.set_option("max_colwidth", None)
nltk.download("stopwords")
```
# Table of Contents
- [Step A.1](#Step-A.1)
* [Substep: EDA](#Substep:-EDA)
- [Step A.2](#Step-A.2)
- [Step A.3](#Step-A.3)
- [Step A.4](#Step-A.4)
- [Step A.5](#Step-A.5)
- [Step A.6](#Step-A.6)
```
# disclaimer: may not work as is in Windows OS
# download the “small” 5-core dataset for the category "Digital Music"
# dataset source: https://nijianmo.github.io/amazon/index.html
!wget --backups=1 http://deepyeti.ucsd.edu/jianmo/amazon/categoryFilesSmall/Digital_Music_5.json.gz -P data/
# disclaimer: may not work as is in Windows OS
# download the metadata for this dataset
# dataset source: https://nijianmo.github.io/amazon/index.html
!wget --backups=1 http://deepyeti.ucsd.edu/jianmo/amazon/metaFiles2/meta_Digital_Music.json.gz -P data/
```
## Step A.1
The 5-core dataset for the category "Digital Music" subset of the [Amazon Review data](https://nijianmo.github.io/amazon/index.html) in which all users and items have at least 5 reviews.
The format is one-review-per-line in JSON, with the following attributes:
- `reviewerID` - ID of the reviewer, e.g. A2SUAM1J3GNN3B
- `asin` - ID of the product, e.g. 0000013714
- `reviewerName` - name of the reviewer
- `vote` - helpful votes of the review
- `style` - a disctionary of the product metadata, e.g., "Format" is "Hardcover"
- `reviewText` - text of the review
- `overall` - rating of the product
- `summary` - summary of the review
- `verified`- whether the review has been verified (boolean)
- `unixReviewTime` - time of the review (unix time)
- `reviewTime` - time of the review (raw)
- `image` - images that users post after they have received the product
Metadata includes descriptions, price, sales-rank, brand info, and co-purchasing links:
- `asin` - ID of the product, e.g. 0000031852
- `title` - name of the product
- `feature` - bullet-point format features of the product
- `description` - description of the product
- `price` - price in US dollars (at time of crawl)
- `imageURL` - url of the product image
- `imageURLHighRes` - url of the high resolution product image
- `related` - related products (also bought, also viewed, bought together, buy after viewing)
- `salesRank` - sales rank information
- `brand` - brand name
- `categories` - list of categories the product belongs to
- `tech1` - the first technical detail table of the product
- `tech2` - the second technical detail table of the product
- `similar_item` - similar product table
- $\dots$
```
def inspect_df(df: pd.DataFrame, n: int = 5) -> pd.DataFrame:
"""Helper method to easily inspect DataFrames."""
print(f"shape: {df.shape}")
return df.head(n)
def parse(filepath: str) -> Iterator[dict]:
file_obj = gzip.open(filepath, "rb")
for line in file_obj:
yield json.loads(line)
def file_to_dataframe(filepath: str) -> pd.DataFrame:
i = 0
df = {}
for d in parse(filepath):
df[i] = d
i += 1
return pd.DataFrame.from_dict(df, orient="index")
review_data = file_to_dataframe("data/Digital_Music_5.json.gz")
inspect_df(review_data)
list(review_data.columns)
review_data.loc[2]
metadata = file_to_dataframe("data/meta_Digital_Music.json.gz")
inspect_df(metadata)
list(metadata.columns)
metadata[metadata["asin"] == review_data.loc[2]["asin"]]
metadata["asin"].value_counts()
metadata.drop_duplicates(subset="asin", keep="first", inplace=True)
# for the content-based RecSys, we need both the review rating & title, description attrs - so an inner join
data = pd.merge(review_data, metadata, how="inner", on="asin")
inspect_df(data)
metadata[metadata["asin"] == "B000091JWJ"]
```
### Substep: EDA
```
data["created_at"] = pd.to_datetime(data["unixReviewTime"], unit="s")
data["created_at"] = pd.to_datetime(data["created_at"], format="%Y-%m")
(
ggplot(data, aes(x="overall"))
+ geom_bar(color="black")
+ labs(x="rating", title="distribution of ratings")
)
(
ggplot(data.groupby(pd.Grouper(key="created_at", freq="M")).count().reset_index())
+ geom_line(aes(x="created_at", y="asin"), color="crimson")
+ labs(x="date", y="count", title="#reviews: evolution through the years")
+ scale_x_datetime(breaks=date_breaks("6 months"), labels=date_format("%Y/%m"))
+ theme(axis_text_x=element_text(rotation=45, hjust=1), figure_size=(14, 7))
)
(
ggplot(
data.groupby([pd.Grouper(key="created_at", freq="M"), "overall"])
.count()
.reset_index()
)
+ geom_line(aes(x="created_at", y="asin"))
+ facet_wrap("overall", ncol=1)
+ labs(
x="date", y="count", title="per-rating #reviews: evolution through the years"
)
+ scale_x_datetime(breaks=date_breaks("6 months"), labels=date_format("%Y/%m"))
+ theme(axis_text_x=element_text(rotation=45, hjust=1), figure_size=(10, 7))
)
(
ggplot(data)
+ geom_smooth(aes(x="created_at", y="overall"))
+ labs(x="date", y="rating", title="rating trend through the years")
+ scale_x_datetime(breaks=date_breaks("6 months"), labels=date_format("%Y/%m"))
+ theme(axis_text_x=element_text(rotation=45, hjust=1), figure_size=(14, 7))
)
(
ggplot(data)
+ geom_smooth(
aes(x="created_at", y="overall"), method="mavg", method_args={"window": 10}
)
+ labs(x="date", y="rating", title="moving average of ratings")
+ scale_x_datetime(breaks=date_breaks("6 months"), labels=date_format("%Y/%m"))
+ theme(axis_text_x=element_text(rotation=45, hjust=1), figure_size=(14, 7))
)
```
## Step A.2
Our objective is to construct “item profiles” for the items, based on information available on their metadata.
```
content = data.copy()
content.drop_duplicates("asin", inplace=True)
content["title"].map(lambda x: isinstance(x, str)).value_counts()
content["description"].map(lambda x: isinstance(x, list)).value_counts()
content["description"]
content[content["description"].map(len) > 1]["description"]
content["rank"].map(lambda x: isinstance(x, list)).value_counts()
def concatenate_list_field(field: list) -> str:
if not isinstance(field, list):
return field
return " ".join(field)
def extract_rank(field: str) -> int:
found = re.search("[0-9]+(,[0-9]+)+", field)
rank = found.group(0) if found else None
return int(rank.replace(",", "")) if rank else None
transformer = FeatureUnion(
[
(
"title_preprocessing",
Pipeline(
[
(
"transform_field",
FunctionTransformer(lambda x: x["title"], validate=False),
),
(
"tfidf",
TfidfVectorizer(
ngram_range=(1, 2),
stop_words=stopwords.words("english"),
strip_accents="unicode",
),
),
]
),
),
(
"description_preprocessing",
Pipeline(
[
(
"transform_field",
FunctionTransformer(
lambda x: x["description"].map(concatenate_list_field),
validate=False,
),
),
(
"tfidf",
TfidfVectorizer(
ngram_range=(1, 2),
stop_words=stopwords.words("english"),
strip_accents="unicode",
),
),
]
),
),
(
"rank_preprocessing",
Pipeline(
[
(
"transform_field",
FunctionTransformer(
lambda x: x["rank"]
.map(concatenate_list_field)
.map(extract_rank)
.fillna(0)
.to_numpy()
.reshape((len(x.index), 1)),
validate=False,
),
),
],
),
),
]
)
transformer.fit(content)
title_vocab = transformer.transformer_list[0][1].steps[1][1].get_feature_names_out()
description_vocab = (
transformer.transformer_list[1][1].steps[1][1].get_feature_names_out()
)
len(title_vocab), len(description_vocab)
transformed_content = transformer.transform(content).toarray()
transformed_content.shape
```
## Step A.3
```
# note that the TF-IDF functionality in sklearn.feature_extraction.text can produce
# normalized vectors, in which case cosine_similarity is equivalent to linear_kernel,
# only slower.
cosine_sim_matrix = linear_kernel(transformed_content, transformed_content)
cosine_sim_matrix.shape
cosine_sim_matrix = pd.DataFrame(
cosine_sim_matrix, index=content["asin"], columns=content["asin"]
)
inspect_df(cosine_sim_matrix)
jaccard_sim_matrix = 1 - pairwise_distances(transformed_content, metric="jaccard")
jaccard_sim_matrix.shape
jaccard_sim_matrix = pd.DataFrame(
jaccard_sim_matrix, index=content["asin"], columns=content["asin"]
)
inspect_df(jaccard_sim_matrix)
```
## Step A.4
```
def top_n_products_per_user(data, n: int = 5):
return (
data.sort_values(["reviewerID", "overall"], ascending=False)
.groupby("reviewerID")
.head(n)
.groupby("reviewerID")["asin"]
.apply(list)
.reset_index()
)
top_5_products_per_user = top_n_products_per_user(content, n=5)
top_5_products_per_user
```
## Step A.5
```
def top_n_recommendations_per_user(
data: pd.DataFrame, sim_matrix: pd.DataFrame, n: int = 5
):
top_products_per_user = top_n_products_per_user(data, n)
recommendations = pd.DataFrame(columns=["reviewerID", "asin"])
recommendations["reviewerID"] = top_products_per_user["reviewerID"]
recommendations["asin"] = np.empty((len(recommendations.index), 0)).tolist()
for index, row in top_products_per_user.iterrows():
similar_movies = []
for column in sim_matrix.loc[row["asin"]].T:
similar_movies.extend(
sim_matrix.loc[row["asin"]]
.T[column]
.drop(
review_data.loc[review_data["reviewerID"] == row["reviewerID"]][
"asin"
].values,
errors="ignore",
)
.nlargest(n + 1)
.tail(n)
.index.values
)
recommendations.at[index, "asin"] = similar_movies
return recommendations
top_5_recommendations_per_user = top_n_recommendations_per_user(
data=content, sim_matrix=cosine_sim_matrix, n=5
)
top_5_recommendations_per_user
# keep also the entire ranking for each user, to be used later on the hybrid context
rankings_per_user = top_n_recommendations_per_user(
data=content, sim_matrix=cosine_sim_matrix, n=len(content["asin"].unique())
)
```
## Step A.6
```
def compare_recommendations_for_user(id_: str) -> None:
viewable_cols = ["title", "asin"]
content_based = content.loc[
content["asin"].isin(
top_5_recommendations_per_user.loc[
top_5_recommendations_per_user["reviewerID"] == id_
]["asin"].values[0]
)
]
naive = content.loc[
content["asin"].isin(
top_5_products_per_user.loc[top_5_products_per_user["reviewerID"] == id_][
"asin"
].values[0]
)
]
print("CONTENT-BASED\n")
print(content_based[viewable_cols])
print("=" * 60)
print("NAIVE\n")
print(naive[viewable_cols])
compare_recommendations_for_user("A15OXG4V7IK2D9")
compare_recommendations_for_user("A1352I3HWDQCZH")
compare_recommendations_for_user("AYPCUQS6ARWFH")
compare_recommendations_for_user("A17B6IPLJ964N0")
compare_recommendations_for_user("AWJ9J0JAHN6PQ")
compare_recommendations_for_user("AXOO7BPM22BDO")
compare_recommendations_for_user("A15GBKY0IPZJI3")
compare_recommendations_for_user("AWMORCDEUIBFO")
compare_recommendations_for_user("AWG2O9C42XW5G")
compare_recommendations_for_user("A15JTJXQXO22JJ")
top_5_products_per_user.to_pickle("data/naive-RS.pkl")
rankings_per_user.to_pickle("data/content-based-RS.pkl")
```
| github_jupyter |
```
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
from IPython.core.debugger import set_trace
import collections
import json
import math
import os
import random
import modeling
import optimization
import tokenization
import six
import os
import tensorflow as tf
import logging
logging.getLogger('tensorflow').disabled = True
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
import warnings
warnings.filterwarnings("ignore")
import time
from pandas import Series
from nltk.tokenize import sent_tokenize
import gensim.downloader as api
from gensim.parsing.preprocessing import remove_stopwords
word_vectors = api.load("glove-wiki-gigaword-100") # load pre-trained word-vectors from gensim-data
# flags = tf.flags
# FLAGS = flags.FLAGS
# ## Required parameters
# flags.DEFINE_string(
# "bert_config_file", None,
# "The config json file corresponding to the pre-trained BERT model. "
# "This specifies the model architecture.")
# flags.DEFINE_string("vocab_file", None,
# "The vocabulary file that the BERT model was trained on.")
# flags.DEFINE_string(
# "output_dir", None,
# "The output directory where the model checkpoints will be written.")
# ## Other parameters
# flags.DEFINE_string("train_file", None,
# "SQuAD json for training. E.g., train-v1.1.json")
# flags.DEFINE_string(
# "predict_file", None,
# "SQuAD json for predictions. E.g., dev-v1.1.json or test-v1.1.json")
# flags.DEFINE_string(
# "init_checkpoint", None,
# "Initial checkpoint (usually from a pre-trained BERT model).")
# flags.DEFINE_bool(
# "do_lower_case", True,
# "Whether to lower case the input text. Should be True for uncased "
# "models and False for cased models.")
# flags.DEFINE_integer(
# "max_seq_length", 384,
# "The maximum total input sequence length after WordPiece tokenization. "
# "Sequences longer than this will be truncated, and sequences shorter "
# "than this will be padded.")
# flags.DEFINE_integer(
# "doc_stride", 128,
# "When splitting up a long document into chunks, how much stride to "
# "take between chunks.")
# flags.DEFINE_integer(
# "max_query_length", 64,
# "The maximum number of tokens for the question. Questions longer than "
# "this will be truncated to this length.")
# flags.DEFINE_bool("do_train", False, "Whether to run training.")
# flags.DEFINE_bool("do_predict", False, "Whether to run eval on the dev set.")
# flags.DEFINE_integer("train_batch_size", 32, "Total batch size for training.")
# flags.DEFINE_integer("predict_batch_size", 8,
# "Total batch size for predictions.")
# flags.DEFINE_float("learning_rate", 5e-5, "The initial learning rate for Adam.")
# flags.DEFINE_float("num_train_epochs", 3.0,
# "Total number of training epochs to perform.")
# flags.DEFINE_float(
# "warmup_proportion", 0.1,
# "Proportion of training to perform linear learning rate warmup for. "
# "E.g., 0.1 = 10% of training.")
# flags.DEFINE_integer("save_checkpoints_steps", 1000,
# "How often to save the model checkpoint.")
# flags.DEFINE_integer("iterations_per_loop", 1000,
# "How many steps to make in each estimator call.")
# flags.DEFINE_integer(
# "n_best_size", 20,
# "The total number of n-best predictions to generate in the "
# "nbest_predictions.json output file.")
# flags.DEFINE_integer(
# "max_answer_length", 30,
# "The maximum length of an answer that can be generated. This is needed "
# "because the start and end predictions are not conditioned on one another.")
# flags.DEFINE_bool("use_tpu", False, "Whether to use TPU or GPU/CPU.")
# tf.flags.DEFINE_string(
# "tpu_name", None,
# "The Cloud TPU to use for training. This should be either the name "
# "used when creating the Cloud TPU, or a grpc://ip.address.of.tpu:8470 "
# "url.")
# tf.flags.DEFINE_string(
# "tpu_zone", None,
# "[Optional] GCE zone where the Cloud TPU is located in. If not "
# "specified, we will attempt to automatically detect the GCE project from "
# "metadata.")
# tf.flags.DEFINE_string(
# "gcp_project", None,
# "[Optional] Project name for the Cloud TPU-enabled project. If not "
# "specified, we will attempt to automatically detect the GCE project from "
# "metadata.")
# tf.flags.DEFINE_string("master", None, "[Optional] TensorFlow master URL.")
# flags.DEFINE_integer(
# "num_tpu_cores", 8,
# "Only used if `use_tpu` is True. Total number of TPU cores to use.")
# flags.DEFINE_bool(
# "verbose_logging", False,
# "If true, all of the warnings related to data processing will be printed. "
# "A number of warnings are expected for a normal SQuAD evaluation.")
# flags.DEFINE_bool(
# "version_2_with_negative", False,
# "If true, the SQuAD examples contain some that do not have an answer.")
# flags.DEFINE_float(
# "null_score_diff_threshold", 0.0,
# "If null_score - best_non_null is greater than the threshold predict null.")
# tf.flags.DEFINE_bool(
# "interact", False,
# "to Initiate interactive question and answers chatbot")
# tf.flags.DEFINE_string(
# "context", None,
# "context file path of Text summary to feed the BERT model ")
# tf.flags.DEFINE_string(
# "question", None,
# "Questioning sentence for the Text summary ")
class Flags:
def __init__(self):
self.bert_config_file = r'BERT_LARGE_DIR/bert_config.json'
self.vocab_file = r'BERT_LARGE_DIR/vocab.txt'
self.output_dir = 'squad_base'
self.train_file = 'train-v1.2.json'
self.predict_file = 'dev-v1.4.json'
self.init_checkpoint = r'BERT_LARGE_DIR/bert_model.ckpt'
self.do_lower_case = True
self.max_seq_length = 384
self.doc_stride = 128
self.max_query_length = 64
self.do_train = False
self.do_predict = True
self.train_batch_size = 12
self.predict_batch_size = 8
self.learning_rate = 3e-5
self.num_train_epochs = 3
self.warmup_proportion = 0.1
self.save_checkpoints_steps = 1000
self.iterations_per_loop = 1000
self.n_best_size = 20
self.max_answer_length = 30
self.use_tpu = False
self.tpu_name = None
self.tpu_zone = None
self.gcp_project = None
self.master = None
self.num_tpu_cores = 8
self.verbose_logging = False
self.version_2_with_negative = False
self.null_score_diff_threshold = 0.0
self.context = None
self.question = None
self.interact = True
FLAGS = Flags()
class SquadExample(object):
"""A single training/test example for simple sequence classification.
For examples without an answer, the start and end position are -1.
"""
def __init__(self,
qas_id,
question_text,
doc_tokens,
orig_answer_text=None,
start_position=None,
end_position=None,
is_impossible=False):
self.qas_id = qas_id
self.question_text = question_text
self.doc_tokens = doc_tokens
self.orig_answer_text = orig_answer_text
self.start_position = start_position
self.end_position = end_position
self.is_impossible = is_impossible
def __str__(self):
return self.__repr__()
def __repr__(self):
s = ""
s += "qas_id: %s" % (tokenization.printable_text(self.qas_id))
s += ", question_text: %s" % (
tokenization.printable_text(self.question_text))
s += ", doc_tokens: [%s]" % (" ".join(self.doc_tokens))
if self.start_position:
s += ", start_position: %d" % (self.start_position)
if self.start_position:
s += ", end_position: %d" % (self.end_position)
if self.start_position:
s += ", is_impossible: %r" % (self.is_impossible)
return s
class InputFeatures(object):
"""A single set of features of data."""
def __init__(self,
unique_id,
example_index,
doc_span_index,
tokens,
token_to_orig_map,
token_is_max_context,
input_ids,
input_mask,
segment_ids,
start_position=None,
end_position=None,
is_impossible=None):
self.unique_id = unique_id
self.example_index = example_index
self.doc_span_index = doc_span_index
self.tokens = tokens
self.token_to_orig_map = token_to_orig_map
self.token_is_max_context = token_is_max_context
self.input_ids = input_ids
self.input_mask = input_mask
self.segment_ids = segment_ids
self.start_position = start_position
self.end_position = end_position
self.is_impossible = is_impossible
def read_squad_examples(data, is_training):
"""Read a SQuAD json file into a list of SquadExample."""
# with tf.gfile.Open(input_file, "r") as reader:
# input_data = json.load(reader)["data"]
input_data = data['data']
def is_whitespace(c):
if c == " " or c == "\t" or c == "\r" or c == "\n" or ord(c) == 0x202F:
return True
return False
examples = []
for entry in input_data[:2]:
for paragraph in entry["paragraphs"]:
paragraph_text = paragraph["context"]
doc_tokens = []
char_to_word_offset = []
prev_is_whitespace = True
for c in paragraph_text:
if is_whitespace(c):
prev_is_whitespace = True
else:
if prev_is_whitespace:
doc_tokens.append(c)
else:
doc_tokens[-1] += c
prev_is_whitespace = False
char_to_word_offset.append(len(doc_tokens) - 1)
for qa in paragraph["qas"]:
qas_id = qa["id"]
question_text = qa["question"]
start_position = None
end_position = None
orig_answer_text = None
is_impossible = False
if is_training:
if FLAGS.version_2_with_negative:
is_impossible = qa["is_impossible"]
if (len(qa["answers"]) != 1) and (not is_impossible):
raise ValueError(
"For training, each question should have exactly 1 answer.")
if not is_impossible:
answer = qa["answers"][0]
orig_answer_text = answer["text"]
answer_offset = answer["answer_start"]
answer_length = len(orig_answer_text)
start_position = char_to_word_offset[answer_offset]
end_position = char_to_word_offset[answer_offset + answer_length -
1]
# Only add answers where the text can be exactly recovered from the
# document. If this CAN'T happen it's likely due to weird Unicode
# stuff so we will just skip the example.
#
# Note that this means for training mode, every example is NOT
# guaranteed to be preserved.
actual_text = " ".join(
doc_tokens[start_position:(end_position + 1)])
cleaned_answer_text = " ".join(
tokenization.whitespace_tokenize(orig_answer_text))
if actual_text.find(cleaned_answer_text) == -1:
# tf.logging.warning("Could not find answer: '%s' vs. '%s'",
# actual_text, cleaned_answer_text)
continue
else:
start_position = -1
end_position = -1
orig_answer_text = ""
example = SquadExample(
qas_id=qas_id,
question_text=question_text,
doc_tokens=doc_tokens,
orig_answer_text=orig_answer_text,
start_position=start_position,
end_position=end_position,
is_impossible=is_impossible)
examples.append(example)
return examples
def convert_examples_to_features(examples, tokenizer, max_seq_length,
doc_stride, max_query_length, is_training,
output_fn):
"""Loads a data file into a list of `InputBatch`s."""
unique_id = 1000000000
for (example_index, example) in enumerate(examples):
query_tokens = tokenizer.tokenize(example.question_text)
if len(query_tokens) > max_query_length:
query_tokens = query_tokens[0:max_query_length]
tok_to_orig_index = []
orig_to_tok_index = []
all_doc_tokens = []
for (i, token) in enumerate(example.doc_tokens):
orig_to_tok_index.append(len(all_doc_tokens))
sub_tokens = tokenizer.tokenize(token)
for sub_token in sub_tokens:
tok_to_orig_index.append(i)
all_doc_tokens.append(sub_token)
tok_start_position = None
tok_end_position = None
if is_training and example.is_impossible:
tok_start_position = -1
tok_end_position = -1
if is_training and not example.is_impossible:
tok_start_position = orig_to_tok_index[example.start_position]
if example.end_position < len(example.doc_tokens) - 1:
tok_end_position = orig_to_tok_index[example.end_position + 1] - 1
else:
tok_end_position = len(all_doc_tokens) - 1
(tok_start_position, tok_end_position) = _improve_answer_span(
all_doc_tokens, tok_start_position, tok_end_position, tokenizer,
example.orig_answer_text)
# The -3 accounts for [CLS], [SEP] and [SEP]
max_tokens_for_doc = max_seq_length - len(query_tokens) - 3
# We can have documents that are longer than the maximum sequence length.
# To deal with this we do a sliding window approach, where we take chunks
# of the up to our max length with a stride of `doc_stride`.
_DocSpan = collections.namedtuple( # pylint: disable=invalid-name
"DocSpan", ["start", "length"])
doc_spans = []
start_offset = 0
while start_offset < len(all_doc_tokens):
length = len(all_doc_tokens) - start_offset
if length > max_tokens_for_doc:
length = max_tokens_for_doc
doc_spans.append(_DocSpan(start=start_offset, length=length))
if start_offset + length == len(all_doc_tokens):
break
start_offset += min(length, doc_stride)
for (doc_span_index, doc_span) in enumerate(doc_spans):
tokens = []
token_to_orig_map = {}
token_is_max_context = {}
segment_ids = []
tokens.append("[CLS]")
segment_ids.append(0)
for token in query_tokens:
tokens.append(token)
segment_ids.append(0)
tokens.append("[SEP]")
segment_ids.append(0)
for i in range(doc_span.length):
split_token_index = doc_span.start + i
token_to_orig_map[len(tokens)] = tok_to_orig_index[split_token_index]
is_max_context = _check_is_max_context(doc_spans, doc_span_index,
split_token_index)
token_is_max_context[len(tokens)] = is_max_context
tokens.append(all_doc_tokens[split_token_index])
segment_ids.append(1)
tokens.append("[SEP]")
segment_ids.append(1)
input_ids = tokenizer.convert_tokens_to_ids(tokens)
# The mask has 1 for real tokens and 0 for padding tokens. Only real
# tokens are attended to.
input_mask = [1] * len(input_ids)
# Zero-pad up to the sequence length.
while len(input_ids) < max_seq_length:
input_ids.append(0)
input_mask.append(0)
segment_ids.append(0)
assert len(input_ids) == max_seq_length
assert len(input_mask) == max_seq_length
assert len(segment_ids) == max_seq_length
start_position = None
end_position = None
if is_training and not example.is_impossible:
# For training, if our document chunk does not contain an annotation
# we throw it out, since there is nothing to predict.
doc_start = doc_span.start
doc_end = doc_span.start + doc_span.length - 1
out_of_span = False
if not (tok_start_position >= doc_start and
tok_end_position <= doc_end):
out_of_span = True
if out_of_span:
start_position = 0
end_position = 0
else:
doc_offset = len(query_tokens) + 2
start_position = tok_start_position - doc_start + doc_offset
end_position = tok_end_position - doc_start + doc_offset
if is_training and example.is_impossible:
start_position = 0
end_position = 0
# if example_index < 20:
# tf.logging.info("*** Example ***")
# tf.logging.info("unique_id: %s" % (unique_id))
# tf.logging.info("example_index: %s" % (example_index))
# tf.logging.info("doc_span_index: %s" % (doc_span_index))
# tf.logging.info("tokens: %s" % " ".join(
# [tokenization.printable_text(x) for x in tokens]))
# tf.logging.info("token_to_orig_map: %s" % " ".join(
# ["%d:%d" % (x, y) for (x, y) in six.iteritems(token_to_orig_map)]))
# tf.logging.info("token_is_max_context: %s" % " ".join([
# "%d:%s" % (x, y) for (x, y) in six.iteritems(token_is_max_context)
# ]))
# tf.logging.info("input_ids: %s" % " ".join([str(x) for x in input_ids]))
# tf.logging.info(
# "input_mask: %s" % " ".join([str(x) for x in input_mask]))
# tf.logging.info(
# "segment_ids: %s" % " ".join([str(x) for x in segment_ids]))
# if is_training and example.is_impossible:
# tf.logging.info("impossible example")
# if is_training and not example.is_impossible:
# answer_text = " ".join(tokens[start_position:(end_position + 1)])
# tf.logging.info("start_position: %d" % (start_position))
# tf.logging.info("end_position: %d" % (end_position))
# tf.logging.info(
# "answer: %s" % (tokenization.printable_text(answer_text)))
feature = InputFeatures(
unique_id=unique_id,
example_index=example_index,
doc_span_index=doc_span_index,
tokens=tokens,
token_to_orig_map=token_to_orig_map,
token_is_max_context=token_is_max_context,
input_ids=input_ids,
input_mask=input_mask,
segment_ids=segment_ids,
start_position=start_position,
end_position=end_position,
is_impossible=example.is_impossible)
# Run callback
output_fn(feature)
unique_id += 1
def _improve_answer_span(doc_tokens, input_start, input_end, tokenizer,
orig_answer_text):
"""Returns tokenized answer spans that better match the annotated answer."""
# The SQuAD annotations are character based. We first project them to
# whitespace-tokenized words. But then after WordPiece tokenization, we can
# often find a "better match". For example:
#
# Question: What year was John Smith born?
# Context: The leader was John Smith (1895-1943).
# Answer: 1895
#
# The original whitespace-tokenized answer will be "(1895-1943).". However
# after tokenization, our tokens will be "( 1895 - 1943 ) .". So we can match
# the exact answer, 1895.
#
# However, this is not always possible. Consider the following:
#
# Question: What country is the top exporter of electornics?
# Context: The Japanese electronics industry is the lagest in the world.
# Answer: Japan
#
# In this case, the annotator chose "Japan" as a character sub-span of
# the word "Japanese". Since our WordPiece tokenizer does not split
# "Japanese", we just use "Japanese" as the annotation. This is fairly rare
# in SQuAD, but does happen.
tok_answer_text = " ".join(tokenizer.tokenize(orig_answer_text))
for new_start in range(input_start, input_end + 1):
for new_end in range(input_end, new_start - 1, -1):
text_span = " ".join(doc_tokens[new_start:(new_end + 1)])
if text_span == tok_answer_text:
return (new_start, new_end)
return (input_start, input_end)
def _check_is_max_context(doc_spans, cur_span_index, position):
"""Check if this is the 'max context' doc span for the token."""
# Because of the sliding window approach taken to scoring documents, a single
# token can appear in multiple documents. E.g.
# Doc: the man went to the store and bought a gallon of milk
# Span A: the man went to the
# Span B: to the store and bought
# Span C: and bought a gallon of
# ...
#
# Now the word 'bought' will have two scores from spans B and C. We only
# want to consider the score with "maximum context", which we define as
# the *minimum* of its left and right context (the *sum* of left and
# right context will always be the same, of course).
#
# In the example the maximum context for 'bought' would be span C since
# it has 1 left context and 3 right context, while span B has 4 left context
# and 0 right context.
best_score = None
best_span_index = None
for (span_index, doc_span) in enumerate(doc_spans):
end = doc_span.start + doc_span.length - 1
if position < doc_span.start:
continue
if position > end:
continue
num_left_context = position - doc_span.start
num_right_context = end - position
score = min(num_left_context, num_right_context) + 0.01 * doc_span.length
if best_score is None or score > best_score:
best_score = score
best_span_index = span_index
return cur_span_index == best_span_index
def create_model(bert_config, is_training, input_ids, input_mask, segment_ids,
use_one_hot_embeddings):
"""Creates a classification model."""
model = modeling.BertModel(
config=bert_config,
is_training=is_training,
input_ids=input_ids,
input_mask=input_mask,
token_type_ids=segment_ids,
use_one_hot_embeddings=use_one_hot_embeddings)
final_hidden = model.get_sequence_output()
final_hidden_shape = modeling.get_shape_list(final_hidden, expected_rank=3)
batch_size = final_hidden_shape[0]
seq_length = final_hidden_shape[1]
hidden_size = final_hidden_shape[2]
output_weights = tf.get_variable(
"cls/squad/output_weights", [2, hidden_size],
initializer=tf.truncated_normal_initializer(stddev=0.02))
output_bias = tf.get_variable(
"cls/squad/output_bias", [2], initializer=tf.zeros_initializer())
final_hidden_matrix = tf.reshape(final_hidden,
[batch_size * seq_length, hidden_size])
logits = tf.matmul(final_hidden_matrix, output_weights, transpose_b=True)
logits = tf.nn.bias_add(logits, output_bias)
logits = tf.reshape(logits, [batch_size, seq_length, 2])
logits = tf.transpose(logits, [2, 0, 1])
unstacked_logits = tf.unstack(logits, axis=0)
(start_logits, end_logits) = (unstacked_logits[0], unstacked_logits[1])
return (start_logits, end_logits)
def model_fn_builder(bert_config, init_checkpoint, learning_rate,
num_train_steps, num_warmup_steps, use_tpu,
use_one_hot_embeddings):
"""Returns `model_fn` closure for TPUEstimator."""
def model_fn(features, labels, mode, params): # pylint: disable=unused-argument
"""The `model_fn` for TPUEstimator."""
# tf.logging.info("*** Features ***")
# for name in sorted(features.keys()):
# tf.logging.info(" name = %s, shape = %s" % (name, features[name].shape))
unique_ids = features["unique_ids"]
input_ids = features["input_ids"]
input_mask = features["input_mask"]
segment_ids = features["segment_ids"]
is_training = (mode == tf.estimator.ModeKeys.TRAIN)
(start_logits, end_logits) = create_model(
bert_config=bert_config,
is_training=is_training,
input_ids=input_ids,
input_mask=input_mask,
segment_ids=segment_ids,
use_one_hot_embeddings=use_one_hot_embeddings)
tvars = tf.trainable_variables()
initialized_variable_names = {}
scaffold_fn = None
if init_checkpoint:
(assignment_map, initialized_variable_names
) = modeling.get_assignment_map_from_checkpoint(tvars, init_checkpoint)
if use_tpu:
def tpu_scaffold():
tf.train.init_from_checkpoint(init_checkpoint, assignment_map)
return tf.train.Scaffold()
scaffold_fn = tpu_scaffold
else:
tf.train.init_from_checkpoint(init_checkpoint, assignment_map)
# tf.logging.info("**** Trainable Variables ****")
for var in tvars:
init_string = ""
if var.name in initialized_variable_names:
init_string = ", *INIT_FROM_CKPT*"
# tf.logging.info(" name = %s, shape = %s%s", var.name, var.shape,
# init_string)
output_spec = None
if mode == tf.estimator.ModeKeys.TRAIN:
seq_length = modeling.get_shape_list(input_ids)[1]
def compute_loss(logits, positions):
one_hot_positions = tf.one_hot(
positions, depth=seq_length, dtype=tf.float32)
log_probs = tf.nn.log_softmax(logits, axis=-1)
loss = -tf.reduce_mean(
tf.reduce_sum(one_hot_positions * log_probs, axis=-1))
return loss
start_positions = features["start_positions"]
end_positions = features["end_positions"]
start_loss = compute_loss(start_logits, start_positions)
end_loss = compute_loss(end_logits, end_positions)
total_loss = (start_loss + end_loss) / 2.0
train_op = optimization.create_optimizer(
total_loss, learning_rate, num_train_steps, num_warmup_steps, use_tpu)
output_spec = tf.contrib.tpu.TPUEstimatorSpec(
mode=mode,
loss=total_loss,
train_op=train_op,
scaffold_fn=scaffold_fn)
elif mode == tf.estimator.ModeKeys.PREDICT:
predictions = {
"unique_ids": unique_ids,
"start_logits": start_logits,
"end_logits": end_logits,
}
output_spec = tf.contrib.tpu.TPUEstimatorSpec(
mode=mode, predictions=predictions, scaffold_fn=scaffold_fn)
else:
raise ValueError(
"Only TRAIN and PREDICT modes are supported: %s" % (mode))
return output_spec
return model_fn
def input_fn_builder(input_file, seq_length, is_training, drop_remainder):
"""Creates an `input_fn` closure to be passed to TPUEstimator."""
name_to_features = {
"unique_ids": tf.FixedLenFeature([], tf.int64),
"input_ids": tf.FixedLenFeature([seq_length], tf.int64),
"input_mask": tf.FixedLenFeature([seq_length], tf.int64),
"segment_ids": tf.FixedLenFeature([seq_length], tf.int64),
}
if is_training:
name_to_features["start_positions"] = tf.FixedLenFeature([], tf.int64)
name_to_features["end_positions"] = tf.FixedLenFeature([], tf.int64)
def _decode_record(record, name_to_features):
"""Decodes a record to a TensorFlow example."""
example = tf.parse_single_example(record, name_to_features)
# tf.Example only supports tf.int64, but the TPU only supports tf.int32.
# So cast all int64 to int32.
for name in list(example.keys()):
t = example[name]
if t.dtype == tf.int64:
t = tf.to_int32(t)
example[name] = t
return example
def input_fn(params):
"""The actual input function."""
batch_size = params["batch_size"]
# For training, we want a lot of parallel reading and shuffling.
# For eval, we want no shuffling and parallel reading doesn't matter.
d = tf.data.TFRecordDataset(input_file)
if is_training:
d = d.repeat()
d = d.shuffle(buffer_size=100)
d = d.apply(
tf.contrib.data.map_and_batch(
lambda record: _decode_record(record, name_to_features),
batch_size=batch_size,
drop_remainder=drop_remainder))
return d
return input_fn
RawResult = collections.namedtuple("RawResult",
["unique_id", "start_logits", "end_logits"])
def write_predictions(all_examples, all_features, all_results, n_best_size,
max_answer_length, do_lower_case, output_prediction_file,
output_nbest_file, output_null_log_odds_file):
"""Write final predictions to the json file and log-odds of null if needed."""
# tf.logging.info("Writing predictions to: %s" % (output_prediction_file))
# tf.logging.info("Writing nbest to: %s" % (output_nbest_file))
example_index_to_features = collections.defaultdict(list)
for feature in all_features:
example_index_to_features[feature.example_index].append(feature)
unique_id_to_result = {}
for result in all_results:
unique_id_to_result[result.unique_id] = result
_PrelimPrediction = collections.namedtuple( # pylint: disable=invalid-name
"PrelimPrediction",
["feature_index", "start_index", "end_index", "start_logit", "end_logit"])
all_predictions = collections.OrderedDict()
all_nbest_json = collections.OrderedDict()
scores_diff_json = collections.OrderedDict()
for (example_index, example) in enumerate(all_examples):
features = example_index_to_features[example_index]
prelim_predictions = []
# keep track of the minimum score of null start+end of position 0
score_null = 1000000 # large and positive
min_null_feature_index = 0 # the paragraph slice with min mull score
null_start_logit = 0 # the start logit at the slice with min null score
null_end_logit = 0 # the end logit at the slice with min null score
for (feature_index, feature) in enumerate(features):
result = unique_id_to_result[feature.unique_id]
start_indexes = _get_best_indexes(result.start_logits, n_best_size)
end_indexes = _get_best_indexes(result.end_logits, n_best_size)
# if we could have irrelevant answers, get the min score of irrelevant
if FLAGS.version_2_with_negative:
feature_null_score = result.start_logits[0] + result.end_logits[0]
if feature_null_score < score_null:
score_null = feature_null_score
min_null_feature_index = feature_index
null_start_logit = result.start_logits[0]
null_end_logit = result.end_logits[0]
for start_index in start_indexes:
for end_index in end_indexes:
# We could hypothetically create invalid predictions, e.g., predict
# that the start of the span is in the question. We throw out all
# invalid predictions.
if start_index >= len(feature.tokens):
continue
if end_index >= len(feature.tokens):
continue
if start_index not in feature.token_to_orig_map:
continue
if end_index not in feature.token_to_orig_map:
continue
if not feature.token_is_max_context.get(start_index, False):
continue
if end_index < start_index:
continue
length = end_index - start_index + 1
if length > max_answer_length:
continue
prelim_predictions.append(
_PrelimPrediction(
feature_index=feature_index,
start_index=start_index,
end_index=end_index,
start_logit=result.start_logits[start_index],
end_logit=result.end_logits[end_index]))
if FLAGS.version_2_with_negative:
prelim_predictions.append(
_PrelimPrediction(
feature_index=min_null_feature_index,
start_index=0,
end_index=0,
start_logit=null_start_logit,
end_logit=null_end_logit))
prelim_predictions = sorted(
prelim_predictions,
key=lambda x: (x.start_logit + x.end_logit),
reverse=True)
_NbestPrediction = collections.namedtuple( # pylint: disable=invalid-name
"NbestPrediction", ["text", "start_logit", "end_logit"])
seen_predictions = {}
nbest = []
for pred in prelim_predictions:
if len(nbest) >= n_best_size:
break
feature = features[pred.feature_index]
if pred.start_index > 0: # this is a non-null prediction
tok_tokens = feature.tokens[pred.start_index:(pred.end_index + 1)]
orig_doc_start = feature.token_to_orig_map[pred.start_index]
orig_doc_end = feature.token_to_orig_map[pred.end_index]
orig_tokens = example.doc_tokens[orig_doc_start:(orig_doc_end + 1)]
tok_text = " ".join(tok_tokens)
# De-tokenize WordPieces that have been split off.
tok_text = tok_text.replace(" ##", "")
tok_text = tok_text.replace("##", "")
# Clean whitespace
tok_text = tok_text.strip()
tok_text = " ".join(tok_text.split())
orig_text = " ".join(orig_tokens)
final_text = get_final_text(tok_text, orig_text, do_lower_case)
if final_text in seen_predictions:
continue
seen_predictions[final_text] = True
else:
final_text = ""
seen_predictions[final_text] = True
nbest.append(
_NbestPrediction(
text=final_text,
start_logit=pred.start_logit,
end_logit=pred.end_logit))
# if we didn't inlude the empty option in the n-best, inlcude it
if FLAGS.version_2_with_negative:
if "" not in seen_predictions:
nbest.append(
_NbestPrediction(
text="", start_logit=null_start_logit,
end_logit=null_end_logit))
# In very rare edge cases we could have no valid predictions. So we
# just create a nonce prediction in this case to avoid failure.
if not nbest:
nbest.append(
_NbestPrediction(text="empty", start_logit=0.0, end_logit=0.0))
assert len(nbest) >= 1
total_scores = []
best_non_null_entry = None
for entry in nbest:
total_scores.append(entry.start_logit + entry.end_logit)
if not best_non_null_entry:
if entry.text:
best_non_null_entry = entry
probs = _compute_softmax(total_scores)
nbest_json = []
for (i, entry) in enumerate(nbest):
output = collections.OrderedDict()
output["text"] = entry.text
output["probability"] = probs[i]
output["start_logit"] = entry.start_logit
output["end_logit"] = entry.end_logit
nbest_json.append(output)
assert len(nbest_json) >= 1
if not FLAGS.version_2_with_negative:
all_predictions[example.qas_id] = nbest_json[0]["text"]
else:
# predict "" iff the null score - the score of best non-null > threshold
score_diff = score_null - best_non_null_entry.start_logit - (
best_non_null_entry.end_logit)
scores_diff_json[example.qas_id] = score_diff
if score_diff > FLAGS.null_score_diff_threshold:
all_predictions[example.qas_id] = ""
else:
all_predictions[example.qas_id] = best_non_null_entry.text
all_nbest_json[example.qas_id] = nbest_json
if FLAGS.interact:
return all_predictions
with tf.gfile.GFile(output_prediction_file, "w") as writer:
writer.write(json.dumps(all_predictions, indent=4) + "\n")
with tf.gfile.GFile(output_nbest_file, "w") as writer:
writer.write(json.dumps(all_nbest_json, indent=4) + "\n")
if FLAGS.version_2_with_negative:
with tf.gfile.GFile(output_null_log_odds_file, "w") as writer:
writer.write(json.dumps(scores_diff_json, indent=4) + "\n")
def get_final_text(pred_text, orig_text, do_lower_case):
"""Project the tokenized prediction back to the original text."""
# When we created the data, we kept track of the alignment between original
# (whitespace tokenized) tokens and our WordPiece tokenized tokens. So
# now `orig_text` contains the span of our original text corresponding to the
# span that we predicted.
#
# However, `orig_text` may contain extra characters that we don't want in
# our prediction.
#
# For example, let's say:
# pred_text = steve smith
# orig_text = Steve Smith's
#
# We don't want to return `orig_text` because it contains the extra "'s".
#
# We don't want to return `pred_text` because it's already been normalized
# (the SQuAD eval script also does punctuation stripping/lower casing but
# our tokenizer does additional normalization like stripping accent
# characters).
#
# What we really want to return is "Steve Smith".
#
# Therefore, we have to apply a semi-complicated alignment heruistic between
# `pred_text` and `orig_text` to get a character-to-charcter alignment. This
# can fail in certain cases in which case we just return `orig_text`.
def _strip_spaces(text):
ns_chars = []
ns_to_s_map = collections.OrderedDict()
for (i, c) in enumerate(text):
if c == " ":
continue
ns_to_s_map[len(ns_chars)] = i
ns_chars.append(c)
ns_text = "".join(ns_chars)
return (ns_text, ns_to_s_map)
# We first tokenize `orig_text`, strip whitespace from the result
# and `pred_text`, and check if they are the same length. If they are
# NOT the same length, the heuristic has failed. If they are the same
# length, we assume the characters are one-to-one aligned.
tokenizer = tokenization.BasicTokenizer(do_lower_case=do_lower_case)
tok_text = " ".join(tokenizer.tokenize(orig_text))
start_position = tok_text.find(pred_text)
if start_position == -1:
# if FLAGS.verbose_logging:
# tf.logging.info(
# "Unable to find text: '%s' in '%s'" % (pred_text, orig_text))
return orig_text
end_position = start_position + len(pred_text) - 1
(orig_ns_text, orig_ns_to_s_map) = _strip_spaces(orig_text)
(tok_ns_text, tok_ns_to_s_map) = _strip_spaces(tok_text)
if len(orig_ns_text) != len(tok_ns_text):
# if FLAGS.verbose_logging:
# tf.logging.info("Length not equal after stripping spaces: '%s' vs '%s'",
# orig_ns_text, tok_ns_text)
return orig_text
# We then project the characters in `pred_text` back to `orig_text` using
# the character-to-character alignment.
tok_s_to_ns_map = {}
for (i, tok_index) in six.iteritems(tok_ns_to_s_map):
tok_s_to_ns_map[tok_index] = i
orig_start_position = None
if start_position in tok_s_to_ns_map:
ns_start_position = tok_s_to_ns_map[start_position]
if ns_start_position in orig_ns_to_s_map:
orig_start_position = orig_ns_to_s_map[ns_start_position]
if orig_start_position is None:
# if FLAGS.verbose_logging:
# tf.logging.info("Couldn't map start position")
return orig_text
orig_end_position = None
if end_position in tok_s_to_ns_map:
ns_end_position = tok_s_to_ns_map[end_position]
if ns_end_position in orig_ns_to_s_map:
orig_end_position = orig_ns_to_s_map[ns_end_position]
if orig_end_position is None:
# if FLAGS.verbose_logging:
# tf.logging.info("Couldn't map end position")
return orig_text
output_text = orig_text[orig_start_position:(orig_end_position + 1)]
return output_text
def _get_best_indexes(logits, n_best_size):
"""Get the n-best logits from a list."""
index_and_score = sorted(enumerate(logits), key=lambda x: x[1], reverse=True)
best_indexes = []
for i in range(len(index_and_score)):
if i >= n_best_size:
break
best_indexes.append(index_and_score[i][0])
return best_indexes
def _compute_softmax(scores):
"""Compute softmax probability over raw logits."""
if not scores:
return []
max_score = None
for score in scores:
if max_score is None or score > max_score:
max_score = score
exp_scores = []
total_sum = 0.0
for score in scores:
x = math.exp(score - max_score)
exp_scores.append(x)
total_sum += x
probs = []
for score in exp_scores:
probs.append(score / total_sum)
return probs
class FeatureWriter(object):
"""Writes InputFeature to TF example file."""
def __init__(self, filename, is_training):
self.filename = filename
self.is_training = is_training
self.num_features = 0
self._writer = tf.python_io.TFRecordWriter(filename)
def process_feature(self, feature):
"""Write a InputFeature to the TFRecordWriter as a tf.train.Example."""
self.num_features += 1
def create_int_feature(values):
feature = tf.train.Feature(
int64_list=tf.train.Int64List(value=list(values)))
return feature
features = collections.OrderedDict()
features["unique_ids"] = create_int_feature([feature.unique_id])
features["input_ids"] = create_int_feature(feature.input_ids)
features["input_mask"] = create_int_feature(feature.input_mask)
features["segment_ids"] = create_int_feature(feature.segment_ids)
if self.is_training:
features["start_positions"] = create_int_feature([feature.start_position])
features["end_positions"] = create_int_feature([feature.end_position])
impossible = 0
if feature.is_impossible:
impossible = 1
features["is_impossible"] = create_int_feature([impossible])
tf_example = tf.train.Example(features=tf.train.Features(feature=features))
self._writer.write(tf_example.SerializeToString())
def close(self):
self._writer.close()
def validate_flags_or_throw(bert_config):
"""Validate the input FLAGS or throw an exception."""
tokenization.validate_case_matches_checkpoint(FLAGS.do_lower_case,
FLAGS.init_checkpoint)
if not FLAGS.do_train and not FLAGS.do_predict:
raise ValueError("At least one of `do_train` or `do_predict` must be True.")
if FLAGS.do_train:
if not FLAGS.train_file:
raise ValueError(
"If `do_train` is True, then `train_file` must be specified.")
if FLAGS.do_predict:
if not FLAGS.predict_file:
raise ValueError(
"If `do_predict` is True, then `predict_file` must be specified.")
if FLAGS.max_seq_length > bert_config.max_position_embeddings:
raise ValueError(
"Cannot use sequence length %d because the BERT model "
"was only trained up to sequence length %d" %
(FLAGS.max_seq_length, bert_config.max_position_embeddings))
if FLAGS.max_seq_length <= FLAGS.max_query_length + 3:
raise ValueError(
"The max_seq_length (%d) must be greater than max_query_length "
"(%d) + 3" % (FLAGS.max_seq_length, FLAGS.max_query_length))
def initializingModels():
# tf.logging.set_verbosity(tf.logging.INFO)
bert_config = modeling.BertConfig.from_json_file(FLAGS.bert_config_file)
validate_flags_or_throw(bert_config)
tf.gfile.MakeDirs(FLAGS.output_dir)
tokenizer = tokenization.FullTokenizer(
vocab_file=FLAGS.vocab_file, do_lower_case=FLAGS.do_lower_case)
tpu_cluster_resolver = None
if FLAGS.use_tpu and FLAGS.tpu_name:
tpu_cluster_resolver = tf.contrib.cluster_resolver.TPUClusterResolver(
FLAGS.tpu_name, zone=FLAGS.tpu_zone, project=FLAGS.gcp_project)
is_per_host = tf.contrib.tpu.InputPipelineConfig.PER_HOST_V2
run_config = tf.contrib.tpu.RunConfig(
cluster=tpu_cluster_resolver,
master=FLAGS.master,
model_dir=FLAGS.output_dir,
save_checkpoints_steps=FLAGS.save_checkpoints_steps,
tpu_config=tf.contrib.tpu.TPUConfig(
iterations_per_loop=FLAGS.iterations_per_loop,
num_shards=FLAGS.num_tpu_cores,
per_host_input_for_training=is_per_host))
return bert_config, run_config, tokenizer
def model_definition(bert_config, run_config):
train_examples = None
num_train_steps = None
num_warmup_steps = None
if FLAGS.do_train:
train_examples = read_squad_examples(
input_file=FLAGS.train_file, is_training=True)
num_train_steps = int(
len(train_examples) / FLAGS.train_batch_size * FLAGS.num_train_epochs)
num_warmup_steps = int(num_train_steps * FLAGS.warmup_proportion)
# Pre-shuffle the input to avoid having to make a very large shuffle
# buffer in in the `input_fn`.
rng = random.Random(12345)
rng.shuffle(train_examples)
model_fn = model_fn_builder(
bert_config=bert_config,
init_checkpoint=FLAGS.init_checkpoint,
learning_rate=FLAGS.learning_rate,
num_train_steps=num_train_steps,
num_warmup_steps=num_warmup_steps,
use_tpu=FLAGS.use_tpu,
use_one_hot_embeddings=FLAGS.use_tpu)
# If TPU is not available, this will fall back to normal Estimator on CPU
# or GPU.
estimator = tf.contrib.tpu.TPUEstimator(
use_tpu=FLAGS.use_tpu,
model_fn=model_fn,
config=run_config,
train_batch_size=FLAGS.train_batch_size,
predict_batch_size=FLAGS.predict_batch_size)
return model_fn, estimator
def testing_model(eval_examples, tokenizer, estimator):
eval_writer = FeatureWriter(
filename=os.path.join(FLAGS.output_dir, "eval.tf_record"),
is_training=False)
eval_features = []
def append_feature(feature):
eval_features.append(feature)
eval_writer.process_feature(feature)
convert_examples_to_features(
examples=eval_examples,
tokenizer=tokenizer,
max_seq_length=FLAGS.max_seq_length,
doc_stride=FLAGS.doc_stride,
max_query_length=FLAGS.max_query_length,
is_training=False,
output_fn=append_feature)
eval_writer.close()
# tf.logging.info("***** Running predictions *****")
# tf.logging.info(" Num orig examples = %d", len(eval_examples))
# tf.logging.info(" Num split examples = %d", len(eval_features))
# tf.logging.info(" Batch size = %d", FLAGS.predict_batch_size)
all_results = []
predict_input_fn = input_fn_builder(
input_file=eval_writer.filename,
seq_length=FLAGS.max_seq_length,
is_training=False,
drop_remainder=False)
# If running eval on the TPU, you will need to specify the number of
# steps.
all_results = []
t1 = time.time()
for result in estimator.predict(
predict_input_fn, yield_single_examples=True):
# print(time.time() - t1)
# if len(all_results) % 1000 == 0:
# tf.logging.info("Processing example: %d" % (len(all_results)))
unique_id = int(result["unique_ids"])
start_logits = [float(x) for x in result["start_logits"].flat]
end_logits = [float(x) for x in result["end_logits"].flat]
all_results.append(
RawResult(
unique_id=unique_id,
start_logits=start_logits,
end_logits=end_logits))
return eval_features, all_results
def PredictAnswer(ContextSummary, question, tokenizer, estimator):
data = {'data': [{'title': 'Random',
'paragraphs': [{'context': ContextSummary,
'qas': [{'answers': [],
'question': question,
'id': '56be4db0acb8001400a502ec'}]}]}],
'version': '1.1'}
FLAGS.interact = True
eval_examples = read_squad_examples(data = data, is_training=False)
eval_features, all_results = testing_model(eval_examples, tokenizer, estimator)
output_prediction_file = os.path.join(FLAGS.output_dir, "predictions.json")
output_nbest_file = os.path.join(FLAGS.output_dir, "nbest_predictions.json")
output_null_log_odds_file = os.path.join(FLAGS.output_dir, "null_odds.json")
Prediction = write_predictions(eval_examples, eval_features, all_results,
FLAGS.n_best_size, FLAGS.max_answer_length,
FLAGS.do_lower_case, output_prediction_file,
output_nbest_file, output_null_log_odds_file)
return list(Prediction.values())[0]
def main(_):
if FLAGS.interact:
bert_config, run_config, tokenizer = initializingModels()
model_fn, estimator = model_definition(bert_config, run_config)
def GetTextSummary(SummaryText, question):
S = SummaryText.apply(lambda x: max([word_vectors.wmdistance(remove_stopwords(sent).lower().split(),remove_stopwords(question).lower().split()) for sent in x]))
Summary = ' \n\n'.join(SummaryText[S.sort_values(ascending=False)[:2].index].apply(','.join).tolist())
return Summary
# with open(r'Data\RPA_data_summary.txt') as f:
# text = f.read()
with open(FLAGS.context,'r') as file:
text = file.read()
SummaryText = Series(text.split('\n\n\n'))
SummaryText = SummaryText.apply(lambda x:[i for i in sent_tokenize(x) if len(i)>2])
while True:
raw_text = input(">>> ")
while not raw_text:
print('Prompt should not be empty!')
raw_text = input(">>> ")
if raw_text.lower() == 'quit':
return
ContextSummary = GetTextSummary(SummaryText,raw_text)
out_text = PredictAnswer(ContextSummary,raw_text, tokenizer, estimator)
print(out_text)
if FLAGS.question != None:
bert_config, run_config, tokenizer = initializingModels()
model_fn, estimator = model_definition(bert_config, run_config)
with open(FLAGS.context,'r') as file:
context = file.read()
Answer = PredictAnswer(SummaryText=context,question=FLAGS.question, tokenizer=tokenizer, estimator=estimator)
print(Answer)
return
tf.logging.set_verbosity(tf.logging.INFO)
bert_config = modeling.BertConfig.from_json_file(FLAGS.bert_config_file)
validate_flags_or_throw(bert_config)
tf.gfile.MakeDirs(FLAGS.output_dir)
tokenizer = tokenization.FullTokenizer(
vocab_file=FLAGS.vocab_file, do_lower_case=FLAGS.do_lower_case)
tpu_cluster_resolver = None
if FLAGS.use_tpu and FLAGS.tpu_name:
tpu_cluster_resolver = tf.contrib.cluster_resolver.TPUClusterResolver(
FLAGS.tpu_name, zone=FLAGS.tpu_zone, project=FLAGS.gcp_project)
is_per_host = tf.contrib.tpu.InputPipelineConfig.PER_HOST_V2
run_config = tf.contrib.tpu.RunConfig(
cluster=tpu_cluster_resolver,
master=FLAGS.master,
model_dir=FLAGS.output_dir,
save_checkpoints_steps=FLAGS.save_checkpoints_steps,
tpu_config=tf.contrib.tpu.TPUConfig(
iterations_per_loop=FLAGS.iterations_per_loop,
num_shards=FLAGS.num_tpu_cores,
per_host_input_for_training=is_per_host))
train_examples = None
num_train_steps = None
num_warmup_steps = None
if FLAGS.do_train:
train_examples = read_squad_examples(
input_file=FLAGS.train_file, is_training=True)
num_train_steps = int(
len(train_examples) / FLAGS.train_batch_size * FLAGS.num_train_epochs)
num_warmup_steps = int(num_train_steps * FLAGS.warmup_proportion)
# Pre-shuffle the input to avoid having to make a very large shuffle
# buffer in in the `input_fn`.
rng = random.Random(12345)
rng.shuffle(train_examples)
model_fn = model_fn_builder(
bert_config=bert_config,
init_checkpoint=FLAGS.init_checkpoint,
learning_rate=FLAGS.learning_rate,
num_train_steps=num_train_steps,
num_warmup_steps=num_warmup_steps,
use_tpu=FLAGS.use_tpu,
use_one_hot_embeddings=FLAGS.use_tpu)
# If TPU is not available, this will fall back to normal Estimator on CPU
# or GPU.
estimator = tf.contrib.tpu.TPUEstimator(
use_tpu=FLAGS.use_tpu,
model_fn=model_fn,
config=run_config,
train_batch_size=FLAGS.train_batch_size,
predict_batch_size=FLAGS.predict_batch_size)
if FLAGS.do_train:
# We write to a temporary file to avoid storing very large constant tensors
# in memory.
train_writer = FeatureWriter(
filename=os.path.join(FLAGS.output_dir, "train.tf_record"),
is_training=True)
convert_examples_to_features(
examples=train_examples,
tokenizer=tokenizer,
max_seq_length=FLAGS.max_seq_length,
doc_stride=FLAGS.doc_stride,
max_query_length=FLAGS.max_query_length,
is_training=True,
output_fn=train_writer.process_feature)
train_writer.close()
tf.logging.info("***** Running training *****")
tf.logging.info(" Num orig examples = %d", len(train_examples))
tf.logging.info(" Num split examples = %d", train_writer.num_features)
tf.logging.info(" Batch size = %d", FLAGS.train_batch_size)
tf.logging.info(" Num steps = %d", num_train_steps)
del train_examples
train_input_fn = input_fn_builder(
input_file=train_writer.filename,
seq_length=FLAGS.max_seq_length,
is_training=True,
drop_remainder=True)
estimator.train(input_fn=train_input_fn, max_steps=num_train_steps)
if FLAGS.do_predict:
eval_examples = read_squad_examples(
input_file=FLAGS.predict_file, is_training=False)
eval_writer = FeatureWriter(
filename=os.path.join(FLAGS.output_dir, "eval.tf_record"),
is_training=False)
eval_features = []
def append_feature(feature):
eval_features.append(feature)
eval_writer.process_feature(feature)
convert_examples_to_features(
examples=eval_examples,
tokenizer=tokenizer,
max_seq_length=FLAGS.max_seq_length,
doc_stride=FLAGS.doc_stride,
max_query_length=FLAGS.max_query_length,
is_training=False,
output_fn=append_feature)
eval_writer.close()
tf.logging.info("***** Running predictions *****")
tf.logging.info(" Num orig examples = %d", len(eval_examples))
tf.logging.info(" Num split examples = %d", len(eval_features))
tf.logging.info(" Batch size = %d", FLAGS.predict_batch_size)
all_results = []
predict_input_fn = input_fn_builder(
input_file=eval_writer.filename,
seq_length=FLAGS.max_seq_length,
is_training=False,
drop_remainder=False)
# If running eval on the TPU, you will need to specify the number of
# steps.
all_results = []
for result in estimator.predict(
predict_input_fn, yield_single_examples=True):
if len(all_results) % 1000 == 0:
tf.logging.info("Processing example: %d" % (len(all_results)))
unique_id = int(result["unique_ids"])
start_logits = [float(x) for x in result["start_logits"].flat]
end_logits = [float(x) for x in result["end_logits"].flat]
all_results.append(
RawResult(
unique_id=unique_id,
start_logits=start_logits,
end_logits=end_logits))
output_prediction_file = os.path.join(FLAGS.output_dir, "predictions.json")
output_nbest_file = os.path.join(FLAGS.output_dir, "nbest_predictions.json")
output_null_log_odds_file = os.path.join(FLAGS.output_dir, "null_odds.json")
write_predictions(eval_examples, eval_features, all_results,
FLAGS.n_best_size, FLAGS.max_answer_length,
FLAGS.do_lower_case, output_prediction_file,
output_nbest_file, output_null_log_odds_file)
if __name__ == "__main__":
flags.mark_flag_as_required("vocab_file")
flags.mark_flag_as_required("bert_config_file")
flags.mark_flag_as_required("output_dir")
tf.app.run()
FLAGS.interact =True
FLAGS.context = r'Data\delete.txt'
tf.app.run()
```
| github_jupyter |
Calculation of Design strength of Lag Screws using CSA O86-09
```
import pint
unit = pint.UnitRegistry()
# define synanyms for common units
inch = unit.inch; mm = unit.mm; m = unit.m; kPa = unit.kPa; MPa = unit.MPa;
psi = unit.psi; kN = unit.kN; ksi = unit.ksi;
dimensionless = unit.dimensionless; s = unit.second; kg = unit.kg
def u_round(Q, digits=3):
"""
Takes a Pint.py quantity and returns same rounded to digits,
default is 3 digits
"""
try:
unit.check(Q.units)
magnitude = Q.magnitude
units = Q.units
return round(magnitude,digits) * units
except:
print('ERROR: u_round() first arguement must be a Pint.py quantity')
return NaN
# Define case parameters
G = 0.50 # mean density for Douglas-Fir-Larch from Table A10.1
t_1 = 38*mm # side member thickness
t_2 = 76*mm # main member thickness
d_F = (3/8) * inch # lag screw shank diameter
lag_L = 100*mm
d_F.ito(mm)
lag_E = 6.4*mm # length of tip, from Wood Design Manual 2010 Table 7.15
L_p = lag_L - (t_1 + lag_E)
K_D = 1.15 # Duration factor, short-term, less than 7 days
K_SF = 1.0 # Dry service conditions
K_T = 1.0 # untreated wood
J_E = 1.0 # screws installed in side grain; not end grain
Kprime = K_D * K_SF * K_T
print('Case parameters:')
print('Mean density of Douglas-Fir-Larch from Table A10.1, G =',round(G,2))
print('Thickness of side member, t_1 =',u_round(t_1,1))
print('Thickness of main member, t_2 =',u_round(t_2,1))
print('Lag Screw diameter, d_F =',u_round(d_F,1))
print('Lag Screw length, L =',u_round(lag_L,1))
print('Lag Screw tip length, E =',u_round(lag_E,1))
print('Lag Screw Penetration, L_p = L - (t_1 + E) =',u_round(L_p,1))
print('')
print('Modification Factors')
print('Duration Factor, K_D =',round(K_D,2))
print('Service Condition Factor, K_SF =',round(K_SF,1))
print('Fire-retardant Treatment Factor, K_T =',round(K_T,1))
print('Grain Factor, J_E =',round(J_E,1))
print('K` = K_D * K_SF * K_T =',round(Kprime,2))
Qprime_r = 1.08*kN
Q_r = Qprime_r * Kprime
print('Per WDM2010 Leg Screw Selection Table, Q`_r = ',u_round(Qprime_r,2))
print('Q_r = Q`_r * K`')
print('Q_r =',u_round(Q_r,2),'per lag screw')
```
**Given:** 2 kPa Live Load
**Given:** Max 1.3m width of platform
**Given:** Lag Screw capacity above
**Given:** Safety Factor of 1.5:1
**Find:** Max on-centre spacing of lag screws
$1.24 kN = (2 kPa)(1.5)(1.3 m / 2)(x)$
$x = 1.24 kN / ((2 kPa)(1.5)(0.65 m))$
```
x = u_round((Q_r / ((2*kPa)*(1.5)*(0.65*m))).to(m),3)
print('Given conditions above and 3/8" x 4" Lag screws',
'screws should be a max',x,'on-centre.')
```
**Given:** Conditions stated above
**Find:** Max on-centre spacing when 3/8" x 3" lag screws are used
```
lag_L = 76*mm
lag_E = 6.4*mm
L_p = lag_L - (t_1 + lag_E)
print('Lag Screw Penetration, L_p = L - (t_1 + E) =',u_round(L_p,1))
# O86-09 10.6.3.3
"""
For lag screws loaded laterally, the minimum length of penetration into the
main member (t_2 in Clause 10.6.6.1.2) shall be not less than five times the
shank diameter, d_F
"""
print('5 x shank diameter =',u_round(5*d_F,1))
```
Leg Screw Penetration is less than minimum required penetration. Therefore, 3/8" x 3" Lag Screws are NFG
| github_jupyter |
```
# Install tf-transformers from github
import json
import tensorflow as tf
import time
import glob
import collections
from tf_transformers.utils.tokenization import BasicTokenizer, ROBERTA_SPECIAL_PEICE
from tf_transformers.utils import fast_sp_alignment
from tf_transformers.data.squad_utils_sp import (
read_squad_examples,
post_clean_train_squad,
example_to_features_using_fast_sp_alignment_train,
example_to_features_using_fast_sp_alignment_test,
_get_best_indexes, evaluate_v1
)
from tf_transformers.data import TFWriter, TFReader, TFProcessor
from tf_transformers.models import RobertaModel
from tf_transformers.core import optimization, SimpleTrainer
from tf_transformers.tasks import Span_Selection_Model
from transformers import RobertaTokenizer
from absl import logging
logging.set_verbosity("INFO")
from tf_transformers.pipeline.span_extraction_pipeline import Span_Extraction_Pipeline
```
### Load Tokenizer
```
tokenizer = RobertaTokenizer.from_pretrained('roberta-base')
basic_tokenizer = BasicTokenizer(do_lower_case=False)
```
### Convert train data to Features
* using Fast Sentence Piece Alignment, we convert text to features (text -> list of sub words)
```
input_file_path = '/mnt/home/PRE_MODELS/HuggingFace_models/datasets/squadv1.1/train-v1.1.json'
is_training = True
# 1. Read Examples
start_time = time.time()
train_examples = read_squad_examples(
input_file=input_file_path,
is_training=is_training,
version_2_with_negative=False
)
end_time = time.time()
print('Time taken {}'.format(end_time-start_time))
# 2.Postprocess (clean text to avoid some unwanted unicode charcaters)
train_examples_processed, failed_examples = post_clean_train_squad(train_examples, basic_tokenizer, is_training=is_training)
# 3.Convert question, context and answer to proper features (tokenized words) not word indices
feature_generator = example_to_features_using_fast_sp_alignment_train(tokenizer, train_examples_processed, max_seq_length = 384,
max_query_length=64, doc_stride=128, SPECIAL_PIECE=ROBERTA_SPECIAL_PEICE)
all_features = []
for feature in feature_generator:
all_features.append(feature)
end_time = time.time()
print("time taken {} seconds".format(end_time-start_time))
```
### Convert features to TFRecords using TFWriter
```
# Convert tokens to id and add type_ids
# input_mask etc
# This is user specific/ tokenizer specific
# Eg: Roberta has input_type_ids = 0, BERT has input_type_ids = [0, 1]
def parse_train():
result = {}
for f in all_features:
input_ids = tokenizer.convert_tokens_to_ids(f['input_ids'])
input_type_ids = tf.zeros_like(input_ids).numpy().tolist()
input_mask = tf.ones_like(input_ids).numpy().tolist()
result['input_ids'] = input_ids
result['input_type_ids'] = input_type_ids
result['input_mask'] = input_mask
result['start_position'] = f['start_position']
result['end_position'] = f['end_position']
yield result
# Lets write using TF Writer
# Use TFProcessor for smaller data
schema = {'input_ids': ("var_len", "int"),
'input_type_ids': ("var_len", "int"),
'input_mask': ("var_len", "int"),
'start_position': ("var_len", "int"),
'end_position': ("var_len", "int")}
tfrecord_train_dir = '../OFFICIAL_TFRECORDS/squad/train'
tfrecord_filename = 'squad'
tfwriter = TFWriter(schema=schema,
file_name=tfrecord_filename,
model_dir=tfrecord_train_dir,
tag='train',
overwrite=True
)
tfwriter.process(parse_fn=parse_train())
```
### Read TFRecords using TFReader
```
# Read Data
schema = json.load(open("{}/schema.json".format(tfrecord_train_dir)))
all_files = glob.glob("{}/*.tfrecord".format(tfrecord_train_dir))
tf_reader = TFReader(schema=schema,
tfrecord_files=all_files)
x_keys = ['input_ids', 'input_type_ids', 'input_mask']
y_keys = ['start_position', 'end_position']
batch_size = 16
train_dataset = tf_reader.read_record(auto_batch=True,
keys=x_keys,
batch_size=batch_size,
x_keys = x_keys,
y_keys = y_keys,
shuffle=True,
drop_remainder=True
)
```
### Load Roberta base Model
```
model_layer, model, config = RobertaModel(model_name='roberta-base', return_all_layer_token_embeddings=False)
model.load_checkpoint("/mnt/home/PRE_MODELS/LegacyAI_models/checkpoints/roberta-base/")
```
### Load Span Selection Model
```
tf.keras.backend.clear_session()
span_selection_layer = Span_Selection_Model(model=model,
use_all_layers=False,
is_training=True)
span_selection_model = span_selection_layer.get_model()
# Delete to save up memory
del model
del model_layer
del span_selection_layer
```
### Define Loss
Loss function is simple.
* labels: 1D (batch_size) # start or end positions
* logits: 2D (batch_size x sequence_length)
```
# Cross Entropy
def span_loss(position, logits):
loss = tf.reduce_mean(tf.nn.sparse_softmax_cross_entropy_with_logits(
logits=logits, labels=tf.squeeze(position, axis=1)))
return loss
# Start logits loss
def start_loss(y_true_dict, y_pred_dict):
start_loss = span_loss(y_true_dict['start_position'], y_pred_dict['start_logits'])
return start_loss
# End logits loss
def end_loss(y_true_dict, y_pred_dict):
end_loss = span_loss(y_true_dict['end_position'], y_pred_dict['end_logits'])
return end_loss
# (start_loss + end_loss) / 2.0
def joint_loss(y_true_dict, y_pred_dict):
sl = start_loss(y_true_dict, y_pred_dict)
el = end_loss(y_true_dict, y_pred_dict)
return (sl + el)/2.0
for (batch_inputs, batch_labels) in train_dataset.take(1):
print(batch_inputs, batch_labels)
```
### Define Optimizer
```
train_data_size = 89000
learning_rate = 2e-5
steps_per_epoch = int(train_data_size / batch_size)
EPOCHS = 3
num_train_steps = steps_per_epoch * EPOCHS
warmup_steps = int(0.1 * num_train_steps)
# creates an optimizer with learning rate schedule
optimizer_type = 'adamw'
optimizer, learning_rate_fn = optimization.create_optimizer(learning_rate,
steps_per_epoch * EPOCHS,
warmup_steps,
optimizer_type)
```
### Train Using Keras :-)
- ```compile2``` allows you to have directly use model outputs as well batch dataset outputs into the loss function, without any further complexity.
Note: For ```compile2```, loss_fn must be None, and custom_loss_fn must be active. Metrics are not supprted for time being.
```
# Keras Fit
keras_loss_fn = {'start_logits': start_loss,
'end_logits': end_loss}
span_selection_model.compile2(optimizer=tf.keras.optimizers.Adam(),
loss=None,
custom_loss=keras_loss_fn
)
history = span_selection_model.fit(train_dataset, epochs=2, steps_per_epoch=10)
```
### Train using SimpleTrainer (part of tf-transformers)
```
# Custom training
history = SimpleTrainer(model = span_selection_model,
optimizer = optimizer,
loss_fn = joint_loss,
dataset = train_dataset.repeat(EPOCHS+1), # This is important
epochs = EPOCHS,
num_train_examples = train_data_size,
batch_size = batch_size,
steps_per_call=100,
gradient_accumulation_steps=None)
```
### Save Models
You can save models as checkpoints using ```.save_checkpoint``` attribute, which is a part of all ```LegacyModels```
```
model_save_dir = '../OFFICIAL_MODELS/squad/roberta_base2'
span_selection_model.save_checkpoint(model_save_dir)
```
### Parse validation data
We use ```TFProcessor``` to create validation data, because dev data is small
```
dev_input_file_path = '/mnt/home/PRE_MODELS/HuggingFace_models/datasets/squadv1.1/dev-v1.1.json'
is_training = False
start_time = time.time()
dev_examples = read_squad_examples(
input_file=dev_input_file_path,
is_training=is_training,
version_2_with_negative=False
)
end_time = time.time()
print('Time taken {}'.format(end_time-start_time))
dev_examples_cleaned = post_clean_train_squad(dev_examples, basic_tokenizer, is_training=False)
qas_id_info, dev_features = example_to_features_using_fast_sp_alignment_test(tokenizer, dev_examples_cleaned, max_seq_length = 384,
max_query_length=64, doc_stride=128, SPECIAL_PIECE=ROBERTA_SPECIAL_PEICE)
def parse_dev():
result = {}
for f in dev_features:
input_ids = tokenizer.convert_tokens_to_ids(f['input_ids'])
input_type_ids = tf.zeros_like(input_ids).numpy().tolist()
input_mask = tf.ones_like(input_ids).numpy().tolist()
result['input_ids'] = input_ids
result['input_type_ids'] = input_type_ids
result['input_mask'] = input_mask
yield result
tf_processor = TFProcessor()
dev_dataset = tf_processor.process(parse_fn=parse_dev())
dev_dataset = tf_processor.auto_batch(dev_dataset, batch_size=32)
```
### Evaluate Exact Match
* Make Predictions
* Extract Answers
* Evaluate
### Make Batch Predictions
```
def extract_from_dict(dict_items, key):
holder = []
for item in dict_items:
holder.append(item[key])
return holder
qas_id_list = extract_from_dict(dev_features, 'qas_id')
doc_offset_list = extract_from_dict(dev_features, 'doc_offset')
# Make batch predictions
per_layer_start_logits = []
per_layer_end_logits = []
start_time = time.time()
for (batch_inputs) in dev_dataset:
model_outputs = span_selection_model(batch_inputs)
per_layer_start_logits.append(model_outputs['start_logits'])
per_layer_end_logits.append(model_outputs['end_logits'])
end_time = time.time()
print('Time taken {}'.format(end_time-start_time))
```
### Extract Answers (text) from Predictions
* Its little tricky as there will be multiple features for one example, if it is longer than max_seq_length
```
# Make batch predictions
n_best_size = 20
max_answer_length = 30
squad_dev_data = json.load(open(dev_input_file_path))['data']
predicted_results = []
# Unstack (matrxi tensor) into list arrays (list)
start_logits_unstcaked = []
end_logits_unstacked = []
for batch_start_logits in per_layer_start_logits:
start_logits_unstcaked.extend(tf.unstack(batch_start_logits))
for batch_end_logits in per_layer_end_logits:
end_logits_unstacked.extend(tf.unstack(batch_end_logits))
# Group (multiple predictions) of one example, due to big passage/context
# We need to choose the best anser of all the chunks of a examples
qas_id_logits = {}
for i in range(len(qas_id_list)):
qas_id = qas_id_list[i]
example = qas_id_info[qas_id]
feature = dev_features[i]
assert qas_id == feature['qas_id']
if qas_id not in qas_id_logits:
qas_id_logits[qas_id] = {'tok_to_orig_index': example['tok_to_orig_index'],
'aligned_words': example['aligned_words'],
'feature_length': [len(feature['input_ids'])],
'doc_offset': [doc_offset_list[i]],
'passage_start_pos': [feature['input_ids'].index(tokenizer.sep_token) + 1],
'start_logits': [start_logits_unstcaked[i]],
'end_logits': [end_logits_unstacked[i]]}
else:
qas_id_logits[qas_id]['start_logits'].append(start_logits_unstcaked[i])
qas_id_logits[qas_id]['end_logits'].append(end_logits_unstacked[i])
qas_id_logits[qas_id]['feature_length'].append(len(feature['input_ids']))
qas_id_logits[qas_id]['doc_offset'].append(doc_offset_list[i])
qas_id_logits[qas_id]['passage_start_pos'].append(feature['input_ids'].index(tokenizer.sep_token) + 1)
# Extract answer assoiate it with single (qas_id) unique identifier
qas_id_answer = {}
skipped = []
skipped_null = []
global_counter = 0
for qas_id in qas_id_logits:
current_example = qas_id_logits[qas_id]
_PrelimPrediction = collections.namedtuple( # pylint: disable=invalid-name
"PrelimPrediction",
["feature_index", "start_index", "end_index",
"start_log_prob", "end_log_prob"])
prelim_predictions = []
example_features = []
for i in range(len( current_example['start_logits'])):
f = dev_features[global_counter]
assert f['qas_id'] == qas_id
example_features.append(f)
global_counter += 1
passage_start_pos = current_example['passage_start_pos'][i]
feature_length = current_example['feature_length'][i]
start_log_prob_list = current_example['start_logits'][i].numpy().tolist()[:feature_length]
end_log_prob_list = current_example['end_logits'][i].numpy().tolist()[:feature_length]
start_indexes = _get_best_indexes(start_log_prob_list, n_best_size)
end_indexes = _get_best_indexes(end_log_prob_list, n_best_size)
for start_index in start_indexes:
for end_index in end_indexes:
# We could hypothetically create invalid predictions, e.g., predict
# that the start of the span is in the question. We throw out all
# invalid predictions.
if start_index < passage_start_pos or end_index < passage_start_pos:
continue
if end_index < start_index:
continue
length = end_index - start_index + 1
if length > max_answer_length:
continue
start_log_prob = start_log_prob_list[start_index]
end_log_prob = end_log_prob_list[end_index]
start_idx = start_index - passage_start_pos
end_idx = end_index - passage_start_pos
prelim_predictions.append(
_PrelimPrediction(
feature_index=i,
start_index=start_idx,
end_index=end_idx,
start_log_prob=start_log_prob,
end_log_prob=end_log_prob))
prelim_predictions = sorted(
prelim_predictions,
key=lambda x: (x.start_log_prob + x.end_log_prob),
reverse=True)
if prelim_predictions:
best_index = prelim_predictions[0].feature_index
aligned_words = current_example['aligned_words']
try:
tok_to_orig_index = current_example['tok_to_orig_index']
reverse_start_index_align = tok_to_orig_index[prelim_predictions[0].start_index + example_features[best_index]['doc_offset']] # aligned index
reverse_end_index_align = tok_to_orig_index[prelim_predictions[0].end_index + example_features[best_index]['doc_offset']]
predicted_words = [w for w in aligned_words[reverse_start_index_align: reverse_end_index_align + 1] if w != ROBERTA_SPECIAL_PEICE]
predicted_text = ' '.join(predicted_words)
qas_id_answer[qas_id] = predicted_text
except:
qas_id_answer[qas_id] = ""
skipped.append(qas_id)
else:
qas_id_answer[qas_id] = ""
skipped_null.append(qas_id)
eval_results = evaluate_v1(squad_dev_data, qas_id_answer)
# {'exact_match': 81.46641438032167, 'f1': 89.72853269935702}
```
### Save as Serialized version
- Now we can use ```save_as_serialize_module``` to save a model directly to saved_model
```
# Save as optimized version
span_selection_model.save_as_serialize_module("{}/saved_model".format(model_save_dir), overwrite=True)
# Load optimized version
span_selection_model_serialized = tf.saved_model.load("{}/saved_model".format(model_save_dir))
```
### TFLite Conversion
TFlite conversion requires:
- static batch size
- static sequence length
```
# Sequence_length = 384
# batch_size = 1
# Lets convert it to a TFlite model
# Load base model with specified sequence length and batch_size
model_layer, model, config = RobertaModel(model_name='roberta-base',
sequence_length=384, # Fix a sequence length for TFlite (it shouldnt be None)
batch_size=1,
use_dropout=False) # batch_size=1
# Disable dropout (important) for TFlite
tf.keras.backend.clear_session()
span_selection_layer = Span_Selection_Model(model=model,
is_training=False)
span_selection_model = span_selection_layer.get_model()
span_selection_model.load_checkpoint(model_save_dir)
# Save to .pb format , we need it for tflite
span_selection_model.save_as_serialize_module("{}/saved_model_for_tflite".format(model_save_dir))
converter = tf.lite.TFLiteConverter.from_saved_model("{}/saved_model_for_tflite".format(model_save_dir)) # path to the SavedModel directory
converter.experimental_new_converter = True
tflite_model = converter.convert()
open("{}/converted_model.tflite".format(model_save_dir), "wb").write(tflite_model)
```
### **In production**
- We can use either ```tf.keras.Model``` or ```saved_model```. I recommend saved_model, which is much much faster and no hassle of having architecture code
```
def tokenizer_fn(features):
"""
features: dict of tokenized text
Convert them into ids
"""
result = {}
input_ids = tokenizer.convert_tokens_to_ids(features['input_ids'])
input_type_ids = tf.zeros_like(input_ids).numpy().tolist()
input_mask = tf.ones_like(input_ids).numpy().tolist()
result['input_ids'] = input_ids
result['input_type_ids'] = input_type_ids
result['input_mask'] = input_mask
return result
# Span Extraction Pipeline
pipeline = Span_Extraction_Pipeline(model = span_selection_model_serialized,
tokenizer = tokenizer,
tokenizer_fn = tokenizer_fn,
SPECIAL_PIECE = ROBERTA_SPECIAL_PEICE,
n_best_size = 20,
n_best = 5,
max_answer_length = 30,
max_seq_length = 384,
max_query_length=64,
doc_stride=20)
questions = ['What was prominent in Kerala?']
questions = ['When was Kerala formed?']
questions = ['How many districts are there in Kerala']
contexts = ['''Kerala (English: /ˈkɛrələ/; Malayalam: [ke:ɾɐɭɐm] About this soundlisten (help·info)) is a state on the southwestern Malabar Coast of India. It was formed on 1 November 1956, following the passage of the States Reorganisation Act, by combining Malayalam-speaking regions of the erstwhile states of Travancore-Cochin and Madras. Spread over 38,863 km2 (15,005 sq mi), Kerala is the twenty-first largest Indian state by area. It is bordered by Karnataka to the north and northeast, Tamil Nadu to the east and south, and the Lakshadweep Sea[14] to the west. With 33,387,677 inhabitants as per the 2011 Census, Kerala is the thirteenth-largest Indian state by population. It is divided into 14 districts with the capital being Thiruvananthapuram. Malayalam is the most widely spoken language and is also the official language of the state.[15]
The Chera Dynasty was the first prominent kingdom based in Kerala. The Ay kingdom in the deep south and the Ezhimala kingdom in the north formed the other kingdoms in the early years of the Common Era (CE). The region had been a prominent spice exporter since 3000 BCE. The region's prominence in trade was noted in the works of Pliny as well as the Periplus around 100 CE. In the 15th century, the spice trade attracted Portuguese traders to Kerala, and paved the way for European colonisation of India. At the time of Indian independence movement in the early 20th century, there were two major princely states in Kerala-Travancore State and the Kingdom of Cochin. They united to form the state of Thiru-Kochi in 1949. The Malabar region, in the northern part of Kerala, had been a part of the Madras province of British India, which later became a part of the Madras State post-independence. After the States Reorganisation Act, 1956, the modern-day state of Kerala was formed by merging the Malabar district of Madras State (excluding Gudalur taluk of Nilgiris district, Lakshadweep Islands, Topslip, the Attappadi Forest east of Anakatti), the state of Thiru-Kochi (excluding four southern taluks of Kanyakumari district, Shenkottai and Tenkasi taluks), and the taluk of Kasaragod (now Kasaragod District) in South Canara (Tulunad) which was a part of Madras State.''']
result = pipeline(questions=questions, contexts=contexts)
```
### Sanity Check TFlite
```
#### lets do a sanity check
sample_inputs = {}
input_ids = tf.random.uniform(minval=0, maxval=100, shape=(1, 384), dtype=tf.int32)
sample_inputs['input_ids'] = input_ids
sample_inputs['input_type_ids'] = tf.zeros_like(sample_inputs['input_ids'])
sample_inputs['input_mask'] = tf.ones_like(sample_inputs['input_ids'])
model_outputs = span_selection_model(sample_inputs)
# Load the TFLite model and allocate tensors.
interpreter = tf.lite.Interpreter(model_path="{}/converted_model.tflite".format(model_save_dir))
interpreter.allocate_tensors()
# Get input and output tensors.
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
interpreter.set_tensor(input_details[0]['index'], sample_inputs['input_ids'])
interpreter.set_tensor(input_details[1]['index'], sample_inputs['input_mask'])
interpreter.set_tensor(input_details[2]['index'], sample_inputs['input_type_ids'])
interpreter.invoke()
end_logits = interpreter.get_tensor(output_details[0]['index'])
start_logits = interpreter.get_tensor(output_details[1]['index'])
# Assertion
print("Start logits", tf.reduce_sum(model_outputs['start_logits']), tf.reduce_sum(start_logits))
print("End logits", tf.reduce_sum(model_outputs['end_logits']), tf.reduce_sum(end_logits))
# We are good :-)
```
| github_jupyter |
# Comparison of approximate robustness curves for different sample sizes (1k vs. 10k points)
```
import os
os.chdir("../")
import sys
import copy
import json
from argparse import Namespace
import numpy as np
import scipy.io
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(context='paper')
import provable_robustness_max_linear_regions.data as dt
from generate_robustness_curves import generate_curve_data
from utils import NumpyEncoder
```
## Plot settings:
```
SMALL_SIZE = 14
MEDIUM_SIZE = 18
BIGGER_SIZE = 26
plt.rc('font', size=SMALL_SIZE) # controls default text sizes
plt.rc('axes', titlesize=MEDIUM_SIZE) # fontsize of the axes title
plt.rc('axes', labelsize=MEDIUM_SIZE) # fontsize of the x and y labels
plt.rc('xtick', labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc('ytick', labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc('legend', fontsize=SMALL_SIZE) # legend fontsize
plt.rc('figure', titlesize=BIGGER_SIZE) # fontsize of the figure title
plt.rc('text', usetex=True)
# dictionary that maps color string to 'good looking' seaborn colors that are easily distinguishable
colors = {
"orange": sns.xkcd_rgb["yellowish orange"],
"red": sns.xkcd_rgb["pale red"],
"green": sns.xkcd_rgb["medium green"],
"blue": sns.xkcd_rgb["denim blue"],
"yellow": sns.xkcd_rgb["amber"],
"purple": sns.xkcd_rgb["dusty purple"],
"cyan": sns.xkcd_rgb["cyan"]
}
```
## Calculate robustness curves:
Estimated runtime (if no file with data is present): 3 days
```
def load_from_json(file_name):
if not os.path.exists("res/" + file_name + ".json"):
return None
else:
with open("res/" + file_name + ".json", 'r') as fp:
loaded_json = json.load(fp)
for key in loaded_json.keys():
loaded_json[key]["x"] = np.array(loaded_json[key]["x"])
loaded_json[key]["y"] = np.array(loaded_json[key]["y"])
loaded_json[key]["y"][np.isnan(loaded_json[key]["x"])] = 1.0
loaded_json[key]["x"] = np.nan_to_num(loaded_json[key]["x"], nan = np.nanmax(loaded_json[key]["x"]))
return loaded_json
def save_to_json(dictionary, file_name):
if not os.path.exists("res"):
os.makedirs("res")
with open("res/" + file_name + ".json", 'w') as fp:
json.dump(dictionary, fp, cls = NumpyEncoder)
n_points = 10000
model_path = "provable_robustness_max_linear_regions/models/mmr+at/provable_robustness_max_linear_regions/models/mmr+at/2019-02-17 01:54:16 dataset=mnist nn_type=cnn_lenet_small p_norm=inf lmbd=0.5 gamma_rb=0.2 gamma_db=0.2 ae_frac=0.5 epoch=100.mat"
robustness_curve_data = dict()
robustness_curve_data = load_from_json("rob_curve_data_MMR+AT_l_inf_n_points={}".format(n_points))
if not robustness_curve_data:
args = Namespace()
args.dataset = "mnist"
args.n_points = n_points
args.model_path = model_path
args.nn_type = "cnn"
args.norms = ["2"]
args.save = False
args.plot = False
robustness_curve_data = generate_curve_data(args)
save_to_json(robustness_curve_data, "rob_curve_data_MMR+AT_l_inf_n_points={}".format(n_points))
```
## Plot:
```
# name to save the plot
save_name = "fig_rc__1k_vs_10k"
tenksample = robustness_curve_data["2"]["x"]
oneksample = np.sort(np.random.choice(robustness_curve_data["2"]["x"], size = 1000))
# number of model types and parameter combinations
n_cols = 1
n_rows = 1
fig, ax = plt.subplots(n_rows,
n_cols,
figsize=(6 * n_cols, 5 * n_rows),
dpi=400)
ax.plot(tenksample,
np.linspace(0, 1, tenksample.shape[0]),
c=colors["blue"],
label="number of samples = 10000")
ax.plot(oneksample,
np.linspace(0, 1, oneksample.shape[0]),
c=colors["red"],
label="number of samples = 1000")
ax.legend()
ax.set_ylabel("test set loss in $\%$")
ax.set_xlabel("perturbation size in $\ell_2$")
ax.set_title("appr. $\ell_2$ robustness curves\nfor different sample sizes")
fig.tight_layout()
fig.savefig('res/{}.pdf'.format(save_name))
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
import os
import numpy as np
import pandas as pd
# manually add a2e-mmc repos to PYTHONPATH if needed
import sys
module_path = os.path.join(os.environ['HOME'],'a2e-mmc')
if module_path not in sys.path:
sys.path.append(module_path)
# wrapper around reader functions to return a single combined dataframe
from mmctools.dataloaders import read_date_dirs
from mmctools.plotting import plot_timeheight
```
# Example: Data processing (of data organized into subdirectories)
written by [Eliot Quon](mailto:eliot.quon@nrel.gov)
- Combine a series of data files, located within a collection of date-time subdirectories, into one dataframe.
- Do a quick data check of the data at the end (make a time-height plot)
Note: This *may* be no longer needed since the `dap-py` and the online [DAP](https://a2e.energy.gov/data) interfaces both appear to consistently download files into a single directory. In the past, some datasets had data organized into subdirectories. This also introduced an additional processing step to parse date-time information from the subdirectory name and then add it to the timestamps in each subdirectory datafile.
```
# dataset name format: project/class.instance.level
dataset = 'wfip2/sodar.z12.b0'
startdate = pd.to_datetime('2016-10-30')
enddate = pd.to_datetime('2016-11-02')
# optional dataset file specs
dataext = 'txt' # file type, dictated by extension
dataext1 = None # 'winds' # e.g., *.winds.txt
download_path = 'data'
overwrite_files = False # force download even if files already exist
```
## Collect downloaded data from the DAP
```
datapath = os.path.join(download_path, dataset.replace('/','.'))
print('Data path:',datapath)
ls $datapath
```
## Process the downloaded files
```
%%time
df = read_date_dirs(datapath, # default reader=pd.read_csv,
expected_date_format='%Y%m', ext='txt',
# additional reader arguments:
parse_dates=['date_time'])
df.head()
```
## Standardization
```
# create multi-indexed dataframe with standard names
df = df.rename(columns={
'date_time':'datetime',
'speed':'wspd',
'direction':'wdir',
'vert':'w',
})
df = df.set_index(['datetime','height'])
df.head()
df.tail()
```
## Check the data
See [example_plotting.ipynb](https://github.com/a2e-mmc/mmctools/blob/dev/example_plotting.ipynb) for more plotting-specific examples.
```
fig,ax,cbars = plot_timeheight(df, fields=['wspd','wdir'])
```
| github_jupyter |
# Saving and Loading Models
In this notebook, I'll show you how to save and load models with PyTorch. This is important because you'll often want to load previously trained models to use in making predictions or to continue training on new data.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms
import helper
import fc_model # should be in your directory
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,))])
# Download and load the training data
trainset = datasets.FashionMNIST('~/.pytorch/F_MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
# Download and load the test data
testset = datasets.FashionMNIST('~/.pytorch/F_MNIST_data/', download=True, train=False, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=64, shuffle=True)
```
Here we can see one of the images.
```
image, label = next(iter(trainloader))
helper.imshow(image[0,:]);
image, label = next(iter(testloader))
helper.imshow(image[0,:]);
```
# Train a network
To make things more concise here, I moved the model architecture and training code from the last part to a file called `fc_model`. Importing this, we can easily create a fully-connected network with `fc_model.Network`, and train the network using `fc_model.train`. I'll use this model (once it's trained) to demonstrate how we can save and load models.
```
# Create the network, define the criterion and optimizer
model = fc_model.Network(784, 10, [512, 256, 128])
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.parameters(), lr=0.001)
fc_model.train(model, trainloader, testloader, criterion, optimizer, epochs=2)
```
## Saving and loading networks
As you can imagine, it's impractical to train a network every time you need to use it. Instead, we can save trained networks then load them later to train more or use them for predictions.
The parameters for PyTorch networks are stored in a model's `state_dict`. We can see the state dict contains the weight and bias matrices for each of our layers.
```
print("Our model: \n\n", model, '\n')
print("The state dict keys: \n\n", model.state_dict().keys())
```
The simplest thing to do is simply save the state dict with `torch.save`. For example, we can save it to a file `'checkpoint.pth'`.
```
torch.save(model.state_dict(), 'checkpoint.pth')
```
Then we can load the state dict with `torch.load`.
```
state_dict = torch.load('checkpoint.pth')
print(state_dict.keys())
```
And to load the state dict in to the network, you do `model.load_state_dict(state_dict)`.
```
model.load_state_dict(state_dict)
```
Seems pretty straightforward, but as usual it's a bit more complicated. Loading the state dict works only if the model architecture is exactly the same as the checkpoint architecture. If I create a model with a different architecture, this fails.
```
# Try this
model = fc_model.Network(784, 10, [400, 200, 100])
# This will throw an error because the tensor sizes are wrong!
model.load_state_dict(state_dict)
```
This means we need to rebuild the model exactly as it was when trained. Information about the model architecture needs to be saved in the checkpoint, along with the state dict. To do this, you build a dictionary with all the information you need to compeletely rebuild the model.
```
checkpoint = {'input_size': 784,
'output_size': 10,
'hidden_layers': [each.out_features for each in model.hidden_layers],
'state_dict': model.state_dict()}
torch.save(checkpoint, 'checkpoint.pth')
```
Now the checkpoint has all the necessary information to rebuild the trained model. You can easily make that a function if you want. Similarly, we can write a function to load checkpoints.
```
def load_checkpoint(filepath):
checkpoint = torch.load(filepath)
model = fc_model.Network(checkpoint['input_size'],
checkpoint['output_size'],
checkpoint['hidden_layers'])
model.load_state_dict(checkpoint['state_dict'])
return model
model = load_checkpoint('checkpoint.pth')
print(model)
```
| github_jupyter |
# Fraction field
```
from konfoo import Index, Byteorder, Fraction
```
## Item
Item type of the `field` class.
```
Fraction.item_type
```
Checks if the `field` class is a `bit` field.
```
Fraction.is_bit()
```
Checks if the `field` class is a `boolean` field.
```
Fraction.is_bool()
```
Checks if the `field` class is a `decimal` number field.
```
Fraction.is_decimal()
```
Checks if the `field` class is a `floating point` number field.
```
Fraction.is_float()
```
Checks if the `field` class is a `pointer` field.
```
Fraction.is_pointer()
```
Checks if the `field` class is a `stream` field.
```
Fraction.is_stream()
```
Checks if the `field` class is a `string` field.
```
Fraction.is_string()
```
## Field
```
fraction = Fraction(bits_integer=2, bit_size=16, align_to=None, signed=False, byte_order='auto')
fraction = Fraction(2, 16)
```
### Field view
```
fraction
str(fraction)
repr(fraction)
```
### Field name
```
fraction.name
```
### Field index
```
fraction.index
```
Byte `index` of the `field` within the `byte stream`.
```
fraction.index.byte
```
Bit offset relative to the byte `index` of the `field` within the `byte stream`.
```
fraction.index.bit
```
Absolute address of the `field` within the `data source`.
```
fraction.index.address
```
Base address of the `byte stream` within the `data source`.
```
fraction.index.base_address
```
Indexes the `field` and returns the `index` after the `field`.
```
fraction.index_field(index=Index())
```
### Field alignment
```
fraction.alignment
```
Byte size of the `field group` which the `field` is *aligned* to.
```
fraction.alignment.byte_size
```
Bit offset of the `field` within its *aligned* `field group`.
```
fraction.alignment.bit_offset
```
### Field size
```
fraction.bit_size
```
### Field byte order
```
fraction.byte_order
fraction.byte_order.value
fraction.byte_order.name
fraction.byte_order = 'auto'
fraction.byte_order = Byteorder.auto
```
### Field value
Checks if the decimal `field` is signed or unsigned.
```
fraction.signed
```
Maximal decimal `field` value.
```
fraction.max()
```
Minimal decimal `field` value.
```
fraction.min()
```
Returns the fraction `field` value as an floating point number.
```
fraction.value
```
Returns the decimal `field` value *aligned* to its `field group` as a number of bytes.
```
bytes(fraction)
bytes(fraction).hex()
```
Returns the decimal `field` value as an integer number.
```
int(fraction)
```
Returns the decimal `field` value as an floating point number.
```
float(fraction)
```
Returns the decimal `field` value as a lowercase hexadecimal string prefixed with `0x`.
```
hex(fraction)
```
Returns the decimal `field` value as a binary string prefixed with `0b`.
```
bin(fraction)
```
Returns the decimal `field` value as an octal string prefixed with `0o`.
```
oct(fraction)
```
Returns the decimal `field` value as a boolean value.
```
bool(fraction)
```
Returns the decimal `field` value as a signed integer number.
```
fraction.as_signed()
```
Returns the decimal `field` value as an unsigned integer number.
```
fraction.as_unsigned()
```
### Field metadata
Returns the ``meatadata`` of the ``field`` as an ordered dictionary.
```
fraction.describe()
```
### Deserialize
```
fraction.deserialize(bytes.fromhex('0100'), byte_order='little')
fraction.value
bytes(fraction)
bytes(fraction).hex()
int(fraction)
float(fraction)
hex(fraction)
bin(fraction)
oct(fraction)
bool(fraction)
```
### Serialize
```
buffer = bytearray()
fraction.value = 1
fraction.value = 1.0
fraction.value = 0x1
fraction.value = 0b1
fraction.value = 0o1
fraction.value = True
fraction.value = 1.0
fraction.serialize(buffer, byte_order='little')
buffer.hex()
bytes(fraction).hex()
```
| github_jupyter |
# AUDIOMATE Processing
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import IPython
import audiomate
from audiomate import containers
from audiomate.processing import pipeline
```
First the dataset is loaded. It is in the format of the UrbanSound8k dataset, but just 3 files of it.
```
urbansound8k_subset = audiomate.Corpus.load('data/urbansound_subset', reader='urbansound8k')
```
## Extracting Mel-Spectrogram Features with deltas
```
feature_path = 'output/mel_features.h5py'
feature_online_path = 'output/mel_features_online.h5py'
frame_size = 1024
hop_size = 512
sampling_rate = 16000
```
In order to compute the features with deltas multiple steps have to be executed. These steps are arranged in a pipeline using the `parent` and `parents` attributes of the different steps. The steps itself mostly just wrap corresponding functions of the `librosa` library.
```
mel_extractor = pipeline.MelSpectrogram(n_mels=60)
power_to_db = pipeline.PowerToDb(ref=np.max, parent=mel_extractor)
deltas = pipeline.Delta(parent=power_to_db)
stack = pipeline.Stack(parents=[power_to_db, deltas])
```
Now we have defined the pipeline. Now we can execute some processing function on the last step to execute the full pipeline. The default methods are running in offline mode. This means all frames of an utterance are processed at once.
```
feat_container = stack.process_corpus(urbansound8k_subset,
feature_path,
frame_size=frame_size,
hop_size=hop_size,
sr=sampling_rate)
```
To process the frames in streaming mode, we have to use the corresponding `_online` method. Now we can define via ``chunk_size`` how much frames at a time are processed. Depending on the pipeline steps we can define if context has to be added for processing. For example when computing deltas some context is required. There is no automatic check implemented if a step is only working in online or offline mode. It is up to the user to apply it correctly.
```
# The online processing would work the same way
# Here the problem is, that currently resampling in online mode is not supported
# Therefore we get an error when processing utterances with different sampling-rates
try:
feat_container_online = stack.process_corpus_online(
urbansound8k_subset,
feature_online_path,
frame_size=frame_size,
hop_size=hop_size
)
except ValueError as e:
print(e)
```
### Reading features
The features are written to a HDF5 file. These can be read using the `FeatureContainer` class.
```
feat_container = containers.FeatureContainer(feature_path)
with feat_container:
for utterance in urbansound8k_subset.utterances.values():
samples_of_utterance = utterance.read_samples(sr=sampling_rate)
features_of_utterance = feat_container.get(utterance.idx, mem_map=False)
print('{} ({}) ({} s)'.format(utterance.idx,
utterance.label_lists[audiomate.corpus.LL_SOUND_CLASS][0].value,
utterance.duration))
print('-'*100)
# Display audio player
IPython.display.display(IPython.display.Audio(samples_of_utterance, rate=sampling_rate) )
# Display Features
fig, ax = plt.subplots(figsize=(30,7))
im = plt.imshow(features_of_utterance.T)
ax.set_xlabel('Frame')
ax.set_ylabel('Bin')
plt.show()
```
| github_jupyter |
```
import os
import numpy as np
import sys
import torch
import pandas as pd
df_dataset = pd.read_json (r'all_data\cardiacSA_withOmmisions.json')
df_extract = df_dataset[['ID_XNAT','joint_self','frames']]
df_extract
# writing down all landmarks to a text file.
#be sure you know the what code is doing before you try it.
for i in range(df_extract.shape[0]):
filename = 'C:/Users/Alvin/Desktop/dissertation/landmark_touse/'+ str(df_extract['ID_XNAT'][i]) + '.txt'
f = open(filename, 'a')
for item in df_extract['joint_self'][i]:
f.writelines([str(item[0]),' ',str(item[1]),' ',str(item[2]),'\n'])
f.close()
# writing down landmarks and center landmarks to a text file
# only used when adding center point
for i in range(df_extract.shape[0]):
filename = 'C:/Users/Alvin/Desktop/dissertation/landmark_touse/'+ str(df_extract['ID_XNAT'][i]) + '.txt'
f = open(filename, 'a')
sum_x = 0
sum_y = 0
for item in df_extract['joint_self'][i]:
sum_x += item[0]
sum_y += item[1]
f.writelines([str(item[0]),' ',str(item[1]),' ',str(item[2]),'\n'])
sum_x = sum_x//3
sum_y = sum_y//3
f.writelines([str(sum_x),' ',str(sum_y),' ',str(item[2]),'\n'])
f.close()
# split the dataset by writing data and test data id into two text files
train_name = 'C:/Users/Alvin/Desktop/dissertation/data/'+'data_train.txt'
test_name = 'C:/Users/Alvin/Desktop/dissertation/data/'+'data_test.txt'
for i in range(df_extract.shape[0]):
if i < 280:
f_1 = open(train_name, 'a')
f_1.writelines([str(df_extract['ID_XNAT'][i]),'\n'])
if i >= 280:
f_2 = open(test_name, 'a')
f_2.writelines([str(df_extract['ID_XNAT'][i]),'\n'])
f_1.close()
f_2.close()
# copy image to the destination file location
import shutil
input_path = 'C:/Users/Alvin/Desktop/dissertation/all_data/'
out_path = 'C:/Users/Alvin/Desktop/dissertation/data/data'
df_extract['ID_XNAT'][i]
for i in range(df_extract.shape[0]):
input_data = 'C:/Users/Alvin/Desktop/dissertation/all_data/'+str(df_extract['frames'][i]['1'])
out_path = 'C:/Users/Alvin/Desktop/dissertation/data/data/'+str(df_extract['ID_XNAT'][i])+'.png'
shutil.copy(input_data,out_path)
from skimage import io
from skimage import transform as sk_transform
from skimage.transform import rescale, resize, downscale_local_mean
import torch
import seaborn as sns
import matplotlib.pyplot as plt
from scipy import stats
import glob
import torchvision.transforms as transforms
from PIL import Image
# load image as image object
def load_image(img_path):
im = io.imread(img_path, as_gray=True)
return im
# load image as image with shape of 101*101, downsized
def load_image_101(img_path):
im = io.imread(img_path, as_gray=True)
image_resized = resize(im,(101,101), anti_aliasing=True)
return image_resized
#load image as image object and using mean-std normalization
def image_reshape_to_numpy(img):
img = np.asarray(img)
img = image_normalized(img)
img_arr = img.reshape(img.shape[0], img.shape[1],1).astype('float32')
return img_arr
# load image as image object and using min-max normalization
def image_reshape_to_numpy_v2(img):
img = np.asarray(img)
img = image_normalized_v2(img)
img_arr = img.reshape(img.shape[0], img.shape[1],1).astype('float32')
return img_arr
# mean-std normalization
def image_normalized(ndarray):
arr_mean = np.mean(ndarray)
arr_std = np.std(ndarray)
result = (ndarray-arr_mean)/arr_std
return result
#min-max normalization
def image_normalized_v2(ndarray):
arr_min = np.min(ndarray).astype('float32')
arr_max = np.max(ndarray).astype('float32')
result = (ndarray.astype('float32')-arr_min)/(arr_max-arr_min)
return result
img_path = 'C:/Users/Alvin/Desktop/dissertation/data/data/PHD_78.png'
img_ = load_image(img_path)
norm = image_normalized_v2(img_)
plt.figure(figsize=(10,5))
plt.subplot(2,2,1)
plt.title("Before Normalisation")
plt.plot(img_)
plt.subplot(2,2,2)
plt.title("After Normalisation")
plt.plot(norm)
plt.subplot(2,2,3)
plt.imshow(img_)
plt.subplot(2,2,4)
plt.imshow(norm)
plt.show()
def extract_landmark_3D(filepath):
f= open(filepath, 'r')
line = f.readlines()
landmark_list = []
for l in line:
row = l.split(' ')
row[0], row[1], row[2] = row[0].replace('\'',''), row[1].replace('\'',''),row[2].replace('\'','')
landmark_list.append([int(float(row[0])),int(float(row[1])),int(float(row[2]))])
f.close()
return landmark_list
import matplotlib.pyplot as plt
from skimage import io
from skimage import transform as sk_transform
from skimage.transform import rescale, resize, downscale_local_mean
import torch
import glob
test_path = 'C:/Users/Alvin/Desktop/dissertation/result/model2_train_290_4/landmarks/test/'
gt_path = 'C:/Users/Alvin/Desktop/dissertation/Landmark_detection_PIN_center/landmark_detection_PIN/data_2d/landmarks/'
img_file = 'PHD_78_ps.txt'
gt_file = 'PHD_78.txt'
img_path = 'C:/Users/Alvin/Desktop/dissertation/data/data/PHD_78.png'
# load image as image object
def load_image(img_path):
im = io.imread(img_path, as_gray=True)
return im
# print all landmarks
def show_landmarks(image, landmarks,color):
"""Show image with landmarks"""
plt.imshow(image)
for i in range(len(landmarks)):
plt.scatter(landmarks[i][0],landmarks[i][1],s=100,c=color)
# print landmarks with center point landmark
def show_landmarks_center(image, landmarks,color):
"""Show image with landmarks"""
plt.imshow(image)
for i in range(len(landmarks)):
plt.scatter(landmarks[i][0],landmarks[i][1],s=100,c=color)
plt.scatter(landmarks[len(landmarks)-1][0],landmarks[len(landmarks)-1][1],s=100,c='c')
# print landmarks excluding the center point
def show_landmarks_test(image, landmarks,color):
"""Show image with landmarks"""
plt.imshow(image)
for i in range(len(landmarks)-1):
plt.scatter(landmarks[i][0],landmarks[i][1],s=100,c=color)
def extract_landmark_2d(filepath):
f= open(filepath, 'r')
line = f.readlines()
landmark_list = []
for l in line:
row = l.split(' ')
row[0], row[1] = row[0].replace('\'',''), row[1].replace('\'','')
landmark_list.append([int(float(row[0])),int(float(row[1]))])
f.close()
return landmark_list
landmark_list_true = extract_landmark_2d(gt_path+ gt_file)
landmark_list_pred = extract_landmark_2d(test_path+ img_file)
plt.figure(figsize=(15,15))
plt.subplot(1,2,1)
plt.title("Ground True")
show_landmarks(load_image(img_path),
landmark_list_true,'r')
plt.subplot(1,2,2)
plt.title("Prediction")
show_landmarks(load_image(img_path),
landmark_list_pred,'y')
plt.show()
# calculate result
test_path = 'C:/Users/Alvin/Desktop/dissertation/result/model2_train_290_4/error.txt'
img_file = 'PHD_6243_ps.txt'
def extract_landmark(filepath):
f= open(filepath, 'r')
line = f.readlines()
landmark_list = []
for l in line:
row = l.split(' ')
row[0], row[1] = row[0].replace('\'',''), row[1].replace('\'','')
landmark_list.append([int(float(row[0])),int(float(row[1]))])
f.close()
return landmark_list
# extract the original 3 landmarks for calculation.
#
f= open(test_path, 'r')
line = f.readlines()
error_list = []
error_list_full = []
error_list_x = []
error_list_y = []
error_list_z = []
for l in line:
row = l.split(' ')
row[1], row[2],row[3] = row[1].replace('\'',''), row[2].replace('\'',''),row[3].replace('\'','')
error_list.append([float(row[1]),float(row[2]),float(row[3])])
error_list_full.append(float(row[1]))
error_list_full.append(float(row[2]))
error_list_full.append(float(row[3]))
error_list_x.append(float(row[1]))
error_list_y.append(float(row[2]))
error_list_z.append(float(row[3]))
f.close()
# used in center point method
# calculate the std and mean based on results with the center point method applied from the original 3 landmarks
import statistics
def calculate_std_mean(input_list):
std_result = statistics.stdev(input_list)
mean_result = statistics.mean(input_list)
return std_result,mean_result
std_,mean_ = calculate_std_mean(error_list_full)
print('std:',std_)
print('mean:',mean_)
std_,mean_ = calculate_std_mean(error_list_x)
print('std:',std_)
print('mean:',mean_)
std_,mean_ = calculate_std_mean(error_list_y)
print('std:',std_)
print('mean:',mean_)
std_,mean_ = calculate_std_mean(error_list_z)
print('std:',std_)
print('mean:',mean_)
import glob
import re
test_file = glob.glob(r'C:/Users/Alvin/Desktop/dissertation/result/model2_train_290_4/landmarks/test/*.txt')
gt_path = 'C:/Users/Alvin/Desktop/dissertation/Landmark_detection_PIN_center/landmark_detection_PIN/data_2d/landmarks_4/'
img_path = 'C:/Users/Alvin/Desktop/dissertation/data/data/'
test_print =[]
gt_print = []
img_print = []
for i in test_file:
a = re.split('\\\\', i)
b = a[-1].replace('_ps','')
c = b.replace('.txt','.png')
test_print.append(a[-1])
gt_print.append(b)
img_print.append(c)
def print_results(img_gt,img_test,img_name):
landmark_list_true = extract_landmark_2d(gt_path+ img_gt)
landmark_list_pred = extract_landmark_2d(test_path+ img_test)
plt.figure(figsize=(15,15))
plt.subplot(1,2,1)
plt.title("Ground True")
show_landmarks_center(load_image(img_path+img_name),
landmark_list_true,'r')
plt.subplot(1,2,2)
plt.title("Prediction")
show_landmarks_test(load_image(img_path+img_name),
landmark_list_pred,'y')
plt.show()
for i in range(len(test_print)):
print_results(gt_print[i],test_print[i],img_print[i])
```
| github_jupyter |
## K26 - Heated Wall - Part 3: Results
Interface at 90°.
Equal fluid densities
Also no Heat capacity => infinitely fast heat conduction
Height of the domain is reduced
#### Instructions
This worksheet serves as a basis to conduct various parameter studies for the Heated Wall setup.
### Step 1 - Initialization
Load the BoSSS code, do not change
```
//#r "..\..\..\src\L4-application\BoSSSpad\bin\Release\net5.0\BoSSSpad.dll"
#r "BoSSSpad.dll"
using System;
using System.Collections.Generic;
using System.Linq;
using ilPSP;
using ilPSP.Utils;
using BoSSS.Platform;
using BoSSS.Foundation;
using BoSSS.Foundation.XDG;
using BoSSS.Foundation.Grid;
using BoSSS.Foundation.Grid.Classic;
using BoSSS.Foundation.IO;
using BoSSS.Solution;
using BoSSS.Solution.Control;
using BoSSS.Solution.GridImport;
using BoSSS.Solution.Statistic;
using BoSSS.Solution.Utils;
using BoSSS.Solution.AdvancedSolvers;
using BoSSS.Solution.Gnuplot;
using BoSSS.Application.BoSSSpad;
using BoSSS.Application.XNSE_Solver;
using static BoSSS.Application.BoSSSpad.BoSSSshell;
Init();
```
### Step 2 - Load postprocessed data
```
using System.Data;
using System.IO;
DataSet DataSet = new DataSet();
using(var tr = new StringReader(File.ReadAllText(".\\HeatedWall_Validation_SessionTable.json"))) {
var obj = TableExtensions.Deserialize(tr.ReadToEnd());
obj.TableName = "sessionTable";
DataSet.Tables.Add(obj);
}
using(var tr = new StringReader(File.ReadAllText(".\\HeatedWall_Validation_StudyTable.json"))) {
var obj = TableExtensions.Deserialize(tr.ReadToEnd());
obj.TableName = "studyTable";
DataSet.Tables.Add(obj);
}
```
### Step 3 - Data Visualization
#### Step 3.1 - Visualize various quantities of all studies
```
// Define which values are to be plotted
// =====================================
List<(string column, int index, string value)> vals2Plt = new List<(string column, int index, string value)>(); // (Column with Plot2Ddata[], Entry of Plot2Ddata)
vals2Plt.Add(("Contactline Plot Data", 1, "Contactline Position")); // contact line position
vals2Plt.Add(("Contactline Plot Data", 4, "Contact Angle")); // contact angle
vals2Plt.Add(("Convergence Data", 0, "Convergence"));
vals2Plt.Add(("Convergence Data", 1, "Convergence"));
vals2Plt.Add(("Convergence Data", 2, "Convergence"));
vals2Plt.Add(("Convergence Data", 3, "Convergence"));
vals2Plt.Add(("Convergence Data", 4, "Convergence"));
// =====================================
Gnuplot gp = new Gnuplot();
int plotCount = vals2Plt.Count;
int studyCount = DataSet.Tables["studyTable"].Rows.Count;
gp.SetMultiplot(studyCount, plotCount);
for(int i = 0; i < studyCount; i++){
for(int j = 0; j < plotCount; j++){
var row = DataSet.Tables["studyTable"].Rows[i];
gp.SetSubPlot(i, j, row["Study Description"].ToString());
Plot2Ddata[] plotData = row.Field<Plot2Ddata[]>(vals2Plt[j].column);
var data = plotData[vals2Plt[j].index];
data.ModFormat();
data.Title = "Study#" + i + " - " + vals2Plt[j].value;
data.TitleFont = 24;
data.LabelFont = 14;
data.LegendAlignment = new string[] {"o","c","b"};
data.ToGnuplot(gp);
}
}
gp.PlotSVG(800 * plotCount, 800 * studyCount)
```
| github_jupyter |
# Prova I- Computação Centífica II
> Aluno: Gil Miranda<br>
> DRE: 118037119<br>
> E-mail: gil.neto@ufrj.br; gilsmneto@gmail.com
## Set-up of dependencies and libraries
```
import numpy as np
import matplotlib.pyplot as plt
##### Vectorized 4th Order Runge Kutta
### Input: F -> Differential equation;
### y0 -> list or scalar for initial condition;
### ts -> list of points on time to evaluate the equation;
### p -> list or scalar for parameters for F, default is set to 0 if F has no extra parameters;
### Output: ys -> numpy array with all solutions for each step t, ys is a Matrix
##### Gil Miranda - last revision 26/10/2019
def rk_4(F, y0, ts, p = 0):
ys = [y0]
t = ts[0]
h = ts[1] - ts[0]
for tnext in ts:
k1 = h*F(t, ys[-1], p)
k2 = h*F(t + h/2, ys[-1] + k1/2, p)
k3 = h*F(t + h/2, ys[-1] + k2/2, p)
k4 = h*F(t + h, ys[-1] + k3)
ynext = ys[-1] + (k1/6+k2/3+k3/3+k4/6)
ys.append(ynext)
t = tnext
return ys[:-1]
```
---
## Creating the 4th and 3rd order Adams-Bashforth Method
The Adams-Bashforth method can be written as
$$
u_{k} = u_{k-1} + h\left(f_{k-1} + \frac{1}{2}\nabla f_{k-1}+ \frac{5}{12}\nabla^2 f_{k-1}+ \frac{9}{24}\nabla^3 f_{k-1} + \dots \right)
$$
And we can aproximate
$$
\begin{align}
\nabla f_{k-1} &= f_{k-1} - f_{k-2}\\
\nabla^2 f_{k-1} &= f_{k-1} - 2f_{k-2} + f_{k-3}\\
\nabla^3 f_{k-1} &= f_{k-1} - 3f_{k-2} + 3f_{k-3} - f_{k-4}\\
\end{align}
$$
So we have
$$
u_{k} = u_{k-1} + h\left(f_{k-1} + \frac{1}{2}(f_{k-1} - f_{k-2})+ \frac{5}{12}(f_{k-1} - 2f_{k-2} + f_{k-3}) + \frac{9}{24}(f_{k-1} - 3f_{k-2} + 3f_{k-3} - f_{k-4})\right)
$$
And collecting terms for the 3rd order and 4th order method, we can end with
$$
u_{k} = u_{k-1} + h\left(\frac{23}{12}f_{k-1} - \frac{16}{12}f_{k-2} + \frac{5}{12}f_{k-3}\right)
$$
for the 3rd order, and
$$
u_{k} = u_{k-1} + h\left(\frac{55}{24}f_{k-1} - \frac{59}{24}f_{k-2} + \frac{37}{24}f_{k-3} - \frac{9}{24}f_{k-4}\right)
$$
for the 4th order method<br>
We have to know the $f_{k-1},\dots, f_{k-n}$ points to solve using AB nth order method , we can solve for this $n$ points using Runge-Kutta
```
##### Adams-Bashfort 4th & 3rd order
### Input: F -> Differential equation;
### t0 -> initial point in time;
### tf -> final point in time;
### y0 -> initial condition;
### h -> step size;
### order -> order of the method;
### Output: ys -> list with all solutions for each step t;
### ts -> list with all points in time;
##### Gil Miranda - last revision 29/10/2019
def adams_bash(F, t0, tf, y0, h, order = 3):
if order == 4:
ws = [55/24, -59/24, 37/24, -9/24] ## setting the weights for 4th order
else:
order = 3
ws = [23/12, -16/12, 5/12] ## setting the weights for 3rd order
### initializing the list of points in time
ts = [t0]
for i in range(order-1):
ts.append(ts[-1] + h)
### solving for the first n points with runge-kutta 4th order, so we can initiliaze AB method
first_ys = rk_4(F, y0, ts)
n = len(first_ys)
ys = first_ys[:] ## list of solutions, initiliazed with rk4
### Adams-Bashfort
while(ts[-1] <= tf):
fs = [F(ts[-i], ys[-i]) for i in range(1,n+1)] ## list with f_k-1,...,f_k-n
ynew = ys[-1]
for (wi,fi) in zip(ws,fs):
ynew += h*(wi*fi)
ys.append(ynew)
ts.append(ts[-1] + h)
return ys, ts
```
#### The differential Equation
Here is the function that returns the differential equation
$$
\frac{\mathrm{d}u}{\mathrm{d}t} = 10e^{-\frac{(t-2)^2}{2(0.075)^2}} - 0.6u
$$
```
def eq_1(t,u, p=0):
a = -(t-2)**2/(2*(0.075)**2)
b = 10*np.e**a
c = 0.6*u
return b-c
```
#### Solving the Differential Equation with Adams-Bashforth 4th and 3rd Order
```
y0 = 0.5 # Initial Condition
hs = [0.01, 0.04, 0.06, 0.08] ## step size
hs_name = [hs[0], hs[0], hs[1], hs[1], hs[2], hs[2], hs[3], hs[3]] ## step size names for plotting
ys_3 = []
ts_3 = []
ys_4 = []
ts_4 = []
for h in hs:
y_3, t_3 = adams_bash(eq_1, 0, 4, y0, h) ## solving order 3 for each step size
y_4, t_4 = adams_bash(eq_1, 0, 4, y0, h, order = 4) ## solving order 4 for each step size
ys_3.append(y_3)
ts_3.append(t_3)
ys_4.append(y_4)
ts_4.append(t_4)
## plotting time
fig, ((ax1,ax2), (ax3,ax4), (ax5, ax6), (ax7, ax8)) = plt.subplots(nrows=4, ncols=2, figsize=(14,18))
fig.suptitle('Numerical Solution with Adams-Bashforth', weight='bold')
names = ['3rd', '4th', '3rd', '4th', '3rd', '4th', '3rd', '4th']
axs = [ax1, ax2, ax3, ax4, ax5, ax6, ax7, ax8]
for i,j,h in zip(axs,names,hs_name):
i.set_title(j + ' Order AB Method - step = $' + str(h) + '$')
for a in axs:
a.grid(alpha = 0.5)
ax1.plot(ts_3[0],ys_3[0], color = 'red')
ax2.plot(ts_4[0],ys_3[0], color = 'red')
ax3.plot(ts_3[1],ys_3[1], color = 'green')
ax4.plot(ts_4[1],ys_4[1], color = 'green')
ax5.plot(ts_3[2],ys_3[2], color = 'blue')
ax6.plot(ts_4[2],ys_4[2], color = 'blue')
ax7.plot(ts_3[3],ys_3[3], color = 'black')
ax8.plot(ts_4[3],ys_4[3], color = 'black')
plt.show()
```
#### Visualizing error between 3rd and 4th orders
```
## calculating the local error
err = []
for j in range(3):
e = [abs(ys_4[j][i] - abs(ys_3[j][i])) for i in range(len(ys_4[j]))]
err.append(e)
## plotting time
fig, ((ax1), (ax2), (ax3)) = plt.subplots(nrows=1, ncols=3, figsize=(20,5))
fig.suptitle('Local error between 3rd and 4th order methods')
axs = [ax1, ax2, ax3]
for i,h in zip(axs,hs):
i.set_title('step = $' + str(h) + '$')
for a in axs:
a.grid(alpha = 0.5)
ax1.plot(ts_3[0],err[0], color = 'red')
ax2.plot(ts_4[1],err[1], color = 'green')
ax3.plot(ts_3[2],err[2], color = 'blue')
plt.show()
```
---
## Adaptative Adams-Bashforth Method
```
##### Adaptative Adams-Bashfort 4th & 3rd order
### Input: F -> Differential equation;
### t0 -> initial point in time;
### tf -> final point in time;
### y0 -> initial condition;
### h -> step size;
### tol -> error tolerance;
### Output: ys -> list with all solutions for each step t;
### ts -> list with all points in time;
##### Gil Miranda - last revision 30/10/2019
def adams_bash_adp(F, t0, tf, y0, h, tol = 1e-3):
ws_4 = [55/24, -59/24, 37/24, -9/24] ## setting the weights for 4th order
ws_3 = [23/12, -16/12, 5/12] ## setting the weights for 3rd order
### initializing the list of points in time
ts = [t0]
for i in range(4):
ts.append(ts[-1] + h)
### solving for the first n points with runge-kutta 4th order, so we can initiliaze AB method
first_ys = rk_4(F, y0, ts)
ys = first_ys[:] ## list of solutions, initiliazed with rk4
### Adptative Adams-Bashfort
while(ts[-1] <= tf):
fs = [F(ts[-i], ys[-i]) for i in range(1,5)] ## list with f_k-1,...,f_k-n
## Solving with 3rd order
ynew = ys[-1]
for (wi,i) in zip(ws_3, range(3)):
ynew += h*(wi* fs[i])
## Solving with 4th order (using 3rd order points)
ynew_til = ys[-1]
for (wi, i) in zip(ws_4, range(4)):
ynew_til += h*(wi * fs[i])
## Local error
err = abs(ynew_til - ynew)
## if err < tol we accept the solution
if err <= tol:
ys.append(ynew)
tnext = ts[-1] + h
ts.append(tnext)
h = h*(tol/err)**(1/4)
else:
## else, we get a new step
h = h/2
return ys, ts
```
### Visualizing Solutions
```
t0 = 0
tf = 4
h = 0.1
y0 = 0.5
ys_1, ts_1 = adams_bash_adp(eq_1, t0, tf, y0, h, tol = 1e-3)
ys_2, ts_2 = adams_bash_adp(eq_1, t0, tf, y0, h, tol = 1e-4)
## plotting time
fig, (ax1, ax2) = plt.subplots(nrows=1, ncols=2, figsize=(14,5))
fig.suptitle('Numerical Solution with Adaptative Adams-Bashforth, initial step size = $' + str(h) + '$', weight='bold')
# names = ['3rd', '4th', '3rd', '4th', '3rd', '4th']
axs = [ax1, ax2]
for a in axs:
a.grid(alpha = 0.5)
ax1.plot(ts_1,ys_1, color = 'black')
ax1.plot(ts_1,ys_1, 'o', color = 'red')
ax1.set_title('tol = $10^{-3}$')
ax2.plot(ts_2,ys_2, color = 'black')
ax2.plot(ts_2,ys_2, 'o', color = 'blue')
ax2.set_title('tol = $10^{-4}$')
plt.show()
```
---
### Bonus Method: Predictor-Corrector Adams-Bashforth-Moulton 4th order Method
Predictor equation (4th order AB)
$$
u_{k} = u_{k-1} + h\left(\frac{55}{24}f_{k-1} - \frac{59}{24}f_{k-2} + \frac{37}{24}f_{k-3} - \frac{9}{24}f_{k-4}\right)
$$
Corrector equation (4th order AM)
$$
u_{k} = u_{k-1} + h\left(\frac{9}{24}f_{k} + \frac{19}{24}f_{k-1} - \frac{5}{24}f_{k-2} + \frac{1}{24}f_{k-3}\right)
$$
We initialize the AB with RK4, and then uses AB $f_k$ to initialize the AM, and then compute the error between the steps, if $err > tol$, tol is given tolerance, calculate again
```
##### Adams-Bashfort-Moulton 4th order
### Input: F -> Differential equation;
### t0 -> initial point in time;
### tf -> final point in time;
### y0 -> initial condition;
### h -> step size;
### tol -> error tolerance;
### Output: ys -> list with all solutions for each step t;
### ts -> list with all points in time;
##### Gil Miranda - last revision 30/10/2019
def adams_bash_mou(F, t0, tf, y0, h, tol = 1e-12):
ws_ab = [55/24, -59/24, 37/24, -9/24] ## setting the weights for adams bashforth
ws_am = [9/24, 19/24, -5/24, 1/24] ## setting the weights for adams moulton
### initializing the list of points in time
ts = [t0]
for i in range(3):
ts.append(ts[-1] + h)
### solving for the first n points with runge-kutta 4th order, so we can initiliaze AB method
first_ys = rk_4(F, y0, ts)
n = len(first_ys)
ys = first_ys[:] ## list of solutions, initiliazed with rk4
while(ts[-1] <= tf):
## Calculating the AB solution
fs_ab = [F(ts[-i], ys[-i]) for i in range(1,n+1)] ## list with f_k-1,...,f_k-n
y_ab = ys[-1]
for (wi,fi) in zip(ws_ab,fs_ab):
y_ab += h*(wi*fi)
## Generating next point in time
tnext = ts[-1] + h
## Generating the f(ti,yi) point to use with AM
f1_am = F(tnext, y_ab)
fs_am = [f1_am] + fs_ab[:-1]
## Calculating the AM solution
y_am = ys[-1]
for (wi,fi) in zip(ws_am,fs_am):
y_am += h*(wi*fi)
## error
err = abs(y_am - y_ab)
y_old = y_am
y_new = y_am
while(err > tol):
## Calculating another solution with AM using last AM solution
f1_new = F(tnext, y_old)
fs_new = [f1_new] + fs_ab[:-1]
y_new = ys[-1]
for (wi,fi) in zip(ws_am,fs_new):
y_new += h*(wi*fi)
err = abs(y_new - y_old)
y_old = y_new
ys.append(y_new)
ts.append(tnext)
return ys, ts
y0 = 0.5 # Initial Condition
hs = [0.01, 0.04, 0.06, 0.08] ## step size
hs_name = [hs[0], hs[1], hs[2], hs[3]] ## step size names for plotting
ys_5 = []
ts_5 = []
for h in hs:
y_5, t_5 = adams_bash_mou(eq_1, 0, 4, y0, h) ## solving for each step with predictor-corrector
ys_5.append(y_5)
ts_5.append(t_5)
## plotting time
fig, ((ax1,ax2), (ax3,ax4)) = plt.subplots(nrows=2, ncols=2, figsize=(14,10))
fig.suptitle('Numerical Solution with Adams-Bashforth-Moulton', weight='bold')
axs = [ax1, ax2, ax3, ax4]
for i,h in zip(axs,hs_name):
i.set_title('h = $' + str(h) + '$')
for a in axs:
a.grid(alpha = 0.5)
ax1.plot(ts_5[0],ys_5[0], color = 'red')
ax2.plot(ts_5[1],ys_5[1], color = 'blue')
ax3.plot(ts_5[2],ys_5[2], color = 'green')
ax4.plot(ts_5[3],ys_5[3], color = 'orange')
plt.show()
```
Unlike the AB 3rd and 4th order methods, we can get a decent solution even with step size $h = 0.08$ with the ABM method<br>
Plotting the solutions for each step-size with AB4, AB Adaptative, ABM
```
## plotting time
fig, ((ax1,ax2), (ax3,ax4)) = plt.subplots(nrows=2, ncols=2, figsize=(14,10))
fig.suptitle('Comparison of solutions for all methods', weight='bold')
axs = [ax1, ax2, ax3, ax4]
for i,h in zip(axs,hs_name):
i.set_title('h = $' + str(h) + '$')
for a in axs:
a.grid(alpha = 0.5)
ax1.plot(ts_5[0],ys_5[0], color = 'red', label = 'ABM')
ax1.plot(ts_4[0], ys_4[0], label = 'AB 4')
ax1.plot(ts_2, ys_2, label = 'AB Adp')
ax2.plot(ts_5[1],ys_5[1], color = 'blue', label = 'ABM')
ax2.plot(ts_4[1], ys_4[1], color = 'red', label = 'AB 4')
ax3.plot(ts_4[2], ys_4[2], color = 'orange', label = 'AB 4')
ax3.plot(ts_5[2],ys_5[2], color = 'green', label = 'ABM')
ax4.plot(ts_5[3],ys_5[3], color = 'orange', label = 'ABM')
ax4.plot(ts_4[3], ys_4[3], color = 'green', label = 'AB 4')
ax1.legend()
ax2.legend()
ax3.legend()
ax4.legend()
plt.show()
```
| github_jupyter |

<hr>
# Recurrent Neural Networks
Recurrent neural networks (RNN) are an extremely powerful tool in deep learning. These models quite accurately mimic how humans process information and learn. Unlike traditional feedforward neural networks, RNNs have memory. That is, information fed into them persists and the network is able to draw on this to make inferences.
##### Long Short-term Memory
Long Short-term Memory (LSTM) is a type of recurrent neural network. Instead of one layer, LSTM cells generally have four, three of which are part of "gates" -- ways to optionally let information through. The three gates are commonly referred to as the forget, input, and output gates. The forget gate layer is where the model decides what information to keep from prior states. At the input gate layer, the model decides which values to update. Finally, the output gate layer is where the final output of the cell state is decided. Essentially, LSTM separately decides what to remember and the rate at which it should update.
##### Financial Applications
LSTM models have produced some great results when applied to time-series prediction. One of the central challenges with conventional time-series models is that, despite trying to account for trends or other non-stationary elements, it is almost impossible to truly predict an outlier like a recession, flash crash, liquidity crisis, etc. By having a long memory, LSTM models are better able to capture these difficult trends in the data without suffering from the level of overfitting a conventional model would need in order to capture the same data.
For a very basic application, we're going to use a LSTM model to predict the price movement, a non-stationary time-series, of SPY.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.preprocessing import MinMaxScaler
# Import keras modules
from keras.layers import LSTM
from keras.layers import Dense
from keras.layers import Dropout
from keras.models import Sequential
qb = QuantBook()
symbol = qb.AddEquity("SPY").Symbol
# Fetch history
history = qb.History([symbol], 1280, Resolution.Daily)
# Fetch price
total_price = history.loc[symbol].close
training_price = history.loc[symbol].close[:1260]
test_price = history.loc[symbol].close[1260:]
# Transform price
price_array = np.array(training_price).reshape((len(training_price), 1))
# Scale data onto [0,1]
scaler = MinMaxScaler(feature_range = (0, 1))
# Transform our data
spy_training_scaled = scaler.fit_transform(price_array)
# Build feauture and label sets (using number of steps 60, batch size 1200, and hidden size 1)
features_set = []
labels = []
for i in range(60, 1260):
features_set.append(spy_training_scaled[i-60:i, 0])
labels.append(spy_training_scaled[i, 0])
features_set, labels = np.array(features_set), np.array(labels)
features_set = np.reshape(features_set, (features_set.shape[0], features_set.shape[1], 1))
features_set.shape
# Build a Sequential keras model
model = Sequential()
# Add our first LSTM layer - 50 nodes
model.add(LSTM(units = 50, return_sequences=True, input_shape=(features_set.shape[1], 1)))
# Add Dropout layer to avoid overfitting
model.add(Dropout(0.2))
# Add additional layers
model.add(LSTM(units=50, return_sequences=True))
model.add(Dropout(0.2))
model.add(LSTM(units=50, return_sequences=True))
model.add(Dropout(0.2))
model.add(LSTM(units=50))
model.add(Dropout(0.2))
model.add(Dense(units = 1))
# Compile the model
model.compile(optimizer = 'adam', loss = 'mean_squared_error', metrics=['mae', 'acc'])
# Fit the model to our data, running 50 training epochs
model.fit(features_set, labels, epochs = 50, batch_size = 32)
# Get and transform inputs for testing our predictions
test_inputs = total_price[-80:].values
test_inputs = test_inputs.reshape(-1,1)
test_inputs = scaler.transform(test_inputs)
# Get test features
test_features = []
for i in range(60, 80):
test_features.append(test_inputs[i-60:i, 0])
test_features = np.array(test_features)
test_features = np.reshape(test_features, (test_features.shape[0], test_features.shape[1], 1))
# Make predictions
predictions = model.predict(test_features)
# Transform predictions back to original data-scale
predictions = scaler.inverse_transform(predictions)
# Plot our results!
plt.figure(figsize=(10,6))
plt.plot(test_price.values, color='blue', label='Actual')
plt.plot(predictions , color='red', label='Prediction')
plt.title('Price vs Predicted Price ')
plt.legend()
plt.show()
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.