text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
```
#nuclio: ignore
import nuclio
%%nuclio cmd -c
pip install opencv-contrib-python
pip install pandas
pip install v3io_frames
import nuclio_sdk
import json
import os
import v3io_frames as v3f
from requests import post
import base64
import numpy as np
import pandas as pd
import cv2
import random
import string
from datetime import datetime
%%nuclio env
DATA_PATH = /User/demos/demos/realtime-face-recognition/dataset/
V3IO_ACCESS_KEY=${V3IO_ACCESS_KEY}
is_partitioned = True #os.environ['IS_PARTITIONED']
def generate_file_name(current_time, is_partitioned):
filename_str = current_time + '.jpg'
if is_partitioned == "true":
filename_str = current_time[:-4] + "/" + filename_str
return filename_str
def generate_image_path(filename, is_unknown):
file_name = filename
if is_unknown:
pathTuple = (os.environ['DATA_PATH'] + 'label_pending', file_name)
else:
pathTuple = (os.environ['DATA_PATH'] + 'images', file_name)
path = "/".join(pathTuple)
return path
def jpg_str_to_frame(encoded):
jpg_original = base64.b64decode(encoded)
jpg_as_np = np.frombuffer(jpg_original, dtype=np.uint8)
img = cv2.imdecode(jpg_as_np, flags=1)
return img
def save_image(encoded_img, path):
frame = jpg_str_to_frame(encoded_img)
directory = '/'.join(path.split('/')[:-1])
if not os.path.exists(directory):
os.mkdir(directory)
cv2.imwrite(path, frame)
def write_to_kv(client, face, path, camera, time):
rnd_tag = ''.join(random.choices(string.ascii_uppercase + string.digits, k=5))
name = face['name']
label = face['label']
encoding = face['encoding']
new_row = {}
new_row = {'c' + str(i).zfill(3): encoding[i] for i in range(128)}
if name != 'unknown':
new_row['label'] = label
new_row['fileName'] = name.replace(' ', '_') + '_' + rnd_tag
else:
new_row['label'] = -1
new_row['fileName'] = 'unknown_' + rnd_tag
new_row['imgUrl'] = path
new_row['camera'] = camera
new_row['time'] = datetime.strptime(time, '%Y%m%d%H%M%S')
new_row_df = pd.DataFrame(new_row, index=[0])
new_row_df = new_row_df.set_index('fileName')
print(new_row['fileName'])
client.write(backend='kv', table='iguazio/demos/demos/realtime-face-recognition/artifacts/encodings', dfs=new_row_df) #, save_mode='createNewItemsOnly')
def init_context(context):
setattr(context.user_data, 'client', v3f.Client("framesd:8081", container="users"))
def handler(context, event):
context.logger.info('extracting metadata')
body = json.loads(event.body)
time = body['time']
camera = body['camera']
encoded_img = body['content']
content = {'img': encoded_img}
context.logger.info('calling model server')
resp = context.platform.call_function('recognize-faces', event)
faces = json.loads(resp.body)
context.logger.info('going through discovered faces')
for face in faces:
is_unknown = face['name'] == 'unknown'
file_name = generate_file_name(time, is_partitioned)
path = generate_image_path(file_name, is_unknown)
context.logger.info('saving image to file system')
save_image(encoded_img, path)
context.logger.info('writing data to kv')
write_to_kv(context.user_data.client, face, path, camera, time)
return faces
#nuclio: end-code
# converts the notebook code to deployable function with configurations
from mlrun import code_to_function, mount_v3io
fn = code_to_function('video-api-server', kind='nuclio')
# set the API/trigger, attach the home dir to the function
fn.with_http(workers=2).apply(mount_v3io())
# set environment variables
fn.set_env('DATA_PATH', '/User/demos/demos/realtime-face-recognition/dataset/')
fn.set_env('V3IO_ACCESS_KEY', os.environ['V3IO_ACCESS_KEY'])
addr = fn.deploy(project='default')
```
| github_jupyter |
### Setup some basic stuff
```
import logging
logging.getLogger().setLevel(logging.DEBUG)
import folium
import folium.features as fof
import folium.utilities as ful
import branca.element as bre
import json
import geojson as gj
import arrow
import shapely.geometry as shpg
import pandas as pd
import geopandas as gpd
def lonlat_swap(lon_lat):
return list(reversed(lon_lat))
def get_row_count(n_maps, cols):
rows = (n_maps / cols)
if (n_maps % cols != 0):
rows = rows + 1
return rows
def get_marker(loc, disp_color):
if loc["geometry"]["type"] == "Point":
curr_latlng = lonlat_swap(loc["geometry"]["coordinates"])
return folium.Marker(curr_latlng, icon=folium.Icon(color=disp_color),
popup="%s" % loc["properties"]["name"])
elif loc["geometry"]["type"] == "Polygon":
assert len(loc["geometry"]["coordinates"]) == 1,\
"Only simple polygons supported!"
curr_latlng = [lonlat_swap(c) for c in loc["geometry"]["coordinates"][0]]
# print("Returning polygon for %s" % curr_latlng)
return folium.PolyLine(curr_latlng, color=disp_color, fill=disp_color,
popup="%s" % loc["properties"]["name"])
```
### Read the data
```
spec_to_validate = json.load(open("final_sfbayarea_filled/train_bus_ebike_mtv_ucb.filled.json"))
sensing_configs = json.load(open("sensing_regimes.all.specs.json"))
```
### Validating the time range
```
print("Experiment runs from %s -> %s" % (arrow.get(spec_to_validate["start_ts"]), arrow.get(spec_to_validate["end_ts"])))
start_fmt_time_to_validate = arrow.get(spec_to_validate["start_ts"]).format("YYYY-MM-DD")
end_fmt_time_to_validate = arrow.get(spec_to_validate["end_ts"]).format("YYYY-MM-DD")
if (start_fmt_time_to_validate != spec_to_validate["start_fmt_date"]):
print("VALIDATION FAILED, got start %s, expected %s" % (start_fmt_time_to_validate, spec_to_validate["start_fmt_date"]))
if (end_fmt_time_to_validate != spec_to_validate["end_fmt_date"]):
print("VALIDATION FAILED, got end %s, expected %s" % (end_fmt_time_to_validate, spec_to_validate["end_fmt_date"]))
```
### Validating calibration trips
```
def get_map_for_calibration_test(trip):
curr_map = folium.Map()
if trip["start_loc"] is None or trip["end_loc"] is None:
return curr_map
curr_start = lonlat_swap(trip["start_loc"]["coordinates"])
curr_end = lonlat_swap(trip["end_loc"]["coordinates"])
folium.Marker(curr_start, icon=folium.Icon(color="green"),
popup="Start: %s" % trip["start_loc"]["name"]).add_to(curr_map)
folium.Marker(curr_end, icon=folium.Icon(color="red"),
popup="End: %s" % trip["end_loc"]["name"]).add_to(curr_map)
folium.PolyLine([curr_start, curr_end], popup=trip["id"]).add_to(curr_map)
curr_map.fit_bounds([curr_start, curr_end])
return curr_map
calibration_tests = spec_to_validate["calibration_tests"]
rows = get_row_count(len(calibration_tests), 4)
calibration_maps = bre.Figure((rows,4))
for i, t in enumerate(calibration_tests):
if t["config"]["sensing_config"] != sensing_configs[t["config"]["id"]]["sensing_config"]:
print("Mismatch in config for test" % t)
curr_map = get_map_for_calibration_test(t)
calibration_maps.add_subplot(rows, 4, i+1).add_child(curr_map)
calibration_maps
```
### Validating evaluation trips
```
def get_map_for_travel_leg(trip):
curr_map = folium.Map()
get_marker(trip["start_loc"], "green").add_to(curr_map)
get_marker(trip["end_loc"], "red").add_to(curr_map)
# trips from relations won't have waypoints
if "waypoint_coords" in trip:
for i, wpc in enumerate(trip["waypoint_coords"]["geometry"]["coordinates"]):
folium.map.Marker(
lonlat_swap(wpc), popup="%d" % i,
icon=fof.DivIcon(class_name='leaflet-div-icon')).add_to(curr_map)
print("Found %d coordinates for the route" % (len(trip["route_coords"]["geometry"]["coordinates"])))
latlng_route_coords = [lonlat_swap(rc) for rc in trip["route_coords"]["geometry"]["coordinates"]]
folium.PolyLine(latlng_route_coords,
popup="%s: %s" % (trip["mode"], trip["name"])).add_to(curr_map)
for i, c in enumerate(latlng_route_coords):
folium.CircleMarker(c, radius=5, popup="%d: %s" % (i, c)).add_to(curr_map)
curr_map.fit_bounds(ful.get_bounds(trip["route_coords"]["geometry"]["coordinates"], lonlat=True))
return curr_map
def get_map_for_shim_leg(trip):
curr_map = folium.Map()
mkr = get_marker(trip["loc"], "purple")
mkr.add_to(curr_map)
curr_map.fit_bounds(mkr.get_bounds())
return curr_map
evaluation_trips = spec_to_validate["evaluation_trips"]
map_list = []
for t in evaluation_trips:
for l in t["legs"]:
if l["type"] == "TRAVEL":
curr_map = get_map_for_travel_leg(l)
map_list.append(curr_map)
else:
curr_map = get_map_for_shim_leg(l)
map_list.append(curr_map)
rows = get_row_count(len(map_list), 2)
evaluation_maps = bre.Figure(ratio="{}%".format((rows/2) * 100))
for i, curr_map in enumerate(map_list):
evaluation_maps.add_subplot(rows, 2, i+1).add_child(curr_map)
evaluation_maps
```
### Validating start and end polygons
```
def check_start_end_contains(leg):
points = gpd.GeoSeries([shpg.Point(p) for p in leg["route_coords"]["geometry"]["coordinates"]])
start_loc = shpg.shape(leg["start_loc"]["geometry"])
end_loc = shpg.shape(leg["end_loc"]["geometry"])
start_contains = points.apply(lambda p: start_loc.contains(p))
print(points[start_contains])
end_contains = points.apply(lambda p: end_loc.contains(p))
print(points[end_contains])
# Some of the points are within the start and end polygons
assert start_contains.any()
assert end_contains.any()
# The first and last point are within the start and end polygons
assert start_contains.iloc[0], points.head()
assert end_contains.iloc[-1], points.tail()
# The points within the polygons are contiguous
max_index_diff_start = pd.Series(start_contains[start_contains == True].index).diff().max()
max_index_diff_end = pd.Series(end_contains[end_contains == True].index).diff().max()
assert pd.isnull(max_index_diff_start) or max_index_diff_start == 1, "Max diff in index = %s for points %s" % (gpd.GeoSeries(start_contains[end_contains == True].index).diff().max(), points.head())
assert pd.isnull(max_index_diff_end) or max_index_diff_end == 1, "Max diff in index = %s for points %s" % (gpd.GeoSeries(end_contains[end_contains == True].index).diff().max(), points.tail())
invalid_legs = []
for t in evaluation_trips:
for l in t["legs"]:
if l["type"] == "TRAVEL" and l["id"] not in invalid_legs:
print("Checking leg %s, %s" % (t["id"], l["id"]))
check_start_end_contains(l)
```
### Validating sensing settings
```
for ss in spec_to_validate["sensing_settings"]:
for phoneOS, compare_map in ss.items():
compare_list = compare_map["compare"]
for i, ssc in enumerate(compare_map["sensing_configs"]):
if ssc["id"] != compare_list[i]:
print("Mismatch in sensing configurations for %s" % ss)
```
| github_jupyter |
```
%pylab inline
import numpy as np
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
from torch.optim.lr_scheduler import StepLR,MultiStepLR
import math
import torch.nn.functional as F
from torch.utils import data
from sklearn.model_selection import train_test_split
from torch.utils.data import Dataset
from ssc_dataset_f import my_Dataset
import os
torch.manual_seed(0)
batch_size = 32
train_dir = './data/train/'
train_files = [train_dir+i for i in os.listdir(train_dir)]
valid_dir = './data/valid/'
valid_files = [valid_dir+i for i in os.listdir(valid_dir)]
test_dir = './data/test/'
test_files = [test_dir+i for i in os.listdir(test_dir)]
train_dataset = my_Dataset(train_files)
valid_dataset = my_Dataset(valid_files)
test_dataset = my_Dataset(test_files)
train_loader = data.DataLoader(train_dataset,batch_size=batch_size,shuffle=True)#,num_workers=10)
valid_loader = data.DataLoader(valid_dataset,batch_size=batch_size,shuffle=True)#,num_workers=5)
test_loader = data.DataLoader(test_dataset,batch_size=batch_size,shuffle=True)#,num_workers=5)
'''
STEP 2: MAKING DATASET ITERABLE
'''
decay = 0.1 # neuron decay rate
thresh = 0.5 # neuronal threshold
lens = 0.5 # hyper-parameters of approximate function
num_epochs = 150 # 150 # n_iters / (len(train_dataset) / batch_size)
num_epochs = int(num_epochs)
'''
STEP 3a: CREATE spike MODEL CLASS
'''
b_j0 = 0.01 # neural threshold baseline
R_m = 1 # membrane resistance
dt = 1 #
gamma = .5 # gradient scale
gradient_type = 'MG'
print('gradient_type: ',gradient_type)
def gaussian(x, mu=0., sigma=.5):
return torch.exp(-((x - mu) ** 2) / (2 * sigma ** 2)) / torch.sqrt(2 * torch.tensor(math.pi)) / sigma
# define approximate firing function
class ActFun_adp(torch.autograd.Function):
@staticmethod
def forward(ctx, input): # input = membrane potential- threshold
ctx.save_for_backward(input)
return input.gt(0).float() # is firing ???
@staticmethod
def backward(ctx, grad_output): # approximate the gradients
input, = ctx.saved_tensors
grad_input = grad_output.clone()
# temp = abs(input) < lens
scale = 6.0
hight = .15
if gradient_type == 'G':
temp = torch.exp(-(input**2)/(2*lens**2))/torch.sqrt(2*torch.tensor(math.pi))/lens
elif gradient_type == 'MG':
temp = gaussian(input, mu=0., sigma=lens) * (1. + hight) \
- gaussian(input, mu=lens, sigma=scale * lens) * hight \
- gaussian(input, mu=-lens, sigma=scale * lens) * hight
elif gradient_type =='linear':
temp = F.relu(1-input.abs())
elif gradient_type == 'slayer':
temp = torch.exp(-5*input.abs())
return grad_input * temp.float() * gamma
act_fun_adp = ActFun_adp.apply
# tau_m = torch.FloatTensor([tau_m])
def mem_update_adp(inputs, mem, spike, tau_adp, b, tau_m, dt=1, isAdapt=1):
alpha = torch.exp(-1. * dt / tau_m).cuda()
ro = torch.exp(-1. * dt / tau_adp).cuda()
if isAdapt:
beta = 1.8
else:
beta = 0.
b = ro * b + (1 - ro) * spike
B = b_j0 + beta * b
mem = mem * alpha + (1 - alpha) * R_m * inputs - B * spike * dt
inputs_ = mem - B
spike = act_fun_adp(inputs_)
#spike = F.relu(inputs_)# # act_fun : approximation firing function
return mem, spike, B, b
# LIF neuron
def mem_update_adp1(inputs, mem, spike, tau_adp, b, tau_m, dt=1, isAdapt=1):
b = 0
B = .5
alpha = torch.exp(-1. * dt / tau_adp).cuda()
mem = mem * .7 + inputs#(1-alpha)*inputs
inputs_ = mem - B
spike = act_fun_adp(inputs_) # act_fun : approximation firing function
mem = (1-spike)*mem
return mem, spike, B, b
def output_Neuron(inputs, mem, tau_m, dt=1):
"""
The read out neuron is leaky integrator without spike
"""
# alpha = torch.exp(-1. * dt / torch.FloatTensor([30.])).cuda()
alpha = torch.exp(-1. * dt / tau_m).cuda()
mem = mem * alpha + (1. - alpha) * R_m * inputs
return mem
def output_Neuron1(inputs, mem, tau_m, dt=1):
"""
The read out neuron is leaky integrator without spike
"""
# alpha = torch.exp(-1. * dt / torch.FloatTensor([30.])).cuda()
alpha = torch.exp(-1. * dt / tau_m).cuda()
mem = mem * 0.7 + R_m * inputs
return mem
class RNN_custom(nn.Module):
def __init__(self, input_size, hidden_size, output_size):
super(RNN_custom, self).__init__()
self.hidden_size = hidden_size
# self.hidden_size = input_size
self.i_2_h1 = nn.Linear(input_size, hidden_size[0])
self.h1_2_h1 = nn.Linear(hidden_size[0], hidden_size[0])
self.h1_2_h2 = nn.Linear(hidden_size[0], hidden_size[1])
self.h2_2_h2 = nn.Linear(hidden_size[1], hidden_size[1])
self.h2o = nn.Linear(hidden_size[1], output_size)
self.tau_adp_h1 = nn.Parameter(torch.Tensor(hidden_size[0]))
self.tau_adp_h2 = nn.Parameter(torch.Tensor(hidden_size[1]))
self.tau_adp_o = nn.Parameter(torch.Tensor(output_size))
self.tau_m_h1 = nn.Parameter(torch.Tensor(hidden_size[0]))
self.tau_m_h2 = nn.Parameter(torch.Tensor(hidden_size[1]))
self.tau_m_o = nn.Parameter(torch.Tensor(output_size))
nn.init.orthogonal_(self.h1_2_h1.weight)
nn.init.orthogonal_(self.h2_2_h2.weight)
nn.init.xavier_uniform_(self.i_2_h1.weight)
nn.init.xavier_uniform_(self.h1_2_h2.weight)
nn.init.xavier_uniform_(self.h2_2_h2.weight)
nn.init.xavier_uniform_(self.h2o.weight)
nn.init.constant_(self.i_2_h1.bias, 0)
nn.init.constant_(self.h1_2_h2.bias, 0)
nn.init.constant_(self.h2_2_h2.bias, 0)
nn.init.constant_(self.h1_2_h1.bias, 0)
# saved
# nn.init.normal_(self.tau_adp_h1, 50,10)
# nn.init.normal_(self.tau_adp_h2, 50,10)
# nn.init.normal_(self.tau_adp_o, 50,10)
# nn.init.normal_(self.tau_m_h1, 20.,5)
# nn.init.normal_(self.tau_m_h2, 20.,5)
# nn.init.normal_(self.tau_m_o, 3.,1)
nn.init.normal_(self.tau_adp_h1, 200,50)
nn.init.normal_(self.tau_adp_h2, 200,50)
nn.init.normal_(self.tau_adp_o, 150,50)
nn.init.normal_(self.tau_m_h1, 20.,5)
nn.init.normal_(self.tau_m_h2, 20.,5)
nn.init.normal_(self.tau_m_o, 3.,1)
self.b_h1 = self.b_h2 = self.b_o = 0
def forward(self, input):
batch_size, seq_num, input_dim = input.shape
self.b_h1 = self.b_h2 = self.b_o = b_j0
mem_layer1 = spike_layer1 = torch.rand(batch_size, self.hidden_size[0]).cuda()
mem_layer2 = spike_layer2 = torch.rand(batch_size, self.hidden_size[1]).cuda()
mem_output = torch.zeros(batch_size,output_dim).cuda()
# mem_output_tmp = torch.rand(batch_size, output_dim).cuda()
output = torch.zeros(batch_size, output_dim).cuda()
hidden_spike_ = []
hidden_spike2_ = []
h2o_mem_ = []
fr = 0
for i in range(seq_num):
input_x = input[:, i, :]
h_input = self.i_2_h1(input_x.float()) + self.h1_2_h1(spike_layer1)
mem_layer1, spike_layer1, theta_h1, self.b_h1 = mem_update_adp(h_input, mem_layer1, spike_layer1,
self.tau_adp_h1, self.b_h1,self.tau_m_h1)
h2_input = self.h1_2_h2(spike_layer1) + self.h2_2_h2(spike_layer2)
mem_layer2, spike_layer2, theta_h2, self.b_h2 = mem_update_adp(h2_input, mem_layer2, spike_layer2,
self.tau_adp_h2, self.b_h2, self.tau_m_h2)
mem_output = output_Neuron(self.h2o(spike_layer2), mem_output, self.tau_m_o)
# mem_output[:,i,:] = mem_output_tmp
if i > 0:#40
output= output + mem_output
hidden_spike_.append(spike_layer1.data.cpu().numpy())
hidden_spike2_.append(spike_layer2.data.cpu().numpy())
# h2o_mem_.append(output.data.cpu().numpy())
h2o_mem_.append(spike_layer2.data.cpu().numpy())
output = F.log_softmax(output/seq_num, dim=1)
hidden_spike_ = np.array(hidden_spike_)
hidden_spike2_ = np.array(hidden_spike2_)
fr = (np.mean(hidden_spike_)+np.mean(hidden_spike2_))/2.
return output, hidden_spike_, fr, h2o_mem_
'''
STEP 4: INSTANTIATE MODEL CLASS
'''
input_dim = 700
hidden_dim = [400,400] # 128
output_dim = 35
seq_dim = 250 # Number of steps to unroll
num_encode = 700
total_steps = seq_dim
model = RNN_custom(input_dim, hidden_dim, output_dim)
model = torch.load('./model/model_74.18800902757334-v3-[400, 400]-2layer_MG.pth')
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print("device:", device)
model.to(device)
criterion = nn.CrossEntropyLoss()
learning_rate = 1e-2 # 1e-2
# optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
base_params = [model.i_2_h1.weight, model.i_2_h1.bias,
model.h1_2_h1.weight, model.h1_2_h1.bias,
model.h1_2_h2.weight, model.h1_2_h2.bias,
model.h2_2_h2.weight, model.h2_2_h2.bias,
model.h2o.weight, model.h2o.bias]
optimizer1 = torch.optim.Adam([
{'params': base_params},
{'params': model.tau_adp_h1, 'lr': learning_rate * 10},
{'params': model.tau_adp_h2, 'lr': learning_rate * 10},
{'params': model.tau_adp_o, 'lr': learning_rate * 10},
{'params': model.tau_m_h1, 'lr': learning_rate * 10},
{'params': model.tau_m_h2, 'lr': learning_rate * 10},
{'params': model.tau_m_o, 'lr': learning_rate * 10}],
lr=learning_rate)
optimizer = torch.optim.Adam([
{'params': base_params},
{'params': model.tau_adp_h1, 'lr': learning_rate * 5},
{'params': model.tau_adp_h2, 'lr': learning_rate *5},
{'params': model.tau_m_h1, 'lr': learning_rate * 2.5},
{'params': model.tau_m_h2, 'lr': learning_rate * 2.5},
{'params': model.tau_m_o, 'lr': learning_rate * 2.5}],
lr=learning_rate)
scheduler = StepLR(optimizer, step_size=5, gamma=.5)
def test(model, dataloader=test_loader):
correct = 0
total = 0
fr_list = []
# Iterate through test dataset
for images, labels in dataloader:
images = images.view(-1, seq_dim, input_dim).to(device)
labels = labels.view(-1,)
outputs, hidden_spike_,fr,output_mem = model(images)
# fr = np.mean(hidden_spike_)
fr_list.append(fr)
_, predicted = torch.max(outputs, 1)
total += labels.size(0)
if torch.cuda.is_available():
correct += (predicted.cpu() == labels.long().cpu()).sum()
else:
correct += (predicted == labels).sum()
accuracy = 100. * correct.numpy() / total
print('avg firing rate: ',np.mean(fr_list))
return accuracy
def predict(model):
# Iterate through test dataset
result = np.zeros(1)
for images, labels in test_loader:
images = images.view(-1, seq_dim, input_dim).to(device)
outputs, _,_,_ = model(images)
# _, Predicted = torch.max(outputs.data, 1)
# result.append(Predicted.data.cpu().numpy())
predicted_vec = outputs.data.cpu().numpy()
Predicted = predicted_vec.argmax(axis=1)
result = np.append(result,Predicted)
return np.array(result[1:]).flatten()
accuracy = test(model,test_loader)
print(' Accuracy: ', accuracy)
i = 1
for images, labels in test_loader:
if i ==1 :
images = images.view(-1, seq_dim, input_dim).to(device)
outputs, _,_,output_mem = model(images)
output_mem = np.array(output_mem).reshape(batch_size,250, 400)
for i in range(20):
plt.plot(output_mem[1,:, i], label=str(i))
# plt.legend()
plt.show()
else:
break
i = 1
spike_count = {'total':[],'fr':[],'per step':[]}
for images, labels in test_loader:
if i>0 :
i+=1
images = images.view(-1, seq_dim, input_dim).to(device)
outputs, spike1,_,spike2 = model(images)
b = images.shape[0]
spike1 = np.array(spike1).reshape(b,250, 400)
spike2 = np.array(spike2).reshape(b,250, 400)
spikes = np.zeros((b,250,800))
spikes[:,:,:400] = spike1
spikes[:,:,400:] = spike2
sum_spike= np.sum(spikes,axis=(1,2))
spike_count['total'].append([np.mean(sum_spike),np.max(sum_spike),np.min(sum_spike)])
spike_count['fr'].append(np.mean(spikes))
spike_count['per step'].append([np.mean(np.sum(spikes,axis=(2))),np.max(np.sum(spikes,axis=(2))),np.min(np.sum(spikes,axis=(2)))])
# else:
# break
spike_total_npy = np.array(spike_count['total'])
np.mean(spike_total_npy[0,:]),np.max(spike_total_npy[1,:]),np.min(spike_total_npy[2,:])
spike_total_npy = np.array(spike_count['fr'])
np.mean(spike_total_npy)
spike_total_npy = np.array(spike_count['per step'])
np.mean(spike_total_npy[0,:]),np.max(spike_total_npy[1,:]),np.min(spike_total_npy[2,:])
```
| github_jupyter |
_Lambda School Data Science, Unit 2_
# Regression & Classification Sprint Challenge
To demonstrate mastery on your Sprint Challenge, do all the required, numbered instructions in this notebook.
To earn a score of "3", also do all the stretch goals.
You are permitted and encouraged to do as much data exploration as you want.
### Part 1, Classification
- 1.1. Begin with baselines for classification
- 1.2. Do train/test split. Arrange data into X features matrix and y target vector
- 1.3. Use scikit-learn to fit a logistic regression model
- 1.4. Report classification metric: accuracy
### Part 2, Regression
- 2.1. Begin with baselines for regression
- 2.2. Do train/validate/test split.
- 2.3. Make visualizations to explore relationships between features and target
- 2.4. Arrange data into X features matrix and y target vector
- 2.5. Do one-hot encoding
- 2.6. Use scikit-learn to fit a linear regression model
- 2.7. Report regression metrics: MAE, $R^2$
- 2.8. Get coefficients of a linear model
### Stretch Goals, Regression
- Try at least 3 feature combinations. You may select features manually, or automatically
- Report train & validation RMSE, MAE, $R^2$ for each feature combination you try
- Report test RMSE, MAE, $R^2$ for your final model
```
'''
# If you're in Colab...
import os, sys
in_colab = 'google.colab' in sys.modules
if in_colab:
!pip install --upgrade category_encoders pandas-profiling plotly
'''
```
# Part 1, Classification: Predict Blood Donations 🚑
Our dataset is from a mobile blood donation vehicle in Taiwan. The Blood Transfusion Service Center drives to different universities and collects blood as part of a blood drive.
The goal is to predict whether the donor made a donation in March 2007, using information about each donor's history.
Good data-driven systems for tracking and predicting donations and supply needs can improve the entire supply chain, making sure that more patients get the blood transfusions they need.
```
import pandas as pd
donors = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/blood-transfusion/transfusion.data')
assert donors.shape == (748,5)
donors = donors.rename(columns={
'Recency (months)': 'months_since_last_donation',
'Frequency (times)': 'number_of_donations',
'Monetary (c.c. blood)': 'total_volume_donated',
'Time (months)': 'months_since_first_donation',
'whether he/she donated blood in March 2007': 'made_donation_in_march_2007'
})
```
## 1.1. Begin with baselines
What accuracy score would you get here with a "majority class baseline"?
(You don't need to split the data into train and test sets yet. You can answer this question either with a scikit-learn function or with a pandas function.)
```
donors.head()
# Can always predict the mean.
df = donors.copy()
df.made_donation_in_march_2007.mean()
```
**Can assume that any random donor has a ~24% chance of voting. This also means the majority class is 0(did not donate)**
## 1.2. Do train/test split. Arrange data into X features matrix and y target vector
You may do these steps in either order.
Split randomly. Use scikit-learn's train/test split function. Include 75% of the data in the train set, and hold out 25% for the test set.
```
from sklearn.model_selection import train_test_split
X = df.drop(columns = 'made_donation_in_march_2007')
y = df['made_donation_in_march_2007']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42)
X_train.shape, X_test.shape, y_train.shape, y_test.shape
import numpy as np
# Build baseline test array
y_pred_naive = np.array([0]*len(y_train))
len(y_pred_naive)
```
## 1.3. Use scikit-learn to fit a logistic regression model
You may use any number of features
```
# Check dtypes for numeric/non-numeric
from IPython.display import display
display(df.dtypes, df.isnull().sum())
```
**All features should be ready to use immediately**
```
from sklearn.linear_model import LogisticRegression
# fit model,
model = LogisticRegression(max_iter=50000, n_jobs=-1, solver='saga')
model.fit(X_train, y_train)
```
## 1.4. Report classification metric: accuracy
What is your model's accuracy on the test set?
Don't worry if your model doesn't beat the majority class baseline. That's okay!
_"The combination of some data and an aching desire for an answer does not ensure that a reasonable answer can be extracted from a given body of data."_ —[John Tukey](https://en.wikiquote.org/wiki/John_Tukey)
```
# Accuracy of majority class baseline (should be the mean above)
from sklearn.metrics import accuracy_score
accuracy_score(y_train, y_pred_naive)
# Accuracy of model
y_pred = model.predict(X_test)
accuracy_score(y_test, y_pred)
```
**Significantly beat by majority class prediction!**
# Part 2, Regression: Predict home prices in Ames, Iowa 🏠
You'll use historical housing data. There's a data dictionary at the bottom of the notebook.
Run this code cell to load the dataset:
```
import pandas as pd
URL = 'https://drive.google.com/uc?export=download&id=1522WlEW6HFss36roD_Cd9nybqSuiVcCK'
homes = pd.read_csv(URL)
assert homes.shape == (2904, 47)
```
## 2.1. Begin with baselines
What is the Mean Absolute Error and R^2 score for a mean baseline?
```
df2 = homes.copy()
from sklearn.metrics import mean_squared_error, r2_score
y_pred_naive = np.array(
[df2.SalePrice.mean()]*df2.shape[0]
)
target = df2.SalePrice.to_numpy()
display(mean_squared_error(target, y_pred_naive), r2_score(target, y_pred_naive))
```
## 2.2. Do train/test split
Train on houses sold in the years 2006 - 2008. (1,920 rows)
Validate on house sold in 2009. (644 rows)
Test on houses sold in 2010. (340 rows)
```
# Split into train, test, and validation sets
train = df2[(df2.Yr_Sold>2005) & (df2.Yr_Sold<2009)]
test = df2[df2.Yr_Sold==2010]
val = df2[df2.Yr_Sold==2009]
# Break into feature and target sets
def return_test_train(df):
return df.drop(columns='SalePrice'), df.SalePrice
X_train, y_train = return_test_train(train)
X_test, y_test = return_test_train(test)
X_val, y_val = return_test_train(val)
display(X_train.shape, X_test.shape, X_val.shape)
```
## 2.3. Make visualizations to explore relationships between features and target
You can visualize many features, or just a few.
You can try Seaborn's ["Categorical estimate" plots](https://seaborn.pydata.org/tutorial/categorical.html) and/or [linear model plots](https://seaborn.pydata.org/tutorial/regression.html).
You do _not_ need to use Seaborn, but it's nice because it includes confidence intervals to visualize uncertainty.
Plotly and Pandas are also great for exploratory plots.
```
# Print the column names for easier use
print(train.columns)
import matplotlib.pyplot as plt
import seaborn as sns
fig, ax = plt.subplots()
ax = sns.barplot(x='Overall_Cond', y='SalePrice', data=df2)
```
**Overall condition could be a great indicator, but ordinality is an issue here for linear regression. Need to re-encode either one-hot or manually**
```
ax = sns.barplot(x='Sale_Condition', y='SalePrice', data=df2)
ax = sns.regplot(x='Full_Bath', y='SalePrice', data=df2)
ax = sns.barplot(x='Central_Air', y='SalePrice', data=df2)
```
## 2.4. Arrange data into X features matrix and y target vector
Select at least one numeric feature and at least one categorical feature.
Otherwise, you many choose whichever features and however many you want.
```
#See Above
```
## 2.5. Do one-hot encoding
Encode your categorical feature(s).
```
from category_encoders.one_hot import OneHotEncoder
cat_vars = ['Central_Air', 'Sale_Condition', 'Overall_Cond']
num_vars = ['Full_Bath']
target = 'SalePrice'
def encode_cat(master, subset, cat_vars):
# Initialize encoder
encoder = OneHotEncoder(cols=cat_vars)
encoder.fit(master)
return encoder.transform(subset)
# Reduce feature sets to features in model
features = cat_vars + num_vars
X_train = X_train[features]
X_test = X_test[features]
X_val = X_val[features]
# Encode categorical variables and return full featureset
X_train = encode_cat(df2[features], X_train, cat_vars)
X_test = encode_cat(df2[features], X_test, cat_vars)
X_val = encode_cat(df2[features], X_val, cat_vars)
```
## 2.6. Use scikit-learn to fit a linear regression model
Fit your model.
```
from sklearn import linear_model
regr = linear_model.LinearRegression()
regr.fit(X_train, y_train)
```
## 2.7. Report regression metrics: Mean Absolute Error, $R^2$
What is your model's Mean Absolute Error and $R^2$ score on the validation set?
```
from sklearn.metrics import mean_squared_error, r2_score, mean_absolute_error
y_pred = regr.predict(X_val)
eval_info = {
'coefficients': regr.coef_,
'mae': mean_absolute_error(y_val, y_pred),
'r2': r2_score(y_val, y_pred)
}
print(eval_info)
# That is pretty bad!
```
## 2.8. Get coefficients of a linear model
Print or plot the coefficients for the features in your model.
```
# See agove
```
## Stretch Goals, Regression
- Try at least 3 feature combinations. You may select features manually, or automatically.
- Report train & validation RMSE, MAE, $R^2$ for each feature combination you try
- Report test RMSE, MAE, $R^2$ for your final model
```
# Kinda hand-made model pipeline
import warnings
from sklearn import linear_model
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import f_regression
from sklearn.preprocessing import RobustScaler
from sklearn.metrics import mean_squared_error, r2_score
from math import sqrt
warnings.filterwarnings(action='ignore', category=RuntimeWarning)
def linear_pipe(X_train, y_train, X_test, y_test, num_features):
'''
Select a number of features for SelectKBest to return.
Scale features with RobustScalar
Passes scaled features to linear model and fitted model
Inputs:
X_train, y_train: Pandas DataFrame of feature training set and corresponding target set
X_test, y_test: Pandas DataFrame of feature test set and corresponding target set
num_features: integer number of features to pass to model
Return fitted model
'''
def select_features(**kw):
# Create Selector and fit to training data
selector = SelectKBest(f_regression, k=num_features)
selector.fit(X_train, y_train)
# Get columns to keep
cols = selector.get_support(indices=True)
# Create subsets of training and test data, return values
return X_train.iloc[:, lambda df: cols], X_test.iloc[:, lambda df: cols]
def scale_features(X_train_selected, X_test_selected):
# Create scalar
scaler = RobustScaler()
# Scale & Transform Features, return value
scaler.fit(X_train_selected.to_numpy())
return scaler.transform(X_train_selected.to_numpy()), scaler.transform(X_test_selected.to_numpy())
def generate_model(features, target):
regr = linear_model.LinearRegression()
return regr.fit(features, target)
def evaluate_model(model, X_test_selected):
y_pred = model.predict(X_test_selected)
eval_info = {
'coefficients': model.coef_,
'rmse': sqrt(mean_squared_error(y_test, y_pred)),
'r2': r2_score(y_test, y_pred)
}
return eval_info
X_train_selected, X_test_selected = select_features()
X_train_scaled, X_test_scaled = scale_features(X_train_selected, X_test_selected)
model = generate_model(features=X_train_scaled, target=y_train)
eval_info = evaluate_model(model, X_test_scaled)
return model, eval_info
model_evals = []
for i in range(1,len(X_train.columns)):
_, info = linear_pipe(X_train, y_train, X_val, y_val, i)
model_evals.append([info['r2'], info['rmse']])
model_evals = pd.DataFrame(model_evals)
# Plot r squared metric
model_evals[0].plot()
# Plot RMSE metric
model_evals[1].plot()
```
**I think it's safe to say that +/- $63,000 is not particularly good! But with the pipeline setup, it wouldn't be too difficult to further explore features**
```
# compare to mean prediction
y_pred_naive = np.array([y_val.mean()]*len(y_val))
sqrt(
mean_squared_error(y_pred_naive, y_val)
)
```
**At least it's a $17,000 improvement over baseline. That's not insignificant**
## Data Dictionary
Here's a description of the data fields:
```
1st_Flr_SF: First Floor square feet
Bedroom_AbvGr: Bedrooms above grade (does NOT include basement bedrooms)
Bldg_Type: Type of dwelling
1Fam Single-family Detached
2FmCon Two-family Conversion; originally built as one-family dwelling
Duplx Duplex
TwnhsE Townhouse End Unit
TwnhsI Townhouse Inside Unit
Bsmt_Half_Bath: Basement half bathrooms
Bsmt_Full_Bath: Basement full bathrooms
Central_Air: Central air conditioning
N No
Y Yes
Condition_1: Proximity to various conditions
Artery Adjacent to arterial street
Feedr Adjacent to feeder street
Norm Normal
RRNn Within 200' of North-South Railroad
RRAn Adjacent to North-South Railroad
PosN Near positive off-site feature--park, greenbelt, etc.
PosA Adjacent to postive off-site feature
RRNe Within 200' of East-West Railroad
RRAe Adjacent to East-West Railroad
Condition_2: Proximity to various conditions (if more than one is present)
Artery Adjacent to arterial street
Feedr Adjacent to feeder street
Norm Normal
RRNn Within 200' of North-South Railroad
RRAn Adjacent to North-South Railroad
PosN Near positive off-site feature--park, greenbelt, etc.
PosA Adjacent to postive off-site feature
RRNe Within 200' of East-West Railroad
RRAe Adjacent to East-West Railroad
Electrical: Electrical system
SBrkr Standard Circuit Breakers & Romex
FuseA Fuse Box over 60 AMP and all Romex wiring (Average)
FuseF 60 AMP Fuse Box and mostly Romex wiring (Fair)
FuseP 60 AMP Fuse Box and mostly knob & tube wiring (poor)
Mix Mixed
Exter_Cond: Evaluates the present condition of the material on the exterior
Ex Excellent
Gd Good
TA Average/Typical
Fa Fair
Po Poor
Exter_Qual: Evaluates the quality of the material on the exterior
Ex Excellent
Gd Good
TA Average/Typical
Fa Fair
Po Poor
Exterior_1st: Exterior covering on house
AsbShng Asbestos Shingles
AsphShn Asphalt Shingles
BrkComm Brick Common
BrkFace Brick Face
CBlock Cinder Block
CemntBd Cement Board
HdBoard Hard Board
ImStucc Imitation Stucco
MetalSd Metal Siding
Other Other
Plywood Plywood
PreCast PreCast
Stone Stone
Stucco Stucco
VinylSd Vinyl Siding
Wd Sdng Wood Siding
WdShing Wood Shingles
Exterior_2nd: Exterior covering on house (if more than one material)
AsbShng Asbestos Shingles
AsphShn Asphalt Shingles
BrkComm Brick Common
BrkFace Brick Face
CBlock Cinder Block
CemntBd Cement Board
HdBoard Hard Board
ImStucc Imitation Stucco
MetalSd Metal Siding
Other Other
Plywood Plywood
PreCast PreCast
Stone Stone
Stucco Stucco
VinylSd Vinyl Siding
Wd Sdng Wood Siding
WdShing Wood Shingles
Foundation: Type of foundation
BrkTil Brick & Tile
CBlock Cinder Block
PConc Poured Contrete
Slab Slab
Stone Stone
Wood Wood
Full_Bath: Full bathrooms above grade
Functional: Home functionality (Assume typical unless deductions are warranted)
Typ Typical Functionality
Min1 Minor Deductions 1
Min2 Minor Deductions 2
Mod Moderate Deductions
Maj1 Major Deductions 1
Maj2 Major Deductions 2
Sev Severely Damaged
Sal Salvage only
Gr_Liv_Area: Above grade (ground) living area square feet
Half_Bath: Half baths above grade
Heating: Type of heating
Floor Floor Furnace
GasA Gas forced warm air furnace
GasW Gas hot water or steam heat
Grav Gravity furnace
OthW Hot water or steam heat other than gas
Wall Wall furnace
Heating_QC: Heating quality and condition
Ex Excellent
Gd Good
TA Average/Typical
Fa Fair
Po Poor
House_Style: Style of dwelling
1Story One story
1.5Fin One and one-half story: 2nd level finished
1.5Unf One and one-half story: 2nd level unfinished
2Story Two story
2.5Fin Two and one-half story: 2nd level finished
2.5Unf Two and one-half story: 2nd level unfinished
SFoyer Split Foyer
SLvl Split Level
Kitchen_AbvGr: Kitchens above grade
Kitchen_Qual: Kitchen quality
Ex Excellent
Gd Good
TA Typical/Average
Fa Fair
Po Poor
LandContour: Flatness of the property
Lvl Near Flat/Level
Bnk Banked - Quick and significant rise from street grade to building
HLS Hillside - Significant slope from side to side
Low Depression
Land_Slope: Slope of property
Gtl Gentle slope
Mod Moderate Slope
Sev Severe Slope
Lot_Area: Lot size in square feet
Lot_Config: Lot configuration
Inside Inside lot
Corner Corner lot
CulDSac Cul-de-sac
FR2 Frontage on 2 sides of property
FR3 Frontage on 3 sides of property
Lot_Shape: General shape of property
Reg Regular
IR1 Slightly irregular
IR2 Moderately Irregular
IR3 Irregular
MS_SubClass: Identifies the type of dwelling involved in the sale.
20 1-STORY 1946 & NEWER ALL STYLES
30 1-STORY 1945 & OLDER
40 1-STORY W/FINISHED ATTIC ALL AGES
45 1-1/2 STORY - UNFINISHED ALL AGES
50 1-1/2 STORY FINISHED ALL AGES
60 2-STORY 1946 & NEWER
70 2-STORY 1945 & OLDER
75 2-1/2 STORY ALL AGES
80 SPLIT OR MULTI-LEVEL
85 SPLIT FOYER
90 DUPLEX - ALL STYLES AND AGES
120 1-STORY PUD (Planned Unit Development) - 1946 & NEWER
150 1-1/2 STORY PUD - ALL AGES
160 2-STORY PUD - 1946 & NEWER
180 PUD - MULTILEVEL - INCL SPLIT LEV/FOYER
190 2 FAMILY CONVERSION - ALL STYLES AND AGES
MS_Zoning: Identifies the general zoning classification of the sale.
A Agriculture
C Commercial
FV Floating Village Residential
I Industrial
RH Residential High Density
RL Residential Low Density
RP Residential Low Density Park
RM Residential Medium Density
Mas_Vnr_Type: Masonry veneer type
BrkCmn Brick Common
BrkFace Brick Face
CBlock Cinder Block
None None
Stone Stone
Mo_Sold: Month Sold (MM)
Neighborhood: Physical locations within Ames city limits
Blmngtn Bloomington Heights
Blueste Bluestem
BrDale Briardale
BrkSide Brookside
ClearCr Clear Creek
CollgCr College Creek
Crawfor Crawford
Edwards Edwards
Gilbert Gilbert
IDOTRR Iowa DOT and Rail Road
MeadowV Meadow Village
Mitchel Mitchell
Names North Ames
NoRidge Northridge
NPkVill Northpark Villa
NridgHt Northridge Heights
NWAmes Northwest Ames
OldTown Old Town
SWISU South & West of Iowa State University
Sawyer Sawyer
SawyerW Sawyer West
Somerst Somerset
StoneBr Stone Brook
Timber Timberland
Veenker Veenker
Overall_Cond: Rates the overall condition of the house
10 Very Excellent
9 Excellent
8 Very Good
7 Good
6 Above Average
5 Average
4 Below Average
3 Fair
2 Poor
1 Very Poor
Overall_Qual: Rates the overall material and finish of the house
10 Very Excellent
9 Excellent
8 Very Good
7 Good
6 Above Average
5 Average
4 Below Average
3 Fair
2 Poor
1 Very Poor
Paved_Drive: Paved driveway
Y Paved
P Partial Pavement
N Dirt/Gravel
Roof_Matl: Roof material
ClyTile Clay or Tile
CompShg Standard (Composite) Shingle
Membran Membrane
Metal Metal
Roll Roll
Tar&Grv Gravel & Tar
WdShake Wood Shakes
WdShngl Wood Shingles
Roof_Style: Type of roof
Flat Flat
Gable Gable
Gambrel Gabrel (Barn)
Hip Hip
Mansard Mansard
Shed Shed
SalePrice: the sales price for each house
Sale_Condition: Condition of sale
Normal Normal Sale
Abnorml Abnormal Sale - trade, foreclosure, short sale
AdjLand Adjoining Land Purchase
Alloca Allocation - two linked properties with separate deeds, typically condo with a garage unit
Family Sale between family members
Partial Home was not completed when last assessed (associated with New Homes)
Sale_Type: Type of sale
WD Warranty Deed - Conventional
CWD Warranty Deed - Cash
VWD Warranty Deed - VA Loan
New Home just constructed and sold
COD Court Officer Deed/Estate
Con Contract 15% Down payment regular terms
ConLw Contract Low Down payment and low interest
ConLI Contract Low Interest
ConLD Contract Low Down
Oth Other
Street: Type of road access to property
Grvl Gravel
Pave Paved
TotRms_AbvGrd: Total rooms above grade (does not include bathrooms)
Utilities: Type of utilities available
AllPub All public Utilities (E,G,W,& S)
NoSewr Electricity, Gas, and Water (Septic Tank)
NoSeWa Electricity and Gas Only
ELO Electricity only
Year_Built: Original construction date
Year_Remod/Add: Remodel date (same as construction date if no remodeling or additions)
Yr_Sold: Year Sold (YYYY)
```
| github_jupyter |
# Build Up My Own Recommend Playlist from Scratch
Collaborative Filtering is usually the first type of method for recommender system. It has two approachs, the first one is user-user based model, which use the similiarity between user to recommend new items, another one is item-item based model. Instead of using the similarity between users, it uses that between items to recommend new items.
Here, we are going to go beyond collaborative filtering and introduce latent factor model in the application of recommender system. In this article, we are going to use [Million Song Dataset](https://labrosa.ee.columbia.edu/millionsong/). It contains users and song data. The main motivation behind is that when we using music streaming service like Spotify, kkbox, and youtube, the recommended songs often catch my eye. Take Spotify for example, there is a feature called *Discover Weekly*, which automatically generate a recommended playlist weekly. Very often, I enjoyed listening to the recommended songs. Therefore, I think it will be a great idea if I can build up a recommend playlist or songs using different methods, and to see what the result will be.
Here are the steps that I take for this experiment:
* Take [Million Song Dataset](https://labrosa.ee.columbia.edu/millionsong/)
* Use user-user based collaborative filtering to build up a recommended playlist
* Use item-item based collaborative filtering to build up a recommended playlist
* Use Latent Factor Model to build up a recommended playlist
* Measure the performance using Root Mean Square Error(RMSE)
* Compare the result of different approachs
For collaborative filtering, we follow the same step as the [previous notebook for recommending movies](https://github.com/johnnychiuchiu/Machine-Learning/blob/master/RecommenderSystem/collaborative_filtering.ipynb).
Firstly, in order to calculate the similarity, we need to get a utility matrix using the song dataframe. For illustration purpose, I also manually append three rows into the utility matrix. Each row represent a person with some specific music taste.
We use two method to compare the measure the result of different approachs. The first approach is calculate the Mean Square Error. Since both collaborative filtering and latent factor model need all the dataset to calcualte the predicted result. The way we generate train and test different from the way we usually use, that is randomly select some row to be test.
In the song data, we randomly take 3 listen_count of each user out and place it in the test dataset. Then we use only train dataset to predict the recommended playlist. After have the predicted score for all the songs, we then compare the nonzero values in test data set with the corresponding value in the train dataset and calcualte the MSE of it. Also, I have make sure each user has at least listened to 5 different songs in the song data.
---
## Implementing Collaborative Filtering to build up Recommeded Playlist
```
%matplotlib inline
import pandas as pd
from sklearn.cross_validation import train_test_split
import numpy as np
import os
from sklearn.metrics import mean_squared_error
def compute_mse(y_true, y_pred):
"""ignore zero terms prior to comparing the mse"""
mask = np.nonzero(y_true)
mse = mean_squared_error(y_true[mask], y_pred[mask])
return mse
def create_train_test(ratings):
"""
split into training and test sets,
remove 3 ratings from each user
and assign them to the test set
"""
test = np.zeros(ratings.shape)
train = ratings.copy()
for user in range(ratings.shape[0]):
test_index = np.random.choice(
np.flatnonzero(ratings[user]), size=3, replace=False)
train[user, test_index] = 0.0
test[user, test_index] = ratings[user, test_index]
# assert that training and testing set are truly disjoint
assert np.all(train * test == 0)
return (train, test)
class collaborativeFiltering():
def __init__(self):
pass
def readSongData(self, top):
"""
Read song data from targeted url
"""
if 'song.pkl' in os.listdir('_data/'):
song_df = pd.read_pickle('_data/song.pkl')
else:
# Read userid-songid-listen_count triplets
# This step might take time to download data from external sources
triplets_file = 'https://static.turi.com/datasets/millionsong/10000.txt'
songs_metadata_file = 'https://static.turi.com/datasets/millionsong/song_data.csv'
song_df_1 = pd.read_table(triplets_file, header=None)
song_df_1.columns = ['user_id', 'song_id', 'listen_count']
# Read song metadata
song_df_2 = pd.read_csv(songs_metadata_file)
# Merge the two dataframes above to create input dataframe for recommender systems
song_df = pd.merge(song_df_1, song_df_2.drop_duplicates(['song_id']), on="song_id", how="left")
# Merge song title and artist_name columns to make a merged column
song_df['song'] = song_df['title'].map(str) + " - " + song_df['artist_name']
n_users = song_df.user_id.unique().shape[0]
n_items = song_df.song_id.unique().shape[0]
print(str(n_users) + ' users')
print(str(n_items) + ' items')
song_df.to_pickle('_data/song.pkl')
# keep top_n rows of the data
song_df = song_df.head(top)
song_df = self.drop_freq_low(song_df)
return(song_df)
def drop_freq_low(self, song_df):
freq_df = song_df.groupby(['user_id']).agg({'song_id': 'count'}).reset_index(level=['user_id'])
below_userid = freq_df[freq_df.song_id <= 5]['user_id']
new_song_df = song_df[~song_df.user_id.isin(below_userid)]
return(new_song_df)
def utilityMatrix(self, song_df):
"""
Transform dataframe into utility matrix, return both dataframe and matrix format
:param song_df: a dataframe that contains user_id, song_id, and listen_count
:return: dataframe, matrix
"""
song_reshape = song_df.pivot(index='user_id', columns='song_id', values='listen_count')
song_reshape = song_reshape.fillna(0)
ratings = song_reshape.as_matrix()
return(song_reshape, ratings)
def fast_similarity(self, ratings, kind='user', epsilon=1e-9):
"""
Calculate the similarity of the rating matrix
:param ratings: utility matrix
:param kind: user-user sim or item-item sim
:param epsilon: small number for handling dived-by-zero errors
:return: correlation matrix
"""
if kind == 'user':
sim = ratings.dot(ratings.T) + epsilon
elif kind == 'item':
sim = ratings.T.dot(ratings) + epsilon
norms = np.array([np.sqrt(np.diagonal(sim))])
return (sim / norms / norms.T)
def predict_fast_simple(self, ratings, kind='user'):
"""
Calculate the predicted score of every song for every user.
:param ratings: utility matrix
:param kind: user-user sim or item-item sim
:return: matrix contains the predicted scores
"""
similarity = self.fast_similarity(ratings, kind)
if kind == 'user':
return similarity.dot(ratings) / np.array([np.abs(similarity).sum(axis=1)]).T
elif kind == 'item':
return ratings.dot(similarity) / np.array([np.abs(similarity).sum(axis=1)])
def get_overall_recommend(self, ratings, song_reshape, user_prediction, top_n=10):
"""
get the top_n predicted result of every user. Notice that the recommended item should be the song that the user
haven't listened before.
:param ratings: utility matrix
:param song_reshape: utility matrix in dataframe format
:param user_prediction: matrix with predicted score
:param top_n: the number of recommended song
:return: a dict contains recommended songs for every user_id
"""
result = dict({})
for i, row in enumerate(ratings):
user_id = song_reshape.index[i]
result[user_id] = {}
zero_item_list = np.where(row == 0)[0]
prob_list = user_prediction[i][np.where(row == 0)[0]]
song_id_list = np.array(song_reshape.columns)[zero_item_list]
result[user_id]['recommend'] = sorted(zip(song_id_list, prob_list), key=lambda item: item[1], reverse=True)[
0:top_n]
return (result)
def get_user_recommend(self, user_id, overall_recommend, song_df):
"""
Get the recommended songs for a particular user using the song information from the song_df
:param user_id:
:param overall_recommend:
:return:
"""
user_score = pd.DataFrame(overall_recommend[user_id]['recommend']).rename(columns={0: 'song_id', 1: 'score'})
user_recommend = pd.merge(user_score,
song_df[['song_id', 'title', 'release', 'artist_name', 'song']].drop_duplicates(),
on='song_id', how='left')
return (user_recommend)
def createNewObs(self, artistName, song_reshape, index_name):
"""
Append a new row with userId 0 that is interested in some specific artists
:param artistName: a list of artist names
:return: dataframe, matrix
"""
interest = []
for i in song_reshape.columns:
if i in song_df[song_df.artist_name.isin(artistName)]['song_id'].unique():
interest.append(10)
else:
interest.append(0)
print(pd.Series(interest).value_counts())
newobs = pd.DataFrame([interest],
columns=song_reshape.columns)
newobs.index = [index_name]
new_song_reshape = pd.concat([song_reshape, newobs])
new_ratings = new_song_reshape.as_matrix()
return (new_song_reshape, new_ratings)
```
## Take Million Song Dataset
We only keep the first 50000 rows for this notebook. Otherwise it will take too long to execute it. As following, we can see that there are around **17k** users and **93k** different songs out of the first 50k rows.
```
cf = collaborativeFiltering()
song_df = cf.readSongData(100000)
song_df.head()
artist_df= song_df.groupby(['artist_name']).agg({'song_id':'count'}).reset_index(level=['artist_name']).sort_values(by='song_id',ascending=False).head(100)
n_users = song_df.user_id.unique().shape[0]
n_items = song_df.song_id.unique().shape[0]
print(str(n_users) + ' users')
print(str(n_items) + ' songs')
```
• **Get the utility matrix**
```
song_reshape, ratings = cf.utilityMatrix(song_df)
```
• **Append new rows to simulate a users who love different kinds of musicians**
```
song_reshape, ratings = cf.createNewObs(['Beyoncé', 'Katy Perry', 'Alicia Keys'], song_reshape, 'GirlFan')
song_reshape, ratings = cf.createNewObs(['Metallica', 'Guns N\' Roses', 'Linkin Park', 'Red Hot Chili Peppers'],
song_reshape, 'HeavyFan')
song_reshape, ratings = cf.createNewObs(['Daft Punk','John Mayer','Hot Chip','Coldplay'],
song_reshape, 'Johnny')
```
• **Create train test dataset**
```
train, test = create_train_test(ratings)
song_reshape.shape
```
## Calculate user-user collaborative filtering
```
user_prediction = cf.predict_fast_simple(train, kind='user')
user_overall_recommend = cf.get_overall_recommend(train, song_reshape, user_prediction, top_n=10)
user_recommend_girl = cf.get_user_recommend('GirlFan', user_overall_recommend, song_df)
user_recommend_heavy = cf.get_user_recommend('HeavyFan', user_overall_recommend, song_df)
user_recommend_johnny = cf.get_user_recommend('Johnny', user_overall_recommend, song_df)
```
## Calculate item-item collaborative filtering
```
item_prediction = cf.predict_fast_simple(train, kind='item')
item_overall_recommend = cf.get_overall_recommend(train, song_reshape, item_prediction, top_n=10)
item_recommend_girl = cf.get_user_recommend('GirlFan', item_overall_recommend, song_df)
item_recommend_heavy = cf.get_user_recommend('HeavyFan', item_overall_recommend, song_df)
item_recommend_johnny = cf.get_user_recommend('Johnny', item_overall_recommend, song_df)
```
---
The main idea behind Latent Factor Model is that we can transform our utility matrix into the multiple of two lower rank matrix. For example, if we have 5 users and 10 songs, then our utility matrix is 5 * 10. We can transform the matrix in to two matrixs, say 5 x 3 (say Q) and 3 x 10 (say P). Each user can be represented by a vector in 3 dimension, and each song can als obe represented by a vector in 3 dimension. The meaning of each dimension for Q can be, for example, do the user like jazz related music; each dimension for P can be, for example, is it a jazz song. The picture copied from google search result visualize it more clearly:

In order to get all the values in the Q and P, we need some optimization method to help us. The optimization method suggested by the winner of Netflix is called **Alternating Least Squares with Weighted Regularization (ALS-WR)**.
Our cost function is as follows:
$$ \begin{align} L &= \sum\limits_{u,i \in S}( r_{ui} - \textbf{x}_{u} \textbf{y}_{i}^{T} )^{2} + \lambda \big( \sum\limits_{u} \left\Vert \textbf{x}_{u} \right\Vert^{2} + \sum\limits_{i} \left\Vert \textbf{y}_{i} \right\Vert^{2} \big) \end{align} $$
We will try to minimize the loss function to get our optimal $x_u$ and $y_i$ vectors. The main idea behind ALS-WR method is that we try to get the optimal Q and P matrix by holding one vector to be fixed at a time. We alternate back and forth until the value of Q and P converges. The reason why we don't optimize both vector at the same time is that it is hard to get optimal vectors at the same time. By holding one vector to be fixed and optimize another vector alternately, we can find the optimal Q and P more efficiently.
For a detailed explaination on how to
please check [Ethen's Alternating Least Squares with Weighted Regularization (ALS-WR) from scratch](http://nbviewer.jupyter.org/github/ethen8181/machine-learning/blob/master/recsys/1_ALSWR.ipynb).
## Recommend using Latent Factor Model
```
class ExplicitMF:
"""
This function is directly taken from Ethen's github (http://nbviewer.jupyter.org/github/ethen8181/machine-learning/blob/master/recsys/1_ALSWR.ipynb)
Train a matrix factorization model using Alternating Least Squares
to predict empty entries in a matrix
Parameters
----------
n_iters : int
number of iterations to train the algorithm
n_factors : int
number of latent factors to use in matrix
factorization model, some machine-learning libraries
denote this as rank
reg : float
regularization term for item/user latent factors,
since lambda is a keyword in python we use reg instead
"""
def __init__(self, n_iters, n_factors, reg):
self.reg = reg
self.n_iters = n_iters
self.n_factors = n_factors
def fit(self, train):#, test
"""
pass in training and testing at the same time to record
model convergence, assuming both dataset is in the form
of User x Item matrix with cells as ratings
"""
self.n_user, self.n_item = train.shape
self.user_factors = np.random.random((self.n_user, self.n_factors))
self.item_factors = np.random.random((self.n_item, self.n_factors))
# record the training and testing mse for every iteration
# to show convergence later (usually, not worth it for production)
# self.test_mse_record = []
# self.train_mse_record = []
for _ in range(self.n_iters):
self.user_factors = self._als_step(train, self.user_factors, self.item_factors)
self.item_factors = self._als_step(train.T, self.item_factors, self.user_factors)
predictions = self.predict()
# test_mse = self.compute_mse(test, predictions)
# train_mse = self.compute_mse(train, predictions)
# self.test_mse_record.append(test_mse)
# self.train_mse_record.append(train_mse)
return self
def _als_step(self, ratings, solve_vecs, fixed_vecs):
"""
when updating the user matrix,
the item matrix is the fixed vector and vice versa
"""
A = fixed_vecs.T.dot(fixed_vecs) + np.eye(self.n_factors) * self.reg
b = ratings.dot(fixed_vecs)
A_inv = np.linalg.inv(A)
solve_vecs = b.dot(A_inv)
return solve_vecs
def predict(self):
"""predict ratings for every user and item"""
pred = self.user_factors.dot(self.item_factors.T)
return pred
```
• **Fit using Alternating Least Square Method**
```
als = ExplicitMF(n_iters=200, n_factors=10, reg=0.01)
als.fit(train)
latent_prediction = als.predict()
latent_overall_recommend = cf.get_overall_recommend(train, song_reshape, latent_prediction, top_n=10)
latent_recommend_girl = cf.get_user_recommend('GirlFan', latent_overall_recommend, song_df)
latent_recommend_heavy = cf.get_user_recommend('HeavyFan', latent_overall_recommend, song_df)
latent_recommend_johnny = cf.get_user_recommend('Johnny', latent_overall_recommend, song_df)
```
## Measure the performance using Root Mean Square Error(RMSE)
```
user_mse = compute_mse(test, user_prediction)
item_mse = compute_mse(test, item_prediction)
latent_mse = compute_mse(test, latent_prediction)
print("MSE for user-user approach: "+str(user_mse))
print("MSE for item-item approach: "+str(item_mse))
print("MSE for latent factor model: "+str(latent_mse))
```
We can see that even though latent factor model is somewhat a more advanced model, the MSE not the lowerest for some reason. It is something that I should keep in mind.
## Compare the result of different approachs
### > Recommend Playlist for someone who is a big fan of *Beyoncé*, *Katy Perry* and *Alicia Keys*
• **User-user approach**
```
user_recommend_girl
```
• **Item-item approach**
```
item_recommend_girl
```
• **Latent Factor Model**
```
latent_recommend_girl
```
### > Recommend Playlist for someone who is a big fan of *Metallica*, *Guns N' Roses*, *Linkin Park* and *Red Hot Chili Peppers*
```
user_recommend_heavy
item_recommend_heavy
latent_recommend_heavy
```
### > Recommend Playlist for myself, I like *Daft Punk*, *John Mayer*, *Hot Chip* and *Coldplay*
```
user_recommend_johnny
item_recommend_johnny
latent_recommend_johnny
```
We see that the all the recommended playlist are actually kind of make sense. In the following notebook, I will continue to try some other methods to build up custom recommended playlists.
### Reference
* http://nbviewer.jupyter.org/github/ethen8181/machine-learning/blob/master/recsys/1_ALSWR.ipynb
* https://github.com/dvysardana/RecommenderSystems_PyData_2016/blob/master/Song%20Recommender_Python.ipynb
| github_jupyter |
# NYAAPOR Text Analytics Tutorial
## Loading in the data
First, download the Kaggle zip file (https://www.kaggle.com/snap/amazon-fine-food-reviews). And unpack it in this repository's root folder
```
import pandas as pd
df = pd.read_csv("../amazon-fine-food-reviews/Reviews.csv")
print(len(df))
```
Wow, that's a lot of data. Let's see what's in here.
```
df.head()
```
Let's just use a sample for now, so things run faster
```
sample = df.sample(10000).reset_index()
```
### Examine the data
Run the cell below a few times, let's take a look at our text and see what it looks like. Always take a look at your raw data.
```
sample.sample(10)['Text'].values
```
I don't know about you, but I noticed some junk in our data - HTML and URLs. Let's clear that out first.
```
import re
def clean_text(text):
text = re.sub(r'http[a-zA-Z0-9\&\?\=\?\/\:\.]+\b', ' ', text)
text = re.sub(r'\<[^\<\>]+\>', ' ', text)
return text
df['Text'] = df['Text'].map(clean_text)
```
## TF-IDF Vectorization (Feature Extraction)
Okay, now let's tokenize our text and turn it into numbers
```
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
tfidf_vectorizer = TfidfVectorizer(
max_df=0.9,
min_df=5,
ngram_range=(1, 1),
stop_words='english',
max_features=2500
)
tfidf = tfidf_vectorizer.fit_transform(sample['Text'])
ngrams = tfidf_vectorizer.get_feature_names()
tfidf
```
Because words are really big, by default we work with sparse matrices. We can expand the sparse matrix with `.todense()` and compute sums like a normal dataframe. Let's check out the top 20 words.
```
ngram_df = pd.DataFrame(tfidf.todense(), columns=ngrams)
ngram_df.sum().sort_values(ascending=False)[:20]
```
## Classification
Let's make an outcome variable. How about we try to predict 5-star reviews, and then maybe helpfulness?
```
sample['good_score'] = sample['Score'].map(lambda x: 1 if x == 5 else 0)
sample['was_helpful'] = ((sample['HelpfulnessNumerator'] / sample['HelpfulnessDenominator']).fillna(0.0) > .80).astype(int)
column_to_predict = 'good_score'
from sklearn.model_selection import StratifiedKFold
from sklearn import svm
from sklearn import metrics
results = []
kfolds = StratifiedKFold(n_splits=5)
```
We just created an object that'll split the data into fifths, and then iterate over it five times, holding out one-fifth each time for testing. Let's do that now. Each "fold" contains an index for training rows, and one for testing rows. For each fold, we'll train a basic linear Support Vector Machine, and evaluate its performance.
```
for i, fold in enumerate(kfolds.split(tfidf, sample[column_to_predict])):
train, test = fold
print("Running new fold, {} training cases, {} testing cases".format(len(train), len(test)))
clf = svm.LinearSVC(
max_iter=1000,
penalty='l2',
class_weight='balanced',
loss='squared_hinge'
)
# We picked some decent starting parameters, but encourage you to try out different ones
# http://scikit-learn.org/stable/modules/generated/sklearn.svm.LinearSVC.html
# If you're ambitious - check out the Scikit-Learn documentation and test out different models
# http://scikit-learn.org/stable/supervised_learning.html
training_text = tfidf[train]
training_outcomes = sample[column_to_predict].loc[train]
clf.fit(training_text, training_outcomes) # Train the classifier on the training data
test_text = tfidf[test]
test_outcomes = sample[column_to_predict].loc[test]
predictions = clf.predict(test_text) # Get predictions for the test data
precision, recall, fscore, support = metrics.precision_recall_fscore_support(
test_outcomes, # Compare the predictions against the true outcomes
predictions
)
results.append({
"fold": i,
"outcome": 0,
"precision": precision[0],
"recall": recall[0],
"fscore": fscore[0],
"support": support[0]
})
results.append({
"fold": i,
"outcome": 1,
"precision": precision[1],
"recall": recall[1],
"fscore": fscore[1],
"support": support[1]
})
results = pd.DataFrame(results)
```
How'd we do?
```
print(results.groupby("outcome").mean()[['precision', 'recall']])
print(results.groupby("outcome").std()[['precision', 'recall']])
```
Now we know that our model is pretty stable and reasonably performant, we can fit and transform the full dataset.
```
clf.fit(tfidf, sample[column_to_predict])
print(metrics.classification_report(sample[column_to_predict].loc[test], predictions))
print(metrics.confusion_matrix(sample[column_to_predict].loc[test], predictions))
```
And now we can see what the most predictive features are.
```
import numpy as np
ngram_coefs = sorted(zip(ngrams, clf.coef_[0]), key=lambda x: x[1], reverse=True)
ngram_coefs[:10]
```
What happens if you change the outcome column to "was_helpful" and re-run it again? Can you think of ways to improve this? Add stopwords? Bigrams?
## Topic Modeling
```
from sklearn.decomposition import NMF, LatentDirichletAllocation
def print_top_words(model, feature_names, n_top_words):
for topic_idx, topic in enumerate(model.components_):
print("Topic #{}: {}".format(
topic_idx,
", ".join([feature_names[i] for i in topic.argsort()[:-n_top_words - 1:-1]])
))
```
Let's find some topics. We'll check out non-negative matrix factorization (NMF) first.
```
nmf = NMF(n_components=10, random_state=42, alpha=.1, l1_ratio=.5).fit(tfidf)
# Try out different numbers of topics (change n_components)
# Documentation: http://scikit-learn.org/stable/modules/generated/sklearn.decomposition.NMF.html
print("\nTopics in NMF model:")
print_top_words(nmf, ngrams, 10)
```
LDA is an other popular topic modeling technique
```
lda = LatentDirichletAllocation(n_topics=10, random_state=42).fit(tfidf)
# Documentation: http://scikit-learn.org/stable/modules/generated/sklearn.decomposition.LatentDirichletAllocation.html
# doc_topic_prior (alpha) - lower alpha means documents will be composed of fewer topics (higher means a more uniform distriution across all topics)
# topic_word_prior (beta) - lower beta means topis will be composed of fewer words (higher means a more uniform distribution across all words)
print("\nTopics in LDA model:")
print_top_words(lda, ngrams, 10)
```
We can use the topic models the same way we did our classifier - everything in Scikit-Learn follows the same fit/transform paradigm. So, let's get the topics for our documents.
```
doc_topics = pd.DataFrame(lda.transform(tfidf))
doc_topics.head()
topic_column_names = ["topic_{}".format(c) for c in doc_topics.columns]
doc_topics.columns = topic_column_names
```
Next we use Pandas to join the topics with the original sample dataframe
```
sample_with_topics = pd.concat([sample, doc_topics], axis=1)
```
Let's look for patterns by running some means and correlations
```
sample_with_topics.groupby("Score").mean()
for topic in topic_column_names:
print "{}: {}".format(topic, sample_with_topics[topic].corr(sample_with_topics['Score']))
```
Here's an example of a linear regression
```
from sklearn import datasets, linear_model
from sklearn.metrics import mean_squared_error, r2_score
training_data = sample_with_topics[topic_column_names[:-1]] # We're leaving a column out to prevent multicollinearity
regression = linear_model.LinearRegression()
# Train the model using the training sets
regression.fit(training_data, sample_with_topics['Score'])
coefficients = regression.coef_
print zip(topic_column_names[:-1], coefficients)
```
Sadly Scikit-Learn doesn't make it easy to get p-values or a regression report like you'd normally expect of something like R or Stata. Scikit-Learn is more about prediction than statistical analysis; for the latter, we can use Statsmodels.
```
import statsmodels.api as sm
regression = sm.OLS(training_data, sample_with_topics['Score'])
results = regression.fit()
print(results.summary())
```
## Clustering
We can also check out other unsupervised methods like clustering. I borrowed/modified some of this code from http://brandonrose.org/clustering
### K-Means Clustering
```
from sklearn.cluster import KMeans
kmeans = KMeans(n_clusters=10, max_iter=50, tol=.01)
# http://scikit-learn.org/stable/modules/generated/sklearn.cluster.KMeans.html
kmeans.fit(tfidf)
clusters = kmeans.labels_.tolist() # You can merge these back into the data if you want
centroids = kmeans.cluster_centers_.argsort()[:, ::-1]
for i, closest_ngrams in enumerate(centroids):
print "Cluster #{}: {}".format(i, np.array(ngrams)[closest_ngrams[:8]])
```
### Agglomerative/Hierarchical Clustering
```
%matplotlib inline
import matplotlib.pyplot as plt
from scipy.cluster.hierarchy import linkage, dendrogram
from sklearn.metrics.pairwise import cosine_similarity
# Uses cosine similarity to get word similarities based on document overlap
# To get this for document similarities in terms of word overlap, just drop the .transpose()!
similarities = cosine_similarity(tfidf.transpose())
distances = 1 - similarities # Converts to distances
clusters = linkage(distances, method='ward') # Run hierarchical clustering on the distances
fig, ax = plt.subplots(figsize=(15, min([len(ngrams)/10.0, 300])))
ax = dendrogram(clusters, labels=ngrams, orientation="left")
plt.tight_layout()
```
| github_jupyter |
```
import csv
import os
from os import listdir
import datetime
from datetime import datetime
import pandas as pd
import pathlib
import sort_bus_by_date
directory = sort_bus_by_date.find_directory()
bus_150_dates = sort_bus_by_date.sort_bus_by_date(directory, 'bus_150/')
bus_150_dates
n = 2
start_row = 51+ (11+47)*n
end_row = start_row + 12
row_list = list(range(start_row)) + list(range(end_row, 960))
start_row
test_file_dir = directory + 'all_data/' + 'False_files/' + 'bus_150/' + bus_150_dates['Filename'].loc[0]
test_file_dir
test_df = pd.read_csv(test_file_dir, header=None, skiprows=row_list)
index_range = list(range(50)) + list(range(51,960))
#df_index = pd.read_csv(test_file_dir, header=0, skiprows = index_range)
#test_df.columns = df_index.columns
test_df
test_df_index = pd.read_csv(test_file_dir, header=0, skiprows=index_range)
test_df
test_df_index
test_df = test_df.dropna(axis=1)
test_df
test_df = test_df.drop(0, axis=1)
test_df
dates = bus_150_dates['DateRetrieved'].astype(str)
test_df_dates = pd.concat([test_df, dates], axis=1)
test_df_dates
ave = test_df.mean()
test_ave_df = pd.DataFrame()
test_ave_df = test_ave_df.append(ave, ignore_index=True)
test_ave_df
def build_module_df(directory, bus_num, module_num):
bus_dates = sort_bus_by_date.sort_bus_by_date(directory, bus_num)
start_row = 51 + (11+47)* (module_num-1)
end_row = start_row + 12
row_list = list(range(start_row)) + list(range(end_row, 960))
index_range = list(range(50)) + list(range(51,960))
module_df = pd.DataFrame()
for i in range(len(bus_dates)):
file = bus_dates['Filename'].loc[i]
file_dir = directory + 'all_data/' + 'False_files/' + bus_num + file
tmp = pd.read_csv(file_dir, header=None, skiprows=row_list)
module_df = module_df.append(tmp)
df_index = pd.read_csv(file_dir, header=0, skiprows = index_range)
module_df.columns = df_index.columns
module_df = module_df.loc[:, ~module_df.columns.str.contains('^Unnamed')]
module_df.reset_index(drop = True, inplace=True)
return module_df
bus_1_mod_1 = build_module_df(directory, 'bus_1/', 1)
bus_1_mod_1
bus_1_mod_1.head(26)
bus_1_mod_1[:12]
bus_1_mod_1[12:24]
bus_1_mod_1[24:36]
import matplotlib
from matplotlib import pyplot as plt
len(bus_1_dates)
plt.hist(bus_1_mod_1[12:24])
fig, ax = plt.subplots(nrows = 4, ncols = 4)
for i in range(len(bus_1_dates)):
row_start = 12*i
row_end = row_start + 12
ax[i] = plt.hist(bus_1_mod_1[row_start:row_end])
def build_module_average_df(directory, bus_num, module_num):
bus_dates = sort_bus_by_date.sort_bus_by_date(directory, bus_num)
start_row = 51 + (11+47)* (module_num-1)
end_row = start_row + 12
row_list = list(range(start_row)) + list(range(end_row, 960))
index_range = list(range(50)) + list(range(51,960))
module_average_df = pd.DataFrame()
for i in range(len(bus_dates)):
file = bus_dates['Filename'].loc[i]
print(file)
file_dir = directory + 'all_data/' + 'False_files/' + bus_num + file
tmp = pd.read_csv(file_dir, header=None, skiprows=row_list)
print(tmp)
tmp = tmp.dropna(axis=1)
print(tmp)
tmp = tmp.drop(0, axis=1)
tmp_ave = tmp.mean()
module_average_df = module_average_df.append(tmp_ave, ignore_index = True)
df_index = pd.read_csv(file_dir, header=0, skiprows = index_range)
df_index = df_index.loc[:, ~df_index.columns.str.contains('^Unnamed')]
module_average_df.columns = df_index.columns
module_average_df.reset_index(drop = True, inplace=True)
module_average_df_final = pd.concat([module_average_df, bus_dates['DateRetrieved']], axis=1)
return module_average_df_final
bus_150_mod_14_ave = build_module_average_df(directory, 'bus_150/', 14)
bus_1_mod_1_ave.columns
bus_1_mod_1_ave = bus_1_mod_1_ave.drop('TOTAL',axis=1)
bus_1_mod_1_ave
bus_1_mod_1_ave[0:1]
y = []
for i in range(len(bus_1_dates)):
x = bus_1_mod_1_ave.iloc[i].values
y.append(x)
x = [1.9, 2.1, 2.3, 2.5, 2.7, 2.9, 3.1, 3.3, 3.5, 3.7, 3.9, 4.1] #voltage bins
bus_1_dates
fig, axs = plt.subplots(9,2, figsize=(20, 40), facecolor='w', edgecolor='k')
axs = axs.ravel()
for i in range(len(bus_1_dates)):
axs[i].bar(x, y[i])
axs[i].set_title('Module 1 ' + str(bus_1_dates['DateRetrieved'].iloc[i]))
axs[i].set_xlabel('Voltage')
axs[i].set_xticks([2.0, 2.2, 2.4, 2.6, 2.8, 3.0, 3.2, 3.4, 3.6, 3.8, 4.0])
axs[i].set_ylabel('Time (s)')
fig.tight_layout()
bus_20_mod_6_ave = build_module_average_df(directory, 'bus_20/', 6)
bus_20_mod_6_ave
```
| github_jupyter |
```
import sys
import os
import glob
import pandas as pd
import numpy as np
import text_extensions_for_pandas as tp
from download_and_correct_corpus import Dataset
files = {
'csv_files' : ["../corrected_labels/all_conll_corrections_combined.csv"],
'dev' : "../original_corpus/eng.testa",
'test' : "../original_corpus/eng.testb",
'train' : "../original_corpus/eng.train"
}
columns = ['doc_offset', 'corpus_span', 'correct_span']
test_df = pd.DataFrame(columns = columns)
dev_df = pd.DataFrame(columns = columns)
train_df = pd.DataFrame(columns = columns)
for f in files['csv_files']:
current_df = pd.read_csv(os.path.abspath(f))
test_df = test_df.append(current_df[(current_df["error_type"]=="Sentence") & (current_df["fold"]=="test")][columns], ignore_index=True)
dev_df = dev_df.append(current_df[(current_df["error_type"]=="Sentence") & (current_df["fold"]=="dev")][columns], ignore_index=True)
train_df = train_df.append(current_df[(current_df["error_type"]=="Sentence") & (current_df["fold"]=="train")][columns], ignore_index=True)
test_df.to_csv("../corrected_labels/sentence_corection_test.csv")
dev_df.to_csv("../corrected_labels/sentence_corection_dev.csv")
train_df.to_csv("../corrected_labels/sentence_corection_train.csv")
correction_df = {
'dev' : dev_df,
'test' : test_df,
'train': train_df
}
splits = ['dev', 'test', 'train']
lines_to_delete = {
'dev' : [],
'test' : [],
'train': []
}
for split in splits:
# Read the raw corpus file lines
f = open(files[split])
lines = f.readlines()
# Create a dataframe for the corpus file and process our corrections csv
dataset = Dataset(files[split])
current_df = correction_df[split]
for i, row in current_df.iterrows():
if split == 'test' and i >= 59:
continue
try:
candidate_lines = dataset.find(row["correct_span"], int(row["doc_offset"]))
except:
candidate_lines = dataset.find(row["corpus_span"], int(row["doc_offset"]))
candidate_lines = (candidate_lines[0]-1, candidate_lines[1]+1)
print("The correct_span did not match lines, using corpus span instead at {}, {}".format(split, i))
appended = 0
for c in range(candidate_lines[0], candidate_lines[1]+1):
if lines[c] == "\n":
lines_to_delete[split].append(c)
appended += 1
if appended == 0:
print("Nothing to append here! Check {}, {} again".format(split, i))
for l in lines_to_delete:
lines_to_delete[l] = list(dict.fromkeys(lines_to_delete[l]))
lines_to_delete[l].sort(reverse=True)
import pprint
pprint.pprint(lines_to_delete)
import json
json = json.dumps(lines_to_delete, indent=4, sort_keys=True)
f = open("../corrected_labels/sentence_corrections.json","w")
f.write(json)
f.close()
```
| github_jupyter |
```
import nltk
from nltk.corpus import twitter_samples
nltk.download('twitter_samples')
positive_tweets = twitter_samples.strings('positive_tweets.json')
negative_tweets = twitter_samples.strings('negative_tweets.json')
len(positive_tweets)
len(negative_tweets)
positive_tweets[:10]
import re
tweet = positive_tweets[5]
tweet
tweet = re.sub(r'(http|https)?:\/\/.*[\r\n]*', '', tweet)
tweet
tweet = re.sub(r'#', '', tweet)
tweet
tweet = re.sub(r'\n', ' ', tweet)
tweet
tweet = re.sub(r'@\w+ ', '', tweet)
tweet
tweet = tweet.lower()
tweet
```
## tweet tokenizer
```
from nltk.tokenize import TweetTokenizer
tokenizer = TweetTokenizer()
tokens = tokenizer.tokenize(tweet)
tokens
from nltk.corpus import stopwords
nltk.download('stopwords')
print(stopwords.words('english'))
tokens = [token for token in tokens if token not in stopwords.words('english')]
tokens
from nltk.stem import PorterStemmer
stemmer = PorterStemmer()
tokens = [stemmer.stem(token) for token in tokens]
tokens
import string
print (string.punctuation)
def process_tweet(tweet, tokenizer,stemmer):
tweet = re.sub(r'(http|https)?:\/\/.*[\r\n]*', '', tweet)
tweet = re.sub(r'#','', tweet)
tweet = re.sub(r'\n', ' ', tweet)
tweet = re.sub(r'@\w+ ', '', tweet)
tweet = tweet.lower()
tokens = tokenizer.tokenize(tweet)
tokens = [token for token in tokens if token not in stopwords.words('english')]
tokens = [token for token in tokens if token not in string.punctuation]
tokens = [stemmer.stem(token) for token in tokens]
return tokens
process_tweet(positive_tweets[2], tokenizer, stemmer)
import numpy as np
np.ones(10)
np.zeros(5)
labels = np.append(np.ones(5000), np.zeros(5000))
len(labels)
labels
labels = np.array([labels])
labels
labels = labels.T
labels
tweets = positive_tweets + negative_tweets
len(tweets)
tweets = np.array([tweets])
tweets = tweets.T
tweets
tweets.shape
labels.shape
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(tweets, labels, test_size=0.3)
X_train
y_train
y_train.shape
frequncies = {('happy', 1.0) : 50, ('happy', 0.0) : 10 }
frequncies.get(('happy', 1.0))
frequncies[('happy', 1.0)] = 65
frequncies.get(('happy', 1.0))
frequncies[('sad', 1.0)] = 5
frequncies[('sad', 0.0)] = 55
frequncies
frequncies = {}
for tweet, label in zip(X_train, y_train):
for token in process_tweet(tweet[0], tokenizer, stemmer):
pair = (token, label[0])
frequncies[pair] = frequncies.get(pair, 0) + 1
frequncies
frequncies.get(('😘', 0.0))
frequncies.get(('😘', 1.0))
frequncies.get(("sad", 1.0))
frequncies.get(('happi', 0.0))
X_train_features = np.zeros((7000, 3))
X_train_features
X_train_features.shape
def generate_features(X):
for i in range(len(X_train)):
pos_points = 0
neg_points = 0
tokens = process_tweet(X_train[i][0], tokenizer, stemmer)
tokens = list(dict.fromkeys(tokens))
for token in tokens:
pos_points += frequncies.get((token, 1.0), 0)
neg_points += frequncies.get((token, 0.0), 0)
X_train_features[i, :] = np.array((pos_points, neg_points, 1))
X_train_features
```
| github_jupyter |
# Visualization with Seaborn
Matplotlib has proven to be an incredibly useful and popular visualization tool, but even avid users will admit it often leaves much to be desired.
There are several valid complaints about Matplotlib that often come up:
- Prior to version 2.0, Matplotlib's defaults are not exactly the best choices. It was based off of MATLAB circa 1999, and this often shows.
- Matplotlib's API is relatively low level. Doing sophisticated statistical visualization is possible, but often requires a *lot* of boilerplate code.
- Matplotlib predated Pandas by more than a decade, and thus is not designed for use with Pandas ``DataFrame``s. In order to visualize data from a Pandas ``DataFrame``, you must extract each ``Series`` and often concatenate them together into the right format. It would be nicer to have a plotting library that can intelligently use the ``DataFrame`` labels in a plot.
An answer to these problems is [Seaborn](http://seaborn.pydata.org/). Seaborn provides an API on top of Matplotlib that offers sane choices for plot style and color defaults, defines simple high-level functions for common statistical plot types, and integrates with the functionality provided by Pandas ``DataFrame``s.
To be fair, the Matplotlib team is addressing this: it has recently added the ``plt.style`` tools and is starting to handle Pandas data more seamlessly.
The newer releases of the library include a default stylesheet that has improved on the visualization. But for all the reasons just discussed, Seaborn remains an extremely useful addon.
## Updating Seaborn on Colab
Colab uses an older version of Seaborn as of writing this notebook. Let's first update it. **Uncomment the cell below and execute it** before continuing on the notebook, if using Colab:
```
#!pip install seaborn --upgrade
```
Please note that you must restart the runtime in order to use newly installed versions. From the menu select Runtime > Restart Runtime.
```
import seaborn
seaborn.__version__
from IPython.display import Pretty as disp
hint = 'https://raw.githubusercontent.com/soltaniehha/Business-Analytics/master/docs/hints/' # path to hints on GitHub
import matplotlib.pyplot as plt
#plt.style.use('classic')
%matplotlib inline
import numpy as np
import pandas as pd
import seaborn as sns
sns.set(rc={'figure.figsize':(10,8)}) # Figure size
```
## Exploring Seaborn Plots
The main idea of Seaborn is that it provides high-level commands to create a variety of plot types useful for statistical data exploration, and even some statistical model fitting.
Let's take a look at a few of the datasets and plot types available in Seaborn. Note that all of the following *could* be done using raw Matplotlib commands (this is, in fact, what Seaborn does under the hood) but the Seaborn API is much more convenient.
## Visualizing statistical **rel**ationships
Statistical analysis is a process of understanding how variables in a dataset relate to each other and how those relationships depend on other variables. Visualization can be a core component of this process because, when data are visualized properly, the human visual system can see trends and patterns that indicate a relationship.
### Relating variables with scatter plots
There are several ways to draw a scatter plot in seaborn. The most basic, which should be used when both variables are numeric, is the `scatterplot()` function.
Let's first load some data from Seaborn's available datasets:
```
tips = sns.load_dataset("tips")
tips.head()
sns.scatterplot(x="total_bill", y="tip", data=tips);
plt.axvline(25, color="k", linestyle="--", alpha=0.5);
```
**Note:** We have combined a *plt* plot with an *sns* one. But in reality they both are Matplotlib plots.
While the points are plotted in two dimensions, another dimension can be added to the plot by coloring the points according to a third variable. In seaborn, this is referred to as using a “hue semantic”, because the color of the point gains meaning:
```
sns.scatterplot(x="total_bill", y="tip", hue="smoker", data=tips);
```
It’s also possible to represent four variables by changing the hue and style of each point independently. But this should be done carefully, because the eye is much less sensitive to shape than to color:
```
sns.scatterplot(x="total_bill", y="tip", hue="smoker", style="time", data=tips);
```
### Showing multiple relationships with facets
We can repeat the previous plots by using `relplot()` to get even more functionality such as a FacetGrid. `relplot()` can be used for scatter and line plots but the default kind is `kind='scatter'`.
```
sns.relplot(x="total_bill", y="tip", hue="time", col="day", data=tips);
```
For more examples of `relplot()` visit [Seaborn tutorial: Visualizing statistical **rel**ationships](https://seaborn.pydata.org/tutorial/relational.html).
## Plotting with categorical data
If one of the main variables is “categorical” (divided into discrete groups) it may be helpful to use a more specialized approach to visualization.
In seaborn, there are several different ways to visualize a relationship involving categorical data. Similar to the relationship between `relplot()` and either `scatterplot()` or `lineplot()`, there are two ways to make these plots. There are a number of axes-level functions for plotting categorical data in different ways and a figure-level interface, `catplot()`, that gives unified higher-level access to them.
It’s helpful to think of the different categorical plot kinds as belonging to three different families, which we’ll discuss in detail below. They are:
Categorical scatterplots:
* `stripplot()` (with kind="strip"; the default)
* `swarmplot()` (with kind="swarm")
Categorical distribution plots:
* `boxplot()` (with kind="box")
* `violinplot()` (with kind="violin")
Categorical estimate plots:
* `pointplot()` (with kind="point")
* `barplot()` (with kind="bar")
* `countplot()` (with kind="count")
Here we’ll mostly focus on the figure-level interface, `catplot()`. Remember that this function is a higher-level interface each of the functions above, so we’ll reference them when we show each kind of plot, keeping the more verbose kind-specific API documentation at hand.
### Categorical scatterplots
The default representation of the data in `catplot()` uses a scatterplot (kind='strip').
```
sns.catplot(x="day", y="total_bill", data=tips);
```
Change the order:
```
sns.catplot(x="day", y="total_bill", data=tips, order=['Sat','Sun','Thur','Fri']);
```
With **jitter** off:
```
sns.catplot(x="day", y="total_bill", jitter=False, data=tips);
```
**Swarm plots:** adjusts the points along the categorical axis using an algorithm that prevents them from overlapping.
### Your turn
Set kind="swarm" to plot a swarm plot. Can you also set the "height" to a larger value? What else do you need to change to obtain the chart below?
```
# Your answer goes here
# Don't run this cell to keep the outcome as your frame of reference
# SOLUTION: Uncomment and execute the cell below to get help
#disp(hint + '09-02-swarm')
```
### Boxplots
```
sns.catplot(x="day", y="total_bill", kind="box", data=tips);
sns.catplot(x="day", y="total_bill", hue="smoker", kind="box", data=tips);
```
Showing multiple relationships with facets by adding *col* or *row*:
```
sns.catplot(x="day", y="total_bill", hue="smoker", col='sex', kind="box", data=tips);
```
### Your turn
Can you reproduce the following plot?
```
# Your answer goes here
# Don't run this cell to keep the outcome as your frame of reference
# SOLUTION: Uncomment and execute the cell below to get help
#disp(hint + '09-02-box')
```
### Violinplots
combines a boxplot with the kernel density estimation:
```
sns.catplot(y="total_bill", x="day", kind="violin", data=tips);
```
`cut` is used to extend the density past the extreme datapoints. Set to 0 to limit the violin range within the range of the observed data:
```
sns.catplot(y="total_bill", x="day", kind="violin", cut=0, data=tips);
```
It’s also possible to “split” the violins when the hue parameter has only two levels, which can allow for a more efficient use of space. Note that in the plot below we have also used a inner="stick":
```
sns.catplot(x="day", y="total_bill", hue="sex", kind="violin", split=True, inner="stick", data=tips);
```
In order to get access to more specific arguments related to violinplots read the help document: `sns.violinplot?`
### Bar plots
In seaborn, the `barplot()` function operates on a full dataset and applies a function to obtain the estimate (taking the mean by default). When there are multiple observations in each category, it also uses bootstrapping to compute a confidence interval around the estimate and plots that using error bars:
```
sns.catplot(x="day", y="size", kind="bar", data=tips);
```
### `countplot()`
```
sns.catplot(x="day", kind="count", hue='sex', data=tips);
```
For more examples of `catplot()` visit [Seaborn tutorial: Plotting with categorical data](https://seaborn.pydata.org/tutorial/categorical.html).
## Visualizing the distribution of a dataset
### Plotting univariate distributions
```
fig, ax = plt.subplots(2,1)
sns.distplot(tips['tip'], ax=ax[0]);
sns.distplot(tips['tip'], kde=False, bins=10, color='r', ax=ax[1]);
```
### Plotting bivariate distributions
```
sns.jointplot(x="tip", y="total_bill", data=tips);
```
### Visualizing pairwise relationships in a dataset
```
sns.pairplot(tips, hue='sex');
```
For more examples of this kind visit [Seaborn tutorial: Visualizing the distribution of a dataset](https://seaborn.pydata.org/tutorial/distributions.html).
## Visualizing linear relationships
### Functions to draw linear regression models
```
sns.lmplot(x="total_bill", y="tip", data=tips);
```
It’s possible to fit a linear regression when one of the variables takes discrete values, however, the simple scatterplot produced by this kind of dataset is often not optimal:
```
sns.lmplot(x="size", y="tip", data=tips);
```
We can collapse over the observations in each discrete bin to plot an estimate of central tendency along with a confidence interval:
```
sns.lmplot(x="size", y="tip", data=tips, x_estimator=np.mean);
```
### Fitting different kinds of models
```
anscombe = sns.load_dataset("anscombe")
sns.lmplot(x="x", y="y", data=anscombe.query("dataset == 'II'"), ci=None, scatter_kws={"s": 80});
```
In the presence of these kind of higher-order relationships, `lmplot()` and `regplot()` can fit a polynomial regression model to explore simple kinds of nonlinear trends in the dataset:
```
sns.lmplot(x="x", y="y", data=anscombe.query("dataset == 'II'"), order=2, ci=None, scatter_kws={"s": 80});
sns.lmplot(x="x", y="y", data=anscombe.query("dataset == 'III'"), ci=None, scatter_kws={"s": 80});
```
In the presence of outliers, it can be useful to fit a robust regression, which uses a different loss function to downweight relatively large residuals:
```
sns.lmplot(x="x", y="y", data=anscombe.query("dataset == 'III'"), robust=True, ci=None, scatter_kws={"s": 80});
tips["big_tip"] = (tips.tip / tips.total_bill) > .15
sns.lmplot(x="total_bill", y="big_tip", data=tips, y_jitter=.03);
plt.title('Linear regression fit');
```
The solution in this case is to fit a logistic regression, such that the regression line shows the estimated probability of y = 1 for a given value of x:
```
sns.lmplot(x="total_bill", y="big_tip", data=tips, logistic=True, y_jitter=.03);
plt.title('Logistic regression fit');
```
For more examples of `regplot()` and `lmplot()` visit [Seaborn tutorial: Visualizing linear relationships](https://seaborn.pydata.org/tutorial/regression.html).
For more information on plotting with Seaborn, see the [Seaborn documentation](http://seaborn.pydata.org/), a [tutorial](http://seaborn.pydata.org/
tutorial.htm), and the [Seaborn gallery](http://seaborn.pydata.org/examples/index.html).
## Further Reading
* [Official Seaborn tutorial](https://seaborn.pydata.org/tutorial.html)
* [Seaborn tutorial: Visualizing statistical relationships](https://seaborn.pydata.org/tutorial/relational.html)
* [Seaborn tutorial: Plotting with categorical data](https://seaborn.pydata.org/tutorial/categorical.html)
* [Seaborn tutorial: Visualizing the distribution of a dataset](https://seaborn.pydata.org/tutorial/distributions.html)
* [Seaborn tutorial: Visualizing linear relationships](https://seaborn.pydata.org/tutorial/regression.html)
* [Building structured multi-plot grids](https://seaborn.pydata.org/tutorial/axis_grids.html)
* [Controlling figure aesthetics](https://seaborn.pydata.org/tutorial/aesthetics.html)
* [Choosing color palettes](https://seaborn.pydata.org/tutorial/color_palettes.html)
* [Seaborn gallery](http://seaborn.pydata.org/examples/index.html)
* [Python Data Science Handbook: Visualization with Seaborn](https://github.com/jakevdp/PythonDataScienceHandbook/blob/8a34a4f653bdbdc01415a94dc20d4e9b97438965/notebooks/04.14-Visualization-With-Seaborn.ipynb)
## Other Python Graphics Libraries
Although Matplotlib is the most prominent Python visualization library, there are other more modern tools that are worth exploring as well.
I'll mention a few of them briefly here:
- [Bokeh](http://bokeh.pydata.org) is a visualization library with a Python frontend that creates highly interactive visualizations capable of handling very large and/or streaming datasets. The Python front-end outputs a JSON data structure that can be interpreted by the Bokeh JS engine.
- Check out these [live tutorial notebooks](https://mybinder.org/v2/gh/bokeh/bokeh-notebooks/master?filepath=tutorial%2F00%20-%20Introduction%20and%20Setup.ipynb).
- [Plotly](http://plot.ly) is the eponymous open source product of the Plotly company, and is similar in spirit to Bokeh. Because Plotly is the main product of a startup, it is receiving a high level of development effort. Use of the library is entirely free.
- [Vispy](http://vispy.org/) is an actively developed project focused on dynamic visualizations of very large datasets. Because it is built to target OpenGL and make use of efficient graphics processors in your computer, it is able to render some quite large and stunning visualizations.
- [Vega](https://vega.github.io/) and [Vega-Lite](https://vega.github.io/vega-lite) are declarative graphics representations, and are the product of years of research into the fundamental language of data visualization. The reference rendering implementation is JavaScript, but the API is language agnostic. There is a Python API under development in the [Altair](https://altair-viz.github.io/) package. Though as of summer 2016 it's not yet fully mature, I'm quite excited for the possibilities of this project to provide a common reference point for visualization in Python and other languages.
The visualization space in the Python community is very dynamic, and I fully expect this list to be out of date as soon as it is published.
Keep an eye out for what's coming in the future!
### An Example of Plotly Express
More examples can be found [here](https://plotly.com/python/plotly-express/).
```
# Install and import the package
!pip install plotly_express -q
import plotly_express as px
df = px.data.iris()
fig = px.scatter(df, x="sepal_width", y="sepal_length", color="species")
fig.show()
df = px.data.gapminder()
fig = px.choropleth(df, locations="iso_alpha", color="lifeExp", hover_name="country", animation_frame="year", range_color=[20,80])
fig.show()
```
| github_jupyter |
# Computer Vision Nanodegree
## Project: Image Captioning
---
In this notebook, you will train your CNN-RNN model.
You are welcome and encouraged to try out many different architectures and hyperparameters when searching for a good model.
This does have the potential to make the project quite messy! Before submitting your project, make sure that you clean up:
- the code you write in this notebook. The notebook should describe how to train a single CNN-RNN architecture, corresponding to your final choice of hyperparameters. You should structure the notebook so that the reviewer can replicate your results by running the code in this notebook.
- the output of the code cell in **Step 2**. The output should show the output obtained when training the model from scratch.
This notebook **will be graded**.
Feel free to use the links below to navigate the notebook:
- [Step 1](#step1): Training Setup
- [Step 2](#step2): Train your Model
- [Step 3](#step3): (Optional) Validate your Model
<a id='step1'></a>
## Step 1: Training Setup
In this step of the notebook, you will customize the training of your CNN-RNN model by specifying hyperparameters and setting other options that are important to the training procedure. The values you set now will be used when training your model in **Step 2** below.
You should only amend blocks of code that are preceded by a `TODO` statement. **Any code blocks that are not preceded by a `TODO` statement should not be modified**.
### Task #1
Begin by setting the following variables:
- `batch_size` - the batch size of each training batch. It is the number of image-caption pairs used to amend the model weights in each training step.
- `vocab_threshold` - the minimum word count threshold. Note that a larger threshold will result in a smaller vocabulary, whereas a smaller threshold will include rarer words and result in a larger vocabulary.
- `vocab_from_file` - a Boolean that decides whether to load the vocabulary from file.
- `embed_size` - the dimensionality of the image and word embeddings.
- `hidden_size` - the number of features in the hidden state of the RNN decoder.
- `num_epochs` - the number of epochs to train the model. We recommend that you set `num_epochs=3`, but feel free to increase or decrease this number as you wish. [This paper](https://arxiv.org/pdf/1502.03044.pdf) trained a captioning model on a single state-of-the-art GPU for 3 days, but you'll soon see that you can get reasonable results in a matter of a few hours! (_But of course, if you want your model to compete with current research, you will have to train for much longer._)
- `save_every` - determines how often to save the model weights. We recommend that you set `save_every=1`, to save the model weights after each epoch. This way, after the `i`th epoch, the encoder and decoder weights will be saved in the `models/` folder as `encoder-i.pkl` and `decoder-i.pkl`, respectively.
- `print_every` - determines how often to print the batch loss to the Jupyter notebook while training. Note that you **will not** observe a monotonic decrease in the loss function while training - this is perfectly fine and completely expected! You are encouraged to keep this at its default value of `100` to avoid clogging the notebook, but feel free to change it.
- `log_file` - the name of the text file containing - for every step - how the loss and perplexity evolved during training.
If you're not sure where to begin to set some of the values above, you can peruse [this paper](https://arxiv.org/pdf/1502.03044.pdf) and [this paper](https://arxiv.org/pdf/1411.4555.pdf) for useful guidance! **To avoid spending too long on this notebook**, you are encouraged to consult these suggested research papers to obtain a strong initial guess for which hyperparameters are likely to work best. Then, train a single model, and proceed to the next notebook (**3_Inference.ipynb**). If you are unhappy with your performance, you can return to this notebook to tweak the hyperparameters (and/or the architecture in **model.py**) and re-train your model.
### Question 1
**Question:** Describe your CNN-RNN architecture in detail. With this architecture in mind, how did you select the values of the variables in Task 1? If you consulted a research paper detailing a successful implementation of an image captioning model, please provide the reference.
**Answer:**
I left the encoder CNN architecture as it is, it produces a vector with the same length as the embedding vectors to be used in our decoder.
The decoder is 2 layer lstm.I arrived at this number by observing outputs from trying out with 1 and 3 layers(i trained them for one epoch before testing to save gpu time,so might be wrong in drawing a conclusion too soon) .
I have added dropout in the LSTM cell as well as a seperate layer before the linear layer.The output of the LSTM is passed through a linear layer.
Referring to a Stanford paper on CNN-LSTM architecture for caption generation(https://cs224d.stanford.edu/reports/msoh.pdf), I tried incorporating a softmax layer after my linear layer. But I was getting a loss of around 6 in a couple of epochs i tried it , i was getting it around 2 without it hence i dropped it.
I tried batch sizes of 64,128,256. batch size of 256 was pretty slow to train in my system ,taking around 2 hours 40 mins for one epoch, 128 was slightly better without much difference in the results. Batch size of 64 didnt also give enough different in results to opt it against 128. Once again i arrived these conclusions on just a couple of epochs i tried it on ,hence might not have been a better choice in longer training times.
I trained my final model for 4 epochs, i stopped it midway the 5th epoch as is saw it was giving me results and the loss had smoothened out somewhat and it did not seems worth more gpu time to train for another epoch.
I tried out cross entropy loss and nll loss.Cross entropy gave me better reuslts.
### (Optional) Task #2
Note that we have provided a recommended image transform `transform_train` for pre-processing the training images, but you are welcome (and encouraged!) to modify it as you wish. When modifying this transform, keep in mind that:
- the images in the dataset have varying heights and widths, and
- if using a pre-trained model, you must perform the corresponding appropriate normalization.
### Question 2
**Question:** How did you select the transform in `transform_train`? If you left the transform at its provided value, why do you think that it is a good choice for your CNN architecture?
**Answer:**
The transforms were not altered. With a random crop of a 256 resized image we ensure images with typical aspect ratios retain most of the central part of the image. If the pre trained model already had been trained with flip images horizontal flip might not have been very useful.
### Task #3
Next, you will specify a Python list containing the learnable parameters of the model. For instance, if you decide to make all weights in the decoder trainable, but only want to train the weights in the embedding layer of the encoder, then you should set `params` to something like:
```
params = list(decoder.parameters()) + list(encoder.embed.parameters())
```
### Question 3
**Question:** How did you select the trainable parameters of your architecture? Why do you think this is a good choice?
**Answer:**
I decided to train the decoder parameters here. I was not sure if pre trained ResNet couldve been trained effectively in the GPU with the timing contraints.
### Task #4
Finally, you will select an [optimizer](http://pytorch.org/docs/master/optim.html#torch.optim.Optimizer).
### Question 4
**Question:** How did you select the optimizer used to train your model?
**Answer:** I experimented with Adam ,RMSprop & SGD. The default training rates were high for all these optimizers so reduced those. Adam gave me the best results RMSprop being close (loss of around 4 at learning rate of 0.004).
```
import nltk
nltk.download('punkt')
import torch
import torch.nn as nn
from torchvision import transforms
import sys
sys.path.append('/opt/cocoapi/PythonAPI')
from pycocotools.coco import COCO
from data_loader import get_loader
from model import EncoderCNN, DecoderRNN
import math
## TODO #1: Select appropriate values for the Python variables below.
batch_size = 128 # batch size
vocab_threshold = 6 # minimum word count threshold
vocab_from_file = True # if True, load existing vocab file
embed_size = 256 # dimensionality of image and word embeddings
hidden_size = 512 # number of features in hidden state of the RNN decoder
num_epochs = 5 # number of training epochs
save_every = 1 # determines frequency of saving model weights
print_every = 400 # determines window for printing average loss
log_file = 'training_log.txt' # name of file with saved training loss and perplexity
# (Optional) TODO #2: Amend the image transform below.
transform_train = transforms.Compose([
transforms.Resize(256), # smaller edge of image resized to 256
transforms.RandomCrop(224), # get 224x224 crop from random location
transforms.RandomHorizontalFlip(), # horizontally flip image with probability=0.5
transforms.ToTensor(), # convert the PIL Image to a tensor
transforms.Normalize((0.485, 0.456, 0.406), # normalize image for pre-trained model
(0.229, 0.224, 0.225))])
# Build data loader.
data_loader = get_loader(transform=transform_train,
mode='train',
batch_size=batch_size,
vocab_threshold=vocab_threshold,
vocab_from_file=vocab_from_file)
# The size of the vocabulary.
vocab_size = len(data_loader.dataset.vocab)
# Initialize the encoder and decoder.
encoder = EncoderCNN(embed_size)
decoder = DecoderRNN(embed_size, hidden_size, vocab_size)
# Move models to GPU if CUDA is available.
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
encoder.to(device)
decoder.to(device)
# Define the loss function.
criterion = nn.CrossEntropyLoss().cuda() if torch.cuda.is_available() else nn.CrossEntropyLoss()
# TODO #3: Specify the learnable parameters of the model.
params = list(decoder.parameters())
# TODO #4: Define the optimizer.
optimizer = torch.optim.Adam(params, lr=0.002)
# Set the total number of training steps per epoch.
total_step = math.ceil(len(data_loader.dataset.caption_lengths) / data_loader.batch_sampler.batch_size)
```
<a id='step2'></a>
## Step 2: Train your Model
Once you have executed the code cell in **Step 1**, the training procedure below should run without issue.
It is completely fine to leave the code cell below as-is without modifications to train your model. However, if you would like to modify the code used to train the model below, you must ensure that your changes are easily parsed by your reviewer. In other words, make sure to provide appropriate comments to describe how your code works!
You may find it useful to load saved weights to resume training. In that case, note the names of the files containing the encoder and decoder weights that you'd like to load (`encoder_file` and `decoder_file`). Then you can load the weights by using the lines below:
```python
# Load pre-trained weights before resuming training.
encoder.load_state_dict(torch.load(os.path.join('./models', encoder_file)))
decoder.load_state_dict(torch.load(os.path.join('./models', decoder_file)))
```
While trying out parameters, make sure to take extensive notes and record the settings that you used in your various training runs. In particular, you don't want to encounter a situation where you've trained a model for several hours but can't remember what settings you used :).
### A Note on Tuning Hyperparameters
To figure out how well your model is doing, you can look at how the training loss and perplexity evolve during training - and for the purposes of this project, you are encouraged to amend the hyperparameters based on this information.
However, this will not tell you if your model is overfitting to the training data, and, unfortunately, overfitting is a problem that is commonly encountered when training image captioning models.
For this project, you need not worry about overfitting. **This project does not have strict requirements regarding the performance of your model**, and you just need to demonstrate that your model has learned **_something_** when you generate captions on the test data. For now, we strongly encourage you to train your model for the suggested 3 epochs without worrying about performance; then, you should immediately transition to the next notebook in the sequence (**3_Inference.ipynb**) to see how your model performs on the test data. If your model needs to be changed, you can come back to this notebook, amend hyperparameters (if necessary), and re-train the model.
That said, if you would like to go above and beyond in this project, you can read about some approaches to minimizing overfitting in section 4.3.1 of [this paper](http://ieeexplore.ieee.org/stamp/stamp.jsp?arnumber=7505636). In the next (optional) step of this notebook, we provide some guidance for assessing the performance on the validation dataset.
```
import torch.utils.data as data
import numpy as np
import os
import requests
import time
# Open the training log file.
f = open(log_file, 'w')
old_time = time.time()
response = requests.request("GET",
"http://metadata.google.internal/computeMetadata/v1/instance/attributes/keep_alive_token",
headers={"Metadata-Flavor":"Google"})
for epoch in range(1, num_epochs+1):
for i_step in range(1, total_step+1):
if time.time() - old_time > 60:
old_time = time.time()
requests.request("POST",
"https://nebula.udacity.com/api/v1/remote/keep-alive",
headers={'Authorization': "STAR " + response.text})
# Randomly sample a caption length, and sample indices with that length.
indices = data_loader.dataset.get_train_indices()
# Create and assign a batch sampler to retrieve a batch with the sampled indices.
new_sampler = data.sampler.SubsetRandomSampler(indices=indices)
data_loader.batch_sampler.sampler = new_sampler
# Obtain the batch.
images, captions = next(iter(data_loader))
# Move batch of images and captions to GPU if CUDA is available.
images = images.to(device)
captions = captions.to(device)
# Zero the gradients.
decoder.zero_grad()
encoder.zero_grad()
# Pass the inputs through the CNN-RNN model.
features = encoder(images)
outputs = decoder(features, captions)
# Calculate the batch loss.
loss = criterion(outputs.view(-1, vocab_size), captions.view(-1))
# Backward pass.
loss.backward()
# Update the parameters in the optimizer.
optimizer.step()
# Get training statistics.
stats = 'Epoch [%d/%d], Step [%d/%d], Loss: %.4f, Perplexity: %5.4f' % (epoch, num_epochs, i_step, total_step, loss.item(), np.exp(loss.item()))
# Print training statistics (on same line).
print('\r' + stats, end="")
sys.stdout.flush()
# Print training statistics to file.
f.write(stats + '\n')
f.flush()
# Print training statistics (on different line).
if i_step % print_every == 0:
print('\r' + stats)
# Save the weights.
if epoch % save_every == 0:
torch.save(decoder.state_dict(), os.path.join('./models', 'decoder-%d.pkl' % epoch))
torch.save(encoder.state_dict(), os.path.join('./models', 'encoder-%d.pkl' % epoch))
# Close the training log file.
f.close()
```
<a id='step3'></a>
## Step 3: (Optional) Validate your Model
To assess potential overfitting, one approach is to assess performance on a validation set. If you decide to do this **optional** task, you are required to first complete all of the steps in the next notebook in the sequence (**3_Inference.ipynb**); as part of that notebook, you will write and test code (specifically, the `sample` method in the `DecoderRNN` class) that uses your RNN decoder to generate captions. That code will prove incredibly useful here.
If you decide to validate your model, please do not edit the data loader in **data_loader.py**. Instead, create a new file named **data_loader_val.py** containing the code for obtaining the data loader for the validation data. You can access:
- the validation images at filepath `'/opt/cocoapi/images/train2014/'`, and
- the validation image caption annotation file at filepath `'/opt/cocoapi/annotations/captions_val2014.json'`.
The suggested approach to validating your model involves creating a json file such as [this one](https://github.com/cocodataset/cocoapi/blob/master/results/captions_val2014_fakecap_results.json) containing your model's predicted captions for the validation images. Then, you can write your own script or use one that you [find online](https://github.com/tylin/coco-caption) to calculate the BLEU score of your model. You can read more about the BLEU score, along with other evaluation metrics (such as TEOR and Cider) in section 4.1 of [this paper](https://arxiv.org/pdf/1411.4555.pdf). For more information about how to use the annotation file, check out the [website](http://cocodataset.org/#download) for the COCO dataset.
```
# (Optional) TODO: Validate your model.
```
| github_jupyter |
# parkinsons-detection
```
%matplotlib inline
import numpy as np
import pandas as pd
import os
import seaborn as sns
from time import time
from math import sqrt
```
#### Timing decorator
```
def timeit(fn):
def wrapper(*args, **kwargs):
start=time()
res=fn(*args, **kwargs)
print(fn.__name__, "took", time()-start, "seconds.")
return res
return wrapper
```
## Data paths
```
control_data_path=os.path.join('data', 'control')
parkinson_data_path=os.path.join('data', 'parkinson')
control_file_list=[os.path.join(control_data_path, x) for x in os.listdir(control_data_path)]
parkinson_file_list=[os.path.join(parkinson_data_path, x) for x in os.listdir(parkinson_data_path)]
```
### Features
1. No of strokes
2. Stroke speed
3. Velocity
4. Acceleration
5. Jerk
6. Horizontal velocity/acceleration/jerk
7. Vertical velocity/acceleration/jerk
8. Number of changes in velocity direction
9. Number of changes in acceleration direction
10. Relative NCV
11. Relative NCA
12. in-air time
13. on-surface time
14. normalized in-air time
15. normalized on-surface time
16. in-air/on-surface ratio
## Feature Extraction
```
header_row=["X", "Y", "Z", "Pressure" , "GripAngle" , "Timestamp" , "Test_ID"]
#@timeit
def get_no_strokes(df):
pressure_data=df['Pressure'].to_numpy()
on_surface = (pressure_data>600).astype(int)
return ((np.roll(on_surface, 1) - on_surface) != 0).astype(int).sum()
#@timeit
def get_speed(df):
total_dist=0
duration=df['Timestamp'].to_numpy()[-1]
coords=df[['X', 'Y', 'Z']].to_numpy()
for i in range(10, df.shape[0]):
temp=np.linalg.norm(coords[i, :]-coords[i-10, :])
total_dist+=temp
speed=total_dist/duration
return speed
#@timeit
def get_in_air_time(data):
data=data['Pressure'].to_numpy()
return (data<600).astype(int).sum()
#@timeit
def get_on_surface_time(data):
data=data['Pressure'].to_numpy()
return (data>600).astype(int).sum()
def find_velocity(f):
'''
change in direction and its position
'''
data_pat=f
Vel = []
horz_Vel = []
horz_vel_mag = []
vert_vel_mag = []
vert_Vel = []
magnitude = []
timestamp_diff = []
t = 0
for i in range(len(data_pat)-2):
if t+10 <= len(data_pat)-1:
Vel.append(((data_pat['X'].to_numpy()[t+10] - data_pat['X'].to_numpy()[t])/(data_pat['Timestamp'].to_numpy()[t+10]-data_pat['Timestamp'].to_numpy()[t]) , (data_pat['Y'].to_numpy()[t+10]-data_pat['Y'].to_numpy()[t])/(data_pat['Timestamp'].to_numpy()[t+10]-data_pat['Timestamp'].to_numpy()[t])))
horz_Vel.append((data_pat['X'].to_numpy()[t+10] - data_pat['X'].to_numpy()[t])/(data_pat['Timestamp'].to_numpy()[t+10]-data_pat['Timestamp'].to_numpy()[t]))
vert_Vel.append((data_pat['Y'].to_numpy()[t+10] - data_pat['Y'].to_numpy()[t])/(data_pat['Timestamp'].to_numpy()[t+10]-data_pat['Timestamp'].to_numpy()[t]))
magnitude.append(sqrt(((data_pat['X'].to_numpy()[t+10]-data_pat['X'].to_numpy()[t])/(data_pat['Timestamp'].to_numpy()[t+10]-data_pat['Timestamp'].to_numpy()[t]))**2 + (((data_pat['Y'].to_numpy()[t+10]-data_pat['Y'].to_numpy()[t])/(data_pat['Timestamp'].to_numpy()[t+10]-data_pat['Timestamp'].to_numpy()[t]))**2)))
timestamp_diff.append(data_pat['Timestamp'].to_numpy()[t+10]-data_pat['Timestamp'].to_numpy()[t])
horz_vel_mag.append(abs(horz_Vel[len(horz_Vel)-1]))
vert_vel_mag.append(abs(vert_Vel[len(vert_Vel)-1]))
t = t+10
else:
break
magnitude_vel = np.mean(magnitude)
magnitude_horz_vel = np.mean(horz_vel_mag)
magnitude_vert_vel = np.mean(vert_vel_mag)
return Vel,magnitude,timestamp_diff,horz_Vel,vert_Vel,magnitude_vel,magnitude_horz_vel,magnitude_vert_vel
def find_acceleration(f):
'''
change in direction and its velocity
'''
Vel,magnitude,timestamp_diff,horz_Vel,vert_Vel,magnitude_vel,magnitude_horz_vel,magnitude_vert_vel = find_velocity(f)
accl = []
horz_Accl = []
vert_Accl = []
magnitude = []
horz_acc_mag = []
vert_acc_mag = []
for i in range(len(Vel)-2):
accl.append(((Vel[i+1][0]-Vel[i][0])/timestamp_diff[i] , (Vel[i+1][1]-Vel[i][1])/timestamp_diff[i]))
horz_Accl.append((horz_Vel[i+1]-horz_Vel[i])/timestamp_diff[i])
vert_Accl.append((vert_Vel[i+1]-vert_Vel[i])/timestamp_diff[i])
horz_acc_mag.append(abs(horz_Accl[len(horz_Accl)-1]))
vert_acc_mag.append(abs(vert_Accl[len(vert_Accl)-1]))
magnitude.append(sqrt(((Vel[i+1][0]-Vel[i][0])/timestamp_diff[i])**2 + ((Vel[i+1][1]-Vel[i][1])/timestamp_diff[i])**2))
magnitude_acc = np.mean(magnitude)
magnitude_horz_acc = np.mean(horz_acc_mag)
magnitude_vert_acc = np.mean(vert_acc_mag)
return accl,magnitude,horz_Accl,vert_Accl,timestamp_diff,magnitude_acc,magnitude_horz_acc,magnitude_vert_acc
def find_jerk(f):
accl,magnitude,horz_Accl,vert_Accl,timestamp_diff,magnitude_acc,magnitude_horz_acc,magnitude_vert_acc = find_acceleration(f)
jerk = []
hrz_jerk = []
vert_jerk = []
magnitude = []
horz_jerk_mag = []
vert_jerk_mag = []
for i in range(len(accl)-2):
jerk.append(((accl[i+1][0]-accl[i][0])/timestamp_diff[i] , (accl[i+1][1]-accl[i][1])/timestamp_diff[i]))
hrz_jerk.append((horz_Accl[i+1]-horz_Accl[i])/timestamp_diff[i])
vert_jerk.append((vert_Accl[i+1]-vert_Accl[i])/timestamp_diff[i])
horz_jerk_mag.append(abs(hrz_jerk[len(hrz_jerk)-1]))
vert_jerk_mag.append(abs(vert_jerk[len(vert_jerk)-1]))
magnitude.append(sqrt(((accl[i+1][0]-accl[i][0])/timestamp_diff[i])**2 + ((accl[i+1][1]-accl[i][1])/timestamp_diff[i])**2))
magnitude_jerk = np.mean(magnitude)
magnitude_horz_jerk = np.mean(horz_jerk_mag)
magnitude_vert_jerk = np.mean(vert_jerk_mag)
return jerk,magnitude,hrz_jerk,vert_jerk,timestamp_diff,magnitude_jerk,magnitude_horz_jerk,magnitude_vert_jerk
def NCV_per_halfcircle(f):
data_pat=f
Vel = []
ncv = []
temp_ncv = 0
basex = data_pat['X'].to_numpy()[0]
for i in range(len(data_pat)-2):
if data_pat['X'].to_numpy()[i] == basex:
ncv.append(temp_ncv)
temp_ncv = 0
continue
Vel.append(((data_pat['X'].to_numpy()[i+1] - data_pat['X'].to_numpy()[i])/(data_pat['Timestamp'].to_numpy()[i+1]-data_pat['Timestamp'].to_numpy()[i]) , (data_pat['Y'].to_numpy()[i+1]-data_pat['Y'].to_numpy()[i])/(data_pat['Timestamp'].to_numpy()[i+1]-data_pat['Timestamp'].to_numpy()[i])))
if Vel[len(Vel)-1] != (0,0):
temp_ncv+=1
ncv.append(temp_ncv)
#ncv = list(filter((2).__ne__, ncv))
ncv_Val = np.sum(ncv)/np.count_nonzero(ncv)
return ncv,ncv_Val
def NCA_per_halfcircle(f):
data_pat=f
Vel,magnitude,timestamp_diff,horz_Vel,vert_Vel,magnitude_vel,magnitude_horz_vel,magnitude_vert_vel = find_velocity(f)
accl = []
nca = []
temp_nca = 0
basex = data_pat['X'].to_numpy()[0]
for i in range(len(Vel)-2):
if data_pat['X'].to_numpy()[i] == basex:
nca.append(temp_nca)
#print ('tempNCa::',temp_nca)
temp_nca = 0
continue
accl.append(((Vel[i+1][0]-Vel[i][0])/timestamp_diff[i] , (Vel[i+1][1]-Vel[i][1])/timestamp_diff[i]))
if accl[len(accl)-1] != (0,0):
temp_nca+=1
nca.append(temp_nca)
nca = list(filter((2).__ne__, nca))
nca_Val = np.sum(nca)/np.count_nonzero(nca)
return nca,nca_Val
#@timeit
def get_features(f, parkinson_target):
global header_row
df=pd.read_csv(f, sep=';', header=None, names=header_row)
df_static=df[df["Test_ID"]==0] # static test
df_dynamic=df[df["Test_ID"]==1] # dynamic test
df_stcp=df[df["Test_ID"]==2] # STCP
#df_static_dynamic=pd.concat([df_static, df_dynamic])
initial_timestamp=df['Timestamp'][0]
df['Timestamp']=df['Timestamp']- initial_timestamp # offset timestamps
duration_static = df_static['Timestamp'].to_numpy()[-1] if df_static.shape[0] else 1
duration_dynamic = df_dynamic['Timestamp'].to_numpy()[-1] if df_dynamic.shape[0] else 1
duration_STCP = df_stcp['Timestamp'].to_numpy()[-1] if df_stcp.shape[0] else 1
data_point=[]
data_point.append(get_no_strokes(df_static) if df_static.shape[0] else 0) # no. of strokes for static test
data_point.append(get_no_strokes(df_dynamic) if df_dynamic.shape[0] else 0) # no. of strokes for dynamic test
data_point.append(get_speed(df_static) if df_static.shape[0] else 0) # speed for static test
data_point.append(get_speed(df_dynamic) if df_dynamic.shape[0] else 0) # speed for dynamic test
Vel,magnitude,timestamp_diff,horz_Vel,vert_Vel,magnitude_vel,magnitude_horz_vel,magnitude_vert_vel = find_velocity(df_static) if df_static.shape[0] else (0,0,0,0,0,0,0,0) # magnitudes of velocity, horizontal velocity and vertical velocity for static test
data_point.extend([magnitude_vel, magnitude_horz_vel,magnitude_vert_vel])
Vel,magnitude,timestamp_diff,horz_Vel,vert_Vel,magnitude_vel,magnitude_horz_vel,magnitude_vert_vel = find_velocity(df_dynamic) if df_dynamic.shape[0] else (0,0,0,0,0,0,0,0) # magnitudes of velocity, horizontal velocity and vertical velocity for dynamic test
data_point.extend([magnitude_vel, magnitude_horz_vel,magnitude_vert_vel])
accl,magnitude,horz_Accl,vert_Accl,timestamp_diff,magnitude_acc,magnitude_horz_acc,magnitude_vert_acc=find_acceleration(df_static) if df_static.shape[0] else (0,0,0,0,0,0,0,0)# magnitudes of acceleration, horizontal acceleration and vertical acceleration for static test
data_point.extend([magnitude_acc,magnitude_horz_acc,magnitude_vert_acc])
accl,magnitude,horz_Accl,vert_Accl,timestamp_diff,magnitude_acc,magnitude_horz_acc,magnitude_vert_acc=find_acceleration(df_dynamic) if df_dynamic.shape[0] else (0,0,0,0,0,0,0,0)# magnitudes of acceleration, horizontal acceleration and vertical acceleration for dynamic test
data_point.extend([magnitude_acc,magnitude_horz_acc,magnitude_vert_acc])
jerk,magnitude,hrz_jerk,vert_jerk,timestamp_diff,magnitude_jerk,magnitude_horz_jerk,magnitude_vert_jerk=find_jerk(df_static) if df_static.shape[0] else (0,0,0,0,0,0,0,0) # magnitudes of jerk, horizontal jerk and vertical jerk for static test
data_point.extend([magnitude_jerk,magnitude_horz_jerk,magnitude_vert_jerk])
jerk,magnitude,hrz_jerk,vert_jerk,timestamp_diff,magnitude_jerk,magnitude_horz_jerk,magnitude_vert_jerk=find_jerk(df_dynamic) if df_dynamic.shape[0] else (0,0,0,0,0,0,0,0) # magnitudes of jerk, horizontal jerk and vertical jerk for dynamic test
data_point.extend([magnitude_jerk,magnitude_horz_jerk,magnitude_vert_jerk])
ncv,ncv_Val=NCV_per_halfcircle(df_static) if df_static.shape[0] else (0,0) # NCV for static test
data_point.append(ncv_Val)
ncv,ncv_Val=NCV_per_halfcircle(df_dynamic) if df_dynamic.shape[0] else (0,0) # NCV for dynamic test
data_point.append(ncv_Val)
nca,nca_Val=NCA_per_halfcircle(df_static) if df_static.shape[0] else (0,0) # NCA for static test
data_point.append(nca_Val)
nca,nca_Val=NCA_per_halfcircle(df_dynamic) if df_dynamic.shape[0] else (0,0) # NCA for dynamic test
data_point.append(nca_Val)
data_point.append(get_in_air_time(df_stcp) if df_stcp.shape[0] else 0) # in air time for STCP
data_point.append(get_on_surface_time(df_static) if df_static.shape[0] else 0) # on surface time for static test
data_point.append(get_on_surface_time(df_dynamic) if df_dynamic.shape[0] else 0) # on surface time for dynamic test
data_point.append(parkinson_target) # traget. 1 for parkinson. 0 for control.
return data_point
temp_feat=get_features(parkinson_file_list[35], 1)
print(temp_feat)
raw=[]
for x in parkinson_file_list:
raw.append(get_features(x, 1))
for x in control_file_list:
raw.append(get_features(x, 0))
raw=np.array(raw)
features_headers=['no_strokes_st', 'no_strokes_dy', 'speed_st', 'speed_dy', 'magnitude_vel_st' , 'magnitude_horz_vel_st' , 'magnitude_vert_vel_st', 'magnitude_vel_dy', 'magnitude_horz_vel_dy' , 'magnitude_vert_vel_dy', 'magnitude_acc_st' , 'magnitude_horz_acc_st' , 'magnitude_vert_acc_st','magnitude_acc_dy' , 'magnitude_horz_acc_dy' , 'magnitude_vert_acc_dy', 'magnitude_jerk_st', 'magnitude_horz_jerk_st' , 'magnitude_vert_jerk_st', 'magnitude_jerk_dy', 'magnitude_horz_jerk_dy' , 'magnitude_vert_jerk_dy', 'ncv_st', 'ncv_dy', 'nca_st', 'nca_dy', 'in_air_stcp','on_surface_st', 'on_surface_dy', 'target']
data=pd.DataFrame(raw, columns=features_headers)
data.tail()
data.to_csv('data.csv')
print('data shape', data.shape)
```
## Classification
```
pos=data[data['target']==1]
neg=data[data['target']==0]
train_pos=pos.head(pos.shape[0]-5)
train_neg=neg.head(pos.shape[0]-5)
train=pd.concat([train_pos, train_neg])
print('train shape', train.shape)
test_pos=pos.tail(5)
test_neg=neg.tail(5)
test=pd.concat([test_pos, test_neg])
train_y=train['target']
train_x=train.drop(['target'], axis=1)
test_y=test['target']
test_x=test.drop(['target'], axis=1)
def accuracy(prediction,actual):
correct = 0
not_correct = 0
for i in range(len(prediction)):
if prediction[i] == actual[i]:
correct+=1
else:
not_correct+=1
return (correct*100)/(correct+not_correct)
def metrics(prediction,actual):
tp = 0
tn = 0
fp = 0
fn = 0
for i in range(len(prediction)):
if prediction[i] == actual[i] and actual[i]==1:
tp+=1
if prediction[i] == actual[i] and actual[i]==0:
tn+=1
if prediction[i] != actual[i] and actual[i]==0:
fp+=1
if prediction[i] != actual[i] and actual[i]==1:
fn+=1
metrics = {'Precision':(tp/(tp+fp+tn+fn)),'Recall':(tp/(tp+fn)),'F1':(2*(tp/(tp+fp+tn+fn))*(tp/(tp+fn)))/((tp/(tp+fp+tn+fn))+(tp/(tp+fn)))}
return (metrics)
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
```
### Logistic Regression
```
clf=LogisticRegression()
clf.fit(train_x, train_y)
preds=clf.predict(test_x)
print('accuracy:',accuracy(test_y.tolist(), preds.tolist()), '%')
print(metrics(test_y.tolist(), preds.tolist()))
```
### Random Forest
```
clf=RandomForestClassifier()
clf.fit(train_x, train_y)
preds=clf.predict(test_x)
print('accuracy:',accuracy(test_y.tolist(), preds.tolist()), '%')
print(metrics(test_y.tolist(), preds.tolist()))
```
### Support Vector Machine
```
clf=SVC()
clf.fit(train_x, train_y)
preds=clf.predict(test_x)
print('accuracy:',accuracy(test_y.tolist(), preds.tolist()), '%')
print(metrics(test_y.tolist(), preds.tolist()))
```
### Decision Tree
```
clf=DecisionTreeClassifier()
clf.fit(train_x, train_y)
preds=clf.predict(test_x)
print('accuracy:',accuracy(test_y.tolist(), preds.tolist()), '%')
print(metrics(test_y.tolist(), preds.tolist()))
```
### K-Nearest Neighbors
```
clf=KNeighborsClassifier()
clf.fit(train_x, train_y)
preds=clf.predict(test_x)
print('accuracy:',accuracy(test_y.tolist(), preds.tolist()), '%')
print(metrics(test_y.tolist(), preds.tolist()))
```
## Some Plots
```
def plot(f, plot_func, t_id=0, x=None, y=None, reg_plot=True):
global header_row
df=pd.read_csv(f, sep=';', header=None, names=header_row)
df=df[df["Test_ID"]==t_id]
initial_timestamp=df['Timestamp'][0]
df['Timestamp']=df['Timestamp']- initial_timestamp
if reg_plot:
plot_func(data=df, x=x, y=y, fit_reg=False, scatter_kws={"s": 0.5})
else:
plot_func(data=df, x=x, y=y)
print(metrics(test_y.tolist(), preds.tolist()))
```
#### Pressure (Parkinsons)
```
plot(f=parkinson_file_list[1], plot_func=sns.regplot, t_id=0, x='Timestamp', y='Pressure')
```
#### Pressure (Control)
```
plot(control_file_list[2], plot_func=sns.regplot, t_id=0, x='Timestamp', y='Pressure')
plot(f=parkinson_file_list[35], plot_func=sns.barplot, t_id=0, x='Timestamp', y='Pressure', reg_plot=False)
```
| github_jupyter |
```
import re
m = re.search(r'(?<=-)\w+', 'spam-egg')
m.group(0)
re.sub(r'def\s+([a-zA-Z_][a-zA-Z_0-9]*)\s*\(\s*\):',
r'static PyObject*\npy_\1(void)\n{',
'def myfunc():')
class GenProp(object):
def __init__(self, prop_type, prop_name):
self.prop_type=prop_type
self.prop_name=prop_name
class GenCollection(object):
def __init__(self, collection_type, element_type, prop_name):
self.prop_name=prop_name
self.collection_type=collection_type
self.element_type=element_type
class GenMeta(object):
def __init__(self):
self.class_name=''
self.base_name=''
self.attributes=[]
code="""class Schedule extends Equatable {
DateTime fromDate= DateTime.now();
TimeOfDay fromTime = const TimeOfDay(hour: 7, minute: 28);
DateTime toDate = DateTime.now();
TimeOfDay toTime = const TimeOfDay(hour: 7, minute: 28);
// final List<String> allActivities = <String>['hiking', 'swimming', 'boating', 'fishing'];
List<String> allActivities= <String>['hiking', 'swimming'];
String activity = 'fishing';
"""
meta=GenMeta()
for line in code.splitlines():
# print(line)
match_obj=re.match(r'class (.*) extends (.*) {', line, re.M|re.I)
if match_obj:
print("@class", match_obj.group(1),"*", match_obj.group(2))
meta.class_name=match_obj.group(1)
meta.base_name=match_obj.group(2)
else:
pass
match_obj=re.match(r'\s*([a-zA-Z_][a-zA-Z_0-9]*)\s*([a-zA-Z_][a-zA-Z_0-9]*)\s*=(.*);', line, re.M|re.I)
# match_obj=re.search(r'([a-zA-Z_]+) ([a-zA-Z_]+) = ([a-zA-Z_]+);', line)
if match_obj:
print("@prop", match_obj.group(1),'*', match_obj.group(2))
meta.attributes.append(GenProp(match_obj.group(1), match_obj.group(2)))
else:
pass
# process collections
match_obj=re.match(r'\s*([a-zA-Z_][a-zA-Z_0-9]*)<(.*)>\s*([a-zA-Z_][a-zA-Z_0-9]*)\s*=(.*);', line, re.M|re.I)
if match_obj:
print("@collection", match_obj.group(1),"-", match_obj.group(2), match_obj.group(3))
meta.attributes.append(GenCollection(match_obj.group(1),match_obj.group(2), match_obj.group(3)))
else:
pass
print(meta)
import json
# json.dumps(meta.__dict__)
from json import JSONEncoder
class MyEncoder(JSONEncoder):
def default(self, o):
return o.__dict__
# data=(MyEncoder().encode(meta))
# print(data)
jd=json.dumps(meta, cls=MyEncoder, indent=2)
print(jd)
def gen_constructor(meta):
lines=[]
lines.append(' %s({'%meta.class_name)
attr_defs=[]
attr_equals=[]
for attr in meta.attributes:
attr_defs.append('this.%s'%attr.prop_name)
attr_equals.append(attr.prop_name)
lines.append(' '+', '.join(attr_defs))
lines.append(' }) : super([')
lines.append(' '+', '.join(attr_equals))
lines.append(' ]);')
print('\n'.join(lines))
gen_constructor(meta)
lines=[]
lines.append(' static %s overrides%s(Map<String, dynamic> map) {'%(meta.class_name, meta.class_name))
lines.append(' return %s('%meta.class_name)
setters=[]
for attr in meta.attributes:
setters.append(" %s: map['%s']"%(attr.prop_name, attr.prop_name))
lines.append(",\n".join(setters))
lines.append(' }')
print('\n'.join(lines))
lines=[]
lines.append(' Map<String, dynamic> asMap() {')
setters=[]
for attr in meta.attributes:
setters.append("'%s':%s"%(attr.prop_name, attr.prop_name))
lines.append(" return {%s};"%(', '.join(setters)))
lines.append(' }')
print('\n'.join(lines))
```
| github_jupyter |
# 先导入 Torch 版本的模型
```
# 把 ChineseBert 源码的根目录,加入到 python sys.path
import sys
sys.path.append('/data1/workspace/research/ChineseBERT-Paddle/ChineseBert')
# 把 clone paddlenlp 源码,加入到 python sys.path
import sys
sys.path.append('/data1/workspace/research/ChineseBERT-Paddle/Paddle_ChineseBert/PaddleNLP')
# ChineseBert Model
from datasets.bert_dataset import BertDataset
from models.modeling_glycebert import GlyceBertModel
CHINESEBERT_PATH='./pretrain_models/torch/ChineseBERT-base/'
tokenizer = BertDataset(CHINESEBERT_PATH)
chinese_bert = GlyceBertModel.from_pretrained(CHINESEBERT_PATH)
config = chinese_bert.config
sentence = '我喜欢猫'
input_ids, pinyin_ids = tokenizer.tokenize_sentence(sentence)
length = input_ids.shape[0]
input_ids = input_ids.view(1, length)
pinyin_ids = pinyin_ids.view(1, length, 8)
output_hidden = chinese_bert.forward(input_ids, pinyin_ids)[0]
print(output_hidden.shape)
```
## 对比 ChineseBert 和 Bert 的网络参数
```
from transformers import BertModel, AutoModelForMaskedLM
torch_chinese_bert_keys = chinese_bert.state_dict().keys()
torch_bert_keys = BertModel.from_pretrained("bert-base-chinese").state_dict().keys()
# 全部的 ChineseBert 的参数
list(torch_chinese_bert_keys)
# ChineseBert 多出来的参数(embedding 层)
set(list(torch_chinese_bert_keys)) - set(list(torch_bert_keys))
# ChineseBert 少的参数
set(list(torch_bert_keys)) - set(list(torch_chinese_bert_keys))
```
## 模型参数 torch 转换 paddle
```
import paddle
import torch
import numpy as np
# ChineseBERT-base: ./pretrain_models/torch/ChineseBERT-base/pytorch_model.bin
# ChineseBERT-large: ./pretrain_models/torch/ChineseBERT-large/pytorch_model.bin
torch_model_path = "./pretrain_models/torch/ChineseBERT-large/pytorch_model.bin"
torch_state_dict = torch.load(torch_model_path)
paddle_model_path = "./pretrain_models/paddle/ChineseBERT-large/model_state.pdparams"
paddle_state_dict = {}
# State_dict's keys mapping: from torch to paddle
keys_dict = {
# about encoder layer
'encoder.layer': 'encoder.layers',
'attention.self.query': 'self_attn.q_proj', # 需要转置
'attention.self.key': 'self_attn.k_proj', # 需要转置
'attention.self.value': 'self_attn.v_proj', # 需要转置
'attention.output.dense': 'self_attn.out_proj', # 需要转置
'attention.output.LayerNorm': 'norm1', # 需要转置
'intermediate.dense': 'linear1', # 需要转置
'output.dense': 'linear2', # 需要转置
'output.LayerNorm': 'norm2', # 需要转置
}
for torch_key in torch_state_dict:
paddle_key = torch_key
for k in keys_dict:
if k in paddle_key:
paddle_key = paddle_key.replace(k, keys_dict[k])
if ('map_fc' in paddle_key) or ('glyph_map' in paddle_key) or ('linear' in paddle_key) or ('proj' in paddle_key) or ('vocab' in paddle_key and 'weight' in paddle_key) or ("dense.weight" in paddle_key) or ('transform.weight' in paddle_key) or ('seq_relationship.weight' in paddle_key):
print("transpose(permute) ---------->")
paddle_state_dict[paddle_key] = paddle.to_tensor(torch_state_dict[torch_key].cpu().numpy().transpose())
else:
paddle_state_dict[paddle_key] = paddle.to_tensor(torch_state_dict[torch_key].cpu().numpy())
print("t: ", torch_key,"\t", torch_state_dict[torch_key].shape)
print("p: ", paddle_key, "\t", paddle_state_dict[paddle_key].shape, "\n")
paddle.save(paddle_state_dict, paddle_model_path)
```
# 使用 Paddle 转写 forward
* 我们按照源码的组成部分分批转写
## Tokenizer
### 对齐 Bert tokenizer
```
# =========================================
# 对齐 paddle 和 torch 的 bert tokenizer
# =========================================
from tokenizers import BertWordPieceTokenizer
from paddlenlp.transformers import BertTokenizer
torch_token = BertWordPieceTokenizer('./pretrain_models/torch/ChineseBERT-base/vocab.txt')
berttokenizer = BertTokenizer('./pretrain_models/torch/ChineseBERT-base/vocab.txt')
sentence="我喜欢猫"
# torch
print("============ PyTorch ==================")
bert_tokens = torch_token.encode(sentence)
print(bert_tokens.ids)
print(bert_tokens.tokens)
print(bert_tokens.offsets)
print("============= Paddle =================")
# paddle
# ids
p_bert_tokens = berttokenizer.encode(sentence)
p_bert_tokens_ids = p_bert_tokens['input_ids']
print(p_bert_tokens_ids)
# tokens
p_bert_tokens_tokens = berttokenizer.tokenize(sentence)
p_bert_tokens_tokens.insert(0, '[CLS]')
p_bert_tokens_tokens.append('[SEP]')
print(p_bert_tokens_tokens)
# offsets
p_bert_tokens_offsets = berttokenizer.get_offset_mapping(sentence)
p_bert_tokens_offsets.insert(0, (0, 0))
p_bert_tokens_offsets.append((0, 0))
print(p_bert_tokens_offsets)
bert_tokens = torch_token.encode(sentence)
bert_tokens, bert_tokens.ids
bert_tokens_t = torch_token.encode(sentence, add_special_tokens=False)
bert_tokens_t, bert_tokens_t.ids, bert_tokens_t.tokens, bert_tokens_t.offsets
```
### 对齐 ChineseBert tokenizer
```
# =================================================
# PyTorch 的 ChineseBert BertDataset (tokenizer)
# =================================================
import json
import os
from typing import List
import tokenizers
import torch
from pypinyin import pinyin, Style
from tokenizers import BertWordPieceTokenizer
class BertDataset(object):
def __init__(self, bert_path, max_length: int = 512):
super().__init__()
vocab_file = os.path.join(bert_path, 'vocab.txt')
config_path = os.path.join(bert_path, 'config')
self.max_length = max_length
self.tokenizer = BertWordPieceTokenizer(vocab_file)
# load pinyin map dict
with open(os.path.join(config_path, 'pinyin_map.json'), encoding='utf8') as fin:
self.pinyin_dict = json.load(fin)
# load char id map tensor
with open(os.path.join(config_path, 'id2pinyin.json'), encoding='utf8') as fin:
self.id2pinyin = json.load(fin)
# load pinyin map tensor
with open(os.path.join(config_path, 'pinyin2tensor.json'), encoding='utf8') as fin:
self.pinyin2tensor = json.load(fin)
def tokenize_sentence(self, sentence):
# convert sentence to ids
tokenizer_output = self.tokenizer.encode(sentence)
bert_tokens = tokenizer_output.ids
pinyin_tokens = self.convert_sentence_to_pinyin_ids(sentence, tokenizer_output)
# assert,token nums should be same as pinyin token nums
assert len(bert_tokens) <= self.max_length
assert len(bert_tokens) == len(pinyin_tokens)
# convert list to tensor
input_ids = torch.LongTensor(bert_tokens)
pinyin_ids = torch.LongTensor(pinyin_tokens).view(-1)
return input_ids, pinyin_ids
def convert_sentence_to_pinyin_ids(self, sentence: str, tokenizer_output: tokenizers.Encoding) -> List[List[int]]:
# get pinyin of a sentence
pinyin_list = pinyin(sentence, style=Style.TONE3, heteronym=True, errors=lambda x: [['not chinese'] for _ in x])
pinyin_locs = {}
# get pinyin of each location
for index, item in enumerate(pinyin_list):
pinyin_string = item[0]
# not a Chinese character, pass
if pinyin_string == "not chinese":
continue
if pinyin_string in self.pinyin2tensor:
pinyin_locs[index] = self.pinyin2tensor[pinyin_string]
else:
ids = [0] * 8
for i, p in enumerate(pinyin_string):
if p not in self.pinyin_dict["char2idx"]:
ids = [0] * 8
break
ids[i] = self.pinyin_dict["char2idx"][p]
pinyin_locs[index] = ids
# find chinese character location, and generate pinyin ids
pinyin_ids = []
for idx, (token, offset) in enumerate(zip(tokenizer_output.tokens, tokenizer_output.offsets)):
if offset[1] - offset[0] != 1:
pinyin_ids.append([0] * 8)
continue
if offset[0] in pinyin_locs:
pinyin_ids.append(pinyin_locs[offset[0]])
else:
pinyin_ids.append([0] * 8)
return pinyin_ids
# =================================================
# Paddle 的 ChineseBert BertDataset (tokenizer)
# =================================================
import json
import os
from typing import List
from pypinyin import pinyin, Style
import paddle
from paddlenlp.transformers import BertTokenizer
class PaddleBertDataset(object):
def __init__(self, bert_path, max_length: int = 512):
super().__init__()
vocab_file = os.path.join(bert_path, 'vocab.txt')
config_path = os.path.join(bert_path, 'config')
self.max_length = max_length
self.tokenizer = BertTokenizer(vocab_file)
# load pinyin map dict
with open(os.path.join(config_path, 'pinyin_map.json'), encoding='utf8') as fin:
self.pinyin_dict = json.load(fin)
# load char id map tensor
with open(os.path.join(config_path, 'id2pinyin.json'), encoding='utf8') as fin:
self.id2pinyin = json.load(fin)
# load pinyin map tensor
with open(os.path.join(config_path, 'pinyin2tensor.json'), encoding='utf8') as fin:
self.pinyin2tensor = json.load(fin)
def tokenize_sentence(self, sentence):
# convert sentence to ids
tokenizer_output = self.tokenizer.encode(sentence)
bert_tokens = tokenizer_output['input_ids']
pinyin_tokens = self.convert_sentence_to_pinyin_ids(sentence)
# assert,token nums should be same as pinyin token nums
assert len(bert_tokens) <= self.max_length
assert len(bert_tokens) == len(pinyin_tokens)
# convert list to tensor
input_ids = paddle.to_tensor(bert_tokens)
pinyin_ids = paddle.to_tensor(pinyin_tokens).reshape([-1])
return input_ids, pinyin_ids
def convert_sentence_to_pinyin_ids(self, sentence: str) -> List[List[int]]:
# get offsets
bert_tokens_offsets = self.tokenizer.get_offset_mapping(sentence)
bert_tokens_offsets.insert(0, (0, 0))
bert_tokens_offsets.append((0, 0))
# get tokens
bert_tokens_tokens = self.tokenizer.tokenize(sentence)
bert_tokens_tokens.insert(0, '[CLS]')
bert_tokens_tokens.append('[SEP]')
# get pinyin of a sentence
pinyin_list = pinyin(sentence, style=Style.TONE3, heteronym=True, errors=lambda x: [['not chinese'] for _ in x])
pinyin_locs = {}
# get pinyin of each location
for index, item in enumerate(pinyin_list):
pinyin_string = item[0]
# not a Chinese character, pass
if pinyin_string == "not chinese":
continue
if pinyin_string in self.pinyin2tensor:
pinyin_locs[index] = self.pinyin2tensor[pinyin_string]
else:
ids = [0] * 8
for i, p in enumerate(pinyin_string):
if p not in self.pinyin_dict["char2idx"]:
ids = [0] * 8
break
ids[i] = self.pinyin_dict["char2idx"][p]
pinyin_locs[index] = ids
# find chinese character location, and generate pinyin ids
pinyin_ids = []
for idx, (token, offset) in enumerate(zip(bert_tokens_tokens, bert_tokens_offsets)):
if offset[1] - offset[0] != 1:
pinyin_ids.append([0] * 8)
continue
if offset[0] in pinyin_locs:
pinyin_ids.append(pinyin_locs[offset[0]])
else:
pinyin_ids.append([0] * 8)
return pinyin_ids
# torch
sentence = '我喜欢猫'
tokenizer = BertDataset(CHINESEBERT_PATH)
input_ids, pinyin_ids = tokenizer.tokenize_sentence(sentence)
print("============================== torch =============================")
print(input_ids.cpu().detach().numpy(), pinyin_ids.cpu().detach().numpy())
print()
# paddle
sentence = '我喜欢猫'
paddle_tokenizer = PaddleBertDataset(CHINESEBERT_PATH)
paddle_input_ids, paddle_pinyin_ids = paddle_tokenizer.tokenize_sentence(sentence)
print("============================== paddle =============================")
print(paddle_input_ids.cpu().detach().numpy(), paddle_pinyin_ids.cpu().detach().numpy())
```
## PinyinEmbedding
```
# Torch PinyinEmbedding
import json
import os
from torch import nn
from torch.nn import functional as F
class PinyinEmbedding(nn.Module):
def __init__(self, embedding_size: int, pinyin_out_dim: int, config_path):
"""
Pinyin Embedding Module
Args:
embedding_size: the size of each embedding vector
pinyin_out_dim: kernel number of conv
"""
super(PinyinEmbedding, self).__init__()
with open(os.path.join(config_path, 'pinyin_map.json')) as fin:
pinyin_dict = json.load(fin)
self.pinyin_out_dim = pinyin_out_dim
self.embedding = nn.Embedding(len(pinyin_dict['idx2char']), embedding_size)
self.conv = nn.Conv1d(in_channels=embedding_size,
out_channels=self.pinyin_out_dim,
kernel_size=2,
stride=1,
padding=0)
def forward(self, pinyin_ids):
"""
Args:
pinyin_ids: (bs*sentence_length*pinyin_locs)
Returns:
pinyin_embed: (bs,sentence_length,pinyin_out_dim)
"""
# input pinyin ids for 1-D conv
embed = self.embedding(pinyin_ids) # [bs,sentence_length,pinyin_locs,embed_size]
bs, sentence_length, pinyin_locs, embed_size = embed.shape
view_embed = embed.view(-1, pinyin_locs, embed_size) # [(bs*sentence_length),pinyin_locs,embed_size]
input_embed = view_embed.permute(0, 2, 1) # [(bs*sentence_length), embed_size, pinyin_locs]
# conv + max_pooling
pinyin_conv = self.conv(input_embed) # [(bs*sentence_length),pinyin_out_dim,H]
pinyin_embed = F.max_pool1d(pinyin_conv, pinyin_conv.shape[-1]) # [(bs*sentence_length),pinyin_out_dim,1]
return pinyin_embed.view(bs, sentence_length, self.pinyin_out_dim) # [bs,sentence_length,pinyin_out_dim]
# Paddle PinyinEmbedding
import json
import os
import paddle
class PaddlePinyinEmbedding(paddle.nn.Layer):
def __init__(self, embedding_size: int, pinyin_out_dim: int, config_path):
"""
Pinyin Embedding Module
Args:
embedding_size: the size of each embedding vector
pinyin_out_dim: kernel number of conv
"""
super(PaddlePinyinEmbedding, self).__init__()
with open(os.path.join(config_path, 'pinyin_map.json')) as fin:
pinyin_dict = json.load(fin)
self.pinyin_out_dim = pinyin_out_dim
self.embedding = paddle.nn.Embedding(len(pinyin_dict['idx2char']), embedding_size)
self.conv = paddle.nn.Conv1D(in_channels=embedding_size,
out_channels=self.pinyin_out_dim,
kernel_size=2,
stride=1,
padding=0,
bias_attr=True)
def forward(self, pinyin_ids):
"""
Args:
pinyin_ids: (bs*sentence_length*pinyin_locs)
Returns:
pinyin_embed: (bs,sentence_length,pinyin_out_dim)
"""
# input pinyin ids for 1-D conv
embed = self.embedding(pinyin_ids) # [bs,sentence_length,pinyin_locs,embed_size]
bs, sentence_length, pinyin_locs, embed_size = embed.shape
view_embed = embed.reshape((-1, pinyin_locs, embed_size)) # [(bs*sentence_length),pinyin_locs,embed_size]
input_embed = view_embed.transpose([0, 2, 1]) # [(bs*sentence_length), embed_size, pinyin_locs]
# conv + max_pooling
pinyin_conv = self.conv(input_embed) # [(bs*sentence_length),pinyin_out_dim,H]
pinyin_embed = paddle.nn.functional.max_pool1d(pinyin_conv, pinyin_conv.shape[-1]) # [(bs*sentence_length),pinyin_out_dim,1]
return pinyin_embed.reshape((bs, sentence_length, self.pinyin_out_dim)) # [bs,sentence_length,pinyin_out_dim]
# torch
print("============================== torch =============================")
sentence = '我喜欢猫'
tokenizer = BertDataset(CHINESEBERT_PATH)
input_ids, pinyin_ids = tokenizer.tokenize_sentence(sentence)
length = input_ids.shape[0]
print(f"length: {length}")
print("torch size:", input_ids.size())
print(pinyin_ids)
pinyin_ids = pinyin_ids.view(1, length, 8)
print(pinyin_ids, pinyin_ids.shape)
hidden_size = 768
config_path='./pretrain_models/torch/ChineseBERT-base/config/'
pinyin_embeddings = PinyinEmbedding(embedding_size=128,
pinyin_out_dim=hidden_size,
config_path=config_path)
torch_pinyin_emb = pinyin_embeddings(pinyin_ids)
print(">>>>torch_pinyin_emb<<<<")
print(torch_pinyin_emb, torch_pinyin_emb.shape)
print()
# paddle
sentence = '我喜欢猫'
paddle_tokenizer = PaddleBertDataset(CHINESEBERT_PATH)
paddle_input_ids, paddle_pinyin_ids = paddle_tokenizer.tokenize_sentence(sentence)
print("============================== paddle =============================")
length = paddle_input_ids.shape[0]
print(f"length: {length}")
print("paddle size(shape)", paddle_input_ids.shape)
print(paddle_pinyin_ids)
paddle_pinyin_ids = paddle_pinyin_ids.reshape((1, length, 8))
print(paddle_pinyin_ids, paddle_pinyin_ids.shape)
hidden_size = 768
config_path='./pretrain_models/torch/ChineseBERT-base/config/'
paddle_pinyin_embeddings = PaddlePinyinEmbedding(embedding_size=128,
pinyin_out_dim=hidden_size,
config_path=config_path)
paddle_pinyin_emb = paddle_pinyin_embeddings(paddle_pinyin_ids)
print(">>>>paddle_pinyin_emb<<<<")
print(paddle_pinyin_emb, paddle_pinyin_emb.shape)
print(f"torch pinyin_embeddings: {pinyin_embeddings}")
print(f"paddle paddle_pinyin_embeddings: {paddle_pinyin_embeddings}")
```
## GlyphEmbedding
```
# torch
from typing import List
import numpy as np
import torch
from torch import nn
class GlyphEmbedding(nn.Module):
"""Glyph2Image Embedding"""
def __init__(self, font_npy_files: List[str]):
super(GlyphEmbedding, self).__init__()
font_arrays = [
np.load(np_file).astype(np.float32) for np_file in font_npy_files
]
self.vocab_size = font_arrays[0].shape[0]
self.font_num = len(font_arrays)
self.font_size = font_arrays[0].shape[-1]
# N, C, H, W
font_array = np.stack(font_arrays, axis=1)
self.embedding = nn.Embedding(
num_embeddings=self.vocab_size,
embedding_dim=self.font_size ** 2 * self.font_num,
_weight=torch.from_numpy(font_array.reshape([self.vocab_size, -1]))
)
def forward(self, input_ids):
"""
get glyph images for batch inputs
Args:
input_ids: [batch, sentence_length]
Returns:
images: [batch, sentence_length, self.font_num*self.font_size*self.font_size]
"""
# return self.embedding(input_ids).view([-1, self.font_num, self.font_size, self.font_size])
return self.embedding(input_ids)
# paddle
from typing import List
import numpy as np
import paddle
class PaddleGlyphEmbedding(paddle.nn.Layer):
"""Glyph2Image Embedding"""
def __init__(self, font_npy_files: List[str]):
super(PaddleGlyphEmbedding, self).__init__()
font_arrays = [
np.load(np_file).astype(np.float32) for np_file in font_npy_files
]
self.vocab_size = font_arrays[0].shape[0]
self.font_num = len(font_arrays)
self.font_size = font_arrays[0].shape[-1]
# N, C, H, W
font_array = np.stack(font_arrays, axis=1)
self.embedding = paddle.nn.Embedding(
num_embeddings=self.vocab_size,
embedding_dim=self.font_size ** 2 * self.font_num
)
self.embedding.weight.set_value(font_array.reshape([self.vocab_size, -1]))
def forward(self, input_ids):
"""
get glyph images for batch inputs
Args:
input_ids: [batch, sentence_length]
Returns:
images: [batch, sentence_length, self.font_num*self.font_size*self.font_size]
"""
# return self.embedding(input_ids).view([-1, self.font_num, self.font_size, self.font_size])
return self.embedding(input_ids)
config_path='./pretrain_models/torch/ChineseBERT-base/config/'
font_files = []
for file in os.listdir(config_path):
if file.endswith(".npy"):
font_files.append(os.path.join(config_path, file))
print(font_files)
# torch
print("============================== torch =============================")
glyph_embeddings = GlyphEmbedding(font_npy_files=font_files)
t_emb_w = glyph_embeddings.state_dict()['embedding.weight'].cpu().detach().numpy()
print(f"torch glyph_embeddings: {glyph_embeddings}")
# paddle
print("============================== paddle =============================")
paddle_glyph_embeddings = PaddleGlyphEmbedding(font_npy_files=font_files)
p_emb_w = paddle_glyph_embeddings.state_dict()['embedding.weight'].cpu().detach().numpy()
print(f"paddle paddle_glyph_embeddings: {paddle_glyph_embeddings}")
(p_emb_w == t_emb_w).all()
paddle_glyph_embeddings.state_dict().keys()
```
## FusionBertEmbeddings
```
# torch
import os
import torch
from torch import nn
class FusionBertEmbeddings(nn.Module):
"""
Construct the embeddings from word, position, glyph, pinyin and token_type embeddings.
"""
def __init__(self, config):
super(FusionBertEmbeddings, self).__init__()
config_path = os.path.join(config.name_or_path, 'config')
font_files = []
for file in os.listdir(config_path):
if file.endswith(".npy"):
font_files.append(os.path.join(config_path, file))
self.word_embeddings = nn.Embedding(config.vocab_size, config.hidden_size, padding_idx=0)
self.position_embeddings = nn.Embedding(config.max_position_embeddings, config.hidden_size)
self.token_type_embeddings = nn.Embedding(config.type_vocab_size, config.hidden_size)
self.pinyin_embeddings = PinyinEmbedding(embedding_size=128, pinyin_out_dim=config.hidden_size,
config_path=config_path)
self.glyph_embeddings = GlyphEmbedding(font_npy_files=font_files)
# self.LayerNorm is not snake-cased to stick with TensorFlow models variable name and be able to load
# any TensorFlow checkpoint file
self.glyph_map = nn.Linear(1728, config.hidden_size)
self.map_fc = nn.Linear(config.hidden_size * 3, config.hidden_size)
self.LayerNorm = nn.LayerNorm(config.hidden_size, eps=config.layer_norm_eps)
self.dropout = nn.Dropout(config.hidden_dropout_prob)
# position_ids (1, len position emb) is contiguous in memory and exported when serialized
self.register_buffer("position_ids", torch.arange(config.max_position_embeddings).expand((1, -1)))
def forward(self, input_ids=None, pinyin_ids=None, token_type_ids=None, position_ids=None, inputs_embeds=None):
if input_ids is not None:
input_shape = input_ids.size()
else:
input_shape = inputs_embeds.size()[:-1]
seq_length = input_shape[1]
if position_ids is None:
position_ids = self.position_ids[:, :seq_length]
if token_type_ids is None:
token_type_ids = torch.zeros(input_shape, dtype=torch.long, device=self.position_ids.device)
if inputs_embeds is None:
inputs_embeds = self.word_embeddings(input_ids)
# get char embedding, pinyin embedding and glyph embedding
word_embeddings = inputs_embeds # [bs,l,hidden_size]
pinyin_embeddings = self.pinyin_embeddings(pinyin_ids) # [bs,l,hidden_size]
glyph_embeddings = self.glyph_map(self.glyph_embeddings(input_ids)) # [bs,l,hidden_size]
# fusion layer
concat_embeddings = torch.cat((word_embeddings, pinyin_embeddings, glyph_embeddings), 2)
inputs_embeds = self.map_fc(concat_embeddings)
position_embeddings = self.position_embeddings(position_ids)
token_type_embeddings = self.token_type_embeddings(token_type_ids)
embeddings = inputs_embeds + position_embeddings + token_type_embeddings
embeddings = self.LayerNorm(embeddings)
embeddings = self.dropout(embeddings)
return embeddings
# paddle
import os
import paddle
class PaddleFusionBertEmbeddings(paddle.nn.Layer):
"""
Construct the embeddings from word, position, glyph, pinyin and token_type embeddings.
"""
def __init__(self, config):
super(PaddleFusionBertEmbeddings, self).__init__()
config_path = os.path.join(config.name_or_path, 'config')
font_files = []
for file in os.listdir(config_path):
if file.endswith(".npy"):
font_files.append(os.path.join(config_path, file))
self.word_embeddings = paddle.nn.Embedding(config.vocab_size, config.hidden_size, padding_idx=0)
self.position_embeddings = paddle.nn.Embedding(config.max_position_embeddings, config.hidden_size)
self.token_type_embeddings = paddle.nn.Embedding(config.type_vocab_size, config.hidden_size)
self.pinyin_embeddings = PaddlePinyinEmbedding(embedding_size=128,
pinyin_out_dim=config.hidden_size,
config_path=config_path)
self.glyph_embeddings = PaddleGlyphEmbedding(font_npy_files=font_files)
# self.LayerNorm is not snake-cased to stick with TensorFlow models variable name and be able to load
# any TensorFlow checkpoint file
self.glyph_map = paddle.nn.Linear(1728, config.hidden_size, bias_attr=True)
self.map_fc = paddle.nn.Linear(config.hidden_size * 3, config.hidden_size, bias_attr=True)
self.LayerNorm = paddle.nn.LayerNorm(config.hidden_size, epsilon=config.layer_norm_eps)
self.dropout = paddle.nn.Dropout(config.hidden_dropout_prob)
# position_ids (1, len position emb) is contiguous in memory and exported when serialized
self.register_buffer("position_ids", paddle.arange(config.max_position_embeddings).expand((1, -1)))
def forward(self, input_ids=None, pinyin_ids=None, token_type_ids=None, position_ids=None, inputs_embeds=None):
if input_ids is not None:
input_shape = input_ids.shape
else:
input_shape = inputs_embeds.shape[:-1]
seq_length = input_shape[1]
if position_ids is None:
position_ids = self.position_ids[:, :seq_length]
if token_type_ids is None:
token_type_ids = paddle.zeros(input_shape, dtype='int64')
if inputs_embeds is None:
inputs_embeds = self.word_embeddings(input_ids)
# get char embedding, pinyin embedding and glyph embedding
word_embeddings = inputs_embeds # [bs,l,hidden_size]
pinyin_embeddings = self.pinyin_embeddings(pinyin_ids) # [bs,l,hidden_size]
glyph_embeddings = self.glyph_map(self.glyph_embeddings(input_ids)) # [bs,l,hidden_size]
# fusion layer
concat_embeddings = paddle.concat((word_embeddings, pinyin_embeddings, glyph_embeddings), 2)
inputs_embeds = self.map_fc(concat_embeddings)
position_embeddings = self.position_embeddings(position_ids)
token_type_embeddings = self.token_type_embeddings(token_type_ids)
embeddings = inputs_embeds + position_embeddings + token_type_embeddings
embeddings = self.LayerNorm(embeddings)
embeddings = self.dropout(embeddings)
return embeddings
config
# torch
print("============================== torch =============================")
torch_fusion_emb = FusionBertEmbeddings(config)
print(torch_fusion_emb)
# paddle
print("============================== paddle =============================")
paddle_fusion_emb = PaddleFusionBertEmbeddings(config)
print(paddle_fusion_emb)
```
## GlyceBertModel
```
# torch
import warnings
import torch
from torch import nn
from transformers.modeling_bert import BertEncoder, BertPooler
from transformers.modeling_bert import BertModel
from transformers.modeling_outputs import BaseModelOutputWithPooling
class GlyceBertModel(BertModel):
r"""
Outputs: `Tuple` comprising various elements depending on the configuration (config) and inputs:
**last_hidden_state**: ``torch.FloatTensor`` of shape ``(batch_size, sequence_length, hidden_size)``
Sequence of hidden-states at the output of the last layer of the models.
**pooler_output**: ``torch.FloatTensor`` of shape ``(batch_size, hidden_size)``
Last layer hidden-state of the first token of the sequence (classification token)
further processed by a Linear layer and a Tanh activation function. The Linear
layer weights are trained from the next sentence prediction (classification)
objective during Bert pretraining. This output is usually *not* a good summary
of the semantic content of the input, you're often better with averaging or pooling
the sequence of hidden-states for the whole input sequence.
**hidden_states**: (`optional`, returned when ``config.output_hidden_states=True``)
list of ``torch.FloatTensor`` (one for the output of each layer + the output of the embeddings)
of shape ``(batch_size, sequence_length, hidden_size)``:
Hidden-states of the models at the output of each layer plus the initial embedding outputs.
**attentions**: (`optional`, returned when ``config.output_attentions=True``)
list of ``torch.FloatTensor`` (one for each layer) of shape ``(batch_size, num_heads, sequence_length, sequence_length)``:
Attentions weights after the attention softmax, used to compute the weighted average in the self-attention heads.
Examples::
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased')
models = BertModel.from_pretrained('bert-base-uncased')
input_ids = torch.tensor(tokenizer.encode("Hello, my dog is cute", add_special_tokens=True)).unsqueeze(0) # Batch size 1
outputs = models(input_ids)
last_hidden_states = outputs[0] # The last hidden-state is the first element of the output tuple
"""
def __init__(self, config):
super(GlyceBertModel, self).__init__(config)
self.config = config
self.embeddings = FusionBertEmbeddings(config)
self.encoder = BertEncoder(config)
self.pooler = BertPooler(config)
self.init_weights()
def forward(
self,
input_ids=None,
pinyin_ids=None,
attention_mask=None,
token_type_ids=None,
position_ids=None,
head_mask=None,
inputs_embeds=None,
encoder_hidden_states=None,
encoder_attention_mask=None,
output_attentions=None,
output_hidden_states=None,
return_dict=None,
):
r"""
encoder_hidden_states (:obj:`torch.FloatTensor` of shape :obj:`(batch_size, sequence_length, hidden_size)`, `optional`):
Sequence of hidden-states at the output of the last layer of the encoder. Used in the cross-attention
if the models is configured as a decoder.
encoder_attention_mask (:obj:`torch.FloatTensor` of shape :obj:`(batch_size, sequence_length)`, `optional`):
Mask to avoid performing attention on the padding token indices of the encoder input. This mask
is used in the cross-attention if the models is configured as a decoder.
Mask values selected in ``[0, 1]``:
- 1 for tokens that are **not masked**,
- 0 for tokens that are **masked**.
"""
output_attentions = output_attentions if output_attentions is not None else self.config.output_attentions
output_hidden_states = (
output_hidden_states if output_hidden_states is not None else self.config.output_hidden_states
)
return_dict = return_dict if return_dict is not None else self.config.use_return_dict
if input_ids is not None and inputs_embeds is not None:
raise ValueError("You cannot specify both input_ids and inputs_embeds at the same time")
elif input_ids is not None:
input_shape = input_ids.size()
elif inputs_embeds is not None:
input_shape = inputs_embeds.size()[:-1]
else:
raise ValueError("You have to specify either input_ids or inputs_embeds")
device = input_ids.device if input_ids is not None else inputs_embeds.device
if attention_mask is None:
attention_mask = torch.ones(input_shape, device=device)
if token_type_ids is None:
token_type_ids = torch.zeros(input_shape, dtype=torch.long, device=device)
# We can provide a self-attention mask of dimensions [batch_size, from_seq_length, to_seq_length]
# ourselves in which case we just need to make it broadcastable to all heads.
extended_attention_mask: torch.Tensor = self.get_extended_attention_mask(attention_mask, input_shape, device)
# If a 2D or 3D attention mask is provided for the cross-attention
# we need to make broadcastable to [batch_size, num_heads, seq_length, seq_length]
if self.config.is_decoder and encoder_hidden_states is not None:
encoder_batch_size, encoder_sequence_length, _ = encoder_hidden_states.size()
encoder_hidden_shape = (encoder_batch_size, encoder_sequence_length)
if encoder_attention_mask is None:
encoder_attention_mask = torch.ones(encoder_hidden_shape, device=device)
encoder_extended_attention_mask = self.invert_attention_mask(encoder_attention_mask)
else:
encoder_extended_attention_mask = None
# Prepare head mask if needed
# 1.0 in head_mask indicate we keep the head
# attention_probs has shape bsz x n_heads x N x N
# input head_mask has shape [num_heads] or [num_hidden_layers x num_heads]
# and head_mask is converted to shape [num_hidden_layers x batch x num_heads x seq_length x seq_length]
head_mask = self.get_head_mask(head_mask, self.config.num_hidden_layers)
embedding_output = self.embeddings(
input_ids=input_ids, pinyin_ids=pinyin_ids, position_ids=position_ids, token_type_ids=token_type_ids,
inputs_embeds=inputs_embeds
)
encoder_outputs = self.encoder(
embedding_output,
attention_mask=extended_attention_mask,
head_mask=head_mask,
encoder_hidden_states=encoder_hidden_states,
encoder_attention_mask=encoder_extended_attention_mask,
output_attentions=output_attentions,
output_hidden_states=output_hidden_states,
return_dict=return_dict,
)
sequence_output = encoder_outputs[0]
pooled_output = self.pooler(sequence_output) if self.pooler is not None else None
if not return_dict:
return (sequence_output, pooled_output) + encoder_outputs[1:]
return BaseModelOutputWithPooling(
last_hidden_state=sequence_output,
pooler_output=pooled_output,
hidden_states=encoder_outputs.hidden_states,
attentions=encoder_outputs.attentions,
)
glyce_bert_model = GlyceBertModel(config)
list(glyce_bert_model.state_dict().keys())[:10]
# paddle
import warnings
import paddle
from paddlenlp.transformers import BertModel
class BertPooler(paddle.nn.Layer):
"""
Pool the result of BertEncoder.
"""
def __init__(self, hidden_size, pool_act="tanh"):
super(BertPooler, self).__init__()
self.dense = paddle.nn.Linear(hidden_size, hidden_size)
self.activation = paddle.nn.Tanh()
self.pool_act = pool_act
def forward(self, hidden_states):
# We "pool" the model by simply taking the hidden state corresponding
# to the first token.
first_token_tensor = hidden_states[:, 0]
pooled_output = self.dense(first_token_tensor)
if self.pool_act == "tanh":
pooled_output = self.activation(pooled_output)
return pooled_output
class PaddleGlyceBertModel(BertModel):
r"""
PaddleGlyceBertModel
"""
def __init__(self, config):
super(PaddleGlyceBertModel, self).__init__(vocab_size=config.vocab_size)
self.embeddings = PaddleFusionBertEmbeddings(config)
encoder_layer = paddle.nn.TransformerEncoderLayer(
config.hidden_size,
config.num_attention_heads,
config.intermediate_size,
dropout=config.hidden_dropout_prob,
activation=config.hidden_act,
attn_dropout=config.attention_probs_dropout_prob,
act_dropout=0)
self.encoder = paddle.nn.TransformerEncoder(encoder_layer, config.num_hidden_layers)
self.pooler = BertPooler(config.hidden_size)
self.apply(self.init_weights)
def forward(
self,
input_ids=None,
pinyin_ids=None,
attention_mask=None,
token_type_ids=None,
position_ids=None,
head_mask=None,
inputs_embeds=None,
encoder_hidden_states=None,
encoder_attention_mask=None,
output_attentions=None,
output_hidden_states=None,
return_dict=None,
):
if attention_mask is None:
attention_mask = paddle.unsqueeze(
(input_ids == self.pad_token_id
).astype(self.pooler.dense.weight.dtype) * -1e9,
axis=[1, 2])
embedding_output = self.embeddings(
input_ids=input_ids, pinyin_ids=pinyin_ids, position_ids=position_ids, token_type_ids=token_type_ids,
inputs_embeds=inputs_embeds
)
if output_hidden_states:
output = embedding_output
encoder_outputs = []
for mod in self.encoder.layers:
output = mod(output, src_mask=attention_mask)
encoder_outputs.append(output)
if self.encoder.norm is not None:
encoder_outputs[-1] = self.encoder.norm(encoder_outputs[-1])
pooled_output = self.pooler(encoder_outputs[-1])
else:
sequence_output = self.encoder(embedding_output, attention_mask)
pooled_output = self.pooler(sequence_output)
if output_hidden_states:
return encoder_outputs, pooled_output
else:
return sequence_output, pooled_output
paddle_glyce_bert_model = PaddleGlyceBertModel(config)
list(paddle_state_dict.keys())[:10]
```
# 计算前项精度比较的方法
* 两个框架对于同一个模型的前项输出,最大误差应该控制在 10^-4,即,说明复现成功
```
import sys
sys.path.append('/data1/workspace/research/ChineseBERT-Paddle/ChineseBert')
# torch
from datasets.bert_dataset import BertDataset
from models.modeling_glycebert import GlyceBertModel
sentence = '我喜欢猫'
CHINESEBERT_PATH='./pretrain_models/torch/ChineseBERT-large/'
tokenizer = BertDataset(CHINESEBERT_PATH)
chinese_bert = GlyceBertModel.from_pretrained(CHINESEBERT_PATH)
chinese_bert.eval()
input_ids, pinyin_ids = tokenizer.tokenize_sentence(sentence)
length = input_ids.shape[0]
input_ids = input_ids.view(1, length)
pinyin_ids = pinyin_ids.view(1, length, 8)
print(input_ids)
print(pinyin_ids)
torch_output_hidden = chinese_bert.forward(input_ids, pinyin_ids)[0]
torch_output_hidden, torch_output_hidden.shape
#chinese_bert.state_dict()
import sys
sys.path.append('/data1/workspace/research/ChineseBERT-Paddle/Paddle_ChineseBert/PaddleNLP')
# paddle
import paddle
from paddlenlp.transformers import ChineseBertTokenizer
from paddlenlp.transformers import GlyceBertModel
sentence = '我喜欢猫'
CHINESEBERT_PADDLE_PATH = "./pretrain_models/paddle/ChineseBERT-large/"
tokenizer = ChineseBertTokenizer(CHINESEBERT_PADDLE_PATH)
glyce_bert_model = GlyceBertModel.from_pretrained(CHINESEBERT_PADDLE_PATH)
glyce_bert_model.eval()
token_input = tokenizer.tokenize_sentence(sentence)
input_ids = paddle.to_tensor(token_input['input_ids'])
pinyin_ids = paddle.to_tensor(token_input['pinyin_ids'])
input_ids = input_ids.unsqueeze(0)
pinyin_ids = pinyin_ids.unsqueeze(0)
print(input_ids)
print(pinyin_ids)
paddle_output_hidden = glyce_bert_model.forward(input_ids, pinyin_ids)[0]
paddle_output_hidden
import paddle
import torch
import numpy as np
# torch
t_output_hidden = torch_output_hidden.cpu().detach().numpy()
# paddle
p_output_hidden = paddle_output_hidden.cpu().detach().numpy()
diff = t_output_hidden - p_output_hidden
error = np.max(abs(diff))
print("最大误差:", error)
total_params = 0
for p in glyce_bert_model.parameters():
if len(p.shape) == 2:
_p = p.shape[0] * p.shape[1]
else:
_p = p.shape[0]
total_params += _p
print(total_params)
```
## QuestionAnswering
### torch
```
qa_outputs = torch.nn.Linear(1024, 2)
logits = qa_outputs(torch_output_hidden)
logits, logits.shape
start_logits, end_logits = logits.split(1, dim=-1)
start_logits, start_logits.shape
start_logits = start_logits.squeeze(-1)
start_logits, start_logits.shape
end_logits = end_logits.squeeze(-1)
end_logits
```
### paddle
```
p_classifier = paddle.nn.Linear(1024, 2)
p_logits = classifier(paddle_output_hidden)
p_logits
p_logits.transpose([2, 0, 1])
paddle.unstack(p_logits.transpose([2, 0, 1]), axis=0)
```
# 下游任务
## ChnSetiCorp
* **注意: 参数较大,建议使用 v100 32GB 的 GPU 执行**
```
from functools import partial
import argparse
import os
import random
import time
import numpy as np
import paddle
import paddle.nn.functional as F
import paddlenlp as ppnlp
from paddlenlp.data import Stack, Tuple, Pad
from paddlenlp.datasets import load_dataset
from paddlenlp.transformers import LinearDecayWithWarmup
def convert_example(example, tokenizer, max_seq_length=512, is_test=False):
# 【FOCUS】 --> https://github.com/ShannonAI/ChineseBert/blob/f6b4cd901e8f8b3ef2340ce2a8685b41df9bc261/datasets/chn_senti_corp_dataset.py#L33
text = example["text"][:max_seq_length-2]
encoded_inputs = tokenizer.tokenize_sentence(text)
input_ids = encoded_inputs["input_ids"]
pinyin_ids = encoded_inputs["pinyin_ids"]
if is_test:
return input_ids, pinyin_ids
label = np.array([example["label"]], dtype="int64")
return input_ids, pinyin_ids, label
def create_dataloader(dataset,
mode='train',
batch_size=1,
batchify_fn=None,
trans_fn=None):
if trans_fn:
dataset = dataset.map(trans_fn)
# shuffle = True if mode == 'train' else False
shuffle = False
if mode == 'train':
batch_sampler = paddle.io.DistributedBatchSampler(
dataset, batch_size=batch_size, shuffle=shuffle)
else:
batch_sampler = paddle.io.BatchSampler(
dataset, batch_size=batch_size, shuffle=shuffle)
return paddle.io.DataLoader(
dataset=dataset,
batch_sampler=batch_sampler,
collate_fn=batchify_fn,
return_list=True)
def set_seed(seed):
"""sets random seed"""
random.seed(seed)
np.random.seed(seed)
paddle.seed(seed)
@paddle.no_grad()
def evaluate(model, criterion, metric, data_loader):
"""
Given a dataset, it evals model and computes the metric.
Args:
model(obj:`paddle.nn.Layer`): A model to classify texts.
data_loader(obj:`paddle.io.DataLoader`): The dataset loader which generates batches.
criterion(obj:`paddle.nn.Layer`): It can compute the loss.
metric(obj:`paddle.metric.Metric`): The evaluation metric.
"""
model.eval()
metric.reset()
losses = []
for batch in data_loader:
input_ids, pinyin_ids, labels = batch
logits = model(input_ids, pinyin_ids)
loss = criterion(logits, labels)
losses.append(loss.numpy())
correct = metric.compute(logits, labels)
metric.update(correct)
accu = metric.accumulate()
print("eval loss: %.5f, accu: %.5f" % (np.mean(losses), accu))
model.train()
metric.reset()
def do_train(model, tokenizer):
paddle.set_device(args.device)
rank = paddle.distributed.get_rank()
if paddle.distributed.get_world_size() > 1:
paddle.distributed.init_parallel_env()
set_seed(args.seed)
train_ds, dev_ds = load_dataset("chnsenticorp", splits=["train", "dev"])
trans_func = partial(
convert_example,
tokenizer=tokenizer)
batchify_fn = lambda samples, fn=Tuple(
Pad(axis=0, pad_val=tokenizer.pad_token_id), # input
Pad(axis=0, pad_val=tokenizer.pad_token_type_id), # segment
Stack(dtype="int64") # label
): [data for data in fn(samples)]
train_data_loader = create_dataloader(
train_ds,
mode='train',
batch_size=args.batch_size,
batchify_fn=batchify_fn,
trans_fn=trans_func)
dev_data_loader = create_dataloader(
dev_ds,
mode='dev',
batch_size=args.batch_size,
batchify_fn=batchify_fn,
trans_fn=trans_func)
if args.init_from_ckpt and os.path.isfile(args.init_from_ckpt):
state_dict = paddle.load(args.init_from_ckpt)
model.set_dict(state_dict)
model = paddle.DataParallel(model)
num_training_steps = len(train_data_loader) * args.epochs
lr_scheduler = LinearDecayWithWarmup(args.learning_rate, num_training_steps,
args.warmup_proportion)
# Generate parameter names needed to perform weight decay.
# All bias and LayerNorm parameters are excluded.
decay_params = [
p.name for n, p in model.named_parameters()
if not any(nd in n for nd in ["bias", "norm"])
]
optimizer = paddle.optimizer.AdamW(
learning_rate=lr_scheduler,
parameters=model.parameters(),
weight_decay=args.weight_decay,
apply_decay_param_fun=lambda x: x in decay_params)
criterion = paddle.nn.loss.CrossEntropyLoss()
metric = paddle.metric.Accuracy()
global_step = 0
tic_train = time.time()
for epoch in range(1, args.epochs + 1):
for step, batch in enumerate(train_data_loader, start=1):
input_ids, pinyin_ids, labels = batch
logits = model(input_ids, pinyin_ids)
loss = criterion(logits, labels)
probs = F.softmax(logits, axis=1)
correct = metric.compute(probs, labels)
metric.update(correct)
acc = metric.accumulate()
global_step += 1
if global_step % 10 == 0 and rank == 0:
print(
"global step %d, epoch: %d, batch: %d, loss: %.5f, accu: %.5f, speed: %.2f step/s"
% (global_step, epoch, step, loss, acc,
10 / (time.time() - tic_train)))
tic_train = time.time()
loss.backward()
optimizer.step()
lr_scheduler.step()
optimizer.clear_grad()
if global_step % 100 == 0 and rank == 0:
save_dir = os.path.join(args.save_dir, "model_%d" % global_step)
if not os.path.exists(save_dir):
os.makedirs(save_dir)
evaluate(model, criterion, metric, dev_data_loader)
# model._layers.save_pretrained(save_dir)
# tokenizer.save_pretrained(save_dir)
```
### ChineseBERT-base
```
parser = argparse.ArgumentParser()
parser.add_argument("--save_dir", default='./checkpoint', type=str, help="The output directory where the model checkpoints will be written.")
parser.add_argument("--max_seq_length", default=512, type=int, help="The maximum total input sequence length after tokenization. "
"Sequences longer than this will be truncated, sequences shorter will be padded.")
parser.add_argument("--batch_size", default=16, type=int, help="Batch size per GPU/CPU for training.")
parser.add_argument("--learning_rate", default=2e-5, type=float, help="The initial learning rate for Adam.")
parser.add_argument("--weight_decay", default=0.0001, type=float, help="Weight decay if we apply some.")
parser.add_argument("--epochs", default=10, type=int, help="Total number of training epochs to perform.")
parser.add_argument("--warmup_proportion", default=0.1, type=float, help="Linear warmup proption over the training process.")
parser.add_argument("--init_from_ckpt", type=str, default=None, help="The path of checkpoint to be loaded.")
parser.add_argument("--seed", type=int, default=1000, help="random seed for initialization")
parser.add_argument('--device', choices=['cpu', 'gpu', 'xpu'], default="gpu", help="Select which device to train model, defaults to gpu.")
args = parser.parse_args(args=[])
# ChineseBertModel
#【注意】该路径下的 model_config.json 中的 `name_or_path` 必须和该路径一致
CHINESEBERT_PADDLE_PATH = "./pretrain_models/paddle/ChineseBERT-base"
model = ppnlp.transformers.GlyceBertForSequenceClassification.from_pretrained(CHINESEBERT_PADDLE_PATH, num_class=2)
# ChineseBertTokenizer
tokenizer = ppnlp.transformers.ChineseBertTokenizer(CHINESEBERT_PADDLE_PATH)
do_train(model, tokenizer)
```
### ChineseBERT-large
```
parser = argparse.ArgumentParser()
parser.add_argument("--save_dir", default='./checkpoint', type=str, help="The output directory where the model checkpoints will be written.")
parser.add_argument("--max_seq_length", default=512, type=int, help="The maximum total input sequence length after tokenization. "
"Sequences longer than this will be truncated, sequences shorter will be padded.")
parser.add_argument("--batch_size", default=8, type=int, help="Batch size per GPU/CPU for training.")
parser.add_argument("--learning_rate", default=1e-5, type=float, help="The initial learning rate for Adam.")
parser.add_argument("--weight_decay", default=0.0001, type=float, help="Weight decay if we apply some.")
parser.add_argument("--epochs", default=10, type=int, help="Total number of training epochs to perform.")
parser.add_argument("--warmup_proportion", default=0.1, type=float, help="Linear warmup proption over the training process.")
parser.add_argument("--init_from_ckpt", type=str, default=None, help="The path of checkpoint to be loaded.")
parser.add_argument("--seed", type=int, default=1000, help="random seed for initialization")
parser.add_argument('--device', choices=['cpu', 'gpu', 'xpu'], default="gpu", help="Select which device to train model, defaults to gpu.")
args = parser.parse_args(args=[])
# ChineseBertModel
#【注意】该路径下的 model_config.json 中的 `name_or_path` 必须和该路径一致
CHINESEBERT_PADDLE_PATH = "./pretrain_models/paddle/ChineseBERT-large"
model = ppnlp.transformers.GlyceBertForSequenceClassification.from_pretrained(CHINESEBERT_PADDLE_PATH, num_class=2)
# ChineseBertTokenizer
tokenizer = ppnlp.transformers.ChineseBertTokenizer(CHINESEBERT_PADDLE_PATH)
do_train(model, tokenizer)
```
# 附录
## 对比 paddlenlp 和 huggingface 的 `bert-base-chinese` 模型
* paddlenlp 文档:(这个文档有些老,但总体上是对的,需要注意 huggingface 现在模型参数已经弃用了 `attention.output.LayerNorm.gamma` 等)
https://paddlenlp.readthedocs.io/zh/latest/community/contribute_models/convert_pytorch_to_paddle.html
### paddlenlp
```
from paddlenlp.transformers import BertModel
p = BertModel.from_pretrained("bert-base-chinese")
p
p.state_dict()['encoder.layers.1.norm2.weight']
for k in p.state_dict().keys():
print(k, p.state_dict()[k].shape)
```
### huggingface
```
from transformers import BertModel
h = BertModel.from_pretrained('bert-base-chinese')
h
h.state_dict()['encoder.layer.0.attention.self.query.weight']
for k in h.state_dict().keys():
print(k, h.state_dict()[k].shape)
from paddlenlp.transformers import BertModel
bm = BertModel.from_pretrained('bert-base-chinese')
bm.save_pretrained('./output/bert-base-chinese')
```
| github_jupyter |
+ This notebook is part of lecture 19 *Determinant formulas and cofactors* in the OCW MIT course 18.06 by Prof Gilbert Strang [1]
+ Created by me, Dr Juan H Klopper
+ Head of Acute Care Surgery
+ Groote Schuur Hospital
+ University Cape Town
+ <a href="mailto:juan.klopper@uct.ac.za">Email me with your thoughts, comments, suggestions and corrections</a>
<a rel="license" href="http://creativecommons.org/licenses/by-nc/4.0/"><img alt="Creative Commons Licence" style="border-width:0" src="https://i.creativecommons.org/l/by-nc/4.0/88x31.png" /></a><br /><span xmlns:dct="http://purl.org/dc/terms/" href="http://purl.org/dc/dcmitype/InteractiveResource" property="dct:title" rel="dct:type">Linear Algebra OCW MIT18.06</span> <span xmlns:cc="http://creativecommons.org/ns#" property="cc:attributionName">IPython notebook [2] study notes by Dr Juan H Klopper</span> is licensed under a <a rel="license" href="http://creativecommons.org/licenses/by-nc/4.0/">Creative Commons Attribution-NonCommercial 4.0 International License</a>.
+ [1] <a href="http://ocw.mit.edu/courses/mathematics/18-06sc-linear-algebra-fall-2011/index.htm">OCW MIT 18.06</a>
+ [2] Fernando Pérez, Brian E. Granger, IPython: A System for Interactive Scientific Computing, Computing in Science and Engineering, vol. 9, no. 3, pp. 21-29, May/June 2007, doi:10.1109/MCSE.2007.53. URL: http://ipython.org
```
from IPython.core.display import HTML, Image
css_file = 'style.css'
HTML(open(css_file, 'r').read())
from sympy import init_printing, symbols, Matrix
from warnings import filterwarnings
init_printing(use_latex = 'mathjax')
filterwarnings('ignore')
x, y = symbols('x y')
```
# Determinant formulas and cofactors
# Tridiagonal matrices
## Creating an equation for the determinant of a 2×2 matrix
* Using just the three main properties from the previous lecture and knowing that the determinant of a matrix with a column of zero's is zero we have the following
$$ \begin{vmatrix} a & b \\ c & d \end{vmatrix}=\begin{vmatrix} a & 0 \\ c & d \end{vmatrix}+\begin{vmatrix} 0 & b \\ c & d \end{vmatrix}\\ =\begin{vmatrix} a & 0 \\ c & 0 \end{vmatrix}+\begin{vmatrix} a & 0 \\ 0 & d \end{vmatrix}+\begin{vmatrix} 0 & b \\ c & 0 \end{vmatrix}+\begin{vmatrix} 0 & b \\ 0 & d \end{vmatrix}\\ \because \quad \begin{vmatrix} a & 0 \\ c & 0 \end{vmatrix}=\begin{vmatrix} 0 & b \\ 0 & d \end{vmatrix}=0\\ \begin{vmatrix} a & 0 \\ 0 & d \end{vmatrix}+\begin{vmatrix} 0 & b \\ c & 0 \end{vmatrix}\\ =\begin{vmatrix} a & 0 \\ 0 & d \end{vmatrix}-\begin{vmatrix} c & 0 \\ 0 & b \end{vmatrix}\\ =ad-bc $$
## Creating an equation for the determinant of a 3×3 matrix
* By the method above, this will create a lot of matrices
* We need to figure out which ones remain, i.e. do not have columns of zeros
* Note carefully that we just keep those with at least one element from each row and column
$$ \begin{vmatrix} { a }_{ 11 } & { a }_{ 12 } & { a }_{ 13 } \\ { a }_{ 21 } & { a }_{ 22 } & { a }_{ 23 } \\ { a }_{ 31 } & { a }_{ 32 } & { a }_{ 33 } \end{vmatrix} \\ =\begin{vmatrix} { a }_{ 11 } & 0 & 0 \\ 0 & { a }_{ 22 } & 0 \\ 0 & 0 & { a }_{ 33 } \end{vmatrix}+\begin{vmatrix} { a }_{ 11 } & 0 & 0 \\ 0 & 0 & { a }_{ 23 } \\ 0 & { a }_{ 32 } & 0 \end{vmatrix}+\begin{vmatrix} 0 & { a }_{ 12 } & 0 \\ { a }_{ 21 } & 0 & 0 \\ 0 & 0 & { a }_{ 33 } \end{vmatrix}+\begin{vmatrix} 0 & { a }_{ 12 } & 0 \\ 0 & 0 & { a }_{ 23 } \\ { a }_{ 31 } & 0 & 0 \end{vmatrix}+\begin{vmatrix} 0 & 0 & { a }_{ 13 } \\ { a }_{ 21 } & 0 & 0 \\ 0 & { a }_{ 32 } & 0 \end{vmatrix}+\begin{vmatrix} 0 & 0 & { a }_{ 13 } \\ 0 & { a }_{ 22 } & 0 \\ { a }_{ 31 } & 0 & 0 \end{vmatrix}\\ ={ a }_{ 11 }{ a }_{ 22 }{ a }_{ 33 }-{ a }_{ 11 }{ a }_{ 23 }{ a }_{ 32 }-{ a }_{ 12 }{ a }_{ 21 }{ a }_{ 33 }+{ a }_{ 12 }{ a }_{ 23 }{ a }_{ 31 }+{ a }_{ 13 }{ a }_{ 21 }{ a }_{ 32 }-{ a }_{ 13 }{ a }_{ 22 }{ a }_{ 31 } $$
## Creating an equation for the determinant of a *n* × *n* matrix
* We will have *n*! terms, half of which is positive and the other half negative
* We have *n* because for the first row we have *n* positions to choose from, the for the second lot we have *n*-1 and so on
$$ \left| A \right| =\sum { \pm { a }_{ 1\alpha }{ a }_{ 2\beta }{ a }_{ 3\gamma }...{ a }_{ n\omega } } $$
* This holds for permuations of the columns (each used only once)
$$ \left( \alpha ,\beta ,\gamma ,\delta ,\dots ,\omega \right) =\left( 1,2,3,4,\dots ,n \right) $$
* Consider this example
$$ \begin{bmatrix} 0 & 0 & 1 & 1 \\ 0 & 1 & 1 & 0 \\ 1 & 1 & 0 & 0 \\ 1 & 0 & 0 & 1 \end{bmatrix} $$
* Successively choosing a single element from each column (using column numbers for the Greek symbols above), we get the following permutations (note their sign as we interchange the numbers to follow in order 1 2 3 4
* (4,3,2,1) = (1,2,3,4) Two *swaps*
* (3,2,1,4) = -(1,2,3,4) One *swap*
* That is it!
* So we have 1 - 1 = 0
* Note that in this example of a 4×4 matrix a lot of the permutations would have a zero in the, so we won't end up with 4! = 24 permutations
```
A = Matrix([[0, 0, 1, 1], [0, 1, 1, 0], [1, 1, 0, 0], [1, 0, 0, 1]])
A
A.det()
```
* We could have seen that this matrix is singular by noting that some combination of rows give identical rows and then by subtraction, a row of zero
```
A.rref()
```
## Cofactors of a 3×3 matrix
* Start with the equation above
$$ { a }_{ 11 }{ a }_{ 22 }{ a }_{ 33 }-{ a }_{ 11 }{ a }_{ 23 }{ a }_{ 32 }-{ a }_{ 12 }{ a }_{ 21 }{ a }_{ 33 }+{ a }_{ 12 }{ a }_{ 23 }{ a }_{ 31 }+{ a }_{ 13 }{ a }_{ 21 }{ a }_{ 32 }-{ a }_{ 13 }{ a }_{ 22 }{ a }_{ 31 } \\ ={ a }_{ 11 }\left( { a }_{ 22 }{ a }_{ 33 }-{ a }_{ 23 }{ a }_{ 32 } \right) +{ a }_{ 12 }\left( -{ a }_{ 21 }{ a }_{ 33 }+{ a }_{ 23 }{ a }_{ 31 } \right) +{ a }_{ 13 }\left( { a }_{ 21 }{ a }_{ 32 }-{ a }_{ 22 }{ a }_{ 31 } \right) $$
* The cofactors are in parentheses and are the 2×2 submatrix determinants
* They signify the determinant of a smaller (*n*-1) matrix with some sign problems, i.e. some are positive the determinant and some are negative the determinant
* We are especially interested here in row one, but any row (or even column) will do
* So for any *a*<sub>ij</sub> the cofactor is the ± determinant of the *n*-1 matrix with its *i* row and *j* column erased
* For the sign, if *i* + *j* is even, the sign is positive and if it is odd, then the sign is negative
* So the cofactor of *a*<sub>ij</sub> = C<sub>ij</sub>
* For rows we have
$$ { \left| A \right| }_{ i }=\sum _{ k=1 }^{ n }{ { a }_{ ik }{ C }_{ ik } } $$
## Diagonal matrices
* Calculate
$$ \left| { A }_{ 1 } \right| $$
```
A = Matrix([1])
A
A.det()
```
* Calculate
$$ \left| { A }_{ 2 } \right| $$
```
A = Matrix([[1, 1], [1, 1]])
A
A.det()
```
* Calculate
$$ \left| { A }_{ 3 } \right| $$
```
A = Matrix([[1, 1, 0], [1, 1, 1], [0, 1, 1]])
A
```
* By the cofactor equation above
$$ { \left| A \right| }_{ i }=\sum _{ k=1 }^{ n }{ { a }_{ ik }{ C }_{ ik } } \\ { \left| A \right| }_{ 1 }={ a }_{ 11 }{ C }_{ 11 }+{ a }_{ 12 }{ C }_{ 12 }+{ a }_{ 13 }{ C }_{ 13 }\\ { C }_{ ij }\rightarrow +;\left( i+j \right) \in 2n\\ { C }_{ ij }\rightarrow -;\left( i+j \right) \in 2n+1\\ { \left| A \right| }_{ 1 }=1\left( 0 \right) -1\left( 1 \right) +0\left( 1 \right) =-1 $$
```
A.det()
```
* Calculate
$$ \left| { A }_{ 4 } \right| $$
```
A = Matrix([[1, 1, 0, 0], [1, 1, 1, 0], [0, 1, 1, 1], [0, 0, 1, 1]])
A
A.det()
```
* Continuing on this path of tridiagonal matrices we have
$$ \left| { A }_{ n } \right| =\left| { A }_{ n-1 } \right| -\left| { A }_{ n-2 } \right| $$
* We would thus have
$$ \left| { A }_{ 5 } \right| =\left| { A }_{ 4 } \right| -\left| { A }_{ 3 } \right| \\ \left| { A }_{ 5 } \right| =-1-\left( -1 \right) =0 \\ \left| { A }_{ 6 } \right| =\left| { A }_{ 5 } \right| -\left| { A }_{ 4 } \right| \\ \left| { A }_{ 6 } \right| =0-\left( -1 \right) =1 $$
* We note that A<sub>7</sub> starts the sequence all over again
* Tridiagonal matrices have determinants of period 6
## Example problems
### Example problem 1
* Calculate the determinant of the following matrix
```
A = Matrix([[x, y, 0, 0, 0,], [0, x, y ,0 ,0 ], [0, 0, x, y, 0], [0, 0, 0, x, y], [y, 0, 0, 0, x]])
A
A.det()
```
#### Solution
* Note how first selecting row 1's *x* and the *y* leaves triangular matrices in the remaining (*n*-1)×(*n*-1) matrix
* These form cofactors and their determinant are simply the product of the entries along the main diagonal
* We simply have to remember the sign rule, which well be (-1)<sup>(5+1)</sup>
$$ \left| { A } \right| =x\left( { x }^{ 4 } \right) +y\left( { y }^{ 4 } \right) ={ x }^{ 5 }+{ y }^{ 5 } $$
### Example problem 2
```
A = Matrix([[x, y, y, y, y], [y, x, y, y, y], [y, y, x, y, y], [y, y, y, x, y], [y, y, y, y, x]])
A
```
#### Solution
```
A.det()
(A.det()).factor()
```
* Note that we can introduce many zero entry by the elementary row operation of subtracting one row from another
* Let's subtract row 4 from row 5
$$ \begin{bmatrix} x & y & y & y & y \\ y & x & y & y & y \\ y & y & x & y & y \\ y & y & y & x & y \\ 0 & 0 & 0 & y-x & x - y \end{bmatrix} $$
* Now subtract row 3 from 4
$$ \begin{bmatrix} x & y & y & y & y \\ y & x & y & y & y \\ y & y & x & y & y \\ 0 & 0 & y-x & x-y & 0 \\ 0 & 0 & 0 & y-x & x - y \end{bmatrix} $$
* Subtract 2 from 3
$$ \begin{bmatrix} x & y & y & y & y \\ y & x & y & y & y \\ 0 & y-x & x-y & 0 & 0 \\ 0 & 0 & y-x & x-y & 0 \\ 0 & 0 & 0 & y-x & x - y \end{bmatrix} $$
* ... and 1 from 2
$$ \begin{bmatrix} x & y & y & y & y \\ y-x & x-y & 0 & 0 & 0 \\ 0 & y-x & x-y & 0 & 0 \\ 0 & 0 & y-x & x-y & 0 \\ 0 & 0 & 0 & y-x & x - y \end{bmatrix} $$
* Now consider some column operations, adding the 5th column to the fourth column and then 4<sup>th</sup> to 3<sup>rd</sup> etc...
* This will introduce new non-zero entries, though
* These can be changed back to a zero by adding the 5<sup>/th</sup> column and the 4<sup>th</sup> to the 3<sup>rd</sup>
* Then columns 5, 4, 3 to 2, etc...
$$ \begin{bmatrix} x+4y & 4y & 3y & 2y & y \\ 0 & x-y & 0 & 0 & 0 \\ 0 & 0 & x-y & 0 & 0 \\ 0 & 0 & & x-y & 0 \\ 0 & 0 & 0 & 0 & x - y \end{bmatrix} $$
* This is upper triangular and the determinant is the product of the entries on the main diagonal
```
(x + 4 * y) * (x - y) ** 4
((x + 4 * y) * (x - y) ** 4).expand()
```
| github_jupyter |
```
# Create an empty list
ALLdata=[]
URL ="https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_US.csv"
resp = req.get(URL)
content = resp.text
#clean the content, then break the content into lines
content=content.replace(",,",",Ex,")
content=content.replace("(","")
content=content.replace(")","")
content=content.replace("\"","")
lines= content.splitlines()
print (len(lines))
# loop the lines one line at a time
# split each line at the delimiter ` , `
# then append the empty list 'ALLdata' with the line (which is now a list): [line]
for line in lines:
#convert the splitlines to strings
line= str(line).split(",")
ALLdata.append(line)
"""
Finding counties with a "threshhold" increase in deaths
Take the last four entries of data to see if the number of deaths has increased for the last three days:
To make it easy to understand lets use dates instead of data.
May10 May11 May12 May13
subract May10 from May11 check if the result is above the 'Threshhold'
subract May11 from May12 check if the result is above the 'Threshhold'
subract May12 from May13 check if the result is above the 'Threshhold'
if all three conditions are met, print the location and information.
"""
Threshhold = 10
count = 0
STATE =[]
COUNTY =[]
Points =[]
HISTORY=[]
lat=[]
long=[]
# Check each line of data, county by county.
for i in range(1,len(ALLdata)):
# Increase a counter for every line - this will allow further investigation into the data
# as demonstarted in the next four cells.
count=count+1
# subtract the last four days of data to see if it has increased by the minimum of the Threshhold each day
if int(ALLdata[i][-3])-int(ALLdata[i][-4]) >Threshhold and int(ALLdata[i][-2])-int(ALLdata[i][-3]) >Threshhold and int(ALLdata[i][-1])-int(ALLdata[i][-2]) >Threshhold:
# if they do increase as specified, define the line as a variable called history
history=[int(ALLdata[i][-3])-int(ALLdata[i][-4]),int(ALLdata[i][-2])-int(ALLdata[i][-3]),int(ALLdata[i][-1])-int(ALLdata[i][-2])]
HISTORY.append(history)
# The total amount of deaths in the specific county
deaths = int(ALLdata[i][-1])
# The county's name
county = ALLdata[i][5]
# The State the county is located in
state = ALLdata[i][6]
STATE.append(state)
COUNTY.append(county)
# The longitude and latitude of the county
longitude = ALLdata[i][9]
latitude = ALLdata[i][8]
long.append(float(longitude))
lat.append(float(latitude))
Points.append([float(longitude),float(latitude)])
#print the data line by line
print ("i="+str(count),deaths,county,state,longitude,latitude,history)
import matplotlib.pyplot as plt
from matplotlib.legend_handler import HandlerLine2D, HandlerTuple
for i in range(0,len(lat)):
#p1, = plt.scatter(long,lat, s=1, color='blue')
p2, = plt.plot(long[i],lat[i], s=20, color='red')
l = plt.legend([(p1, p2)], ['Two keys'], numpoints=1,
handler_map={tuple: HandlerTuple(ndivide=None)})
import matplotlib.pyplot as plt
import numpy as np
c = np.char.array(COUNTY)
S = np.char.array(STATE)
y = np.array(long)
x =np.array(lat)
inc= len(COUNTY)
colorZ = ['yellowgreen','red','gold','lightskyblue','white','lightcoral','blue','pink', 'darkgreen','yellow','grey','violet','magenta','cyan','yellowgreen','red','gold','lightskyblue','white','lightcoral','blue','pink', 'darkgreen','yellow','grey','violet','magenta','cyan','navy']
colors = colorZ[-inc:]
plt.scatter(x,y, s=40, color=colors)
labels = ['{0}, {1}'.format(i,j) for i,j in zip(c, S)]
sortlegend0 = [colors[0], labels[0],]
sortlegend1 = [colors[1], labels[1],]
print (sortlegend0,sortlegend1)
plt.legend([(sortlegend0,sortlegend1)], loc='left center', bbox_to_anchor=(-0.1, 1.),
fontsize=8)
"""
plt.legend(handles=sortlegend, loc='left center', bbox_to_anchor=(-0.1, 1.),
fontsize=8)
"""
plt.savefig('piechart.png', bbox_inches='tight')
#plt.legend(handles=[line_up, line_down])
text0 = COUNTY
text = STATE
for i in range(len(Points)):
x = Points[i][0]
y = Points[i][1]
tx = text0[i]+", "+text[i]
xx, yy = m(x,y)
print(xx, yy)
#plt.plot(xx, yy, 'bo')
plt.scatter(xx, yy, s=20, color='red', zorder=5, alpha=0.6)
plt.text(xx, yy, tx, fontsize=16, color="white")
import warnings
warnings.filterwarnings("ignore")
import matplotlib.pyplot as plt
from matplotlib.collections import PatchCollection
from matplotlib.patches import Polygon
import numpy as np
from PIL import Image,ImageFont,ImageDraw,ImageFilter,ImageChops
from random import randint
from mpl_toolkits.basemap import Basemap
import requests as req
import time
DATE = time.strftime("%m-%d-%H_")
# Create an empty list
ALLdata=[]
URL ="https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_US.csv"
resp = req.get(URL)
content = resp.text
#clean the content, then break the content into lines
content=content.replace(",,",",Ex,")
content=content.replace("(","")
content=content.replace(")","")
content=content.replace("\"","")
lines= content.splitlines()
print (len(lines))
# loop the lines one line at a time
# split each line at the delimiter ` , `
# then append the empty list 'ALLdata' with the line (which is now a list): [line]
for line in lines:
#convert the splitlines to strings
line= str(line).split(",")
ALLdata.append(line)
"""
Finding counties with a "threshhold" increase in deaths
Take the last four entries of data to see if the number of deaths has increased for the last three days:
To make it easy to understand lets use dates instead of data.
May10 May11 May12 May13
subract May10 from May11 check if the result is above the 'Threshhold'
subract May11 from May12 check if the result is above the 'Threshhold'
subract May12 from May13 check if the result is above the 'Threshhold'
if all three conditions are met, print the location and information.
"""
Threshhold = 10
count = 0
STATE =[]
COUNTY =[]
Points =[]
lat=[]
long=[]
# Check each line of data, county by county.
for i in range(1,len(ALLdata)):
# Increase a counter for every line - this will allow further investigation into the data
# as demonstarted in the next four cells.
count=count+1
# subtract the last four days of data to see if it has increased by the minimum of the Threshhold each day
if int(ALLdata[i][-3])-int(ALLdata[i][-4]) >Threshhold and int(ALLdata[i][-2])-int(ALLdata[i][-3]) >Threshhold and int(ALLdata[i][-1])-int(ALLdata[i][-2]) >Threshhold:
# if they do increase as specified, define the line as a variable called history
history=[int(ALLdata[i][-3])-int(ALLdata[i][-4]),int(ALLdata[i][-2])-int(ALLdata[i][-3]),int(ALLdata[i][-1])-int(ALLdata[i][-2])]
# The total amount of deaths in the specific county
deaths = int(ALLdata[i][-1])
# The county's name
county = ALLdata[i][5]
# The State the county is located in
state = ALLdata[i][6]
STATE.append(state)
COUNTY.append(county)
# The longitude and latitude of the county
longitude = ALLdata[i][9]
latitude = ALLdata[i][8]
long.append(float(longitude))
lat.append(float(latitude))
Points.append([float(longitude),float(latitude)])
#print the data line by line
print ("i="+str(count),deaths,county,state,longitude,latitude,history)
"""
Plot the data on a basemape with annotations of the County namea
"""
fig = plt.figure(num=None, figsize=(12, 8), dpi=120 )
m = Basemap(width=6000000,height=4500000,resolution='h',projection='aea',lat_1=35.,lat_2=45,lon_0=-100,lat_0=40)
m.drawcoastlines(linewidth=0.5)
m.fillcontinents(color='tan',lake_color='lightblue')
# draw parallels and meridians.
m.drawparallels(np.arange(-90.,91.,10.),labels=[True,True,False,False],dashes=[2,2])
m.drawmeridians(np.arange(-180.,181.,10.),labels=[False,False,False,True],dashes=[2,2])
m.drawmapboundary(fill_color='lightblue')
m.drawcountries(linewidth=2, linestyle='solid', color='k' )
m.drawstates(linewidth=0.5, linestyle='solid', color='k')
m.drawrivers(linewidth=0.5, linestyle='solid', color='blue')
#-- Place the text in the upper left hand corner of the axes
# The basemap instance doesn't have an annotate method, so we'll use the pyplot
# interface instead. (This is one of the many reasons to use cartopy instead.)
#plt.annotate('Jul-24-2012', xy=(0, 1), xycoords='axes fraction')
text0 = COUNTY
text = STATE
for i in range(len(Points)):
x = Points[i][0]
y = Points[i][1]
tx = str(text0[i])+", "+str(text[i])
xx, yy = m(x,y)
print(xx, yy)
#plt.plot(xx, yy, 'bo')
plt.scatter(xx, yy, s=20, color='red', zorder=5, alpha=0.6)
plt.text(xx, yy, tx, fontsize=16, color="white")
filename = "BaseMap/Hotspots__.png"
plt.savefig(filename, dpi=120, facecolor='salmon', edgecolor='b',
orientation='portrait', papertype=None, format=None,
transparent=False, bbox_inches="tight", pad_inches=0.2,
frameon=None, metadata=None)
plt.show()
```
| github_jupyter |
# Scientific Python
Python has a large number of tools available for doing data science, sometimes referred to as 'scientific Python'.
The scientific Python ecosystem revolves around some a set of core modules, including:
- `scipy`
- `numpy`
- `pandas`
- `matplotlib`
- `scikit-learn`
Here we will explore the basics of these modules and what they do, starting with scipy.
<br>
<br>
<img src="https://raw.githubusercontent.com/COGS108/Tutorials/master/img/scipy.png" width="200px">
<br>
<br>
<div class="alert alert-success">
Scipy is an 'ecosystem', including a collection of open-source packages for scientific computing in Python.
</div>
<div class="alert alert-info">
The scipy organization website is
<a href="https://www.scipy.org/" class="alert-link">here</a>,
including a
<a href="https://www.scipy.org/about.html" class="alert-link">description</a>
of the 'ecosystem', materials for
<a href="https://www.scipy.org/getting-started.html" class="alert-link">getting started</a>,
and extensive
<a href="https://docs.scipy.org/doc/scipy/reference/tutorial/" class="alert-link">tutorials</a>.
</div>
```
# You can import the full scipy package, typically shortened to 'sp'
import scipy as sp
# However, it is perhaps more common to import particular submodules
# For example, let's import the stats submodule
import scipy.stats as sts
```
Scipy has a broad range of functionality.
For a simple / random example, let's use it's stats module to model flipping a coin with [Bernouilli Distribution](https://en.wikipedia.org/wiki/Bernoulli_distribution), which is a distribution that can model a random variable that can be either 0 (call it Tails) or 1 (call it Heads).
```
# Let's model a fair coin - with 0.5 probability of being Heads
sts.bernoulli.rvs(0.5)
# Let's flip a bunch of coins!
coin_flips = [sts.bernoulli.rvs(0.5) for ind in range(100)]
print('The first ten coin flips are: ', coin_flips[:10])
print('The percent of heads from this sample is: ', sum(coin_flips) / len(coin_flips) * 100, '%')
```
<br>
<br>
<img src="https://raw.githubusercontent.com/COGS108/Tutorials/master/img/numpy.png" width="300px">
<br>
<br>
<div class="alert alert-success">
Numpy contains an array object (for multi-dimensional data, typically of uniform type), and operations for linear algrebra and analysis on arrays.
</div>
<div class="alert alert-info">
The numpy website is
<a href="http://www.numpy.org/" class="alert-link">here</a>,
including their official
<a href="https://docs.scipy.org/doc/numpy-dev/user/quickstart.html" class="alert-link">quickstart tutorial</a>.
</div>
Note:
An array is a 'a systematic arrangement of similar objects, usually in rows and columns' (definition from [Wikipedia](https://en.wikipedia.org/wiki/Array))
```
# Numpy is standardly imported as 'np'
import numpy as np
# Numpy's specialty is linear algebra and arrays of (uniform) data
# Define some arrays
# Arrays can have different types, but all the data within an array needs to be the same type
arr_1 = np.array([1, 2, 3])
arr_2 = np.array([4, 5, 6])
bool_arr = np.array([True, False, True])
str_arr = np.array(['a', 'b', 'c'])
# Note that if you try to make a mixed-data-type array, numpy won't fail,
# but it will (silently)
arr = np.array([1, 'b', True])
# Check the type of array items
print(type(arr[0]))
print(type(arr[2]))
# These array will therefore not act like you might expect
# The last item looks like a Boolen
print(arr[2])
# However, since it's actually a string, it won't evaluate like a Boolean
print(arr[2] == True)
```
<div class="alert alert-info">
For more practice with numpy, check out the collection
<a href="https://github.com/rougier/numpy-100" class="alert-link">numpy exercises</a>.
</div>
<br>
<br>
<img src="https://raw.githubusercontent.com/COGS108/Tutorials/master/img/pandas.png" width="400px">
<br>
<br>
<div class="alert alert-success">
Pandas is a package for organizing data in data structures, and performing data analysis upon them.
</div>
<div class="alert alert-info">
The official pandas website is
<a href="http://pandas.pydata.org/" class="alert-link">here</a>,
including materials such as
<a href="http://pandas.pydata.org/pandas-docs/version/0.17.0/10min.html" class="alert-link">10 minutes to pandas</a>
and a tutorial on
<a href="http://pandas.pydata.org/pandas-docs/version/0.17.0/basics.html" class="alert-link">essential basic functionality</a>.
</div>
Pandas main data object is the DataFrame, which is a powerful data object to store mixed data types together with labels.
Pandas dataframes also offer a large range of available methods for processing and analyzing data.
If you are familiar with R, pandas dataframes object and approaches are quite similar to R.
```
# Pandas is standardly imported as pd
import pandas as pd
# Let's start with an array of data, but we also have a label for each data item
dat_1 = np.array(['London', 'Washington', 'London', 'Budapest'])
labels = ['Ada', 'Alonzo', 'Alan', 'John']
# Pandas offers the 'Series' data object to store 1d data with axis labels
pd.Series?
# Let's make a Series with out data, and check it out
ser_1 = pd.Series(dat_1, labels)
ser_1.head()
# If we have some different data (with the same labels) we can make another Series
dat_2 = [36, 92, 41, 53]
ser_2 = pd.Series(dat_2, labels)
ser_2.head()
# However, having a collection of series can quickly get quite messy
# Pandas therefore offer the dataframe - a powerful data object to store mixed type data with labels
pd.DataFrame?
# There are several ways to initialize a dataframe
# Here, we provide a dictionary made up of our series
df = pd.DataFrame(data={'Col-A': ser_1, 'Col-B':ser_2}, index=labels)
# For categorical data, we can check how many of each value there are
df['Col-A'].value_counts()
# Note that dataframes are actually collections of Series
# When we index the df, as above, we actually pull out a Series
# So, the '.value_counts()' is actually a Series method
type(df['Col-A'])
# Pandas also gives us tons an ways to directly explore and analyze data in dataframes
# For example, the mean for all numberic data columns
df.mean()
```
<div class="alert alert-info">
For more practice with pandas, you can try some collections of exercises, including
<a href="https://github.com/guipsamora/pandas_exercises" class="alert-link">this one</a>
and
<a href="https://github.com/ajcr/100-pandas-puzzles" class="alert-link"> this one</a>.
</div>
<br>
<br>
<img src="https://raw.githubusercontent.com/COGS108/Tutorials/master/img/matplotlib.png" width="500px">
<br>
<br>
<div class="alert alert-success">
Matplotlib is a library for plotting, in particular for 2D plots.
</div>
<div class="alert alert-info">
The official numpy
<a href="http://matplotlib.org/" class="alert-link">website</a>
includes the official
<a href="http://matplotlib.org/users/pyplot_tutorial.html" class="alert-link">tutorial</a>
as well as a
<a href="https://matplotlib.org/gallery.html" class="alert-link">gallery</a>
of examples that you can start from and modify.
</div>
```
# This magic command is used to plot all figures inline in the notebook
%matplotlib inline
# Matplotlib is standardly imported as plt
import matplotlib.pyplot as plt
# Plot a basic line graph
plt.plot([1, 2, 3], [4, 6, 8])
```
<div class="alert alert-info">
There are also many external materials for using matplotlib, including
<a href="https://github.com/rougier/matplotlib-tutorial" class="alert-link">this one</a>.
</div>
<br>
<br>
<img src="https://raw.githubusercontent.com/COGS108/Tutorials/master/img/sklearn.png" width="250px">
<br>
<br>
<div class="alert alert-success">
Scikit-Learn is a packages for data mining, data analysis, and machine learning.
</div>
<div class="alert alert-info">
Here is the official scikit-learn
<a href="http://scikit-learn.org/" class="alert-link">website</a>
including their official
<a href="http://scikit-learn.org/stable/tutorial/basic/tutorial.html" class="alert-link">tutorial</a>.
</div>
```
# Import sklearn
import sklearn as skl
# Check out module description
skl?
```
We will get into machine learning and working with sklearn later on in the tutorials.
## External Resources
There are many, many resources to learn how to use those packages.
The links above include the official documentation and tutorials, which are the best place to start.
You can also search google for other resources and exercises.
<div class="alert alert-info">
A particularly good (and free) resource, covering all these tools is the
<a href="https://github.com/jakevdp/PythonDataScienceHandbook/" class="alert-link">Data Science Handbook </a>
by
<a href="https://github.com/jakevdp" class="alert-link">Jake Vanderplas</a>.
</div>
| github_jupyter |
<a href="https://colab.research.google.com/github/anirbrhm/lenet5-mnist/blob/main/Implementation_of_LeNet_5.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Digit Recognition using LeNet-5
```
import torch
import matplotlib.pyplot as plt
import numpy as np
import torchvision
import torchvision.transforms as transforms
```
## Data Loading
```
# adding this as downloading the dataset directly shows 403 error
from six.moves import urllib
opener = urllib.request.build_opener()
opener.addheaders = [('User-agent', 'Mozilla/5.0')]
urllib.request.install_opener(opener)
trainset = torchvision.datasets.MNIST(root = "./data", train = True, download = True,transform=transforms.ToTensor())
classes = ("0","1","2","3","4","5","6","7","8","9")
trainloader = torch.utils.data.DataLoader(trainset, batch_size=4, shuffle=True) # gives me 4 images everytime, shuffle means randomly.
dataiter = iter(trainloader) # create an iterator
images, labels = dataiter.next() # gives me one instance of trainloader (4 images)
print(images.shape)
print(images[1].shape)
print(labels[1].item())
```
The images are 1x28x28 shaped images.
## Visualize Data
```
img = images[1]
print(type(img))
npimg = img.numpy()
print(npimg.shape) # in the shape of (z,x,y) , but to plot it we need (x,y,z)
npimg = np.transpose(npimg, (1,2,0)).reshape(28,28) # for higher dimension matrices , transpose is any permutation.
print(npimg.shape)
plt.figure(figsize=(2,2))
plt.imshow(npimg) # image plotting on matplotlib
plt.show()
```
We can automate this process by programming a function to do this for us
```
def imshow(img):
npimg = img.numpy()
plt.figure(figsize=(2,2))
plt.imshow(np.transpose(npimg,(1,2,0)).reshape(28,28))
plt.show()
f, axarr = plt.subplots(2,2)
axarr[0,0].imshow(images[0].reshape(28,28))
axarr[0,1].imshow(images[1].reshape(28,28))
axarr[1,0].imshow(images[2].reshape(28,28))
axarr[1,1].imshow(images[3].reshape(28,28))
```
## LeNet Architecture
```
import torch.nn as nn
class LeNet(nn.Module) :
def __init__(self):
super(LeNet,self).__init__()
self.cnn_model = nn.Sequential(
nn.Conv2d(1,6,5), # (N,1,28,28) -> (N,6,24,24)
nn.Tanh(), # (N,6,24,24) -> (N,6,24,24)
nn.AvgPool2d(2,stride = 2), # (N,6,24,24) -> (N,6,12,12)
nn.Conv2d(6,16,5), # (N,6,12,12) -> (N,16,8,8)
nn.Tanh(), # (N,16,8,8) -> (N,16,8,8)
nn.AvgPool2d(2, stride = 2), # (N,16,8,8) -> (N,16,4,4)
)
# we flatten the tensor here to be a 256 dimensional vector
self.fc_model = nn.Sequential(
nn.Linear(256,120), # (N, 256) -> (N, 120)
nn.Tanh(),
nn.Linear(120,84), # (N, 120) -> (N, 84)
nn.Tanh(),
nn.Linear(84,10), # (N, 84) -> (N, 10)
nn.Softmax(1) # along the right dimension
)
def forward(self,x):
# print(x.shape)
x = self.cnn_model(x)
# print(x.shape)
x = x.view(x.size(0),-1)
# print(x.shape)
x = self.fc_model(x)
# print(x.shape)
return x
```
Loading the whole MNIST Data Set
```
batch_size = 128
trainset = torchvision.datasets.MNIST(root = "./data", train = True, download = True,transform=transforms.ToTensor())
trainloader = torch.utils.data.DataLoader(trainset, batch_size=batch_size, shuffle=True)
testset = torchvision.datasets.MNIST(root = "./data", train = False, download = True,transform=transforms.ToTensor())
testloader = torch.utils.data.DataLoader(testset, batch_size=batch_size, shuffle=False) # no need for shuffling
```
## Move to GPU
```
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
def evaluation(dataloader):
total , correct = 0 , 0
for data in dataloader:
inputs, labels = data
inputs, labels = inputs.to(device), labels.to(device)
outputs = net(inputs)
_, pred = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (pred == labels).sum().item()
return 100 * correct / total
import torch.optim as optim
net = LeNet().to(device)
loss_fn = nn.CrossEntropyLoss()
opt = optim.Adam(net.parameters())
%time
max_epochs = 16
for epoch in range(max_epochs):
for i, data in enumerate(trainloader, 0):
inputs, labels = data
inputs, labels = inputs.to(device), labels.to(device)
opt.zero_grad()
outputs = net(inputs)
loss = loss_fn(outputs, labels)
loss.backward()
opt.step()
print("Epoch : %d/%d" %(epoch,max_epochs))
print("Test acc : %0.2f, Train acc : %0.2f" %(evaluation(testloader),evaluation(trainloader)))
```
| github_jupyter |
# Query Classifier Tutorial
[](https://colab.research.google.com/github/deepset-ai/haystack/blob/master/tutorials/Tutorial14_Query_Classifier.ipynb)
In this tutorial we introduce the query classifier the goal of introducing this feature was to optimize the overall flow of Haystack pipeline by detecting the nature of user queries. Now, the Haystack can detect primarily three types of queries using both light-weight SKLearn Gradient Boosted classifier or Transformer based more robust classifier. The three categories of queries are as follows:
### 1. Keyword Queries:
Such queries don't have semantic meaning and merely consist of keywords. For instance these three are the examples of keyword queries.
* arya stark father
* jon snow country
* arya stark younger brothers
### 2. Interrogative Queries:
In such queries users usually ask a question, regardless of presence of "?" in the query the goal here is to detect the intent of the user whether any question is asked or not in the query. For example:
* who is the father of arya stark ?
* which country was jon snow filmed ?
* who are the younger brothers of arya stark ?
### 3. Declarative Queries:
Such queries are variation of keyword queries, however, there is semantic relationship between words. Fo example:
* Arya stark was a daughter of a lord.
* Jon snow was filmed in a country in UK.
* Bran was brother of a princess.
In this tutorial, you will learn how the `TransformersQueryClassifier` and `SklearnQueryClassifier` classes can be used to intelligently route your queries, based on the nature of the user query. Also, you can choose between a lightweight Gradients boosted classifier or a transformer based classifier.
Furthermore, there are two types of classifiers you can use out of the box from Haystack.
1. Keyword vs Statement/Question Query Classifier
2. Statement vs Question Query Classifier
As evident from the name the first classifier detects the keywords search queries and semantic statements like sentences/questions. The second classifier differentiates between question based queries and declarative sentences.
### Prepare environment
#### Colab: Enable the GPU runtime
Make sure you enable the GPU runtime to experience decent speed in this tutorial.
**Runtime -> Change Runtime type -> Hardware accelerator -> GPU**
<img src="https://raw.githubusercontent.com/deepset-ai/haystack/master/docs/_src/img/colab_gpu_runtime.jpg">
These lines are to install Haystack through pip
```
# Install the latest release of Haystack in your own environment
#! pip install farm-haystack
# Install the latest master of Haystack
!pip install grpcio-tools==1.34.1
!pip install --upgrade git+https://github.com/deepset-ai/haystack.git
# Install pygraphviz
!apt install libgraphviz-dev
!pip install pygraphviz
# If you run this notebook on Google Colab, you might need to
# restart the runtime after installing haystack.
# In Colab / No Docker environments: Start Elasticsearch from source
! wget https://artifacts.elastic.co/downloads/elasticsearch/elasticsearch-7.9.2-linux-x86_64.tar.gz -q
! tar -xzf elasticsearch-7.9.2-linux-x86_64.tar.gz
! chown -R daemon:daemon elasticsearch-7.9.2
import os
from subprocess import Popen, PIPE, STDOUT
es_server = Popen(['elasticsearch-7.9.2/bin/elasticsearch'],
stdout=PIPE, stderr=STDOUT,
preexec_fn=lambda: os.setuid(1) # as daemon
)
# wait until ES has started
! sleep 30
```
If running from Colab or a no Docker environment, you will want to start Elasticsearch from source
## Initialization
Here are some core imports
Then let's fetch some data (in this case, pages from the Game of Thrones wiki) and prepare it so that it can
be used indexed into our `DocumentStore`
```
from haystack.utils import print_answers
from haystack.preprocessor.utils import fetch_archive_from_http, convert_files_to_dicts
from haystack.preprocessor.cleaning import clean_wiki_text
from haystack import Pipeline
from haystack.pipeline import TransformersQueryClassifier, SklearnQueryClassifier, RootNode
from haystack.utils import launch_es
from haystack.document_store import ElasticsearchDocumentStore
from haystack.retriever.sparse import ElasticsearchRetriever
from haystack.retriever.dense import DensePassageRetriever
from haystack.reader import FARMReader
#Download and prepare data - 517 Wikipedia articles for Game of Thrones
doc_dir = "data/article_txt_got"
s3_url = "https://s3.eu-central-1.amazonaws.com/deepset.ai-farm-qa/datasets/documents/wiki_gameofthrones_txt.zip"
fetch_archive_from_http(url=s3_url, output_dir=doc_dir)
# convert files to dicts containing documents that can be indexed to our datastore
got_dicts = convert_files_to_dicts(
dir_path=doc_dir,
clean_func=clean_wiki_text,
split_paragraphs=True
)
# Initialize DocumentStore and index documents
launch_es()
document_store = ElasticsearchDocumentStore()
document_store.delete_documents()
document_store.write_documents(got_dicts)
# Initialize Sparse retriever
es_retriever = ElasticsearchRetriever(document_store=document_store)
# Initialize dense retriever
dpr_retriever = DensePassageRetriever(document_store)
document_store.update_embeddings(dpr_retriever, update_existing_embeddings=False)
reader = FARMReader(model_name_or_path="deepset/roberta-base-squad2")
```
## Keyword vs Question/Statement Classifier
The keyword vs question/statement query classifier essentially distinguishes between the keyword queries and statements/questions. So you can intelligently route to different retrieval nodes based on the nature of the query. Using this classifier can potentially yield the following benefits:
* Getting better search results (e.g. by routing only proper questions to DPR / QA branches and not keyword queries)
* Less GPU costs (e.g. if 50% of your traffic is only keyword queries you could just use elastic here and save the GPU resources for the other 50% of traffic with semantic queries)

Below, we define a `SklQueryClassifier` and show how to use it:
Read more about the trained model and dataset used [here](https://ext-models-haystack.s3.eu-central-1.amazonaws.com/gradboost_query_classifier/readme.txt)
```
# Here we build the pipeline
sklearn_keyword_classifier = Pipeline()
sklearn_keyword_classifier.add_node(component=SklearnQueryClassifier(), name="QueryClassifier", inputs=["Query"])
sklearn_keyword_classifier.add_node(component=dpr_retriever, name="DPRRetriever", inputs=["QueryClassifier.output_1"])
sklearn_keyword_classifier.add_node(component=es_retriever, name="ESRetriever", inputs=["QueryClassifier.output_2"])
sklearn_keyword_classifier.add_node(component=reader, name="QAReader", inputs=["ESRetriever", "DPRRetriever"])
sklearn_keyword_classifier.draw("pipeline_classifier.png")
# Run only the dense retriever on the full sentence query
res_1 = sklearn_keyword_classifier.run(
query="Who is the father of Arya Stark?"
)
print("DPR Results" + "\n" + "="*15)
print_answers(res_1)
# Run only the sparse retriever on a keyword based query
res_2 = sklearn_keyword_classifier.run(
query="arya stark father"
)
print("ES Results" + "\n" + "="*15)
print_answers(res_2)
# Run only the dense retriever on the full sentence query
res_3 = sklearn_keyword_classifier.run(
query="which country was jon snow filmed ?"
)
print("DPR Results" + "\n" + "="*15)
print_answers(res_3)
# Run only the sparse retriever on a keyword based query
res_4 = sklearn_keyword_classifier.run(
query="jon snow country"
)
print("ES Results" + "\n" + "="*15)
print_answers(res_4)
# Run only the dense retriever on the full sentence query
res_5 = sklearn_keyword_classifier.run(
query="who are the younger brothers of arya stark ?"
)
print("DPR Results" + "\n" + "="*15)
print_answers(res_5)
# Run only the sparse retriever on a keyword based query
res_6 = sklearn_keyword_classifier.run(
query="arya stark younger brothers"
)
print("ES Results" + "\n" + "="*15)
print_answers(res_6)
```
## Transformer Keyword vs Question/Statement Classifier
Firstly, it's essential to understand the trade-offs between SkLearn and Transformer query classifiers. The transformer classifier is more accurate than SkLearn classifier however, it requires more memory and most probably GPU for faster inference however the transformer size is roughly `50 MBs`. Whereas, SkLearn is less accurate however is much more faster and doesn't require GPU for inference.
Below, we define a `TransformersQueryClassifier` and show how to use it:
Read more about the trained model and dataset used [here](https://huggingface.co/shahrukhx01/bert-mini-finetune-question-detection)
```
# Here we build the pipeline
transformer_keyword_classifier = Pipeline()
transformer_keyword_classifier.add_node(component=TransformersQueryClassifier(), name="QueryClassifier", inputs=["Query"])
transformer_keyword_classifier.add_node(component=dpr_retriever, name="DPRRetriever", inputs=["QueryClassifier.output_1"])
transformer_keyword_classifier.add_node(component=es_retriever, name="ESRetriever", inputs=["QueryClassifier.output_2"])
transformer_keyword_classifier.add_node(component=reader, name="QAReader", inputs=["ESRetriever", "DPRRetriever"])
transformer_keyword_classifier.draw("pipeline_classifier.png")
# Run only the dense retriever on the full sentence query
res_1 = transformer_keyword_classifier.run(
query="Who is the father of Arya Stark?"
)
print("DPR Results" + "\n" + "="*15)
print_answers(res_1)
# Run only the sparse retriever on a keyword based query
res_2 = transformer_keyword_classifier.run(
query="arya stark father"
)
print("ES Results" + "\n" + "="*15)
print_answers(res_2)
# Run only the dense retriever on the full sentence query
res_3 = transformer_keyword_classifier.run(
query="which country was jon snow filmed ?"
)
print("DPR Results" + "\n" + "="*15)
print_answers(res_3)
# Run only the sparse retriever on a keyword based query
res_4 = transformer_keyword_classifier.run(
query="jon snow country"
)
print("ES Results" + "\n" + "="*15)
print_answers(res_4)
# Run only the dense retriever on the full sentence query
res_5 = transformer_keyword_classifier.run(
query="who are the younger brothers of arya stark ?"
)
print("DPR Results" + "\n" + "="*15)
print_answers(res_5)
# Run only the sparse retriever on a keyword based query
res_6 = transformer_keyword_classifier.run(
query="arya stark younger brothers"
)
print("ES Results" + "\n" + "="*15)
print_answers(res_6)
```
## Question vs Statement Classifier
One possible use case of this classifier could be to route queries after the document retrieval to only send questions to QA reader and in case of declarative sentence, just return the DPR/ES results back to user to enhance user experience and only show answers when user explicitly asks it.

Below, we define a `TransformersQueryClassifier` and show how to use it:
Read more about the trained model and dataset used [here](https://huggingface.co/shahrukhx01/question-vs-statement-classifier)
```
# Here we build the pipeline
transformer_question_classifier = Pipeline()
transformer_question_classifier.add_node(component=dpr_retriever, name="DPRRetriever", inputs=["Query"])
transformer_question_classifier.add_node(component=TransformersQueryClassifier(model_name_or_path="shahrukhx01/question-vs-statement-classifier"), name="QueryClassifier", inputs=["DPRRetriever"])
transformer_question_classifier.add_node(component=reader, name="QAReader", inputs=["QueryClassifier.output_1"])
transformer_question_classifier.draw("question_classifier.png")
# Run only the QA reader on the question query
res_1 = transformer_question_classifier.run(
query="Who is the father of Arya Stark?"
)
print("DPR Results" + "\n" + "="*15)
print_answers(res_1)
# Show only DPR results
res_2 = transformer_question_classifier.run(
query="Arya Stark was the daughter of a Lord."
)
print("ES Results" + "\n" + "="*15)
res_2
```
## Standalone Query Classifier
Below we run queries classifiers standalone to better understand their outputs on each of the three types of queries
```
# Here we create the keyword vs question/statement query classifier
from haystack.pipeline import TransformersQueryClassifier
queries = ["arya stark father","jon snow country",
"who is the father of arya stark","which country was jon snow filmed?"]
keyword_classifier = TransformersQueryClassifier()
for query in queries:
result = keyword_classifier.run(query=query)
if result[1] == "output_1":
category = "question/statement"
else:
category = "keyword"
print(f"Query: {query}, raw_output: {result}, class: {category}")
# Here we create the question vs statement query classifier
from haystack.pipeline import TransformersQueryClassifier
queries = ["Lord Eddard was the father of Arya Stark.","Jon Snow was filmed in United Kingdom.",
"who is the father of arya stark?","Which country was jon snow filmed in?"]
question_classifier = TransformersQueryClassifier(model_name_or_path="shahrukhx01/question-vs-statement-classifier")
for query in queries:
result = question_classifier.run(query=query)
if result[1] == "output_1":
category = "question"
else:
category = "statement"
print(f"Query: {query}, raw_output: {result}, class: {category}")
```
## Conclusion
The query classifier gives you more possibility to be more creative with the pipelines and use different retrieval nodes in a flexible fashion. Moreover, as in the case of Question vs Statement classifier you can also choose the queries which you want to send to the reader.
Finally, you also have the possible of bringing your own classifier and plugging it into either `TransformersQueryClassifier(model_name_or_path="<huggingface_model_name_or_file_path>")` or using the `SklearnQueryClassifier(model_name_or_path="url_to_classifier_or_file_path_as_pickle", vectorizer_name_or_path="url_to_vectorizer_or_file_path_as_pickle")`
## About us
This [Haystack](https://github.com/deepset-ai/haystack/) notebook was made with love by [deepset](https://deepset.ai/) in Berlin, Germany
We bring NLP to the industry via open source!
Our focus: Industry specific language models & large scale QA systems.
Some of our other work:
- [German BERT](https://deepset.ai/german-bert)
- [GermanQuAD and GermanDPR](https://deepset.ai/germanquad)
- [FARM](https://github.com/deepset-ai/FARM)
Get in touch:
[Twitter](https://twitter.com/deepset_ai) | [LinkedIn](https://www.linkedin.com/company/deepset-ai/) | [Slack](https://haystack.deepset.ai/community/join) | [GitHub Discussions](https://github.com/deepset-ai/haystack/discussions) | [Website](https://deepset.ai)
By the way: [we're hiring!](https://apply.workable.com/deepset/)
| github_jupyter |
# KubeFlow Pipeline: Github Issue Summarization using Tensor2Tensor
This notebook assumes that you have already set up a GKE cluster with Kubeflow installed as per this codelab: [g.co/codelabs/kubecon18](g.co/codelabs/kubecon18). Currently, this notebook must be run from the Kubeflow JupyterHub installation, as described in the codelab.
In this notebook, we will show how to:
* Interactively define a KubeFlow Pipeline using the Pipelines Python SDK
* Submit and run the pipeline
* Add a step in the pipeline
This example pipeline trains a [Tensor2Tensor](https://github.com/tensorflow/tensor2tensor/) model on Github issue data, learning to predict issue titles from issue bodies. It then exports the trained model and deploys the exported model to [Tensorflow Serving](https://github.com/tensorflow/serving).
The final step in the pipeline launches a web app which interacts with the TF-Serving instance in order to get model predictions.
## Setup
Do some installations and imports, and set some variables. Set the `WORKING_DIR` to a path under the Cloud Storage bucket you created earlier. The Pipelines SDK is bundled with the notebook server image, but we'll make sure that we're using the most current version for this example. You may need to restart your kernel after the SDK update.
```
!pip install -U kfp
import kfp # the Pipelines SDK.
from kfp import compiler
import kfp.dsl as dsl
import kfp.gcp as gcp
import kfp.components as comp
from kfp.dsl.types import Integer, GCSPath, String
import kfp.notebook
# Define some pipeline input variables.
WORKING_DIR = 'gs://YOUR_GCS_BUCKET/t2t/notebooks' # Such as gs://bucket/object/path
PROJECT_NAME = 'YOUR_PROJECT'
GITHUB_TOKEN = 'YOUR_GITHUB_TOKEN' # needed for prediction, to grab issue data from GH
DEPLOY_WEBAPP = 'false'
```
## Create an *Experiment* in the Kubeflow Pipeline System
The Kubeflow Pipeline system requires an "Experiment" to group pipeline runs. You can create a new experiment, or call `client.list_experiments()` to get existing ones.
```
# Note that this notebook should be running in JupyterHub in the same cluster as the pipeline system.
# Otherwise, additional config would be required to connect.
client = kfp.Client()
client.list_experiments()
exp = client.create_experiment(name='t2t_notebook')
```
## Define a Pipeline
Authoring a pipeline is like authoring a normal Python function. The pipeline function describes the topology of the pipeline. The pipeline components (steps) are container-based. For this pipeline, we're using a mix of predefined components loaded from their [component definition files](https://www.kubeflow.org/docs/pipelines/sdk/component-development/), and some components defined via [the `dsl.ContainerOp` constructor](https://www.kubeflow.org/docs/pipelines/sdk/build-component/). For this codelab, we've prebuilt all the components' containers.
While not shown here, there are other ways to build Kubeflow Pipeline components as well, including converting stand-alone python functions to containers via [`kfp.components.func_to_container_op(func)`](https://www.kubeflow.org/docs/pipelines/sdk/lightweight-python-components/). You can read more [here](https://www.kubeflow.org/docs/pipelines/sdk/).
This pipeline has several steps:
- An existing model checkpoint is copied to your bucket.
- Dataset metadata is logged to the Kubeflow metadata server.
- A [Tensor2Tensor](https://github.com/tensorflow/tensor2tensor/) model is trained using preprocessed data. (Training starts from the existing model checkpoint copied in the first step, then trains for a few more hundred steps-- it would take too long to fully train it now). When it finishes, it exports the model in a form suitable for serving by [TensorFlow serving](https://github.com/tensorflow/serving/).
- Training metadata is logged to the metadata server.
- The next step in the pipeline deploys a TensorFlow-serving instance using that model.
- The last step launches a web app for interacting with the served model to retrieve predictions.
We'll first define some constants and load some components from their definition files.
```
COPY_ACTION = 'copy_data'
TRAIN_ACTION = 'train'
WORKSPACE_NAME = 'ws_gh_summ'
DATASET = 'dataset'
MODEL = 'model'
copydata_op = comp.load_component_from_url(
'https://raw.githubusercontent.com/kubeflow/examples/master/github_issue_summarization/pipelines/components/t2t/datacopy_component.yaml' # pylint: disable=line-too-long
)
train_op = comp.load_component_from_url(
'https://raw.githubusercontent.com/kubeflow/examples/master/github_issue_summarization/pipelines/components/t2t/train_component.yaml' # pylint: disable=line-too-long
)
metadata_log_op = comp.load_component_from_url(
'https://raw.githubusercontent.com/kubeflow/examples/master/github_issue_summarization/pipelines/components/t2t/metadata_log_component.yaml' # pylint: disable=line-too-long
)
```
Next, we'll define the pipeline itself.
```
@dsl.pipeline(
name='Github issue summarization',
description='Demonstrate Tensor2Tensor-based training and TF-Serving'
)
def gh_summ( #pylint: disable=unused-argument
train_steps: 'Integer' = 2019300,
project: String = 'YOUR_PROJECT_HERE',
github_token: String = 'YOUR_GITHUB_TOKEN_HERE',
working_dir: GCSPath = 'YOUR_GCS_DIR_HERE',
checkpoint_dir: GCSPath = 'gs://aju-dev-demos-codelabs/kubecon/model_output_tbase.bak2019000/',
deploy_webapp: String = 'true',
data_dir: GCSPath = 'gs://aju-dev-demos-codelabs/kubecon/t2t_data_gh_all/'
):
copydata = copydata_op(
data_dir=data_dir,
checkpoint_dir=checkpoint_dir,
model_dir='%s/%s/model_output' % (working_dir, dsl.RUN_ID_PLACEHOLDER),
action=COPY_ACTION,
).apply(gcp.use_gcp_secret('user-gcp-sa'))
log_dataset = metadata_log_op(
log_type=DATASET,
workspace_name=WORKSPACE_NAME,
run_name=dsl.RUN_ID_PLACEHOLDER,
data_uri=data_dir
)
train = train_op(
data_dir=data_dir,
model_dir=copydata.outputs['copy_output_path'],
action=TRAIN_ACTION, train_steps=train_steps,
deploy_webapp=deploy_webapp
).apply(gcp.use_gcp_secret('user-gcp-sa'))
log_model = metadata_log_op(
log_type=MODEL,
workspace_name=WORKSPACE_NAME,
run_name=dsl.RUN_ID_PLACEHOLDER,
model_uri=train.outputs['train_output_path']
)
serve = dsl.ContainerOp(
name='serve',
image='gcr.io/google-samples/ml-pipeline-kubeflow-tfserve',
arguments=["--model_name", 'ghsumm-%s' % (dsl.RUN_ID_PLACEHOLDER,),
"--model_path", train.outputs['train_output_path']
]
)
log_dataset.after(copydata)
log_model.after(train)
train.set_gpu_limit(1)
train.set_memory_limit('48G')
with dsl.Condition(train.outputs['launch_server'] == 'true'):
webapp = dsl.ContainerOp(
name='webapp',
image='gcr.io/google-samples/ml-pipeline-webapp-launcher:v2ap',
arguments=["--model_name", 'ghsumm-%s' % (dsl.RUN_ID_PLACEHOLDER,),
"--github_token", github_token]
)
webapp.after(serve)
```
## Submit an experiment *run*
```
compiler.Compiler().compile(gh_summ, 'ghsumm.tar.gz')
```
The call below will run the compiled pipeline. We won't actually do that now, but instead we'll add a new step to the pipeline, then run it.
```
# You'd uncomment this call to actually run the pipeline.
# run = client.run_pipeline(exp.id, 'ghsumm', 'ghsumm.tar.gz',
# params={'working-dir': WORKING_DIR,
# 'github-token': GITHUB_TOKEN,
# 'project': PROJECT_NAME})
```
## Add a step to the pipeline
Next, let's add a new step to the pipeline above. As currently defined, the pipeline accesses a directory of pre-processed data as input to training. Let's see how we could include the pre-processing as part of the pipeline.
We're going to cheat a bit, as processing the full dataset will take too long for this workshop, so we'll use a smaller sample. For that reason, you won't actually make use of the generated data from this step (we'll stick to using the full dataset for training), but this shows how you could do so if we had more time.
First, we'll define the new pipeline step. Note the last line of this new function, which gives this step's pod the credentials to access GCP.
```
# defining the new data preprocessing pipeline step.
# Note the last line, which gives this step's pod the credentials to access GCP
def preproc_op(data_dir, project):
return dsl.ContainerOp(
name='datagen',
image='gcr.io/google-samples/ml-pipeline-t2tproc',
arguments=[ "--data-dir", data_dir, "--project", project]
).apply(gcp.use_gcp_secret('user-gcp-sa'))
```
### Modify the pipeline to add the new step
Now, we'll redefine the pipeline to add the new step. We're reusing the component ops defined above.
```
# Then define a new Pipeline. It's almost the same as the original one,
# but with the addition of the data processing step.
@dsl.pipeline(
name='Github issue summarization',
description='Demonstrate Tensor2Tensor-based training and TF-Serving'
)
def gh_summ2(
train_steps: 'Integer' = 2019300,
project: String = 'YOUR_PROJECT_HERE',
github_token: String = 'YOUR_GITHUB_TOKEN_HERE',
working_dir: GCSPath = 'YOUR_GCS_DIR_HERE',
checkpoint_dir: GCSPath = 'gs://aju-dev-demos-codelabs/kubecon/model_output_tbase.bak2019000/',
deploy_webapp: String = 'true',
data_dir: GCSPath = 'gs://aju-dev-demos-codelabs/kubecon/t2t_data_gh_all/'
):
# The new pre-processing op.
preproc = preproc_op(project=project,
data_dir=('%s/%s/gh_data' % (working_dir, dsl.RUN_ID_PLACEHOLDER)))
copydata = copydata_op(
data_dir=data_dir,
checkpoint_dir=checkpoint_dir,
model_dir='%s/%s/model_output' % (working_dir, dsl.RUN_ID_PLACEHOLDER),
action=COPY_ACTION,
).apply(gcp.use_gcp_secret('user-gcp-sa'))
log_dataset = metadata_log_op(
log_type=DATASET,
workspace_name=WORKSPACE_NAME,
run_name=dsl.RUN_ID_PLACEHOLDER,
data_uri=data_dir
)
train = train_op(
data_dir=data_dir,
model_dir=copydata.outputs['copy_output_path'],
action=TRAIN_ACTION, train_steps=train_steps,
deploy_webapp=deploy_webapp
).apply(gcp.use_gcp_secret('user-gcp-sa'))
log_dataset.after(copydata)
train.after(preproc)
log_model = metadata_log_op(
log_type=MODEL,
workspace_name=WORKSPACE_NAME,
run_name=dsl.RUN_ID_PLACEHOLDER,
model_uri=train.outputs['train_output_path']
)
serve = dsl.ContainerOp(
name='serve',
image='gcr.io/google-samples/ml-pipeline-kubeflow-tfserve',
arguments=["--model_name", 'ghsumm-%s' % (dsl.RUN_ID_PLACEHOLDER,),
"--model_path", train.outputs['train_output_path']
]
)
log_model.after(train)
train.set_gpu_limit(1)
train.set_memory_limit('48G')
with dsl.Condition(train.outputs['launch_server'] == 'true'):
webapp = dsl.ContainerOp(
name='webapp',
image='gcr.io/google-samples/ml-pipeline-webapp-launcher:v2ap',
arguments=["--model_name", 'ghsumm-%s' % (dsl.RUN_ID_PLACEHOLDER,),
"--github_token", github_token]
)
webapp.after(serve)
```
### Compile the new pipeline definition and submit the run
```
compiler.Compiler().compile(gh_summ2, 'ghsumm2.tar.gz')
run = client.run_pipeline(exp.id, 'ghsumm2', 'ghsumm2.tar.gz',
params={'working-dir': WORKING_DIR,
'github-token': GITHUB_TOKEN,
'deploy-webapp': DEPLOY_WEBAPP,
'project': PROJECT_NAME})
```
You should be able to see your newly defined pipeline run in the dashboard:

The new pipeline has the following structure:

Below is a screenshot of the pipeline running.

When this new pipeline finishes running, you'll be able to see your generated processed data files in GCS under the path: `WORKING_DIR/<pipeline_name>/gh_data`. There isn't time in the workshop to pre-process the full dataset, but if there had been, we could have defined our pipeline to read from that generated directory for its training input.
-----------------------------
Copyright 2018 Google LLC
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
| github_jupyter |
## Import Necessary Libraries and Packages
```
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import zipfile
import os
import statistics
import seaborn as sns
from google.colab import files
%matplotlib inline
import six
import sys
sys.modules['sklearn.externals.six'] = six
```
## Load the Preprocessed Data
```
uploaded = files.upload() # 'lin_filtered.csv'
# load preprocessed data file as a pandas dataframe
df = pd.read_csv('lin_filtered.csv', low_memory=False)
df.columns
df.drop(columns='Unnamed: 0', inplace=True)
df.head()
```
#### Pop Out Nametags Column
```
nametags = df.pop('nametags').astype(str)
print(nametags)
df.head()
```
### Create Train/Test Split
```
# import from sklearn
from sklearn.metrics import precision_recall_curve
from sklearn.metrics import f1_score
from sklearn.metrics import auc
from sklearn.metrics import classification_report,confusion_matrix,roc_curve,roc_auc_score
from sklearn.metrics import accuracy_score,log_loss
from sklearn.model_selection import train_test_split
# dont forget to define your X and y
X = df.iloc[:, 0:342] # features
y = df.activity # target variable
feature_list = list(X.columns) # save feature names for later
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.2, random_state=420)
```
### Scale Data to Prepare for Model Creation
```
from sklearn.preprocessing import MaxAbsScaler
# build scaler based on training data and apply it to test data to then also scale the test data
scaler = MaxAbsScaler().fit(X_train)
X_train_scaled=scaler.transform(X_train)
X_test_scaled=scaler.transform(X_test)
```
# Create Baseline Models and Compare
```
!pip install lazypredict
from lazypredict.Supervised import LazyClassifier
lazy_clf = LazyClassifier(verbose=0,ignore_warnings=True, custom_metric=None, random_state=420)
models,predictions = lazy_clf.fit(X_train_scaled, X_test_scaled, y_train, y_test)
models
```
Of the four different performance metrics presented in this report, I will be focusing on the "Balanced Accuracy", a particularly useful metric for evaluating imbalanced classifiers. Balanced accuracy is the arithmetic mean of the sensitivity and specificity.
A naive classifier, one that always predicts the majority (negative) class, would have a balanced accuracy of 0.50, or 50%. We can tell how much better any of the classification algorithms above are performing compared to a naive classifier by how much greater their balanced accuracy is than 50%.

The models with the highest balanced accuracy are:
1. Nearest Centroid
2. Gaussian Naive Bayes
In addition to these models, I will be training a Random Forest model and Logistic Regression model.
I chose Random Forest because it is highly optimizable and interpretable. However the report shows RandomForestClassifier has a balanced accuracy of only 52%, so it is barely doing better than a naive classifier that always predicts negative. Hyperparameter tuning can have a huge effect on model performance, though, so I will still give this one a chance.
I chose Logistic Regression Classifier because it is also very efficient to train, interpretable, and easy to implement. I expect this algorithm to perform better than the Random Forest, since the LogisticRegressionClassifier has a 61% balanced accuracy compared to the RandomForestClassifier's 52% balanced accuracy.
## Balance the Imbalanced Classes with SMOTETomek
SMOTE is an oversampling method that interpolates synthetic observations between the existing ones in the dataset. Tomek Links are pairs of nearest neighbors that have different classes, which in this case, are instances of the majority and minority class that are next to each other. Tomek Links are used for undersampling the majority class by getting rid of those observations which are next to the minority class examples, thus creating a more distinct decision boundary for the classifier.
SMOTETomek is a combined method that oversamples the minority class and undersamples the majority class. This method is preferable to using SMOTE by itself since it cleans up the noise that SMOTE tends to create in the sample space.
```
!pip install imblearn
# import imblearn libraries for SMOTETomek
from imblearn.over_sampling import SMOTE
from imblearn.under_sampling import TomekLinks
from imblearn.combine import SMOTETomek
# define the resampler, smt
# oversample minority class, undersample majority class
smt = SMOTETomek(random_state=420, smote=SMOTE(sampling_strategy='minority'), tomek=TomekLinks(sampling_strategy='majority'))
# resample the training dataset
X_res, y_res = smt.fit_resample(X_train_scaled, y_train) # only for training data!!
# visualize the two classes with a barchart to confirm balance
y_res_ser = pd.Series(y_res) #resampled training labels
y_tr_ser = pd.Series(y_train) #original training labels
plt.figure(figsize=(10, 8))
plt.subplot(2, 1, 1)
plt.tight_layout()
_ = y_tr_ser.value_counts().plot(kind='barh', color=['lightcoral', 'cornflowerblue'])
_ = plt.xlabel('Number of Observations')
_ = plt.ylabel('Classes')
_ = plt.title('Class Balance of Original Training Data')
plt.subplot(2, 1, 2)
plt.tight_layout()
_ = y_res_ser.value_counts().plot(kind='barh', color=['lightcoral', 'cornflowerblue'])
_ = plt.xlabel('Number of Observations')
_ = plt.ylabel('Classes')
_ = plt.title('Class Balance of Resampled Training Data')
plt.show()
```
# Random Forest Classifier
The first classification model that I'll be using is a Random Forest Classifier. I will be comparing the performance of the Random Forest Classifier on the unbalanced train and test sets with a Random Forest Classifier trained on the balanced train and test sets.
### Train and Fit the RandomForestClassifier Model
```
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier(n_estimators = 500, random_state=420, n_jobs=-1)
model1 = clf.fit(X_train_scaled, y_train)
y_pred = model1.predict(X_test_scaled)
y_pred_prob = model1.predict_proba(X_test_scaled)
rf_probs = y_pred_prob[:,1]
ac = accuracy_score(y_test, y_pred)
f1 = f1_score(y_test, y_pred, average='weighted')
print('Random Forest: Accuracy=%.3f' % (ac))
print('Random Forest: f1-score=%.3f' % (f1))
```
The Random Forest classifier model shows an overall accuracy of 99.4% and an F1-score of 99.1% - both of which look very good at first glance, but are misleading!
My guess is that this model has a high accuracy score due to overfitting. Random Forest Classifiers have a tendency to overfit, especially when there is a large number of trees (n_estimators). For this reason, I will use two different methods for hyperparameter optimization - RandomizedSearchCV and Bayesian Optimization - to train Random Forest Classifiers and then compare the results with this baseline model. In the next section, I will take a closer look at the model's performance.
### Evaluate Model Performance
To evaluate my Random Forest Classifier, I will be visualizing the model's performance using a confusion matrix, and then use the classification report to view the precision, recall, and f1-scores for each category. The confusion matrix will give me an overall idea of the model's performance, while the classification report will help me understand my model's performance in a more nuanced way.
```
conf = confusion_matrix(y_test, y_pred).astype('float')
conf = conf / conf.sum(axis=1)[:, np.newaxis]
#plot confusion matrix
plt.figure(figsize=(16, 10))
sns.heatmap(conf, annot=True, annot_kws={'size':10}, cmap=plt.cm.Greens, linewidths=0.2)
sns.set(font_scale=1.5)
_ = plt.xlabel('Predicted Activity')
_ = plt.ylabel('True Activity')
_ = plt.title('Confusion Matrix for Random Forest Model')
plt.show()
```
The 0's correspond to **inactive p53 proteins**, and the 1's correspond to **active p53 proteins**.
This confusion matrix shows that my classifier had trouble correctly classifying the active p53 proteins as active, as seen by the large proportion of false negatives. On the other hand, my classifier did very well at correctly labeling *inactive* p53 proteins.
From looking at the confusion matrix, I can tell that this model is very close to a *naive classifier* - it nearly always predicts "0", or inactive, for the protein. Statistically, this is the best way for the model to maximize the chances of making a correct prediction, since there is a huge class imbalance in the training data. This confirms my previous hypothesis regarding the Random Forest Classifier's performance for this dataset.
This makes sense, as the data is very imbalanced with regards to inactive vs active protein data. There is much more data for inactive proteins in the dataset than for active proteins. I saw this imbalance in the data during the exploratory data analysis that was performed earlier.
```
print(classification_report(y_test, y_pred))
```
The imbalance in the data is further confirmed by the low recall and low f1-scores for the "True" (aka "Active" p53) category. This is due to the low amount of "True" observations within the dataset, which does not allow the classifier enough data to learn how to predict the "True" class. This is a common problem with classifiers made with imbalanced data.
To address this problem, I will oversample my minority class by generating synthetic data (SMOTE), which will create a more balanced dataset for my classifier model to learn from. I will also be combining this method with the Tomek Links undersampling method, which will create a clearer decision boundary for my binary classifier.
Before I move on, however, I am going to visualize the top 10 feature importances from this Random Forest classifier, so that I can compare these results with those of following models:
```
feature_importances = pd.Series(model1.feature_importances_, index=feature_list)
_ = feature_importances.nlargest(10).plot(kind='barh', color=['royalblue', 'orange', 'green', 'tab:red', 'blueviolet', 'tab:brown', 'tab:pink', 'tab:gray', 'tab:olive', 'tab:cyan']).invert_yaxis()
_ = plt.ylabel('Feature Numbers')
_ = plt.xlabel('Importances')
_ = plt.title('Top 10 Feature Importances for Random Forest Classifier Model')
plt.show()
```
Looks like the **most importance features** for this Random Forest Classifier model are: **3991, 2827, 1674, 2826, 4028, 2833, 4120, 4134, 2767, and 370**.
All of the Top 10 Most Important Features are **Electrostatic and Surface-Based Features**! This is very interesting, because I would have thought that the Distance-Based Features would be more important for indicating wild-type p53 function. However, this feature list is not reliable due to the extremely low recall for the positive class. Thus, I will be comparing these Top Features with those of following models after resampling the data and hyperparameter tuning my model algorithms.
## Random Forest with Oversampling/Undersampling and Randomized Search
```
# run a Random Forest model with the resampled data and call it model2
model2 = clf.fit(X_res, y_res) # use X_res and y_res instead of X_train_scaled and y_train
y_pred = model2.predict(X_test_scaled)
y_pred_prob = model2.predict_proba(X_test_scaled)
rf_probs2 = y_pred_prob[:,1]
ac = accuracy_score(y_test, y_pred)
f1 = f1_score(y_test, y_pred, average='weighted')
print(f1)
# classification report for model 2
print(classification_report(y_test, y_pred))
```
<div class="alert-warning">
Resampling the training data with SMOTETomek helped increase the recall for the positive class from 0.05 to 0.27 and the f1-score from 0.09 to 0.35! That's a significant improvement, and a great starting point. I want to continue to maximize the f1-score, which effectively balances the precision and recall, in further models.
</div>
```
# plot confusion matrix for model 2
conf2 = confusion_matrix(y_test, y_pred).astype('float')
conf2 = conf2 / conf2.sum(axis=1)[:, np.newaxis]
plt.figure(figsize=(10, 8))
sns.heatmap(conf2, annot=True, annot_kws={'size':10}, cmap=plt.cm.Greens, linewidths=0.2)
sns.set(font_scale=1.5)
_ = plt.xlabel('Predicted Activity')
_ = plt.ylabel('True Activity')
_ = plt.title('Confusion Matrix for Random Forest Model 2')
plt.show()
```
### Increase Weights on the Minority Class
```
# add class weights in favor of the minority (False/Inactive) class and run Random Forest model
# model3, clf2
clf2 = RandomForestClassifier(n_estimators = 500, class_weight={0:1, 1:10}, random_state=420, n_jobs=-1)
model3 = clf2.fit(X_res, y_res) # use X_res and y_res instead of X_train_scaled and y_train
y_pred = model3.predict(X_test_scaled)
y_pred_prob = model3.predict_proba(X_test_scaled)
rf_probs3 = y_pred_prob[:,1]
ac = accuracy_score(y_test, y_pred)
f1 = f1_score(y_test, y_pred, average='weighted')
# classification report for model 3
print(classification_report(y_test, y_pred))
```
<div class="alert-warning">
By increasing the class weight on the positive class, the random forest algorithm now penalizes incorrect classifications of the positive class ten times more than incorrect classifications of the negative class.
This change decreased the f1-score to 0.27, but it did even out the precision and recall for the positive class. This tells me that changing the class weights in this way makes the model "pickier" - it does not classify very many of the observations as "active", but out of the ones it *does* classify as "active", it is usually correct. I want to explore this hyperparameter further.
</div>
## Hyperparameter Tuning with Randomized Search
Let's start by taking a look at the parameters used by the previous Random Forest model:
```
print('Current Random Forest Parameters:\n ', clf.get_params())
```
<div class="alert-danger">
Out of these parameters, I will be focusing on the tuning the following:
+ max_features: max number of features considered when splitting a node
+ min_samples_split: minimum number of samples required to split a node
+ min_samples_leaf: minimum number of samples required at each leaf
+ n_estimators: number of trees in the forest
</div>
```
from sklearn.model_selection import RandomizedSearchCV
# max_features
max_features=['auto', 'sqrt']
# min_samples_split
min_samples_split = [i for i in np.linspace(2, 40, 20)]
# min_samples_leaf
min_samples_leaf = [1, 2, 4]
# n_estimators
n_estimators = [i for i in np.linspace(200, 2000, 20)]
# create the random parameter grid
random_grid = {'n_estimators': n_estimators
'max_features': max_features,
'min_samples_split': min_samples_split,
'min_samples_leaf': min_samples_leaf}
# create a RandomForestClassifier to tune
clf = RandomForestClassifier(random_state=420)
# create random search model
model4 = RandomizedSearchCV(estimator=clf, param_distributions=random_grid, n_iter=100, cv=3, verbose=2, random_state=420, n_jobs=-1)
# fit random search model on resampled data
model4.fit(X_res, y_res)
# view best hyperparameters from fitting the random search
print(model4.best_params_)
```
<div class="alert-danger">
The best parameters from the Randomized Search are:
{'n_estimators': 673, 'min_samples_split': 2, 'min_samples_leaf': 1, 'max_features': 'sqrt'}
To deal with the imbalance in the dataset, I will also be adding the class_weights parameter to the Random Forest Classifier and assigning more weights to the "active" class.
+ class_weight: class weights affect how the model is penalized for misclassifying examples of each class
</div>
```
# compute class_weights based on original y_train
from sklearn.utils.class_weight import compute_class_weight
weights = compute_class_weight(class_weight='balanced', classes=np.unique(y_res), y=y_train)
weights # weights for original training data
# compute class_weights based on resampled y_train
weights_res = compute_class_weight(class_weight='balanced', classes=np.unique(y_res), y=y_res)
weights_res # weights for resampled y_train labels
# Random Forest Classifier with tuned hyperparameters
# min_samples_split': 2, 'min_samples_leaf': 1, 'max_features': 'sqrt'
clf3 = RandomForestClassifier(n_estimators = 673, class_weight={0:0.50456204, 1:55.3}, min_samples_split=2, min_samples_leaf=1, max_features='sqrt', random_state=420, n_jobs=-1)
model_tuned = clf3.fit(X_res, y_res) # use X_res and y_res instead of X_train_scaled and y_train
y_pred = model_tuned.predict(X_test_scaled)
y_pred_prob = model_tuned.predict_proba(X_test_scaled)
rf_probs4 = y_pred_prob[:,1]
ac = accuracy_score(y_test, y_pred)
f1 = f1_score(y_test, y_pred, average='weighted')
# print classification report for tuned Random Forest model
print(classification_report(y_test, y_pred))
```
# Logistic Regression Classifier
Logistic Regression is a solid algorithm choice for modeling this data for a few reasons:
+ The dependent (target) variable is binary
+ The features have already been filtered using linear correlation, so there should be little collinearity between variables
+ My sample sizes are large
I will be training my model on the rescaled, resampled data that I used for my Random Forest Classifiers.
```
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression(penalty='l1', class_weight='balanced', random_state=420, solver='saga', multi_class='ovr', n_jobs=-1, warm_start=True)
logreg.fit(X_res, y_res) # fit on training data
y_pred = logreg.predict(X_test_scaled)
y_pred_prob = logreg.predict_proba(X_test_scaled)
logreg_probs = y_pred_prob[:,1]
```
I chose the hyperparameters for this model:
* penalty='l1'
* class_weight='balanced'
* solver='saga'
* multi_class='ovr'
* n_jobs=-1
* warm_start=True
```
# plot confusion matrix for logistic regression model
conf_logreg = confusion_matrix(y_test, y_pred).astype('float')
conf_logreg = conf_logreg / conf_logreg.sum(axis=1)[:, np.newaxis]
plt.figure(figsize=(10, 8))
sns.heatmap(conf_logreg, annot=True, annot_kws={'size':10}, cmap=plt.cm.Greens, linewidths=0.2)
sns.set(font_scale=1.5)
_ = plt.xlabel('Predicted Activity')
_ = plt.ylabel('True Activity')
_ = plt.title('Confusion Matrix for Logistic Regression Binary Classifier')
plt.show()
# show the classification report for logistic regression model
print(classification_report(y_test, y_pred))
```
The Logistic Regression model shows a high recall for both classes, with the recall for the positive class being 73%! A high recall indicates a low false negative rate, which is exactly what I want. Since the positive class, which are transcriptionally active p53 mutants, are so rare, I want to make sure that as many of them as possible are correctly classified as being "active".
On the other hand, this model has very low precision for the positive class, and I can see from the confusion matrix that some of the "active" p53 mutants were incorrectly classified as being "inactive". This is okay, since minimizing the false negatives is more important for my classification problem.
The Logistic Regression classifier performed better than any of the Random Forest Classifiers, which agrees with my initial hypothesis.
# Gaussian Naive Bayes Classifier
This classifier assumes the likelihood of the features is Gaussian:

```
from sklearn.naive_bayes import GaussianNB
# GaussianNB model here
gnb = GaussianNB()
gnb.fit(X_res, y_res)
y_pred = gnb.predict(X_test_scaled)
gnb_probs = gnb.predict_proba(X_test_scaled)[:,1] # use to plot calibration curve
# plot confusion matrix for Gaussian Naive Bayes model
conf_gnb = confusion_matrix(y_test, y_pred).astype('float')
conf_gnb = conf_gnb / conf_gnb.sum(axis=1)[:, np.newaxis]
plt.figure(figsize=(10, 8))
sns.heatmap(conf_gnb, annot=True, annot_kws={'size':10}, cmap=plt.cm.Greens, linewidths=0.2)
sns.set(font_scale=1.5)
_ = plt.xlabel('Predicted Activity')
_ = plt.ylabel('True Activity')
_ = plt.title('Confusion Matrix for Gaussian Naive Bayes Classifier')
plt.show()
print(classification_report(y_test, y_pred))
```
# Nearest Centroid Classifier
```
from sklearn.neighbors import NearestCentroid
nc_clf = NearestCentroid()
nc_clf.fit(X_res, y_res)
y_pred = nc_clf.predict(X_test_scaled)
nc_centroids = nc_clf.centroids_
# plot confusion matrix nearest centroid model
conf_nc = confusion_matrix(y_test, y_pred).astype('float')
conf_nc = conf_nc / conf_nc.sum(axis=1)[:, np.newaxis]
plt.figure(figsize=(10, 8))
sns.heatmap(conf_nc, annot=True, annot_kws={'size':10}, cmap=plt.cm.Greens, linewidths=0.2)
sns.set(font_scale=1.5)
_ = plt.xlabel('Predicted Activity')
_ = plt.ylabel('True Activity')
_ = plt.title('Confusion Matrix for Nearest Centroid Classifier')
plt.show()
print(classification_report(y_test, y_pred))
```
The Nearest Centroid model has high recall but low precision, which means that it casts a wide net - a third of the "inactive" proteins were classified as "active". However, this model does correctly classify 86% of the "active" proteins, which is better than the other models perform.
# Comparison of Probability Calibration Curves
```
from sklearn.calibration import calibration_curve
from sklearn.metrics import (brier_score_loss, precision_score, recall_score,
f1_score)
# plot calibration curves using: rf_prob, logreg_prob, gnb_prob, nc_prob
plt.figure(figsize=(10, 8))
ax1 = plt.subplot2grid((3, 1), (0, 0), rowspan=2)
ax2 = plt.subplot2grid((3, 1), (2, 0))
probas = [(rf_probs, 'Baseline Random Forest'),
(rf_probs2, 'Random Forest with Balanced Data'),
(rf_probs3, 'Random Forest with 1:10 Class Weights'),
(rf_probs4, 'Optimized Random Forest'),
(logreg_probs, 'Logistic Regression'),
(gnb_probs, 'Gaussian Naive Bayes')]
for prob, name in probas:
fraction_of_positives, mean_predicted_value = \
calibration_curve(y_test, prob, n_bins=5)
ax1.plot(mean_predicted_value, fraction_of_positives, "s-",
label="%s" % (name, ))
ax2.hist(prob, range=(0, 1), bins=10, label=name,
histtype="step", lw=2)
ax1.set_ylabel("Fraction of positives")
ax1.set_ylim([-0.05, 1.05])
ax1.legend(loc="upper left", fontsize='xx-small')
ax1.set_title('Calibration plots (reliability curve)')
ax2.set_xlabel("Mean predicted value")
ax2.set_ylabel("Count")
ax2.legend(loc="upper center", ncol=2, fontsize='xx-small')
plt.tight_layout()
plt.show()
```
| github_jupyter |
# Run Torch2TRT from Arachne
The [torch2trt](https://github.com/NVIDIA-AI-IOT/torch2trt>) is a PyTorch to TensorRT converter.
## Prepare a Model
First, we have to prepare a model to be used in this tutorial.
Here, we will use a pre-trained model of the ResNet-18 from `torchvision.models`.
```
import torch
import torchvision
resnet18 = torchvision.models.resnet18(pretrained=True)
torch.save(resnet18, f="/tmp/resnet18.pt")
```
## Run Torch2TRT from Arachne
Now, let's optimize the model with the torch2trt by Arachne.
To use the tool, we have to specify `+tools=torch2trt` to `arachne.driver.cli`.
Available options can be seen by adding `--help`.
```
%%bash
python -m arachne.driver.cli +tools=torch2trt --help
```
### Optimize with FP32 precision
First, we will start with the simplest case.
You can optimize a TF model with FP32 precision by the following command.
Note that, the Pytorch model does not include the information about tensor specification.
So, we need to pass the YAML file indicating the shape information.
```
yml = """
inputs:
- dtype: float32
name: input
shape:
- 1
- 3
- 224
- 224
outputs:
- dtype: float32
name: output
shape:
- 1
- 16
- 224
- 224
"""
open("/tmp/resnet18.yaml", "w").write(yml)
%%bash
python -m arachne.driver.cli +tools=torch2trt model_file=/tmp/resnet18.pt model_spec_file=/tmp/resnet18.yaml output_path=/tmp/output.tar
```
### Optimize with FP16 precision
To optimize with FP16 precision, set `true` to the `tools.torch2trt.fp16_mode` option.
```
%%bash
python -m arachne.driver.cli +tools=torch2trt model_file=/tmp/resnet18.pt model_spec_file=/tmp/resnet18.yaml output_path=/tmp/output.tar tools.torch2trt.fp16_mode=true
```
### Optimize with INT8 Precision
To convert with INT8 precision, we need calibrate or estimate the range of all floating-point tensors in the model.
We provide an interface to feed the dataset to be used in the calibration.
First, we have to prepare a NPY file that contains a list of `np.ndarray` which is a dataset used for calibration.
Here, we use a dummy dataset for explanation because the IMAGENET dataset requires manual setups for users.
```
import numpy as np
datasets = []
shape = [1, 3, 224, 224]
dtype = "float32"
for _ in range(100):
datasets.append(np.random.rand(*shape).astype(np.dtype(dtype))) # type: ignore
np.save("/tmp/calib_dataset.npy", datasets)
```
Next, specify `true` to the `tools.torch2trt.int8_mode` option and pass the NPY file to the `tools.torch2trt.int8_calib_dataset`.
```
%%bash
python -m arachne.driver.cli +tools=torch2trt model_file=/tmp/resnet18.pt model_spec_file=/tmp/resnet18.yaml output_path=/tmp/output.tar \
tools.torch2trt.int8_mode=true tools.torch2trt.int8_calib_dataset=/tmp/calib_dataset.npy
```
## Run Torch2TRT from Arachne Python Interface
The following code shows an example of using the tool from Arachne Python interface.
```
from arachne.data import ModelSpec, TensorSpec
from arachne.utils.model_utils import init_from_file, save_model
from arachne.tools.torch2trt import Torch2TRT, Torch2TRTConfig
model_file_path = "/tmp/resnet18.pt"
input = init_from_file(model_file_path)
spec = ModelSpec(
inputs=[TensorSpec(name="input0", shape=[1, 3, 224, 224], dtype="float32")],
outputs=[TensorSpec(name="output0", shape=[1, 1000], dtype="float32")],
)
input.spec = spec
cfg = Torch2TRTConfig()
# cfg.fp16_mode = True
output = Torch2TRT.run(input, cfg)
save_model(model=output, output_path="/tmp/output.tar")
```
| github_jupyter |
<span style="font-size:36px"><b>Foundation of Statistics</b></span>
Copyright 2019 Gunawan Lumban Gaol
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language overning permissions and limitations under the License.
# Import Packages
```
import math
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from edapy.edapy import transformation
from edapy.edapy import plotting
# Tukey's fences outlier removal using k=3
def outlier_removal(X, method='Tukey', k=3):
Q3 = X.quantile(0.75)
Q1 = X.quantile(0.25)
IQR = Q3 - Q1
upper = Q3 + 3*IQR
lower = Q1 - 3*IQR
res = (X < lower) | (X > upper)
return res
def z_score(X):
mean = X.mean()
std = X.std()
z = (X - mean) / std
return z
```
# Import Data
The original dataset can be found here at [Google Drive](https://drive.google.com/drive/folders/1Rih-ODbqoIeWsWyTKbVaJTnNjUqBb_w2). THe followings are the columns in the data.
* Kontrak : nomor kontrak
* Tipe_Kendaraan : tipe kendaraan yang dibeli
* Pekerjaan : jenis pekerjaan
* Area : area kantor tempat kredit diberikan
* Tenor : jangka waktu hutang yang diajukan (bulan)
* Bucket_Collectability : keterlambatan membayar hutang (hari)
* NPF : non performing financing
* Total_AR : jumlah pinjaman yang diterima (juta rupiah)
* LTV : loan to value (%)
* OTR : harga beli mobil setelah ditambahkan dengan seluruh pajak dan berbagai dokumen lainnya (puluh juta rupiah)
* Age : usia (tahun)
* Interest_Rate : suku bunga bank indonesia (%)
* Income : pendapatan (juta rupiah)
* DP : uang muka (juta rupiah)
* AF : amount financed (juta rupiah)
* Tujuan : tujuan pembelian
* Pendidikan : tingkat pendidikan
* Tanggungan : jumlah tanggungan keluarga
* Status : status pernikahan peminjam
* Gender : gender peminjam
* Paket : paket kredit
* Inflasi : tingkat inflasi (%)
* Segmen : segmentasi transaksi
Date accessed: October 25, 2019
```
data = pd.read_excel("xlsx/data_loan.xlsx", index_col='Kontrak')
data.shape
```
# Task
1. Ambil sample secara acak dari data loan sebanyak 80% dari banyakanya data.
2. Lakukan analisis statistik menggunakan metode-metode yang telah dipelajari di kelas ini. Analisis dilakukan sesuai kreativitas anda dengan mempertimba waktu yang tersedia.
<span style="font-size:30px">Analysis: **Predicting Amount of Total_AR application**.</span>
<br><span style="font-size:16px"><br>Model: **Linear Regression (Ordinary Least Square)**</span>
<br><span style="font-size:16px"><br>Metrics: **RMSE (Root Mean Square Error)**</span>
## Data Preprocessing
* Transform numerical datatype that is actually categorical.
* Separate numerical and categorical column
* Perform random sample of 80% of data
```
transformation.convert_to_categorical(data)
```
Create target columns
```
col_target = 'Total_AR'
cols_num = data.select_dtypes(include=[np.number]).columns
cols_cat = data.select_dtypes(exclude=[np.number]).columns
```
Sample 80% of data.
```
sample_size = int(0.8 * data.shape[0])
data_s = data.sample(sample_size, random_state=42)
print("Sample size of {}, with population size of {}. Ratio of {:2f}".format(sample_size, data.shape[0], sample_size/data.shape[0]))
```
## Data Understanding
* Check missing & null data, remove them
* Check duplicated data, remove them
```
data_s.info()
```
No missing values for current data, which is good. Now check for duplicated data.
```
data_s.index.duplicated().sum()
```
No duplicated data, we can proceed.
We see that our target variable has a looks like normal distribution, no need for extra transformation of our target variable.
### Numerical Columns
See how our variable correlates with our target variable.
```
plt.figure(figsize=(7, 7))
sns.heatmap(data_s[cols_num].corr(), annot=True, fmt='.2f')
plt.show()
filt_cols = list(cols_num[1:]) # exlude our target column
filt_cols
```
From the correlation heatmap, we can conclude, for feature engineering, decide to remove attributes which has **more than 0.7** in correlation score
* LTV vs DP, **we remove DP** since LTV has higher correlation for our target variable `Total_AR`
* OTR vs AF, **we remove OTR** since AF has higher correlation for our target variable `Total_AR`
```
cols_remove = ['DP', 'OTR']
filt_cols = [x for x in filt_cols if x not in cols_remove]
```
Reassign filt_cols to cols_num
```
cols_num = filt_cols
```
Next we want to see the distribution of attributes, for not overfitting linear regression, it is considerably best to work with attributes that has normal distribution.
* Perform log transformation for skewed variable distribution
```
plotting.distplot_numerical(data_s, filt_cols)
```
* We need to do log transformation on column: `Income`
* We need to remove outlier for column `AF`
* We **may** need to remove outlier for column `LTV`, but for now, decide not to remove it.
```
# Log transformation
data_s['Income'] = np.log10(data_s['Income'])
# Outlier removal
AF_outlier = outlier_removal(data_s['AF']) # 135 total outlier
# Removes outliers
data_s = data_s[~AF_outlier]
```
Redraw the transformed and removed outliers data.
```
plotting.distplot_numerical(data, filt_cols)
```
### Target Column Distribution
See our target column distribution after outlier removal.
```
fig, ax = plt.subplots(figsize=(15, 6))
sns.distplot(data_s[col_target], ax=ax)
plt.show()
```
### Categorical Columns
See distribution of `col_target` along each of our unique values in `cols_num`.
```
plotting.distplot_categorical(data_s, cols_cat, col_target)
```
* Potential strong predictor: `Tipe_Kendaraan`, `Area`, `Tenor`, `Bucket_Collectability`, `NPF`, `Tujuan`, `Paket`
* Potential good predictor: `Pekerjaan`, `Tanggungan`
* Not potential predictor: `Pendidikan`, `Status`, `Gender`, `Segmen` **we remove these variables**
```
cols_remove_cat = ['Pendidikan', 'Status', 'Gender', 'Segmen']
cols_cat = [x for x in cols_cat if x not in cols_remove_cat]
```
Replot to make sure of the results.
```
plotting.distplot_categorical(data_s, cols_cat, col_target)
```
## Data Processing
* Transform numerical variable and target variable to z-score
* Transform categorical variable into dummy variable, don't forget to remove one of the result from dummy variable
```
for col in cols_num+[col_target]:
data_s['z_'+col] = z_score(data_s[col])
cols_num_z = ['z_'+x for x in cols_num]
for col in cols_cat:
data_s = pd.concat([data_s, pd.get_dummies(data_s[col], prefix='d_'+col, drop_first=True)], axis=1)
cols_cat_d = [c for c in data_s.columns if 'd_' in c]
```
## Modelling
```
from sklearn.linear_model import LinearRegression
from sklearn import metrics
cols_feature = cols_num_z+cols_cat_d
X = data_s[[c for c in cols_feature]]
y = data_s['z_Total_AR']
reg = LinearRegression().fit(X, y)
coef = reg.coef_
intercept = reg.intercept_
print("y = {} + {} x".format(intercept, coef))
preds = reg.predict(X)
rmse = math.sqrt(metrics.mean_squared_error(y.values, preds))
print("Root mean squared error: {:.3f}".format(rmse))
r_square = metrics.r2_score(y.values, preds)
print("R-square: {:.3f}".format(r_square))
```
Final predictors:
```
print(cols_num_z)
print(cols_cat)
```
For a data which scales from 0 to 1, **rmse of 0.114** and **R-square of 0.987** is quite good. But we need to perform validation on the other dataset, this will not be done in this notebook.
<hr>
# Class Exercise
Below is combination of what we do during the class and some improvements in visualization.
## Data Understanding
Separate numerical columns and categorical columns for analysis
```
cols_num = data.select_dtypes(include=[np.number]).columns
cols_cat = data.select_dtypes(exclude=[np.number]).columns
```
### Detecting Outliers
```
fig, ax = plt.subplots(1, 2, figsize=(16, 3))
sns.boxplot((data['Income']), ax=ax[0])
sns.boxplot(np.log10(data['Income']), ax=ax[1])
plt.show()
```
Below use the rule of outlier > 3 z-score.
```
z_score = standardized_income
data[['Kontrak', 'Income']][np.abs(z_score) > 3].head()
```
Below example of using 1.5\*IQR rule to detect outlier.
```
Q1 = data['Income'].quantile(0.25)
Q3 = data['Income'].quantile(0.75)
IQR = Q3 - Q1
outlier_mask = (data['Income'] < (Q1 - 1.5*IQR)) | (data['Income'] > (Q3 + 1.5*IQR))
data[['Kontrak', 'Income']][outlier_mask].head()
```
### Hypothesis testing
```
from scipy.stats import ttest_ind
print(x1.std(), x2.std())
x1 = data['Income'][data['NPF'] == 'Ya']
x2 = data['Income'][data['NPF'] != 'Ya']
ttest_ind(x1, x2, equal_var=False)
fig, ax = plt.subplots(figsize=(14, 6))
sns.distplot(np.log10(x1))
sns.distplot(np.log10(x2))
plt.legend(["Performing", "NPF"])
plt.show()
```
## Data Transformation Excercises
### Edit `Tipe_Kendaraan`, flagging `Tipe_Kendaraan` as MINIBUS and NONMINIBUS.
```
data['Tipe_Kendaraan'] = ['MINIBUS' if x == 'MINIBUS' else 'NON_MINIBUS' for x in data['Tipe_Kendaraan']]
```
### Create dummy variable from `Area` columns, dropping first columns.
```
pd.concat([data, pd.get_dummies(data['Area'], prefix='Area', drop_first=True)], axis=1).head()
```
### Bin `LTV` variable into 5 bins with equal range bin.
```
pd.cut(data['LTV'], 5).head()
```
### Transformation `Income` to standard distribution, then normalized it to log scale.
```
mean = data['Income'].mean()
std = data['Income'].std() # Normalized by N-1 by default
standardized_income = np.array([(x-mean)/std for x in data['Income']])
data['Income'].describe()
pd.Series(standardized_income).describe()
def log10_negative(x, e=0.001):
return np.log10(x - min(x) + e) + min(x)
import matplotlib.pyplot as plt
import seaborn as sns
fig, ax = plt.subplots(1, 2, figsize=(14, 6))
sns.distplot(np.log10(data['Income']), ax=ax[0])
ax[0].set_title('Before log(base10) transformation')
sns.distplot(log10_negative(standardized_income), ax=ax[1])
ax[1].set_title('After log(base10) transformation')
plt.show()
```
## Modelling
```
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn import metrics
x = (data['Income'])
y = data['Total_AR']
reg = LinearRegression().fit(np.array(x).reshape(-1, 1), np.array(y).reshape(-1, 1))
coef = reg.coef_
intercept = reg.intercept_
preds = (intercept[0] + x*coef[0][0])
print("y = {} + {} x".format(intercept[0], coef[0][0]))
fig, ax = plt.subplots(figsize=(15, 8))
sns.scatterplot(x=x, y=y, ax=ax)
plt.plot(x, preds)
plt.show()
import statsmodels.formula.api as smf
mc = pd.read_excel('xlsx/data_microchip.xlsx')
mc_stats = smf.ols('number_of_component ~ year_since_1959', data=mc).fit()
print(mc_stats.summary())
```
| github_jupyter |
# Getting started with Coiled
Welcome to the getting started guide for Coiled! This notebook covers installing and setting up Coiled as well as running your first computation using Coiled.
## Launch a cluster
The first step is to spin up a Dask Cluster. In Coiled, this is done by creating a `coiled.Cluster` instance, there are [several keyword arguments](https://docs.coiled.io/user_guide/api.html#coiled.Cluster) you can use to specify the details of your cluster further. Please read the [cluster creation documentation](https://docs.coiled.io/user_guide/cluster_creation.html) to know more.
Note that we will give a name to this cluster, if you don't specify this keyword argument, clusters will be given a unique randomly generated name.
```
import coiled
cluster = coiled.Cluster(name="quickstart-example", n_workers=10)
```
Once a cluster has been created (you can see the status on your [Coiled dashboard](https://cloud.coiled.io/)), you can connect Dask to the cluster by creating a `distributed.Client` instance.
```
from dask.distributed import Client
client = Client(cluster)
client
```
## Analyze data in the cloud
Now that we have our cluster running and Dask connected to it, let's run a computation. This example will run the computation on about 84 million rows.
```
import dask.dataframe as dd
df = dd.read_csv(
"s3://nyc-tlc/trip data/yellow_tripdata_2019-*.csv",
dtype={
"payment_type": "UInt8",
"VendorID": "UInt8",
"passenger_count": "UInt8",
"RatecodeID": "UInt8",
},
storage_options={"anon": True},
blocksize="16 MiB",
).persist()
df.groupby("passenger_count").tip_amount.mean().compute()
```
## Stop a cluster
By default, clusters will shutdown after 20 minutes of inactivity. You can stop a cluster by pressing the stop button on the [Coiled dashboard](https://cloud.coiled.io/). Alternatively, we can get a list of all running clusters and use the cluster name to stop it.
```
coiled.list_clusters()
```
The command `list_clusters` returns a dictionary with the cluster name used as the key. We can grab that and then call the command `coiled.delete_cluster()` to stop the running cluster, and `client.close()` to close the client.
```
coiled.delete_cluster(name="quickstart-example")
client.close()
```
You can now go back to the [Coiled dashboard](https://cloud.coiled.io/) and you will see that the cluster is now stopping/stopped
# Software Environments
Software Environments are Docker images that contain all your dependencies and files that you might need to run your computations. If you don't specify a software environment to the `coiled.Cluster` constructor, we will use Coiled's default software environment. You can learn more about software environments in our [documentation](https://docs.coiled.io/user_guide/software_environment.html).
## Create a software environment
When creating software environments, there are [several keyword arguments](https://docs.coiled.io/user_guide/api.html#coiled.create_software_environment) that you can use to create a custom environment for your work.
```
coiled.create_software_environment(
name="quickstart",
conda={
"channels": ["conda-forge"],
"dependencies": ["coiled"]
}
)
```
We can now follow our previous workflow of creating a cluster - this time, we will use our newly created software environment - connect the cluster to Dask and then running the same example.
```
cluster = coiled.Cluster(n_workers=10, software="quickstart")
client = Client(cluster)
client
```
If you go to the [Coiled dashboard](https://cloud.coiled.io/), under the **Software Environment** column, you can see that we are using the quickstart software environment we have just created. Note also that this time, the cluster will have a randomly generated name.
Let's now run the same computation as before, but using the cluster that is running with the software environment that we have recently created.
```
df = dd.read_csv(
"s3://nyc-tlc/trip data/yellow_tripdata_2019-*.csv",
dtype={
"payment_type": "UInt8",
"VendorID": "UInt8",
"passenger_count": "UInt8",
"RatecodeID": "UInt8",
},
storage_options={"anon": True},
blocksize="16 MiB",
).persist()
df.groupby("passenger_count").tip_amount.mean().compute()
```
| github_jupyter |

<a href="https://hub.callysto.ca/jupyter/hub/user-redirect/git-pull?repo=https%3A%2F%2Fgithub.com%2Fcallysto%2Fhackathon&branch=master&subPath=ColonizingMars/AccessingData/nasa-pictures-of-mars.ipynb&depth=1" target="_parent"><img src="https://raw.githubusercontent.com/callysto/curriculum-notebooks/master/open-in-callysto-button.svg?sanitize=true" width="123" height="24" alt="Open in Callysto"/></a>
# Pictures of Mars from NASA
Source: https://api.nasa.gov/
NASA API: The objective of this site is to make NASA data, including imagery, accessible to application developers
## Mars Rover Photos
[Text taken directly from their dataset description]
This API is designed to collect image data gathered by NASA's Curiosity, Opportunity, and Spirit rovers on Mars and make it more easily available to other developers, educators, and citizen scientists. This API is maintained by Chris Cerami.
Each rover has its own set of photos stored in the database, which can be queried separately. There are several possible queries that can be made against the API. Photos are organized by the sol (Martian rotation or day) on which they were taken, counting up from the rover's landing date. A photo taken on Curiosity's 1000th Martian sol exploring Mars, for example, will have a sol attribute of 1000. If instead you prefer to search by the Earth date on which a photo was taken, you can do that too.
Along with querying by date, results can also be filtered by the camera with which it was taken and responses will be limited to 25 photos per call. Queries that should return more than 25 photos will be split onto several pages, which can be accessed by adding a 'page' param to the query.
```
camera_list = ['FHAZ', 'RHAZ', 'MAST', 'CHEMCAM', 'NAVCAM']
camera_name = camera_list[0]
DEMO_KEY='MCPiQTUOtrm8K1bZAXY2UIYVB3JNcunfwkEd2IYm'
base_url = "https://api.nasa.gov/mars-photos/api/v1/rovers/curiosity/"
build_query = "photos?sol=1000&camera=" + camera_name + "&api_key=" + DEMO_KEY
full_url = base_url + build_query
import requests
resp = requests.get(url=full_url)
data = resp.json()
import pandas as pd
df = pd.io.json.json_normalize(data,record_path="photos") # or pd.json_normalize with Pandas > 1.0
df
from IPython.display import Image
for image_url in df['img_src']:
print(image_url)
image = Image(url=image_url)
display(image)
```
[](https://github.com/callysto/curriculum-notebooks/blob/master/LICENSE.md)
| github_jupyter |
## Fixed End Forces
This module computes the fixed end forces (moments and shears) due to transverse loads
acting on a 2-D planar structural member.
```
import numpy as np
import sys
from salib import extend
```
### Class EF
Instances of class **EF** represent the 6 end-forces for a 2-D planar beam element.
The forces (and local degrees of freedom) are numbered 0 through 5, and are shown here in their
positive directions on a beam-element of length **L**. The 6 forces are labelled by prefixing the number with a letter to suggest the normal interpretation of that force: **c** for axial force,
**v** for shear force, and **m** for moment.

For use in this module, the end forces will be *fixed-end-forces*.
```
class EF(object):
"""Class EF represents the 6 end forces acting on a 2-D, planar, beam element."""
def __init__(self,c0=0.,v1=0.,m2=0.,c3=0.,v4=0.,m5=0.):
"""Initialize an instance with the 6 end forces. If the first
argument is a 6-element array, initialize from a copy of that
array and ignore any other arguments."""
if np.isscalar(c0):
self.fefs = np.matrix([c0,v1,m2,c3,v4,m5],dtype=np.float64).T
else:
self.fefs = c0.copy()
def __getitem__(self,ix):
"""Retreive one of the forces by numer. This allows allows unpacking
of all 6 end forces into 6 variables using something like:
c0,v1,m2,c3,v4,m5 = self
"""
return self.fefs[ix,0]
def __add__(self,other):
"""Add this set of end forces to another, returning the sum."""
assert type(self) is type(other)
new = self.__class__(self.fefs+other.fefs)
return new
def __sub__(self,other):
"""Subtract the other from this set of forces, returning the difference."""
assert type(self) is type(other)
new = self.__class__(self.fefs-other.fefs)
return new
def __mul__(self,scale):
"""Multiply this set of forces by the scalar value, returning the product."""
if scale == 1.0:
return self
return self.__class__(self.fefs*scale)
__rmul__ = __mul__
def __repr__(self):
return '{}({},{},{},{},{},{})'.format(self.__class__.__name__,*list(np.array(self.fefs.T)[0]))
##test:
f = EF(1,2,0,4,1,6)
f
##test:
g = f+f+f
g
##test:
f[1]
##test:
f[np.ix_([3,0,1])]
##test:
g[(3,0,1)]
##test:
f0,f1,f2,f3,f4,f5 = g
f3
##test:
g, g*5, 5*g
```
Now define properties so that the individual components can be accessed like name atrributes,
eg: '`ef.m3`' or '`ef.m5 = 100.`'.
```
@extend
class EF:
@property
def c0(self):
return self.fefs[0,0]
@c0.setter
def c0(self,v):
self.fefs[0,0] = v
@property
def v1(self):
return self.fefs[1,0]
@v1.setter
def v1(self,v):
self.fefs[1,0] = v
@property
def m2(self):
return self.fefs[2,0]
@m2.setter
def m2(self,v):
self.fefs[2,0] = v
@property
def c3(self):
return self.fefs[3,0]
@c3.setter
def c3(self,v):
self.fefs[3,0] = v
@property
def v4(self):
return self.fefs[4,0]
@v4.setter
def v4(self,v):
self.fefs[4,0] = v
@property
def m5(self):
return self.fefs[5,0]
@m5.setter
def m5(self,v):
self.fefs[5,0] = v
##test:
f = EF(10.,11,12,13,15,15)
f, f.c0, f.v1, f.m2, f.c3, f.v4, f.m5
##test:
f.c0 *= 2
f.v1 *= 3
f.m2 *= 4
f.c3 *= 5
f.v4 *= 6
f.m5 *= 7
f
```
## Class MemberLoad
This is the base class for all the different types of member loads (point loads, UDLs, etc.)
of 2D planar beam elements.
The main purpose is to calculate the fixed-end member forces, but we will also supply
logic to enable calculation of internal shears and moments at any point along the span.
All types of member loads will be input using a table containing five data columns:
**W1**, **W2**, **A**, **B**, and **C**. Each load type contains a '**TABLE_MAP**'
that specifies the mapping between attribute name and column name in the table.
```
class MemberLoad(object):
TABLE_MAP = {} # map from load parameter names to column names in table
def fefs(self):
"""Return the complete set of 6 fixed end forces produced by the load."""
raise NotImplementedError()
def shear(self,x):
"""Return the shear force that is in equilibrium with that
produced by the portion of the load to the left of the point at
distance 'x'. 'x' may be a scalar or a 1-dimensional array
of values."""
raise NotImplementedError()
def moment(self,x):
"""Return the bending moment that is in equilibrium with that
produced by the portion of the load to the left of the point at
distance 'x'. 'x' may be a scalar or a 1-dimensional array
of values."""
raise NotImplementedError()
@extend
class MemberLoad:
@property
def vpts(self):
"""Return a descriptor of the points at which the shear force must
be evaluated in order to draw a proper shear force diagram for this
load. The descriptor is a 3-tuple of the form: (l,r,d) where 'l'
is the leftmost point, 'r' is the rightmost point and 'd' is the
degree of the curve between. One of 'r', 'l' may be None."""
raise NotImplementedError()
@property
def mpts(self):
"""Return a descriptor of the points at which the moment must be
evaluated in order to draw a proper bending moment diagram for this
load. The descriptor is a 3-tuple of the form: (l,r,d) where 'l'
is the leftmost point, 'r' is the rightmost point and 'd' is the
degree of the curve between. One of 'r', 'l' may be None."""
raise NotImplementedError()
```
#### Load Type PL
Load type **PL** represents a single concentrated force, of magnitude **P**, at a distance **a** from the *j-end*:

```
class PL(MemberLoad):
TABLE_MAP = {'P':'W1','a':'A'}
def __init__(self,L,P,a):
self.L = L
self.P = P
self.a = a
def fefs(self):
P = self.P
L = self.L
a = self.a
b = L-a
m2 = -P*a*b*b/(L*L)
m5 = P*a*a*b/(L*L)
v1 = (m2 + m5 - P*b)/L
v4 = -(m2 + m5 + P*a)/L
return EF(0.,v1,m2,0.,v4,m5)
def shear(self,x):
return -self.P*(x>self.a)
def moment(self,x):
return self.P*(x-self.a)*(x>self.a)
def __repr__(self):
return '{}(L={},P={},a={})'.format(self.__class__.__name__,self.L,self.P,self.a)
##test:
p = PL(1000.,300.,400.)
p, p.fefs()
@extend
class MemberLoad:
EPSILON = 1.0E-6
@extend
class PL:
@property
def vpts(self):
return (self.a-self.EPSILON,self.a+self.EPSILON,0)
@property
def mpts(self):
return (self.a,None,1)
##test:
p = PL(1000.,300.,400.)
p.vpts
##test:
p.mpts
```
#### Load Type PLA
Load type **PLA** represents a single concentrated force applied parallel to the length
of the segment (producing only axial forces).

```
class PLA(MemberLoad):
TABLE_MAP = {'P':'W1','a':'A'}
def __init__(self,L,P,a):
self.L = L
self.P = P
self.a = a
def fefs(self):
P = self.P
L = self.L
a = self.a
c0 = -P*(L-a)/L
c3 = -P*a/L
return EF(c0=c0,c3=c3)
def shear(self,x):
return 0.
def moment(self,x):
return 0.
def __repr__(self):
return '{}(L={},P={},a={})'.format(self.__class__.__name__,self.L,self.P,self.a)
##test:
p = PLA(10.,P=100.,a=4.)
p.fefs()
@extend
class PLA:
@property
def vpts(self):
return (0.,self.L,0)
@property
def mpts(self):
return (0.,self.L,0)
```
#### Load Type UDL
Load type **UDL** represents a uniformly distributed load, of magnitude **w**, over the complete length of the element.

```
class UDL(MemberLoad):
TABLE_MAP = {'w':'W1'}
def __init__(self,L,w):
self.L = L
self.w = w
def __repr__(self):
return '{}(L={},w={})'.format(self.__class__.__name__,self.L,self.w)
def fefs(self):
L = self.L
w = self.w
return EF(0.,-w*L/2., -w*L*L/12., 0., -w*L/2., w*L*L/12.)
def shear(self,x):
l = x*(x>0.)*(x<=self.L) + self.L*(x>self.L) # length of loaded portion
return -(l*self.w)
def moment(self,x):
l = x*(x>0.)*(x<=self.L) + self.L*(x>self.L) # length of loaded portion
d = (x-self.L)*(x>self.L) # distance from loaded portion to x: 0 if x <= L else x-L
return self.w*l*(l/2.+d)
@property
def vpts(self):
return (0.,self.L,1)
@property
def mpts(self):
return (0.,self.L,2)
##test:
w = UDL(12,10)
w,w.fefs()
```
#### Load Type LVL
Load type **LVL** represents a linearly varying distributed load actiong over a portion of the span:

```
class LVL(MemberLoad):
TABLE_MAP = {'w1':'W1','w2':'W2','a':'A','b':'B','c':'C'}
def __init__(self,L,w1,w2=None,a=None,b=None,c=None):
if a is not None and b is not None and c is not None and L != (a+b+c):
raise Exception('Cannot specify all of a, b & c')
if a is None:
if b is not None and c is not None:
a = L - (b+c)
else:
a = 0.
if c is None:
if b is not None:
c = L - (a+b)
else:
c = 0.
if b is None:
b = L - (a+c)
if w2 is None:
w2 = w1
self.L = L
self.w1 = w1
self.w2 = w2
self.a = a
self.b = b
self.c = c
def fefs(self):
"""This mess was generated via sympy. See:
../../examples/cive3203-notebooks/FEM-2-Partial-lvl.ipynb """
L = float(self.L)
a = self.a
b = self.b
c = self.c
w1 = self.w1
w2 = self.w2
m2 = -b*(15*a*b**2*w1 + 5*a*b**2*w2 + 40*a*b*c*w1 + 20*a*b*c*w2 + 30*a*c**2*w1 + 30*a*c**2*w2 + 3*b**3*w1 + 2*b**3*w2 + 10*b**2*c*w1 + 10*b**2*c*w2 + 10*b*c**2*w1 + 20*b*c**2*w2)/(60.*(a + b + c)**2)
m5 = b*(20*a**2*b*w1 + 10*a**2*b*w2 + 30*a**2*c*w1 + 30*a**2*c*w2 + 10*a*b**2*w1 + 10*a*b**2*w2 + 20*a*b*c*w1 + 40*a*b*c*w2 + 2*b**3*w1 + 3*b**3*w2 + 5*b**2*c*w1 + 15*b**2*c*w2)/(60.*(a + b + c)**2)
v4 = -(b*w1*(a + b/2.) + b*(a + 2*b/3.)*(-w1 + w2)/2. + m2 + m5)/L
v1 = -b*(w1 + w2)/2. - v4
return EF(0.,v1,m2,0.,v4,m5)
def __repr__(self):
return '{}(L={},w1={},w2={},a={},b={},c={})'\
.format(self.__class__.__name__,self.L,self.w1,self.w2,self.a,self.b,self.c)
def shear(self,x):
c = (x>self.a+self.b) # 1 if x > A+B else 0
l = (x-self.a)*(x>self.a)*(1.-c) + self.b*c # length of load portion to the left of x
return -(self.w1 + (self.w2-self.w1)*(l/self.b)/2.)*l
def moment(self,x):
c = (x>self.a+self.b) # 1 if x > A+B else 0
# note: ~c doesn't work if x is scalar, thus we use 1-c
l = (x-self.a)*(x>self.a)*(1.-c) + self.b*c # length of load portion to the left of x
d = (x-(self.a+self.b))*c # distance from right end of load portion to x
return ((self.w1*(d+l/2.)) + (self.w2-self.w1)*(l/self.b)*(d+l/3.)/2.)*l
@property
def vpts(self):
return (self.a,self.a+self.b,1 if self.w1==self.w2 else 2)
@property
def mpts(self):
return (self.a,self.a+self.b,2 if self.w1==self.w2 else 3)
```
#### Load Type CM
Load type **CM** represents a single concentrated moment of magnitude **M** a distance **a** from the j-end:

```
class CM(MemberLoad):
TABLE_MAP = {'M':'W1','a':'A'}
def __init__(self,L,M,a):
self.L = L
self.M = M
self.a = a
def fefs(self):
L = float(self.L)
A = self.a
B = L - A
M = self.M
m2 = B*(2.*A - B)*M/L**2
m5 = A*(2.*B - A)*M/L**2
v1 = (M + m2 + m5)/L
v4 = -v1
return EF(0,v1,m2,0,v4,m5)
def shear(self,x):
return x*0.
def moment(self,x):
return -self.M*(x>self.A)
@property
def vpts(self):
return (None,None,0)
@property
def mpts(self):
return (self.A-self.EPSILON,self.A+self.EPSILON,1)
def __repr__(self):
return '{}(L={},M={},a={})'.format(self.__class__.__name__,self.L,self.M,self.a)
```
### makeMemberLoad() factory function
Finally, the function `makeMemberLoad()` will create a load object of the correct type from
the data in dictionary `data`. That dictionary would normally containing the data from one
row ov the input data file table.
```
def makeMemberLoad(L,data,ltype=None):
def all_subclasses(cls):
_all_subclasses = []
for subclass in cls.__subclasses__():
_all_subclasses.append(subclass)
_all_subclasses.extend(all_subclasses(subclass))
return _all_subclasses
if ltype is None:
ltype = data.get('TYPE',None)
for c in all_subclasses(MemberLoad):
if c.__name__ == ltype and hasattr(c,'TABLE_MAP'):
MAP = c.TABLE_MAP
argv = {k:data[MAP[k]] for k in MAP.keys()}
return c(L,**argv)
raise Exception('Invalid load type: {}'.format(ltype))
##test:
ml = makeMemberLoad(12,{'TYPE':'UDL', 'W1':10})
ml, ml.fefs()
def unmakeMemberLoad(load):
type = load.__class__.__name__
ans = {'TYPE':type}
for a,col in load.TABLE_MAP.items():
ans[col] = getattr(load,a)
return ans
##test:
unmakeMemberLoad(ml)
```
| github_jupyter |
<a name="building-language-model"></a>
# Building the language model
<a name="count-matrix"></a>
### Count matrix
To calculate the n-gram probability, you will need to count frequencies of n-grams and n-gram prefixes in the training dataset. In some of the code assignment exercises, you will store the n-gram frequencies in a dictionary.
In other parts of the assignment, you will build a count matrix that keeps counts of (n-1)-gram prefix followed by all possible last words in the vocabulary.
The following code shows how to check, retrieve and update counts of n-grams in the word count dictionary.
```
# manipulate n_gram count dictionary
n_gram_counts = {
('i', 'am', 'happy'): 2,
('am', 'happy', 'because'): 1}
# get count for an n-gram tuple
print(f"count of n-gram {('i', 'am', 'happy')}: {n_gram_counts[('i', 'am', 'happy')]}")
# check if n-gram is present in the dictionary
if ('i', 'am', 'learning') in n_gram_counts:
print(f"n-gram {('i', 'am', 'learning')} found")
else:
print(f"n-gram {('i', 'am', 'learning')} missing")
# update the count in the word count dictionary
n_gram_counts[('i', 'am', 'learning')] = 1
if ('i', 'am', 'learning') in n_gram_counts:
print(f"n-gram {('i', 'am', 'learning')} found")
else:
print(f"n-gram {('i', 'am', 'learning')} missing")
```
The next code snippet shows how to merge two tuples in Python. That will be handy when creating the n-gram from the prefix and the last word.
```
# concatenate tuple for prefix and tuple with the last word to create the n_gram
prefix = ('i', 'am', 'happy')
word = 'because'
# note here the syntax for creating a tuple for a single word
n_gram = prefix + (word,)
print(n_gram)
```
In the lecture, you've seen that the count matrix could be made in a single pass through the corpus. Here is one approach to do that.
```
import numpy as np
import pandas as pd
from collections import defaultdict
def single_pass_trigram_count_matrix(corpus):
"""
Creates the trigram count matrix from the input corpus in a single pass through the corpus.
Args:
corpus: Pre-processed and tokenized corpus.
Returns:
bigrams: list of all bigram prefixes, row index
vocabulary: list of all found words, the column index
count_matrix: pandas dataframe with bigram prefixes as rows,
vocabulary words as columns
and the counts of the bigram/word combinations (i.e. trigrams) as values
"""
bigrams = []
vocabulary = []
count_matrix_dict = defaultdict(dict)
# go through the corpus once with a sliding window
for i in range(len(corpus) - 3 + 1):
# the sliding window starts at position i and contains 3 words
trigram = tuple(corpus[i : i + 3])
bigram = trigram[0 : -1]
if not bigram in bigrams:
bigrams.append(bigram)
last_word = trigram[-1]
if not last_word in vocabulary:
vocabulary.append(last_word)
if (bigram,last_word) not in count_matrix_dict:
count_matrix_dict[bigram,last_word] = 0
count_matrix_dict[bigram,last_word] += 1
# convert the count_matrix to np.array to fill in the blanks
count_matrix = np.zeros((len(bigrams), len(vocabulary)))
for trigram_key, trigam_count in count_matrix_dict.items():
count_matrix[bigrams.index(trigram_key[0]), \
vocabulary.index(trigram_key[1])]\
= trigam_count
# np.array to pandas dataframe conversion
count_matrix = pd.DataFrame(count_matrix, index=bigrams, columns=vocabulary)
return bigrams, vocabulary, count_matrix
corpus = ['i', 'am', 'happy', 'because', 'i', 'am', 'learning', '.']
bigrams, vocabulary, count_matrix = single_pass_trigram_count_matrix(corpus)
print(count_matrix)
```
<a name="probability-matrix"></a>
### Probability matrix
The next step is to build a probability matrix from the count matrix.
You can use an object dataframe from library pandas and its methods [sum](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.sum.html?highlight=sum#pandas.DataFrame.sum) and [div](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.div.html) to normalize the cell counts with the sum of the respective rows.
```
# create the probability matrix from the count matrix
row_sums = count_matrix.sum(axis=1)
# delete each row by its sum
prob_matrix = count_matrix.div(row_sums, axis=0)
print(prob_matrix)
```
The probability matrix now helps you to find a probability of an input trigram.
```
# find the probability of a trigram in the probability matrix
trigram = ('i', 'am', 'happy')
# find the prefix bigram
bigram = trigram[:-1]
print(f'bigram: {bigram}')
# find the last word of the trigram
word = trigram[-1]
print(f'word: {word}')
# we are using the pandas dataframes here, column with vocabulary word comes first, row with the prefix bigram second
trigram_probability = prob_matrix[word][bigram]
print(f'trigram_probability: {trigram_probability}')
```
In the code assignment, you will be searching for the most probable words starting with a prefix. You can use the method [str.startswith](https://docs.python.org/3/library/stdtypes.html#str.startswith) to test if a word starts with a prefix.
Here is a code snippet showing how to use this method.
```
# lists all words in vocabulary starting with a given prefix
vocabulary = ['i', 'am', 'happy', 'because', 'learning', '.', 'have', 'you', 'seen','it', '?']
starts_with = 'ha'
print(f'words in vocabulary starting with prefix: {starts_with}\n')
for word in vocabulary:
if word.startswith(starts_with):
print(word)
```
<a name="language-model-evaluation"></a>
## Language model evaluation
<a name="train-validation-test-split"></a>
### Train/validation/test split
In the videos, you saw that to evaluate language models, you need to keep some of the corpus data for validation and testing.
The choice of the test and validation data should correspond as much as possible to the distribution of the data coming from the actual application. If nothing but the input corpus is known, then random sampling from the corpus is used to define the test and validation subset.
Here is a code similar to what you'll see in the code assignment. The following function allows you to randomly sample the input data and return train/validation/test subsets in a split given by the method parameters.
```
# we only need train and validation %, test is the remainder
import random
def train_validation_test_split(data, train_percent, validation_percent):
"""
Splits the input data to train/validation/test according to the percentage provided
Args:
data: Pre-processed and tokenized corpus, i.e. list of sentences.
train_percent: integer 0-100, defines the portion of input corpus allocated for training
validation_percent: integer 0-100, defines the portion of input corpus allocated for validation
Note: train_percent + validation_percent need to be <=100
the reminder to 100 is allocated for the test set
Returns:
train_data: list of sentences, the training part of the corpus
validation_data: list of sentences, the validation part of the corpus
test_data: list of sentences, the test part of the corpus
"""
# fixed seed here for reproducibility
random.seed(87)
# reshuffle all input sentences
random.shuffle(data)
train_size = int(len(data) * train_percent / 100)
train_data = data[0:train_size]
validation_size = int(len(data) * validation_percent / 100)
validation_data = data[train_size:train_size + validation_size]
test_data = data[train_size + validation_size:]
return train_data, validation_data, test_data
data = [x for x in range (0, 100)]
train_data, validation_data, test_data = train_validation_test_split(data, 80, 10)
print("split 80/10/10:\n",f"train data:{train_data}\n", f"validation data:{validation_data}\n",
f"test data:{test_data}\n")
train_data, validation_data, test_data = train_validation_test_split(data, 98, 1)
print("split 98/1/1:\n",f"train data:{train_data}\n", f"validation data:{validation_data}\n",
f"test data:{test_data}\n")
```
<a name="perplexity"></a>
### Perplexity
In order to implement the perplexity formula, you'll need to know how to implement m-th order root of a variable.
\begin{equation*}
PP(W)=\sqrt[M]{\prod_{i=1}^{m}{\frac{1}{P(w_i|w_{i-1})}}}
\end{equation*}
Remember from calculus:
\begin{equation*}
\sqrt[M]{\frac{1}{x}} = x^{-\frac{1}{M}}
\end{equation*}
Here is a code that will help you with the formula.
```
# to calculate the exponent, use the following syntax
p = 10 ** (-250)
M = 100
perplexity = p ** (-1 / M)
print(perplexity)
```
| github_jupyter |
```
import os
from collections import *
from numpy import *
import re
import json
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName("read_csv").getOrCreate()
sc = spark.sparkContext
data_dir = '../data_test/'
test_file_dir=os.path.join(data_dir, '20200614.csv')
result_file_dir=os.path.join(data_dir, '20200614-result.csv')
sliceNum=2
lines = sc.textFile(test_file_dir,sliceNum) # 对大文件 进行切片 sliceNum=8,否则报错
# 20/05/30 11:54:28 ERROR PythonRunner: This may have been caused by a prior exception:
# java.net.SocketException: Connection reset by peer: socket write error
print(lines.take(10))
col_num=3
# 自定义 Map 函数 ,
def process_oneline(line,col_num):
line_array=line.split("\x01") # "\x01" 为字节表示
length=len(line_array)
line_array.append(length)
res=None
if length!=col_num : # 找到 字段数目 不符合 col_num 的
res=line_array
return res # 每一行 必须都 要有 返回
linesByMap=lines.map(lambda line :process_oneline(line ,col_num ))
print(linesByMap.take(10)) # [None, ['1', ' abc \x03 ', ' 超哥 ', ' 666 ', 4], None]
# 自定义 mapPartitions
# 找到 字段数目 不匹配的行 并输出
def process_oneslice(lines_slice,col_num):
res=[]
for line in lines_slice:
line_array=line.split("\x01")
length=len(line_array)
line_array.append(length) # 记录 总的字段数目
if length!=col_num : # 找到 字段数目 不符合 col_num 的
res.append( line + str(line_array) )
return res
linesByMapPartitions=lines.mapPartitions(lambda lines_slice :process_oneslice(lines_slice ,col_num ))
# print(linesByMapPartitions.take(10)) #
# 分区合并 ,最后只写出 一个文件
one_slice=linesByMapPartitions.coalesce(1, shuffle=True)
one_slice.saveAsTextFile(result_file_dir)
# 将所有 不可见的字符 写到 文本中
data_dir = '../data_test/'
test_file_dir = os.path.join(data_dir, 'all_invisible_characters.csv') #
encoding='utf-8'
control_chars = ''.join(map(chr, list(range(0,32)) + list(range(127,160)))) # 不可见字符的 范围 为 0-32 和 127-160
# control_chars=re.escape(control_chars) #对文本(字符串)中所有可能被解释为正则运算符的字符进行转义
with open(test_file_dir , "wb+") as f_test:
f_test.seek(0, 0) # 指向 切片文件的 首位
row_text = control_chars
row_text_bytes = row_text.encode(encoding)
f_test.write(row_text_bytes)
# 字符串的 转义
s='ABC\-001'
print(s)
s = r'ABC\-001'
# 对应的正则表达式字符串不变:
print (s) #'ABC\-001'
orgin = " '!"#$%&'()*+,-./:;<=>?@[\]^_`{|}~ "
print('[{0}]'.format(re.escape(orgin)))
# 查找 文本中的 所有 不可见字符
import re
STX=chr(0x02)
SOH=chr(0x01)
text='a0'+STX+SOH
print(text)
reg_illegal = re.compile("\x01") # 字节表示
# reg_illegal = re.compile(SOH) # 字符表示
print(reg_illegal.search(text))
control_chars = ''.join(map(chr, list(range(0,32)) + list(range(127,160)))) # 不可见字符的 范围 为 0-32 和 127-160
reg_illegal= re.compile('[%s]' % re.escape(control_chars))
print(reg_illegal.findall(text))
# 清除 文本中的 所有 不可见字符
STX=chr(0x02)
SOH=chr(0x01)
original_json='a0'+STX+SOH
def remove_control_chars(s):
control_chars = ''.join(map(chr, list(range(0,32)) + list(range(127,160)))) # 不可见字符的 范围 为 0-32 和 127-160
control_chars = re.compile('[%s]' % re.escape(control_chars))
return control_chars.sub('', s)
cleaned_json = remove_control_chars(original_json)
print(cleaned_json)
# 自定义 mapPartitions
# 找到 字段 中出现 非法字符 的行 并输出
anti_Slash=chr(0x5C) #反斜杠
# print(anti_Slash)
SOH=chr(0x01)
STX=chr(0x02)
reg_illegal = re.compile("\x02")
def process_oneslice_find_illegalChar(lines_slice,reg_illegal):
res=[]
for line in lines_slice:
line_array=line.split("\x01")
exist=False
for field in line_array: # 遍历 一行中的 每一个字段
if reg_illegal.search(field) !=None:
res.append( line )
break
return res
linesByMapPartitions=lines.mapPartitions(lambda lines_slice :process_oneslice_find_illegalChar(lines_slice ,reg_illegal ))
# 分区合并
one_slice=linesByMapPartitions.coalesce(1, shuffle=True)
one_slice.saveAsTextFile(result_file_dir)
# python 实现 flatMap
def flatten_single_dim(mapped):
for item in mapped:
for subitem in item:
yield subitem
table_a=[['A', '1'], ['B', '2'], ['C', '3'], ['D', '4'],['E','5']]
list(flatten_single_dim(table_a))
#利用 groupByKey 算子 实现 mapreduce 的 join
# table_a=[['A', '1'], ['B', '2'], ['C', '3'], ['D', '4'],['E','5']]
# table_b=[['A', 'a'], ['B', 'b'], ['C', 'c'], ['D', 'd']]
# table_a = sc.parallelize(table_a,3)
# table_b = sc.parallelize(table_b,2)
data_dir = '../data_test/'
table_a_dir=os.path.join(data_dir, 'table_A')
table_b_dir=os.path.join(data_dir, 'table_B')
table_dir=os.path.join(data_dir, 'table')
sliceNum=2
table_a = sc.textFile(table_a_dir,sliceNum)
table_b = sc.textFile(table_b_dir,sliceNum)
table_a=table_a.map(lambda line : line.split(','))
table_b=table_b.map(lambda line : line.split(','))
table_a=table_a.map(lambda line: (line[0],line[1:])) # 只能有 两个元素 ,第1个为 Key; 否则后面的 groupByKey() 报错
table_b=table_b.map(lambda line: (line[0],line[1:]))
table=table_a.union(table_b) # 合并后 分区 数目 也是 两个 RDD 的分区的和
table=table.groupByKey() # all the *byKey methods(reduce) operate on PairwiseRDDs. In Python it means RDD which contains tuples of length 2.
# partitionBy requires a PairwiseRDD which in Python is equivalent to RDD of tuples (lists) of length 2 where the first element is a key and the second one is a value.
# table.take(10)
def flatten_single_dim(mapped):
for item in mapped:
for subitem in item:
yield subitem
def process_oneslice(lines_slice,col_num):
res=[]
for line in lines_slice:
key=line[0]
values=[]
for col in line[1]:
values.append(col)
if len(values)== col_num-1: # col 匹配 说明 关联成功
res.append( [key]+ list(flatten_single_dim(values)) )
return res
col_num=3
table=table.mapPartitions(lambda lines_slice : process_oneslice(lines_slice,col_num))
# print(table.take(10))
table_one_slice=table.map(lambda line : ",".join(line) ).coalesce(1, shuffle=True) # 输出为 一个切片
table_one_slice.saveAsTextFile(table_dir)
# 利用 基本 算子 实现 mapreduce 的 join
data_dir = '../data_test/'
table_a_dir=os.path.join(data_dir, 'table_A')
table_b_dir=os.path.join(data_dir, 'table_B')
table_dir=os.path.join(data_dir, 'table')
sliceNum=2
table_a = sc.textFile(table_a_dir,sliceNum)
table_b = sc.textFile(table_b_dir,sliceNum)
table_a=table_a.map(lambda line : line.split(','))
table_b=table_b.map(lambda line : line.split(','))
table_a=table_a.map(lambda line: (line[0],line[1:])) # 只能有 两个元素 ,第1个为 Key; 否则后面的 groupByKey() 报错
table_b=table_b.map(lambda line: (line[0],line[1:]))
table=table_a.union(table_b) # 合并后 分区 数目 也是 两个 RDD 的分区的和
# table.glom().collect() # 输出 各个分区 的元素 列表
# [[('1', ['a', '27']), ('2', ['b', '24']), ('3', ['c', '23'])],
# [('4', ['d', '21']), ('5', ['e', '22']), ('6', ['f', '20'])],
# [('1', ['male']), ('2', ['female'])],
# [('4', ['female']), ('5', ['male'])]]
table=table.partitionBy(2)
# table.glom().collect()
# [[('1', ['a', '27']), ('4', ['d', '21']), ('1', ['male']), ('4', ['female'])],
# [('2', ['b', '24']),
# ('3', ['c', '23']),
# ('5', ['e', '22']),
# ('6', ['f', '20']),
# ('2', ['female']),
# ('5', ['male'])]]
def process_oneslice(one_slice,col_num):
res=[]
hash_table={}
for line in one_slice:
key=line[0]
value=line[1]
if key not in hash_table:
hash_table[key]=value
else:
hash_table[key]= hash_table[key]+value
for key, value in hash_table.items():
if len(value) == col_num: # 这一行的 col 个数 匹配 说明 关联成功
res.append([key]+value)
return res
col_num=3 # 最终表 除了 Key 之外 应该有 3 个列(字段)
table=table.mapPartitions(lambda one_slice : process_oneslice(one_slice,col_num))
# table.glom().collect()
table_one_slice=table.map(lambda line : ",".join(line) ).coalesce(1, shuffle=True) # 输出为 一个切片
table_one_slice.saveAsTextFile(table_dir)
# 利用 基本 算子 实现 hash join
data_dir = '../data_test/'
table_a_dir=os.path.join(data_dir, 'table_A')
table_b_dir=os.path.join(data_dir, 'table_B')
table_dir=os.path.join(data_dir, 'table')
sliceNum = 2
table_a = sc.textFile(table_a_dir, sliceNum)
table_b = sc.textFile(table_b_dir, sliceNum)
table_a = table_a.map(lambda line: line.split(',')) # 大表
table_b = table_b.map(lambda line: line.split(',')) # 小表
table_a = table_a.map(lambda line: (line[0], line[1:])) # 只能有 两个元素 ,第1个为 Key; 否则后面的 groupByKey() 报错
table_b = table_b.map(lambda line: (line[0], line[1:]))
table_b=table_b.collect() #[('1', ['male']), ('2', ['female']), ('4', ['female']), ('5', ['male'])]
hash_table_b={}
for line in table_b:
hash_table_b[line[0]]=line[1][0]
# 把小表 作为 广播变量 分发到各个 计算节点上
broadcast_table_b = sc.broadcast(hash_table_b) # SPARK-5063: RDD 不能被广播
def process_oneslice(big_table_slice):
res=[]
for line in big_table_slice:
key=line[0]
values=line[1]
if key in hash_table_b:
res.append( [key]+[hash_table_b[key]] + values )
return res
table=table_a.mapPartitions(lambda big_table_slice : process_oneslice(big_table_slice))
# table.collect()
table_one_slice=table.map(lambda line : ",".join(line) ).coalesce(1, shuffle=True) # 输出为 一个切片
table_one_slice.saveAsTextFile(table_dir)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
#import lightgbm as lgb
from sklearn.model_selection import KFold
import warnings
import gc
import time
import sys
import datetime
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import mean_squared_error
warnings.simplefilter(action='ignore', category=FutureWarning)
warnings.filterwarnings('ignore')
from sklearn import metrics
import scipy.stats as stats
from sklearn.model_selection import permutation_test_score
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer
from sklearn.base import BaseEstimator, ClassifierMixin
from sklearn.preprocessing import FunctionTransformer
from sklearn.preprocessing import OneHotEncoder
from sklearn.impute import SimpleImputer
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
plt.style.use('seaborn')
sns.set(font_scale=2)
pd.set_option('display.max_columns', 500)
def analysis(col, tops = 10):
temp = train[col].value_counts()
temp = temp.iloc[:tops].index
#temp = train.index
temp_df = train[train[col].isin(temp)]
# prob = temp_df[col].value_counts(normalize=True)
# draw = np.random.choice(prob.index, p=prob, size=len(temp_df))
# output = pd.Series(draw).value_counts(normalize=True).rename('simulated')
# zeros = set(temp_df[col].dropna().unique()).difference(set(output.index))
# output = output.append(pd.Series([0 for i in zeros], index = zeros)) / (temp_df[col].value_counts())
temp_df['shuffle'] = temp_df['HasDetections'].sample(replace=False, n=len(temp_df)).reset_index(drop=True)
output = temp_df[temp_df['shuffle'] == 1][col].value_counts() / temp_df[col].value_counts()
pd.DataFrame({'train_data': temp_df[temp_df['HasDetections'] == 1][col].value_counts()/ temp_df[col].value_counts(),
'random_data': output}).plot(kind = 'bar', figsize=(20,10))
plt.title('Percent of Has detections by {} (most of the catogaries)'.format(col))
display(pd.DataFrame({'train_data': temp_df[temp_df['HasDetections'] == 1][col].value_counts()/ temp_df[col].value_counts(),
'random_data': output}))
return stats.ks_2samp(temp_df[temp_df['HasDetections'] == 1][col].value_counts(normalize = True),
output)
#stats.chi2_contingency([temp_df.groupby(col).HasDetections.mean(),
# temp_df.groupby(col).random_data.mean()])
COLS = [
'HasDetections',
'AVProductStatesIdentifier','AVProductsInstalled', 'AVProductsEnabled'
]
train = pd.read_csv("train.csv", sep=',', engine='c', usecols=COLS)
train.head()
#General analysis
#1.1 AVProductStatesIdentifier
#Top 20 categories detection
analysis(COLS[1], 100)
# hypothesis: Different Antivirius product will have different performance over the virius detection
train[COLS[2]].value_counts()
# hypothesis: Different Antivirius product installed will have different performance over the virius detection
analysis(COLS[2], 3)
#Need deep analysis
# hypothesis: Different Antivirius product installed will have different performance over the virius detection
analysis(COLS[3], 2)
#Need deep analysis
# trial w/ random forest
def skl(col):
nominal_transformer = Pipeline(steps=[
('onehot', OneHotEncoder(handle_unknown='ignore'))
])
preproc = ColumnTransformer(transformers=[('onehot', nominal_transformer, col)],\
remainder='drop')
clf = RandomForestClassifier(n_estimators=7, max_depth=60)
pl = Pipeline(steps=[('preprocessor', preproc),
('clf', clf)
])
return pl
X_train, X_test, y_train, y_test = train_test_split(train.dropna().drop('HasDetections',axis = 1)\
, train.dropna()['HasDetections'], test_size=0.25)
N = len(y_test)
y_random = y_test.sample(replace=False, frac = 1)
output = pd.DataFrame(columns = ['Observation accuracy', 'Random_Data accuracy'], index = COLS[1:])
for i in COLS[1:]:
pl = skl([i])
pl.fit(X_train, y_train)
pred_score = pl.score(X_test, y_test)
rand_score = pl.score(X_test, y_random)
output.loc[i, 'Observation accuracy'] = pred_score
output.loc[i, 'Random_Data accuracy'] = rand_score
pl = skl(COLS[1:])
pl.fit(X_train, y_train)
pred_score = pl.score(X_test, y_test)
rand_score = pl.score(X_test, y_random)
output.loc['combined', 'Observation accuracy'] = pred_score
output.loc['combined', 'Random_Data accuracy'] = rand_score
output
output.plot(kind = 'bar', ylim = (0.45, 0.65))
#Conclusion, when using random forest clustering, 'AVProductStatesIdentifier' will dominate the performance
#of prediction, compare the comparison with random data, 'AVProductStatesIdentifier' have a significant imporvement
#in identifying malware.
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Automated Machine Learning
**Continuous retraining using Pipelines and Time-Series TabularDataset**
## Contents
1. [Introduction](#Introduction)
2. [Setup](#Setup)
3. [Compute](#Compute)
4. [Run Configuration](#Run-Configuration)
5. [Data Ingestion Pipeline](#Data-Ingestion-Pipeline)
6. [Training Pipeline](#Training-Pipeline)
7. [Publish Retraining Pipeline and Schedule](#Publish-Retraining-Pipeline-and-Schedule)
8. [Test Retraining](#Test-Retraining)
## Introduction
In this example we use AutoML and Pipelines to enable contious retraining of a model based on updates to the training dataset. We will create two pipelines, the first one to demonstrate a training dataset that gets updated over time. We leverage time-series capabilities of `TabularDataset` to achieve this. The second pipeline utilizes pipeline `Schedule` to trigger continuous retraining.
Make sure you have executed the [configuration notebook](../../../configuration.ipynb) before running this notebook.
In this notebook you will learn how to:
* Create an Experiment in an existing Workspace.
* Configure AutoML using AutoMLConfig.
* Create data ingestion pipeline to update a time-series based TabularDataset
* Create training pipeline to prepare data, run AutoML, register the model and setup pipeline triggers.
## Setup
As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments.
```
import logging
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
from sklearn import datasets
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.train.automl import AutoMLConfig
```
This sample notebook may use features that are not available in previous versions of the Azure ML SDK.
```
print("This notebook was created using version 1.29.0 of the Azure ML SDK")
print("You are currently using version", azureml.core.VERSION, "of the Azure ML SDK")
```
Accessing the Azure ML workspace requires authentication with Azure.
The default authentication is interactive authentication using the default tenant. Executing the ws = Workspace.from_config() line in the cell below will prompt for authentication the first time that it is run.
If you have multiple Azure tenants, you can specify the tenant by replacing the ws = Workspace.from_config() line in the cell below with the following:
```
from azureml.core.authentication import InteractiveLoginAuthentication
auth = InteractiveLoginAuthentication(tenant_id = 'mytenantid')
ws = Workspace.from_config(auth = auth)
```
If you need to run in an environment where interactive login is not possible, you can use Service Principal authentication by replacing the ws = Workspace.from_config() line in the cell below with the following:
```
from azureml.core.authentication import ServicePrincipalAuthentication
auth = auth = ServicePrincipalAuthentication('mytenantid', 'myappid', 'mypassword')
ws = Workspace.from_config(auth = auth)
```
For more details, see aka.ms/aml-notebook-auth
```
ws = Workspace.from_config()
dstor = ws.get_default_datastore()
# Choose a name for the run history container in the workspace.
experiment_name = 'retrain-noaaweather'
experiment = Experiment(ws, experiment_name)
output = {}
output['Subscription ID'] = ws.subscription_id
output['Workspace'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Run History Name'] = experiment_name
pd.set_option('display.max_colwidth', -1)
outputDf = pd.DataFrame(data = output, index = [''])
outputDf.T
```
## Compute
#### Create or Attach existing AmlCompute
You will need to create a compute target for your AutoML run. In this tutorial, you create AmlCompute as your training compute resource.
> Note that if you have an AzureML Data Scientist role, you will not have permission to create compute resources. Talk to your workspace or IT admin to create the compute targets described in this section, if they do not already exist.
#### Creation of AmlCompute takes approximately 5 minutes.
If the AmlCompute with that name is already in your workspace this code will skip the creation process.
As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.
```
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Choose a name for your CPU cluster
amlcompute_cluster_name = "cont-cluster"
# Verify that cluster does not exist already
try:
compute_target = ComputeTarget(workspace=ws, name=amlcompute_cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2',
max_nodes=4)
compute_target = ComputeTarget.create(ws, amlcompute_cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
```
## Run Configuration
```
from azureml.core.runconfig import CondaDependencies, RunConfiguration
# create a new RunConfig object
conda_run_config = RunConfiguration(framework="python")
# Set compute target to AmlCompute
conda_run_config.target = compute_target
conda_run_config.environment.docker.enabled = True
cd = CondaDependencies.create(pip_packages=['azureml-sdk[automl]', 'applicationinsights', 'azureml-opendatasets', 'azureml-defaults'],
conda_packages=['numpy==1.16.2'],
pin_sdk_version=False)
conda_run_config.environment.python.conda_dependencies = cd
print('run config is ready')
```
## Data Ingestion Pipeline
For this demo, we will use NOAA weather data from [Azure Open Datasets](https://azure.microsoft.com/services/open-datasets/). You can replace this with your own dataset, or you can skip this pipeline if you already have a time-series based `TabularDataset`.
```
# The name and target column of the Dataset to create
dataset = "NOAA-Weather-DS4"
target_column_name = "temperature"
```
### Upload Data Step
The data ingestion pipeline has a single step with a script to query the latest weather data and upload it to the blob store. During the first run, the script will create and register a time-series based `TabularDataset` with the past one week of weather data. For each subsequent run, the script will create a partition in the blob store by querying NOAA for new weather data since the last modified time of the dataset (`dataset.data_changed_time`) and creating a data.csv file.
```
from azureml.pipeline.core import Pipeline, PipelineParameter
from azureml.pipeline.steps import PythonScriptStep
ds_name = PipelineParameter(name="ds_name", default_value=dataset)
upload_data_step = PythonScriptStep(script_name="upload_weather_data.py",
allow_reuse=False,
name="upload_weather_data",
arguments=["--ds_name", ds_name],
compute_target=compute_target,
runconfig=conda_run_config)
```
### Submit Pipeline Run
```
data_pipeline = Pipeline(
description="pipeline_with_uploaddata",
workspace=ws,
steps=[upload_data_step])
data_pipeline_run = experiment.submit(data_pipeline, pipeline_parameters={"ds_name":dataset})
data_pipeline_run.wait_for_completion(show_output=False)
```
## Training Pipeline
### Prepare Training Data Step
Script to check if new data is available since the model was last trained. If no new data is available, we cancel the remaining pipeline steps. We need to set allow_reuse flag to False to allow the pipeline to run even when inputs don't change. We also need the name of the model to check the time the model was last trained.
```
from azureml.pipeline.core import PipelineData
# The model name with which to register the trained model in the workspace.
model_name = PipelineParameter("model_name", default_value="noaaweatherds")
data_prep_step = PythonScriptStep(script_name="check_data.py",
allow_reuse=False,
name="check_data",
arguments=["--ds_name", ds_name,
"--model_name", model_name],
compute_target=compute_target,
runconfig=conda_run_config)
from azureml.core import Dataset
train_ds = Dataset.get_by_name(ws, dataset)
train_ds = train_ds.drop_columns(["partition_date"])
```
### AutoMLStep
Create an AutoMLConfig and a training step.
```
from azureml.train.automl import AutoMLConfig
from azureml.pipeline.steps import AutoMLStep
automl_settings = {
"iteration_timeout_minutes": 10,
"experiment_timeout_hours": 0.25,
"n_cross_validations": 3,
"primary_metric": 'r2_score',
"max_concurrent_iterations": 3,
"max_cores_per_iteration": -1,
"verbosity": logging.INFO,
"enable_early_stopping": True
}
automl_config = AutoMLConfig(task = 'regression',
debug_log = 'automl_errors.log',
path = ".",
compute_target=compute_target,
training_data = train_ds,
label_column_name = target_column_name,
**automl_settings
)
from azureml.pipeline.core import PipelineData, TrainingOutput
metrics_output_name = 'metrics_output'
best_model_output_name = 'best_model_output'
metrics_data = PipelineData(name='metrics_data',
datastore=dstor,
pipeline_output_name=metrics_output_name,
training_output=TrainingOutput(type='Metrics'))
model_data = PipelineData(name='model_data',
datastore=dstor,
pipeline_output_name=best_model_output_name,
training_output=TrainingOutput(type='Model'))
automl_step = AutoMLStep(
name='automl_module',
automl_config=automl_config,
outputs=[metrics_data, model_data],
allow_reuse=False)
```
### Register Model Step
Script to register the model to the workspace.
```
register_model_step = PythonScriptStep(script_name="register_model.py",
name="register_model",
allow_reuse=False,
arguments=["--model_name", model_name, "--model_path", model_data, "--ds_name", ds_name],
inputs=[model_data],
compute_target=compute_target,
runconfig=conda_run_config)
```
### Submit Pipeline Run
```
training_pipeline = Pipeline(
description="training_pipeline",
workspace=ws,
steps=[data_prep_step, automl_step, register_model_step])
training_pipeline_run = experiment.submit(training_pipeline, pipeline_parameters={
"ds_name": dataset, "model_name": "noaaweatherds"})
training_pipeline_run.wait_for_completion(show_output=False)
```
### Publish Retraining Pipeline and Schedule
Once we are happy with the pipeline, we can publish the training pipeline to the workspace and create a schedule to trigger on blob change. The schedule polls the blob store where the data is being uploaded and runs the retraining pipeline if there is a data change. A new version of the model will be registered to the workspace once the run is complete.
```
pipeline_name = "Retraining-Pipeline-NOAAWeather"
published_pipeline = training_pipeline.publish(
name=pipeline_name,
description="Pipeline that retrains AutoML model")
published_pipeline
from azureml.pipeline.core import Schedule
schedule = Schedule.create(workspace=ws, name="RetrainingSchedule",
pipeline_parameters={"ds_name": dataset, "model_name": "noaaweatherds"},
pipeline_id=published_pipeline.id,
experiment_name=experiment_name,
datastore=dstor,
wait_for_provisioning=True,
polling_interval=1440)
```
## Test Retraining
Here we setup the data ingestion pipeline to run on a schedule, to verify that the retraining pipeline runs as expected.
Note:
* Azure NOAA Weather data is updated daily and retraining will not trigger if there is no new data available.
* Depending on the polling interval set in the schedule, the retraining may take some time trigger after data ingestion pipeline completes.
```
pipeline_name = "DataIngestion-Pipeline-NOAAWeather"
published_pipeline = training_pipeline.publish(
name=pipeline_name,
description="Pipeline that updates NOAAWeather Dataset")
published_pipeline
from azureml.pipeline.core import Schedule
schedule = Schedule.create(workspace=ws, name="RetrainingSchedule-DataIngestion",
pipeline_parameters={"ds_name":dataset},
pipeline_id=published_pipeline.id,
experiment_name=experiment_name,
datastore=dstor,
wait_for_provisioning=True,
polling_interval=1440)
```
| github_jupyter |
```
import math
import os
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import torch
import pyro
import pyro.infer
import pyro.optim
from pyro.optim import Adam
import pyro.distributions as dist
import torch.distributions.constraints as constraints
from pyro.infer import SVI, Trace_ELBO
torch.manual_seed(101);
# N(0, 1)
loc = 0. # mean zero
scale = 1. # unit variance
normal = dist.Normal(loc, scale) # create a normal distribution object
x = normal.sample() # draw a sample from N(0,1)
print("sample", x.data.numpy())
print("log prob", normal.log_prob(x).data.numpy()) # score the sample from N(0,1)
x = pyro.sample("my_sample", dist.Normal(loc, scale))
print(x)
def weather():
cloudy = pyro.sample('cloudy', dist.Bernoulli(0.3))
cloudy = 'cloudy' if cloudy.item() == 1.0 else 'sunny'
mean_temp = {'cloudy': 55.0, 'sunny': 75.0}[cloudy]
scale_temp = {'cloudy': 10.0, 'sunny': 15.0}[cloudy]
temp = pyro.sample('temp', dist.Normal(mean_temp, scale_temp))
return cloudy, temp.item()
def ice_cream_sales():
cloudy, temp = weather()
expected_sales = 200. if cloudy == 'sunny' and temp > 80.0 else 50.
ice_cream = pyro.sample('ice_cream', dist.Normal(expected_sales, 10.0))
return ice_cream
for _ in range(3):
print(weather())
print (ice_cream_sales())
def geometric(p, t=None):
if t is None:
t = 0
x = pyro.sample("x_{}".format(t), dist.Bernoulli(p))
if x.item() == 0:
return x
else:
return x + geometric(p, t + 1)
print(geometric(0.5))
def normal_product(loc, scale):
z1 = pyro.sample("z1", dist.Normal(loc, scale))
z2 = pyro.sample("z2", dist.Normal(loc, scale))
y = z1 * z2
return y
def make_normal_normal():
mu_latent = pyro.sample("mu_latent", dist.Normal(0, 1))
fn = lambda scale: normal_product(mu_latent, scale)
return fn
print(make_normal_normal()(1.))
```
## Inference
```
def _scale(guess):
# The prior over weight encodes our uncertainty about our guess
weight = pyro.sample("weight", dist.Normal(guess, 1.0))
# This encodes our belief about the noisiness of the scale:
# the measurement fluctuates around the true weight
return pyro.sample("measurement", dist.Normal(weight, 0.75))
posterior = pyro.infer.Importance(_scale, num_samples=100)
guess = 8.5
marginal = pyro.infer.EmpiricalMarginal(posterior.run(guess))
print(marginal())
plt.figure()
plt.hist([marginal().item() for _ in range(100)], range=(5.0, 12.0))
plt.title("P(measuremet | guess)")
plt.xlabel("weight")
plt.ylabel("#")
plt.show()
conditioned_scale = pyro.condition(
_scale,
data={"measurement": 9.5}
)
def deferred_conditioned_scale(measurement, *args, **kwargs):
return pyro.condition(_scale, data={"measurement": measurement})(*args, **kwargs)
def scale2(guess):
weight = pyro.sample("weight", dist.Normal(guess, 1.))
tolerance = torch.abs(pyro.sample("tolerance", dist.Normal(0., 1.)))
return pyro.sample("measurement", dist.Normal(weight, tolerance))
# conditioning composes:
# the following are all equivalent and do not interfere with each other
conditioned_scale2_1 = pyro.condition(
pyro.condition(scale2, data={"weight": 9.2}),
data={"measurement": 9.5})
conditioned_scale2_2 = pyro.condition(
pyro.condition(scale2, data={"measurement": 9.5}),
data={"weight": 9.2})
conditioned_scale2_3 = pyro.condition(
scale2, data={"weight": 9.2, "measurement": 9.5})
guess = 8.5
measurement = 9.5
conditioned_scale = pyro.condition(_scale, data={"measurement": measurement})
marginal = pyro.infer.EmpiricalMarginal(
pyro.infer.Importance(conditioned_scale, num_samples=100).run(guess), sites="weight")
# The marginal distribution concentrates around the data
print(marginal())
plt.hist([marginal().item() for _ in range(100)], range=(5.0, 12.0))
plt.title("P(weight | measurement, guess)")
plt.xlabel("weight")
plt.ylabel("#");
def scale_prior_guide(guess):
return pyro.sample("weight", dist.Normal(guess, 1.))
posterior = pyro.infer.Importance(conditioned_scale,
guide=scale_prior_guide,
num_samples=10)
marginal = pyro.infer.EmpiricalMarginal(posterior.run(guess), sites="weight")
def scale_posterior_guide(measurement, guess):
# note that torch.size(measurement, 0) is the total number of measurements
# that we're conditioning on
a = (guess + torch.sum(measurement)) / (measurement.size(0) + 1.0)
b = 1. / (measurement.size(0) + 1.0)
return pyro.sample("weight", dist.Normal(a, b))
posterior = pyro.infer.Importance(deferred_conditioned_scale,
guide=scale_posterior_guide,
num_samples=20)
marginal = pyro.infer.EmpiricalMarginal(posterior.run(torch.tensor([measurement]), guess), sites="weight")
plt.hist([marginal().item() for _ in range(100)], range=(5.0, 12.0))
plt.title("P(weight | measurement, guess)")
plt.xlabel("weight")
plt.ylabel("#");
simple_param_store = {}
a = simple_param_store.setdefault("a", torch.randn(1))
def scale_parametrized_guide(guess):
a = pyro.param("a", torch.tensor(torch.randn(1) + guess))
b = pyro.param("b", torch.randn(1))
return pyro.sample("weight", dist.Normal(a, torch.abs(b)))
pyro.clear_param_store()
svi = pyro.infer.SVI(model=conditioned_scale,
guide=scale_parametrized_guide,
optim=pyro.optim.SGD({"lr": 0.001}),
loss=pyro.infer.Trace_ELBO())
losses = []
for t in range(1000):
losses.append(svi.step(guess))
plt.plot(losses)
plt.title("ELBO")
plt.xlabel("step")
plt.ylabel("loss");
```
## SVI
```
# this is for running the notebook in our testing framework
smoke_test = ('CI' in os.environ)
n_steps = 2 if smoke_test else 2000
# enable validation (e.g. validate parameters of distributions)
pyro.enable_validation(True)
# clear the param store in case we're in a REPL
pyro.clear_param_store()
# create some data with 6 observed heads and 4 observed tails
data = []
for _ in range(6):
data.append(torch.tensor(1.0))
for _ in range(4):
data.append(torch.tensor(0.0))
def model(data):
# define the hyperparameters that control the beta prior
alpha0 = torch.tensor(10.0)
beta0 = torch.tensor(10.0)
# sample f from the beta prior
f = pyro.sample("latent_fairness", dist.Beta(alpha0, beta0))
# loop over the observed data [WE ONLY CHANGE THE NEXT LINE]
for i in pyro.irange("data_loop", len(data)):
# observe datapoint i using the bernoulli likelihood
pyro.sample("obs_{}".format(i), dist.Bernoulli(f), obs=data[i])
def guide(data):
# register the two variational parameters with Pyro
# - both parameters will have initial value 15.0.
# - because we invoke constraints.positive, the optimizer
# will take gradients on the unconstrained parameters
# (which are related to the constrained parameters by a log)
alpha_q = pyro.param("alpha_q", torch.tensor(15.0),
constraint=constraints.positive)
beta_q = pyro.param("beta_q", torch.tensor(15.0),
constraint=constraints.positive)
# sample latent_fairness from the distribution Beta(alpha_q, beta_q)
pyro.sample("latent_fairness", dist.Beta(alpha_q, beta_q))
# setup the optimizer
adam_params = {"lr": 0.0005, "betas": (0.90, 0.999)}
optimizer = Adam(adam_params)
# setup the inference algorithm
svi = SVI(model, guide, optimizer, loss=Trace_ELBO())
# do gradient steps
for step in range(n_steps):
svi.step(data)
if step % 100 == 0:
print('.', end='')
# grab the learned variational parameters
alpha_q = pyro.param("alpha_q").item()
beta_q = pyro.param("beta_q").item()
# here we use some facts about the beta distribution
# compute the inferred mean of the coin's fairness
inferred_mean = alpha_q / (alpha_q + beta_q)
# compute inferred standard deviation
factor = beta_q / (alpha_q * (1.0 + alpha_q + beta_q))
inferred_std = inferred_mean * math.sqrt(factor)
print("\nbased on the data and our prior belief, the fairness " +
"of the coin is %.3f +- %.3f" % (inferred_mean, inferred_std))
```
| github_jupyter |
# 机器学习练习 5 - 偏差和方差
```
import numpy as np
import scipy.io as scio
import scipy.optimize as opt
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
def load_data():
d = scio.loadmat('ex5data1.mat')
return map(np.ravel,[d['X'],d['y'],d['Xval'],d['yval'],d['Xtest'],d['ytest']])
X,y,Xval,yval,Xtest,ytest = load_data()
df = pd.DataFrame({'water_level':X,'flow':y })
sns.lmplot(x='water_level',y = 'flow',data=df,fit_reg=False,height=7)
plt.show()
X,Xval,Xtest = [np.insert(x.reshape(x.shape[0],1),0,np.ones(x.shape[0]),axis=1) for x in (X,Xval,Xtest)]
```
## 代价函数
```
def cost(theta,X,y):
m = X.shape[0]
inner = X @ theta - y
square_sum = inner.T @ inner
cost = square_sum/(2*m)
return cost
theta = np.ones(X.shape[1])
cost(theta,X,y)
```
## 梯度
```
def gradient(theta,X,y):
m = X.shape[0]
inner = X.T @ (X @ theta -y)
return inner / m
gradient(theta,X,y)
```
## 正则化梯度
```
def regularized_gradient(theta,X,y,l=1):
m = X.shape[0]
regularized_term = theta.copy()
regularized_term[0] = 0
regularized_term = (l/m) * regularized_term
return gradient(theta,X,y) + regularized_term
regularized_gradient(theta, X, y,0)
```
## 拟合数据
```
def linear_regression_np(X,y,l=1):
theta = np.ones(X.shape[1])
res = opt.minimize(fun = regularized_cost,
x0 = theta,
args = (X,y,l),
method='TNC',
jac=regularized_gradient,
options={'disp':True})
return res
def regularized_cost(theta, X, y,l=1):
m = X.shape[0]
regularized_term = (l / (2 * m)) * np.power(theta[1:], 2).sum()
return cost(theta, X, y) + regularized_term
theta = np.ones(X.shape[0])
final_theta = linear_regression_np(X, y, l=0).get('x')
final_theta
b = final_theta[0] # intercept
m = final_theta[1] # slope
plt.scatter(X[:,1], y, label="Training data")
plt.plot(X[:, 1], X[:, 1]*m + b, label="Prediction")
plt.legend(loc=2)
plt.show()
training_cost, cv_cost = [], []
m = X.shape[0]
for i in range(1,m+1):
res = linear_regression_np(X[:i,:],y[:i],l=0)
tc = regularized_cost(res.x,X[:i,:],y[:i],l = 0)
cv = regularized_cost(res.x,Xval,yval,l = 0)
training_cost.append(tc)
cv_cost.append(cv)
plt.plot(np.arange(1, m+1), training_cost, label='training cost')
plt.plot(np.arange(1, m+1), cv_cost, label='cv cost')
plt.legend(loc=1)
plt.show()
```
这个模型拟合不太好, 欠拟合了
## 创建多项式特征
```
def prepare_poly_data(*args,power):
def prepare(x):
df = poly_features(x,power)
ndarr = normalize_feature(df).values
return np.insert(ndarr,0,np.ones(ndarr.shape[0]),axis=1)
return [prepare(x) for x in args]
def poly_features(x,power,as_ndarray=False):
data = {'f{}'.format(i): np.power(x,i) for i in range(1,power+1)}
df = pd.DataFrame(data)
return df.values if as_ndarray else df
X, y, Xval, yval, Xtest, ytest = load_data()
poly_features(X, power=3,as_ndarray=True)
```
## 准备多项式回归数据
```
def normalize_feature(df):
return df.apply(lambda column : (column - column.mean())/column.std())
X_poly, Xval_poly, Xtest_poly= prepare_poly_data(X, Xval, Xtest, power=8)
X_poly[:3, :]
```
## 画出学习曲线
```
def plot_learning_curve(X, y, Xval, yval, l=0):
training_cost, cv_cost = [], []
m = X.shape[0]
for i in range(1, m + 1):
# regularization applies here for fitting parameters
res = linear_regression_np(X[:i, :], y[:i], l)
# remember, when you compute the cost here, you are computing
# non-regularized cost. Regularization is used to fit parameters only
tc = cost(res.x, X[:i, :], y[:i])
cv = cost(res.x, Xval, yval)
training_cost.append(tc)
cv_cost.append(cv)
plt.plot(np.arange(1, m + 1), training_cost, label='training cost')
plt.plot(np.arange(1, m + 1), cv_cost, label='cv cost')
plt.legend(loc=1)
return res
res = plot_learning_curve(X_poly, y, Xval_poly, yval, l=0)
plt.show()
plt.scatter(X, y, label="Training data")
x = np.linspace(-60,60)
xploy= prepare_poly_data(x, power=8)
plt.plot(x,(xploy @ res.x).reshape(-1), label='training cost')
plt.legend(loc=2)
plt.show()
```
你可以看到训练的代价太低了,不真实. 这是 过拟合了
## try 𝜆=1
```
plot_learning_curve(X_poly, y, Xval_poly, yval, l=1)
plt.show()
```
训练代价增加了些,不再是0了。 也就是说我们减轻过拟合
## try 𝜆=100
```
plot_learning_curve(X_poly, y, Xval_poly, yval, l=100)
plt.show()
```
太多正则化了.
变成 欠拟合状态
## 找到最佳的 𝜆
```
l_candidate = [0, 0.001, 0.003, 0.01, 0.03, 0.1, 0.3, 1, 3, 10]
training_cost, cv_cost = [], []
for l in l_candidate:
res = linear_regression_np(X_poly, y, l)
tc = cost(res.x, X_poly, y)
cv = cost(res.x, Xval_poly, yval)
training_cost.append(tc)
cv_cost.append(cv)
plt.plot(l_candidate, training_cost, label='training')
plt.plot(l_candidate, cv_cost, label='cross validation')
plt.legend(loc=2)
plt.xlabel('lambda')
plt.ylabel('cost')
plt.show()
# best cv I got from all those candidates
l_candidate[np.argmin(cv_cost)]
# use test data to compute the cost
for l in l_candidate:
theta = linear_regression_np(X_poly, y, l).x
print('test cost(l={}) = {}'.format(l, cost(theta, Xtest_poly, ytest)))
```
调参后, 𝜆=0.3 是最优选择,这个时候测试代价最小
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Examining the TensorFlow Graph
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tensorboard/graphs"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/tensorboard/blob/master/docs/graphs.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/tensorboard/blob/master/docs/graphs.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
## Overview
TensorBoard’s **Graphs dashboard** is a powerful tool for examining your TensorFlow model. You can quickly view a conceptual graph of your model’s structure and ensure it matches your intended design. You can also view a op-level graph to understand how TensorFlow understands your program. Examining the op-level graph can give you insight as to how to change your model. For example, you can redesign your model if training is progressing slower than expected.
This tutorial presents a quick overview of how to generate graph diagnostic data and visualize it in TensorBoard’s Graphs dashboard. You’ll define and train a simple Keras Sequential model for the Fashion-MNIST dataset and learn how to log and examine your model graphs. You will also use a tracing API to generate graph data for functions created using the new `tf.function` annotation.
## Setup
```
# Load the TensorBoard notebook extension.
%load_ext tensorboard
from datetime import datetime
from packaging import version
import tensorflow as tf
from tensorflow import keras
print("TensorFlow version: ", tf.__version__)
assert version.parse(tf.__version__).release[0] >= 2, \
"This notebook requires TensorFlow 2.0 or above."
import tensorboard
tensorboard.__version__
# Clear any logs from previous runs
!rm -rf ./logs/
```
## Define a Keras model
In this example, the classifier is a simple four-layer Sequential model.
```
# Define the model.
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(32, activation='relu'),
keras.layers.Dropout(0.2),
keras.layers.Dense(10, activation='softmax')
])
model.compile(
optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
```
Download and prepare the training data.
```
(train_images, train_labels), _ = keras.datasets.fashion_mnist.load_data()
train_images = train_images / 255.0
```
## Train the model and log data
Before training, define the [Keras TensorBoard callback](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/TensorBoard), specifying the log directory. By passing this callback to Model.fit(), you ensure that graph data is logged for visualization in TensorBoard.
```
# Define the Keras TensorBoard callback.
logdir="logs/fit/" + datetime.now().strftime("%Y%m%d-%H%M%S")
tensorboard_callback = keras.callbacks.TensorBoard(log_dir=logdir)
# Train the model.
model.fit(
train_images,
train_labels,
batch_size=64,
epochs=5,
callbacks=[tensorboard_callback])
```
## Op-level graph
Start TensorBoard and wait a few seconds for the UI to load. Select the Graphs dashboard by tapping “Graphs” at the top.
```
%tensorboard --logdir logs
```
By default, TensorBoard displays the **op-level graph**. (On the left, you can see the “Default” tag selected.) Note that the graph is inverted; data flows from bottom to top, so it’s upside down compared to the code. However, you can see that the graph closely matches the Keras model definition, with extra edges to other computation nodes.
Graphs are often very large, so you can manipulate the graph visualization:
* Scroll to **zoom** in and out
* Drag to **pan**
* Double clicking toggles **node expansion** (a node can be a container for other nodes)
You can also see metadata by clicking on a node. This allows you to see inputs, outputs, shapes and other details.
<img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/graphs_computation.png?raw=1"/>
<img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/graphs_computation_detail.png?raw=1"/>
## Conceptual graph
In addition to the execution graph, TensorBoard also displays a **conceptual graph**. This is a view of just the Keras model. This may be useful if you’re reusing a saved model and you want to examine or validate its structure.
To see the conceptual graph, select the “keras” tag. For this example, you’ll see a collapsed **Sequential** node. Double-click the node to see the model’s structure:
<img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/graphs_tag_selection.png?raw=1"/> <br/>
<img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/graphs_conceptual.png?raw=1"/>
## Graphs of tf.functions
The examples so far have described graphs of Keras models, where the graphs have been created by defining Keras layers and calling Model.fit().
You may encounter a situation where you need to use the `tf.function` annotation to ["autograph"](https://www.tensorflow.org/guide/function), i.e., transform, a Python computation function into a high-performance TensorFlow graph. For these situations, you use **TensorFlow Summary Trace API** to log autographed functions for visualization in TensorBoard.
To use the Summary Trace API:
* Define and annotate a function with `tf.function`
* Use `tf.summary.trace_on()` immediately before your function call site.
* Add profile information (memory, CPU time) to graph by passing `profiler=True`
* With a Summary file writer, call `tf.summary.trace_export()` to save the log data
You can then use TensorBoard to see how your function behaves.
```
# The function to be traced.
@tf.function
def my_func(x, y):
# A simple hand-rolled layer.
return tf.nn.relu(tf.matmul(x, y))
# Set up logging.
stamp = datetime.now().strftime("%Y%m%d-%H%M%S")
logdir = 'logs/func/%s' % stamp
writer = tf.summary.create_file_writer(logdir)
# Sample data for your function.
x = tf.random.uniform((3, 3))
y = tf.random.uniform((3, 3))
# Bracket the function call with
# tf.summary.trace_on() and tf.summary.trace_export().
tf.summary.trace_on(graph=True, profiler=True)
# Call only one tf.function when tracing.
z = my_func(x, y)
with writer.as_default():
tf.summary.trace_export(
name="my_func_trace",
step=0,
profiler_outdir=logdir)
%tensorboard --logdir logs/func
```
<img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/graphs_autograph.png?raw=1"/>
You can now see the structure of your function as understood by TensorBoard. Click on the "Profile" radiobutton to see CPU and memory statistics.
| github_jupyter |
# Altair in `JupyterLite`
**Altair** is a declarative statistical visualization library for Python.
Most of the examples below are from: https://altair-viz.github.io/gallery
## Import the dependencies:
```
import micropip
# Last version of jsonschema before it added the pyrsistent dependency (native code, no wheel)
await micropip.install("https://files.pythonhosted.org/packages/77/de/47e35a97b2b05c2fadbec67d44cfcdcd09b8086951b331d82de90d2912da/jsonschema-2.6.0-py2.py3-none-any.whl")
await micropip.install("altair")
```
## Simple Bar Chart
```
import altair as alt
import pandas as pd
source = pd.DataFrame({
'a': ['A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I'],
'b': [28, 55, 43, 91, 81, 53, 19, 87, 52]
})
alt.Chart(source).mark_bar().encode(
x='a',
y='b'
)
```
## Simple Heatmap
```
import altair as alt
import numpy as np
import pandas as pd
# Compute x^2 + y^2 across a 2D grid
x, y = np.meshgrid(range(-5, 5), range(-5, 5))
z = x ** 2 + y ** 2
# Convert this grid to columnar data expected by Altair
source = pd.DataFrame({'x': x.ravel(),
'y': y.ravel(),
'z': z.ravel()})
alt.Chart(source).mark_rect().encode(
x='x:O',
y='y:O',
color='z:Q'
)
```
## Install the Vega Dataset
```
await micropip.install('vega_datasets')
```
## Interactive Average
```
import altair as alt
from vega_datasets import data
source = data.seattle_weather()
brush = alt.selection(type='interval', encodings=['x'])
bars = alt.Chart().mark_bar().encode(
x='month(date):O',
y='mean(precipitation):Q',
opacity=alt.condition(brush, alt.OpacityValue(1), alt.OpacityValue(0.7)),
).add_selection(
brush
)
line = alt.Chart().mark_rule(color='firebrick').encode(
y='mean(precipitation):Q',
size=alt.SizeValue(3)
).transform_filter(
brush
)
alt.layer(bars, line, data=source)
```
## Locations of US Airports
```
import altair as alt
from vega_datasets import data
airports = data.airports.url
states = alt.topo_feature(data.us_10m.url, feature='states')
# US states background
background = alt.Chart(states).mark_geoshape(
fill='lightgray',
stroke='white'
).properties(
width=500,
height=300
).project('albersUsa')
# airport positions on background
points = alt.Chart(airports).transform_aggregate(
latitude='mean(latitude)',
longitude='mean(longitude)',
count='count()',
groupby=['state']
).mark_circle().encode(
longitude='longitude:Q',
latitude='latitude:Q',
size=alt.Size('count:Q', title='Number of Airports'),
color=alt.value('steelblue'),
tooltip=['state:N','count:Q']
).properties(
title='Number of airports in US'
)
background + points
```
| github_jupyter |
# Two Loop FDEM
```
from geoscilabs.base import widgetify
import geoscilabs.em.InductionLoop as IND
from ipywidgets import interact, FloatSlider, FloatText
```
## Parameter Descriptions
<img style="float: right; width: 500px" src="https://github.com/geoscixyz/geosci-labs/blob/main/images/em/InductionLoop.png?raw=true">
Below are the adjustable parameters for widgets within this notebook:
* $I_p$: Transmitter current amplitude [A]
* $a_{Tx}$: Transmitter loop radius [m]
* $a_{Rx}$: Receiver loop radius [m]
* $x_{Rx}$: Receiver x position [m]
* $z_{Rx}$: Receiver z position [m]
* $\theta$: Receiver normal vector relative to vertical [degrees]
* $R$: Resistance of receiver loop [$\Omega$]
* $L$: Inductance of receiver loop [H]
* $f$: Specific frequency [Hz]
* $t$: Specific time [s]
## Background Theory: Induced Currents due to a Harmonic Primary Signal
Consider the case in the image above, where a circular loop of wire ($Tx$) caries a harmonic current $I_p (\omega)$. According to the Biot-Savart law, this produces a harmonic primary magnetic field. The harmonic nature of the corresponding magnetic flux which passes through the receiver coil ($Rx$) generates an induced secondary current $I_s (\omega)$, which depends on the coil's resistance ($R$) and inductance ($L$). Here, we will provided final analytic results associated with the app below. Full derivations can be found at the bottom of the page.
### Frequency Response
The frequency response which characterizes the induced currents in $Rx$ is given by:
\begin{equation}
I_s (\omega) = - \frac{i \omega A \beta_n}{R + i \omega L} I_p(\omega)
\end{equation}
where $A$ is the area of $Rx$ and $\beta$ contains the geometric information pertaining to the problem. The induced current has both in-phase and quadrature components. These are given by:
\begin{align}
I_{Re} (\omega) &= - \frac{i \omega A \beta_n R}{R^2 + (\omega L)^2} I_p(\omega) \\
I_{Im} (\omega) &= - \frac{ \omega^2 A \beta_n L}{R^2 + (\omega L)^2} I_p(\omega)
\end{align}
### Time-Harmonic Response
In the time domain, let us consider a time-harmonic primary current of the form $I_p(t) = I_0 \textrm{cos}(\omega t)$. In this case, the induced currents within $Rx$ are given by:
\begin{equation}
I_s (t) = - \Bigg [ \frac{\omega I_0 A \beta_n}{R \, \textrm{sin} \phi + \omega L \, \textrm{cos} \phi} \Bigg ] \, \textrm{cos} (\omega t -\phi) \;\;\;\;\; \textrm{where} \;\;\;\;\; \phi = \textrm{tan}^{-1} \Bigg ( \frac{\omega L}{R} \Bigg ) \, \in \, [\pi/2, \pi ]
\end{equation}
The phase-lag between the primary and secondary currents is represented by $\phi$. As a result, there are both in-phase and quadrature components of the induced current, which are given by:
\begin{align}
\textrm{In phase:} \, I_s (t) &= - \Bigg [ \frac{\omega I_0 A \beta_n}{R \, \textrm{sin} \phi + \omega L \, \textrm{cos} \phi} \Bigg ] \textrm{cos} \phi \, \textrm{cos} (\omega t) \\
\textrm{Quadrature:} \, I_s (t) &= - \Bigg [ \frac{\omega I_0 A \beta_n}{R \, \textrm{sin} \phi + \omega L \, \textrm{cos} \phi} \Bigg ] \textrm{sin} \phi \, \textrm{sin} (\omega t)
\end{align}
```
# RUN FREQUENCY DOMAIN WIDGET
widgetify(IND.fcn_FDEM_Widget,I=FloatSlider(min=1, max=10., value=1., step=1., continuous_update=False, description = "$I_0$"),\
a1=FloatSlider(min=1., max=20., value=10., step=1., continuous_update=False, description = "$a_{Tx}$"),\
a2=FloatSlider(min=1., max=20.,value=5.,step=1.,continuous_update=False,description = "$a_{Rx}$"),\
xRx=FloatSlider(min=-15., max=15., value=0., step=1., continuous_update=False, description = "$x_{Rx}$"),\
zRx=FloatSlider(min=-15., max=15., value=-8., step=1., continuous_update=False, description = "$z_{Rx}$"),\
azm=FloatSlider(min=-90., max=90., value=0., step=10., continuous_update=False, description = "$\\theta$"),\
logR=FloatSlider(min=0., max=6., value=2., step=1., continuous_update=False, description = "$log_{10}(R)$"),\
logL=FloatSlider(min=-7., max=-2., value=-4., step=1., continuous_update=False, description = "$log_{10}(L)$"),\
logf=FloatSlider(min=0., max=8., value=5., step=1., continuous_update=False, description = "$log_{10}(f)$"))
```
## Supporting Derivation for the Frequency Response
Consider a transmitter loop which carries a harmonic primary current $I_p(\omega)$. According to the Biot-Savart law, this results in a primary magnetic field:
\begin{equation}
\mathbf{B_p} (\mathbf{r},\omega) = \boldsymbol{\beta} \, I_p(\omega) \;\;\;\; \textrm{where} \;\;\;\;\; \boldsymbol{\beta} = \frac{\mu_0}{4 \pi} \int_C \frac{d \mathbf{l} \times \mathbf{r'}}{|\mathbf{r'}|^2}
\end{equation}
where $\boldsymbol{\beta}$ contains the problem geometry. Assume the magnetic field is homogeneous through the receiver loop. The primary field generates an EMF within the receiver loop equal to:
\begin{equation}
EMF = - i\omega \Phi \;\;\;\;\; \textrm{where} \;\;\;\;\; \Phi = A \beta_n I_p(\omega)
\end{equation}
where $A$ is the area of the receiver loop and $\beta_n$ is the component of $\boldsymbol{\beta}$ along $\hat n$. The EMF induces a secondary current $I_s(\omega)$ within the receiver loop. The secondary current is defined by the following expression:
\begin{equation}
V = - i \omega A \beta_n I_p (\omega) = \big (R + i\omega L \big )I_s(\omega)
\end{equation}
Rearranging this expression to solve for the secondary current we obtain
\begin{equation}
I_s (\omega) = - \frac{i \omega A \beta_n}{R + i \omega L} I_p(\omega)
\end{equation}
The secondary current has both real and imaginary components. These are given by:
\begin{equation}
I_{Re} (\omega) = - \frac{i \omega A \beta_n R}{R^2 + (\omega L)^2} I_p(\omega)
\end{equation}
and
\begin{equation}
I_{Im} (\omega) = - \frac{ \omega^2 A \beta_n L}{R^2 + (\omega L)^2} I_p(\omega)
\end{equation}
## Supporting Derivation for the Time-Harmonic Response
Consider a transmitter loop which carries a harmonic primary current of the form:
\begin{equation}
I_p(t) = I_0 \textrm{cos} (\omega t)
\end{equation}
According to the Biot-Savart law, this results in a primary magnetic field:
\begin{equation}
\mathbf{B_p} (\mathbf{r},t) = \boldsymbol{\beta} \, I_0 \, \textrm{cos} (\omega t) \;\;\;\; \textrm{where} \;\;\;\;\; \boldsymbol{\beta} = \frac{\mu_0}{4 \pi} \int_C \frac{d \mathbf{l} \times \mathbf{r'}}{|\mathbf{r'}|^2}
\end{equation}
where $\boldsymbol{\beta}$ contains the problem geometry. If the magnetic field is homogeneous through the receiver loop, the primary field generates an EMF within the receiver loop equal to:
\begin{equation}
EMF = - \frac{\partial \Phi}{\partial t} \;\;\;\;\; \textrm{where} \;\;\;\;\; \Phi = A\hat n \cdot \mathbf{B_p} = I_0 A \beta_n \, \textrm{cos} (\omega t)
\end{equation}
where $A$ is the area of the receiver loop and $\beta_n$ is the component of $\boldsymbol{\beta}$ along $\hat n$. The EMF induces a secondary current $I_s$ within the receiver loop. The secondary current is defined by the following ODE:
\begin{equation}
V = \omega I_0 A \beta_n \, \textrm{sin} (\omega t) = I_s R + L \frac{dI_s}{dt}
\end{equation}
The ODE has a solution of the form:
\begin{equation}
I_s (t) = \alpha \, \textrm{cos} (\omega t - \phi)
\end{equation}
where $\alpha$ is the amplitude of the secondary current and $\phi$ is the phase lag. By solving the ODE, the secondary current induced in the receiver loop is given by:
\begin{equation}
I_s (t) = - \Bigg [ \frac{\omega I_0 A \beta_n}{R \, \textrm{sin} \phi + \omega L \, \textrm{cos} \phi} \Bigg ] \, \textrm{cos} (\omega t -\phi) \;\;\;\;\; \textrm{where} \;\;\;\;\; \phi = \textrm{tan}^{-1} \Bigg ( \frac{\omega L}{R} \Bigg ) \, \in \, [\pi/2, \pi ]
\end{equation}
The secondary current has both in-phase and quadrature components, these are given by:
\begin{equation}
\textrm{In phase:} \, I_s (t) = - \Bigg [ \frac{\omega I_0 A \beta_n}{R \, \textrm{sin} \phi + \omega L \, \textrm{cos} \phi} \Bigg ] \textrm{cos} \phi \, \textrm{cos} (\omega t)
\end{equation}
and
\begin{equation}
\textrm{Quadrature:} \, I_s (t) = - \Bigg [ \frac{\omega I_0 A \beta_n}{R \, \textrm{sin} \phi + \omega L \, \textrm{cos} \phi} \Bigg ] \textrm{sin} \phi \, \textrm{sin} (\omega t)
\end{equation}
| github_jupyter |
# Pandas support
It is convenient to use the Pandas package when dealing with numerical data, so Pint provides PintArray. A PintArray is a Pandas Extension Array, which allows Pandas to recognise the Quantity and store it in Pandas DataFrames and Series.
## Basic example
This example will show the simplist way to use pandas with pint and the underlying objects. It's slightly fiddly as you are not reading from a file. A more normal use case is given in Reading a csv.
First some imports
```
import pandas as pd
import pint
```
Next, we create a DataFrame with PintArrays as columns.
```
df = pd.DataFrame({
"torque": pd.Series([1, 2, 2, 3], dtype="pint[lbf ft]"),
"angular_velocity": pd.Series([1, 2, 2, 3], dtype="pint[rpm]"),
})
df
```
Operations with columns are units aware so behave as we would intuitively expect.
```
df['power'] = df['torque'] * df['angular_velocity']
df
```
We can see the columns' units in the dtypes attribute
```
df.dtypes
```
Each column can be accessed as a Pandas Series
```
df.power
```
Which contains a PintArray
```
df.power.values
```
The PintArray contains a Quantity
```
df.power.values.quantity
```
Pandas Series accessors are provided for most Quantity properties and methods, which will convert the result to a Series where possible.
```
df.power.pint.units
df.power.pint.to("kW").values
```
## Reading from csv
Reading from files is the far more standard way to use pandas. To facilitate this, DataFrame accessors are provided to make it easy to get to PintArrays.
```
import pandas as pd
import pint
import io
```
Here's the contents of the csv file.
```
test_data = '''speed,mech power,torque,rail pressure,fuel flow rate,fluid power
rpm,kW,N m,bar,l/min,kW
1000.0,,10.0,1000.0,10.0,
1100.0,,10.0,100000000.0,10.0,
1200.0,,10.0,1000.0,10.0,
1200.0,,10.0,1000.0,10.0,'''
```
Let's read that into a DataFrame.
Here io.StringIO is used in place of reading a file from disk, whereas a csv file path would typically be used and is shown commented.
```
df = pd.read_csv(io.StringIO(test_data),header=[0,1])
# df = pd.read_csv("/path/to/test_data.csv",header=[0,1])
df
```
Then use the DataFrame's pint accessor's quantify method to convert the columns from `np.ndarray`s to PintArrays, with units from the bottom column level.
```
df.dtypes
df_ = df.pint.quantify(level=-1)
df_
```
As previously, operations between DataFrame columns are unit aware
```
df_.speed*df_.torque
df_
df_['mech power'] = df_.speed*df_.torque
df_['fluid power'] = df_['fuel flow rate'] * df_['rail pressure']
df_
```
The DataFrame's `pint.dequantify` method then allows us to retrieve the units information as a header row once again.
```
df_.pint.dequantify()
```
This allows for some rather powerful abilities. For example, to change single column units
```
df_['fluid power'] = df_['fluid power'].pint.to("kW")
df_['mech power'] = df_['mech power'].pint.to("kW")
df_.pint.dequantify()
```
The units are harder to read than they need be, so lets change pints default format for displaying units.
```
pint.PintType.ureg.default_format = "~P"
df_.pint.dequantify()
```
or the entire table's units
```
df_.pint.to_base_units().pint.dequantify()
```
## Advanced example
This example shows alternative ways to use pint with pandas and other features.
Start with the same imports.
```
import pandas as pd
import pint
```
We'll be use a shorthand for PintArray
```
PA_ = pint.PintArray
```
And set up a unit registry and quantity shorthand.
```
ureg=pint.UnitRegistry()
Q_=ureg.Quantity
```
Operations between PintArrays of different unit registry will not work. We can change the unit registry that will be used in creating new PintArrays to prevent this issue.
```
pint.PintType.ureg = ureg
```
These are the possible ways to create a PintArray.
Note that pint[unit] must be used for the Series constuctor, whereas the PintArray constructor allows the unit string or object.
```
df = pd.DataFrame({
"length" : pd.Series([1,2], dtype="pint[m]"),
"width" : PA_([2,3], dtype="pint[m]"),
"distance" : PA_([2,3], dtype="m"),
"height" : PA_([2,3], dtype=ureg.m),
"depth" : PA_.from_1darray_quantity(Q_([2,3],ureg.m)),
})
df
df.length.values.units
```
| github_jupyter |
```
# default_exp models.layers
```
# Layers
> Helper function used to build PyTorch timeseries models.
```
#export
from tsai.imports import *
from tsai.utils import *
from torch.nn.init import normal_
from fastai.torch_core import Module
from fastai.layers import *
from torch.nn.utils import weight_norm, spectral_norm
#export
def noop(x): return x
#export
def init_lin_zero(m):
if isinstance(m, (nn.Linear)):
if getattr(m, 'bias', None) is not None: nn.init.constant_(m.bias, 0)
nn.init.constant_(m.weight, 0)
for l in m.children(): init_lin_zero(l)
lin_zero_init = init_lin_zero
#export
class SwishBeta(Module):
def __init__(self, beta=1.):
self.sigmoid = torch.sigmoid
self.beta = nn.Parameter(torch.Tensor(1).fill_(beta))
def forward(self, x): return x.mul(self.sigmoid(x*self.beta))
#export
class Chomp1d(nn.Module):
def __init__(self, chomp_size):
super(Chomp1d, self).__init__()
self.chomp_size = chomp_size
def forward(self, x):
return x[:, :, :-self.chomp_size].contiguous()
#export
def same_padding1d(seq_len, ks, stride=1, dilation=1):
"Same padding formula as used in Tensorflow"
p = (seq_len - 1) * stride + (ks - 1) * dilation + 1 - seq_len
return p // 2, p - p // 2
class Pad1d(nn.ConstantPad1d):
def __init__(self, padding, value=0.):
super().__init__(padding, value)
@delegates(nn.Conv1d)
class SameConv1d(Module):
"Conv1d with padding='same'"
def __init__(self, ni, nf, ks=3, stride=1, dilation=1, **kwargs):
self.ks, self.stride, self.dilation = ks, stride, dilation
self.conv1d_same = nn.Conv1d(ni, nf, ks, stride=stride, dilation=dilation, **kwargs)
self.weight = self.conv1d_same.weight
self.bias = self.conv1d_same.bias
self.pad = Pad1d
def forward(self, x):
self.padding = same_padding1d(x.shape[-1], self.ks, dilation=self.dilation) #stride=self.stride not used in padding calculation!
return self.conv1d_same(self.pad(self.padding)(x))
#export
def same_padding2d(H, W, ks, stride=(1, 1), dilation=(1, 1)):
"Same padding formula as used in Tensorflow"
if isinstance(ks, Integral): ks = (ks, ks)
if ks[0] == 1: p_h = 0
else: p_h = (H - 1) * stride[0] + (ks[0] - 1) * dilation[0] + 1 - H
if ks[1] == 1: p_w = 0
else: p_w = (W - 1) * stride[1] + (ks[1] - 1) * dilation[1] + 1 - W
return (p_w // 2, p_w - p_w // 2, p_h // 2, p_h - p_h // 2)
class Pad2d(nn.ConstantPad2d):
def __init__(self, padding, value=0.):
super().__init__(padding, value)
@delegates(nn.Conv2d)
class Conv2dSame(Module):
"Conv2d with padding='same'"
def __init__(self, ni, nf, ks=(3, 3), stride=(1, 1), dilation=(1, 1), **kwargs):
if isinstance(ks, Integral): ks = (ks, ks)
if isinstance(stride, Integral): stride = (stride, stride)
if isinstance(dilation, Integral): dilation = (dilation, dilation)
self.ks, self.stride, self.dilation = ks, stride, dilation
self.conv2d_same = nn.Conv2d(ni, nf, ks, stride=stride, dilation=dilation, **kwargs)
self.weight = self.conv2d_same.weight
self.bias = self.conv2d_same.bias
self.pad = Pad2d
def forward(self, x):
self.padding = same_padding2d(x.shape[-2], x.shape[-1], self.ks, dilation=self.dilation) #stride=self.stride not used in padding calculation!
return self.conv2d_same(self.pad(self.padding)(x))
@delegates(nn.Conv2d)
def Conv2d(ni, nf, kernel_size=None, ks=None, stride=1, padding='same', dilation=1, init='auto', bias_std=0.01, **kwargs):
"conv1d layer with padding='same', 'valid', or any integer (defaults to 'same')"
assert not (kernel_size and ks), 'use kernel_size or ks but not both simultaneously'
assert kernel_size is not None or ks is not None, 'you need to pass a ks'
kernel_size = kernel_size or ks
if padding == 'same':
conv = Conv2dSame(ni, nf, kernel_size, stride=stride, dilation=dilation, **kwargs)
elif padding == 'valid': conv = nn.Conv2d(ni, nf, kernel_size, stride=stride, padding=0, dilation=dilation, **kwargs)
else: conv = nn.Conv2d(ni, nf, kernel_size, stride=stride, padding=padding, dilation=dilation, **kwargs)
init_linear(conv, None, init=init, bias_std=bias_std)
return conv
bs = 2
c_in = 3
c_out = 5
h = 16
w = 20
t = torch.rand(bs, c_in, h, w)
test_eq(Conv2dSame(c_in, c_out, ks=3, stride=1, dilation=1, bias=False)(t).shape, (bs, c_out, h, w))
test_eq(Conv2dSame(c_in, c_out, ks=(3, 1), stride=1, dilation=1, bias=False)(t).shape, (bs, c_out, h, w))
test_eq(Conv2dSame(c_in, c_out, ks=3, stride=(1, 1), dilation=(2, 2), bias=False)(t).shape, (bs, c_out, h, w))
test_eq(Conv2dSame(c_in, c_out, ks=3, stride=(2, 2), dilation=(1, 1), bias=False)(t).shape, (bs, c_out, h//2, w//2))
test_eq(Conv2dSame(c_in, c_out, ks=3, stride=(2, 2), dilation=(2, 2), bias=False)(t).shape, (bs, c_out, h//2, w//2))
test_eq(Conv2d(c_in, c_out, ks=3, padding='same', stride=1, dilation=1, bias=False)(t).shape, (bs, c_out, h, w))
#export
class CausalConv1d(torch.nn.Conv1d):
def __init__(self, ni, nf, ks, stride=1, dilation=1, groups=1, bias=True):
super(CausalConv1d, self).__init__(ni, nf, kernel_size=ks, stride=stride, padding=0, dilation=dilation, groups=groups, bias=bias)
self.__padding = (ks - 1) * dilation
def forward(self, input):
return super(CausalConv1d, self).forward(F.pad(input, (self.__padding, 0)))
#export
@delegates(nn.Conv1d)
def Conv1d(ni, nf, kernel_size=None, ks=None, stride=1, padding='same', dilation=1, init='auto', bias_std=0.01, **kwargs):
"conv1d layer with padding='same', 'causal', 'valid', or any integer (defaults to 'same')"
assert not (kernel_size and ks), 'use kernel_size or ks but not both simultaneously'
assert kernel_size is not None or ks is not None, 'you need to pass a ks'
kernel_size = kernel_size or ks
if padding == 'same':
if kernel_size%2==1:
conv = nn.Conv1d(ni, nf, kernel_size, stride=stride, padding=kernel_size//2 * dilation, dilation=dilation, **kwargs)
else:
conv = SameConv1d(ni, nf, kernel_size, stride=stride, dilation=dilation, **kwargs)
elif padding == 'causal': conv = CausalConv1d(ni, nf, kernel_size, stride=stride, dilation=dilation, **kwargs)
elif padding == 'valid': conv = nn.Conv1d(ni, nf, kernel_size, stride=stride, padding=0, dilation=dilation, **kwargs)
else: conv = nn.Conv1d(ni, nf, kernel_size, stride=stride, padding=padding, dilation=dilation, **kwargs)
init_linear(conv, None, init=init, bias_std=bias_std)
return conv
bs = 2
c_in = 3
c_out = 5
seq_len = 512
t = torch.rand(bs, c_in, seq_len)
dilation = 1
test_eq(CausalConv1d(c_in, c_out, ks=3, dilation=dilation)(t).shape, Conv1d(c_in, c_out, ks=3, padding="same", dilation=dilation)(t).shape)
dilation = 2
test_eq(CausalConv1d(c_in, c_out, ks=3, dilation=dilation)(t).shape, Conv1d(c_in, c_out, ks=3, padding="same", dilation=dilation)(t).shape)
bs = 2
ni = 3
nf = 5
seq_len = 6
ks = 3
t = torch.rand(bs, c_in, seq_len)
test_eq(Conv1d(ni, nf, ks, padding=0)(t).shape, (bs, c_out, seq_len - (2 * (ks//2))))
test_eq(Conv1d(ni, nf, ks, padding='valid')(t).shape, (bs, c_out, seq_len - (2 * (ks//2))))
test_eq(Conv1d(ni, nf, ks, padding='same')(t).shape, (bs, c_out, seq_len))
test_eq(Conv1d(ni, nf, ks, padding='causal')(t).shape, (bs, c_out, seq_len))
test_error('use kernel_size or ks but not both simultaneously', Conv1d, ni, nf, kernel_size=3, ks=3)
test_error('you need to pass a ks', Conv1d, ni, nf)
conv = Conv1d(ni, nf, ks, padding='same')
init_linear(conv, None, init='auto', bias_std=.01)
conv
conv = Conv1d(ni, nf, ks, padding='causal')
init_linear(conv, None, init='auto', bias_std=.01)
conv
conv = Conv1d(ni, nf, ks, padding='valid')
init_linear(conv, None, init='auto', bias_std=.01)
weight_norm(conv)
conv
conv = Conv1d(ni, nf, ks, padding=0)
init_linear(conv, None, init='auto', bias_std=.01)
weight_norm(conv)
conv
#export
class SeparableConv1d(Module):
def __init__(self, ni, nf, ks, stride=1, padding='same', dilation=1, bias=True, bias_std=0.01):
self.depthwise_conv = Conv1d(ni, ni, ks, stride=stride, padding=padding, dilation=dilation, groups=ni, bias=bias)
self.pointwise_conv = nn.Conv1d(ni, nf, 1, stride=1, padding=0, dilation=1, groups=1, bias=bias)
if bias:
if bias_std != 0:
normal_(self.depthwise_conv.bias, 0, bias_std)
normal_(self.pointwise_conv.bias, 0, bias_std)
else:
self.depthwise_conv.bias.data.zero_()
self.pointwise_conv.bias.data.zero_()
def forward(self, x):
x = self.depthwise_conv(x)
x = self.pointwise_conv(x)
return x
bs = 64
c_in = 6
c_out = 5
seq_len = 512
t = torch.rand(bs, c_in, seq_len)
test_eq(SeparableConv1d(c_in, c_out, 3)(t).shape, (bs, c_out, seq_len))
#export
class AddCoords1d(Module):
"""Add coordinates to ease position identification without modifying mean and std"""
def forward(self, x):
bs, _, seq_len = x.shape
cc = torch.linspace(-1,1,x.shape[-1], device=x.device).repeat(bs, 1, 1)
cc = (cc - cc.mean()) / cc.std()
x = torch.cat([x, cc], dim=1)
return x
bs = 2
c_in = 3
c_out = 5
seq_len = 50
t = torch.rand(bs, c_in, seq_len)
t = (t - t.mean()) / t.std()
test_eq(AddCoords1d()(t).shape, (bs, c_in + 1, seq_len))
new_t = AddCoords1d()(t)
test_close(new_t.mean(),0, 1e-2)
test_close(new_t.std(), 1, 1e-2)
#export
class ConvBlock(nn.Sequential):
"Create a sequence of conv1d (`ni` to `nf`), activation (if `act_cls`) and `norm_type` layers."
def __init__(self, ni, nf, kernel_size=None, ks=3, stride=1, padding='same', bias=None, bias_std=0.01, norm='Batch', zero_norm=False, bn_1st=True,
act=nn.ReLU, act_kwargs={}, init='auto', dropout=0., xtra=None, coord=False, separable=False, **kwargs):
kernel_size = kernel_size or ks
ndim = 1
layers = [AddCoords1d()] if coord else []
norm_type = getattr(NormType,f"{snake2camel(norm)}{'Zero' if zero_norm else ''}") if norm is not None else None
bn = norm_type in (NormType.Batch, NormType.BatchZero)
inn = norm_type in (NormType.Instance, NormType.InstanceZero)
if bias is None: bias = not (bn or inn)
if separable: conv = SeparableConv1d(ni + coord, nf, ks=kernel_size, bias=bias, stride=stride, padding=padding, **kwargs)
else: conv = Conv1d(ni + coord, nf, ks=kernel_size, bias=bias, stride=stride, padding=padding, **kwargs)
act = None if act is None else act(**act_kwargs)
if not separable: init_linear(conv, act, init=init, bias_std=bias_std)
if norm_type==NormType.Weight: conv = weight_norm(conv)
elif norm_type==NormType.Spectral: conv = spectral_norm(conv)
layers += [conv]
act_bn = []
if act is not None: act_bn.append(act)
if bn: act_bn.append(BatchNorm(nf, norm_type=norm_type, ndim=ndim))
if inn: act_bn.append(InstanceNorm(nf, norm_type=norm_type, ndim=ndim))
if bn_1st: act_bn.reverse()
if dropout: layers += [nn.Dropout(dropout)]
layers += act_bn
if xtra: layers.append(xtra)
super().__init__(*layers)
Conv = named_partial('Conv', ConvBlock, norm=None, act=None)
ConvBN = named_partial('ConvBN', ConvBlock, norm='Batch', act=None)
CoordConv = named_partial('CoordConv', ConvBlock, norm=None, act=None, coord=True)
SepConv = named_partial('SepConv', ConvBlock, norm=None, act=None, separable=True)
#export
class ResBlock1dPlus(Module):
"Resnet block from `ni` to `nh` with `stride`"
@delegates(ConvLayer.__init__)
def __init__(self, expansion, ni, nf, coord=False, stride=1, groups=1, reduction=None, nh1=None, nh2=None, dw=False, g2=1,
sa=False, sym=False, norm='Batch', zero_norm=True, act_cls=defaults.activation, ks=3,
pool=AvgPool, pool_first=True, **kwargs):
if nh2 is None: nh2 = nf
if nh1 is None: nh1 = nh2
nf,ni = nf*expansion,ni*expansion
k0 = dict(norm=norm, zero_norm=False, act=act_cls, **kwargs)
k1 = dict(norm=norm, zero_norm=zero_norm, act=None, **kwargs)
convpath = [ConvBlock(ni, nh2, ks, coord=coord, stride=stride, groups=ni if dw else groups, **k0),
ConvBlock(nh2, nf, ks, coord=coord, groups=g2, **k1)
] if expansion == 1 else [
ConvBlock(ni, nh1, 1, coord=coord, **k0),
ConvBlock(nh1, nh2, ks, coord=coord, stride=stride, groups=nh1 if dw else groups, **k0),
ConvBlock(nh2, nf, 1, coord=coord, groups=g2, **k1)]
if reduction: convpath.append(SEModule(nf, reduction=reduction, act_cls=act_cls))
if sa: convpath.append(SimpleSelfAttention(nf,ks=1,sym=sym))
self.convpath = nn.Sequential(*convpath)
idpath = []
if ni!=nf: idpath.append(ConvBlock(ni, nf, 1, coord=coord, act=None, **kwargs))
if stride!=1: idpath.insert((1,0)[pool_first], pool(stride, ndim=1, ceil_mode=True))
self.idpath = nn.Sequential(*idpath)
self.act = defaults.activation(inplace=True) if act_cls is defaults.activation else act_cls()
def forward(self, x): return self.act(self.convpath(x) + self.idpath(x))
#export
def SEModule1d(ni, reduction=16, act=nn.ReLU, act_kwargs={}):
"Squeeze and excitation module for 1d"
nf = math.ceil(ni//reduction/8)*8
assert nf != 0, 'nf cannot be 0'
return SequentialEx(nn.AdaptiveAvgPool1d(1),
ConvBlock(ni, nf, ks=1, norm=None, act=act, act_kwargs=act_kwargs),
ConvBlock(nf, ni, ks=1, norm=None, act=nn.Sigmoid), ProdLayer())
t = torch.rand(8, 32, 12)
test_eq(SEModule1d(t.shape[1], 16, act=nn.ReLU, act_kwargs={})(t).shape, t.shape)
#export
def Norm(nf, ndim=1, norm='Batch', zero_norm=False, init=True, **kwargs):
"Norm layer with `nf` features and `ndim` with auto init."
assert 1 <= ndim <= 3
nl = getattr(nn, f"{snake2camel(norm)}Norm{ndim}d")(nf, **kwargs)
if nl.affine and init:
nl.bias.data.fill_(1e-3)
nl.weight.data.fill_(0. if zero_norm else 1.)
return nl
BN1d = partial(Norm, ndim=1, norm='Batch')
IN1d = partial(Norm, ndim=1, norm='Instance')
bs = 2
ni = 3
nf = 5
sl = 4
ks = 5
t = torch.rand(bs, ni, sl)
test_eq(ConvBlock(ni, nf, ks)(t).shape, (bs, nf, sl))
test_eq(ConvBlock(ni, nf, ks, padding='causal')(t).shape, (bs, nf, sl))
test_eq(ConvBlock(ni, nf, ks, coord=True)(t).shape, (bs, nf, sl))
test_eq(BN1d(ni)(t).shape, (bs, ni, sl))
test_eq(BN1d(ni).weight.data.mean().item(), 1.)
test_eq(BN1d(ni, zero_norm=True).weight.data.mean().item(), 0.)
test_eq(ConvBlock(ni, nf, ks, norm='batch', zero_norm=True)[1].weight.data.unique().item(), 0)
test_ne(ConvBlock(ni, nf, ks, norm='batch', zero_norm=False)[1].weight.data.unique().item(), 0)
test_eq(ConvBlock(ni, nf, ks, bias=False)[0].bias, None)
ConvBlock(ni, nf, ks, act=Swish, coord=True)
#export
class LinLnDrop(nn.Sequential):
"Module grouping `LayerNorm1d`, `Dropout` and `Linear` layers"
def __init__(self, n_in, n_out, ln=True, p=0., act=None, lin_first=False):
layers = [nn.LayerNorm(n_out if lin_first else n_in)] if ln else []
if p != 0: layers.append(nn.Dropout(p))
lin = [nn.Linear(n_in, n_out, bias=not ln)]
if act is not None: lin.append(act)
layers = lin+layers if lin_first else layers+lin
super().__init__(*layers)
LinLnDrop(2, 3, p=.5)
#export
class LambdaPlus(Module):
def __init__(self, func, *args, **kwargs): self.func,self.args,self.kwargs=func,args,kwargs
def forward(self, x): return self.func(x, *self.args, **self.kwargs)
#export
class Squeeze(Module):
def __init__(self, dim=-1): self.dim = dim
def forward(self, x): return x.squeeze(dim=self.dim)
def __repr__(self): return f'{self.__class__.__name__}(dim={self.dim})'
class Unsqueeze(Module):
def __init__(self, dim=-1): self.dim = dim
def forward(self, x): return x.unsqueeze(dim=self.dim)
def __repr__(self): return f'{self.__class__.__name__}(dim={self.dim})'
class Add(Module):
def forward(self, x, y): return x.add(y)
def __repr__(self): return f'{self.__class__.__name__}'
class Concat(Module):
def __init__(self, dim=1): self.dim = dim
def forward(self, *x): return torch.cat(*x, dim=self.dim)
def __repr__(self): return f'{self.__class__.__name__}(dim={self.dim})'
class Permute(Module):
def __init__(self, *dims): self.dims = dims
def forward(self, x): return x.permute(self.dims)
def __repr__(self): return f"{self.__class__.__name__}(dims={', '.join([str(d) for d in self.dims])})"
class Transpose(Module):
def __init__(self, *dims, contiguous=False): self.dims, self.contiguous = dims, contiguous
def forward(self, x):
if self.contiguous: return x.transpose(*self.dims).contiguous()
else: return x.transpose(*self.dims)
def __repr__(self):
if self.contiguous: return f"{self.__class__.__name__}(dims={', '.join([str(d) for d in self.dims])}).contiguous()"
else: return f"{self.__class__.__name__}({', '.join([str(d) for d in self.dims])})"
class View(Module):
def __init__(self, *shape): self.shape = shape
def forward(self, x): return x.view(x.shape[0], *self.shape)
def __repr__(self): return f"{self.__class__.__name__}({', '.join(['bs'] + [str(s) for s in self.shape])})"
class Reshape(Module):
def __init__(self, *shape): self.shape = shape
def forward(self, x): return x.reshape(x.shape[0], *self.shape)
def __repr__(self): return f"{self.__class__.__name__}({', '.join(['bs'] + [str(s) for s in self.shape])})"
class Max(Module):
def __init__(self, dim=None, keepdim=False): self.dim, self.keepdim = dim, keepdim
def forward(self, x): return x.max(self.dim, keepdim=self.keepdim)[0]
def __repr__(self): return f'{self.__class__.__name__}(dim={self.dim}, keepdim={self.keepdim})'
class LastStep(Module):
def forward(self, x): return x[..., -1]
def __repr__(self): return f'{self.__class__.__name__}()'
class SoftMax(Module):
"SoftMax layer"
def __init__(self, dim=-1):
self.dim = dim
def forward(self, x):
return F.softmax(x, dim=self.dim)
def __repr__(self): return f'{self.__class__.__name__}(dim={self.dim})'
class Clamp(Module):
def __init__(self, min=None, max=None):
self.min, self.max = min, max
def forward(self, x):
return x.clamp(min=self.min, max=self.max)
def __repr__(self): return f'{self.__class__.__name__}(min={self.min}, max={self.max})'
class Clip(Module):
def __init__(self, min=None, max=None):
self.min, self.max = min, max
def forward(self, x):
if self.min is not None:
x = torch.maximum(x, self.min)
if self.max is not None:
x = torch.minimum(x, self.max)
return x
def __repr__(self): return f'{self.__class__.__name__}()'
class ReZero(Module):
def __init__(self, module):
self.module = module
self.alpha = nn.Parameter(torch.zeros(1))
def forward(self, x):
return x + self.alpha * self.module(x)
Noop = nn.Sequential()
bs = 2
nf = 5
sl = 4
t = torch.rand(bs, nf, sl)
test_eq(Permute(0,2,1)(t).shape, (bs, sl, nf))
test_eq(Max(1)(t).shape, (bs, sl))
test_eq(Transpose(1,2)(t).shape, (bs, sl, nf))
test_eq(Transpose(1,2, contiguous=True)(t).shape, (bs, sl, nf))
test_eq(View(-1, 2, 10)(t).shape, (bs, 1, 2, 10))
test_eq(Reshape(-1, 2, 10)(t).shape, (bs, 1, 2, 10))
Transpose(1,2), Permute(0,2,1), View(-1, 2, 10), Transpose(1,2, contiguous=True), Reshape(-1, 2, 10), Noop
# export
class DropPath(nn.Module):
"""Drop paths (Stochastic Depth) per sample (when applied in main path of residual blocks).
It's similar to Dropout but it drops individual connections instead of nodes.
Original code in https://github.com/rwightman/pytorch-image-models (timm library)
"""
def __init__(self, p=None):
super().__init__()
self.p = p
def forward(self, x):
if self.p == 0. or not self.training: return x
keep_prob = 1 - self.p
shape = (x.shape[0],) + (1,) * (x.ndim - 1)
random_tensor = keep_prob + torch.rand(shape, dtype=x.dtype, device=x.device)
random_tensor.floor_()
output = x.div(keep_prob) * random_tensor
# output = x.div(random_tensor.mean()) * random_tensor # divide by the actual mean to mantain the input mean?
return output
t = torch.ones(100,2,3)
test_eq(DropPath(0.)(t), t)
assert DropPath(0.5)(t).max() >= 1
#export
class Sharpen(Module):
"This is used to increase confidence in predictions - MixMatch paper"
def __init__(self, T=.5): self.T = T
def forward(self, x):
x = x**(1. / self.T)
return x / x.sum(dim=1, keepdims=True)
n_samples = 1000
n_classes = 3
t = (torch.rand(n_samples, n_classes) - .5) * 10
probas = F.softmax(t, -1)
sharpened_probas = Sharpen()(probas)
plt.plot(probas.flatten().sort().values, color='r')
plt.plot(sharpened_probas.flatten().sort().values, color='b')
plt.show()
test_gt(sharpened_probas[n_samples//2:].max(-1).values.sum().item(), probas[n_samples//2:].max(-1).values.sum().item())
#export
class Sequential(nn.Sequential):
"""Class that allows you to pass one or multiple inputs"""
def forward(self, *x):
for i, module in enumerate(self._modules.values()):
x = module(*x) if isinstance(x, (list, tuple, L)) else module(x)
return x
#export
class TimeDistributed(nn.Module):
def __init__(self, module, batch_first=False):
super(TimeDistributed, self).__init__()
self.module = module
self.batch_first = batch_first
def forward(self, x):
if len(x.size()) <= 2:
return self.module(x)
# Squash samples and timesteps into a single axis
x_reshape = x.contiguous().view(-1, x.size(-1)) # (samples * timesteps, input_size)
y = self.module(x_reshape)
# We have to reshape Y
if self.batch_first:
y = y.contiguous().view(x.size(0), -1, y.size(-1)) # (samples, timesteps, output_size)
else:
y = y.view(-1, x.size(1), y.size(-1)) # (timesteps, samples, output_size)
return y
#export
class Temp_Scale(Module):
"Used to perform Temperature Scaling (dirichlet=False) or Single-parameter Dirichlet calibration (dirichlet=True)"
def __init__(self, temp=1., dirichlet=False):
self.weight = nn.Parameter(tensor(temp))
self.bias = None
self.log_softmax = dirichlet
def forward(self, x):
if self.log_softmax: x = F.log_softmax(x, dim=-1)
return x.div(self.weight)
class Vector_Scale(Module):
"Used to perform Vector Scaling (dirichlet=False) or Diagonal Dirichlet calibration (dirichlet=True)"
def __init__(self, n_classes=1, dirichlet=False):
self.weight = nn.Parameter(torch.ones(n_classes))
self.bias = nn.Parameter(torch.zeros(n_classes))
self.log_softmax = dirichlet
def forward(self, x):
if self.log_softmax: x = F.log_softmax(x, dim=-1)
return x.mul(self.weight).add(self.bias)
class Matrix_Scale(Module):
"Used to perform Matrix Scaling (dirichlet=False) or Dirichlet calibration (dirichlet=True)"
def __init__(self, n_classes=1, dirichlet=False):
self.ms = nn.Linear(n_classes, n_classes)
self.ms.weight.data = nn.Parameter(torch.eye(n_classes))
nn.init.constant_(self.ms.bias.data, 0.)
self.weight = self.ms.weight
self.bias = self.ms.bias
self.log_softmax = dirichlet
def forward(self, x):
if self.log_softmax: x = F.log_softmax(x, dim=-1)
return self.ms(x)
def get_calibrator(calibrator=None, n_classes=1, **kwargs):
if calibrator is None or not calibrator: return noop
elif calibrator.lower() == 'temp': return Temp_Scale(dirichlet=False, **kwargs)
elif calibrator.lower() == 'vector': return Vector_Scale(n_classes=n_classes, dirichlet=False, **kwargs)
elif calibrator.lower() == 'matrix': return Matrix_Scale(n_classes=n_classes, dirichlet=False, **kwargs)
elif calibrator.lower() == 'dtemp': return Temp_Scale(dirichlet=True, **kwargs)
elif calibrator.lower() == 'dvector': return Vector_Scale(n_classes=n_classes, dirichlet=True, **kwargs)
elif calibrator.lower() == 'dmatrix': return Matrix_Scale(n_classes=n_classes, dirichlet=True, **kwargs)
else: assert False, f'please, select a correct calibrator instead of {calibrator}'
bs = 2
c_out = 3
t = torch.rand(bs, c_out)
for calibrator, cal_name in zip(['temp', 'vector', 'matrix'], ['Temp_Scale', 'Vector_Scale', 'Matrix_Scale']):
cal = get_calibrator(calibrator, n_classes=c_out)
# print(calibrator)
# print(cal.weight, cal.bias, '\n')
test_eq(cal(t), t)
test_eq(cal.__class__.__name__, cal_name)
for calibrator, cal_name in zip(['dtemp', 'dvector', 'dmatrix'], ['Temp_Scale', 'Vector_Scale', 'Matrix_Scale']):
cal = get_calibrator(calibrator, n_classes=c_out)
# print(calibrator)
# print(cal.weight, cal.bias, '\n')
test_eq(cal(t), F.log_softmax(t, dim=1))
test_eq(cal.__class__.__name__, cal_name)
bs = 2
c_out = 3
t = torch.rand(bs, c_out)
test_eq(Temp_Scale()(t).shape, t.shape)
test_eq(Vector_Scale(c_out)(t).shape, t.shape)
test_eq(Matrix_Scale(c_out)(t).shape, t.shape)
test_eq(Temp_Scale(dirichlet=True)(t).shape, t.shape)
test_eq(Vector_Scale(c_out, dirichlet=True)(t).shape, t.shape)
test_eq(Matrix_Scale(c_out, dirichlet=True)(t).shape, t.shape)
test_eq(Temp_Scale()(t), t)
test_eq(Vector_Scale(c_out)(t), t)
test_eq(Matrix_Scale(c_out)(t), t)
bs = 2
c_out = 5
t = torch.rand(bs, c_out)
test_eq(Vector_Scale(c_out)(t), t)
test_eq(Vector_Scale(c_out).weight.data, torch.ones(c_out))
test_eq(Vector_Scale(c_out).weight.requires_grad, True)
test_eq(type(Vector_Scale(c_out).weight), torch.nn.parameter.Parameter)
bs = 2
c_out = 3
weight = 2
bias = 1
t = torch.rand(bs, c_out)
test_eq(Matrix_Scale(c_out)(t).shape, t.shape)
test_eq(Matrix_Scale(c_out).weight.requires_grad, True)
test_eq(type(Matrix_Scale(c_out).weight), torch.nn.parameter.Parameter)
#export
class LogitAdjustmentLayer(Module):
"Logit Adjustment for imbalanced datasets"
def __init__(self, class_priors):
self.class_priors = class_priors
def forward(self, x):
return x.add(self.class_priors)
LogitAdjLayer = LogitAdjustmentLayer
bs, n_classes = 16, 3
class_priors = torch.rand(n_classes)
logits = torch.randn(bs, n_classes) * 2
test_eq(LogitAdjLayer(class_priors)(logits), logits + class_priors)
#export
class PPV(Module):
def __init__(self, dim=-1):
self.dim = dim
def forward(self, x):
return torch.gt(x, 0).sum(dim=self.dim).float() / x.shape[self.dim]
def __repr__(self): return f'{self.__class__.__name__}(dim={self.dim})'
class PPAuc(Module):
def __init__(self, dim=-1):
self.dim = dim
def forward(self, x):
x = F.relu(x).sum(self.dim) / (abs(x).sum(self.dim) + 1e-8)
return x
def __repr__(self): return f'{self.__class__.__name__}(dim={self.dim})'
class MaxPPVPool1d(Module):
"Drop-in replacement for AdaptiveConcatPool1d - multiplies nf by 2"
def forward(self, x):
_max = x.max(dim=-1).values
_ppv = torch.gt(x, 0).sum(dim=-1).float() / x.shape[-1]
return torch.cat((_max, _ppv), dim=-1).unsqueeze(2)
bs = 2
nf = 5
sl = 4
t = torch.rand(bs, nf, sl)
test_eq(MaxPPVPool1d()(t).shape, (bs, nf*2, 1))
test_eq(MaxPPVPool1d()(t).shape, AdaptiveConcatPool1d(1)(t).shape)
#export
class AdaptiveWeightedAvgPool1d(Module):
'''Global Pooling layer that performs a weighted average along the temporal axis
It can be considered as a channel-wise form of local temporal attention. Inspired by the paper:
Hyun, J., Seong, H., & Kim, E. (2019). Universal Pooling--A New Pooling Method for Convolutional Neural Networks. arXiv preprint arXiv:1907.11440.'''
def __init__(self, n_in, seq_len, mult=2, n_layers=2, ln=False, dropout=0.5, act=nn.ReLU(), zero_init=True):
layers = nn.ModuleList()
for i in range(n_layers):
inp_mult = mult if i > 0 else 1
out_mult = mult if i < n_layers -1 else 1
p = dropout[i] if is_listy(dropout) else dropout
layers.append(LinLnDrop(seq_len * inp_mult, seq_len * out_mult, ln=False, p=p,
act=act if i < n_layers-1 and n_layers > 1 else None))
self.layers = layers
self.softmax = SoftMax(-1)
if zero_init: init_lin_zero(self)
def forward(self, x):
wap = x
for l in self.layers: wap = l(wap)
wap = self.softmax(wap)
return torch.mul(x, wap).sum(-1)
#export
class GAP1d(Module):
"Global Adaptive Pooling + Flatten"
def __init__(self, output_size=1):
self.gap = nn.AdaptiveAvgPool1d(output_size)
self.flatten = Flatten()
def forward(self, x):
return self.flatten(self.gap(x))
class GACP1d(Module):
"Global AdaptiveConcatPool + Flatten"
def __init__(self, output_size=1):
self.gacp = AdaptiveConcatPool1d(output_size)
self.flatten = Flatten()
def forward(self, x):
return self.flatten(self.gacp(x))
class GAWP1d(Module):
"Global AdaptiveWeightedAvgPool1d + Flatten"
def __init__(self, n_in, seq_len, n_layers=2, ln=False, dropout=0.5, act=nn.ReLU(), zero_init=False):
self.gacp = AdaptiveWeightedAvgPool1d(n_in, seq_len, n_layers=n_layers, ln=ln, dropout=dropout, act=act, zero_init=zero_init)
self.flatten = Flatten()
def forward(self, x):
return self.flatten(self.gacp(x))
# export
class GlobalWeightedAveragePool1d(Module):
""" Global Weighted Average Pooling layer
Inspired by Building Efficient CNN Architecture for Offline Handwritten Chinese Character Recognition
https://arxiv.org/pdf/1804.01259.pdf
"""
def __init__(self, n_in, seq_len):
self.weight = nn.Parameter(torch.ones(1, n_in, seq_len))
self.bias = nn.Parameter(torch.zeros(1, n_in, seq_len))
def forward(self, x):
α = F.softmax(torch.sigmoid(x * self.weight + self.bias), dim=-1)
return (x * α).sum(-1)
GWAP1d = GlobalWeightedAveragePool1d
def gwa_pool_head(n_in, c_out, seq_len, bn=True, fc_dropout=0.):
return nn.Sequential(GlobalWeightedAveragePool1d(n_in, seq_len), Flatten(), LinBnDrop(n_in, c_out, p=fc_dropout, bn=bn))
t = torch.randn(16, 64, 50)
head = gwa_pool_head(64, 5, 50)
test_eq(head(t).shape, (16, 5))
#export
class AttentionalPool1d(Module):
"""Global Adaptive Pooling layer inspired by Attentional Pooling for Action Recognition https://arxiv.org/abs/1711.01467"""
def __init__(self, n_in, c_out, bn=False):
store_attr()
self.bn = nn.BatchNorm1d(n_in) if bn else None
self.conv1 = Conv1d(n_in, 1, 1)
self.conv2 = Conv1d(n_in, c_out, 1)
def forward(self, x):
if self.bn is not None: x = self.bn(x)
return (self.conv1(x) @ self.conv2(x).transpose(1,2)).transpose(1,2)
class GAttP1d(nn.Sequential):
def __init__(self, n_in, c_out, bn=False):
super().__init__(AttentionalPool1d(n_in, c_out, bn=bn), Flatten())
def attentional_pool_head(n_in, c_out, seq_len=None, bn=True, **kwargs):
return nn.Sequential(AttentionalPool1d(n_in, c_out, bn=bn, **kwargs), Flatten())
bs, c_in, seq_len = 16, 1, 50
c_out = 3
t = torch.rand(bs, c_in, seq_len)
test_eq(GAP1d()(t).shape, (bs, c_in))
test_eq(GACP1d()(t).shape, (bs, c_in*2))
bs, c_in, seq_len = 16, 4, 50
t = torch.rand(bs, c_in, seq_len)
test_eq(GAP1d()(t).shape, (bs, c_in))
test_eq(GACP1d()(t).shape, (bs, c_in*2))
test_eq(GAWP1d(c_in, seq_len, n_layers=2, ln=False, dropout=0.5, act=nn.ReLU(), zero_init=False)(t).shape, (bs, c_in))
test_eq(GAWP1d(c_in, seq_len, n_layers=2, ln=False, dropout=0.5, act=nn.ReLU(), zero_init=False)(t).shape, (bs, c_in))
test_eq(GAWP1d(c_in, seq_len, n_layers=1, ln=False, dropout=0.5, zero_init=False)(t).shape, (bs, c_in))
test_eq(GAWP1d(c_in, seq_len, n_layers=1, ln=False, dropout=0.5, zero_init=True)(t).shape, (bs, c_in))
test_eq(AttentionalPool1d(c_in, c_out)(t).shape, (bs, c_out, 1))
bs, c_in, seq_len = 16, 128, 50
c_out = 14
t = torch.rand(bs, c_in, seq_len)
attp = attentional_pool_head(c_in, c_out)
test_eq(attp(t).shape, (bs, c_out))
#export
class GEGLU(Module):
def forward(self, x):
x, gates = x.chunk(2, dim=-1)
return x * F.gelu(gates)
class ReGLU(Module):
def forward(self, x):
x, gates = x.chunk(2, dim=-1)
return x * F.relu(gates)
#export
pytorch_acts = [nn.ELU, nn.LeakyReLU, nn.PReLU, nn.ReLU, nn.ReLU6, nn.SELU, nn.CELU, nn.GELU, nn.Sigmoid, Mish, nn.Softplus,
nn.Tanh, nn.Softmax, GEGLU, ReGLU]
pytorch_act_names = [a.__name__.lower() for a in pytorch_acts]
def get_act_fn(act, **act_kwargs):
if act is None: return
elif isinstance(act, nn.Module): return act
elif callable(act): return act(**act_kwargs)
idx = pytorch_act_names.index(act.lower())
return pytorch_acts[idx](**act_kwargs)
test_eq(get_act_fn(nn.ReLU).__repr__(), "ReLU()")
test_eq(get_act_fn(nn.ReLU()).__repr__(), "ReLU()")
test_eq(get_act_fn(nn.LeakyReLU, negative_slope=0.05).__repr__(), "LeakyReLU(negative_slope=0.05)")
test_eq(get_act_fn('reglu').__repr__(), "ReGLU()")
test_eq(get_act_fn('leakyrelu', negative_slope=0.05).__repr__(), "LeakyReLU(negative_slope=0.05)")
#export
def create_pool_head(n_in, c_out, seq_len=None, concat_pool=False, fc_dropout=0., bn=False, y_range=None, **kwargs):
if kwargs: print(f'{kwargs} not being used')
if concat_pool: n_in*=2
layers = [GACP1d(1) if concat_pool else GAP1d(1)]
layers += [LinBnDrop(n_in, c_out, bn=bn, p=fc_dropout)]
if y_range: layers += [SigmoidRange(*y_range)]
return nn.Sequential(*layers)
pool_head = create_pool_head
average_pool_head = partial(pool_head, concat_pool=False)
setattr(average_pool_head, "__name__", "average_pool_head")
concat_pool_head = partial(pool_head, concat_pool=True)
setattr(concat_pool_head, "__name__", "concat_pool_head")
bs = 16
nf = 12
c_out = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
test_eq(create_pool_head(nf, c_out, seq_len, fc_dropout=0.5)(t).shape, (bs, c_out))
test_eq(create_pool_head(nf, c_out, seq_len, concat_pool=True, fc_dropout=0.5)(t).shape, (bs, c_out))
create_pool_head(nf, c_out, seq_len, concat_pool=True, bn=True, fc_dropout=.5)
#export
def max_pool_head(n_in, c_out, seq_len, fc_dropout=0., bn=False, y_range=None, **kwargs):
if kwargs: print(f'{kwargs} not being used')
layers = [nn.MaxPool1d(seq_len, **kwargs), Flatten()]
layers += [LinBnDrop(n_in, c_out, bn=bn, p=fc_dropout)]
if y_range: layers += [SigmoidRange(*y_range)]
return nn.Sequential(*layers)
bs = 16
nf = 12
c_out = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
test_eq(max_pool_head(nf, c_out, seq_len, fc_dropout=0.5)(t).shape, (bs, c_out))
#export
def create_pool_plus_head(*args, lin_ftrs=None, fc_dropout=0., concat_pool=True, bn_final=False, lin_first=False, y_range=None):
nf = args[0]
c_out = args[1]
if concat_pool: nf = nf * 2
lin_ftrs = [nf, 512, c_out] if lin_ftrs is None else [nf] + lin_ftrs + [c_out]
ps = L(fc_dropout)
if len(ps) == 1: ps = [ps[0]/2] * (len(lin_ftrs)-2) + ps
actns = [nn.ReLU(inplace=True)] * (len(lin_ftrs)-2) + [None]
pool = AdaptiveConcatPool1d() if concat_pool else nn.AdaptiveAvgPool1d(1)
layers = [pool, Flatten()]
if lin_first: layers.append(nn.Dropout(ps.pop(0)))
for ni,no,p,actn in zip(lin_ftrs[:-1], lin_ftrs[1:], ps, actns):
layers += LinBnDrop(ni, no, bn=True, p=p, act=actn, lin_first=lin_first)
if lin_first: layers.append(nn.Linear(lin_ftrs[-2], c_out))
if bn_final: layers.append(nn.BatchNorm1d(lin_ftrs[-1], momentum=0.01))
if y_range is not None: layers.append(SigmoidRange(*y_range))
return nn.Sequential(*layers)
pool_plus_head = create_pool_plus_head
bs = 16
nf = 12
c_out = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
test_eq(create_pool_plus_head(nf, c_out, seq_len, fc_dropout=0.5)(t).shape, (bs, c_out))
test_eq(create_pool_plus_head(nf, c_out, concat_pool=True, fc_dropout=0.5)(t).shape, (bs, c_out))
create_pool_plus_head(nf, c_out, seq_len, fc_dropout=0.5)
#export
def create_conv_head(*args, adaptive_size=None, y_range=None):
nf = args[0]
c_out = args[1]
layers = [nn.AdaptiveAvgPool1d(adaptive_size)] if adaptive_size is not None else []
for i in range(2):
if nf > 1:
layers += [ConvBlock(nf, nf // 2, 1)]
nf = nf//2
else: break
layers += [ConvBlock(nf, c_out, 1), GAP1d(1)]
if y_range: layers += [SigmoidRange(*y_range)]
return nn.Sequential(*layers)
conv_head = create_conv_head
bs = 16
nf = 12
c_out = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
test_eq(create_conv_head(nf, c_out, seq_len)(t).shape, (bs, c_out))
test_eq(create_conv_head(nf, c_out, adaptive_size=50)(t).shape, (bs, c_out))
create_conv_head(nf, c_out, 50)
#export
def create_mlp_head(nf, c_out, seq_len=None, flatten=True, fc_dropout=0., bn=False, lin_first=False, y_range=None):
if flatten: nf *= seq_len
layers = [Flatten()] if flatten else []
layers += [LinBnDrop(nf, c_out, bn=bn, p=fc_dropout, lin_first=lin_first)]
if y_range: layers += [SigmoidRange(*y_range)]
return nn.Sequential(*layers)
mlp_head = create_mlp_head
bs = 16
nf = 12
c_out = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
test_eq(create_mlp_head(nf, c_out, seq_len, fc_dropout=0.5)(t).shape, (bs, c_out))
t = torch.rand(bs, nf, seq_len)
create_mlp_head(nf, c_out, seq_len, bn=True, fc_dropout=.5)
#export
def create_fc_head(nf, c_out, seq_len=None, flatten=True, lin_ftrs=None, y_range=None, fc_dropout=0., bn=False, bn_final=False, act=nn.ReLU(inplace=True)):
if flatten: nf *= seq_len
layers = [Flatten()] if flatten else []
lin_ftrs = [nf, 512, c_out] if lin_ftrs is None else [nf] + lin_ftrs + [c_out]
if not is_listy(fc_dropout): fc_dropout = [fc_dropout]*(len(lin_ftrs) - 1)
actns = [act for _ in range(len(lin_ftrs) - 2)] + [None]
layers += [LinBnDrop(lin_ftrs[i], lin_ftrs[i+1], bn=bn and (i!=len(actns)-1 or bn_final), p=p, act=a) for i,(p,a) in enumerate(zip(fc_dropout+[0.], actns))]
if y_range is not None: layers.append(SigmoidRange(*y_range))
return nn.Sequential(*layers)
fc_head = create_fc_head
bs = 16
nf = 12
c_out = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
test_eq(create_fc_head(nf, c_out, seq_len, fc_dropout=0.5)(t).shape, (bs, c_out))
create_mlp_head(nf, c_out, seq_len, bn=True, fc_dropout=.5)
#export
def create_rnn_head(*args, fc_dropout=0., bn=False, y_range=None):
nf = args[0]
c_out = args[1]
layers = [LastStep()]
layers += [LinBnDrop(nf, c_out, bn=bn, p=fc_dropout)]
if y_range: layers += [SigmoidRange(*y_range)]
return nn.Sequential(*layers)
rnn_head = create_rnn_head
bs = 16
nf = 12
c_out = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
test_eq(create_rnn_head(nf, c_out, seq_len, fc_dropout=0.5)(t).shape, (bs, c_out))
create_rnn_head(nf, c_out, seq_len, bn=True, fc_dropout=.5)
# export
def imputation_head(c_in, c_out, seq_len=None, ks=1, y_range=None, fc_dropout=0.):
layers = [nn.Dropout(fc_dropout), nn.Conv1d(c_in, c_out, ks)]
if y_range is not None:
y_range = (tensor(y_range[0]), tensor(y_range[1]))
layers += [SigmoidRange(*y_range)]
return nn.Sequential(*layers)
bs = 16
nf = 12
ni = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
head = imputation_head(nf, ni, seq_len=None, ks=1, y_range=None, fc_dropout=0.)
test_eq(head(t).shape, (bs, ni, seq_len))
head = imputation_head(nf, ni, seq_len=None, ks=1, y_range=(.3,.7), fc_dropout=0.)
test_ge(head(t).min(), .3)
test_le(head(t).max(), .7)
y_range = (tensor([0.1000, 0.1000, 0.1000, 0.1000, 0.2000, 0.2000, 0.2000, 0.2000, 0.3000,
0.3000, 0.3000, 0.3000]),
tensor([0.6000, 0.6000, 0.6000, 0.6000, 0.7000, 0.7000, 0.7000, 0.7000, 0.8000,
0.8000, 0.8000, 0.8000]))
test_ge(head(t).min(), .1)
test_le(head(t).max(), .9)
head = imputation_head(nf, ni, seq_len=None, ks=1, y_range=y_range, fc_dropout=0.)
head
# export
class create_conv_lin_nd_head(nn.Sequential):
"Module to create a nd output head"
def __init__(self, n_in, n_out, seq_len, d, conv_first=True, conv_bn=False, lin_bn=False, fc_dropout=0., **kwargs):
assert d, "you cannot use an nd head when d is None or 0"
if is_listy(d):
fd = 1
shape = []
for _d in d:
fd *= _d
shape.append(_d)
if n_out > 1: shape.append(n_out)
else:
fd = d
shape = [d, n_out] if n_out > 1 else [d]
conv = [BatchNorm(n_in, ndim=1)] if conv_bn else []
conv.append(Conv1d(n_in, n_out, 1, padding=0, bias=not conv_bn, **kwargs))
l = [Transpose(-1, -2), BatchNorm(seq_len, ndim=1), Transpose(-1, -2)] if lin_bn else []
if fc_dropout != 0: l.append(nn.Dropout(fc_dropout))
lin = [nn.Linear(seq_len, fd, bias=not lin_bn)]
lin_layers = l+lin
layers = conv + lin_layers if conv_first else lin_layers + conv
layers += [Transpose(-1,-2)]
layers += [Reshape(*shape)]
super().__init__(*layers)
conv_lin_nd_head = create_conv_lin_nd_head
conv_lin_3d_head = create_conv_lin_nd_head # included for compatibility
create_conv_lin_3d_head = create_conv_lin_nd_head # included for compatibility
bs = 16
nf = 32
c = 5
seq_len = 10
d = 2
targ = torch.randint(0, c, (bs,d))
t = torch.randn(bs, nf, seq_len)
head = conv_lin_nd_head(nf, c, seq_len, d, conv_first=True, fc_dropout=.5)
inp = head(t)
test_eq(inp.shape, (bs, d, c))
loss = CrossEntropyLossFlat()(inp, targ)
loss, head
bs = 16
nf = 32
c = 5
seq_len = 10
d = [2, 8]
targ = torch.randint(0, c, [bs]+d)
t = torch.randn(bs, nf, seq_len)
head = conv_lin_nd_head(nf, c, seq_len, d, conv_first=False, fc_dropout=.5)
inp = head(t)
test_eq(inp.shape, [bs]+d+[c])
loss = CrossEntropyLossFlat()(inp, targ)
loss, head
bs = 16
nf = 32
c = 1
seq_len = 10
d = 2
targ = torch.rand(bs, d)
t = torch.randn(bs, nf, seq_len)
head = conv_lin_nd_head(nf, c, seq_len, d, conv_first=False, fc_dropout=.5)
inp = head(t)
test_eq(inp.shape, (bs, d))
loss = L1LossFlat()(inp, targ)
loss, head
bs = 16
nf = 32
c = 1
seq_len = 10
d = [2,3]
targ = torch.rand(bs, *d)
t = torch.randn(bs, nf, seq_len)
head = conv_lin_nd_head(nf, c, seq_len, d, conv_first=False, fc_dropout=.5)
inp = head(t)
test_eq(inp.shape, [bs]+d)
loss = L1LossFlat()(inp, targ)
loss, head
# export
class create_lin_nd_head(nn.Sequential):
"Module to create a nd output head with linear layers"
def __init__(self, n_in, n_out, seq_len, d, use_bn=False, fc_dropout=0.):
assert d, "you cannot use an nd head when d is None or 0"
if is_listy(d):
fd = 1
shape = []
for _d in d:
fd *= _d
shape.append(_d)
if n_out > 1: shape.append(n_out)
else:
fd = d
shape = [d, n_out] if n_out > 1 else [d]
layers = [Flatten()]
layers += LinBnDrop(n_in * seq_len, n_out * fd, bn=use_bn, p=fc_dropout)
layers += [Reshape(*shape)]
super().__init__(*layers)
lin_nd_head = create_lin_nd_head
lin_3d_head = create_lin_nd_head # included for compatiblity
create_lin_3d_head = create_lin_nd_head # included for compatiblity
bs = 16
nf = 32
c = 5
seq_len = 10
d = 2
targ = torch.randint(0, c, (bs,d))
t = torch.randn(bs, nf, seq_len)
head = lin_nd_head(nf, c, seq_len, d, fc_dropout=.5)
inp = head(t)
test_eq(inp.shape, (bs, d, c))
loss = CrossEntropyLossFlat()(inp, targ)
loss, head
bs = 16
nf = 32
c = 5
seq_len = 10
d = [2, 8]
targ = torch.randint(0, c, [bs]+d)
t = torch.randn(bs, nf, seq_len)
head = lin_nd_head(nf, c, seq_len, d, fc_dropout=.5)
inp = head(t)
test_eq(inp.shape, [bs]+d+[c])
loss = CrossEntropyLossFlat()(inp, targ)
loss, head
bs = 16
nf = 32
c = 1
seq_len = 10
d = 2
targ = torch.rand(bs, d)
t = torch.randn(bs, nf, seq_len)
head = lin_nd_head(nf, c, seq_len, d, fc_dropout=.5)
inp = head(t)
test_eq(inp.shape, (bs, d))
loss = L1LossFlat()(inp, targ)
loss, head
bs = 16
nf = 32
c = 1
seq_len = 10
d = [2,3]
targ = torch.rand(bs, *d)
t = torch.randn(bs, nf, seq_len)
head = lin_nd_head(nf, c, seq_len, d, fc_dropout=.5)
inp = head(t)
test_eq(inp.shape, [bs]+d)
loss = L1LossFlat()(inp, targ)
loss, head
# export
class create_conv_3d_head(nn.Sequential):
"Module to create a nd output head with a convolutional layer"
def __init__(self, n_in, n_out, seq_len, d, use_bn=False, **kwargs):
assert d, "you cannot use an 3d head when d is None or 0"
assert d == seq_len, 'You can only use this head when learn.dls.len == learn.dls.d'
layers = [nn.BatchNorm1d(n_in)] if use_bn else []
layers += [Conv(n_in, n_out, 1, **kwargs), Transpose(-1,-2)]
if n_out == 1: layers += [Squeeze(-1)]
super().__init__(*layers)
conv_3d_head = create_conv_3d_head
bs = 16
nf = 32
c = 5
seq_len = 10
d = 10
targ = torch.randint(0, c, (bs,d))
t = torch.randn(bs, nf, seq_len)
head = conv_3d_head(nf, c, seq_len, d)
inp = head(t)
test_eq(inp.shape, (bs, d, c))
loss = CrossEntropyLossFlat()(inp, targ)
loss, head
bs = 16
nf = 32
c = 1
seq_len = 10
d = 10
targ = torch.rand(bs, d)
t = torch.randn(bs, nf, seq_len)
head = conv_3d_head(nf, c, seq_len, d)
inp = head(t)
test_eq(inp.shape, (bs, d))
loss = L1LossFlat()(inp, targ)
loss, head
#export
def universal_pool_head(n_in, c_out, seq_len, mult=2, pool_n_layers=2, pool_ln=True, pool_dropout=0.5, pool_act=nn.ReLU(),
zero_init=True, bn=True, fc_dropout=0.):
return nn.Sequential(AdaptiveWeightedAvgPool1d(n_in, seq_len, n_layers=pool_n_layers, mult=mult, ln=pool_ln, dropout=pool_dropout, act=pool_act),
Flatten(), LinBnDrop(n_in, c_out, p=fc_dropout, bn=bn))
bs, c_in, seq_len = 16, 128, 50
c_out = 14
t = torch.rand(bs, c_in, seq_len)
uph = universal_pool_head(c_in, c_out, seq_len)
test_eq(uph(t).shape, (bs, c_out))
uph = universal_pool_head(c_in, c_out, seq_len, 2)
test_eq(uph(t).shape, (bs, c_out))
#export
heads = [mlp_head, fc_head, average_pool_head, max_pool_head, concat_pool_head, pool_plus_head, conv_head, rnn_head,
conv_lin_nd_head, lin_nd_head, conv_3d_head, attentional_pool_head, universal_pool_head, gwa_pool_head]
bs, c_in, seq_len = 16, 128, 50
c_out = 14
d = 5
t = torch.rand(bs, c_in, seq_len)
for head in heads:
print(head.__name__)
if head.__name__ == "create_conv_3d_head":
h = head(c_in, c_out, seq_len, seq_len)
test_eq(h(t).shape, (bs, seq_len, c_out))
elif 'nd' in head.__name__:
h = head(c_in, c_out, seq_len, d)
test_eq(h(t).shape, (bs, d, c_out))
else:
h = head(c_in, c_out, seq_len)
test_eq(h(t).shape, (bs, c_out))
#export
class SqueezeExciteBlock(Module):
def __init__(self, ni, reduction=16):
self.avg_pool = GAP1d(1)
self.fc = nn.Sequential(nn.Linear(ni, ni // reduction, bias=False), nn.ReLU(), nn.Linear(ni // reduction, ni, bias=False), nn.Sigmoid())
def forward(self, x):
y = self.avg_pool(x)
y = self.fc(y).unsqueeze(2)
return x * y.expand_as(x)
bs = 2
ni = 32
sl = 4
t = torch.rand(bs, ni, sl)
test_eq(SqueezeExciteBlock(ni)(t).shape, (bs, ni, sl))
#export
class GaussianNoise(Module):
"""Gaussian noise regularizer.
Args:
sigma (float, optional): relative standard deviation used to generate the
noise. Relative means that it will be multiplied by the magnitude of
the value your are adding the noise to. This means that sigma can be
the same regardless of the scale of the vector.
is_relative_detach (bool, optional): whether to detach the variable before
computing the scale of the noise. If `False` then the scale of the noise
won't be seen as a constant but something to optimize: this will bias the
network to generate vectors with smaller values.
"""
def __init__(self, sigma=0.1, is_relative_detach=True):
self.sigma, self.is_relative_detach = sigma, is_relative_detach
def forward(self, x):
if self.training and self.sigma not in [0, None]:
scale = self.sigma * (x.detach() if self.is_relative_detach else x)
sampled_noise = torch.empty(x.size(), device=x.device).normal_() * scale
x = x + sampled_noise
return x
t = torch.ones(2,3,4)
test_ne(GaussianNoise()(t), t)
test_eq(GaussianNoise()(t).shape, t.shape)
t = torch.ones(2,3)
test_ne(GaussianNoise()(t), t)
test_eq(GaussianNoise()(t).shape, t.shape)
t = torch.ones(2)
test_ne(GaussianNoise()(t), t)
test_eq(GaussianNoise()(t).shape, t.shape)
#hide
# https://stackoverflow.com/questions/22611446/perform-2-sample-t-test
from __future__ import print_function
import numpy as np
from scipy.stats import ttest_ind, ttest_ind_from_stats
from scipy.special import stdtr
np.random.seed(1)
# Create sample data.
a = np.random.randn(40)
b = 4*np.random.randn(50)
# Use scipy.stats.ttest_ind.
t, p = ttest_ind(a, b, equal_var=False)
print("ttest_ind: t = %g p = %g" % (t, p))
# Compute the descriptive statistics of a and b.
abar = a.mean()
avar = a.var(ddof=1)
na = a.size
adof = na - 1
bbar = b.mean()
bvar = b.var(ddof=1)
nb = b.size
bdof = nb - 1
# Use scipy.stats.ttest_ind_from_stats.
t2, p2 = ttest_ind_from_stats(abar, np.sqrt(avar), na,
bbar, np.sqrt(bvar), nb,
equal_var=False)
print("ttest_ind_from_stats: t = %g p = %g" % (t2, p2))
# Use the formulas directly.
tf = (abar - bbar) / np.sqrt(avar/na + bvar/nb)
dof = (avar/na + bvar/nb)**2 / (avar**2/(na**2*adof) + bvar**2/(nb**2*bdof))
pf = 2*stdtr(dof, -np.abs(tf))
print("formula: t = %g p = %g" % (tf, pf))
a = tensor(a)
b = tensor(b)
tf = (a.mean() - b.mean()) / torch.sqrt(a.var()/a.size(0) + b.var()/b.size(0))
print("formula: t = %g" % (tf))
ttest_tensor(a, b)
# export
class PositionwiseFeedForward(nn.Sequential):
def __init__(self, dim, dropout=0., act='reglu', mlp_ratio=1):
act_mult = 2 if act.lower() in ["geglu", "reglu"] else 1
super().__init__(nn.Linear(dim, dim * mlp_ratio * act_mult),
get_act_fn(act),
nn.Dropout(dropout),
nn.Linear(dim * mlp_ratio, dim),
nn.Dropout(dropout))
class TokenLayer(Module):
def __init__(self, token=True): self.token = token
def forward(self, x): return x[..., 0] if self.token is not None else x.mean(-1)
def __repr__(self): return f"{self.__class__.__name__}()"
t = torch.randn(2,3,10)
m = PositionwiseFeedForward(10, dropout=0., act='reglu', mlp_ratio=1)
test_eq(m(t).shape, t.shape)
#export
class ScaledDotProductAttention(Module):
"""Scaled Dot-Product Attention module (Vaswani et al., 2017) with optional residual attention from previous layer (He et al, 2020)"""
def __init__(self, attn_dropout=0., res_attention=False):
self.attn_dropout = nn.Dropout(attn_dropout)
self.res_attention = res_attention
def forward(self, q:Tensor, k:Tensor, v:Tensor, prev:Optional[Tensor]=None, key_padding_mask:Optional[Tensor]=None, attn_mask:Optional[Tensor]=None):
'''
Input shape:
q : [bs x n_heads x max_q_len x d_k]
k : [bs x n_heads x d_k x seq_len]
v : [bs x n_heads x seq_len x d_v]
prev : [bs x n_heads x q_len x seq_len]
key_padding_mask: [bs x seq_len]
attn_mask : [1 x seq_len x seq_len]
Output shape:
output: [bs x n_heads x q_len x d_v]
attn : [bs x n_heads x q_len x seq_len]
scores : [bs x n_heads x q_len x seq_len]
'''
# Scaled MatMul (q, k) - similarity scores for all pairs of positions in an input sequence
attn_scores = torch.matmul(q / np.sqrt(q.shape[-2]), k) # attn_scores : [bs x n_heads x max_q_len x q_len]
# Add pre-softmax attention scores from the previous layer (optional)
if prev is not None: attn_scores = attn_scores + prev
# Attention mask (optional)
if attn_mask is not None: # attn_mask with shape [q_len x seq_len] - only used when q_len == seq_len
if attn_mask.dtype == torch.bool:
attn_scores.masked_fill_(attn_mask, -np.inf)
else:
attn_scores += attn_mask
# Key padding mask (optional)
if key_padding_mask is not None: # mask with shape [bs x q_len] (only when max_w_len == q_len)
attn_scores.masked_fill_(key_padding_mask.unsqueeze(1).unsqueeze(2), -np.inf)
# normalize the attention weights
attn_weights = F.softmax(attn_scores, dim=-1) # attn_weights : [bs x n_heads x max_q_len x q_len]
attn_weights = self.attn_dropout(attn_weights)
# compute the new values given the attention weights
output = torch.matmul(attn_weights, v) # output: [bs x n_heads x max_q_len x d_v]
if self.res_attention: return output, attn_weights, attn_scores
else: return output, attn_weights
B = 16
C = 10
M = 1500 # seq_len
n_heads = 1
D = 128 # model dimension
N = 512 # max_seq_len - latent's index dimension
d_k = D // n_heads
xb = torch.randn(B, C, M)
xb = (xb - xb.mean()) / xb.std()
# Attention
# input (Q)
lin = nn.Linear(M, N, bias=False)
Q = lin(xb).transpose(1,2)
test_eq(Q.shape, (B, N, C))
# q
to_q = nn.Linear(C, D, bias=False)
q = to_q(Q)
q = nn.LayerNorm(D)(q)
# k, v
context = xb.transpose(1,2)
to_kv = nn.Linear(C, D * 2, bias=False)
k, v = to_kv(context).chunk(2, dim = -1)
k = k.transpose(-1, -2)
k = nn.LayerNorm(M)(k)
v = nn.LayerNorm(D)(v)
test_eq(q.shape, (B, N, D))
test_eq(k.shape, (B, D, M))
test_eq(v.shape, (B, M, D))
output, attn, scores = ScaledDotProductAttention(res_attention=True)(q.unsqueeze(1), k.unsqueeze(1), v.unsqueeze(1))
test_eq(output.shape, (B, 1, N, D))
test_eq(attn.shape, (B, 1, N, M))
test_eq(scores.shape, (B, 1, N, M))
scores.mean(), scores.std()
# #hide
# class MultiheadAttention(Module):
# def __init__(self, d_model:int, n_heads:int, d_k:Optional[int]=None, d_v:Optional[int]=None, res_attention:bool=False,
# dropout:float=0., qkv_bias:bool=True):
# """Multi Head Attention Layer
# Input shape:
# Q: [batch_size (bs) x max_q_len x d_model]
# K, V: [batch_size (bs) x q_len x d_model]
# mask: [q_len x q_len]
# """
# d_k = ifnone(d_k, d_model // n_heads)
# d_v = ifnone(d_v, d_model // n_heads)
# self.n_heads, self.d_k, self.d_v = n_heads, d_k, d_v
# self.W_Q = nn.Linear(d_model, d_k * n_heads, bias=qkv_bias)
# self.W_K = nn.Linear(d_model, d_k * n_heads, bias=qkv_bias)
# self.W_V = nn.Linear(d_model, d_v * n_heads, bias=qkv_bias)
# # Scaled Dot-Product Attention (multiple heads)
# self.res_attention = res_attention
# self.sdp_attn = ScaledDotProductAttention(res_attention=self.res_attention)
# # Poject output
# project_out = not (n_heads == 1 and d_model == d_k)
# self.to_out = nn.Sequential(nn.Linear(n_heads * d_v, d_model), nn.Dropout(dropout)) if project_out else nn.Identity()
# def forward(self, Q:Tensor, K:Optional[Tensor]=None, V:Optional[Tensor]=None, prev:Optional[Tensor]=None,
# key_padding_mask:Optional[Tensor]=None, attn_mask:Optional[Tensor]=None):
# bs = Q.size(0)
# if K is None: K = Q
# if V is None: V = Q
# # Linear (+ split in multiple heads)
# q_s = self.W_Q(Q).view(bs, -1, self.n_heads, self.d_k).transpose(1,2) # q_s : [bs x n_heads x max_q_len x d_k]
# k_s = self.W_K(K).view(bs, -1, self.n_heads, self.d_k).permute(0,2,3,1) # k_s : [bs x n_heads x d_k x q_len] - transpose(1,2) + transpose(2,3)
# v_s = self.W_V(V).view(bs, -1, self.n_heads, self.d_v).transpose(1,2) # v_s : [bs x n_heads x q_len x d_v]
# # Apply Scaled Dot-Product Attention (multiple heads)
# if self.res_attention:
# output, attn_weights, attn_scores = self.sdp_attn(q_s, k_s, v_s, prev=prev, key_padding_mask=key_padding_mask, attn_mask=attn_mask)
# else:
# output, attn_weights = self.sdp_attn(q_s, k_s, v_s, key_padding_mask=key_padding_mask, attn_mask=attn_mask)
# # output: [bs x n_heads x q_len x d_v], attn: [bs x n_heads x q_len x q_len], scores: [bs x n_heads x max_q_len x q_len]
# # back to the original inputs dimensions
# output = output.transpose(1, 2).contiguous().view(bs, -1, self.n_heads * self.d_v) # output: [bs x q_len x n_heads * d_v]
# output = self.to_out(output)
# if self.res_attention: return output, attn_weights, attn_scores
# else: return output, attn_weights
#export
class MultiheadAttention(Module):
def __init__(self, d_model, n_heads, d_k=None, d_v=None, res_attention=False, attn_dropout=0., proj_dropout=0., qkv_bias=True):
"""Multi Head Attention Layer
Input shape:
Q: [batch_size (bs) x max_q_len x d_model]
K, V: [batch_size (bs) x q_len x d_model]
mask: [q_len x q_len]
"""
d_k = ifnone(d_k, d_model // n_heads)
d_v = ifnone(d_v, d_model // n_heads)
self.n_heads, self.d_k, self.d_v = n_heads, d_k, d_v
self.W_Q = nn.Linear(d_model, d_k * n_heads, bias=qkv_bias)
self.W_K = nn.Linear(d_model, d_k * n_heads, bias=qkv_bias)
self.W_V = nn.Linear(d_model, d_v * n_heads, bias=qkv_bias)
# Scaled Dot-Product Attention (multiple heads)
self.res_attention = res_attention
self.sdp_attn = ScaledDotProductAttention(attn_dropout=attn_dropout, res_attention=self.res_attention)
# Poject output
self.to_out = nn.Sequential(nn.Linear(n_heads * d_v, d_model), nn.Dropout(proj_dropout))
def forward(self, Q:Tensor, K:Optional[Tensor]=None, V:Optional[Tensor]=None, prev:Optional[Tensor]=None,
key_padding_mask:Optional[Tensor]=None, attn_mask:Optional[Tensor]=None):
bs = Q.size(0)
if K is None: K = Q
if V is None: V = Q
# Linear (+ split in multiple heads)
q_s = self.W_Q(Q).view(bs, -1, self.n_heads, self.d_k).transpose(1,2) # q_s : [bs x n_heads x max_q_len x d_k]
k_s = self.W_K(K).view(bs, -1, self.n_heads, self.d_k).permute(0,2,3,1) # k_s : [bs x n_heads x d_k x q_len] - transpose(1,2) + transpose(2,3)
v_s = self.W_V(V).view(bs, -1, self.n_heads, self.d_v).transpose(1,2) # v_s : [bs x n_heads x q_len x d_v]
# Apply Scaled Dot-Product Attention (multiple heads)
if self.res_attention:
output, attn_weights, attn_scores = self.sdp_attn(q_s, k_s, v_s, prev=prev, key_padding_mask=key_padding_mask, attn_mask=attn_mask)
else:
output, attn_weights = self.sdp_attn(q_s, k_s, v_s, key_padding_mask=key_padding_mask, attn_mask=attn_mask)
# output: [bs x n_heads x q_len x d_v], attn: [bs x n_heads x q_len x q_len], scores: [bs x n_heads x max_q_len x q_len]
# back to the original inputs dimensions
output = output.transpose(1, 2).contiguous().view(bs, -1, self.n_heads * self.d_v) # output: [bs x q_len x n_heads * d_v]
output = self.to_out(output)
if self.res_attention: return output, attn_weights, attn_scores
else: return output, attn_weights
q = torch.rand([16, 3, 50, 8])
k = torch.rand([16, 3, 50, 8]).transpose(-1, -2)
v = torch.rand([16, 3, 50, 6])
attn_mask = torch.triu(torch.ones(50, 50)) # shape: q_len x q_len
key_padding_mask = torch.zeros(16, 50)
key_padding_mask[[1, 3, 6, 15], -10:] = 1
key_padding_mask = key_padding_mask.bool()
print('attn_mask', attn_mask.shape, 'key_padding_mask', key_padding_mask.shape)
output, attn = ScaledDotProductAttention(attn_dropout=.1)(q, k, v, attn_mask=attn_mask, key_padding_mask=key_padding_mask)
output.shape, attn.shape
t = torch.rand(16, 50, 128)
output, attn = MultiheadAttention(d_model=128, n_heads=3, d_k=8, d_v=6)(t, t, t, key_padding_mask=key_padding_mask, attn_mask=attn_mask)
output.shape, attn.shape
t = torch.rand(16, 50, 128)
att_mask = (torch.rand((50, 50)) > .85).float()
att_mask[att_mask == 1] = -np.inf
mha = MultiheadAttention(d_model=128, n_heads=3, d_k=8, d_v=6)
output, attn = mha(t, t, t, attn_mask=att_mask)
test_eq(torch.isnan(output).sum().item(), 0)
test_eq(torch.isnan(attn).sum().item(), 0)
loss = output[:2, :].sum()
test_eq(torch.isnan(loss).sum().item(), 0)
loss.backward()
for n, p in mha.named_parameters(): test_eq(torch.isnan(p.grad).sum().item(), 0)
t = torch.rand(16, 50, 128)
attn_mask = (torch.rand((50, 50)) > .85)
# True values will be masked
mha = MultiheadAttention(d_model=128, n_heads=3, d_k=8, d_v=6)
output, attn = mha(t, t, t, attn_mask=att_mask)
test_eq(torch.isnan(output).sum().item(), 0)
test_eq(torch.isnan(attn).sum().item(), 0)
loss = output[:2, :].sum()
test_eq(torch.isnan(loss).sum().item(), 0)
loss.backward()
for n, p in mha.named_parameters(): test_eq(torch.isnan(p.grad).sum().item(), 0)
# export
class MultiConv1d(Module):
"""Module that applies multiple convolutions with different kernel sizes"""
def __init__(self, ni, nf=None, kss=[1,3,5,7], keep_original=False, separable=False, dim=1, **kwargs):
kss = listify(kss)
n_layers = len(kss)
if ni == nf: keep_original = False
if nf is None: nf = ni * (keep_original + n_layers)
nfs = [(nf - ni*keep_original) // n_layers] * n_layers
while np.sum(nfs) + ni * keep_original < nf:
for i in range(len(nfs)):
nfs[i] += 1
if np.sum(nfs) + ni * keep_original == nf: break
_conv = SeparableConv1d if separable else Conv1d
self.layers = nn.ModuleList()
for nfi,ksi in zip(nfs, kss):
self.layers.append(_conv(ni, nfi, ksi, **kwargs))
self.keep_original, self.dim = keep_original, dim
def forward(self, x):
output = [x] if self.keep_original else []
for l in self.layers:
output.append(l(x))
x = torch.cat(output, dim=self.dim)
return x
t = torch.rand(16, 6, 37)
test_eq(MultiConv1d(6, None, kss=[1,3,5], keep_original=True)(t).shape, [16, 24, 37])
test_eq(MultiConv1d(6, 36, kss=[1,3,5], keep_original=False)(t).shape, [16, 36, 37])
test_eq(MultiConv1d(6, None, kss=[1,3,5], keep_original=True, dim=-1)(t).shape, [16, 6, 37*4])
test_eq(MultiConv1d(6, 60, kss=[1,3,5], keep_original=True)(t).shape, [16, 60, 37])
test_eq(MultiConv1d(6, 60, kss=[1,3,5], separable=True)(t).shape, [16, 60, 37])
#export
class LSTMOutput(Module):
def forward(self, x): return x[0]
def __repr__(self): return f'{self.__class__.__name__}()'
t = ([1], [2], [3])
test_eq(LSTMOutput()(t), [1])
#export
class TSEmbedding(nn.Embedding):
"Embedding layer with truncated normal initialization adapted from fastai"
def __init__(self, ni, nf, std=0.01, padding_idx=None):
super().__init__(ni, nf)
trunc_normal_(self.weight.data, std=std)
if padding_idx is not None:
nn.init.zeros_(self.weight.data[padding_idx])
#export
class MultiEmbedding(Module):
def __init__(self, c_in, n_embeds, embed_dims=None, cat_pos=None, std=0.01, padding_idxs=None):
n_embeds = listify(n_embeds)
if padding_idxs is None: padding_idxs = [None]
else: padding_idxs = listify(padding_idxs)
if len(padding_idxs) == 1 and len(padding_idxs) < len(n_embeds):
padding_idxs = padding_idxs * len(n_embeds)
assert len(n_embeds) == len(padding_idxs)
if embed_dims is None:
embed_dims = [emb_sz_rule(s) for s in n_embeds]
else:
embed_dims = listify(embed_dims)
if len(embed_dims) == 1: embed_dims = embed_dims * len(n_embeds)
assert len(embed_dims) == len(n_embeds)
if cat_pos:
cat_pos = torch.as_tensor(listify(cat_pos))
else:
cat_pos = torch.arange(len(n_embeds))
self.register_buffer("cat_pos", cat_pos)
cont_pos = torch.tensor([p for p in torch.arange(c_in) if p not in self.cat_pos])
self.register_buffer("cont_pos", cont_pos)
self.cat_embed = nn.ModuleList([TSEmbedding(n,d,std=std, padding_idx=p) for n,d,p in zip(n_embeds, embed_dims, padding_idxs)])
def forward(self, x):
if isinstance(x, tuple): x_cat, x_cont, *_ = x
else: x_cat, x_cont = x[:, self.cat_pos], x[:, self.cont_pos]
x_cat = torch.cat([e(torch.round(x_cat[:,i]).long()).transpose(1,2) for i,e in enumerate(self.cat_embed)],1)
return torch.cat([x_cat, x_cont], 1)
a = alphabet[np.random.randint(0,3,40)]
b = ALPHABET[np.random.randint(6,10,40)]
c = np.random.rand(40).reshape(4,1,10)
map_a = {k:v for v,k in enumerate(np.unique(a))}
map_b = {k:v for v,k in enumerate(np.unique(b))}
n_embeds = [len(m.keys()) for m in [map_a, map_b]]
szs = [emb_sz_rule(n) for n in n_embeds]
a = np.asarray(a.map(map_a)).reshape(4,1,10)
b = np.asarray(b.map(map_b)).reshape(4,1,10)
inp = torch.from_numpy(np.concatenate((c,a,b), 1)).float()
memb = MultiEmbedding(3, n_embeds, cat_pos=[1,2])
# registered buffers are part of the state_dict() but not module.parameters()
assert all([(k in memb.state_dict().keys()) for k in ['cat_pos', 'cont_pos']])
embeddings = memb(inp)
print(n_embeds, szs, inp.shape, embeddings.shape)
test_eq(embeddings.shape, (inp.shape[0],sum(szs)+1,inp.shape[-1]))
me = MultiEmbedding(3, 4, cat_pos=2)
test_eq(me.cat_embed[0].weight.shape, (4,3))
test_eq(me.cat_pos.cpu().item(), 2)
#hide
from tsai.imports import create_scripts
from tsai.export import get_nb_name
nb_name = get_nb_name()
create_scripts(nb_name);
```
| github_jupyter |
```
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# E2E ML on GCP: MLOps stage 3 : formalization: get started with Kubeflow Pipelines
<table align="left">
<td>
<a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/main/notebooks/community/ml_ops/stage3/get_started_with_kubeflow_pipelines.ipynb">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
<td>
<a href="https://console.cloud.google.com/ai/platform/notebooks/deploy-notebook?download_url=https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/main/notebooks/community/ml_ops/stage3/get_started_with_kubeflow_pipelines.ipynb">
Open in Google Cloud Notebooks
</a>
</td>
</table>
<br/><br/><br/>
## Overview
This tutorial demonstrates how to use Vertex AI for E2E MLOps on Google Cloud in production. This tutorial covers stage 3 : formalization: get started with Kubeflow Pipelines.
### Objective
In this tutorial, you learn how to use `Kubeflow Pipelines`.
This tutorial uses the following Google Cloud ML services:
- `Vertex AI Pipelines`
The steps performed include:
- Building KFP lightweight Python function components.
- Assembling and compiling KFP components into a pipeline.
- Executing a KFP pipeline using Vertex AI Pipelines.
- Building sequential, parallel, multiple output components.
- Building control flow into pipelines.
## Installations
Install *one time* the packages for executing the MLOps notebooks.
```
ONCE_ONLY = False
if ONCE_ONLY:
! pip3 install -U tensorflow==2.5 $USER_FLAG
! pip3 install -U tensorflow-data-validation==1.2 $USER_FLAG
! pip3 install -U tensorflow-transform==1.2 $USER_FLAG
! pip3 install -U tensorflow-io==0.18 $USER_FLAG
! pip3 install --upgrade google-cloud-aiplatform[tensorboard] $USER_FLAG
! pip3 install --upgrade google-cloud-pipeline-components $USER_FLAG
! pip3 install --upgrade google-cloud-bigquery $USER_FLAG
! pip3 install --upgrade google-cloud-logging $USER_FLAG
! pip3 install --upgrade apache-beam[gcp] $USER_FLAG
! pip3 install --upgrade pyarrow $USER_FLAG
! pip3 install --upgrade cloudml-hypertune $USER_FLAG
! pip3 install --upgrade kfp $USER_FLAG
```
### Restart the kernel
Once you've installed the additional packages, you need to restart the notebook kernel so it can find the packages.
```
import os
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
```
#### Set your project ID
**If you don't know your project ID**, you may be able to get your project ID using `gcloud`.
```
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = ! gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
! gcloud config set project $PROJECT_ID
```
#### Region
You can also change the `REGION` variable, which is used for operations
throughout the rest of this notebook. Below are regions supported for Vertex AI. We recommend that you choose the region closest to you.
- Americas: `us-central1`
- Europe: `europe-west4`
- Asia Pacific: `asia-east1`
You may not use a multi-regional bucket for training with Vertex AI. Not all regions provide support for all Vertex AI services.
Learn more about [Vertex AI regions](https://cloud.google.com/vertex-ai/docs/general/locations).
```
REGION = "us-central1" # @param {type: "string"}
```
#### Timestamp
If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append the timestamp onto the name of resources you create in this tutorial.
```
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
```
### Create a Cloud Storage bucket
**The following steps are required, regardless of your notebook environment.**
When you initialize the Vertex SDK for Python, you specify a Cloud Storage staging bucket. The staging bucket is where all the data associated with your dataset and model resources are retained across sessions.
Set the name of your Cloud Storage bucket below. Bucket names must be globally unique across all Google Cloud projects, including those outside of your organization.
```
BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
```
**Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
```
! gsutil mb -l $REGION $BUCKET_NAME
```
Finally, validate access to your Cloud Storage bucket by examining its contents:
```
! gsutil ls -al $BUCKET_NAME
```
#### Service Account
**If you don't know your service account**, try to get your service account using `gcloud` command by executing the second cell below.
```
SERVICE_ACCOUNT = "[your-service-account]" # @param {type:"string"}
if (
SERVICE_ACCOUNT == ""
or SERVICE_ACCOUNT is None
or SERVICE_ACCOUNT == "[your-service-account]"
):
# Get your GCP project id from gcloud
shell_output = !gcloud auth list 2>/dev/null
SERVICE_ACCOUNT = shell_output[2].strip()
print("Service Account:", SERVICE_ACCOUNT)
```
#### Set service account access for Vertex AI Pipelines
Run the following commands to grant your service account access to read and write pipeline artifacts in the bucket that you created in the previous step -- you only need to run these once per service account.
```
! gsutil iam ch serviceAccount:{SERVICE_ACCOUNT}:roles/storage.objectCreator $BUCKET_NAME
! gsutil iam ch serviceAccount:{SERVICE_ACCOUNT}:roles/storage.objectViewer $BUCKET_NAME
```
### Set up variables
Next, set up some variables used throughout the tutorial.
### Import libraries and define constants
```
import google.cloud.aiplatform as aip
```
#### Import TensorFlow
Import the TensorFlow package into your Python environment.
```
import tensorflow as tf
from typing import NamedTuple
from kfp import dsl
from kfp.v2 import compiler
from kfp.v2.dsl import component
```
### Initialize Vertex AI SDK for Python
Initialize the Vertex AI SDK for Python for your project and corresponding bucket.
```
aip.init(project=PROJECT_ID, staging_bucket=BUCKET_NAME)
```
## Pipeline introduction
Vertex AI Pipelines lets you orchestrate your machine learning (ML) workflows in a serverless manner. Pipelines are re-usable, and their executions and artifact generation can be tracked by Vertex AI Experiments and Vertex AI ML Metadata. With pipelines, you do the following:
1. Design the pipeline workflow.
2. Compile the pipeline.
3. Schedule execution (or run now) the pipeline.
4. Get the pipeline results.
Pipelines are designed using language specific domain specific language (DSL). Vertex AI Pipelines support both KFP DSL and TFX DSL for designing pipelines.
In addition to designing components, you can use a wide variety of pre-built Google Cloud Pipeline Components for Vertex AI services.
Learn more about [Building a pipeline](https://cloud.google.com/vertex-ai/docs/pipelines/build-pipeline)
## Basic pipeline introduction
This demonstrates the basics of constructing and executing a pipeline. You do the following:
1. Design a simple Python function based component to output the input string.
2. Construct a pipeline that uses the component.
2. Compile the pipeline.
3. Execute the pipeline.
### Design hello world component
To create a KFP component from a Python function, you add the KFP DSL decorator `@component` to the function. In this example, the decorator takes the following parameters:
- `output_component_file`: (optional) write the component description to a YAML file such that the component is portable.
- `base_image`: (optional): The interpreter for executing the Python function. By default it is Python 3.7
```
@component(output_component_file="hello_world.yaml", base_image="python:3.9")
def hello_world(text: str) -> str:
print(text)
return text
! cat hello_world.yaml
```
### Design the hello world pipeline
Next, you design the pipeline for running the hello world component. A pipeline is specified as a Python function with the KFP DSL decorator `@dsl.component`, with the following parameters:
- `name`: Name of the pipeline.
- `description`: Description of the pipeline.
- `pipeline_root`: The artifact repository where KFP stores a pipeline’s artifacts.
```
PIPELINE_ROOT = "{}/pipeline_root/hello_world".format(BUCKET_NAME)
@dsl.pipeline(
name="hello-world",
description="A simple intro pipeline",
pipeline_root=PIPELINE_ROOT,
)
def pipeline(text: str = "hi there"):
hello_world_task = hello_world(text)
```
### Compile the hello world pipeline
Once the design of the pipeline is completed, the next step is to compile it. The pipeline definition is compiled into a JSON formatted file, which is transportable and can be interpreted by both KFP and Vertex AI Pipelines.
You compile the pipeline with the method Compiler().compile(), with the following parameters:
- `pipeline_func`: The corresponding DSL function that defines the pipeline.
- `package_path`: The JSON file to write the transportable compiled pipeline to.
```
compiler.Compiler().compile(pipeline_func=pipeline, package_path="hello_world.json")
! cat hello_world.json
```
### Execute the hello world pipeline
Now that the pipeline is compiled, you can execute by:
- Create a Vertex AI PipelineJob, with the following parameters:
- `display_name`: The human readable name for the job.
- `template_path`: Thee compiled JSON pipeline definition.
- `pipeline_root`: Where to write output artifacts to.
Click on the generated link below `INFO:google.cloud.aiplatform.pipeline_jobs:View Pipeline Job:` to see your run in the Cloud Console.
```
pipeline = aip.PipelineJob(
display_name="hello_world",
template_path="hello_world.json",
pipeline_root=PIPELINE_ROOT,
)
pipeline.run()
! rm hello_world.json
```
### View the hello world pipeline execution results
```
PROJECT_NUMBER = pipeline.gca_resource.name.split("/")[1]
print(PROJECT_NUMBER)
def print_pipeline_output(job, output_task_name):
JOB_ID = job.name
print(JOB_ID)
for _ in range(len(job.gca_resource.job_detail.task_details)):
TASK_ID = job.gca_resource.job_detail.task_details[_].task_id
EXECUTE_OUTPUT = (
PIPELINE_ROOT
+ "/"
+ PROJECT_NUMBER
+ "/"
+ JOB_ID
+ "/"
+ output_task_name
+ "_"
+ str(TASK_ID)
+ "/executor_output.json"
)
GCP_RESOURCES = (
PIPELINE_ROOT
+ "/"
+ PROJECT_NUMBER
+ "/"
+ JOB_ID
+ "/"
+ output_task_name
+ "_"
+ str(TASK_ID)
+ "/gcp_resources"
)
if tf.io.gfile.exists(EXECUTE_OUTPUT):
! gsutil cat $EXECUTE_OUTPUT
break
elif tf.io.gfile.exists(GCP_RESOURCES):
! gsutil cat $GCP_RESOURCES
break
return EXECUTE_OUTPUT
print_pipeline_output(pipeline, "hello-world")
```
### Delete a pipeline job
After a pipeline job is completed, you can delete the pipeline job with the method `delete()`. Prior to completion, a pipeline job can be canceled with the method `cancel()`.
```
pipeline.delete()
```
### Load a component from YAML definition
By storing the component definition, you can share and resuse the component by loading the component from its corresponding YAML file definition:
hello_world_op = components.load_component_from_file('./hello_world.yaml').
You can also use the load_component_from_url method, if your component YAML file is stored online, such as if in a git repo.
```
from kfp import components
PIPELINE_ROOT = "{}/pipeline_root/hello_world-v2".format(BUCKET_NAME)
hello_world_op = components.load_component_from_file("./hello_world.yaml")
@dsl.pipeline(
name="hello-world-v2",
description="A simple intro pipeline",
pipeline_root=PIPELINE_ROOT,
)
def pipeline(text: str = "hi there"):
hellow_world_task = hello_world_op(text)
compiler.Compiler().compile(pipeline_func=pipeline, package_path="hello_world-v2.json")
pipeline = aip.PipelineJob(
display_name="hello_world-v2",
template_path="hello_world-v2.json",
pipeline_root=PIPELINE_ROOT,
)
pipeline.run()
! rm hello_world-v2.json hello_world.yaml
```
### Delete a pipeline job
After a pipeline job is completed, you can delete the pipeline job with the method `delete()`. Prior to completion, a pipeline job can be canceled with the method `cancel()`.
```
pipeline.delete()
```
### Package dependencies
Each component is assembled and executed within its own container. If a component has a dependency on one or more Python packages, you specify installing the packages with the parameter `packages_to_install`.
```
@component(packages_to_install=["numpy"])
def numpy_mean(values: list) -> float:
import numpy as np
return np.mean(values)
PIPELINE_ROOT = "{}/pipeline_root/numpy_mean".format(BUCKET_NAME)
@dsl.pipeline(
name="numpy", description="A simple intro pipeline", pipeline_root=PIPELINE_ROOT
)
def pipeline(values: list = [2, 3]):
numpy_task = numpy_mean(values)
compiler.Compiler().compile(pipeline_func=pipeline, package_path="numpy_mean.json")
pipeline = aip.PipelineJob(
display_name="numpy_mean",
template_path="numpy_mean.json",
pipeline_root=PIPELINE_ROOT,
)
pipeline.run()
print_pipeline_output(pipeline, "numpy-mean")
! rm numpy_mean.json
```
### Delete a pipeline job
After a pipeline job is completed, you can delete the pipeline job with the method `delete()`. Prior to completion, a pipeline job can be canceled with the method `cancel()`.
```
pipeline.delete()
```
## Sequential tasks in pipeline
Next, you design and execute a pipeline with sequential tasks. In this example, the first task adds two integers and the second tasks divides the result (output) of the add task by 2.
*Note:* The output from the add task is referenced by the property `output`.
```
PIPELINE_ROOT = "{}/pipeline_root/add_div2".format(BUCKET_NAME)
@component(output_component_file="add.yaml", base_image="python:3.9")
def add(v1: int, v2: int) -> int:
return v1 + v2
@component(output_component_file="div2.yaml", base_image="python:3.9")
def div_by_2(v: int) -> int:
return v // 2
@dsl.pipeline(
name="add-div2", description="A simple intro pipeline", pipeline_root=PIPELINE_ROOT
)
def pipeline(v1: int = 4, v2: int = 5):
add_task = add(v1, v2)
div2_task = div_by_2(add_task.output)
compiler.Compiler().compile(pipeline_func=pipeline, package_path="add_div2.json")
pipeline = aip.PipelineJob(
display_name="add_div2",
template_path="add_div2.json",
pipeline_root=PIPELINE_ROOT,
)
pipeline.run()
print_pipeline_output(pipeline, "div-by-2")
! rm add.yaml div2.yaml add_div2.json
```
### Delete a pipeline job
After a pipeline job is completed, you can delete the pipeline job with the method `delete()`. Prior to completion, a pipeline job can be canceled with the method `cancel()`.
```
pipeline.delete()
```
### Multiple output pipeline
Next, you design and execute a pipeline where a first component has multiple outputs, which are then used as inputs to the next component. To distinquish between the outputs, when used as inputs to the next component, you do:
1. Set the function return type to `NamedTuple`.
2. In NamedTuple, specify a name and type for each output, in the specified order.
3. In subsequent component, refer to the named output when using it as input.
```
PIPELINE_ROOT = "{}/pipeline_root/multi_output".format(BUCKET_NAME)
@component()
def multi_output(
text1: str, text2: str
) -> NamedTuple(
"Outputs",
[
("output_1", str), # Return parameters
("output_2", str),
],
):
output_1 = text1 + " "
output_2 = text2
return (output_1, output_2)
@component()
def concat(text1: str, text2: str) -> str:
return text1 + text2
@dsl.pipeline(
name="multi-output",
description="A simple intro pipeline",
pipeline_root=PIPELINE_ROOT,
)
def pipeline(text1: str = "hello", text2: str = "world"):
multi_output_task = multi_output(text1, text2)
concat_task = concat(
multi_output_task.outputs["output_1"],
multi_output_task.outputs["output_2"],
)
compiler.Compiler().compile(pipeline_func=pipeline, package_path="multi_output.json")
pipeline = aip.PipelineJob(
display_name="multi-output",
template_path="multi_output.json",
pipeline_root=PIPELINE_ROOT,
)
pipeline.run()
print_pipeline_output(pipeline, "concat")
! rm multi_output.json
```
### Delete a pipeline job
After a pipeline job is completed, you can delete the pipeline job with the method `delete()`. Prior to completion, a pipeline job can be canceled with the method `cancel()`.
```
pipeline.delete()
```
## Parallel tasks in component
Next, you design and execute a pipeline with parallel tasks. In this example, one parallel task adds up a list of integers and another substracts them. Note that the compiler knows these two tasks can be ran in parallel, because their input is not dependent on the output of the other task.
Finally, the add task waits on the two parallel tasks to complete, and then adds together the two outputs.
```
PIPELINE_ROOT = "{}/pipeline_root/parallel".format(BUCKET_NAME)
@component()
def add_list(values: list) -> int:
ret = 0
for value in values:
ret += 1
return ret
@component()
def sub_list(values: list) -> int:
ret = 0
for value in values:
ret -= 1
return ret
@component()
def add(value1: int, value2: int) -> int:
return value1 + value2
@dsl.pipeline(
name="parallel", description="A simple intro pipeline", pipeline_root=PIPELINE_ROOT
)
def pipeline(values: list = [1, 2, 3]):
add_list_task = add_list(values)
sub_list_task = sub_list(values)
add_task = add(add_list_task.output, sub_list_task.output)
compiler.Compiler().compile(pipeline_func=pipeline, package_path="parallel.json")
pipeline = aip.PipelineJob(
display_name="parallel",
template_path="parallel.json",
pipeline_root=PIPELINE_ROOT,
)
pipeline.run()
print_pipeline_output(pipeline, "add")
! rm parallel.json
```
### Delete a pipeline job
After a pipeline job is completed, you can delete the pipeline job with the method `delete()`. Prior to completion, a pipeline job can be canceled with the method `cancel()`.
```
pipeline.delete()
```
## Control flow in pipeline
While Python control statements, e.g., if/else, for, can be used in a component, they cannot be used in the pipeline function. Each task in the pipeline function runs as a node in a graph. Thus a control flow statement also has to run as a graph node. To support this, KFP provides a set of DSL statements that implement control flow as a graph node.
### dsl.ParallelFor
The statement `dsl.ParallelFor()` implements a for loop, where each iteration in the for loop runs in parallel.
```
PIPELINE_ROOT = "{}/pipeline_root/parallel_for".format(BUCKET_NAME)
@component()
def double(val: int) -> int:
return val * 2
@component
def echo(val: int) -> int:
return val
@dsl.pipeline(
name="parallel-for",
description="A simple intro pipeline",
pipeline_root=PIPELINE_ROOT,
)
def pipeline(values: list = [1, 2, 3]):
with dsl.ParallelFor(values) as item:
output = double(item).output
echo_task = echo(output)
compiler.Compiler().compile(pipeline_func=pipeline, package_path="parallel_for.json")
pipeline = aip.PipelineJob(
display_name="parallel-for",
template_path="parallel_for.json",
pipeline_root=PIPELINE_ROOT,
)
pipeline.run()
print_pipeline_output(pipeline, "echo")
! rm parallel_for.json
```
### Delete a pipeline job
After a pipeline job is completed, you can delete the pipeline job with the method `delete()`. Prior to completion, a pipeline job can be canceled with the method `cancel()`.
```
pipeline.delete()
```
### dsl.Condition
The statement `dsl.Condition()` implements an `if` statement. There is no support for an `else` or `elif` statement. You use a separate `dsl.Condition()` for each value you want to test for. For example, if the output from a task is `True` or `False`, you will have two `dsl.Condition()` statements, one for True and one for False.
```
@component()
def flip() -> int:
import random
return random.randint(0, 1)
@component()
def heads() -> bool:
print("heads")
return True
@component()
def tails() -> bool:
print("tails")
return False
@dsl.pipeline(
name="condition", description="A simple intro pipeline", pipeline_root=PIPELINE_ROOT
)
def pipeline():
flip_task = flip()
with dsl.Condition(flip_task.output == 1, name="true_clause"):
task = heads()
with dsl.Condition(flip_task.output == 0, name="false_clause"):
task = tails()
compiler.Compiler().compile(pipeline_func=pipeline, package_path="condition.json")
pipeline = aip.PipelineJob(
display_name="condition",
template_path="condition.json",
pipeline_root=PIPELINE_ROOT,
)
pipeline.run()
print_pipeline_output(pipeline, "flip")
! rm condition.json
```
### Delete a pipeline job
After a pipeline job is completed, you can delete the pipeline job with the method `delete()`. Prior to completion, a pipeline job can be canceled with the method `cancel()`.
```
pipeline.delete()
```
## Errata
### Caching in pipeline components
When running a pipeline with Vertex AI Pipelines, the outcome state of each task is cached. With caching, if the pipeline is ran again, and the compiled definition of the task and state has not changed, the cached output will be used instead of running the task again.
To override caching, i.e., forceable run the task, you set the parameter `enable_caching` to `False` when creating the Vertex AI Pipeline job.
```
pipeline = aip.PipelineJob(
display_name="example",
template_path="example.json",
pipeline_root=PIPELINE_ROOT,
enable_caching=False
)
```
### Asynchronous execution of pipeline
When running a pipeline with the method `run()`, the pipeline is ran synchronously. To run asynchronously, you use the method `submit()`. Once the job has started, your Python script can continue to execute. Then when you need to block execution using the method `wait()`.
### Setting machine resources for pipeline steps
By default, Vertex AI Pipelines will automatically find the best matching machine type to run the component. You can override and specify the machine resources on a per component basis, when you invoke the component in a pipeline, as follows:
```
@dsl.pipeline(name='my-pipeline')
def pipeline():
task = taskOp().
set_cpu_limit('CPU_LIMIT').
set_memory_limit('MEMORY_LIMIT').
add_node_selector_constraint(SELECTOR_CONSTRAINT).
set_gpu_limit(GPU_LIMIT))
```
Learn more about [Specifying machine types in pipelines](https://cloud.google.com/vertex-ai/docs/pipelines/machine-types)
# Cleaning up
To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
Otherwise, you can delete the individual resources you created in this tutorial:
- Dataset
- Pipeline
- Model
- Endpoint
- AutoML Training Job
- Batch Job
- Custom Job
- Hyperparameter Tuning Job
- Cloud Storage Bucket
```
delete_all = True
if delete_all:
# Delete the dataset using the Vertex dataset object
try:
if "dataset" in globals():
dataset.delete()
except Exception as e:
print(e)
# Delete the model using the Vertex model object
try:
if "model" in globals():
model.delete()
except Exception as e:
print(e)
# Delete the endpoint using the Vertex endpoint object
try:
if "endpoint" in globals():
endpoint.undeploy_all()
endpoint.delete()
except Exception as e:
print(e)
# Delete the AutoML or Pipeline training job
try:
if "dag" in globals():
dag.delete()
except Exception as e:
print(e)
# Delete the custom training job
try:
if "job" in globals():
job.delete()
except Exception as e:
print(e)
# Delete the batch prediction job using the Vertex batch prediction object
try:
if "batch_predict_job" in globals():
batch_predict_job.delete()
except Exception as e:
print(e)
# Delete the hyperparameter tuning job using the Vertex hyperparameter tuning object
try:
if "hpt_job" in globals():
hpt_job.delete()
except Exception as e:
print(e)
if "BUCKET_NAME" in globals():
! gsutil rm -r $BUCKET_NAME
```
| github_jupyter |
## Hyperparameter Tuning with Chainer
[VGG](https://arxiv.org/pdf/1409.1556v6.pdf) is an architecture for deep convolution networks. In this example, we use convolutional networks to perform image classification using the CIFAR-10 dataset. CIFAR-10 consists of 60000 32x32 colour images in 10 classes, with 6000 images per class. There are 50000 training images and 10000 test images.
We'll use SageMaker's hyperparameter tuning to train multiple convolutional networks, experimenting with different hyperparameter combinations. After that, we'll find the model with the best performance, deploy it to Amazon SageMaker hosting, and then classify images using the deployed model.
This notebook uses the Chainer script and estimator setup from [the "Training with Chainer" notebook](files/chainer_single_machine_cifar10.ipynb).
```
# Setup
from sagemaker import get_execution_role
import sagemaker
sagemaker_session = sagemaker.Session()
# This role retrieves the SageMaker-compatible role used by this notebook instance.
role = get_execution_role()
```
## Downloading training and test data
We use helper functions provided by `chainer` to download and preprocess the CIFAR10 data.
```
import chainer
from chainer.datasets import get_cifar10
train, test = get_cifar10()
```
## Uploading the data
We save the preprocessed data to the local filesystem, and then use the `sagemaker.Session.upload_data` function to upload our datasets to an S3 location. The return value `inputs` identifies the S3 location, which we will use when we start the Training Job.
```
import os
import shutil
import numpy as np
train_data = [element[0] for element in train]
train_labels = [element[1] for element in train]
test_data = [element[0] for element in test]
test_labels = [element[1] for element in test]
try:
os.makedirs('/tmp/data/train_cifar')
os.makedirs('/tmp/data/test_cifar')
np.savez('/tmp/data/train_cifar/train.npz', data=train_data, labels=train_labels)
np.savez('/tmp/data/test_cifar/test.npz', data=test_data, labels=test_labels)
train_input = sagemaker_session.upload_data(
path=os.path.join('/tmp', 'data', 'train_cifar'),
key_prefix='notebook/chainer_cifar/train')
test_input = sagemaker_session.upload_data(
path=os.path.join('/tmp', 'data', 'test_cifar'),
key_prefix='notebook/chainer_cifar/test')
finally:
shutil.rmtree('/tmp/data')
print('training data at %s' % train_input)
print('test data at %s' % test_input)
```
## Writing the Chainer script
We use a single script to train and host a Chainer model. The training part is similar to a script you might run outside of SageMaker.
The hosting part requires implementing certain functions. Here, we've defined only `model_fn()`, which loads model artifacts that were created during training. The other functions will take on default values as described [here](https://github.com/aws/sagemaker-python-sdk#model-serving).
For a more in-depth discussion of this script see [the "Training with Chainer" notebook](files/chainer_single_machine_cifar10.ipynb).
For more on writing Chainer scripts to run on SageMaker, or for more on the Chainer container itself, please see the following repositories:
* For writing Chainer scripts to run on SageMaker: https://github.com/aws/sagemaker-python-sdk
* For more on the Chainer container and default hosting functions: https://github.com/aws/sagemaker-chainer-containers
```
!pygmentize 'src/chainer_cifar_vgg_single_machine.py'
```
## Running hyperparameter tuning jobs on SageMaker
To specify options for a training job using Chainer, we construct a `Chainer` estimator using the [sagemaker-python-sdk](https://github.com/aws/sagemaker-python-sdk). We pass in an `entry_point`, the name of a script that contains a couple of functions with certain signatures (`train()` and `model_fn()`), and a `source_dir`, a directory containing all code to run inside the Chainer container. This script will be run on SageMaker in a container that invokes these functions to train and load Chainer models.
For this example, we're specifying the number of epochs to be 1 for the purposes of demonstration. We suggest at least 50 epochs for a more meaningful result.
```
from sagemaker.chainer.estimator import Chainer
chainer_estimator = Chainer(entry_point='chainer_cifar_vgg_single_machine.py',
source_dir="src",
role=role,
sagemaker_session=sagemaker_session,
train_instance_count=1,
train_instance_type='ml.p2.xlarge',
hyperparameters={'epochs': 1, 'batch-size': 64})
```
We then need to pass this estimator to a `HyperparameterTuner`. For the `HyperparameterTuner` class, we define the following options for running hyperparameter tuning jobs:
* `hyperparameter_ranges`: the hyperparameters we'd like to tune and their possible values. We have three different types of hyperparameters that can be tuned: categorical, continuous, and integer.
* `objective_metric_name`: the objective metric we'd like to tune.
* `metric_definitions`: the name of the objective metric as well as the regular expression (regex) used to extract the metric from the CloudWatch logs of each training job.
* `max_jobs`: number of training jobs to run in total.
* `max_parallel_jobs`: number of training jobs to run simultaneously.
For this example, we are going to tune on learning rate. In general, if possible, it's best to specify a value as the least restrictive type, so we define learning rate as a continuous parameter ranging between 0.5 and 0.6 rather than, say, a categorical parameter with possible values of 0.5, 0.55, and 0.6.
```
from sagemaker.tuner import ContinuousParameter
hyperparameter_ranges = {'learning-rate': ContinuousParameter(0.05, 0.06)}
```
Next, we define our objective metric, which we use to evaluate each training job. This consists of a name and a regex. The training script in this example uses Chainer's [`PrintReport`](https://docs.chainer.org/en/stable/reference/generated/chainer.training.extensions.PrintReport.html) to print out metrics for each epoch, which looks something like this when run for 50 epochs (truncated here):
```
epoch main/loss validation/main/loss main/accuracy validation/main/accuracy elapsed_time
#033[J1 2.33857 1.86438 0.175811 0.254479 47.5526
#033[J2 1.78559 1.59937 0.298095 0.376493 79.5099
#033[J3 1.50956 1.38693 0.422015 0.469646 111.372
...
#033[J48 0.378797 0.573417 0.879842 0.821955 1548.58
#033[J49 0.373226 0.573498 0.879516 0.812201 1580.56
#033[J50 0.369154 0.485158 0.882242 0.843451 1612.49
```
The regex we use captures the fourth number in the last row, which is the validation accuracy for the final epoch in the training job. Because we're using only one epoch for demonstration purposes, our regex has 'J1' in it, but the '1' should be replaced with the number of epochs used for each training job.
```
objective_metric_name = 'Validation-accuracy'
metric_definitions = [{'Name': 'Validation-accuracy', 'Regex': '\[J1\s+\d\.\d+\s+\d\.\d+\s+\d\.\d+\s+(\d\.\d+)'}]
```
Finally, we need to define how many training jobs to run. We recommend you set the parallel jobs value to less than 10% of the total number of training jobs, but we are setting it higher here to keep this example short. We are also setting `max_jobs` to a low value to shorten the time needed for the hyperparameter tuning job to complete, but note that running only two jobs won't demonstrate any meaningful hyperparameter tuning results.
```
max_jobs = 2
max_parallel_jobs = 2
from sagemaker.tuner import HyperparameterTuner
chainer_tuner = HyperparameterTuner(estimator=chainer_estimator,
objective_metric_name=objective_metric_name,
hyperparameter_ranges=hyperparameter_ranges,
metric_definitions=metric_definitions,
max_jobs=max_jobs,
max_parallel_jobs=max_parallel_jobs)
```
With our tuner, we can now invoke `fit()` to start a hyperparameter tuning job:
```
chainer_tuner.fit({'train': train_input, 'test': test_input})
```
## Waiting for the hyperparameter tuning job to complete
Now we wait for the hyperparameter tuning job to complete. We have a convenience method, `wait()`, that will block until the hyperparameter tuning job has completed. We can call that here to see if the hyperparameter tuning job is still running; the cell will finish running when the hyperparameter tuning job has completed.
```
chainer_tuner.wait()
```
## Deploying the Trained Model
After training, we use the tuner object to create and deploy a hosted prediction endpoint with the best training job. We can use a CPU-based instance for inference (in this case an `ml.m4.xlarge`), even though we trained on GPU instances.
The predictor object returned by `deploy()` lets us call the new endpoint and perform inference on our sample images using the model from the best training job found during hyperparameter tuning.
```
predictor = chainer_tuner.deploy(initial_instance_count=1, instance_type='ml.m4.xlarge')
```
### CIFAR10 sample images
We'll use these CIFAR10 sample images to test the service:
<img style="display: inline; height: 32px; margin: 0.25em" src="images/airplane1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/automobile1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/bird1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/cat1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/deer1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/dog1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/frog1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/horse1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/ship1.png" />
<img style="display: inline; height: 32px; margin: 0.25em" src="images/truck1.png" />
## Predicting using SageMaker Endpoint
We batch the images together into a single NumPy array to obtain multiple inferences with a single prediction request.
```
from skimage import io
import numpy as np
def read_image(filename):
img = io.imread(filename)
img = np.array(img).transpose(2, 0, 1)
img = np.expand_dims(img, axis=0)
img = img.astype(np.float32)
img *= 1. / 255.
img = img.reshape(3, 32, 32)
return img
def read_images(filenames):
return np.array([read_image(f) for f in filenames])
filenames = ['images/airplane1.png',
'images/automobile1.png',
'images/bird1.png',
'images/cat1.png',
'images/deer1.png',
'images/dog1.png',
'images/frog1.png',
'images/horse1.png',
'images/ship1.png',
'images/truck1.png']
image_data = read_images(filenames)
```
The predictor runs inference on our input data and returns a list of predictions whose argmax gives the predicted label of the input data.
```
response = predictor.predict(image_data)
for i, prediction in enumerate(response):
print('image {}: prediction: {}'.format(i, prediction.argmax(axis=0)))
```
## Cleanup
After you have finished with this example, remember to delete the prediction endpoint to release the instance(s) associated with it.
```
chainer_tuner.delete_endpoint()
```
| github_jupyter |
# Training Deep Neural Networks on a GPU with PyTorch
### Part 4 of "Deep Learning with Pytorch: Zero to GANs"
This tutorial series is a hands-on beginner-friendly introduction to deep learning using [PyTorch](https://pytorch.org), an open-source neural networks library. These tutorials take a practical and coding-focused approach. The best way to learn the material is to execute the code and experiment with it yourself. Check out the full series here:
1. [PyTorch Basics: Tensors & Gradients](https://jovian.ai/aakashns/01-pytorch-basics)
2. [Gradient Descent & Linear Regression](https://jovian.ai/aakashns/02-linear-regression)
3. [Working with Images & Logistic Regression](https://jovian.ai/aakashns/03-logistic-regression)
4. [Training Deep Neural Networks on a GPU](https://jovian.ai/aakashns/04-feedforward-nn)
5. [Image Classification using Convolutional Neural Networks](https://jovian.ai/aakashns/05-cifar10-cnn)
6. [Data Augmentation, Regularization and ResNets](https://jovian.ai/aakashns/05b-cifar10-resnet)
7. [Generating Images using Generative Adversarial Networks](https://jovian.ai/aakashns/06b-anime-dcgan/)
This tutorial covers the following topics:
* Creating a deep neural network with hidden layers
* Using a non-linear activation function
* Using a GPU (when available) to speed up training
* Experimenting with hyperparameters to improve the model
### How to run the code
This tutorial is an executable [Jupyter notebook](https://jupyter.org) hosted on [Jovian](https://www.jovian.ai). You can _run_ this tutorial and experiment with the code examples in a couple of ways: *using free online resources* (recommended) or *on your computer*.
#### Option 1: Running using free online resources (1-click, recommended)
The easiest way to start executing the code is to click the **Run** button at the top of this page and select **Run on Colab**. [Google Colab](https://colab.research.google.com) is a free online platform for running Jupyter notebooks using Google's cloud infrastructure. You can also select "Run on Binder" or "Run on Kaggle" if you face issues running the notebook on Google Colab.
#### Option 2: Running on your computer locally
To run the code on your computer locally, you'll need to set up [Python](https://www.python.org), download the notebook and install the required libraries. We recommend using the [Conda](https://docs.conda.io/projects/conda/en/latest/user-guide/install/) distribution of Python. Click the **Run** button at the top of this page, select the **Run Locally** option, and follow the instructions.
> **Jupyter Notebooks**: This tutorial is a [Jupyter notebook](https://jupyter.org) - a document made of _cells_. Each cell can contain code written in Python or explanations in plain English. You can execute code cells and view the results, e.g., numbers, messages, graphs, tables, files, etc., instantly within the notebook. Jupyter is a powerful platform for experimentation and analysis. Don't be afraid to mess around with the code & break things - you'll learn a lot by encountering and fixing errors. You can use the "Kernel > Restart & Clear Output" or "Edit > Clear Outputs" menu option to clear all outputs and start again from the top.
### Using a GPU for faster training
You can use a [Graphics Processing Unit](https://en.wikipedia.org/wiki/Graphics_processing_unit) (GPU) to train your models faster if your execution platform is connected to a GPU manufactured by NVIDIA. Follow these instructions to use a GPU on the platform of your choice:
* _Google Colab_: Use the menu option "Runtime > Change Runtime Type" and select "GPU" from the "Hardware Accelerator" dropdown.
* _Kaggle_: In the "Settings" section of the sidebar, select "GPU" from the "Accelerator" dropdown. Use the button on the top-right to open the sidebar.
* _Binder_: Notebooks running on Binder cannot use a GPU, as the machines powering Binder aren't connected to any GPUs.
* _Linux_: If your laptop/desktop has an NVIDIA GPU (graphics card), make sure you have installed the [NVIDIA CUDA drivers](https://docs.nvidia.com/cuda/cuda-installation-guide-linux/index.html).
* _Windows_: If your laptop/desktop has an NVIDIA GPU (graphics card), make sure you have installed the [NVIDIA CUDA drivers](https://docs.nvidia.com/cuda/cuda-installation-guide-microsoft-windows/index.html).
* _macOS_: macOS is not compatible with NVIDIA GPUs
If you do not have access to a GPU or aren't sure what it is, don't worry, you can execute all the code in this tutorial just fine without a GPU.
## Preparing the Data
In [the previous tutorial](https://jovian.ai/aakashns/03-logistic-regression), we trained a logistic regression model to identify handwritten digits from the MNIST dataset with an accuracy of around 86%. The dataset consists of 28px by 28px grayscale images of handwritten digits (0 to 9) and labels for each image indicating which digit it represents. Here are some sample images from the dataset:
We noticed that it's quite challenging to improve the accuracy of a logistic regression model beyond 87%, since the model assumes a linear relationship between pixel intensities and image labels. In this post, we'll try to improve upon it using a *feed-forward neural network* which can capture non-linear relationships between inputs and targets.
Let's begin by installing and importing the required modules and classes from `torch`, `torchvision`, `numpy`, and `matplotlib`.
```
# Uncomment and run the appropriate command for your operating system, if required
# Linux / Binder
# !pip install numpy matplotlib torch==1.7.0+cpu torchvision==0.8.1+cpu torchaudio==0.7.0 -f https://download.pytorch.org/whl/torch_stable.html
# Windows
# !pip install numpy matplotlib torch==1.7.0+cpu torchvision==0.8.1+cpu torchaudio==0.7.0 -f https://download.pytorch.org/whl/torch_stable.html
# MacOS
# !pip install numpy matplotlib torch torchvision torchaudio
import torch
import torchvision
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import torch.nn as nn
import torch.nn.functional as F
from torchvision.datasets import MNIST
from torchvision.transforms import ToTensor
from torchvision.utils import make_grid
from torch.utils.data.dataloader import DataLoader
from torch.utils.data import random_split
%matplotlib inline
# Use a white background for matplotlib figures
matplotlib.rcParams['figure.facecolor'] = '#ffffff'
```
We can download the data and create a PyTorch dataset using the `MNIST` class from `torchvision.datasets`.
```
dataset = MNIST(root='data/', download=True, transform=ToTensor())
```
Let's look at a couple of images from the dataset. The images are converted to PyTorch tensors with the shape `1x28x28` (the dimensions represent color channels, width and height). We can use `plt.imshow` to display the images. However, `plt.imshow` expects channels to be last dimension in an image tensor, so we use the `permute` method to reorder the dimensions of the image.
```
image, label = dataset[0]
print('image.shape:', image.shape)
plt.imshow(image.permute(1, 2, 0), cmap='gray')
print('Label:', label)
image, label = dataset[0]
print('image.shape:', image.shape)
plt.imshow(image.permute(1, 2, 0), cmap='gray')
print('Label:', label)
```
Next, let's use the `random_split` helper function to set aside 10000 images for our validation set.
```
val_size = 10000
train_size = len(dataset) - val_size
train_ds, val_ds = random_split(dataset, [train_size, val_size])
len(train_ds), len(val_ds)
```
We can now create PyTorch data loaders for training and validation.
```
batch_size=128
train_loader = DataLoader(train_ds, batch_size, shuffle=True, num_workers=4, pin_memory=True)
val_loader = DataLoader(val_ds, batch_size*2, num_workers=4, pin_memory=True)
```
Can you figure out the purpose of the arguments `num_workers` and `pin_memory`? Try looking into the documentation: https://pytorch.org/docs/stable/data.html .
Let's visualize a batch of data in a grid using the `make_grid` function from `torchvision`. We'll also use the `.permute` method on the tensor to move the channels to the last dimension, as expected by `matplotlib`.
```
for images, _ in train_loader:
print('images.shape:', images.shape)
plt.figure(figsize=(16,8))
plt.axis('off')
plt.imshow(make_grid(images, nrow=16).permute((1, 2, 0)))
break
```
## Hidden Layers, Activation Functions and Non-Linearity
We'll create a neural network with two layers: a _hidden layer_ and an _output layer_. Additionally, we'll use an _activation function_ between the two layers. Let's look at a step-by-step example to learn how hidden layers and activation functions can help capture non-linear relationships between inputs and outputs.
First, let's create a batch of inputs tensors. We'll flatten the `1x28x28` images into vectors of size `784`, so they can be passed into an `nn.Linear` object.
```
for images, labels in train_loader:
print('images.shape:', images.shape)
inputs = images.reshape(-1, 784)
print('inputs.shape:', inputs.shape)
break
```
Next, let's create a `nn.Linear` object, which will serve as our _hidden_ layer. We'll set the size of the output from the hidden layer to 32. This number can be increased or decreased to change the _learning capacity_ of the model.
```
input_size = inputs.shape[-1]
hidden_size = 32
layer1 = nn.Linear(input_size, hidden_size)
```
We can now compute intermediate outputs for the batch of images by passing `inputs` through `layer1`.
```
inputs.shape
layer1_outputs = layer1(inputs)
print('layer1_outputs.shape:', layer1_outputs.shape)
```
The image vectors of size `784` are transformed into intermediate output vectors of length `32` by performing a matrix multiplication of `inputs` matrix with the transposed weights matrix of `layer1` and adding the bias. We can verify this using `torch.allclose`. For a more detailed explanation, review the tutorial on [linear regression](https://jovian.ai/aakashns/02-linear-regression).
```
layer1_outputs_direct = inputs @ layer1.weight.t() + layer1.bias
layer1_outputs_direct.shape
# torch.allclose is use to check the similarities of two variables
torch.allclose(layer1_outputs, layer1_outputs_direct, 1e-3)
```
Thus, `layer1_outputs` and `inputs` have a linear relationship, i.e., each element of `layer_outputs` is a weighted sum of elements from `inputs`. Thus, even as we train the model and modify the weights, `layer1` can only capture linear relationships between `inputs` and `outputs`.
<img src="https://i.imgur.com/inXsLuq.png" width="360">
Next, we'll use the Rectified Linear Unit (ReLU) function as the activation function for the outputs. It has the formula `relu(x) = max(0,x)` i.e. it simply replaces negative values in a given tensor with the value 0. ReLU is a non-linear function, as seen here visually:
<img src="https://i.imgur.com/yijV4xF.png" width="420">
We can use the `F.relu` method to apply ReLU to the elements of a tensor.
```
F.relu(torch.tensor([[1, -1, 0],
[-0.1, .2, 3]]))
layer1_outputs.shape
```
Let's apply the activation function to `layer1_outputs` and verify that negative values were replaced with 0.
```
relu_outputs = F.relu(layer1_outputs)
print('min(layer1_outputs):', torch.min(layer1_outputs).item())
print('min(relu_outputs):', torch.min(relu_outputs).item())
```
Now that we've applied a non-linear activation function, `relu_outputs` and `inputs` do not have a linear relationship. We refer to `ReLU` as the _activation function_, because for each input certain outputs are activated (those with non-zero values) while others turned off (those with zero values)
Next, let's create an output layer to convert vectors of length `hidden_size` in `relu_outputs` into vectors of length 10, which is the desired output of our model (since there are 10 target labels).
```
output_size = 10
layer2 = nn.Linear(hidden_size, output_size)
relu_outputs.shape
layer2_outputs = layer2(relu_outputs)
print(layer2_outputs.shape)
inputs.shape
```
As expected, `layer2_outputs` contains a batch of vectors of size 10. We can now use this output to compute the loss using `F.cross_entropy` and adjust the weights of `layer1` and `layer2` using gradient descent.
```
F.cross_entropy(layer2_outputs, labels)
```
Thus, our model transforms `inputs` into `layer2_outputs` by applying a linear transformation (using `layer1`), followed by a non-linear activation (using `F.relu`), followed by another linear transformation (using `layer2`). Let's verify this by re-computing the output using basic matrix operations.
```
# Expanded version of layer2(F.relu(layer1(inputs)))
outputs = (F.relu(inputs @ layer1.weight.t() + layer1.bias)) @ layer2.weight.t() + layer2.bias
torch.allclose(outputs, layer2_outputs, 1e-3)
```
Note that `outputs` and `inputs` do not have a linear relationship due to the non-linear activation function `F.relu`. As we train the model and adjust the weights of `layer1` and `layer2`, we can now capture non-linear relationships between the images and their labels. In other words, introducing non-linearity makes the model more powerful and versatile. Also, since `hidden_size` does not depend on the dimensions of the inputs or outputs, we vary it to increase the number of parameters within the model. We can also introduce new hidden layers and apply the same non-linear activation after each hidden layer.
The model we just created is called a neural network. A _deep neural network_ is simply a neural network with one or more hidden layers. In fact, the [Universal Approximation Theorem](http://neuralnetworksanddeeplearning.com/chap4.html) states that a sufficiently large & deep neural network can compute any arbitrary function i.e. it can _learn_ rich and complex non-linear relationships between inputs and targets. Here are some examples:
* Identifying if an image contains a cat or a dog (or [something else](https://machinelearningmastery.com/introduction-to-the-imagenet-large-scale-visual-recognition-challenge-ilsvrc/))
* Identifying the genre of a song using a 10-second sample
* Classifying movie reviews as positive or negative based on their content
* Navigating self-driving cars using a video feed of the road
* Translating sentences from English to French (and hundreds of other languages)
* Converting a speech recording to text and vice versa
* And many more...
It's hard to imagine how the simple process of multiplying inputs with randomly initialized matrices, applying non-linear activations, and adjusting weights repeatedly using gradient descent can yield such astounding results. Deep learning models often contain millions of parameters, which can together capture far more complex relationships than the human brain can comprehend.
If we hadn't included a non-linear activation between the two linear layers, the final relationship between inputs and outputs would still be linear. A simple refactoring of the computations illustrates this.
```
# Same as layer2(layer1(inputs))
outputs2 = (inputs @ layer1.weight.t() + layer1.bias) @ layer2.weight.t() + layer2.bias
# Create a single layer to replace the two linear layers
combined_layer = nn.Linear(input_size, output_size)
combined_layer.weight.data = layer2.weight @ layer1.weight
combined_layer.bias.data = layer1.bias @ layer2.weight.t() + layer2.bias
# Same as combined_layer(inputs)
outputs3 = inputs @ combined_layer.weight.t() + combined_layer.bias
torch.allclose(outputs2, outputs3, 1e-3)
```
### Save and upload your notebook
Whether you're running this Jupyter notebook online or on your computer, it's essential to save your work from time to time. You can continue working on a saved notebook later or share it with friends and colleagues to let them execute your code. [Jovian](https://jovian.ai/platform-features) offers an easy way of saving and sharing your Jupyter notebooks online.
```
# Install the library
!pip install jovian --upgrade --quiet
import jovian
jovian.commit(project='04-feedforward-nn')
```
`jovian.commit` uploads the notebook to your Jovian account, captures the Python environment, and creates a shareable link for your notebook, as shown above. You can use this link to share your work and let anyone (including you) run your notebooks and reproduce your work.
## Model
We are now ready to define our model. As discussed above, we'll create a neural network with one hidden layer. Here's what that means:
* Instead of using a single `nn.Linear` object to transform a batch of inputs (pixel intensities) into outputs (class probabilities), we'll use two `nn.Linear` objects. Each of these is called a _layer_ in the network.
* The first layer (also known as the hidden layer) will transform the input matrix of shape `batch_size x 784` into an intermediate output matrix of shape `batch_size x hidden_size`. The parameter `hidden_size` can be configured manually (e.g., 32 or 64).
* We'll then apply a non-linear *activation function* to the intermediate outputs. The activation function transforms individual elements of the matrix.
* The result of the activation function, which is also of size `batch_size x hidden_size`, is passed into the second layer (also known as the output layer). The second layer transforms it into a matrix of size `batch_size x 10`. We can use this output to compute the loss and adjust weights using gradient descent.
As discussed above, our model will contain one hidden layer. Here's what it looks like visually:
<img src="https://i.imgur.com/eN7FrpF.png" width="480">
Let's define the model by extending the `nn.Module` class from PyTorch.
```
class MnistModel(nn.Module):
"""Feedfoward neural network with 1 hidden layer"""
def __init__(self, in_size, hidden_size, out_size):
super().__init__()
# hidden layer
self.linear1 = nn.Linear(in_size, hidden_size)
# output layer
self.linear2 = nn.Linear(hidden_size, out_size)
def forward(self, xb):
# Flatten the image tensors
xb = xb.view(xb.size(0), -1)
# Get intermediate outputs using hidden layer
out = self.linear1(xb)
# Apply activation function
out = F.relu(out)
# Get predictions using output layer
out = self.linear2(out)
return out
def training_step(self, batch):
images, labels = batch
out = self(images) # Generate predictions
loss = F.cross_entropy(out, labels) # Calculate loss
return loss
def validation_step(self, batch):
images, labels = batch
out = self(images) # Generate predictions
loss = F.cross_entropy(out, labels) # Calculate loss
acc = accuracy(out, labels) # Calculate accuracy
return {'val_loss': loss, 'val_acc': acc}
def validation_epoch_end(self, outputs):
batch_losses = [x['val_loss'] for x in outputs]
epoch_loss = torch.stack(batch_losses).mean() # Combine losses
batch_accs = [x['val_acc'] for x in outputs]
epoch_acc = torch.stack(batch_accs).mean() # Combine accuracies
return {'val_loss': epoch_loss.item(), 'val_acc': epoch_acc.item()}
def epoch_end(self, epoch, result):
print("Epoch [{}], val_loss: {:.4f}, val_acc: {:.4f}".format(epoch, result['val_loss'], result['val_acc']))
```
We also need to define an `accuracy` function which calculates the accuracy of the model's prediction on an batch of inputs. It's used in `validation_step` above.
```
def accuracy(outputs, labels):
_, preds = torch.max(outputs, dim=1)
return torch.tensor(torch.sum(preds == labels).item() / len(preds))
```
We'll create a model that contains a hidden layer with 32 activations.
```
input_size = 784
hidden_size = 32 # you can change this
num_classes = 10
model = MnistModel(input_size, hidden_size=32, out_size=num_classes)
```
Let's take a look at the model's parameters. We expect to see one weight and bias matrix for each of the layers.
```
for t in model.parameters():
print(t.shape)
```
Let's try and generate some outputs using our model. We'll take the first batch of 128 images from our dataset and pass them into our model.
```
for images, labels in train_loader:
outputs = model(images)
loss = F.cross_entropy(outputs, labels)
print('Loss:', loss.item())
break
print('outputs.shape : ', outputs.shape)
print('Sample outputs :\n', outputs[:2].data)
```
## Using a GPU
As the sizes of our models and datasets increase, we need to use GPUs to train our models within a reasonable amount of time. GPUs contain hundreds of cores optimized for performing expensive matrix operations on floating-point numbers quickly, making them ideal for training deep neural networks. You can use GPUs for free on [Google Colab](https://colab.research.google.com/) and [Kaggle](https://www.kaggle.com/kernels) or rent GPU-powered machines on services like [Google Cloud Platform](https://cloud.google.com/gpu/), [Amazon Web Services](https://docs.aws.amazon.com/dlami/latest/devguide/gpu.html), and [Paperspace](https://www.paperspace.com/).
We can check if a GPU is available and the required NVIDIA CUDA drivers are installed using `torch.cuda.is_available`.
```
torch.cuda.is_available()
```
Let's define a helper function to ensure that our code uses the GPU if available and defaults to using the CPU if it isn't.
```
def get_default_device():
"""Pick GPU if available, else CPU"""
if torch.cuda.is_available():
return torch.device('cuda')
else:
return torch.device('cpu')
device = get_default_device()
device
```
Next, let's define a function that can move data and model to a chosen device.
```
def to_device(data, device):
"""Move tensor(s) to chosen device"""
if isinstance(data, (list,tuple)):
return [to_device(x, device) for x in data]
return data.to(device, non_blocking=True)
for images, labels in train_loader:
print(images.shape)
images = to_device(images, device)
print(images.device)
break
```
Finally, we define a `DeviceDataLoader` class to wrap our existing data loaders and move batches of data to the selected device. Interestingly, we don't need to extend an existing class to create a PyTorch datal oader. All we need is an `__iter__` method to retrieve batches of data and an `__len__` method to get the number of batches.
```
class DeviceDataLoader():
"""Wrap a dataloader to move data to a device"""
def __init__(self, dl, device):
self.dl = dl
self.device = device
def __iter__(self):
"""Yield a batch of data after moving it to device"""
for b in self.dl:
yield to_device(b, self.device)
def __len__(self):
"""Number of batches"""
return len(self.dl)
```
The `yield` keyword in Python is used to create a generator function that can be used within a `for` loop, as illustrated below.
```
def some_numbers():
yield 10
yield 20
yield 30
for value in some_numbers():
print(value)
```
We can now wrap our data loaders using `DeviceDataLoader`.
```
train_loader = DeviceDataLoader(train_loader, device)
val_loader = DeviceDataLoader(val_loader, device)
```
Tensors moved to the GPU have a `device` property which includes that word `cuda`. Let's verify this by looking at a batch of data from `valid_dl`.
```
for xb, yb in val_loader:
print('xb.device:', xb.device)
print('yb:', yb)
break
```
## Training the Model
We'll define two functions: `fit` and `evaluate` to train the model using gradient descent and evaluate its performance on the validation set. For a detailed walkthrough of these functions, check out the [previous tutorial](https://jovian.ai/aakashns/03-logistic-regression).
```
def evaluate(model, val_loader):
"""Evaluate the model's performance on the validation set"""
outputs = [model.validation_step(batch) for batch in val_loader]
return model.validation_epoch_end(outputs)
def fit(epochs, lr, model, train_loader, val_loader, opt_func=torch.optim.SGD):
"""Train the model using gradient descent"""
history = []
optimizer = opt_func(model.parameters(), lr)
for epoch in range(epochs):
# Training Phase
for batch in train_loader:
loss = model.training_step(batch)
loss.backward()
optimizer.step()
optimizer.zero_grad()
# Validation phase
result = evaluate(model, val_loader)
model.epoch_end(epoch, result)
history.append(result)
return history
```
Before we train the model, we need to ensure that the data and the model's parameters (weights and biases) are on the same device (CPU or GPU). We can reuse the `to_device` function to move the model's parameters to the right device.
```
# Model (on GPU)
model = MnistModel(input_size, hidden_size=hidden_size, out_size=num_classes)
to_device(model, device)
```
Let's see how the model performs on the validation set with the initial set of weights and biases.
```
history = [evaluate(model, val_loader)]
history
```
The initial accuracy is around 10%, as one might expect from a randomly initialized model (since it has a 1 in 10 chance of getting a label right by guessing randomly).
Let's train the model for five epochs and look at the results. We can use a relatively high learning rate of 0.5.
```
history += fit(5, 0.5, model, train_loader, val_loader)
```
96% is pretty good! Let's train the model for five more epochs at a lower learning rate of 0.1 to improve the accuracy further.
```
history += fit(5, 0.1, model, train_loader, val_loader)
```
We can now plot the losses & accuracies to study how the model improves over time.
```
losses = [x['val_loss'] for x in history]
plt.plot(losses, '-x')
plt.xlabel('epoch')
plt.ylabel('loss')
plt.title('Loss vs. No. of epochs');
accuracies = [x['val_acc'] for x in history]
plt.plot(accuracies, '-x')
plt.xlabel('epoch')
plt.ylabel('accuracy')
plt.title('Accuracy vs. No. of epochs');
```
Our current model outperforms the logistic regression model (which could only achieve around 86% accuracy) by a considerable margin! It quickly reaches an accuracy of 97% but doesn't improve much beyond this. To improve accuracy further, we need to make the model more powerful by increasing the hidden layer's size or adding more hidden layers with activations. I encourage you to try out both these approaches and see which one works better.
As a final step, we can save and commit our work using the `jovian` library.
```
!pip install jovian --upgrade -q
import jovian
jovian.commit(project='04-feedforward-nn', environment=None)
```
## Testing with individual images
While we have been tracking the overall accuracy of a model so far, it's also a good idea to look at model's results on some sample images. Let's test out our model with some images from the predefined test dataset of 10000 images. We begin by recreating the test dataset with the `ToTensor` transform.
```
# Define test dataset
test_dataset = MNIST(root='data/',
train=False,
transform=ToTensor())
```
Let's define a helper function `predict_image`, which returns the predicted label for a single image tensor.
```
def predict_image(img, model):
xb = to_device(img.unsqueeze(0), device)
yb = model(xb)
_, preds = torch.max(yb, dim=1)
return preds[0].item()
```
Let's try it out with a few images.
```
img, label = test_dataset[0]
plt.imshow(img[0], cmap='gray')
print('Label:', label, ', Predicted:', predict_image(img, model))
img, label = test_dataset[1839]
plt.imshow(img[0], cmap='gray')
print('Label:', label, ', Predicted:', predict_image(img, model))
img, label = test_dataset[193]
plt.imshow(img[0], cmap='gray')
print('Label:', label, ', Predicted:', predict_image(img, model))
```
Identifying where our model performs poorly can help us improve the model, by collecting more training data, increasing/decreasing the complexity of the model, and changing the hypeparameters.
As a final step, let's also look at the overall loss and accuracy of the model on the test set.
```
test_loader = DeviceDataLoader(DataLoader(test_dataset, batch_size=256), device)
result = evaluate(model, test_loader)
result
```
We expect this to be similar to the accuracy/loss on the validation set. If not, we might need a better validation set that has similar data and distribution as the test set (which often comes from real world data).
Let's save the model's weights and attach it to the notebook using `jovian.commit`. We will also record the model's performance on the test dataset using `jovian.log_metrics`.
```
jovian.log_metrics(test_loss=result['val_loss'], test_acc=result['val_loss'])
torch.save(model.state_dict(), 'mnist-feedforward.pth')
jovian.commit(project='04-feedforward-nn',
environment=None,
outputs=['mnist-feedforward.pth'])
```
## Exercises
Try out the following exercises to apply the concepts and techniques you have learned so far:
* Coding exercises on end-to-end model training: https://jovian.ai/aakashns/03-cifar10-feedforward
* Starter notebook for deep learning models: https://jovian.ai/aakashns/fashion-feedforward-minimal
Training great machine learning models reliably takes practice and experience. Try experimenting with different datasets, models and hyperparameters, it's the best way to acquire this skill.
## Summary and Further Reading
Here is a summary of the topics covered in this tutorial:
* We created a neural network with one hidden layer to improve upon the logistic regression model from the previous tutorial. We also used the ReLU activation function to introduce non-linearity into the model, allowing it to learn more complex relationships between the inputs (pixel densities) and outputs (class probabilities).
* We defined some utilities like `get_default_device`, `to_device` and `DeviceDataLoader` to leverage a GPU if available, by moving the input data and model parameters to the appropriate device.
* We were able to use the exact same training loop: the `fit` function we had define earlier to train out model and evaluate it using the validation dataset.
There's a lot of scope to experiment here, and I encourage you to use the interactive nature of Jupyter to play around with the various parameters. Here are a few ideas:
* Try changing the size of the hidden layer, or add more hidden layers and see if you can achieve a higher accuracy.
* Try changing the batch size and learning rate to see if you can achieve the same accuracy in fewer epochs.
* Compare the training times on a CPU vs. GPU. Do you see a significant difference. How does it vary with the size of the dataset and the size of the model (no. of weights and parameters)?
* Try building a model for a different dataset, such as the [CIFAR10 or CIFAR100 datasets](https://www.cs.toronto.edu/~kriz/cifar.html).
Here are some references for further reading:
* [A visual proof that neural networks can compute any function](http://neuralnetworksanddeeplearning.com/chap4.html), also known as the Universal Approximation Theorem.
* [But what *is* a neural network?](https://www.youtube.com/watch?v=aircAruvnKk) - A visual and intuitive introduction to what neural networks are and what the intermediate layers represent
* [Stanford CS229 Lecture notes on Backpropagation](http://cs229.stanford.edu/notes/cs229-notes-backprop.pdf) - for a more mathematical treatment of how gradients are calculated and weights are updated for neural networks with multiple layers.
You are now ready to move on to the next tutorial: [Image Classification using Convolutional Neural Networks](https://jovian.ai/aakashns/05-cifar10-cnn).
| github_jupyter |
# GenCode Explore
Explore the human RNA sequences from GenCode.
Assume user downloaded files from GenCode 38 [FTP](http://ftp.ebi.ac.uk/pub/databases/gencode/Gencode_human/release_38/)
to a subdirectory called data.
Improve on GenCode_Explore_101.ipynb
Use ORF_counter.
Use MatPlotLib to make box plots and heat maps.
```
import time
def show_time():
t = time.time()
s = time.strftime('%Y-%m-%d %H:%M:%S %Z', time.localtime(t))
print(s)
show_time()
import numpy as np
import pandas as pd
import gzip
import sys
try:
from google.colab import drive
IN_COLAB = True
print("On Google CoLab, mount cloud-local file, get our code from GitHub.")
PATH='/content/drive/'
#drive.mount(PATH,force_remount=True) # hardly ever need this
drive.mount(PATH) # Google will require login credentials
DATAPATH=PATH+'My Drive/data/' # must end in "/"
import requests
s = requests.get('https://raw.githubusercontent.com/ShepherdCode/Soars2021/master/SimTools/RNA_describe.py')
with open('RNA_describe.py', 'w') as f:
f.write(s.text) # writes to cloud local, delete the file later?
from RNA_describe import *
except:
print("CoLab not working. On my PC, use relative paths.")
IN_COLAB = False
DATAPATH='../data/' # must end in "/"
sys.path.append("..") # append parent dir in order to use sibling dirs
from SimTools.RNA_describe import *
MODELPATH="BestModel" # saved on cloud instance and lost after logout
#MODELPATH=DATAPATH+MODELPATH # saved on Google Drive but requires login
if not assert_imported_RNA_describe():
print("ERROR: Cannot use RNA_describe.")
PC_FILENAME='gencode.v38.pc_transcripts.fa.gz'
NC_FILENAME='gencode.v38.lncRNA_transcripts.fa.gz'
def load_gencode(filename,label):
DEFLINE='>'
DELIM='|'
EMPTY=''
labels=[] # usually 1 for protein-coding or 0 for non-coding
seqs=[] # usually string of ACGT
lens=[] # sequence length
ids=[] # GenCode transcript ID, always starts ENST
one_seq = EMPTY
one_id = None
# Use gzip 'r' mode to open file in read-only mode.
# Use gzip 't' mode to read each line of text as type string.
with gzip.open (filename,'rt') as infile:
for line in infile:
if line[0]==DEFLINE:
# Save the previous sequence if one exists.
if not one_seq == EMPTY:
labels.append(label)
seqs.append(one_seq)
lens.append(len(one_seq))
ids.append(one_id)
# Get ready to read the next sequence.
# Parse a GenCode defline that is formatted like this:
# >transcript_ID|gene_ID|other_fields other_info|other_info
one_id = line[1:].split(DELIM)[0]
one_seq = EMPTY
else:
# Continue loading sequence lines till next defline.
additional = line.rstrip()
one_seq = one_seq + additional
# Don't forget to save the last sequence after end-of-file.
if not one_seq == EMPTY:
labels.append(label)
seqs.append(one_seq)
lens.append(len(one_seq))
ids.append(one_id)
df1=pd.DataFrame(ids,columns=['tid'])
df2=pd.DataFrame(labels,columns=['class'])
df3=pd.DataFrame(seqs,columns=['sequence'])
df4=pd.DataFrame(lens,columns=['seqlen'])
df=pd.concat((df1,df2,df3,df4),axis=1)
return df
def get_the_facts(seqs,verbose=False):
oc = ORF_counter()
count = len(seqs)
max_orf_lengths=np.zeros(count)
for s in range(0,count):
seq = seqs[s]
oc.set_sequence(seq)
max_orf = oc.get_max_orf_len()
max_orf_lengths[s] = max_orf
mean_max_orf = np.mean(max_orf_lengths,axis=0)
std_max_orf = np.std(max_orf_lengths,axis=0)
if verbose:
print("mean longest ORF length:",int(mean_max_orf),"+/-",int(std_max_orf))
return mean_max_orf
```
## Load the GenCode data.
Warning: GenCode has
over 100K protein-coding RNA (mRNA)
and almost 50K non-coding RNA (lncRNA).
```
PC_FULLPATH=DATAPATH+PC_FILENAME
NC_FULLPATH=DATAPATH+NC_FILENAME
show_time()
pcdf=load_gencode(PC_FULLPATH,1)
print("PC seqs loaded:",len(pcdf))
show_time()
ncdf=load_gencode(NC_FULLPATH,0)
print("NC seqs loaded:",len(ncdf))
show_time()
print("Sorting PC...")
pcdf.sort_values('seqlen', ascending=True, inplace=True)
print("Sorting NC...")
ncdf.sort_values('seqlen', ascending=True, inplace=True)
# This is a fast way to slice if you have length thresholds.
# TO DO: choose length thresholds and apply to PC and NC RNA.
# For example: 200, 400, 800, 1600, 3200, 6400 (e.g. 200-399, etc.)
#mask = (ncdf['sequence'].str.len() < 1000)
#subset = ncdf.loc[mask]
# Here is one way to extract a list from a dataframe.
#mylist=subset['sequence'].tolist()
#TODO: filter out sequences less than 200 here?
```
###Bin sequences by length
---
```
def subset_list_by_len_bounds(input_list, min_len, max_len):
return list(filter(lambda x: len(x) > min_len and len(x) < max_len, input_list))
import matplotlib.pyplot as plt
import numpy as np
#Bin the RNA sequences
bins = [(200, 400), (400, 800), (800, 1600), (1600, 3200), (3200, 6400), (6400, 12800)]
num_bins = len(bins)
binned_pc_sequences = []
binned_nc_sequences = []
for i in range(0, num_bins):
bin = bins[i]
binned_pc_sequences.append([])
binned_nc_sequences.append([])
binned_pc_sequences[i] = subset_list_by_len_bounds(pcdf['sequence'].tolist(), bin[0], bin[1])
binned_nc_sequences[i] = subset_list_by_len_bounds(ncdf['sequence'].tolist(), bin[0], bin[1])
show_time()
```
##Gather data on ORF lengths and the number of contained and non-contained ORFs
---
```
pc_max_len_data = []
pc_max_cnt_data = []
pc_contain_data = []
nc_max_len_data = []
nc_max_cnt_data = []
nc_contain_data = []
oc = ORF_counter()
for bin_num in range(0, num_bins):
#Gather protein-coding sequence data
pc_max_len_data.append([])
pc_max_cnt_data.append([])
pc_contain_data.append([])
for seq_num in range(0, len(binned_pc_sequences[bin_num])):
oc.set_sequence(binned_pc_sequences[bin_num][seq_num])
pc_max_len_data[bin_num].append(oc.get_max_orf_len())
pc_max_cnt_data[bin_num].append(oc.count_maximal_orfs())
pc_contain_data[bin_num].append(oc.count_contained_orfs())
#Gather non-coding sequence data
nc_max_len_data.append([])
nc_max_cnt_data.append([])
nc_contain_data.append([])
for seq_num in range(0, len(binned_nc_sequences[bin_num])):
oc.set_sequence(binned_nc_sequences[bin_num][seq_num])
nc_max_len_data[bin_num].append(oc.get_max_orf_len())
nc_max_cnt_data[bin_num].append(oc.count_maximal_orfs())
nc_contain_data[bin_num].append(oc.count_contained_orfs())
show_time()
```
##Prepare data for heatmap
---
```
def mean(data):
return sum(data) / len(data)
#Get the means of all of the data
mean_pc_max_len_data = []
mean_pc_max_cnt_data = []
mean_pc_contain_data = []
mean_nc_max_len_data = []
mean_nc_max_cnt_data = []
mean_nc_contain_data = []
for i in range(0, num_bins):
mean_pc_max_len_data.append(mean(pc_max_len_data[i]))
mean_pc_max_cnt_data.append(mean(pc_max_cnt_data[i]))
mean_pc_contain_data.append(mean(pc_contain_data[i]))
mean_nc_max_len_data.append(mean(nc_max_len_data[i]))
mean_nc_max_cnt_data.append(mean(nc_max_cnt_data[i]))
mean_nc_contain_data.append(mean(nc_contain_data[i]))
#Combine them into a (6,6) shaped data set for the heatmap
hm_data = []
hm_data.append(mean_pc_max_len_data)
hm_data.append(mean_pc_max_cnt_data)
hm_data.append(mean_pc_contain_data)
hm_data.append(mean_nc_max_len_data)
hm_data.append(mean_nc_max_cnt_data)
hm_data.append(mean_nc_contain_data)
show_time()
```
###Prepare data for plot of bin sizes
---
```
pc_bin_sizes = []
nc_bin_sizes = []
for i in range(0, num_bins):
pc_bin_sizes.append(len(binned_pc_sequences[i]))
nc_bin_sizes.append(len(binned_nc_sequences[i]))
show_time()
```
###Prepare data for plot of number of sequences with no ORFs
---
```
"""Counts the number of RNA sequences that fit the given constraints.
Sequence range constraints are exclusive.
Max ORF length constraints are inclusive,
"""
def count_constraint_valid_sequences(data, min_seq_len, max_seq_len, min_max_orf_len, max_max_orf_len):
count = 0
oc = ORF_counter()
sequences = data['sequence'].tolist()
for seq in sequences:
if len(seq) > min_seq_len and len(seq) < max_seq_len:
oc.set_sequence(seq)
max_orf_len = oc.get_max_orf_len()
if max_orf_len >= min_max_orf_len and max_orf_len <= max_max_orf_len:
count += 1
return count
pc_no_orf_count = []
nc_no_orf_count = []
for bin in bins:
pc_no_orf_count.append(count_constraint_valid_sequences(pcdf, bin[0], bin[1], 0, 0))
nc_no_orf_count.append(count_constraint_valid_sequences(ncdf, bin[0], bin[1], 0, 0))
show_time()
```
## Plot the data
---
```
def bar_plot_compare(data_a, data_b, title, x_label, y_label, x_axis_labels, a_name, b_name):
x = np.arange(0, len(data_a), 1)
plt.figure()
bar_a = plt.bar(x - 0.25, data_a, color='r', width=0.5)
bar_b = plt.bar(x + 0.25, data_b, color='b', width=0.5)
plt.title(title)
plt.xlabel(x_label)
plt.ylabel(y_label)
plt.xticks(x, labels=x_axis_labels, rotation=45, ha='right')
plt.legend(labels=[a_name, b_name])
plt.show()
def box_plot_compare(data_a, data_b, title, x_label, y_label, x_axis_labels, fliers, y_scale, y_base):
a_pos = []
b_pos = []
x_ticks = []
for i in range(0 , len(data_a) + len(data_b)):
x_ticks.append(i)
for i in range(0, len(data_a)):
a_pos.append(i*2)
for i in range(0, len(data_b)):
b_pos.append(i*2+1)
plt.figure()
box_a = plt.boxplot(data_a, patch_artist=True, positions=a_pos, showfliers=fliers)
for box in box_a['boxes']:
box.set(color='red', linewidth=1)
box.set(facecolor='white')
for flier in box_a['fliers']:
flier.set(markersize=1)
box_b = plt.boxplot(data_b, patch_artist=True, positions=b_pos, showfliers=fliers)
for box in box_b['boxes']:
box.set(color='blue', linewidth=1)
box.set(facecolor='white')
for flier in box_b['fliers']:
flier.set(markersize=0.5, alpha=0.5)
plt.yscale(y_scale, basey=y_base)
plt.xticks(x_ticks, labels=x_axis_labels, rotation=45, ha='right')
plt.xlabel(x_label)
plt.ylabel(y_label)
plt.title(title)
plt.show()
```
###Options for plots
```
#Box plots
show_outliers=True
y_scale = 'log'
y_base = 2
#Heatmap
```
###Generate the plots
```
#Generate x-axis labels
bp_x_axis_labels = []
hm_x_axis_labels = []
for bin in bins:
bp_x_axis_labels.append(str(bin[0]) + "-" + str(bin[1]) + " (mRNA)")
bp_x_axis_labels.append(str(bin[0]) + "-" + str(bin[1]) + " (lncRNA)")
hm_x_axis_labels.append(str(bin[0]) + "-" + str(bin[1]))
bar_x_axis_labels = hm_x_axis_labels
#Bar plots
bar_plot_compare(pc_bin_sizes, nc_bin_sizes, "Number of Sequences per Sequence Length Range", "Sequence Length Ranges", "Number of Sequences", bar_x_axis_labels, "mRNA", "lncRNA")
bar_plot_compare(pc_no_orf_count, nc_no_orf_count, "Number of Sequences without ORFs", "Sequence Length Ranges", "Number of Sequences", bar_x_axis_labels, "mRNA", "lncRNA")
#Heatmap
plt.imshow(hm_data, cmap='hot', interpolation='nearest')
plt.xlabel('Sequence Length Ranges')
plt.xticks(np.arange(6), hm_x_axis_labels, rotation=45, ha='right')
plt.yticks(np.arange(6), ["mRNA Mean Longest ORF Length", "mRNA Mean Number of Non-contained ORFs", "mRNA Mean Number of Contained ORFs", "lncRNA Mean Longest ORF Length", "lncRNA Mean Number of Non-contained ORFs", "lncRNA Mean Number of Contained ORFs"])
plt.show()
#Box plots
box_plot_compare(pc_max_len_data, nc_max_len_data, "Length of Longest ORF in RNA Sequences", "Sequence Length Ranges", "ORF Length", bp_x_axis_labels, show_outliers, y_scale, y_base)
box_plot_compare(pc_max_cnt_data, nc_max_cnt_data, "Number of Non-contained ORFs in RNA Sequences", "Sequence Length Ranges", "Number of Non-contained ORFs", bp_x_axis_labels, show_outliers, y_scale, y_base)
box_plot_compare(pc_contain_data, nc_contain_data, "Number of Contained ORFs in RNA Sequences", "Sequence Length Ranges", "Number of Contained ORFs", bp_x_axis_labels, show_outliers, y_scale, y_base)
```
## Plotting examples
[boxplot doc](https://matplotlib.org/stable/api/_as_gen/matplotlib.pyplot.boxplot.html)
[boxplot demo](https://matplotlib.org/stable/gallery/pyplots/boxplot_demo_pyplot.html)
[heatmap examples](https://stackoverflow.com/questions/33282368/plotting-a-2d-heatmap-with-matplotlib) - scroll down!
```
```
| github_jupyter |
```
%reload_ext autoreload
%autoreload 2
%matplotlib inline
from fastai.model import fit
from fastai.dataset import *
import torchtext
from torchtext import vocab, data
from torchtext.datasets import language_modeling
from fastai.rnn_reg import *
from fastai.rnn_train import *
from fastai.nlp import *
from fastai.lm_rnn import *
import dill as pickle
import random
bs,bptt = 64,70
```
## Language modeling
### Data
```
import os, requests, time
# feedparser isn't a fastai dependency so you may need to install it.
import feedparser
import pandas as pd
class GetArXiv(object):
def __init__(self, pickle_path, categories=list()):
"""
:param pickle_path (str): path to pickle data file to save/load
:param pickle_name (str): file name to save pickle to path
:param categories (list): arXiv categories to query
"""
if os.path.isdir(pickle_path):
pickle_path = f"{pickle_path}{'' if pickle_path[-1] == '/' else '/'}all_arxiv.pkl"
if len(categories) < 1:
categories = ['cs*', 'cond-mat.dis-nn', 'q-bio.NC', 'stat.CO', 'stat.ML']
# categories += ['cs.CV', 'cs.AI', 'cs.LG', 'cs.CL']
self.categories = categories
self.pickle_path = pickle_path
self.base_url = 'http://export.arxiv.org/api/query'
@staticmethod
def build_qs(categories):
"""Build query string from categories"""
return '+OR+'.join(['cat:'+c for c in categories])
@staticmethod
def get_entry_dict(entry):
"""Return a dictionary with the items we want from a feedparser entry"""
try:
return dict(title=entry['title'], authors=[a['name'] for a in entry['authors']],
published=pd.Timestamp(entry['published']), summary=entry['summary'],
link=entry['link'], category=entry['category'])
except KeyError:
print('Missing keys in row: {}'.format(entry))
return None
@staticmethod
def strip_version(link):
"""Strip version number from arXiv paper link"""
return link[:-2]
def fetch_updated_data(self, max_retry=5, pg_offset=0, pg_size=1000, wait_time=15):
"""
Get new papers from arXiv server
:param max_retry: max number of time to retry request
:param pg_offset: number of pages to offset
:param pg_size: num abstracts to fetch per request
:param wait_time: num seconds to wait between requests
"""
i, retry = pg_offset, 0
df = pd.DataFrame()
past_links = []
if os.path.isfile(self.pickle_path):
df = pd.read_pickle(self.pickle_path)
df.reset_index()
if len(df) > 0: past_links = df.link.apply(self.strip_version)
while True:
params = dict(search_query=self.build_qs(self.categories),
sortBy='submittedDate', start=pg_size*i, max_results=pg_size)
response = requests.get(self.base_url, params='&'.join([f'{k}={v}' for k, v in params.items()]))
entries = feedparser.parse(response.text).entries
if len(entries) < 1:
if retry < max_retry:
retry += 1
time.sleep(wait_time)
continue
break
results_df = pd.DataFrame([self.get_entry_dict(e) for e in entries])
max_date = results_df.published.max().date()
new_links = ~results_df.link.apply(self.strip_version).isin(past_links)
print(f'{i}. Fetched {len(results_df)} abstracts published {max_date} and earlier')
if not new_links.any():
break
df = pd.concat((df, results_df.loc[new_links]), ignore_index=True)
i += 1
retry = 0
time.sleep(wait_time)
print(f'Downloaded {len(df)-len(past_links)} new abstracts')
df.sort_values('published', ascending=False).groupby('link').first().reset_index()
df.to_pickle(self.pickle_path)
return df
@classmethod
def load(cls, pickle_path):
"""Load data from pickle and remove duplicates"""
return pd.read_pickle(cls(pickle_path).pickle_path)
@classmethod
def update(cls, pickle_path, categories=list(), **kwargs):
"""
Update arXiv data pickle with the latest abstracts
"""
cls(pickle_path, categories).fetch_updated_data(**kwargs)
return True
PATH='data/arxiv/'
ALL_ARXIV = f'{PATH}all_arxiv.pkl'
# all_arxiv.pkl: if arxiv hasn't been downloaded yet, it'll take some time to get it - go get some coffee
if not os.path.exists(ALL_ARXIV): GetArXiv.update(ALL_ARXIV)
# arxiv.csv: see dl1/nlp-arxiv.ipynb to get this one
df_mb = pd.read_csv(f'{PATH}arxiv.csv')
df_all = pd.read_pickle(ALL_ARXIV)
def get_txt(df):
return '<CAT> ' + df.category.str.replace(r'[\.\-]','') + ' <SUMM> ' + df.summary + ' <TITLE> ' + df.title
df_mb['txt'] = get_txt(df_mb)
df_all['txt'] = get_txt(df_all)
n=len(df_all); n
os.makedirs(f'{PATH}trn/yes', exist_ok=True)
os.makedirs(f'{PATH}val/yes', exist_ok=True)
os.makedirs(f'{PATH}trn/no', exist_ok=True)
os.makedirs(f'{PATH}val/no', exist_ok=True)
os.makedirs(f'{PATH}all/trn', exist_ok=True)
os.makedirs(f'{PATH}all/val', exist_ok=True)
os.makedirs(f'{PATH}models', exist_ok=True)
for (i,(_,r)) in enumerate(df_all.iterrows()):
dset = 'trn' if random.random()>0.1 else 'val'
open(f'{PATH}all/{dset}/{i}.txt', 'w').write(r['txt'])
for (i,(_,r)) in enumerate(df_mb.iterrows()):
lbl = 'yes' if r.tweeted else 'no'
dset = 'trn' if random.random()>0.1 else 'val'
open(f'{PATH}{dset}/{lbl}/{i}.txt', 'w').write(r['txt'])
from spacy.symbols import ORTH
# install the 'en' model if the next line of code fails by running:
#python -m spacy download en # default English model (~50MB)
#python -m spacy download en_core_web_md # larger English model (~1GB)
my_tok = spacy.load('en')
my_tok.tokenizer.add_special_case('<SUMM>', [{ORTH: '<SUMM>'}])
my_tok.tokenizer.add_special_case('<CAT>', [{ORTH: '<CAT>'}])
my_tok.tokenizer.add_special_case('<TITLE>', [{ORTH: '<TITLE>'}])
my_tok.tokenizer.add_special_case('<BR />', [{ORTH: '<BR />'}])
my_tok.tokenizer.add_special_case('<BR>', [{ORTH: '<BR>'}])
def my_spacy_tok(x): return [tok.text for tok in my_tok.tokenizer(x)]
TEXT = data.Field(lower=True, tokenize=my_spacy_tok)
FILES = dict(train='trn', validation='val', test='val')
md = LanguageModelData.from_text_files(f'{PATH}all/', TEXT, **FILES, bs=bs, bptt=bptt, min_freq=10)
pickle.dump(TEXT, open(f'{PATH}models/TEXT.pkl','wb'))
len(md.trn_dl), md.nt, len(md.trn_ds), len(md.trn_ds[0].text)
TEXT.vocab.itos[:12]
' '.join(md.trn_ds[0].text[:150])
```
### Train
```
em_sz = 200
nh = 500
nl = 3
opt_fn = partial(optim.Adam, betas=(0.7, 0.99))
learner = md.get_model(opt_fn, em_sz, nh, nl,
dropout=0.05, dropouth=0.1, dropouti=0.05, dropoute=0.02, wdrop=0.2)
# dropout=0.4, dropouth=0.3, dropouti=0.65, dropoute=0.1, wdrop=0.5
# dropouti=0.05, dropout=0.05, wdrop=0.1, dropoute=0.02, dropouth=0.05)
learner.reg_fn = partial(seq2seq_reg, alpha=2, beta=1)
learner.clip=0.3
learner.fit(3e-3, 1, wds=1e-6)
learner.fit(3e-3, 3, wds=1e-6, cycle_len=1, cycle_mult=2)
learner.save_encoder('adam2_enc')
learner.fit(3e-3, 10, wds=1e-6, cycle_len=5, cycle_save_name='adam3_10')
learner.save_encoder('adam3_10_enc')
learner.fit(3e-3, 8, wds=1e-6, cycle_len=10, cycle_save_name='adam3_5')
learner.fit(3e-3, 1, wds=1e-6, cycle_len=20, cycle_save_name='adam3_20')
learner.save_encoder('adam3_20_enc')
learner.save('adam3_20')
```
### Test
```
def proc_str(s): return TEXT.preprocess(TEXT.tokenize(s))
def num_str(s): return TEXT.numericalize([proc_str(s)])
m=learner.model
s="""<CAT> cscv <SUMM> algorithms that"""
def sample_model(m, s, l=50):
t = num_str(s)
m[0].bs=1
m.eval()
m.reset()
res,*_ = m(t)
print('...', end='')
for i in range(l):
n=res[-1].topk(2)[1]
n = n[1] if n.data[0]==0 else n[0]
word = TEXT.vocab.itos[n.data[0]]
print(word, end=' ')
if word=='<eos>': break
res,*_ = m(n[0].unsqueeze(0))
m[0].bs=bs
sample_model(m,"<CAT> csni <SUMM> algorithms that")
sample_model(m,"<CAT> cscv <SUMM> algorithms that")
sample_model(m,"<CAT> cscv <SUMM> algorithms. <TITLE> on ")
sample_model(m,"<CAT> csni <SUMM> algorithms. <TITLE> on ")
sample_model(m,"<CAT> cscv <SUMM> algorithms. <TITLE> towards ")
sample_model(m,"<CAT> csni <SUMM> algorithms. <TITLE> towards ")
```
### Sentiment
```
TEXT = pickle.load(open(f'{PATH}models/TEXT.pkl','rb'))
class ArxivDataset(torchtext.data.Dataset):
def __init__(self, path, text_field, label_field, **kwargs):
fields = [('text', text_field), ('label', label_field)]
examples = []
for label in ['yes', 'no']:
fnames = glob(os.path.join(path, label, '*.txt'));
assert fnames, f"can't find 'yes.txt' or 'no.txt' under {path}/{label}"
for fname in fnames:
with open(fname, 'r') as f: text = f.readline()
examples.append(data.Example.fromlist([text, label], fields))
super().__init__(examples, fields, **kwargs)
@staticmethod
def sort_key(ex): return len(ex.text)
@classmethod
def splits(cls, text_field, label_field, root='.data',
train='train', test='test', **kwargs):
return super().splits(
root, text_field=text_field, label_field=label_field,
train=train, validation=None, test=test, **kwargs)
ARX_LABEL = data.Field(sequential=False)
splits = ArxivDataset.splits(TEXT, ARX_LABEL, PATH, train='trn', test='val')
md2 = TextData.from_splits(PATH, splits, bs)
# dropout=0.3, dropouti=0.4, wdrop=0.3, dropoute=0.05, dropouth=0.2)
from sklearn.metrics import precision_recall_curve
import matplotlib.pyplot as plt
def prec_at_6(preds,targs):
precision, recall, _ = precision_recall_curve(targs==2, preds[:,2])
print(recall[precision>=0.6][0])
return recall[precision>=0.6][0]
# dropout=0.4, dropouth=0.3, dropouti=0.65, dropoute=0.1, wdrop=0.5
m3 = md2.get_model(opt_fn, 1500, bptt, emb_sz=em_sz, n_hid=nh, n_layers=nl,
dropout=0.1, dropouti=0.65, wdrop=0.5, dropoute=0.1, dropouth=0.3)
m3.reg_fn = partial(seq2seq_reg, alpha=2, beta=1)
m3.clip=25.
# this notebook has a mess of some things going under 'all/' others not, so a little hack here
!ln -sf ../all/models/adam3_20_enc.h5 {PATH}models/adam3_20_enc.h5
m3.load_encoder(f'adam3_20_enc')
lrs=np.array([1e-4,1e-3,1e-3,1e-2,3e-2])
m3.freeze_to(-1)
m3.fit(lrs/2, 1, metrics=[accuracy])
m3.unfreeze()
m3.fit(lrs, 1, metrics=[accuracy], cycle_len=1)
m3.fit(lrs, 2, metrics=[accuracy], cycle_len=4, cycle_save_name='imdb2')
prec_at_6(*m3.predict_with_targs())
m3.fit(lrs, 4, metrics=[accuracy], cycle_len=2, cycle_save_name='imdb2')
prec_at_6(*m3.predict_with_targs())
```
| github_jupyter |
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
### DEFINE USEFUL FUNCTIONS
def fk(x, Ap):
"A useful function"
fk = 0.2*x**5 * (1 + 5/7*(2+4*Ap)*x**2 + 5/9 * (3 + 10*Ap + 4*Ap**2)*x**4)
return fk
def fgam(x, rc, Lp, Ap):
"Annother useful function"
D = (rc/Lp)**2 * (1-0.3*(rc/Lp)**2)
fgam = x**3 * (-D/3 + ((1+D)/5)*x**2 + ((Ap*D-1.3)/7)*x**4)
return fgam
def fchi(x, ri, Lp):
"Yet annother useful function"
fchi = x**3 * (-1/3*(ri/Lp)**2 + 0.2*(1+(ri/Lp)**2)*x**2 - 13/70*x**4)
return fchi
def fc(x, delta, Ap):
"The last useful function, for now"
fc = x**3 * (1 - 0.6*(delta+1)*x**2 - 3/14*(delta+1)*(2*Ap-delta)*x**4)
return fc
def runEarthModel(Qbmo0, QbmoNow, H0M, h, TC, core_Kppm, Ppc):
### DEFINE CONSTANTS
## Fundamental
mu0 = 4e-7 * np.pi # Vacuum pearmeability (SI)
G = 6.67e-11 # Gravitational constant (SI)
y2s = 3.154e7 # Seconds per Earth year
R = 8.3145 # Universal gas constant (J/K/mol)
## Planet
rp = 6371e3 # Radius of the planet (m)
## Core
ri = 0 # Radius of the inner core (m)
rc = 3480e3 # Radius of the core (m)
kc = 40 # Thermal conductivity (W/m/K)
K0 = 1403e9 # Effective modulus (Pa)
K1 = 3.567 # Effective derivative of effective modulus
rho0 = 12451 # Central density (kg/m^3)
Lp = np.sqrt(3*K0/(2*np.pi*G*rho0**2)) # Length scale (m)
Ap = 0.1*(5*K1-13) # Constant in density profile
Pc = 125e9 # Pressure at the core/mantle boundary (Pa)
P0 = Pc + K0*((rc/Lp)**2 + 0.8*(rc/Lp)**4) # Central pressure (Pa)
Cc = 750 # Specific heat of the core (J/kg/K)
bet = 0.83 # Coefficient of compositional expansion for inner core
alp = 0.8 # Coefficient of compositional expansion for magnesium precipitation
dTLdc = -1e4 # Compositional dependence of liquidus temperature (K)
dTLdP = 9e-9 # Pressure dependence of liquidus temperature (K/Pa)
c0 = 0.056 # Initial mass fraction of light elements
gamm = 1.5 # Gruneisen parameter
TL0 = 5806 # Liquidus temperature at the center (K)
DSc = 127 # Entropy of melting (J/K/kg)
TcIC = TL0*(1 - (rc/Lp)**2 - Ap*(rc/Lp)**4)**gamm; # T @ CMB when IC nucleates (K)
Mc = 4/3*np.pi*rho0*Lp**3*fc(rc/Lp, 0, Ap); # Mass of the core (kg)
g = 4/3*np.pi*G*rho0*rc*(1-0.6*(rc/Lp)**2 - 3/7*Ap*(rc/Lp)**4) # Gravitational acceleration near the CMB (m/s^2)
HlamC = 1.76e-17 # Average decay constant (1/s)
h0C = 4.1834e-14 # Heating per unit mass per ppm of K in the core (W/kg/ppm)
## Basal Magma Ocean (BMO)
DSm = 300 # Entropy of melting (J/K/kg) =652 in Stixrude et al. but =300 in Labrosse et al.
rhoM = 5500 # Density of the basal mantle (kg/m^3)
Cm = 1000 # Specific heat of the basal mantle (J/kg/K)
Dphi = 0.088 # Mass fraction change of FeO-rich component upon freezing
TA = 5500 # Melt temperature of the MgO-rich component (K)
TB = 3500 # Melt temperature of the FeO-rich component (K)
alphT = 1.25e-5 # Coefficient of thermal expansion in the BMO (1/K)
OmegaE = 2*np.pi/(24*3600) # Rotation rate of Earth
HT = Cm/(alphT*g) # Thermal scale height for the BMO (m)
sigBMO = 2e4 # Electrical conductivity of the BMO (S/M)
c = 0.63 # Constant prefactor in the scaling law for B-field strength
fohm = 0.9 # Fraction of available power converted into magnetic field energy
HlamM = 1.38e-17 # Average decay constant (1/s)
rb = rc + h # Initial radius of BMO (m)
### RUN THE MODEL
## Define timesteps
NN = 9000 # Number of timesteps
tend = 4.5e9 * y2s # Duration of model (s)
dt = tend/(NN-1) # Duration of timestep (s)
t_all = np.linspace(0,tend,NN)
# Create empty arrays to store parameters
rb_all = np.zeros(NN) # Radius of the BMO upper boundary (m)
ri_all = np.zeros(NN) # Radius of the inner core (m)
TM_all = np.zeros(NN) # Temperature of the solid mantle (K)
TC_all = np.zeros(NN) # Temperature of the BMO and core (K)
TLi_all = np.zeros(NN) # Temperature at the inner core boundary (K)
Qsm_all = np.zeros(NN) # Secular cooling of the BMO (W)
Qlm_all = np.zeros(NN) # Latent heat in the BMO (W)
Qsc_all = np.zeros(NN) # Secular cooling of the core (W)
Qpc_all = np.zeros(NN) # Precipitation of light elements from the core (W)
Qgc_all = np.zeros(NN) # Gravitational energy from inner core growth (W)
Qlc_all = np.zeros(NN) # Latent heat from inner core growth (W)
Qic_all = np.zeros(NN) # Conductive cooling of the inner core (W)
# Initialize radiogenic heating and the heat flow into the solid mantle
Qbmo_all = np.linspace(Qbmo0,QbmoNow,NN) # Heat flow into the base of the solid mantle (W)
Qrm_all = H0M*np.exp(-HlamM*t_all) # Radiogenic heat in the BMO (W)
Qrc_all = Mc*h0C*core_Kppm*np.exp(-HlamC*t_all) # Radiogenic heating in the core (W)
## Begin the time loop
for ii, t in enumerate(t_all):
# Calculate heat flow out of the BMO
Qbmo = Qbmo_all[ii]
# Calculate radiogenic heating in the BMO and core
Qrm = Qrm_all[ii]
Qrc = Qrc_all[ii]
# Calculate proportionalities for BMO
Mm = (4/3)*np.pi*(rb**3-rc**3)*rhoM # Mass of the BMO (kg)
BigTerm = (rb**3-rc**3)/(3*rb**2*Dphi*(TA-TB)) # Gather some constants...
Qsm_til = -Mm*Cm
Qlm_til = -4*np.pi*rb**2*DSm*rhoM*TC*BigTerm
# Calculate proportionalities for core
Qpc_til = 8/3*(np.pi**2*G*rho0**2*Lp**5*alp*Ppc *
(fgam(rc/Lp,rc,Lp,Ap) - fgam(ri/Lp,rc,Lp,Ap)))
if TC > TcIC: # No inner core!
Qsc_til = -4/3*(np.pi*rho0*Cc*Lp**3 * fc(rc/Lp, gamm, Ap) *
(1-(rc/Lp)**2 - Ap*(rc/Lp)**4)**(-gamm))
Qgc_til = 0
Qlc_til = 0
Qic_til = 0
dridt = 0
TLi = TL0
else: # Yes inner core!
if ri < 2e4:
ri = 2e4 # Avoid dividing by zero (even once...)
Mic = Mc - 4/3*np.pi*rho0*Lp**3*(fc(rc/Lp,0,Ap) - fc(ri/Lp,0,Ap))
TLi = TL0 - K0*dTLdP*(ri/Lp)**2 + dTLdc*c0*ri**3/(Lp**3*fc(rc/Lp,0,Ap))
dTLdri = -2*K0*dTLdP*ri/Lp**2 + 3*dTLdc*c0*ri**2 / (Lp**3*fc(rc/Lp,0,Ap))
rhoi = rho0 * (1 -(ri/Lp)**2 - Ap*(ri/Lp)**4)
gi = 4/3*np.pi*G*rho0*ri*(1-0.6*(ri/Lp)**2 - 3/7*Ap*(ri/Lp)**4)
dTadP = gamm*TLi/K0
dridTC = -(1/(dTLdP-dTadP)) * TLi/(TC*rhoi*gi)
Psc = (-4/3*np.pi*rho0*Cc*Lp**3 *
(1-(ri/Lp)**2-Ap*(ri/Lp)**4)**(-gamm) *
(dTLdri+2*gamm*TLi*ri/Lp**2 *
(1+2*Ap*(ri/Lp)**2)/(1-(ri/Lp)**2-Ap*(ri/Lp)**4)) *
(fc(rc/Lp, gamm, Ap) - fc(ri/Lp, gamm, Ap)))
Pgc = (8*np.pi**2*c0*G*rho0**2*bet*ri**2*Lp**2 / fc(rc/Lp,0,Ap) *
(fchi(rc/Lp,ri,Lp) - fchi(ri/Lp,ri,Lp)))
Plc = 4*np.pi*ri**2*rhoi*TLi*DSc
Pic = Cc*Mic*dTLdP*K0*(2*ri/Lp**2 + 3.2*ri/Lp**5)
Qsc_til = Psc * dridTC
Qgc_til = Pgc * dridTC
Qlc_til = Plc * dridTC
Qic_til = Pic * dridTC
# Calculate cooling rate
dTCdt = (Qbmo - Qrm - Qrc)/(Qsm_til + Qlm_til + Qsc_til + Qpc_til + Qgc_til + Qlc_til + Qic_til)
drbdt = BigTerm*dTCdt
if TC < TcIC:
dridt = dridTC * dTCdt
else:
dridt = 0
# Calculate all energetic terms
Qsm = Qsm_til * dTCdt
Qlm = Qlm_til * dTCdt
Qsc = Qsc_til * dTCdt
Qpc = Qpc_til * dTCdt
Qgc = Qgc_til * dTCdt
Qlc = Qlc_til * dTCdt
Qic = Qic_til * dTCdt
# Store output
rb_all[ii] = rb
ri_all[ii] = ri
TC_all[ii] = TC
TLi_all[ii] = TLi
Qbmo_all[ii] = Qbmo
Qsm_all[ii] = Qsm
Qlm_all[ii] = Qlm
Qsc_all[ii] = Qsc
Qpc_all[ii] = Qpc
Qgc_all[ii] = Qgc
Qlc_all[ii] = Qlc
Qic_all[ii] = Qic
# Advance parameters one step
TC = TC + dTCdt*dt
rb = rb + drbdt*dt
ri = ri + dridt*dt
### POST-PROCESSING
## Dynamo in BMO?
# Flow velocities
h_all = rb_all - rc
qsm = Qbmo_all/(4*np.pi*rb_all**2)
v_mix = (h_all*qsm/(rhoM*HT))**(1/3) # Mixing length theory
v_CIA = (qsm/(rhoM*HT))**(2/5) * (h_all/OmegaE)**(1/5) # CIA balance
v_MAC = (qsm/(rhoM*OmegaE*HT))**(1/2) # MAC balance
# Magnetic Reynolds numbers
Rm_mix = mu0*sigBMO*h_all*v_mix
Rm_CIA = mu0*sigBMO*h_all*v_CIA
Rm_MAC = mu0*sigBMO*h_all*v_MAC
# Magnetic field strength at the BMO
Bm_mix = np.sqrt(2*mu0*fohm*c*rhoM*v_mix**2)
Bm_CIA = np.sqrt(2*mu0*fohm*c*rhoM*v_CIA**2)
Bm_MAC = np.sqrt(2*mu0*fohm*c*rhoM*v_MAC**2)
# Magnetic field strength at the surface
Bs_mix = 1/7*Bm_mix*(rb_all/rp)**3
Bs_CIA = 1/7*Bm_CIA*(rb_all/rp)**3
Bs_MAC = 1/7*Bm_MAC*(rb_all/rp)**3
## Dynamo in core?
EK_all = 16*np.pi*gamm**2*kc*Lp*(fk(rc/Lp,Ap)-fk(ri_all/Lp,Ap))
Tbot_all = np.zeros(NN)
for ii, ri in enumerate(ri_all):
if ri > 0:
Tbot_all[ii] = TLi_all[ii]
else:
Tbot_all[ii] = TC_all[ii]*(1-(rc/Lp)**2 - Ap*(rc/Lp)**4)**(-gamm)
TS_all = Tbot_all*((1-(ri_all/Lp)**2-Ap*(ri_all/Lp)**4)**(-gamm) *
(fc(rc/Lp,gamm,Ap) - fc(ri_all/Lp,gamm,Ap)) /
(fc(rc/Lp,0,Ap) - fc(ri_all/Lp,0,Ap)))
Tdis = ((Tbot_all / (1 - (ri_all/Lp)**2 - Ap*(ri_all/Lp)**4)**gamm) *
((fc(rc/Lp,0,Ap) - fc(ri_all/Lp,0,Ap)) / (fc(rc/Lp,-gamm,Ap) - fc(ri_all/Lp,-gamm,Ap))))
Plc_all = (Tdis*(TLi_all-TC_all)/(TLi_all*TC_all))*Qlc_all
Pic_all = (Tdis*(TLi_all-TC_all)/(TLi_all*TC_all))*Qic_all
Pgc_all = (Tdis/TC_all)*Qgc_all
Qcc_all = Qsc_all + Qpc_all + Qgc_all + Qlc_all + Qic_all
Prc_all = ((Tdis-TC_all)/TC_all)*Qrc_all
Psc_all = (Tdis*(TS_all-TC_all)/(TS_all*TC_all))*Qsc_all
Ppc_all = (Tdis/TC_all)*Qpc_all
Pk_all = Tdis*EK_all
P_inner = Plc_all + Pic_all + Pgc_all
P_outer = Prc_all + Psc_all + Ppc_all - Pk_all
Vc = 4/3*np.pi*(rc-ri_all)**3 # Volume of the core (m^3)
rho_av = Mc/Vc # Average density of the core (kg/m^3)
D_all = rc-ri_all # Thickness of the outer core (m)
phi_outer = rc*g/2 # Gravitational potential at the CMB (m^2/s^2)
phi_inner = ri_all**2/rc * g/2 # Gravitational potential at the inner core boundary (m^2/s^2)
phi_mean = 0.3*g/rc*((rc**5-ri_all**5)/(rc**3-ri_all**3)) # Average grav. potential in the outer core (m^2/s^2)
P_total = P_inner + P_outer
TDM_all = np.zeros(NN)
for ii, P in enumerate(P_total):
if P > 0:
F_inner = P_inner[ii]/(phi_mean[ii] - phi_inner[ii])
F_outer = P_outer[ii]/(phi_outer - phi_mean[ii])
f_rat = F_inner/(F_outer+F_inner)
powB = (P_inner[ii] + P_outer[ii])/(Vc[ii]*rho_av[ii]*OmegaE**3*D_all[ii]**2)
b_dip = 7.3*(1-ri_all[ii]/rc)*(1+f_rat)
B_rms = powB**0.34*np.sqrt(rho_av[ii]*mu0)*OmegaE*D_all[ii]
TDM = np.maximum(4*np.pi*rc**3/(np.sqrt(2)*mu0) * B_rms/b_dip, 0)
TDM_all[ii] = TDM
else:
TDM_all[ii] = 0
magicConstant = 7.94e22/TDM_all[NN-1]
TDM_all = magicConstant * TDM_all
Bs_core = mu0*TDM_all/(4*np.pi*rp**3)
return (
t_all, h_all, ri_all, TC_all, TS_all, TLi_all, Tdis, Tbot_all,
Qbmo_all, Qsm_all, Qlm_all, Qrm_all, Qcc_all,
Qsc_all, Qrc_all, Qpc_all, Qgc_all, Qlc_all, Qic_all, TDM_all, magicConstant,
Psc_all, Prc_all, Ppc_all, Pgc_all, Plc_all, Pic_all, Pk_all, P_total, Bs_core,
v_mix, v_CIA, v_MAC, Rm_mix, Rm_CIA, Rm_MAC, Bs_mix, Bs_CIA, Bs_MAC,
)
def postprocessEarth(Qbmo0, QbmoNow, H0M, h, TC, core_Kppm, Ppc):
(t_all, h_all, ri_all, TC_all, TS_all, TLi_all, Tdis, Tbot_all,
Qbmo_all, Qsm_all, Qlm_all, Qrm_all, Qcc_all,
Qsc_all, Qrc_all, Qpc_all, Qgc_all, Qlc_all, Qic_all, TDM_all, magicConstant,
Psc_all, Prc_all, Ppc_all, Pgc_all, Plc_all, Pic_all, Pk_all, P_total, Bs_core,
v_mix, v_CIA, v_MAC, Rm_mix, Rm_CIA, Rm_MAC, Bs_mix, Bs_CIA, Bs_MAC,
) = runEarthModel(Qbmo0, QbmoNow, H0M, h, TC, core_Kppm, Ppc)
if np.argmin(Rm_mix>40) > 0:
i_mix = np.argmin(Rm_mix>40)
Bs_mix_now = 0
t_mix = t_all[i_mix]
else:
Bs_mix_now = Bs_mix[-1]
t_mix = t_all[-1]
if np.argmin(Rm_CIA>40) > 0:
i_CIA = np.argmin(Rm_CIA>40)
Bs_CIA_now = 0
t_CIA = t_all[i_CIA]
else:
Bs_CIA_now = Bs_CIA[-1]
t_CIA = t_all[-1]
if np.argmin(Rm_MAC>40) > 0:
i_MAC = np.argmin(Rm_MAC>40)
Bs_MAC_now = 0
t_MAC = t_all[i_MAC]
else:
Bs_MAC_now = Bs_MAC[-1]
t_MAC = t_all[-1]
h_now = h_all[-1]
Qcc_now = Qcc_all[-1]
Bs_core_now = Bs_core[-1]
y2s = 3.154e7
return (h_now/1e3, Qcc_now/1e12, 1e6*Bs_core_now,
1e6*Bs_mix_now, 1e6*Bs_CIA_now, 1e6*Bs_MAC_now,
t_mix/(y2s*1e9), t_CIA/(y2s*1e9), t_MAC/(y2s*1e9))
# "Constants"
QbmoNow = 15e12 # Present-day value of Qbmo (W)
H0M = 20e12 # Initial radiogenic heating (W)
core_Kppm = 50 # Amount of K in the core (ppm)
Ppc = 5e-6 # Precipitation rate of light elements (1/K)
# Reference values
Qbmo0_ref = 55e12
h0_ref = 750e3
TC0_ref = 5230
(h_now_ref, Qcc_now_ref, Bs_core_now_ref,
Bs_mix_now_ref, Bs_CIA_now_ref, Bs_MAC_now_ref,
t_mix_ref, t_CIA_ref, t_MAC_ref) = postprocessEarth(Qbmo0_ref, QbmoNow, H0M, h0_ref, TC0_ref, core_Kppm, Ppc)
# Sensitivity test
N_T0s = 10
N_Q0s = 11
N_h0s = 12
Qbmo0s = 1e12*np.linspace(35,60,N_Q0s) # Initial value of Qbmo (W)
h0s = 1e3*np.linspace(600,1500,N_h0s)
TC0s = np.linspace(4400,6100,N_T0s)
h_h0 = np.zeros((N_h0s,N_Q0s))
h_TC = np.zeros((N_T0s,N_Q0s))
td_h0 = np.zeros((N_h0s,N_Q0s))
td_TC = np.zeros((N_T0s,N_Q0s))
for ii, Qbmo0 in enumerate(Qbmo0s):
for jj, h0 in enumerate(h0s):
(h_now, Qcc_now, Bs_core_now,
Bs_mix_now, Bs_CIA_now, Bs_MAC_now,
t_mix, t_CIA, t_MAC) = postprocessEarth(Qbmo0, QbmoNow, H0M, h0, TC0_ref, core_Kppm, Ppc)
h_h0[jj,ii] = h_now
td_h0[jj,ii] = t_CIA
for kk, TC0 in enumerate(TC0s):
(h_now, Qcc_now, Bs_core_now,
Bs_mix_now, Bs_CIA_now, Bs_MAC_now,
t_mix, t_CIA, t_MAC) = postprocessEarth(Qbmo0, QbmoNow, H0M, h0_ref, TC0, core_Kppm, Ppc)
h_TC[kk,ii] = h_now
td_TC[kk,ii] = t_CIA
fig, axs = plt.subplots(2,1,figsize=(9,9))
fn = 'Arial'
fs = 18
lw = 3
ax1 = plt.subplot(211)
CP = plt.contourf(Qbmo0s/1e12, h0s/1e3, h_h0, 50, vmin=0, vmax=950)
plt.contour(Qbmo0s/1e12, h0s/1e3, h_h0, levels = [10], colors = ['w'], linewidths = [lw], linestyles='dashed')
plt.ylabel('Initial BMO thickness (km)',fontname=fn,fontsize=fs)
plt.xticks(np.linspace(35,60,6),fontname=fn, fontsize=fs)
plt.yticks(np.linspace(750,1500,4),fontname=fn, fontsize=fs)
plt.scatter(Qbmo0_ref/1e12,h0_ref/1e3,s=300,c='white',marker='*')
ax1.spines["top"].set_visible(False)
ax1.spines["right"].set_visible(False)
plt.minorticks_on()
cbar = fig.colorbar(CP, ticks=np.linspace(0,800,5))
cbar.ax.set_ylabel('Present-day thickness\nof the BMO (km)',fontname=fn,fontsize=fs)
cbar.ax.tick_params(labelsize=fs)
for c in CP.collections:
c.set_edgecolor("face")
plt.text(0.95,0.93,'a',color='white',fontname=fn,fontsize=fs,fontweight='bold',transform=ax1.transAxes)
ax2 = plt.subplot(212)
CP2 = plt.contourf(Qbmo0s/1e12, h0s/1e3, td_h0, 50, vmin=0.34, vmax=4.5, cmap='magma_r')
plt.contour(Qbmo0s/1e12, h0s/1e3, h_h0, levels = [10], colors = ['w'], linewidths = [lw], linestyles='dashed')
plt.ylabel('Initial BMO thickness (km)',fontname=fn,fontsize=fs)
plt.xticks(np.linspace(35,60,6),fontname=fn, fontsize=fs)
plt.yticks(np.linspace(750,1500,4),fontname=fn, fontsize=fs)
plt.scatter(Qbmo0_ref/1e12,h0_ref/1e3,s=300,c='white',marker='*')
ax2.spines["top"].set_visible(False)
ax2.spines["right"].set_visible(False)
plt.minorticks_on()
cbar2 = fig.colorbar(CP2, ticks=np.linspace(0,4,5))
cbar2.ax.set_ylabel('Lifetime of the BMO\ndynamo (Gyr)',fontname=fn,fontsize=fs)
cbar2.ax.tick_params(labelsize=fs)
for c in CP2.collections:
c.set_edgecolor("face")
plt.text(0.95,0.93,'b',color='white',fontname=fn,fontsize=fs,fontweight='bold',transform=ax2.transAxes)
plt.savefig('sensitiveEarth.pdf')
plt.show()
```
| github_jupyter |
# MNIST
## 1. as a image
```
import torch
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from sklearn.model_selection import train_test_split
from torch import nn,functional
import torch.nn.functional as F
np.random.seed(1) # seed
df_train = pd.read_csv(r"./mnist_as_csv/train.csv")
df_train.head(5)
X = df_train.loc[:,df_train.columns !="label"]
Y = df_train.loc[:,df_train.columns =="label"]
X = X/255
X =X.to_numpy()
Y = Y.to_numpy()
x_train, x_test, y_train, y_test = train_test_split(
X, Y, test_size=0.2, random_state=0)
BATCH_SIZE =16
X_train = torch.from_numpy(x_train)
X_test = torch.from_numpy(x_test)
Y_train = torch.from_numpy(y_train).type(torch.LongTensor)
Y_test = torch.from_numpy(y_test).type(torch.LongTensor)
train = torch.utils.data.TensorDataset(X_train,Y_train)
test = torch.utils.data.TensorDataset(X_test,Y_test)
train_loader = torch.utils.data.DataLoader(train, batch_size = BATCH_SIZE, shuffle = True)
test_loader = torch.utils.data.DataLoader(test, batch_size = BATCH_SIZE, shuffle = True)
```
# model
## 1.1DNN
```
class DNN(nn.Module):
def __init__(self):
super(DNN,self).__init__()
self.fc1 = nn.Linear(28 * 28, 128)
self.fc2 = nn.Linear(128, 64)
self.fc3 = nn.Linear(64, 10)
self.relu= nn.ReLU()
def forward(self,x):
x = self.fc1(x)
x = self.relu(x)
x = self.fc2(x)
x = self.relu(x)
x = self.fc3(x)
return F.log_softmax(x, dim=1)
class DNN_drop(nn.Module):
def __init__(self):
super(DNN_drop,self).__init__()
self.fc1 = nn.Linear(28 * 28, 128)
self.dropout= nn.Dropout(0.1)
self.fc2 = nn.Linear(128, 64)
self.fc3 = nn.Linear(64, 10)
self.relu= nn.ReLU()
def forward(self,x):
x = self.fc1(x)
x = self.relu(x)
x = self.dropout(x)
x = self.fc2(x)
x = self.relu(x)
x = self.fc3(x)
return F.log_softmax(x, dim=1)
class CNN(nn.Module):
def __init__(self):
super(CNN,self).__init__()
self.conv = nn.Conv2d(in_channels = 1, out_channels = 6, kernel_size = 2)
self.relu = nn.ReLU()
self.fc1 = nn.Linear(4374, 128)
self.fc2 = nn.Linear(128, 64)
self.out = nn.Linear(64, 10)
def forward(self,x):
out = self.conv(x)
out = self.relu(out)
out = out.view(out.size(0), -1)
out = self.fc1(out)
out = self.fc2(out)
out = self.out(out)
return F.log_softmax(out, dim=1)
from torchsummary import summary
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
model_1 = DNN().to(device)
model_2 = DNN_drop().to(device)
model_3 = CNN().to(device)
# model_1=model_1.double()
# model_2=model_2.double()
# model_3=model_3.double()
summary(model_1, (1, 28*28))
summary(model_2, (1, 28*28))
summary(model_3, (1, 28, 28))
# criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model_1.parameters(),lr=0.001)
```
# dataset
### custom dataset과 image folder를 활용해보자
```
# class MNIST_CSV(Dataset):
# def __init__(self, csv_file, transform=None):
# self.mnist = pd.read_csv(csv_file)
# self.transform = transform
# def __len__(self):
# return len(self.mnist)
# def __getitem__(self, idx):
# sample = self.mnist[idx]
# if self.transform:
# sample = self.transform(sample)
# return sample
# from torchvision.datasets import ImageFolder
# # #dataset 먼저 선언 후 loader에 맵핑
# train_dataset = ImageFolder(root='./mnist_as_image', transform=None)
# train_loader = torch.utils.data.DataLoader(train_dataset, batch_size=16, shuffle=True, num_workers=4)
```
## without dropout
```
total_step = len(train_loader)
for epoch in range(15):
model_1.train()
for i, (images, labels) in enumerate(train_loader):
# Move tensors to the configured device
images = images.to(device).float()
labels = labels.to(device)
labels = labels.squeeze(1)
optimizer.zero_grad()
outputs = model_1(images)
loss = F.nll_loss(outputs, labels)
loss.backward()
optimizer.step()
with torch.no_grad():
correct = 0
total = 0
model_1.eval()
for images, labels in test_loader:
images = images.to(device).float()
labels = labels.to(device)
labels = labels.squeeze(1)
outputs = model_1(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: {} %'.format(100 * correct / total))
optimizer = torch.optim.Adam(model_2.parameters(),lr=0.001)
```
## with dropout
```
total_step = len(train_loader)
for epoch in range(15):
model_2.train()
for i, (images, labels) in enumerate(train_loader):
images = images.to(device).float()
labels = labels.to(device)
labels = labels.squeeze(1)
optimizer.zero_grad()
outputs = model_2(images)
loss = F.nll_loss(outputs, labels)
# print("loss is :%f" % loss.item())
loss.backward()
optimizer.step()
with torch.no_grad():
correct = 0
total = 0
model_2.eval()
for images, labels in test_loader:
images = images.to(device).float()
labels = labels.to(device)
labels = labels.squeeze(1)
outputs = model_2(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: {} %'.format(100 * correct / total))
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model_3.parameters(),lr=0.001)
total_step = len(train_loader)
for epoch in range(15):
model_3.train()
for i, (images, labels) in enumerate(train_loader):
# Move tensors to the configured device
images = images.reshape(-1,1,28,28).to(device).float()
labels = labels.to(device)
labels = labels.squeeze(1)
optimizer.zero_grad()
outputs = model_3(images)
loss = F.nll_loss(outputs, labels)
loss.backward()
optimizer.step()
if (i+1) % 100 == 0:
print ('Epoch [{}/{}], Step [{}/{}], Loss: {:.4f}'
.format(epoch+1, 15, i+1, total_step, loss.item()))
with torch.no_grad():
correct = 0
total = 0
model_3.eval()
for images, labels in test_loader:
images = images.reshape(-1,1,28,28).to(device).float()
labels = labels.to(device)
labels = labels.squeeze(1)
outputs = model_3(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: {} %'.format(100 * correct / total))
```
| github_jupyter |
# Setup
```
%matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
data = pd.read_csv("articles_2017-09-01_2017-09-30.csv", index_col="id", \
parse_dates=["published", "discovered"])
data.head()
```
# Response Score
The response score is a number between 0 and 50 that indicates the level of response to an article.
Perhaps in the future we may choose to include other factors, but for now we just include engagements on Facebook. The maximum score of 50 should be achieved by an article that does really well compared with others.
```
pd.options.display.float_format = '{:.2f}'.format
data.fb_engagements.describe([0.5, 0.75, 0.9, 0.95, 0.99, 0.995, 0.999])
```
There's a few articles there with 1 million plus engagements, let's just double check that.
```
data[data.fb_engagements > 1000000]
data.fb_engagements.mode()
```
Going back to the enagement counts, we see the mean is 1,315, mode is zero, median is 26, 90th percentile is 1,717, 99th percentile is 24,236, 99.5th percentile is 42,666. The standard deviation is 9,625, significantly higher than the mean, so this is not a normal distribution.
Key publishers stats
```
data.groupby("publisher_id").agg({'url': 'count', 'fb_engagements': ['sum', 'median', 'mean']})
mean = data.fb_engagements.mean()
median = data.fb_engagements.median()
non_zero_fb_enagagements = data.fb_engagements[data.fb_engagements > 0]
```
That's a bit better, but still way too clustered at the low end. Let's look at a log normal distribution.
```
mean = data.fb_engagements.mean()
median = data.fb_engagements.median()
ninety = data.fb_engagements.quantile(.90)
ninetyfive = data.fb_engagements.quantile(.95)
ninetynine = data.fb_engagements.quantile(.99)
plt.figure(figsize=(12,4.5))
plt.hist(np.log(non_zero_fb_enagagements + median), bins=50)
plt.axvline(np.log(mean), linestyle=':', label=f'Mean ({mean:,.0f})', color='green')
plt.axvline(np.log(median), label=f'Median ({median:,.0f})', color='green')
plt.axvline(np.log(ninety), linestyle='--', label=f'90% percentile ({ninety:,.0f})', color='red')
plt.axvline(np.log(ninetyfive), linestyle='-.', label=f'95% percentile ({ninetyfive:,.0f})', color='red')
plt.axvline(np.log(ninetynine), linestyle=':', label=f'99% percentile ({ninetynine:,.0f})', color='red')
leg = plt.legend()
log_engagements = (non_zero_fb_enagagements
.clip_upper(data.fb_engagements.quantile(.999))
.apply(lambda x: np.log(x + median))
)
log_engagements.describe()
```
Use standard feature scaling to bring that to a 1 to 50 range
```
def scale_log_engagements(engagements_logged):
return np.ceil(
50 * (engagements_logged - log_engagements.min()) / (log_engagements.max() - log_engagements.min())
)
def scale_engagements(engagements):
return scale_log_engagements(np.log(engagements + median))
scaled_non_zero_engagements = scale_log_engagements(log_engagements)
scaled_non_zero_engagements.describe()
# add in the zeros, as zero
scaled_engagements = pd.concat([scaled_non_zero_engagements, data.fb_engagements[data.fb_engagements == 0]])
proposed = pd.DataFrame({"fb_engagements": data.fb_engagements, "response_score": scaled_engagements})
proposed.response_score.plot.hist(bins=50)
```
Looks good to me, lets save that.
```
data["response_score"] = proposed.response_score
```
### Proposal
The maximum of 50 points is awarded when the engagements are greater than the 99.9th percentile, rolling over the last month.
i.e. where $limit$ is the 99.5th percentile of engagements calculated over the previous month, the response score for article $a$ is:
\begin{align}
basicScore_a & =
\begin{cases}
0 & \text{if } engagements_a = 0 \\
\log(\min(engagements_a,limit) + median(engagements)) & \text{if } engagements_a > 0
\end{cases} \\
responseScore_a & =
\begin{cases}
0 & \text{if } engagements_a = 0 \\
50 \cdot \frac{basicScore_a - \min(basicScore)}{\max(basicScore) - \min(basicScore)} & \text{if } engagements_a > 0
\end{cases} \\
\\
\text{The latter equation can be expanded to:} \\
responseScore_a & =
\begin{cases}
0 & \text{if } engagements_a = 0 \\
50 \cdot
\frac{\log(\min(engagements_a,limit) + median(engagements)) - \log(1 + median(engagements))}
{\log(limit + median(engagements)) - \log(1 + median(engagements))} & \text{if } engagements_a > 0
\end{cases} \\
\end{align}
# Promotion Score
The aim of the promotion score is to indicate how important the article was to the publisher, by tracking where they chose to promote it. This is a number between 0 and 50 comprised of:
- 20 points based on whether the article was promoted as the "lead" story on the publisher's home page
- 15 points based on how long the article was promoted anywhere on the publisher's home page
- 15 points based on whether the article was promoted on the publisher's main facebook brand page
The first two should be scaled by the popularity/reach of the home page, for which we use the alexa page rank as a proxy.
The last should be scaled by the popularity/reach of the brand page, for which we use the number of likes the brand page has.
### Lead story (20 points)
```
data.mins_as_lead.describe([0.5, 0.75, 0.9, 0.95, 0.99, 0.995, 0.999])
```
As expected, the vast majority of articles don't make it as lead. Let's explore how long typically publishers put something as lead for.
```
lead_articles = data[data.mins_as_lead > 0]
lead_articles.mins_as_lead.describe([0.25, 0.5, 0.75, 0.9, 0.95, 0.99, 0.995, 0.999])
lead_articles.mins_as_lead.plot.hist(bins=50)
```
For lead, it's a significant thing for an article to be lead at all, so although we want to penalise articles that were lead for a very short time, mostly we want to score the maximum even if it wasn't lead for ages. So we'll give maximum points when something has been lead for an hour.
```
lead_articles.mins_as_lead.clip_upper(60).plot.hist(bins=50)
```
We also want to scale this by the alexa page rank, such that the maximum score of 20 points is for an article that was on the front for 4 hours for the most popular site.
So lets explore the alexa nunbers.
```
alexa_ranks = data.groupby(by="publisher_id").alexa_rank.mean().sort_values()
alexa_ranks
alexa_ranks.plot.bar(figsize=[10,5])
```
Let's try the simple option first: just divide the number of minutes as lead by the alexa rank. What's the scale of numbers we get then.
```
lead_proposal_1 = lead_articles.mins_as_lead.clip_upper(60) / lead_articles.alexa_rank
lead_proposal_1.plot.hist()
```
Looks like there's too much of a cluster around 0. Have we massively over penalised the publishers with a high alexa rank?
```
lead_proposal_1.groupby(data.publisher_id).mean().plot.bar(figsize=[10,5])
```
Yes. Let's try taking the log of the alexa rank and see if that looks better.
```
lead_proposal_2 = (lead_articles.mins_as_lead.clip_upper(60) / np.log(lead_articles.alexa_rank))
lead_proposal_2.plot.hist()
lead_proposal_2.groupby(data.publisher_id).describe()
lead_proposal_2.groupby(data.publisher_id).min().plot.bar(figsize=[10,5])
```
That looks about right, as long as the smaller publishers were closer to zero. So let's apply feature scaling to this, to give a number between 1 and 20. (Anything not as lead will pass though as zero.)
```
def rescale(series):
return (series - series.min()) / (series.max() - series.min())
lead_proposal_3 = np.ceil(20 * rescale(lead_proposal_2))
lead_proposal_2.min(), lead_proposal_2.max()
lead_proposal_3.plot.hist()
lead_proposal_3.groupby(data.publisher_id).median().plot.bar(figsize=[10,5])
data["lead_score"] = pd.concat([lead_proposal_3, data.mins_as_lead[data.mins_as_lead==0]])
data.lead_score.value_counts().sort_index()
data.lead_score.groupby(data.publisher_id).max()
```
In summary then, score for article $a$ is:
$$
unscaledLeadScore_a = \frac{\min(minsAsLead_a, 60)}{\log(alexaRank_a)}\\
leadScore_a = 19 \cdot
\frac{unscaledLeadScore_a - \min(unscaledLeadScore)}
{\max(unscaledLeadScore) - \min(unscaledLeadScore)}
+ 1
$$
Since the minium value of $minsAsLead$ is 1, $\min(unscaledLeadScore)$ is pretty insignificant. So we can simplify this to:
$$
leadScore_a = 20 \cdot
\frac{unscaledLeadScore_a }
{\max(unscaledLeadScore)}
$$
or:
$$
leadScore_a = 20 \cdot
\frac{\frac{\min(minsAsLead_a, 60)}{\log(alexaRank_a)} }
{\frac{60}{\log(\max(alexaRank))}}
$$
$$
leadScore_a = \left( 20 \cdot
\frac{\min(minsAsLead_a, 60)}{\log(alexaRank_a)} \cdot
{\frac{\log(\max(alexaRank))}{60}} \right)
$$
## Time on front score (15 points)
This is similar to time as lead, so lets try doing the same calculation, except we also want to factor in the number of slots on the front:
$$frontScore_a =
15
\left(\frac{\min(minsOnFront_a, 1440)}{alexaRank_a \cdot numArticlesOnFront_a}\right)
\left( \frac{\min(alexaRank \cdot numArticlesOnFront)}{1440} \right)$$
```
(data.alexa_rank * data.num_articles_on_front).min() / 1440
time_on_front_proposal_1 = np.ceil(data.mins_on_front.clip_upper(1440) / (data.alexa_rank * data.num_articles_on_front) * (2.45) * 15)
time_on_front_proposal_1.plot.hist(figsize=(15, 7), bins=15)
time_on_front_proposal_1.value_counts().sort_index()
time_on_front_proposal_1.groupby(data.publisher_id).sum()
```
That looks good to me.
```
data["front_score"] = np.ceil(data.mins_on_front.clip_upper(1440) / (data.alexa_rank * data.num_articles_on_front) * (2.45) * 15).fillna(0)
data.front_score
```
## Facebook brand page promotion (15 points)
One way a publisher has of promoting content is to post to their brand page. The significance of doing so is stronger when the brand page has more followers (likes).
$$ facebookPromotionProposed1_a = 15 \left( \frac {brandPageLikes_a} {\max(brandPageLikes)} \right) $$
Now lets explore the data to see if that makes sense. **tr;dr the formula above is incorrect**
```
data.fb_brand_page_likes.max()
facebook_promotion_proposed_1 = np.ceil((15 * (data.fb_brand_page_likes / data.fb_brand_page_likes.max())).fillna(0))
facebook_promotion_proposed_1.value_counts().sort_index().plot.bar()
facebook_promotion_proposed_1.groupby(data.publisher_id).describe()
```
That's too much variation: sites like the Guardian, which have a respectable 7.5m likes, should not be scoring a 3. Lets try applying a log to it, and then standard feature scaling again.
```
data.fb_brand_page_likes.groupby(data.publisher_id).max()
np.log(2149)
np.log(data.fb_brand_page_likes.groupby(data.publisher_id).max())
```
That's more like it, but the lower numbers should be smaller.
```
np.log(data.fb_brand_page_likes.groupby(data.publisher_id).max() / 1000)
scaled_fb_brand_page_likes = (data.fb_brand_page_likes / 1000)
facebook_promotion_proposed_2 = np.ceil(\
(15 * \
(np.log(scaled_fb_brand_page_likes) / np.log(scaled_fb_brand_page_likes.max()))\
)\
).fillna(0)
facebook_promotion_proposed_2.groupby(data.publisher_id).max()
```
LGTM. So the equation is
$$ facebookPromotion_a = 15 \left(
\frac {\log(\frac {brandPageLikes_a}{1000})}
{\log(\frac {\max(brandPageLikes)}{1000}))} \right) $$
Now, let's try applying standard feature scaling approch to this, rather than using a magic number of 1,000. That equation would be:
\begin{align}
unscaledFacebookPromotion_a &=
\log(brandPageLikes_a) \\
facebookPromotion_a &=
15 \cdot \frac{unscaledFacebookPromotion_a - \min(unscaledFacebookPromotion)}{\max(unscaledFacebookPromotion) - \min(unscaledFacebookPromotion)} \\
\\
\text{The scaling can be simplified to:} \\
facebookPromotion_a &=
15 \cdot \frac{unscaledFacebookPromotion_a - \log(\min(brandPageLikes))}{\log(\max(brandPageLikes)) - \log(\min(brandPageLikes))} \\
\\
\text{Meaning the overall equation becomes:} \\
facebookPromotion_a &=
15 \cdot \frac{\log(brandPageLikes_a) - \log(\min(brandPageLikes))}{\log(\max(brandPageLikes)) - \log(\min(brandPageLikes))}
\end{align}
```
facebook_promotion_proposed_3 = np.ceil(
(14 *
(
(np.log(data.fb_brand_page_likes) - np.log(data.fb_brand_page_likes.min()) ) /
(np.log(data.fb_brand_page_likes.max()) - np.log(data.fb_brand_page_likes.min()))
)
) + 1
)
facebook_promotion_proposed_3.groupby(data.publisher_id).max()
data["facebook_promotion_score"] = facebook_promotion_proposed_3.fillna(0.0)
```
# Review
```
data["promotion_score"] = (data.lead_score + data.front_score + data.facebook_promotion_score)
data["attention_index"] = (data.promotion_score + data.response_score)
data.promotion_score.plot.hist(bins=np.arange(50), figsize=(15,6))
data.attention_index.plot.hist(bins=np.arange(100), figsize=(15,6))
data.attention_index.value_counts().sort_index()
# and lets see the articles with the biggest attention index
data.sort_values("attention_index", ascending=False)
data["score_diff"] = data.promotion_score - data.response_score
# promoted but low response
data.sort_values("score_diff", ascending=False).head(25)
# high response but not promoted
data.sort_values("score_diff", ascending=True).head(25)
```
Write that data to a file. Note that the scores here are provisional for two reasons:
1. they should be using a rolling-month based on the article publication date to calculate medians/min/max etc, whereas in this workbook we as just using values for the month of May
2. for analysis, we've rounded the numbers; we don't expect to do that for the actual scores
```
data.to_csv("articles_with_provisional_scores_2017-09-01_2017-09-30.csv")
```
# Summary
The attention index of an article is comprised of four components:
- *lead score* (max 20 points) based on how long an article was the lead story on the publisher's home page, scaled by the traffic to that publisher
- *front score* (max 15 points) based on how long an article was present on the publisher's home page, scaled by traffic to that publisher
- *Facebook promotion score* (max 15 points) based on whether the article was promoted to the publisher's Facebook brand page, scaled by the reach of that brand page
- *response score* (max 50 points) based on the number of Facebook engagements the article received, relative to other articles
Or, in other words:
\begin{align}
attentionIndex_a &= leadScore_a + frontScore_a + facebookPromotionScore_a + responseScore_a \\
leadScore_a &= 20 \cdot \left(\frac{\min(minsAsLead_a, 60)}{alexaRank_a}\right) \cdot \left( \frac{\min(alexaRank)}{60} \right) \\
frontScore_a &=
15 \cdot
\left(\frac{\min(minsOnFront_a, 1440)}{alexaRank_a \cdot numArticlesOnFront_a}\right) \cdot
\left( \frac{\min(alexaRank \cdot numArticlesOnFront)}{1440} \right) \\
facebookPromotion_a &=
\begin{cases}
0 \text{ if not shared on brand page }\\
15 \cdot \frac{\log(brandPageLikes_a) - \log(\min(brandPageLikes))}{\log(\max(brandPageLikes)) - \log(\min(brandPageLikes))} \text{ otherwise }
\end{cases}
\\
responseScore_a &=
\begin{cases}
0 \text{ if } engagements_a = 0 \\
50 \cdot
\frac{\log(\min(engagements_a,limit) + median(engagements)) - \log(1 + median(engagements))}
{\log(limit + median(engagements)) - \log(1 + median(engagements))} \text{ if } engagements_a > 0
\end{cases} \\
\end{align}
| github_jupyter |
# <center/>加载文本数据集
## 概述
MindSpore提供的`mindspore.dataset`模块可以帮助用户构建数据集对象,分批次地读取文本数据。同时,在各个数据集类中还内置了数据处理和数据分词算子,使得数据在训练过程中能够像经过pipeline管道的水一样源源不断地流向训练系统,提升数据训练效果。
此外,MindSpore还支持分布式场景数据加载,用户可以在加载数据集时指定分片数目,具体用法参见[数据并行模式加载数据集](https://www.mindspore.cn/tutorial/training/zh-CN/master/advanced_use/distributed_training_ascend.html#id6)。
下面,本教程将简要演示如何使用MindSpore加载和处理文本数据。
## 整体流程
- 准备环节。
- 加载数据集。
- 数据处理。
- 数据分词。
## 准备环节
### 导入模块
导入`mindspore.dataset`和`mindspore.dataset.text`模块。
```
import mindspore.dataset as ds
import mindspore.dataset.text as text
```
### 准备所需数据集
创建文本数据,内容如下:
```
Welcome to Beijing
北京欢迎您!
我喜欢English!
```
```
import os
if not os.path.exists('./datasets'):
os.mkdir('./datasets')
file_handle=open('./datasets/tokenizer.txt',mode='w')
file_handle.write('Welcome to Beijing \n北京欢迎您! \n我喜欢English! \n')
file_handle.close()
! tree ./datasets
```
## 加载数据集
MindSpore目前支持加载文本领域常用的经典数据集和多种数据存储格式下的数据集,用户也可以通过构建自定义数据集类实现自定义方式的数据集加载。各种数据集的详细加载方法,可参考编程指南中[数据集加载](https://www.mindspore.cn/doc/programming_guide/zh-CN/master/dataset_loading.html)章节。
下面演示使用`MindSpore.dataset`模块中的`TextFileDataset`类加载数据集。
1. 配置数据集目录,创建数据集对象。
```
DATA_FILE = './datasets/tokenizer.txt'
dataset = ds.TextFileDataset(DATA_FILE, shuffle=False)
```
2. 创建字典迭代器,通过迭代器获取数据。
```
for data in dataset.create_dict_iterator(output_numpy=True):
print(text.to_str(data['text']))
```
## 数据处理
MindSpore目前支持的数据处理算子及其详细使用方法,可参考编程指南中[数据处理](https://www.mindspore.cn/doc/programming_guide/zh-CN/master/pipeline.html)章节。
在生成`dataset`对象后可对其进行数据处理操作,比如`SlidingWindow`、`shuffle`等。
- SlidingWindow
下面演示使用`SlidingWindow`对文本数据进行切片操作。
1. 加载数据集。
```
inputs = [["大","家","早","上","好"]]
dataset_slide = ds.NumpySlicesDataset(inputs, column_names=['text'], shuffle=False)
```
2. 原始数据输出效果。
```
for data in dataset_slide.create_dict_iterator(output_numpy=True):
print(text.to_str(data['text']).tolist())
```
3. 执行切片操作。
```
dataset_slide = dataset_slide.map(operations=text.SlidingWindow(2,0),input_columns=['text'])
```
4. 执行之后输出效果。
```
for data in dataset_slide.create_dict_iterator(output_numpy=True):
print(text.to_str(data['text']).tolist())
```
- shuffle
下面演示在加载数据集时使用`shuffle`对文本数据进行混洗操作。
1. 加载数据集。
```
inputs = ["a","b","c","d"]
dataset_shuffle = ds.NumpySlicesDataset(inputs, column_names=['text'], shuffle=True)
```
2. 数据输出效果。
```
for data in dataset_shuffle.create_dict_iterator(output_numpy=True):
print(text.to_str(data['text']).tolist())
```
## 数据分词
MindSpore目前支持的数据分词算子及其详细使用方法,可参考编程指南中[分词器](https://www.mindspore.cn/doc/programming_guide/zh-CN/master/tokenizer.html)章节。
下面演示使用`WhitespaceTokenizer`分词器来分词,该分词是按照空格来进行分词。
1. 创建`tokenizer`。
```
tokenizer = text.WhitespaceTokenizer()
```
2. 执行操作`tokenizer`。
```
dataset = dataset.map(operations=tokenizer)
```
3. 创建字典迭代器,通过迭代器获取数据。
```
for data in dataset.create_dict_iterator(num_epochs=1,output_numpy=True):
print(text.to_str(data['text']).tolist())
```
| github_jupyter |
# Binary extension MLP for ordinal regression and deep learning -- cement strength dataset
This tutorial explains how to train a deep neural network (here: multilayer perceptron) with the binary extension method by Niu at al. 2016 for ordinal regression.
**Paper reference:**
- Niu, Zhenxing, Mo Zhou, Le Wang, Xinbo Gao, and Gang Hua. "[Ordinal regression with multiple output cnn for age estimation](https://openaccess.thecvf.com/content_cvpr_2016/papers/Niu_Ordinal_Regression_With_CVPR_2016_paper.pdf)." In Proceedings of the IEEE conference on computer vision and pattern recognition.
## 0 -- Obtaining and preparing the cement_strength dataset
We will be using the cement_strength dataset from [https://github.com/gagolews/ordinal_regression_data/blob/master/cement_strength.csv](https://github.com/gagolews/ordinal_regression_data/blob/master/cement_strength.csv).
First, we are going to download and prepare the and save it as CSV files locally. This is a general procedure that is not specific to CORN.
This dataset has 5 ordinal labels (1, 2, 3, 4, and 5). Note that the method requires labels to be starting at 0, which is why we subtract "1" from the label column.
```
import pandas as pd
import numpy as np
data_df = pd.read_csv("https://raw.githubusercontent.com/gagolews/ordinal_regression_data/master/cement_strength.csv")
data_df["response"] = data_df["response"]-1 # labels should start at 0
data_labels = data_df["response"]
data_features = data_df.loc[:, ["V1", "V2", "V3", "V4", "V5", "V6", "V7", "V8"]]
print('Number of features:', data_features.shape[1])
print('Number of examples:', data_features.shape[0])
print('Labels:', np.unique(data_labels.values))
```
### Split into training and test data
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
data_features.values,
data_labels.values,
test_size=0.2,
random_state=1,
stratify=data_labels.values)
```
### Standardize features
```
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train_std = sc.fit_transform(X_train)
X_test_std = sc.transform(X_test)
```
## 1 -- Setting up the dataset and dataloader
In this section, we set up the data set and data loaders using PyTorch utilities. This is a general procedure that is not specific to the method.
```
import torch
##########################
### SETTINGS
##########################
# Hyperparameters
random_seed = 1
learning_rate = 0.05
num_epochs = 50
batch_size = 128
# Architecture
NUM_CLASSES = 10
# Other
DEVICE = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print('Training on', DEVICE)
from torch.utils.data import Dataset
class MyDataset(Dataset):
def __init__(self, feature_array, label_array, dtype=np.float32):
self.features = feature_array.astype(np.float32)
self.labels = label_array
def __getitem__(self, index):
inputs = self.features[index]
label = self.labels[index]
return inputs, label
def __len__(self):
return self.labels.shape[0]
import torch
from torch.utils.data import DataLoader
# Note transforms.ToTensor() scales input images
# to 0-1 range
train_dataset = MyDataset(X_train_std, y_train)
test_dataset = MyDataset(X_test_std, y_test)
train_loader = DataLoader(dataset=train_dataset,
batch_size=batch_size,
shuffle=True, # want to shuffle the dataset
num_workers=0) # number processes/CPUs to use
test_loader = DataLoader(dataset=test_dataset,
batch_size=batch_size,
shuffle=False,
num_workers=0)
# Checking the dataset
for inputs, labels in train_loader:
print('Input batch dimensions:', inputs.shape)
print('Input label dimensions:', labels.shape)
break
```
## 2 - Equipping MLP with a modified output layer
In this section, we are using the output layer as it was originally implemented by Niu et al. This is actually very similar to a standard output layer as we can see below:
```
class MLP(torch.nn.Module):
def __init__(self, in_features, num_classes, num_hidden_1=300, num_hidden_2=300):
super().__init__()
self.num_classes = num_classes
self.my_network = torch.nn.Sequential(
# 1st hidden layer
torch.nn.Linear(in_features, num_hidden_1, bias=False),
torch.nn.LeakyReLU(),
torch.nn.Dropout(0.2),
torch.nn.BatchNorm1d(num_hidden_1),
# 2nd hidden layer
torch.nn.Linear(num_hidden_1, num_hidden_2, bias=False),
torch.nn.LeakyReLU(),
torch.nn.Dropout(0.2),
torch.nn.BatchNorm1d(num_hidden_2),
)
### Specify Niu et al. layer
self.fc = torch.nn.Linear(num_hidden_2, (num_classes-1)*2)
###--------------------------------------------------------------------###
def forward(self, x):
x = self.my_network(x)
logits = self.fc(x)
### Reshape logits for Niu et al. loss
logits = logits.view(-1, (self.num_classes-1), 2)
###--------------------------------------------------------------------###
return logits
torch.manual_seed(random_seed)
model = MLP(in_features=8, num_classes=NUM_CLASSES)
model.to(DEVICE)
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
```
## 3 - Using the extended binary loss for model training
During training, all you need to do is to
1) convert the integer class labels into the extended binary label format using the `levels_from_labelbatch` provided via `coral_pytorch`:
```python
levels = levels_from_labelbatch(class_labels,
num_classes=NUM_CLASSES)
```
2) Apply the extended binary loss:
```python
def niu_et_al_loss(logits, levels):
val = (-torch.sum((F.log_softmax(logits, dim=2)[:, :, 1]*levels
+ F.log_softmax(logits, dim=2)[:, :, 0]*(1-levels)), dim=1))
return torch.mean(val)
loss = niu_et_al_loss(logits, levels)
```
```
#!pip install coral-pytorch
import torch.nn.functional as F
from coral_pytorch.dataset import levels_from_labelbatch
def niu_et_al_loss(logits, levels):
val = (-torch.sum((F.log_softmax(logits, dim=2)[:, :, 1]*levels
+ F.log_softmax(logits, dim=2)[:, :, 0]*(1-levels)), dim=1))
return torch.mean(val)
for epoch in range(num_epochs):
model = model.train()
for batch_idx, (features, class_labels) in enumerate(train_loader):
##### Convert class labels for extended binary loss
levels = levels_from_labelbatch(class_labels,
num_classes=model.num_classes)
###--------------------------------------------------------------------###
features = features.to(DEVICE)
levels = levels.to(DEVICE)
logits = model(features)
#### Niu et al. loss loss
loss = niu_et_al_loss(logits, levels)
###--------------------------------------------------------------------###
optimizer.zero_grad()
loss.backward()
optimizer.step()
### LOGGING
if not batch_idx % 200:
print ('Epoch: %03d/%03d | Batch %03d/%03d | Loss: %.4f'
%(epoch+1, num_epochs, batch_idx,
len(train_loader), loss))
```
## 4 -- Evaluate model
Finally, after model training, we can evaluate the performance of the model. For example, via the mean absolute error and mean squared error measures.
For this, we are going to use the `niu_logits_to_label` utility function below.
```
def niu_logits_to_labels(logits):
probas = F.softmax(logits, dim=2)[:, :, 1]
predict_levels = probas > 0.5
predicted_labels = torch.sum(predict_levels, dim=1)
return predicted_labels
def compute_mae_and_mse(model, data_loader, device):
with torch.no_grad():
mae, mse, acc, num_examples = 0., 0., 0., 0
for i, (features, targets) in enumerate(data_loader):
features = features.to(device)
targets = targets.float().to(device)
logits = model(features)
predicted_labels = niu_logits_to_labels(logits)
num_examples += targets.size(0)
mae += torch.sum(torch.abs(predicted_labels - targets))
mse += torch.sum((predicted_labels - targets)**2)
mae = mae / num_examples
mse = mse / num_examples
return mae, mse
train_mae, train_mse = compute_mae_and_mse(model, train_loader, DEVICE)
test_mae, test_mse = compute_mae_and_mse(model, test_loader, DEVICE)
print(f'Mean absolute error (train/test): {train_mae:.2f} | {test_mae:.2f}')
print(f'Mean squared error (train/test): {train_mse:.2f} | {test_mse:.2f}')
```
| github_jupyter |
```
from datetime import date
from gs_quant.instrument import EqOption, OptionType, OptionStyle, UnderlierType
from gs_quant.session import Environment, GsSession
# external users should substitute their client id and secret; please skip this step if using internal jupyterhub
GsSession.use(Environment.PROD, client_id=None, client_secret=None, scopes=('run_analytics',))
# create a .STOXX50E 3m call option striking at-the-money spot
eq_option = EqOption('.STOXX50E', expiration_date='3m', strike_price='ATMS', option_type=OptionType.Call,
option_style=OptionStyle.European)
# calculate local price and dollar price
print('Local price: {:,.4f}'.format(eq_option.price()))
print('Dollar price: {:,.4f}'.format(eq_option.dollar_price()))
```
#### Underlier Syntax
The underlier accepts an underlier as a RIC or BBID identifier. The default is RIC.
| Syntax | Defintion |
|---------|---------------------|
| 'RIC' | Reuters identifier |
| 'BBID' | Bloomberg identifier |
```
# resolve using a Bloomberg ID
eq_option_bbid = EqOption('SX5E', underlier_type=UnderlierType.BBID, expiration_date='3m', strike_price='ATMS', option_type=OptionType.Call,
option_style=OptionStyle.European)
eq_option_bbid.resolve()
eq_option_bbid.as_dict()
```
#### Strike Syntax
The strike_price syntax allows for an int or a string. The absolute level can be specified using an integer.
The following solver keys using a string format are accepted:
| Syntax | Defintion |
|---------|---------------------|
| '%' | Percent of Spot |
| 'ATMS' | At the Money |
| 'ATMF' | At the Money Forward|
| 'D' | Delta Strikes |
| 'P' | Premium |
- For ATM, ATMF: '1.05*ATMF+.01'
- For Delta Strikes, specify the option delta: '25D', '20D-.01', etc.
- You can also solve for Premium: P=,<target>%
```
# resolve with strike at 110% of spot
eq_atm_solver = EqOption('.STOXX50E', expiration_date='3m', strike_price='ATMS+10%', option_type=OptionType.Put,
option_style=OptionStyle.European)
eq_atm_solver.resolve()
eq_atm_solver.strike_price
# resolve with strike at 94.5% of spot
eq_spot_pct = EqOption('.STOXX50E', expiration_date='3m', strike_price='94.5%', option_type=OptionType.Put,
option_style=OptionStyle.European)
eq_spot_pct.resolve()
eq_spot_pct.strike_price
# resolve with strike at spot minus 10
eq_atmf_solver = EqOption('.STOXX50E', expiration_date='1m', strike_price='ATMF-10', option_type=OptionType.Put,
option_style=OptionStyle.European)
eq_atmf_solver.resolve()
eq_atmf_solver.strike_price
# resolve with strike solving for 10% premium
eq_10x = EqOption('.STOXX50E', expiration_date='6m', strike_price='P=10%', option_type=OptionType.Put,
option_style=OptionStyle.European)
eq_10x.resolve()
eq_10x.strike_price
```
| github_jupyter |
# Train deep model + own emmbedings
```
import numpy as np
import pandas as pd
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.layers import (
Dense,
LSTM,
Embedding,
SpatialDropout1D,
)
from tensorflow.keras.models import (
Model,
load_model,
Sequential
)
from tensorflow.keras.callbacks import ModelCheckpoint
from ast import literal_eval
from sklearn.model_selection import train_test_split
df = pd.read_csv('../data/pandas_data_frame.csv', index_col=0)
all_data = df.where((pd.notnull(df)), '')
all_data['hashtag'] = all_data['hashtag'].apply(literal_eval)
full_text = all_data['tidy_tweet'][(all_data['label']=='1.0') | (all_data['label']=='0.0')]
y = all_data['label'][(all_data['label']=='1.0') | (all_data['label']=='0.0')]
tk = Tokenizer(lower=True, filters='')
tk.fit_on_texts(full_text)
max_len = 120 # Calculate as max in dataset see 1.data_process.ipynb
train_tokenized = tk.texts_to_sequences(full_text)
X = pad_sequences(train_tokenized, maxlen=max_len)
x_train, x_val, y_train, y_val = train_test_split(X, y, random_state=1992, test_size=0.2)
print(x_train.shape,y_train.shape)
print(x_val.shape,y_val.shape)
import pickle
# saving
with open('../model_weights/tokenizer.pickle', 'wb') as handle:
pickle.dump(tk, handle, protocol=pickle.HIGHEST_PROTOCOL)
import sys
sys.path.append("../")
from personal_library.sce_keras.loss_functions import f1_loss
from personal_library.sce_keras.metrics_functions import f1
from personal_library.sce_keras.callbacks import (
LearningRateDecay,
WarmUpCosineDecayScheduler
)
epochs = 200
batch_size = 128
embed_dim = 150
lstm_out = 200
max_fatures = X.max() + 1
learnRate = 0.001
warmup_epoch = 20
lrate_decay = LearningRateDecay(epochs, learnRate).step_decay
warm_up_lr = WarmUpCosineDecayScheduler(learning_rate_base=learnRate,
warmup_learning_rate=0,
warmup_epoch=warmup_epoch,
hold_base_rate_steps=5,
verbose=0)
checkpoint_path = "../model_weights/5_w.hdf5"
checkpoint_path1 = "../model_weights/5_ch.hdf5"
checkpointer = ModelCheckpoint(filepath=checkpoint_path,
monitor='val_loss', verbose=2,
save_best_only=True, mode='min')
checkpointer1 = ModelCheckpoint(filepath=checkpoint_path1,
monitor='val_loss', verbose=2,
save_best_only=False, mode='min')
model = Sequential()
model.add(Embedding(max_fatures, embed_dim, input_length = X.shape[1]))
model.add(SpatialDropout1D(0.3))
model.add(LSTM(lstm_out, dropout=0.5, recurrent_dropout=0.5))
model.add(Dense(1,activation='sigmoid'))
model.compile(loss = f1_loss,
optimizer='adam',
metrics = ['accuracy', f1])
model.summary()
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_val, y_val),
callbacks=[checkpointer, checkpointer1, warm_up_lr])
from sklearn.metrics import f1_score, accuracy_score
#Load best model
model.load_weights(checkpoint_path)
y_pred = model.predict(x_val, batch_size=1)
y_pred = np.where(y_pred > 0.5, 1, 0)
print("Own emmbeding f1_sklearn: {}".format(f1_score(y_val.astype(float), y_pred)))
print("Own emmbeding accuracy: {}".format(accuracy_score(y_val.astype(float), y_pred)))
```
| github_jupyter |
```
import solver
import numpy as np
```
## Generalized PA equation
This solver implements the generalized PA equation
$$N_k(t+1) = N_k(t) + B\delta_{k,1} + \frac{G}{(B+G)t + K_0} [(k-1)N_{k-1}(t) - k N_k(t)],$$
where $N_k(t)$ counts the number of elements with $k$ shares of the total resource $K(t)=t + K_0$ at time $t$, where $B$ is the birth rate and $G$ is the growth rate.
This equation works for abritrary $B,G,K_0$.
So we can test the equation right away, starting from the condition $N_0(t=0)=0$ and $N_1(t=0)=1$.
```
state = np.array([0, 1])
birth_rate = 2
growth_rate = 1
for t in range(1, 10):
state = solver.update_preferential_attachment(state, birth_rate, growth_rate, t)
```
Let's plot the (unormalized) mean field solution after 10 time steps.
```
import matplotlib.pyplot as plt
import seaborn as sns # for style, comment if unavailable
%matplotlib inline
sns.set_style('ticks')
plt.loglog(state)
```
And a few more iterations will allows us to converge nicely
```
for t in range(10, 100):
state = solver.update_preferential_attachment(state, birth_rate, growth_rate, t)
plt.loglog(state);
```
## Automated rate computation
Now, the interesting part of the `solver` module is all the birth / growth computations.
For example, if we have a $d=3$ HPA structure with the parameters used in Fig. 4 of Hébert-Dufresne et al. PRE 92, 062809 (2016), $\vec{p} = (0, 0.0005, 0.185, 0.385, 0)$ and $\vec{q} = (0.8, 0.6, 0.5, 1, 0)$, then the structural / nodal rates of growh / birth are immediately given by the module:
```
p = np.array([0, 0.0005, 0.185, 0.385, 1])
q = np.array([0.80, 0.60, 0.50, 1, 0])
sb = solver.get_sb(p)
sg = solver.get_sg(p)
q_prime = solver.get_q_prime(q, sb, sg)
r = solver.get_r(q_prime, p)
ng = solver.get_ng(r)
nb = solver.get_nb(r, q)
print("Structural birth rate\n", sb)
print("Structural growth rate\n", sg)
print("Nodal birth rate\n", nb)
print("Nodal growth rate\n", ng)
```
## Complete HPA solutions
The above rates can be We can solve the generalized PA equations for level $k=1,2,3$ separately and re-obtain Fig.4.
```
init = np.array([0,1])
n = [init.copy() for i in range(3)]
s = [init.copy() for i in range(3)]
t_max = 1000
for t in range(1, t_max):
for k in [1, 2, 3]:
s[k - 1] = solver.update_preferential_attachment(s[k - 1], sb[k], sg[k], t)
n[k - 1] = solver.update_preferential_attachment(n[k - 1], nb[k], ng[k], t)
plt.figure(figsize=(10,3))
plt.subplot(1,2,1)
for k in [1,2,3]:
plt.loglog(n[k - 1] / sum(n[k - 1]), label="$k=" + str(k) + "$")
plt.xlabel('$m$')
plt.ylabel('$\\tilde{N}_{k,m}$')
plt.ylim(1e-6,1)
plt.legend(loc=1)
plt.subplot(1,2,2)
for k in [1,2,3]:
plt.loglog(s[k - 1] / sum(s[k - 1]), label="$k=" + str(k) + "$")
plt.xlabel('$n$')
plt.ylabel('$\\tilde{S}_{k,n}$')
plt.ylim(1e-6,1)
plt.legend(loc=3)
plt.tight_layout(pad=2)
```
Et *voilà*!
For ease of use, we also provide wrapper functions that perform all the computations at once.
The first is a function that computes all the birth / growth rates.
```
(sb, sg, ng, nb) = solver.get_all_rates(p, q)
print("Structural birth rate\n", sb)
print("Structural growth rate\n", sg)
print("Nodal birth rate\n", nb)
print("Nodal growth rate\n", ng)
```
And the second iterates depth $d$ system of equations for the user.
```
n, s = solver.solve_hpa(p, q, 100)
plt.figure(figsize=(10,3))
plt.subplot(1,2,1)
for k in [1,2,3]:
plt.loglog(n[k - 1] / sum(n[k - 1]), label="$k=" + str(k) + "$")
plt.xlabel('$m$')
plt.ylabel('$\\tilde{N}_{k,m}$')
plt.ylim(1e-6,1)
plt.legend(loc=1)
plt.subplot(1,2,2)
for k in [1,2,3]:
plt.loglog(s[k - 1] / sum(s[k - 1]), label="$k=" + str(k) + "$")
plt.xlabel('$n$')
plt.ylabel('$\\tilde{S}_{k,n}$')
plt.ylim(1e-6,1)
plt.legend(loc=3)
plt.tight_layout(pad=2)
```
This would be what the system looks like after 100 iterations instead of 1000.
| github_jupyter |
```
# look at tools/set_up_magics.ipynb
yandex_metrica_allowed = True ; get_ipython().run_cell('# one_liner_str\n\nget_ipython().run_cell_magic(\'javascript\', \'\', \'// setup cpp code highlighting\\nIPython.CodeCell.options_default.highlight_modes["text/x-c++src"] = {\\\'reg\\\':[/^%%cpp/]} ;\')\n\n# creating magics\nfrom IPython.core.magic import register_cell_magic, register_line_magic\nfrom IPython.display import display, Markdown, HTML\nimport argparse\nfrom subprocess import Popen, PIPE\nimport random\nimport sys\nimport os\nimport re\nimport signal\nimport shutil\nimport shlex\nimport glob\n\n@register_cell_magic\ndef save_file(args_str, cell, line_comment_start="#"):\n parser = argparse.ArgumentParser()\n parser.add_argument("fname")\n parser.add_argument("--ejudge-style", action="store_true")\n args = parser.parse_args(args_str.split())\n \n cell = cell if cell[-1] == \'\\n\' or args.no_eof_newline else cell + "\\n"\n cmds = []\n with open(args.fname, "w") as f:\n f.write(line_comment_start + " %%cpp " + args_str + "\\n")\n for line in cell.split("\\n"):\n line_to_write = (line if not args.ejudge_style else line.rstrip()) + "\\n"\n if line.startswith("%"):\n run_prefix = "%run "\n if line.startswith(run_prefix):\n cmds.append(line[len(run_prefix):].strip())\n f.write(line_comment_start + " " + line_to_write)\n continue\n run_prefix = "%# "\n if line.startswith(run_prefix):\n f.write(line_comment_start + " " + line_to_write)\n continue\n raise Exception("Unknown %%save_file subcommand: \'%s\'" % line)\n else:\n f.write(line_to_write)\n f.write("" if not args.ejudge_style else line_comment_start + r" line without \\n")\n for cmd in cmds:\n display(Markdown("Run: `%s`" % cmd))\n get_ipython().system(cmd)\n\n@register_cell_magic\ndef cpp(fname, cell):\n save_file(fname, cell, "//")\n\n@register_cell_magic\ndef asm(fname, cell):\n save_file(fname, cell, "//")\n \n@register_cell_magic\ndef makefile(fname, cell):\n assert not fname\n save_file("makefile", cell.replace(" " * 4, "\\t"))\n \n@register_line_magic\ndef p(line):\n try:\n expr, comment = line.split(" #")\n display(Markdown("`{} = {}` # {}".format(expr.strip(), eval(expr), comment.strip())))\n except:\n display(Markdown("{} = {}".format(line, eval(line))))\n \ndef show_file(file, clear_at_begin=True, return_html_string=False):\n if clear_at_begin:\n get_ipython().system("truncate --size 0 " + file)\n obj = file.replace(\'.\', \'_\').replace(\'/\', \'_\') + "_obj"\n html_string = \'\'\'\n <!--MD_BEGIN_FILTER-->\n <script type=text/javascript>\n var entrance___OBJ__ = 0;\n var errors___OBJ__ = 0;\n function refresh__OBJ__()\n {\n entrance___OBJ__ -= 1;\n var elem = document.getElementById("__OBJ__");\n if (elem) {\n var xmlhttp=new XMLHttpRequest();\n xmlhttp.onreadystatechange=function()\n {\n var elem = document.getElementById("__OBJ__");\n console.log(!!elem, xmlhttp.readyState, xmlhttp.status, entrance___OBJ__);\n if (elem && xmlhttp.readyState==4) {\n if (xmlhttp.status==200)\n {\n errors___OBJ__ = 0;\n if (!entrance___OBJ__) {\n elem.innerText = xmlhttp.responseText;\n entrance___OBJ__ += 1;\n console.log("req");\n window.setTimeout("refresh__OBJ__()", 300); \n }\n return xmlhttp.responseText;\n } else {\n errors___OBJ__ += 1;\n if (errors___OBJ__ < 10 && !entrance___OBJ__) {\n entrance___OBJ__ += 1;\n console.log("req");\n window.setTimeout("refresh__OBJ__()", 300); \n }\n }\n }\n }\n xmlhttp.open("GET", "__FILE__", true);\n xmlhttp.setRequestHeader("Cache-Control", "no-cache");\n xmlhttp.send(); \n }\n }\n \n if (!entrance___OBJ__) {\n entrance___OBJ__ += 1;\n refresh__OBJ__(); \n }\n </script>\n \n <font color="white"> <tt>\n <p id="__OBJ__" style="font-size: 16px; border:3px #333333 solid; background: #333333; border-radius: 10px; padding: 10px; "></p>\n </tt> </font>\n <!--MD_END_FILTER-->\n <!--MD_FROM_FILE __FILE__ -->\n \'\'\'.replace("__OBJ__", obj).replace("__FILE__", file)\n if return_html_string:\n return html_string\n display(HTML(html_string))\n \nBASH_POPEN_TMP_DIR = "./bash_popen_tmp"\n \ndef bash_popen_terminate_all():\n for p in globals().get("bash_popen_list", []):\n print("Terminate pid=" + str(p.pid), file=sys.stderr)\n p.terminate()\n globals()["bash_popen_list"] = []\n if os.path.exists(BASH_POPEN_TMP_DIR):\n shutil.rmtree(BASH_POPEN_TMP_DIR)\n\nbash_popen_terminate_all() \n\ndef bash_popen(cmd):\n if not os.path.exists(BASH_POPEN_TMP_DIR):\n os.mkdir(BASH_POPEN_TMP_DIR)\n h = os.path.join(BASH_POPEN_TMP_DIR, str(random.randint(0, 1e18)))\n stdout_file = h + ".out.html"\n stderr_file = h + ".err.html"\n run_log_file = h + ".fin.html"\n \n stdout = open(stdout_file, "wb")\n stdout = open(stderr_file, "wb")\n \n html = """\n <table width="100%">\n <colgroup>\n <col span="1" style="width: 70px;">\n <col span="1">\n </colgroup> \n <tbody>\n <tr> <td><b>STDOUT</b></td> <td> {stdout} </td> </tr>\n <tr> <td><b>STDERR</b></td> <td> {stderr} </td> </tr>\n <tr> <td><b>RUN LOG</b></td> <td> {run_log} </td> </tr>\n </tbody>\n </table>\n """.format(\n stdout=show_file(stdout_file, return_html_string=True),\n stderr=show_file(stderr_file, return_html_string=True),\n run_log=show_file(run_log_file, return_html_string=True),\n )\n \n cmd = """\n bash -c {cmd} &\n pid=$!\n echo "Process started! pid=${{pid}}" > {run_log_file}\n wait ${{pid}}\n echo "Process finished! exit_code=$?" >> {run_log_file}\n """.format(cmd=shlex.quote(cmd), run_log_file=run_log_file)\n # print(cmd)\n display(HTML(html))\n \n p = Popen(["bash", "-c", cmd], stdin=PIPE, stdout=stdout, stderr=stdout)\n \n bash_popen_list.append(p)\n return p\n\n\n@register_line_magic\ndef bash_async(line):\n bash_popen(line)\n \n \ndef show_log_file(file, return_html_string=False):\n obj = file.replace(\'.\', \'_\').replace(\'/\', \'_\') + "_obj"\n html_string = \'\'\'\n <!--MD_BEGIN_FILTER-->\n <script type=text/javascript>\n var entrance___OBJ__ = 0;\n var errors___OBJ__ = 0;\n function halt__OBJ__(elem, color)\n {\n elem.setAttribute("style", "font-size: 14px; background: " + color + "; padding: 10px; border: 3px; border-radius: 5px; color: white; "); \n }\n function refresh__OBJ__()\n {\n entrance___OBJ__ -= 1;\n if (entrance___OBJ__ < 0) {\n entrance___OBJ__ = 0;\n }\n var elem = document.getElementById("__OBJ__");\n if (elem) {\n var xmlhttp=new XMLHttpRequest();\n xmlhttp.onreadystatechange=function()\n {\n var elem = document.getElementById("__OBJ__");\n console.log(!!elem, xmlhttp.readyState, xmlhttp.status, entrance___OBJ__);\n if (elem && xmlhttp.readyState==4) {\n if (xmlhttp.status==200)\n {\n errors___OBJ__ = 0;\n if (!entrance___OBJ__) {\n if (elem.innerHTML != xmlhttp.responseText) {\n elem.innerHTML = xmlhttp.responseText;\n }\n if (elem.innerHTML.includes("Process finished.")) {\n halt__OBJ__(elem, "#333333");\n } else {\n entrance___OBJ__ += 1;\n console.log("req");\n window.setTimeout("refresh__OBJ__()", 300); \n }\n }\n return xmlhttp.responseText;\n } else {\n errors___OBJ__ += 1;\n if (!entrance___OBJ__) {\n if (errors___OBJ__ < 6) {\n entrance___OBJ__ += 1;\n console.log("req");\n window.setTimeout("refresh__OBJ__()", 300); \n } else {\n halt__OBJ__(elem, "#994444");\n }\n }\n }\n }\n }\n xmlhttp.open("GET", "__FILE__", true);\n xmlhttp.setRequestHeader("Cache-Control", "no-cache");\n xmlhttp.send(); \n }\n }\n \n if (!entrance___OBJ__) {\n entrance___OBJ__ += 1;\n refresh__OBJ__(); \n }\n </script>\n\n <p id="__OBJ__" style="font-size: 14px; background: #000000; padding: 10px; border: 3px; border-radius: 5px; color: white; ">\n </p>\n \n </font>\n <!--MD_END_FILTER-->\n <!--MD_FROM_FILE __FILE__.md -->\n \'\'\'.replace("__OBJ__", obj).replace("__FILE__", file)\n if return_html_string:\n return html_string\n display(HTML(html_string))\n\n \nclass TInteractiveLauncher:\n tmp_path = "./interactive_launcher_tmp"\n def __init__(self, cmd):\n try:\n os.mkdir(TInteractiveLauncher.tmp_path)\n except:\n pass\n name = str(random.randint(0, 1e18))\n self.inq_path = os.path.join(TInteractiveLauncher.tmp_path, name + ".inq")\n self.log_path = os.path.join(TInteractiveLauncher.tmp_path, name + ".log")\n \n os.mkfifo(self.inq_path)\n open(self.log_path, \'w\').close()\n open(self.log_path + ".md", \'w\').close()\n\n self.pid = os.fork()\n if self.pid == -1:\n print("Error")\n if self.pid == 0:\n exe_cands = glob.glob("../tools/launcher.py") + glob.glob("../../tools/launcher.py")\n assert(len(exe_cands) == 1)\n assert(os.execvp("python3", ["python3", exe_cands[0], "-l", self.log_path, "-i", self.inq_path, "-c", cmd]) == 0)\n self.inq_f = open(self.inq_path, "w")\n interactive_launcher_opened_set.add(self.pid)\n show_log_file(self.log_path)\n\n def write(self, s):\n s = s.encode()\n assert len(s) == os.write(self.inq_f.fileno(), s)\n \n def get_pid(self):\n n = 100\n for i in range(n):\n try:\n return int(re.findall(r"PID = (\\d+)", open(self.log_path).readline())[0])\n except:\n if i + 1 == n:\n raise\n time.sleep(0.1)\n \n def input_queue_path(self):\n return self.inq_path\n \n def close(self):\n self.inq_f.close()\n os.waitpid(self.pid, 0)\n os.remove(self.inq_path)\n # os.remove(self.log_path)\n self.inq_path = None\n self.log_path = None \n interactive_launcher_opened_set.remove(self.pid)\n self.pid = None\n \n @staticmethod\n def terminate_all():\n if "interactive_launcher_opened_set" not in globals():\n globals()["interactive_launcher_opened_set"] = set()\n global interactive_launcher_opened_set\n for pid in interactive_launcher_opened_set:\n print("Terminate pid=" + str(pid), file=sys.stderr)\n os.kill(pid, signal.SIGKILL)\n os.waitpid(pid, 0)\n interactive_launcher_opened_set = set()\n if os.path.exists(TInteractiveLauncher.tmp_path):\n shutil.rmtree(TInteractiveLauncher.tmp_path)\n \nTInteractiveLauncher.terminate_all()\n \nyandex_metrica_allowed = bool(globals().get("yandex_metrica_allowed", False))\nif yandex_metrica_allowed:\n display(HTML(\'\'\'<!-- YANDEX_METRICA_BEGIN -->\n <script type="text/javascript" >\n (function(m,e,t,r,i,k,a){m[i]=m[i]||function(){(m[i].a=m[i].a||[]).push(arguments)};\n m[i].l=1*new Date();k=e.createElement(t),a=e.getElementsByTagName(t)[0],k.async=1,k.src=r,a.parentNode.insertBefore(k,a)})\n (window, document, "script", "https://mc.yandex.ru/metrika/tag.js", "ym");\n\n ym(59260609, "init", {\n clickmap:true,\n trackLinks:true,\n accurateTrackBounce:true\n });\n </script>\n <noscript><div><img src="https://mc.yandex.ru/watch/59260609" style="position:absolute; left:-9999px;" alt="" /></div></noscript>\n <!-- YANDEX_METRICA_END -->\'\'\'))\n\ndef make_oneliner():\n html_text = \'("В этот ноутбук встроен код Яндекс Метрики для сбора статистики использований. Если вы не хотите, чтобы по вам собиралась статистика, исправьте: yandex_metrica_allowed = False" if yandex_metrica_allowed else "")\'\n html_text += \' + "<""!-- MAGICS_SETUP_PRINTING_END -->"\'\n return \'\'.join([\n \'# look at tools/set_up_magics.ipynb\\n\',\n \'yandex_metrica_allowed = True ; get_ipython().run_cell(%s);\' % repr(one_liner_str),\n \'display(HTML(%s))\' % html_text,\n \' #\'\'MAGICS_SETUP_END\'\n ])\n \n\n');display(HTML(("В этот ноутбук встроен код Яндекс Метрики для сбора статистики использований. Если вы не хотите, чтобы по вам собиралась статистика, исправьте: yandex_metrica_allowed = False" if yandex_metrica_allowed else "") + "<""!-- MAGICS_SETUP_PRINTING_END -->")) #MAGICS_SETUP_END
```
# Инструментирование в linux и mmap
## Инструментирование
Что это такое? Можно считать, что отладка. Получение информации о работающей программе и вмешательство в ее работу. Или не о программе, а о ядре системы. Известный вам пример - gdb.
Инструментарий для отладки
* Статический - прям в коде: счетчики, метрики (пример: санитайзеры)
* Оверхед
* Занимают место в коде
* Много может, так как имеет много возможностей
* Динамический - над кодом (примеры: gdb, strace, [eBPF](https://habr.com/ru/post/435142/))
* Динамически можно выбирать, на что смотреть
* ...
Разные подходы к инструментированию:
* Трейсинг - обрабатывать события. (пример: strace)
* Семплинг - условно смотреть состояние системы 100 раз в секунду. (пример: perf)
### Примеры использования для обнаружения проезда по памяти
```
%%cpp segfault.cpp
#include<stdio.h>
int main() {
int a[2];
printf("%d\n", a[100500]); // проезд по памяти
}
# компилируем и запускаем как обычно
!gcc segfault.cpp -o segfault.exe
!./segfault.exe
# компилируем с санитайзером и запускаем как обычно (семинарист рекомендует)
!gcc -fsanitize=address segfault.cpp -o segfault.exe
!./segfault.exe
# комбинируем санитайзер и gdb (это семинарист рекомендует, если вы хотите больше подробностей)
# по идее это должно находить больше косяков, чем вариант в следующей ячейке
!gcc -g -fsanitize=address segfault.cpp -o segfault.exe
!gdb -ex=r -batch --args ./segfault.exe
# компилируем с отладочной информацией и запускаем под gdb
!gcc -g segfault.cpp -o segfault.exe
!gdb -ex=r -batch --args ./segfault.exe
# компилируем как обычно и запускаем с valgrind
!gcc segfault.cpp -o segfault.exe
!valgrind --tool=memcheck ./segfault.exe 2>&1 | head -n 8 # берем только первые 8 строк выхлопа, а то там много
```
### Примеры использования для обнаружения утечек памяти
```
%%cpp memory_leak.cpp
#include<stdlib.h>
int main() {
malloc(16);
}
# компилируем как обычно и запускаем с valgrind
!gcc memory_leak.cpp -o memory_leak.exe
!valgrind --tool=memcheck --leak-check=full ./memory_leak.exe 2>&1
# компилируем с санитайзером и запускаем как обычно
!gcc -fsanitize=leak memory_leak.cpp -o memory_leak.exe
!./memory_leak.exe
```
### Примеры использования для отладки системных вызовов
```
%%cpp printing.cpp
#include<stdio.h>
int main() {
printf("Hello, world!");
}
# компилируем как обычно и запускаем с strace
!gcc printing.cpp -o printing.exe
!strace ./printing.exe > out.txt
!echo "Program output:"
!cat out.txt
%%asm printing_asm.S
.intel_syntax noprefix
.text
.global _start
_start:
mov eax, 4
mov ebx, 1
mov ecx, hello_world_ptr
mov edx, 14
int 0x80
mov eax, 1
mov ebx, 1
int 0x80
.data
hello_world:
.string "Hello, World!\n"
hello_world_ptr:
.long hello_world
# компилируем как обычно и запускаем с strace
!gcc -m32 -nostdlib printing_asm.S -o printing_asm.exe
!strace ./printing_asm.exe > out.txt
!echo "Program output:"
!cat out.txt
```
### Примеры использования для CPU профайлинга
```
%%cpp work_hard.cpp
int work_hard_1(int n) {
int ret = 0;
for (int i = 0; i < n; i++) ret ^= i;
return ret;
}
int work_hard_2(int n) {
int ret = 0;
for (int i = 0; i < n; i++) {
ret ^= work_hard_1(i * 3);
for (int j = 0; j < i * 2; ++j) {
ret ^= j;
}
}
return ret;
}
int main() {
return work_hard_2(10000);
}
# компилируем как обычно и запускаем с perf stat
!gcc work_hard.cpp -o work_hard.exe
!echo $PASSWORD | sudo -S perf stat ./work_hard.exe
# сравним с программой без stdlib
!echo $PASSWORD | sudo -S perf stat ./printing_asm.exe
```
Результат perf report можно отрисовывать в более красивом виде с помощью [flamegraph](https://github.com/brendangregg/FlameGraph)
```
# компилируем как обычно и запускаем с perf record и потом смотрим результаты
!gcc work_hard.cpp -o work_hard.exe
!echo $PASSWORD | sudo -S perf record ./work_hard.exe 2>&1 > perf.log
!echo $PASSWORD | sudo -S chmod 0666 perf.data
!perf report | cat
```
Есть еще вариант посчитать количество выполненных программой/функцией инструкций. Оно не всегда хорошо коррелирует со временем выполнения, зато является стабильным значением от запуска к запуску.
Здесь приводить способ не буду, если интересно, связывайтесь со мной отдельно :)
### Примеры использования для поиска UB (undefined behaviour)
```
%%cpp ub.c
int main(int argc, char** argv) {
return -argc << 31;
// return (argc * (int)((1ull << 31) - 1)) + 1;
}
# компилируем с санитайзером и запускаем как обычно (семинарист рекомендует)
# к сожалению у меня ни gcc, ни g++ не работают с -fsanitize=undefined
# а разбираться не хочется, поэтому clang
!clang -fsanitize=undefined ub.c -o ub.exe
!./ub.exe
```
## MMAP
(Копия [ридинга от Яковлева](https://github.com/victor-yacovlev/mipt-diht-caos/tree/master/practice/mmap))
```c
#include <sys/mman.h>
void *mmap(
void *addr, /* рекомендуемый адрес отображения */
size_t length, /* размер отображения */
int prot, /* аттрибуты доступа */
int flags, /* флаги совместного отображения */
int fd, /* файловый декскриптор файла */
off_t offset /* смещение относительно начала файла */
);
int munmap(void *addr, size_t length) /* освободить отображение */
```
Системный вызов `mmap` предназначен для создания в виртуальном адресном пространстве процесса доступной области по определенному адресу. Эта область может быть как связана с определенным файлом (ранее открытым), так и располагаться в оперативной памяти. Второй способ использования обычно реализуется в функциях `malloc`/`calloc`.
Память можно выделять только постранично. Для большинства архитектур размер одной страницы равен 4Кб, хотя процессоры архитектуры x86_64 поддерживают страницы большего размера: 2Мб и 1Гб.
В общем случае, никогда нельзя полагаться на то, что размер страницы равен 4096 байт. Его можно узнать с помощью команды `getconf` или функции `sysconf`:
```bash
# Bash
> getconf PAGE_SIZE
4096
```
```c
/* Си */
#include <unistd.h>
long page_size = sysconf(_SC_PAGE_SIZE);
```
Параметр `offset` (если используется файл) обязан быть кратным размеру страницы; параметр `length` - нет, но ядро системы округляет это значение до размера страницы в большую сторону. Параметр `addr` (рекомендуемый адрес) может быть равным `NULL`, - в этом случае ядро само назначает адрес в виртуальном адресном пространстве.
При использовании отображения на файл, параметр `length` имеет значение длины отображаемых данных; в случае, если размер файла меньше размера страницы, или отображается его последний небольшой фрагмент, то оставшаяся часть страницы заполняется нулями.
Страницы памяти могут флаги аттрибутов доступа:
* чтение `PROT_READ`;
* запись `PROT_WRITE`;
* выполнение `PROT_EXE`;
* ничего `PROT_NONE`.
В случае использования отображения на файл, он должен быть открыт на чтение или запись в соответствии с требуемыми аттрибутами доступа.
Флаги `mmap`:
* `MAP_FIXED` - требует, чтобы память была выделена по указаному в первом аргументе адресу; без этого флага ядро может выбрать адрес, наиболее близкий к указанному.
* `MAP_ANONYMOUS` - выделить страницы в оперативной памяти, а не связать с файлом.
* `MAP_SHARED` - выделить страницы, разделяемые с другими процессами; в случае с отображением на файл - синхронизировать изменения так, чтобы они были доступны другим процессам.
* `MAP_PRIVATE` - в противоположность `MAP_SHARED`, не делать отображение доступным другим процессам. В случае отображения на файл, он доступен для чтения, а созданные процессом изменения, в файл не сохраняются.
Зачем mmap?
* Дешевая переподгрузка файла-ресурса (если файл не изменился, то не будет выделяться лишняя память).
* ...
Пример с mmap (и с ftruncate). Простенькая программа, делающая циклический сдвиг (как цифры) второго символа в файле.
**Тут была некритичная ошибка так как по размеру страницы выравнивалась length, а должны быть выровнены только addr и offset**
```
%%cpp mmap_example.c
%run gcc mmap_example.c -o mmap_example.exe
%run echo "000" > buf.txt && ./mmap_example.exe && cat buf.txt
%run echo "79" > buf.txt && ./mmap_example.exe && cat buf.txt
%run echo "xxx" > buf.txt && ./mmap_example.exe && cat buf.txt
%run echo -n "S" > buf.txt && ./mmap_example.exe && cat buf.txt
#include <stdio.h>
#include <stdlib.h>
#include <unistd.h>
#include <sys/types.h>
#include <sys/stat.h>
#include <sys/mman.h>
#include <fcntl.h>
#include <assert.h>
int get_page_size() {
static int page_size = 0;
return page_size = page_size ?: sysconf(_SC_PAGE_SIZE);
}
int upper_round_to_page_size(int sz) {
return (sz + get_page_size() - 1) / get_page_size() * get_page_size();
}
int main() {
printf("page size = %d\n", get_page_size());
int fd = open("buf.txt", O_RDWR);
struct stat s;
assert(fstat(fd, &s) == 0);
printf("file size = %d\n", (int)s.st_size);
int old_st_size = s.st_size;
if (s.st_size < 2) {
const int new_size = 10;
assert(ftruncate(fd, new_size) == 0); // изменяем размер файла
assert(fstat(fd, &s) == 0);
printf("new file size = %d\n", (int)s.st_size);
}
void* mapped = mmap(
/* desired addr, addr = */ NULL,
/* length = */ s.st_size,
/* access attributes, prot = */ PROT_READ | PROT_WRITE,
/* flags = */ MAP_SHARED,
/* fd = */ fd,
/* offset in file, offset = */ 0
);
assert(mapped != MAP_FAILED);
char* buf = mapped;
if (old_st_size != s.st_size) {
for (int j = old_st_size; j < s.st_size; ++j) {
buf[j] = '_';
}
}
buf[1] = ('0' <= buf[1] && buf[1] <= '9') ? ((buf[1] - '0' + 1) % 10 + '0') : '0';
assert(munmap(
/* mapped addr, addr = */ mapped,
/* length = */ s.st_size
) == 0);
assert(close(fd) == 0); // Не забываем закрывать файл
return 0;
}
```
Еще один пример по мотивам advanced-1. Интерпретируем байты как функцию и выполняем.
```
%%cpp func.c
%run gcc -g -fPIC func.c -c -o func.o
%run objdump -F -d func.o | grep my_func
int my_func(int a, int b) {
return a + b + 1;
}
```
Замечаем, что `File Offset: 0x40`. То есть в объектном файле функция лежит начиная с `0x40` позиции.
```
%%cpp mmap_exec_example.c
%run gcc -g mmap_exec_example.c -o mmap_exec_example.exe
%run ./mmap_exec_example.exe
#include <stdio.h>
#include <stdlib.h>
#include <unistd.h>
#include <sys/types.h>
#include <sys/stat.h>
#include <sys/mman.h>
#include <fcntl.h>
#include <assert.h>
int main() {
int fd = open("func.o", O_RDWR);
struct stat s;
assert(fstat(fd, &s) == 0);
printf("file size = %d\n", (int)s.st_size);
void* mapped = mmap(
/* desired addr, addr = */ NULL,
/* length = */ s.st_size,
/* access attributes, prot = */ PROT_READ | PROT_EXEC | PROT_WRITE, // обратите внимание на PROT_EXEC
/* flags = */ MAP_SHARED,
/* fd = */ fd,
/* offset in file, offset = */ 0
);
assert(close(fd) == 0); // Не забываем закрывать файл (при закрытии регион памяти остается доступным)
if (mapped == MAP_FAILED) {
perror("Can't mmap");
return -1;
}
int (*func)(int, int) = (void*)((char*)mapped + 0x40); // 0x40 - тот самый оффсет
printf("func(1, 1) = %d\n", func(1, 1));
printf("func(10, 100) = %d\n", func(10, 100));
printf("func(40, 5000) = %d\n", func(40, 5000));
assert(munmap(
/* mapped addr, addr = */ mapped,
/* length = */ s.st_size
) == 0);
return 0;
}
```
# Рекомендации по контесту inf08
* inf08-1
```c
char* buf = ...;
...
struct Item* item = (void*)(buf + offset);
```
* inf08-2 <br>
mmap, ftruncate, snprintf
* advanced-3 <br>
Можно попробовать поддерживать список свободных сегментов памяти.
При аллокации выбирать наименьший достаточного объема и отрезать от него.
При деаллокации добавлять новый свободный сегмент и склеивать смежные.
| github_jupyter |
# **KModes Algorithm to predict Shelf-Life Clusters**
> Following Clusters are classified into :
1. Poor - 2
2. Average - 1
3. Good - 0
**Importing all the crucial libraries**
```
import pandas as pd
import numpy as np
!pip install kmodes
from kmodes.kmodes import KModes
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('darkgrid')
%matplotlib inline
! git clone https://github.com/vaishakhshetty/STEER-ML.git
ls
```
## Data Preprocessing
```
data = pd.read_csv('/content/STEER-ML/data/V5_Dummy-Data.csv')
data.head(5)
data = data.drop(['Unnamed: 0','Device_ID'], axis = 1)
data.head()
# join two attribute columns - Device Brand and Device Model >
data['Device_list'] = data[['Device_brand', 'Device_model']].values.tolist()
data['Device'] = data['Device_list'].apply(' '.join)
data = data.drop(['Device_list', 'Device_brand', 'Device_model'], axis = 1)
data.head(5)
data['Usage_range'] = pd.cut(data['Usage(hrs)'], [0, 3, 5, 7, 11, 13, 20], labels=['<3', '3-5', '5-7', '7-11', '11-13', '13>'])
data = data.drop('Usage(hrs)',axis = 1)
data.head(5)
data['Age_days'] = pd.cut(data['Age(days)'], [0, 300, 500, 700, 1100, 1300, 2000, 2500, 3000, 3500, 4000, 4500, 5000, 10000], labels=['<300', '300-500', '500-700', '700-1100', '1100-1300', '1300-1700', '1700-2000', '2000-2500', '2500-3000', '3000-3500', '4000-4500', '4500-5000', '5000>'])
data = data.drop('Age(days)',axis = 1)
data.head(5)
#Adding a checkpoint
data_copy = data.copy()
```
## Exploratory Data Analysis
```
data.head(10)
data.tail(10)
data.info()
```
## Modelling
```
z = data['Device']
z
data = data.drop(['Device'], axis = 1)
from sklearn import preprocessing
le = preprocessing.LabelEncoder()
data = data.apply(le.fit_transform)
data.head()
data = data.drop(['Manufacture_Date', 'Allocation_Date'], axis = 1)
data.head(2)
data = data.drop(['Assessment_Date'], axis = 1)
data.head(2)
# Elbow curve to find optimal K
%%time
cost = []
K = range(1,5)
for num_clusters in list(K):
kmode = KModes(n_clusters=num_clusters, init = "random", n_init = 10, verbose=1, random_state=42)
kmode.fit_predict(data)
cost.append(kmode.cost_)
plt.plot(K, cost, 'bx-')
plt.xlabel('No. of clusters')
plt.ylabel('Cost')
plt.grid(True)
plt.title('Elbow Method For Optimal k')
plt.show()
# Building the model with 3 clusters
kmode = KModes(n_clusters=3, init = "random", n_init = 10, verbose=1, random_state=42)
clusters = kmode.fit_predict(data)
clusters
data.insert(0, "Cluster", clusters, True)
data.head()
data_copy.info()
data = data_copy.reset_index()
clustersDf = pd.DataFrame(clusters)
clustersDf.columns = ['cluster_predicted']
combinedDf = pd.concat([data, clustersDf], axis = 1).reset_index()
combinedDf = combinedDf.drop(['index', 'level_0'], axis = 1)
combinedDf.info()
combinedDf.head(10)
all_features = combinedDf[['device_in_use', 'water_damage', 'device_responsive',
'battery_health', 'Latency(ms)', 'Usage_range', 'Age_days']]
for col in all_features:
plt.subplots(figsize = (15,5))
sns.countplot(x='cluster_predicted',hue=col, data = combinedDf)
plt.grid(True)
plt.show()
#end
```
| github_jupyter |
```
from time import sleep
import matplotlib.pyplot as plt
import sys
import os
from scipy.interpolate import spline
import numpy as np
sys.path.append('rrr')
import decoy_mnist
from rfft.multilayer_perceptron import *
from rfft.hypothesis import Hypothesis
import autograd.numpy as np
import pdb
import pickle
Xr, X, y, E, Xtr, Xt, yt, Et = decoy_mnist.generate_dataset()
indices, hypothesis = decoy_mnist.load_hypothesis(X)
hypothesis.weight = 200000
accuracies = []
def score_model(mlp):
print('Train: {0}, Test: {1}'.format(mlp.score(X, y), mlp.score(Xt, yt)))
f = MultilayerPerceptron()
annotated = np.array(range(0, len(indices), len(indices)//10))
accuracies = []
for x in annotated:
print("Taking %d annotated samples" %x)
f.fit(X, y , hypothesis = hypothesis, num_epochs = 10, always_include = indices[:x], show_progress=False)
score_model(f)
accuracies.append(f.score(Xt, yt))
xnew = np.linspace(annotated.min(),annotated.max(), 300)
power_smooth = spline(annotated,accuracies,xnew)
plt.plot(xnew,power_smooth, label = 'Test Accucarcy')
plt.xlabel('Number of annotations')
plt.ylabel('Accuracy')
plt.show()
```
```
print('Training f0')
if os.path.exists('models/1.pkl'):
f0 = pickle.load(open('models/1.pkl', 'rb'))
else:
f0 = MultilayerPerceptron()
f0.fit(X, y, hypothesis=hypothesis,
num_epochs=16, always_include=indices)
pickle.dump(f0, open('models/1.pkl', 'wb'))
score_model(f0)
l2_grads = np.array(range(0, 10000, 1000))
accuracies = []
for l2 in l2_grads:
print('l2 =', l2)
f1 = MultilayerPerceptron(l2_grads=l2)
print('Training f1')
M0 = f0.largest_gradient_mask(X)
f1.fit(X, y, hypothesis=Hypothesis(M0), num_epochs=10)
score_model(f1)
accuracies.append(f1.score(Xt, yt))
```
### Accuracy vs L2 norm
```
xnew = np.linspace(l2_grads.min(), l2_grads.max(), 500)
power_smooth = spline(l2_grads, accuracies, xnew)
plt.plot(xnew, power_smooth, label='Test Accucarcy')
plt.xlabel('L2 grad')
plt.ylabel('Accuracy')
plt.show()
weights = np.array(range(0, 100000, 10000))
accuracies = []
l2 = 1000
for weight in weights:
hypothesis.weight = weight
print('l2 =', l2)
f1 = MultilayerPerceptron(l2_grads=l2)
print('Training f1')
M0 = f0.largest_gradient_mask(X)
f1.fit(X, y, hypothesis=Hypothesis(M0), num_epochs=10)
score_model(f1)
accuracies.append(f1.score(Xt, yt))
```
### Accuracy vs Hypothesis weight
```
xnew = np.linspace(weights.min(),weights.max(),300)
power_smooth = spline(weights,accuracies,xnew)
plt.plot(xnew,power_smooth, label = 'Test Accucarcy')
plt.xlabel('Hypothesis.weight')
plt.ylabel('Accuracy')
plt.show()
```
| github_jupyter |
```
import getpass
import os
if( os.system('klist | grep Default | grep mbonanom@CERN.CH') ):
os.system('echo %s | kinit mbonanom' % getpass.getpass() )
v10_prefix = 'root://eoscms.cern.ch//eos/cms/store/group/dpg_hgcal/tb_hgcal/2018/cern_h2_october/offline_analysis/ntuples/v10/'
v11_prefix = 'root://eoscms.cern.ch//eos/cms/store/group/dpg_hgcal/tb_hgcal/2018/cern_h2_october/offline_analysis/ntuples/v11/'
v9_prefix = 'root://eoscms.cern.ch//eos/cms/store/group/dpg_hgcal/tb_hgcal/2018/cern_h2_october/offline_analysis/ntuples/v9/'
v8_prefix = 'root://eoscms.cern.ch//eos/cms/store/group/dpg_hgcal/tb_hgcal/2018/cern_h2_october/offline_analysis/ntuples/v8/'
import pandas as pd
import numpy as np
from scipy.stats import norm
import matplotlib.mlab as mlab
from root_pandas import read_root
import itertools
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
gamma = 0.3
#from mpl_toolkits.mplot3d import Axes3D
from matplotlib.colors import LogNorm
mpl.rcParams['image.cmap'] = 'viridis'
%matplotlib inline
from scipy.optimize import curve_fit
from scipy.stats import binned_statistic#, binned_statistic_2d
from scipy.special import erf
from scipy.stats import iqr
from collections import defaultdict
import os.path
font = {'family' : 'DejaVu Sans',
'weight' : 'normal',
'size' : 16}
mpl.rc('font', **font)
def line(x, m, q):
return m*x + q
def E_res(x, p0, p1, p2):
return np.sqrt((p0/np.sqrt(x))**2 + (p1/x)**2 + (p2)**2)
def err_prop(sigma, E, err_sigma, err_E):
n = sigma/E
dval1 = err_sigma/sigma
dval2 = err_E/E
return n*np.sqrt((dval1)**2 + (dval2)**2)
def gaussian(x, amplitude, mean, standard_deviation):
return amplitude * np.exp( - 0.5*((x - mean) / standard_deviation) ** 2)
def gausExp(x, xb, sig, k, N):
if k < 0:
k = -k
total = 0.*x
total += ((x-xb)/sig > -k) * N * np.exp(- (x-xb)**2/(2.*sig**2))
total += ((x-xb)/sig <= -k) * N * np.exp(k**2/2. + k*((x-xb)/sig))
return total
from scipy.special import erf
def crystal_ball(x, a, n, xb, sig):
x = x+0j
if a < 0:
a = -a
if n < 0:
n = -n
aa = abs(a)
A = (n/aa)**n * np.exp(- aa**2 / 2)
B = n/aa - aa
C = n/aa*1/(n-1)*np.exp(- aa**2 / 2)
D = np.sqrt(np.pi/2)*(1+erf(aa/np.sqrt(2)))
N = 1./(sig*(C+D))
total = 0.*x
total += ((x-xb)/sig > -a) * N * np.exp(- (x-xb)**2/(2.*sig**2))
total += ((x-xb)/sig <= -a) * N * A * (B - (x-xb)/sig)**(-n)
try:
return total.real
except:
return total
return total
def CBFit(histogram, energy):
binning = 100
if (energy > 100):
binning = 200
upLim = 8
dLim = 12
# n, bins, patch = plt.hist(histogram, density=1, color = 'cyan', histtype='step', linewidth=2, bins=binning)
n, bins = np.histogram(histogram, density=1, bins=binning)
bins = 0.5*(bins[1:]+bins[:-1])
popt, pcov = curve_fit(crystal_ball, bins, n)
h_std = np.sqrt(Noise*Noise + Sampl*Sampl/energy + Linear*Linear/(energy**2))
sel = (bins > histogram.median()*0.6) & (bins < histogram.median()*1.2)
xmed = bins[np.argmax(n)]
popt, pcov = curve_fit(crystal_ball, bins[sel], n[sel], p0=[ 1.1, 3, xmed, h_std])
E = popt[2]
sig = popt[3]
errors = np.sqrt(np.diag(pcov))
err_E = errors[2]
err_sig = errors[3]
resolution = sig/E
err_res = err_prop(sig, E, err_sig, err_E)
xmax = bins[np.argmax(n)]
xdraw = np.linspace(0, histogram.max(), 1000)
plt.plot(xdraw, crystal_ball(xdraw, *popt), linewidth=2, color='cyan', label = r'$\sigma$/E (CB) = %.3f' %resolution)
# plt.xlim(xmax - dLim*sig, xmax + upLim*sig)
plt.legend()#title="CB fit")
return resolution, E, err_res
Sampl = 2.198e-01
Noise = 8.210e-03
Linear = 3e-6
def GausExpFit(histogram, energy):
binning = 100
if (energy > 100):
binning = 200
upLim = 8
dLim = 12
# n, bins, patch = plt.hist(histogram, density=1, color = 'red', alpha=0.6, bins=binning)
n, bins = np.histogram(histogram, density = 1, bins=binning)
bins = 0.5*(bins[1:]+bins[:-1])
xmax = bins[np.argmax(n)]
ymax = n.max()
sel = (bins > histogram.median()*0.5) & (bins < histogram.median()*1.5)
popt, pcov = curve_fit(gausExp, bins[sel], n[sel], p0 = [xmax, 0.054, 0.5, ymax])
popt, pcov = curve_fit(gausExp, bins[sel], n[sel], p0 = popt)
E = popt[0]
sig = popt[1]
errors = np.sqrt(np.diag(pcov))
err_E = errors[0]
err_sig = errors[1]
resolution = sig/E
err_res = err_prop(sig, E, err_sig, err_E)
xmax = bins[np.argmax(n)]
xdraw = np.linspace(0, histogram.max(), 1000)
plt.plot(xdraw, gausExp(xdraw, *popt), linewidth=2, color = 'red', label = r'$\sigma$/E (GE) = %.3f' %resolution)
# plt.xlim(xmax - dLim*sig, xmax + upLim*sig)
plt.legend()#title="gausExp fit")
return resolution, E, err_res
Sampl = 2.198e-01
Noise = 8.210e-03
Linear = 3e-6
def repeatedGausFit(histogram, energy, rangeInSigmaLeft = 1, rangeInSigmaRight = 2.5):
binning = 300
n, bins, patch = plt.hist(histogram, density = 1, alpha =0.6, bins=binning)
n, bins = np.histogram(histogram, density = 1, bins=binning)
bins = 0.5*(bins[1:]+bins[:-1])
h_med = bins[np.argmax(n)]
h_std = np.sqrt(Noise*Noise + Sampl*Sampl/energy + Linear*Linear/(energy**2))
s_fit = (bins > h_med - rangeInSigmaLeft*h_std)& (bins < (h_med + rangeInSigmaRight*h_std))
ymaximum = n.max()
plt.xlim(histogram.median()*0.6, histogram.median()*1.5)
popt, pcov = curve_fit(gaussian, bins[s_fit], n[s_fit], p0=[ymaximum, h_med, h_std])
# popt, pcov = curve_fit(lambda x, mean, st_dev: gaussian(x, ymaximum, mean, st_dev), bins[s_fit], n[s_fit],
# p0=[h_med, h_std])
# print popt
mu_tmp = popt[1]
std_tmp = popt[2]
ymaximum_tmp = popt[0]
s_fit = (bins > mu_tmp- rangeInSigmaLeft*std_tmp) & (bins < mu_tmp + rangeInSigmaRight*std_tmp)
popt, pcov = curve_fit(gaussian, bins[s_fit], n[s_fit], p0=[ymaximum_tmp, mu_tmp, std_tmp])
# popt, pcov = curve_fit(lambda x, mean, st_dev: gaussian(x, ymaximum, mean, st_dev), bins[s_fit], n[s_fit],
# p0=[mu_tmp, std_tmp])
# print popt
e_reco = popt[1]
sigma = popt[2]
err = np.sqrt(np.diag(pcov))
E_err = err[1]
sigma_err = err[2]
resolution = sigma/e_reco
res = ((n[s_fit] - gaussian(bins[s_fit], *popt))/gaussian(bins[s_fit], *popt))**2
chi2 = np.sum(res)/(len(bins[s_fit])-len(popt))
xdraw = np.linspace(popt[1]-rangeInSigmaLeft*popt[2],popt[1]+rangeInSigmaRight*popt[2], 1000)
plt.plot(xdraw, gaussian(xdraw, popt[0], popt[1], popt[2]), linewidth=2, label=r'$\sigma/E (Gaus)$ = %.3f' %resolution)
return e_reco, sigma, E_err, sigma_err, popt[0], chi2
def is_electron(df):
if df.pdgID[0] == 11: return True
else: return False
w = [10.199, 9.851, 9.851, 9.851, 9.851, 9.851, 9.851, 9.851, 9.851, 9.851, 9.851, 9.851,
9.851, 9.851, 9.851, 9.851, 9.851, 9.851, 9.851, 9.851, 11.360, 11.360, 11.360, 11.360,
10.995, 10.995, 11.153, 36.139, 57.496, 57.654, 57.654, 57.654, 56.884, 38.620, 39.390,
57.654, 58.083, 57.757, 56.574, 37.755]
w_tot = sum(w)
X0 = [0.943, 1.923, 2.858, 3.838, 4.773, 5.753, 6.688, 7.668, 8.603, 9.583,
10.518, 11.498, 12.433, 13.413, 14.348, 15.328, 16.263, 17.243, 18.178,
19.157, 20.092, 21.240, 22.174, 23.322, 24.256, 25.505, 26.440,
27.725, 30.566, 33.501, 36.437, 39.373, 42.308, 45.159, 46.275, 49.211,
52.146, 55.276, 58.322, 61.175]
weights = dict()
lay_X0 = dict()
su = 0
for i in range(len(w)):
weights[i+1] = (w[i]/w_tot) #dEdx are averaged in MeV/MIP --> convert to GeV
lay_X0[i+1] = (X0[i])
simul_reso = [0.05252533050053361, 0.04246531378576422, 0.03316893269421308, 0.026026766200077246, 0.023539708490593735,
0.020451829612688918, 0.01862616723089308, 0.017134081339427978, 0.016346269539945477]
E_MC = [0.17205952777473252, 0.25891686515687895, 0.4322641161581555, 0.6915440898971871, 0.8641409222828782, 1.2897407103365943,
1.7044081463354082, 2.103598722522489, 2.4799942424055996]
#1 MIP = 84.9 keV for all layers (see: https://gitlab.cern.ch/cms-hgcal-tb/TestBeam/wikis/samples/October2018-CERN)
MIPtoGeV = 84.9*1e-6
```
<h1> STABILITY FOR SAME ENERGY RUNS </h1>
```
_20GeV = range(436, 456)
_30GeV = range(594, 608)
_50GeV = range(456, 466) + range(608, 620)
_80GeV = range(466, 476) + range(655, 664)
_100GeV = range(477, 492)
_120GeV = range(361, 372) + range(382, 384) + range(620, 645)
_150GeV = range(303, 319) + range(337, 361) + range(493, 510)
_200GeV = range(372, 382) + range(664, 677)
_250GeV = range(384, 405) + range(645, 655)
_300GeV = range(405, 435)
branches = [u'event', 'beamEnergy', u'rechit_module',u'rechit_chip', # 'pdgID', u'rechit_chip', u'rechit_channel',
u'rechit_layer', u'rechit_energy']
rh_branches = [branch for branch in branches if 'rechit' in branch]
energies = [20, 30, 50, 80, 100, 120, 150, 200, 250, 300]
medians = defaultdict(list)
medians_w = defaultdict(list)
run_id = defaultdict(list)
%%time
for i in range(len(energies)):
energy = energies[i]
runs = '_%iGeV' %energy
runs = eval(runs)
for j in range(len(runs)):
run = runs[j]
try:
fname = v11_prefix + 'ntuple_%i.root' %run
df = read_root(fname,key='rechitntupler/hits',columns = branches, flatten = rh_branches)
df['rechit_energy_w'] = df['rechit_energy']*MIPtoGeV
run_id[energy].append(run)
sel_en = (df.rechit_energy>0.5) #& (df.rechit_chip != 3) #& (df.rechit_layer <= 28)
sel_en &= ~((df.rechit_module == 78) & (df.rechit_chip == 0))
#sel_en &= ~((df.rechit_chip == 3) & (df.rechit_channel == 22))
df_sel = df[sel_en]
sum_en = df_sel.groupby('event')['rechit_energy'].sum()
medians[energy].append(sum_en.median())
sum_en = df_sel.groupby('event')['rechit_energy_w'].sum()
medians_w[energy].append(sum_en.median()/energy)
except IOError:
continue
plt.figure(figsize=(10,8))
for i in range(len(energies)):
energy = energies[i]
# if energy > 120:
plt.plot(run_id[energy], medians_w[energy], 'o', label = '%i GeV' %energy)
plt.legend()
plt.ylim(0.007, 0.009)
plt.grid()
plt.ylabel(r'Median E/$E_{Beam}$')
plt.xlabel('run number')
plt.axvline(x=589, color='black', linestyle = '-.')
energy = 200
run_bad = 376
branches = [u'event', 'beamEnergy', u'rechit_module', u'rechit_chip', u'rechit_channel',
u'rechit_layer', u'rechit_energy']
rh_branches = [branch for branch in branches if 'rechit' in branch]
fname = v9_prefix + 'ntuple_%i.root' %run_bad
df = read_root(fname,key='rechitntupler/hits',columns = branches, flatten = rh_branches)
df['rechit_energy_w'] = df['rechit_energy']*(df['rechit_layer'].map(weights))
sel_en = (df.rechit_energy > 0.5) #& (df.rechit_layer > 28)
sel_en &= ~((df.rechit_chip == 3) & (df.rechit_channel == 22))
df_sel = df[sel_en]
sum_en_bad = df_sel.groupby('event')['rechit_energy_w'].sum()
sel_en = (df.rechit_energy > 0.5) & (df.rechit_layer > 28)
df_sel = df[sel_en]
sum_en_FH_bad = df_sel.groupby('event')['rechit_energy_w'].sum()
sel_en = (df.rechit_energy > 0.5) & (df.rechit_layer <= 28)
df_sel = df[sel_en]
sum_en_EE_bad = df_sel.groupby('event')['rechit_energy_w'].sum()
plt.figure(figsize=(10,8))
plt.title('Run %i. %i GeV' %(run_bad, energy))
sel = sum_en_FH_bad<100
r = plt.hist2d(sum_en_EE_bad[sel], sum_en_FH_bad[sel], 300, cmap='viridis', norm=LogNorm())#mcolors.PowerNorm(gamma))
plt.xlabel('EE energy [GeV]')
plt.ylabel('FH energy [GeV]')
run_good = 672
fname = v9_prefix + 'ntuple_%i.root' %run_good
df = read_root(fname,key='rechitntupler/hits',columns = branches, flatten = rh_branches)
df['rechit_energy_w'] = df['rechit_energy']*(df['rechit_layer'].map(weights))
sel_en = (df.rechit_energy > 0.5) #& (df.rechit_layer > 28)
sel_en &= ~((df.rechit_chip == 3) & (df.rechit_channel == 22))
df_sel = df[sel_en]
sum_en_good = df_sel.groupby('event')['rechit_energy_w'].sum()
sel_en = (df.rechit_energy > 0.5) & (df.rechit_layer > 28)
df_sel = df[sel_en]
sum_en_FH_good = df_sel.groupby('event')['rechit_energy_w'].sum()
sel_en = (df.rechit_energy > 0.5) & (df.rechit_layer <= 28)
df_sel = df[sel_en]
sum_en_EE_good = df_sel.groupby('event')['rechit_energy_w'].sum()
plt.figure(figsize=(10,8))
plt.title('Run %i. %i GeV' %(run_good, energy))
sel = sum_en_FH_good<100
r = plt.hist2d(sum_en_EE_good[sel], sum_en_FH_good[sel], 300, cmap='viridis', norm=LogNorm())#mcolors.PowerNorm(gamma))
plt.xlabel('EE energy [GeV]')
plt.ylabel('FH energy [GeV]')
plt.figure(figsize=(20,10))
plt.subplot(121)
plt.title('All Layers')
r= plt.hist(sum_en_good, normed=1, bins=500, label='Run %i' %run_good)
r= plt.hist(sum_en_bad, normed=1, bins=500, alpha=0.7, label='Run %i' %run_bad)
plt.xlabel('E dE/dX [GeV]')
plt.legend()
plt.subplot(122)
plt.title('EE layers')
r= plt.hist(sum_en_EE_good, normed=1, bins=500, label='Run %i' %run_good)
r= plt.hist(sum_en_EE_bad, normed=1, bins=500, alpha=0.7, label='Run %i' %run_bad)
plt.xlabel('E dE/dX [GeV]')
plt.legend()
plt.figure(figsize=(20,10))
plt.subplot(121)
plt.title('RUN %i' %run_bad)
# r= plt.hist(sum_en_bad, normed=1, bins=500, label='Run %i, all layers' %run_bad)
# r= plt.hist(sum_en_EE_bad, normed=1, bins=500, alpha=0.7, label='Run %i, EE' %run_bad)
r=plt.hist2d(sum_en_EE_bad, sum_en_bad, 300, cmap='viridis', norm=LogNorm())#mcolors.PowerNorm(gamma))
plt.xlabel('E in EE, dE/dX [GeV]')
plt.ylabel('E tot, dE/dX [GeV]')
# plt.legend()
plt.subplot(122)
plt.title('RUN %i' %run_good)
# r= plt.hist(sum_en_good, normed=1, bins=500, label='Run %i, all layers' %run_good)
# r= plt.hist(sum_en_EE_good, normed=1, bins=500, alpha=0.7, label='Run %i, EE' %run_good)
r=plt.hist2d(sum_en_EE_good, sum_en_good, 300, cmap='viridis', norm=LogNorm())#mcolors.PowerNorm(gamma))
plt.xlabel('E in EE, dE/dX [GeV]')
plt.ylabel('E tot, dE/dX [GeV]')
# plt.legend()
```
<h1> ENERGY RESOLUTION </h1>
```
# energies = [20, 30, 50, 80, 100, 120, 150, 200, 250, 300]
energies = [20, 30, 49.99, 79.93, 99.83, 119.65, 149.14, 197.32, 243.61, 287.18]
runs = [455, 607, 458, 469, 490, 641, 494, 664, 653, 435]
# runs = [455, 607, 457, 469, 477, 382, 494, 664, 653, 405]
branches = [u'event', 'beamEnergy', 'pdgID', u'rechit_chip', u'rechit_channel', 'rechit_module'
u'rechit_layer', u'rechit_energy', u'rechit_energy_noHG', u'rechit_x', u'rechit_y']
dwc_branches = [u'event', u'ntracks']
rh_branches = [branch for branch in branches if 'rechit' in branch]
def add_dwc(run, location, version):
branches = [u'event', u'rechit_chip', u'rechit_channel', 'rechit_module',
u'rechit_energy', u'rechit_x', u'rechit_y']
dwc_branches = [u'event', u'ntracks', u'b_x', u'b_y']
rh_branches = [branch for branch in branches if 'rechit' in branch]
fname = location + 'ntuple_%i.root' %run
df = read_root(fname,key='rechitntupler/hits',columns = branches, flatten = rh_branches)
if (version == 9):
df['rechit_energy_w'] = df['rechit_energy']*MIPtoGeV*1.14
if (version == 8):
df['rechit_energy_w'] = df['rechit_energy']*MIPtoGeV*1.105
if ((version == 10) | (version == 11)):
df['rechit_energy_w'] = df['rechit_energy']*MIPtoGeV*1.06
dwc = read_root(fname, key='trackimpactntupler/impactPoints', columns=dwc_branches)
df.set_index('event', inplace=True)
dwc.set_index('event', inplace=True)
df_dwc = df.add(dwc, fill_value=0)
return df_dwc
def do_Esum(df, version):
sel_en = (df.rechit_energy>0.5)
sel_en &= ~((df.rechit_chip == 3) & (df.rechit_channel == 22))
if (version == 11):
sel_en &= ~((df.rechit_chip == 0) & (df.rechit_module == 78))
sel_en &= (df.ntracks == 1)
# In the analysis was used the cut above, but it was shown that does not select the central beam spot
# When v10 ready try the comparison using the cut below: (should select better the beam spot)
if (version == 8):#((version == 9) | (version == 8)):
# sel_en &= ((df.b_x > -2) & (df.b_x < 2))
# sel_en &= ((df.b_y > -1) & (df.b_y < 2))
# sel_en &= ((df.b_x > -0.5) & (df.b_x < 3.5))
# sel_en &= ((df.b_y > -1.5) & (df.b_y < 2))
sel_en &= ((df.b_x > 1) & (df.b_x < 2.5))
sel_en &= ((df.b_y > -1) & (df.b_y < 1.5))
if ((version == 10) | (version == 11)):
# sel_en = ((df.b_x > -4.5) & (df.b_x < -1.5))
# sel_en &= ((df.b_y > -0.5) & (df.b_y < 3))
sel_en &= ((df.b_x > -2.5) & (df.b_x < -0.5))
sel_en &= ((df.b_y > 0) & (df.b_y < 2.5))
df_sel = df[sel_en]
sum_en = df_sel.groupby('event')['rechit_energy_w'].sum()
return sum_en
def do_Esum_Noise(df):
sel_en = (df.rechit_energy>0.5) & (df.rechit_noise_flag == False)
# sel_en &= ~((df.rechit_chip == 3) & (df.rechit_channel == 22))
sel_en &= (df.ntracks == 1)
sel_en &= ((df.b_x > -2) & (df.b_x < 2))
sel_en &= ((df.b_y > -1) & (df.b_y < 2))
df_sel = df[sel_en]
sum_en = df_sel.groupby('event')['rechit_energy_w'].sum()
return sum_en
%%time
df_v10 = []
for run in runs:
df_v10.append(add_dwc(run, v10_prefix, 10))
%%time
df_v11 = []
for run in runs:
df_v11.append(add_dwc(run, v11_prefix, 11))
%%time
df_v8 = []
for run in runs:
df_v8.append(add_dwc(run, v8_prefix, 8))
%%time
sum_en_v10 = []
for df in df_v10:
sum_en_v10.append(do_Esum(df, 10))
%%time
sum_en_v11 = []
for df in df_v11:
sum_en_v11.append(do_Esum(df, 11))
%%time
sum_en_v8 = []
for df in df_v8:
sum_en_v8.append(do_Esum(df, 8))
fig, (ax1,ax2) = plt.subplots(1, 2, figsize=(20, 10))
# sel_en = ((df_v8[9].b_x > -2) & (df_v8[9].b_x < 2))
# sel_en &= ((df_v8[9].b_y > -1) & (df_v8[9].b_y < 2))
sel_en = ((df_v8[9].b_x > -0.5) & (df_v8[9].b_x < 3.5))
sel_en &= ((df_v8[9].b_y > -1.5) & (df_v8[9].b_y < 2))
r=ax1.hist2d(df_v8[9][sel_en].b_x, df_v8[9][sel_en].b_y, 50, norm=LogNorm())
ax1.set_title('RUN %i. %.2f GeV. v8, DWC cuts' %(runs[9], energies[9]))
ax1.set_xlabel('b_x DWC')
ax1.set_ylabel('b_y DWC')
plt.colorbar(r[3], ax=ax1)
for i, energy in zip(range(len(runs)), energies):
h = ax2.hist(sum_en_v8[i], density = 1, bins=200, label = '%.2f GeV' %energy)
ax2.set_title('Reco. E sums')
ax2.set_xlabel('visible E [GeV]')
plt.legend()
plt.xlim(0., 3.)
fig, (ax1,ax2) = plt.subplots(1, 2, figsize=(20, 10))
sel_en = ((df_v11[9].b_x > -4) & (df_v11[9].b_x < 1))
sel_en &= ((df_v11[9].b_y > -1) & (df_v11[9].b_y < 3))
# sel_en = ((df_v11[9].b_x > -4.5) & (df_v11[9].b_x < -1.5))
# sel_en &= ((df_v11[9].b_y > -0.5) & (df_v11[9].b_y < 2.5))
r=ax1.hist2d(df_v11[9][sel_en].b_x, df_v11[9][sel_en].b_y, 50, norm=LogNorm())
ax1.axvline(x = -2.5, color = 'red')
ax1.axvline(x = -0.5, color = 'red')
ax1.axhline(y = 0, color = 'red')
ax1.axhline(y = 2.5, color = 'red')
ax1.set_title('RUN %i. %.2f GeV. v11, DWC cuts' %(runs[9], energies[9]))
ax1.set_xlabel('b_x DWC')
ax1.set_ylabel('b_y DWC')
plt.colorbar(r[3], ax=ax1)
for i, energy in zip(range(len(runs)), energies):
h = ax2.hist(sum_en_v11[i], density = 1, bins=200, label = '%.2f GeV' %energy)
ax2.set_title('Reco. E sums')
ax2.set_xlabel('visible E [GeV]')
plt.legend()
plt.xlim(0., 3.)
fig, (ax1, ax2) = plt.subplots(1,2, figsize=(20, 8))
plt.suptitle('RUN %i. %.2f GeV. v11' %(runs[9], energies[9]))
df_v11[9]['Ex'] = df_v11[9].rechit_energy_w * df_v11[9].b_x
df_v11[9]['Ey'] = df_v11[9].rechit_energy_w * df_v11[9].b_y
sel_en = ((df_v11[9].b_x > -10) & (df_v11[9].b_x < 10))
sel_en &= ((df_v11[9].b_y > -10) & (df_v11[9].b_y < 10))
esums = df_v11[9][sel_en].groupby('event')['rechit_energy_w'].sum()
bary_x = df_v11[9][sel_en].groupby('event')['Ex'].sum()/esums
bary_y = df_v11[9][sel_en].groupby('event')['Ey'].sum()/esums
dx = df_v11[9][sel_en].b_x
dy = df_v11[9][sel_en].b_y
x_med, ex_med = binned_statistic(bary_x, [bary_x, esums], bins=50, statistic='median').statistic
x_iqr, ex_iqr = binned_statistic(bary_x, [bary_x, esums], bins=50, statistic=iqr).statistic
y_med, ey_med = binned_statistic(bary_y, [bary_y, esums], bins=50, statistic='median').statistic
y_iqr, ey_iqr = binned_statistic(bary_y, [bary_y, esums], bins=50, statistic=iqr).statistic
r=ax1.hist2d(bary_x, esums, 50, norm=LogNorm())
ax1.errorbar(x_med, ex_med, xerr=x_iqr, yerr=ex_iqr, marker='o', color='tab:red', linestyle='None')
ax1.set_title('Correlation E visible, bary. x')
ax1.set_xlabel('barycentre x')
ax1.set_ylabel('E visible [GeV]')
plt.colorbar(r[3], ax=ax1)
r=ax2.hist2d(bary_y, esums, 50, norm=LogNorm())
ax2.errorbar(y_med, ey_med, xerr=y_iqr, yerr=ey_iqr, marker='o', color='tab:red', linestyle='None')
ax2.set_title('Correlation E visible, bary. y')
ax2.set_xlabel('barycentre y')
ax2.set_ylabel('E visible [GeV]')
plt.colorbar(r[3], ax=ax2)
plt.figure(figsize=(10,8))
dr = np.hypot(dx,dy)
E = df_v11[9][sel_en].rechit_energy_w/esums
r = plt.hist2d(dr, E, 50, norm = LogNorm())
plt.xlabel('dr')
plt.ylabel('E visible [GeV]')
fig, (ax1, ax2) = plt.subplots(1,2, figsize=(20, 8))
plt.suptitle('RUN %i. %.2f GeV. v8' %(runs[3], energies[3]))
df_v8[3]['Ex'] = df_v8[3].rechit_energy_w * df_v8[3].b_x
df_v8[3]['Ey'] = df_v8[3].rechit_energy_w * df_v8[3].b_y
sel_en = ((df_v8[3].b_x > -2) & (df_v8[3].b_x < 2))
sel_en &= ((df_v8[3].b_y > -1) & (df_v8[3].b_y < 2))
esums = df_v8[3][sel_en].groupby('event')['rechit_energy_w'].sum()
bary_x = df_v8[3][sel_en].groupby('event')['Ex'].sum()/esums
bary_y = df_v8[3][sel_en].groupby('event')['Ey'].sum()/esums
x_med, ex_med = binned_statistic(bary_x, [bary_x, esums], bins=50, statistic='median').statistic
x_iqr, ex_iqr = binned_statistic(bary_x, [bary_x, esums], bins=50, statistic=iqr).statistic
y_med, ey_med = binned_statistic(bary_y, [bary_y, esums], bins=50, statistic='median').statistic
y_iqr, ey_iqr = binned_statistic(bary_y, [bary_y, esums], bins=50, statistic=iqr).statistic
r=ax1.hist2d(bary_x, esums, 50, norm=LogNorm())
ax1.errorbar(x_med, ex_med, xerr=x_iqr, yerr=ex_iqr, marker='o', color='tab:red', linestyle='None')
ax1.set_title('Correlation E visible, bary. x')
ax1.set_xlabel('barycentre x')
ax1.set_ylabel('E visible [GeV]')
plt.colorbar(r[3], ax=ax1)
r=ax2.hist2d(bary_y, esums, 50, norm=LogNorm())
ax2.errorbar(y_med, ey_med, xerr=y_iqr, yerr=ey_iqr, marker='o', color='tab:red', linestyle='None')
ax2.set_title('Correlation E visible, bary. y')
ax2.set_xlabel('barycentre y')
ax2.set_ylabel('E visible [GeV]')
plt.colorbar(r[3], ax=ax2)
plt.figure(figsize=(20,10))
plt.subplot(121)
plt.title('RUN %i. %i GeV. Reco. E sums' %(runs[0], energies[0]))
# r=repeatedGausFit(sum_en_v11[0], 20)
r=plt.hist(sum_en_v11[0], density=1, color = 'tab:blue', bins=200)#, label=r'reco E v11. $\sigma = $ %.3f $E =$ %.3f' %(r[1], r[0]))
# r=repeatedGausFit(sum_en_v8[0], 20)
r=plt.hist(sum_en_v8[0], density=1, bins=200, color='darkorange')#, label=r'reco E v8. $\sigma = $ %.3f $E =$ %.3f' %(r[1], r[0]))
plt.legend()
plt.subplot(122)
plt.title('RUN %i. %i GeV. Reco. E sums' %(runs[1], energies[1]))
# r=repeatedGausFit(sum_en_v11[1], 30)
r=plt.hist(sum_en_v11[1], density=1, color = 'tab:blue', bins=200)#, label=r'reco E v11. $\sigma = $ %.3f $E =$ %.3f' %(r[1], r[0]))
# r=repeatedGausFit(sum_en_v8[1], 30)
r=plt.hist(sum_en_v8[1], density=1, bins=200, color='darkorange')#, label=r'reco E v8. $\sigma = $ %.3f $E =$ %.3f' %(r[1], r[0]))
# plt.yscale('log')
plt.legend()
energies = [20, 30, 50, 80, 100, 120, 150, 200, 250, 300]
resolution_gE = []; res_err_gE = []; MIP_gE = []
resolution_CB = []; res_err_CB = []; MIP_CB = []
resolution_G = []; res_err_G = []; MIP_G = []
# plt.figure(figsize=(20,10))
for sum_v11, energy in zip(sum_en_v11,energies):
if energy == 120: continue
plt.figure(figsize=(10,8))
plt.title('v11 %i GeV' %energy)
r2 = repeatedGausFit(sum_v11, energy)
resolution_G.append(r2[1]/r2[0])
error = err_prop(r2[1],r2[0],r2[3],r2[2])
res_err_G.append(error)
MIP_G.append(r2[0])
r2 = GausExpFit(sum_v11, energy)
resolution_gE.append(r2[0])
res_err_gE.append(r2[2])
MIP_gE.append(r2[1])
r2 = CBFit(sum_v11, energy)
resolution_CB.append(r2[0])
res_err_CB.append(r2[2])
MIP_CB.append(r2[1])
plt.savefig('v11_cutsDWC_MultiFit_%iGeV.png' %energy)
energies = [20, 30, 49.99, 79.93, 99.83, 119.65, 149.14, 197.32, 243.61, 287.18]
energies = [energy for energy in energies if (energy != 119.65)]
plt.figure(figsize=(10,8))
plt.title('Resolution comparison for different fitting methods' '\n' 'Using DWC cuts')
sys = np.asarray([0.05*res for res in resolution_G])
plt.errorbar(energies, np.asarray(resolution_G), xerr=0.002*np.asarray(energies),
yerr = np.sqrt(sys**2 + np.asarray(res_err_G)**2), marker = 'o', linestyle = 'None', label = 'Gaus fit')
sys = np.asarray([0.05*res for res in resolution_CB])
plt.errorbar(energies, np.asarray(resolution_CB), xerr=0.002*np.asarray(energies),
yerr = np.sqrt(sys**2 + np.asarray(res_err_CB)**2), marker = 'o', linestyle = 'None', label = 'CB fit')
sys = np.asarray([0.05*res for res in resolution_gE])
plt.errorbar(energies, np.asarray(resolution_gE), xerr=0.002*np.asarray(energies),
yerr = np.sqrt(sys**2 + np.asarray(res_err_gE)**2), marker = 'o', linestyle = 'None', label = 'GausExp fit')
plt.plot(energies, simul_reso, marker='o', mfc = 'None', linestyle='None', color='red', label ='MC')
plt.ylim(0, 0.06)
plt.legend()
plt.grid()
fig_1 = plt.figure(figsize=(10,8))
frame1 = fig_1.add_axes((.1,.3,.8,.6))
plt.title('Linearity comparison for different fitting methods' '\n' 'Using DWC cuts')
plt.plot(energies, MIP_G, 'o', label = 'Gaus')
plt.plot(energies, MIP_CB, '*', label = 'CB')
plt.plot(energies, MIP_gE, '^', label = 'GausExp')
plt.plot(energies, E_MC, marker='o', linestyle='None', mfc = 'None', color='red', label = 'MC')
plt.ylabel(r'$\left\langle E_{fit} \right\rangle$ visible')
plt.ylim(0., 3.)
plt.grid()
plt.legend(title = 'HGCAL TB2018 Data/MC')
frame1.set_xticklabels([])
frame2=fig_1.add_axes((.1,.1,.8,.2))
diff_GE = (np.asarray(MIP_gE)-np.asarray(E_MC))/np.asarray(E_MC)
diff_CB = (np.asarray(MIP_CB)-np.asarray(E_MC))/np.asarray(E_MC)
diff_Gaus = (np.asarray(MIP_G)-np.asarray(E_MC))/np.asarray(E_MC)
plt.errorbar(energies, diff_Gaus, xerr=0.002*np.asarray(energies), yerr=0,
marker='o', linestyle='None',label='<E> res. Gaus')
plt.errorbar(energies, diff_GE, xerr=0.002*np.asarray(energies), yerr=0,
marker='o', linestyle='None',label='<E> res. GausExp')
plt.errorbar(energies, diff_CB, xerr=0.002*np.asarray(energies), yerr=0,
marker='o', linestyle='None',label='<E> res. Crystal Ball')
plt.ylabel(r'$\left\langle E\right\rangle$ [MIP]')
plt.xlabel(r'$E_{beam}$ [GeV]')
plt.ylabel(r'($E_{fit}$-$E_{MC}$)/$E_{MC}$')
plt.ylim(-0.08, 0.08)
plt.grid()
resolution_new = []; resolution_old = []; en_fit = []; res_err_new = []; res_err_old = []; e_lin = []; e_lin_old=[]
MIP_new = []; MIP_old = []; err_MIP_new = []; err_MIP_old = []
energies = [20, 30, 50, 80, 100, 120, 150, 200, 250, 300]
resolution_11 = []; res_err_11 = []; MIP_11 = []
plt.figure(figsize=(20,10))
# plt.subplot(121)
# plt.title('v9 fits')
# for sum_v9, energy in zip(sum_en_v9, energies):
# if energy == 120: continue
# r2 = repeatedGausFit(sum_v9, energy)
# resolution_new.append(r2[1]/r2[0])
# error = err_prop(r2[1],r2[0],r2[3],r2[2])
# res_err_new.append(error)
# MIP_new.append(r2[0])
# plt.xlabel('E sum [GeV]')
plt.subplot(121)
plt.title('v8 fits')
for sum_v8, energy in zip(sum_en_v8,energies):
# plt.figure(figsize=(10,8))
if energy == 120: continue
r2 = repeatedGausFit(sum_v8, energy)
# plt.title('Data (v8) %i GeV' %energy)
# a = plt.hist(sum_v8, bins=300, normed=1)
resolution_old.append(r2[1]/r2[0])
error = err_prop(r2[1],r2[0],r2[3],r2[2])
res_err_old.append(error)
MIP_old.append(r2[0])
plt.xlim(0., 3.0)
plt.subplot(122)
plt.title('v11 fits')
for sum_v11, energy in zip(sum_en_v11,energies):
if energy == 120: continue
r2 = repeatedGausFit(sum_v11, energy)
resolution_11.append(r2[1]/r2[0])
error = err_prop(r2[1],r2[0],r2[3],r2[2])
res_err_11.append(error)
MIP_11.append(r2[0])
resolution_11 = [0.05509654028934182,
0.048416831183358836,
0.036802084835793086,
0.030230580062532198,
0.027763607459974998,
0.022211176199090948,
0.02041216657438586,
0.018305488347147493,
0.01724068354471974]
res_err_11 = [0.001038942172852401,
0.0010491398030856168,
0.0009532104540543196,
0.0008421136042935656,
0.0009105000867031456,
0.0005300965539583607,
0.0005066785701384369,
0.00037238852587587624,
0.000592254750896224]
MIP_11 =[0.16928161145257495,
0.2531743242030127,
0.4251338470661471,
0.6816991088977058,
0.8510718944104344,
1.2707809023945833,
1.6780642077063601,
2.066967093295746,
2.444514904087021]
resolution_old = [0.05246849411574991,
0.04356579361980387,
0.03632208007443212,
0.029958395913805447,
0.02530352238094564,
0.022427495545102396,
0.0187401627143027,
0.017819044938413693,
0.016865872898028156]
res_err_old = [0.0014072224607638963,
0.001124067458120093,
0.0008543703439377619,
0.0008318049333382934,
0.0009267709664638018,
0.0009136234433585624,
0.0007944884503405302,
0.0003716883142259822,
0.000785111101665239]
MIP_old = [0.17353610429150784,
0.25847092741828015,
0.4319115554318758,
0.690278299772658,
0.8586427611874371,
1.2814253028723628,
1.6856248528595612,
2.067077924249674,
2.42826522259055]
energies = [20, 30, 49.99, 79.93, 99.83, 119.65, 149.14, 197.32, 243.61, 287.18]
energies = [energy for energy in energies if (energy != 119.65)]
plt.figure(figsize=(10,8))
sys_old = [0.05*res for res in resolution_old]
plt.errorbar(energies, np.asarray(resolution_old), xerr=0.002*np.asarray(energies),
yerr=np.sqrt(np.asarray(res_err_old)**2+np.asarray(sys_old)**2), marker='o', linestyle='None',label='Data v8')
# popt, pcov = curve_fit(E_res, energies, np.asarray(resolution_old))
# plt.plot(energies, E_res(energies, *popt), '-.', label='Stoc. = %.2f, N = %.2f, C = %.2f' %(popt[0], popt[1], popt[2]))
# print popt, np.sqrt(np.diag(pcov))
sys_11 = [0.05*res for res in resolution_11]
plt.errorbar(energies, np.asarray(resolution_11), xerr=0.002*np.asarray(energies),
yerr=np.sqrt(np.asarray(res_err_11)**2+np.asarray(sys_11)**2), marker='o', linestyle='None',label=' Data v11')
# popt, pcov = curve_fit(E_res, energies, resolution_11, p0=[ 2.5e-01, 1.3e-05, -9.6e-03])
# plt.plot(energies, E_res(energies, *popt), '-.', label='Stoc. = %.2f, N = %.2f, C = %.2f' %(popt[0], popt[1], popt[2]))
# print popt, pcov
plt.plot(energies, simul_reso, 'o', mfc='None', color='red',label = 'MC')
# popt, pcov = curve_fit(E_res, energies, simul_reso)
# plt.plot(energies, E_res(energies, *popt), '-.', label='Stoc. = %.2f, N = %.2f, C = %.2f' %(popt[0], popt[1], popt[2]))
# print popt, np.sqrt(np.diag(pcov))
plt.title('Resolution comparison w/ new DWC cuts')
plt.legend(title = 'HGCAL TB2018 Data/MC')
plt.xlabel(r'$E_{beam}$ [GeV]')
plt.ylabel(r'Gaus $\sigma$/$\left\langle E\right\rangle$')
plt.grid()
plt.ylim(0., 0.06)
fig_1 = plt.figure(figsize=(10,8))
frame1 = fig_1.add_axes((.1,.3,.8,.6))
plt.title('Linearity comparison w/ new DWC cuts')
plt.errorbar(energies, MIP_old, xerr=0.002*np.asarray(energies), yerr=0,
marker='^', linestyle='None',label='Data v8')
plt.errorbar(energies, MIP_11, xerr=0.002*np.asarray(energies), yerr=0,
marker='^', linestyle='None',label='Data v11')
plt.errorbar(energies, E_MC, xerr=0.002*np.asarray(energies), yerr=0,
marker='o', color='red', mfc='None',linestyle='None',label='MC')
plt.legend(title = 'HGCAL TB2018 Data/MC')
plt.ylabel(r'$\left\langle E_{fit} \right\rangle$ visible')
plt.ylim(0., 3.)
plt.grid()
frame1.set_xticklabels([])
frame2=fig_1.add_axes((.1,.1,.8,.2))
diff_v8 = (np.asarray(MIP_old)-np.asarray(E_MC))/np.asarray(E_MC)
diff_v11 = (np.asarray(MIP_11)-np.asarray(E_MC))/np.asarray(E_MC)
plt.errorbar(energies, diff_v8, xerr=0.002*np.asarray(energies), yerr=0,
marker='o', linestyle='None',label='<E> res. v8')
plt.errorbar(energies, diff_v11, xerr=0.002*np.asarray(energies), yerr=0,
marker='o', linestyle='None',label='<E> res. v11')
plt.ylabel(r'$\left\langle E\right\rangle$ [MIP]')
plt.legend(title = 'HGCAL TB2018 Data/MC')
plt.xlabel(r'$E_{beam}$ [GeV]')
plt.ylabel(r'($E_{fit}$-$E_{MC}$)/$E_{MC}$')
plt.ylim(-0.08, 0.08)
plt.grid()
plt.legend(ncol=2)
plt.figure(figsize=(10,8))
# sys_new = [0.05*res for res in resolution_new]
# plt.errorbar(energies, np.asarray(resolution_new), xerr=0.002*np.asarray(energies),
# yerr=np.sqrt(np.asarray(res_err_new)**2+np.asarray(sys_new)**2), marker='o', linestyle='None',label=' Data v9')
# plt.errorbar(79.93, res_v9_470, xerr=[0.002*79.93],yerr=[err_v9_470], marker='o', linestyle='None',label='467 v9')
sys_old = [0.05*res for res in resolution_old]
plt.errorbar(energies, np.asarray(resolution_old), xerr=0.002*np.asarray(energies),
yerr=np.sqrt(np.asarray(res_err_old)**2+np.asarray(sys_old)**2), marker='o', linestyle='None',label='Data')
# plt.errorbar(79.93, res_v8_470, xerr=[0.002*79.93], yerr=[err_v8_470], marker='o', linestyle='None',label='467 v8')
plt.plot(energies, simul_reso, 'o', mfc='None', color='red',label = 'MC')
plt.legend(title = 'HGCAL TB2018 Data/MC')
plt.xlabel(r'$E_{beam}$ [GeV]')
plt.ylabel(r'Gaus $\sigma$/$\left\langle E\right\rangle$')
plt.grid()
plt.ylim(0., 0.06)
fig1 = plt.figure(figsize=(10,8))
frame1=fig1.add_axes((.1,.3,.8,.6))
# plt.title('Reco. energy linearity from rechit_energy')#\n Using median and IQR')
# diff_v9 = (np.asarray(MIP_new)-np.asarray(E_MC))/np.asarray(E_MC)
# plt.errorbar(energies, diff_v9, xerr=0.002*np.asarray(energies), yerr=0,
# marker='o', linestyle='None',label='v9')
diff_v8 = (np.asarray(MIP_old)-np.asarray(E_MC))/np.asarray(E_MC)
plt.errorbar(energies, diff_v8, xerr=0.002*np.asarray(energies), yerr=0,
marker='o', linestyle='None',label='Data')
plt.ylabel(r'$\left\langle E\right\rangle$ [MIP]')
plt.legend(title = 'HGCAL TB2018 Data/MC')
plt.xlabel(r'$E_{beam}$ [GeV]')
plt.ylabel(r'($E_{fit}$-$E_{MC}$)/$E_{MC}$')
plt.ylim(-0.08, 0.08)
plt.grid()
fig1 = plt.figure(figsize=(10,8))
frame1=fig1.add_axes((.1,.3,.8,.6))
# plt.title('Reco. energy linearity from rechit_energy')#\n Using median and IQR')
# plt.errorbar(energies, E_new, xerr=0.002*np.asarray(energies), yerr=0,
# marker='o', linestyle='None',label='v9')
plt.errorbar(energies, MIP_old, xerr=0.002*np.asarray(energies), yerr=0,
marker='^', linestyle='None',label='Data')
plt.errorbar(energies, E_MC, xerr=0.002*np.asarray(energies), yerr=0,
marker='o', color='red', mfc='None',linestyle='None',label='MC')
plt.legend(title = 'HGCAL TB2018 Data/MC')
plt.xlabel(r'$E_{beam}$ [GeV]')
plt.ylabel(r'$\left\langle E_{fit} \right\rangle$ visible')
plt.ylim(0., 3.)
plt.grid()
fig_1 = plt.figure(figsize=(10,8))
frame1 = fig_1.add_axes((.1,.3,.8,.6))
plt.errorbar(energies, MIP_old, xerr=0.002*np.asarray(energies), yerr=0,
marker='^', linestyle='None',label='Data')
plt.errorbar(energies, E_MC, xerr=0.002*np.asarray(energies), yerr=0,
marker='o', color='red', mfc='None',linestyle='None',label='MC')
plt.legend(title = 'HGCAL TB2018 Data/MC')
# plt.xlabel(r'$E_{beam}$ [GeV]')
plt.ylabel(r'$\left\langle E_{fit} \right\rangle$ visible')
plt.ylim(0., 3.)
plt.grid()
frame1.set_xticklabels([])
frame2=fig_1.add_axes((.1,.1,.8,.2))
diff_v8 = (np.asarray(MIP_old)-np.asarray(E_MC))/np.asarray(E_MC)
plt.errorbar(energies, diff_v8, xerr=0.002*np.asarray(energies), yerr=0,
marker='o', linestyle='None',label='<E> res.')
plt.ylabel(r'$\left\langle E\right\rangle$ [MIP]')
plt.legend(title = 'HGCAL TB2018 Data/MC')
plt.xlabel(r'$E_{beam}$ [GeV]')
plt.ylabel(r'($E_{fit}$-$E_{MC}$)/$E_{MC}$')
plt.ylim(-0.08, 0.08)
plt.grid()
plt.legend()
```
<h1> Resolution comparison flagging noisy cells </h1>
```
%%time
# 80 GeV runs
_80GeV = range(466, 476)
_100GeV = range(477, 492)
_150GeV = range(493, 510)#range(303, 319) + range(337, 361) + range(493, 510)
run_real = []
df_v8 = []
for run in _150GeV:
try:
df_v8.append(add_dwc(run, v8_prefix, 8))
run_real.append(run)
except IOError:
continue
%%time
sum_en_v8 = []
for df in df_v8:
sum_en_v8.append(do_Esum(df))
%%time
#Create E sums flagging noisy cells
sum_en_v8_NF = []
for df in df_v8:
sum_en_v8_NF.append(do_Esum_Noise(df))
resolution_old = []; res_err_old = []; MIP_old = []
resolution_old_NF = []; res_err_old_NF = []; MIP_old_NF = []
Es = []; sigmas = []
energies = [20, 30, 50, 80, 100, 120, 150, 200, 250, 300]
for sum_v8 in sum_en_v8:
energy = 150
# if energy == 20: continue
r2 = repeatedGausFit(sum_v8, energy)
resolution_old.append(r2[1]/r2[0])
Es.append(r2[0])
sigmas.append(r2[1])
error = err_prop(r2[1],r2[0],r2[3],r2[2])
res_err_old.append(error)
MIP_old.append(r2[0])
for sum_v8 in sum_en_v8_NF:
energy = 150
# if energy == 20: continue
r2 = repeatedGausFit(sum_v8, energy)
resolution_old_NF.append(r2[1]/r2[0])
error = err_prop(r2[1],r2[0],r2[3],r2[2])
res_err_old_NF.append(error)
MIP_old_NF.append(r2[0])
import itertools
from collections import defaultdict
resos = defaultdict(list)
for i in range(len(sum_en_v8)):
resos[i].append(sum_en_v8[i])
for i in range(len(sum_en_v8)):
ss = pd.Series(list(itertools.chain.from_iterable(resos[i])))
r = repeatedGausFit(ss, 150)
print r[1]/r[0]
plt.figure(figsize=(10,8))
sys_old = [0.05*res for res in resolution_old]
plt.errorbar(run_real, np.asarray(resolution_old), xerr=0,
yerr=np.sqrt(np.asarray(res_err_old)**2+np.asarray(sys_old)**2), marker='o', linestyle='None',label='Data')
sys_old_NF = [0.05*res for res in resolution_old_NF]
plt.errorbar(run_real, np.asarray(resolution_old_NF), xerr=0,
yerr=np.sqrt(np.asarray(res_err_old_NF)**2+np.asarray(sys_old_NF)**2), marker='o', linestyle='None',label='Data (no noisy cells)')
# plt.plot(energies, simul_reso, 'o', mfc='None', color='red',label = 'MC')
plt.xlabel(r'Run ID')
plt.ylabel(r'Gaus $\sigma$/$\left\langle E\right\rangle$')
plt.grid()
# plt.axvline(x=466, color='red', linestyle = '-.')
# plt.axvline(x=472, color='red', linestyle = '-.')
# plt.axvline(x=468, color='red', linestyle = '-.')
plt.axhline(y=0.020451829612688918, color='green', linestyle='--', label='MC res.')
plt.axhline(y=resolution_old[1], color='red', linestyle='-.', label='Run 494 res.')
plt.axhline(y=np.median(resolution_old), color='purple', linestyle='-.', label='median 150GeV res.')
plt.axhline(y=np.mean(resolution_old), color='cyan', linestyle='-.', label='mean 150GeV res.')
plt.axhline(y=0.022761034755404596, color='black', linestyle='-', label='150GeV res. combined')
plt.legend(title = 'HGCAL TB2018 Data, 150GeV', ncol=2)
plt.title('150GeV runs 493-509 (with DWC)')
plt.ylim(0.015, 0.03)
```
<h1> Shower profiles and comparison </h1>
```
branches = [u'event',
u'rechit_layer','rechit_module', u'rechit_chip', u'rechit_channel', u'rechit_x', u'rechit_y', u'rechit_z',
u'rechit_energy', u'rechit_amplitudeHigh', u'rechit_amplitudeLow', u'rechit_Tot', 'rechit_toaRise',
'rechit_TS3High','rechit_TS3Low', 'rechit_TS2High','rechit_TS2Low']
rh_branches = [branch for branch in branches if "rechit" in branch]
# Adding entries in df for energy bary. components calculation
def do_df(run, v_prefix):
fname = v_prefix + "ntuple_%i.root" %run
df = read_root(fname,key='rechitntupler/hits',columns = branches, flatten = rh_branches)
df['e_w'] = df['rechit_energy']*MIPtoGeV#(df['rechit_layer'].map(weights)+df['rechit_layer'].map(weights)*MIPtoGeV)
df['lay_X0'] = df['rechit_layer'].map(lay_X0)
df['Ex'] = df.rechit_energy * df.rechit_x
df['Ey'] = df.rechit_energy * df.rechit_y
df['Ez'] = df.rechit_energy * df.rechit_z
return df
def eProf(df):
sel = (df.rechit_energy>0.5) #& (df.rechit_layer < 28)
sel &= ~((df.rechit_chip == 3) & (df.rechit_channel == 22))
df_sel = df[sel]
e_sum = df_sel.groupby('event')['e_w'].sum()
lay_sum = df_sel.groupby(['event','lay_X0'])['e_w'].sum()
e_frac = (lay_sum/e_sum).reset_index().groupby('lay_X0')['e_w'].mean()
e_lay = df_sel.groupby('lay_X0')['e_w'].size()
xbar = df_sel.groupby('event')['Ex'].sum()/e_sum
ybar = df_sel.groupby('event')['Ey'].sum()/e_sum
zbar = df_sel.groupby('event')['Ez'].sum()/e_sum
bary = [xbar, ybar, zbar]
return lay_sum, e_lay, bary
from scipy.special import gamma
def eShow(t, E, a, b):
return E*((b*t)**(a-1)*b*np.exp(-b*t))/gamma(a)
energies = [20, 30, 50, 80, 100, 120, 150, 200, 250, 300]
runs = [452, 607, 457, 467, 477, 621, 496, 664, 653, 420]
eFracs = []
for i in range(len(runs)):
run = runs[i]
print energies[i],
df = do_df(run, v9_prefix)
shower = eProf(df)
eFracs.append(shower[0])
plt.figure(figsize=(20,10))
for i in range(len(eFracs)):
energy = energies[i]
plt.plot(eFracs[i].index, eFracs[i], 'o--', alpha=0.6, label = '%i GeV' %energy)
sel = eFracs[i].index < 28
popt, pcov = curve_fit(eShow, eFracs[i].index, eFracs[i])
T = (popt[0]-1)/popt[1]
t = popt[1]/popt[0]
E_c = 40.19*1e-3
eT = np.exp(T)
# plt.axvline(x=T, color='red')
E_beam = eT*E_c
print E_beam
plt.plot(eFracs[i].index, eShow(eFracs[i].index, *popt), linewidth=2, label = r'$E_{fit}$ = %i GeV' %E_beam)
plt.legend()
dd = do_df(1008, v11_prefix)
e11 = eProf(dd)
plt.plot(e11[0][3279].index, e11[0][3279], 'o--')
sel = (e11[0][3279].index < 29)
popt, pcov = curve_fit(eShow, e11[0][3279][sel].index, e11[0][3279][sel])
plt.plot(e11[0][3279].index, eShow(e11[0][3279].index, *popt))
print popt[0]#*MIPtoGeV
# Create df for ntuples
run = 672
df_v9 = do_df(run, v9_prefix); df_v8 = do_df(run, v8_prefix)
etot_v9 = df_v9.groupby('event').e_w.sum(); etot_v8 = df_v8.groupby('event').e_w.sum()
e_v9 = eProf(df_v9); e_v8 = eProf(df_v8)
eFrac_v9 = e_v9[0]; eFrac_v8 = e_v8[0]
eLay_v9 = e_v9[1]; eLay_v8 = e_v8[1]
bar_v9 = e_v9[2]; bar_v8 = e_v8[2]
# Energy fraction profiles comparison
plt.figure(figsize=(20, 10))
plt.subplot(121)
plt.title('RUN %i' %run)
plt.plot(eFrac_v9.index, eFrac_v9, 'o-', label = 'v9')
plt.plot(eFrac_v8, '^-', label = 'v8')
plt.xlabel('Layer [X0]')
plt.ylabel(r'$E_{dep}/Lay.$')
plt.legend()
plt.subplot(122)
plt.title('RUN %i' %run)
plt.plot(eLay_v9, 'o-', label = 'v9')
plt.plot(eLay_v8, '^-', label = 'v8')
plt.xlabel('Layer [X0]')
plt.ylabel(r'E/Lay.')
plt.legend()
z_med_v9, e_med_v9 = binned_statistic(bar_v9[2], [bar_v9[2], etot_v9], bins=50, statistic='median').statistic
z_iqr_v9, e_iqr_v9 = binned_statistic(bar_v9[2], [bar_v9[2], etot_v9], bins=50, statistic=iqr).statistic
z_med_v8, e_med_v8 = binned_statistic(bar_v8[2], [bar_v8[2], etot_v8], bins=50, statistic='median').statistic
z_iqr_v8, e_iqr_v8 = binned_statistic(bar_v8[2], [bar_v8[2], etot_v8], bins=50, statistic=iqr).statistic
plt.figure(figsize = (30, 20))
plt.subplot(221)
plt.title('Shower profile v9. RUN %i' %run)
r =plt.hist2d(bar_v9[2], etot_v9, 50, cmap = 'viridis', norm = LogNorm())
plt.plot(z_med_v9, e_med_v9, 'ko', color='red')
plt.xlabel('bary z')
plt.ylabel('E sum')
plt.subplot(222)
plt.title('Shower profile v8. RUN %i' %run)
r =plt.hist2d(bar_v8[2], etot_v8, 50, cmap = 'viridis', norm = LogNorm())
plt.plot(z_med_v8, e_med_v8, 'ko', color='red')
plt.xlabel('bary z')
plt.ylabel('E sum [GeV]')
sel = df_v9.rechit_layer < 29
bary_z_EE_v9 = df_v9[sel].groupby('event')['Ez'].sum()/etot_v9
sel = df_v8.rechit_layer < 29
bary_z_EE_v8 = df_v8[sel].groupby('event')['Ez'].sum()/etot_v8
z_med_v9_EE, e_med_v9_EE = binned_statistic(bary_z_EE_v9, [bary_z_EE_v9, etot_v9], bins=50, statistic='median').statistic
z_med_v8_EE, e_med_v8_EE = binned_statistic(bary_z_EE_v8, [bary_z_EE_v8, etot_v8], bins=50, statistic='median').statistic
z_iqr_v9_EE, e_iqr_v9_EE = binned_statistic(bary_z_EE_v9, [bary_z_EE_v9, etot_v9], bins=50, statistic=iqr).statistic
z_iqr_v8_EE, e_iqr_v8_EE = binned_statistic(bary_z_EE_v8, [bary_z_EE_v8, etot_v8], bins=50, statistic=iqr).statistic
plt.subplot(223)
plt.title('Shower profile in EE v9. RUN %i' %run)
r =plt.hist2d(bary_z_EE_v9, etot_v9, 50, cmap = 'viridis', norm = LogNorm())
plt.plot(z_med_v9_EE, e_med_v9_EE, 'ko', color='red')
plt.xlabel('bary z')
plt.ylabel('E sum [GeV]')
plt.subplot(224)
plt.title('Shower profile in EE v8. RUN %i' %run)
r =plt.hist2d(bary_z_EE_v8, etot_v8, 50, cmap = 'viridis', norm = LogNorm())
plt.plot(z_med_v8_EE, e_med_v8_EE, 'ko', color='red')
plt.xlabel('bary z')
plt.ylabel('E sum [GeV]')
plt.figure(figsize=(20,10))
plt.subplot(121)
plt.title('Shower energy deposit profile. RUN %i' %run)
plt.errorbar(z_med_v9, e_med_v9, xerr=z_iqr_v9, yerr=e_iqr_v9, marker='o', linestyle='None', label = 'v9')
plt.errorbar(z_med_v8, e_med_v8, xerr=z_iqr_v8, yerr=e_iqr_v8, marker='o', linestyle='None', label = 'v8')
plt.xlabel(r'$\left\langle bary. z\right\rangle$')
plt.ylabel(r'$\left\langle E sum\right\rangle$')
plt.legend()
plt.subplot(122)
plt.title('Shower energy deposit profile in EE. RUN %i' %run)
plt.errorbar(z_med_v9_EE, e_med_v9_EE, xerr=z_iqr_v9_EE, yerr=e_iqr_v9_EE, marker='o', linestyle='None', label = 'v9')
plt.errorbar(z_med_v8_EE, e_med_v8_EE, xerr=z_iqr_v8_EE, yerr=e_iqr_v8_EE, marker='o', linestyle='None', label = 'v8')
plt.xlabel(r'$\left\langle bary. z\right\rangle$')
plt.ylabel(r'$\left\langle E sum\right\rangle$')
plt.legend()
v9_layer = df_v9.groupby(['event']).lay_X0.mean()
v8_layer = df_v8.groupby(['event']).lay_X0.mean()
v9_lay_med, e_v9_med = binned_statistic(v9_layer, [v9_layer, etot_v9], bins=50, statistic='median').statistic
v8_lay_med, e_v8_med = binned_statistic(v8_layer, [v8_layer, etot_v8], bins=50, statistic='median').statistic
v9_lay_iqr, e_v9_iqr = binned_statistic(v9_layer, [v9_layer, etot_v9], bins=50, statistic=iqr).statistic
v8_lay_iqr, e_v8_iqr = binned_statistic(v8_layer, [v8_layer, etot_v8], bins=50, statistic=iqr).statistic
plt.figure(figsize=(20, 10))
plt.subplot(121)
plt.title('Energy deposit per layer profile, v9. RUN %i' %run)
r =plt.hist2d(v9_layer, etot_v9, 50, cmap = 'viridis', norm = LogNorm())
plt.plot(v9_lay_med, e_v9_med, 'ko', color='red')
plt.xlabel('Layer')
plt.ylabel('E sum [GeV]')
plt.subplot(122)
plt.title('Energy deposit per layer profile, v8. RUN %i' %run)
r =plt.hist2d(v8_layer, etot_v8, 50, cmap = 'viridis', norm=LogNorm())
plt.plot(v8_lay_med, e_v8_med, 'ko', color='red')
plt.xlabel('Layer')
plt.ylabel('E sum [GeV]')
plt.figure(figsize=(10,8))
plt.title('Energy per layer profiles. RUN %i' %run)
plt.errorbar(v9_lay_med, e_v9_med, xerr=v9_lay_iqr, yerr=e_iqr_v9, marker='o', linestyle = 'None', label = 'v9')
plt.errorbar(v8_lay_med, e_v8_med, xerr=v8_lay_iqr, yerr=e_iqr_v8, marker='o', linestyle = 'None', label = 'v8')
plt.legend()
plt.xlabel('Layer')
plt.ylabel('E sum [GeV]')
```
<h1> Comparison for different energies </h1>
```
def compare_Showers(df_v9, df_v8):
etot_v9 = df_v9.groupby('event').rechit_energy.sum(); etot_v8 = df_v8.groupby('event').rechit_energy.sum()
e_v9 = eProf(df_v9); e_v8 = eProf(df_v8)
eFrac_v9 = e_v9[0]; eFrac_v8 = e_v8[0]
eLay_v9 = e_v9[1]; eLay_v8 = e_v8[1]
bar_v9 = e_v9[2]; bar_v8 = e_v8[2]
# Energy fraction profiles comparison
plt.figure(figsize=(20, 10))
plt.subplot(121)
plt.title('RUN %i' %run)
plt.plot(eFrac_v9, 'o-', label = 'v9')
plt.plot(eFrac_v8, '^-', label = 'v8')
plt.xlabel('Layer')
plt.ylabel(r'$E_{dep}/Lay.$')
plt.legend()
plt.subplot(122)
plt.title('RUN %i' %run)
plt.plot(eLay_v9, 'o-', label = 'v9')
plt.plot(eLay_v8, '^-', label = 'v8')
plt.xlabel('Layer')
plt.ylabel(r'E/Lay.')
plt.legend()
z_med_v9, e_med_v9 = binned_statistic(bar_v9[2], [bar_v9[2], etot_v9], bins=50, statistic='median').statistic
z_iqr_v9, e_iqr_v9 = binned_statistic(bar_v9[2], [bar_v9[2], etot_v9], bins=50, statistic=iqr).statistic
z_med_v8, e_med_v8 = binned_statistic(bar_v8[2], [bar_v8[2], etot_v8], bins=50, statistic='median').statistic
z_iqr_v8, e_iqr_v8 = binned_statistic(bar_v8[2], [bar_v8[2], etot_v8], bins=50, statistic=iqr).statistic
plt.figure(figsize = (30, 20))
plt.subplot(221)
plt.title('Shower profile v9. RUN %i' %run)
r =plt.hist2d(bar_v9[2], etot_v9, 50, cmap = 'viridis', norm = LogNorm())
plt.plot(z_med_v9, e_med_v9, 'ko', color='red')
plt.xlabel('bary z')
plt.ylabel('E sum')
plt.subplot(222)
plt.title('Shower profile v8. RUN %i' %run)
r =plt.hist2d(bar_v8[2], etot_v8, 50, cmap = 'viridis', norm = LogNorm())
plt.plot(z_med_v8, e_med_v8, 'ko', color='red')
plt.xlabel('bary z')
plt.ylabel('E sum')
sel = df_v9.rechit_layer < 28
bary_z_EE_v9 = df_v9[sel].groupby('event')['Ez'].sum()/etot_v9
sel = df_v8.rechit_layer < 28
bary_z_EE_v8 = df_v8[sel].groupby('event')['Ez'].sum()/etot_v8
z_med_v9_EE, e_med_v9_EE = binned_statistic(bary_z_EE_v9, [bary_z_EE_v9, etot_v9], bins=50, statistic='median').statistic
z_med_v8_EE, e_med_v8_EE = binned_statistic(bary_z_EE_v8, [bary_z_EE_v8, etot_v8], bins=50, statistic='median').statistic
z_iqr_v9_EE, e_iqr_v9_EE = binned_statistic(bary_z_EE_v9, [bary_z_EE_v9, etot_v9], bins=50, statistic=iqr).statistic
z_iqr_v8_EE, e_iqr_v8_EE = binned_statistic(bary_z_EE_v8, [bary_z_EE_v8, etot_v8], bins=50, statistic=iqr).statistic
plt.subplot(223)
plt.title('Shower profile in EE v9. RUN %i' %run)
r =plt.hist2d(bary_z_EE_v9, etot_v9, 50, cmap = 'viridis', norm = LogNorm())
plt.plot(z_med_v9_EE, e_med_v9_EE, 'ko', color='red')
plt.xlabel('bary z')
plt.ylabel('E sum')
plt.subplot(224)
plt.title('Shower profile in EE v8. RUN %i' %run)
r =plt.hist2d(bary_z_EE_v8, etot_v8, 50, cmap = 'viridis', norm = LogNorm())
plt.plot(z_med_v8_EE, e_med_v8_EE, 'ko', color='red')
plt.xlabel('bary z')
plt.ylabel('E sum')
plt.figure(figsize=(20,10))
plt.subplot(121)
plt.title('Shower energy deposit profile. RUN %i' %run)
plt.errorbar(z_med_v9, e_med_v9, xerr=z_iqr_v9, yerr=e_iqr_v9, marker='o', linestyle='None', label = 'v9')
plt.errorbar(z_med_v8, e_med_v8, xerr=z_iqr_v8, yerr=e_iqr_v8, marker='o', linestyle='None', label = 'v8')
plt.xlabel(r'$\left\langle bary. z\right\rangle$')
plt.ylabel(r'$\left\langle E sum\right\rangle$')
plt.legend()
plt.subplot(122)
plt.title('Shower energy deposit profile in EE. RUN %i' %run)
plt.errorbar(z_med_v9_EE, e_med_v9_EE, xerr=z_iqr_v9, yerr=e_iqr_v9, marker='o', linestyle='None', label = 'v9')
plt.errorbar(z_med_v8_EE, e_med_v8_EE, xerr=z_iqr_v8, yerr=e_iqr_v8, marker='o', linestyle='None', label = 'v8')
plt.xlabel(r'$\left\langle bary. z\right\rangle$')
plt.ylabel(r'$\left\langle E sum\right\rangle$')
plt.legend()
v9_layer = df_v9.groupby(['event']).rechit_layer.mean()
v8_layer = df_v8.groupby(['event']).rechit_layer.mean()
v9_lay_med, e_v9_med = binned_statistic(v9_layer, [v9_layer, etot_v9], bins=50, statistic='median').statistic
v8_lay_med, e_v8_med = binned_statistic(v8_layer, [v8_layer, etot_v8], bins=50, statistic='median').statistic
v9_lay_iqr, e_v9_iqr = binned_statistic(v9_layer, [v9_layer, etot_v9], bins=50, statistic=iqr).statistic
v8_lay_iqr, e_v8_iqr = binned_statistic(v8_layer, [v8_layer, etot_v8], bins=50, statistic=iqr).statistic
plt.figure(figsize=(20, 10))
plt.subplot(121)
plt.title('Energy deposit per layer profile, v9. RUN %i' %run)
r =plt.hist2d(v9_layer, etot_v9, 50, cmap = 'viridis', norm = LogNorm())
plt.plot(v9_lay_med, e_v9_med, 'ko', color='red')
plt.xlabel('Layer')
plt.ylabel('E sum [MIP]')
plt.subplot(122)
plt.title('Energy deposit per layer profile, v8. RUN %i' %run)
r =plt.hist2d(v8_layer, etot_v8, 50, cmap = 'viridis', norm=LogNorm())
plt.plot(v8_lay_med, e_v8_med, 'ko', color='red')
plt.xlabel('Layer')
plt.ylabel('E sum [MIP]')
plt.figure(figsize=(10,8))
plt.title('Energy per layer profiles. RUN %i' %run)
plt.errorbar(v9_lay_med, e_v9_med, xerr=v9_lay_iqr, yerr=e_iqr_v9, marker='o', linestyle = 'None', label = 'v9')
plt.errorbar(v8_lay_med, e_v8_med, xerr=v8_lay_iqr, yerr=e_iqr_v8, marker='o', linestyle = 'None', label = 'v8')
plt.legend()
plt.xlabel('Layer')
plt.ylabel('E sum [MIP]')
energies = [20, 30, 50, 80, 100, 120, 150, 200, 250, 300]
runs = [441, 594, 599, 456, 469, 480, 622, 502, 381, 390, 416]
run = 441
df_v9 = do_df(run, v9_prefix); df_v8 = do_df(run, v8_prefix)
compare_Showers(df_v9, df_v8)
run = 549
df_v9 = do_df(run, v9_prefix); df_v8 = do_df(run, v8_prefix)
compare_Showers(df_v9, df_v8)
run = 469
df_v9 = do_df(run, v9_prefix); df_v8 = do_df(run, v8_prefix)
compare_Showers(df_v9, df_v8)
run = 381
df_v9 = do_df(run, v9_prefix); df_v8 = do_df(run, v8_prefix)
compare_Showers(df_v9, df_v8)
run = 502
df_v9 = do_df(run, v9_prefix); df_v8 = do_df(run, v8_prefix)
compare_Showers(df_v9, df_v8)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import torch
import torchvision
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from matplotlib import pyplot as plt
from myrmsprop import MyRmsprop
%matplotlib inline
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
train_data = np.load("train_type4_data.npy",allow_pickle=True)
test_data = np.load("test_type4_data.npy",allow_pickle=True)
mosaic_list_of_images = train_data[0]["mosaic_list"]
mosaic_label = train_data[0]["mosaic_label"]
fore_idx = train_data[0]["fore_idx"]
test_mosaic_list_of_images = test_data[0]["mosaic_list"]
test_mosaic_label = test_data[0]["mosaic_label"]
test_fore_idx = test_data[0]["fore_idx"]
class MosaicDataset1(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list, mosaic_label,fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx] , self.fore_idx[idx]
batch = 3000
train_dataset = MosaicDataset1(mosaic_list_of_images, mosaic_label, fore_idx)
train_loader = DataLoader( train_dataset,batch_size= batch ,shuffle=False)
#batch = 2000
#test_dataset = MosaicDataset1(test_mosaic_list_of_images, test_mosaic_label, test_fore_idx)
#test_loader = DataLoader(test_dataset,batch_size= batch ,shuffle=False)
n_batches = 3000//batch
bg = []
for i in range(n_batches):
torch.manual_seed(i)
betag = torch.randn(3000,9)#torch.ones((250,9))/9
bg.append( betag.requires_grad_() )
len(bg)
#H = np.zeros((27000,27000))
for i, data in enumerate(train_loader, 0):
print(i) # only one batch
inputs,_,_ = data
inputs = torch.reshape(inputs,(27000,2))
dis = torch.cdist(inputs,inputs)**2
gamma = -1/torch.median(dis)
print(gamma)
H = torch.exp(gamma*dis)
H.shape
class Module2(nn.Module):
def __init__(self):
super(Module2, self).__init__()
self.linear1 = nn.Linear(2,100)
self.linear2 = nn.Linear(100,3)
def forward(self,x):
x = F.relu(self.linear1(x))
x = self.linear2(x)
return x
torch.manual_seed(1234)
what_net = Module2().double()
what_net.load_state_dict(torch.load("type4_what_net.pt"))
what_net = what_net.to("cuda")
def attn_avg(x,beta):
y = torch.zeros([batch,2], dtype=torch.float64)
y = y.to("cuda")
alpha = F.softmax(beta,dim=1) # alphas
#print(alpha[0],x[0,:])
for i in range(9):
alpha1 = alpha[:,i]
y = y + torch.mul(alpha1[:,None],x[:,i])
return y,alpha
def calculate_attn_loss(dataloader,what,criter):
what.eval()
r_loss = 0
alphas = []
lbls = []
pred = []
fidices = []
correct = 0
tot = 0
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels,fidx= data
lbls.append(labels)
fidices.append(fidx)
inputs = inputs.double()
beta = bg[i] # beta for ith batch
inputs, labels,beta = inputs.to("cuda"),labels.to("cuda"),beta.to("cuda")
avg,alpha = attn_avg(inputs,beta)
alpha = alpha.to("cuda")
outputs = what(avg)
_, predicted = torch.max(outputs.data, 1)
correct += sum(predicted == labels)
tot += len(predicted)
pred.append(predicted.cpu().numpy())
alphas.append(alpha.cpu().numpy())
loss = criter(outputs, labels)
r_loss += loss.item()
alphas = np.concatenate(alphas,axis=0)
pred = np.concatenate(pred,axis=0)
lbls = np.concatenate(lbls,axis=0)
fidices = np.concatenate(fidices,axis=0)
#print(alphas.shape,pred.shape,lbls.shape,fidices.shape)
analysis = analyse_data(alphas,lbls,pred,fidices)
return r_loss/(i+1),analysis,correct.item(),tot,correct.item()/tot
for param in what_net.parameters():
param.requires_grad = False
def analyse_data(alphas,lbls,predicted,f_idx):
'''
analysis data is created here
'''
batch = len(predicted)
amth,alth,ftpt,ffpt,ftpf,ffpf = 0,0,0,0,0,0
for j in range (batch):
focus = np.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
amth +=1
else:
alth +=1
if(focus == f_idx[j] and predicted[j] == lbls[j]):
ftpt += 1
elif(focus != f_idx[j] and predicted[j] == lbls[j]):
ffpt +=1
elif(focus == f_idx[j] and predicted[j] != lbls[j]):
ftpf +=1
elif(focus != f_idx[j] and predicted[j] != lbls[j]):
ffpf +=1
#print(sum(predicted==lbls),ftpt+ffpt)
return [ftpt,ffpt,ftpf,ffpf,amth,alth]
optim1 = []
H= H.to("cpu")
for i in range(n_batches):
optim1.append(MyRmsprop([bg[i]],H=H,lr=0.01))
#optim1.append(optim.RMSprop([bg[i]],lr=0.1))
# instantiate optimizer
#optimizer_what = optim.RMSprop(what_net.parameters(), lr=0.001)#, momentum=0.9)#,nesterov=True)
criterion = nn.CrossEntropyLoss()
acti = []
analysis_data_tr = []
analysis_data_tst = []
loss_curi_tr = []
loss_curi_tst = []
epochs = 200
# calculate zeroth epoch loss and FTPT values
running_loss,anlys_data,correct,total,accuracy = calculate_attn_loss(train_loader,what_net,criterion)
print('training epoch: [%d ] loss: %.3f correct: %.3f, total: %.3f, accuracy: %.3f' %(0,running_loss,correct,total,accuracy))
loss_curi_tr.append(running_loss)
analysis_data_tr.append(anlys_data)
# training starts
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
#what_net.train()
for i, data in enumerate(train_loader, 0):
# get the inputs
inputs, labels,_ = data
inputs = inputs.double()
beta = bg[i] # alpha for ith batch
#print(labels)
inputs, labels,beta = inputs.to("cuda"),labels.to("cuda"),beta.to("cuda")
# zero the parameter gradients
#optimizer_what.zero_grad()
optim1[i].zero_grad()
# forward + backward + optimize
avg,alpha = attn_avg(inputs,beta)
outputs = what_net(avg)
loss = criterion(outputs, labels)
# print statistics
running_loss += loss.item()
#alpha.retain_grad()
loss.backward(retain_graph=False)
#optimizer_what.step()
optim1[i].step()
running_loss_tr,anls_data,correct,total,accuracy = calculate_attn_loss(train_loader,what_net,criterion)
analysis_data_tr.append(anls_data)
loss_curi_tr.append(running_loss_tr) #loss per epoch
print('training epoch: [%d ] loss: %.3f correct: %.3f, total: %.3f, accuracy: %.3f' %(epoch+1,running_loss_tr,correct,total,accuracy))
if running_loss_tr<=0.08:
break
print('Finished Training run ')
analysis_data_tr = np.array(analysis_data_tr)
columns = ["epochs", "argmax > 0.5" ,"argmax < 0.5", "focus_true_pred_true", "focus_false_pred_true", "focus_true_pred_false", "focus_false_pred_false" ]
df_train = pd.DataFrame()
df_test = pd.DataFrame()
df_train[columns[0]] = np.arange(0,epoch+2)
df_train[columns[1]] = analysis_data_tr[:,-2]/30
df_train[columns[2]] = analysis_data_tr[:,-1]/30
df_train[columns[3]] = analysis_data_tr[:,0]/30
df_train[columns[4]] = analysis_data_tr[:,1]/30
df_train[columns[5]] = analysis_data_tr[:,2]/30
df_train[columns[6]] = analysis_data_tr[:,3]/30
df_train
fig= plt.figure(figsize=(6,6))
plt.plot(df_train[columns[0]],df_train[columns[3]], label ="focus_true_pred_true ")
plt.plot(df_train[columns[0]],df_train[columns[4]], label ="focus_false_pred_true ")
plt.plot(df_train[columns[0]],df_train[columns[5]], label ="focus_true_pred_false ")
plt.plot(df_train[columns[0]],df_train[columns[6]], label ="focus_false_pred_false ")
plt.title("On Train set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("percentage of data")
plt.xticks([0,50,100,150,200])
#plt.vlines(vline_list,min(min(df_train[columns[3]]/300),min(df_train[columns[4]]/300),min(df_train[columns[5]]/300),min(df_train[columns[6]]/300)), max(max(df_train[columns[3]]/300),max(df_train[columns[4]]/300),max(df_train[columns[5]]/300),max(df_train[columns[6]]/300)),linestyles='dotted')
plt.show()
fig.savefig("train_analysis.pdf")
fig.savefig("train_analysis.png")
aph = []
for i in bg:
aph.append(F.softmax(i,dim=1).detach().numpy())
aph = np.concatenate(aph,axis=0)
# torch.save({
# 'epoch': 500,
# 'model_state_dict': what_net.state_dict(),
# #'optimizer_state_dict': optimizer_what.state_dict(),
# "optimizer_alpha":optim1,
# "FTPT_analysis":analysis_data_tr,
# "alpha":aph
# }, "type4_what_net_500.pt")
aph[0]
xx,yy= np.meshgrid(np.arange(1,8,0.01),np.arange(2,9,0.01))
X = np.concatenate((xx.reshape(-1,1),yy.reshape(-1,1)),axis=1)
X = torch.Tensor(X).double().to("cuda")
Y1 = what_net(X)
Y1 = Y1.to("cpu")
Y1 = Y1.detach().numpy()
Y1 = torch.softmax(torch.Tensor(Y1),dim=1)
_,Z4= torch.max(Y1,1)
Z1 = Y1[:,0]
Z2 = Y1[:,1]
Z3 = Y1[:,2]
X = X.to("cpu")
data = np.load("type_4_data.npy",allow_pickle=True)
x = data[0]["X"]
y = data[0]["Y"]
idx= []
for i in range(10):
print(i,sum(y==i))
idx.append(y==i)
avrg = []
with torch.no_grad():
for i, data in enumerate(train_loader):
inputs , labels , fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
beta = bg[i]
beta = beta.to("cuda")
avg,alpha = attn_avg(inputs,beta)
avrg.append(avg.detach().cpu().numpy())
avrg= np.concatenate(avrg,axis=0)
fig = plt.figure(figsize=(6,6))
#plt.scatter(X[:,0],X[:,1],c=Z4)
Z4 = Z4.reshape(xx.shape)
plt.contourf(xx, yy, Z4, alpha=0.4)
for i in range(3):
plt.scatter(x[idx[i],0],x[idx[i],1],label="class_"+str(i),alpha=0.8)
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.scatter(avrg[:,0],avrg[:,1],alpha=0.2)
plt.savefig("decision_boundary.png",bbox_inches="tight")
```
| github_jupyter |
# Synthetic Radiographs with Custom Source Profiles
[Tracker]: ../../api/plasmapy.diagnostics.charged_particle_radiography.Tracker.rst#plasmapy.diagnostics.charged_particle_radiography.Tracker
In real charged particle radiography experiments, the finite size and distribution of the particle source limits the resolution of the radiograph. Some realistic sources produce particles with a non-uniform angular distribution that then superimposes a large scale "source profile" on the radiograph. For these reasons, the
[Tracker] particle tracing class allows users to specify their own initial particle positions and velocities. This example will demonstrate how to use this functionality to create a more realistic synthetic radiograph that includes the effects from a non-uniform, finite source profile.
```
import astropy.constants as const
import astropy.units as u
import matplotlib.pyplot as plt
import numpy as np
import warnings
from mpl_toolkits.mplot3d import Axes3D
from plasmapy.diagnostics import charged_particle_radiography as cpr
from plasmapy.formulary.mathematics import rot_a_to_b
from plasmapy.particles import Particle
from plasmapy.plasma.grids import CartesianGrid
```
## Contents
1. [Creating Particles](#Creating-Particles)
1. [Creating the Initial Particle Velocities](#Creating-the-Initial-Particle-Velocities)
1. [Creating the Initial Particle Positions](#Creating-the-Initial-Particle-Positions)
1. [Creating a Synthetic Radiograph](#Creating-a-Synthetic-Radiograph)
## Creating Particles
In this example we will create a source of 1e5 protons with a 5% variance in energy, a non-uniform angular velocity distribution, and a finite size.
```
nparticles = 1e5
particle = Particle("p+")
```
We will choose a setup in which the source-detector axis is parallel to the $y$-axis.
```
# define location of source and detector plane
source = (0 * u.mm, -10 * u.mm, 0 * u.mm)
detector = (0 * u.mm, 100 * u.mm, 0 * u.mm)
```
### Creating the Initial Particle Velocities
We will create the source distribution by utilizing the method of separation of variables,
$$f(v, \theta, \phi)=u(v)g(\theta)h(\phi)$$
and separately define the distribution component for each independent variable, $u(v)$, $g(\theta)$, and $h(\phi)$. For geometric convenience, we will generate the velocity vector distribution around the $z$-axis and then rotate the final velocities to be parallel to the source-detector axis (in this case the $y$-axis).
<img src="proton_radiography_source_profile_setup_graphic.png">
First we will create the orientation angles polar ($\theta$) and azimuthal ($\phi$) for each particle. Generating $\phi$ is simple: we will choose the azimuthal angles to just be uniformly distributed
```
phi = np.random.uniform(high=2 * np.pi, size=int(nparticles))
```
However, choosing $\theta$ is more complicated. Since the solid angle $d\Omega = sin \theta d\theta d\phi$, if we draw a uniform distribution of $\theta$ we will create a non-uniform distribution of particles in solid angle. This will create a sharp central peak on the detector plane.
```
theta = np.random.uniform(high=np.pi / 2, size=int(nparticles))
fig, ax = plt.subplots(figsize=(6, 6))
theta_per_sa, bins = np.histogram(theta, bins=100, weights=1 / np.sin(theta))
ax.set_xlabel("$\\theta$ (rad)", fontsize=14)
ax.set_ylabel("N/N$_0$ per d$\\Omega$", fontsize=14)
ax.plot(bins[1:], theta_per_sa / np.sum(theta_per_sa))
ax.set_title(f"N$_0$ = {nparticles:.0e}", fontsize=14)
ax.set_yscale("log")
ax.set_xlim(0, np.pi / 2)
ax.set_ylim(None, 1);
```
[np.random.choice()]: https://numpy.org/doc/stable/reference/random/generated/numpy.random.choice.html
To create a uniform distribution in solid angle, we need to draw values of $\theta$ with a probability distribution weighted by $\sin \theta$. This can be done using the [np.random.choice()] function, which draws `size` elements from a distribution `arg` with a probability distribution `prob`. Setting the `replace` keyword allows the same arguments to be drawn multiple times.
```
arg = np.linspace(0, np.pi / 2, num=int(1e5))
prob = np.sin(arg)
prob *= 1 / np.sum(prob)
theta = np.random.choice(arg, size=int(nparticles), replace=True, p=prob)
fig, ax = plt.subplots(figsize=(6, 6))
theta_per_sa, bins = np.histogram(theta, bins=100, weights=1 / np.sin(theta))
ax.plot(bins[1:], theta_per_sa / np.sum(theta_per_sa))
ax.set_xlabel("$\\theta$ (rad)", fontsize=14)
ax.set_ylabel("N/N$_0$ per d$\\Omega$", fontsize=14)
ax.set_title(f"N$_0$ = {nparticles:.0e}", fontsize=14)
ax.set_yscale("log")
ax.set_xlim(0, np.pi / 2)
ax.set_ylim(None, 0.1);
```
[np.random.choice()]: https://numpy.org/doc/stable/reference/random/generated/numpy.random.choice.html
[create_particles()]: ../../api/plasmapy.diagnostics.charged_particle_radiography.Tracker.rst#plasmapy.diagnostics.charged_particle_radiography.Tracker.create_particles
Now that we have a $\theta$ distribution that is uniform in solid angle, we can perturb it by adding additional factors to the probability distribution used in [np.random.choice()]. For this case, let's create a Gaussian distribution in solid angle.
Since particles moving at large angles will not be seen in the synthetic radiograph, we will set an upper bound $\theta_{max}$ on the argument here. This is equivalent to setting the `max_theta` keyword in [create_particles()]
```
arg = np.linspace(0, np.pi / 8, num=int(1e5))
prob = np.sin(arg) * np.exp(-(arg ** 2) / 0.1 ** 2)
prob *= 1 / np.sum(prob)
theta = np.random.choice(arg, size=int(nparticles), replace=True, p=prob)
fig, ax = plt.subplots(figsize=(6, 6))
theta_per_sa, bins = np.histogram(theta, bins=100, weights=1 / np.sin(theta))
ax.plot(bins[1:], theta_per_sa / np.sum(theta_per_sa))
ax.set_title(f"N$_0$ = {nparticles:.0e}", fontsize=14)
ax.set_xlabel("$\\theta$ (rad)", fontsize=14)
ax.set_ylabel("N/N$_0$ per d$\\Omega$", fontsize=14)
ax.set_yscale("log")
ax.set_xlim(0, np.pi / 2)
ax.set_ylim(None, 1);
```
Now that the angular distributions are done, we will determine the energy (speed) for each particle. For this example, we will assume that the particle energy distribution is not a function of angle. We will create a Gaussian distribution of speeds with ~5% variance centered on a particle energy of 15 MeV.
```
v_cent = np.sqrt(2 * 15 * u.MeV / particle.mass).to(u.m / u.s).value
v0 = np.random.normal(loc=v_cent, scale=1e6, size=int(nparticles))
v0 *= u.m / u.s
fig, ax = plt.subplots(figsize=(6, 6))
v_per_bin, bins = np.histogram(v0.si.value, bins=100)
ax.plot(bins[1:], v_per_bin / np.sum(v_per_bin))
ax.set_title(f"N$_0$ = {nparticles:.0e}", fontsize=14)
ax.set_xlabel("v0 (m/s)", fontsize=14)
ax.set_ylabel("N/N$_0$", fontsize=14)
ax.axvline(x=1.05 * v_cent, label="+5%", color="C1")
ax.axvline(x=0.95 * v_cent, label="-5%", color="C2")
ax.legend(fontsize=14, loc="upper right");
```
Next, we will construct velocity vectors centered around the z-axis for each particle.
```
vel = np.zeros([int(nparticles), 3]) * u.m / u.s
vel[:, 0] = v0 * np.sin(theta) * np.cos(phi)
vel[:, 1] = v0 * np.sin(theta) * np.sin(phi)
vel[:, 2] = v0 * np.cos(theta)
```
[rot_a_to_b()]: ../../api/plasmapy.formulary.mathematics.rot_a_to_b.rst#plasmapy.formulary.mathematics.rot_a_to_b
Finally, we will use the function [rot_a_to_b()] to create a rotation matrix that will rotate the `vel` distribution so the distribution is centered about the $y$ axis instead of the $z$ axis.
```
a = np.array([0, 0, 1])
b = np.array([0, 1, 0])
R = rot_a_to_b(a, b)
vel = np.matmul(vel, R)
```
Since the velocity vector distribution should be symmetric about the $y$ axis, we can confirm this by checking that the normalized average velocity vector is close to the $y$ unit vector.
```
avg_v = np.mean(vel, axis=0)
print(avg_v / np.linalg.norm(avg_v))
```
### Creating the Initial Particle Positions
For this example, we will create an initial position distribution representing a laser spot centered on the source location defined above as `source`. The distribution will be cylindrical (oriented along the $y$-axis) with a uniform distribution in y and a Gaussian distribution in radius (in the xz plane). We therefore need to create distributions in $y$, $\theta$, and $r$, and then transform those into Cartesian positions.
Just as we previously weighted the $\theta$ distribution with a $sin \theta$ probability distribution to generate a uniform distribution in solid angle, we need to weight the $r$ distribution with a $r$ probability distribution so that the particles are uniformly distributed over the area of the disk.
```
dy = 300 * u.um
y = np.random.uniform(
low=(source[1] - dy).to(u.m).value,
high=(source[1] + dy).to(u.m).value,
size=int(nparticles),
)
arg = np.linspace(1e-9, 1e-3, num=int(1e5))
prob = arg * np.exp(-((arg / 3e-4) ** 2))
prob *= 1 / np.sum(prob)
r = np.random.choice(arg, size=int(nparticles), replace=True, p=prob)
theta = np.random.uniform(low=0, high=2 * np.pi, size=int(nparticles))
x = r * np.cos(theta)
z = r * np.sin(theta)
hist, xpos, zpos = np.histogram2d(
x * 1e6, z * 1e6, bins=[100, 100], range=np.array([[-5e2, 5e2], [-5e2, 5e2]])
)
hist2, xpos2, ypos = np.histogram2d(
x * 1e6,
(y - source[1].to(u.m).value) * 1e6,
bins=[100, 100],
range=np.array([[-5e2, 5e2], [-5e2, 5e2]]),
)
fig, ax = plt.subplots(ncols=2, figsize=(12, 6))
fig.subplots_adjust(wspace=0.3, right=0.8)
fig.suptitle("Initial Particle Position Distribution", fontsize=14)
vmax = np.max([np.max(hist), np.max(hist2)])
p1 = ax[0].pcolormesh(xpos, zpos, hist.T, vmax=vmax)
ax[0].set_xlabel("x ($\\mu m$)", fontsize=14)
ax[0].set_ylabel("z ($\\mu m$)", fontsize=14)
ax[0].set_aspect("equal")
p2 = ax[1].pcolormesh(xpos2, ypos, hist2.T, vmax=vmax)
ax[1].set_xlabel("x ($\\mu m$)", fontsize=14)
ax[1].set_ylabel("y - $y_0$ ($\\mu m$)", fontsize=14)
ax[1].set_aspect("equal")
cbar_ax = fig.add_axes([0.85, 0.2, 0.03, 0.6])
cbar_ax.set_title("# Particles")
fig.colorbar(p2, cax=cbar_ax);
```
Finally we will combine these position arrays into an array with units.
```
pos = np.zeros([int(nparticles), 3]) * u.m
pos[:, 0] = x * u.m
pos[:, 1] = y * u.m
pos[:, 2] = z * u.m
```
## Creating a Synthetic Radiograph
To create an example synthetic radiograph, we will first create a field grid representing the analytical electric field produced by a sphere of Gaussian potential.
```
# Create a Cartesian grid
L = 1 * u.mm
grid = CartesianGrid(-L, L, num=100)
# Create a spherical potential with a Gaussian radial distribution
radius = np.linalg.norm(grid.grid, axis=3)
arg = (radius / (L / 3)).to(u.dimensionless_unscaled)
potential = 6e5 * np.exp(-(arg ** 2)) * u.V
# Calculate E from the potential
Ex, Ey, Ez = np.gradient(potential, grid.dax0, grid.dax1, grid.dax2)
mask = radius < L / 2
Ex = -np.where(mask, Ex, 0)
Ey = -np.where(mask, Ey, 0)
Ez = -np.where(mask, Ez, 0)
# Add those quantities to the grid
grid.add_quantities(E_x=Ex, E_y=Ey, E_z=Ez, phi=potential)
# Plot the E-field
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(111, projection="3d")
ax.view_init(30, 30)
# skip some points to make the vector plot intelligable
s = tuple([slice(None, None, 6)] * 3)
ax.quiver(
grid.pts0[s].to(u.mm).value,
grid.pts1[s].to(u.mm).value,
grid.pts2[s].to(u.mm).value,
grid["E_x"][s],
grid["E_y"][s],
grid["E_z"][s],
length=5e-7,
)
ax.set_xlabel("X (mm)", fontsize=14)
ax.set_ylabel("Y (mm)", fontsize=14)
ax.set_zlabel("Z (mm)", fontsize=14)
ax.set_xlim(-1, 1)
ax.set_ylim(-1, 1)
ax.set_zlim(-1, 1)
ax.set_title("Gaussian Potential Electric Field", fontsize=14);
```
We will then create the synthetic radiograph object. The warning filter ignores a warning that arises because $B_x$, $B_y$, $B_z$ are not provided in the grid (they will be assumed to be zero).
```
with warnings.catch_warnings():
warnings.simplefilter("ignore")
sim = cpr.Tracker(grid, source, detector, verbose=False)
```
[create_particles()]: ../../api/plasmapy.diagnostics.charged_particle_radiography.Tracker.rst#plasmapy.diagnostics.charged_particle_radiography.Tracker.create_particles
[load_particles()]: ../../api/plasmapy.diagnostics.charged_particle_radiography.Tracker.rst#plasmapy.diagnostics.charged_particle_radiography.Tracker.load_particles
Now, instead of using [create_particles()] to create the particle distribution, we will use the [load_particles()] function to use the particles we have created above.
```
sim.load_particles(pos, vel, particle=particle)
```
Now the particle radiograph simulation can be run as usual.
```
sim.run();
size = np.array([[-1, 1], [-1, 1]]) * 1.5 * u.cm
bins = [200, 200]
hax, vax, intensity = cpr.synthetic_radiograph(sim, size=size, bins=bins)
fig, ax = plt.subplots(figsize=(8, 8))
plot = ax.pcolormesh(
hax.to(u.cm).value, vax.to(u.cm).value, intensity.T, cmap="Blues_r", shading="auto",
)
cb = fig.colorbar(plot)
cb.ax.set_ylabel("Intensity", fontsize=14)
ax.set_aspect("equal")
ax.set_xlabel("X (cm), Image plane", fontsize=14)
ax.set_ylabel("Z (cm), Image plane", fontsize=14)
ax.set_title("Synthetic Proton Radiograph", fontsize=14);
```
[synthetic_radiograph()]: ../../api/plasmapy.diagnostics.charged_particle_radiography.synthetic_radiograph.rst#plasmapy.diagnostics.charged_particle_radiography.synthetic_radiograph
Calling the [synthetic_radiograph()] function with the `ignore_grid` keyword will produce the synthetic radiograph corresponding to the source profile propagated freely through space (i.e. in the absence of any grid fields).
```
hax, vax, intensity = cpr.synthetic_radiograph(
sim, size=size, bins=bins, ignore_grid=True
)
fig, ax = plt.subplots(figsize=(8, 8))
plot = ax.pcolormesh(
hax.to(u.cm).value, vax.to(u.cm).value, intensity.T, cmap="Blues_r", shading="auto",
)
cb = fig.colorbar(plot)
cb.ax.set_ylabel("Intensity", fontsize=14)
ax.set_aspect("equal")
ax.set_xlabel("X (cm), Image plane", fontsize=14)
ax.set_ylabel("Z (cm), Image plane", fontsize=14)
ax.set_title("Source Profile", fontsize=14);
fig, ax = plt.subplots(figsize=(6, 6))
ax.plot(hax.to(u.cm).value, np.mean(intensity, axis=0))
ax.set_xlabel("X (cm), Image plane", fontsize=14)
ax.set_ylabel("Mean intensity", fontsize=14)
ax.set_title("Mean source profile", fontsize=14);
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.
# Form Recognizer Scoring Pipeline
This notebook covers the process of setting up and running an AML pipeline that will score a custom form recognizer model.
Steps in this notebook include:
- Clapperboard Selection Step using OCR
- Form Extraction step (scoring)
- Postprocessing Step
## Import Dependencies
```
#Load dotenv extension
%load_ext dotenv
%dotenv
import os
from os.path import join
import sys
sys.path.append("../")
import pandas as pd
import numpy as np
from azureml.core import Environment, Datastore, Workspace, Experiment
from azureml.core.conda_dependencies import CondaDependencies
from azureml.data.data_reference import DataReference
from azureml.pipeline.core import Pipeline, PipelineData, PipelineParameter
from azureml.pipeline.steps import PythonScriptStep
from azureml.core.runconfig import RunConfiguration
from mlops.common.attach_compute import get_compute
from mlops.common.get_datastores import get_blob_datastore
```
## Configure Workspace and Set Compute Target
```
# Restore AML workspace from config.json file (can be downloaded through the portal)
ws = Workspace.from_config()
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep = '\n')
# Set compute target
compute_target = get_compute(
workspace=ws,
compute_name=os.getenv("AML_CLUSTER_NAME"),
vm_size=os.getenv("AML_CLUSTER_CPU_SKU"),
vm_priority=os.environ.get("AML_CLUSTER_PRIORITY", 'lowpriority'),
min_nodes=int(os.environ.get("AML_CLUSTER_MIN_NODES", 0)),
max_nodes=int(os.environ.get("AML_CLUSTER_MAX_NODES", 4)),
scale_down=int(os.environ.get("AML_CLUSTER_SCALE_DOWN", 600)),
)
```
## Configure Datastores
```
#create root datastore
#datastore should represent storage container where data will be accessed
root_datastore = get_blob_datastore(ws, os.getenv("BLOB_DATASTORE_NAME"), os.getenv("STORAGE_NAME"),
os.getenv("STORAGE_KEY"), os.getenv("STORAGE_CONTAINER"))
# Create input and output data references
# WARNING! DataReference works up to 12x times faster than Dataset
root_dir = DataReference(
datastore=root_datastore,
data_reference_name="form_data_ref",
mode="mount"
)
```
## Set Keyvault Secrets
```
#Set pipeline secrets using defauly keyvault every AML workspace comes with
print("Setting Pipeline Secrets in Azure Key Vault")
key_vault = ws.get_default_keyvault()
key_vault.set_secret(name="formkey", value=os.getenv('FORM_RECOGNIZER_KEY'))
key_vault.set_secret(name="formendpoint", value=os.getenv('FORM_RECOGNIZER_ENDPOINT'))
key_vault.set_secret(name="formmodelid", value=os.getenv('FORM_RECOGNIZER_CUSTOM_MODEL_ID'))
key_vault.set_secret(name="ocrkey", value=os.getenv('OCR_KEY'))
key_vault.set_secret(name="ocrendpoint", value=os.getenv('OCR_ENDPOINT'))
```
## Define Pipeline Parameters
```
# Just an example how we can use parameters to provide different input folders and values
input_dir = PipelineParameter(name="input_dir", default_value="val/clapperboard")
ocr_output_dir = PipelineParameter(name="ocr_output_dir", default_value="selected_clapperboards")
form_output_dir = PipelineParameter(name="form_output_dir", default_value="form_scoring_test_run")
form_labels = PipelineParameter(name="form_labels",
default_value="filename, roll, scene, take, title, director, camera, description")
```
## Build and set up dependencies for task-specific environment
```
# Build task-specific environment
from azureml.core.runconfig import RunConfiguration
from azureml.core.conda_dependencies import CondaDependencies
# Create Pipeline run configuration
run_config = RunConfiguration()
run_config.environment.docker.enabled = True
run_config.environment.python.conda_dependencies = CondaDependencies.create(
pip_packages=[
'argparse==1.4.0',
'azureml-sdk==1.18.0',
'azure-storage-blob==12.5.0',
'azure-identity==1.4.1',
'azure-mgmt-resource==10.2.0',
'azure-mgmt-network==16.0.0',
'azure-mgmt-compute==17.0.0',
'pyjwt==1.7.1',
'numpy==1.18.5',
'pandas==1.1.3',
'pillow==7.2.0',
'pyarrow==1.0.1',
'scikit-image==0.17.2',
'scikit-learn==0.23.2',
'scipy==1.5.2',
'tqdm==4.48.2',
'opencv-python-headless',
'tensorflow==2.3.0',
'azure-cognitiveservices-vision-customvision==3.0.0',
'PyYAML==5.3.1',
'ipywidgets==7.5.1',
'click==7.1.2',
'python-dotenv==0.10.3'
]
)
#specify docker image that will be used
dockerfile = r"""
FROM mcr.microsoft.com/azureml/intelmpi2018.3-ubuntu16.04:20200821.v1
"""
run_config.environment.docker.base_image = None
run_config.environment.docker.base_dockerfile = dockerfile
```
## Configure and instantiate pipeline steps
```
# Create and configure Form Recognizer Training pipeline steps
source_directory = "../"
clapperboard_selection_step = PythonScriptStep(
name="clapperboard selection",
script_name="mlops/form_scoring_pipeline/steps/select_clapperboards.py",
arguments=[
"--root_dir",
root_dir,
"--input_dir",
input_dir,
"--output_dir",
ocr_output_dir,
"--force",
True,
],
inputs=[root_dir],
outputs=[],
compute_target=compute_target,
source_directory=source_directory,
runconfig=run_config,
allow_reuse=False,
)
form_extraction_step = PythonScriptStep(
name="form_recognizer",
script_name="mlops/form_scoring_pipeline/steps/extract_forms.py",
arguments=[
"--root_dir",
root_dir,
"--input_dir",
input_dir,
"--clapperboard_dir",
ocr_output_dir,
"--output_dir",
form_output_dir,
"--labels",
form_labels,
"--force",
True,
],
inputs=[root_dir],
outputs=[],
compute_target=compute_target,
source_directory=source_directory,
runconfig=run_config,
allow_reuse=False,
)
form_extraction_step.run_after(clapperboard_selection_step)
form_postprocessing_step = PythonScriptStep(
name="postprocessing_form_recognizer",
script_name="mlops/form_scoring_pipeline/steps/postprocess.py",
arguments=[
"--root_dir",
root_dir,
"--input_dir",
form_output_dir,
"--output_dir",
form_output_dir,
"--force",
True,
],
inputs=[root_dir],
compute_target=compute_target,
source_directory=source_directory,
outputs=[],
runconfig=run_config,
allow_reuse=False,
)
form_postprocessing_step.run_after(form_extraction_step)
print("Pipeline Steps Created")
```
## Configure and publish pipeline to AML
```
# Create pipeline using existing steps
scoring_pipeline = Pipeline(workspace=ws, steps=[clapperboard_selection_step,
form_extraction_step,
form_postprocessing_step])
# Check if the pipeline is consistent
scoring_pipeline.validate()
# Publish pipeline
published_pipeline = scoring_pipeline.publish(
name = "form_scoring_pipeline",
description = "Pipeline to score a Custom Form Recognizer model"
)
```
## Submit and run pipeline in AML
```
# Submit the pipeline
pipeline_run = Experiment(ws, 'form-score-pipeline').submit(scoring_pipeline)
pipeline_run.wait_for_completion()
```
| github_jupyter |
# Clustering
In contrast to *supervised* machine learning, *unsupervised* learning is used when there is no "ground truth" from which to train and validate label predictions. The most common form of unsupervised learning is *clustering*, which is simllar conceptually to *classification*, except that the the training data does not include known values for the class label to be predicted. Clustering works by separating the training cases based on similarities that can be determined from their feature values. Think of it this way; the numeric features of a given entity can be though of as vector coordinates that define the entity's position in n-dimensional space. What a clustering model seeks to do is to identify groups, or *clusters*, of entities that are close to one another while being separated from other clusters.
For example, let's take a look at a dataset that contains measurements of different species of wheat seed.
> **Citation**: The seeds dataset used in the this exercise was originally published by the Institute of Agrophysics of the Polish Academy of Sciences in Lublin, and can be downloaded from the UCI dataset repository (Dua, D. and Graff, C. (2019). UCI Machine Learning Repository [http://archive.ics.uci.edu/ml]. Irvine, CA: University of California, School of Information and Computer Science).
```
import pandas as pd
# load the training dataset
data = pd.read_csv('data/seeds.csv')
# Display a random sample of 10 observations (just the features)
features = data[data.columns[0:6]]
features.sample(10)
```
As you can see, the dataset contains six data points (or *features*) for each instance (*observation*) of a seed. So you could interpret these as coordinates that describe each instance's location in six-dimensional space.
Now, of course six-dimensional space is difficult to visualise in a three-dimensional world, or on a two-dimensional plot; so we'll take advantage of a mathematical technique called *Principal Component Analysis* (PCA) to analyze the relationships between the features and summarize each observation as coordinates for two principal components - in other words, we'll translate the six-dimensional feature values into two-dimensional coordinates.
```
from sklearn.preprocessing import MinMaxScaler
from sklearn.decomposition import PCA
# Normalize the numeric features so they're on the same scale
scaled_features = MinMaxScaler().fit_transform(features[data.columns[0:6]])
# Get two principal components
pca = PCA(n_components=2).fit(scaled_features)
features_2d = pca.transform(scaled_features)
features_2d[0:10]
```
Now that we have the data points translated to two dimensions, we can visualize them in a plot:
```
import matplotlib.pyplot as plt
%matplotlib inline
plt.scatter(features_2d[:,0],features_2d[:,1])
plt.xlabel('Dimension 1')
plt.ylabel('Dimension 2')
plt.title('Data')
plt.show()
```
Hopefully you can see at least two, arguably three, reasonably distinct groups of data points; but here lies one of the fundamental problems with clustering - without known class labels, how do you know how many clusters to separate your data into?
One way we can try to find out is to use a data sample to create a series of clustering models with an incrementing number of clusters, and measure how tightly the data points are grouped within each cluster. A metric often used to measure this tightness is the *within cluster sum of squares* (WCSS), with lower values meaning that the data points are closer. You can then plot the WCSS for each model.
```
#importing the libraries
import numpy as np
import matplotlib.pyplot as plt
from sklearn.cluster import KMeans
%matplotlib inline
# Create 10 models with 1 to 10 clusters
wcss = []
for i in range(1, 11):
kmeans = KMeans(n_clusters = i)
# Fit the data points
kmeans.fit(features.values)
# Get the WCSS (inertia) value
wcss.append(kmeans.inertia_)
#Plot the WCSS values onto a line graph
plt.plot(range(1, 11), wcss)
plt.title('WCSS by Clusters')
plt.xlabel('Number of clusters')
plt.ylabel('WCSS')
plt.show()
```
The plot shows a large reduction in WCSS (so greater *tightness*) as the number of clusters increases from one to two, and a further noticable reduction from two to three clusters. After that, the reduction is less pronounced, resulting in an "elbow" in the chart at around three clusters. This is a good indication that there are two to three reasonably well separated clusters of data points.
## K-Means Clustering
The algorithm we used to create our test clusters is *K-Means*. This is a commonly used clustering algorithm that separates a dataset into *K* clusters of equal variance. The number of clusters, *K*, is user defined. The basic algorithm has the following steps:
1. A set of K centroids are randomly chosen.
2. Clusters are formed by assigning the data points to their closest centroid.
3. The means of each cluster is computed and the centroid is moved to the mean.
4. Steps 2 and 3 are repeated until a stopping criteria is met. Typically, the algorithm terminates when each new iteration results in negligable movement of centroids and the clusters become static.
5. When the clusters stop changing, the algorithm has *converged*, defining the locations of the clusters - note that the random starting point for the centroids means that re-running the algorithm could result in slightly different clusters, so training usually involves multiple iterations, reinitializing the centroids each time, and the model with the best WCSS is selected.
Let's try using K-Means on our seeds data with a K value of 3.
```
from sklearn.cluster import KMeans
# Create a model based on 3 centroids
model = KMeans(n_clusters=3, init='k-means++', n_init=100, max_iter=1000)
# Fit to the data and predict the cluster assignments for each data point
km_clusters = model.fit_predict(features.values)
# View the cluster assignments
km_clusters
```
Let's see those cluster assignments with the two-dimensional data points.
```
def plot_clusters(samples, clusters):
col_dic = {0:'blue',1:'green',2:'orange'}
mrk_dic = {0:'*',1:'x',2:'+'}
colors = [col_dic[x] for x in clusters]
markers = [mrk_dic[x] for x in clusters]
for sample in range(len(clusters)):
plt.scatter(samples[sample][0], samples[sample][1], color = colors[sample], marker=markers[sample], s=100)
plt.xlabel('Dimension 1')
plt.ylabel('Dimension 2')
plt.title('Assignments')
plt.show()
plot_clusters(features_2d, km_clusters)
```
Hopefully, the the data has been separated into three distinct clusters.
So what's the practical use of clustering? In some cases, you may have data that you need to group into distict clusters without knowing how many clusters there are or what they indicate. For example a marketing organization might want to separate customers into distinct segments, and then investigate how those segments exhibit different purchasing behaviors.
Sometimes, clustering is used as an initial step towards creating a classification model. You start by identifying distinct groups of data points, and then assign class labels to those clusters. You can then use this labelled data to train a classification model.
In the case of the seeds data, the different species of seed are already known and encoded as 0 (*Kama*), 1 (*Rosa*), or 2 (*Canadian*), so we can use these identifiers to compare the species classifications to the clusters identified by our unsupervised algorithm
```
seed_species = data[data.columns[7]]
plot_clusters(features_2d, seed_species.values)
```
There may be some differences between the cluster assignments and class labels, but the K-Means model should have done a reasonable job of clustering the observations so that seeds of the same species are generally in the same cluster.
## Hierarchical Clustering
Hierarchical clustering methods make fewer distributional assumptions when compared to K-means methods. However, K-means methods are generally more scalable, sometimes very much so.
Hierarchical clustering creates clusters by either a *divisive* method or *agglomerative* method. The divisive method is a "top down" approach starting with the entire dataset and then finding partitions in a stepwise manner. Agglomerative clustering is a "bottom up** approach. In this lab you will work with agglomerative clustering which roughly works as follows:
1. The linkage distances between each of the data points is computed.
2. Points are clustered pairwise with their nearest neighbor.
3. Linkage distances between the clusters are computed.
4. Clusters are combined pairwise into larger clusters.
5. Steps 3 and 4 are repeated until all data points are in a single cluster.
The linkage function can be computed in a number of ways:
- Ward linkage measures the increase in variance for the clusters being linked,
- Average linkage uses the mean pairwise distance between the members of the two clusters,
- Complete or Maximal linkage uses the maximum distance between the members of the two clusters.
Several different distance metrics are used to compute linkage functions:
- Euclidian or l2 distance is the most widely used. This metric is only choice for the Ward linkage method.
- Manhattan or l1 distance is robust to outliers and has other interesting properties.
- Cosine similarity, is the dot product between the location vectors divided by the magnitudes of the vectors. Notice that this metric is a measure of similarity, whereas the other two metrics are measures of difference. Similarity can be quite useful when working with data such as images or text documents.
### Agglomerative Clustering
Let's see an example of clustering the seeds data using an agglomerative clustering algorithm.
```
from sklearn.cluster import AgglomerativeClustering
agg_model = AgglomerativeClustering(n_clusters=3)
agg_clusters = agg_model.fit_predict(features.values)
agg_clusters
```
So what do the agglomerative cluster assignments look like?
```
import matplotlib.pyplot as plt
%matplotlib inline
def plot_clusters(samples, clusters):
col_dic = {0:'blue',1:'green',2:'orange'}
mrk_dic = {0:'*',1:'x',2:'+'}
colors = [col_dic[x] for x in clusters]
markers = [mrk_dic[x] for x in clusters]
for sample in range(len(clusters)):
plt.scatter(samples[sample][0], samples[sample][1], color = colors[sample], marker=markers[sample], s=100)
plt.xlabel('Dimension 1')
plt.ylabel('Dimension 2')
plt.title('Assignments')
plt.show()
plot_clusters(features_2d, agg_clusters)
```
In this notebook, you've explored clustering; an unsupervised form of machine learning.
To learn more about clustering with scikit-learn, see the [scikit-learn documentation](https://scikit-learn.org/stable/modules/clustering.html).
| github_jupyter |
# Fonctions
Une **fonction** est une séquence d'instructions nommée, qui est définie avant son utilisation, et qui peut être appelée multiple fois. Une fonction peut avoir des valeurs, et elle peut retourner des valeurs.
## Fonctions de conversion de type
Nous avons vu la fonction `type` qui retourne le type d'une valeur donnée, tel que:
* `int` (entier)
* `float` (virgule flottante)
* `str` (chaîne de caractères)
```
type(1), type(1.1), type('12')
```
Des **fonctions de conversion de type** permettent de changer un type de donnée vers un autre, si la forme le permet.
```
int(1.23), int('123')
float(1), float('1.23')
str(123), str(1.23)
```
## Fonctions mathématiques
Python seulement propose les 4 opérations arithmétique de base
* addition (+)
* soustraction (-)
* multiplication (*)
* division (/)
Ainsi que
* division entières (//)
* modulo (%)
Le module **math** fournit les operations trigonométriques.
```
import math
m = dir(math)
for m in dir(math): print(m, end=', ')
```
Nous y trouvons quelques constantes:
```
math.pi, math.e
```
Egalement les fonctions trigonométriques.
```
math.sin(1), math.cos(1)
math.sqrt(2)
math.factorial(30)
```
## Composition
Les fonctions et expressions peuvent être composé. Par exemple l'hypotenuse d'un triangle peut être calculée à partir de ses deux côtés.
```
a = 3
b = 4
math.sqrt(a**2 + b**2)
```
## Créer une nouvelle fonction
Dans Python il est possible de définir ses propore fonctions qui consistent d'un **en-tête** d'une ligne composé de:
* le mot-clé `def`
* un nom de fonction
* une paire de parenthèses `()`
* des arguments optionel
* un double-point `:`
L'en-tête de fonction est suivi du **corps**
* une ou multiple instructions
* indenté (de 4 espaces)
* terminé par le mot-clé `return` suivi d'une valeur de retour
Les règles pour former une nom de fonction: des lettres, des chiffres, un soulignement. Le première caractère ne peu pas être un chiffre.
Voici trois définitions de fonctions qui calcuent le diamètre, la circonférence et la surface d'un cercle à partir de son rayon **r**.
```
def diameter(r):
d = 2 * r
return d
def circonference(r):
c = 2 * r * math.pi
return c
def surface(r):
s = r**2 * math.pi
return s
```
Ces nouveaux fonctions peuvent être appelé avec un argument.
```
surface(2)
```
Les fonctions peuvent également être appelé à l'intérieur d'une fonction `print`.
```
r = 2
print('radius =', r)
print('diameter =', diameter(r))
print('circonference =', circonference(r))
print('surface =', surface(r))
surface = 12
surface
surface
```
Un retour d'indention termine la fonction. Par contre des lignes vides à l'interieur du corps peuvent servir pour séparer des sections.
```
def f():
print('hello')
print('world')
print('hello')
f()
```
## Définition et utilisation
L'effet d'une définition de fonction est de créer un objet fonction. La définition de fonction ne génère pas de sortie.
```
def surface(r):
s = r**2 * math.pi
return s
surface
```
Une fonction doit être défini plus haut dans le programme avant de pouvoir l'utiliser. On parle de **paramètre** dans la définition de la fonction, et d'**argument** dans l'appel de fonction.
```
surface(2)
```
## Paramètres et arguments
Certaines fonctions prennent un argument.
```
math.sin(1)
```
Certaines fonctions prennent deux arguments.
```
math.pow(2, 8)
```
Vous pouvez utiliser des variables comme argument
```
base = 2
exposant = 8
math.pow(base, exposant)
```
## Fonction productives et fonctions vides
Les fonctions qui retournent une valeur avec le mot-clé `return` sont appelé productives
```
def f_prod(a):
return a*5
f_prod(10)
f_prod('ha')
```
D'autre fonctions commme la fonction print ne retournent rien, mais ont un effet secondaire: imprimer quelque chose dans la console
```
def f_vide(a):
print('side effect =', a*5)
```
La fonction `f_vide` imprime comme effet secondaire vers la console, mais sa valeur de retour est `None`
```
print(f_prod(10), f_vide(10))
```
La valeur `None` n'est pas la même chose que la chaîne 'None'. La première est du type **NoneType**, la deuxième du type **str**
```
print(type(None), type('None'))
```
## Exercices
### Ex 1: aligner à droite
Ecriver une fonction `aligner_a_droite` qui prend comme paramètre une chaine nommée `s` et affiche avec suffisamment de caractères espaces pour que la dernière lettre de la chaine se trouve dans la colonne 70 de l'écran.
```
def aligner_a_droite(s):
n = 70
spaces = ' '*(n - len(s))
print(spaces+s)
aligner_a_droite('hello world')
aligner_a_droite('spam')
aligner_a_droite('Python is a programming language')
```
### Ex 2: afficher une carré
Imprimer une grille qui cons qui utilise seulement
* `print`
* la répétition de caractères `*`
* la concaténation de caractères `+`
* le caractère de retour à la ligne `\n`
Voici le code pour imprimer le première ligne
```
n = 3
print('+ ' + '- '*n + '+')
```
Voici le code pour imprimer la deuxième ligne:
```
print('| ' + ' '*n + '|')
```
Normalement `print` imprime une espace entre les arguments et termine avec un retour à la ligne.
```
print(1, 2, 3)
print(1, 2, 3)
```
En utilisant les paramètres nommés `sep` et `end` ces deux caractères peuvent être changé. Par exemple:
* `sep='++'` imprime deux + au lieu d'un espace
* `end='--'` imprime deux - au lieu d'un retour à la ligne
```
print(1, 2, 3, sep='++', end='--')
print(1, 2, 3, sep='++', end='--')
```
Voici la definition de la fonction pour imprimer un carré. Nous ajoutons le caractère de retour à la ligne, pour pouvoir imprimer toutes les lignes d'un seul coup.
```
def square(n):
line1 = '+ ' + '- '*n + '+\n'
line2 = '| ' + ' '*n + '|\n'
print(line1 + line2 * n + line1, end="")
square(2)
square(5)
```
### Ex 3: afficher une grille
```
def grid(n, m):
line1 = ('+ ' + '- '*n)*m + '+\n'
line2 = ('| ' + ' '*n)*m + '|\n'
print((line1 + line2 * n) * m + line1, end="")
grid(3, 2)
grid(2, 4)
```
| github_jupyter |
# PPO for transformer models
> A Pytorch implementation of Proximal Policy Optimization for transfomer models.
This follows the language model approach proposed in paper ["Fine-Tuning Language Models from Human Preferences"](
https://arxiv.org/pdf/1909.08593.pdf) and is similar to the [original implementation](https://github.com/openai/lm-human-preferences). The two main differences are 1) the method is implemented in Pytorch and 2) works with the `transformer` library by Hugging Face.
```
# default_exp ppo
# export
import numpy as np
import torch.nn.functional as F
from torch.optim import Adam
import torch
import collections
import time
import random
from trl.core import (logprobs_from_logits,
whiten,
clip_by_value,
entropy_from_logits,
flatten_dict,
average_torch_dicts,
stats_to_np,
stack_dicts,
add_suffix)
```
## KL-controllers
To ensure that the learned policy does not deviate to much from the original language model the KL divergence between the policy and a reference policy (the language model before PPO training) is used as an additional reward signal. Large KL-divergences are punished and staying close to the reference is rewarded.
Two controllers are presented in the paper: an adaptive log-space proportional controller and a fixed controller.
```
# exports
class AdaptiveKLController:
"""
Adaptive KL controller described in the paper:
https://arxiv.org/pdf/1909.08593.pdf
"""
def __init__(self, init_kl_coef, target, horizon):
self.value = init_kl_coef
self.target = target
self.horizon = horizon
def update(self, current, n_steps):
target = self.target
proportional_error = np.clip(current / target - 1, -0.2, 0.2)
mult = 1 + proportional_error * n_steps / self.horizon
self.value *= mult
# exports
class FixedKLController:
"""Fixed KL controller."""
def __init__(self, kl_coef):
self.value = kl_coef
def update(self, current, n_steps):
pass
# exports
class PPOTrainer:
"""
The PPO_trainer uses Proximal Policy Optimization to optimise language models.
"""
default_params = {
"lr": 1.41e-5,
"adap_kl_ctrl": True,
"init_kl_coef":0.2,
"target": 6,
"horizon":10000,
"gamma":1,
"lam":0.95,
"cliprange": .2,
"cliprange_value":.2,
"vf_coef":.1,
"batch_size": 256,
"forward_batch_size": 16,
"ppo_epochs": 4,
}
def __init__(self, model, ref_model, **ppo_params):
"""
Initialize PPOTrainer.
Args:
model (torch.model): Hugging Face transformer GPT2 model with value head
ref_model (torch.model): Hugging Face transformer GPT2 refrence model used for KL penalty
ppo_params (dict or None): PPO parameters for training. Can include following keys:
'lr' (float): Adam learning rate, default: 1.41e-5
'batch_size' (int): Number of samples per optimisation step, default: 256
'forward_batch_size' (int): Number of samples forward passed through model at a time, default: 16
'ppo_epochs' (int): Number of optimisation epochs per batch of samples, default: 4
'gamma' (float)): Gamma parameter for advantage calculation, default: 1.
'lam' (float): Lambda parameter for advantage calcualation, default: 0.95
'cliprange_value' (float): Range for clipping values in loss calculation, default: 0.2
'cliprange' (float): Range for clipping in PPO policy gradient loss, default: 0.2
'vf_coef' (float): Scaling factor for value loss, default: 0.1
'adap_kl_ctrl' (bool): Use adaptive KL control, otherwise linear, default: True
'init_kl_coef' (float): Initial KL penalty coefficient (used for adaptive and linear control), default: 0.2
'target' (float): Target KL value for adaptive KL control, default: 6.0
'horizon' (float): Horizon for adaptive KL control, default: 10000
"""
self.ppo_params = self.default_params
self.ppo_params.update(ppo_params)
self.ref_model = ref_model
self.model = model
self.optimizer = Adam(model.parameters(), lr=self.ppo_params['lr'])
self.kl_ctl = AdaptiveKLController(self.ppo_params['init_kl_coef'],
self.ppo_params['target'],
self.ppo_params['horizon'])
def step(self, query, response, scores):
"""
Run a PPO optimisation step.
args:
query (torch.tensor): tensor containing the encoded queries, shape [batch_size, query_length]
response (torch.tensor): tensor containing the encoded responses, shape [batch_size, response_length]
scores (torch.tensor): tensor containing the scores, shape [batch_size]
returns:
train_stats (dict): a summary of the training statistics
"""
bs = self.ppo_params['batch_size']
timing = dict()
t0 = time.time()
gen_len = response.shape[1]
model_input = torch.cat((query, response), axis=1)
t = time.time()
logprobs, ref_logprobs, values = self.batched_forward_pass(model_input, gen_len)
timing['time/ppo/forward_pass'] = time.time()-t
t = time.time()
rewards, non_score_reward, kl_coef = self.compute_rewards(scores, logprobs, ref_logprobs)
timing['time/ppo/compute_rewards'] = time.time()-t
t = time.time()
all_stats = []
idxs = list(range(bs))
for _ in range(self.ppo_params['ppo_epochs']):
random.shuffle(idxs)
for i in range(bs):
idx = idxs[i]
train_stats = self.train_minibatch(logprobs[idx:idx+1], values[idx:idx+1],
rewards[idx:idx+1], query[idx:idx+1],
response[idx:idx+1], model_input[idx:idx+1])
all_stats.append(train_stats)
timing['time/ppo/optimize_step'] = time.time()-t
t = time.time()
train_stats = stack_dicts(all_stats)
# reshape advantages/ratios such that they are not averaged.
train_stats['policy/advantages'] = torch.flatten(train_stats['policy/advantages']).unsqueeze(0)
train_stats['policy/ratio'] = torch.flatten(train_stats['policy/ratio']).unsqueeze(0)
stats = self.record_step_stats(scores=scores, logprobs=logprobs, ref_logprobs=ref_logprobs,
non_score_reward=non_score_reward, train_stats=train_stats,
kl_coef=kl_coef)
stats = stats_to_np(stats)
timing['time/ppo/calc_stats'] = time.time()-t
self.kl_ctl.update(stats['objective/kl'], self.ppo_params['batch_size'])
timing['time/ppo/total'] = time.time()-t0
stats.update(timing)
return stats
def batched_forward_pass(self, model_input, gen_len):
"""Calculate model outputs in multiple batches."""
bs = self.ppo_params['batch_size']
fbs = self.ppo_params['forward_batch_size']
logprobs = []
ref_logprobs = []
values = []
for i in range(int(self.ppo_params['batch_size']/fbs)):
m_input = model_input[i*fbs:(i+1)*fbs]
logits, _, v = self.model(m_input)
ref_logits, _, _ = self.ref_model(m_input)
values.append(v[:, -gen_len-1:-1].detach())
logprobs.append(logprobs_from_logits(logits[:,:-1,:], m_input[:,1:])[:, -gen_len:].detach())
ref_logprobs.append(logprobs_from_logits(ref_logits[:,:-1,:], m_input[:,1:])[:, -gen_len:].detach())
return torch.cat(logprobs), torch.cat(ref_logprobs), torch.cat(values)
def train_minibatch(self, logprobs, values, rewards, query, response, model_input):
"""Train one PPO minibatch"""
loss_p, loss_v, train_stats = self.loss(logprobs, values, rewards, query, response, model_input)
loss = loss_p + loss_v
self.optimizer.zero_grad()
loss.backward()
self.optimizer.step()
return train_stats
def compute_rewards(self, scores, logprobs, ref_logprobs):
"""Compute per token rewards from scores and KL-penalty."""
kl = logprobs - ref_logprobs
non_score_reward = -self.kl_ctl.value * kl
rewards = non_score_reward.clone().detach()
rewards[:, -1] += scores
return rewards, non_score_reward, self.kl_ctl.value
def loss(self, old_logprobs, values, rewards, query, response, model_input):
"""Calculate policy and value losses."""
lastgaelam = 0
advantages_reversed = []
gen_len = response.shape[1]
for t in reversed(range(gen_len)):
nextvalues = values[:, t + 1] if t < gen_len - 1 else 0.0
delta = rewards[:, t] + self.ppo_params['gamma'] * nextvalues - values[:, t]
lastgaelam = delta + self.ppo_params['gamma'] * self.ppo_params['lam'] * lastgaelam
advantages_reversed.append(lastgaelam)
advantages = torch.stack(advantages_reversed[::-1]).transpose(0, 1)
returns = advantages + values
advantages = whiten(advantages)
advantages = advantages.detach()
logits, _, vpred = self.model(model_input)
logprob = logprobs_from_logits(logits[:,:-1,:], model_input[:, 1:])
#only the generation part of the values/logprobs is needed
logprob, vpred = logprob[:, -gen_len:], vpred[:,-gen_len-1:-1]
vpredclipped = clip_by_value(vpred,
values - self.ppo_params["cliprange_value"],
values + self.ppo_params["cliprange_value"])
vf_losses1 = (vpred - returns)**2
vf_losses2 = (vpredclipped - returns)**2
vf_loss = .5 * torch.mean(torch.max(vf_losses1, vf_losses2))
vf_clipfrac = torch.mean(torch.gt(vf_losses2, vf_losses1).double())
ratio = torch.exp(logprob - old_logprobs)
pg_losses = -advantages * ratio
pg_losses2 = -advantages * torch.clamp(ratio,
1.0 - self.ppo_params['cliprange'],
1.0 + self.ppo_params['cliprange'])
pg_loss = torch.mean(torch.max(pg_losses, pg_losses2))
pg_clipfrac = torch.mean(torch.gt(pg_losses2, pg_losses).double())
loss = pg_loss + self.ppo_params['vf_coef'] * vf_loss
entropy = torch.mean(entropy_from_logits(logits))
approxkl = .5 * torch.mean((logprob - old_logprobs)**2)
policykl = torch.mean(logprob - old_logprobs)
return_mean, return_var = torch.mean(returns), torch.var(returns)
value_mean, value_var = torch.mean(values), torch.var(values)
stats = dict(
loss=dict(policy=pg_loss, value=vf_loss, total=loss),
policy=dict(entropy=entropy, approxkl=approxkl,policykl=policykl, clipfrac=pg_clipfrac,
advantages=advantages, advantages_mean=torch.mean(advantages), ratio=ratio),
returns=dict(mean=return_mean, var=return_var),
val=dict(vpred=torch.mean(vpred), error=torch.mean((vpred - returns) ** 2),
clipfrac=vf_clipfrac, mean=value_mean, var=value_var),
)
return pg_loss, self.ppo_params['vf_coef'] * vf_loss, flatten_dict(stats)
def record_step_stats(self, kl_coef, **data):
"""Record training step statistics."""
kl = data['logprobs'] - data['ref_logprobs']
mean_kl = torch.mean(torch.sum(kl, axis=-1))
mean_entropy = torch.mean(torch.sum(-data['logprobs'], axis=1))
mean_non_score_reward =torch.mean(torch.sum(data['non_score_reward'], axis=1))
stats = {
'objective/kl': mean_kl,
'objective/kl_dist': kl,
'objective/logprobs': data['logprobs'],
'objective/ref_logprobs': data['ref_logprobs'],
'objective/kl_coef': kl_coef,
'objective/entropy': mean_entropy,
'ppo/mean_non_score_reward': mean_non_score_reward,
}
for k, v in data['train_stats'].items():
stats[f'ppo/{k}'] = torch.mean(v, axis=0)
stats['ppo/val/var_explained'] = 1 - stats['ppo/val/error'] / stats['ppo/returns/var']
return stats
```
## Tensor shapes and contents
Debugging tensor shapes and contents usually involves inserting a lot of print statements in the code. To avoid this in the future I add a list of the tensor shapes and contents for reference. If the tensors are sliced or reshaped I list the last shape.
| Name | Shape | Content |
|-------|---------|---------|
| `query` | `[batch_size, query_length]`| contains token ids of query|
| `response`| `[batch_size, response_length]`| contains token ids of responses|
| `scores`| `[batch_size]`| rewards of each query/response pair|
| `model_input`| `[batch_size, query_length + response_length]`| combined query and response tokens|
| `m_input`|`[forward_batch_size, query_length + response_length]`| small forward batch of model_input|
| `logits` | `[forward_batch_size, query_length + response_length, vocab_size]`| logits from model outputs|
| `ref_logits`|`[forward_batch_size, query_length + response_length, vocab_size]`| logits from ref_model outputs|
| `logprobs`| `[batch_size, response_length]`| log-probabilities of response tokens |
| `ref_logprobs`| `[batch_size, response_length]`| reference log-probabilities of response tokens |
| `rewards`| `[batch_size, response_length]`| the model rewards incl. kl-score for each token|
| `non_score_reward`| `[batch_size, response_length]`| the model kl-score for each token|
## Model output alignments
Some notes on output alignments, since I spent a considerable time debugging this. All model outputs are shifted by 1 to the model inputs. That means that the logits are shifted by one as well as values. For this reason the logits and values are always shifted one step to the left. This also means we don't have logits for the first input element and so we delete the first input token when calculating the softmax, since we don't have logits predictions. The same applies for the values and we shift them by index one to the left.
## KL-divergence
One question that came up during the implementation was "Why is the KL-divergence just the difference of the log-probs? Where is the probability in front of the log term?". The answer can be found in Sergey Levine's [lecture slides](http://rll.berkeley.edu/deeprlcourse/docs/week_3_lecture_1_dynamics_learning.pdf): To calculate the KL divergence we calculate the expected value of the log term. The probability usually in front of the log-term comes from that expected value and for a set of trajectories we can simply take the mean over the sampled trajectories.
| github_jupyter |
# Welter issue #9
## Generate synthetic, noised-up two-temperature model spectra, then naively fit a single temperature model to it.
### Part 3- Iterate
Michael Gully-Santiago
Friday, January 8, 2015
We now need to iterate.
```
import warnings
warnings.filterwarnings("ignore")
import numpy as np
from astropy.io import fits
import matplotlib.pyplot as plt
% matplotlib inline
% config InlineBackend.figure_format = 'retina'
import seaborn as sns
sns.set_context('notebook')
```
See the previous notebook for the theory and background.
```
import os
import json
import pandas as pd
import yaml
```
## Make mixture model
Assumes you have run splot.py and renamed the model json file to model_a and model_b.
```
def make_mixture_model():
try:
with open('model_A_spec.json') as f:
ma_raw = json.load(f)
with open('model_B_spec.json') as f:
mb_raw = json.load(f)
except FileNotFoundError:
print("You need to have the model_A and model_B spectral json files in place first.")
r_factor = 1.0/2.39894
wl = np.array(ma_raw['wl'])
fl_ma = np.array(ma_raw['model'])
nd_ma = np.array(ma_raw['noise_draw'])
fl_mb_raw = np.array(mb_raw['model'])
nd_mb_raw = np.array(mb_raw['noise_draw'])
#Scale the flux (and noise spectrum) by the cofactor $r$ as discussed above.
fl_mb = fl_mb_raw * r_factor
nd_mb = nd_mb_raw * r_factor
# Mixture model
c = 0.70
mmix = c * fl_ma + (1 - c) * fl_mb
ndmix = 0.5 * (c * nd_ma + (1 - c) * nd_mb) # Remove the factor of 2.
mmix_noised = mmix + ndmix
return {"wl":wl.tolist(), "mmix":mmix.tolist(), "mmix_noised":mmix_noised.tolist()}
```
## Automate the mixture model making
```
os.chdir('/Users/gully/GitHub/welter/notebooks/')
for i in range(26, 27):
print(i)
os.chdir('../sf/eo{:03d}/'.format(i))
# Make model A
with open('config.yaml', mode='r') as f:
config = yaml.load(f)
config['Theta']['grid'] = [4100.0, 3.5, 0.0]
with open('config.yaml', mode='w') as f:
f.write(yaml.dump(config))
os.system('star.py --generate')
os.system('splot.py s0_o0spec.json --matplotlib --noise --save_draw')
os.system('mv model_spec.json model_A_spec.json')
# Make model B
with open('config.yaml', mode='r') as f:
config = yaml.load(f)
config['Theta']['grid'] = [3300.0, 3.5, 0.0]
with open('config.yaml', mode='w') as f:
f.write(yaml.dump(config))
os.system('star.py --generate')
os.system('splot.py s0_o0spec.json --matplotlib --noise --save_draw')
os.system('mv model_spec.json model_B_spec.json')
# Make mixture model
my_dict = make_mixture_model()
# Save mixture model
with open("mixture_model.json", mode='w') as f:
json.dump(my_dict, f, indent=2, sort_keys=True)
os.chdir('/Users/gully/GitHub/welter/notebooks/')
```
| github_jupyter |
# 4 Modeling<a id='4'></a>
## 4.1 Contents<a id='4.1'></a>
* [4 Preprocessing Data](#4)
* [4.1 Contents](#4.1)
* [4.2 Introduction](#4.2)
* [4.3 Imports](#4.3)
* [4.4 Load the Cleaned Data](#4.4)
* [4.5 Train Test Split](#4.5)
* [4.6 Preprocessing Columns](#4.6)
* [4.7 Define Column groups](#4.7)
* [4.6.1 Column Selector](#4.6.1)
* [4.6.2 Mode Imputer](#4.6.2)
* [4.7 Pipeline](#4.7)
* [4.7.1 Categorical Pipeline](#4.7.1)
* [4.7.2 Ordinal Pipeline](#4.7.2)
* [4.7.3 Age Distance Pipeline](#4.7.3)
* [4.7.4 Wait Times Pipeline](#4.7.4)
* [4.8 Feature union](#4.8)
* [4.9 Checking Steps](#4.9)
* [4.9.1 Fit Transform Training Data Check](#4.9.1)
* [4.9.2 Transform Test Data Check](#4.9.2)
* [4.10 Modeling](#4.10)
* [4.10.1 Baseline Model](#4.10.1)
* [4.10.2 Logistic Regression Model](#4.10.2)
## 4.2 Introduction<a id='4.2'></a>
We've divided the data into Categorical, Ordinal Continuous and Continuous Variables
## 4.3 Imports<a id='4.3'></a>
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn import __version__ as sklearn_version
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import RandomizedSearchCV
from sklearn.model_selection import GridSearchCV
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.preprocessing import OneHotEncoder, MinMaxScaler, StandardScaler, PowerTransformer
from sklearn.impute import SimpleImputer
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn.dummy import DummyClassifier
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score
from sklearn.metrics import roc_curve
from sklearn.metrics import roc_auc_score
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
import pickle
import datetime
```
#### 4.4 Load the Cleaned data<a id='4.4'></a>
```
raw_data_clean = pd.read_csv('../data/interim/raw_data_clean.csv')
#Preview the data
pd.set_option('display.max_columns', None)
with pd.option_context('display.max_rows',10):
display(raw_data_clean)
```
## 4.5 Train Test Split<a id='4.5'></a>
```
X_train, X_test, y_train, y_test = train_test_split(raw_data_clean.drop(columns = ['satisfaction', 'id']),
raw_data_clean.satisfaction,
test_size = 0.2,
random_state = 42)
print(X_train.shape, y_train.shape)
print(X_test.shape, y_test.shape)
```
## 4.6 Preprocessing Columns<a id='4.6'></a>
We need to create pipelines for the categorical columns, ordinal columns and continuous columns since each one of them will have different transformations
```
categorical_cols = ['Gender', 'Type of Travel', 'Class', 'Customer Type']
ordinal_cols = list(raw_data_clean.loc[:, 'Inflight wifi service': 'Cleanliness'].columns)
continuous_cols_1 = ['Age', 'Flight Distance']
continuous_cols_2 = ['Departure Delay in Minutes', 'Arrival Delay in Minutes']
```
### 4.6.1 Column Selector<a id='4.6.1'></a>
```
# Define custom transformer
class ColumnSelector(BaseEstimator, TransformerMixin):
"""Select only specified columns."""
def __init__(self, columns):
self.columns = columns
def fit(self, X, y=None):
return self
def transform(self, X):
return X[self.columns]
```
### 4.6.2 Mode Imputer<a id='4.6.2'></a>
We need to define a mode imputer to impute Categorical variables with the most frequent occurance
This solution has been adapted from the stackoverflow link: <a href = 'https://stackoverflow.com/questions/25239958/impute-categorical-missing-values-in-scikit-learn'>Mode Imputer Stack Over Flow</a>
```
class ModeImputer(TransformerMixin):
def __init__(self):
"""Impute missing values.
Columns of dtype object are imputed with the most frequent value
in column.
Columns of other types are imputed with mean of column.
"""
def fit(self, X, y=None):
self.fill = pd.Series([X[c].value_counts().index[0]
if X[c].dtype == np.dtype('O') else X[c].mean() for c in X],
index=X.columns)
return self
def transform(self, X, y=None):
return X.fillna(self.fill)
```
## 4.7 Pipeline<a id='4.7'></a>
### 4.7.1 Categorical Pipeline<a id='4.7.1'></a>
```
cat_pipe = Pipeline([('selector', ColumnSelector(categorical_cols)),
('imputer', ModeImputer()),
('encoder', OneHotEncoder(handle_unknown='ignore', sparse=False))])
```
### 4.7.2 Ordinal Pipeline<a id='4.7.2'></a>
```
ord_pipe = Pipeline([('selector', ColumnSelector(ordinal_cols)),
('imputer', SimpleImputer(strategy = 'constant', fill_value = 0)),
('scaler', MinMaxScaler())])
```
### 4.7.3 Continous Cols Age Distance Pipeline<a id='4.7.3'></a>
```
cont_1_pipe = Pipeline([('selector', ColumnSelector(continuous_cols_1)),
('imputer', SimpleImputer(strategy = 'mean')),
('scaler', StandardScaler())])
```
#### 4.7.4 Continous Cols Wait Times Pipeline<a id='4.7.4'></a>
```
cont_2_pipe = Pipeline([('selector', ColumnSelector(continuous_cols_2)),
('imputer', SimpleImputer(strategy = 'constant', fill_value = 0)),
('scaler', PowerTransformer())])
```
## 4.8 Feature union<a id='4.8'></a>
```
# Fit feature union to training data
preprocessor = FeatureUnion(transformer_list=[('cat', cat_pipe),
('ord', ord_pipe),
('cont_1', cont_1_pipe),
('cont_2', cont_2_pipe)])
preprocessor.fit(X_train)
```
## 4.9 Checking Steps<a id='4.9'></a>
```
# Prepare column names
cat_columns = preprocessor.transformer_list[0][1]['encoder'].get_feature_names(categorical_cols)
columns = np.append(np.append(np.append(cat_columns, ordinal_cols), continuous_cols_1), continuous_cols_2)
```
### 4.9.1 Fit Transform Training Data Check<a id='4.9.1'></a>
```
# Inspect training data before and after
print("******************** Training data ********************")
display(X_train.head())
display(pd.DataFrame(preprocessor.transform(X_train), columns=columns).head())
```
### 4.9.2 Transform Test Data Check<a id='4.9.2'></a>
```
# Inspect test data before and after
print("******************** Test data ********************")
display(X_test.head())
print("******************** Transformed Test data ********************")
display(pd.DataFrame(preprocessor.transform(X_test), columns=columns).head())
```
## 4.10 Modeling<a id='4.10'></a>
### 4.10.1 Baseline Model<a id='4.10.1'></a>
```
# Combine categorical and numerical pipeline
pipe = Pipeline(steps=[('preprocessor', preprocessor),
('model', DummyClassifier())])
pipe.fit(X_train, y_train)
# Predict training data
y_train_pred = pipe.predict(X_train)
print(f"Predictions on training data: {y_train_pred}")
# Predict test data
y_test_pred = pipe.predict(X_test)
print(f"Predictions on test data: {y_test_pred}")
print(confusion_matrix(y_test, y_test_pred))
print(f'Baseline Model score {round(accuracy_score(y_test, y_test_pred) * 100, 2)}%' )
```
### 4.10.2 LogisticRegression Model<a id='4.10.2'></a>
```
# Combine categorical and numerical pipeline
pipe_logreg = Pipeline(steps=[('preprocessor', preprocessor),
('logreg', LogisticRegression())])
pipe_logreg.fit(X_train, y_train)
# Predict training data
y_train_pred = pipe_logreg.predict(X_train)
print(f"Predictions on training data: {y_train_pred}")
# Predict test data
y_test_pred = pipe_logreg.predict(X_test)
print(f"Predictions on test data: {y_test_pred}")
print(confusion_matrix(y_test, y_test_pred))
print(f'Logistic Regression Model score {round(accuracy_score(y_test, y_test_pred) * 100, 2)}%' )
print(classification_report(y_test, y_test_pred))
```
### Model function
```
# Combine categorical and numerical pipeline
def tune_clf_func(model, X_train, X_test, y_test, y_train, param_grid):
pipe = Pipeline(steps=[('preprocessor', preprocessor), ('model', model)])
grid_cv = GridSearchCV(pipe, param_grid, cv = 5, n_jobs = -1, scoring = 'roc_auc')
grid_cv.fit(X_train, y_train)
# Print the tuned parameters and score
print('----------------------Hyper Parameter Tuning---------------------------\n')
print(f"Tuned {model} Parameters: {grid_cv.best_params_}")
print(f"Best score is {grid_cv.best_score_}")
print('-----------------------------------------------------------------------\n')
best_model = grid_cv.best_estimator_
best_model.fit(X_train, y_train)
# Predict training data
y_train_pred = best_model.predict(X_train)
print(f"Predictions on training data: {y_train_pred}")
# Predict test data
y_pred = best_model.predict(X_test)
print(f"Predictions on test data: {y_pred}")
print(confusion_matrix(y_test, y_pred))
print(f'{best_model} Model AUC score {round(roc_auc_score(y_test, y_pred) * 100, 2)}%' )
print(classification_report(y_test, y_pred))
return best_model, round(roc_auc_score(y_test, y_pred) * 100, 4)
```
#### 4.10.2.1 Hyper Parameter Tuning Logistic Regression Model<a id='4.10.3.1'></a>
```
scores = {}
c_space = [100, 10, 1.0, 0.1, 0.01]
penalty = ['l2']
param_grid = {'model__C': c_space, 'model__max_iter': [1000], 'model__penalty':penalty}
best_model, score = tune_clf_func(LogisticRegression(), X_train, X_test, y_test, y_train, param_grid)
scores[str(best_model[1])] = score
scores
```
#### 4.10.2.2 Hyper Parameter Tuning KNN Model<a id='4.10.3.2'></a>
```
n_neighbors = np.arange(1, 9)
param_grid = {'model__n_neighbors': n_neighbors}
best_model, score = tune_clf_func(KNeighborsClassifier(), X_train, X_test, y_test, y_train, param_grid)
scores[str(best_model[1])] = score
```
#### 4.10.2.3 Hyper Parameter Tuning RandomForest Model<a id='4.10.3.3'></a>
```
n_estimators = [10, 100, 300, 500]
max_depth = [None, 5, 8, 15, 25, 30]
min_samples_split = [2, 5, 10, 15, 100]
min_samples_leaf = [1, 2, 5, 10]
param_grid = {'model__n_estimators': n_estimators, 'model__max_depth': max_depth, 'model__min_samples_split': min_samples_split,
'model__min_samples_leaf': min_samples_leaf}
best_model, score = tune_clf_func(RandomForestClassifier(), X_train, X_test, y_test, y_train, param_grid)
scores[str(best_model[1])] = score
scores
```
### 4.11 Final Model<a id='4.11'></a>
```
X = raw_data_clean.drop(columns = ['satisfaction', 'id'])
y = raw_data_clean.satisfaction
n_estimators = [500]
max_depth = [30]
min_samples_split = [5]
param_grid = {'model__n_estimators': n_estimators, 'model__max_depth': max_depth, 'model__min_samples_split': min_samples_split}
pipe_final = Pipeline(steps=[('preprocessor', preprocessor), ('model', RandomForestClassifier())])
grid_cv_final = GridSearchCV(pipe_final, param_grid, cv = 5, n_jobs = -1, scoring = 'roc_auc')
grid_cv_final.fit(X, y)
#Let's call this model version '1.0'
best_model = grid_cv_final.best_estimator_
best_model.version = '1.0'
best_model.pandas_version = pd.__version__
best_model.numpy_version = np.__version__
best_model.sklearn_version = sklearn_version
best_model.X_columns = [col for col in X_train.columns]
best_model.build_datetime = datetime.datetime.now()
filename = '../models/predict_customer_sat_model.pkl'
pickle.dump(best_model, open(filename, 'wb'))
```
| github_jupyter |
# Including Bus Routes
Purpose: Identify bus stop locations along routes in order to estimate the vehicle drive cycle.
Work Flow: Isolate bus stops along desired route. Calculate distance between route points and bus stops.
```
import os
import sys
module_path = os.path.abspath(os.path.join('..'))
sys.path.append(module_path)
import route_dynamics.route_elevation.base as base
import geopandas as gpd
import pandas as pd
import branca.colormap as cm
import folium
import rasterstats
import matplotlib.pyplot as plt
from folium.features import GeoJson
from shapely.geometry import mapping
from shapely.geometry import LineString
from shapely.geometry import Polygon
from rasterio.mask import mask
from geopy.distance import geodesic
```
First, input data and select bus route.
```
routes_shp = '../data/six_routes.shp'
stops_shp = '../data/Transit_Stops_for_King_County_Metro__transitstop_point.shp'
route_num = 45
```
Use base functions, `read_shape` and `extract_point_df` to create a dataframe of route points.
```
route = base.read_shape(routes_shp, route_num)
points = base.extract_point_df(route)
points.head()
```
Isolate bus stops that service desired route number.
```
stops = gpd.read_file(stops_shp)
```
First, need to remove some pesky 'none' types...
```
stops['ROUTE_LIST'].fillna(value=str(0), inplace=True)
#stops['ROUTE_LIST'].head(10)
stops_list = pd.DataFrame()
for i in range(0, len(stops)):
if str(route_num) in (stops['ROUTE_LIST'][i]):
for x in stops['ROUTE_LIST'][i].split(' '):
if str(route_num) == x:
stops_list = stops_list.append(stops.iloc[i])
else:
pass
#stops_list = pd.concat([stops_list, stops.iloc[i]], axis = 0)
else:
pass
stops_list.head()
```
Create a dataframe of bus stop points.
```
geometry = stops_list.geometry.values
xy = []
for i in range(len(geometry)):
dic = mapping(geometry[i])
coords = dic['coordinates']
xy.append(coords)
xy_df = pd.DataFrame()
xy_df['coordinates'] = xy
xy_df.head()
def make_lines(gdf, idx):
coordinate_1 = gdf.loc[idx]['coordinates']
coordinate_2 = gdf.loc[idx + 1]['coordinates']
line = LineString([coordinate_1, coordinate_2])
data = {'geometry':[line]}
df_line = pd.DataFrame(data, columns = ['geometry'])
return df_line
def make_route_lines(gdf):
df_route = pd.DataFrame(columns = ['geometry'])
for idx in range(len(gdf) - 1):
df_linestring = make_lines(gdf, idx)
df_route = pd.concat([df_route, df_linestring])
gdf_route = gpd.GeoDataFrame(df_route)
return gdf_route
gdf_route = make_route_lines(points)
gdf_stops = pd.DataFrame(xy, columns = ['Lat','Long'])
```
Show map:
```
UW_coords = [47.655548, -122.303200]
figure_size = folium.Figure(height = 400)
route_map = folium.Map(location = UW_coords, zoom_start = 12)
route_json = gdf_route.to_json()
route_layer = folium.GeoJson(route_json)
route_layer.add_child
route_map.add_child(route_layer)
route_map.add_to(figure_size)
fg = folium.FeatureGroup()
for i in range(len(gdf_stops)):
fg.add_child(folium.Circle(location=[gdf_stops['Long'][i],gdf_stops['Lat'][i]], radius=8, color='red'))
route_map.add_child(fg)
route_map
```
| github_jupyter |
# Exploratory Data Analysis (EDA) for Natural Language Processing using WordCloud
## What is WordCloud?
<img src="https://altoona.psu.edu/sites/default/files/styles/photo_gallery_large/http/news.psu.edu/sites/default/files/success-word-cloud.jpg?itok=4_HTmhRg+" alt="Drawing" style="width: 600;"/>
Many times you might have seen a cloud filled with lots of words in different sizes, which represent the frequency or the importance of each word. This is called [Tag Cloud](https://en.wikipedia.org/wiki/Tag_cloud) or WordCloud. For this tutorial, you will learn how to create a WordCloud of your own in Python and customize it as you see fit. This tool will be quite handy for exploring text data and making your report more lively.
In this tutorial we will use a wine review dataset taking from [Wine Enthusiast website](https://www.winemag.com/?s=&drink_type=wine) to learn:
- How to create a basic wordcloud from one to several text document
- Adjust color, size and number of text inside your wordcloud
- Mask your wordcloud into any shape of your choice
- Mask your wordcloud into any color pattern of your choice

### Prerequisites
You will need to install some packages below:
- [numpy](http://www.numpy.org/)
- [pandas](https://pandas.pydata.org/)
- [matplotlib](https://matplotlib.org/index.html)
- [pillow](https://pillow.readthedocs.io/en/5.1.x/)
- [wordcloud](https://github.com/amueller/word_cloud)
The `numpy` library is one of the most popular and helpful library that is used for handling multi-dimensional arrays and matrices. It is also used in combination with `Pandas` library to perform data analysis.
The Python `os` module is a built-in library so you don't have to install it. To read more about handling files with os module, [this DataCamp tutorial](https://www.datacamp.com/community/tutorials/reading-writing-files-python#os) will be helpful.
For visualization, `matplotlib` is a basic library that enable many other libraries to run and plot on its base including [`seaborn`](https://seaborn.pydata.org/) or `wordcloud` that you will use in this tutorial. The `pillow` library is a package that enable image reading. Its tutorial can be found [here](https://pillow.readthedocs.io/en/5.1.x/handbook/tutorial.html). Pillow is a wrapper for PIL - Python Imaging Library. You will need this library to read in image as the mask for the wordcloud.
`wordcloud` can be a little tricky to install. If you only need it for plotting a basic wordcloud, then `pip install wordcloud` or `conda install -c conda-forge wordcloud` would be sufficient. However, the latest version with the ability to mask the cloud into any shape of your choice requires a different method of installation as below:
```
git clone https://github.com/amueller/word_cloud.git
cd word_cloud
pip install .
```
### Dataset:
This tutorial uses the [wine review dataset](https://www.kaggle.com/zynicide/wine-reviews/data) from [Kaggle](https://www.kaggle.com). This collection is a great dataset for learning with no missing values (which will take time to handle) and a lot of text (wine reviews), categorical, and numerical data.
### Now let's get started!
First thing first, you load all the necessary libraries:
```
import warnings
warnings.filterwarnings("ignore")
# Start with loading all necessary libraries
import numpy as np
import pandas as pd
from os import path
from PIL import Image
#from wordcloud import WordCloud, STOPWORDS, ImageColorGenerator
import matplotlib.pyplot as plt
% matplotlib inline
```
If you have more than 10 libraries, organize them by sections (such as basic libs, visualization, models, etc.) using comment in code will make your code clean and easy to follow.
Now, using pandas `read_csv` to load in the dataframe. Notice the use of `index_col=0` meaning we don't read in row name (index) as a separated column.
```
# Load in the dataframe
df = pd.read_csv("data/winemag-data-130k-v2.csv", index_col=0)
# Looking at first 5 rows of the dataset
df.head()
```
You can print out some basic information about the dataset using `print()` combined with `.format()` to have a nice print out.
```
print("There are {} observations and {} features in this dataset. \n".format(df.shape[0],df.shape[1]))
print("There are {} types of wine in this dataset such as {}... \n".format(len(df.variety.unique()),
", ".join(df.variety.unique()[0:5])))
print("There are {} countries producing wine in this dataset such as {}... \n".format(len(df.country.unique()),
", ".join(df.country.unique()[0:5])))
df[["country", "description","points"]].head()
```
To make comparisons between groups of a feature, you can use `groupby()` and compute summary statistics.
With the wine dataset, you can group by country and look at either the summary statistics for all countries' points and price or select the most popular and expensive ones.
```
# Groupby by country
country = df.groupby("country")
# Summary statistic of all countries
country.describe().head()
```
This selects the top 5 highest average points among all 44 countries:
```
country.mean().sort_values(by="points",ascending=False).head()
```
You can plot the number of wines by country using the plot method of Pandas DataFrame and Matplotlib. If you are not familiar with Matplotlib, I suggested to take a quick look at [this tutorial](https://www.datacamp.com/community/tutorials/matplotlib-tutorial-python).
```
plt.figure(figsize=(15,10))
country.size().sort_values(ascending=False).plot.bar()
plt.xticks(rotation=50)
plt.xlabel("Country of Origin")
plt.ylabel("Number of Wines")
plt.show()
```
Among 44 countries producing wine, US has more than 50,000 types of wine in the wine review dataset, twice as much as the next one in the rank: France - the country famous for its wine. Italy also produces a lot of quality wine, having nearly 20,000 wines open to review.
##### Does quantity over quality?
Let's now take a look at the plot of all 44 countries by its highest rated wine, using the same plotting technique as above:
```
plt.figure(figsize=(15,10))
country.max().sort_values(by="points",ascending=False)["points"].plot.bar()
plt.xticks(rotation=50)
plt.xlabel("Country of Origin")
plt.ylabel("Highest point of Wines")
plt.show()
```
Australia, US, Portugal, Italy and France all have 100 points wine. If you notice, Portugal ranks 5th and Australia rankes 9th in the number of wines produces in the dataset, and both countries have less than 8000 types of wine.
That's a little bit of data exploration to get to know the dataset that you are using today. Now you will start dive into the main course of the meal: **WordCloud**.
### Set up a basic WordCloud
[WordCloud](https://amueller.github.io/word_cloud/index.html) is a technique to show which words are the most frequent among the given text. The first thing you may want to do before using any functions is check out the docstring of the function, and see all required and optional arguments. To do so, type `?function` and run it to get all information.
```
?WordCloud
```
You can see that the only required argument for a WordCloud object is the **text**, while all others are optional.
So let's start with a simple example: using the first observation description as the input for the wordcloud. The three steps are:
- Extract the review (text document)
- Create and generate a wordcloud image
- Display the cloud using matplotlib
```
# Start with one review:
text = df.description[0]
# Create and generate a word cloud image:
wordcloud = WordCloud().generate(text)
# Display the generated image:
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
plt.show()
```
Great! You can see that the first review mentioned a lot about dried flavours, and the aromas of the wine.
Now, change some optional arguments of the WordCloud like `max_font_size`, `max_word` and `background_color`.
```
# lower max_font_size, change the maximum number of word and lighten the background:
wordcloud = WordCloud(max_font_size=50, max_words=100, background_color="white").generate(text)
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
```
Ugh, seems like `max_font_size` here might not be a good idea. It makes it more difficult to see the differences between word frequencies. However, brightening the background makes the cloud easier to read.
If you want to save the image, WordCloud provides a function `to_file`
```
# Save the image in the img folder:
wordcloud.to_file("img/first_review.png")
```
The result will look like this when you load them in:

You've probably noticed the argument `interpolation="bilinear"` in the `plt.imshow()`. This is to make the displayed image appear more smoothly. For more information about the choice, [here](https://matplotlib.org/gallery/images_contours_and_fields/interpolation_methods.html) is a helpful link to explore more about this choice.
So now you'll combine all wine reviews into one big text and create a big fat cloud to see which characteristics are most common in these wines.
```
text = " ".join(review for review in df.description)
print ("There are {} words in the combination of all review.".format(len(text)))
# Create stopword list:
stopwords = set(STOPWORDS)
stopwords.update(["drink", "now", "wine", "flavor", "flavors"])
# Generate a word cloud image
wordcloud = WordCloud(stopwords=stopwords, background_color="white").generate(text)
# Display the generated image:
# the matplotlib way:
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
plt.show()
```
Ohhh, it seems like black cherry and full-bodied are the most mentioned characteristics and Cabernet Sauvignon is the most popular of them all. This aligns with the fact that Cabernet Sauvignon "is one of the world's most widely recognized red wine grape varieties. It is grown in nearly every major wine producing country among a diverse spectrum of climates from Canada's Okanagan Valley to Lebanon's Beqaa Valley".<sup>[[1]](https://en.wikipedia.org/wiki/Cabernet_Sauvignon)</sup>
Now, let's pour these words into a cup of wine!
Seriously,
Even a bottle of wine if you wish!
In order to create a shape for your wordcloud, first you need to find a PNG file to become the mask. Below is a nice one that is available on the internet:

Not all mask images have the same format resulting in different outcomes, hence making the WordCloud function not working properly. To make sure that your mask works, let's take a look at it in the numpy array form:
```
wine_mask = np.array(Image.open("img/wine_mask.png"))
wine_mask
```
The way the masking functions works is that it requires all white part of the mask should be 255 not 0 (integer type). This value represents the "intensity" of the pixel. Values of 255 are pure white, whereas values of 1 are black. Here, you can use the provided function below to transform your mask if your mask has the same format as above. Notice if you have a mask that the background is not 0, but 1 or 2, adjust the function to match your mask.
First, you use the `transform_format()` function to swap number 0 to 255.
```
def transform_format(val):
if val == 0:
return 255
else:
return val
```
Then, create a new mask with the same shape as the mask you have in hand and apply the function `transform_format()` to each value in each row of the previous mask.
```
# Transform your mask into a new one that will work with the function:
transformed_wine_mask = np.ndarray((wine_mask.shape[0],wine_mask.shape[1]), np.int32)
for i in range(len(wine_mask)):
transformed_wine_mask[i] = list(map(transform_format, wine_mask[i]))
```
Now, you have a new mask in the correct form. Print out the transformed mask is the best way to check if the function works fine.
```
# Check the expected result of your mask
transformed_wine_mask
```
Okay! With the right mask, you can start making the wordcloud with your selected shape. Notice in the `WordCloud` function, there is a `mask` argument that take in the transformed mask that you created above. The `contour_width` and `contour_color` are, as their name, arguments to adjust the outline characteristics of the cloud. The wine bottle you have here is a redwine bottle, so firebrick seems like a good choice for contour color. For more choice of color, you can take a look at this [color code table](https://matplotlib.org/2.0.0/examples/color/named_colors.html)
```
# Create a word cloud image
wc = WordCloud(background_color="white", max_words=1000, mask=transformed_wine_mask,
stopwords=stopwords, contour_width=3, contour_color='firebrick')
# Generate a wordcloud
wc.generate(text)
# store to file
wc.to_file("img/wine.png")
# show
plt.figure(figsize=[20,10])
plt.imshow(wc, interpolation='bilinear')
plt.axis("off")
plt.show()
```
Voila! You created a wordcloud in the shape of a wine bottle! It seems like wine descriptions most often mention about black cherry, fruit flavours and full bodied characteristics of the wine. Now let's take a closer look at the reviews for each country and plot the wordcloud using each country flag. For you easy to imagine, this is an example that you will create soon:
<img src="img/us_wine.png" alt="Drawing" style="width: 650px;"/>
### Creating wordcloud following a color pattern
You can combine all the reviews of five countries that have the most wines. To find those countries, you can either look at the plot *country vs number* of wine above or use the group that you got above to find the number of observations for each country (each group) and `sort_values()` with argument `ascending=False` to sort descending.
```
country.size().sort_values(ascending=False).head()
```
So now you have 5 top countries: US, France, Italy, Spain and Portugal. You can change the number of countries by putting your choice number insider `head()` like below
```
country.size().sort_values(ascending=False).head(10)
```
For now, 5 countries should be enough.
To get all review for each country, you can concatenate all of the reviews using the `" ".join(list)` syntax, which joins all elements in a list separating them by whitespace.
```
# Join all reviews of each country:
usa = " ".join(review for review in df[df["country"]=="US"].description)
fra = " ".join(review for review in df[df["country"]=="France"].description)
ita = " ".join(review for review in df[df["country"]=="Italy"].description)
spa = " ".join(review for review in df[df["country"]=="Spain"].description)
por = " ".join(review for review in df[df["country"]=="Portugal"].description)
```
Then, creating the wordcloud as above. You can combine the two steps of creating and generate into one as below. The color mapping is done right before you plot the cloud using the [ImageColorGenerator](https://amueller.github.io/word_cloud/generated/wordcloud.ImageColorGenerator.html) function from WordCloud library.
```
# Generate a word cloud image
mask = np.array(Image.open("img/us.png"))
wordcloud_usa = WordCloud(stopwords=stopwords, background_color="white", mode="RGBA", max_words=1000, mask=mask).generate(usa)
# create coloring from image
image_colors = ImageColorGenerator(mask)
plt.figure(figsize=[7,7])
plt.imshow(wordcloud_usa.recolor(color_func=image_colors), interpolation="bilinear")
plt.axis("off")
# store to file
plt.savefig("img/us_wine.png", format="png")
plt.show()
```
Looks good! Now let's repeat with review from France.
```
# Generate a word cloud image
mask = np.array(Image.open("img/france.png"))
wordcloud_fra = WordCloud(stopwords=stopwords, background_color="white", mode="RGBA", max_words=1000, mask=mask).generate(fra)
# create coloring from image
image_colors = ImageColorGenerator(mask)
plt.figure(figsize=[7,7])
plt.imshow(wordcloud_fra.recolor(color_func=image_colors), interpolation="bilinear")
plt.axis("off")
# store to file
plt.savefig("img/fra_wine.png", format="png")
#plt.show()
```
Please note that you should save the image after plotting to have the wordcloud with desired color pattern.
```
# Generate a word cloud image
mask = np.array(Image.open("img/italy.png"))
wordcloud_ita = WordCloud(stopwords=stopwords, background_color="white", max_words=1000, mask=mask).generate(ita)
# create coloring from image
image_colors = ImageColorGenerator(mask)
plt.figure(figsize=[7,7])
plt.imshow(wordcloud_ita.recolor(color_func=image_colors), interpolation="bilinear")
plt.axis("off")
# store to file
plt.savefig("img/ita_wine.png", format="png")
#plt.show()
```
Following Italy is Spain:
```
# Generate a word cloud image
mask = np.array(Image.open("img/spain.png"))
wordcloud_spa = WordCloud(stopwords=stopwords, background_color="white", max_words=1000, mask=mask).generate(spa)
# create coloring from image
image_colors = ImageColorGenerator(mask)
plt.figure(figsize=[7,7])
plt.imshow(wordcloud_spa.recolor(color_func=image_colors), interpolation="bilinear")
plt.axis("off")
# store to file
plt.savefig("img/spa_wine.png", format="png")
#plt.show()
```
Lastly, Portugal:
```
# Generate a word cloud image
mask = np.array(Image.open("img/portugal.png"))
wordcloud_por = WordCloud(stopwords=stopwords, background_color="white", max_words=1000, mask=mask).generate(por)
# create coloring from image
image_colors = ImageColorGenerator(mask)
plt.figure(figsize=[7,7])
plt.imshow(wordcloud_por.recolor(color_func=image_colors), interpolation="bilinear")
plt.axis("off")
# store to file
plt.savefig("img/por_wine.png", format="png")
#plt.show()
```
The end result are in the below table to compare between the mask and the wordcloud. Which one is your favorite?
|Mask|Result|
|:--:| :--: |
|<img src="img/us.png" alt="Drawing" style="width: 500px;"/>|<img src="img/us_wine.png" alt="Drawing" style="width: 650px;"/>|
|<img src="img/france.png" alt="Drawing" style="width: 500px;"/>|<img src="img/fra_wine.png" alt="Drawing" style="width: 650px;"/>|
|<img src="img/italy.png" alt="Drawing" style="width: 500px;"/>|<img src="img/ita_wine.png" alt="Drawing" style="width: 650px;"/>|
|<img src="img/spain.png" alt="Drawing" style="width: 500px;"/>|<img src="img/spa_wine.png" alt="Drawing" style="width: 650px;"/>|
|<img src="img/portugal.png" alt="Drawing" style="width: 500px;"/>|<img src="img/por_wine.png" alt="Drawing" style="width: 650px;"/>|
## Congratulations!
You made it! You have learned several ways to draw a WordCloud that would be helpful for visualization any text analysis. You also learn how to mask the cloud into any shape, using any color of your choice. If you want to practice your skills, consider the DataCamp's project: [The Hottest Topics in Machine Learning](https://www.datacamp.com/projects/158)
If you'd like to get in touch with me, you can drop me an e-mail at dqvu.ubc@gmail.com or connect with me via [LinkedIn](https://www.linkedin.com/in/duongqvu/).
| github_jupyter |
# 패키지 불러오기
```
import tensorflow as tf
import numpy as np
import os
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
import matplotlib.pyplot as plt
from preprocess import *
```
# 시각화 함수
```
def plot_graphs(history, string):
plt.plot(history.history[string])
plt.plot(history.history['val_'+string], '')
plt.xlabel("Epochs")
plt.ylabel(string)
plt.legend([string, 'val_'+string])
plt.show()
```
# 학습 데이터 경로 정의
```
DATA_IN_PATH = './data_in/'
DATA_OUT_PATH = './data_out/'
TRAIN_INPUTS = 'train_inputs.npy'
TRAIN_OUTPUTS = 'train_outputs.npy'
TRAIN_TARGETS = 'train_targets.npy'
DATA_CONFIGS = 'data_configs.json'
```
# 랜덤 시드 고정
```
SEED_NUM = 1234
tf.random.set_seed(SEED_NUM)
```
# 파일 로드
```
index_inputs = np.load(open(DATA_IN_PATH + TRAIN_INPUTS, 'rb'))
index_outputs = np.load(open(DATA_IN_PATH + TRAIN_OUTPUTS , 'rb'))
index_targets = np.load(open(DATA_IN_PATH + TRAIN_TARGETS , 'rb'))
prepro_configs = json.load(open(DATA_IN_PATH + DATA_CONFIGS, 'r'))
# Show length
print(len(index_inputs), len(index_outputs), len(index_targets))
```
## 모델 만들기에 필요한 값 선언
```
MODEL_NAME = 'seq2seq_kor'
BATCH_SIZE = 2
MAX_SEQUENCE = 25
EPOCH = 30
UNITS = 1024
EMBEDDING_DIM = 256
VALIDATION_SPLIT = 0.1
char2idx = prepro_configs['char2idx']
idx2char = prepro_configs['idx2char']
std_index = prepro_configs['std_symbol']
end_index = prepro_configs['end_symbol']
vocab_size = prepro_configs['vocab_size']
```
# 모델
## 인코더
```
class Encoder(tf.keras.layers.Layer):
def __init__(self, vocab_size, embedding_dim, enc_units, batch_sz):
super(Encoder, self).__init__()
self.batch_sz = batch_sz
self.enc_units = enc_units
self.vocab_size = vocab_size
self.embedding_dim = embedding_dim
self.embedding = tf.keras.layers.Embedding(self.vocab_size, self.embedding_dim)
self.gru = tf.keras.layers.GRU(self.enc_units,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
def call(self, x, hidden):
x = self.embedding(x)
output, state = self.gru(x, initial_state = hidden)
return output, state
def initialize_hidden_state(self, inp):
return tf.zeros((tf.shape(inp)[0], self.enc_units))
```
## 어텐션
```
class BahdanauAttention(tf.keras.layers.Layer):
def __init__(self, units):
super(BahdanauAttention, self).__init__()
self.W1 = tf.keras.layers.Dense(units)
self.W2 = tf.keras.layers.Dense(units)
self.V = tf.keras.layers.Dense(1)
def call(self, query, values):
hidden_with_time_axis = tf.expand_dims(query, 1)
score = self.V(tf.nn.tanh(
self.W1(values) + self.W2(hidden_with_time_axis)))
attention_weights = tf.nn.softmax(score, axis=1)
context_vector = attention_weights * values
context_vector = tf.reduce_sum(context_vector, axis=1)
return context_vector, attention_weights
```
## 디코더
```
class Decoder(tf.keras.layers.Layer):
def __init__(self, vocab_size, embedding_dim, dec_units, batch_sz):
super(Decoder, self).__init__()
self.batch_sz = batch_sz
self.dec_units = dec_units
self.vocab_size = vocab_size
self.embedding_dim = embedding_dim
self.embedding = tf.keras.layers.Embedding(self.vocab_size, self.embedding_dim)
self.gru = tf.keras.layers.GRU(self.dec_units,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
self.fc = tf.keras.layers.Dense(self.vocab_size)
self.attention = BahdanauAttention(self.dec_units)
def call(self, x, hidden, enc_output):
context_vector, attention_weights = self.attention(hidden, enc_output)
x = self.embedding(x)
x = tf.concat([tf.expand_dims(context_vector, 1), x], axis=-1)
output, state = self.gru(x)
output = tf.reshape(output, (-1, output.shape[2]))
x = self.fc(output)
return x, state, attention_weights
optimizer = tf.keras.optimizers.Adam()
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True, reduction='none')
train_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='accuracy')
def loss(real, pred):
mask = tf.math.logical_not(tf.math.equal(real, 0))
loss_ = loss_object(real, pred)
mask = tf.cast(mask, dtype=loss_.dtype)
loss_ *= mask
return tf.reduce_mean(loss_)
def accuracy(real, pred):
mask = tf.math.logical_not(tf.math.equal(real, 0))
mask = tf.expand_dims(tf.cast(mask, dtype=pred.dtype), axis=-1)
pred *= mask
acc = train_accuracy(real, pred)
return tf.reduce_mean(acc)
```
## 시퀀스 투 스퀀스 모델
```
class seq2seq(tf.keras.Model):
def __init__(self, vocab_size, embedding_dim, enc_units, dec_units, batch_sz, end_token_idx=2):
super(seq2seq, self).__init__()
self.end_token_idx = end_token_idx
self.encoder = Encoder(vocab_size, embedding_dim, enc_units, batch_sz)
self.decoder = Decoder(vocab_size, embedding_dim, dec_units, batch_sz)
def call(self, x):
inp, tar = x
enc_hidden = self.encoder.initialize_hidden_state(inp)
enc_output, enc_hidden = self.encoder(inp, enc_hidden)
dec_hidden = enc_hidden
predict_tokens = list()
for t in range(0, tar.shape[1]):
dec_input = tf.dtypes.cast(tf.expand_dims(tar[:, t], 1), tf.float32)
predictions, dec_hidden, _ = self.decoder(dec_input, dec_hidden, enc_output)
predict_tokens.append(tf.dtypes.cast(predictions, tf.float32))
return tf.stack(predict_tokens, axis=1)
def inference(self, x):
inp = x
enc_hidden = self.encoder.initialize_hidden_state(inp)
enc_output, enc_hidden = self.encoder(inp, enc_hidden)
dec_hidden = enc_hidden
dec_input = tf.expand_dims([char2idx[std_index]], 1)
predict_tokens = list()
for t in range(0, MAX_SEQUENCE):
predictions, dec_hidden, _ = self.decoder(dec_input, dec_hidden, enc_output)
predict_token = tf.argmax(predictions[0])
if predict_token == self.end_token_idx:
break
predict_tokens.append(predict_token)
dec_input = tf.dtypes.cast(tf.expand_dims([predict_token], 0), tf.float32)
return tf.stack(predict_tokens, axis=0).numpy()
model = seq2seq(vocab_size, EMBEDDING_DIM, UNITS, UNITS, BATCH_SIZE, char2idx[end_index])
model.compile(loss=loss, optimizer=tf.keras.optimizers.Adam(1e-3), metrics=[accuracy])
#model.run_eagerly = True
```
## 학습 진행
```
PATH = DATA_OUT_PATH + MODEL_NAME
if not(os.path.isdir(PATH)):
os.makedirs(os.path.join(PATH))
checkpoint_path = DATA_OUT_PATH + MODEL_NAME + '/weights.h5'
cp_callback = ModelCheckpoint(
checkpoint_path, monitor='val_accuracy', verbose=1, save_best_only=True, save_weights_only=True)
earlystop_callback = EarlyStopping(monitor='val_accuracy', min_delta=0.0001, patience=10)
history = model.fit([index_inputs, index_outputs], index_targets,
batch_size=BATCH_SIZE, epochs=EPOCH,
validation_split=VALIDATION_SPLIT, callbacks=[earlystop_callback, cp_callback])
```
## 결과 플롯
```
plot_graphs(history, 'accuracy')
plot_graphs(history, 'loss')
```
### 결과 확인
```
SAVE_FILE_NM = "weights.h5"
model.load_weights(os.path.join(DATA_OUT_PATH, MODEL_NAME, SAVE_FILE_NM))
query = "남자친구 승진 선물로 뭐가 좋을까?"
test_index_inputs, _ = enc_processing([query], char2idx)
predict_tokens = model.inference(test_index_inputs)
print(predict_tokens)
print(' '.join([idx2char[str(t)] for t in predict_tokens]))
```
| github_jupyter |
# Calculating and plotting degassing paths
## Calculate degassing paths
A degassing path is a series of volatile concentrations both in the liquid and fluid that a magma will follow during decompression. In the calculation, the saturation pressure is computed, and then the system is equilibrated along a trajectory of decreasing pressure values at steps of 100 bars (or 10 bars if the starting pressure is <500 bars). If so desired, this calculation can be performed for any initial pressure, but the default is the saturation pressure. If a pressure is specified that is above the saturation pressure, the calculation will simlpy proceed from the saturation pressure, since the magma cannot degas until it reaches saturation.
Completely open-system, completely closed-system or partially open-system degassing paths can be calculated by specifying what proportion of the fluid to fractionate. The fluid fractionation value can range between 0 (closed-system: no fluid is removed, all is retained at each pressure step) and 1 (open-system: all fluid is removed, none is retained at each pressure step). Closed and partially open-system runs allow the user to speficy the initial presence of exsolved fluid that is in equilirium with the melt at the starting pressure.
**Method structure:**<br>
>*Only single-sample calculations.* `def calculate_degassing_path(self, sample, temperature, pressure='saturation', fractionate_vapor=0.0, init_vapor=0.0).result`
**Required inputs:**<br>
>`sample`: The sample composition, as a dictionary with values in wt%
>`temperature`: The temperature in $^{\circ}$C.
**Optional inputs:**<br>
>`pressure`: The perssure at which to begin the degassing calculations, in bars. Default value is 'saturation', which runs the calculation with the initial pressure at the saturation pressure. If a pressure greater than the saturation pressure is input, the calculation will start at saturation, since this is the first pressure at which any degassing will occur.
>`fractionate_vapor`: Proportion of vapor removed at each pressure step. Default value is 0.0 (completely closed-system degassing). Specifies the type of calculation performed, either closed system (0.0) or open system (1.0) degassing. If any value between <1.0 is chosen, user can also specify the 'init_vapor' argument (see below). A value in between 0 and 1 will remove that proportion of vapor at each step. For example, for a value of 0.2, the calculation will remove 20% of the vapor and retain 80% of the vapor at each pressure step.
>`init_vapor`: Default value is 0.0. Specifies the amount of vapor (in wt%) coexisting with the melt before degassing.
**Calculated outputs:**
>The function returns a pandas DataFrame with columns as: 'Pressure_bars', 'H2O_liq' and 'CO2_liq' (the concentration of H$_2$O and CO$_2$ in the liquid, in wt%), 'XH2O_fl' and 'XCO2_fl' (the composition of the H$_2$O-CO$_2$ fluid, in mol fraction), and 'FluidProportion_wt' (the proportion of fluid in the fluid-melt system, in wt%).
```
import sys
sys.path.insert(0, '../')
import VESIcal as v
```
### Import an Excel file and extract a single sample
```
myfile = v.ExcelFile('../manuscript/example_data.xlsx')
SampleName = 'BT-ex'
extracted_bulk_comp = myfile.get_sample_oxide_comp(SampleName)
```
### Open system degassing calculation
```
open_df = v.calculate_degassing_path(sample=extracted_bulk_comp, temperature=900.0, fractionate_vapor=1.0).result
```
### Closed system degassing calculation
```
closed_df = v.calculate_degassing_path(sample=extracted_bulk_comp, temperature=900.0).result
```
### Partially closed system degassing calculation
Here we will fractionate 50% of the fluid produced at each calculation step.
```
half_df = v.calculate_degassing_path(sample=extracted_bulk_comp, temperature=900.0, fractionate_vapor=0.5).result
```
### Closed system with initial fluid
Here we will calculate a closed system degassing path where the initial magma contains 2 wt% exsolved fluid.
```
exsolved_df = v.calculate_degassing_path(sample=extracted_bulk_comp, temperature=900.0, init_vapor=2.0).result
```
### Calculate from initial pressure below saturation
By default, degassing paths are calculated from the computed saturation pressure for the sample. The user can instead pass their own pressure. If the user supplied pressure is below the saturation pressure computed by the model, the degassing path will be computed from the user supplied pressure.
```
start2000_df = v.calculate_degassing_path(sample=extracted_bulk_comp, temperature=900.0, pressure=2000.0).result
```
## Plotting degassing paths
Once degassing paths are calcualted, they may be easily plotted using VESIcal's built in `plot_degassing_paths` method. The user can plot multiple degassing paths on one plot. Optionally, labels in the plot legend can be specified.
**Method structure:**<br>
>`def plot_degassing_paths(degassing_paths, labels=None)`
**Required inputs:**<br>
>`degassing_paths`: A list of DataFrames with degassing information as generated by calculate_degassing_path().
**Optional inputs:**<br>
>`labels`: Labels for the plot legend. Default is None, in which case each plotted line will be given the generic legend name of "Pathn", with n referring to the nth degassing path passed. The user can pass their own labels as a list of strings.
**Calculated outputs:**
>The function returns a matplotlib object with the x-axis as H$_2$O, wt% and the y-axis as CO$_2$, wt%. All degassing paths passed are plotted on one figure.
### Plotting degassig paths from saturation pressure
```
v.plot_degassing_paths([open_df, half_df, closed_df, exsolved_df],
labels=["Open", "Half", "Closed", "Exsolved"])
```
### Plotting degassing path from 2000 bars
```
v.plot_degassing_paths([start2000_df], labels=["2000 bars"])
```
| github_jupyter |
# CHURN PREDICTION - TOP BANK COMPANY
```
Image('img/capa.jpg')
```
While getting new customers is an obvious win, many businesses often forget about an important, and potentially bigger win: keeping the customers you already have.
Customer churn – the loss of customers – is a big business killer. Even small increases in churn can cut your revenues in half. There’s no question about it: churn can crush your business, if you let it.
## **The Four Most Common Causes of Customer Churn**
### 1) Bad Customer Service
Many companies think of customer service as a cost to be minimized, rather than an investment to be maximized. Here’s the issue with that: if you think of support as a cost center, then it will be. That is, if you don’t prioritize support and work to deliver excellent service to your customers, then it’s only going to cost you money…and customers.
In fact, one Oracle study found that almost 9 in 10 customers have abandoned a business because of a poor experience. But just as bad customer service can be a huge loss for your business, the same study found that great customer service can be a huge win, with 86% of customers willing to pay more for a better customer experience.
### 2) Bad Onboarding
To your business, two of the most important milestones in the life of a customer are:
- The moment they sign up for your product, and…
- The moment they achieve their first “success” with your product
```
Image("img/bad_unboarding.jpg")
```
A disproportionate amount of your customer churn will take place between (1) and (2).
That’s where customers abandon your product because they get lost, don’t understand something, don’t get value from the product, or simply lose interest.
Bad onboarding – the process by which you help a customer go from (1) to (2) – can crush your retention rate, and undo all of that hard work you did to get your customers to convert in the first place.
It’s your job to make that transition as fast and smooth as possible for your customer, and that’s where great onboarding comes in.
### 3) Lack of Ongoing Customer Success
While onboarding gets your customer to their initial success, your job isn’t done there.
Hundreds of variables – including changing needs, confusion about new features and product updates, extended absences from the product and competitor marketing – could lead your customers away.
```
Image("img/3.png")
```
If your customers stop hearing from you, and you stop helping them get value from your product throughout their entire lifecycle, then you risk making that lifecycle much, much shorter.
### 4) Natural Causes
Not every customer that abandons you does so because you failed.
Sometimes, customers go out of business.
Sometimes, operational or staff changes lead to vendor switches.
Sometimes, they simply outgrow your product or service.
And that’s okay. It’ll happen to every business.
But it’s still churn, and you can get value from acting on it.
**REFERENCES** https://www.groovehq.com/blog/reduce-customer-churn
## Business Problem
TopBank is a large banking services company. It operates mainly in European countries offering financial products, from bank accounts to investments, including some types of insurance and investment products.
The company's business model is a service type, that is, it sells banking services to its customers through physical branches and an online portal.
The company's main product is a bank account, in which the customer can deposit his salary, make withdrawals, deposits and transfer to other accounts. This bank account has no cost to the customer and is valid for 12 months, that is, the customer needs to renew the contract of that account to continue using it for the next 12 months.
According to the TopBank Analytics team, each customer who has this bank account returns a monetary value of 15 % of the value of their estimated salary, if it is less than the average and 20 % if this salary is higher than the average, during the current period of your account. This value is calculated annually.
For example, if a customer's monthly salary is R $ 1,000.00 and the average of all bank wages is R $ 800. The company, therefore, invoices R $ 200 annually with this client. If this customer has been in the bank for 10 years, the company has already earned R $ 2,000.00 from its transactions and account usage.
In recent months, the Analytics team realized that the rate of customers canceling their accounts and leaving the bank, reached unprecedented numbers in the company. Concerned about the increase in this rate, the team devised an action plan to reduce the rate of customer evasion.
Concerned about the drop in this metric, TopBottom's Analytics team hired you as a Data Science consultant to create an action plan, with the objective of reducing customer evasion, that is, preventing the customer from canceling his contract and not renew it for another 12 months. This evasion, in business metrics, is known as Churn.
In general, Churn is a metric that indicates the number of customers who have canceled the contract or have stopped buying your product within a certain period of time. For example, customers who canceled the service contract or after it expired, did not renew it, they are considered churn customers.
Another example would be customers who have not made a purchase for more than 60 days. These customers can be considered churn customers until a purchase is made. The 60-day period is completely arbitrary and varies between companies.
We were hired as a Data Science Consultant to create a high performance model for identifying churn customers.
At the end of your consultancy, you need to deliver to the TopBank CEO a model in production, which will receive a customer base via API and develop this base "scorada", that is, one more column with the probability of each customer going into churn .
In addition, you will need to deliver a report reporting your model's performance and results in relation to
- What is TopBank's current Churn rate? How does it vary monthly?
- What is the model's performance in classifying customers as churns?
- What is the expected return, in terms of revenue, if the company uses its model to avoid churn from customers?
A possible action to prevent the customer from churning is to offer a discount coupon, or some other financial incentive for him to renew his contract for another 12 months.
Which customers would you give the financial incentive to and what would that amount be, in order to maximize ROI (Return on Investment). Recalling that the sum of incentives for each client cannot exceed R $ 10,000.00
# 0.0. IMPORTS
```
#data manipulation
import pandas as pd
import numpy as np
#data visualization
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
#data preparation
from sklearn.model_selection import train_test_split
#machine learning algorithms
from sklearn.linear_model import LogisticRegression
#metrics
from sklearn.metrics import accuracy_score, classification_report,cohen_kappa_score,recall_score,f1_score,roc_auc_score, plot_precision_recall_curve, precision_score,roc_curve
from scikitplot import metrics as mt
#auxiliar packages
from IPython.display import Image
from IPython.core.display import HTML
import inflection
import warnings
import random
warnings.filterwarnings('ignore')
```
## 0.1. Helper Functions
```
def jupyter_settings():
%matplotlib inline
%pylab inline
plt.style.use( 'bmh' )
plt.rcParams['figure.figsize'] = [25, 30]
plt.rcParams['font.size'] = 25
display( HTML( '<style>.container { width:100% !important; }</style>') )
pd.options.display.max_columns = None
pd.options.display.max_rows = None
pd.set_option( 'display.expand_frame_repr', False )
sns.set()
jupyter_settings()
```
## 0.2. Loading Data
The data set that will be used to create the solution for TopBottom, is available on the Kaggle platform. This is the link: [ChurnDataset]("https://www.kaggle.com/mervetorkan/churndataset")
Each row represents a customer and each column contains some attributes that describe that customer. The data set includes information about:
- **RowNumber**: The column number
- **CustomerID**: Unique customer identifier
- **Surname**: Last name of the customer.
- **CreditScore**: The customer's Credit score for the consumer market.
- **Geography**: The state where the customer resides.
- **Gender**: The gender of the customer.
- **Age**: The age of the customer.
- **Tenure**: Number of months that the customer has remained active.
- **Balance**: The amount spent by the customer with the company TopBottom.
- **NumOfProducts**: The number of products purchased by the customer.
- **HasCrCard**: Flag indicating whether the customer has a credit card or not.
- **IsActiveMember**: Flag indicating whether the customer still has an active registration at TopBottom.
- **EstimateSalary**: Estimated monthly salary of the client.
- **Exited**: Flag indicating whether or not the customer is in Churn
```
#load data as a dataframe
df_raw = pd.read_csv("data/churn.csv")
```
# 1.0. DATA DESCRIPTION
```
#copy data
df1 = df_raw.copy()
```
## 1.1. Rename Columns
```
cols_old = ['RowNumber', 'CustomerId', 'Surname', 'CreditScore', 'Geography',
'Gender', 'Age', 'Tenure', 'Balance', 'NumOfProducts', 'HasCrCard',
'IsActiveMember', 'EstimatedSalary', 'Exited']
#apply snakecase
snakecase = lambda x: inflection.underscore(x)
cols_new = list(map(snakecase, cols_old))
#rename columns
df1.columns = cols_new
df1.head()
```
## 1.2. Data Dimensions
```
#print shape
print("Number of rows: {}".format(df1.shape[0]))
print("Number of columns: {}".format(df1.shape[1]))
```
## 1.3. Data Types
```
#check data types
df1.dtypes
```
## 1.4. Check NA
```
#check if there are null values in our dataframe
df1.isnull().sum()
```
## 1.5. Descriptive Statistical
### 1.5.1. Numerical Attributes
```
#select only numerical features
num_attributes = df1.select_dtypes(include = ['int64','float64'])
num_attributes.drop(columns = ['row_number','customer_id','is_active_member','has_cr_card','exited'], axis = 1, inplace = True)
#central tendency - mean and median
ct1 = pd.DataFrame(np.round(num_attributes.apply(np.mean),2)).T
ct2 = pd.DataFrame(np.round(num_attributes.apply(np.median),2)).T
#dipersion - std, max, min, range, skew, kurtosis
d1 = pd.DataFrame(np.round(num_attributes.apply(np.std))).T
d2 = pd.DataFrame(np.round(num_attributes.apply(min))).T
d3 = pd.DataFrame(np.round(num_attributes.apply(max))).T
d4 = pd.DataFrame(np.round(num_attributes.apply(lambda x: x.max() - x.min()))).T
d5 = pd.DataFrame(np.round(num_attributes.apply(lambda x: x.skew()),2)).T
d6 = pd.DataFrame(np.round(num_attributes.apply(lambda x: x.kurtosis()),2)).T
#concat numerical attributes
m = pd.concat([d2,d3,d4,ct1,ct2,d1,d5,d6]).T.reset_index()
m.columns = ['attributes','min','max','range','mean','median','std','skew','kurtosis']
m
#plot each graphic to visualize if there are outliers
plt.subplot(6,2,1)
sns.boxplot(df1['credit_score'])
plt.subplot(6,2,2)
sns.distplot(df1['credit_score'])
plt.subplot(6,2,3)
sns.boxplot(df1['age'])
plt.subplot(6,2,4)
sns.distplot(df1['age'])
plt.subplot(6,2,5)
sns.boxplot(df1['tenure'])
plt.subplot(6,2,6)
sns.distplot(df1['tenure'])
plt.subplot(6,2,7)
sns.boxplot(df1['balance'])
plt.subplot(6,2,8)
sns.distplot(df1['balance'])
plt.subplot(6,2,9)
sns.boxplot(df1['num_of_products'])
plt.subplot(6,2,10)
sns.distplot(df1['num_of_products'])
plt.subplot(6,2,11)
sns.boxplot(df1['estimated_salary'])
plt.subplot(6,2,12)
sns.distplot(df1['estimated_salary'])
```
### 1.5.2. Categorical and Binary Attributes
```
#select only categorical features and binary features
categorical_features = ['geography','gender','has_cr_card','is_active_member','exited']
cat_attributes = df1[categorical_features]
cat_attributes.head()
#plot each graphic to our categorical variables
plt.subplot(2,3,1)
sns.countplot(cat_attributes['geography'])
plt.subplot(2,3,2)
sns.countplot(cat_attributes['gender'])
plt.subplot(2,3,3)
sns.countplot(cat_attributes['has_cr_card'])
plt.subplot(2,3,4)
sns.countplot(cat_attributes['is_active_member'])
plt.subplot(2,3,5)
sns.countplot(cat_attributes['exited'])
```
# 2.0. FEATURE ENGINEERING
```
df2 = df1.copy()
df2.head()
```
## 2.1. MindMap Hypothesis
```
Image("img/mindmap.png")
```
## 2.2. Hypothesis List
### Age
- Among the people who are in churn, the elderly are the majority.
- Young people have less credit on the card
- There are young people more active than older.
### Balance
- People who spend more are not churning.
### Is Active Member
- There are young people more active than older.
- Active customers spend more.
- The higher the salary, the greater the probability of staying
- People with a credit card are more active.
- People who buy multiple products are more active.
### Salary
- The higher the salary, the greater the probability of staying
- The lower the salary, the greater the debt on the card
- The lower the salary, the lower the credit.
- Young people have a lower salary.
- The lower the salary, the greater the chance of churning
### Gender
- Gender does not influence the likelihood that someone will churn.
### Have a credit card
- People with higher expenses have a credit card.
- People with debt cards are less active.
### Num of products
- The greater the quantity of products purchased, the less chance of churning
- People with a higher salary buy more products
### Location
- The country does not influence the likelihood of the customer churning
### Tenure
- The longer the customer remained active, the less chance of churning.
## 2.3. Final Hypothesis List
**H1** Among the people who are in churn, the elderly are the majority.
**H2** Young people have less credit on the card
**H3** There are young people more active than older.
**H4** People who spend more are not churning.
**H5** There are young people more active than older.
**H6** Active customers spend more.
**H7** The higher the salary, the greater the probability of staying
**H8** People with a credit card are more active.
**H9** People who buy multiple products are more active.
**H10** The lower the salary, the greater the debt on the card
**H11** The lower the salary, the lower the credit.
**H12** Young people have a lower salary.
**H13** The lower the salary, the greater the chance of churning
**H14** Gender does not influence the likelihood that someone will churn.
**H15** People with higher expenses have a credit card.
**H16** People with debt cards are less active.
**H17** The greater the quantity of products purchased, the less chance of churning
**H18** People with a higher salary buy more products
**H19** The country does not influence the likelihood of the customer churning
**H20** The longer the customer remained active, the less chance of churning.
## 2.4. Feature Engineering
```
#age_rate
df2['age_rate'] = df2['age'].apply(lambda x: "Young" if (x >= 18) & (x <= 24) else "Adult" if (x > 24) & (x < 60) else "Old")
#salary
df2['salary_rate'] = df2['estimated_salary'].apply(lambda x: "Below Average" if x < 100000 else "Above average salary" if (x > 100000) & (x < 150000) else 'Above average salary')
```
# 4.0. EXPLORATORY DATA ANALYSIS
```
df4 = df2.copy()
```
## 4.1. Univariate Analysis
### 4.1.1. Response Variable
```
sns.countplot(df4['exited'])
```
### 4.1.2. Numerical Variables
```
num_attributes = df4.select_dtypes(include = ['int64','float64'])
num_attributes.drop(columns = ['row_number','customer_id','has_cr_card','is_active_member','exited'], axis = 1, inplace = True)
num_attributes.hist();
```
### 4.1.3. Categorical Variables
## 4.2. Bivariate Analysis
## 4.3. Multivariate Analysis
# 3.0. DATA PREPARATION
```
df3 = df2.copy()
```
## 3.1. Split data into training and test
```
X = df3.drop(columns = 'exited', axis = 1)
y = df3['exited']
X_train,X_test,y_train,y_test = train_test_split(X,y, test_size = 0.2, random_state = 42)
```
# 4.0. FEATURE SELECTION
## 4.1. Manual Feature Selection
```
cols_selected = ['credit_score', 'geography',
'gender', 'age', 'tenure', 'balance', 'num_of_products', 'has_cr_card',
'is_active_member', 'estimated_salary']
X_train = X_train[cols_selected]
X_test = X_test[cols_selected]
X_train = pd.get_dummies(X_train)
X_test = pd.get_dummies(X_test)
```
# 5.0. MACHINE LEARNING MODELLING
## 5.1. Baseline Model
```
import random
churn = y_test.drop_duplicates().sort_values().tolist()
random_test = X_test.shape[0]
churn_weights = df3['exited'].value_counts( normalize=True ).sort_index().tolist()
# prediction
yhat_random = random.choices( churn, k=random_test,
weights=churn_weights )
# Accuracy
acc_random = accuracy_score( y_test, yhat_random )
print( 'Accuracy: {}'.format( acc_random ) )
# Kappa Metrics
kappa_random = cohen_kappa_score( y_test, yhat_random )
print( 'Kappa Score: {}'.format( kappa_random ) )
# Classification report
print( classification_report( y_test, yhat_random ) )
# Confusion Matrix
mt.plot_confusion_matrix( y_test, yhat_random, normalize=False, figsize=(12,12))
```
## 5.2. Logistic Regression
```
#define model
logreg = LogisticRegression()
#fit model
logreg.fit(X_train,y_train)
#predict model
yhat_logreg = logreg.predict(X_test)
# Accuracy
acc_logreg = accuracy_score( y_test, yhat_logreg )
print( 'Accuracy: {}'.format( acc_logreg ) )
# Kappa Metrics
kappa_logreg = cohen_kappa_score( y_test, yhat_logreg )
print( 'Kappa Score: {}'.format( kappa_logreg ) )
# Classification report
print( classification_report( y_test, yhat_logreg ) )
# Confusion Matrix
mt.plot_confusion_matrix( y_test, yhat_logreg, normalize=False, figsize=(12,12))
print( 'Number of Rows: {}'.format( X_train.shape[0] ) )
print( 'Number of Features: {}'.format( X_train.shape[1] ) )
print( 'Number of Classes: {}'.format( y_train.nunique() ) )
```
| github_jupyter |
---
syncID: a9ef8a3acfb841e2b77b1f6360e22648
title: "Unsupervised Spectral Classification in Python: Endmember Extraction"
description: "Learn to classify spectral data using Endmember Extraction, Spectral Information Divergence, and Spectral Angle Mapping."
dateCreated: 2018-07-10
authors: Bridget Hass
contributors: Donal O'Leary
estimatedTime: 1 hour
packagesLibraries: numpy, gdal, matplotlib, matplotlib.pyplot
topics: hyperspectral-remote-sensing, HDF5, remote-sensing
languagesTool: python
dataProduct: NEON.DP1.30006, NEON.DP3.30006, NEON.DP1.30008
code1: https://raw.githubusercontent.com/NEONScience/NEON-Data-Skills/main/tutorials/Python/Hyperspectral/hyperspectral-classification/classification_endmember_extraction_py/classification_endmember_extraction_py.ipynb
tutorialSeries: intro-hsi-py-series
urlTitle: classification-endmember-python
---
This tutorial runs through an example of spectral unmixing to carry out unsupervised classification of a SERC hyperspectral data file using the <a href="https://pysptools.sourceforge.io/index.html" target="_blank">PySpTools package</a> to carry out **endmember extraction**, plot **abundance maps** of the spectral endmembers, and use **Spectral Angle Mapping** and **Spectral Information Divergence** to classify the SERC tile.
<div id="ds-objectives" markdown="1">
### Objectives
After completing this tutorial, you will be able to:
* Classify spectral remote sensing data.
### Install Python Packages
* **numpy**
* **gdal**
* **matplotlib**
* **matplotlib.pyplot**
### Download Data
This tutorial uses a 1km AOP Hyperspectral Reflectance 'tile' from the SERC site. <a href="https://ndownloader.figshare.com/files/25752665">
Download the spectral classification teaching data subset here</a>.
<a href="https://ndownloader.figshare.com/files/25752665" class="link--button link--arrow">
Download Dataset</a>
</div>
This tutorial runs through an example of spectral unmixing to carry out unsupervised classification of a SERC hyperspectral data file using the <a href="https://pysptools.sourceforge.io/index.html" target="_blank">PySpTools package</a> to carry out **endmember extraction**, plot **abundance maps** of the spectral endmembers, and use **Spectral Angle Mapping** and **Spectral Information Divergence** to classify the SERC tile.
Since spectral data is so large in size, it is often useful to remove any unncessary or redundant data in order to save computational time. In this example, we will remove the water vapor bands, but you can also take a subset of bands, depending on your research application.
## Set up
To run this notebook, the following Python packages need to be installed.
You can install required packages from command line `pip install pysptools scikit-learn cvxopt`.
or if already in a Jupyter Notebook:
1. PySpTools: Download <a href="https://pypi.python.org/pypi/pysptools" target="_blank">pysptools-0.14.2.tar.gz</a>.
2. Run the following code in a Notebook code cell.
```
import sys
# You will need to download the package using the link above
# and re-point the filepath to the tar.gz file below
!{sys.executable} -m pip install "/Users/olearyd/Downloads/pysptools-0.15.0.tar.gz"
!conda install --yes --prefix {sys.prefix} scikit-learn
!conda install --yes --prefix {sys.prefix} cvxopt
```
We will also use the following user-defined functions:
* **`read_neon_reflh5`**: function to read in NEON AOP Hyperspectral Data file (in hdf5 format)
* **`clean_neon_refl_data`**: function to clean NEON hyperspectral data, including applying the data ignore value and reflectance scale factor, and removing water vapor bands
* **`plot_aop_refl`**: function to plot a band of NEON hyperspectral data for reference
Once PySpTools is installed, import the following packages.
```
import h5py, os, copy
import matplotlib.pyplot as plt
import numpy as np
import pysptools.util as util
import pysptools.eea as eea #endmembers extraction algorithms
import pysptools.abundance_maps as amap
import pysptools.classification as cls
import pysptools.material_count as cnt
%matplotlib inline
#for clean output, to not print warnings, don't use when developing script
import warnings
warnings.filterwarnings('ignore')
```
Define the function `read_neon_reflh5` to read in the h5 file, without cleaning it (applying the no-data value and scale factor); we will do that with a separate function that also removes the water vapor bad band windows.
```
def read_neon_reflh5(refl_filename):
"""read in a NEON AOP reflectance hdf5 file and returns
reflectance array, and metadata dictionary containing metadata
(similar to envi header format)
--------
Parameters
refl_filename -- full or relative path and name of reflectance hdf5 file
--------
Returns
--------
reflArray:
array of reflectance values
metadata:
dictionary containing the following metadata (all strings):
bad_band_window1: min and max wavelenths of first water vapor window (tuple)
bad_band_window2: min and max wavelenths of second water vapor window (tuple)
bands: # of bands (float)
coordinate system string: coordinate system information (string)
data ignore value: value corresponding to no data (float)
interleave: 'BSQ' (string)
reflectance scale factor: factor by which reflectance is scaled (float)
wavelength: wavelength values (float)
wavelength unit: 'm' (string)
spatial extent: extent of tile [xMin, xMax, yMin, yMax], UTM meters
--------
Example Execution:
--------
sercRefl, sercMetadata = h5refl2array('NEON_D02_SERC_DP1_20160807_160559_reflectance.h5') """
#Read in reflectance hdf5 file
hdf5_file = h5py.File(refl_filename,'r')
#Get the site name
file_attrs_string = str(list(hdf5_file.items()))
file_attrs_string_split = file_attrs_string.split("'")
sitename = file_attrs_string_split[1]
#Extract the reflectance & wavelength datasets
refl = hdf5_file[sitename]['Reflectance']
reflData = refl['Reflectance_Data']
reflArray = refl['Reflectance_Data'].value
#Create dictionary containing relevant metadata information
metadata = {}
metadata['map info'] = refl['Metadata']['Coordinate_System']['Map_Info'].value
metadata['wavelength'] = refl['Metadata']['Spectral_Data']['Wavelength'].value
#Extract no data value & set no data value to NaN
metadata['data ignore value'] = float(reflData.attrs['Data_Ignore_Value'])
metadata['reflectance scale factor'] = float(reflData.attrs['Scale_Factor'])
metadata['interleave'] = reflData.attrs['Interleave']
#Extract spatial extent from attributes
metadata['spatial extent'] = reflData.attrs['Spatial_Extent_meters']
#Extract bad band windows
metadata['bad_band_window1'] = (refl.attrs['Band_Window_1_Nanometers'])
metadata['bad_band_window2'] = (refl.attrs['Band_Window_2_Nanometers'])
#Extract projection information
metadata['projection'] = refl['Metadata']['Coordinate_System']['Proj4'].value
metadata['epsg'] = int(refl['Metadata']['Coordinate_System']['EPSG Code'].value)
#Extract map information: spatial extent & resolution (pixel size)
mapInfo = refl['Metadata']['Coordinate_System']['Map_Info'].value
hdf5_file.close
return reflArray, metadata
```
Now that the function is defined, we can call it to read in the sample reflectance file. Note that if your data is stored in a different location, you'll have to change the relative path, or include the absolute path.
```
# You will need to download the example dataset using the link above,
# then update the filepath below to fit your local file structure
h5refl_filename = '/Users/olearyd/Git/data/NEON_D02_SERC_DP3_368000_4306000_reflectance.h5'
data,metadata = read_neon_reflh5(h5refl_filename)
```
Let's take a quick look at the data contained in the `metadata` dictionary with a `for loop`:
```
for key in sorted(metadata.keys()):
print(key)
```
bad_band_window1
bad_band_window2
data ignore value
epsg
interleave
map info
projection
reflectance scale factor
spatial extent
wavelength
Now we can define a function that cleans the reflectance cube. Note that this also removes the water vapor bands, stored in the metadata as `bad_band_window1` and `bad_band_window2`, as well as the last 10 bands, which tend to be noisy. It is important to remove these values before doing classification or other analysis.
```
def clean_neon_refl_data(data,metadata):
"""Clean h5 reflectance data and metadata
1. set data ignore value (-9999) to NaN
2. apply reflectance scale factor (10000)
3. remove bad bands (water vapor band windows + last 10 bands):
Band_Window_1_Nanometers = 1340,1445
Band_Window_2_Nanometers = 1790,1955
"""
# use copy so original data and metadata doesn't change
data_clean = data.copy().astype(float)
metadata_clean = metadata.copy()
#set data ignore value (-9999) to NaN:
if metadata['data ignore value'] in data:
nodata_ind = np.where(data_clean==metadata['data ignore value'])
data_clean[nodata_ind]=np.nan
#apply reflectance scale factor (divide by 10000)
data_clean = data_clean/metadata['reflectance scale factor']
#remove bad bands
#1. define indices corresponding to min/max center wavelength for each bad band window:
bb1_ind0 = np.max(np.where((np.asarray(metadata['wavelength'])<float(metadata['bad_band_window1'][0]))))
bb1_ind1 = np.min(np.where((np.asarray(metadata['wavelength'])>float(metadata['bad_band_window1'][1]))))
bb2_ind0 = np.max(np.where((np.asarray(metadata['wavelength'])<float(metadata['bad_band_window2'][0]))))
bb2_ind1 = np.min(np.where((np.asarray(metadata['wavelength'])>float(metadata['bad_band_window2'][1]))))
bb3_ind0 = len(metadata['wavelength'])-10
#define valid band ranges from indices:
vb1 = list(range(0,bb1_ind0));
vb2 = list(range(bb1_ind1,bb2_ind0))
vb3 = list(range(bb2_ind1,bb3_ind0))
valid_band_range = [i for j in (range(0,bb1_ind0),
range(bb1_ind1,bb2_ind0),
range(bb2_ind1,bb3_ind0)) for i in j]
data_clean = data_clean[:,:,vb1+vb2+vb3]
metadata_clean['wavelength'] = [metadata['wavelength'][i] for i in valid_band_range]
return data_clean, metadata_clean
```
Now, use this function to pre-process the data:
```
data_clean,metadata_clean = clean_neon_refl_data(data,metadata)
```
Let's see the dimensions of the data before and after cleaning:
```
print('Raw Data Dimensions:',data.shape)
print('Cleaned Data Dimensions:',data_clean.shape)
```
Raw Data Dimensions: (1000, 1000, 426)
Cleaned Data Dimensions: (1000, 1000, 360)
Note that we have retained 360 of the 426 bands. This still contains plenty of information, in your processing, you may wish to subset even further. Let's take a look at a histogram of the cleaned data:
```
plt.hist(data_clean[~np.isnan(data_clean)],50);
```
Lastly, let's take a look at the data using the function `plot_aop_refl` function:
```
def plot_aop_refl(band_array,
refl_extent,
colorlimit=(0,1),
ax=plt.gca(),
title='',
cbar ='on',
cmap_title='',
colormap='Greys'):
plot = plt.imshow(band_array,extent=refl_extent,clim=colorlimit);
if cbar == 'on':
cbar = plt.colorbar(plot,aspect=40); plt.set_cmap(colormap);
cbar.set_label(cmap_title,rotation=90,labelpad=20);
plt.title(title); ax = plt.gca();
ax.ticklabel_format(useOffset=False, style='plain');
rotatexlabels = plt.setp(ax.get_xticklabels(),rotation=90);
plot_aop_refl(data_clean[:,:,0],
metadata_clean['spatial extent'],
(0,0.2))
```

## Unsupervised Classification with Spectral Unmixing:
### Endmember Extraction and Abundance Mapping
**Spectral Unmixing** allows pixels to be composed of fractions or abundances of each class.**Spectral Endmembers** can be thought of as the basis spectra of an image. Once these endmember spectra are determined, the image cube can be 'unmixed' into the fractional abundance of each material in each pixel (Winter, 1999).
**Spectral Angle Mapper (SAM):** is a physically-based spectral classification that uses an n-D angle to match pixels to reference spectra. The algorithm determines the spectral similarity between two spectra by calculating the angle between the spectra and treating them as vectors in a space with dimensionality equal to the number of bands. This technique, when used on calibrated reflectance data, is relatively insensitive to illumination and albedo effects. Endmember spectra used by SAM in this example are extracted from the NFINDR algorithm. SAM compares the angle between the endmember spectrum vector and each pixel vector in n-D space. Smaller angles represent closer matches to the reference spectrum. Pixels further away than the specified maximum angle threshold in radians are not classified.
Read more on Spectral Angle Mapper from
<a href="http://www.harrisgeospatial.com/docs/SpectralAngleMapper.html" target="_blank">Harris Geospatial</a>.
**Spectral Information Divergence (SID):** is a spectral classification method that uses a divergence measure to match pixels to reference spectra. The smaller the divergence, the more likely the pixels are similar. Pixels with a measurement greater than the specified maximum divergence threshold are not classified. Endmember spectra used by SID in this example are extracted from the NFINDR endmembor extraction algorithm.
Read more on Spectral Information Divergence from
<a href="http://www.harrisgeospatial.com/docs/SpectralInformationDivergence.html" target="_blank">Harris Geospatial</a>.
First we need to define the endmember extraction algorithm, and use the `extract` method to extract the endmembers from our data cube. You have to specify the # of endmembers you want to find, and can optionally specify a maximum number of iterations (by default it will use 3p, where p is the 3rd dimension of the HSI cube (m x n x p). For this example, we will specify a small # of iterations in the interest of time.
```
#eea = data_clean
ee = eea.NFINDR()
U = ee.extract(data_clean,4,maxit=5,
normalize=False,ATGP_init=True)
```
In order to display these endmember spectra, we need to define the endmember axes `dictionary`. Specifically we want to show the wavelength values on the x-axis. The `metadata['wavelength']` is a `list`, but the ee_axes requires a `float` data type, so we have to cast it to the right data type.
```
type(metadata_clean['wavelength'])
```
list
```
ee_axes = {} # set ee_axes data type to dictionary
# cast wavelength values to float to apply to ee_axes for display purposes
ee_axes['wavelength'] = [float(i) for i in metadata_clean['wavelength']]
ee_axes['x']='Wavelength, nm' #x axis label
ee_axes['y']='Reflectance' #y axis label
```
Now that the axes are defined, we can display the spectral endmembers with `ee.display`:
```
ee.display(axes=ee_axes,suffix='SERC')
```
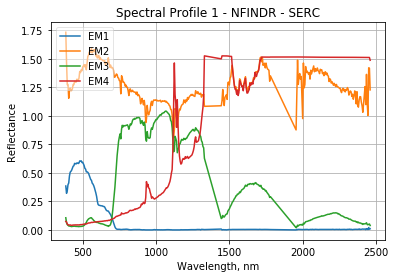
Now that we have extracted the spectral endmembers, we can take a look at the abundance maps for each member. These show the fractional components of each of the endmembers.
```
am = amap.FCLS() #define am object using the amap
amaps = am.map(data_clean,U,normalize=False) #create abundance maps for the HSI cubems
```
Use `am.display` to plot these abundance maps:
```
am.display(colorMap='jet',columns=4,suffix='SERC')
```
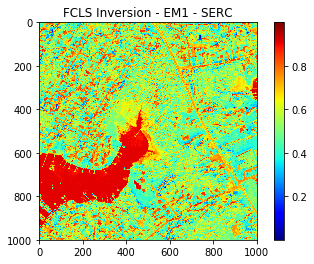
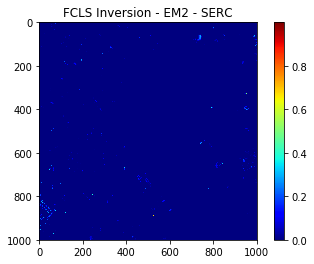
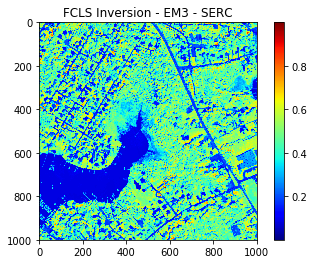
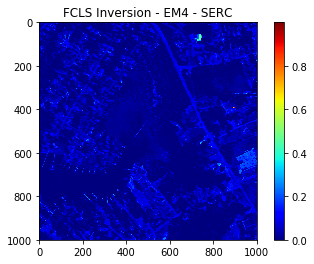
<Figure size 432x288 with 0 Axes>
Print mean values of each abundance map to better estimate thresholds to use in the classification routines.
```
print('Abundance Map Mean Values:')
print('EM1:',np.mean(amaps[:,:,0]))
print('EM2:',np.mean(amaps[:,:,1]))
print('EM3:',np.mean(amaps[:,:,2]))
print('EM4:',np.mean(amaps[:,:,3]))
```
Abundance Map Mean Values:
EM1: 0.59177357
EM2: 0.00089541974
EM3: 0.3809638
EM4: 0.026367119
You can also look at histogram of each abundance map:
```
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(18,8))
ax1 = fig.add_subplot(2,4,1); plt.title('EM1')
amap1_hist = plt.hist(np.ndarray.flatten(amaps[:,:,0]),bins=50,range=[0,1.0])
ax2 = fig.add_subplot(2,4,2); plt.title('EM2')
amap1_hist = plt.hist(np.ndarray.flatten(amaps[:,:,1]),bins=50,range=[0,0.001])
ax3 = fig.add_subplot(2,4,3); plt.title('EM3')
amap1_hist = plt.hist(np.ndarray.flatten(amaps[:,:,2]),bins=50,range=[0,0.5])
ax4 = fig.add_subplot(2,4,4); plt.title('EM4')
amap1_hist = plt.hist(np.ndarray.flatten(amaps[:,:,3]),bins=50,range=[0,0.05])
```
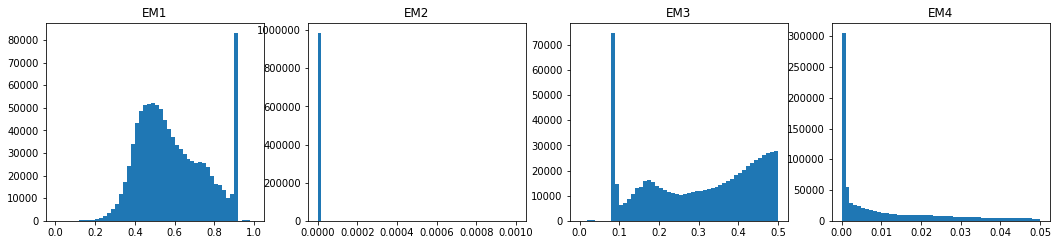
Below we define a function to compute and display Spectral Information Diverngence (SID):
```
def SID(data,E,thrs=None):
sid = cls.SID()
cmap = sid.classify(data,E,threshold=thrs)
sid.display(colorMap='tab20b',suffix='SERC')
```
Now we can call this function using the three endmembers (classes) that contain the most information:
```
U2 = U[[0,2,3],:]
SID(data_clean, U2, [0.8,0.3,0.03])
```
From this map we can see that SID did a pretty good job of identifying the water (dark blue), roads/buildings (orange), and vegetation (blue). We can compare it to the <a href="https://viewer.nationalmap.gov/" target="_blank">USA Topo Base map</a>.
<figure>
<a href="https://raw.githubusercontent.com/NEONScience/NEON-Data-Skills/main/graphics/hyperspectral-general/SERC_368000_4307000_UStopo.png">
<img src="https://raw.githubusercontent.com/NEONScience/NEON-Data-Skills/main/graphics/hyperspectral-general/SERC_368000_4307000_UStopo.png" width="300" height="300"></a>
<figcaption> The NEON SJER field site. Source: National Ecological Observatory Network (NEON)
</figcaption>
</figure>
## Challenges
1. On your own, try the Spectral Angle Mapper. If you aren't sure where to start, refer to
<a href="https://pysptools.sourceforge.io/classification.html#spectral-angle-mapper-sam" target="_blank">PySpTools SAM documentation</a>
and the
<a href="https://pysptools.sourceforge.io/examples_front.html#examples-using-the-ipython-notebook" target="_blank">Pine Creek example 1</a>.
**Hint**: use the SAM function below, and refer to the SID syntax used above.
```
def SAM(data,E,thrs=None):
sam = cls.SAM()
cmap = sam.classify(data,E,threshold=thrs)
sam.display(colorMap='Paired')
```
2. Experiment with different settings with SID and SAM (e.g., adjust the # of endmembers, thresholds, etc.)
3. Determine which algorithm (SID, SAM) you think does a better job classifying the SERC data tile. Synthesize your results in a markdown cell.
4. Take a subset of the bands before running endmember extraction. How different is the classification if you use only half the data points? How much faster does the algorithm run? When running analysis on large data sets, it is useful to
**Hints**:
* To extract every 10th element from the array `A`, use `A[0::10]`
* Import the package `time` to track the amount of time it takes to run a script.
```
#start_time = time.time()
# code
#elapsed_time = time.time() - start_time
```
## What Next?
`PySpTools` has an alpha interface with the Python machine learning package `scikit-learn`. To apply more advanced machine learning techniques, you may wish to explore some of these algorithms.
* <a href="https://pysptools.sourceforge.io/skl.html" target="_blank">Scikit-learn documentation on SourceForge</a>.
* <a href="http://scikit-learn.org/stable/" target="_blank">Scikit-learn website</a>.
| github_jupyter |
Maddie's portions of the final presentation notebook
```
import numpy as np
import matplotlib.pyplot as plt
from processing import *
from visualization import *
from sample import sample
image = np.load('image2.npy')
im = sample(image, 32)
W, H = run_nmf(im, 2)
```
## Motivations
Photo-induced force microscopy is a tool often used to visualize a variety of materials and how those materials' topographies, absorptions, and emissions behave under different excitation wavelengths.
Analysis of PiFM images relies on major assumptions:
- The ratio of the prepared solutions or samples is maintained post-preparation and during measurement
- Spectra of individual components can be parsed out of a spectrum of the mixed sample
A variety of techniques have been used to probe these assumptions, including Principal Component Analaysis and Principal Component Regression. These techniques allow us to better understand if intended ratios are maintained, how well mixed samples behave, and how sample preparation or sample interactions might affect spectra.
## So, what is NMF?
Non-negative Matrix Factorization (NMF) is a technique that has a variety of applications. It's most common application is in image analysis.
NMF approximates matrix $X$ by decomposing it into component matrices $W$ and $H$.
<h1><center>$X = WH$</center></h1>
The $W$ matrix is often considered the *weight* matrix while the $H$ matrix contains the given number of *components*.
```
fig, ax = plt.subplots(figsize = (18,7))
figure = plt.imread('nmf.PNG')
ax.imshow(figure)
ax.axis('off')
```
[Source for image.](https://blog.acolyer.org/2019/02/18/the-why-and-how-of-nonnegative-matrix-factorization/)
Our goal: Given a hyperspectral image (stacks of many images taken at different excitation wavenumbers), what are the weight matrices and component spectra of a mixed polymer sample?
How NMF helps:
- Non-negativitiy applied to all components (no such thing as a negative polymer!)
- Highly custommizable for >2 polymer blends
## Datasets
We'll be focusing on *hyperspectral* data sets. We often refer to these as data cubes. Essentially, for each excitation wavenumber of a given range, we take an image of our sample. You can imagine these images stacked on top of each other with the same x,y coordinates but different intensity values.
```
interactive_hyperimage(im)
```
## check_comps function
How can we assign NMF component spectra to ground truths?
First, let's get initialize a **dictionary** of our ground truth spectra.
```
ps_truth = np.load('ps_ground_truth.npy')
dppdtt_truth = np.load('dppdtt_ground_truth.npy')
ps = np.interp(ps_truth, (ps_truth.min(), ps_truth.max()), (0,1))
dppdtt = np.interp(dppdtt_truth, (dppdtt_truth.min(), dppdtt_truth.max()), (0,1))
truths = {'ps': ps, 'dppdtt': dppdtt}
```
Let's look at the logic inside `check_comps(comps, truths)`:
`matches = {}
for i in range(comps.shape[0]):
comps[i,:] = np.interp(comps[i,:], (comps[i,:].min(), comps[i,:].max()), (0,1))
for key in truths:
matches[key] = {'truth':truths[key],
'NMF':None,
'RMSE':None,
'Index':None}
rmse_vals = {}
for key in truths:
rmse_vals[key] = {}
for i in range(comps.shape[0]):
rmse_vals[key][i] = mean_squared_error(truths[key], comps[i,:], squared = False)
for i in range(comps.shape[0]):
val = 1000000000
min_key = None
for key in truths:
if val > rmse_vals[key][i]:
val = rmse_vals[key][i]
min_key = key
else:
pass
matches[min_key]['NMF'] = comps[i,:]
matches[min_key]['RMSE'] = val
matches[min_key]['Index'] = i
return matches`
So, with each step, we are minimizing the RMSE between the NMF component and ground truth.
```
matches = check_comps(H, truths)
fig, ax = plt.subplots(ncols = 2, figsize = (19,5))
x = np.arange(750,1878)
ax[0].plot(x, matches['ps']['NMF'], color = 'red', label = 'NMF Component')
ax[0].plot(x, matches['ps']['truth'], color = 'red', alpha = 0.7, label = 'Ground Truth')
ax[0].set_xlabel('Wavenumber $cm^{-1}$')
ax[0].set_ylabel('Intensity')
ax[0].set_title('PS')
ax[0].legend()
ax[1].plot(x, matches['dppdtt']['NMF'], color = 'blue', label = 'NMF Component')
ax[1].plot(x, matches['dppdtt']['truth'], color = 'blue', alpha = 0.7, label = 'Ground Truth')
ax[1].set_xlabel('Wavenumber $cm^{-1}$')
ax[1].set_ylabel('Intensity')
ax[1].set_title('DPPDTT')
ax[1].legend()
plt.show()
```
Our spectra match! Looks like NMF did a good job decomposing the original image and our `check_comps` function was able to correctly identify which is which!
```
plt.bar([0.5, 1.5],[matches['dppdtt']['RMSE'], matches['ps']['RMSE']])
plt.xticks([0.5,1.5], ['DPPDTT', 'PS'])
plt.title('RMSE for NMF Fits')
plt.show()
```
In addition to the ground truths, NMF components, and RMSE values saved in the `matches` dictionary, we also have a key for the *index* of the matched component. We can use that same index to parse through the W matrix.
## Building on this Project in the *Future*!
- modifying the `check_comps` function to take .json files instead of dictionaries of ground truths
- "Classifying" the package, or turning hyperspectral images into Classes with the analyses functions given above acting as methods
- making a pip installable package
- internally normalizing for intensity markers
| github_jupyter |
<a href="https://colab.research.google.com/github/kylemath/eeg-notebooks/blob/master/notebooks/CueingGroupAnalysis_Colab_Winter2019.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# CueingGroupAnalysis_Colab_Winter2019
The cueing task can ellicit a number of reliable changes. A central cue indicates the location of an upcoming target onset. Here the task can be changed to be perfectly predictive, or have some level of cue validity. Task is to indicate the orientation of a spatial grating on the target, up for vertical, right for horizontal.
ERP - Validly cued targets ellict larger ERP's than invalidly cued targets
Response ERPs - Validly cued targets are more quickly identified and better identified
Oscillations - Alpha power lateralizes after a spatial cue onset preceeding the upcoming onset of a target. Alpha power becomes smaller contraleral to the target side, and larger ipsilateral with the target.
```
!git clone https://github.com/kylemath/eeg-notebooks --recurse-submodules
%cd eeg-notebooks/notebooks
!pip install mne
from mne import Epochs, find_events, concatenate_raws
from mne.time_frequency import tfr_morlet
import numpy as np
import os
from utils import utils
from collections import OrderedDict
import warnings
warnings.filterwarnings('ignore')
from matplotlib import pyplot as plt
import matplotlib.patches as patches
```
# Load data into MNE objects
MNE is a very powerful Python library for analyzing EEG data. It provides helpful functions for performing key tasks such as filtering EEG data, rejecting artifacts, and grouping EEG data into chunks (epochs).
The first step after loading dependencies is use MNE to read the data we've collected into an MNE Raw object
```
# Fall 2018
# subs = [101, 102, 103, 104, 105, 106, 108, 109, 110, 111, 112,
# 202, 203, 204, 205, 207, 208, 209, 210, 211,
# 301, 302, 303, 304, 305, 306, 307, 308, 309]
# Winter 2019
subs = [1101, 1102, 1103, 1104, 1105, 1106, 1108, 1109, 1110,
1202, 1203, 1205, 1206, 1209, 1210, 1211, 1215,
1301, 1302, 1313,
1401, 1402, 1403, 1404, 1405, 1408, 1410, 1411, 1412, 1413, 1413, 1414, 1415, 1416]
# Both
# subs = [101, 102, 103, 104, 105, 106, 108, 109, 110, 111, 112,
# 202, 203, 204, 205, 207, 208, 209, 210, 211,
# 301, 302, 303, 304, 305, 306, 307, 308, 309,
# 1101, 1102, 1103, 1104, 1105, 1106, 1108, 1109, 1110,
# 1202, 1203, 1205, 1206, 1209, 1210, 1211, 1215,
# 1301, 1302, 1313,
# 1401, 1402, 1403, 1404, 1405, 1408, 1410, 1411, 1412, 1413, 1413, 1414, 1415, 1416]
# placeholders to add to for each subject
diff_out = []
Ipsi_out = []
Contra_out = []
Ipsi_spectra_out = []
Contra_spectra_out = []
diff_spectra_out = []
ERSP_diff_out = []
ERSP_Ipsi_out = []
ERSP_Contra_out = []
frequencies = np.linspace(6, 30, 100, endpoint=True)
wave_cycles = 6
# time frequency window for analysis
f_low = 7 # Hz
f_high = 10
f_diff = f_high-f_low
t_low = 0 # s
t_high = 1
t_diff = t_high-t_low
bad_subs= [6, 7, 13, 26]
really_bad_subs = [11, 12, 19]
sub_count = 0
for sub in subs:
print(sub)
sub_count += 1
if (sub_count in really_bad_subs):
rej_thresh_uV = 90
elif (sub_count in bad_subs):
rej_thresh_uV = 90
else:
rej_thresh_uV = 90
rej_thresh = rej_thresh_uV*1e-6
# Load both sessions
raw = utils.load_data('visual/cueing', sfreq=256.,
subject_nb=sub, session_nb=1)
raw.append( utils.load_data('visual/cueing', sfreq=256.,
subject_nb=sub, session_nb=2) )
# Filter Raw Data
raw.filter(1,30, method='iir')
#Select Events
events = find_events(raw)
event_id = {'LeftCue': 1, 'RightCue': 2}
epochs = Epochs(raw, events=events, event_id=event_id,
tmin=-1, tmax=2, baseline=(-1, 0),
reject={'eeg':rej_thresh}, preload=True,
verbose=False, picks=[0, 3])
print('Trials Remaining: ' + str(len(epochs.events)) + '.')
# Compute morlet wavelet
# Left Cue
tfr, itc = tfr_morlet(epochs['LeftCue'], freqs=frequencies,
n_cycles=wave_cycles, return_itc=True)
tfr = tfr.apply_baseline([-1,-.5],mode='mean')
power_Ipsi_TP9 = tfr.data[0,:,:]
power_Contra_TP10 = tfr.data[1,:,:]
# Right Cue
tfr, itc = tfr_morlet(epochs['RightCue'], freqs=frequencies,
n_cycles=wave_cycles, return_itc=True)
tfr = tfr.apply_baseline([-1,-.5],mode='mean')
power_Contra_TP9 = tfr.data[0,:,:]
power_Ipsi_TP10 = tfr.data[1,:,:]
# Compute averages Differences
power_Avg_Ipsi = (power_Ipsi_TP9+power_Ipsi_TP10)/2;
power_Avg_Contra = (power_Contra_TP9+power_Contra_TP10)/2;
power_Avg_Diff = power_Avg_Ipsi-power_Avg_Contra;
#output data into array
times = epochs.times
Ipsi_out.append(np.mean(power_Avg_Ipsi[np.argmax(frequencies>f_low):
np.argmax(frequencies>f_high)-1,
np.argmax(times>t_low):np.argmax(times>t_high)-1 ]
)
)
Ipsi_spectra_out.append(np.mean(power_Avg_Ipsi[:,np.argmax(times>t_low):
np.argmax(times>t_high)-1 ],1
)
)
Contra_out.append(np.mean(power_Avg_Contra[np.argmax(frequencies>f_low):
np.argmax(frequencies>f_high)-1,
np.argmax(times>t_low):np.argmax(times>t_high)-1 ]
)
)
Contra_spectra_out.append(np.mean(power_Avg_Contra[:,np.argmax(times>t_low):
np.argmax(times>t_high)-1 ],1))
diff_out.append(np.mean(power_Avg_Diff[np.argmax(frequencies>f_low):
np.argmax(frequencies>f_high)-1,
np.argmax(times>t_low):np.argmax(times>t_high)-1 ]
)
)
diff_spectra_out.append(np.mean(power_Avg_Diff[:,np.argmax(times>t_low):
np.argmax(times>t_high)-1 ],1
)
)
#save the spectrograms to average over after
ERSP_diff_out.append(power_Avg_Diff)
ERSP_Ipsi_out.append(power_Avg_Ipsi)
ERSP_Contra_out.append(power_Avg_Contra)
```
# Combine Subjects
```
#average spectrograms
GrandAvg_diff = np.nanmean(ERSP_diff_out,0)
GrandAvg_Ipsi = np.nanmean(ERSP_Ipsi_out,0)
GrandAvg_Contra = np.nanmean(ERSP_Contra_out,0)
#average spectra
GrandAvg_spec_Ipsi = np.nanmean(Ipsi_spectra_out,0)
GrandAvg_spec_Contra = np.nanmean(Contra_spectra_out,0)
GrandAvg_spec_diff = np.nanmean(diff_spectra_out,0)
#error bars for spectra (standard error)
num_good = len(diff_out) - sum(np.isnan(diff_out))
GrandAvg_spec_Ipsi_ste = np.nanstd(Ipsi_spectra_out,0)/np.sqrt(num_good)
GrandAvg_spec_Contra_ste = np.nanstd(Contra_spectra_out,0)/np.sqrt(num_good)
GrandAvg_spec_diff_ste = np.nanstd(diff_spectra_out,0)/np.sqrt(num_good)
##
#Plot Spectra error bars
fig, ax = plt.subplots(1)
plt.errorbar(frequencies,GrandAvg_spec_Ipsi,yerr=GrandAvg_spec_Ipsi_ste)
plt.errorbar(frequencies,GrandAvg_spec_Contra,yerr=GrandAvg_spec_Contra_ste)
plt.legend(('Ipsi','Contra'))
plt.xlabel('Frequency (Hz)')
plt.ylabel('Power (uV^2)')
plt.hlines(0,3,33)
#Plot Spectra Diff error bars
fig, ax = plt.subplots(1)
plt.errorbar(frequencies,GrandAvg_spec_diff,yerr=GrandAvg_spec_diff_ste)
plt.legend('Ipsi-Contra')
plt.xlabel('Frequency (Hz)')
plt.ylabel('Power (uV^2)')
plt.hlines(0,3,33)
##
#Grand Average Ipsi
plot_max = np.max([np.max(np.abs(GrandAvg_Ipsi)), np.max(np.abs(GrandAvg_Contra))])
fig, ax = plt.subplots(1)
im = plt.imshow(GrandAvg_Ipsi,
extent=[times[0], times[-1], frequencies[0], frequencies[-1]],
aspect='auto', origin='lower', cmap='coolwarm', vmin=-plot_max, vmax=plot_max)
plt.xlabel('Time (sec)')
plt.ylabel('Frequency (Hz)')
plt.title('Power Ipsi')
cb = fig.colorbar(im)
cb.set_label('Power')
# Create a Rectangle patch
rect = patches.Rectangle((t_low,f_low),t_diff,f_diff,linewidth=1,edgecolor='k',facecolor='none')
# Add the patch to the Axes
ax.add_patch(rect)
#Grand Average Contra
fig, ax = plt.subplots(1)
im = plt.imshow(GrandAvg_Contra,
extent=[times[0], times[-1], frequencies[0], frequencies[-1]],
aspect='auto', origin='lower', cmap='coolwarm', vmin=-plot_max, vmax=plot_max)
plt.xlabel('Time (sec)')
plt.ylabel('Frequency (Hz)')
plt.title('Power Contra')
cb = fig.colorbar(im)
cb.set_label('Power')
# Create a Rectangle patch
rect = patches.Rectangle((t_low,f_low),t_diff,f_diff,linewidth=1,edgecolor='k',facecolor='none')
# Add the patch to the Axes
ax.add_patch(rect)
#Grand Average Ipsi-Contra Difference
plot_max_diff = np.max(np.abs(GrandAvg_diff))
fig, ax = plt.subplots(1)
im = plt.imshow(
,
extent=[times[0], times[-1], frequencies[0], frequencies[-1]],
aspect='auto', origin='lower', cmap='coolwarm', vmin=-plot_max_diff, vmax=plot_max_diff)
plt.xlabel('Time (sec)')
plt.ylabel('Frequency (Hz)')
plt.title('Power Difference Ipsi-Contra')
cb = fig.colorbar(im)
cb.set_label('Ipsi-Contra Power')
# Create a Rectangle patch
rect = patches.Rectangle((t_low,f_low),t_diff,f_diff,linewidth=1,edgecolor='k',facecolor='none')
# Add the patch to the Axes
ax.add_patch(rect)
```
# Compute T-test
```
import scipy
num_good = len(diff_out) - sum(np.isnan(diff_out))
[tstat, pval] = scipy.stats.ttest_ind(diff_out,np.zeros(len(diff_out)),nan_policy='omit')
print('Ipsi Mean: '+ str(np.nanmean(Ipsi_out)))
print('Contra Mean: '+ str(np.nanmean(Contra_out)))
print('Mean Diff: '+ str(np.nanmean(diff_out)))
print('t(' + str(num_good-1) + ') = ' + str(round(tstat,3)))
print('p = ' + str(round(pval,3)))
```
# Save average powers ipsi and contra
```
import pandas as pd
print(diff_out)
raw_data = {'Ipsi Power': Ipsi_out,
'Contra Power': Contra_out}
df = pd.DataFrame(raw_data, columns = ['Ipsi Power', 'Contra Power'])
print(df)
df.to_csv('375CueingEEG.csv')
print('Saved subject averages for each condition to 375CueingEEG.csv file in present directory')
```
# Save Spectra
```
df = pd.DataFrame(Ipsi_spectra_out,columns=frequencies)
print(df)
df.to_csv('375CueingIpsiSpec.csv')
df = pd.DataFrame(Contra_spectra_out,columns=frequencies)
df.to_csv('375CueingContraSpec.csv')
print('Saved Spectra to 375CueingContraSpec.csv file in present directory')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/mrk-W2D1/tutorials/W2D1_BayesianStatistics/student/W2D1_Tutorial2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Neuromatch Academy: Week 2, Day 1, Tutorial 2
# Causal inference with mixture of Gaussians
__Content creators:__ Vincent Valton, Konrad Kording, with help from Matt Krause
__Content reviewers:__ Matt Krause, Jesse Livezey, Karolina Stosio, Saeed Salehi, Michael Waskom
# Tutorial Objectives
The previous notebook introduced Gaussians and Bayes' rule, allowing us to model very simple combinations of auditory and visual input. In this and the following notebook, we will use those building blocks to explore more complicated sensory integration and ventriloquism!
In this notebook, you will:
1. Learn more about the problem setting, which we wil also use in Tutorial 3,
2. Implement a mixture-of-Gaussian prior, and
3. Explore how that prior produces more complex posteriors.
```
# @title Video 1: Introduction
from IPython.display import YouTubeVideo
video = YouTubeVideo(id='GdIwJWsW9-s', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
---
##Setup
Please execute the cells below to initialize the notebook environment.
```
# imports
import numpy as np
import matplotlib.pyplot as plt
#@title Figure Settings
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
#@title Helper functions
def my_gaussian(x_points, mu, sigma):
"""
DO NOT EDIT THIS FUNCTION !!!
Returns normalized Gaussian estimated at points `x_points`, with parameters `mu` and `sigma`
Args:
x_points (numpy array of floats) - points at which the gaussian is evaluated
mu (scalar) - mean of the Gaussian
sigma (scalar) - standard deviation of the gaussian
Returns:
(numpy array of floats): normalized Gaussian (i.e. without constant) evaluated at `x`
"""
px = np.exp(- 1/2/sigma**2 * (mu - x_points) ** 2)
px = px / px.sum() # this is the normalization part with a very strong assumption, that
# x_points cover the big portion of probability mass around the mean.
# Please think/discuss when this would be a dangerous assumption.
return px
def plot_mixture_prior(x, gaussian1, gaussian2, combined):
"""
DO NOT EDIT THIS FUNCTION !!!
Plots a prior made of a mixture of gaussians
Args:
x (numpy array of floats): points at which the likelihood has been evaluated
gaussian1 (numpy array of floats): normalized probabilities for Gaussian 1 evaluated at each `x`
gaussian2 (numpy array of floats): normalized probabilities for Gaussian 2 evaluated at each `x`
posterior (numpy array of floats): normalized probabilities for the posterior evaluated at each `x`
Returns:
Nothing
"""
fig, ax = plt.subplots()
ax.plot(x, gaussian1, '--b', LineWidth=2, label='Gaussian 1')
ax.plot(x, gaussian2, '-.b', LineWidth=2, label='Gaussian 2')
ax.plot(x, combined, '-r', LineWidth=2, label='Gaussian Mixture')
ax.legend()
ax.set_ylabel('Probability')
ax.set_xlabel('Orientation (Degrees)')
```
# Section 1: Motivating example
Ventriloquists produce the illusion that their puppets are talking because:
1. We observe the visual input of the puppet moving its mouth, as if speaking.
2. The speech that the puppeteer generates originates near the puppet's mouth.
Since we are accustomed to voices coming from moving mouths, we tend to interpret the voice as coming directly from the puppet itself rather than from the puppeteer (who is also hiding his/her own mouth movements). In the remaining tutorials, we will study how this illusion breaks down as the distance between the visual stimulus (the puppet's mouth) and the auditory stimulus (the puppeter's concealed speech) changes.
Imagine an experiment where participants are shown a puppet moving its mouth at a location directly in front of them (at position 0˚). The subjects are told that 75% of the time, the voice they hear originates from the puppet. On the remaining 25% of trials, sounds come from elsewhere. Participants learn this over multiple trials, after which a curtain is dropped in front of the puppeteer and the puppet.
Next, we present only the auditory stimulus at varying locations and we ask participants to report where the source of the sound is located. The participants have access to two pieces of information:
* The prior information about sound localization, learned during the trials before the curtain fell.
* Their noisy sensory estimates about where a particular sound originates.
Our eventual goal, which we achieve in Tutorial 3, is to predict the subjects' responses: when do subjects ascribe a sound to the puppet, and when do they believe it originated elsewhere? Doing so requires building a prior that captures the participant's knowledge and expectations, which we wil do in the exercises that follow here.
# Section 2: Mixture-of-Gaussians Prior
In the previous tutorial, you learned how to create a single Gaussian prior that could represent one of these possibilties. A broad Gaussian with a large $\sigma$ could represent sounds originating from nearly anywhere, while a narrow Gaussian with $\mu$ near zero could represent sounds orginating from the puppet.
Here, we will combine those into a mixure-of-Gaussians probability density function (PDF) that captures both possibilties. We will control how the Gaussians are mixed by summing them together with a 'mixing' or weight parameter $p_{common}$, set to a value between 0 and 1, like so:
\begin{eqnarray}
\text{Mixture} = \bigl[\; p_{common} \times \mathcal{N}(\mu_{common},\sigma_{common}) \; \bigr] + \bigl[ \;\underbrace{(1-p_{common})}_{p_{independent}} \times \mathcal{N}(\mu_{independent},\sigma_{independent}) \; \bigr]
\end{eqnarray}
$p_{common}$ denotes the probability that auditory stimulus shares a "common" source with the learnt visual input; in other words, the probability that the "puppet" is speaking. You might think that we need to include a separate weight for the possibility that sound is "independent" from the puppet. nHowever, since there are only two, mutually-exclusive possibilties, we can replace $p_{independent}$ with $(1 - p_{common})$ since, by the law of total probability, $p_{common} + p_{independent}$ must equal one.
Using the formula above, complete the code to build this mixture-of-Gaussians PDF:
* Generate a Gaussian with mean 0 and standard deviation 0.5 to be the 'common' part of the Gaussian mixture prior. (This is already done for you below).
* Generate another Gaussian with mean 0 and standard deviation 3 to serve as the 'independent' part.
* Combine the two Gaussians to make a new prior by mixing the two Gaussians with mixing parameter $p_{common}$ = 0.75 so that the peakier "common-cause" Gaussian has 75% of the weight. Don't forget to normalize afterwards!
Hints:
* Code for the `my_gaussian` function from Tutorial 1 is available for you to use. Its documentation is below.
**Helper function(s)**
```
help(my_gaussian)
```
## Exercise 1: Implement the prior
```
def mixture_prior(x, mean=0, sigma_common=0.5, sigma_independent=3, p_common=0.75):
###############################################################################
## Insert your code here to:
# * Create a second gaussian representing the independent-cause component
# * Combine the two priors, using the mixing weight p_common. Don't forget
# to normalize the result so it remains a proper probability density function
#
# * Comment the line below to test out your function
raise NotImplementedError("Please complete Exercise 1")
###############################################################################
gaussian_common = my_gaussian(x, mean, sigma_common)
gaussian_independent = ...
mixture = ...
return gaussian_common, gaussian_independent, mixture
x = np.arange(-10, 11, 0.1)
# Uncomment the lines below to visualize out your solution
# common, independent, mixture = mixture_prior(x)
# plot_mixture_prior(x, common, independent, mixture)
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D1_BayesianStatistics/solutions/W2D1_Tutorial2_Solution_01b53c96.py)
*Example output:*
<img alt='Solution hint' align='left' width=424 height=280 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D1_BayesianStatistics/static/W2D1_Tutorial2_Solution_01b53c96_2.png>
# Section 3: Bayes Theorem with Complex Posteriors
```
#@title Video 2: Mixture-of-Gaussians and Bayes' Theorem
video = YouTubeVideo(id='LWKM35te0WI', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
Now that we have created a mixture of Gaussians prior that embodies the participants' expectations about sound location, we want to compute the posterior probability, which represents the subjects' beliefs about a specific sound's origin.
To do so we will compute the posterior by using *Bayes Theorem* to combine the mixture-of-gaussians prior and varying auditory Gaussian likelihood. This works exactly the same as in Tutorial 1: we simply multiply the prior and likelihood pointwise, then normalize the resulting distribution so it sums to 1. (The closed-form solution from Exercise 2B, however, no longer applies to this more complicated prior).
Here, we provide you with the code mentioned in the video (lucky!). Instead, use the interactive demo to explore how a mixture-of-Gaussians prior and Gaussian likelihood interact. For simplicity, we have fixed the prior mean to be zero. We also recommend starting with same other prior parameters used in Exercise 1: $\sigma_{common} = 0.5, \sigma_{independent} = 3, p_{common}=0.75$; vary the likelihood instead.
Unlike the demo in Tutorial 1, you should see several qualitatively different effects on the posterior, depending on the relative position and width of likelihood. Pay special attention to both the overall shape of the posterior and the location of the peak. What do you see?
## Interactive Demo 1: Mixture-of-Gaussian prior and the posterior
```
#@title
#@markdown Make sure you execute this cell to enable the widget!
fig_domain = np.arange(-10, 11, 0.1)
import ipywidgets as widgets
def refresh(sigma_common=0.5, sigma_independent=3, p_common=0.75, mu_auditory=3, sigma_auditory=1.5):
_, _, prior = mixture_prior(fig_domain, 0, sigma_common, sigma_independent, p_common)
likelihood = my_gaussian(fig_domain, mu_auditory, sigma_auditory)
posterior = prior * likelihood
posterior /= posterior.sum()
plt.plot(fig_domain, prior, label="Mixture Prior")
plt.plot(fig_domain, likelihood, label="Likelihood")
plt.plot(fig_domain, posterior, label="Posterior")
plt.legend()
plt.show()
style = {'description_width': 'initial'}
_ = widgets.interact(refresh,
sigma_common=widgets.FloatSlider(value=0.5, min=0.01, max=10, step=0.5, description="sigma_common", style=style),
sigma_independent=widgets.FloatSlider(value=3, min=0.01, max=10, step=0.5, description="sigma_independent:", style=style),
p_common=widgets.FloatSlider(value=0.75, min=0, max=1, steps=0.01, description="p_common"),
mu_auditory=widgets.FloatSlider(value=2, min=-10, max=10, step=0.1, description="mu_auditory:", style=style),
sigma_auditory=widgets.FloatSlider(value=0.5, min=0.01, max=10, step=0.5, description="sigma_auditory:", style=style),
)
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D1_BayesianStatistics/solutions/W2D1_Tutorial2_Solution_9569e15c.py)
# Section 3: Conclusion
```
#@title Video 3: Outro
video = YouTubeVideo(id='UgeAtE8xZT8', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
In this tutorial, we introduced the ventriloquism setting that will form the basis of Tutorials 3 and 4 as well. We built a mixture-of-Gaussians prior that captures the participants' subjective experiences. In the next tutorials, we will use these to perform causal inference and predict the subject's responses to indvidual stimuli.
| github_jupyter |
# Hyper-parameter tuning for GANITE
This notebook presents the solution for hyper-parameter searching for the __GANITE__(for both Tensorflow and PyTorch version) algorithm over the [Twins](https://bitbucket.org/mvdschaar/mlforhealthlabpub/src/master/data/twins/) dataset.
For details about each algorithm, please refer to their dedicated notebooks:
- [GANITE(Tensorflow) notebook](https://github.com/bcebere/ite-api/blob/main/notebooks/ganite_train_evaluation.ipynb).
- [GANITE(PyTorch) notebook](https://github.com/bcebere/ite-api/blob/main/notebooks/ganite_pytorch_train_evaluation.ipynb).
## Hyper-parameter tuning
Hyperparameter tuning refers to performing a search to discover the model parameters that result in the model's best performance on a specific dataset.
One algorithm for performing hyperparameter optimization is the [__Bayesian Optimization__](https://en.wikipedia.org/wiki/Bayesian_optimization).
__Bayesian Optimization__ provides a principled technique based on [Bayes Theorem](https://en.wikipedia.org/wiki/Bayes%27_theorem) to direct a search of a global optimization problem that is efficient and effective. It works by building a probabilistic model of the objective function, called the surrogate function, that is then searched efficiently with an acquisition function before candidate samples are chosen to evaluate the real objective function.
For the tuning, we use the [__Scikit-Optimize__](https://scikit-optimize.github.io/stable/) library, which provides a general toolkit for [Bayesian Optimization](https://en.wikipedia.org/wiki/Bayesian_optimization) that can be used for hyperparameter tuning.
For __GANITE__, we try to optimize the following hyperparameters using the ranges suggested in [[3] Table 6](https://openreview.net/forum?id=ByKWUeWA-):
| Hyperparameter | Search area | Description |
| --- | --- | --- |
| dim_hidden | {dim, int(dim/2), int(dim/3), int(dim/4), int(dim/5)} | the size of the hidden layers. |
| depth |{1, 3, 5, 7, 9} | the number of hidden layers in the generator and inference blocks. |
| alpha | {0, 0.1, 0.5, 1, 2, 5, 10} | weight for the Generator block loss. |
| beta | {0, 0.1, 0.5, 1, 2, 5, 10} | weight for the ITE block loss. |
| num_discr_iterations | [3, 10] | number of iterations executed by the Counterfactual discriminator. |
| minibatch_size | {32, 64, 128, 256} | the size of the dataset batches. |
You can find the __GANITE__ hyperparameter tuning implementation [here](https://github.com/bcebere/ite-api/blob/main/src/ite/algs/hyperparam_tuning.py).
## Setup
First, make sure that all the depends are installed in the current environment.
```
pip install -r requirements.txt
pip install .
```
Next, we import all the dependencies necessary for the task.
```
import ite.algs.hyperparam_tuning as tuning
from IPython.display import HTML, display
import tabulate
param_search_names = ["num_discr_iterations", "minibatch_size", "dim_hidden", "alpha", "beta", "depth"]
```
### GANITE(Tensorflow)
```
tf_best_params = tuning.search("GANITE", iterations=5000)
```
### Hyper-parameter tuning for GANITE(Tensorflow)
```
display(HTML(tabulate.tabulate([tf_best_params], headers=param_search_names, tablefmt='html')))
```
### GANITE (PyTorch)
```
torch_best_params = tuning.search("GANITE_TORCH")
```
### Hyper-parameter tuning results for GANITE(PyTorch)
```
display(HTML(tabulate.tabulate([torch_best_params], headers=param_search_names, tablefmt='html')))
```
## References
1. [Scikit-Optimize for Hyperparameter Tuning in Machine Learning](https://machinelearningmastery.com/scikit-optimize-for-hyperparameter-tuning-in-machine-learning).
2. [scikit-optimize](https://scikit-optimize.github.io/).
3. Jinsung Yoon, James Jordon, Mihaela van der Schaar, "GANITE: Estimation of Individualized Treatment Effects using Generative Adversarial Nets", International Conference on Learning Representations (ICLR), 2018 ([Paper](https://openreview.net/forum?id=ByKWUeWA-)).
| github_jupyter |
# Using Amazon SageMaker with Public Datasets
__*Clustering Gene Variants into Geographic Populations*__
## Introduction
Amazon SageMaker allows you to bring powerful machine learning workflows to data that is already in the cloud. In this example, we will do just that - combining Amazon SageMaker with data from the [1000 Genomes Project] which is hosted by AWS as a [public dataset]. Specifically, we will perform unsupervised learning using Amazon SageMaker's KMeans algorithm to see if we can predict the geographic population for a set of single nucleotide polymorphisms.
Single nucleotide polymorphisms or SNPs (pronounced "snips") are single base-pair changes to DNA. DNA is a long chain molecule that is used to store the "source code" for all living organisms and is "read" as a sequence of four nucleotides: A, T, C, and G. A single letter is called a "base". SNPs occur when one of these bases in the sequence changes due to environmental causes or random replication errors during cell division in germ cells (eggs and sperm). Sometimes these changes are harmless, and sometimes they can cause serious diseases.
Here we are going to cluster high frequency SNPs found on Chromosome 6
### Attribution
This notebook is based on work previously described by [Databricks using Spark][databricks blog]
[1000 Genomes Project]: https://aws.amazon.com/1000genomes/
[public dataset]: https://aws.amazon.com/public-datasets/
[databricks blog]: https://databricks.com/blog/2016/05/24/predicting-geographic-population-using-genome-variants-and-k-means.html
## Setup
> This notebook was created and tested on an `ml.m4.2xlarge` notebook instance
Let's start by:
1. Downloading the data we need from S3
1. Installing some utility packages for processing the data
### Data sources
We can get variant call data (which describes SNPs, and other kinds of DNA sequence modifications) from the publicly hosted 1000 Genomes dataset on AWS. We are need the "\*.vcf" file corresponding to Chromosome 6 from the 20130502 release of the data.
```
%%bash
aws s3 ls --human-readable s3://1000genomes/release/20130502/ | grep chr6
```
The data for Chromosome 6 is nearly 1GB in size. For the purpose of this exercise and to be conservative of space (the scratch area for sagemaker notebooks only have about 5GB of space) we are going to use a sub-sample of the data. To generate that, we can use `tabix` a bioinformatics command line utility found in the [htslib] set of tools.
[htslib]: https://github.com/samtools/htslib
The current version of `tabix` (1.8) has been containerized and is hosted on Amazon ECR. We'll pull down the docker container image into our SageMaker environment and use it to sample the Chromosome 6 VCF file __*directly on S3*__ and create a data file we can use here for model training. Here we've reduced the data to entries found between positions 1000000-1250000.
```
%%bash
docker run --rm -i \
733263974272.dkr.ecr.us-west-2.amazonaws.com/htslib:latest \
tabix -h \
s3://1000genomes/release/20130502/ALL.chr6.phase3_shapeit2_mvncall_integrated_v5a.20130502.genotypes.vcf.gz \
6:1000000-1250000 > 6-sample.vcf
```
Now let's grab metadata - information about geographic locations of where sample sequences came from - to use as labels in our model training.
```
%%bash
aws s3 cp s3://1000genomes/release/20130502/integrated_call_samples_v3.20130502.ALL.panel .
```
## Exploration
To make exploring and processing the data a little easier, we'll use the `scikit-allel` package. While this package does not come included with the SageMaker environment, it is easy to install.
More information about it can be found at:
http://scikit-allel.readthedocs.io/en/latest/index.html
It has good utilities for reading VCF files
http://alimanfoo.github.io/2017/06/14/read-vcf.html
```
!conda install -c conda-forge -y scikit-allel
```
Now we can easily read in our sampled VCF file for data exploration
```
import allel
callset = allel.read_vcf('6-sample.vcf')
```
The above returns a Python dictionary with keys that represent parts of the data. These are specific parts of the VCF file that are useful for analysis. For example the `calldata/GT` key contains and array of all the genotype calls for each variant for each sample.
```
callset.keys()
```
How many variants and samples are in this data?
```
print("Samples: {samples}, Variants: {variants}".format(
variants=len(callset['calldata/GT']),
samples=len(callset['samples']))
)
```
The data comes from human genome sequences. Humans have 23 pairs of chromosomes, and hence are "diploid" - meaning they should have two copies of any given DNA sequence (with a couple exceptions - e.g. genes in the XY chromosomes).
A variant in a copy of a DNA sequence is called an "allele". At minimum, there is at least one allele - the DNA sequence that matches the human reference genome. Alleles that do not match the reference are called "alternates".
There appear to be up to 6 alternate alleles for each variant.
```
import numpy as np
np.unique(callset['calldata/GT'])
```
A genotype is a combination of variants for a DNA sequence position, over all copies. For example, let's say that the reference for a DNA position is 'A', and a variant for the position is 'T'. The possible genotypes for this position would be:
* REF / REF - "homozygous" for the reference
* REF / ALT - "heterozygous"
* ALT / REF - "heterozygous"
* ALT / ALT - "homozygous" for the alternate
Typical genotype calls use integer IDs to represent the REF and ALT alleles, with REF always being '0'. Alternative alleles start at '1' and count up to the total number of alternative alleles for the variant. So the possible genotypes for the example above would be:
* 0 / 0
* 0 / 1
* 1 / 0
* 1 / 1
In cases where there is more than one alternate allele, you might see genotypes like '0 / 2' or '1 / 2'.
All the genotypes in the data can be collected in a `GenotypeArray` object
```
gt = allel.GenotypeArray(callset['calldata/GT'])
gt
```
Sometimes sequence isn't perfect - base calling can produce ambiguous results. This results in missing variant calls and incomplete genotypes. For any machine learning task, it is important to know if there is missing data and deal with it accordingly.
The `GenotypeArray` also tells us that there are no missing calls
```
gt.count_missing()
```
As mentioned above, variants are a combination of SNPs, InDels, and Copy Number variants (duplications of DNA, beyond the chromosome count). We can see that in the data.
```
np.unique(callset['variants/ALT'])
```
## The Modeling Problem
### Feature selection
For this modeling exercise, we are going to use the variant "ID" - combination of the chromosome position, the reference allele, and the alternative allele as features for K-Means clustering. Above we saw that there were about 8300 variants. We want to reduce this down to a more manageable set, which can improve our clustering performance.
To start, we'll just focus on SNPs that are bi-allelic variants. That is a SNP with only one alternate allele. To do this, we filter for entries where there is only 1 nucleotide in the REF and ALT lists.
There are 8090 variants with single nucleotide reference alleles
```
REF = callset['variants/REF']
REF
is_1bp_ref = np.array(list(map(lambda r: len(r) == 1, REF)), dtype=bool)
sum(is_1bp_ref)
```
There are 8147 variants with single nucleotide alternate alleles
```
# the default import considers only 3 allele alternatives
ALT = callset['variants/ALT']
ALT
is_1bp_alt = np.array(list(map(lambda a: len(a[0]) == 1 and not any(a[1:]), ALT)), dtype=bool)
sum(is_1bp_alt)
```
The intersection of the above yields 7946 SNP variants
```
is_snp_var = is_1bp_ref & is_1bp_alt
sum(is_snp_var)
```
We can reduce the feature set further by only considering variants with alternate allele frequencies > 30%. This will eliminate rare variants that won't help our clustering.
```
# get a count of alleles for each variant
ac = gt.count_alleles()
ac
# since we're only looking for only bi-allelic SNPs, we're only concerned with
# the reference and first alternative allele
is_hifreq_snp_var = is_snp_var & (ac[:, 1] / (ac[:, 0] + ac[:, 1]) > .30)
sum(is_hifreq_snp_var)
```
We are now down to ~376 features, which is certainly more manageable than the ~8300 we started with
### Data transformation
Machine learning algorithms work best on numerical data.
We can convert the genotypes into integer values easily using bit-packing provided by the `GenotypeArray.to_packed` method
```
xx = allel.GenotypeArray([
[[0,0], [0,1], [1,0], [1,1]]
])
xx
xx.to_packed()
```
From the above we see that a values of 1 and 16 are effectively equivalent - they correspond to the same genotype: heterozygous for the alt allele. Where 17 is homozygous for the alt allele. We can recode these to:
* 16 --> 1
* 17 --> 2
We'll apply this transformation to our GenoTypeArray, which we'll use as our data for training, and apply the filter for only high frequency SNPs that we generated above. We also want the data entries to list samples along the rows and variants along the columns, so we'll use the transpose of the coded GenoTypeArray.
```
gt_coded = gt.to_packed()[is_hifreq_snp_var,:]
gt_coded[gt_coded == 16] = 1
gt_coded[gt_coded == 17] = 2
gt_coded.transpose()
```
Use `CHROM`, `POS`, `REF`, and `ALT` fields from the variant data to create variant feature IDs
```
features = [
'{}-{}-{}-{}'.format(c, p, r, a[0])
for c, p, r, a in zip(
callset['variants/CHROM'],
callset['variants/POS'],
callset['variants/REF'],
callset['variants/ALT']
)]
features = np.array(features)[is_hifreq_snp_var]
features[:10]
```
Let's read in the panel metadata to get class labels - the geographic location that each sample originated from. There are many popluations in the data set. For this example, we'll only focus on the following populations:
* GBR: British from England and Scotland
* ASW: African Ancestry in Southwest US
* CHB: Han Chinese in Bejing, China
Here we'll use `pandas` to process the metadata panel into classes we can use
```
import pandas as pd
classes = pd.read_table('integrated_call_samples_v3.20130502.ALL.panel', usecols=['sample', 'pop'])
classes = classes[classes['pop'].isin(['GBR', 'ASW', 'CHB'])].copy()
classes.head()
```
Let's see how the data is distributed across each of our target populations:
```
%matplotlib inline
classes.groupby('pop').count().reset_index().plot.bar('pop')
```
Based on the chart above, the distribution of samples across these three populations looks reasonable - i.e. each group has roughly the same number of samples.
Let's now create the data frame to feed into model training
```
data = classes.merge(
pd.concat((
pd.Series(callset['samples'], name='sample'),
pd.DataFrame(gt_coded.transpose(), columns=features)),
axis=1),
on='sample'
)
data.sample(10)
```
After all of our processing, we have a data set with 255 samples and ~376 features
```
data.shape
```
## Training
This dataset is small, only 255 observations, but should give an idea of how to use the built-in KMeans algorithm.
Let's use an 80/20 ratio a train/test split. We'll train the KMeans clustering model with the `train` set and use the `test` set to evaluate predictions. To prepare for training, we need to remove all non-numeric values, so below we'll drop the `pop` field from the coded genotype data and store it with labels that we can use later.
```
from math import ceil, floor
train_data = data.sample(frac=.8, random_state=1024)
test_data = data[~data['sample'].isin(train_data['sample'])].copy()
train_labels = train_data[['sample', 'pop']].copy().set_index('sample')
train_labels['pop'] = pd.Categorical(train_labels['pop'])
train_data = train_data.drop(columns='pop').set_index('sample')
test_labels = test_data[['sample', 'pop']].copy().set_index('sample')
test_labels['pop'] = pd.Categorical(test_labels['pop'])
test_data = test_data.drop(columns='pop').set_index('sample')
print('Observations')
print(f'training: {train_data.shape[0]}')
print(f'test: {test_data.shape[0]}')
```
Here's the standard setup for SageMaker training using KMeans.
Be sure to set the `bucket` name to something you have access to.
The fitting process will upload the training data to this bucket for the training instance(s) to access. Once training is done, a model will be uploaded to the bucket.
```
from sagemaker import KMeans, get_execution_role
role = get_execution_role()
bucket = '<your_s3_bucket_name_here>'
data_location = 's3://{}/sagemaker/genome-kmeans/data'.format(bucket)
output_location = 's3://{}/sagemaker/genome-kmeans/output'.format(bucket)
print('training data will be uploaded to: {}'.format(data_location))
print('training artifacts will be uploaded to: {}'.format(output_location))
kmeans = KMeans(role=role,
train_instance_count=2,
train_instance_type='ml.c4.8xlarge',
output_path=output_location,
k=3,
data_location=data_location)
```
Time to train the model. This should take only about 5-9 minutes.
```
%%time
kmeans.fit(kmeans.record_set(np.float32(train_data.values)))
```
## Inference Endpoint Deployment
Now, let's deploy the model behind an endpoint we can use for predictions. This process takes about 5-9 mins.
```
%%time
kmeans_predictor = kmeans.deploy(initial_instance_count=1,
instance_type='ml.m4.xlarge')
```
Let's use our newly deployed endpoint to test the model.
The predictor will return a results object from which we can extract the cluster assignments for each sample.
```
%%time
result = kmeans_predictor.predict(np.float32(train_data))
clusters = np.int0([r.label['closest_cluster'].float32_tensor.values[0] for r in result])
```
Let's see how these predicted clusters map to the real classes
First, how well did the training set cluster?
```
pd.crosstab(train_labels['pop'], columns=clusters, colnames=['cluster'])
```
From this cross tabulation we see that there are clusters with majority membership in each of our populations.
What do the clusters look like visually? To answer this question, we'll generate a force weighted graph of the clusters and color code them by their original population code.
To accomplish this, well use the [lightning-viz](http://lightning-viz.org/) package for Python.
```
!pip install lightning-python
from lightning import Lightning
lgn = Lightning(ipython=True, local=True)
graph_data = [
{
'cluster': int(r.label['closest_cluster'].float32_tensor.values[0]),
'distance': float(r.label['distance_to_cluster'].float32_tensor.values[0])
}
for r in result
]
gg = pd.concat(
(train_labels.reset_index(),
pd.DataFrame(graph_data)),
axis=1
)
gg['code'] = pd.np.NaN # place holder for population category codes
gg = pd.concat(
(pd.DataFrame({
'cluster': [0,1,2],
'distance': 0,
'sample': ['0', '1', '2'],
'pop': ''}
), gg)).reset_index().drop(columns='index')
gg['code'] = pd.Categorical(gg['pop']).codes
# generate the network links and plot
nn = [(r[0], r[1], r[2]) for r in gg.to_records()]
lgn.force(nn, group=gg['code'], labels=gg['sample'] + '\n' + gg['pop'])
```
The clustering results are roughly the same on the test data
```
result = kmeans_predictor.predict(np.float32(test_data))
clusters = np.int0([r.label['closest_cluster'].float32_tensor.values[0] for r in result])
pd.crosstab(test_labels['pop'], columns=clusters, colnames=['cluster'])
```
## Bottom Line
The mixture of populations in the clusters may be interpretted as individuals with mixed ancestry. Also, the clustering could be improved further if there was additional dimensionality reduction (e.g. via PCA), more samples, or both.
### (Optional) Delete the Endpoint
If you're ready to be done with this notebook, make sure run the cell below. This will remove the hosted endpoint you created and avoid any charges from a stray instance being left on.
```
print(kmeans_predictor.endpoint)
import sagemaker
sagemaker.Session().delete_endpoint(kmeans_predictor.endpoint)
```
| github_jupyter |
```
#导入matplotlib
import matplotlib.pyplot as plt
import numpy as np
n = 1024
X = np.random.normal(10,10,n)
Y = np.random.normal(20,10,n)
plt.scatter(X,Y) #绘制散点图
plt.show() #显示图像
import matplotlib.pyplot as plt
import numpy as np
def f(x,y): return (1-x/2+x**5+y**3)*np.exp(-x**2-y**2)
n = 10
x = np.linspace(-3,3,n)
y = np.linspace(-3,3,n)
X,Y = np.meshgrid(x,y)
plt.scatter(X,Y) #绘制散点图
plt.show()
Y
import numpy as np
from numpy import *
import pandas as pd
df = pd.read_csv('data.csv',encoding='gbk')
df.head(35)
df.info()
#数据清洗
df=df.dropna(how="all")
df=df.drop([0])#删除年份及空行这两个无关行
df
todel=[]
for i in range(df.shape[0]):
sum = 0
for j in range(df.shape[1]):
if pd.isnull(df.iloc[i,j]):
sum+=1
if sum>=2:
todel.append(i+1)
break
df=df.drop(todel)
#删除缺失值超过1的行
# todel=[]
# for index, row in df.iterrows():
# lie=0
# sum = 0
# for i in row:
# if pd.isnull(i):
# sum+=1
# if sum>=2:
# todel.append(lie)
# break
# lie+=1
# print(todel[0])
df
type(sum(pd.isnull(df[0:1])))
df[1:2].count()
len(df)
pd.isnull(df[0:1])
df[0:1]
len(df.index)
df.shape[1]
pd.isnull(df.iloc[3,3])
df.iloc[26,2]
df.to_excel('datawash.xls')#之后在datawash中手动修改标号
df = pd.read_excel('datawash.xls',encoding='gbk')
df
#缺失值处理
from scipy.interpolate import lagrange
def ploy(s,n,k=6):
y=s[list(range(n-k,n))+list(range(n+1,n+1+k))]#取数
y=y[y.notnull()]
return lagrange(y.index,list(y))(n)
for i in df.columns:
for j in range(len(df)):
if(df[i].isnull())[j]:
df[i][j]=ploy(df[i],j)
df.to_excel('data222.xls')
(df["农村居民家庭人均可支配收入(元/人)"].isnull())[1]
df = pd.read_excel('data222.xls',encoding='gbk')
import numpy as np
import math as math
dataset = np.array([[3,5,1,4,1],
[4,4,3,5,3],
[3,4,4,4,4],
[3,3,5,2,1],
[3,4,5,4,3]])
def corr(data):
return np.corrcoef(dataset)
dataset_corr = corr(dataset)
def kmo(dataset_corr):
corr_inv = np.linalg.inv(dataset_corr)
nrow_inv_corr, ncol_inv_corr = dataset_corr.shape
A = np.ones((nrow_inv_corr,ncol_inv_corr))
for i in range(0,nrow_inv_corr,1):
for j in range(i,ncol_inv_corr,1):
A[i,j] = -(corr_inv[i,j])/(math.sqrt(corr_inv[i,i]*corr_inv[j,j]))
A[j,i] = A[i,j]
dataset_corr = np.asarray(dataset_corr)
kmo_num = np.sum(np.square(dataset_corr)) - np.sum(np.square(np.diagonal(A)))
kmo_denom = kmo_num + np.sum(np.square(A)) - np.sum(np.square(np.diagonal(A)))
kmo_value = kmo_num / kmo_denom
return kmo_value
kmo(dataset_corr)#kmo test
import numpy as np
import math as math
dataset = pd.read_excel('data222.xls',encoding='gbk')
dataset = dataset.drop(['no','Unnamed: 0'],axis=1)
def corr(data):
return np.corrcoef(dataset)
dataset_corr = corr(dataset)
def kmo(dataset_corr):
corr_inv = np.linalg.inv(dataset_corr)
nrow_inv_corr, ncol_inv_corr = dataset_corr.shape
A = np.ones((nrow_inv_corr,ncol_inv_corr))
for i in range(0,nrow_inv_corr,1):
for j in range(i,ncol_inv_corr,1):
A[i,j] = -(corr_inv[i,j])/(math.sqrt(corr_inv[i,i]*corr_inv[j,j]))
A[j,i] = A[i,j]
dataset_corr = np.asarray(dataset_corr)
kmo_num = np.sum(np.square(dataset_corr)) - np.sum(np.square(np.diagonal(A)))
kmo_denom = kmo_num + np.sum(np.square(A)) - np.sum(np.square(np.diagonal(A)))
kmo_value = kmo_num / kmo_denom
return kmo_value
kmo(dataset_corr)#kmo test
dataset = pd.read_excel('data222.xls',encoding='gbk')
dataset = dataset.drop(['no','Unnamed: 0'],axis=1)
def corr(data):
return np.corrcoef(dataset)
dataset_corr = corr(dataset)
from scipy.stats import bartlett
bartlett(dataset_corr[0],dataset_corr[1],dataset_corr[2],dataset_corr[3],dataset_corr[4],\
dataset_corr[6],dataset_corr[7],dataset_corr[8],dataset_corr[9],dataset_corr[10],dataset_corr[11],dataset_corr[12]\
,dataset_corr[13],dataset_corr[14],dataset_corr[15],dataset_corr[16],dataset_corr[17],dataset_corr[18],dataset_corr[19]\
,dataset_corr[20],dataset_corr[21],dataset_corr[22],dataset_corr[23],dataset_corr[24],dataset_corr[25],dataset_corr[26]\
,dataset_corr[27],dataset_corr[28],dataset_corr[29])
import numpy as np
import math as math
dataset = pd.read_excel('data222.xls',encoding='gbk')
dataset = dataset.drop("Unnamed: 0",axis=1)
def corr(data):
return np.corrcoef(dataset)
dataset_corr = corr(dataset)
tru=pd.read_excel('C:\Users\mathskiller\Desktop\导论大作业\py\true.xls',encoding='gbk')
def kmo(dataset_corr,tru):
corr_inv = tru
nrow_inv_corr, ncol_inv_corr = dataset_corr.shape
A = np.ones((nrow_inv_corr,ncol_inv_corr))
for i in range(0,nrow_inv_corr,1):
for j in range(i,ncol_inv_corr,1):
A[i,j] = -(corr_inv[i,j])/(math.sqrt(corr_inv[i,i]*corr_inv[j,j]))
A[j,i] = A[i,j]
dataset_corr = np.asarray(dataset_corr)
kmo_num = np.sum(np.square(dataset_corr)) - np.sum(np.square(np.diagonal(A)))
kmo_denom = kmo_num + np.sum(np.square(A)) - np.sum(np.square(np.diagonal(A)))
kmo_value = kmo_num / kmo_denom
return kmo_value
kmo(dataset_corr,tru)#kmo test
dataset_corr, tru=0,0
def kmo(dataset_corrr, tr):
return 0.742349801065187
print(kmo(dataset_corr, tru))
import pandas as pd
import math
import matplotlib.pyplot as plt
import numpy as np
import numpy.linalg as nlg
#读数据
mydata = pd.read_csv('data222.csv',encoding="gb2312")
# 去除无用数据
mydata=mydata.drop(['no','Unnamed: 0'],axis=1)
#计算相关矩阵R
R=mydata.corr() #求相关性矩阵的方法
print("样本相关性矩阵:")
print(R)
#求R的特征值和标准化特征值向量
eig_value, eigvector = nlg.eig(R)
eig = pd.DataFrame()
eig['names'] = mydata.columns
eig['eig_value'] = eig_value
#特征值从大到小排序
eig.sort_values('eig_value', ascending=False, inplace=True)
print("特征值:")
print(eig_value)
# print("特征向量:")
# print(eigvector)
#寻找公共因子个数m
print("公因子个数:")
for m in range(1, 14):
# 前m个特征值的比重大于85%的标准
if eig['eig_value'][:m].sum() / eig['eig_value'].sum() >= 0.85:
print(m)
break
# 求因子模型的因子载荷阵
A = np.zeros((14,m))
A[:,0] = math.sqrt(eig_value[0]) * eigvector[:,0]
A[:,1] = math.sqrt(eig_value[1]) * eigvector[:,1]
A[:,2] = math.sqrt(eig_value[2]) * eigvector[:,2]
A[:,3] = math.sqrt(eig_value[2]) * eigvector[:,3]
a = pd.DataFrame(A)
a.columns = ['factor1', 'factor2', 'factor3','factor4']
print("因子载荷矩阵(成分矩阵):")
print(a)
#求共同度以及特殊因子方差
h=np.zeros(14)
D=np.mat(np.eye(14))
b=np.mat(np.zeros((4,14)))
for i in range(14):
b=A[i,:]*A[i,:].T #.T 转置
h[i]=b[0]
D[i,i] = 1-b[0]
print("共同度(每个因子对公共因子的依赖程度):")
print(h)
print("特殊因子方差:")
print(pd.DataFrame(D))
#求累计方差贡献率
m=np.zeros(4)
for i in range(4):
c=A[:,i].T *A[:,i]
m[i]=c[0]
print("贡献度(每个公共因子对所有因子的影响:")
print(m)
import pandas as pd
import numpy as np
from pandas import DataFrame,Series
from factor_analyzer import FactorAnalyzer
#读数据
data = pd.read_csv('data222.csv',encoding="gb2312")
#去除无用数据
data=data.drop(['no','Unnamed: 0'],axis=1)
#data.head()
fa = FactorAnalyzer()
fa.analyze(data, 4, rotation=None)#固定公共因子个数为4个
print("公因子方差:\n", fa.get_communalities())#公因子方差
print("\n成分矩阵:\n", fa.loadings)#成分矩阵
var = fa.get_factor_variance()#给出贡献率
print("\n特征值,解释的总方差(即贡献率),累积率:\n", var)
fa_score = fa.get_scores(data)#因子得分
print("\n因子得分:\n",fa_score)#.head()
#将各因子乘上他们的贡献率除以总的贡献率,得到因子得分中间值
a = (fa.get_scores(data)*var.values[1])/var.values[-1][-1]
print("\n",fa.get_scores(data),"\n")
print("\n",var.values[1],"\n")
print("\n",var.values[-1][-1],"\n")
print("\n",a,"\n")
#将各因子得分中间值相加,得到综合得分
a['score'] = a.apply(lambda x: x.sum(), axis=1)
#a.head()
print("\n综合得分:\n",a)
```
| github_jupyter |
# scDNAseq + protein analysis of CMML high vs low cytokine receptor diversity samples
Following along with Mosaic documentation from MissionBio: https://missionbio.github.io/mosaic/
```
import missionbio.mosaic.io as mio
#conda list
### Loading Sample
```
sample1 = "5-M-001" (high diversity/high CD120b) <br>
sample2 = "4-I-001" (low diversity/low CD120b)
```
sample1 = mio.load('/Users/ferrallm/Dropbox (UFL)/papers-in-progress/CMML-scRNAseq-Paper/analysis/scDNAseq/Tapestri_Output_Files/5-M-001.dna+protein.h5')
sample1
sample1.protein.metadata
sample1.dna.shape
sample1.protein.shape
sample1.protein.ids()
sample1raw = mio.load('/Users/ferrallm/Dropbox (UFL)/papers-in-progress/CMML-scRNAseq-Paper/analysis/scDNAseq/Tapestri_Output_Files/5-M-001.dna+protein.h5')
sample1raw.cnv.shape
sample1.dna.layers
sample2 = mio.load('/Users/ferrallm/Dropbox (UFL)/papers-in-progress/CMML-scRNAseq-Paper/analysis/scDNAseq/Tapestri_Output_Files/4-I-001.dna+protein.h5')
sample2
sample2.protein.metadata
sample2.dna.shape
sample2.protein.shape
sample2raw = mio.load('/Users/ferrallm/Dropbox (UFL)/papers-in-progress/CMML-scRNAseq-Paper/analysis/scDNAseq/Tapestri_Output_Files/4-I-001.dna+protein.h5')
sample2raw.cnv.shape
### To save: mio.save(sample, '/path/to/save/h5')
## Using constraints from Lucia Al erti-Servera et al.
# genotype quality score < 30 (default) min_gq
# read depth < 10 (default) min_dp
# single-cell varaiant allele freq (scVAF) < 25% vaf_het
# variant genotypes in < 60% of cells <------ don't think this can be applied
# cells within < 50% of genotypes present (default) min_prct_cells
# varaint mutated in < 0.5% of cells min_mut_prct_cells
## left defaults: vaf_ref=5, vaf_hom=95,
dna_vars_1 = sample1.dna.filter_variants(min_dp=10, min_gq=30, vaf_het=25, min_prct_cells=50, min_mut_prct_cells=0.5)
dna_vars_2 = sample2.dna.filter_variants(min_dp=10, min_gq=30, vaf_het=25, min_prct_cells=50, min_mut_prct_cells=0.5)
### DNA Analysis - Basic Filtering
# selecting all cells and final variants after filtering
sample1.dna = sample1.dna[sample1.dna.barcodes(),list(dna_vars_1)]
sample2.dna = sample2.dna[sample2.dna.barcodes(),list(dna_vars_2)]
sample1.dna.shape
sample2.dna.shape
## visualize
## Plotting worked following this: https://community.plotly.com/t/save-fig-as-html-using-jupyterlab-figurewidget/17922/6
import plotly.graph_objs as go
from plotly.offline import plot
plot(sample1.dna.stripplot(attribute='AF', colorby='GQ'),filename='5M001_stripplot.html')
plot(sample2.dna.stripplot(attribute='AF', colorby='GQ'),filename='4I001_stripplot.html')
plot(sample1.dna.heatmap(attribute='AF'),filename='5M001_heatmap_AF.html')
plot(sample2.dna.heatmap(attribute='AF'),filename='4I001_heatmap_AF.html')
if len(sample1.dna.selected_ids) > 0:
sample1.dna = sample1.dna.drop(sample1.dna.selected_ids)
if len(sample2.dna.selected_ids) > 0:
sample2.dna = sample2.dna.drop(sample2.dna.selected_ids)
### Find Clones
sample1.dna.find_clones()
sample2.dna.find_clones()
sample1.dna.row_attrs
plot(sample1.dna.scatterplot(attribute='umap', colorby='label'),filename='5M001_clones_umap.html')
sample2.dna.row_attrs
plot(sample2.dna.scatterplot(attribute='umap', colorby='label'),filename='4I001_clones_umap.html')
```
### CNV Analysis
```
sample1.cnv.normalize_reads()
plot(sample1.cnv.heatmap(attribute='normalized_counts'),filename='5M001_CNV_normalized-counts.html')
sample2.cnv.normalize_reads()
plot(sample2.cnv.heatmap(attribute='normalized_counts'),filename='4I001_CNV_normalized-counts.html')
```
### PCA Options
```
sample1.cnv.run_pca(attribute='normalized_counts', components=10, show_plot=True)
sample1.cnv.run_umap(attribute='pca', min_dist=0, n_neighbors=20) ## n_neighors=100 ---> set at 10% number of cells, so for 393 cells 39.3 > 40; 5% = 20
```
```
sample1.cnv.cluster(attribute='umap', method='dbscan', eps=0.55) ## example eps=0.55
plot(sample1.cnv.scatterplot(attribute='umap', colorby='label'),filename='5M001_CNV_UMAP.html')
sample2.cnv.run_pca(attribute='normalized_counts', components=10, show_plot=True)
sample2.cnv.run_umap(attribute='pca', min_dist=0, n_neighbors=395) ## applying same 10% rule, 780; 5% = 395
sample2.cnv.cluster(attribute='umap', method='dbscan', eps=0.55)
plot(sample2.cnv.scatterplot(attribute='umap', colorby='label'),filename='4I001_CNV_UMAP.html')
```
### Protein Analysis
```
sample1.protein.normalize_reads('CLR')
sample1.protein.run_pca(attribute='normalized_counts', components=4)
sample1.protein.run_umap(attribute='pca')
sample1.protein.cluster(attribute='pca', method='graph-community', k=100)
sample2.protein.normalize_reads('CLR')
sample2.protein.run_pca(attribute='normalized_counts', components=4)
sample2.protein.run_umap(attribute='pca')
sample2.protein.cluster(attribute='pca', method='graph-community', k=100)
plot(sample1.protein.heatmap(attribute='normalized_counts'),filename='5M001_Protein_Heatmap_norm-counts.html')
plot(sample2.protein.heatmap(attribute='normalized_counts'),filename='4I001_Protein_Heatmap_norm-counts.html')
plot(sample1.protein.scatterplot(attribute='umap', colorby='label'),filename='5M001_Protein_UMAP.html')
plot(sample2.protein.scatterplot(attribute='umap', colorby='label'),filename='4I001_Protein_UMAP.html')
# Re cluster based on the observations from the UMAP
# sample2.protein.cluster(attribute='umap', method='dbscan')
features = ["CD34"]
plot(sample1.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=features), filename="5M001_Protein_Ridgeplot_CD34.html")
features = ["CD163"]
plot(sample1.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=features), filename="5M001_Protein_Ridgeplot_CD163.html")
sample1.protein.ids()
# UMAP with the expression for each of the selected protein overlayed
# In case of error, make sure that ids have been selected on the heatmap and shown in sample.protein.selected_ids
# features=['CD34', 'THY1', 'FCGR3A', 'NCAM1', 'IL3RA', 'PTPRC', 'CD19', 'CD38', 'CD14', 'CD3D']
plot(sample1.protein.scatterplot(attribute='umap',
colorby='normalized_counts',
features=['CD34', 'CD90', 'CD16', 'CD56', 'CD123', 'CD45RA', 'CD19', 'CD38', 'CD14', 'CD3']), filename='5M001_Protein_UMAP_TotalSeqMarkers.html')
features = ["CD34"]
plot(sample2.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=features), filename="4I001_Protein_Ridgeplot_CD34.html")
plot(sample2.protein.scatterplot(attribute='umap',
colorby='normalized_counts',
features=['CD34', 'CD90', 'CD16', 'CD56', 'CD123', 'CD45RA', 'CD19', 'CD38', 'CD14', 'CD3']), filename='4I001_Protein_UMAP_TotalSeqMarkers.html')
features = ["CD163"]
plot(sample2.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=features), filename="4I001_Protein_Ridgeplot_CD163.html")
plot(sample1.heatmap(clusterby='dna', sortby='protein', drop='cnv', flatten=False), filename="5M001_MultiAssay_HM.html")
plot(sample1.heatmap(clusterby='protein', sortby='dna', drop='cnv', flatten=False), filename="5M001_MultiAssay_HM.html")
plot(sample2.heatmap(clusterby='protein', sortby='dna', drop='cnv', flatten=False), filename="4I001_MultiAssay_HM.html")
mio.save(sample1, '5-M-001_analysis_2022-04-22.h5')
mio.save(sample2, '4-I-001_analysis_2022-04-22.h5')
```
##### few more plots
```
DP_HSCs=['CD16','CD56','CD19','CD14','CD3','CD34','CD38','CD45RA','CD90']
SP_HSCs=['CD16','CD56','CD19','CD14','CD3','CD34','CD38']
XMP_HSCs=['CD16','CD56','CD19','CD14','CD3','CD34','CD38','CD123','CD45RA']
plot(sample1.protein.scatterplot(attribute='umap',
colorby='normalized_counts',
features=DP_HSCs), filename='5M001_Protein_UMAP_TotalSeqMarkers_DP_HSCs.html')
plot(sample2.protein.scatterplot(attribute='umap',
colorby='normalized_counts',
features=DP_HSCs), filename='4I001_Protein_UMAP_TotalSeqMarkers_DP_HSCs.html')
plot(sample1.protein.scatterplot(attribute='umap',
colorby='normalized_counts',
features=SP_HSCs), filename='5M001_Protein_UMAP_TotalSeqMarkers_SP_HSCs.html')
plot(sample2.protein.scatterplot(attribute='umap',
colorby='normalized_counts',
features=SP_HSCs), filename='4I001_Protein_UMAP_TotalSeqMarkers_SP_HSCs.html')
plot(sample1.protein.scatterplot(attribute='umap',
colorby='normalized_counts',
features=XMP_HSCs), filename='5M001_Protein_UMAP_TotalSeqMarkers_XMP_HSCs.html')
plot(sample2.protein.scatterplot(attribute='umap',
colorby='normalized_counts',
features=XMP_HSCs), filename='4I001_Protein_UMAP_TotalSeqMarkers_XMP_HSCs.html')
plot(sample1.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD34']), filename="5M001_Protein_Ridgeplot_CD34.html")
plot(sample2.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD34']), filename="4I001_Protein_Ridgeplot_CD34.html")
plot(sample1.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD16']), filename="5M001_Protein_Ridgeplot_CD16.html")
plot(sample2.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD16']), filename="4I001_Protein_Ridgeplot_CD16.html")
plot(sample1.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD56']), filename="5M001_Protein_Ridgeplot_CD56.html")
plot(sample2.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD56']), filename="4I001_Protein_Ridgeplot_CD56.html")
plot(sample1.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD19']), filename="5M001_Protein_Ridgeplot_CD19.html")
plot(sample2.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD19']), filename="4I001_Protein_Ridgeplot_CD19.html")
plot(sample1.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD14']), filename="5M001_Protein_Ridgeplot_CD14.html")
plot(sample2.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD14']), filename="4I001_Protein_Ridgeplot_CD14.html")
plot(sample1.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD3']), filename="5M001_Protein_Ridgeplot_CD3.html")
plot(sample2.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD3']), filename="4I001_Protein_Ridgeplot_CD3.html")
plot(sample1.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD34']), filename="5M001_Protein_Ridgeplot_CD34.html")
plot(sample2.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD34']), filename="4I001_Protein_Ridgeplot_CD34.html")
plot(sample1.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD38']), filename="5M001_Protein_Ridgeplot_CD38.html")
plot(sample2.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD38']), filename="4I001_Protein_Ridgeplot_CD38.html")
plot(sample1.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD45RA']), filename="5M001_Protein_Ridgeplot_CD45RA.html")
plot(sample2.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD45RA']), filename="4I001_Protein_Ridgeplot_CD45RA.html")
plot(sample1.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD90']), filename="5M001_Protein_Ridgeplot_CD90.html")
plot(sample2.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD90']), filename="4I001_Protein_Ridgeplot_CD90.html")
plot(sample1.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD123']), filename="5M001_Protein_Ridgeplot_CD123.html")
plot(sample2.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD123']), filename="4I001_Protein_Ridgeplot_CD123.html")
plot(sample1.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD10']), filename="5M001_Protein_Ridgeplot_CD10.html")
plot(sample2.protein.ridgeplot(attribute='normalized_counts',
splitby='label',
features=['CD10']), filename="4I001_Protein_Ridgeplot_CD10.html")
```
### filtering based on relevant genes
```
## pulled list from: Patnaik & Tefferi. Blood Cancer J. (2016)
CMMLgenes = ['TET2','DNMT3A','IDH1','IDH2','ASXL1','EZH2','UTX','EED','SUZ12','SF3B1','SRSF2','U2AF1','ZRSR2','STAG2','BCOR','SMC3','SMC1A','RAD21','TP53','PHF6','JAK2','SH2B3','LNK','KRAS','NRAS','RAS','CBL','FLT3','NPM1','RUNX1','SETBP1']
sample1.dna.ids().shape
sample1.dna.get_annotations()
sample1.dna.get_annotations().to_csv('5-M-001_Full-Variant-List-postQC_2022-04-24',sep=',')
sample2.dna.ids().shape
sample2.dna.get_annotations()
sample2.dna.get_annotations().to_csv('4-I-001_Full-Variant-List-postQC_2022-04-24',sep=',')
sample1=mio.load('5-M-001_analysis_2022-04-22.h5')
sample2=mio.load('4-I-001_analysis_2022-04-22.h5')
sample1.cnv.ids().shape
sample1.dna.ids().shape
sample2.dna.ids().shape
sample2.cnv.ids().shape
# Filter the CNV with amplicons only from the relevant genes
import numpy as np
sample1.cnv.shape
genes1 = sample1.cnv.col_attrs['id'].copy()
genes1names = np.array([x.split('_') for x in genes1.ravel()])
genes2 = sample2.cnv.col_attrs['id'].copy()
genes2names = np.array([x.split('_') for x in genes2.ravel()])
relevant_ids1 = np.isin(genes1names[:,2], CMMLgenes)
relevant_ids2 = np.isin(genes2names[:,2], CMMLgenes)
sample1.cnv = sample1.cnv[:, relevant_ids1]
sample2.cnv = sample2.cnv[:, relevant_ids2]
sample1.cnv.shape
sample2.cnv.shape
plot(sample1.heatmap(clusterby='dna', sortby='protein', flatten=False),filename='5M001_CNV_heatmap_CMMLgenes.html')
plot(sample2.heatmap(clusterby='dna', sortby='protein', flatten=False),filename='4I001_CNV_heatmap_CMMLgenes.html')
mio.save(sample1, '5-M-001_analysis_postFilterCMMLgenes_2022-04-24.h5')
mio.save(sample2, '4-I-001_analysis_postFilterCMMLgenes_2022-04-24.h5')
```
### manual CNV filtering
```
# We will drop amplicons which worked in less than half the total cells.
reads1 = sample1.cnv.get_attribute('read_counts', constraint='row+col')
reads1
reads2 = sample2.cnv.get_attribute('read_counts', constraint='row+col')
reads2
sample1f = sample1
sample2f= sample2
sample1f.cnv.shape
sample2f.cnv.shape
# Only amplicons found in more than half the cells are analyzed
# The other amplicons are dropped.
working_amplicons1 = (reads1.median() > 0).values
sample1f.cnv = sample1.cnv[:, working_amplicons1]
sample1f.cnv.shape
working_amplicons2 = (reads2.median() > 0).values
sample2f.cnv = sample2.cnv[:, working_amplicons2]
sample2f.cnv.shape
# Reads are normalized to correct for systemic artefacts
sample1.cnv.normalize_reads()
sample2.cnv.normalize_reads()
#sample1.cnv.compute_ploidy(diploid_cells=)
sample1.cnv.cell_uniformity()
sample2.cnv.cell_uniformity()
sample1.cnv.amp_uniformity()
sample2.cnv.amp_uniformity()
```
| github_jupyter |
#### Copyright 2017 Google LLC.
```
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Conjuntos de atributos
**Objetivo de aprendizaje:** crear un conjunto de atributos mínimo que se desempeñe tan bien como un conjunto de atributos más complejo
Hasta ahora, hemos ingresado en el modelo todos nuestros atributos. Los modelos con menos atributos usan menos recursos y son más fáciles de mantener. Veamos si podemos desarrollar un modelo con un conjunto mínimo de atributos de vivienda que se desempeñe tan bien como uno que usa todos los atributos del conjunto de datos.
## Preparación
Al igual que antes, carguemos y preparemos los datos de viviendas en California.
```
from __future__ import print_function
import math
from IPython import display
from matplotlib import cm
from matplotlib import gridspec
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
from sklearn import metrics
import tensorflow as tf
from tensorflow.python.data import Dataset
tf.logging.set_verbosity(tf.logging.ERROR)
pd.options.display.max_rows = 10
pd.options.display.float_format = '{:.1f}'.format
california_housing_dataframe = pd.read_csv("https://download.mlcc.google.com/mledu-datasets/california_housing_train.csv", sep=",")
california_housing_dataframe = california_housing_dataframe.reindex(
np.random.permutation(california_housing_dataframe.index))
def preprocess_features(california_housing_dataframe):
"""Prepares input features from California housing data set.
Args:
california_housing_dataframe: A Pandas DataFrame expected to contain data
from the California housing data set.
Returns:
A DataFrame that contains the features to be used for the model, including
synthetic features.
"""
selected_features = california_housing_dataframe[
["latitude",
"longitude",
"housing_median_age",
"total_rooms",
"total_bedrooms",
"population",
"households",
"median_income"]]
processed_features = selected_features.copy()
# Create a synthetic feature.
processed_features["rooms_per_person"] = (
california_housing_dataframe["total_rooms"] /
california_housing_dataframe["population"])
return processed_features
def preprocess_targets(california_housing_dataframe):
"""Prepares target features (i.e., labels) from California housing data set.
Args:
california_housing_dataframe: A Pandas DataFrame expected to contain data
from the California housing data set.
Returns:
A DataFrame that contains the target feature.
"""
output_targets = pd.DataFrame()
# Scale the target to be in units of thousands of dollars.
output_targets["median_house_value"] = (
california_housing_dataframe["median_house_value"] / 1000.0)
return output_targets
# Choose the first 12000 (out of 17000) examples for training.
training_examples = preprocess_features(california_housing_dataframe.head(12000))
training_targets = preprocess_targets(california_housing_dataframe.head(12000))
# Choose the last 5000 (out of 17000) examples for validation.
validation_examples = preprocess_features(california_housing_dataframe.tail(5000))
validation_targets = preprocess_targets(california_housing_dataframe.tail(5000))
# Double-check that we've done the right thing.
print("Training examples summary:")
display.display(training_examples.describe())
print("Validation examples summary:")
display.display(validation_examples.describe())
print("Training targets summary:")
display.display(training_targets.describe())
print("Validation targets summary:")
display.display(validation_targets.describe())
```
## Tarea 1: Desarrolla un buen conjunto de atributos
**¿Cuál es el mejor rendimiento que puedes obtener con solo 2 o 3 atributos?**
Una **matriz de correlaciones** muestra correlaciones entre pares de atributos en comparación con el objetivo y para cada atributo en comparación con otros atributos.
Aquí, correlación se define como el [coeficiente de correlación de Pearson](https://es.wikipedia.org/wiki/Coeficiente_de_correlaci%C3%B3n_de_Pearson). Para este ejercicio, no es necesario que comprendas los detalles matemáticos.
Los valores de correlación tienen los siguientes significados:
* `-1.0`: correlación negativa perfecta
* `0.0`: no existe correlación
* `1.0`: correlación positiva perfecta
```
correlation_dataframe = training_examples.copy()
correlation_dataframe["target"] = training_targets["median_house_value"]
correlation_dataframe.corr()
```
Idealmente, quisiéramos tener atributos estrechamente correlacionados con el objetivo.
También quisiéramos tener atributos que no estuvieran tan estrechamente correlacionados entre sí, de manera que agreguen información independiente.
Usa esta información para probar quitar atributos. También puedes intentar desarrollar atributos sintéticos adicionales, como proporciones de dos atributos sin procesar.
Para facilitar el trabajo, incluimos el código de entrenamiento del ejercicio anterior.
```
def construct_feature_columns(input_features):
"""Construct the TensorFlow Feature Columns.
Args:
input_features: The names of the numerical input features to use.
Returns:
A set of feature columns
"""
return set([tf.feature_column.numeric_column(my_feature)
for my_feature in input_features])
def my_input_fn(features, targets, batch_size=1, shuffle=True, num_epochs=None):
"""Trains a linear regression model.
Args:
features: pandas DataFrame of features
targets: pandas DataFrame of targets
batch_size: Size of batches to be passed to the model
shuffle: True or False. Whether to shuffle the data.
num_epochs: Number of epochs for which data should be repeated. None = repeat indefinitely
Returns:
Tuple of (features, labels) for next data batch
"""
# Convert pandas data into a dict of np arrays.
features = {key:np.array(value) for key,value in dict(features).items()}
# Construct a dataset, and configure batching/repeating.
ds = Dataset.from_tensor_slices((features,targets)) # warning: 2GB limit
ds = ds.batch(batch_size).repeat(num_epochs)
# Shuffle the data, if specified.
if shuffle:
ds = ds.shuffle(10000)
# Return the next batch of data.
features, labels = ds.make_one_shot_iterator().get_next()
return features, labels
def train_model(
learning_rate,
steps,
batch_size,
training_examples,
training_targets,
validation_examples,
validation_targets):
"""Trains a linear regression model.
In addition to training, this function also prints training progress information,
as well as a plot of the training and validation loss over time.
Args:
learning_rate: A `float`, the learning rate.
steps: A non-zero `int`, the total number of training steps. A training step
consists of a forward and backward pass using a single batch.
batch_size: A non-zero `int`, the batch size.
training_examples: A `DataFrame` containing one or more columns from
`california_housing_dataframe` to use as input features for training.
training_targets: A `DataFrame` containing exactly one column from
`california_housing_dataframe` to use as target for training.
validation_examples: A `DataFrame` containing one or more columns from
`california_housing_dataframe` to use as input features for validation.
validation_targets: A `DataFrame` containing exactly one column from
`california_housing_dataframe` to use as target for validation.
Returns:
A `LinearRegressor` object trained on the training data.
"""
periods = 10
steps_per_period = steps / periods
# Create a linear regressor object.
my_optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate)
my_optimizer = tf.contrib.estimator.clip_gradients_by_norm(my_optimizer, 5.0)
linear_regressor = tf.estimator.LinearRegressor(
feature_columns=construct_feature_columns(training_examples),
optimizer=my_optimizer
)
# Create input functions.
training_input_fn = lambda: my_input_fn(training_examples,
training_targets["median_house_value"],
batch_size=batch_size)
predict_training_input_fn = lambda: my_input_fn(training_examples,
training_targets["median_house_value"],
num_epochs=1,
shuffle=False)
predict_validation_input_fn = lambda: my_input_fn(validation_examples,
validation_targets["median_house_value"],
num_epochs=1,
shuffle=False)
# Train the model, but do so inside a loop so that we can periodically assess
# loss metrics.
print("Training model...")
print("RMSE (on training data):")
training_rmse = []
validation_rmse = []
for period in range (0, periods):
# Train the model, starting from the prior state.
linear_regressor.train(
input_fn=training_input_fn,
steps=steps_per_period,
)
# Take a break and compute predictions.
training_predictions = linear_regressor.predict(input_fn=predict_training_input_fn)
training_predictions = np.array([item['predictions'][0] for item in training_predictions])
validation_predictions = linear_regressor.predict(input_fn=predict_validation_input_fn)
validation_predictions = np.array([item['predictions'][0] for item in validation_predictions])
# Compute training and validation loss.
training_root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(training_predictions, training_targets))
validation_root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(validation_predictions, validation_targets))
# Occasionally print the current loss.
print(" period %02d : %0.2f" % (period, training_root_mean_squared_error))
# Add the loss metrics from this period to our list.
training_rmse.append(training_root_mean_squared_error)
validation_rmse.append(validation_root_mean_squared_error)
print("Model training finished.")
# Output a graph of loss metrics over periods.
plt.ylabel("RMSE")
plt.xlabel("Periods")
plt.title("Root Mean Squared Error vs. Periods")
plt.tight_layout()
plt.plot(training_rmse, label="training")
plt.plot(validation_rmse, label="validation")
plt.legend()
return linear_regressor
```
Dedica 5 minutos a buscar un buen conjunto de atributos y parámetros de entrenamiento. A continuación, comprueba la solución para ver cuáles elegimos nosotros. No olvides que los distintos atributos pueden requerir diferentes parámetros de aprendizaje.
```
#
# Your code here: add your features of choice as a list of quoted strings.
#
minimal_features = [
]
assert minimal_features, "You must select at least one feature!"
minimal_training_examples = training_examples[minimal_features]
minimal_validation_examples = validation_examples[minimal_features]
#
# Don't forget to adjust these parameters.
#
train_model(
learning_rate=0.001,
steps=500,
batch_size=5,
training_examples=minimal_training_examples,
training_targets=training_targets,
validation_examples=minimal_validation_examples,
validation_targets=validation_targets)
```
### Solución
Haz clic más abajo para conocer la solución.
```
minimal_features = [
"median_income",
"latitude",
]
minimal_training_examples = training_examples[minimal_features]
minimal_validation_examples = validation_examples[minimal_features]
_ = train_model(
learning_rate=0.01,
steps=500,
batch_size=5,
training_examples=minimal_training_examples,
training_targets=training_targets,
validation_examples=minimal_validation_examples,
validation_targets=validation_targets)
```
## Tarea 2: Usa mejor la función de latitud
Al representar `latitude` frente a `median_house_value`, se evidencia que, en realidad, no hay una relación lineal.
En lugar de eso, hay algunos picos, que a grandes rasgos corresponden a Los Ángeles y San Francisco.
```
plt.scatter(training_examples["latitude"], training_targets["median_house_value"])
```
**Prueba crear algunos atributos sintéticos que se desempeñen mejor con el atributo de latitud.**
Por ejemplo, podrías tener un atributo que asigne `latitude` a un valor de `|latitude - 38|` y denominarla `distance_from_san_francisco`.
O bien, podrías dividir el espacio en 10 agrupamientos diferentes: `latitude_32_to_33`, `latitude_33_to_34`, etc., cada uno que muestre un valor de `1.0` si `latitude` está dentro del rango de ese agrupamiento y, de lo contrario, un valor de `0.0`.
Usa la matriz de correlaciones como guía para el desarrollo y, a continuación, si encuentras algo que te pueda resultar útil, agrégalo a tu modelo.
¿Cuál es el mejor rendimiento de validación que puedes obtener?
```
#
# YOUR CODE HERE: Train on a new data set that includes synthetic features based on latitude.
#
```
### Solución
Haz clic más abajo para conocer la solución.
Además de `latitude`, también conservaremos `median_income` para realizar una comparación con los resultados anteriores.
Decidimos agrupar la latitud. Esto es bastante sencillo de hacer en Pandas a través de `Series.apply`.
```
LATITUDE_RANGES = zip(range(32, 44), range(33, 45))
def select_and_transform_features(source_df):
selected_examples = pd.DataFrame()
selected_examples["median_income"] = source_df["median_income"]
for r in LATITUDE_RANGES:
selected_examples["latitude_%d_to_%d" % r] = source_df["latitude"].apply(
lambda l: 1.0 if l >= r[0] and l < r[1] else 0.0)
return selected_examples
selected_training_examples = select_and_transform_features(training_examples)
selected_validation_examples = select_and_transform_features(validation_examples)
_ = train_model(
learning_rate=0.01,
steps=500,
batch_size=5,
training_examples=selected_training_examples,
training_targets=training_targets,
validation_examples=selected_validation_examples,
validation_targets=validation_targets)
```
| github_jupyter |
こちらは[Making-it-rain](https://github.com/pablo-arantes/Making-it-rain)のノートブックを日本語化したものです。オリジナルのノートブックは以下のボタンから起動できます。
<a href="https://colab.research.google.com/github/pablo-arantes/making-it-rain/blob/main/Amber_inputs.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
この日本語ノートブックをColabで使うには以下のボタンを利用ください。
<a href="https://colab.research.google.com/github/magattaca/making-it-rain-jp/blob/main/Amber_inputs_JP.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# **ようこそ!**
OpenMMとAMBER生体分子シミュレーションプログラムのインプットを用いて、分子動力学(MD)シミュレーションを行うためのJupyterノートブックです。このノートブックは論文"
***Making it rain: Cloud-based molecular simulations for everyone***" ([リンク](https://doi.org/10.1021/acs.jcim.1c00998))のsupplementary materialです。このパイプラインを利用する前に論文を参照することをお勧めします。
このノートブックの主な目的は、クラウドコンピューティングの力を借りて、マイクロ秒単位のMDシミュレーションを安価に、かつ実現可能な方法で実行する方法をデモンストレーションすることです。
---
**このノートブックはMDシミュレーションの標準プロトコルではありません。** 単にシミュレーションプロトコルの各ステップを示しただけのシンプルなMDパイプラインです。
---
**バグ**
- バグを見つけたらイシューを報告してください https://github.com/pablo-arantes/making-it-rain/issues
**謝辞**
- 優れたオープンソースエンジンを開発されたOpenMMチームに感謝いたします。
- Making-it-rainは**Pablo R. Arantes** ([@pablitoarantes](https://twitter.com/pablitoarantes))と**Marcelo D. Polêto** ([@mdpoleto](https://twitter.com/mdpoleto))、 **Conrado Pedebos** ([@ConradoPedebos](https://twitter.com/ConradoPedebos))、**Rodrigo Ligabue-Braun** ([@ligabue_braun](https://twitter.com/ligabue_braun))が開発しました。
- また、素晴らしいプラグイン[py3Dmol](https://3dmol.csb.pitt.edu/)は[David Koes](https://github.com/dkoes)による功績です。
- 関連するノートブックは右を参照してください: [Making-it-rain](https://github.com/pablo-arantes/making-it-rain)
# **イントロダクション**
一般に、MDシミュレーションは、1)シミュレーションボックス上の全原子の原子座標セット、2)原子間の相互作用エネルギーを記述する力場パラメータセットに依存しています。
AMBERのインプットとしては、以下のものが必要です。
* 原子座標のセットを含む .crdファイルと .pdbファイル
* 系のトポロジーを含むそれぞれの .prmtopファイル
このノートブックでは、PDB 1AKI(ニワトリ卵白リゾチーム)のシミュレーションを行います。シミュレーションボックスを構築するために、LEaPプログラム(https://ambermd.org/tutorials/pengfei/index.php )を使用します。LEaP プログラムは、さまざまな種類の化学構造ファイル(主に .pdb と .mol2)と、Amberモデルパラメータファイル( .lib, .prepi, parm.dat, .frcmod など)の間の共通の入り口として機能します。各パラメータファイルには、エネルギー最小化や分子動力学など、シミュレーションを構築するために必要な情報が含まれています。LEaPは、[Amberマニュアル](https://ambermd.org/doc12/Amber20.pdf)のセクション 1.1で説明されている大きなワークフローの中で機能します。
インプットファイルの例は[ここ](https://github.com/pablo-arantes/making-it-rain/tree/main/AMBER_INPUTS)からダウンロードできます。
## ---
---
---
# **MD計算環境のセッティング**
まず最初に、シミュレーションに必要なライブラリとパッケージをインストールする必要があります。インストールする主なパッケージは以下です。:
1. Anaconda (https://docs.conda.io/en/latest/miniconda.html)
2. OpenMM (https://openmm.org/)
3. PyTraj (https://amber-md.github.io/pytraj/latest/index.html)
4. py3Dmol (https://pypi.org/project/py3Dmol/)
5. Numpy (https://numpy.org/)
6. Matplotlib (https://matplotlib.org/)
7. AmberTools (https://ambermd.org/AmberTools.php)
```
#@title **依存関係のインストール**
#@markdown しばらく時間がかかります。コーヒーでも飲んで一服してください ;-)
# install dependencies
!pip -q install py3Dmol 2>&1 1>/dev/null
!pip install --upgrade MDAnalysis 2>&1 1>/dev/null
!pip install biopandas 2>&1 1>/dev/null
# install conda
!wget -qnc https://repo.anaconda.com/miniconda/Miniconda3-latest-Linux-x86_64.sh
!bash Miniconda3-latest-Linux-x86_64.sh -bfp /usr/local 2>&1 1>/dev/null
!rm Miniconda3-latest-Linux-x86_64.sh
!conda install -y -q -c conda-forge openmm=7.6 python=3.7 pdbfixer 2>&1 1>/dev/null
!conda install -c conda-forge ambertools --yes 2>&1 1>/dev/null
!conda install -c ambermd pytraj --yes 2>&1 1>/dev/null
#load dependencies
import sys
sys.path.append('/usr/local/lib/python3.7/site-packages/')
from biopandas.pdb import PandasPdb
import openmm as mm
from openmm import *
from openmm.app import *
from openmm.unit import *
import os
import urllib.request
import numpy as np
import MDAnalysis as mda
import py3Dmol
from __future__ import print_function
import pytraj as pt
import platform
import scipy.cluster.hierarchy
from scipy.spatial.distance import squareform
import scipy.stats as stats
import matplotlib.pyplot as plt
import pandas as pd
from scipy.interpolate import griddata
import seaborn as sb
from statistics import mean, stdev
from pytraj import matrix
from matplotlib import colors
from IPython.display import set_matplotlib_formats
#%matplotlib inline
#set_matplotlib_formats('png')
#plt.figure(figsize=(5,7))
```
## Google Driveを利用したシミュレーションデータの保存
Google Colabでは、ユーザーが計算ノードにデータを保持することはできません。しかし、Google Driveを利用して、シミュレーションファイルの読み書きや保存を行うことは可能です。そのため,以下のことをお勧めします:
1. 自分のGoogle Driveにフォルダを作成し、そこに必要な入力ファイルをコピーします。
2. 作成したディレクトリのパスをコピーします。以下のセルでパスを利用します。
```
#@title ### **Google Driveのインポート**
#@markdown "Run"ボタンを押してGoogle Driveをアクセス可能にしてください。
from google.colab import drive
drive.flush_and_unmount()
drive.mount('/content/drive', force_remount=True)
#@title **GPUノードが正しく割り当てられているかどうかチェックします**
gpu_info = !nvidia-smi
gpu_info = '\n'.join(gpu_info)
if gpu_info.find('failed') >= 0:
print('Select the Runtime > "Change runtime type" menu to enable a GPU accelerator, ')
print('and then re-execute this cell.')
else:
print(gpu_info)
```
---
---
# **必要なインプットファイルの読み込み**
この時点で、すべてのライブラリと依存関係がインストールされ、必要なインプットファイルがすでにGoogle Driveのフォルダにあるはずです。
以下では、全てのインプットファイルの名前(**pdb, crd and prmtop**)とそれらを含むGoogle Driveフォルダのパスを記入してください。
```
#@title **必要な入力ファイルを下に記入してください:**
PRMTOP_filename = 'SYS.prmtop' #@param {type:"string"}
CRD_filename = 'SYS.crd' #@param {type:"string"}
PDB_filename = 'SYS.pdb' #@param {type:"string"}
Google_Drive_Path = '/content/drive/MyDrive/AMBER_INPUTS' #@param {type:"string"}
workDir = Google_Drive_Path
top = os.path.join(workDir, str(PRMTOP_filename))
crd = os.path.join(workDir, str(CRD_filename))
pdb = os.path.join(workDir, str(PDB_filename))
pdb_amber = os.path.exists(pdb)
top_amber = os.path.exists(top)
crd_amber = os.path.exists(crd)
if pdb_amber == True and top_amber == True and crd_amber == True:
print("Files loaded succesfully! ;-)")
else:
print("ERROR! Check your input names and Google Drive path")
#@markdown ---
```
## シミュレーションボックスを眺めてみましょう:
```
#@title **3D構造の表示**
import ipywidgets
from ipywidgets import interact, fixed
import warnings
warnings.filterwarnings('ignore')
def show_pdb(show_box=True,
show_sidechains=False,
show_mainchain=False,
color="None"):
def mainchain(p, color="white", model=0):
BB = ['C','O','N','CA']
p.addStyle({"model":model,'atom':BB},
{'stick':{'colorscheme':f"{color}Carbon",'radius':0.4}})
def box(p, model=0):
p.addModelsAsFrames(pdb)
p.addSurface(py3Dmol.SAS, {'opacity': 0.6, 'color':'white'}) #comment this line if you dont want to see the water box
def sidechain(p, model=0):
HP = ["ALA","GLY","VAL","ILE","LEU","PHE","MET","PRO","TRP","CYS","TYR"]
BB = ['C','O','N']
p.addStyle({"model":model,'and':[{'resn':HP},{'atom':BB,'invert':True}]},
{'stick':{'colorscheme':"whiteCarbon",'radius':0.4}})
p.addStyle({"model":model,'and':[{'resn':"GLY"},{'atom':'CA'}]},
{'sphere':{'colorscheme':"whiteCarbon",'radius':0.4}})
p.addStyle({"model":model,'and':[{'resn':"PRO"},{'atom':['C','O'],'invert':True}]},
{'stick':{'colorscheme':"whiteCarbon",'radius':0.4}})
p.addStyle({"model":model,'and':[{'resn':HP,'invert':True},{'atom':BB,'invert':True}]},
{'stick':{'colorscheme':"whiteCarbon",'radius':0.4}})
p = py3Dmol.view(js='https://3dmol.org/build/3Dmol.js')
p.addModel(open(pdb,'r').read(),'pdb')
if color == "rainbow":
p.setStyle({'cartoon': {'color':'spectrum'}})
else:
p.setStyle({'cartoon':{}})
if show_sidechains: sidechain(p)
if show_mainchain: mainchain(p)
if show_box: box(p)
p.zoomTo()
return p.show()
interact(show_pdb,
show_box=ipywidgets.Checkbox(value=True),
show_sidechains=ipywidgets.Checkbox(value=False),
show_mainchain=ipywidgets.Checkbox(value=False),
color=ipywidgets.Dropdown(options=['None', 'rainbow'], value='rainbow'))
```
---
---
# **シミュレーションボックスの平衡化**
適切なMD平衡化プロトコルは、タンパク質の実験的なコンフォメーションを維持しながら、シミュレーションボックス全体で温度と圧力の両方を平衡化するように設計されています。さらに、溶媒がタンパク質の周りに馴染むようにし、適切な溶媒和層を形成します。
以下では、温度、圧力、シミュレーション時間などのMD平衡化パラメータを設定します。また、タンパク質の重原子をその場に拘束しておくための力定数(force constant)や、原子座標をトラジェクトリファイル(.dcd)に保存する頻度も定義します。
設定が終わったら、次の2つのセルを実行して系を平衡化することができます。
```
#@title ### **MD平衡化プロトコルのパラメータ:**
# remove whitespaces
Jobname = '1AKI_equil' #@param {type:"string"}
Minimization_steps = "1000" #@param ["1000", "5000", "10000", "20000", "50000", "100000"]
#@markdown シミュレーション時間(ナノ秒)と積分時間(フェムト秒):
Time = "2" #@param {type:"string"}
stride_time_eq = Time
Integration_timestep = "2" #@param ["0.5", "1", "2", "3", "4"]
dt_eq = Integration_timestep
#@markdown 温度(ケルビン)と圧力(バール)
Temperature = 298 #@param {type:"string"}
temperature_eq = Temperature
Pressure = 1 #@param {type:"string"}
pressure_eq = Pressure
#@markdown 位置拘束の力定数(kJ/mol):
Force_constant = 800 #@param {type:"slider", min:0, max:2000, step:100}
#@markdown トラジェクトリファイルを書き出す頻度(ピコ秒):
Write_the_trajectory = "10" #@param ["10", "100", "200", "500", "1000"]
write_the_trajectory_eq = Write_the_trajectory
#@markdown ログファイルを書き出す頻度(ピコ秒):
Write_the_log = "10" #@param ["10", "100", "200", "500", "1000"]
write_the_log_eq = Write_the_log
#@markdown ---
#@title **平衡化MDシミュレーション(NPTアンサンブル)の実行**
#@markdown さあ、系を平衡化しましょう!
###########################################
import openmm as mm
from openmm import *
from openmm.app import *
from openmm.unit import *
import pytraj as pt
from sys import stdout, exit, stderr
import os, math, fnmatch
#############################################
# Defining MD simulation parameters
jobname = os.path.join(workDir, Jobname)
coordinatefile = os.path.join(workDir, str(CRD_filename))
pdbfile = os.path.join(workDir, str(PDB_filename))
topologyfile = os.path.join(workDir, str(PRMTOP_filename))
time_ps = float(Time)*1000
simulation_time = float(time_ps)*picosecond # in ps
dt = int(dt_eq)*femtosecond
temperature = float(temperature_eq)*kelvin
savcrd_freq = int(write_the_trajectory_eq)*picosecond
print_freq = int(write_the_log_eq)*picosecond
pressure = float(pressure_eq)*bar
restraint_fc = int(Force_constant) # kJ/mol
nsteps = int(simulation_time.value_in_unit(picosecond)/dt.value_in_unit(picosecond))
nprint = int(print_freq.value_in_unit(picosecond)/dt.value_in_unit(picosecond))
nsavcrd = int(savcrd_freq.value_in_unit(picosecond)/dt.value_in_unit(picosecond))
#############################################
# Defining functions to use below:
def backup_old_log(pattern, string):
result = []
for root, dirs, files in os.walk("./"):
for name in files:
if fnmatch.fnmatch(name, pattern):
try:
number = int(name[-2])
avail = isinstance(number, int)
#print(name,avail)
if avail == True:
result.append(number)
except:
pass
if len(result) > 0:
maxnumber = max(result)
else:
maxnumber = 0
backup_file = "\#" + string + "." + str(maxnumber + 1) + "#"
os.system("mv " + string + " " + backup_file)
return backup_file
def restraints(system, crd, fc, restraint_array):
boxlx = system.getDefaultPeriodicBoxVectors()[0][0].value_in_unit(nanometers)
boxly = system.getDefaultPeriodicBoxVectors()[1][1].value_in_unit(nanometers)
boxlz = system.getDefaultPeriodicBoxVectors()[2][2].value_in_unit(nanometers)
if fc > 0:
# positional restraints for all heavy-atoms
posresPROT = CustomExternalForce('k*periodicdistance(x, y, z, x0, y0, z0)^2;')
posresPROT.addPerParticleParameter('k')
posresPROT.addPerParticleParameter('x0')
posresPROT.addPerParticleParameter('y0')
posresPROT.addPerParticleParameter('z0')
for atom1 in restraint_array:
atom1 = int(atom1)
xpos = crd.positions[atom1].value_in_unit(nanometers)[0]
ypos = crd.positions[atom1].value_in_unit(nanometers)[1]
zpos = crd.positions[atom1].value_in_unit(nanometers)[2]
posresPROT.addParticle(atom1, [fc, xpos, ypos, zpos])
system.addForce(posresPROT)
return system
##############################################
#############################################
print("\n> Simulation details:\n")
print("\tJob name = " + jobname)
print("\tCoordinate file = " + str(coordinatefile))
print("\tPDB file = " + str(pdbfile))
print("\tTopology file = " + str(topologyfile))
print("\n\tSimulation_time = " + str(simulation_time))
print("\tIntegration timestep = " + str(dt))
print("\tTotal number of steps = " + str(nsteps))
print("\n\tSave coordinates each " + str(savcrd_freq))
print("\tPrint in log file each " + str(print_freq))
print("\n\tTemperature = " + str(temperature))
print("\tPressure = " + str(pressure))
#############################################
print("\n> Setting the system:\n")
print("\t- Reading topology and structure file...")
prmtop = AmberPrmtopFile(topologyfile)
inpcrd = AmberInpcrdFile(coordinatefile)
print("\t- Creating system and setting parameters...")
nonbondedMethod = PME
nonbondedCutoff = 1.0*nanometers
ewaldErrorTolerance = 0.0005
constraints = HBonds
rigidWater = True
constraintTolerance = 0.000001
friction = 1.0
system = prmtop.createSystem(nonbondedMethod=nonbondedMethod, nonbondedCutoff=nonbondedCutoff,
constraints=constraints, rigidWater=rigidWater, ewaldErrorTolerance=ewaldErrorTolerance)
print("\t- Applying restraints. Force Constant = " + str(Force_constant) + "kJ/mol")
pt_system = pt.iterload(coordinatefile, topologyfile)
pt_topology = pt_system.top
restraint_array = pt.select_atoms('!(:H*) & !(:WAT) & !(:Na+) & !(:Cl-) & !(:Mg+) & !(:K+)', pt_topology)
system = restraints(system, inpcrd, restraint_fc, restraint_array)
print("\t- Setting barostat...")
system.addForce(MonteCarloBarostat(pressure, temperature))
print("\t- Setting integrator...")
integrator = LangevinIntegrator(temperature, friction, dt)
integrator.setConstraintTolerance(constraintTolerance)
simulation = Simulation(prmtop.topology, system, integrator)
simulation.context.setPositions(inpcrd.positions)
if inpcrd.boxVectors is not None:
simulation.context.setPeriodicBoxVectors(*inpcrd.boxVectors)
print("\t- Energy minimization: " + str(Minimization_steps) + " steps")
simulation.minimizeEnergy(tolerance=10*kilojoule/mole, maxIterations=int(Minimization_steps))
print("\t-> Potential Energy = " + str(simulation.context.getState(getEnergy=True).getPotentialEnergy()))
print("\t- Setting initial velocities...")
simulation.context.setVelocitiesToTemperature(temperature)
#############################################
# Running Equilibration on NPT ensemble
dcd_file = jobname + ".dcd"
log_file = jobname + ".log"
rst_file = jobname + ".rst"
prv_rst_file = jobname + ".rst"
pdb_file = jobname + ".pdb"
# Creating a trajectory file and reporters
dcd = DCDReporter(dcd_file, nsavcrd)
firstdcdstep = (nsteps) + nsavcrd
dcd._dcd = DCDFile(dcd._out, simulation.topology, simulation.integrator.getStepSize(), firstdcdstep, nsavcrd) # charmm doesn't like first step to be 0
simulation.reporters.append(dcd)
simulation.reporters.append(StateDataReporter(stdout, nprint, step=True, speed=True, progress=True, totalSteps=nsteps, remainingTime=True, separator='\t\t'))
simulation.reporters.append(StateDataReporter(log_file, nprint, step=True, kineticEnergy=True, potentialEnergy=True, totalEnergy=True, temperature=True, volume=True, speed=True))
print("\n> Simulating " + str(nsteps) + " steps...")
simulation.step(nsteps)
simulation.reporters.clear() # remove all reporters so the next iteration don't trigger them.
##################################
# Writing last frame information of stride
print("\n> Writing state file (" + str(rst_file) + ")...")
state = simulation.context.getState( getPositions=True, getVelocities=True )
with open(rst_file, 'w') as f:
f.write(XmlSerializer.serialize(state))
last_frame = int(nsteps/nsavcrd)
print("> Writing coordinate file (" + str(pdb_file) + ", frame = " + str(last_frame) + ")...")
positions = simulation.context.getState(getPositions=True).getPositions()
PDBFile.writeFile(simulation.topology, positions, open(pdb_file, 'w'))
print("\n> Finished!\n")
```
---
---
# **MDシミュレーション本番の実行(Production)**
最後に、平衡化された系の座標を入力構造として、シミュレーション本番(Production simulation)そのものを進めます。
ここでは、熱力学的に平衡化された系から本番のシミュレーションを開始することを保証するために、平衡化シミュレーションの最終フレームの原子の位置と速度を含む*.rst 状態ファイル*を使用することに注意してください。
ここでもう一つの重要な情報は**Number_of_strides**と**Stride_Time**。このノートブックでは指定した*stride*数のシミュレーションを行うので、**simulation time = Number_of_strides*Stride_Time**となります。例えば、*Number_of_strides=10* と*Stride_Time=10 ns*と設定することで100nsシミュレーションできます。
**重要:Productionシミュレーションの最後に、すべてのstrideを連結して完全なトラジェクトリファイルを作成し、可視化および分析することができます。**
この方法の背景にあるアイデアは、Google ColabでGPUを使える断続的な時間(12h/24h)をうまく利用することです。
```
#@markdown ### **インプットファイルの名前を下に記入してください:**
Equilibrated_PDB = '1AKI_equil.pdb' #@param {type:"string"}
State_file = '1AKI_equil.rst' #@param {type:"string"}
#@markdown ---
#@markdown ### **MD Prodcutionプロトコルのパラメータ:**
# remove whitespaces
Jobname = '1AKI_prod' #@param {type:"string"}
#@markdown シミュレーション時間(ナノ秒)、stride数(整数)と積分時間(フェムト秒):
Stride_Time = "5" #@param {type:"string"}
stride_time_prod = Stride_Time
Number_of_strides = "1" #@param {type:"string"}
nstride = Number_of_strides
Integration_timestep = "2" #@param ["0.5", "1", "2", "3", "4"]
dt_prod = Integration_timestep
#@markdown 温度(ケルビン)と圧力(バール)
Temperature = 298 #@param {type:"string"}
temperature_prod = Temperature
Pressure = 1 #@param {type:"string"}
pressure_prod = Pressure
#@markdown トラジェクトリファイルを書き出す頻度(ピコ秒):
Write_the_trajectory = "10" #@param ["10", "100", "200", "500", "1000"]
write_the_trajectory_prod = Write_the_trajectory
#@markdown ログファイルを書き出す頻度(ピコ秒):
Write_the_log = "10" #@param ["10", "100", "200", "500", "1000"]
write_the_log_prod = Write_the_log
#@markdown ---
#@title **平衡化した後のMDシミュレーション本番(Production)(NPTアンサンブル)**
#
###########################################
import openmm as mm
from openmm import *
from openmm.app import *
from openmm.unit import *
from sys import stdout, exit, stderr
import os, math, fnmatch
#############################################
# Defining MD simulation parameters
jobname = os.path.join(workDir, str(Jobname))
coordinatefile = os.path.join(workDir, str(CRD_filename))
pdbfile = os.path.join(workDir, Equilibrated_PDB)
topologyfile = os.path.join(workDir, str(PRMTOP_filename))
equil_rst_file = os.path.join(workDir, State_file)
stride_time_ps = float(stride_time_prod)*1000
stride_time = float(stride_time_ps)*picosecond
nstride = int(Number_of_strides)
dt = int(dt_prod)*femtosecond
temperature = float(temperature_prod)*kelvin
savcrd_freq = int(write_the_trajectory_prod)*picosecond
print_freq = int(write_the_log_prod)*picosecond
pressure = float(pressure_prod)*bar
simulation_time = stride_time*nstride
nsteps = int(stride_time.value_in_unit(picosecond)/dt.value_in_unit(picosecond))
nprint = int(print_freq.value_in_unit(picosecond)/dt.value_in_unit(picosecond))
nsavcrd = int(savcrd_freq.value_in_unit(picosecond)/dt.value_in_unit(picosecond))
firststride = 1 # must be integer
#############################################
# Defining functions to use below:
def backup_old_log(pattern, string):
result = []
for root, dirs, files in os.walk("./"):
for name in files:
if fnmatch.fnmatch(name, pattern):
try:
number = int(name[-2])
avail = isinstance(number, int)
#print(name,avail)
if avail == True:
result.append(number)
except:
pass
if len(result) > 0:
maxnumber = max(result)
else:
maxnumber = 0
backup_file = "\#" + string + "." + str(maxnumber + 1) + "#"
os.system("mv " + string + " " + backup_file)
return backup_file
##############################################
#############################################
print("\n> Simulation details:\n")
print("\tJob name = " + jobname)
print("\tCoordinate file = " + str(coordinatefile))
print("\tPDB file = " + str(pdbfile))
print("\tTopology file = " + str(topologyfile))
print("\n\tSimulation_time = " + str(stride_time*nstride))
print("\tIntegration timestep = " + str(dt))
print("\tTotal number of steps = " + str(nsteps*nstride))
print("\tNumber of strides = " + str(nstride) + " (" + str(stride_time) + " in each stride)")
print("\n\tSave coordinates each " + str(savcrd_freq))
print("\tSave checkpoint each " + str(savcrd_freq))
print("\tPrint in log file each " + str(print_freq))
print("\n\tTemperature = " + str(temperature))
print("\tPressure = " + str(pressure))
#############################################
print("\n> Setting the system:\n")
print("\t- Reading topology and structure file...")
prmtop = AmberPrmtopFile(topologyfile)
inpcrd = AmberInpcrdFile(coordinatefile)
print("\t- Creating system and setting parameters...")
nonbondedMethod = PME
nonbondedCutoff = 1.0*nanometers
ewaldErrorTolerance = 0.0005
constraints = HBonds
rigidWater = True
constraintTolerance = 0.000001
friction = 1.0
system = prmtop.createSystem(nonbondedMethod=nonbondedMethod, nonbondedCutoff=nonbondedCutoff,
constraints=constraints, rigidWater=rigidWater, ewaldErrorTolerance=ewaldErrorTolerance)
print("\t- Setting barostat...")
system.addForce(MonteCarloBarostat(pressure, temperature))
print("\t- Setting integrator...")
integrator = LangevinIntegrator(temperature, friction, dt)
integrator.setConstraintTolerance(constraintTolerance)
simulation = Simulation(prmtop.topology, system, integrator)
simulation.context.setPositions(inpcrd.positions)
if inpcrd.boxVectors is not None:
simulation.context.setPeriodicBoxVectors(*inpcrd.boxVectors)
#############################################
# Opening a loop of extension NSTRIDE to simulate the entire STRIDE_TIME*NSTRIDE
for n in range(1, nstride + 1):
print("\n\n>>> Simulating Stride #" + str(n) + " <<<")
dcd_file = jobname + "_" + str(n) + ".dcd"
log_file = jobname + "_" + str(n) + ".log"
rst_file = jobname + "_" + str(n) + ".rst"
prv_rst_file = jobname + "_" + str(n-1) + ".rst"
pdb_file = jobname + "_" + str(n) + ".pdb"
if os.path.exists(rst_file):
print("> Stride #" + str(n) + " finished (" + rst_file + " present). Moving to next stride... <")
continue
if n == 1:
print("\n> Loading previous state from equilibration > " + equil_rst_file + " <")
with open(equil_rst_file, 'r') as f:
simulation.context.setState(XmlSerializer.deserialize(f.read()))
currstep = int((n-1)*nsteps)
currtime = currstep*dt.in_units_of(picosecond)
simulation.currentStep = currstep
simulation.context.setTime(currtime)
print("> Current time: " + str(currtime) + " (Step = " + str(currstep) + ")")
else:
print("> Loading previous state from > " + prv_rst_file + " <")
with open(prv_rst_file, 'r') as f:
simulation.context.setState(XmlSerializer.deserialize(f.read()))
currstep = int((n-1)*nsteps)
currtime = currstep*dt.in_units_of(picosecond)
simulation.currentStep = currstep
simulation.context.setTime(currtime)
print("> Current time: " + str(currtime) + " (Step = " + str(currstep) + ")")
dcd = DCDReporter(dcd_file, nsavcrd)
firstdcdstep = (currstep) + nsavcrd
dcd._dcd = DCDFile(dcd._out, simulation.topology, simulation.integrator.getStepSize(), firstdcdstep, nsavcrd) # first step should not be 0
simulation.reporters.append(dcd)
simulation.reporters.append(StateDataReporter(stdout, nprint, step=True, speed=True, progress=True, totalSteps=(nsteps*nstride), remainingTime=True, separator='\t\t'))
simulation.reporters.append(StateDataReporter(log_file, nprint, step=True, kineticEnergy=True, potentialEnergy=True, totalEnergy=True, temperature=True, volume=True, speed=True))
print("\n> Simulating " + str(nsteps) + " steps... (Stride #" + str(n) + ")")
simulation.step(nsteps)
simulation.reporters.clear() # remove all reporters so the next iteration don't trigger them.
##################################
# Writing last frame information of stride
print("\n> Writing state file (" + str(rst_file) + ")...")
state = simulation.context.getState( getPositions=True, getVelocities=True )
with open(rst_file, 'w') as f:
f.write(XmlSerializer.serialize(state))
last_frame = int(nsteps/nsavcrd)
print("> Writing coordinate file (" + str(pdb_file) + ", frame = " + str(last_frame) + ")...")
positions = simulation.context.getState(getPositions=True).getPositions()
PDBFile.writeFile(simulation.topology, positions, open(pdb_file, 'w'))
print("\n> Finished!\n")
#@title **トラジェクトリを連結し整列する**
Skip = "1" #@param ["1", "2", "5", "10", "20", "50"]
stride_traj = Skip
Output_format = "dcd" #@param ["dcd", "pdb", "trr", "xtc"]
#@markdown **注意:** フレーム数が大きすぎるとColabのメモリ許容範囲を超えてしまいます。5000フレーム以下なら十分です。
simulation_time_analysis = stride_time_ps*nstride
simulation_ns = float(Stride_Time)*int(Number_of_strides)
number_frames = int(simulation_time_analysis)/int(Write_the_trajectory)
number_frames_analysis = number_frames/int(stride_traj)
traj_end = os.path.join(workDir, str(Jobname) + "_all.dcd")
traj_end2 = os.path.join(workDir, str(Jobname) + "_all." + str(Output_format))
template = os.path.join(workDir, str(Jobname) + '_%s.dcd')
flist = [template % str(i) for i in range(1, nstride + 1)]
#print(flist)
trajlist = pt.load(flist, pdb, stride=stride_traj)
traj_image = trajlist.iterframe(autoimage=True, rmsfit=0)
traj_write = pt.write_traj(traj_end, traj_image, overwrite=True)
traj_load = pt.load(traj_end, pdb)
traj_align = pt.align(traj_load, mask="@CA", ref=0)
traj_write = pt.write_traj(traj_end, traj_align, overwrite=True, options='dcd')
traj_write = pt.write_traj(traj_end2, traj_align, overwrite=True, options=Output_format)
traj_load = pt.load(traj_end, pdb)
print(traj_load)
traj_end_check = os.path.exists(traj_end2)
if traj_end_check == True:
print("Trajectory concatenated successfully! :-)")
else:
print("ERROR: Check your inputs! ")
#@title **トラジェクトリの読み込み、可視化と確認*
#@markdown しばらく時間がかかります。コーヒーをもう一杯どうでしょう? :-)
#@markdown **注意:** もし系の原子数が100K以上なら、このステップをスキップしてください。原子数が多いとColabのメモリ制限を超えて止まる可能性があります。
import warnings
warnings.filterwarnings('ignore')
!rm *.pdb 2> /dev/null
#py3dmol functions
class Atom(dict):
def __init__(self, line):
self["type"] = line[0:6].strip()
self["idx"] = line[6:11].strip()
self["name"] = line[12:16].strip()
self["resname"] = line[17:20].strip()
self["resid"] = int(int(line[22:26]))
self["x"] = float(line[30:38])
self["y"] = float(line[38:46])
self["z"] = float(line[46:54])
self["sym"] = line[76:78].strip()
def __str__(self):
line = list(" " * 80)
line[0:6] = self["type"].ljust(6)
line[6:11] = self["idx"].ljust(5)
line[12:16] = self["name"].ljust(4)
line[17:20] = self["resname"].ljust(3)
line[22:26] = str(self["resid"]).ljust(4)
line[30:38] = str(self["x"]).rjust(8)
line[38:46] = str(self["y"]).rjust(8)
line[46:54] = str(self["z"]).rjust(8)
line[76:78] = self["sym"].rjust(2)
return "".join(line) + "\n"
class Molecule(list):
def __init__(self, file):
for line in file:
if "ATOM" in line or "HETATM" in line:
self.append(Atom(line))
def __str__(self):
outstr = ""
for at in self:
outstr += str(at)
return outstr
if number_frames_analysis > 10:
stride_animation = number_frames_analysis/10
else:
stride_animation = 1
u = mda.Universe(top, traj_end)
# Write out frames for animation
protein = u.select_atoms('not (resname WAT)')
i = 0
for ts in u.trajectory[0:len(u.trajectory):int(stride_animation)]:
if i > -1:
with mda.Writer('' + str(i) + '.pdb', protein.n_atoms) as W:
W.write(protein)
i = i + 1
# Load frames as molecules
molecules = []
for i in range(int(len(u.trajectory)/int(stride_animation))):
with open('' + str(i) + '.pdb') as ifile:
molecules.append(Molecule(ifile))
models = ""
for i in range(len(molecules)):
models += "MODEL " + str(i) + "\n"
for j,mol in enumerate(molecules[i]):
models += str(mol)
models += "ENDMDL\n"
#view.addModelsAsFrames(models)
# Animation
view = py3Dmol.view(width=800, height=600)
view.addModelsAsFrames(models)
for i, at in enumerate(molecules[0]):
default = {"cartoon": {'color': 'spectrum'}}
view.setStyle({'model': -1, 'serial': i+1}, at.get("pymol", default))
view.zoomTo()
view.animate({'loop': "forward"})
view.show()
```
---
---
# **解析**
トラジェクトリを可視化することは非常に有効ですが、より定量的なデータも時には必要です。
MDトラジェクトリの解析は多岐にわたるので、ここですべてを網羅するつもりはありません。しかし、MDanalysisやPyTraj を利用することで、簡単にシミュレーションを解析することができます。
以下では、シミュレーションの挙動を解明するのに光を当てるのに役立つコードスニペットの例をいくつか示します。
```
#@title **タンパク質CA原子のRMSDを計算**
#@markdown **出力ファイルの名前を下に記入してください:**
Output_name = 'rmsd_ca' #@param {type:"string"}
rmsd = pt.rmsd(traj_load, ref = 0, mask = "@CA")
time = len(rmsd)*int(Write_the_trajectory)/1000
time_array = np.arange(0,time,int(Write_the_trajectory)/1000)*int(stride_traj)
# Plotting:
ax = plt.plot(time_array, rmsd, alpha=0.6, color = 'blue', linewidth = 1.0)
plt.xlim(0, simulation_ns)
#plt.ylim(2, 6)
plt.xlabel("Time (ns)", fontsize = 14, fontweight = 'bold')
plt.ylabel("RMSD [$\AA$]", fontsize = 14, fontweight = 'bold')
plt.xticks(fontsize = 12)
plt.yticks(fontsize = 12)
plt.savefig(os.path.join(workDir, Output_name + ".png"), dpi=600, bbox_inches='tight')
raw_data=pd.DataFrame(rmsd)
raw_data.to_csv(os.path.join(workDir, Output_name + ".csv"))
#@title **RMSDを分布としてプロット**
#@markdown **出力ファイルの名前を下に記入してください:**
Output_name = 'rmsd_dist' #@param {type:"string"}
ax = sb.kdeplot(rmsd, color="blue", shade=True, alpha=0.2, linewidth=0.5)
plt.xlabel('RMSD [$\AA$]', fontsize = 14, fontweight = 'bold')
plt.xticks(fontsize = 12)
plt.yticks([])
plt.ylabel('')
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['bottom'].set_visible(True)
ax.spines['left'].set_visible(False)
plt.savefig(os.path.join(workDir, Output_name + ".png"), dpi=600, bbox_inches='tight')
#@title **タンパク質CA原子の慣性半径(radius of gyration )を計算**
#@markdown **出力ファイルの名前を下に記入してください:**
Output_name = 'radius_gyration' #@param {type:"string"}
radgyr = pt.radgyr(traj_load, mask = "@CA")
time = len(rmsd)*int(Write_the_trajectory)/1000
time_array = np.arange(0,time,int(Write_the_trajectory)/1000)*int(stride_traj)
# Plotting:
plt.plot(time_array, radgyr, alpha=0.6, color = 'green', linewidth = 1.0)
plt.xlim(0, simulation_ns)
#plt.ylim(2, 6)
plt.xlabel("Time (ns)", fontsize = 14, fontweight = 'bold')
plt.ylabel("Radius of gyration ($\AA$)", fontsize = 14, fontweight = 'bold')
plt.xticks(fontsize = 12)
plt.yticks(fontsize = 12)
plt.savefig(os.path.join(workDir, Output_name + ".png"), dpi=600, bbox_inches='tight')
raw_data=pd.DataFrame(radgyr)
raw_data.to_csv(os.path.join(workDir, Output_name + ".csv"))
#@title **慣性半径を分布としてプロット**
#@markdown **出力ファイルの名前を下に記入してください:**
Output_name = 'radius_gyration_dist' #@param {type:"string"}
ax = sb.kdeplot(radgyr, color="green", shade=True, alpha=0.2, linewidth=0.5)
plt.xlabel('Radius of gyration ($\AA$)', fontsize = 14, fontweight = 'bold')
plt.xticks(fontsize = 12)
plt.yticks([])
plt.ylabel('')
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['bottom'].set_visible(True)
ax.spines['left'].set_visible(False)
plt.savefig(os.path.join(workDir, Output_name + ".png"), dpi=600, bbox_inches='tight')
#@title **タンパク質CA原子のRMSFを計算**
#@markdown **出力ファイルの名前を下に記入してください:**
Output_name = 'rmsf_ca' #@param {type:"string"}
rmsf = pt.rmsf(traj_load, "@CA")
bfactor = pt.bfactors(traj_load, byres=True)
# Plotting:
plt.plot(rmsf[:,1], alpha=1.0, color = 'red', linewidth = 1.0)
plt.xlabel("Residue", fontsize = 14, fontweight = 'bold')
plt.ylabel("RMSF ($\AA$)", fontsize = 14, fontweight = 'bold')
plt.xticks(fontsize = 12)
plt.xlim(0, len(rmsf[:-1]))
#plt.xticks(np.arange(min(rmsf[:1]), max(rmsf[:1])))
plt.yticks(fontsize = 12)
plt.savefig(os.path.join(workDir, Output_name + ".png"), dpi=600, bbox_inches='tight')
raw_data=pd.DataFrame(rmsf)
raw_data.to_csv(os.path.join(workDir, Output_name + ".csv"))
#@title **2D RMSD**
#@markdown **出力ファイルの名前を下に記入してください:**
Output_name = '2D_rmsd' #@param {type:"string"}
last_frame = len(time_array)
stride_ticks_f = (last_frame)/5
ticks_frame = np.arange(0,(len(time_array) + float(stride_ticks_f)), float(stride_ticks_f))
a = ticks_frame.astype(float)
stride_ticks_t = (simulation_ns)/5
tick_time = np.arange(0,(float(simulation_ns) + float(stride_ticks_t)), float(stride_ticks_t))
b = tick_time.astype(float)
mat1 = pt.pairwise_rmsd(traj_load, mask="@CA", frame_indices=range(int(number_frames_analysis)))
ax = plt.imshow(mat1, cmap = 'PRGn', origin='lower', interpolation = 'bicubic')
plt.title('2D RMSD')
plt.xlabel('Time (ns)', fontsize = 14, fontweight = 'bold')
plt.ylabel('Time (ns)', fontsize = 14, fontweight = 'bold')
# plt.xticks(fontsize = 12)
# plt.yticks(fontsize = 12)
plt.xticks(a, b.round(decimals=3), fontsize = 12)
plt.yticks(a, b.round(decimals=3), fontsize = 12)
# plt.xlim(0, a[-1])
# plt.ylim(0, a[-1])
cbar1 = plt.colorbar()
cbar1.set_label("RMSD ($\AA$)", fontsize = 14, fontweight = 'bold')
plt.savefig(os.path.join(workDir, Output_name + ".png"), dpi=600, bbox_inches='tight')
raw_data=pd.DataFrame(mat1)
raw_data.to_csv(os.path.join(workDir, Output_name + ".csv"))
#@title **主成分分析(PCA)の固有ベクトルを計算**
data = pt.pca(traj_load, fit=True, ref=0, mask='@CA', n_vecs=2)
#print('projection values of each frame to first mode = {} \n'.format(data[0][0]))
#print('projection values of each frame to second mode = {} \n'.format(data[0][1]))
#print('eigvenvalues of first two modes', data[1][0])
#print("")
#print('eigvenvectors of first two modes: \n', data[1][1])
last_frame = len(time_array)
stride_ticks_f = (last_frame)/5
ticks_frame = np.arange(0,(len(time_array) + float(stride_ticks_f)), float(stride_ticks_f))
a = ticks_frame.astype(float)
a2 = a.tolist()
stride_ticks_t = (simulation_ns)/5
tick_time = np.arange(0,(float(simulation_ns) + float(stride_ticks_t)), float(stride_ticks_t))
b = tick_time.astype(float)
#@markdown **出力ファイルの名前を下に記入してください:**
Output_name = 'PCA' #@param {type:"string"}
Output_PC1 = 'PC1' #@param {type:"string"}
Output_PC2 = 'PC2' #@param {type:"string"}
%matplotlib inline
%config InlineBackend.figure_format = 'retina' # high resolution
projection_data = data[0]
plt.title(r'PCA of C-$\alpha$')
PC1 = data[0][0]
PC2 = data[0][1]
a = plt.scatter(PC1,PC2, c=range(int(number_frames_analysis)), cmap='Greens', marker='o',s=8, alpha=1)
plt.clim(0, last_frame)
plt.xlabel('PC1', fontsize = 14, fontweight = 'bold')
plt.ylabel('PC2', fontsize = 14, fontweight = 'bold')
plt.xticks(fontsize = 12)
plt.yticks(fontsize = 12)
# N = len(number_frames)
# x2 = np.arange(N)
cbar1 = plt.colorbar(a, orientation="vertical")
cbar1.set_label('Time(ns)', fontsize = 14, fontweight = 'bold')
cbar1.set_ticks(a2)
cbar1.set_ticklabels(b.round(decimals=3))
plt.savefig(os.path.join(workDir, Output_name + ".png"), dpi=600, bbox_inches='tight')
pc1=pd.DataFrame(PC1)
pc1.to_csv(os.path.join(workDir, Output_PC1 + ".csv"))
pc2=pd.DataFrame(PC2)
pc2.to_csv(os.path.join(workDir, Output_PC2 + ".csv"))
#@title **主成分1(PC1)と主成分2(PC2)を分布としてプロット**
Output_name = 'PCA_dist' #@param {type:"string"}
fig = plt.figure(figsize=(9,5))
plt.subplot(1, 2, 1)
ax = sb.kdeplot(PC1, color="green", shade=True, alpha=0.2, linewidth=0.5)
plt.xlabel('PC1', fontsize = 14, fontweight = 'bold')
plt.xticks(fontsize = 12)
plt.yticks([])
plt.ylabel('')
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['bottom'].set_visible(True)
ax.spines['left'].set_visible(False)
plt.subplot(1, 2, 2)
ax2 = sb.kdeplot(PC2, color="purple", shade=True, alpha=0.2, linewidth=0.5)
plt.xlabel('PC2', fontsize = 14, fontweight = 'bold')
plt.xticks(fontsize = 12)
plt.yticks([])
plt.ylabel('')
ax2.spines['top'].set_visible(False)
ax2.spines['right'].set_visible(False)
ax2.spines['bottom'].set_visible(True)
ax2.spines['left'].set_visible(False)
plt.savefig(os.path.join(workDir, Output_name + ".png"), dpi=600, bbox_inches='tight')
#@title **Pearson's Cross Correlation (CC)**
#@markdown **出力ファイルの名前を下に記入してください:**
Output_name = 'cross_correlation' #@param {type:"string"}
traj_align = pt.align(traj_load, mask='@CA', ref=0)
mat_cc = matrix.correl(traj_align, '@CA')
ax = plt.imshow(mat_cc, cmap = 'PiYG_r', interpolation = 'bicubic', vmin = -1, vmax = 1, origin='lower')
plt.xlabel('Residues', fontsize = 14, fontweight = 'bold')
plt.ylabel('Residues', fontsize = 14, fontweight = 'bold')
plt.xticks(fontsize = 12)
plt.yticks(fontsize = 12)
cbar1 = plt.colorbar()
cbar1.set_label('$CC_ij$', fontsize = 14, fontweight = 'bold')
plt.savefig(os.path.join(workDir, Output_name + ".png"), dpi=600, bbox_inches='tight')
raw_data=pd.DataFrame(mat_cc)
raw_data.to_csv(os.path.join(workDir, Output_name + ".csv"))
```
| github_jupyter |
```
%%file sumArraysOnGPU.cu
#include <stdlib.h>
#include <string.h>
#include <time.h>
#include <stdio.h>
#include <cuda_runtime.h>
#define CHECK(call) \
{ \
const cudaError_t error = call; \
if (error != cudaSuccess) \
{ \
fprintf(stderr, "Error: %s:%d, ", __FILE__, __LINE__); \
fprintf(stderr, "code: %d, reason: %s\n", error, \
cudaGetErrorString(error)); \
exit(1); \
} \
}
__global__ void sumArraysOnDevice(float *A, float *B, float *C){
int idx = blockIdx.x * blockDim.x + threadIdx.x;
C[idx] = A[idx] + B[idx];
}
void initialData(float *ip, int size){
// generate different seed for random number
time_t t;
srand((unsigned int) time (&t));
for (int i=0; i<size; i++){
ip[i] = (float)(rand() & 0xFF) / 10.0f;
}
}
void sumArraysOnHost(float *A, float *B, float *C, const int N){
for (int idx=0; idx<N; idx++){
C[idx] = A[idx] + B[idx];
}
}
void checkResult(float *hostRef, float *gpuRef, const int N){
double epsilon = 1.0E-8;
int match = 1;
for (int i = 0; i < N; i++){
if (abs(hostRef[i] - gpuRef[i]) > epsilon){
match = 0;
printf("Arrays do not match!\n");
printf("host %5.2f gpu %5.2f at current %d\n",
hostRef[i], gpuRef[i], i);
break;
}
}
if (match) printf("Arrays match. \n\n");
}
int main(int argc, char **argv){
printf("%s Starting...\n", argv[0]);
// malloc host memory
int nElem = 10000;
size_t nBytes = nElem * sizeof(float);
// initialize data at host side
float *h_A, *h_B, *hostRef, *gpuRef;
h_A = (float *)malloc(nBytes);
h_B = (float *)malloc(nBytes);
hostRef = (float *)malloc(nBytes);
gpuRef = (float *)malloc(nBytes);
// initialize data at host side
initialData(h_A, nElem);
initialData(h_B, nElem);
memset(hostRef, 0, nBytes);
memset(gpuRef, 0, nBytes);
// malloc device global memory
float *d_A, *d_B, *d_C;
cudaMalloc((float**)&d_A, nBytes);
cudaMalloc((float**)&d_B, nBytes);
cudaMalloc((float**)&d_C, nBytes);
// Use cudaMemcpy to transfer the data from the host memory to the GPU global memory with the
// parameter cudaMemcpyHostToDevice specifying the transfer direction.
CHECK(cudaMemcpy(d_A, h_A, nBytes, cudaMemcpyHostToDevice));
CHECK(cudaMemcpy(d_B, h_B, nBytes, cudaMemcpyHostToDevice));
// invoke kernel at host side
dim3 block(100);
dim3 grid(nElem / block.x);
sumArraysOnDevice<<<grid, block>>>(d_A, d_B, d_C);
printf("Execution configuration <<<%d, %d>>>\n", grid.x, block.x);
// copy kernel result back to host side
cudaMemcpy(gpuRef, d_C, nBytes, cudaMemcpyDeviceToHost);
// add vector at host side for result checks
sumArraysOnHost(h_A, h_B, hostRef, nElem);
for (int i=0; i<10; i++){
printf("%f + %f = %f \n", h_A[i], h_B[i], hostRef[i]);
}
// check device results
checkResult(hostRef, gpuRef, nElem);
free(h_A);
free(h_B);
free(hostRef);
free(gpuRef);
// use cudaFree to release the memory used on the GPU
cudaFree(d_A);
cudaFree(d_B);
cudaFree(d_C);
cudaDeviceReset();
return (0);
}
%%bash
nvcc sumArraysOnGPU.cu -o addvector
nvprof --unified-memory-profiling off ./addvector
```
| github_jupyter |
# FloPy
## MODFLOW-USG CLN package demo
This example problem demonstrates use of the CLN process for simulating flow to a well pumping from two aquifers seperated by an impermeable confining unit. A structured finite-difference grid with 100 rows and 100 columns was used. Each square cells is 470m by 470 m in extent. Initial heads are 10m in aquifer 1 and 30m in aquifer 2.
```
import os, shutil
import numpy as np
import matplotlib.pyplot as plt
import flopy
cln_ws= os.path.join('../data', 'usg_test')
if not os.path.exists(cln_ws):
os.mkdir(cln_ws)
```
## Loading Example 03_conduit_confined
The elevation of the top of layer 1 is -100 m, the bottom of layer 1 is -110 m, the top of layer 2 is -120 m and the bottom of layer 2 is -130 m. The confining unit between layers1 and 2 is impermeable and is represented using a quasi-three-dimensional approach. The only way a stress from one aquifer can be propagated to another is through the cln well that penetrates both aquifers.
The hydraulic conductivity values of the upper and lower aquifers are 100 and 400 m/d, respectively. Both aquifers have a primaey storage coefficient of 0.0001 and a specific yield of 0.01.
A vertical conduit well is located at the center of the domain and has a radius of 0.5 m. The well pumps 62,840 m3/d and is open fully to both aquifers from top to bottom. The CLN Process was used with a circular conduit geometry type to discretize the well bore with two conduit cells, one in each layer. The WEL Package was used to pump from the bottom CLN cell.
```
model_ws = os.path.join('../data/mfusg_test', '03_conduit_confined')
mf = flopy.mfusg.MfUsg.load(
'ex3.nam', model_ws=model_ws, exe_name="mfusg", check=False,verbose=True)
# output control
mf.remove_package('OC')
spd = {}
for i in range(mf.nper):
for j in range(mf.dis.nstp[i]):
spd[(i,j)] = ['save head', 'save budget']
oc = flopy.modflow.ModflowOc(mf, stress_period_data=spd, unitnumber=[22,30,31,50])
model_ws= os.path.join(cln_ws, 'ex03')
if os.path.exists(model_ws):
shutil.rmtree(model_ws)
os.mkdir(model_ws)
mf.model_ws=model_ws
mf.write_input()
mf.run_model()
head_file = os.path.join(mf.model_ws, 'ex3.clnhds')
headobj = flopy.utils.HeadFile(head_file)
simtimes = headobj.get_times()
nper=len(simtimes)
nnode= mf.cln.nclnnds
simhead = np.zeros((1,1,nnode,nper))
for i in range(nper):
simhead[:,:,:,i] = headobj.get_data(kstpkper = (i,0))
simhead=np.squeeze(simhead)
fig = plt.figure(figsize=(8, 5), dpi=150)
ax = fig.add_subplot(111)
ax.plot(simtimes, simhead[0], label='CLN Layer 1')
ax.plot(simtimes, simhead[1], label='CLN Layer 2')
ax.set_xlabel('Time, in days')
ax.set_ylabel('Simulated Head in pumping well, in meters')
ax.set_title("MODFLOW USG Ex3 Conduit Confined")
ax.legend()
cbb_file = os.path.join(mf.model_ws, 'ex3.clncbb')
cbb = flopy.utils.CellBudgetFile(cbb_file)
#cbb.list_records()
simflow = cbb.get_data(kstpkper=(0, 0), text='GWF')[0]
for i in range(nper-1):
simflow=np.append(simflow, cbb.get_data(kstpkper = (i+1,0), text = 'GWF')[0])
simflow1=simflow[simflow['node']==1]['q']
simflow2=simflow[simflow['node']==2]['q']
fig = plt.figure(figsize=(8, 11), dpi=150)
ax1 = fig.add_subplot(211)
ax1.plot(simtimes, simflow1, label='CLN Layer 1')
ax1.set_xlabel('Time, in days')
ax1.set_ylabel('Discharge to well, in cubic meters per day')
ax1.set_title("MODFLOW USG Ex3 Conduit Confined")
ax1.legend()
ax2 = fig.add_subplot(212)
ax2.plot(simtimes, simflow2, label='CLN Layer 2')
ax2.set_xlabel('Time, in days')
ax2.set_ylabel('Discharge to well, in cubic meters per day')
ax2.legend()
```
## Create example 03A_conduit_unconfined of mfusg 1.5
An unconfined example was simulated with this problem setup. The same simulation setup of previous example is used. However, the elevation of the top of layer 1 is 10 m, the bottom of layer 1 is 0 m, the top of layer 2 is -10 m, and the bottom of layer 2 is -20 m. The CLN domain is discretized using two vertical conduit cells. It depicts the behavior of unconfined flow in the conduit when the CLN cell in layer 1 becomes dry.
```
modelname='ex03a'
model_ws = os.path.join(cln_ws, modelname)
if os.path.exists(model_ws):
shutil.rmtree(model_ws)
os.mkdir(model_ws)
mf = flopy.mfusg.MfUsg(modelname=modelname, model_ws=model_ws,
exe_name="mfusg", verbose=True)
ipakcb=50
nlay = 2
nrow = 100
ncol = 100
laycbd = [1, 0]
delr = 470.000
delc = 470.000
ztop = 10.0
botm = [0.0, -10.0, -20.0]
perlen = 160
nstp = 160
dis = flopy.modflow.ModflowDis(mf, nlay, nrow, ncol, delr=delr, delc=delc,
laycbd=laycbd, top=ztop, botm=botm,
perlen=perlen, nstp=nstp, steady=False, lenuni=0)
bas = flopy.modflow.ModflowBas(mf, ibound=1, strt=[10.,30.])
bcf = flopy.mfusg.MfUsgBcf(mf, ipakcb=ipakcb, laycon=4, wetfct=1.0, iwetit=5,
hy=[100.0, 400.0], vcont=0.0, sf1=1e-4, sf2=0.01)
sms = flopy.mfusg.MfUsgSms(mf, hclose=1.0E-3, hiclose=1.0E-5, mxiter=220,
iter1=600, iprsms=1, nonlinmeth=2, linmeth=1,
theta=0.9, akappa=0.07, gamma=0.1, amomentum=0.0,
numtrack=200, btol=1.1, breduc=0.2, reslim=1.0,
iacl=2, norder=1, level=3, north=14)
# output control
spd = {}
for i in range(mf.nper):
for j in range(mf.dis.nstp[i]):
spd[(i,j)] = ['save head', 'save budget']
oc = flopy.modflow.ModflowOc(mf, stress_period_data=spd)
unitnumber = [71, 35, 36, 0, 0, 0, 0]
node_prop = [[1,1,0,10.0,0.0,1.57,0,0],[2,1,0,10.0,-20.0,1.57,0,0]]
cln_gwc = [[1,1,50,50,0,0,10.0,1.0,0],[2,2,50,50,0,0,10.0,1.0,0]]
nconduityp=1
cln_circ=[[1,0.5,3.23e10]]
strt = [10.0, 30.0]
cln = flopy.mfusg.MfUsgCln(mf, ncln=1, iclnnds=-1, nndcln=2, nclngwc = 2,
node_prop =node_prop, cln_gwc =cln_gwc,
cln_circ=cln_circ, strt =strt,
unitnumber=unitnumber)
options = []
options.append('autoflowreduce')
cln_stress_period_data = {0:[[1, -62840.]]}
wel = flopy.mfusg.MfUsgWel(mf, ipakcb=ipakcb,options=options,
cln_stress_period_data=cln_stress_period_data)
wel.cln_stress_period_data.data
mf.write_input()
mf.run_model()
head_file = os.path.join(mf.model_ws, modelname+'.clnhd')
headobj = flopy.utils.HeadFile(head_file)
simtimes = headobj.get_times()
nper=len(simtimes)
nnode= mf.cln.nclnnds
simhead = np.zeros((1,1,nnode,nper))
for i in range(nper):
simhead[:,:,:,i] = headobj.get_data(kstpkper = (i,0))
head_case1=np.squeeze(simhead)
fig = plt.figure(figsize=(8, 5), dpi=150)
ax = fig.add_subplot(111)
ax.plot(simtimes, head_case1[0], label='CLN Layer 1')
ax.plot(simtimes, head_case1[1], label='CLN Layer 2')
ax.set_xlabel('Time, in days')
ax.set_ylabel('Simulated Head in pumping well, in meters')
ax.set_title("MODFLOW USG Ex3a Conduit Unconfined")
ax.legend()
cbb_file = os.path.join(mf.model_ws, modelname+'.clncb')
cbb = flopy.utils.CellBudgetFile(cbb_file)
#cbb.list_records()
simflow = cbb.get_data(kstpkper=(0, 0), text='GWF')[0]
for i in range(nper-1):
simflow=np.append(simflow, cbb.get_data(kstpkper = (i+1,0), text = 'GWF')[0])
flow_case1=simflow
```
## Modify CLN amd WEL package to example create 03B_conduit_unconfined of mfusg 1.5
The problem is solved using only one CLN conduit cell to represent the well connecting both aquifer layers. This is conceptually equivalent to the MNW methodology and does not colve for flow within the well. The behavior of unconfined flow between the well and layer 1 negalects the dry-cell condition whereby head in the well is below the bottom of layer 1.
```
modelname='ex03b'
model_ws = os.path.join(cln_ws, modelname)
if os.path.exists(model_ws):
shutil.rmtree(model_ws)
os.mkdir(model_ws)
mf.model_ws = model_ws
mf._set_name(modelname)
for i, fname in enumerate(mf.output_fnames):
mf.output_fnames[i] = modelname + os.path.splitext(fname)[1]
mf.remove_package('CLN')
node_prop = [[1,1,0,30.0,-20.,1.57]]
cln_gwc = [[1,1,50,50,0,0,10.0,1.0,0],[1,2,50,50,0,0,10.0,1.0,0]]
strt = 20.0
cln = flopy.mfusg.MfUsgCln(mf, ncln=1, iclnnds=-1, nndcln=1, nclngwc = 2,
node_prop =node_prop, cln_gwc =cln_gwc,
cln_circ=cln_circ, strt =strt,
unitnumber =unitnumber)
mf.remove_package('WEL')
options = []
options.append('autoflowreduce')
options.append('iunitafr 55')
cln_stress_period_data = {0:[[0, -62840.]]}
wel = flopy.mfusg.MfUsgWel(mf, ipakcb=ipakcb,options=options,
cln_stress_period_data=cln_stress_period_data)
mf.write_input()
mf.run_model(silent=True)
head_file = os.path.join(mf.model_ws, modelname+'.clnhd')
headobj = flopy.utils.HeadFile(head_file)
simtimes = headobj.get_times()
nper=len(simtimes)
nnode= mf.cln.nclnnds
simhead = np.zeros((1,1,nnode,nper))
for i in range(nper):
simhead[:,:,:,i] = headobj.get_data(kstpkper = (i,0))
head_case2=np.squeeze(simhead)
fig = plt.figure(figsize=(8, 5), dpi=150)
ax = fig.add_subplot(111)
ax.plot(simtimes, head_case2)
ax.set_xlabel('Time, in days')
ax.set_ylabel('Simulated Head in pumping well, in meters')
ax.set_title("MODFLOW USG Ex3b Conduit Unconfined")
cbb_file = os.path.join(mf.model_ws, modelname+'.clncb')
cbb = flopy.utils.CellBudgetFile(cbb_file)
#cbb.list_records()
simflow = cbb.get_data(kstpkper=(0, 0), text='GWF')[0]
for i in range(nper-1):
simflow=np.append(simflow, cbb.get_data(kstpkper = (i+1,0), text = 'GWF')[0])
flow_case2=simflow
```
## Modify CLN amd WEL package to example create 03C_conduit_unconfined of mfusg 1.5
The CLN well is discretized using two conduit geometry CLN cells but with the confined option for flow winthin the conduit and between CLN and GWF domains.
```
modelname='ex03c'
model_ws = os.path.join(cln_ws, modelname)
if os.path.exists(model_ws):
shutil.rmtree(model_ws)
os.mkdir(model_ws)
mf.model_ws=model_ws
mf._set_name(modelname)
for i, fname in enumerate(mf.output_fnames):
mf.output_fnames[i] = modelname + os.path.splitext(fname)[1]
mf.remove_package('CLN')
node_prop = [[1,1,0,10.0,0.0,1.57,1,0],[2,1,0,10.0,-20.0,1.57,1,0]]
cln_gwc = [[1,1,50,50,0,0,10.0,1.0,0],[2,2,50,50,0,0,10.0,1.0,0]]
strt = [10.0, 30.0]
cln = flopy.mfusg.MfUsgCln(mf, ncln=1, iclnnds=-1, nndcln=2, nclngwc = 2,
node_prop =node_prop, cln_gwc =cln_gwc,
cln_circ=cln_circ, strt =strt,
unitnumber =unitnumber)
mf.remove_package('WEL')
cln_stress_period_data = {0:[[1, -62840.]]}
wel = flopy.mfusg.MfUsgWel(mf, ipakcb=ipakcb,options=options,
cln_stress_period_data=cln_stress_period_data)
mf.write_input()
mf.run_model(silent=True)
head_file = os.path.join(mf.model_ws, modelname+'.clnhd')
headobj = flopy.utils.HeadFile(head_file)
simtimes = headobj.get_times()
nper=len(simtimes)
nnode= mf.cln.nclnnds
simhead = np.zeros((1,1,nnode,nper))
for i in range(nper):
simhead[:,:,:,i] = headobj.get_data(kstpkper = (i,0))
head_case3=np.squeeze(simhead)
fig = plt.figure(figsize=(8, 5), dpi=150)
ax = fig.add_subplot(111)
ax.plot(simtimes, head_case3[0], label='CLN Layer 1')
ax.plot(simtimes, head_case3[1], label='CLN Layer 2')
ax.set_xlabel('Time, in days')
ax.set_ylabel('Simulated Head in pumping well, in meters')
ax.set_title("MODFLOW USG Ex3c Conduit Unconfined")
ax.legend()
cbb_file = os.path.join(mf.model_ws, modelname+'.clncb')
cbb = flopy.utils.CellBudgetFile(cbb_file)
#cbb.list_records()
simflow = cbb.get_data(kstpkper=(0, 0), text='GWF')[0]
for i in range(nper-1):
simflow=np.append(simflow, cbb.get_data(kstpkper = (i+1,0), text = 'GWF')[0])
flow_case3=simflow
```
## Modify CLN amd WEL package to example create 03D_conduit_unconfined of mfusg 1.5
Only one CLN cell to discretize the well but includes the "flow-to-dry-cell" option to limit flow in layer 1 when the head in the CLN cell is below the bottom of the layer.
```
modelname='ex03d'
model_ws = os.path.join(cln_ws, modelname)
if os.path.exists(model_ws):
shutil.rmtree(model_ws)
os.mkdir(model_ws)
mf.model_ws=model_ws
mf._set_name(modelname)
for i, fname in enumerate(mf.output_fnames):
mf.output_fnames[i] = modelname + os.path.splitext(fname)[1]
mf.remove_package('CLN')
node_prop = [[1,1,0,30.0,-20.,1.57]]
cln_gwc = [[1,1,50,50,0,0,10.0,1.0,1],[1,2,50,50,0,0,10.0,1.0,1]]
strt = 20.0
cln = flopy.mfusg.MfUsgCln(mf, ncln=1, iclnnds=-1, nndcln=1, nclngwc = 2,
node_prop =node_prop, cln_gwc =cln_gwc,
cln_circ=cln_circ, strt =strt,
unitnumber =unitnumber)
mf.remove_package('WEL')
cln_stress_period_data = {0:[[0, -62840.]]}
wel = flopy.mfusg.MfUsgWel(mf, ipakcb=ipakcb,options=options,
cln_stress_period_data=cln_stress_period_data)
mf.write_input()
mf.run_model(silent=True)
head_file = os.path.join(mf.model_ws, modelname+'.clnhd')
headobj = flopy.utils.HeadFile(head_file)
simtimes = headobj.get_times()
nper=len(simtimes)
nnode= mf.cln.nclnnds
simhead = np.zeros((1,1,nnode,nper))
for i in range(nper):
simhead[:,:,:,i] = headobj.get_data(kstpkper = (i,0))
head_case4=np.squeeze(simhead)
cbb_file = os.path.join(mf.model_ws, modelname+'.clncb')
cbb = flopy.utils.CellBudgetFile(cbb_file)
#cbb.list_records()
simflow = cbb.get_data(kstpkper=(0, 0), text='GWF')[0]
for i in range(nper-1):
simflow=np.append(simflow, cbb.get_data(kstpkper = (i+1,0), text = 'GWF')[0])
flow_case4=simflow
```
## Comparing four cases
```
fig = plt.figure(figsize=(8, 5), dpi=150)
ax = fig.add_subplot(111)
ax.plot(simtimes, head_case1[1], label='Case A')
ax.plot(simtimes, head_case2, label='Case B')
ax.plot(simtimes, head_case3[1], dashes=[6, 2],label='Case C')
ax.plot(simtimes, head_case4, dashes=[6, 2],label='Case D')
ax.set_xlabel('Time, in days')
ax.set_ylabel('Simulated Head in pumping well, in meters')
ax.legend()
fig = plt.figure(figsize=(8, 11), dpi=150)
ax1 = fig.add_subplot(211)
ax1.plot(simtimes, flow_case1[::2,]['q'], label='Case A')
ax1.plot(simtimes, flow_case2[::2,]['q'], label='Case B')
ax1.plot(simtimes, flow_case3[::2,]['q'], dashes=[6, 2], label='Case C')
ax1.plot(simtimes, flow_case4[::2,]['q'], dashes=[6, 2], label='Case D')
ax1.set_xlabel('Time, in days')
ax1.set_ylabel('Layer 1 flow to well')
ax1.legend()
ax2 = fig.add_subplot(212)
ax2.plot(simtimes, flow_case1[1::2,]['q'], label='Case A')
ax2.plot(simtimes, flow_case2[1::2,]['q'], label='Case B')
ax2.plot(simtimes, flow_case3[1::2,]['q'], dashes=[6, 2], label='Case C')
ax2.plot(simtimes, flow_case4[1::2,]['q'], dashes=[6, 2], label='Case D')
ax2.set_xlabel('Time, in days')
ax2.set_ylabel('Layer 2 flow to well')
ax2.legend()
```
| github_jupyter |
Examples require an initialized GsSession and relevant entitlements. External clients need to substitute thier own client id and client secret below. Please refer to [Authentication](https://developer.gs.com/p/docs/institutional/platform/authentication/) for details.
```
from gs_quant.session import GsSession
GsSession.use(client_id=None, client_secret=None, scopes=('read_product_data',))
```
## How to query data
The Data APIs support many ways to query datasets to intuitively fetch only the data users need.
More details on [Querying Data](https://developer.gs.com/p/docs/services/data/data-access/query-data/) can be found in the documentation
```
from datetime import date, timedelta, datetime
from gs_quant.data import Dataset
import pydash
```
Data in Marquee is available in the form of Datasets (collections of homogenous data). Each Dataset has a set of entitlements, a fixed schema, and assets in coverage.
```
dataset_id = 'FXIVOL_STANDARD' # https://marquee.gs.com/s/developer/datasets/FXIVOL_STANDARD
ds = Dataset(dataset_id)
```
Data for limited number of assets or spanning a small time frame can be queried in one go by specifying the assets to query and date/time range.
```
start_date = date(2019, 1, 15)
end_date = date(2019, 1, 18)
data = ds.get_data(start_date, end_date, bbid=['EURCAD'])
data.head()
```
Instead of a range, one can also specify a set of date/times to get data for just those specific date/times
```
data = ds.get_data(dates=[date(2019, 1, 15), date(2019, 1, 18)],
bbid=['EURCAD'])
data.head()
```
For larger number of assets or for longer time ranges,
we recommend iterating over assets and time to avoid hitting API query limits.
```
# loop over assets
def iterate_over_assets(dataset, coverage, start, end, batch_size=5, query_dimension='assetId', delta=timedelta(days=6)):
for ids in pydash.chunk(coverage[query_dimension].tolist(), size=batch_size):
print('iterate over assets', ids)
iterate_over_time(start, end, ids, dataset, delta=delta, query_dimension=query_dimension)
# loop over time
def iterate_over_time(start, end, ids, dataset, delta=timedelta(days=6), query_dimension='assetId'):
iter_start = start
while iter_start < end:
iter_end = min(iter_start + delta, end)
print('time iteration since', iter_start, 'until', iter_end)
data = dataset.get_data(iter_start, iter_end, **{query_dimension: ids})
# Add your code here to make use of fetched data
iter_start = iter_end
dataset_id = 'EDRVOL_PERCENT_STANDARD' # https://marquee.gs.com/s/developer/datasets/EDRVOL_PERCENT_STANDARD
ds = Dataset(dataset_id)
coverage = ds.get_coverage()
iterate_over_assets(ds, coverage, date(2021, 5, 1), date(2021, 5, 31), batch_size=5)
```
Similar approach can be used to download all data of a dataset
```
coverage = ds.get_coverage(include_history=True)
coverage = coverage.sort_values(by='historyStartDate', axis=0)
start_date = datetime.strptime(coverage['historyStartDate'].values[0], '%Y-%m-%d').date()
# warning: long running operation
iterate_over_assets(ds, coverage, start_date, date.today())
```
| github_jupyter |
```
import copy
import os, sys
os.chdir('..')
import numpy as np
import pandas as pd
import matplotlib
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from mpl_toolkits.axes_grid1 import make_axes_locatable
from scipy.stats.mstats import zscore
from sklearn.linear_model import LinearRegression
# loa my modules
from src.utils import load_pkl, unflatten
from src.visualise import *
from src.models import clean_confound
import joblib
import pickle
# Built-in modules #
import random
# Third party modules #
import numpy, scipy, matplotlib, pandas
from matplotlib import pyplot
import scipy.cluster.hierarchy as sch
import scipy.spatial.distance as dist
model_path = './models/SCCA_Yeo7nodes_revision_4_0.80_0.50.pkl'
label_path = './references/names.csv'
dat_path = './data/processed/dict_SCCA_data_prepro_revision1.pkl'
# load data
model = joblib.load(model_path)
dataset = load_pkl(dat_path)
df_label = pd.read_csv(label_path)
#df = pd.read_pickle(df_path)
u, v = model.u * [1, 1, -1, 1] , model.v * [1, 1, -1, 1]
n = model.n_components
# create labels for the nodes
seed_names = df_label.iloc[:, 0].apply(str) + '-' + df_label.iloc[:, -2] + '-' + df_label.iloc[:, -3] + ' ' + df_label.iloc[:, -1]
# unflatten the functional corr coeff
u_mat = []
for i in range(4):
u_mat.append(unflatten(u[:, i]))
FC_nodes = dataset['FC_nodes']
MRIQ = dataset['MRIQ']
mot = dataset['Motion_Jenkinson']
sex = dataset['Gender']
age = dataset['Age']
confound_raw = np.hstack((mot, sex, age))
X, Y, R = clean_confound(FC_nodes, MRIQ, confound_raw)
X_scores = zscore(X).dot(u)
Y_scores = zscore(Y).dot(v)
canpair_score = zscore(X_scores) + zscore(Y_scores)
model.cancorr_
np.corrcoef(X_scores, rowvar=False)
np.corrcoef(Y_scores, rowvar=False)
np.corrcoef(canpair_score, rowvar=False)
```
# variance inflation factor
```
def vif(X):
corr = np.corrcoef(X, rowvar=0, bias=True)
minv = np.linalg.inv(corr)
vif = minv.dot(corr).dot(minv)
vif = np.diag(vif)
return vif
print sum(vif(X) > 10)
from sklearn.decomposition import PCA
pca = PCA(n_components=200)
pca.fit(X)
pca.explained_variance_
```
# CC explained variance of the original data
```
def _Rsquare(X, P):
'''
calculate the coefficent of determination (R square):
the ratio of the explained variation to the total variation.
'''
lr = LinearRegression(fit_intercept=False)
lr.fit(P, X.T)
rec_ = lr.coef_.dot(P.T)
return 1 - (np.var(X - rec_) / np.var(X))
print 'The canonical vectors explained {0:.1f}% of the original connectivity data'.format(_Rsquare(X, u)*100)
print 'The canonical vectors explained {0:.1f}% of the original self-reports'.format(_Rsquare(Y, v)*100)
```
# Visualise the components
```
df_yeo7color = pd.read_csv('./references/yeo7_color.csv', index_col=0)
c_label = []
for l in df_label.iloc[:, -2].values:
cur_color = df_yeo7color[l].values
hex_c = '#%02x%02x%02x' % tuple(cur_color)
c_label.append(mpl.colors.to_rgb(hex_c))
plt.close()
for i in range(4):
set_text_size(8)
fig = plt.figure(figsize=(20, 20))
ax = fig.add_subplot(111)
max = np.abs(u_mat[i]).max()
m = ax.matshow(u_mat[i], vmax=max, vmin=-max, cmap='RdBu_r')
ax.set_xticks(np.arange(u_mat[i].shape[1]))
ax.set_yticks(np.arange(u_mat[i].shape[0]))
ax.set_xticklabels(seed_names, rotation='vertical')
ax.set_yticklabels(seed_names)
for xtick, color in zip(ax.get_xticklabels(), c_label):
xtick.set_color(color)
for ytick, color in zip(ax.get_yticklabels(), c_label):
ytick.set_color(color)
fig.colorbar(m)
plt.savefig("reports/plots/yeo7node_{}.png".format(i + 1), dpi=300, tight_layout=True)
```
# try to average the correlation coefficents
```
yeo7_names = ['VIS', 'S-M', 'VAN', 'DAN', 'LIM', 'FPN', 'DMN']
yeo7_fullnames = ['Visual', 'Somatomotor', 'VentralAttention', 'DorsalAttention', 'Limbic', 'Frontoparietal', 'Default']
import numpy as np
summary_mat = np.zeros((7, 7, 4))
for k in range(n):
df = pd.DataFrame(u_mat[k], columns=df_label.iloc[:, -2].values, index=df_label.iloc[:, -2].values)
for i, x in enumerate(yeo7_fullnames):
for j, y in enumerate(yeo7_fullnames):
mat = df.loc[x, y].values.mean()
summary_mat[i, j, k] = mat
from src.visualise import rank_labels
df_v = pd.DataFrame(v, index=dataset['MRIQ_labels'])
def sparse_row(seri_v):
vi, cur_v_labels = rank_labels(seri_v)
idx = np.isnan(vi).reshape((vi.shape[0]))
vi = vi[~idx]
vi = vi.reshape((vi.shape[0], 1))
cur_v_labels = np.array(cur_v_labels)[~idx]
return vi, cur_v_labels
u_max = np.abs(summary_mat).max()
v_max = np.abs(v).max()
set_text_size(11)
for i in range(n):
# thought probe
vi, cur_v_labels = sparse_row(df_v.iloc[:, i])
# between networks
mat = np.tril(summary_mat[..., i], 0)
mat[np.triu_indices(mat.shape[0], 0)] = np.nan
cur_df = pd.DataFrame(mat, columns=yeo7_names, index=yeo7_names)
# within networks
within_net = summary_mat[..., i].diagonal().reshape((7,1))
fig = plt.figure(figsize=(6, 2.5))
ax = fig.add_subplot(131)
t = ax.matshow(vi, vmax=v_max, vmin=-v_max, cmap='RdBu_r')
ax.set_xticks(np.arange(vi.shape[1]))
ax.set_yticks(np.arange(vi.shape[0]))
ax.set_xticklabels([' '])
ax.set_yticklabels(cur_v_labels)
ax.set_title('Thoughts', fontsize=16)
ax = fig.add_subplot(132)
m1 = ax.matshow(cur_df.values, vmax=u_max, vmin=-u_max, cmap='RdBu_r')
ax.set_xticks(np.arange(cur_df.shape[1]))
ax.set_yticks(np.arange(cur_df.shape[0]))
ax.set_xticklabels(yeo7_names, rotation=45)
ax.set_yticklabels(yeo7_names)
ax.set_title('Between', fontsize=16)
ax.set_frame_on(False)
ax.plot([-0.5, -0.5], [-0.5, 6.5], ls='-', c='.1')
ax.plot([-0.5, 6.5], [6.5, 6.5], ls='-', c='.1')
ax.xaxis.set_ticks_position('bottom')
ax = fig.add_subplot(133)
m2 = ax.matshow(within_net, vmax=u_max, vmin=-u_max, cmap='RdBu_r')
ax.set_xticks(np.arange(within_net.shape[1]))
ax.set_yticks(np.arange(within_net.shape[0]))
ax.set_xticklabels(' ')
ax.set_yticklabels(yeo7_names)
ax.set_title('Within', fontsize=16)
plt.tight_layout()
plt.savefig('./reports/plots/yeo7nodes_bwsummary_{}.png'.format(i + 1), dpi=300)
plt.show()
master = []
vmax = np.abs(v).max()
vmin = -vmax
for i in range(4):
rescale = (v[:,i] - vmin) / (vmax - vmin)
colors_hex = []
for c in cm.RdBu_r(rescale):
colors_hex.append(matplotlib.colors.to_hex(c))
master.append(colors_hex)
colors_hex = np.array(master).T
df_v_color = pd.DataFrame(colors_hex, index=dataset['MRIQ_labels'])
df_v_color.to_csv('./reports/plots/wordcloud_colors.csv')
df_v.to_csv('./reports/plots/v.csv')
# word cloud colorbar
set_text_size(10)
fig = plt.figure(figsize=(2, 0.7))
ax = fig.add_subplot(111)
cmap = cm.RdBu_r
norm = matplotlib.colors.Normalize(vmin=vmin, vmax=vmax)
cb1 = mpl.colorbar.ColorbarBase(ax, cmap=cmap,
norm=norm, orientation='horizontal')
plt.tight_layout()
plt.savefig('./reports/plots/wordcloud_cb.png', transparent=True, dpi=300)
plt.show()
# word cloud colorbar
set_text_size(10)
fig = plt.figure(figsize=(4, 0.8))
ax = fig.add_subplot(111)
cmap = cm.RdBu_r
norm = matplotlib.colors.Normalize(vmin=-u_max, vmax=u_max)
cb1 = mpl.colorbar.ColorbarBase(ax, cmap=cmap,
norm=norm, orientation='horizontal')
plt.tight_layout()
plt.savefig('./reports/plots/fc_cb.png', transparent=True, dpi=300)
plt.show()
```
# within network connectivity
```
for k in range(n):
df = pd.DataFrame(u_mat[k], columns=df_label.iloc[:, -2].values, index=df_label.iloc[:, -2].values)
for net in yeo7_names:
mat = df.loc[net, net].values
label_idx = df_label.loc[:, 'Yeo7'] == net
label_l = df_label.iloc[:, -3][label_idx]
label_r = df_label.iloc[:, -1][label_idx]
label = list((label_l + " - " +label_r).values)
set_text_size(8)
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(111)
max_val = np.abs(mat).max()
m = ax.matshow(mat, vmax=max_val, vmin=-max_val, cmap='RdBu_r')
ax.set_xticks(np.arange(mat.shape[1]))
ax.set_yticks(np.arange(mat.shape[0]))
ax.set_xticklabels(label, rotation='vertical')
ax.set_yticklabels(label)
# ax.set_frame_on(False)
# ax.plot([-0.5, -0.5], [-0.5, 6.5], ls='-', c='.1')
# ax.plot([-0.5, 6.5], [6.5, 6.5], ls='-', c='.1')
ax.xaxis.set_ticks_position('bottom')
ax.set_title('Component {} - {}'.format(k + 1, net))
fig.colorbar(m)
plt.savefig('./reports/plots/withinNetworks/com{}.png'.format(k + 1, net))
for k in range(n):
df = pd.DataFrame(u_mat[k], columns=df_label.iloc[:, -2].values, index=df_label.iloc[:, -2].values)
for net in yeo7_names:
mat = df.loc[net, net].values
label_idx = df_label.loc[:, 'Yeo7'] == net
label_l = df_label.iloc[:, -3][label_idx]
label_r = df_label.iloc[:, -1][label_idx]
label = list((label_l + " - " +label_r).values)
set_text_size(8)
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(111)
max_val = np.abs(mat).max()
m = ax.matshow(mat, vmax=max_val, vmin=-max_val, cmap='RdBu_r')
ax.set_xticks(np.arange(mat.shape[1]))
ax.set_yticks(np.arange(mat.shape[0]))
ax.set_xticklabels(label, rotation='vertical')
ax.set_yticklabels(label)
# ax.set_frame_on(False)
# ax.plot([-0.5, -0.5], [-0.5, 6.5], ls='-', c='.1')
# ax.plot([-0.5, 6.5], [6.5, 6.5], ls='-', c='.1')
ax.xaxis.set_ticks_position('bottom')
ax.set_title('Component {} - {}'.format(k + 1, net))
fig.colorbar(m)
plt.savefig('./reports/plots/withinNetworks/com{}_{}.png'.format(k + 1, net))
```
| github_jupyter |
# Networking, HTTP, web services
## ISO/OSI model

## TCP/IP model in relation to ISO

As application developers, we are interested in:
- Transport protocols (TCP or UPD) - implemented by OPERATING SYSTEM libraries and kernel
- Application protocols (like HTTP or HTTPs) - implemented by our APPLICATION
## TCP and UDP
- both are associated with IP address and port number
- UDP messages (also called datagrams) are "fire and forget". delivery of messages are not guaranteed
- TCP controls order of messages and deliverability (error checking)
Cases for using UDP:
- Gaming network code
- Telemetry data collection from thin (IoT) devices
- Gathering image frames from monitoring cameras
- Other cases when deliverability / order of messages is not critical, but performance is

## Addressing
IPv4 addresses:
- dotted decimal notation - denotes a single address
* 192.168.1.1
* 127.0.0.1
* 10.10.1.55
- prefix notation - denotes a group of addresses (subnetwork)
* 192.147.0.0/24
IPv6 addresses:
- hexadecimal notation
* 2001:db8:85a3:8d3:1319:8a2e:370:7348
- prefix notation
* 2001:db8:1234::/48
Port number:
- 16-bit unsigned number (0-65535).
Host names, like "google.com" are NOT IP addresses. They are resolved by an application-level protocol DNS.
Connection are ALWAYS made to ip ADDRESS (and port).
### Network interfaces
- a piece of hardware (can be virtual or emulated) that provides network communication
- network address (including IP addresses) belong to that interface
- single interface can have more than 1 addresses
- an address will always have a network interface where it belongs to.
- single machine can have multiple interfaces
### LOOPBACK INTERFACE (LOCALHOST)
- 127.0.0.1
- ::1
- localhost (can have more aliases, like localhost.localdomain)
a special address (and hostname) referencing to current machine.
Important that it's a separate _interface_.
One common error related to that is when running containers or virtual machines on your local PC, they cannot connect to the host machine by specifying "localhost" or "127.0.0.1" - that address will reference themselves instead of host.
### PORT NUMBERS
- 0 - 65535
- under 1024 - reserved ("well-known" or system ports) - do not use them for your application
- higher than 1024 - some ports are "registered" in IANA.
## SOCKETS
Abstraction of a "data tunnel" between network endpoints.
- server "listens" for accepting connections, client "connects" to remote address and port
- after communication is established, both sides can read from socket and send data to it.
- communication is bidirectional

Note that we did not specify what is actually sent and written. It's up to application and is denoted with _application-level protocol_.
more details:
- by default, data sent and received is "raw"
- reads can be "blocking" and "non-blocking"
- anothe important parameter for sockets is "timeout"
```
# in python:
# server:
import socket
s = socket.socket(socket.AF_INET, socket.SOCK_STREAM)
s.bind(("0.0.0.0", 9000))
s.listen(1)
while True:
conn, addr = s.accept()
print("Received connection from", addr)
conn.send(f"Hello {addr}\n".encode())
conn.close()
# client:
import socket
s = socket.socket()
s.connect(("127.0.0.1", 9000))
print(s.recv(1024))
s.close()
```
To be able to serve multiple connections at once, in python we can
- .accept() connections in loop
- open a new thread with connection handler for processing
But usually is better to use higher-level frameworks, as they tend to be more optimized. Operating systems have more mechanisms of controlling and checking communication state.
- whether new connections arrived
- whether new data appeared available to read, etc.
Usually as python programmers we dont need to go that deep. Raw socket programming with python is rare.
### Possible problems with sockets and troubleshooting
Debugging and troubleshooting tools:
- `ping`
- `traceroute`
- `netstat -p`
- `lsof -i` or `lsof -i -n`
- `telnet` command
- netcat (`nc`) command
possible problems:
- sockets in TIME_WAIT state - usually because of connection drops because of server exceptions
- number of open files / socket exceeded: check `ulimit -a`. default limit of open files is 1024.
- "blocking" connection - check socket timeouts. Very widespread problem. default is NO TIMEOUT.
- small packets not arriving at once (for 1-byte packets for example): use TCP_NODELAY flag
## NAME RESOLUTION (DNS)
from hostnames to IP addresses.
- socket.gethostbyname()
- socket.getaddrinfo()

### Important tools for name resolution troubleshooting
- ping
- nslookup (can point to specific name resolution server)
- hosts file (/etc/hosts, c:\Windows\System32\drivers\etc\hosts)
### Modern DNS
Due to privacy concerns a number of tools and standards are emerging related to name resolution
- DNS over TLS (DoT)
- DNS over HTTPS (DoH)
- DNSCrypt (prevents forgery, but still visible)
# HTTP
Application-level protocol for serving hypertext content.
## Time for demo!
## Basic request components
- outside of request itself
* server network address (host IP address and port)
* schema (https / http)
- inside http request:
* method (GET, POST, DELETE, OPTIONS, etc)
* request path
* request query string
* request headers
* request body
- inside http response:
* status code
* status message
* response headers
* response body
## Cookies
Responses sometimes contain header `set-cookie`. This information is stored in browser and later reused for subsequent request to the same website (or its part).
This is the main identification mechanism that is implemented in internet.
```
# in python:
import requests
result = requests.get('https://google.com').text
```
## content-types and encodings
for responses:
- text/plain
- text/html
- binary/octet-stream
- application/json
- image/jpeg
- many others
for requests:
- multipart/form-data
- application/x-www-form-urlencoded
- application/json
# SSL (TLS) and HTTPs
- HTTPS is a transport layer wrapper ON TOP of HTTP
- TLS (SSL) can be generally used not only for HTTP, but for any other socket-based communication.
- Using PKI concepts and infrastructure
Steps:
- Server acquires _certificate_ from CERTIFICATE AUTHORITY (CA)
- Server certificate is _*SERVER PUBLIC KEY SIGNED BY CA*_ with additional information (CN = server name)
- Client already has ALL CA certificates (ROOT CAs). (provided by OS or separate package. In python: certifi).
- When connection is established, server supplies its certificate. Client checks that server name matches to certificate's CN and verifies that server certificate is valid.
- Client and server negotiate a session-level encryption protocol, generate symmetric session encryption key
- All further communication between server and client is made by using negotiated session key
### HTTPS in python:
Most frameworks does NOT support HTTPs directly (and is actually discouraged to use certificate with applications directly).
Usually web frameworks will receive unwrapped HTTP requests.
Process of unwrapping SSL to underlying protocol is called SSL termination.
SSL is usually performed at generic webserver (nginx) or load balancing level (gunicorn, haproxy, or container orchestration framework). Reasons: multiple HOSTs at singe webserver, load balancing, centralized webserver log collection, DDOS prevention, etc.
### Typical webservice stack for python webserver frameworks
- load balancer (haproxy)
- generic webserver (NGINX, apache, lighthttpd). Usually it also server static files.
- python fastcgi / http server (uwsgi, gunicorn) that preforks python application workers
- python web application processes
### CGI, FASTCGI, WSGI
Represent an evolution of web servers
- Static pages
- Dynamic pages ( CGI )
- mod_python - embeds python into webserver code to run python applications (almost the same as CGI)
- Dedicated processes serving dynamic content on-demand (fastcgi)
- WSGI - python-specific interface standard similar to fastcgi ( PEP 3333 )
- Modern async frameworks handle HTTP requests themselves
EXAMPLE OF CGI environment variables (from CGI wiki page). Some are common and defined by OS, some are set by server
```
COMSPEC="C:\Windows\system32\cmd.exe"
DOCUMENT_ROOT="C:/Program Files (x86)/Apache Software Foundation/Apache2.4/htdocs"
GATEWAY_INTERFACE="CGI/1.1"
HOME="/home/SYSTEM"
HTTP_ACCEPT="text/html,application/xhtml+xml,application/xml;q=0.9,*/*;q=0.8"
HTTP_ACCEPT_CHARSET="ISO-8859-1,utf-8;q=0.7,*;q=0.7"
HTTP_ACCEPT_ENCODING="gzip, deflate, br"
HTTP_ACCEPT_LANGUAGE="en-us,en;q=0.5"
HTTP_CONNECTION="keep-alive"
HTTP_HOST="example.com"
HTTP_USER_AGENT="Mozilla/5.0 (Windows NT 6.1; WOW64; rv:67.0) Gecko/20100101 Firefox/67.0"
PATH="/home/SYSTEM/bin:/bin:/cygdrive/c/progra~2/php:/cygdrive/c/windows/system32:..."
PATHEXT=".COM;.EXE;.BAT;.CMD;.VBS;.VBE;.JS;.JSE;.WSF;.WSH;.MSC"
PATH_INFO="/foo/bar"
PATH_TRANSLATED="C:\Program Files (x86)\Apache Software Foundation\Apache2.4\htdocs\foo\bar"
QUERY_STRING="var1=value1&var2=with%20percent%20encoding"
REMOTE_ADDR="127.0.0.1"
REMOTE_PORT="63555"
REQUEST_METHOD="GET"
REQUEST_URI="/cgi-bin/printenv.pl/foo/bar?var1=value1&var2=with%20percent%20encoding"
SCRIPT_FILENAME="C:/Program Files (x86)/Apache Software Foundation/Apache2.4/cgi-bin/printenv.pl"
SCRIPT_NAME="/cgi-bin/printenv.pl"
SERVER_ADDR="127.0.0.1"
SERVER_ADMIN="(server admin's email address)"
SERVER_NAME="127.0.0.1"
SERVER_PORT="80"
SERVER_PROTOCOL="HTTP/1.1"
SERVER_SIGNATURE=""
SERVER_SOFTWARE="Apache/2.4.39 (Win32) PHP/7.3.7"
SYSTEMROOT="C:\Windows"
TERM="cygwin"
WINDIR="C:\Windows"
```
```
# WSGI application in python is just a callable with 2 positional parameters.
def web_app(env, start_response):
print(env)
status = '200 OK'
response_headers = [('Content-type', 'text/plain')]
start_response(status, response_headers)
return [b"Welcome to the machine\n"]
class WebApp:
def __init__(self, env, start_response):
print(env)
self.env = env
self.callback = start_response
def __iter__(self):
status = '200 OK'
response_headers = [('Content-type', 'text/plain')]
self.callback(status, response_headers)
return iter([b"Have a cigar\n"])
```
WSGI environement variables:
```
{
'wsgi.errors': <gunicorn.http.wsgi.WSGIErrorsWrapper object at 0x7f2734f35a60>,
'wsgi.version': (1, 0),
'wsgi.multithread': False,
'wsgi.multiprocess': False,
'wsgi.run_once': False,
'wsgi.file_wrapper': <class 'gunicorn.http.wsgi.FileWrapper'>,
'wsgi.input_terminated': True,
'SERVER_SOFTWARE': 'gunicorn/20.0.4',
'wsgi.input': <gunicorn.http.body.Body object at 0x7f2734f35f70>,
'gunicorn.socket':
<socket.socket fd=12, family=AddressFamily.AF_INET, type=SocketKind.SOCK_STREAM, proto=0, laddr=('127.0.0.1', 8000), raddr=('127.0.0.1', 55766)>,
'REQUEST_METHOD': 'GET',
'QUERY_STRING': '',
'RAW_URI': '/test',
'SERVER_PROTOCOL': 'HTTP/1.1',
'HTTP_HOST': '127.0.0.1:8000',
'HTTP_USER_AGENT': 'curl/7.68.0',
'HTTP_ACCEPT': '*/*',
'wsgi.url_scheme': 'http',
'REMOTE_ADDR': '127.0.0.1',
'REMOTE_PORT': '55766',
'SERVER_NAME': '127.0.0.1',
'SERVER_PORT': '8000',
'PATH_INFO': '/test',
'SCRIPT_NAME': ''
}
```
Note that all this data is pretty basic, you will still need to parse query string yourself, and even request headers are included _inside_ request body.
# WEBSERVER FRAMEWORKS
- Prepare request data for consumption
- Organize your request handling code in structured way
- Supply additional batteries to shorten and reuse common request-related tasks
popular frameworks:
- synchronous (django, pyramid, flask, bottle, falcon, etc.)
- asynchronous (aiohttp, fastapi, sanic, tornado, etc)
And MANY MANY different standalone libraries that supply specific pluggable functionality for each specific task.
Examples: marshmallow, sqlalchemy, itsdangerous, deform, jinja, genshi, many others.
## Anatomy of web framework
- request parser
- routing
- template engine
- modularity and code organization
- data validation and XSS attack prevention
- session handling
- configuration management
- built-in ORM
ORM + request handler + template engine combination implements paradigm of MVC (model-view-controller).
MVC frameworks usually aim to be a generic solution for classic single-server web applications.
At the other end of spectrum we have microframeworks as bottle and frameworks tailored for specific tasks (like microservices: vivid example would be fastapi that is specifically designed to be a API framework, or DRF, that is tailored to be REST services provider).
```
from bottle import route, request, default_app
@route('/')
def index():
name = request.query.get('name', 'anonymous')
return f'Hello, {name}!' # do not do things like that in production, XSS!
# Use templating engine with safeguards
app = default_app() # you can create application object explicitly and add routes on top of it
```
## Useful tools
- CURL (the best tool for web request analysis)
- python and requests library
- web browser in developer mode.
- various http bins (be wary about passwords and sensitive data though!, better to self-host them)
- load testing: ab (Apache bench), gobench, locust, yandex-tank
# THE END
| github_jupyter |
```
import datetime
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
```
To be able to process locally, we will use 1% of data. After that, we still have a large number of 241,179 records.
```
df = pd.read_csv('train.csv.gz', sep=',').dropna()
df = df.sample(frac=0.01, random_state=99)
df.shape
df.head()
count_classes = pd.value_counts(df['is_booking'], sort = True).sort_index()
count_classes.plot(kind = 'bar')
plt.title("Booking or Not booking")
plt.xlabel("Class")
plt.ylabel("Frequency")
```
It is obvious that our data is very imbalanced. We will have to deal with it.
Data quality check
```
df.isnull().sum()
df.dtypes
```
### Feature engineering
The process includes create new columns such as year, month, plan time and hotel nights. And remove the columns we do not need anymore afterwards.
```
df["date_time"] = pd.to_datetime(df["date_time"])
df["year"] = df["date_time"].dt.year
df["month"] = df["date_time"].dt.month
df['srch_ci']=pd.to_datetime(df['srch_ci'],infer_datetime_format = True,errors='coerce')
df['srch_co']=pd.to_datetime(df['srch_co'],infer_datetime_format = True,errors='coerce')
df['plan_time'] = ((df['srch_ci']-df['date_time'])/np.timedelta64(1,'D')).astype(float)
df['hotel_nights']=((df['srch_co']-df['srch_ci'])/np.timedelta64(1,'D')).astype(float)
cols_to_drop = ['date_time', 'srch_ci', 'srch_co', 'user_id']
df.drop(cols_to_drop, axis=1, inplace=True)
df.head()
```
Plot a correlation matrix using a heatmap to explore the correlation between features. Nothing really exciting here.
```
correlation = df.corr()
plt.figure(figsize=(18, 18))
sns.heatmap(correlation, vmax=1, square=True,annot=True,cmap='viridis')
plt.title('Correlation between different fearures')
```
Look at the correlation of each column compared to the other one. We do not see any two variables are very closely correlated.
### Dealing with imbalanced data
```
booking_indices = df[df.is_booking == 1].index
random_indices = np.random.choice(booking_indices, len(df.loc[df.is_booking == 1]), replace=False)
booking_sample = df.loc[random_indices]
not_booking = df[df.is_booking == 0].index
random_indices = np.random.choice(not_booking, sum(df['is_booking']), replace=False)
not_booking_sample = df.loc[random_indices]
df_new = pd.concat([not_booking_sample, booking_sample], axis=0)
print("Percentage of not booking clicks: ", len(df_new[df_new.is_booking == 0])/len(df_new))
print("Percentage of booking clicks: ", len(df_new[df_new.is_booking == 1])/len(df_new))
print("Total number of records in resampled data: ", len(df_new))
```
Shuffle the resampled dataframe.
```
df_new = df_new.sample(frac=1).reset_index(drop=True)
df_new.shape
```
Assign features and label from the new dataframe.
```
X = df_new.loc[:, df_new.columns != 'is_booking']
y = df_new.loc[:, df_new.columns == 'is_booking']
X.head()
```
## PCA
Principal component analysis, or PCA, is a statistical technique to convert high dimensional data to low dimensional data by selecting the most important features that capture maximum information about the dataset.
Standardize the dataset
```
scaler = StandardScaler()
X=scaler.fit_transform(X)
X
```
Apply PCA. And we have 23 features in our data.
```
pca = PCA(n_components=23)
pca.fit(X)
```
Calculate Eigenvalues
```
var=np.cumsum(np.round(pca.explained_variance_ratio_, decimals=3)*100)
var
```
In the above array we see that the first feature explains 9.3% of the variance within our data set while the first two explain 17.3% and so on. If we employ all features we capture 99.9% of the variance within the dataset, thus we gain some by implementing an additional feature. No any single feature outstanding.
Step 4: Sort & Select
```
plt.ylabel('% Variance Explained')
plt.xlabel('# of Features')
plt.title('PCA Analysis')
plt.style.context('seaborn-whitegrid')
plt.plot(var)
```
Based on the plot above it's clear we should keep all 23 features.
### Train, predict and performance evaluation
```
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=1)
X_train.shape
pca = PCA()
X_train = pca.fit_transform(X_train)
X_test = pca.transform(X_test)
classifier = RandomForestClassifier(max_depth=2, random_state=0)
classifier.fit(X_train, y_train)
y_pred = classifier.predict(X_test)
cm = confusion_matrix(y_test, y_pred)
print(cm)
print('Accuracy', accuracy_score(y_test, y_pred))
```
### Logistic Regression
```
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import Pipeline
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=1)
pca = PCA(n_components=23)
logReg = LogisticRegression()
pipe = Pipeline([('pca', pca), ('logistic', logReg)])
pipe.fit(X_train, y_train)
y_pred = pipe.predict(X_test)
cm = confusion_matrix(y_test, y_pred)
print(cm)
print('Accuracy', accuracy_score(y_test, y_pred))
```
| github_jupyter |
```
%matplotlib inline
import matplotlib.cm as cm
import matplotlib.pyplot as plt
import numpy as np
from mpl_toolkits.mplot3d import Axes3D
from numpy import linalg
from sklearn.datasets import load_iris
from sklearn.cluster import KMeans
from sklearn.metrics import accuracy_score, silhouette_samples, silhouette_score
```
---
## [1] DATA
```
iris = load_iris()
X = iris.data
y = iris.target
iris.target_names
iris.feature_names
```
---
## [2] Plot Iris in 3D
```
fig = plt.figure(1, figsize=(8, 6))
ax = Axes3D(fig, elev=-160, azim=130)
ax.scatter(X[:, 0], X[:, 1], X[:, 2], c=y, cmap=plt.cm.Paired)
ax.set_title("Iris Dataset in 3D")
ax.set_xlabel("sepal length")
ax.w_xaxis.set_ticklabels([])
ax.set_ylabel("sepal width")
ax.w_yaxis.set_ticklabels([])
ax.set_zlabel("petal length")
ax.w_zaxis.set_ticklabels([])
plt.show()
```
---
## [3] K-means
We happen to know there are 3 clusters because we have ground truth labels. However, with true unsupervised learning we don't have labels so finding the "best" number of clusters is context-driven and often challenging.
```
kmeans = KMeans(n_clusters=3, random_state=42, n_jobs=-1)
kmeans.fit(X)
fig = plt.figure(1, figsize=(8, 6))
ax = Axes3D(fig, elev=-160, azim=130)
ax.scatter(X[:, 0], X[:, 1], X[:, 2], c=kmeans.labels_, cmap=plt.cm.Paired)
ax.set_title("Iris Dataset in 3D (kmeans)")
ax.set_xlabel("sepal length")
ax.w_xaxis.set_ticklabels([])
ax.set_ylabel("sepal width")
ax.w_yaxis.set_ticklabels([])
ax.set_zlabel("petal length")
ax.w_zaxis.set_ticklabels([])
plt.show()
```
## [4] Are My Clusters Any Good?
In the case where I don't actually have a priori knowledge about the number of clusters, how do I know how many clusters to choose? There are a number of methods to address this challenge.
### Method 1: Silhouette Plot
From sklearn:
Silhouette analysis can be used to study the separation distance between the resulting clusters. The silhouette plot displays a measure of how close each point in one cluster is to points in the neighboring clusters and thus provides a way to assess parameters like number of clusters visually. This measure has a range of [-1, 1].
Silhouette coefficients (as these values are referred to as) near +1 indicate that the sample is far away from the neighboring clusters. A value of 0 indicates that the sample is on or very close to the decision boundary between two neighboring clusters and negative values indicate that those samples might have been assigned to the wrong cluster.
$s_i = \frac{b(i)-a(i)}{max(a_i, b_i)}$
```
n_clusters = 3
y_lower = 10
cluster_labels = kmeans.fit_predict(X)
silhouette_avg = silhouette_score(X, cluster_labels)
silhouette_values = silhouette_samples(X, cluster_labels)
fig = plt.figure(1, figsize=(8, 6))
ax = fig.add_subplot(1, 1, 1)
for i in range(n_clusters):
# Aggregate the silhouette scores for samples belonging to cluster i and sort them
ith_cluster_silhouette_values = silhouette_values[cluster_labels == i]
ith_cluster_silhouette_values.sort()
size_cluster_i = ith_cluster_silhouette_values.shape[0]
y_upper = y_lower + size_cluster_i
color = cm.spectral(float(i) / n_clusters)
ax.fill_betweenx(np.arange(y_lower, y_upper),
0, ith_cluster_silhouette_values,
facecolor=color, edgecolor=color, alpha=0.7)
# Label the silhouette plots with their cluster numbers at the middle
ax.text(-0.05, y_lower + 0.5 * size_cluster_i, str(i))
# Compute the new y_lower for next plot
y_lower = y_upper + 10 # 10 for the 0 samples
ax.set_title("Silhouette Plot")
ax.set_xlabel("Silhouette Coefficient Values")
ax.set_ylabel("Cluster label")
# The vertical line for average silhouette score of all the values
ax.axvline(x=silhouette_avg, color="red", linestyle="--")
ax.set_yticks([]) # Clear the yaxis labels / ticks
#ax.set_xticks([-0.1, 0, 0.1, 0.2, 0.3, 0.4, 0.6, 0.8, 1]);
ax.set_xticks(np.arange(-0.1, 1, 0.1));
```
---
Pretend we didn't know how many clusters to choose. How would we approach it? Answer: for loop + avg(silhouette score)!
```
for i in range(2, 11):
kmeans = KMeans(n_clusters=i, random_state=42, n_jobs=-1)
cluster_labels = kmeans.fit_predict(X)
silhouette_avg = silhouette_score(X, cluster_labels)
print(i, silhouette_avg)
type(X)
def optimal_clusterer(X, max_clusters=10):
'''
Iterative K-means clustering that finds optimal # of clusters based on silhouette score.
Parameters
----------
X: numpy ndarray
data to cluster on
max_clusters: int, optional, default: 10
max number of clusters to try
Attributes
----------
best number of clusters
silhouette score
'''
score = []
for i in range(2, max_clusters+1):
kmeans = KMeans(n_clusters=i, random_state=42, n_jobs=-1)
cluster_labels = kmeans.fit_predict(X)
score.append(silhouette_score(X, cluster_labels))
return np.argmax(score)+2, round(max(score), 4)
optimal_clusterer(X)
```
| github_jupyter |
# Density of States 3D Example
This notebook computes the density of states for the same example gyroid system shown in the original Scheme/MATLAB implementation.
The parameters used for computing the density of states and plotting variables are chosen to match the example plot in the original implementation.
This notebook imports material properties from `material.py`, integrator routines from `integrators.py`, and utility functions/classes from `util.py`, so those files must be contained in the same directory.
Please refer to those files for docstrings and citation information where applicable.
Both the tetrahedron (Tr) and Generalized Gilat-Raubenheimer (GGR) methods are used to determine the density of states.
```
import os
from contextlib import redirect_stdout
from functools import partial
import ase
import matplotlib.pyplot as plt
import meep
import numpy as np
from meep import mpb
from tqdm import tqdm
from tqdm.contrib import tzip
from tqdm.contrib.concurrent import process_map
from integrators import ggr_integration, tetrahedron_integration
from material import (
default_material,
eps_func,
gyroid_material,
k_point_band_path,
k_point_interpolation,
k_point_labels,
k_point_sequence,
lattice,
)
from util import TqdmWrappedIterable, _array_to_vec3s
```
Choose parameters for MPB and evaluate the bands over the k-path defined in `material.py`.
```
resolution = 16
mesh_size = 2
num_bands = 20
k_path_solver = mpb.ModeSolver(
resolution=resolution,
mesh_size=mesh_size,
num_bands=num_bands,
k_points=TqdmWrappedIterable(k_point_band_path),
default_material=default_material,
geometry_lattice=lattice,
)
k_path_solver.verbose = False
with open("mpb_output_band_path.txt", "w") as mpb_out, redirect_stdout(mpb_out):
k_path_solver.run()
```
Plot the band structure.
```
plt.figure(figsize=(8, 5), dpi=100)
labels_x = np.arange(len(k_point_labels)) * k_point_interpolation
plt.xticks(ticks=labels_x, labels=k_point_labels)
for x in labels_x:
plt.axvline(x, color="#bbb")
for i in range(k_path_solver.all_freqs.shape[1]):
plt.plot(k_path_solver.all_freqs[:, i])
plt.title("Band structure")
plt.ylabel("Frequency $\omega a$/(2$\pi$c)")
plt.ylim(0, np.max(k_path_solver.all_freqs) * 1.05)
plt.xlim(0, np.max(labels_x))
plt.savefig("band_structure.png")
plt.show()
```
Determine k-points for evaluating the density of states.
```
num_points = (10, 10, 10)
for x in num_points:
assert x % 2 == 0
k_points = _array_to_vec3s(ase.dft.kpoints.monkhorst_pack(num_points))
```
Here, we provide `append_velocities` as a "band function." Band functions must be callables with only one argument, the `ModeSolver` instance. In an ideal world, I would prefer to use a method of a class, or even a callable class instance, but MPB does some kind of Python introspection on the provided function that prevents those approaches from working. Instead, we create a function `append_velocities` that binds a variable `velocities` from outside the function scope and appends the group velocities for each k-point as the `ModeSolver` evaluates them. Finally, we use `velocities_as_array` to convert the group velocites into a NumPy array.
See the MPB docs for additional information about running with a band function: https://mpb.readthedocs.io/en/latest/Python_User_Interface/#run-functions
```
def append_velocities(solver):
velocities.append(solver.compute_group_velocities())
def velocities_as_array():
return np.asarray([[[v[0], v[1], v[2]] for v in band] for band in velocities])
solver = mpb.ModeSolver(
resolution=resolution,
mesh_size=mesh_size,
num_bands=num_bands,
k_points=TqdmWrappedIterable(k_points),
default_material=default_material,
geometry_lattice=lattice,
)
solver.verbose = False
velocities = []
with open("mpb_output.txt", "w") as mpb_out, redirect_stdout(mpb_out):
solver.run(append_velocities)
velocities = velocities_as_array()
```
The tetrahedron integration method performs best if the tetrahedra are formed by splitting along the shortest cell diagonal. This cell arranges the frequencies in an order with the shortest diagonal.
```
reciprocal_vectors = np.array([[0, 1, 1], [1, 0, 1], [1, 1, 0]])
diagonal_lengths = np.array(
[
np.linalg.norm(reciprocal_vectors @ [1, 1, 1]),
np.linalg.norm(reciprocal_vectors @ [-1, 1, 1]),
np.linalg.norm(reciprocal_vectors @ [1, -1, 1]),
np.linalg.norm(reciprocal_vectors @ [1, 1, -1]),
]
)
min_diagonal_length = np.argmin(diagonal_lengths)
w_grid = solver.all_freqs.reshape(num_points + (num_bands,))
w_grid = np.flip(w_grid, axis=min_diagonal_length)
```
Perform the tetrahedron integration.
```
w_bins_Tr, DOS_Tr = tetrahedron_integration(
num_points, num_bands, w_grid, w_min=0, w_max=None, num_bins=20000
)
```
Perform the Generalized Gilat-Raubenheimer integration.
```
w_bins_ggr, DOS_ggr = ggr_integration(
num_points, solver.all_freqs, velocities, num_bins=20000
)
```
Plot the densities of states obtained from each integration method.
```
plt.figure(figsize=(8, 5), dpi=100)
plt.plot(DOS_ggr, w_bins_ggr, label="GGR")
plt.plot(DOS_Tr, w_bins_Tr, label="Tr")
plt.xlim(0, 100)
plt.xlabel("per cell")
plt.ylabel("Frequency $\omega a$/(2$\pi$c)")
plt.legend()
plt.savefig("density_of_states.png")
plt.show()
```
Plot the band structure and GGR density of states.
```
plt.figure(figsize=(8, 5), dpi=150)
labels_x = np.arange(len(k_point_labels)) * k_point_interpolation
plt.xticks(ticks=labels_x, labels=k_point_labels)
for x in labels_x:
plt.axvline(x, color="#bbb")
for i in range(k_path_solver.all_freqs.shape[1]):
plt.plot(k_path_solver.all_freqs[:, i])
x_start_dos = (len(k_point_labels) - 1) * k_point_interpolation
dos_width = 2 * k_point_interpolation / 100
plt.gca().fill_betweenx(w_bins_ggr, x_start_dos, x_start_dos + DOS_ggr * dos_width)
plt.xlim(0, x_start_dos + dos_width * 100)
plt.title("Band structure and Density of States D2$\pi$c/a")
plt.ylabel("Frequency $\omega a$/(2$\pi$c)")
plt.ylim(0, np.max(k_path_solver.all_freqs) * 0.95)
plt.savefig("band_structure_and_density_of_states.png")
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/lmoroney/dlaicourse/blob/master/TensorFlow%20In%20Practice/Course%204%20-%20S%2BP/S%2BP%20Week%202%20Lesson%202.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
RunningInCOLAB = 'google.colab' in str(get_ipython())
RunningInCOLAB
if RunningInCOLAB:
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
else:
import tensorflow as tf
tf_version = tf.__version__
print(tf_version)
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
def trend(time, slope=0):
return slope * time
time = np.arange(4 * 365 + 1, dtype="float32")
baseline = 10
series = trend(time, 0.1)
series
def plot_series(time, series, format="-", start=0, end=None):
plt.plot(time[start:end], series[start:end], format)
plt.xlabel("Time")
plt.ylabel("Value")
plt.grid(True)
plot_series(time,series)
amplitude = 50
slope = 0.05
noise_level = 10
def seasonal_pattern(season_time):
return np.where(season_time < 0.4,
#np.cos(season_time * 2 * np.pi),
np.sin(season_time * 2),
1 / np.exp(3 * season_time))
def seasonality(time, period, amplitude=1, phase=0):
season_time = ((time + phase) % period) / period
return amplitude * seasonal_pattern(season_time)
def noise(time, noise_level=1, seed=None):
rnd = np.random.RandomState(seed)
return rnd.randn(len(time)) * noise_level
series = baseline + trend(time, slope)
plot_series(time,series)
series = baseline + trend(time, slope) + seasonality(time, period=365, amplitude=amplitude)
plot_series(time,series)
series += noise(time, noise_level, seed=42)
plot_series(time,series)
series
split_time = 1000
time_train = time[:split_time]
x_train = series[:split_time]
time_valid = time[split_time:]
x_valid = series[split_time:]
window_size = 20
batch_size = 32
shuffle_buffer_size = 1000
def windowed_dataset(series, window_size, batch_size, shuffle_buffer):
dataset = tf.data.Dataset.from_tensor_slices(series)
dataset = dataset.window(window_size + 1, shift=1, drop_remainder=True)
dataset = dataset.flat_map(lambda window: window.batch(window_size + 1))
dataset = dataset.shuffle(shuffle_buffer).map(lambda window: (window[:-1], window[-1]))
dataset = dataset.batch(batch_size).prefetch(1)
return dataset
dataset = windowed_dataset(x_train, window_size, batch_size, shuffle_buffer_size)
print(dataset)
layer1 = tf.keras.layers.Dense(1, input_shape=[window_size])
model = tf.keras.models.Sequential([layer1])
model.compile(loss="mse", optimizer=tf.keras.optimizers.SGD(lr=1e-6, momentum=0.9))
model.fit(dataset,epochs=100,verbose=0)
print("Layer weights {}".format(layer1.get_weights()))
forecast = []
for time in range(len(series) - window_size):
forecast.append(model.predict(series[time:time + window_size][np.newaxis]))
forecast = forecast[split_time-window_size:]
results = np.array(forecast)[:, 0, 0]
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, results)
tf.keras.metrics.mean_absolute_error(x_valid, results).numpy()
tf.keras.metrics.mean_squared_error(x_valid, results).numpy()
```
| github_jupyter |
```
import pickle
import subprocess
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import make_axes_locatable
matplotlib.rc('pdf', fonttype=42) # Enable pdf compatible fonts
import scipy.stats
from sklearn.neighbors import KernelDensity
from sklearn.mixture import GaussianMixture
import Bio.SeqIO
# Sample information
ortholog = 'Cas13bt-3'
samples = ['Bt3_BR1_S4_R1_001spacers.p', 'Bt3_BR2_S5_R1_001spacers.p', 'Bt3_BR3_S6_R1_001spacers.p', 'Bt3ctrl_BR1_S1_R1_001spacers.p', 'Bt3ctrl_BR2_S2_R1_001spacers.p', 'Bt3ctrl_BR3_S3_R1_001spacers.p', 'Bt3_input_S7_R1_001spacers.p']
samples_to_names = {samples[0] : 'Cas13bt3 Experiment Rep 1',
samples[1] : 'Cas13bt3 Experiment Rep 2',
samples[2] : 'Cas13bt3 Experiment Rep 3',
samples[3] : 'Cas13bt3 Control Rep 1',
samples[4] : 'Cas13bt3 Control Rep 2',
samples[5] : 'Cas13bt3 Control Rep 3',
samples[6] : 'Cas13bt3 Input Library',
}
# Map sample ids tosample names and filepaths
sample_pair = (samples[0], samples[2])
pair_names = [samples_to_names[sample_pair[0]], samples_to_names[sample_pair[1]]]
# Load non-targeting spacers list
nt_spacers = []
with open('nt_spacers.csv', 'r') as f:
for line in f:
nt_spacers.append(line.strip())
# Obtain the experiment condition sample information
e_N_avg = {}
e_Ns = []
es = []
for e_name in samples[0:3]:
e = pickle.load(open(e_name, 'rb'), encoding='latin1')
# Get sum of all read counts
e_sum = sum([v for v in e.values()])
# Normalize individual spacer count by sum of all read counts in sample
e_N = {u : float(v)/e_sum for u,v in e.items()}
e_Ns.append(e_N)
es.append(e)
for u in e_Ns[0]:
e_N_avg[u] = ((e_Ns[0][u], e_Ns[1][u], e_Ns[2][u]), (es[0][u], es[1][u], es[2][u]))
# Obtain the control condition sample information
c_N_avg = {}
c_Ns = []
cs = []
for c_name in samples[3:6]:
c = pickle.load(open(c_name, 'rb'), encoding='latin1')
# Get sum of all read counts
c_sum = sum([v for v in c.values()])
# Normalize individual spacer count by sum of all read counts in sample
c_N = {u : float(v)/c_sum for u,v in c.items()}
c_Ns.append(c_N)
cs.append(c)
for u in c_Ns[0]:
c_N_avg[u] = ((c_Ns[0][u], c_Ns[1][u], c_Ns[2][u]), (cs[0][u], cs[1][u], cs[2][u]))
# Compute the ratios between the average experimental condition abundance and average control condition abundance
ratios = {}
for u in c_N_avg:
# Keep track of total read counts across replicates
c_total_count = np.sum(c_N_avg[u][1])
e_total_count = np.sum(e_N_avg[u][1])
c_abundance = np.average(c_N_avg[u][0])
e_abundance = np.average(e_N_avg[u][0])
# Use 1e-9 to avoid divsion by near zero
ratios[u] = (c_total_count, e_total_count, c_abundance, e_abundance, e_abundance / (c_abundance+1e-9))
eps = 1e-12 # Additive constant to avoid division by small numbers
min_read_count = 100 # Minimum read count for analysis
sigma = 5 # Number of standard deviations away from mean to establish significance
# Obtain targeting and non-targeting experiment (Y) vs control (X) average abundances.
X,Y = zip(*[(v[2]+eps, v[3]+eps) for u,v in ratios.items() if v[0] >= min_read_count and not u in nt_spacers])
X_nt, Y_nt = zip(*[(v[2]+eps, v[3]+eps) for u,v in ratios.items() if v[0] >= min_read_count and u in nt_spacers])
# Obtain mean, medan and all log depletion ratios of non-targeting spacers
mean = np.mean(np.array(np.log10(Y_nt)) - np.log10(np.array(X_nt)))
median = np.median(np.array(np.log10(Y_nt)) - np.log10(np.array(X_nt)))
# Get the spacers depletion ratios of the non-targets
dep = np.log10(np.array(Y_nt)) - np.log10(np.array(X_nt))
# Perform fit on two component Gaussian mixture model
x_d = np.linspace(-4,2, 200)
m = GaussianMixture(n_components=2)
m.fit(dep[:, None])
m_m = m.means_[0]
m_std = np.sqrt(m.covariances_[0])
logprob1 = scipy.stats.norm(m_m,m_std).logpdf(x_d)[0,:]
m_m = m.means_[1]
m_std = np.sqrt(m.covariances_[1])
logprob2 = scipy.stats.norm(m_m,m_std).logpdf(x_d)[0,:]
hi_idx = np.argsort(m.means_.flatten())[-1]
print(m.means_)
high_mean = m.means_[hi_idx]
high_std = np.sqrt(m.covariances_[hi_idx])
# Renormalize targeting and non-targeting conditions by the control median (which is in log10 space)
# Normalization parameter for all experimental conditions (to keep depletions of non-target with no offtarget
# centered at 1)
median = high_mean
Y = np.array(Y) / np.power(10, median)
Y_nt = np.array(Y_nt) / np.power(10, median)
# Redo the GMM fit using the renormalized data
dep = np.log10(np.array(Y_nt)) - np.log10(np.array(X_nt))
x_d = np.linspace(-4,2, 200)
m = GaussianMixture(n_components=2)
m.fit(dep[:, None])
m_m = m.means_[0]
m_std = np.sqrt(m.covariances_[0])
logprob1 = scipy.stats.norm(m_m,m_std).logpdf(x_d)[0,:]
m_m = m.means_[1]
m_std = np.sqrt(m.covariances_[1])
logprob2 = scipy.stats.norm(m_m,m_std).logpdf(x_d)[0,:]
hi_idx = np.argsort(m.means_.flatten())[-1]
print(m.means_)
high_mean = m.means_[hi_idx]
high_std = np.sqrt(m.covariances_[hi_idx])
depletion_thresh = float(np.power(10, high_mean - sigma*high_std))
print(depletion_thresh)
import json
"""
with open('./randoms.json', 'w') as f:
data = {'median' : float(median), 'high_mean' : float(high_mean),
'high_std' : float(high_std), 'depletion_thresh' : float(depletion_thresh)}
json.dump(data, f, sort_keys=True, indent=4)
"""
with open('./randoms.json', 'r') as f:
d = json.load(f)
print(d)
plt.figure(figsize=(3,2))
plt.axvspan(np.log10(depletion_thresh),high_mean+10,color='k',alpha=0.03)
plt.hist(dep,density=True, bins=100, color=[193/255,195/255,200/255],label='_nolegend_')
plt.plot(x_d, m.weights_[0]*np.exp(logprob1), color=[241/255,97/255,121/255], lw=2)
plt.plot(x_d, m.weights_[1]*np.exp(logprob2), color=[74/255,121/255,188/255], lw=2)
plt.axvline(np.log10(depletion_thresh), c='k',label='_nolegend_', lw=0.5)
plt.axvline(high_mean, c='k', ls='--', lw=1)
plt.xlim([-2,1])
plt.ylabel('Normalized counts')
plt.xlabel('NT spacer abundance')
plt.legend(['NT without off-target','NT with Off-target','Baseline mean',r'5$\sigma$ of baseline'], prop={'size': 6.5})
ax = plt.gca()
for item in ([] +
ax.get_xticklabels() + ax.get_yticklabels()):
item.set_fontsize(6.5)
for item in [ax.title, ax.xaxis.label, ax.yaxis.label]:
item.set_fontsize(7)
plt.savefig('./generated_data_and_data/'+ortholog+' nt GMM.pdf')
bins = np.linspace(-2,1,100)
plt.figure(figsize=(3,2))
u = np.histogram(np.log10(np.array(Y_nt) / np.array(X_nt)), bins=bins, density=True)
plt.fill_between(u[1][1:],u[0], step="pre", color=[[255/255, 81/255, 101/255]], lw=0, alpha=0.5)
u = np.histogram(np.log10(np.array(Y) / np.array(X)), bins=bins, density=True)
plt.fill_between(u[1][1:],u[0], step="pre", color=[[0.2, 0.25, 0.3]], lw=0, alpha=0.5)
plt.xlim([-2, 1])
plt.axvline(np.log10(depletion_thresh), linestyle='-', color=[0.05, 0.05, 0.1],lw=1)
plt.axvline(high_mean, c='k', ls='--', lw=1)
plt.xlabel('Log Depletion Ratio')
plt.ylabel('Normalized Counts')
plt.legend(['5$\sigma$','GMM Mean','NT', 'EG'], loc='upper left', frameon=False, prop={'size' : 6.5})
plt.ylim([0,4.8])
ax = plt.gca()
for item in ([] +
ax.get_xticklabels() + ax.get_yticklabels()):
item.set_fontsize(6.5)
for item in [ax.title, ax.xaxis.label, ax.yaxis.label]:
item.set_fontsize(7)
plt.savefig('./generated_data_and_data/'+ortholog + ' Depletion Ratios.pdf')
plt.show()
# Plot experimental vs control abundance
fig = plt.figure(figsize=(5,3.2))
ax = plt.gca()
x_l, x_r = min([min(X), min(X_nt)])/10*9, 3e-4
X_line = np.linspace(x_l, x_r, 1000)
Y_line = depletion_thresh*X_line
Y_middle_line = X_line
plt.plot(X_line, Y_middle_line, '--', c=[0.2, 0.2, 0.2], linewidth=1, zorder=3)
plt.plot(X_line, Y_line, '-', c=[0.2, 0.2, 0.2], linewidth=1, zorder=3)
plt.scatter(X, Y, color=[[255/255, 81/255, 101/255]], marker='o', s=5, alpha=0.1, rasterized=True,lw=None)
plt.scatter(X_nt, Y_nt, color='k', marker='o', s=5, alpha=0.5, rasterized=True, lw=None)
plt.xlim([x_l, x_r])
plt.ylim([1e-8, 1e-3])
ax.set_yscale('log')
ax.set_xscale('log')
plt.title(ortholog + ' Depletion')
plt.xlabel('Average control spacer abundance')
plt.ylabel('Average adjusted \nexperimental spacer abundance')
plt.legend(['x=y','5$\sigma$', 'EG', 'NT'], loc='lower right', frameon=False,prop={'size' : 6.5})
[i.set_linewidth(1) for i in ax.spines.values()]
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
plt.gcf().subplots_adjust(bottom=0.15, left=0.2)
ax = plt.gca()
for item in ([] +
ax.get_xticklabels() + ax.get_yticklabels()):
item.set_fontsize(6.5)
for item in [ax.title, ax.xaxis.label, ax.yaxis.label]:
item.set_fontsize(7)
plt.tight_layout()
plt.savefig('./generated_data_and_data/'+ortholog + ' Average Abundance Depletion.pdf', dpi=900)
plt.show()
combined_rep = [(u, (v[3]+eps) / (v[2]+eps) / np.power(10, median), v[0], v[1]) for u,v in ratios.items() if v[0] >= min_read_count and (v[3]+eps) / (v[2]+eps) / np.power(10, median) < depletion_thresh]
all_rep = [(u, (v[3]+eps) / (v[2]+eps) / np.power(10, median), v[0], v[1]) for u,v in ratios.items() if v[0] >= min_read_count]
non_depleted_rep = [(u, (v[3]+eps) / (v[2]+eps) / np.power(10, median), v[0], v[1]) for u,v in ratios.items() if v[0] >= min_read_count and (v[3]+eps) / (v[2]+eps) / np.power(10, median) >= depletion_thresh]
len(combined_rep), len(all_rep), len(non_depleted_rep)
# Index of the CDS sequences used in the experiment for targeting
cds_ids_in_exp = [
169,
46,
336,
1222,
793,
157,
156,
3994,
136,
1471,
1797,
2695,
2906,
2882,
3984,
3236,
2608,
2376,
3780,
179,
159,
28,
1018,
502,
3495,
2824,
448,
4592,
2903,
4399,
1056,
2685,
3751,
155,
1464,
1560,
2164,
1223,
1981,
2119,
447,
1484,
442,
3319,
2130,
]
# Get the e coli transcripts (CDS)
spacer_len = 30
records = list(Bio.SeqIO.parse(open('e_coli.gbk'), 'genbank'))
genome_seq = records[0].seq
cds_orig = []
flank = 500
for i,feature in enumerate(records[0].features):
if feature.type != 'CDS':
continue
loc = feature.location
feature_seq = genome_seq[loc.start-flank:loc.end+flank]
# Get the sense strand
if feature.strand == -1:
feature_seq = Bio.Seq.reverse_complement(feature_seq)
cds_orig.append((feature.qualifiers['product'][0], feature_seq))
# Filter cds to only be those from
cds = [cds_orig[i] for i in cds_ids_in_exp]
spacer_to_target_map = {}
for i,(u,v) in enumerate(ratios.items()):
if v[0] < min_read_count:
continue
search = Bio.Seq.reverse_complement(u)
s = ''
coords = (None,None)
for j,(name, seq) in enumerate(cds):
idx = seq.find(search)
if idx >= 6:
s = seq[idx-6:idx+spacer_len+6]
coords = (idx, (idx - flank) / (len(seq)-2*flank), 1)
# Add to CDS
break
if s == '':
continue
if len(s) < spacer_len+12:
continue
spacer_to_target_map[u] = s
# Identfy weblogos (requires weblogo to be installed)
with open('./generated_data_and_data/for_logo_control.fa', 'w') as f:
for i,(v) in enumerate(all_rep):
if not v[0] in spacer_to_target_map:
continue
f.write('>'+str(i)+'\n')
f.write(str(spacer_to_target_map[v[0]]).replace('T','U')+'\n')
subprocess.call(['weblogo',
'-s', 'small',
'-n', '42',
'-S', '0.05',
'--ticmarks','0.05',
'-W', '4.8',
'-F','pdf',
'-D','fasta',
'--color', '#FAA51A', 'G', 'Guanidine',
'--color', '#0F8140', 'A', 'Adenosine',
'--color', '#ED2224', 'U', 'Uracil',
'--color','#3A53A4', 'C', 'Cytidine',
'-f', './generated_data_and_data/for_logo_control.fa',
'-o', './generated_data_and_data/'+ortholog+'_weblogo_control.pdf'])
with open('./generated_data_and_data/for_logo_non_depleted.fa', 'w') as f:
for i,(v) in enumerate(non_depleted_rep):
if not v[0] in spacer_to_target_map:
continue
f.write('>'+str(i)+'\n')
f.write(str(spacer_to_target_map[v[0]]).replace('T','U')+'\n')
subprocess.call(['weblogo',
'-s', 'small',
'-n', '42',
'-S', '0.1',
'--ticmarks','0.1',
'-W', '4.8',
'-F','pdf',
'-D','fasta',
'--color', '#FAA51A', 'G', 'Guanidine',
'--color', '#0F8140', 'A', 'Adenosine',
'--color', '#ED2224', 'U', 'Uracil',
'--color','#3A53A4', 'C', 'Cytidine',
'-f', './generated_data_and_data/for_logo_non_depleted.fa',
'-o', './generated_data_and_data/'+ortholog+'_weblogo_non_depleted.pdf'])
with open('./generated_data_and_data/for_logo_depleted.fa', 'w') as f:
for i,(v) in enumerate(combined_rep):
if not v[0] in spacer_to_target_map:
continue
f.write('>'+str(i)+'\n')
f.write(str(spacer_to_target_map[v[0]]).replace('T','U')+'\n')
subprocess.call(['weblogo',
'-s', 'small',
'-n', '42',
'-S', '0.3',
'--ticmarks','0.15',
'-W', '4.8',
'-F','pdf',
'-D','fasta',
'--color', '#FAA51A', 'G', 'Guanidine',
'--color', '#0F8140', 'A', 'Adenosine',
'--color', '#ED2224', 'U', 'Uracil',
'--color','#3A53A4', 'C', 'Cytidine',
'-f', './generated_data_and_data/for_logo_depleted.fa',
'-o', './generated_data_and_data/'+ortholog+'_weblogo_depleted.pdf'])
all_rep_sorted = sorted(all_rep,key=lambda x: x[1])
one_perc = int(0.01 * len(all_rep_sorted))
with open('./generated_data_and_data/for_logo_top_one_perc.fa', 'w') as f:
for i,(v) in enumerate(all_rep_sorted):
if i > one_perc:
break
if not v[0] in spacer_to_target_map:
continue
f.write('>'+str(i)+'\n')
f.write(str(spacer_to_target_map[v[0]]).replace('T','U')+'\n')
subprocess.call(['weblogo',
'-s', 'small',
'-n', '42',
'-S', '0.5',
'-W', '4.8',
'-F','pdf',
'-D','fasta',
'--color', '#FAA51A', 'G', 'Guanidine',
'--color', '#0F8140', 'A', 'Adenosine',
'--color', '#ED2224', 'U', 'Uracil',
'--color','#3A53A4', 'C', 'Cytidine',
'-f', './generated_data_and_data/for_logo_top_one_perc.fa',
'-o', './generated_data_and_data/'+ortholog+'_weblogo_top_one_perc.pdf'])
offtargets = [(u, (v[3]+eps) / (v[2]+eps) / np.power(10, median), v[0], v[1]) for u,v in ratios.items()
if v[0] >= min_read_count
and (v[3]+eps) / (v[2]+eps) / np.power(10, median) < depletion_thresh
and u in nt_spacers]
print(len(offtargets))
with open('./generated_data_and_data/for_logo_offtargets.fa', 'w') as f:
for i,(v) in enumerate(offtargets):
# Offtargets do not have any genomematch
f.write('>'+str(i)+'\n')
f.write(v[0].replace('T','U')+'\n')
subprocess.call(['weblogo',
'-s', 'small',
'-n', '42',
'-S', '1.0',
'-W', '4.8',
'-F','pdf',
'-D','fasta',
'--color', '#FAA51A', 'G', 'Guanidine',
'--color', '#0F8140', 'A', 'Adenosine',
'--color', '#ED2224', 'U', 'Uracil',
'--color','#3A53A4', 'C', 'Cytidine',
'-f', './generated_data_and_data/for_logo_offtargets.fa',
'-o', './generated_data_and_data/'+ortholog+'_weblogo_offtargets.pdf'])
# Group spacers into +PFS or -PFS
X_has_pfs = []
Y_has_pfs = []
X_no_pfs = []
Y_no_pfs = []
for i,(u,v) in enumerate(ratios.items()):
if not u in spacer_to_target_map:
continue
s = spacer_to_target_map[u]
# NOTICE - Cas13b-t3 specific PAM
if s[5] != 'C':# and s[16] in ['A','T']:
X_has_pfs.append(v[2]+eps)
Y_has_pfs.append(v[3]+eps)
else:
X_no_pfs.append(v[2]+eps)
Y_no_pfs.append(v[3]+eps)
# Normalize by nontarget median
Y_has_pfs = np.array(Y_has_pfs) / np.power(10, median)
Y_no_pfs = np.array(Y_no_pfs) / np.power(10, median)
# Plot abundance histogram
plt.rcParams.update({'font.size': 6})
bins = np.linspace(-1.5,0.5,100)
plt.figure(figsize=(1.1,0.7))
plt.subplot(1,2,1)
u = np.histogram(np.log10(np.array(Y_no_pfs) / np.array(X_no_pfs)), bins=bins, density=True)
m = np.mean(np.log10(np.array(Y_no_pfs) / np.array(X_no_pfs)))
plt.axvline(m, color=[0.2, 0.25, 0.3], lw=0.5)
plt.fill_between(u[1][1:],u[0], step="pre", color=[[0.2, 0.25, 0.3]], lw=0, alpha=0.25)
plt.xlim([-1.5, 0.5])
ax = plt.gca()
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
u = np.histogram(np.log10(np.array(Y_has_pfs) / np.array(X_has_pfs)), bins=bins, density=True)
m = np.mean(np.log10(np.array(Y_has_pfs) / np.array(X_has_pfs)))
plt.axvline(m, color=[241/255, 95/255, 121/255], lw=0.5)
plt.fill_between(u[1][1:],u[0], step="pre", color=[[241/255, 95/255, 121/255]], lw=0, alpha=0.5)
plt.xlim([-1.5, 0.5])
plt.ylim([0,5])
plt.yticks([0,5])
ax = plt.gca()
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(0.25)
ax.tick_params(width=0.25)
plt.gcf().subplots_adjust(bottom=0.25, right=1)
plt.subplot(1,2,2)
u = np.histogram(np.log10(np.array(Y_nt) / np.array(X_nt)), bins=bins, density=True)
plt.fill_between(u[1][1:],u[0], step="pre", color=[[188/255, 230/255, 250/255]], lw=0)
plt.xlim([-1.5, 0.5])
ax = plt.gca()
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
plt.ylim([0,5])
plt.yticks([0,5])
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(0.25)
ax.tick_params(width=0.25)
plt.savefig('./generated_data_and_data/'+ortholog + ' Depletion Ratios With PFS.pdf')
plt.show()
# PFS Efficacy in prediction
pfs_eff = np.sum(np.array(Y_has_pfs) / np.array(X_has_pfs) < depletion_thresh) / len(Y_has_pfs)
off_target = np.sum(np.array(Y_nt) / np.array(X_nt) < depletion_thresh) / len(Y_nt)
eff = np.sum(np.array(Y) / np.array(X) < depletion_thresh) / len(Y)
print(pfs_eff, eff, off_target)
# Get coordinates and guide information on a per CDS basis
spacer_len = 30
counts = {}
guides = {}
for i in range(len(all_rep)):
search = Bio.Seq.reverse_complement(all_rep[i][0])
s = ''
coords = (None,None)
for j,(name, seq) in enumerate(cds):
# Find match
idx = seq.find(search)
if idx > 0:
# If match, extract match sequence and coordinates
s = seq[idx-6:idx+spacer_len+6]
coords = (idx, (idx - flank) / (len(seq)-2*flank), 1)
# Count number of guides mapping to the CDS j
counts[j] = counts.get(j,0) + 1
if not j in guides:
guides[j] = []
# Append all the guides matching to this CDS
guides[j].append(search)
if i % 1000 == 0:
print(i)
# Get depletion information on a per CDS basis
depletion_info = []
depletion_no_pam_info = []
depletion_nt_info = []
all_js = set()
spacer_len = 30
for i in range(len(all_rep)):
search = Bio.Seq.reverse_complement(all_rep[i][0])
# Get the normalized depletion (NT median divided off)
d = all_rep[i][1]
s = ''
coords = (None,None)
for j,(name, seq) in enumerate(cds):
if not j in guides:
continue
rc = Bio.Seq.reverse_complement(seq)
idx = seq.find(search)
if idx >= 6:
s = seq[idx-6:idx+spacer_len+6]
coords = (idx, (idx - flank) / (len(seq)-2*flank), 1)
break
if all_rep[i][0] in nt_spacers:
depletion_nt_info.append((j, coords, s, d))
continue
if s == '':
continue
if len(s) < spacer_len+12:
print(s)
continue
# Cas13b-t3 specific conditions
if s[5] != 'C':# and s[16] in ['A','T']:
depletion_info.append((j, coords, s, d))
else:
depletion_no_pam_info.append((j, coords, s, d))
import itertools
delta = 0.025
# Create a coordinate line linspace
L = np.arange(-0.05,1.05+delta/2,delta)
V = []
V_no_pam = []
for i in range(len(L)-1):
l = L[i]
u = L[i+1]
kv = [(v[0],v[3]) for v in depletion_info if (not v[1][1] is None) and l <= v[1][1] and v[1][1] < u]
# Groupby
gb = {}
for j,d in kv:
if not j in gb:
gb[j] = []
gb[j].append(d)
# Calculate mean across each cds
mean_by_cds = list(map(lambda x: (x[0], np.mean(x[1])),gb.items()))
# Take mean of means
v = np.mean([m for j,m in mean_by_cds])
V.append(v)
kv = [(v[0],v[3]) for v in depletion_no_pam_info if (not v[1][1] is None) and l <= v[1][1] and v[1][1] < u]
# Groupby
gb = {}
for j,d in kv:
if not j in gb:
gb[j] = []
gb[j].append(d)
# Calculate mean across each cds
mean_by_cds = list(map(lambda x: (x[0], np.mean(x[1])),gb.items()))
# Take mean of means
v = np.mean([m for j,m in mean_by_cds])
V_no_pam.append(v)
W = [v[3] for v in depletion_no_pam_info + depletion_info if not v[1][1] is None and l <= v[1][1] and v[1][1] < u]
div_factor = 1
plt.figure(figsize=(0.5,0.6))
plt.plot(L[:-1]+delta/2,np.array(V_no_pam),color=[0.2, 0.25, 0.3], lw=0.5)
plt.plot(L[:-1]+delta/2,np.array(V),color=[241/255, 95/255, 121/255], lw=0.5)
plt.ylim([0.0,1.0])
plt.yticks([0.0,0.5,1.0])
ax = plt.gca()
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(0.25)
ax.tick_params(width=0.25)
ax.tick_params(axis='x',direction='out', length=1.44, width=0.25)
ax.tick_params(axis='y',direction='out', length=1.80, width=0.25)
plt.xlim([-0.05,1.05])
plt.savefig('./generated_data_and_data/'+ortholog+' positional_preference.pdf')
pos = [u[1][1] for u in depletion_info if not u[1][1] is None]
plt.figure(figsize=(4,2))
plt.hist(pos, 100, color=[0.2, 0.25, 0.3], density=True)
plt.xlabel('Normalized position along gene')
plt.ylabel('Normalized \nguide count')
plt.title(ortholog)
plt.xlim([-0.2, 1.2])
plt.gcf().subplots_adjust(bottom=0.25, left=0.2)
plt.savefig('./generated_data_and_data/'+ortholog + ' Gene Position Distribution.pdf')
plt.show()
depletion_info = []
all_depletion_info = []
all_js = set()
spacer_len = 30
for i in range(len(all_rep)):
search = Bio.Seq.reverse_complement(all_rep[i][0])
d = all_rep[i][1]
s = ''
coords = (None,None)
for j,(name, seq) in enumerate(cds):
if not j in guides:
continue
rc = Bio.Seq.reverse_complement(seq)
idx = seq.find(search)
if idx >= 6:
s = seq[idx-6:idx+spacer_len+6]
coords = (idx, (idx - flank) / (len(seq)-2*flank), 1)
break
if all_rep[i][0] in nt_spacers:
depletion_nt_info.append((j, coords, s, d))
continue
if s == '':
continue
if len(s) < spacer_len+12:
print(s)
continue
if d < depletion_thresh:
depletion_info.append((j, coords, s, d))
all_depletion_info.append((j, coords, s, d))
print(len(depletion_info))
print(len(all_depletion_info))
# Multi positional preferences
bases = ['A', 'T', 'G', 'C']
p1 = 5
p2 = 37
p3 = 38
tokens = {(a,b,c) : 0 for a in bases for b in bases for c in bases}
tokens_all = {(a,b,c) : 0 for a in bases for b in bases for c in bases}
for i in range(len(depletion_info)):
try:
token = (str(depletion_info[i][2][p1]), str(depletion_info[i][2][p2]), str(depletion_info[i][2][p3]))
tokens[token] += 1
except:
pass
for i in range(len(all_depletion_info)):
try:
token = (str(all_depletion_info[i][2][p1]), str(all_depletion_info[i][2][p2]), str(all_depletion_info[i][2][p3]))
tokens_all[token] += 1
except:
pass
token_depletion = {u : 1-tokens[u] / (tokens_all[u]+0.001) for u in tokens.keys()}
dual_bases = [(a,b) for a in bases for b in bases]
dual_bases_labels = [a+b for a,b in dual_bases]
Z = np.zeros((4,16))
for i,a in enumerate(bases):
for j,(b,c) in enumerate(dual_bases):
token = (a,b,c)
depletion = token_depletion[token]
Z[i,j] = depletion
plt.figure(figsize=(4,1.75))
cm = plt.cm.get_cmap('magma_r')
ax = plt.gca()
im = plt.imshow(Z,cmap=cm, vmax=1)
ax.set_xticks(np.arange(len(dual_bases_labels)))
ax.set_yticks(np.arange(len(bases)))
# ... and label them with the respective list entries
ax.set_xticklabels(dual_bases_labels, rotation=-60, fontdict={'fontfamily' : 'Andale Mono'})
ax.set_yticklabels(bases,rotation=0, fontdict={'fontfamily' : 'Andale Mono'})
plt.xlabel('3\' PFS (+2, +3)')
plt.ylabel('5\' PFS (-1)')
plt.title(ortholog)
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.05)
plt.colorbar(im,cax=cax)
plt.savefig('./generated_data_and_data/'+ortholog + ' pfs map.pdf')
plt.show()
```
| github_jupyter |
# Finding Contours
### Import resources and display image
```
import numpy as np
import matplotlib.pyplot as plt
import cv2
%matplotlib inline
# Read in the image
image = cv2.imread('images/thumbs_up_down.jpg')
# Change color to RGB (from BGR)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
plt.imshow(image)
```
### Produce a binary image for finding contours
```
# Convert to grayscale
gray = cv2.cvtColor(image,cv2.COLOR_RGB2GRAY)
# Create a binary thresholded image
retval, binary = cv2.threshold(gray, 230, 255, cv2.THRESH_BINARY_INV)
#retval, binary = cv2.threshold(gray, 230, 255, cv2.THRESH_BINARY)
plt.imshow(binary, cmap='gray')
```
### Find and draw the contours
```
# Find contours from thresholded, binary image
contours, hierarchy = cv2.findContours(binary, cv2.RETR_TREE, cv2.CHAIN_APPROX_SIMPLE)
# Draw all contours on a copy of the original image
contours_image = np.copy(image)
contours_image = cv2.drawContours(contours_image, contours, -1, (0,255,0), 3)
plt.imshow(contours_image)
print(contours[1])
#print(contours.lenght)
```
## Contour Features
Every contour has a number of features that you can calculate, including the area of the contour, it's orientation (the direction that most of the contour is pointing in), it's perimeter, and many other properties outlined in [OpenCV documentation, here](http://opencv-python-tutroals.readthedocs.io/en/latest/py_tutorials/py_imgproc/py_contours/py_contour_properties/py_contour_properties.html).
In the next cell, you'll be asked to identify the orientations of both the left and right hand contours. The orientation should give you an idea of which hand has its thumb up and which one has its thumb down!
### Orientation
The orientation of an object is the angle at which an object is directed. To find the angle of a contour, you should first find an ellipse that fits the contour and then extract the `angle` from that shape.
```python
# Fit an ellipse to a contour and extract the angle from that ellipse
(x,y), (MA,ma), angle = cv2.fitEllipse(selected_contour)
```
**Orientation values**
These orientation values are in degrees measured from the x-axis. A value of zero means a flat line, and a value of 90 means that a contour is pointing straight up!
So, the orientation angles that you calculated for each contour should be able to tell us something about the general position of the hand. The hand with it's thumb up, should have a higher (closer to 90 degrees) orientation than the hand with it's thumb down.
### TODO: Find the orientation of each contour
```
## TODO: Complete this function so that
## it returns the orientations of a list of contours
## The list should be in the same order as the contours
## i.e. the first angle should be the orientation of the first contour
def orientations(contours):
"""
Orientation
:param contours: a list of contours
:return: angles, the orientations of the contours
"""
# Create an empty list to store the angles in
# Tip: Use angles.append(value) to add values to this list
angles = []
for selected_contour in contours:
(x,y), (MA,ma), angle = cv2.fitEllipse(selected_contour)
angles.append(angle)
return angles
# ---------------------------------------------------------- #
# Print out the orientation values
angles = orientations(contours)
print('Angles of each contour (in degrees): ' + str(angles))
```
### Bounding Rectangle
In the next cell, you'll be asked to find the bounding rectangle around the *left* hand contour, which has its thumb up, then use that bounding rectangle to crop the image and better focus on that one hand!
```python
# Find the bounding rectangle of a selected contour
x,y,w,h = cv2.boundingRect(selected_contour)
# Draw the bounding rectangle as a purple box
box_image = cv2.rectangle(contours_image, (x,y), (x+w,y+h), (200,0,200),2)
```
And to crop the image, select the correct width and height of the image to include.
```python
# Crop using the dimensions of the bounding rectangle (x, y, w, h)
cropped_image = image[y: y + h, x: x + w]
```
### TODO: Crop the image around a contour
```
## TODO: Complete this function so that
## it returns a new, cropped version of the original image
def left_hand_crop(image, selected_contour):
"""
Left hand crop
:param image: the original image
:param selectec_contour: the contour that will be used for cropping
:return: cropped_image, the cropped image around the left hand
"""
## TODO: Detect the bounding rectangle of the left hand contour
x,y,w,h = cv2.boundingRect(selected_contour)
## TODO: Crop the image using the dimensions of the bounding rectangle
# Make a copy of the image to crop
cropped_image = np.copy(image)
cropped_image = cropped_image[y: y + h, x: x + w]
return cropped_image
## TODO: Select the left hand contour from the list
## Replace this value
selected_contour = contours[1]
# ---------------------------------------------------------- #
# If you've selected a contour
if(selected_contour is not None):
# Call the crop function with that contour passed in as a parameter
cropped_image = left_hand_crop(image, selected_contour)
plt.imshow(cropped_image)
```
| github_jupyter |

## Table of contents
1) [Introduction](#introduction)
2) [Circuit Depth](#depth)
3) [Circuit Unitary Factors](#unitary)
4) [Circuits with Classical Registers](#classical)
5) [Gate Set Dependence of Depth](#gate_dependence)
```
from qiskit import *
%matplotlib inline
```
## Introduction <a name='introduction'></a>
When constructing quantum circuits, there are several properties that help quantify the "size" of the circuits, and their ability to be run on a noisy quantum device. Some of these, like number of qubits, are straightforward to understand, while others like depth and number of tensor components require a bit more explanation. Here we will explain all of these properties, and, in preparation for understanding how circuits change when run on actual devices, highlight the conditions under which they change.
### Basics
Consider the following circuit:
```
qc = QuantumCircuit(12)
for idx in range(5):
qc.h(idx)
qc.cx(idx, idx+5)
qc.cx(1, 7)
qc.x(8)
qc.cx(1, 9)
qc.x(7)
qc.cx(1, 11)
qc.swap(6, 11)
qc.swap(6, 9)
qc.swap(6, 10)
qc.x(6)
qc.draw()
```
From the plot, it is easy to see that this circuit has 12 qubits, and a collection of Hadamard, CNOT, X, and SWAP gates. But how to quantify this programmatically? Because we can do single-qubit gates on all the qubits simultaneously, the number of qubits in this circuit is equal to the **width** of the circuit:
```
qc.width()
```
<div class="alert alert-block alert-warning">
<b>Warning:</b> For a quantum circuit composed from just qubits, the circuit width is equal to the number of qubits. This is the definition used in quantum computing. However, for more complicated circuits with classical registers, and classically controlled gates, this equivalence breaks down. As such, from now on we will <b>not</b> refer to the number of qubits in a quantum circuit as the width.
</div>
We can also just get the number of qubits directly:
```
qc.n_qubits
```
It is also straightforward to get the number and type of the gates in a circuit using `count_ops()`:
```
qc.count_ops()
```
We can also get just the raw count of operations by computing the circuits **size**:
```
qc.size()
```
## Quantum Circuit Depth <a name="depth"></a>
A particularly important circuit property is known as the **depth**. The depth of a quantum circuit is a measure of how many "layers" of quantum gates, executed in parallel, it takes to complete the computation defined by the circuit. Because quantum gates take time to implement, the depth of a circuit roughly corresponds to the amount of time it takes the quantum computer to execute the circuit. Thus, the depth of a circuit is one important quantity used to measure if a quantum circuit can be run on a device.
The depth of a quantum circuit has a mathematical definition as the longest path in a directed acyclic graph (DAG). However, such a definition is a bit hard to grasp, even for experts. Fortunately, the depth of a circuit can be easily understood by anyone familiar with playing [Tetris](https://en.wikipedia.org/wiki/Tetris). To understand this, let us redraw the circuit with the first five CNOT gates colored differently for clarity:

To compute the depth, we turn the circuit counterclockwise so that the beginning of the circuit is at the bottom. We then let the gates fall to the bottom. Later gates stack on earlier gates, and multi-qubit gate components must be stacked at the same height. For the current circuit of interest, this stacking looks like:

We can see that the first five CNOT gates all collapsed down on top of each other, and are stacked on top of the initial set of Hadamard gates. The remaining gates stack on top of the CNOT layer as shown. The stack of gates can be partitioned into "layers", where each layer represents a set of gates that can be executed in parallel on a quantum device (Hardware limitations may restrict the number and/or type of gates that can be run in parallel). The **depth** of the circuit is just the number of layers in the circuit; The depth is equal to the height of the stack of gates. This computation is done for you in qiskit, and we can verify our visual method:
```
qc.depth()
```
## Unitary Factors <a name="unitary"></a>
The circuit we are focusing on here is a 12-qubit circuit. However, does this circuit actually require a 12-qubit quantum computer to run? That is to say, can we compute the same result by running a collection of smaller circuits individually?
In the limit where only single-qubit gates are performed, it should be clear that each qubit is controlled independently of the rest, and thus we can run each qubit independently and still get the desired result. Thus, the question becomes are there enough entangling gates in the circuit to have all qubits interacting? Again, this is best understood in terms of diagrams. Below, we track the sets of qubits that interact amongst themselves via CNOT gates at each layer in the circuit.

We can see that at the end of the computation there are three independent sets of qubits. Thus, our 12-qubit computation is actual two two-qubit calculations and a single eight-qubit computation. We can verify this via qiskit:
```
qc.num_unitary_factors()
```
## Circuits with Classical Registers and Measurements <a name="classical"></a>
Several of the circuit properties introduced so far change when adding classical registers and measurements.
Let's add measurements to the circuit above.
```
qc2 = QuantumCircuit(12, 12)
for idx in range(5):
qc2.h(idx)
qc2.cx(idx, idx+5)
qc2.cx(1, 7)
qc2.x(8)
qc2.cx(1, 9)
qc2.x(7)
qc2.cx(1, 11)
qc2.swap(6, 11)
qc2.swap(6, 9)
qc2.swap(6, 10)
qc2.x(6)
qc2.barrier()
qc2.measure(range(12), range(12))
qc2.draw()
```
The **width** of the circuit now includes the number of qubits _and_ number of classical bits:
```
qc2.width()
```
The number of operations has increased because of the measurements and the barrier we used:
```
qc2.count_ops()
```
and the **size** of the circuit has grown:
```
qc2.size()
```
The **depth** of the circuit has now increased because measurements are included in the depth computation as they perform physical operations:
```
qc2.depth()
```
Barriers and other special commands like snapshots do not count towards the depth directly. However, gates cannot pass through them and must therefore start stacking on top.
## Dependence of Depth on Gate Selection <a name="gate_dependence"></a>
We close by highlighting a very important point. The **depth** of a quantum circuit, and thus the ability to run said circuit on noisy quantum hardware depends on the choice of gates used to implement that circuit. The original circuit used in this tutorial had a depth of 9.
```
qc.depth()
```
However, the SWAP gates used in the construction of that circuit are not native to the IBM Q devices. A decomposition that runs on the devices is a decomposition in terms of three CNOT gates:
```
qc3 = QuantumCircuit(2)
qc3.swap(0,1)
qc3.decompose().draw() # This decomposes the swap gate into the gates we want
```
So the exact same circuit could be written as:
```
qc4 = QuantumCircuit(12)
for idx in range(5):
qc4.h(idx)
qc4.cx(idx, idx+5)
qc4.cx(1, 7)
qc4.x(8)
qc4.cx(1, 9)
qc4.x(7)
qc4.cx(1, 11)
qc4.cx(6, 11)
qc4.cx(11, 6)
qc4.cx(6, 11)
qc4.cx(6, 9)
qc4.cx(9, 6)
qc4.cx(6, 9)
qc4.cx(6, 10)
qc4.cx(10, 6)
qc4.cx(6, 10)
qc4.x(6)
qc4.draw()
```
That has a depth approaching twice that of the original circuit
```
qc4.depth()
```
This simple example is meant to illustrate a very important point: **When running circuits on actual quantum devices, the circuit that gets run is in general not the same circuit that you constructed**. In addition, the depth of that new circuit is likely to be larger, and in some cases much larger, than the original one. Fortunately, often times one can reduce this overhead through smart circuit rewriting toolchains.
```
import qiskit.tools.jupyter
%qiskit_version_table
%qiskit_copyright
```
| github_jupyter |
# Country Happiness Score Assessment
## By Chad Goldberg
I created this particular dataset to compare with a similarly structured dataset comprised of values from 2015, while mine uses values from 2017. The original dataset was used to analyze the relationship between foreign investment and the happiness of the countries being invested in. Overall, the majority of this project was spent exploring and learning how to merge data from different sources with some exploratory data analysis sprinkled in throughout. Though I feel more confident now in my ability to locate, import, and merge data, there are still many areas that I need to work in in the future to make myself a more effective data scientist.
```
# Load the Pandas libraries with alias 'pd'
import pandas as pd
import geopandas as gpd
import numpy as np
import seaborn as sns
import statsmodels.formula.api as smf
from sklearn.linear_model import Lasso
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestRegressor
import matplotlib.pyplot as plt
!pip install geopandas
!pip install descartes
# import preliminary dataset and display first 5 rows
happiness = pd.read_csv("happiness.csv")
happiness.head()
# a simple histogram to see if i can visualize something
happiness.score.hist()
happiness.sort_values(by='score', ascending=True)
```
## The above histogram is fairly interesting because it shows that the largest score bucket is that with happiness ratings between 4.5 & 5 (roughly 25). Additionally, there are only 13 countries with happiness scores over 7, and luckily only 8 with scores 3.5 or lower. The sorted printed data additionally helps us confirm and view those countries with the lowest and highest scores, showing only 1 below 3 (Burundi), and Finland being the highest on the list.
```
# basic statistics in pandas
happiness.score.describe()
```
## Though not incredibly useful for my purposes, these basic statistics describing the happiness scores offer insights into mean happiness score for the world as will as specific values for min & max again.
```
happiness.investment.hist()
happiness.sort_values(by='investment', ascending=True)
```
## Similarly, I sorted the dataset by foreign direct investment (least to greatest). Foreign direct investment (FDI), in this context, is described as net inflows coming into a country from all outside (foreign) sources. Clearly there is missing data, as evidenced by the NaN values towards the end, but the dataset leads to some interesting questions when looking at other values. For example, it's curious to me that the Netherlands have the second largest foreign direct investment of all countries in the world. Further research suggests that this number is accurate, citing a competitive tax structure as well as liberal policy towards foreign investment. It's also interesting to note that there are only 12 countries with negative FDI, an indication the capital outflow, or divestment, exceeds the country's inflow. It was an initial hypothesis that countries with higher FDIs would have higher happiness scores, though that doesn't necessarily seem to be the case as Iceland, one of the happiest countries, has one of the lowest FDI values and there are multiple countries with high FDIs that have happiness scores below 6 (India,China,Russia).
```
# scatterplot
f, ax = plt.subplots(figsize=(6.5, 6.5))
happiness.plot.scatter('investment', 'score', alpha=0.15,ax=ax)
```
## The above is another attempt to view a correlation between happiness score and FDI, though the resulting scatterplot does not seem to suggest any obvious relationship.
```
# scatterplot...potentially good because it doesn't necessarily correlate population with happiness score
happiness.plot.scatter('score', 'population')
```
## I then wanted to look into the relationship between population and happiness score, having no real idea as to whether there would be one or not. Again, the scatterplot does not necessarily indicate a correlation between happiness score and country population.
```
# scatterplot...no real relationship between population & investment
happiness.plot.scatter('investment', 'population')
```
## A scatterplot assessing the relationship between FDI and population shows the potential for a slight positive correlation, though I ran across some issues when trying to actually calculate correlation. I tried using np.corrcoef(happiness.investment, happiness.population) but the results came back as NaN for each results. Given more time, I would work to actually calculate it.
# Importing & Merging New Data
## The following section begins by importing a new dataset that contains a list of countries with accompanying scores for Political Rights (PR) and Civil Liberties (CL).
```
# import secondary dataset /political rights scores/ and display first 5 rows to see if it worked
political_rights = pd.read_csv("prscores.csv")
political_rights.head()
```
## The import seemed to work but there are additional columns (w,x,y,z) that are unncessary. I ended up saving that problem until later so that I could focus on merging this new dataset with the old.
```
# merge happiness dataset with new political rights dataset
merged = pd.merge(happiness, political_rights, on='country')
merged.head()
```
## With a seemingly good merge, I print the whole list to get an idea of the whole dataset and see what's potentially missing or still out of place.
```
print(merged)
```
## Dropping the extra columns ended up being simple with the following code:
```
# trying to figure out how to drop the unfilled columns
merged.drop(["w","x","y","z"],axis=1, inplace=True)
print(merged)
#update: successfully dropped unnamed columns
```
## I then wanted to see if I could visualize some of the new data with the old so I took to more EDA:
```
#merged.plot.scatter('score','PR')
type(merged.PR)
```
## I first tried to do a scatterplot of happiness scores vs PR, though I ran into an interesting issue.
## I was presented with the following "ValueError: scatter requires x column to be numeric", so I checked
## to see what datatype PR was and it was listed as a series, so I went through the steps to convert it
# to a numeric using the following code.
```
s = pd.Series(merged.PR)
>>> pd.to_numeric(s)
```
## I assumed it would be fine so I tried it again but got the same error. I then tried to convert it again to a "float64" just to make sure
```
s.astype('float64')
```
## Again I got the same error. I then looked it up online and the problem seems to be unresolved amongst many users so I figured I could do without visualizing that relationship.
## Because of that failed visualization I wanted to import more data. I ended up going with data that shows the "average annual hours worked per person per country" as I figured this would be interesting to compare with the other variables and I assumed potential relationship between hrs worked, happiness,and foreign investment. Unfortunately the dataset only accounted for OECD countries so there are only 38 datapoints which will obviously provide for limited analysis.
```
# import dataset /avg annual hrs worked/ and display first 5 rows
hrsworked = pd.read_csv("hrsworked.csv")
hrsworked.head()
# merge hrsworked dataset with current dataset to be able to compare all countries that have data on avg annual hrs worked
merged2 = pd.merge(merged, hrsworked, on='country')
merged2.head()
```
## Import and merge went well so I now wanted to sort (ascending) based on hrs.
```
# want to sort top and bottom 5 avg annual work hour countries
merged2.sort_values(by=['hrs'])
#merged2.plot.scatter('investment', 'hrs')
area = np.pi*3
plt.scatter(merged2.score, merged2.hrs, s=area, alpha=0.5)
```
## As expected, the relationship between hrs worked and happiness score does not necessarily show a correlation and seems somewhat random. With more time and resources I would like to know the average hrs worked from people in other countries as I feel that may tell a different story based on the fact that Most OECD members are high-income economies with a very high Human Development Index (HDI) and are regarded as developed countries.It would be interesting to compare lower income economies.
## That prompted the idea of importing new data that shows country GDP. The following code is my importing of the new datafram followed by a merge with my initial merged dataset.
```
# import dataset /GDP per capita/ and display first 5 rows
GDP = pd.read_csv("GDP.csv")
GDP.head()
merged3 = pd.merge(merged, GDP, on='country')
merged3.head()
area = np.pi*3
plt.scatter(merged3.score, merged3.GDP, s=area, alpha=0.5)
```
## The above scatterplot is pretty interesting, as it shows a fairly clear relationship (seemingly exponential) between happiness score and GDP.
# Attempts At Making Choropleth Maps
## The following section outlines my ambitions to use geopandas and geographic data to be able to visually assess the countries and their data from a world view.
## To start, I read in a publicly available dataset that has country/region names with the accompanying geometry necessary to draw the areas on the map.
```
# reading in
world = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
#world.plot()
world
# In anticipation of a merge, I changed the "name" column to "country" so I can merge new dataset with the old on country
world.columns = ['pop_est','continent','country','iso_a3','gdp_md_est','geometry']
world.plot(column='pop_est')
```
## I was luckily able to plot a sample with the given 'pop_est' data the came with the public dataset. Next I needed to merge with my old data. I decided to do a right merge so that I would keep all of the extra countries that did not match with my old dataset. Given more time, the next order of business would be to properly clean the data and make a dataframe with all the relevant and matching countries.
```
# merge new country geometry data with initial dataframe
merged4 = pd.merge(merged, world, on='country', how='right')
#displays the type of dataframe
type(merged4), type(world)
crs = {'init': 'epsg:4326'}
merged4_gdf = gpd.GeoDataFrame(merged4, crs=crs, geometry=merged4.geometry)
```
## I ran into trouble plotting the map after I merged but after some exploring I found that the merged dataset was no longer a GeoDataFrame, but a regular dataframe. The above code changed the merged4 accordingly. I then wanted to plot a visual of world happiness scores.
```
fig, ax = plt.subplots(1, 1)
merged4_gdf.plot('score', ax=ax, legend=True)
```
## This ended up working fairly well, though some obvious problems arose. It seems as though some of the data is corrupted as there seems to be too many places where the scores are 3.5 or lower, and clearly the United States is not one of them. Again, I would need to go through and thoroughly clean the new merged data and figure out how to properly account for the desired variables.
## There are some positives, however, as one can clearly confirm some of the happiest countries are those in Scandinavia.
# Conclusion
## Overall, I'd say the most interesting find was that there seemed to be an exponential relationship between GDP and happiness score. The data suggests that GDP increases as country happiness scores increase.
## I also feel with more time and practice I could create some pretty good looking geographic visuals, though I clearly would need more practice with data cleaning. I'm a bit frustrated at the problem that arose with some of the coding that I couldn't figure out, so there was some visualization left to be desired and some analysis that I could not do. I feel there are many more variables that I could potentially look at with regards to countries around the world but the hardest part in synthesizing it all would be to match each country correctly with corresponding data.
# Data Sheet Information
## Happiness Investment Datasheet
## Owner: Chad Goldberg
## Motivation for Dataset Creation
## Why was the dataset created? (e.g., were there specific tasks in mind, or a specific gap that needed to be filled?)
## I created this particular dataset to compare with a similarly structured dataset comprised of values from 2015, while mine uses values from 2017. The original dataset was used to analyze the relationship between foreign investment and the happiness of the countries being invested in. I intend to compare these results and map changes over the years.
## What (other) tasks could the dataset be used for? Are there obvious tasks for which it should not be used?
## Though this dataset may be used in conjunction with many other values for numerous tasks, the values I chose to use only make sense for my particular narrative.
## I have included in this dataset a few instances (political rights, civil liberties) that one may use for additional tasks, however I did not utilize them in my analyses and included them merely to replicate the previous dataset construction as closely as possible.
## Has the dataset been used for any tasks already? If so, where are the results so others can compare (e.g., links to published papers)?
## This particular dataset has not been used, though the same variables were used to create a dataset based on 2015 values. (https://www.kaggle.com/andreyka2/happiness-and-investment)
## Who funded the creation of the dataset? If there is an associated grant, provide the grant number.
## There was no funding for the creation of this dataset as it is being used strictly for practice in an applied data science course at Tufts University.
## Any other comments?
## Dataset Composition
## What are the instances? (that is, examples; e.g., documents, images, people, countries) Are there multiple types of instances? (e.g., movies, users, ratings; people, interactions between them; nodes, edges)
## Instances include multiple values for: ‘country’, ‘happiness score’, ‘happiness rank’, ‘foreign investment’ (in US dollars), ‘population’, ‘political rights’, ‘civil liberties’
## Are relationships between instances made explicit in the data (e.g., social network links, user/movie ratings, etc.)?
## Our data displays each country as it relates to the accompanying values and is fairly intuitive to understand.
## How many instances of each type are there?
## The unmerged and unclean data has the following number of instances for each type:
## ‘country’: 156
## ‘happiness score’: 156 (scores ranging from 2.905-7.632)
## ‘happiness rank’: 156 (ranks 1-156)
## ‘foreign investment’: 243
## ‘population’: 262
## ‘political rights’: 195 (scores ranging from 1-7)
## ‘civil liberties’: 195 (scores ranging from 1-7)
## ‘hrs’
## ‘GDP’
## The final merged and cleaned data will accommodate the lowest number of instances so as to avoid large amounts of missing data, which would come from choosing the largest number. In this case, we intend our dataset to have 156 instances for each type.
## What data does each instance consist of? “Raw” data (e.g., unprocessed text or images)? Features/attributes? Is there a label/target associated with instances? If the instances are related to people, are subpopulations identified (e.g., by age, gender, etc.) and what is their distribution?
## Raw data of the instances include scores based on survey responses (‘happiness score’, ‘political rights’, ‘civil liberties’) and cumulative values from the International Monetary Fund (‘foreign investment’).
## Is everything included or does the data rely on external resources? (e.g., websites, tweets, datasets) If external resources, a) are there guarantees that they will exist, and remain constant, over time; b) is there an official archival version. Are there licenses, fees or rights associated with any of the data?
## The data relies on the following external collection sources:
## For ‘happiness score’ & ‘happiness rank’:
## http://worldhappiness.report/ed/2018/
## For ‘foreign investment’:
## https://data.worldbank.org/indicator/BX.KLT.DINV.CD.WD?end=2017&start=2017
## For ‘population’:
## https://data.worldbank.org/indicator/sp.pop.totl
## For ‘political rights’ & ‘civil liberties’:
## https://freedomhouse.org/report/fiw-2017-table-country-scores
## For ‘average work hours’
## https://stats.oecd.org/Index.aspx?DataSetCode=ANHRS
## For ‘GDP per capita’
## https://data.worldbank.org/indicator/ny.gdp.pcap.cd
## Though there is no guarantee that the data will exist in the future or remain constant, the data is currently drawn from sources that try to maintain and update the values annually.
## Are there recommended data splits or evaluation measures? (e.g., training, development, testing; accuracy/AUC)
## There are no recommended data splits or evaluation measures.
## What experiments were initially run on this dataset?
## Have a summary of those results and, if available, provide the link to a paper with more information here.
## There were no experiments initially run on this dataset.
## Any other comments?
## Data Collection Process
## How was the data collected? (e.g., hardware apparatus/sensor, manual human curation, software pro- gram, software interface/API; how were these con- structs/measures/methods validated?)
## The data was collected from three main public sources: World Happiness report, World Bank, and Freedom House (links to the sources were previously referenced). After downloading the relevant data files from each source and isolating values for 2017, I manually consolidated values for each instance into one excel spreadsheet.
## Who was involved in the data collection process? (e.g., students, crowdworkers) How were they compensated? (e.g., how much were crowdworkers paid?)
## I was solely involved in the data collection process and was not compensated, as the project is for learning purposes and academic in nature.
## Over what time-frame was the data collected? Does the collection time-frame match the creation time-frame?
## The data collection and creation time-frames were equal in nature, each spanning a period of roughly two days.
## How was the data associated with each instance acquired? Was the data directly observable (e.g., raw text, movie ratings), reported by subjects (e.g., survey responses), or indirectly inferred/derived from other data (e.g., part of speech tags; model-based guesses for age or language)? If the latter two, were they validated/verified and if so how?
## Not all the data was directly observable, as much was the result of pooled values stemming from previous analysis (‘happiness score’, ‘foreign investment’, ‘civil liberties’, ‘political liberties’).
## Does the dataset contain all possible instances? Or is it, for instance, a sample (not necessarily random) from a larger set of instances?
## This dataset merely contains samples from larger sets of instances. I simply pulled desired instances and values from the sources referenced above.
## If the dataset is a sample, then what is the population?
## What was the sampling strategy (e.g., deterministic, probabilistic with specific sampling probabilities)? Is the sample representative of the larger set (e.g., geographic coverage)? If not, why not (e.g., to cover a more diverse range of in- stances)? How does this affect possible uses?
## Though this dataset is a combined sample larger datasets, the instance (country) acts as the factor that relates all other instances. That is, all the datasets I pulled from were utilizing data representative of my main population.
## Is there information missing from the dataset and why?
(this does not include intentionally dropped instances; it might include, e.g., redacted text, withheld documents) Is this data missing because it was unavailable?
## There are missing values in each individual instance but it Is unclear as to why. The missing values seem to be sporadic in nature and the original datasets offer no reason for empty cells.
## Are there any known errors, sources of noise, or redundancies in the data?
## I have not noticed any such cases.
## Any other comments?
## n/a
## Data Preprocessing
## What preprocessing/cleaning was done? (e.g., discretization or bucketing, tokenization, part-of-speech tagging, SIFT feature extraction, removal of instances, processing of missing values, etc.)
## Some of the country names were different across each set of files (for example, Laos and Lao PDR, South Korea and Republic of Korea) so I had to manually change each different value. Additionally, I had to ensure that missing values were removed.
## Was the “raw” data saved in addition to the preprocessed/cleaned data? (e.g., to support unanticipated future uses)
## Yes, I saved each individual “raw” data file as separate original entities and have saved my data at multiple stages to support future uses as well as enable the ability to start fresh from certain processing points.
## Is the preprocessing software available?
## No it is not.
## Does this dataset collection/processing procedure achieve the motivation for creating the dataset stated in the first section of this datasheet?
## While I am closer to achieving the motivation for creating the dataset, I am still in early stages of processing and analysis. I will feel more excited and accomplished once I begin to visualize the data.
## Any other comments?
## Dataset Distribution
## How is the dataset distributed? (e.g., website, API, etc.; does the data have a DOI; is it archived redundantly?)
## The dataset will be published on the owner’s GitHub account.
## When will the dataset be released/first distributed? (Is there a canonical paper/reference for this dataset?)
## The dataset will be released in the coming months, at the completion of visualization and analysis.
## What license (if any) is it distributed under? Are there any copyrights on the data?
## There are no copyrights on the data.
## Are there any fees or access/export restrictions? Any other comments?
## There are no fees or restrictions.
## Any other comments?
## Dataset Maintenance
## Who is supporting/hosting/maintaining the dataset?
## How does one contact the owner/curator/manager of the dataset (e.g. email address, or other contact info)?
## The owner may be contacted at the following email address: chad.goldberg@tufts.edu
## Will the dataset be updated? How often and by whom? How will updates/revisions be documented and communicated (e.g., mailing list, GitHub)? Is there an erratum?
## I do not anticipate updating the dataset as this intends to be a one-time academic assignment.
## If the dataset becomes obsolete how will this be communicated?
## There will be no way of communicating this other than through replication or similar analysis. However, because this dataset represents reported values from 2017, it represents a mere snapshot of the particular instances associated with that year. For that reason, I do not see the data set becoming “obsolete” in the traditional sense, as may be utilized when comparing future replications or similar analysis.
## Is there a repository to link to any/all papers/systems that use this dataset?
## There will eventually be a link to the owner’s GitHub account.
## If others want to extend/augment/build on this dataset, is there a mechanism for them to do so? If so, is there a process for tracking/assessing the quality of those contributions. What is the process for communicating/distributing these contributions to users?
## The dataset will be available to download as a .csv file so any and all augmentations will be individual to each new use case. Similarly, communication or distribution of said contributions will be on an individual basis as well.
## Any other comments?
## Legal and Ethical Considerations
## If the dataset relates to people (e.g., their attributes) or was generated by people, were they informed about the data collection? (e.g., datasets that collect writing, photos, interactions, transactions, etc.)
## This does not apply to my dataset.
## If it relates to other ethically protected subjects, have appropriate obligations been met? (e.g., medical data might include information collected from animals)
## This does not apply to my dataset.
## If it relates to people, were there any ethical review applications/reviews/approvals? (e.g. Institutional Review Board applications)
## This does not apply to my dataset.
## If it relates to people, were they told what the dataset would be used for and did they consent? What community norms exist for data collected from human communications? If consent was obtained, how? Were the people provided with any mechanism to revoke their consent in the future or for certain uses?
## This does not apply to my dataset.
## If it relates to people, could this dataset expose people to harm or legal action? (e.g., financial social or otherwise) What was done to mitigate or reduce the potential for harm?
## This does not apply to my dataset.
## If it relates to people, does it unfairly advantage or dis- advantage a particular social group? In what ways? How was this mitigated?
## This does not apply to my dataset.
## If it relates to people, were they provided with privacy guarantees? If so, what guarantees and how are these ensured?
## This does not apply to my dataset.
## Does the dataset comply with the EU General Data Protection Regulation (GDPR)? Does it comply with any other standards, such as the US Equal Employment Opportunity Act?
## Yes, as far as I know. The data is collected from trusted associations or government initiatives that are known to comply with GDPR.
## Does the dataset contain information that might be considered sensitive or confidential? (e.g., personally identifying information)
## No it does not.
## Does the dataset contain information that might be considered inappropriate or offensive?
## No it does not.
| github_jupyter |
# Time Series Prediction with BQML and AutoML
**Objectives**
1. Learn how to use BQML to create a classification time-series model using `CREATE MODEL`.
2. Learn how to use BQML to create a linear regression time-series model.
3. Learn how to use AutoML Tables to build a time series model from data in BigQuery.
## Set up environment variables and load necessary libraries
```
PROJECT = !(gcloud config get-value core/project)
PROJECT = PROJECT[0]
%env PROJECT = {PROJECT}
%env REGION = "us-central1"
```
## Create the dataset
```
from google.cloud import bigquery
from IPython import get_ipython
bq = bigquery.Client(project=PROJECT)
def create_dataset():
dataset = bigquery.Dataset(bq.dataset("stock_market"))
try:
bq.create_dataset(dataset) # Will fail if dataset already exists.
print("Dataset created")
except:
print("Dataset already exists")
def create_features_table():
error = None
try:
bq.query(
"""
CREATE TABLE stock_market.eps_percent_change_sp500
AS
SELECT *
FROM `stock_market.eps_percent_change_sp500`
"""
).to_dataframe()
except Exception as e:
error = str(e)
if error is None:
print("Table created")
elif "Already Exists" in error:
print("Table already exists.")
else:
print(error)
raise Exception("Table was not created.")
create_dataset()
create_features_table()
```
## Review the dataset
In the previous lab we created the data we will use modeling and saved them as tables in BigQuery. Let's examine that table again to see that everything is as we expect. Then, we will build a model using BigQuery ML using this table.
```
%%bigquery --project $PROJECT
#standardSQL
SELECT
*
FROM
stock_market.eps_percent_change_sp500
LIMIT
10
```
## Using BQML
### Create classification model for `direction`
To create a model
1. Use `CREATE MODEL` and provide a destination table for resulting model. Alternatively we can use `CREATE OR REPLACE MODEL` which allows overwriting an existing model.
2. Use `OPTIONS` to specify the model type (linear_reg or logistic_reg). There are many more options [we could specify](https://cloud.google.com/bigquery/docs/reference/standard-sql/bigqueryml-syntax-create#model_option_list), such as regularization and learning rate, but we'll accept the defaults.
3. Provide the query which fetches the training data
Have a look at [Step Two of this tutorial](https://cloud.google.com/bigquery/docs/bigqueryml-natality) to see another example.
**The query will take about two minutes to complete**
We'll start with creating a classification model to predict the `direction` of each stock.
We'll take a random split using the `symbol` value. With about 500 different values, using `ABS(MOD(FARM_FINGERPRINT(symbol), 15)) = 1` will give 30 distinct `symbol` values which corresponds to about 171,000 training examples. After taking 70% for training, we will be building a model on about 110,000 training examples.
```
%%bigquery --project $PROJECT
#standardSQL
CREATE OR REPLACE MODEL
stock_market.direction_model OPTIONS(model_type = "logistic_reg",
input_label_cols = ["direction"]) AS
-- query to fetch training data
SELECT
symbol,
Date,
Open,
close_MIN_prior_5_days,
close_MIN_prior_20_days,
close_MIN_prior_260_days,
close_MAX_prior_5_days,
close_MAX_prior_20_days,
close_MAX_prior_260_days,
close_AVG_prior_5_days,
close_AVG_prior_20_days,
close_AVG_prior_260_days,
close_STDDEV_prior_5_days,
close_STDDEV_prior_20_days,
close_STDDEV_prior_260_days,
direction
FROM
`stock_market.eps_percent_change_sp500`
WHERE
tomorrow_close IS NOT NULL
AND ABS(MOD(FARM_FINGERPRINT(symbol), 15)) = 1
AND ABS(MOD(FARM_FINGERPRINT(symbol), 15 * 100)) <= 15 * 70
```
## Get training statistics and examine training info
After creating our model, we can evaluate the performance using the [`ML.EVALUATE` function](https://cloud.google.com/bigquery-ml/docs/bigqueryml-natality#step_four_evaluate_your_model). With this command, we can find the precision, recall, accuracy F1-score and AUC of our classification model.
```
%%bigquery --project $PROJECT
#standardSQL
SELECT
*
FROM
ML.EVALUATE(MODEL `stock_market.direction_model`,
(
SELECT
symbol,
Date,
Open,
close_MIN_prior_5_days,
close_MIN_prior_20_days,
close_MIN_prior_260_days,
close_MAX_prior_5_days,
close_MAX_prior_20_days,
close_MAX_prior_260_days,
close_AVG_prior_5_days,
close_AVG_prior_20_days,
close_AVG_prior_260_days,
close_STDDEV_prior_5_days,
close_STDDEV_prior_20_days,
close_STDDEV_prior_260_days,
direction
FROM
`stock_market.eps_percent_change_sp500`
WHERE
tomorrow_close IS NOT NULL
AND ABS(MOD(FARM_FINGERPRINT(symbol), 15)) = 1
AND ABS(MOD(FARM_FINGERPRINT(symbol), 15 * 100)) > 15 * 70
AND ABS(MOD(FARM_FINGERPRINT(symbol), 15 * 100)) <= 15 * 85))
```
We can also examine the training statistics collected by Big Query. To view training results we use the [`ML.TRAINING_INFO`](https://cloud.google.com/bigquery/docs/reference/standard-sql/bigqueryml-syntax-train) function.
```
%%bigquery --project $PROJECT
#standardSQL
SELECT
*
FROM
ML.TRAINING_INFO(MODEL `stock_market.direction_model`)
ORDER BY iteration
```
### Compare to simple benchmark
Another way to asses the performance of our model is to compare with a simple benchmark. We can do this by seeing what kind of accuracy we would get using the naive strategy of just predicted the majority class. For the training dataset, the majority class is 'STAY'. The following query we can see how this naive strategy would perform on the eval set.
```
%%bigquery --project $PROJECT
#standardSQL
WITH
eval_data AS (
SELECT
symbol,
Date,
Open,
close_MIN_prior_5_days,
close_MIN_prior_20_days,
close_MIN_prior_260_days,
close_MAX_prior_5_days,
close_MAX_prior_20_days,
close_MAX_prior_260_days,
close_AVG_prior_5_days,
close_AVG_prior_20_days,
close_AVG_prior_260_days,
close_STDDEV_prior_5_days,
close_STDDEV_prior_20_days,
close_STDDEV_prior_260_days,
direction
FROM
`stock_market.eps_percent_change_sp500`
WHERE
tomorrow_close IS NOT NULL
AND ABS(MOD(FARM_FINGERPRINT(symbol), 15)) = 1
AND ABS(MOD(FARM_FINGERPRINT(symbol), 15 * 100)) > 15 * 70
AND ABS(MOD(FARM_FINGERPRINT(symbol), 15 * 100)) <= 15 * 85)
SELECT
direction,
(COUNT(direction)* 100 / (
SELECT
COUNT(*)
FROM
eval_data)) AS percentage
FROM
eval_data
GROUP BY
direction
```
So, the naive strategy of just guessing the majority class would have accuracy of 0.5509 on the eval dataset, just below our BQML model.
### Create regression model for `normalized change`
We can also use BigQuery to train a regression model to predict the normalized change for each stock. To do this in BigQuery we need only change the OPTIONS when calling `CREATE OR REPLACE MODEL`. This will give us a more precise prediction rather than just predicting if the stock will go up, down, or stay the same. Thus, we can treat this problem as either a regression problem or a classification problem, depending on the business needs.
```
%%bigquery --project $PROJECT
#standardSQL
CREATE OR REPLACE MODEL
stock_market.price_model OPTIONS(model_type = "linear_reg",
input_label_cols = ["normalized_change"]) AS
-- query to fetch training data
SELECT
symbol,
Date,
Open,
close_MIN_prior_5_days,
close_MIN_prior_20_days,
close_MIN_prior_260_days,
close_MAX_prior_5_days,
close_MAX_prior_20_days,
close_MAX_prior_260_days,
close_AVG_prior_5_days,
close_AVG_prior_20_days,
close_AVG_prior_260_days,
close_STDDEV_prior_5_days,
close_STDDEV_prior_20_days,
close_STDDEV_prior_260_days,
normalized_change
FROM
`stock_market.eps_percent_change_sp500`
WHERE
normalized_change IS NOT NULL
AND ABS(MOD(FARM_FINGERPRINT(symbol), 15)) = 1
AND ABS(MOD(FARM_FINGERPRINT(symbol), 15 * 100)) <= 15 * 70
```
Just as before we can examine the evaluation metrics for our regression model and examine the training statistics in Big Query
```
%%bigquery --project $PROJECT
#standardSQL
SELECT
*
FROM
ML.EVALUATE(MODEL `stock_market.price_model`,
(
SELECT
symbol,
Date,
Open,
close_MIN_prior_5_days,
close_MIN_prior_20_days,
close_MIN_prior_260_days,
close_MAX_prior_5_days,
close_MAX_prior_20_days,
close_MAX_prior_260_days,
close_AVG_prior_5_days,
close_AVG_prior_20_days,
close_AVG_prior_260_days,
close_STDDEV_prior_5_days,
close_STDDEV_prior_20_days,
close_STDDEV_prior_260_days,
normalized_change
FROM
`stock_market.eps_percent_change_sp500`
WHERE
normalized_change IS NOT NULL
AND ABS(MOD(FARM_FINGERPRINT(symbol), 15)) = 1
AND ABS(MOD(FARM_FINGERPRINT(symbol), 15 * 100)) > 15 * 70
AND ABS(MOD(FARM_FINGERPRINT(symbol), 15 * 100)) <= 15 * 85))
%%bigquery --project $PROJECT
#standardSQL
SELECT
*
FROM
ML.TRAINING_INFO(MODEL `stock_market.price_model`)
ORDER BY iteration
```
## Train a Time Series model using AutoML Tables
### Step 1. Launch AutoML
Within the GCP console, navigate to Tables in the console menu.
<img src='../assets/console_menu_tables.png' width='50%'>
Click **Enable API**, if API is not enabled.
Click **GET STARTED**.
### Step 2. Create a Dataset
Select **New Dataset** and give it a name like `stock_market` and click *Create Dataset*. In the section on Importing data, select the option to import your data from a BigQuery Table. Fill in the details for your project, the dataset ID, and the table ID.
<img src='../assets/import_data_options.png' width='50%'>
### Step 3. Import the Data
Once you have created the dataset you can then import the data. This will take a few minutes.
<img src='../assets/importing_data.png' width='50%'>
### Step 4. Train the model
Once the data has been imported into the dataset. You can examine the Schema of your data, Analyze the properties and values of the features and ultimately Train the model. Here you can also determine the label column and features for training the model. Since we are doing a classifcation model, we'll use `direction` as our target column.
<img src='../assets/schema_analyze_train.png' width='80%'>
Under the `Train` tab, click **Train Model**. You can choose the features to use when training. Select the same features as we used above.
<img src='../assets/train_model.png' width='50%'>
### Step 5. Evaluate your model.
Training can take many hours. But once training is complete you can inspect the evaluation metrics of your model. Since this is a classification task, we can also adjust the threshold and explore how different thresholds will affect your evaluation metrics. Also on that page, we can explore the feature importance of the various features used in the model and view confusion matrix for our model predictions.
<img src='../assets/eval_metrics.png' width='80%'>
### Step 6. Predict with the trained model.
Once the model is done training, navigate to the Models page and Deploy the model, so we can test prediction.
<img src='../assets/deploy_model.png' width='80%'>
When calling predictions, you can call batch prediction jobs by specifying a BigQuery table or csv file. Or you can do online prediction for a single instance.
Copyright 2019 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| github_jupyter |
## GWAS Tutorial
This notebook is designed to provide a broad overview of Hail's functionality, with emphasis on the functionality to manipulate and query a genetic dataset. We walk through a genome-wide SNP association test, and demonstrate the need to control for confounding caused by population stratification.
```
import hail as hl
hl.init()
```
If the above cell ran without error, we're ready to go!
Before using Hail, we import some standard Python libraries for use throughout the notebook.
```
from hail.plot import show
from pprint import pprint
hl.plot.output_notebook()
```
### Download public 1000 Genomes data
We use a small chunk of the public 1000 Genomes dataset, created by downsampling the genotyped SNPs in the full VCF to about 20 MB. We will also integrate sample and variant metadata from separate text files.
These files are hosted by the Hail team in a public Google Storage bucket; the following cell downloads that data locally.
```
hl.utils.get_1kg('data/')
```
### Importing data from VCF
The data in a VCF file is naturally represented as a Hail [MatrixTable](https://hail.is/docs/0.2/hail.MatrixTable.html#hail.MatrixTable). By first importing the VCF file and then writing the resulting MatrixTable in Hail's native file format, all downstream operations on the VCF's data will be MUCH faster.
```
hl.import_vcf('data/1kg.vcf.bgz').write('data/1kg.mt', overwrite=True)
```
Next we read the written file, assigning the variable `mt` (for `m`atrix `t`able).
```
mt = hl.read_matrix_table('data/1kg.mt')
```
### Getting to know our data
It's important to have easy ways to slice, dice, query, and summarize a dataset. Some of this functionality is demonstrated below.
The [rows](https://hail.is/docs/0.2/hail.MatrixTable.html#hail.MatrixTable.rows) method can be used to get a table with all the row fields in our MatrixTable.
We can use `rows` along with [select](https://hail.is/docs/0.2/hail.Table.html#hail.Table.select) to pull out 5 variants. The `select` method takes either a string refering to a field name in the table, or a Hail [Expression](https://hail.is/docs/0.2/hail.expr.Expression.html?#expression). Here, we leave the arguments blank to keep only the row key fields, `locus` and `alleles`.
Use the `show` method to display the variants.
```
mt.rows().select().show(5)
```
Alternatively:
```
mt.row_key.show(5)
```
Here is how to peek at the first few sample IDs:
```
mt.s.show(5)
```
To look at the first few genotype calls, we can use [entries](https://hail.is/docs/0.2/hail.MatrixTable.html#hail.MatrixTable.entries) along with `select` and `take`. The `take` method collects the first n rows into a list. Alternatively, we can use the `show` method, which prints the first n rows to the console in a table format.
Try changing `take` to `show` in the cell below.
```
mt.entry.take(5)
```
### Adding column fields
A Hail MatrixTable can have any number of row fields and column fields for storing data associated with each row and column. Annotations are usually a critical part of any genetic study. Column fields are where you'll store information about sample phenotypes, ancestry, sex, and covariates. Row fields can be used to store information like gene membership and functional impact for use in QC or analysis.
In this tutorial, we demonstrate how to take a text file and use it to annotate the columns in a MatrixTable.
The file provided contains the sample ID, the population and "super-population" designations, the sample sex, and two simulated phenotypes (one binary, one discrete).
This file can be imported into Hail with [import_table](https://hail.is/docs/0.2/methods/impex.html#hail.methods.import_table). This function produces a [Table](https://hail.is/docs/0.2/hail.Table.html#hail.Table) object. Think of this as a Pandas or R dataframe that isn't limited by the memory on your machine -- behind the scenes, it's distributed with Spark.
```
table = (hl.import_table('data/1kg_annotations.txt', impute=True)
.key_by('Sample'))
```
A good way to peek at the structure of a `Table` is to look at its `schema`.
```
table.describe()
```
To peek at the first few values, use the `show` method:
```
table.show(width=100)
```
Now we'll use this table to add sample annotations to our dataset, storing the annotations in column fields in our MatrixTable. First, we'll print the existing column schema:
```
print(mt.col.dtype)
```
We use the [annotate_cols](https://hail.is/docs/0.2/hail.MatrixTable.html#hail.MatrixTable.annotate_cols) method to join the table with the MatrixTable containing our dataset.
```
mt = mt.annotate_cols(pheno = table[mt.s])
mt.col.describe()
```
### Query functions and the Hail Expression Language
Hail has a number of useful query functions that can be used for gathering statistics on our dataset. These query functions take Hail Expressions as arguments.
We will start by looking at some statistics of the information in our table. The [aggregate](https://hail.is/docs/0.2/hail.Table.html#hail.Table.aggregate) method can be used to aggregate over rows of the table.
`counter` is an aggregation function that counts the number of occurrences of each unique element. We can use this to pull out the population distribution by passing in a Hail Expression for the field that we want to count by.
```
pprint(table.aggregate(hl.agg.counter(table.SuperPopulation)))
```
`stats` is an aggregation function that produces some useful statistics about numeric collections. We can use this to see the distribution of the CaffeineConsumption phenotype.
```
pprint(table.aggregate(hl.agg.stats(table.CaffeineConsumption)))
```
However, these metrics aren't perfectly representative of the samples in our dataset. Here's why:
```
table.count()
mt.count_cols()
```
Since there are fewer samples in our dataset than in the full thousand genomes cohort, we need to look at annotations on the dataset. We can use [aggregate_cols](https://hail.is/docs/0.2/hail.MatrixTable.html#hail.MatrixTable.aggregate_cols) to get the metrics for only the samples in our dataset.
```
mt.aggregate_cols(hl.agg.counter(mt.pheno.SuperPopulation))
pprint(mt.aggregate_cols(hl.agg.stats(mt.pheno.CaffeineConsumption)))
```
The functionality demonstrated in the last few cells isn't anything especially new: it's certainly not difficult to ask these questions with Pandas or R dataframes, or even Unix tools like `awk`. But Hail can use the same interfaces and query language to analyze collections that are much larger, like the set of variants.
Here we calculate the counts of each of the 12 possible unique SNPs (4 choices for the reference base * 3 choices for the alternate base).
To do this, we need to get the alternate allele of each variant and then count the occurences of each unique ref/alt pair. This can be done with Hail's `counter` function.
```
snp_counts = mt.aggregate_rows(hl.agg.counter(hl.Struct(ref=mt.alleles[0], alt=mt.alleles[1])))
pprint(snp_counts)
```
We can list the counts in descending order using Python's Counter class.
```
from collections import Counter
counts = Counter(snp_counts)
counts.most_common()
```
It's nice to see that we can actually uncover something biological from this small dataset: we see that these frequencies come in pairs. C/T and G/A are actually the same mutation, just viewed from from opposite strands. Likewise, T/A and A/T are the same mutation on opposite strands. There's a 30x difference between the frequency of C/T and A/T SNPs. Why?
The same Python, R, and Unix tools could do this work as well, but we're starting to hit a wall - the latest [gnomAD release](https://gnomad.broadinstitute.org/) publishes about 250 million variants, and that won't fit in memory on a single computer.
What about genotypes? Hail can query the collection of all genotypes in the dataset, and this is getting large even for our tiny dataset. Our 284 samples and 10,000 variants produce 10 million unique genotypes. The gnomAD dataset has about **5 trillion** unique genotypes.
Hail plotting functions allow Hail fields as arguments, so we can pass in the DP field directly here. If the range and bins arguments are not set, this function will compute the range based on minimum and maximum values of the field and use the default 50 bins.
```
p = hl.plot.histogram(mt.DP, range=(0,30), bins=30, title='DP Histogram', legend='DP')
show(p)
```
### Quality Control
QC is where analysts spend most of their time with sequencing datasets. QC is an iterative process, and is different for every project: there is no "push-button" solution for QC. Each time the Broad collects a new group of samples, it finds new batch effects. However, by practicing open science and discussing the QC process and decisions with others, we can establish a set of best practices as a community.
QC is entirely based on the ability to understand the properties of a dataset. Hail attempts to make this easier by providing the [sample_qc](https://hail.is/docs/0.2/methods/genetics.html#hail.methods.sample_qc) function, which produces a set of useful metrics and stores them in a column field.
```
mt.col.describe()
mt = hl.sample_qc(mt)
mt.col.describe()
```
Plotting the QC metrics is a good place to start.
```
p = hl.plot.histogram(mt.sample_qc.call_rate, range=(.88,1), legend='Call Rate')
show(p)
p = hl.plot.histogram(mt.sample_qc.gq_stats.mean, range=(10,70), legend='Mean Sample GQ')
show(p)
```
Often, these metrics are correlated.
```
p = hl.plot.scatter(mt.sample_qc.dp_stats.mean, mt.sample_qc.call_rate, xlabel='Mean DP', ylabel='Call Rate')
show(p)
```
Removing outliers from the dataset will generally improve association results. We can make arbitrary cutoffs and use them to filter:
```
mt = mt.filter_cols((mt.sample_qc.dp_stats.mean >= 4) & (mt.sample_qc.call_rate >= 0.97))
print('After filter, %d/284 samples remain.' % mt.count_cols())
```
Next is genotype QC. It's a good idea to filter out genotypes where the reads aren't where they should be: if we find a genotype called homozygous reference with >10% alternate reads, a genotype called homozygous alternate with >10% reference reads, or a genotype called heterozygote without a ref / alt balance near 1:1, it is likely to be an error.
In a low-depth dataset like 1KG, it is hard to detect bad genotypes using this metric, since a read ratio of 1 alt to 10 reference can easily be explained by binomial sampling. However, in a high-depth dataset, a read ratio of 10:100 is a sure cause for concern!
```
ab = mt.AD[1] / hl.sum(mt.AD)
filter_condition_ab = ((mt.GT.is_hom_ref() & (ab <= 0.1)) |
(mt.GT.is_het() & (ab >= 0.25) & (ab <= 0.75)) |
(mt.GT.is_hom_var() & (ab >= 0.9)))
fraction_filtered = mt.aggregate_entries(hl.agg.fraction(~filter_condition_ab))
print(f'Filtering {fraction_filtered * 100:.2f}% entries out of downstream analysis.')
mt = mt.filter_entries(filter_condition_ab)
```
Variant QC is a bit more of the same: we can use the [variant_qc](https://hail.is/docs/0.2/methods/genetics.html#hail.methods.variant_qc) function to produce a variety of useful statistics, plot them, and filter.
```
mt = hl.variant_qc(mt)
mt.row.describe()
```
These statistics actually look pretty good: we don't need to filter this dataset. Most datasets require thoughtful quality control, though. The [filter_rows](https://hail.is/docs/0.2/hail.MatrixTable.html#hail.MatrixTable.filter_rows) method can help!
### Let's do a GWAS!
First, we need to restrict to variants that are :
- common (we'll use a cutoff of 1%)
- not so far from [Hardy-Weinberg equilibrium](https://en.wikipedia.org/wiki/Hardy%E2%80%93Weinberg_principle) as to suggest sequencing error
```
mt = mt.filter_rows(mt.variant_qc.AF[1] > 0.01)
mt = mt.filter_rows(mt.variant_qc.p_value_hwe > 1e-6)
print('Samples: %d Variants: %d' % (mt.count_cols(), mt.count_rows()))
```
These filters removed about 15% of sites (we started with a bit over 10,000). This is _NOT_ representative of most sequencing datasets! We have already downsampled the full thousand genomes dataset to include more common variants than we'd expect by chance.
In Hail, the association tests accept column fields for the sample phenotype and covariates. Since we've already got our phenotype of interest (caffeine consumption) in the dataset, we are good to go:
```
gwas = hl.linear_regression_rows(y=mt.pheno.CaffeineConsumption,
x=mt.GT.n_alt_alleles(),
covariates=[1.0])
gwas.row.describe()
```
Looking at the bottom of the above printout, you can see the linear regression adds new row fields for the beta, standard error, t-statistic, and p-value.
Hail makes it easy to visualize results! Let's make a [Manhattan plot](https://en.wikipedia.org/wiki/Manhattan_plot):
```
p = hl.plot.manhattan(gwas.p_value)
show(p)
```
This doesn't look like much of a skyline. Let's check whether our GWAS was well controlled using a [Q-Q (quantile-quantile) plot](https://en.wikipedia.org/wiki/Q–Q_plot).
```
p = hl.plot.qq(gwas.p_value)
show(p)
```
### Confounded!
The observed p-values drift away from the expectation immediately. Either every SNP in our dataset is causally linked to caffeine consumption (unlikely), or there's a confounder.
We didn't tell you, but sample ancestry was actually used to simulate this phenotype. This leads to a [stratified](https://en.wikipedia.org/wiki/Population_stratification) distribution of the phenotype. The solution is to include ancestry as a covariate in our regression.
The [linear_regression_rows](https://hail.is/docs/0.2/methods/stats.html#hail.methods.linear_regression_rows) function can also take column fields to use as covariates. We already annotated our samples with reported ancestry, but it is good to be skeptical of these labels due to human error. Genomes don't have that problem! Instead of using reported ancestry, we will use genetic ancestry by including computed principal components in our model.
The [pca](https://hail.is/docs/0.2/methods/stats.html#hail.methods.pca) function produces eigenvalues as a list and sample PCs as a Table, and can also produce variant loadings when asked. The [hwe_normalized_pca](https://hail.is/docs/0.2/methods/genetics.html#hail.methods.hwe_normalized_pca) function does the same, using HWE-normalized genotypes for the PCA.
```
eigenvalues, pcs, _ = hl.hwe_normalized_pca(mt.GT)
pprint(eigenvalues)
pcs.show(5, width=100)
```
Now that we've got principal components per sample, we may as well plot them! Human history exerts a strong effect in genetic datasets. Even with a 50MB sequencing dataset, we can recover the major human populations.
```
mt = mt.annotate_cols(scores = pcs[mt.s].scores)
p = hl.plot.scatter(mt.scores[0],
mt.scores[1],
label=mt.pheno.SuperPopulation,
title='PCA', xlabel='PC1', ylabel='PC2')
show(p)
```
Now we can rerun our linear regression, controlling for sample sex and the first few principal components. We'll do this with input variable the number of alternate alleles as before, and again with input variable the genotype dosage derived from the PL field.
```
gwas = hl.linear_regression_rows(
y=mt.pheno.CaffeineConsumption,
x=mt.GT.n_alt_alleles(),
covariates=[1.0, mt.pheno.isFemale, mt.scores[0], mt.scores[1], mt.scores[2]])
```
We'll first make a Q-Q plot to assess inflation...
```
p = hl.plot.qq(gwas.p_value)
show(p)
```
That's more like it! This shape is indicative of a well-controlled (but not especially well-powered) study. And now for the Manhattan plot:
```
p = hl.plot.manhattan(gwas.p_value)
show(p)
```
We have found a caffeine consumption locus! Now simply apply Hail's Nature paper function to publish the result.
Just kidding, that function won't land until Hail 1.0!
### Rare variant analysis
Here we'll demonstrate how one can use the expression language to group and count by any arbitrary properties in row and column fields. Hail also implements the sequence kernel association test (SKAT).
```
entries = mt.entries()
results = (entries.group_by(pop = entries.pheno.SuperPopulation, chromosome = entries.locus.contig)
.aggregate(n_het = hl.agg.count_where(entries.GT.is_het())))
results.show()
```
What if we want to group by minor allele frequency bin and hair color, and calculate the mean GQ?
```
entries = entries.annotate(maf_bin = hl.if_else(entries.info.AF[0]<0.01, "< 1%",
hl.if_else(entries.info.AF[0]<0.05, "1%-5%", ">5%")))
results2 = (entries.group_by(af_bin = entries.maf_bin, purple_hair = entries.pheno.PurpleHair)
.aggregate(mean_gq = hl.agg.stats(entries.GQ).mean,
mean_dp = hl.agg.stats(entries.DP).mean))
results2.show()
```
We've shown that it's easy to aggregate by a couple of arbitrary statistics. This specific examples may not provide especially useful pieces of information, but this same pattern can be used to detect effects of rare variation:
- Count the number of heterozygous genotypes per gene by functional category (synonymous, missense, or loss-of-function) to estimate per-gene functional constraint
- Count the number of singleton loss-of-function mutations per gene in cases and controls to detect genes involved in disease
### Epilogue
Congrats! You've reached the end of the first tutorial. To learn more about Hail's API and functionality, take a look at the other tutorials. You can check out the [Python API](https://hail.is/docs/0.2/api.html#python-api) for documentation on additional Hail functions. If you use Hail for your own science, we'd love to hear from you on [Zulip chat](https://hail.zulipchat.com) or the [discussion forum](https://discuss.hail.is).
For reference, here's the full workflow to all tutorial endpoints combined into one cell.
```
table = hl.import_table('data/1kg_annotations.txt', impute=True).key_by('Sample')
mt = hl.read_matrix_table('data/1kg.mt')
mt = mt.annotate_cols(pheno = table[mt.s])
mt = hl.sample_qc(mt)
mt = mt.filter_cols((mt.sample_qc.dp_stats.mean >= 4) & (mt.sample_qc.call_rate >= 0.97))
ab = mt.AD[1] / hl.sum(mt.AD)
filter_condition_ab = ((mt.GT.is_hom_ref() & (ab <= 0.1)) |
(mt.GT.is_het() & (ab >= 0.25) & (ab <= 0.75)) |
(mt.GT.is_hom_var() & (ab >= 0.9)))
mt = mt.filter_entries(filter_condition_ab)
mt = hl.variant_qc(mt)
mt = mt.filter_rows(mt.variant_qc.AF[1] > 0.01)
eigenvalues, pcs, _ = hl.hwe_normalized_pca(mt.GT)
mt = mt.annotate_cols(scores = pcs[mt.s].scores)
gwas = hl.linear_regression_rows(
y=mt.pheno.CaffeineConsumption,
x=mt.GT.n_alt_alleles(),
covariates=[1.0, mt.pheno.isFemale, mt.scores[0], mt.scores[1], mt.scores[2]])
```
| github_jupyter |
# **Distance Based Classification Techniques on IRIS Dataset**
<br>
**Iris Dataset**
The Fisher's Iris data set contains attributes of three Iris species, namely, Iris Setosa, Iris versicolor, and Iris virginica. Each species (class) consists of 50 samples, each having four features, Sepal length, Sepal width, Petal length and Petal width.
<br>
**Distance based classification**
Classification is the process of categorizing an input data using a machine learning model. The Distance based classification technique is a supervised learning process whose goal is to identify a flower's species based on a previously trained dataset. The training takes place by finding the centroid of each class which is later used to assign a label to the data point based on its distance from the centroid. The one with the shortest distance is taken as the label.
## **Preprocessing**
Import all necessary packages and define constants. Numpy and pandas will come handy for matrix operations and data visualization respectively. Matplotlib and scikit-learn will be useful for plotting. Scikit-learn is a free machine learning library for Python.
```
import matplotlib.pyplot as plt
from sklearn import datasets
from sklearn import preprocessing
from sklearn.metrics import mean_squared_error as mse
import statistics as st
import numpy as np
import pandas as pd
import sys
MAX = sys.maxsize
```
## **Types of Distances**
**Euclidean Distance**
>$d(p_{x1,y1},q_{x2,y2}) = \sqrt{(x_1-x_2)^2+(y_1-y_2)^2}$
```
def euclidean_distance(p1,p2):
distance = pow(sum([(a - b) ** 2 for a, b in zip(p1, p2)]),0.5)
return distance
```
**Manhattan Distance or City Block Distance**
>$d(p_{x1,y1},q_{x2,y2}) = |x_1 - x_2| + |y_1 - y_2|$
```
def manhattan_distance(p1,p2):
distance = 0
for i in range(len(p1)):
distance += abs(p1[i] - p2[i])
return distance
```
**Chessboard Distance or Chebyshev Distance**
>$d(p_{x1,y1},q_{x2,y2}) = max(|x_1 - x_2|,|y_1 - y_2|)$
```
def chessboard_distance(p1,p2):
distance = abs(p1[0] - p2[0])
for i in range(1,len(p1)):
distance = max(distance,abs(p1[i] - p2[i]))
return distance
```
**Minkowski Distance**
>$d(u_{x1,y1},v_{x2,y2}) = (|x_1 - x_2|^p + |y_1 - y_2|^p)^{1/p}$
```
def minkowski_distance(p1,p2,p):
s = 0
for i in range(len(p1)):
s += abs(p1[i] - p2[i])**p
distance = s**(1/p)
return distance
```
**Cosine Distance**
>$\text{cosine distance} = 1 - \text{cosine similarity}(A,B)$
>$\text{cosine similarity} = \cos(\theta) = {\mathbf{A} \cdot \mathbf{B} \over \|\mathbf{A}\| \|\mathbf{B}\|} = \frac{ x_1 * x_2 + y_1 * y_2 }{ \sqrt{x_1^2+y_1^2 } \sqrt{x_2^2 + y_2^2}}$
```
def cosine_distance(p1,p2):
norm_p1 = 0
norm_p2 = 0
for i in range(len(p1)):
norm_p1 += p1[i]**2
norm_p2 += p2[i]**2
norm_p1 = norm_p1**0.5
norm_p2 = norm_p2**0.5
s = 0
for i in range(len(p1)):
s += p1[i]*p2[i]
distance = 1 - s/(norm_p1*norm_p2)
return distance
```
**Correlation Distance**
>$CD = 1 - (u - Mean[u]).(v - Mean[v])/
(Norm(u - Mean[u])Norm(v - Mean[v]))$
```
def correlation_distance(p1,p2):
norm_p1 = 0
norm_p2 = 0
for i in range(len(p1)):
norm_p1 += (p1[i] - st.mean(p1))**2
norm_p2 += (p2[i] - st.mean(p2))**2
norm_p1 = norm_p1**0.5
norm_p2 = norm_p2**0.5
s = 0
for i in range(len(p1)):
s += (p1[i] - st.mean(p1))*(p2[i] - st.mean(p2))
distance = 1 - s/(norm_p1*norm_p2)
return distance
```
**Bray Curtis Distance**
>$BCD[u_{a,b},v_{x,y}] = Total[|(a,b) - (x,y)|]/Total[|(a,b) + (x,y)|]$
```
def bray_curtis_distance(p1,p2):
s1 = 0
s2 = 0
for i in range(len(p1)):
s1 += abs(p1[i] - p2[i])
s2 += abs(p1[i] + p2[i])
distance = s1/s2
return distance
```
**Canberra Distance**
$CAD[u_{a,b},v_{x,y}] = Total[|(a,b) - (x,y)|/|(a,b) + (x,y)|]$
```
def canberra_distance(p1,p2):
distance = 0
for i in range(len(p1)):
s1 = abs(p1[i] - p2[i])
s2 = abs(p1[i] + p2[i])
distance += s1/s2
return distance
```
### **Loading Iris Dataset**
```
# iris dataset has three classes
iris = datasets.load_iris()
df = pd.DataFrame(iris.data, columns = [iris.feature_names])
print(df)
```
**Plotting Species based on Petal Length and Petal Width**
```
X_petal = iris.data[:, 2:4];
setosa_plot = plt.scatter(X_petal[0:50,0], X_petal[0:50,1],color='red')
versicolor_plot = plt.scatter(X_petal[50:100,0], X_petal[50:100,1],color='green')
virginica_plot = plt.scatter(X_petal[100:150,0], X_petal[100:150,1],color='blue')
plt.legend((setosa_plot,versicolor_plot,virginica_plot),('Setosa', 'Versicolor','Virginica'),numpoints=1, loc='lower right', ncol=3, fontsize=8)
plt.title("Plotting Species based Petal Length and Petal Width")
plt.xlabel('Petal Length (cm)')
plt.ylabel('Petal Width (cm)')
plt.show()
```
**Plotting Species based on Sepal Length and Sepal Width**
```
X_sepal = iris.data[:, :2];
setosa_plot = plt.scatter(X_sepal[0:50,0], X_sepal[0:50,1],color='red')
versicolor_plot = plt.scatter(X_sepal[50:100,0], X_sepal[50:100,1],color='green')
virginica_plot = plt.scatter(X_sepal[100:150,0], X_sepal[100:150,1],color='blue')
plt.legend((setosa_plot,versicolor_plot,virginica_plot),('Setosa', 'Versicolor','Virginica'),numpoints=1, loc='lower right', ncol=3, fontsize=8)
plt.title("Plotting Species based Sepal Length and Sepal Width")
plt.xlabel('Sepal Length (cm)')
plt.ylabel('Sepal Width (cm)')
plt.show()
```
To start the training process, we will be requiring two features. These features will allow us to plot the sample data as well help in classifying them. We also need true class labels to group these points according to their classes.
```
X = iris.data[:, 2:4] # last two features, petal length and petal width
y = iris.target # contains true class labels
print('True Class Labels')
print(y)
print()
# Class Names 0 Setosa, 1 Versicolor, 2 Virginica
tg_names = iris.target_names;
print('Species')
print(tg_names)
```
## **Training Phase**
```
# Distance Based Classification
########################## Training Phase ##########################
# Provide 60% of instances
# Class 0 has instances from 0 to 49
# We need instances from 0 to 29 for training
C0_x = X[0:30,0]
C0_y = X[0:30,1]
# Class 1 has instances from 50 to 99
# We need instances from 50 to 79 for training
C1_x = X[50:80,0]
C1_y = X[50:80,1]
# Class 2 has instances from 100 to 149
# We need instances from 100 to 129 for training
C2_x = X[100:130,0]
C2_y = X[100:130,1]
# Finding centroid for each class
# Centroid of C0
c0_mean_x = st.mean(C0_x)
c0_mean_y = st.mean(C0_y)
# Centroid of C1
c1_mean_x = st.mean(C1_x)
c1_mean_y = st.mean(C1_y)
# Centroid of C2
c2_mean_x = st.mean(C2_x)
c2_mean_y = st.mean(C2_y)
# Placing all centroids together for simplicity
all_centroid = np.array([[c0_mean_x, c0_mean_y],
[c1_mean_x, c1_mean_y],
[c2_mean_x, c2_mean_y]])
```
## **Plotting centroid for each class**
```
X_petal = iris.data[:, 2:4];
setosa_plot = plt.scatter(X_petal[0:50,0], X_petal[0:50,1],color='red')
versicolor_plot = plt.scatter(X_petal[50:100,0], X_petal[50:100,1],color='green')
virginica_plot = plt.scatter(X_petal[100:150,0], X_petal[100:150,1],color='blue')
centroid_plot = plt.scatter(all_centroid[:,0],all_centroid[:,1],color='black')
plt.legend((setosa_plot,versicolor_plot,virginica_plot,centroid_plot),('Setosa', 'Versicolor','Virginica','Centroid'),numpoints=1, loc='lower right', ncol=3, fontsize=8)
plt.title("Plotting Species based on Petal Length and Petal Width")
plt.xlabel('Petal Length (cm)')
plt.ylabel('Petal Width (cm)')
plt.show()
```
## **Testing Phase**
```
########################## Testing Phase ##########################
# Provide 40% of instances
# Since we don't need class names now, we can simply merge all
# instances together
C_x = np.concatenate((X[30:50,0], X[80:100,0], X[130:150,0] ))
C_y = np.concatenate((X[30:50,1], X[80:100,1], X[130:150,1] ))
# minimum distance
min_dist = MAX
# predicted labels
predicted = [0]*len(C_x);
# actual labels
actual = np.concatenate((y[30:50], y[80:100], y[130:150] ))
for i in range (len(C_x)):
for j in range(0,3):
distance = euclidean_distance([all_centroid[j,0],all_centroid[j,1]],[C_x[i],C_y[i]])
if(min_dist > distance):
min_dist = distance
lbl = j;
predicted[i] = lbl;
#reset min_dist
min_dist = MAX
# Displaying Predicted Labels against True Labels
dict = {'True Labels' : actual,
'Predicted Labels' : predicted}
df2 = pd.DataFrame(dict)
print('Comparision of True Class Labels and Predicted Class Labels\n')
print(df2)
```
## **Distance Based Classifier Code**
In the above code, we have seen classification based on euclidean distance. The following code provides distance based classification based on different types of distance measurements.
The distance_based_classifier function takes in four parameters, feature matrix, true labels, type of distance and training rate. The output of the function is Predicted Labels, MER, Mean Squared Error and Mean Absolute error.
```
# Distance Based Classification
"""
@author: Ajay Biswas
220CS2184
M.Tech Information Security
National Institute of Technology, Rourkela
"""
from sklearn import datasets
import statistics as st
import numpy as np
import pandas as pd
from sklearn.metrics import mean_squared_error as mse
from sklearn.metrics import mean_absolute_error as mae
import sys
MAX = sys.maxsize
def MER_Error(X,Y):
correct_count = 0
for i in range(len(X)):
if(X[i] == Y[i]):
correct_count = correct_count + 1
MER_val = 1 - (correct_count/len(X))
return MER_val
def me(X,Y,d):
error=0
error_dist=0
for i in range(len(X)):
if(X[i] != Y[i]):
error += 1
error_dist+=d[i]
m_error =error_dist/error
return m_error
def euclidean_distance(p1,p2):
distance = pow(sum([(a - b) ** 2 for a, b in zip(p1, p2)]),0.5)
return distance
def manhattan_distance(p1,p2):
distance = 0
for i in range(len(p1)):
distance += abs(p1[i] - p2[i])
return distance
def chessboard_distance(p1,p2):
distance = abs(p1[0] - p2[0])
for i in range(1,len(p1)):
distance = max(distance,abs(p1[i] - p2[i]))
return distance
def correlation_distance(p1,p2):
norm_p1 = 0
norm_p2 = 0
for i in range(len(p1)):
norm_p1 += (p1[i] - st.mean(p1))**2
norm_p2 += (p2[i] - st.mean(p2))**2
norm_p1 = norm_p1**0.5
norm_p2 = norm_p2**0.5
s = 0
for i in range(len(p1)):
s += (p1[i] - st.mean(p1))*(p2[i] - st.mean(p2))
distance = 1 - s/(norm_p1*norm_p2)
return distance
def minkowski_distance(p1,p2,p):
s = 0
for i in range(len(p1)):
s += abs(p1[i] - p2[i])**p
distance = s**(1/p)
return distance
def cosine_distance(p1,p2):
norm_p1 = 0
norm_p2 = 0
for i in range(len(p1)):
norm_p1 += p1[i]**2
norm_p2 += p2[i]**2
norm_p1 = norm_p1**0.5
norm_p2 = norm_p2**0.5
s = 0
for i in range(len(p1)):
s += p1[i]*p2[i]
distance = 1 - s/(norm_p1*norm_p2)
return distance
def bray_curtis_distance(p1,p2):
s1 = 0
s2 = 0
for i in range(len(p1)):
s1 += abs(p1[i] - p2[i])
s2 += abs(p1[i] + p2[i])
distance = s1/s2
return distance
def canberra_distance(p1,p2):
distance = 0
for i in range(len(p1)):
s1 = abs(p1[i] - p2[i])
s2 = abs(p1[i] + p2[i])
distance += s1/s2
return distance
def select_distance(p1,p2,distance_type):
# returns the calculated distance based on the type of distance provided
if(distance_type == "euclidean"):
return euclidean_distance(p1,p2)
elif(distance_type == "manhattan" or distance_type == "cityblock"):
return manhattan_distance(p1,p2)
elif(distance_type == "chessboard" or distance_type == "chebyshev"):
return chessboard_distance(p1,p2)
elif(distance_type == "minkowski"):
return minkowski_distance(p1,p2,3)
elif(distance_type == "correlation"):
return correlation_distance(p1,p2)
elif(distance_type == "cosine"):
return cosine_distance(p1,p2)
elif(distance_type == "bray_curtis"):
return bray_curtis_distance(p1,p2)
elif(distance_type == "canberra"):
return canberra_distance(p1,p2)
else:
return None
def distance_based_classifier(X,y,d_type,tp):
# X is a 2D matrix with two columns as features and rows as instances
# y is the true class labels
# d_type is the type of distance taken for classifying
# tp is the fraction of training dataset, the fractional testing dataset will be (1-tp)
# Placing two features in two separate 2D arrays Species_x and Species_y
# the rows of this 2D array determines each separate class
cols = 1
rows = 0
for i in range(0,len(y)):
if(y[i]!=rows):
cols = 0
rows = rows + 1
cols = cols + 1;
rows = rows+1;
Species_x = np.zeros((rows, cols))
Species_y = np.zeros((rows, cols))
cnt = 0
for i in range(rows):
for j in range(cols):
Species_x[i][j] = X[cnt,0]
Species_y[i][j] = X[cnt,1]
cnt = cnt + 1
# rows = no. of classes
# cols = no. of instances in each class
########################## Training Phase ##########################
# invalid training dataset size
if(tp >= 1 or tp <= 0):
return None
percent = tp*100
# Slicing from beginning
train_range_s = 0
train_range_e = int((cols/100)*percent)
all_centroid = np.zeros((rows, 2))
# Taking mean of all points of each class and finding their centroid
for k in range(rows):
CL_x = Species_x[k][train_range_s:train_range_e]
CL_mean_x = st.mean(CL_x)
CL_y = Species_y[k][train_range_s:train_range_e]
CL_mean_y = st.mean(CL_y)
# (x,y) coordinates of centroid, column 0 - x, 1 - y
all_centroid[k,0] = CL_mean_x;
all_centroid[k,1] = CL_mean_y;
########################## Testing Phase ##########################
# Since we don't need class names now, we can simply merge all instances together
# Slicing after the last training instance
test_range_s = int((cols/100)*percent)
test_range_e = cols
C_x = np.zeros((rows, test_range_e - test_range_s))
C_y = np.zeros((rows, test_range_e - test_range_s))
for k in range(rows):
C_x[k][:] = Species_x[k][test_range_s:test_range_e]
C_y[k][:] = Species_y[k][test_range_s:test_range_e]
# Flattenning numpy array
C_x = C_x.flatten()
C_y = C_y.flatten()
# predicted labels
predicted = [0]*len(C_x);
# actual labels
# initially we keep labels in different rows w.r.t classes
# later we will flatten the array
actual = np.zeros((rows, test_range_e - test_range_s))
beg = test_range_s
end = cols
for k in range(rows):
actual[k][:] = y[beg:end]
beg = beg + cols
end = end + cols
# flatten the array
actual = actual.flatten()
# classifying points by measuring its distance from centroid of each class
distances_predicted = [0]*len(C_x)
min_dist = MAX
for i in range (len(C_x)):
for j in range(0,rows):
distance = select_distance([all_centroid[j,0],all_centroid[j,1]],[C_x[i],C_y[i]],d_type)
# invalid distance
if(distance == None):
return None
# finding the minimum distance
if(min_dist > distance):
min_dist = distance
distances_predicted[i] = distance
lbl = j;
# store predicted label
predicted[i] = lbl;
#reset min_dist
min_dist = MAX
# Calculating actual distances
distances_actual = [0]*len(C_x);
for i in range (len(C_x)):
distances_actual[i] = select_distance([all_centroid[int(actual[i]),0],all_centroid[int(actual[i]),1]],[C_x[i],C_y[i]],d_type)
# Accuracy Calculations
mer_error = MER_Error(actual,predicted)
mse_error = mse(distances_actual,distances_predicted)
mae_error = mae(distances_actual,distances_predicted)
mean_error = me(actual,predicted,distances_predicted)
return [predicted, mer_error,mse_error,mae_error,mean_error]
######################################################################################
# iris dataset has three classes with 50 instances each
iris = datasets.load_iris()
X = iris.data[:, :2] # first two features
y = iris.target # contains true class labels
euclidean = distance_based_classifier(X,y,"euclidean",0.6)
manhattan = distance_based_classifier(X,y,"manhattan",0.6)
chessboard = distance_based_classifier(X,y,"chessboard",0.6)
minkowski = distance_based_classifier(X,y,"minkowski",0.6)
cosine = distance_based_classifier(X,y,"cosine",0.6)
correlation = distance_based_classifier(X,y,"correlation",0.6)
chebyshev = distance_based_classifier(X,y,"chebyshev",0.6)
bray_curtis = distance_based_classifier(X,y,"bray_curtis",0.6)
canberra = distance_based_classifier(X,y,"canberra",0.6)
```
## **Estimated Errors**
```
errors = [ euclidean[1:], manhattan[1:], chessboard[1:], minkowski[1:], cosine[1:], correlation[1:], chebyshev[1:], bray_curtis[1:], canberra[1:]]
dff = pd.DataFrame(errors, columns = ['MER','MSE','MAE','Mean Error'],
index = ['Euclidean','Manhattan','Chessboard','Minkowski','Cosine','Correlation','Chebyshev','Bray Curtis','Canberra'])
print(dff)
```
**Misclassification Error Rate Vs. Classifier**
```
import matplotlib.pyplot as plt
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
error_type = ['Euclidean', 'Manhattan', 'Chessboard', 'Minkowski', 'Cosine','Correlation','Bray Curtis','Canberra']
dt = [euclidean[1],manhattan[1],chessboard[1],minkowski[1],cosine[1],correlation[1],bray_curtis[1],canberra[1]]
ax.bar(error_type,dt)
plt.title("Misclassification Error Rate Vs. Classifier")
plt.xlabel('Classifier Type')
plt.ylabel('MER')
plt.show()
```
**Mean Squared Error Rate Vs. Classifier**
```
import matplotlib.pyplot as plt
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
error_type = ['Euclidean', 'Manhattan', 'Chessboard', 'Minkowski', 'Cosine','Correlation','Bray Curtis','Canberra']
dt = [euclidean[2],manhattan[2],chessboard[2],minkowski[2],cosine[2],correlation[2],bray_curtis[2],canberra[2]]
ax.bar(error_type,dt)
plt.title("Mean Squared Error Rate Vs. Classifier")
plt.xlabel('Classifier Type')
plt.ylabel('Mean Squared Error Rate')
plt.show()
```
**Mean Absolute Rate Vs. Classifier**
```
import matplotlib.pyplot as plt
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
error_type = ['Euclidean', 'Manhattan', 'Chessboard', 'Minkowski', 'Cosine','Correlation','Bray Curtis','Canberra']
dt = [euclidean[3],manhattan[3],chessboard[3],minkowski[3],cosine[3],correlation[3],bray_curtis[3],canberra[3]]
ax.bar(error_type,dt)
plt.title("Mean Absolute Error Rate Vs. Classifier")
plt.xlabel('Classifier Type')
plt.ylabel('Mean Absolute Error Rate')
plt.show()
```
**Mean Error Rate Vs. Classifier**
```
import matplotlib.pyplot as plt
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
error_type = ['Euclidean', 'Manhattan', 'Chessboard', 'Minkowski', 'Cosine','Correlation','Bray Curtis','Canberra']
dt = [euclidean[4],manhattan[4],chessboard[4],minkowski[4],cosine[4],correlation[4],bray_curtis[4],canberra[4]]
ax.bar(error_type,dt)
plt.title("Mean Error Rate Vs. Classifier")
plt.xlabel('Classifier Type')
plt.ylabel('Mean Error Rate')
plt.show()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn import preprocessing
import time
from sklearn.model_selection import train_test_split
font = r'C:\Windows\Fonts\simfang.ttf'
data = pd.read_csv(r"./dataset/data/train.csv")
data
today = pd.to_datetime('2018-01-01')
data['birth_date'] = pd.to_datetime(data['birth_date'])
data['age'] = np.round((today - data['birth_date']).apply(lambda x: x.days) / 365.,1)
data
cols = ['potential','age','y']
data = pd.DataFrame(data,columns = cols)
median_num = data['y'].median()
data.loc[(data['y']>=median_num),'y'] = 1
data.loc[(data['y']!=1),'y'] = -1
data = data.sample(frac=1)
data = data[:1000]
feature = ['potential','age']
data_feature = pd.DataFrame(data,columns = feature)
data_target = pd.DataFrame(data,columns = ['y'])
#Max-Min标准化
#建立MinMaxScaler对象
minmax = preprocessing.MinMaxScaler()
# 标准化处理
data_feature = minmax.fit_transform(data_feature)
data_feature[:10]
X_train,X_test,Y_train,Y_test = train_test_split(data_feature, data_target, test_size=0.2, random_state=0)
X_train[:10]
Y_train = Y_train.values
Y_test = Y_test.values
data = data.values
# data
for i in range(len(data)):
if data[i][-1] == 1:
plt.scatter(data_feature[i][0],data_feature[i][1],c='r',marker='o')
else:
plt.scatter(data_feature[i][0],data_feature[i][1],c='b',marker='x')
plt.show()
# training = data_feature[:200]
# test = data_feature[200:]
# # print(len(test))
# training_cons = data[:200]
# test_cons = data[200:300]
# # training_cons[:10]
for i in range(len(X_train)):
if Y_train[i] == 1:
plt.scatter(X_train[i][0],X_train[i][1],c='r',marker='o')
else:
plt.scatter(X_train[i][0],X_train[i][1],c='b',marker='x')
plt.show()
for i in range(len(X_test)):
if Y_test[i] == 1:
plt.scatter(X_test[i][0],X_test[i][1],c='r',marker='o')
else:
plt.scatter(X_test[i][0],X_test[i][1],c='b',marker='x')
plt.show()
def sign(a):
if a>0:
return 1
else:
return -1
def feeling():
w = np.zeros(2)
lr = 0.01
c = 1
cnt = 0
while True:
cnt += 1
if cnt > 1000:
break
for i in range(len(X_train)):
x = np.array(X_train[i])
y = Y_train[i]
s = np.sum(w*x)+c
z = sign(s)
if y * z <= 0:
w = w+lr*y*x
c = c+lr*y
return cnt,w,c
if __name__=='__main__':
cnt,w,c=feeling()
print("迭代次数:",cnt)
print("超平面法向量:",w)
print("阈值:",c)
count = 0
for i in range(len(X_test)):
target = Y_test[i]
result = sign(np.sum(w*X_test[i])+c)
if(target == result):
count += 1
accuracy = float(count/len(X_test))
for i in range(len(X_test)):
if Y_test[i] == 1:
plt.scatter(X_test[i][0],X_test[i][1],c='r',marker='o')
else:
plt.scatter(X_test[i][0],X_test[i][1],c='b',marker='x')
X = np.random.uniform(0.2,0.8,100)
Y = -(w[0]*X+c)/w[1]
plt.xlabel('potential')
plt.ylabel('age')
plt.plot(X,Y,'g')
plt.show()
print("测试集准确率:",accuracy)
for i in range(len(data)):
if data[i][-1] == 1:
plt.scatter(data_feature[i][0],data_feature[i][1],c='r',marker='o')
else:
plt.scatter(data_feature[i][0],data_feature[i][1],c='b',marker='x')
X = np.random.uniform(0.2,0.8,100)
Y = -(w[0]*X+c)/w[1]
plt.xlabel('potential')
plt.ylabel('age')
plt.plot(X,Y,'g')
plt.show()
```
## 口袋算法
```
def sign(a):
if a>0:
return 1
else:
return -1
def checkErrorRate(X_train,Y_train,w,c):
count=0
for i in range(len(X_train)):
x=np.array(X_train[i])
y=Y_train[i]
if sign(np.sum(w*x)+c)!=sign(y):
count+=1
return count/len(X_train)
def Pocket():
w = np.zeros(2)
lr = 0.01
c = 1
cnt = 0
best_w = w
best_c = c
bestRate = 1
while True:
cnt += 1
if cnt > 1000:
break
for i in range(len(X_train)):
x = np.array(X_train[i])
y = Y_train[i]
s = np.sum(w*x)+c
z = sign(s)
if y * z <= 0:
w = w+lr*y*x
c = c+lr*y
rate = checkErrorRate(X_train,Y_train,w,c)
# print(rate)
if rate < bestRate:
bestRate = rate
print("bestRate update",rate)
best_w = w
best_c = c
return cnt,best_w,best_c
if __name__=='__main__':
cnt,best_w,best_c=Pocket()
print("迭代次数:",cnt)
print("超平面法向量:",w)
print("阈值:",c)
count = 0
for i in range(len(X_test)):
target = Y_test[i]
result = sign(np.sum(best_w*X_test[i])+best_c)
if(target == result):
count += 1
accuracy = float(count/len(X_test))
for i in range(len(X_test)):
if Y_test[i] == 1:
plt.scatter(X_test[i][0],X_test[i][1],c='r',marker='o')
else:
plt.scatter(X_test[i][0],X_test[i][1],c='b',marker='x')
X = np.random.uniform(0.2,0.8,100)
Y = -(best_w[0]*X+best_c)/best_w[1]
plt.xlabel('potential')
plt.ylabel('age')
plt.plot(X,Y,'g')
plt.show()
print("测试集准确率:",accuracy)
for i in range(len(data)):
if data[i][-1] == 1:
plt.scatter(data_feature[i][0],data_feature[i][1],c='r',marker='o')
else:
plt.scatter(data_feature[i][0],data_feature[i][1],c='b',marker='x')
X = np.random.uniform(0.2,0.8,100)
Y = -(best_w[0]*X+best_c)/best_w[1]
plt.xlabel('potential')
plt.ylabel('age')
plt.plot(X,Y,'g')
plt.show()
```
| github_jupyter |
# Convolutional Neural Networks: Step by Step
Welcome to Course 4's first assignment! In this assignment, you will implement convolutional (CONV) and pooling (POOL) layers in numpy, including both forward propagation and (optionally) backward propagation.
**Notation**:
- Superscript $[l]$ denotes an object of the $l^{th}$ layer.
- Example: $a^{[4]}$ is the $4^{th}$ layer activation. $W^{[5]}$ and $b^{[5]}$ are the $5^{th}$ layer parameters.
- Superscript $(i)$ denotes an object from the $i^{th}$ example.
- Example: $x^{(i)}$ is the $i^{th}$ training example input.
- Lowerscript $i$ denotes the $i^{th}$ entry of a vector.
- Example: $a^{[l]}_i$ denotes the $i^{th}$ entry of the activations in layer $l$, assuming this is a fully connected (FC) layer.
- $n_H$, $n_W$ and $n_C$ denote respectively the height, width and number of channels of a given layer. If you want to reference a specific layer $l$, you can also write $n_H^{[l]}$, $n_W^{[l]}$, $n_C^{[l]}$.
- $n_{H_{prev}}$, $n_{W_{prev}}$ and $n_{C_{prev}}$ denote respectively the height, width and number of channels of the previous layer. If referencing a specific layer $l$, this could also be denoted $n_H^{[l-1]}$, $n_W^{[l-1]}$, $n_C^{[l-1]}$.
We assume that you are already familiar with `numpy` and/or have completed the previous courses of the specialization. Let's get started!
## 1 - Packages
Let's first import all the packages that you will need during this assignment.
- [numpy](www.numpy.org) is the fundamental package for scientific computing with Python.
- [matplotlib](http://matplotlib.org) is a library to plot graphs in Python.
- np.random.seed(1) is used to keep all the random function calls consistent. It will help us grade your work.
```
import numpy as np
import h5py
import matplotlib.pyplot as plt
%matplotlib inline
plt.rcParams['figure.figsize'] = (5.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
%load_ext autoreload
%autoreload 2
np.random.seed(1)
```
## 2 - Outline of the Assignment
You will be implementing the building blocks of a convolutional neural network! Each function you will implement will have detailed instructions that will walk you through the steps needed:
- Convolution functions, including:
- Zero Padding
- Convolve window
- Convolution forward
- Convolution backward (optional)
- Pooling functions, including:
- Pooling forward
- Create mask
- Distribute value
- Pooling backward (optional)
This notebook will ask you to implement these functions from scratch in `numpy`. In the next notebook, you will use the TensorFlow equivalents of these functions to build the following model:
<img src="images/model.png" style="width:800px;height:300px;">
**Note** that for every forward function, there is its corresponding backward equivalent. Hence, at every step of your forward module you will store some parameters in a cache. These parameters are used to compute gradients during backpropagation.
## 3 - Convolutional Neural Networks
Although programming frameworks make convolutions easy to use, they remain one of the hardest concepts to understand in Deep Learning. A convolution layer transforms an input volume into an output volume of different size, as shown below.
<img src="images/conv_nn.png" style="width:350px;height:200px;">
In this part, you will build every step of the convolution layer. You will first implement two helper functions: one for zero padding and the other for computing the convolution function itself.
### 3.1 - Zero-Padding
Zero-padding adds zeros around the border of an image:
<img src="images/PAD.png" style="width:600px;height:400px;">
<caption><center> <u> <font color='purple'> **Figure 1** </u><font color='purple'> : **Zero-Padding**<br> Image (3 channels, RGB) with a padding of 2. </center></caption>
The main benefits of padding are the following:
- It allows you to use a CONV layer without necessarily shrinking the height and width of the volumes. This is important for building deeper networks, since otherwise the height/width would shrink as you go to deeper layers. An important special case is the "same" convolution, in which the height/width is exactly preserved after one layer.
- It helps us keep more of the information at the border of an image. Without padding, very few values at the next layer would be affected by pixels as the edges of an image.
**Exercise**: Implement the following function, which pads all the images of a batch of examples X with zeros. [Use np.pad](https://docs.scipy.org/doc/numpy/reference/generated/numpy.pad.html). Note if you want to pad the array "a" of shape $(5,5,5,5,5)$ with `pad = 1` for the 2nd dimension, `pad = 3` for the 4th dimension and `pad = 0` for the rest, you would do:
```python
a = np.pad(a, ((0,0), (1,1), (0,0), (3,3), (0,0)), 'constant', constant_values = (..,..))
```
```
# GRADED FUNCTION: zero_pad
def zero_pad(X, pad):
"""
Pad with zeros all images of the dataset X. The padding is applied to the height and width of an image,
as illustrated in Figure 1.
Argument:
X -- python numpy array of shape (m, n_H, n_W, n_C) representing a batch of m images
pad -- integer, amount of padding around each image on vertical and horizontal dimensions
Returns:
X_pad -- padded image of shape (m, n_H + 2*pad, n_W + 2*pad, n_C)
"""
### START CODE HERE ### (≈ 1 line)
X_pad = np.pad(X,((0,0), (pad,pad), (pad,pad), (0,0)), 'constant')
### END CODE HERE ###
return X_pad
np.random.seed(1)
x = np.random.randn(4, 3, 3, 2)
x_pad = zero_pad(x, 2)
print ("x.shape =", x.shape)
print ("x_pad.shape =", x_pad.shape)
print ("x[1,1] =", x[1,1])
print ("x_pad[1,1] =", x_pad[1,1])
fig, axarr = plt.subplots(1, 2)
axarr[0].set_title('x')
axarr[0].imshow(x[0,:,:,0])
axarr[1].set_title('x_pad')
axarr[1].imshow(x_pad[0,:,:,0])
```
**Expected Output**:
<table>
<tr>
<td>
**x.shape**:
</td>
<td>
(4, 3, 3, 2)
</td>
</tr>
<tr>
<td>
**x_pad.shape**:
</td>
<td>
(4, 7, 7, 2)
</td>
</tr>
<tr>
<td>
**x[1,1]**:
</td>
<td>
[[ 0.90085595 -0.68372786]
[-0.12289023 -0.93576943]
[-0.26788808 0.53035547]]
</td>
</tr>
<tr>
<td>
**x_pad[1,1]**:
</td>
<td>
[[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]]
</td>
</tr>
</table>
### 3.2 - Single step of convolution
In this part, implement a single step of convolution, in which you apply the filter to a single position of the input. This will be used to build a convolutional unit, which:
- Takes an input volume
- Applies a filter at every position of the input
- Outputs another volume (usually of different size)
<img src="images/Convolution_schematic.gif" style="width:500px;height:300px;">
<caption><center> <u> <font color='purple'> **Figure 2** </u><font color='purple'> : **Convolution operation**<br> with a filter of 2x2 and a stride of 1 (stride = amount you move the window each time you slide) </center></caption>
In a computer vision application, each value in the matrix on the left corresponds to a single pixel value, and we convolve a 3x3 filter with the image by multiplying its values element-wise with the original matrix, then summing them up and adding a bias. In this first step of the exercise, you will implement a single step of convolution, corresponding to applying a filter to just one of the positions to get a single real-valued output.
Later in this notebook, you'll apply this function to multiple positions of the input to implement the full convolutional operation.
**Exercise**: Implement conv_single_step(). [Hint](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.sum.html).
```
# GRADED FUNCTION: conv_single_step
def conv_single_step(a_slice_prev, W, b):
"""
Apply one filter defined by parameters W on a single slice (a_slice_prev) of the output activation
of the previous layer.
Arguments:
a_slice_prev -- slice of input data of shape (f, f, n_C_prev)
W -- Weight parameters contained in a window - matrix of shape (f, f, n_C_prev)
b -- Bias parameters contained in a window - matrix of shape (1, 1, 1)
Returns:
Z -- a scalar value, result of convolving the sliding window (W, b) on a slice x of the input data
"""
### START CODE HERE ### (≈ 2 lines of code)
# Element-wise product between a_slice and W. Do not add the bias yet.
s = a_slice_prev*W
# Sum over all entries of the volume s.
Z = s.sum()
# Add bias b to Z. Cast b to a float() so that Z results in a scalar value.
Z = Z+float(b)
### END CODE HERE ###
return Z
np.random.seed(1)
a_slice_prev = np.random.randn(4, 4, 3)
W = np.random.randn(4, 4, 3)
b = np.random.randn(1, 1, 1)
Z = conv_single_step(a_slice_prev, W, b)
print("Z =", Z)
```
**Expected Output**:
<table>
<tr>
<td>
**Z**
</td>
<td>
-6.99908945068
</td>
</tr>
</table>
### 3.3 - Convolutional Neural Networks - Forward pass
In the forward pass, you will take many filters and convolve them on the input. Each 'convolution' gives you a 2D matrix output. You will then stack these outputs to get a 3D volume:
<center>
<video width="620" height="440" src="images/conv_kiank.mp4" type="video/mp4" controls>
</video>
</center>
**Exercise**: Implement the function below to convolve the filters W on an input activation A_prev. This function takes as input A_prev, the activations output by the previous layer (for a batch of m inputs), F filters/weights denoted by W, and a bias vector denoted by b, where each filter has its own (single) bias. Finally you also have access to the hyperparameters dictionary which contains the stride and the padding.
**Hint**:
1. To select a 2x2 slice at the upper left corner of a matrix "a_prev" (shape (5,5,3)), you would do:
```python
a_slice_prev = a_prev[0:2,0:2,:]
```
This will be useful when you will define `a_slice_prev` below, using the `start/end` indexes you will define.
2. To define a_slice you will need to first define its corners `vert_start`, `vert_end`, `horiz_start` and `horiz_end`. This figure may be helpful for you to find how each of the corner can be defined using h, w, f and s in the code below.
<img src="images/vert_horiz_kiank.png" style="width:400px;height:300px;">
<caption><center> <u> <font color='purple'> **Figure 3** </u><font color='purple'> : **Definition of a slice using vertical and horizontal start/end (with a 2x2 filter)** <br> This figure shows only a single channel. </center></caption>
**Reminder**:
The formulas relating the output shape of the convolution to the input shape is:
$$ n_H = \lfloor \frac{n_{H_{prev}} - f + 2 \times pad}{stride} \rfloor +1 $$
$$ n_W = \lfloor \frac{n_{W_{prev}} - f + 2 \times pad}{stride} \rfloor +1 $$
$$ n_C = \text{number of filters used in the convolution}$$
For this exercise, we won't worry about vectorization, and will just implement everything with for-loops.
```
# GRADED FUNCTION: conv_forward
def conv_forward(A_prev, W, b, hparameters):
"""
Implements the forward propagation for a convolution function
Arguments:
A_prev -- output activations of the previous layer, numpy array of shape (m, n_H_prev, n_W_prev, n_C_prev)
W -- Weights, numpy array of shape (f, f, n_C_prev, n_C)
b -- Biases, numpy array of shape (1, 1, 1, n_C)
hparameters -- python dictionary containing "stride" and "pad"
Returns:
Z -- conv output, numpy array of shape (m, n_H, n_W, n_C)
cache -- cache of values needed for the conv_backward() function
"""
### START CODE HERE ###
# Retrieve dimensions from A_prev's shape (≈1 line)
(m, n_H_prev, n_W_prev, n_C_prev) = (A_prev.shape[0],A_prev.shape[1],A_prev.shape[2],A_prev.shape[3])
# Retrieve dimensions from W's shape (≈1 line)
(f, f, n_C_prev, n_C) = (W.shape[0],W.shape[1],W.shape[2],W.shape[3])
# Retrieve information from "hparameters" (≈2 lines)
stride = hparameters["stride"]
pad = hparameters["pad"]
# Compute the dimensions of the CONV output volume using the formula given above. Hint: use int() to floor. (≈2 lines)
n_H = int((n_H_prev-f+2*pad)/stride)+1
n_W = int((n_W_prev-f+2*pad)/stride)+1
# Initialize the output volume Z with zeros. (≈1 line)
Z = np.zeros((m, n_H, n_W, n_C))
# Create A_prev_pad by padding A_prev
A_prev_pad = zero_pad(A_prev, pad)
for i in range(m): # loop over the batch of training examples
a_prev_pad = A_prev_pad[i,:,:,:] # Select ith training example's padded activation
for h in range(n_H): # loop over vertical axis of the output volume
for w in range(n_W): # loop over horizontal axis of the output volume
for c in range(n_C): # loop over channels (= #filters) of the output volume
# Find the corners of the current "slice" (≈4 lines)
vert_start = h*stride
vert_end = h*stride+f
horiz_start = w*stride
horiz_end = w*stride+f
# Use the corners to define the (3D) slice of a_prev_pad (See Hint above the cell). (≈1 line)
a_slice_prev = a_prev_pad[vert_start:vert_end,horiz_start:horiz_end,:]
# Convolve the (3D) slice with the correct filter W and bias b, to get back one output neuron. (≈1 line)
Z[i, h, w, c] = conv_single_step(a_slice_prev, W[:,:,:,c], b[:,:,:,c])
### END CODE HERE ###
# Making sure your output shape is correct
assert(Z.shape == (m, n_H, n_W, n_C))
# Save information in "cache" for the backprop
cache = (A_prev, W, b, hparameters)
return Z, cache
np.random.seed(1)
A_prev = np.random.randn(10,4,4,3)
W = np.random.randn(2,2,3,8)
b = np.random.randn(1,1,1,8)
hparameters = {"pad" : 2,
"stride": 2}
Z, cache_conv = conv_forward(A_prev, W, b, hparameters)
print("Z's mean =", np.mean(Z))
print("Z[3,2,1] =", Z[3,2,1])
print("cache_conv[0][1][2][3] =", cache_conv[0][1][2][3])
```
**Expected Output**:
<table>
<tr>
<td>
**Z's mean**
</td>
<td>
0.0489952035289
</td>
</tr>
<tr>
<td>
**Z[3,2,1]**
</td>
<td>
[-0.61490741 -6.7439236 -2.55153897 1.75698377 3.56208902 0.53036437
5.18531798 8.75898442]
</td>
</tr>
<tr>
<td>
**cache_conv[0][1][2][3]**
</td>
<td>
[-0.20075807 0.18656139 0.41005165]
</td>
</tr>
</table>
Finally, CONV layer should also contain an activation, in which case we would add the following line of code:
```python
# Convolve the window to get back one output neuron
Z[i, h, w, c] = ...
# Apply activation
A[i, h, w, c] = activation(Z[i, h, w, c])
```
You don't need to do it here.
## 4 - Pooling layer
The pooling (POOL) layer reduces the height and width of the input. It helps reduce computation, as well as helps make feature detectors more invariant to its position in the input. The two types of pooling layers are:
- Max-pooling layer: slides an ($f, f$) window over the input and stores the max value of the window in the output.
- Average-pooling layer: slides an ($f, f$) window over the input and stores the average value of the window in the output.
<table>
<td>
<img src="images/max_pool1.png" style="width:500px;height:300px;">
<td>
<td>
<img src="images/a_pool.png" style="width:500px;height:300px;">
<td>
</table>
These pooling layers have no parameters for backpropagation to train. However, they have hyperparameters such as the window size $f$. This specifies the height and width of the fxf window you would compute a max or average over.
### 4.1 - Forward Pooling
Now, you are going to implement MAX-POOL and AVG-POOL, in the same function.
**Exercise**: Implement the forward pass of the pooling layer. Follow the hints in the comments below.
**Reminder**:
As there's no padding, the formulas binding the output shape of the pooling to the input shape is:
$$ n_H = \lfloor \frac{n_{H_{prev}} - f}{stride} \rfloor +1 $$
$$ n_W = \lfloor \frac{n_{W_{prev}} - f}{stride} \rfloor +1 $$
$$ n_C = n_{C_{prev}}$$
```
# GRADED FUNCTION: pool_forward
def pool_forward(A_prev, hparameters, mode = "max"):
"""
Implements the forward pass of the pooling layer
Arguments:
A_prev -- Input data, numpy array of shape (m, n_H_prev, n_W_prev, n_C_prev)
hparameters -- python dictionary containing "f" and "stride"
mode -- the pooling mode you would like to use, defined as a string ("max" or "average")
Returns:
A -- output of the pool layer, a numpy array of shape (m, n_H, n_W, n_C)
cache -- cache used in the backward pass of the pooling layer, contains the input and hparameters
"""
# Retrieve dimensions from the input shape
(m, n_H_prev, n_W_prev, n_C_prev) = A_prev.shape
# Retrieve hyperparameters from "hparameters"
f = hparameters["f"]
stride = hparameters["stride"]
# Define the dimensions of the output
n_H = int(1 + (n_H_prev - f) / stride)
n_W = int(1 + (n_W_prev - f) / stride)
n_C = n_C_prev
# Initialize output matrix A
A = np.zeros((m, n_H, n_W, n_C))
### START CODE HERE ###
for i in range(m): # loop over the training examples
for h in range(n_H): # loop on the vertical axis of the output volume
for w in range(n_W): # loop on the horizontal axis of the output volume
for c in range (n_C): # loop over the channels of the output volume
# Find the corners of the current "slice" (≈4 lines)
vert_start = h*stride
vert_end = h*stride+f
horiz_start = w*stride
horiz_end = W*stride+f
# Use the corners to define the current slice on the ith training example of A_prev, channel c. (≈1 line)
a_prev_slice = (A_prev[i])[vert_start:vert_end,horiz_start:horiz_end,:]
# Compute the pooling operation on the slice. Use an if statment to differentiate the modes. Use np.max/np.mean.
if mode == "max":
A[i, h, w, c] = np.max(a_prev_slice)
elif mode == "average":
A[i, h, w, c] = np.mean(a_prev_slice)
### END CODE HERE ###
# Store the input and hparameters in "cache" for pool_backward()
cache = (A_prev, hparameters)
# Making sure your output shape is correct
assert(A.shape == (m, n_H, n_W, n_C))
return A, cache
np.random.seed(1)
A_prev = np.random.randn(2, 4, 4, 3)
hparameters = {"stride" : 2, "f": 3}
A, cache = pool_forward(A_prev, hparameters)
print("mode = max")
print("A =", A)
print()
A, cache = pool_forward(A_prev, hparameters, mode = "average")
print("mode = average")
print("A =", A)
```
**Expected Output:**
<table>
<tr>
<td>
A =
</td>
<td>
[[[[ 1.74481176 0.86540763 1.13376944]]]
[[[ 1.13162939 1.51981682 2.18557541]]]]
</td>
</tr>
<tr>
<td>
A =
</td>
<td>
[[[[ 0.02105773 -0.20328806 -0.40389855]]]
[[[-0.22154621 0.51716526 0.48155844]]]]
</td>
</tr>
</table>
Congratulations! You have now implemented the forward passes of all the layers of a convolutional network.
The remainer of this notebook is optional, and will not be graded.
## 5 - Backpropagation in convolutional neural networks (OPTIONAL / UNGRADED)
In modern deep learning frameworks, you only have to implement the forward pass, and the framework takes care of the backward pass, so most deep learning engineers don't need to bother with the details of the backward pass. The backward pass for convolutional networks is complicated. If you wish however, you can work through this optional portion of the notebook to get a sense of what backprop in a convolutional network looks like.
When in an earlier course you implemented a simple (fully connected) neural network, you used backpropagation to compute the derivatives with respect to the cost to update the parameters. Similarly, in convolutional neural networks you can to calculate the derivatives with respect to the cost in order to update the parameters. The backprop equations are not trivial and we did not derive them in lecture, but we briefly presented them below.
### 5.1 - Convolutional layer backward pass
Let's start by implementing the backward pass for a CONV layer.
#### 5.1.1 - Computing dA:
This is the formula for computing $dA$ with respect to the cost for a certain filter $W_c$ and a given training example:
$$ dA += \sum _{h=0} ^{n_H} \sum_{w=0} ^{n_W} W_c \times dZ_{hw} \tag{1}$$
Where $W_c$ is a filter and $dZ_{hw}$ is a scalar corresponding to the gradient of the cost with respect to the output of the conv layer Z at the hth row and wth column (corresponding to the dot product taken at the ith stride left and jth stride down). Note that at each time, we multiply the the same filter $W_c$ by a different dZ when updating dA. We do so mainly because when computing the forward propagation, each filter is dotted and summed by a different a_slice. Therefore when computing the backprop for dA, we are just adding the gradients of all the a_slices.
In code, inside the appropriate for-loops, this formula translates into:
```python
da_prev_pad[vert_start:vert_end, horiz_start:horiz_end, :] += W[:,:,:,c] * dZ[i, h, w, c]
```
#### 5.1.2 - Computing dW:
This is the formula for computing $dW_c$ ($dW_c$ is the derivative of one filter) with respect to the loss:
$$ dW_c += \sum _{h=0} ^{n_H} \sum_{w=0} ^ {n_W} a_{slice} \times dZ_{hw} \tag{2}$$
Where $a_{slice}$ corresponds to the slice which was used to generate the acitivation $Z_{ij}$. Hence, this ends up giving us the gradient for $W$ with respect to that slice. Since it is the same $W$, we will just add up all such gradients to get $dW$.
In code, inside the appropriate for-loops, this formula translates into:
```python
dW[:,:,:,c] += a_slice * dZ[i, h, w, c]
```
#### 5.1.3 - Computing db:
This is the formula for computing $db$ with respect to the cost for a certain filter $W_c$:
$$ db = \sum_h \sum_w dZ_{hw} \tag{3}$$
As you have previously seen in basic neural networks, db is computed by summing $dZ$. In this case, you are just summing over all the gradients of the conv output (Z) with respect to the cost.
In code, inside the appropriate for-loops, this formula translates into:
```python
db[:,:,:,c] += dZ[i, h, w, c]
```
**Exercise**: Implement the `conv_backward` function below. You should sum over all the training examples, filters, heights, and widths. You should then compute the derivatives using formulas 1, 2 and 3 above.
```
def conv_backward(dZ, cache):
"""
Implement the backward propagation for a convolution function
Arguments:
dZ -- gradient of the cost with respect to the output of the conv layer (Z), numpy array of shape (m, n_H, n_W, n_C)
cache -- cache of values needed for the conv_backward(), output of conv_forward()
Returns:
dA_prev -- gradient of the cost with respect to the input of the conv layer (A_prev),
numpy array of shape (m, n_H_prev, n_W_prev, n_C_prev)
dW -- gradient of the cost with respect to the weights of the conv layer (W)
numpy array of shape (f, f, n_C_prev, n_C)
db -- gradient of the cost with respect to the biases of the conv layer (b)
numpy array of shape (1, 1, 1, n_C)
"""
### START CODE HERE ###
# Retrieve information from "cache"
(A_prev, W, b, hparameters) = None
# Retrieve dimensions from A_prev's shape
(m, n_H_prev, n_W_prev, n_C_prev) = None
# Retrieve dimensions from W's shape
(f, f, n_C_prev, n_C) = None
# Retrieve information from "hparameters"
stride = None
pad = None
# Retrieve dimensions from dZ's shape
(m, n_H, n_W, n_C) = None
# Initialize dA_prev, dW, db with the correct shapes
dA_prev = None
dW = None
db = None
# Pad A_prev and dA_prev
A_prev_pad = None
dA_prev_pad = None
for i in range(None): # loop over the training examples
# select ith training example from A_prev_pad and dA_prev_pad
a_prev_pad = None
da_prev_pad = None
for h in range(None): # loop over vertical axis of the output volume
for w in range(None): # loop over horizontal axis of the output volume
for c in range(None): # loop over the channels of the output volume
# Find the corners of the current "slice"
vert_start = None
vert_end = None
horiz_start = None
horiz_end = None
# Use the corners to define the slice from a_prev_pad
a_slice = None
# Update gradients for the window and the filter's parameters using the code formulas given above
da_prev_pad[vert_start:vert_end, horiz_start:horiz_end, :] += None
dW[:,:,:,c] += None
db[:,:,:,c] += None
# Set the ith training example's dA_prev to the unpaded da_prev_pad (Hint: use X[pad:-pad, pad:-pad, :])
dA_prev[i, :, :, :] = None
### END CODE HERE ###
# Making sure your output shape is correct
assert(dA_prev.shape == (m, n_H_prev, n_W_prev, n_C_prev))
return dA_prev, dW, db
np.random.seed(1)
dA, dW, db = conv_backward(Z, cache_conv)
print("dA_mean =", np.mean(dA))
print("dW_mean =", np.mean(dW))
print("db_mean =", np.mean(db))
```
** Expected Output: **
<table>
<tr>
<td>
**dA_mean**
</td>
<td>
1.45243777754
</td>
</tr>
<tr>
<td>
**dW_mean**
</td>
<td>
1.72699145831
</td>
</tr>
<tr>
<td>
**db_mean**
</td>
<td>
7.83923256462
</td>
</tr>
</table>
## 5.2 Pooling layer - backward pass
Next, let's implement the backward pass for the pooling layer, starting with the MAX-POOL layer. Even though a pooling layer has no parameters for backprop to update, you still need to backpropagation the gradient through the pooling layer in order to compute gradients for layers that came before the pooling layer.
### 5.2.1 Max pooling - backward pass
Before jumping into the backpropagation of the pooling layer, you are going to build a helper function called `create_mask_from_window()` which does the following:
$$ X = \begin{bmatrix}
1 && 3 \\
4 && 2
\end{bmatrix} \quad \rightarrow \quad M =\begin{bmatrix}
0 && 0 \\
1 && 0
\end{bmatrix}\tag{4}$$
As you can see, this function creates a "mask" matrix which keeps track of where the maximum of the matrix is. True (1) indicates the position of the maximum in X, the other entries are False (0). You'll see later that the backward pass for average pooling will be similar to this but using a different mask.
**Exercise**: Implement `create_mask_from_window()`. This function will be helpful for pooling backward.
Hints:
- [np.max()]() may be helpful. It computes the maximum of an array.
- If you have a matrix X and a scalar x: `A = (X == x)` will return a matrix A of the same size as X such that:
```
A[i,j] = True if X[i,j] = x
A[i,j] = False if X[i,j] != x
```
- Here, you don't need to consider cases where there are several maxima in a matrix.
```
def create_mask_from_window(x):
"""
Creates a mask from an input matrix x, to identify the max entry of x.
Arguments:
x -- Array of shape (f, f)
Returns:
mask -- Array of the same shape as window, contains a True at the position corresponding to the max entry of x.
"""
### START CODE HERE ### (≈1 line)
mask = None
### END CODE HERE ###
return mask
np.random.seed(1)
x = np.random.randn(2,3)
mask = create_mask_from_window(x)
print('x = ', x)
print("mask = ", mask)
```
**Expected Output:**
<table>
<tr>
<td>
**x =**
</td>
<td>
[[ 1.62434536 -0.61175641 -0.52817175] <br>
[-1.07296862 0.86540763 -2.3015387 ]]
</td>
</tr>
<tr>
<td>
**mask =**
</td>
<td>
[[ True False False] <br>
[False False False]]
</td>
</tr>
</table>
Why do we keep track of the position of the max? It's because this is the input value that ultimately influenced the output, and therefore the cost. Backprop is computing gradients with respect to the cost, so anything that influences the ultimate cost should have a non-zero gradient. So, backprop will "propagate" the gradient back to this particular input value that had influenced the cost.
### 5.2.2 - Average pooling - backward pass
In max pooling, for each input window, all the "influence" on the output came from a single input value--the max. In average pooling, every element of the input window has equal influence on the output. So to implement backprop, you will now implement a helper function that reflects this.
For example if we did average pooling in the forward pass using a 2x2 filter, then the mask you'll use for the backward pass will look like:
$$ dZ = 1 \quad \rightarrow \quad dZ =\begin{bmatrix}
1/4 && 1/4 \\
1/4 && 1/4
\end{bmatrix}\tag{5}$$
This implies that each position in the $dZ$ matrix contributes equally to output because in the forward pass, we took an average.
**Exercise**: Implement the function below to equally distribute a value dz through a matrix of dimension shape. [Hint](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ones.html)
```
def distribute_value(dz, shape):
"""
Distributes the input value in the matrix of dimension shape
Arguments:
dz -- input scalar
shape -- the shape (n_H, n_W) of the output matrix for which we want to distribute the value of dz
Returns:
a -- Array of size (n_H, n_W) for which we distributed the value of dz
"""
### START CODE HERE ###
# Retrieve dimensions from shape (≈1 line)
(n_H, n_W) = None
# Compute the value to distribute on the matrix (≈1 line)
average = None
# Create a matrix where every entry is the "average" value (≈1 line)
a = None
### END CODE HERE ###
return a
a = distribute_value(2, (2,2))
print('distributed value =', a)
```
**Expected Output**:
<table>
<tr>
<td>
distributed_value =
</td>
<td>
[[ 0.5 0.5]
<br\>
[ 0.5 0.5]]
</td>
</tr>
</table>
### 5.2.3 Putting it together: Pooling backward
You now have everything you need to compute backward propagation on a pooling layer.
**Exercise**: Implement the `pool_backward` function in both modes (`"max"` and `"average"`). You will once again use 4 for-loops (iterating over training examples, height, width, and channels). You should use an `if/elif` statement to see if the mode is equal to `'max'` or `'average'`. If it is equal to 'average' you should use the `distribute_value()` function you implemented above to create a matrix of the same shape as `a_slice`. Otherwise, the mode is equal to '`max`', and you will create a mask with `create_mask_from_window()` and multiply it by the corresponding value of dZ.
```
def pool_backward(dA, cache, mode = "max"):
"""
Implements the backward pass of the pooling layer
Arguments:
dA -- gradient of cost with respect to the output of the pooling layer, same shape as A
cache -- cache output from the forward pass of the pooling layer, contains the layer's input and hparameters
mode -- the pooling mode you would like to use, defined as a string ("max" or "average")
Returns:
dA_prev -- gradient of cost with respect to the input of the pooling layer, same shape as A_prev
"""
### START CODE HERE ###
# Retrieve information from cache (≈1 line)
(A_prev, hparameters) = None
# Retrieve hyperparameters from "hparameters" (≈2 lines)
stride = None
f = None
# Retrieve dimensions from A_prev's shape and dA's shape (≈2 lines)
m, n_H_prev, n_W_prev, n_C_prev = None
m, n_H, n_W, n_C = None
# Initialize dA_prev with zeros (≈1 line)
dA_prev = None
for i in range(None): # loop over the training examples
# select training example from A_prev (≈1 line)
a_prev = None
for h in range(None): # loop on the vertical axis
for w in range(None): # loop on the horizontal axis
for c in range(None): # loop over the channels (depth)
# Find the corners of the current "slice" (≈4 lines)
vert_start = None
vert_end = None
horiz_start = None
horiz_end = None
# Compute the backward propagation in both modes.
if mode == "max":
# Use the corners and "c" to define the current slice from a_prev (≈1 line)
a_prev_slice = None
# Create the mask from a_prev_slice (≈1 line)
mask = None
# Set dA_prev to be dA_prev + (the mask multiplied by the correct entry of dA) (≈1 line)
dA_prev[i, vert_start: vert_end, horiz_start: horiz_end, c] += None
elif mode == "average":
# Get the value a from dA (≈1 line)
da = None
# Define the shape of the filter as fxf (≈1 line)
shape = None
# Distribute it to get the correct slice of dA_prev. i.e. Add the distributed value of da. (≈1 line)
dA_prev[i, vert_start: vert_end, horiz_start: horiz_end, c] += None
### END CODE ###
# Making sure your output shape is correct
assert(dA_prev.shape == A_prev.shape)
return dA_prev
np.random.seed(1)
A_prev = np.random.randn(5, 5, 3, 2)
hparameters = {"stride" : 1, "f": 2}
A, cache = pool_forward(A_prev, hparameters)
dA = np.random.randn(5, 4, 2, 2)
dA_prev = pool_backward(dA, cache, mode = "max")
print("mode = max")
print('mean of dA = ', np.mean(dA))
print('dA_prev[1,1] = ', dA_prev[1,1])
print()
dA_prev = pool_backward(dA, cache, mode = "average")
print("mode = average")
print('mean of dA = ', np.mean(dA))
print('dA_prev[1,1] = ', dA_prev[1,1])
```
**Expected Output**:
mode = max:
<table>
<tr>
<td>
**mean of dA =**
</td>
<td>
0.145713902729
</td>
</tr>
<tr>
<td>
**dA_prev[1,1] =**
</td>
<td>
[[ 0. 0. ] <br>
[ 5.05844394 -1.68282702] <br>
[ 0. 0. ]]
</td>
</tr>
</table>
mode = average
<table>
<tr>
<td>
**mean of dA =**
</td>
<td>
0.145713902729
</td>
</tr>
<tr>
<td>
**dA_prev[1,1] =**
</td>
<td>
[[ 0.08485462 0.2787552 ] <br>
[ 1.26461098 -0.25749373] <br>
[ 1.17975636 -0.53624893]]
</td>
</tr>
</table>
### Congratulations !
Congratulation on completing this assignment. You now understand how convolutional neural networks work. You have implemented all the building blocks of a neural network. In the next assignment you will implement a ConvNet using TensorFlow.
| github_jupyter |
# Tuning an Image Classification model (MNIST)
In this tutorial we will see how we can use PyTorchWrapper in order to tune an Image Classification model on the MNIST dataset.
#### Additional libraries
First of all we need to install the `torchvision` library in order to download the data.
```
! pip install torchvision
```
#### Import Statements
```
import torch
import torchvision
import math
import random
import hyperopt
import os
from pprint import pprint
from hyperopt import hp
from torch import nn
from torchvision.datasets import MNIST
from torch.utils.data.dataset import Dataset
from torch.utils.data.dataloader import DataLoader
from torch.utils.data.sampler import SubsetRandomSampler
from pytorch_wrapper import modules, System
from pytorch_wrapper import evaluators as evaluators
from pytorch_wrapper.loss_wrappers import GenericPointWiseLossWrapper
from pytorch_wrapper.training_callbacks import EarlyStoppingCriterionCallback
from pytorch_wrapper.tuner import Tuner
```
#### Dataset Definition
Since torchvision provides ready to use `torch.utils.data.Dataset` object for the MNIST Dataset we just need to wrap it with a custom class in order to adhere to the requirements of PyTorchWrapper, i.e. the data loaders must represent a batch as a dictionary.
```
class MNISTDatasetWrapper(Dataset):
def __init__(self, is_train):
self.dataset = MNIST(
'data/mnist/',
train=is_train,
download=True,
transform=torchvision.transforms.ToTensor()
)
def __getitem__(self, index):
return {'input': self.dataset[index][0], 'target': self.dataset[index][1]}
def __len__(self):
return len(self.dataset)
```
#### Model Definition
The model will be CNN based, but the exact architecture will be chosen by the tuner.
```
class Model(nn.Module):
def __init__(self, channels, kernel_size, depth, dp, mlp_depth, mlp_hl):
super(Model, self).__init__()
cnn_list = [
nn.Conv2d(
in_channels=1,
out_channels=channels,
kernel_size=kernel_size,
padding=math.floor(kernel_size / 2)
),
nn.Dropout(p=dp),
nn.MaxPool2d(kernel_size=2),
nn.ReLU()
]
for _ in range(depth - 1):
cnn_list.extend([
nn.Conv2d(
in_channels=channels,
out_channels=channels,
kernel_size=kernel_size,
padding=math.floor(kernel_size / 2)
),
nn.Dropout(p=dp),
nn.MaxPool2d(kernel_size=2),
nn.ReLU()
])
self.cnn = nn.Sequential(*cnn_list)
self.out_mlp = modules.MLP(
input_size=int(pow(int(28 // (math.pow(2, depth))), 2)) * channels,
num_hidden_layers=mlp_depth,
hidden_layer_size=mlp_hl,
hidden_activation=nn.ReLU,
hidden_dp=dp,
output_size=10,
output_activation=None
)
def forward(self, x):
x = self.cnn(x)
x = x.view(x.shape[0], -1)
return self.out_mlp(x)
```
#### Training
First of all we create the dataset objects alongside four data loaders. The train_dataloader will be used for training, the val_dataloader for early stopping, the dev_dataloader for hyperparameter optimization, and the test_dataloader
for the final evaluation.
```
train_val_dev_dataset = MNISTDatasetWrapper(True)
test_dataset = MNISTDatasetWrapper(False)
eval_size = math.floor(0.1 * len(train_val_dev_dataset))
train_val_dev_indexes = list(range(len(train_val_dev_dataset)))
random.seed(12345)
random.shuffle(train_val_dev_indexes)
train_indexes = train_val_dev_indexes[eval_size * 2:]
val_indexes = train_val_dev_indexes[eval_size:eval_size * 2]
dev_indexes = train_val_dev_indexes[:eval_size]
train_dataloader = DataLoader(
train_val_dev_dataset,
sampler=SubsetRandomSampler(train_indexes),
batch_size=128
)
val_dataloader = DataLoader(
train_val_dev_dataset,
sampler=SubsetRandomSampler(val_indexes),
batch_size=128
)
dev_dataloader = DataLoader(
train_val_dev_dataset,
sampler=SubsetRandomSampler(dev_indexes),
batch_size=128
)
test_dataloader = DataLoader(test_dataset, batch_size=128, shuffle=False)
```
Next we define the step function. This function is called in the beginning of each iteration of the tuning process.
This function is responsible for creating, training and evaluating the model given the chosen hyper parameters. The goal of the tuning process is to find the hyper parameters that minimize a chosen metric. In this example we try to minimize
the **negative** f1-score.
```
def step_function(current_params):
model = Model(**current_params['model_params'])
last_activation = nn.Softmax(dim=-1)
if torch.cuda.is_available():
system = System(model, last_activation=last_activation, device=torch.device('cuda'))
else:
system = System(model, last_activation=last_activation, device=torch.device('cpu'))
loss_wrapper = GenericPointWiseLossWrapper(nn.CrossEntropyLoss())
evals = {
'prec': evaluators.MultiClassPrecisionEvaluator(average='macro'),
'rec': evaluators.MultiClassRecallEvaluator(average='macro'),
'f1': evaluators.MultiClassF1Evaluator(average='macro')
}
optimizer = torch.optim.Adam(
filter(lambda p: p.requires_grad, system.model.parameters()),
lr=current_params['training_params']['lr']
)
os.makedirs('tmp', exist_ok=True)
_ = system.train(
loss_wrapper,
optimizer,
train_data_loader=train_dataloader,
evaluators=evals,
evaluation_data_loaders={
'val': val_dataloader
},
callbacks=[
EarlyStoppingCriterionCallback(
3,
'val',
'f1',
'tmp/mnist_tuning_cur_best.weights'
)
]
)
return -system.evaluate(dev_dataloader, evals)['f1'].score
```
Finally we define the hyper_parameter_generators, create the tuner and run it. For more information about the definition of the hyper_parameter_generators check the HyperOpt documentation.
```
hyper_parameter_generators = {
'model_params': {
'channels': hp.choice('channels', [5, 10, 20, 30, 50]),
'kernel_size': hp.choice('kernel_size', [3, 5, 7]),
'depth': hp.choice('depth', [1, 2, 3, 4]),
'dp': hp.uniform('dp', 0, 0.5),
'mlp_depth': hp.choice('mlp_depth', [1, 2, 3, 4]),
'mlp_hl': hp.choice('mlp_hl', [32, 64, 128, 256])
},
'training_params': {
'lr': hp.loguniform('lr', math.log(0.0001), math.log(0.1))
}
}
tuner = Tuner(
hyper_parameter_generators,
step_function=step_function,
algorithm=hyperopt.tpe.suggest,
fit_iterations=20
)
results = tuner.run()
pprint(results[0])
```
| github_jupyter |
```
# Import the necessary libraries
import tensorflow as tf
import tensorflow.keras as keras
# This loads the EfficientNetB0 model from the Keras library
# Input Shape is the shape of the image that is input to the first layer. For example, consider an image with shape (width, height , number of channels)
# 'include_top' is set to 'False' to load the model with out the classification or dense layers. Top layers are not required as this is a segmentation problem.
# 'weights' is set to imagenet, that is, it uses the weight it learnt while training on the imagenet dataset. You can set it to None or your custom_weights.
# IMAGE_WIDTH, IMAGE_HEIGHT and CHANNELS values provided for visualization. Please change to suit your dataset.
IMAGE_WIDTH = 512
IMAGE_HEIGHT = 512
CHANNELS = 3
model = tf.keras.applications.EfficientNetB0(input_shape=(IMAGE_WIDTH, IMAGE_HEIGHT, CHANNELS),
include_top=False, weights="imagenet")
#To see the list of layers and parameters
# For EfficientNetB0, you should see
'''Total params: 4,049,571
Trainable params: 4,007,548
Non-trainable params: 42,023'''
model.summary()
# Importing the layers to create the decoder and complete the network
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Input
from tensorflow.keras.layers import Dropout
from tensorflow.keras.layers import Lambda
from tensorflow.keras.layers import Conv2D, Conv2DTranspose
from tensorflow.keras.layers import MaxPooling2D
from tensorflow.keras.layers import Activation
from tensorflow.keras.layers import Concatenate
from tensorflow.keras import optimizers
from tensorflow.keras.layers import BatchNormalization
from tensorflow.keras.metrics import MeanIoU
from tensorflow.keras.callbacks import ModelCheckpoint, EarlyStopping, ReduceLROnPlateau
from tensorflow.keras.metrics import MeanIoU, Recall, Precision
import tensorflow_addons as tfa
# Defining the Convolution Block
def conv_block(input, num_filters):
x = Conv2D(num_filters, 3, padding="same", kernel_initializer="he_normal")(input)
x = BatchNormalization()(x)
#Used the Mish activation function as it performs better than ReLU (but is computionally expensive)
x = tfa.activations.mish(x)
#Comment the previous line and uncomment the next line if you limited compute resource
#x = Activation("relu")(x)
x = Conv2D(num_filters, 3, padding="same", kernel_initializer="he_normal")(x)
x = BatchNormalization()(x)
x = tfa.activations.mish(x)
#x = x*tf.math.tanh(tf.softplus(x)) #Mish activation in a mathematical form
#x = Activation("relu")(x)
return x
#Defining the Transpose Convolution Block
def decoder_block(input, skip_features, num_filters):
x = Conv2DTranspose(num_filters, (2, 2), strides=2, padding="same")(input)
x = Concatenate()([x, skip_features])
#Use dropout only if the model is overfitting
#x = Dropout(0.05)(x)
x = conv_block(x, num_filters)
return x
#Building the EfficientNetB0_UNet
def build_efficientNetB0_unet(input_shape):
""" Input """
inputs = Input(shape=input_shape, name='input_image')
""" Pre-trained EfficientNetB0 Model """
effNetB0 = tf.keras.applications.EfficientNetB0(input_tensor=inputs, include_top=False,
weights="imagenet")
# This Section will let you freeze and unfreeze layers. Here I have frozen all layer except
# the last convolution block layers starting after layer 16
for layer in effNetB0.layers[:-16]:
layer.trainable = False
for l in effNetB0.layers:
print(l.name, l.trainable)
""" Encoder """
s1 = effNetB0.get_layer("input_image").output ## (512 x 512)
s2 = effNetB0.get_layer("block1a_activation").output ## (256 x 256)
s3 = effNetB0.get_layer("block2a_activation").output ## (128 x 128)
s4 = effNetB0.get_layer("block3a_activation").output ## (64 x 64)
s5 = effNetB0.get_layer("block4a_activation").output ## (32 x 32)
""" Bridge """
b1 = effNetB0.get_layer("block7a_activation").output ## (16 x 16)
""" Decoder """
d1 = decoder_block(b1, s5, 512) ## (32 x 32)
d2 = decoder_block(d1, s4, 256) ## (64 x 64)
d3 = decoder_block(d2, s3, 128) ## (128 x 128)
d4 = decoder_block(d3, s2, 64) ## (256 x 256)
d5 = decoder_block(d4, s1, 32) ## (512 x 512)
""" Output """
outputs = Conv2D(1, 1, padding="same", activation="sigmoid")(d5)
model = Model(inputs, outputs, name="EfficientNetB0_U-Net")
return model
if __name__ == "__main__":
input_shape = (IMAGE_WIDTH, IMAGE_HEIGHT, CHANNELS)
model = build_efficientNetB0_unet(input_shape)
#Shows the entire EfficientNetB0_UNet Model
model.summary()
#Adding Model Checkpoints, Early Stopping based on Validation Loss and LR Reducer
model_path = "path/Model_Name.h5"
checkpointer = ModelCheckpoint(model_path,
monitor="val_loss",
mode="min",
save_best_only = True,
verbose=1)
earlystopper = EarlyStopping(monitor = 'val_loss',
min_delta = 0,
patience = 30,
verbose = 1,
restore_best_weights = True)
lr_reducer = ReduceLROnPlateau(monitor='val_loss',
factor=0.6,
patience=6,
verbose=1,
min_lr=0.0001
#min_delta=5e-5
)
lr_schedule = keras.optimizers.schedules.ExponentialDecay(
initial_learning_rate=1e-3,
decay_steps=6000,
decay_rate=0.9)
optimizer = keras.optimizers.Adam(learning_rate=lr_schedule)
from tensorflow.keras import backend as K
# To calculate Intersection over Union between Predicted Mask and Ground Truth
def iou_coef(y_true, y_pred, smooth=1):
intersection = K.sum(K.abs(y_true * y_pred), axis=[1,2,3])
union = K.sum(y_true,[1,2,3])+K.sum(y_pred,[1,2,3])-intersection
iou = K.mean((intersection + smooth) / (union + smooth), axis=0)
return iou
smooth = 1e-5
# F1 score or Dice Coefficient
def f1_score(y_true, y_pred, smooth = 1):
y_true_f = K.flatten(y_true)
y_pred_f = K.flatten(y_pred)
intersection = K.sum(y_true_f * y_pred_f)
return (2. * intersection + smooth) / (K.sum(y_true_f) + K.sum(y_pred_f) + smooth)
# Soft Dice Loss
def soft_dice_loss(y_true, y_pred):
return 1-dice_coef(y_true, y_pred)
#Compiling the model with Adam Optimizer and Metrics related to segmentation
model.compile(optimizer=optimizer,
loss=soft_dice_loss,
metrics=[iou_coef, Recall(), Precision(), MeanIoU(num_classes=2), f1_score])
# Initiate Model Training
'''history = model.fit(train_images,
train_masks/255,
validation_split=0.10,
epochs=EPOCHS,
batch_size = BATCH_SIZE,
callbacks = [checkpointer, earlystopper, lr_reducer])'''
```
| github_jupyter |
**This notebook is an exercise in the [Intermediate Machine Learning](https://www.kaggle.com/learn/intermediate-machine-learning) course. You can reference the tutorial at [this link](https://www.kaggle.com/alexisbcook/introduction).**
---
As a warm-up, you'll review some machine learning fundamentals and submit your initial results to a Kaggle competition.
# Setup
The questions below will give you feedback on your work. Run the following cell to set up the feedback system.
```
# Set up code checking
import os
if not os.path.exists("../input/train.csv"):
os.symlink("../input/home-data-for-ml-course/train.csv", "../input/train.csv")
os.symlink("../input/home-data-for-ml-course/test.csv", "../input/test.csv")
from learntools.core import binder
binder.bind(globals())
from learntools.ml_intermediate.ex1 import *
print("Setup Complete")
```
You will work with data from the [Housing Prices Competition for Kaggle Learn Users](https://www.kaggle.com/c/home-data-for-ml-course) to predict home prices in Iowa using 79 explanatory variables describing (almost) every aspect of the homes.

Run the next code cell without changes to load the training and validation features in `X_train` and `X_valid`, along with the prediction targets in `y_train` and `y_valid`. The test features are loaded in `X_test`. (_If you need to review **features** and **prediction targets**, please check out [this short tutorial](https://www.kaggle.com/dansbecker/your-first-machine-learning-model). To read about model **validation**, look [here](https://www.kaggle.com/dansbecker/model-validation). Alternatively, if you'd prefer to look through a full course to review all of these topics, start [here](https://www.kaggle.com/learn/machine-learning).)_
```
import pandas as pd
from sklearn.model_selection import train_test_split
# Read the data
X_full = pd.read_csv('../input/train.csv', index_col='Id')
X_test_full = pd.read_csv('../input/test.csv', index_col='Id')
# Obtain target and predictors
y = X_full.SalePrice
features = ['LotArea', 'YearBuilt', '1stFlrSF', '2ndFlrSF', 'FullBath', 'BedroomAbvGr', 'TotRmsAbvGrd']
X = X_full[features].copy()
X_test = X_test_full[features].copy()
# Break off validation set from training data
X_train, X_valid, y_train, y_valid = train_test_split(X, y, train_size=0.8, test_size=0.2,
random_state=0)
```
Use the next cell to print the first several rows of the data. It's a nice way to get an overview of the data you will use in your price prediction model.
```
X_train.head()
```
The next code cell defines five different random forest models. Run this code cell without changes. (_To review **random forests**, look [here](https://www.kaggle.com/dansbecker/random-forests)._)
```
from sklearn.ensemble import RandomForestRegressor
# Define the models
model_1 = RandomForestRegressor(n_estimators=50, random_state=0)
model_2 = RandomForestRegressor(n_estimators=100, random_state=0)
model_3 = RandomForestRegressor(n_estimators=100, criterion='mae', random_state=0)
model_4 = RandomForestRegressor(n_estimators=200, min_samples_split=20, random_state=0)
model_5 = RandomForestRegressor(n_estimators=100, max_depth=7, random_state=0)
models = [model_1, model_2, model_3, model_4, model_5]
```
To select the best model out of the five, we define a function `score_model()` below. This function returns the mean absolute error (MAE) from the validation set. Recall that the best model will obtain the lowest MAE. (_To review **mean absolute error**, look [here](https://www.kaggle.com/dansbecker/model-validation).)_
Run the code cell without changes.
```
from sklearn.metrics import mean_absolute_error
# Function for comparing different models
def score_model(model, X_t=X_train, X_v=X_valid, y_t=y_train, y_v=y_valid):
model.fit(X_t, y_t)
preds = model.predict(X_v)
return mean_absolute_error(y_v, preds)
for i in range(0, len(models)):
mae = score_model(models[i])
print("Model %d MAE: %d" % (i+1, mae))
```
# Step 1: Evaluate several models
Use the above results to fill in the line below. Which model is the best model? Your answer should be one of `model_1`, `model_2`, `model_3`, `model_4`, or `model_5`.
```
# Fill in the best model
best_model = model_3
# Check your answer
step_1.check()
# Lines below will give you a hint or solution code
#step_1.hint()
#step_1.solution()
```
# Step 2: Generate test predictions
Great. You know how to evaluate what makes an accurate model. Now it's time to go through the modeling process and make predictions. In the line below, create a Random Forest model with the variable name `my_model`.
```
# Define a model
my_model = RandomForestRegressor(n_estimators=100, criterion='mae', random_state=0) # Your code here
# Check your answer
step_2.check()
# Lines below will give you a hint or solution code
#step_2.hint()
#step_2.solution()
```
Run the next code cell without changes. The code fits the model to the training and validation data, and then generates test predictions that are saved to a CSV file. These test predictions can be submitted directly to the competition!
```
# Fit the model to the training data
my_model.fit(X, y)
# Generate test predictions
preds_test = my_model.predict(X_test)
# Save predictions in format used for competition scoring
output = pd.DataFrame({'Id': X_test.index,
'SalePrice': preds_test})
output.to_csv('submission.csv', index=False)
```
# Submit your results
Once you have successfully completed Step 2, you're ready to submit your results to the leaderboard! First, you'll need to join the competition if you haven't already. So open a new window by clicking on [this link](https://www.kaggle.com/c/home-data-for-ml-course). Then click on the **Join Competition** button.

Next, follow the instructions below:
1. Begin by clicking on the blue **Save Version** button in the top right corner of the window. This will generate a pop-up window.
2. Ensure that the **Save and Run All** option is selected, and then click on the blue **Save** button.
3. This generates a window in the bottom left corner of the notebook. After it has finished running, click on the number to the right of the **Save Version** button. This pulls up a list of versions on the right of the screen. Click on the ellipsis **(...)** to the right of the most recent version, and select **Open in Viewer**. This brings you into view mode of the same page. You will need to scroll down to get back to these instructions.
4. Click on the **Output** tab on the right of the screen. Then, click on the file you would like to submit, and click on the blue **Submit** button to submit your results to the leaderboard.
You have now successfully submitted to the competition!
If you want to keep working to improve your performance, select the blue **Edit** button in the top right of the screen. Then you can change your code and repeat the process. There's a lot of room to improve, and you will climb up the leaderboard as you work.
# Keep going
You've made your first model. But how can you quickly make it better?
Learn how to improve your competition results by incorporating columns with **[missing values](https://www.kaggle.com/alexisbcook/missing-values)**.
---
*Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum/161289) to chat with other Learners.*
| github_jupyter |
# TensorFlow Datasets
TFDS provides a collection of ready-to-use datasets for use with TensorFlow, Jax, and other Machine Learning frameworks.
It handles downloading and preparing the data deterministically and constructing a `tf.data.Dataset` (or `np.array`).
Note: Do not confuse [TFDS](https://www.tensorflow.org/datasets) (this library) with `tf.data` (TensorFlow API to build efficient data pipelines). TFDS is a high level wrapper around `tf.data`. If you're not familiar with this API, we encourage you to read [the official tf.data guide](https://www.tensorflow.org/guide/data) first.
Copyright 2018 The TensorFlow Datasets Authors, Licensed under the Apache License, Version 2.0
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/datasets/overview"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/datasets/blob/master/docs/overview.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/datasets/blob/master/docs/overview.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/datasets/docs/overview.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
## Installation
TFDS exists in two packages:
* `pip install tensorflow-datasets`: The stable version, released every few months.
* `pip install tfds-nightly`: Released every day, contains the last versions of the datasets.
This colab uses `tfds-nightly`:
```
!pip install -q tfds-nightly tensorflow matplotlib
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
import tensorflow_datasets as tfds
```
## Find available datasets
All dataset builders are subclass of `tfds.core.DatasetBuilder`. To get the list of available builders, use `tfds.list_builders()` or look at our [catalog](https://www.tensorflow.org/datasets/catalog/overview).
```
tfds.list_builders()
```
## Load a dataset
### tfds.load
The easiest way of loading a dataset is `tfds.load`. It will:
1. Download the data and save it as [`tfrecord`](https://www.tensorflow.org/tutorials/load_data/tfrecord) files.
2. Load the `tfrecord` and create the `tf.data.Dataset`.
```
ds = tfds.load('mnist', split='train', shuffle_files=True)
assert isinstance(ds, tf.data.Dataset)
print(ds)
```
Some common arguments:
* `split=`: Which split to read (e.g. `'train'`, `['train', 'test']`, `'train[80%:]'`,...). See our [split API guide](https://www.tensorflow.org/datasets/splits).
* `shuffle_files=`: Control whether to shuffle the files between each epoch (TFDS store big datasets in multiple smaller files).
* `data_dir=`: Location where the dataset is saved (
defaults to `~/tensorflow_datasets/`)
* `with_info=True`: Returns the `tfds.core.DatasetInfo` containing dataset metadata
* `download=False`: Disable download
### tfds.builder
`tfds.load` is a thin wrapper around `tfds.core.DatasetBuilder`. You can get the same output using the `tfds.core.DatasetBuilder` API:
```
builder = tfds.builder('mnist')
# 1. Create the tfrecord files (no-op if already exists)
builder.download_and_prepare()
# 2. Load the `tf.data.Dataset`
ds = builder.as_dataset(split='train', shuffle_files=True)
print(ds)
```
### `tfds build` CLI
If you want to generate a specific dataset, you can use the [`tfds` command line](https://www.tensorflow.org/datasets/cli). For example:
```sh
tfds build mnist
```
See [the doc](https://www.tensorflow.org/datasets/cli) for available flags.
## Iterate over a dataset
### As dict
By default, the `tf.data.Dataset` object contains a `dict` of `tf.Tensor`s:
```
ds = tfds.load('mnist', split='train')
ds = ds.take(1) # Only take a single example
for example in ds: # example is `{'image': tf.Tensor, 'label': tf.Tensor}`
print(list(example.keys()))
image = example["image"]
label = example["label"]
print(image.shape, label)
```
To find out the `dict` key names and structure, look at the dataset documentation in [our catalog](https://www.tensorflow.org/datasets/catalog/overview#all_datasets). For example: [mnist documentation](https://www.tensorflow.org/datasets/catalog/mnist).
### As tuple (`as_supervised=True`)
By using `as_supervised=True`, you can get a tuple `(features, label)` instead for supervised datasets.
```
ds = tfds.load('mnist', split='train', as_supervised=True)
ds = ds.take(1)
for image, label in ds: # example is (image, label)
print(image.shape, label)
```
### As numpy (`tfds.as_numpy`)
Uses `tfds.as_numpy` to convert:
* `tf.Tensor` -> `np.array`
* `tf.data.Dataset` -> `Iterator[Tree[np.array]]` (`Tree` can be arbitrary nested `Dict`, `Tuple`)
```
ds = tfds.load('mnist', split='train', as_supervised=True)
ds = ds.take(1)
for image, label in tfds.as_numpy(ds):
print(type(image), type(label), label)
```
### As batched tf.Tensor (`batch_size=-1`)
By using `batch_size=-1`, you can load the full dataset in a single batch.
This can be combined with `as_supervised=True` and `tfds.as_numpy` to get the the data as `(np.array, np.array)`:
```
image, label = tfds.as_numpy(tfds.load(
'mnist',
split='test',
batch_size=-1,
as_supervised=True,
))
print(type(image), image.shape)
```
Be careful that your dataset can fit in memory, and that all examples have the same shape.
## Benchmark your datasets
Benchmarking a dataset is a simple `tfds.benchmark` call on any iterable (e.g. `tf.data.Dataset`, `tfds.as_numpy`,...).
```
ds = tfds.load('mnist', split='train')
ds = ds.batch(32).prefetch(1)
tfds.benchmark(ds, batch_size=32)
tfds.benchmark(ds, batch_size=32) # Second epoch much faster due to auto-caching
```
* Do not forget to normalize the results per batch size with the `batch_size=` kwarg.
* In the summary, the first warmup batch is separated from the other ones to capture `tf.data.Dataset` extra setup time (e.g. buffers initialization,...).
* Notice how the second iteration is much faster due to [TFDS auto-caching](https://www.tensorflow.org/datasets/performances#auto-caching).
* `tfds.benchmark` returns a `tfds.core.BenchmarkResult` which can be inspected for further analysis.
### Build end-to-end pipeline
To go further, you can look:
* Our [end-to-end Keras example](https://www.tensorflow.org/datasets/keras_example) to see a full training pipeline (with batching, shuffling,...).
* Our [performance guide](https://www.tensorflow.org/datasets/performances) to improve the speed of your pipelines (tip: use `tfds.benchmark(ds)` to benchmark your datasets).
## Visualization
### tfds.as_dataframe
`tf.data.Dataset` objects can be converted to [`pandas.DataFrame`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.html) with `tfds.as_dataframe` to be visualized on [Colab](https://colab.research.google.com).
* Add the `tfds.core.DatasetInfo` as second argument of `tfds.as_dataframe` to visualize images, audio, texts, videos,...
* Use `ds.take(x)` to only display the first `x` examples. `pandas.DataFrame` will load the full dataset in-memory, and can be very expensive to display.
```
ds, info = tfds.load('mnist', split='train', with_info=True)
tfds.as_dataframe(ds.take(4), info)
```
### tfds.show_examples
`tfds.show_examples` returns a `matplotlib.figure.Figure` (only image datasets supported now):
```
ds, info = tfds.load('mnist', split='train', with_info=True)
fig = tfds.show_examples(ds, info)
```
## Access the dataset metadata
All builders include a `tfds.core.DatasetInfo` object containing the dataset metadata.
It can be accessed through:
* The `tfds.load` API:
```
ds, info = tfds.load('mnist', with_info=True)
```
* The `tfds.core.DatasetBuilder` API:
```
builder = tfds.builder('mnist')
info = builder.info
```
The dataset info contains additional informations about the dataset (version, citation, homepage, description,...).
```
print(info)
```
### Features metadata (label names, image shape,...)
Access the `tfds.features.FeatureDict`:
```
info.features
```
Number of classes, label names:
```
print(info.features["label"].num_classes)
print(info.features["label"].names)
print(info.features["label"].int2str(7)) # Human readable version (8 -> 'cat')
print(info.features["label"].str2int('7'))
```
Shapes, dtypes:
```
print(info.features.shape)
print(info.features.dtype)
print(info.features['image'].shape)
print(info.features['image'].dtype)
```
### Split metadata (e.g. split names, number of examples,...)
Access the `tfds.core.SplitDict`:
```
print(info.splits)
```
Available splits:
```
print(list(info.splits.keys()))
```
Get info on individual split:
```
print(info.splits['train'].num_examples)
print(info.splits['train'].filenames)
print(info.splits['train'].num_shards)
```
It also works with the subsplit API:
```
print(info.splits['train[15%:75%]'].num_examples)
print(info.splits['train[15%:75%]'].file_instructions)
```
## Troubleshooting
### Manual download (if download fails)
If download fails for some reason (e.g. offline,...). You can always manually download the data yourself and place it in the `manual_dir` (defaults to `~/tensorflow_datasets/download/manual/`.
To find out which urls to download, look into:
* For new datasets (implemented as folder): [`tensorflow_datasets/`](https://github.com/tensorflow/datasets/tree/master/tensorflow_datasets/)`<type>/<dataset_name>/checksums.tsv`. For example: [`tensorflow_datasets/text/bool_q/checksums.tsv`](https://github.com/tensorflow/datasets/blob/master/tensorflow_datasets/text/bool_q/checksums.tsv).
You can find the dataset source location in [our catalog](https://www.tensorflow.org/datasets/catalog/overview).
* For old datasets: [`tensorflow_datasets/url_checksums/<dataset_name>.txt`](https://github.com/tensorflow/datasets/tree/master/tensorflow_datasets/url_checksums)
### Fixing `NonMatchingChecksumError`
TFDS ensure determinism by validating the checksums of downloaded urls.
If `NonMatchingChecksumError` is raised, might indicate:
* The website may be down (e.g. `503 status code`). Please check the url.
* For Google Drive URLs, try again later as Drive sometimes rejects downloads when too many people access the same URL. See [bug](https://github.com/tensorflow/datasets/issues/1482)
* The original datasets files may have been updated. In this case the TFDS dataset builder should be updated. Please open a new Github issue or PR:
* Register the new checksums with `tfds build --register_checksums`
* Eventually update the dataset generation code.
* Update the dataset `VERSION`
* Update the dataset `RELEASE_NOTES`: What caused the checksums to change ? Did some examples changed ?
* Make sure the dataset can still be built.
* Send us a PR
Note: You can also inspect the downloaded file in `~/tensorflow_datasets/download/`.
## Citation
If you're using `tensorflow-datasets` for a paper, please include the following citation, in addition to any citation specific to the used datasets (which can be found in the [dataset catalog](https://www.tensorflow.org/datasets/catalog/overview)).
```
@misc{TFDS,
title = { {TensorFlow Datasets}, A collection of ready-to-use datasets},
howpublished = {\url{https://www.tensorflow.org/datasets}},
}
```
| github_jupyter |
```
import csv
import tensorflow as tf
import numpy as np
import pandas as pd
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
#!wget --no-check-certificate \
# https://storage.googleapis.com/laurencemoroney-blog.appspot.com/bbc-text.csv \
# -O /tmp/bbc-text.csv
vocab_size = 1000
embedding_dim = 16
max_length = 120
trunc_type = 'post'
padding_type = 'post'
oov_tok = '<OOV>'
training_portion = .8
sentences = []
labels = []
stopwords = [ "a", "about", "above", "after", "again", "against", "all", "am", "an", "and", "any", "are", "as", "at", "be", "because", "been", "before", "being", "below", "between", "both", "but", "by", "could", "did", "do", "does", "doing", "down", "during", "each", "few", "for", "from", "further", "had", "has", "have", "having", "he", "he'd", "he'll", "he's", "her", "here", "here's", "hers", "herself", "him", "himself", "his", "how", "how's", "i", "i'd", "i'll", "i'm", "i've", "if", "in", "into", "is", "it", "it's", "its", "itself", "let's", "me", "more", "most", "my", "myself", "nor", "of", "on", "once", "only", "or", "other", "ought", "our", "ours", "ourselves", "out", "over", "own", "same", "she", "she'd", "she'll", "she's", "should", "so", "some", "such", "than", "that", "that's", "the", "their", "theirs", "them", "themselves", "then", "there", "there's", "these", "they", "they'd", "they'll", "they're", "they've", "this", "those", "through", "to", "too", "under", "until", "up", "very", "was", "we", "we'd", "we'll", "we're", "we've", "were", "what", "what's", "when", "when's", "where", "where's", "which", "while", "who", "who's", "whom", "why", "why's", "with", "would", "you", "you'd", "you'll", "you're", "you've", "your", "yours", "yourself", "yourselves" ]
print(len(stopwords))
# Expected Output
# 153
file = pd.read_csv('./tmp/nlp/bbc-text.csv')
file['text'] = file['text'].apply(lambda x: ' '.join([word for word in x.split() if word not in (stopwords)]))
sentences = file['text'].values.tolist()
labels = file['category'].values.tolist()
print(len(labels))
print(len(sentences))
print(sentences[0])
# Expected Output
# 2225
# 2225
# tv future hands viewers home theatre systems plasma high-definition tvs digital video recorders moving living room way people watch tv will radically different five years time. according expert panel gathered annual consumer electronics show las vegas discuss new technologies will impact one favourite pastimes. us leading trend programmes content will delivered viewers via home networks cable satellite telecoms companies broadband service providers front rooms portable devices. one talked-about technologies ces digital personal video recorders (dvr pvr). set-top boxes like us s tivo uk s sky+ system allow people record store play pause forward wind tv programmes want. essentially technology allows much personalised tv. also built-in high-definition tv sets big business japan us slower take off europe lack high-definition programming. not can people forward wind adverts can also forget abiding network channel schedules putting together a-la-carte entertainment. us networks cable satellite companies worried means terms advertising revenues well brand identity viewer loyalty channels. although us leads technology moment also concern raised europe particularly growing uptake services like sky+. happens today will see nine months years time uk adam hume bbc broadcast s futurologist told bbc news website. likes bbc no issues lost advertising revenue yet. pressing issue moment commercial uk broadcasters brand loyalty important everyone. will talking content brands rather network brands said tim hanlon brand communications firm starcom mediavest. reality broadband connections anybody can producer content. added: challenge now hard promote programme much choice. means said stacey jolna senior vice president tv guide tv group way people find content want watch simplified tv viewers. means networks us terms channels take leaf google s book search engine future instead scheduler help people find want watch. kind channel model might work younger ipod generation used taking control gadgets play them. might not suit everyone panel recognised. older generations comfortable familiar schedules channel brands know getting. perhaps not want much choice put hands mr hanlon suggested. end kids just diapers pushing buttons already - everything possible available said mr hanlon. ultimately consumer will tell market want. 50 000 new gadgets technologies showcased ces many enhancing tv-watching experience. high-definition tv sets everywhere many new models lcd (liquid crystal display) tvs launched dvr capability built instead external boxes. one example launched show humax s 26-inch lcd tv 80-hour tivo dvr dvd recorder. one us s biggest satellite tv companies directtv even launched branded dvr show 100-hours recording capability instant replay search function. set can pause rewind tv 90 hours. microsoft chief bill gates announced pre-show keynote speech partnership tivo called tivotogo means people can play recorded programmes windows pcs mobile devices. reflect increasing trend freeing multimedia people can watch want want.
train_size = int(len(sentences)*training_portion)
train_sentences = sentences[:train_size]
train_labels = labels[:train_size]
validation_sentences = sentences[train_size:]
validation_labels = labels[train_size:]
print(train_size)
print(len(train_sentences))
print(len(train_labels))
print(len(validation_sentences))
print(len(validation_labels))
# Expected output (if training_portion=.8)
# 1780
# 1780
# 1780
# 445
# 445
tokenizer = Tokenizer(num_words = 100, oov_token=oov_tok)
tokenizer.fit_on_texts(sentences)
word_index = tokenizer.word_index
train_sequences = tokenizer.texts_to_sequences(train_sentences)
train_padded = pad_sequences(train_sequences, maxlen=max_length)
print(len(train_sequences[0]))
print(len(train_padded[0]))
print(len(train_sequences[1]))
print(len(train_padded[1]))
print(len(train_sequences[10]))
print(len(train_padded[10]))
# Expected Ouput
# 449
# 120
# 200
# 120
# 192
# 120
validation_sequences = tokenizer.texts_to_sequences(validation_sentences)
validation_padded = pad_sequences(validation_sequences, maxlen=max_length)
print(len(validation_sequences))
print(validation_padded.shape)
# Expected output
# 445
# (445, 120)
label_tokenizer = Tokenizer()
label_tokenizer.fit_on_texts(labels)
training_label_seq = np.asarray(label_tokenizer.texts_to_sequences(train_labels))
validation_label_seq = np.asarray(label_tokenizer.texts_to_sequences(validation_labels))
print(training_label_seq[0])
print(training_label_seq[1])
print(training_label_seq[2])
print(training_label_seq.shape)
print(validation_label_seq[0])
print(validation_label_seq[1])
print(validation_label_seq[2])
print(validation_label_seq.shape)
# Expected output
# [4]
# [2]
# [1]
# (1780, 1)
# [5]
# [4]
# [3]
# (445, 1)
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim, input_length=max_length),
tf.keras.layers.GlobalAveragePooling1D(),
tf.keras.layers.Dense(24, activation = 'relu'),
tf.keras.layers.Dense(6, activation = 'sigmoid')
])
model.compile(loss='sparse_categorical_crossentropy',optimizer='adam',metrics=['accuracy'])
model.summary()
# Expected Output
# Layer (type) Output Shape Param #
# =================================================================
# embedding (Embedding) (None, 120, 16) 16000
# _________________________________________________________________
# global_average_pooling1d (Gl (None, 16) 0
# _________________________________________________________________
# dense (Dense) (None, 24) 408
# _________________________________________________________________
# dense_1 (Dense) (None, 6) 150
# =================================================================
# Total params: 16,558
# Trainable params: 16,558
# Non-trainable params: 0
num_epochs = 30
history = model.fit(x=train_padded,y=training_label_seq, epochs=num_epochs, validation_data=(validation_padded,validation_label_seq))
import matplotlib.pyplot as plt
def plot_graphs(history, string):
plt.plot(history.history[string])
plt.plot(history.history['val_'+string])
plt.xlabel("Epochs")
plt.ylabel(string)
plt.legend([string, 'val_'+string])
plt.show()
plot_graphs(history, "accuracy")
plot_graphs(history, "loss")
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
def decode_sentence(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
e = model.layers[0]
weights = e.get_weights()[0]
print(weights.shape) # shape: (vocab_size, embedding_dim)
# Expected output
# (1000, 16)
import io
out_v = io.open('vecs.tsv', 'w', encoding='utf-8')
out_m = io.open('meta.tsv', 'w', encoding='utf-8')
for word_num in range(1, vocab_size):
word = reverse_word_index[word_num]
embeddings = weights[word_num]
out_m.write(word + "\n")
out_v.write('\t'.join([str(x) for x in embeddings]) + "\n")
out_v.close()
out_m.close()
try:
from google.colab import files
except ImportError:
pass
else:
files.download('vecs.tsv')
files.download('meta.tsv')
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.